An Algorithm for Bayesian Variable Selection in High-dimensional Generalized Linear Models
|
|
|
- Gerald Park
- 5 years ago
- Views:
Transcription
1 Proceedings 59th ISI World Statistics Congress, August 2013, Hong Kong (Session CPS023) p.3938 An Algorithm for Bayesian Variable Selection in High-dimensional Generalized Linear Models Vitara Pungpapong Department of Statistics, Faculty of Commerce and Accountancy Chulalongkorn Unversity, Bangkok, THAILAND Abstract Inspired by analysis of genomic data, the primary quest is to identify associations between studied traits and genetic markers where number of markers is typically much larger than sample size. Bayesian variable selection methods with Markov chain Monte Carlo (MCMC) are extensively applied to analyze such high-dimensional data. However, MCMC is often slow to converge with large number of candidate predictors. In this study, we examine the empirical Bayes variable selection with a sparse prior on the unknown coefficients. An iterated conditional modes/medians (ICM/M) algorithm is proposed for implementation by iteratively minimizing a conditional loss function in high-dimensional linear regression model. Attention is then directed to extend the algorithm to a generalized linear model. The performances of our approach are evaluated through simulation study. Keywords: Bayesian inference, high-dimensional data, sparse variables, generalized linear models 1. Introduction Linear and nonlinear models have been extensively used to identify associations between response and explanatory variables. The advent of high throughput technologies enables the collection of massive and complex data in biomedical science. High-dimensional data (i.e. number of predictors is larger than sample size) have posed a challenge in selecting variables for such models. Tibshirani (1996) proposed lasso by putting an l -penalty on likelihood function producing sparse coefficients. Lasso cannot be applied to only a linear model but has also extended to generalized linear models (GLMs). Methods to determine the entire path of lasso for GLMs were discussed in Efron et. al. (2004), Park and Hastie (2007), Friedman et. al. (2010). Johnstone and Silverman (2004) proposed empirical Bayes thresholding by putting a mixture prior of an atom of probability at zero and a heavy-tailed density. Pungpapong et. al. (2012) presented a contribution of empirical Bayes thresholding to select variables in linear regression framework. An iterated conditional modes/medians (ICM/M) is introduced for fast and easy-to-implement algorithm. The aim of this study is to generalize empirical Bayes variable selection (Pungpapong et. al., 2012) to more class beyond linear model. The details of ICM/M algorithm for GLMs are also described. The ability in selecting variables is demonstrated in simulation study.
2 Proceedings 59th ISI World Statistics Congress, August 2013, Hong Kong (Session CPS023) p Methods Empirical Bayes variable selection in linear model Consider a normal linear regression model, ~ 0,. (1) is a 1 random vector, is a matrix of predictors, and is a identity matrix. is further assumed to be centralized and to be standardized. In order to perform variable selection, Pungpapong et. al. introduced the following independent mixture prior distribution to model the sparsity of : ~ 1, (2) where. is a Dirac delta function at zero and. is a probability function. Under this prior distribution, each is zero with probability 1 and is drawn from the nonzero part of prior. with probability. Johnstone and Silverman suggested a heavy-tailed density such as Laplace distribution for. 1 exp 1 (3) 2, where 0 is a scale parameter. The default value of = 0.5 is used throughout the paper as suggested by Johnstone and Silverman. Jeffrey s prior (Jeffreys, 1946) is taken on. When the information of structural relationship among predictors is available, it is more sensible to incorporate such information in the prior. Pungpapong et. al. (2012) introduced an indicator variable,, where 1 and an underlying structure among can be represented by an undirected graph, comprising a set of vertices and a set of edges. Specifically, the prior distribution is set to be dependent to, ~ 1. (4) The following Ising model (Onsagar, 1943) is then employed to model a priori information on, 1 (5), exp,, where and are two parameters and, is a normalizing constant. ICM/M Algorithm Pungpapong et. al. (2012) presented an iterated conditional modes/medians (ICM/M) algorithm for fast computation of empirical Bayes variable selection. Data-driven optimal values for hyperparameters and auxiliary parameters are obtained as the modes of their full conditional distribution functions. Each regression coefficient is obtained as the median of its full conditional distribution function. With the presented Bayesian framework, the use of conditional medians can enforce the variable selection and estimation simultaneously. The iterative procedure for updating regression coefficients and other parameters is carried out until convergence. The ICM/M algorithm for empirical Bayes variable selection with independent prior on coefficients has the following steps: 1. Obtain initial estimates of all parameters denoted as,.
3 Proceedings 59th ISI World Statistics Congress, August 2013, Hong Kong (Session CPS023) p Estimate hyperparameter with the value of as the mode of its full conditional distribution function. mode,,, 3. Update each regression coefficient as the median of its full conditional distribution function. median,,,,, where,..,,,,. 1,,, 4. Update as the mode of its full conditional distribution function. mode,,, 5. Iterate between steps 2-4 until convergence. With Ising prior for structured predictors, the ICM/M algorithm is implemented involving steps as follows: 1. Obtain initial estimates of all parameters denoted as,. 2. Obtain the indicator variable,, where With current value of, estimate hyperparameters, with the values of, as the mode of its pseudo-likelihood function., mode ;, mode :, ;, The general setup of ICM/M algorithm suggests that, are the modes of their full conditional distribution function which is the prior likelihood. The reason for using the pseudo-likelihood function instead is to avoid computational difficulty in computing an unknown normalizing constant in Ising model. 4. Update each regression coefficient as the median of its full conditional distribution function. median,,,,,, where,..,,,,. 1,,, 5. Update as the mode of its full conditional distribution function. mode,,,, 6. Iterate between steps 2-5 until convergence. Extension to GLMs Generalized linear model (GLM) is an extension of the ordinary linear model. Each component is assumed to be independently distributed with mean and variance. is further assumed to have a distribution in the exponential family taken the form,exp (6),..,., and. are functions which vary according to distributions.,,, is known as the canonical parameter. Here we assume a fixed
4 Proceedings 59th ISI World Statistics Congress, August 2013, Hong Kong (Session CPS023) p.3941 known dispersion parameter as 1. A link function. is introduced to connect the linear predictor to the mean. That is,. A quadratic approximation to the likelihood is employed to construct pseudodata and pseudovariances. The basic idea is to approximate the generalized linear model by a linear model. Such a procedure is equivalent to an iteratively reweighted least squares (IRLS) algorithm. Particularly, based on the current parameter estimates at -th iteration,, pseudodata,, and pseudovariances Σ diag are constructed as (7) and Σ diag diag (8), where and the derivative is evaluated at. The approximated distribution of pseudodata is,σ. Borrowing the idea of IRLS, pseudodata is treated as a working response. After obtaining pseudodata and pseudovariances, the ICM/M algorithm is used to update regression coefficients and other parameters. The procedure consists of the outer and inner loop. The outer loop is taken place to update pseudodata and pseudovariances based on current estimates. The inner loop is where ICM/M is employed to loop through update all parameters. This simple-to-implement algorithm achieves fast computation even in high-dimensional data analysis. 3. Results To demonstrate the proposed methods, we consider four different cases of large small datasets. Our simulations focus on binary and survival data which are particularly interested in analysis of genomic data. Logistic regression GLM was employed to analyze binary data and Cox proportional hazards model (Cox, 1972) was carried out to deal with survival data. In the first three cases, we fixed = 1,000 and = 250. The simulations were run 100 times for each setup. Case 1: Binary data: high/mild correlations in covariates The binary response was simulated with the probability following logistic regression model: exp 1 1 exp. The intercept term is zero and the non-zero regression coefficients are = 10 and = -5. The covariates are partitioned into ten blocks, with each block including 100 covariates and they are serially correlated at the same level of. The values of are {0, 0.5, 0.9}. Case 2: Survival data: high/mild correlations in covariates The covariates were simulated the same way as in Case 1. Survival times were generated from a Cox model and follow Weibull distribution with shape a parameter = 10 and a scale parameter = 1. Among 1,000 predictors, the failure rate is determined by a linear combination of 20 non-zero coefficients: = 5 and = 2. The censoring times were generated randomly to achieve 50% of the observed event times. Case 3: Survival data: Markov chain
5 Proceedings 59th ISI World Statistics Congress, August 2013, Hong Kong (Session CPS023) p.3942 Same as Case 2 except that the covariates were generated from AR(1) with different value of in {0, 0.5, 0.9} and the location of non-zero coefficients follows a Markov chain. Particularly, the indicator variables,, form a Markov chain with the transition probabilities specified as follows: , The first indicator variable was sampled from Bernoulli(0.5). The effect sizes of those non-zero coefficients were drawn from Uniform[0.5,5]. Case 4: Binary data: pathway information Our simulation is based on sampling of 871 individuals from Parkinson's disease (PD) SNP data (dbgap study accession number: phs v3.p2). Here we consider 1,152 SNPs affiliating with 341 genes in PD-related metabolic pathways. The phenotype was simulated from logistic regression with 46 non-zero coefficients resided in 15 genes in PD pathway and the effect sizes were drawn from Uniform[1,10]. Here we evaluate the algorithm in terms of false positive and false negative rates. Two different sets of initial values for EBVS were considered, one with true values and another one with lasso fit. We also compared the performance of EBVS with lasso. EBVS with Ising prior ( ) was applied to only Case 3 and Case 4 assuming known structural relationship among predictors. Table 1: Median false positive rates (with standard errors in parentheses) across 100 datasets. Lasso EBVS EBVS (1e-3) 0.000(7e-4) 0.000(1e-3) - - Case (2e-3) 0.000(0.00) 0.000(1e-3) (3e-3) 0.000(8e-4) 0.000(4e-3) (2e-3) 0.000(3e-3) 0.000(2e-3) - - Case (1e-3) 0.000(2e-3) 0.000(7e-4) (1e-3) 0.000(7e-4) 0.000(3e-3) (2e-3) 0.000(3e-3) 0.000(0.00) 0.000(3e-3) 0.000(9e-4) Case (1e-3) 0.000(3e-3) 0.000(9e-4) 0.000(2e-3) 0.000(3e-3) (2e-3) 0.000(3e-3) 0.053(9e-3) 0.000(8e-4) 0.095(0.01) Case (3e-3) 0.000(1e-3) 0.036(3e-3) 0.000(0.00) 0.038(5e-3) Table 2: Median false negative rates (with standard errors in parentheses) across 100 datasets. Lasso EBVS EBVS (3e-5) 0.000(0.00) 0.002(9e-5) - - Case (3e-5) 0.000(0.00) 0.003(9e-5) (6e-5) 0.000(1e-5) 0.005(7e-5) (1e-4) 0.000(0.00) 0.000(3e-4) - - Case (0.00) 0.000(0.00) 0.000(0.00) (3e-5) 0.000(0.00) 0.000(9e-5) (9e-5) 0.000(0.00) 0.001(1e-4) 0.000(0.00) 0.001(1e-4) Case (5e-5) 0.000(0.00) 0.001(1e-4) 0.000(0.00) 0.000(9e-5) (1e-4) 0.000(0.00) 0.001(1e-4) 0.000(0.00) 0.001(1e-4) Case (3e-4) 0.000(3e-4) 0.020(3e-4) 0.000(0.00) 0.017(3e-4) As shown in Table 1 and Table 2, EBVS and with true coefficients are the clear winners in all cases. However, true coefficients are unknown in practice. Lasso tends to select a large number of non-zero coefficients due to high false positive rates. EBVS and with lasso fit as initial values can reduce false positive rates dramatically.
6 Proceedings 59th ISI World Statistics Congress, August 2013, Hong Kong (Session CPS023) p.3943 The median false positive rates for EBVS are all zeros in both Case 1 and Case 2. While its false negative rates are not zero in Case 1, they are all zeros in Case 2 indicating that our method can pick up the correct model. When comparing the performance between EBVS and, we see that they produce similar results in Case 3 with = 0.00 and = When = 0.09, has higher median false positive rate. The median false positive rate for is also slightly higher than that for n Case 4. However, produces smaller false negative rates implying has more power to identify true variables. 4. Discussion In this study, we have introduced an algorithm for empirical Bayes variable selection in high-dimensional GLMs. We can extend ICM/M algorithm to fit GLMs by adopting pseudodata and pseudovariances as in IRLS. Our algorithm is easy to implement and achieves fast computation empirically. Our results show that our method seems to work better with survival data than binary data in terms of false negative rate. This might be due to the fact that binary data is less informative than continuous data. We notice that has more power to detect non-zero coefficients than EBVS when the structural information among predictors is available. However, it sacrifices with slightly higher false positive rate when correlations among covariates are high. We see that our method with true coefficients as starting values can select the true model for both EBVS and. Therefore, choices of initial values other than lasso should be further studied. References Cox, D.R. (1972) Regression Models and Life-Tables. Journal of the Royal Statistical Society, Series B, 34(2), Efron, B., Hastie, T., Johnstone, I.M., and Tibshirani, R. (2004) Least angle regression. The Annals of Statistics, 32, Friedman, J., Hastie, T., and Tibshirani, R. (2010) Regularization paths for generalized linear models via coordinate descent. Journal of Statistical Software,33, Jeffreys, H. (1946) An invariant form for the prior probability in estimation problems. Proceedings of the Royal Society of London Series A, 196, Johnstone, I.M., and Silverman, B.W. (2004) Needles and straw in haystacks: empirical Bayes estimates of possibly sparse sequence The Annals of Statistics, 32, Onsagar, L. (1944) Crystal statistics. I.A two-dimensional model with an orderdisorder transition. Physical Review, 65(3-4), Park, M.Y., and Hastie, T. (2007). L 1 -regularization path algorithm for generalized linear models. Journal of the Royal Statistical Society Series B, 69, Pungpapong, V., Zhang, M., and Zhang, D. (2012), Empirical Bayes variable selection using iterated conditional modes/medians. Manuscript submitted for publication. Tibshirani, R. (1996) Regression shrinkage and selection via the lasso. Journal of the Royal Statistical Society Series B, 58,
arxiv: v2 [stat.me] 9 Aug 2017
![arxiv: v2 [stat.me] 9 Aug 2017 arxiv: v2 [stat.me] 9 Aug 2017](/thumbs/84/91144446.jpg) Variable Selection for High-dimensional Generalized Linear Models using an Iterated Conditional Modes/Medians Algorithm arxiv:1707.08298v2 [stat.me] 9 Aug 2017 Vitara Pungpapong a,, Min Zhang b, Dabao
Variable Selection for High-dimensional Generalized Linear Models using an Iterated Conditional Modes/Medians Algorithm arxiv:1707.08298v2 [stat.me] 9 Aug 2017 Vitara Pungpapong a,, Min Zhang b, Dabao
Chris Fraley and Daniel Percival. August 22, 2008, revised May 14, 2010
 Model-Averaged l 1 Regularization using Markov Chain Monte Carlo Model Composition Technical Report No. 541 Department of Statistics, University of Washington Chris Fraley and Daniel Percival August 22,
Model-Averaged l 1 Regularization using Markov Chain Monte Carlo Model Composition Technical Report No. 541 Department of Statistics, University of Washington Chris Fraley and Daniel Percival August 22,
Package icmm. October 12, 2017
 Type Package Package icmm October 12, 2017 Title Empirical Bayes Variable Selection via ICM/M Algorithm Version 1.1 Author Vitara Pungpapong [aut, cre], Min Zhang [aut], Dabao Zhang [aut] Maintainer Vitara
Type Package Package icmm October 12, 2017 Title Empirical Bayes Variable Selection via ICM/M Algorithm Version 1.1 Author Vitara Pungpapong [aut, cre], Min Zhang [aut], Dabao Zhang [aut] Maintainer Vitara
Or How to select variables Using Bayesian LASSO
 Or How to select variables Using Bayesian LASSO x 1 x 2 x 3 x 4 Or How to select variables Using Bayesian LASSO x 1 x 2 x 3 x 4 Or How to select variables Using Bayesian LASSO On Bayesian Variable Selection
Or How to select variables Using Bayesian LASSO x 1 x 2 x 3 x 4 Or How to select variables Using Bayesian LASSO x 1 x 2 x 3 x 4 Or How to select variables Using Bayesian LASSO On Bayesian Variable Selection
Fast Regularization Paths via Coordinate Descent
 August 2008 Trevor Hastie, Stanford Statistics 1 Fast Regularization Paths via Coordinate Descent Trevor Hastie Stanford University joint work with Jerry Friedman and Rob Tibshirani. August 2008 Trevor
August 2008 Trevor Hastie, Stanford Statistics 1 Fast Regularization Paths via Coordinate Descent Trevor Hastie Stanford University joint work with Jerry Friedman and Rob Tibshirani. August 2008 Trevor
Selection of Smoothing Parameter for One-Step Sparse Estimates with L q Penalty
 Journal of Data Science 9(2011), 549-564 Selection of Smoothing Parameter for One-Step Sparse Estimates with L q Penalty Masaru Kanba and Kanta Naito Shimane University Abstract: This paper discusses the
Journal of Data Science 9(2011), 549-564 Selection of Smoothing Parameter for One-Step Sparse Estimates with L q Penalty Masaru Kanba and Kanta Naito Shimane University Abstract: This paper discusses the
Pattern Recognition and Machine Learning
 Christopher M. Bishop Pattern Recognition and Machine Learning ÖSpri inger Contents Preface Mathematical notation Contents vii xi xiii 1 Introduction 1 1.1 Example: Polynomial Curve Fitting 4 1.2 Probability
Christopher M. Bishop Pattern Recognition and Machine Learning ÖSpri inger Contents Preface Mathematical notation Contents vii xi xiii 1 Introduction 1 1.1 Example: Polynomial Curve Fitting 4 1.2 Probability
An Iterated Conditional Modes/Medians Algorithm for Empirical Bayes Selection of Massive Variables
 An Iterated Conditional Modes/Medians Algorithm for Empirical Bayes Selection of Massive Variables Vitara Pungpapong, Min Zhang, Dabao Zhang August 22, 2013 Vitara Pungpapong (email: vitara@cbs.chula.ac.th)
An Iterated Conditional Modes/Medians Algorithm for Empirical Bayes Selection of Massive Variables Vitara Pungpapong, Min Zhang, Dabao Zhang August 22, 2013 Vitara Pungpapong (email: vitara@cbs.chula.ac.th)
Bayesian Regression Linear and Logistic Regression
 When we want more than point estimates Bayesian Regression Linear and Logistic Regression Nicole Beckage Ordinary Least Squares Regression and Lasso Regression return only point estimates But what if we
When we want more than point estimates Bayesian Regression Linear and Logistic Regression Nicole Beckage Ordinary Least Squares Regression and Lasso Regression return only point estimates But what if we
Regularization Path Algorithms for Detecting Gene Interactions
 Regularization Path Algorithms for Detecting Gene Interactions Mee Young Park Trevor Hastie July 16, 2006 Abstract In this study, we consider several regularization path algorithms with grouped variable
Regularization Path Algorithms for Detecting Gene Interactions Mee Young Park Trevor Hastie July 16, 2006 Abstract In this study, we consider several regularization path algorithms with grouped variable
STA414/2104 Statistical Methods for Machine Learning II
 STA414/2104 Statistical Methods for Machine Learning II Murat A. Erdogdu & David Duvenaud Department of Computer Science Department of Statistical Sciences Lecture 3 Slide credits: Russ Salakhutdinov Announcements
STA414/2104 Statistical Methods for Machine Learning II Murat A. Erdogdu & David Duvenaud Department of Computer Science Department of Statistical Sciences Lecture 3 Slide credits: Russ Salakhutdinov Announcements
A Survey of L 1. Regression. Céline Cunen, 20/10/2014. Vidaurre, Bielza and Larranaga (2013)
 A Survey of L 1 Regression Vidaurre, Bielza and Larranaga (2013) Céline Cunen, 20/10/2014 Outline of article 1.Introduction 2.The Lasso for Linear Regression a) Notation and Main Concepts b) Statistical
A Survey of L 1 Regression Vidaurre, Bielza and Larranaga (2013) Céline Cunen, 20/10/2014 Outline of article 1.Introduction 2.The Lasso for Linear Regression a) Notation and Main Concepts b) Statistical
Package LBLGXE. R topics documented: July 20, Type Package
 Type Package Package LBLGXE July 20, 2015 Title Bayesian Lasso for detecting Rare (or Common) Haplotype Association and their interactions with Environmental Covariates Version 1.2 Date 2015-07-09 Author
Type Package Package LBLGXE July 20, 2015 Title Bayesian Lasso for detecting Rare (or Common) Haplotype Association and their interactions with Environmental Covariates Version 1.2 Date 2015-07-09 Author
Penalized likelihood logistic regression with rare events
 Penalized likelihood logistic regression with rare events Georg Heinze 1, Angelika Geroldinger 1, Rainer Puhr 2, Mariana Nold 3, Lara Lusa 4 1 Medical University of Vienna, CeMSIIS, Section for Clinical
Penalized likelihood logistic regression with rare events Georg Heinze 1, Angelika Geroldinger 1, Rainer Puhr 2, Mariana Nold 3, Lara Lusa 4 1 Medical University of Vienna, CeMSIIS, Section for Clinical
Variational Bayesian Inference Techniques
 Advanced Signal Processing 2, SE Variational Bayesian Inference Techniques Johann Steiner 1 Outline Introduction Sparse Signal Reconstruction Sparsity Priors Benefits of Sparse Bayesian Inference Variational
Advanced Signal Processing 2, SE Variational Bayesian Inference Techniques Johann Steiner 1 Outline Introduction Sparse Signal Reconstruction Sparsity Priors Benefits of Sparse Bayesian Inference Variational
A Blockwise Descent Algorithm for Group-penalized Multiresponse and Multinomial Regression
 A Blockwise Descent Algorithm for Group-penalized Multiresponse and Multinomial Regression Noah Simon Jerome Friedman Trevor Hastie November 5, 013 Abstract In this paper we purpose a blockwise descent
A Blockwise Descent Algorithm for Group-penalized Multiresponse and Multinomial Regression Noah Simon Jerome Friedman Trevor Hastie November 5, 013 Abstract In this paper we purpose a blockwise descent
The picasso Package for Nonconvex Regularized M-estimation in High Dimensions in R
 The picasso Package for Nonconvex Regularized M-estimation in High Dimensions in R Xingguo Li Tuo Zhao Tong Zhang Han Liu Abstract We describe an R package named picasso, which implements a unified framework
The picasso Package for Nonconvex Regularized M-estimation in High Dimensions in R Xingguo Li Tuo Zhao Tong Zhang Han Liu Abstract We describe an R package named picasso, which implements a unified framework
Ronald Christensen. University of New Mexico. Albuquerque, New Mexico. Wesley Johnson. University of California, Irvine. Irvine, California
 Texts in Statistical Science Bayesian Ideas and Data Analysis An Introduction for Scientists and Statisticians Ronald Christensen University of New Mexico Albuquerque, New Mexico Wesley Johnson University
Texts in Statistical Science Bayesian Ideas and Data Analysis An Introduction for Scientists and Statisticians Ronald Christensen University of New Mexico Albuquerque, New Mexico Wesley Johnson University
Variable Selection in Structured High-dimensional Covariate Spaces
 Variable Selection in Structured High-dimensional Covariate Spaces Fan Li 1 Nancy Zhang 2 1 Department of Health Care Policy Harvard University 2 Department of Statistics Stanford University May 14 2007
Variable Selection in Structured High-dimensional Covariate Spaces Fan Li 1 Nancy Zhang 2 1 Department of Health Care Policy Harvard University 2 Department of Statistics Stanford University May 14 2007
Fast Regularization Paths via Coordinate Descent
 KDD August 2008 Trevor Hastie, Stanford Statistics 1 Fast Regularization Paths via Coordinate Descent Trevor Hastie Stanford University joint work with Jerry Friedman and Rob Tibshirani. KDD August 2008
KDD August 2008 Trevor Hastie, Stanford Statistics 1 Fast Regularization Paths via Coordinate Descent Trevor Hastie Stanford University joint work with Jerry Friedman and Rob Tibshirani. KDD August 2008
Mixture modelling of recurrent event times with long-term survivors: Analysis of Hutterite birth intervals. John W. Mac McDonald & Alessandro Rosina
 Mixture modelling of recurrent event times with long-term survivors: Analysis of Hutterite birth intervals John W. Mac McDonald & Alessandro Rosina Quantitative Methods in the Social Sciences Seminar -
Mixture modelling of recurrent event times with long-term survivors: Analysis of Hutterite birth intervals John W. Mac McDonald & Alessandro Rosina Quantitative Methods in the Social Sciences Seminar -
A New Bayesian Variable Selection Method: The Bayesian Lasso with Pseudo Variables
 A New Bayesian Variable Selection Method: The Bayesian Lasso with Pseudo Variables Qi Tang (Joint work with Kam-Wah Tsui and Sijian Wang) Department of Statistics University of Wisconsin-Madison Feb. 8,
A New Bayesian Variable Selection Method: The Bayesian Lasso with Pseudo Variables Qi Tang (Joint work with Kam-Wah Tsui and Sijian Wang) Department of Statistics University of Wisconsin-Madison Feb. 8,
Model-Free Knockoffs: High-Dimensional Variable Selection that Controls the False Discovery Rate
 Model-Free Knockoffs: High-Dimensional Variable Selection that Controls the False Discovery Rate Lucas Janson, Stanford Department of Statistics WADAPT Workshop, NIPS, December 2016 Collaborators: Emmanuel
Model-Free Knockoffs: High-Dimensional Variable Selection that Controls the False Discovery Rate Lucas Janson, Stanford Department of Statistics WADAPT Workshop, NIPS, December 2016 Collaborators: Emmanuel
Latent Variable Models for Binary Data. Suppose that for a given vector of explanatory variables x, the latent
 Latent Variable Models for Binary Data Suppose that for a given vector of explanatory variables x, the latent variable, U, has a continuous cumulative distribution function F (u; x) and that the binary
Latent Variable Models for Binary Data Suppose that for a given vector of explanatory variables x, the latent variable, U, has a continuous cumulative distribution function F (u; x) and that the binary
Bi-level feature selection with applications to genetic association
 Bi-level feature selection with applications to genetic association studies October 15, 2008 Motivation In many applications, biological features possess a grouping structure Categorical variables may
Bi-level feature selection with applications to genetic association studies October 15, 2008 Motivation In many applications, biological features possess a grouping structure Categorical variables may
Multilevel Statistical Models: 3 rd edition, 2003 Contents
 Multilevel Statistical Models: 3 rd edition, 2003 Contents Preface Acknowledgements Notation Two and three level models. A general classification notation and diagram Glossary Chapter 1 An introduction
Multilevel Statistical Models: 3 rd edition, 2003 Contents Preface Acknowledgements Notation Two and three level models. A general classification notation and diagram Glossary Chapter 1 An introduction
Sparse Linear Models (10/7/13)
 STA56: Probabilistic machine learning Sparse Linear Models (0/7/) Lecturer: Barbara Engelhardt Scribes: Jiaji Huang, Xin Jiang, Albert Oh Sparsity Sparsity has been a hot topic in statistics and machine
STA56: Probabilistic machine learning Sparse Linear Models (0/7/) Lecturer: Barbara Engelhardt Scribes: Jiaji Huang, Xin Jiang, Albert Oh Sparsity Sparsity has been a hot topic in statistics and machine
25 : Graphical induced structured input/output models
 10-708: Probabilistic Graphical Models 10-708, Spring 2013 25 : Graphical induced structured input/output models Lecturer: Eric P. Xing Scribes: Meghana Kshirsagar (mkshirsa), Yiwen Chen (yiwenche) 1 Graph
10-708: Probabilistic Graphical Models 10-708, Spring 2013 25 : Graphical induced structured input/output models Lecturer: Eric P. Xing Scribes: Meghana Kshirsagar (mkshirsa), Yiwen Chen (yiwenche) 1 Graph
Regularization Parameter Selection for a Bayesian Multi-Level Group Lasso Regression Model with Application to Imaging Genomics
 Regularization Parameter Selection for a Bayesian Multi-Level Group Lasso Regression Model with Application to Imaging Genomics arxiv:1603.08163v1 [stat.ml] 7 Mar 016 Farouk S. Nathoo, Keelin Greenlaw,
Regularization Parameter Selection for a Bayesian Multi-Level Group Lasso Regression Model with Application to Imaging Genomics arxiv:1603.08163v1 [stat.ml] 7 Mar 016 Farouk S. Nathoo, Keelin Greenlaw,
Analysis Methods for Supersaturated Design: Some Comparisons
 Journal of Data Science 1(2003), 249-260 Analysis Methods for Supersaturated Design: Some Comparisons Runze Li 1 and Dennis K. J. Lin 2 The Pennsylvania State University Abstract: Supersaturated designs
Journal of Data Science 1(2003), 249-260 Analysis Methods for Supersaturated Design: Some Comparisons Runze Li 1 and Dennis K. J. Lin 2 The Pennsylvania State University Abstract: Supersaturated designs
Prediction & Feature Selection in GLM
 Tarigan Statistical Consulting & Coaching statistical-coaching.ch Doctoral Program in Computer Science of the Universities of Fribourg, Geneva, Lausanne, Neuchâtel, Bern and the EPFL Hands-on Data Analysis
Tarigan Statistical Consulting & Coaching statistical-coaching.ch Doctoral Program in Computer Science of the Universities of Fribourg, Geneva, Lausanne, Neuchâtel, Bern and the EPFL Hands-on Data Analysis
Bayesian Inference: Principles and Practice 3. Sparse Bayesian Models and the Relevance Vector Machine
 Bayesian Inference: Principles and Practice 3. Sparse Bayesian Models and the Relevance Vector Machine Mike Tipping Gaussian prior Marginal prior: single α Independent α Cambridge, UK Lecture 3: Overview
Bayesian Inference: Principles and Practice 3. Sparse Bayesian Models and the Relevance Vector Machine Mike Tipping Gaussian prior Marginal prior: single α Independent α Cambridge, UK Lecture 3: Overview
Smoothly Clipped Absolute Deviation (SCAD) for Correlated Variables
 Smoothly Clipped Absolute Deviation (SCAD) for Correlated Variables LIB-MA, FSSM Cadi Ayyad University (Morocco) COMPSTAT 2010 Paris, August 22-27, 2010 Motivations Fan and Li (2001), Zou and Li (2008)
Smoothly Clipped Absolute Deviation (SCAD) for Correlated Variables LIB-MA, FSSM Cadi Ayyad University (Morocco) COMPSTAT 2010 Paris, August 22-27, 2010 Motivations Fan and Li (2001), Zou and Li (2008)
ESL Chap3. Some extensions of lasso
 ESL Chap3 Some extensions of lasso 1 Outline Consistency of lasso for model selection Adaptive lasso Elastic net Group lasso 2 Consistency of lasso for model selection A number of authors have studied
ESL Chap3 Some extensions of lasso 1 Outline Consistency of lasso for model selection Adaptive lasso Elastic net Group lasso 2 Consistency of lasso for model selection A number of authors have studied
Graphical Model Selection
 May 6, 2013 Trevor Hastie, Stanford Statistics 1 Graphical Model Selection Trevor Hastie Stanford University joint work with Jerome Friedman, Rob Tibshirani, Rahul Mazumder and Jason Lee May 6, 2013 Trevor
May 6, 2013 Trevor Hastie, Stanford Statistics 1 Graphical Model Selection Trevor Hastie Stanford University joint work with Jerome Friedman, Rob Tibshirani, Rahul Mazumder and Jason Lee May 6, 2013 Trevor
Least Absolute Shrinkage is Equivalent to Quadratic Penalization
 Least Absolute Shrinkage is Equivalent to Quadratic Penalization Yves Grandvalet Heudiasyc, UMR CNRS 6599, Université de Technologie de Compiègne, BP 20.529, 60205 Compiègne Cedex, France Yves.Grandvalet@hds.utc.fr
Least Absolute Shrinkage is Equivalent to Quadratic Penalization Yves Grandvalet Heudiasyc, UMR CNRS 6599, Université de Technologie de Compiègne, BP 20.529, 60205 Compiègne Cedex, France Yves.Grandvalet@hds.utc.fr
10708 Graphical Models: Homework 2
 10708 Graphical Models: Homework 2 Due Monday, March 18, beginning of class Feburary 27, 2013 Instructions: There are five questions (one for extra credit) on this assignment. There is a problem involves
10708 Graphical Models: Homework 2 Due Monday, March 18, beginning of class Feburary 27, 2013 Instructions: There are five questions (one for extra credit) on this assignment. There is a problem involves
Outline of GLMs. Definitions
 Outline of GLMs Definitions This is a short outline of GLM details, adapted from the book Nonparametric Regression and Generalized Linear Models, by Green and Silverman. The responses Y i have density
Outline of GLMs Definitions This is a short outline of GLM details, adapted from the book Nonparametric Regression and Generalized Linear Models, by Green and Silverman. The responses Y i have density
Bayesian construction of perceptrons to predict phenotypes from 584K SNP data.
 Bayesian construction of perceptrons to predict phenotypes from 584K SNP data. Luc Janss, Bert Kappen Radboud University Nijmegen Medical Centre Donders Institute for Neuroscience Introduction Genetic
Bayesian construction of perceptrons to predict phenotypes from 584K SNP data. Luc Janss, Bert Kappen Radboud University Nijmegen Medical Centre Donders Institute for Neuroscience Introduction Genetic
Approximating high-dimensional posteriors with nuisance parameters via integrated rotated Gaussian approximation (IRGA)
 Approximating high-dimensional posteriors with nuisance parameters via integrated rotated Gaussian approximation (IRGA) Willem van den Boom Department of Statistics and Applied Probability National University
Approximating high-dimensional posteriors with nuisance parameters via integrated rotated Gaussian approximation (IRGA) Willem van den Boom Department of Statistics and Applied Probability National University
25 : Graphical induced structured input/output models
 10-708: Probabilistic Graphical Models 10-708, Spring 2016 25 : Graphical induced structured input/output models Lecturer: Eric P. Xing Scribes: Raied Aljadaany, Shi Zong, Chenchen Zhu Disclaimer: A large
10-708: Probabilistic Graphical Models 10-708, Spring 2016 25 : Graphical induced structured input/output models Lecturer: Eric P. Xing Scribes: Raied Aljadaany, Shi Zong, Chenchen Zhu Disclaimer: A large
Frequentist Accuracy of Bayesian Estimates
 Frequentist Accuracy of Bayesian Estimates Bradley Efron Stanford University RSS Journal Webinar Objective Bayesian Inference Probability family F = {f µ (x), µ Ω} Parameter of interest: θ = t(µ) Prior
Frequentist Accuracy of Bayesian Estimates Bradley Efron Stanford University RSS Journal Webinar Objective Bayesian Inference Probability family F = {f µ (x), µ Ω} Parameter of interest: θ = t(µ) Prior
Monte Carlo in Bayesian Statistics
 Monte Carlo in Bayesian Statistics Matthew Thomas SAMBa - University of Bath m.l.thomas@bath.ac.uk December 4, 2014 Matthew Thomas (SAMBa) Monte Carlo in Bayesian Statistics December 4, 2014 1 / 16 Overview
Monte Carlo in Bayesian Statistics Matthew Thomas SAMBa - University of Bath m.l.thomas@bath.ac.uk December 4, 2014 Matthew Thomas (SAMBa) Monte Carlo in Bayesian Statistics December 4, 2014 1 / 16 Overview
Chapter 17: Undirected Graphical Models
 Chapter 17: Undirected Graphical Models The Elements of Statistical Learning Biaobin Jiang Department of Biological Sciences Purdue University bjiang@purdue.edu October 30, 2014 Biaobin Jiang (Purdue)
Chapter 17: Undirected Graphical Models The Elements of Statistical Learning Biaobin Jiang Department of Biological Sciences Purdue University bjiang@purdue.edu October 30, 2014 Biaobin Jiang (Purdue)
Aspects of Feature Selection in Mass Spec Proteomic Functional Data
 Aspects of Feature Selection in Mass Spec Proteomic Functional Data Phil Brown University of Kent Newton Institute 11th December 2006 Based on work with Jeff Morris, and others at MD Anderson, Houston
Aspects of Feature Selection in Mass Spec Proteomic Functional Data Phil Brown University of Kent Newton Institute 11th December 2006 Based on work with Jeff Morris, and others at MD Anderson, Houston
Penalized Loss functions for Bayesian Model Choice
 Penalized Loss functions for Bayesian Model Choice Martyn International Agency for Research on Cancer Lyon, France 13 November 2009 The pure approach For a Bayesian purist, all uncertainty is represented
Penalized Loss functions for Bayesian Model Choice Martyn International Agency for Research on Cancer Lyon, France 13 November 2009 The pure approach For a Bayesian purist, all uncertainty is represented
STA 4273H: Statistical Machine Learning
 STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Computer Science! Department of Statistical Sciences! rsalakhu@cs.toronto.edu! h0p://www.cs.utoronto.ca/~rsalakhu/ Lecture 7 Approximate
STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Computer Science! Department of Statistical Sciences! rsalakhu@cs.toronto.edu! h0p://www.cs.utoronto.ca/~rsalakhu/ Lecture 7 Approximate
Proteomics and Variable Selection
 Proteomics and Variable Selection p. 1/55 Proteomics and Variable Selection Alex Lewin With thanks to Paul Kirk for some graphs Department of Epidemiology and Biostatistics, School of Public Health, Imperial
Proteomics and Variable Selection p. 1/55 Proteomics and Variable Selection Alex Lewin With thanks to Paul Kirk for some graphs Department of Epidemiology and Biostatistics, School of Public Health, Imperial
Bayesian linear regression
 Bayesian linear regression Linear regression is the basis of most statistical modeling. The model is Y i = X T i β + ε i, where Y i is the continuous response X i = (X i1,..., X ip ) T is the corresponding
Bayesian linear regression Linear regression is the basis of most statistical modeling. The model is Y i = X T i β + ε i, where Y i is the continuous response X i = (X i1,..., X ip ) T is the corresponding
Bayesian Methods for Machine Learning
 Bayesian Methods for Machine Learning CS 584: Big Data Analytics Material adapted from Radford Neal s tutorial (http://ftp.cs.utoronto.ca/pub/radford/bayes-tut.pdf), Zoubin Ghahramni (http://hunch.net/~coms-4771/zoubin_ghahramani_bayesian_learning.pdf),
Bayesian Methods for Machine Learning CS 584: Big Data Analytics Material adapted from Radford Neal s tutorial (http://ftp.cs.utoronto.ca/pub/radford/bayes-tut.pdf), Zoubin Ghahramni (http://hunch.net/~coms-4771/zoubin_ghahramani_bayesian_learning.pdf),
Generalized Boosted Models: A guide to the gbm package
 Generalized Boosted Models: A guide to the gbm package Greg Ridgeway April 15, 2006 Boosting takes on various forms with different programs using different loss functions, different base models, and different
Generalized Boosted Models: A guide to the gbm package Greg Ridgeway April 15, 2006 Boosting takes on various forms with different programs using different loss functions, different base models, and different
An Introduction to Graphical Lasso
 An Introduction to Graphical Lasso Bo Chang Graphical Models Reading Group May 15, 2015 Bo Chang (UBC) Graphical Lasso May 15, 2015 1 / 16 Undirected Graphical Models An undirected graph, each vertex represents
An Introduction to Graphical Lasso Bo Chang Graphical Models Reading Group May 15, 2015 Bo Chang (UBC) Graphical Lasso May 15, 2015 1 / 16 Undirected Graphical Models An undirected graph, each vertex represents
Density Estimation. Seungjin Choi
 Density Estimation Seungjin Choi Department of Computer Science and Engineering Pohang University of Science and Technology 77 Cheongam-ro, Nam-gu, Pohang 37673, Korea seungjin@postech.ac.kr http://mlg.postech.ac.kr/
Density Estimation Seungjin Choi Department of Computer Science and Engineering Pohang University of Science and Technology 77 Cheongam-ro, Nam-gu, Pohang 37673, Korea seungjin@postech.ac.kr http://mlg.postech.ac.kr/
Machine Learning Linear Classification. Prof. Matteo Matteucci
 Machine Learning Linear Classification Prof. Matteo Matteucci Recall from the first lecture 2 X R p Regression Y R Continuous Output X R p Y {Ω 0, Ω 1,, Ω K } Classification Discrete Output X R p Y (X)
Machine Learning Linear Classification Prof. Matteo Matteucci Recall from the first lecture 2 X R p Regression Y R Continuous Output X R p Y {Ω 0, Ω 1,, Ω K } Classification Discrete Output X R p Y (X)
Generalized Elastic Net Regression
 Abstract Generalized Elastic Net Regression Geoffroy MOURET Jean-Jules BRAULT Vahid PARTOVINIA This work presents a variation of the elastic net penalization method. We propose applying a combined l 1
Abstract Generalized Elastic Net Regression Geoffroy MOURET Jean-Jules BRAULT Vahid PARTOVINIA This work presents a variation of the elastic net penalization method. We propose applying a combined l 1
STA 4273H: Sta-s-cal Machine Learning
 STA 4273H: Sta-s-cal Machine Learning Russ Salakhutdinov Department of Computer Science! Department of Statistical Sciences! rsalakhu@cs.toronto.edu! h0p://www.cs.utoronto.ca/~rsalakhu/ Lecture 2 In our
STA 4273H: Sta-s-cal Machine Learning Russ Salakhutdinov Department of Computer Science! Department of Statistical Sciences! rsalakhu@cs.toronto.edu! h0p://www.cs.utoronto.ca/~rsalakhu/ Lecture 2 In our
Journal of Statistical Software
 JSS Journal of Statistical Software March 2011, Volume 39, Issue 5. http://www.jstatsoft.org/ Regularization Paths for Cox s Proportional Hazards Model via Coordinate Descent Noah Simon Stanford University
JSS Journal of Statistical Software March 2011, Volume 39, Issue 5. http://www.jstatsoft.org/ Regularization Paths for Cox s Proportional Hazards Model via Coordinate Descent Noah Simon Stanford University
On Bayesian Computation
 On Bayesian Computation Michael I. Jordan with Elaine Angelino, Maxim Rabinovich, Martin Wainwright and Yun Yang Previous Work: Information Constraints on Inference Minimize the minimax risk under constraints
On Bayesian Computation Michael I. Jordan with Elaine Angelino, Maxim Rabinovich, Martin Wainwright and Yun Yang Previous Work: Information Constraints on Inference Minimize the minimax risk under constraints
LASSO Review, Fused LASSO, Parallel LASSO Solvers
 Case Study 3: fmri Prediction LASSO Review, Fused LASSO, Parallel LASSO Solvers Machine Learning for Big Data CSE547/STAT548, University of Washington Sham Kakade May 3, 2016 Sham Kakade 2016 1 Variable
Case Study 3: fmri Prediction LASSO Review, Fused LASSO, Parallel LASSO Solvers Machine Learning for Big Data CSE547/STAT548, University of Washington Sham Kakade May 3, 2016 Sham Kakade 2016 1 Variable
Supplementary Materials for Molecular QTL Discovery Incorporating Genomic Annotations using Bayesian False Discovery Rate Control
 Supplementary Materials for Molecular QTL Discovery Incorporating Genomic Annotations using Bayesian False Discovery Rate Control Xiaoquan Wen Department of Biostatistics, University of Michigan A Model
Supplementary Materials for Molecular QTL Discovery Incorporating Genomic Annotations using Bayesian False Discovery Rate Control Xiaoquan Wen Department of Biostatistics, University of Michigan A Model
Exploratory quantile regression with many covariates: An application to adverse birth outcomes
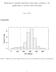 Exploratory quantile regression with many covariates: An application to adverse birth outcomes June 3, 2011 eappendix 30 Percent of Total 20 10 0 0 1000 2000 3000 4000 5000 Birth weights efigure 1: Histogram
Exploratory quantile regression with many covariates: An application to adverse birth outcomes June 3, 2011 eappendix 30 Percent of Total 20 10 0 0 1000 2000 3000 4000 5000 Birth weights efigure 1: Histogram
Package horseshoe. November 8, 2016
 Title Implementation of the Horseshoe Prior Version 0.1.0 Package horseshoe November 8, 2016 Description Contains functions for applying the horseshoe prior to highdimensional linear regression, yielding
Title Implementation of the Horseshoe Prior Version 0.1.0 Package horseshoe November 8, 2016 Description Contains functions for applying the horseshoe prior to highdimensional linear regression, yielding
SCMA292 Mathematical Modeling : Machine Learning. Krikamol Muandet. Department of Mathematics Faculty of Science, Mahidol University.
 SCMA292 Mathematical Modeling : Machine Learning Krikamol Muandet Department of Mathematics Faculty of Science, Mahidol University February 9, 2016 Outline Quick Recap of Least Square Ridge Regression
SCMA292 Mathematical Modeling : Machine Learning Krikamol Muandet Department of Mathematics Faculty of Science, Mahidol University February 9, 2016 Outline Quick Recap of Least Square Ridge Regression
Bayesian Grouped Horseshoe Regression with Application to Additive Models
 Bayesian Grouped Horseshoe Regression with Application to Additive Models Zemei Xu 1,2, Daniel F. Schmidt 1, Enes Makalic 1, Guoqi Qian 2, John L. Hopper 1 1 Centre for Epidemiology and Biostatistics,
Bayesian Grouped Horseshoe Regression with Application to Additive Models Zemei Xu 1,2, Daniel F. Schmidt 1, Enes Makalic 1, Guoqi Qian 2, John L. Hopper 1 1 Centre for Epidemiology and Biostatistics,
Integrated Non-Factorized Variational Inference
 Integrated Non-Factorized Variational Inference Shaobo Han, Xuejun Liao and Lawrence Carin Duke University February 27, 2014 S. Han et al. Integrated Non-Factorized Variational Inference February 27, 2014
Integrated Non-Factorized Variational Inference Shaobo Han, Xuejun Liao and Lawrence Carin Duke University February 27, 2014 S. Han et al. Integrated Non-Factorized Variational Inference February 27, 2014
Bayesian Inference for Discretely Sampled Diffusion Processes: A New MCMC Based Approach to Inference
 Bayesian Inference for Discretely Sampled Diffusion Processes: A New MCMC Based Approach to Inference Osnat Stramer 1 and Matthew Bognar 1 Department of Statistics and Actuarial Science, University of
Bayesian Inference for Discretely Sampled Diffusion Processes: A New MCMC Based Approach to Inference Osnat Stramer 1 and Matthew Bognar 1 Department of Statistics and Actuarial Science, University of
Coupled Hidden Markov Models: Computational Challenges
 .. Coupled Hidden Markov Models: Computational Challenges Louis J. M. Aslett and Chris C. Holmes i-like Research Group University of Oxford Warwick Algorithms Seminar 7 th March 2014 ... Hidden Markov
.. Coupled Hidden Markov Models: Computational Challenges Louis J. M. Aslett and Chris C. Holmes i-like Research Group University of Oxford Warwick Algorithms Seminar 7 th March 2014 ... Hidden Markov
Sparse Gaussian Markov Random Field Mixtures for Anomaly Detection
 Sparse Gaussian Markov Random Field Mixtures for Anomaly Detection Tsuyoshi Idé ( Ide-san ), Ankush Khandelwal*, Jayant Kalagnanam IBM Research, T. J. Watson Research Center (*Currently with University
Sparse Gaussian Markov Random Field Mixtures for Anomaly Detection Tsuyoshi Idé ( Ide-san ), Ankush Khandelwal*, Jayant Kalagnanam IBM Research, T. J. Watson Research Center (*Currently with University
Sparse inverse covariance estimation with the lasso
 Sparse inverse covariance estimation with the lasso Jerome Friedman Trevor Hastie and Robert Tibshirani November 8, 2007 Abstract We consider the problem of estimating sparse graphs by a lasso penalty
Sparse inverse covariance estimation with the lasso Jerome Friedman Trevor Hastie and Robert Tibshirani November 8, 2007 Abstract We consider the problem of estimating sparse graphs by a lasso penalty
Two-stage Adaptive Randomization for Delayed Response in Clinical Trials
 Two-stage Adaptive Randomization for Delayed Response in Clinical Trials Guosheng Yin Department of Statistics and Actuarial Science The University of Hong Kong Joint work with J. Xu PSI and RSS Journal
Two-stage Adaptive Randomization for Delayed Response in Clinical Trials Guosheng Yin Department of Statistics and Actuarial Science The University of Hong Kong Joint work with J. Xu PSI and RSS Journal
CONTRIBUTIONS TO PENALIZED ESTIMATION. Sunyoung Shin
 CONTRIBUTIONS TO PENALIZED ESTIMATION Sunyoung Shin A dissertation submitted to the faculty of the University of North Carolina at Chapel Hill in partial fulfillment of the requirements for the degree
CONTRIBUTIONS TO PENALIZED ESTIMATION Sunyoung Shin A dissertation submitted to the faculty of the University of North Carolina at Chapel Hill in partial fulfillment of the requirements for the degree
Bayesian Analysis of Massive Datasets Via Particle Filters
 Bayesian Analysis of Massive Datasets Via Particle Filters Bayesian Analysis Use Bayes theorem to learn about model parameters from data Examples: Clustered data: hospitals, schools Spatial models: public
Bayesian Analysis of Massive Datasets Via Particle Filters Bayesian Analysis Use Bayes theorem to learn about model parameters from data Examples: Clustered data: hospitals, schools Spatial models: public
Building a Prognostic Biomarker
 Building a Prognostic Biomarker Noah Simon and Richard Simon July 2016 1 / 44 Prognostic Biomarker for a Continuous Measure On each of n patients measure y i - single continuous outcome (eg. blood pressure,
Building a Prognostic Biomarker Noah Simon and Richard Simon July 2016 1 / 44 Prognostic Biomarker for a Continuous Measure On each of n patients measure y i - single continuous outcome (eg. blood pressure,
LARGE NUMBERS OF EXPLANATORY VARIABLES. H.S. Battey. WHAO-PSI, St Louis, 9 September 2018
 LARGE NUMBERS OF EXPLANATORY VARIABLES HS Battey Department of Mathematics, Imperial College London WHAO-PSI, St Louis, 9 September 2018 Regression, broadly defined Response variable Y i, eg, blood pressure,
LARGE NUMBERS OF EXPLANATORY VARIABLES HS Battey Department of Mathematics, Imperial College London WHAO-PSI, St Louis, 9 September 2018 Regression, broadly defined Response variable Y i, eg, blood pressure,
Learning with Sparsity Constraints
 Stanford 2010 Trevor Hastie, Stanford Statistics 1 Learning with Sparsity Constraints Trevor Hastie Stanford University recent joint work with Rahul Mazumder, Jerome Friedman and Rob Tibshirani earlier
Stanford 2010 Trevor Hastie, Stanford Statistics 1 Learning with Sparsity Constraints Trevor Hastie Stanford University recent joint work with Rahul Mazumder, Jerome Friedman and Rob Tibshirani earlier
Association studies and regression
 Association studies and regression CM226: Machine Learning for Bioinformatics. Fall 2016 Sriram Sankararaman Acknowledgments: Fei Sha, Ameet Talwalkar Association studies and regression 1 / 104 Administration
Association studies and regression CM226: Machine Learning for Bioinformatics. Fall 2016 Sriram Sankararaman Acknowledgments: Fei Sha, Ameet Talwalkar Association studies and regression 1 / 104 Administration
STA 4273H: Statistical Machine Learning
 STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 7 Approximate
STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 7 Approximate
Logistic Regression. Seungjin Choi
 Logistic Regression Seungjin Choi Department of Computer Science and Engineering Pohang University of Science and Technology 77 Cheongam-ro, Nam-gu, Pohang 37673, Korea seungjin@postech.ac.kr http://mlg.postech.ac.kr/
Logistic Regression Seungjin Choi Department of Computer Science and Engineering Pohang University of Science and Technology 77 Cheongam-ro, Nam-gu, Pohang 37673, Korea seungjin@postech.ac.kr http://mlg.postech.ac.kr/
Markov Chain Monte Carlo in Practice
 Markov Chain Monte Carlo in Practice Edited by W.R. Gilks Medical Research Council Biostatistics Unit Cambridge UK S. Richardson French National Institute for Health and Medical Research Vilejuif France
Markov Chain Monte Carlo in Practice Edited by W.R. Gilks Medical Research Council Biostatistics Unit Cambridge UK S. Richardson French National Institute for Health and Medical Research Vilejuif France
Bagging During Markov Chain Monte Carlo for Smoother Predictions
 Bagging During Markov Chain Monte Carlo for Smoother Predictions Herbert K. H. Lee University of California, Santa Cruz Abstract: Making good predictions from noisy data is a challenging problem. Methods
Bagging During Markov Chain Monte Carlo for Smoother Predictions Herbert K. H. Lee University of California, Santa Cruz Abstract: Making good predictions from noisy data is a challenging problem. Methods
Multistate Modeling and Applications
 Multistate Modeling and Applications Yang Yang Department of Statistics University of Michigan, Ann Arbor IBM Research Graduate Student Workshop: Statistics for a Smarter Planet Yang Yang (UM, Ann Arbor)
Multistate Modeling and Applications Yang Yang Department of Statistics University of Michigan, Ann Arbor IBM Research Graduate Student Workshop: Statistics for a Smarter Planet Yang Yang (UM, Ann Arbor)
Outlier detection and variable selection via difference based regression model and penalized regression
 Journal of the Korean Data & Information Science Society 2018, 29(3), 815 825 http://dx.doi.org/10.7465/jkdi.2018.29.3.815 한국데이터정보과학회지 Outlier detection and variable selection via difference based regression
Journal of the Korean Data & Information Science Society 2018, 29(3), 815 825 http://dx.doi.org/10.7465/jkdi.2018.29.3.815 한국데이터정보과학회지 Outlier detection and variable selection via difference based regression
Who was Bayes? Bayesian Phylogenetics. What is Bayes Theorem?
 Who was Bayes? Bayesian Phylogenetics Bret Larget Departments of Botany and of Statistics University of Wisconsin Madison October 6, 2011 The Reverand Thomas Bayes was born in London in 1702. He was the
Who was Bayes? Bayesian Phylogenetics Bret Larget Departments of Botany and of Statistics University of Wisconsin Madison October 6, 2011 The Reverand Thomas Bayes was born in London in 1702. He was the
TECHNICAL REPORT NO. 1091r. A Note on the Lasso and Related Procedures in Model Selection
 DEPARTMENT OF STATISTICS University of Wisconsin 1210 West Dayton St. Madison, WI 53706 TECHNICAL REPORT NO. 1091r April 2004, Revised December 2004 A Note on the Lasso and Related Procedures in Model
DEPARTMENT OF STATISTICS University of Wisconsin 1210 West Dayton St. Madison, WI 53706 TECHNICAL REPORT NO. 1091r April 2004, Revised December 2004 A Note on the Lasso and Related Procedures in Model
Bayesian Phylogenetics
 Bayesian Phylogenetics Bret Larget Departments of Botany and of Statistics University of Wisconsin Madison October 6, 2011 Bayesian Phylogenetics 1 / 27 Who was Bayes? The Reverand Thomas Bayes was born
Bayesian Phylogenetics Bret Larget Departments of Botany and of Statistics University of Wisconsin Madison October 6, 2011 Bayesian Phylogenetics 1 / 27 Who was Bayes? The Reverand Thomas Bayes was born
arxiv: v1 [stat.me] 6 Jul 2017
![arxiv: v1 [stat.me] 6 Jul 2017 arxiv: v1 [stat.me] 6 Jul 2017](/thumbs/89/98510504.jpg) Sparsity information and regularization in the horseshoe and other shrinkage priors arxiv:77.694v [stat.me] 6 Jul 7 Juho Piironen and Aki Vehtari Helsinki Institute for Information Technology, HIIT Department
Sparsity information and regularization in the horseshoe and other shrinkage priors arxiv:77.694v [stat.me] 6 Jul 7 Juho Piironen and Aki Vehtari Helsinki Institute for Information Technology, HIIT Department
Part 1: Expectation Propagation
 Chalmers Machine Learning Summer School Approximate message passing and biomedicine Part 1: Expectation Propagation Tom Heskes Machine Learning Group, Institute for Computing and Information Sciences Radboud
Chalmers Machine Learning Summer School Approximate message passing and biomedicine Part 1: Expectation Propagation Tom Heskes Machine Learning Group, Institute for Computing and Information Sciences Radboud
SOLVING NON-CONVEX LASSO TYPE PROBLEMS WITH DC PROGRAMMING. Gilles Gasso, Alain Rakotomamonjy and Stéphane Canu
 SOLVING NON-CONVEX LASSO TYPE PROBLEMS WITH DC PROGRAMMING Gilles Gasso, Alain Rakotomamonjy and Stéphane Canu LITIS - EA 48 - INSA/Universite de Rouen Avenue de l Université - 768 Saint-Etienne du Rouvray
SOLVING NON-CONVEX LASSO TYPE PROBLEMS WITH DC PROGRAMMING Gilles Gasso, Alain Rakotomamonjy and Stéphane Canu LITIS - EA 48 - INSA/Universite de Rouen Avenue de l Université - 768 Saint-Etienne du Rouvray
Probabilistic Graphical Models
 School of Computer Science Probabilistic Graphical Models Gaussian graphical models and Ising models: modeling networks Eric Xing Lecture 0, February 7, 04 Reading: See class website Eric Xing @ CMU, 005-04
School of Computer Science Probabilistic Graphical Models Gaussian graphical models and Ising models: modeling networks Eric Xing Lecture 0, February 7, 04 Reading: See class website Eric Xing @ CMU, 005-04
Fast Regularization Paths via Coordinate Descent
 user! 2009 Trevor Hastie, Stanford Statistics 1 Fast Regularization Paths via Coordinate Descent Trevor Hastie Stanford University joint work with Jerome Friedman and Rob Tibshirani. user! 2009 Trevor
user! 2009 Trevor Hastie, Stanford Statistics 1 Fast Regularization Paths via Coordinate Descent Trevor Hastie Stanford University joint work with Jerome Friedman and Rob Tibshirani. user! 2009 Trevor
Learning in Bayesian Networks
 Learning in Bayesian Networks Florian Markowetz Max-Planck-Institute for Molecular Genetics Computational Molecular Biology Berlin Berlin: 20.06.2002 1 Overview 1. Bayesian Networks Stochastic Networks
Learning in Bayesian Networks Florian Markowetz Max-Planck-Institute for Molecular Genetics Computational Molecular Biology Berlin Berlin: 20.06.2002 1 Overview 1. Bayesian Networks Stochastic Networks
The lasso, persistence, and cross-validation
 The lasso, persistence, and cross-validation Daniel J. McDonald Department of Statistics Indiana University http://www.stat.cmu.edu/ danielmc Joint work with: Darren Homrighausen Colorado State University
The lasso, persistence, and cross-validation Daniel J. McDonald Department of Statistics Indiana University http://www.stat.cmu.edu/ danielmc Joint work with: Darren Homrighausen Colorado State University
Knockoffs as Post-Selection Inference
 Knockoffs as Post-Selection Inference Lucas Janson Harvard University Department of Statistics blank line blank line WHOA-PSI, August 12, 2017 Controlled Variable Selection Conditional modeling setup:
Knockoffs as Post-Selection Inference Lucas Janson Harvard University Department of Statistics blank line blank line WHOA-PSI, August 12, 2017 Controlled Variable Selection Conditional modeling setup:
Expression Data Exploration: Association, Patterns, Factors & Regression Modelling
 Expression Data Exploration: Association, Patterns, Factors & Regression Modelling Exploring gene expression data Scale factors, median chip correlation on gene subsets for crude data quality investigation
Expression Data Exploration: Association, Patterns, Factors & Regression Modelling Exploring gene expression data Scale factors, median chip correlation on gene subsets for crude data quality investigation
Disease mapping with Gaussian processes
 EUROHEIS2 Kuopio, Finland 17-18 August 2010 Aki Vehtari (former Helsinki University of Technology) Department of Biomedical Engineering and Computational Science (BECS) Acknowledgments Researchers - Jarno
EUROHEIS2 Kuopio, Finland 17-18 August 2010 Aki Vehtari (former Helsinki University of Technology) Department of Biomedical Engineering and Computational Science (BECS) Acknowledgments Researchers - Jarno
Bayesian Inference of Interactions and Associations
 Bayesian Inference of Interactions and Associations Jun Liu Department of Statistics Harvard University http://www.fas.harvard.edu/~junliu Based on collaborations with Yu Zhang, Jing Zhang, Yuan Yuan,
Bayesian Inference of Interactions and Associations Jun Liu Department of Statistics Harvard University http://www.fas.harvard.edu/~junliu Based on collaborations with Yu Zhang, Jing Zhang, Yuan Yuan,
Introductory Bayesian Analysis
 Introductory Bayesian Analysis Jaya M. Satagopan Memorial Sloan-Kettering Cancer Center Weill Cornell Medical College (Affiliate) satagopj@mskcc.org March 14, 2013 Bayesian Analysis Fit probability models
Introductory Bayesian Analysis Jaya M. Satagopan Memorial Sloan-Kettering Cancer Center Weill Cornell Medical College (Affiliate) satagopj@mskcc.org March 14, 2013 Bayesian Analysis Fit probability models
Model Selection for Semiparametric Bayesian Models with Application to Overdispersion
 Proceedings 59th ISI World Statistics Congress, 25-30 August 2013, Hong Kong (Session CPS020) p.3863 Model Selection for Semiparametric Bayesian Models with Application to Overdispersion Jinfang Wang and
Proceedings 59th ISI World Statistics Congress, 25-30 August 2013, Hong Kong (Session CPS020) p.3863 Model Selection for Semiparametric Bayesian Models with Application to Overdispersion Jinfang Wang and
Robust Variable Selection Through MAVE
 Robust Variable Selection Through MAVE Weixin Yao and Qin Wang Abstract Dimension reduction and variable selection play important roles in high dimensional data analysis. Wang and Yin (2008) proposed sparse
Robust Variable Selection Through MAVE Weixin Yao and Qin Wang Abstract Dimension reduction and variable selection play important roles in high dimensional data analysis. Wang and Yin (2008) proposed sparse
MS&E 226: Small Data
 MS&E 226: Small Data Lecture 12: Logistic regression (v1) Ramesh Johari ramesh.johari@stanford.edu Fall 2015 1 / 30 Regression methods for binary outcomes 2 / 30 Binary outcomes For the duration of this
MS&E 226: Small Data Lecture 12: Logistic regression (v1) Ramesh Johari ramesh.johari@stanford.edu Fall 2015 1 / 30 Regression methods for binary outcomes 2 / 30 Binary outcomes For the duration of this
