The gpca Package for Identifying Batch Effects in High-Throughput Genomic Data
|
|
|
- Vivien Barnett
- 6 years ago
- Views:
Transcription
1 The gpca Package for Identifying Batch Effects in High-Throughput Genomic Data Sarah Reese July 31, 2013 Batch effects are commonly observed systematic non-biological variation between groups of samples due to experimental artifacts, such as processing date, lab, or technician. Combining samples from multiple batches can cause the true biological variation in a high-throughput experiment to be obscured by variation due to batch. 1 Guided Principal Components Analysis Guided principal components analysis (gpca) is an extension of principal components analysis (PCA) that replaces the data X matrix in the singular value decomposition (SVD) of PCA with Y X such that Y X = UDV where Y is an n b indicator matrix where n denotes sample and b denotes batch. For k = 1,..., b batches, each is comprised of n k observations such that b k=1 n k = n. The indicator matrix consists of b blocks with n k rows for k = 1,..., b, and k columns where, for each block, { 1 if k = b Y k = 0 otherwise. Performing SVD on Y X results in a b b batch loadings matrix U and a p p probe loadings matrix V. Large singular values (the diagonal elements of the q q matrix D where q = min(n, p)) imply that the batch is important for the corresponding principal component. gpca guides the SVD to look for batch effects in the data based on the batch indicator matrix Y, which can be defined to indicate any type of potential batch effect, such as time of hybridization, plate, or other experimental artifact. In Reese et al. [6], we proposed a test statistic δ that quantifies the proportion of variance due to batch effects in experimental genomic data. The proportion of total variance due to batch is taken to be the ratio of the variance of the first principal component from gpca to the variance of the first principal component from unguided PCA reesese@vcu.edu δ = var(xv g 1 ) var(xv u1 ) 1
2 where g indicates gpca and u indicates unguided PCA. V is the matrix of probe loadings resulting from gpca or PCA, respectively. Large values of δ (values near 1) imply that the batch effect is large. To determine whether δ is significantly larger than would be expected by chance, a p- value is estimated using a permutation distribution created by permuting the batch vector M = 1000 times so that δ pm is computed for m = 1,..., M where p indicates the permutation. Here δ pm is the proportion of the total variance due to the first principal component from the m th permutation from gpca to the total variance due to the first principal component from the m th permutation from unguided PCA. A one-sided p-value (testing H 0 : δ pm = δ versus H 1 : δ pm > δ) is estimated as the proportion of times the observed δ was in the extreme tail of the permutation distribution p-value = For more details on gpca see Reese et al. [6]. 2 R Package M m=1 (δ p m > δ). M The gpca package includes four example data sets, the gpca.batchdetect() function that produces the δ statistic and corresponding p-value, and additional visualization functions. 2.1 Data Four data sets are included in the gpca package, three simulated data sets and one case study data set. The case study data (data(casedat)) contains copy number variation data with n = 500 observations and p = 1000 features that were retained after a variance filter was applied. The simulated data represents copy number data under three scenarios: (1) feature data (here, feature denotes probe) with no phenotypic variable (data(nophedat)); (2) feature data with a high variance phenotypic variable (data(highphedat)); and (3) feature data with a low variance phenotypic variable (data(lowphedat)). The feature data were generated independently from a multivariate normal distribution with 1000 features and 90 observations. Data with two batches and two phenotypes were simulated. Batch mean vectors µ b1 = 0 and µ b2 = 1 and batch variance σb 2I where σ2 b = 0.5 were used to simulate the data. The proportion of features affected by batch was bprop = 0.01 for the no phenotype scenario and bprop = 0.05 for the high and low variance phenotype scenarios. For the scenarios with phenotypic effects, the proportion of features affected by phenotype was pprop = 0.1. The phenotypic mean vectors were µ p1 = 0 and µ p2 = 1 and the phenotypic variance was σpi 2 where σp 2 = 2 for the high variance phenotype scenario and σp 2 = 0.2 for the low variance phenotype scenario. Reese et al. [6] provides an in depth description of the data simulations. For all four data sets, the first column of the data frame containing the data contains the batch vector which indicates batch for the n observations. The rest of the data frame contains the uncentered feature data. 2
3 2.2 Application The δ statistic, corresponding p-value from the permutation test, and various other measures are output by the gpca.batchdetect() function. The syntax for this function is > out<-gpca.batchdetect(x=data,batch=batch,center=false, + scaley=false,filt=null,nperm=1000,seed=null) where x is the n p matrix of feature data X, batch is a length n vector indicating batch which is used to calculate the Y matrix for gpca. The option center is a logical indicating whether or not data is centered where center=true if the data x is already centered. scaley is a logical indicating whether the batch indicator matrix Y is to be scaled by the batch sample size n k. nperm indicates how many permutations will be used for calculating the permutation test statistic (defaults to 1000), filt gives the number of features to retain when applying a variance-based filter to the data (defaults to NULL indicating no filter applied), and seed sets set.seed(seed). Note that x must be complete data (i.e. contain no missing values) and the class of x must be "matrix". The function, when run actively, will ask if mean-value imputation should be performed for any missing values, but when run passively will cause an error. The gpca.batchdetect() function outputs the value of the statistic δ, the associated p-value, the batch vector batch, the M values of δ p resulting from the permutation test, the proportion of variance associated with the first principal component from unguided (PCu) and guided (PCg) PCA, as well as the cumulative variance associated with all n principal components resulting from unguided PCA (cumulative.var.x) and the cumulative variance associated with all b principal components resulting from gpca (cumulative.var.g). The gpca package also has three functions to visualize the data. The function gdist produces a density plot of the δ p values output by the gpca.batchdetect function. The function PCplot produces principal component plots of either the unguided or guided principal components and allows for either directly comparing the first two principal components, or comparing the first npcs principal components. Finally, the function CumulativeVarPlot produces a plot of the cumulative variance from guided or unguided PCA. > gdist(out) > PCplot(out,ug="guided",type="1v2") > PCplot(out,ug="guided",type="comp",npcs=3) > CumulativeVarPlot(out,ug="unguided",col="blue") 3 Example We will discuss a brief example using casedat data from the gpca package. We first load the data casedat and assign the first column to batch. The rest of the data frame is the feature data, so we assign that to dat and re-classify it as a matrix. Since the casedat feature data is already centered, we set center=true. The value of the test statistic δ and the corresponding p-value are easily printed and the percent of total variation that is explained by batch is calculated. 3
4 > data(casedat) > batch<-casedat$batch > data<-casedat$data > out<-gpca.batchdetect(x=data,batch=batch,center=true) > out$delta ; out$p.val [1] [1] "<0.001" > ((out$varpcg1-out$varpcu1)/out$varpcg1)*100 [1] We can also plot the distribution of the δ p values from the permutation test and see where our test statistic δ (represented by the red dashed line) falls in comparison (Figure 1). Plots of the first versus the second principal components from gpca can be plotted (Figure 2) as well as a sample of the first few principal comparisons (Figure 3). 4 Conclusion The gpca package provides functionality to test for batch effects in high-throughput genomic data using the function gpca.batchdetect(). The ability to detect batch effects in genomic data allows further batch correction procedures such as batch mean-centering [7], distance weighted discrimination (DWD) [1, 2, 3, 5], or empirical Bayes [4], to be employed to attempt to remove the unwanted variation due to batch effects. However, correcting for batch when there is no significant batch effect may result in removing biological variation instead of the systematic non-biological variation due to batch. This package provides the ability to perform a test to detect batch effects. 5 Session Info > sessioninfo() R version ( ) Platform: x86_64-apple-darwin (64-bit) locale: [1] C/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8 attached base packages: [1] stats graphics grdevices utils datasets methods base other attached packages: 4
5 > gdist(out) Distribution of delta values from permutations (delta=0.572; p value=<0.001) Density N = 1000 Bandwidth = Figure 1: Distribution plot of δ p values 5
6 > par(mai=c(0.8,0.8,0.1,0.1),cex=0.8) > PCplot(out,ug="guided",type="1v2") PC 1 PC 2 Figure 2: Principal components plot of first two principal components from gpca 6
7 > par(mai=c(0.65,0.65,0.1,0.1),cex=0.8) > PCplot(out,ug="guided",type="comp",npcs=3) Density PC 1 PC PC 1 PC3 PC 2 PC Density PC 2 PC PC 3 PC PC 3 PC Density Figure 3: Principal components plots of the first three principal components with density plots of the principal components on the diagonal. 7
8 [1] gpca_1.0 loaded via a namespace (and not attached): [1] tools_3.0.1 References [1] Benito, M., Parker, J., Du, Q., Wu, J., Xiang, D., P., C. M., and Marron, J. S. Adjustment of systematic microarray data biases. Bioinformatics, 20(1): , [2] Huang, H., Lu, X., Liu, Y., Haaland, P., and Marron, J. R/DWD: distance-weighted discrimination for classification, visualization and batch adjustment. Bioinformatics, 28(8): , [3] Huang, H., Liu, Y., Du, Y., Perou, C. M., Hayes, D. N., Todd, M. J., and Marron, J. S. Multiclass distance weighted discrimination. Journal of Computational and Graphical Statistics, [4] Johnson, W. E., Li, C., and Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics, 8(1): , [5] Marron, J. S. and Todd, M. Distance weighted discrimination. Aug [6] Reese, S. E., Archer, K. J., Therneau, T. M., Atkinson, E. J., Vachon, C. M., de Andrade, M., Kocher, J. A., and Eckel-Passow, J. E. A new statistic for identifying batch effects in high-throughput genomic data that uses guided principal components analysis. Bioinformatics, under review. [7] Sims, A. H., Smethurst, G. J., Hey, Y., Okoniewski, M. J., Pepper, S. D., Howell, A., Miller, C. J., and Clarke, R. B. The removal of multiplicative, systematic bias allows integration of breast cancer gene expression datasets improving meta-analysis and prediction of prognosis. BMC Medical Genomics, 1(42),
Lecture 6: Methods for high-dimensional problems
 Lecture 6: Methods for high-dimensional problems Hector Corrada Bravo and Rafael A. Irizarry March, 2010 In this Section we will discuss methods where data lies on high-dimensional spaces. In particular,
Lecture 6: Methods for high-dimensional problems Hector Corrada Bravo and Rafael A. Irizarry March, 2010 In this Section we will discuss methods where data lies on high-dimensional spaces. In particular,
Manual: R package HTSmix
 Manual: R package HTSmix Olga Vitek and Danni Yu May 2, 2011 1 Overview High-throughput screens (HTS) measure phenotypes of thousands of biological samples under various conditions. The phenotypes are
Manual: R package HTSmix Olga Vitek and Danni Yu May 2, 2011 1 Overview High-throughput screens (HTS) measure phenotypes of thousands of biological samples under various conditions. The phenotypes are
Conditional variable importance in R package extendedforest
 Conditional variable importance in R package extendedforest Stephen J. Smith, Nick Ellis, C. Roland Pitcher February 10, 2011 Contents 1 Introduction 1 2 Methods 2 2.1 Conditional permutation................................
Conditional variable importance in R package extendedforest Stephen J. Smith, Nick Ellis, C. Roland Pitcher February 10, 2011 Contents 1 Introduction 1 2 Methods 2 2.1 Conditional permutation................................
Introduction to Machine Learning. PCA and Spectral Clustering. Introduction to Machine Learning, Slides: Eran Halperin
 1 Introduction to Machine Learning PCA and Spectral Clustering Introduction to Machine Learning, 2013-14 Slides: Eran Halperin Singular Value Decomposition (SVD) The singular value decomposition (SVD)
1 Introduction to Machine Learning PCA and Spectral Clustering Introduction to Machine Learning, 2013-14 Slides: Eran Halperin Singular Value Decomposition (SVD) The singular value decomposition (SVD)
Lecture 4: Principal Component Analysis and Linear Dimension Reduction
 Lecture 4: Principal Component Analysis and Linear Dimension Reduction Advanced Applied Multivariate Analysis STAT 2221, Fall 2013 Sungkyu Jung Department of Statistics University of Pittsburgh E-mail:
Lecture 4: Principal Component Analysis and Linear Dimension Reduction Advanced Applied Multivariate Analysis STAT 2221, Fall 2013 Sungkyu Jung Department of Statistics University of Pittsburgh E-mail:
Statistical Methods for Analysis of Genetic Data
 Statistical Methods for Analysis of Genetic Data Christopher R. Cabanski A dissertation submitted to the faculty of the University of North Carolina at Chapel Hill in partial fulfillment of the requirements
Statistical Methods for Analysis of Genetic Data Christopher R. Cabanski A dissertation submitted to the faculty of the University of North Carolina at Chapel Hill in partial fulfillment of the requirements
Principal Component Analysis-I Geog 210C Introduction to Spatial Data Analysis. Chris Funk. Lecture 17
 Principal Component Analysis-I Geog 210C Introduction to Spatial Data Analysis Chris Funk Lecture 17 Outline Filters and Rotations Generating co-varying random fields Translating co-varying fields into
Principal Component Analysis-I Geog 210C Introduction to Spatial Data Analysis Chris Funk Lecture 17 Outline Filters and Rotations Generating co-varying random fields Translating co-varying fields into
Multiclass Distance Weighted Discrimination with Applications to Batch Adjustment
 Multiclass Distance Weighted Discrimination with Applications to Batch Adjustment Hanwen Huang 1, Yufeng Liu 1, Michael J. Todd 2, Ying Du 3, Charles M. Perou 3, D. Neil Hayes 3, and J. S. Marron 1 November
Multiclass Distance Weighted Discrimination with Applications to Batch Adjustment Hanwen Huang 1, Yufeng Liu 1, Michael J. Todd 2, Ying Du 3, Charles M. Perou 3, D. Neil Hayes 3, and J. S. Marron 1 November
Principal Component Analysis. Applied Multivariate Statistics Spring 2012
 Principal Component Analysis Applied Multivariate Statistics Spring 2012 Overview Intuition Four definitions Practical examples Mathematical example Case study 2 PCA: Goals Goal 1: Dimension reduction
Principal Component Analysis Applied Multivariate Statistics Spring 2012 Overview Intuition Four definitions Practical examples Mathematical example Case study 2 PCA: Goals Goal 1: Dimension reduction
Exercises for Applied Predictive Modeling Chapter 6 Linear Regression and Its Cousins
 Exercises for Applied Predictive Modeling Chapter 6 Linear Regression and Its Cousins Max Kuhn, Kjell Johnson Version 1 January 8, 2015 The solutions in this file uses several R packages not used in the
Exercises for Applied Predictive Modeling Chapter 6 Linear Regression and Its Cousins Max Kuhn, Kjell Johnson Version 1 January 8, 2015 The solutions in this file uses several R packages not used in the
Latent Variable Methods for the Analysis of Genomic Data
 John D. Storey Center for Statistics and Machine Learning & Lewis-Sigler Institute for Integrative Genomics Latent Variable Methods for the Analysis of Genomic Data http://genomine.org/talks/ Data m variables
John D. Storey Center for Statistics and Machine Learning & Lewis-Sigler Institute for Integrative Genomics Latent Variable Methods for the Analysis of Genomic Data http://genomine.org/talks/ Data m variables
DISCUSSION OF INFLUENTIAL FEATURE PCA FOR HIGH DIMENSIONAL CLUSTERING. By T. Tony Cai and Linjun Zhang University of Pennsylvania
 Submitted to the Annals of Statistics DISCUSSION OF INFLUENTIAL FEATURE PCA FOR HIGH DIMENSIONAL CLUSTERING By T. Tony Cai and Linjun Zhang University of Pennsylvania We would like to congratulate the
Submitted to the Annals of Statistics DISCUSSION OF INFLUENTIAL FEATURE PCA FOR HIGH DIMENSIONAL CLUSTERING By T. Tony Cai and Linjun Zhang University of Pennsylvania We would like to congratulate the
Principal Component Analysis, A Powerful Scoring Technique
 Principal Component Analysis, A Powerful Scoring Technique George C. J. Fernandez, University of Nevada - Reno, Reno NV 89557 ABSTRACT Data mining is a collection of analytical techniques to uncover new
Principal Component Analysis, A Powerful Scoring Technique George C. J. Fernandez, University of Nevada - Reno, Reno NV 89557 ABSTRACT Data mining is a collection of analytical techniques to uncover new
Introduction to multivariate analysis Outline
 Introduction to multivariate analysis Outline Why do a multivariate analysis Ordination, classification, model fitting Principal component analysis Discriminant analysis, quickly Species presence/absence
Introduction to multivariate analysis Outline Why do a multivariate analysis Ordination, classification, model fitting Principal component analysis Discriminant analysis, quickly Species presence/absence
Design of Microarray Experiments. Xiangqin Cui
 Design of Microarray Experiments Xiangqin Cui Experimental design Experimental design: is a term used about efficient methods for planning the collection of data, in order to obtain the maximum amount
Design of Microarray Experiments Xiangqin Cui Experimental design Experimental design: is a term used about efficient methods for planning the collection of data, in order to obtain the maximum amount
Correcting gene expression data when neither the unwanted variation nor the factor of interest are observed
 Correcting gene expression data when neither the unwanted variation nor the factor of interest are observed Laurent Jacob 1 laurent@stat.berkeley.edu Johann Gagnon-Bartsch 1 johann@berkeley.edu Terence
Correcting gene expression data when neither the unwanted variation nor the factor of interest are observed Laurent Jacob 1 laurent@stat.berkeley.edu Johann Gagnon-Bartsch 1 johann@berkeley.edu Terence
CLASS-SENSITIVE PRINCIPAL COMPONENTS ANALYSIS. Di Miao. Chapel Hill 2015
 CLASS-SENSITIVE PRINCIPAL COMPONENTS ANALYSIS Di Miao A dissertation submitted to the faculty at the University of North Carolina at Chapel Hill in partial fulfillment of the requirements for the degree
CLASS-SENSITIVE PRINCIPAL COMPONENTS ANALYSIS Di Miao A dissertation submitted to the faculty at the University of North Carolina at Chapel Hill in partial fulfillment of the requirements for the degree
Lecture 13. Principal Component Analysis. Brett Bernstein. April 25, CDS at NYU. Brett Bernstein (CDS at NYU) Lecture 13 April 25, / 26
 Principal Component Analysis Brett Bernstein CDS at NYU April 25, 2017 Brett Bernstein (CDS at NYU) Lecture 13 April 25, 2017 1 / 26 Initial Question Intro Question Question Let S R n n be symmetric. 1
Principal Component Analysis Brett Bernstein CDS at NYU April 25, 2017 Brett Bernstein (CDS at NYU) Lecture 13 April 25, 2017 1 / 26 Initial Question Intro Question Question Let S R n n be symmetric. 1
Using the tmle.npvi R package
 Using the tmle.npvi R package Antoine Chambaz Pierre Neuvial Package version 0.10.0 Date 2015-05-13 Contents 1 Citing tmle.npvi 1 2 The non-parametric variable importance parameter 2 3 Using the tmle.npvi
Using the tmle.npvi R package Antoine Chambaz Pierre Neuvial Package version 0.10.0 Date 2015-05-13 Contents 1 Citing tmle.npvi 1 2 The non-parametric variable importance parameter 2 3 Using the tmle.npvi
R-companion to: Estimation of the Thurstonian model for the 2-AC protocol
 R-companion to: Estimation of the Thurstonian model for the 2-AC protocol Rune Haubo Bojesen Christensen, Hye-Seong Lee & Per Bruun Brockhoff August 24, 2017 This document describes how the examples in
R-companion to: Estimation of the Thurstonian model for the 2-AC protocol Rune Haubo Bojesen Christensen, Hye-Seong Lee & Per Bruun Brockhoff August 24, 2017 This document describes how the examples in
The Third International Workshop in Sequential Methodologies
 Area C.6.1: Wednesday, June 15, 4:00pm Kazuyoshi Yata Institute of Mathematics, University of Tsukuba, Japan Effective PCA for large p, small n context with sample size determination In recent years, substantial
Area C.6.1: Wednesday, June 15, 4:00pm Kazuyoshi Yata Institute of Mathematics, University of Tsukuba, Japan Effective PCA for large p, small n context with sample size determination In recent years, substantial
statistical methods for tailoring seasonal climate forecasts Andrew W. Robertson, IRI
 statistical methods for tailoring seasonal climate forecasts Andrew W. Robertson, IRI tailored seasonal forecasts why do we make probabilistic forecasts? to reduce our uncertainty about the (unknown) future
statistical methods for tailoring seasonal climate forecasts Andrew W. Robertson, IRI tailored seasonal forecasts why do we make probabilistic forecasts? to reduce our uncertainty about the (unknown) future
Principal Component Analysis Utilizing R and SAS Software s
 International Journal of Current Microbiology and Applied Sciences ISSN: 2319-7706 Volume 7 Number 05 (2018) Journal homepage: http://www.ijcmas.com Original Research Article https://doi.org/10.20546/ijcmas.2018.705.441
International Journal of Current Microbiology and Applied Sciences ISSN: 2319-7706 Volume 7 Number 05 (2018) Journal homepage: http://www.ijcmas.com Original Research Article https://doi.org/10.20546/ijcmas.2018.705.441
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
High-Throughput Sequencing Course. Introduction. Introduction. Multiple Testing. Biostatistics and Bioinformatics. Summer 2018
 High-Throughput Sequencing Course Multiple Testing Biostatistics and Bioinformatics Summer 2018 Introduction You have previously considered the significance of a single gene Introduction You have previously
High-Throughput Sequencing Course Multiple Testing Biostatistics and Bioinformatics Summer 2018 Introduction You have previously considered the significance of a single gene Introduction You have previously
Lecture 16: Small Sample Size Problems (Covariance Estimation) Many thanks to Carlos Thomaz who authored the original version of these slides
 Lecture 16: Small Sample Size Problems (Covariance Estimation) Many thanks to Carlos Thomaz who authored the original version of these slides Intelligent Data Analysis and Probabilistic Inference Lecture
Lecture 16: Small Sample Size Problems (Covariance Estimation) Many thanks to Carlos Thomaz who authored the original version of these slides Intelligent Data Analysis and Probabilistic Inference Lecture
CMSC858P Supervised Learning Methods
 CMSC858P Supervised Learning Methods Hector Corrada Bravo March, 2010 Introduction Today we discuss the classification setting in detail. Our setting is that we observe for each subject i a set of p predictors
CMSC858P Supervised Learning Methods Hector Corrada Bravo March, 2010 Introduction Today we discuss the classification setting in detail. Our setting is that we observe for each subject i a set of p predictors
Assignment 3. Introduction to Machine Learning Prof. B. Ravindran
 Assignment 3 Introduction to Machine Learning Prof. B. Ravindran 1. In building a linear regression model for a particular data set, you observe the coefficient of one of the features having a relatively
Assignment 3 Introduction to Machine Learning Prof. B. Ravindran 1. In building a linear regression model for a particular data set, you observe the coefficient of one of the features having a relatively
Principal Components Analysis using R Francis Huang / November 2, 2016
 Principal Components Analysis using R Francis Huang / huangf@missouri.edu November 2, 2016 Principal components analysis (PCA) is a convenient way to reduce high dimensional data into a smaller number
Principal Components Analysis using R Francis Huang / huangf@missouri.edu November 2, 2016 Principal components analysis (PCA) is a convenient way to reduce high dimensional data into a smaller number
Single gene analysis of differential expression. Giorgio Valentini
 Single gene analysis of differential expression Giorgio Valentini valenti@disi.unige.it Comparing two conditions Each condition may be represented by one or more RNA samples. Using cdna microarrays, samples
Single gene analysis of differential expression Giorgio Valentini valenti@disi.unige.it Comparing two conditions Each condition may be represented by one or more RNA samples. Using cdna microarrays, samples
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
Quantitative Understanding in Biology Short Course Session 9 Principal Components Analysis
 Quantitative Understanding in Biology Short Course Session 9 Principal Components Analysis Jinhyun Ju Jason Banfelder Luce Skrabanek June 21st, 218 1 Preface For the last session in this course, we ll
Quantitative Understanding in Biology Short Course Session 9 Principal Components Analysis Jinhyun Ju Jason Banfelder Luce Skrabanek June 21st, 218 1 Preface For the last session in this course, we ll
Biochip informatics-(i)
 Biochip informatics-(i) : biochip normalization & differential expression Ju Han Kim, M.D., Ph.D. SNUBI: SNUBiomedical Informatics http://www.snubi snubi.org/ Biochip Informatics - (I) Biochip basics Preprocessing
Biochip informatics-(i) : biochip normalization & differential expression Ju Han Kim, M.D., Ph.D. SNUBI: SNUBiomedical Informatics http://www.snubi snubi.org/ Biochip Informatics - (I) Biochip basics Preprocessing
PCA: Principal Component Analysis
 PCA: Principal Component Analysis Lyron Winderbaum University of Adelaide January 29, 2015 PCA is the vanilla flavour of Component Analysis What is a component? What makes a component principal? A Contrived
PCA: Principal Component Analysis Lyron Winderbaum University of Adelaide January 29, 2015 PCA is the vanilla flavour of Component Analysis What is a component? What makes a component principal? A Contrived
Structure in Data. A major objective in data analysis is to identify interesting features or structure in the data.
 Structure in Data A major objective in data analysis is to identify interesting features or structure in the data. The graphical methods are very useful in discovering structure. There are basically two
Structure in Data A major objective in data analysis is to identify interesting features or structure in the data. The graphical methods are very useful in discovering structure. There are basically two
MLCC 2015 Dimensionality Reduction and PCA
 MLCC 2015 Dimensionality Reduction and PCA Lorenzo Rosasco UNIGE-MIT-IIT June 25, 2015 Outline PCA & Reconstruction PCA and Maximum Variance PCA and Associated Eigenproblem Beyond the First Principal Component
MLCC 2015 Dimensionality Reduction and PCA Lorenzo Rosasco UNIGE-MIT-IIT June 25, 2015 Outline PCA & Reconstruction PCA and Maximum Variance PCA and Associated Eigenproblem Beyond the First Principal Component
Package plw. R topics documented: May 7, Type Package
 Type Package Package plw May 7, 2018 Title Probe level Locally moderated Weighted t-tests. Version 1.40.0 Date 2009-07-22 Author Magnus Astrand Maintainer Magnus Astrand
Type Package Package plw May 7, 2018 Title Probe level Locally moderated Weighted t-tests. Version 1.40.0 Date 2009-07-22 Author Magnus Astrand Maintainer Magnus Astrand
Similarities of Ordered Gene Lists. User s Guide to the Bioconductor Package OrderedList
 for Center Berlin Genome Based Bioinformatics Max Planck Institute for Molecular Genetics Computational Diagnostics Group @ Dept. Vingron Ihnestrasse 63-73, D-14195 Berlin, Germany http://compdiag.molgen.mpg.de/
for Center Berlin Genome Based Bioinformatics Max Planck Institute for Molecular Genetics Computational Diagnostics Group @ Dept. Vingron Ihnestrasse 63-73, D-14195 Berlin, Germany http://compdiag.molgen.mpg.de/
Package LBLGXE. R topics documented: July 20, Type Package
 Type Package Package LBLGXE July 20, 2015 Title Bayesian Lasso for detecting Rare (or Common) Haplotype Association and their interactions with Environmental Covariates Version 1.2 Date 2015-07-09 Author
Type Package Package LBLGXE July 20, 2015 Title Bayesian Lasso for detecting Rare (or Common) Haplotype Association and their interactions with Environmental Covariates Version 1.2 Date 2015-07-09 Author
samplesizelogisticcasecontrol Package
 samplesizelogisticcasecontrol Package January 31, 2017 > library(samplesizelogisticcasecontrol) Random data generation functions Let X 1 and X 2 be two variables with a bivariate normal ditribution with
samplesizelogisticcasecontrol Package January 31, 2017 > library(samplesizelogisticcasecontrol) Random data generation functions Let X 1 and X 2 be two variables with a bivariate normal ditribution with
Package sgpca. R topics documented: July 6, Type Package. Title Sparse Generalized Principal Component Analysis. Version 1.0.
 Package sgpca July 6, 2013 Type Package Title Sparse Generalized Principal Component Analysis Version 1.0 Date 2012-07-05 Author Frederick Campbell Maintainer Frederick Campbell
Package sgpca July 6, 2013 Type Package Title Sparse Generalized Principal Component Analysis Version 1.0 Date 2012-07-05 Author Frederick Campbell Maintainer Frederick Campbell
Dimensionality Reduction
 Lecture 5 1 Outline 1. Overview a) What is? b) Why? 2. Principal Component Analysis (PCA) a) Objectives b) Explaining variability c) SVD 3. Related approaches a) ICA b) Autoencoders 2 Example 1: Sportsball
Lecture 5 1 Outline 1. Overview a) What is? b) Why? 2. Principal Component Analysis (PCA) a) Objectives b) Explaining variability c) SVD 3. Related approaches a) ICA b) Autoencoders 2 Example 1: Sportsball
Package beam. May 3, 2018
 Type Package Package beam May 3, 2018 Title Fast Bayesian Inference in Large Gaussian Graphical Models Version 1.0.2 Date 2018-05-03 Author Gwenael G.R. Leday [cre, aut], Ilaria Speranza [aut], Harry Gray
Type Package Package beam May 3, 2018 Title Fast Bayesian Inference in Large Gaussian Graphical Models Version 1.0.2 Date 2018-05-03 Author Gwenael G.R. Leday [cre, aut], Ilaria Speranza [aut], Harry Gray
Measuring relationships among multiple responses
 Measuring relationships among multiple responses Linear association (correlation, relatedness, shared information) between pair-wise responses is an important property used in almost all multivariate analyses.
Measuring relationships among multiple responses Linear association (correlation, relatedness, shared information) between pair-wise responses is an important property used in almost all multivariate analyses.
Problem Set 2. MAS 622J/1.126J: Pattern Recognition and Analysis. Due: 5:00 p.m. on September 30
 Problem Set MAS 6J/1.16J: Pattern Recognition and Analysis Due: 5:00 p.m. on September 30 [Note: All instructions to plot data or write a program should be carried out using Matlab. In order to maintain
Problem Set MAS 6J/1.16J: Pattern Recognition and Analysis Due: 5:00 p.m. on September 30 [Note: All instructions to plot data or write a program should be carried out using Matlab. In order to maintain
Dimensionality reduction
 Dimensionality Reduction PCA continued Machine Learning CSE446 Carlos Guestrin University of Washington May 22, 2013 Carlos Guestrin 2005-2013 1 Dimensionality reduction n Input data may have thousands
Dimensionality Reduction PCA continued Machine Learning CSE446 Carlos Guestrin University of Washington May 22, 2013 Carlos Guestrin 2005-2013 1 Dimensionality reduction n Input data may have thousands
Asymptotic behavior of Support Vector Machine for spiked population model
 Journal of Machine Learning Research 18 (2017) 1-21 Submitted 11/16; Revised 3/17; Published 4/17 Asymptotic behavior of Support Vector Machine for spiked population model Department of Epidemiology and
Journal of Machine Learning Research 18 (2017) 1-21 Submitted 11/16; Revised 3/17; Published 4/17 Asymptotic behavior of Support Vector Machine for spiked population model Department of Epidemiology and
PCA, Kernel PCA, ICA
 PCA, Kernel PCA, ICA Learning Representations. Dimensionality Reduction. Maria-Florina Balcan 04/08/2015 Big & High-Dimensional Data High-Dimensions = Lot of Features Document classification Features per
PCA, Kernel PCA, ICA Learning Representations. Dimensionality Reduction. Maria-Florina Balcan 04/08/2015 Big & High-Dimensional Data High-Dimensions = Lot of Features Document classification Features per
Revision: Chapter 1-6. Applied Multivariate Statistics Spring 2012
 Revision: Chapter 1-6 Applied Multivariate Statistics Spring 2012 Overview Cov, Cor, Mahalanobis, MV normal distribution Visualization: Stars plot, mosaic plot with shading Outlier: chisq.plot Missing
Revision: Chapter 1-6 Applied Multivariate Statistics Spring 2012 Overview Cov, Cor, Mahalanobis, MV normal distribution Visualization: Stars plot, mosaic plot with shading Outlier: chisq.plot Missing
ABSSeq: a new RNA-Seq analysis method based on modelling absolute expression differences
 ABSSeq: a new RNA-Seq analysis method based on modelling absolute expression differences Wentao Yang October 30, 2018 1 Introduction This vignette is intended to give a brief introduction of the ABSSeq
ABSSeq: a new RNA-Seq analysis method based on modelling absolute expression differences Wentao Yang October 30, 2018 1 Introduction This vignette is intended to give a brief introduction of the ABSSeq
Statistics Toolbox 6. Apply statistical algorithms and probability models
 Statistics Toolbox 6 Apply statistical algorithms and probability models Statistics Toolbox provides engineers, scientists, researchers, financial analysts, and statisticians with a comprehensive set of
Statistics Toolbox 6 Apply statistical algorithms and probability models Statistics Toolbox provides engineers, scientists, researchers, financial analysts, and statisticians with a comprehensive set of
Research Article Sample Size Calculation for Controlling False Discovery Proportion
 Probability and Statistics Volume 2012, Article ID 817948, 13 pages doi:10.1155/2012/817948 Research Article Sample Size Calculation for Controlling False Discovery Proportion Shulian Shang, 1 Qianhe Zhou,
Probability and Statistics Volume 2012, Article ID 817948, 13 pages doi:10.1155/2012/817948 Research Article Sample Size Calculation for Controlling False Discovery Proportion Shulian Shang, 1 Qianhe Zhou,
Principal Component Analysis
 I.T. Jolliffe Principal Component Analysis Second Edition With 28 Illustrations Springer Contents Preface to the Second Edition Preface to the First Edition Acknowledgments List of Figures List of Tables
I.T. Jolliffe Principal Component Analysis Second Edition With 28 Illustrations Springer Contents Preface to the Second Edition Preface to the First Edition Acknowledgments List of Figures List of Tables
hsnim: Hyper Scalable Network Inference Machine for Scale-Free Protein-Protein Interaction Networks Inference
 CS 229 Project Report (TR# MSB2010) Submitted 12/10/2010 hsnim: Hyper Scalable Network Inference Machine for Scale-Free Protein-Protein Interaction Networks Inference Muhammad Shoaib Sehgal Computer Science
CS 229 Project Report (TR# MSB2010) Submitted 12/10/2010 hsnim: Hyper Scalable Network Inference Machine for Scale-Free Protein-Protein Interaction Networks Inference Muhammad Shoaib Sehgal Computer Science
December 20, MAA704, Multivariate analysis. Christopher Engström. Multivariate. analysis. Principal component analysis
 .. December 20, 2013 Todays lecture. (PCA) (PLS-R) (LDA) . (PCA) is a method often used to reduce the dimension of a large dataset to one of a more manageble size. The new dataset can then be used to make
.. December 20, 2013 Todays lecture. (PCA) (PLS-R) (LDA) . (PCA) is a method often used to reduce the dimension of a large dataset to one of a more manageble size. The new dataset can then be used to make
CGEN(Case-control.GENetics) Package
 CGEN(Case-control.GENetics) Package October 30, 2018 > library(cgen) Example of snp.logistic Load the ovarian cancer data and print the first 5 rows. > data(xdata, package="cgen") > Xdata[1:5, ] id case.control
CGEN(Case-control.GENetics) Package October 30, 2018 > library(cgen) Example of snp.logistic Load the ovarian cancer data and print the first 5 rows. > data(xdata, package="cgen") > Xdata[1:5, ] id case.control
Lab 12: Dimension reduction in R
 Lab : Dimension reduction in R June 6, 00 In this lab, we present some dimension reduction algorithms that are available in R and that are useful for classification of microarray data. The first example
Lab : Dimension reduction in R June 6, 00 In this lab, we present some dimension reduction algorithms that are available in R and that are useful for classification of microarray data. The first example
Follow-up data with the Epi package
 Follow-up data with the Epi package Summer 2014 Michael Hills Martyn Plummer Bendix Carstensen Retired Highgate, London International Agency for Research on Cancer, Lyon plummer@iarc.fr Steno Diabetes
Follow-up data with the Epi package Summer 2014 Michael Hills Martyn Plummer Bendix Carstensen Retired Highgate, London International Agency for Research on Cancer, Lyon plummer@iarc.fr Steno Diabetes
Principal Component Analysis
 Principal Component Analysis Yingyu Liang yliang@cs.wisc.edu Computer Sciences Department University of Wisconsin, Madison [based on slides from Nina Balcan] slide 1 Goals for the lecture you should understand
Principal Component Analysis Yingyu Liang yliang@cs.wisc.edu Computer Sciences Department University of Wisconsin, Madison [based on slides from Nina Balcan] slide 1 Goals for the lecture you should understand
Package ShrinkCovMat
 Type Package Package ShrinkCovMat Title Shrinkage Covariance Matrix Estimators Version 1.2.0 Author July 11, 2017 Maintainer Provides nonparametric Steinian shrinkage estimators
Type Package Package ShrinkCovMat Title Shrinkage Covariance Matrix Estimators Version 1.2.0 Author July 11, 2017 Maintainer Provides nonparametric Steinian shrinkage estimators
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Bradley Broom Department of Bioinformatics and Computational Biology UT M. D. Anderson Cancer Center kabagg@mdanderson.org bmbroom@mdanderson.org
GS01 0163 Analysis of Microarray Data Keith Baggerly and Bradley Broom Department of Bioinformatics and Computational Biology UT M. D. Anderson Cancer Center kabagg@mdanderson.org bmbroom@mdanderson.org
Introduction PCA classic Generative models Beyond and summary. PCA, ICA and beyond
 PCA, ICA and beyond Summer School on Manifold Learning in Image and Signal Analysis, August 17-21, 2009, Hven Technical University of Denmark (DTU) & University of Copenhagen (KU) August 18, 2009 Motivation
PCA, ICA and beyond Summer School on Manifold Learning in Image and Signal Analysis, August 17-21, 2009, Hven Technical University of Denmark (DTU) & University of Copenhagen (KU) August 18, 2009 Motivation
Singular Value Decomposition and Principal Component Analysis (PCA) I
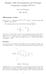 Singular Value Decomposition and Principal Component Analysis (PCA) I Prof Ned Wingreen MOL 40/50 Microarray review Data per array: 0000 genes, I (green) i,i (red) i 000 000+ data points! The expression
Singular Value Decomposition and Principal Component Analysis (PCA) I Prof Ned Wingreen MOL 40/50 Microarray review Data per array: 0000 genes, I (green) i,i (red) i 000 000+ data points! The expression
A direct formulation for sparse PCA using semidefinite programming
 A direct formulation for sparse PCA using semidefinite programming A. d Aspremont, L. El Ghaoui, M. Jordan, G. Lanckriet ORFE, Princeton University & EECS, U.C. Berkeley Available online at www.princeton.edu/~aspremon
A direct formulation for sparse PCA using semidefinite programming A. d Aspremont, L. El Ghaoui, M. Jordan, G. Lanckriet ORFE, Princeton University & EECS, U.C. Berkeley Available online at www.princeton.edu/~aspremon
Supervised Learning: Linear Methods (1/2) Applied Multivariate Statistics Spring 2012
 Supervised Learning: Linear Methods (1/2) Applied Multivariate Statistics Spring 2012 Overview Review: Conditional Probability LDA / QDA: Theory Fisher s Discriminant Analysis LDA: Example Quality control:
Supervised Learning: Linear Methods (1/2) Applied Multivariate Statistics Spring 2012 Overview Review: Conditional Probability LDA / QDA: Theory Fisher s Discriminant Analysis LDA: Example Quality control:
Standardization and Singular Value Decomposition in Canonical Correlation Analysis
 Standardization and Singular Value Decomposition in Canonical Correlation Analysis Melinda Borello Johanna Hardin, Advisor David Bachman, Reader Submitted to Pitzer College in Partial Fulfillment of the
Standardization and Singular Value Decomposition in Canonical Correlation Analysis Melinda Borello Johanna Hardin, Advisor David Bachman, Reader Submitted to Pitzer College in Partial Fulfillment of the
Fundamentals to Biostatistics. Prof. Chandan Chakraborty Associate Professor School of Medical Science & Technology IIT Kharagpur
 Fundamentals to Biostatistics Prof. Chandan Chakraborty Associate Professor School of Medical Science & Technology IIT Kharagpur Statistics collection, analysis, interpretation of data development of new
Fundamentals to Biostatistics Prof. Chandan Chakraborty Associate Professor School of Medical Science & Technology IIT Kharagpur Statistics collection, analysis, interpretation of data development of new
Effective Linear Discriminant Analysis for High Dimensional, Low Sample Size Data
 Effective Linear Discriant Analysis for High Dimensional, Low Sample Size Data Zhihua Qiao, Lan Zhou and Jianhua Z. Huang Abstract In the so-called high dimensional, low sample size (HDLSS) settings, LDA
Effective Linear Discriant Analysis for High Dimensional, Low Sample Size Data Zhihua Qiao, Lan Zhou and Jianhua Z. Huang Abstract In the so-called high dimensional, low sample size (HDLSS) settings, LDA
Processing microarray data with Bioconductor
 Processing microarray data with Bioconductor Statistical analysis of gene expression data with R and Bioconductor University of Copenhagen Copenhagen Biocenter Laurent Gautier 1, 2 August 17-21 2009 Contents
Processing microarray data with Bioconductor Statistical analysis of gene expression data with R and Bioconductor University of Copenhagen Copenhagen Biocenter Laurent Gautier 1, 2 August 17-21 2009 Contents
Dimension reduction, PCA & eigenanalysis Based in part on slides from textbook, slides of Susan Holmes. October 3, Statistics 202: Data Mining
 Dimension reduction, PCA & eigenanalysis Based in part on slides from textbook, slides of Susan Holmes October 3, 2012 1 / 1 Combinations of features Given a data matrix X n p with p fairly large, it can
Dimension reduction, PCA & eigenanalysis Based in part on slides from textbook, slides of Susan Holmes October 3, 2012 1 / 1 Combinations of features Given a data matrix X n p with p fairly large, it can
Basics of Multivariate Modelling and Data Analysis
 Basics of Multivariate Modelling and Data Analysis Kurt-Erik Häggblom 6. Principal component analysis (PCA) 6.1 Overview 6.2 Essentials of PCA 6.3 Numerical calculation of PCs 6.4 Effects of data preprocessing
Basics of Multivariate Modelling and Data Analysis Kurt-Erik Häggblom 6. Principal component analysis (PCA) 6.1 Overview 6.2 Essentials of PCA 6.3 Numerical calculation of PCs 6.4 Effects of data preprocessing
Machine learning comes from Bayesian decision theory in statistics. There we want to minimize the expected value of the loss function.
 Bayesian learning: Machine learning comes from Bayesian decision theory in statistics. There we want to minimize the expected value of the loss function. Let y be the true label and y be the predicted
Bayesian learning: Machine learning comes from Bayesian decision theory in statistics. There we want to minimize the expected value of the loss function. Let y be the true label and y be the predicted
Principal Components Analysis (PCA) and Singular Value Decomposition (SVD) with applications to Microarrays
 Principal Components Analysis (PCA) and Singular Value Decomposition (SVD) with applications to Microarrays Prof. Tesler Math 283 Fall 2015 Prof. Tesler Principal Components Analysis Math 283 / Fall 2015
Principal Components Analysis (PCA) and Singular Value Decomposition (SVD) with applications to Microarrays Prof. Tesler Math 283 Fall 2015 Prof. Tesler Principal Components Analysis Math 283 / Fall 2015
Package SHIP. February 19, 2015
 Type Package Package SHIP February 19, 2015 Title SHrinkage covariance Incorporating Prior knowledge Version 1.0.2 Author Maintainer Vincent Guillemot The SHIP-package allows
Type Package Package SHIP February 19, 2015 Title SHrinkage covariance Incorporating Prior knowledge Version 1.0.2 Author Maintainer Vincent Guillemot The SHIP-package allows
Bayesian Hierarchical Classification. Seminar on Predicting Structured Data Jukka Kohonen
 Bayesian Hierarchical Classification Seminar on Predicting Structured Data Jukka Kohonen 17.4.2008 Overview Intro: The task of hierarchical gene annotation Approach I: SVM/Bayes hybrid Barutcuoglu et al:
Bayesian Hierarchical Classification Seminar on Predicting Structured Data Jukka Kohonen 17.4.2008 Overview Intro: The task of hierarchical gene annotation Approach I: SVM/Bayes hybrid Barutcuoglu et al:
Multivariate analysis of genetic data: an introduction
 Multivariate analysis of genetic data: an introduction Thibaut Jombart MRC Centre for Outbreak Analysis and Modelling Imperial College London XXIV Simposio Internacional De Estadística Bogotá, 25th July
Multivariate analysis of genetic data: an introduction Thibaut Jombart MRC Centre for Outbreak Analysis and Modelling Imperial College London XXIV Simposio Internacional De Estadística Bogotá, 25th July
Regression Model In The Analysis Of Micro Array Data-Gene Expression Detection
 Jamal Fathima.J.I 1 and P.Venkatesan 1. Research Scholar -Department of statistics National Institute For Research In Tuberculosis, Indian Council For Medical Research,Chennai,India,.Department of statistics
Jamal Fathima.J.I 1 and P.Venkatesan 1. Research Scholar -Department of statistics National Institute For Research In Tuberculosis, Indian Council For Medical Research,Chennai,India,.Department of statistics
Principle Components Analysis (PCA) Relationship Between a Linear Combination of Variables and Axes Rotation for PCA
 Principle Components Analysis (PCA) Relationship Between a Linear Combination of Variables and Axes Rotation for PCA Principle Components Analysis: Uses one group of variables (we will call this X) In
Principle Components Analysis (PCA) Relationship Between a Linear Combination of Variables and Axes Rotation for PCA Principle Components Analysis: Uses one group of variables (we will call this X) In
Multivariate Fundamentals: Rotation. Exploratory Factor Analysis
 Multivariate Fundamentals: Rotation Exploratory Factor Analysis PCA Analysis A Review Precipitation Temperature Ecosystems PCA Analysis with Spatial Data Proportion of variance explained Comp.1 + Comp.2
Multivariate Fundamentals: Rotation Exploratory Factor Analysis PCA Analysis A Review Precipitation Temperature Ecosystems PCA Analysis with Spatial Data Proportion of variance explained Comp.1 + Comp.2
Data Preprocessing. Data Preprocessing
 Data Preprocessing 1 Data Preprocessing Normalization: the process of removing sampleto-sample variations in the measurements not due to differential gene expression. Bringing measurements from the different
Data Preprocessing 1 Data Preprocessing Normalization: the process of removing sampleto-sample variations in the measurements not due to differential gene expression. Bringing measurements from the different
Principal Variance Components Analysis for Quantifying Variability in Genomics Data
 Principal Variance Components Analysis for Quantifying Variability in Genomics Data Tzu-Ming Chu SAS Institute Inc. Outlines Motivation PVCA (PCA + VCA) Example Mouse Lung Tumorigenicity Data Grouped Batch
Principal Variance Components Analysis for Quantifying Variability in Genomics Data Tzu-Ming Chu SAS Institute Inc. Outlines Motivation PVCA (PCA + VCA) Example Mouse Lung Tumorigenicity Data Grouped Batch
Statistical Methods for Integration of Multiple Omics Data
 Statistical Methods for Integration of Multiple Omics Data Hae-Won Uh BMTL, October 2014, Naples October 23, 2014 Hae-Won Uh, BMTL, October 2014, Naples Statistical Methods for Integration of Multiple
Statistical Methods for Integration of Multiple Omics Data Hae-Won Uh BMTL, October 2014, Naples October 23, 2014 Hae-Won Uh, BMTL, October 2014, Naples Statistical Methods for Integration of Multiple
Lecture Topic Projects 1 Intro, schedule, and logistics 2 Applications of visual analytics, data types 3 Data sources and preparation Project 1 out 4
 Lecture Topic Projects 1 Intro, schedule, and logistics 2 Applications of visual analytics, data types 3 Data sources and preparation Project 1 out 4 Data reduction, similarity & distance, data augmentation
Lecture Topic Projects 1 Intro, schedule, and logistics 2 Applications of visual analytics, data types 3 Data sources and preparation Project 1 out 4 Data reduction, similarity & distance, data augmentation
One-shot Learning of Poisson Distributions Information Theory of Audic-Claverie Statistic for Analyzing cdna Arrays
 One-shot Learning of Poisson Distributions Information Theory of Audic-Claverie Statistic for Analyzing cdna Arrays Peter Tiňo School of Computer Science University of Birmingham, UK One-shot Learning
One-shot Learning of Poisson Distributions Information Theory of Audic-Claverie Statistic for Analyzing cdna Arrays Peter Tiňo School of Computer Science University of Birmingham, UK One-shot Learning
MACAU 2.0 User Manual
 MACAU 2.0 User Manual Shiquan Sun, Jiaqiang Zhu, and Xiang Zhou Department of Biostatistics, University of Michigan shiquans@umich.edu and xzhousph@umich.edu April 9, 2017 Copyright 2016 by Xiang Zhou
MACAU 2.0 User Manual Shiquan Sun, Jiaqiang Zhu, and Xiang Zhou Department of Biostatistics, University of Michigan shiquans@umich.edu and xzhousph@umich.edu April 9, 2017 Copyright 2016 by Xiang Zhou
Machine Learning. Principal Components Analysis. Le Song. CSE6740/CS7641/ISYE6740, Fall 2012
 Machine Learning CSE6740/CS7641/ISYE6740, Fall 2012 Principal Components Analysis Le Song Lecture 22, Nov 13, 2012 Based on slides from Eric Xing, CMU Reading: Chap 12.1, CB book 1 2 Factor or Component
Machine Learning CSE6740/CS7641/ISYE6740, Fall 2012 Principal Components Analysis Le Song Lecture 22, Nov 13, 2012 Based on slides from Eric Xing, CMU Reading: Chap 12.1, CB book 1 2 Factor or Component
Principal component analysis PCA. Kathleen Marchal Dept Plant Biotechnology and Bioinformatics Department information technology (INTEC)
 Principal component analysis PCA Kathleen Marchal Dept Plant Biotechnology and Bioinformatics Department information technology (INTEC) Overzicht lessen 26/02 13h biostat S3 emile clapeyron 07/03 ma 13
Principal component analysis PCA Kathleen Marchal Dept Plant Biotechnology and Bioinformatics Department information technology (INTEC) Overzicht lessen 26/02 13h biostat S3 emile clapeyron 07/03 ma 13
Expectation Maximization
 Expectation Maximization Machine Learning CSE546 Carlos Guestrin University of Washington November 13, 2014 1 E.M.: The General Case E.M. widely used beyond mixtures of Gaussians The recipe is the same
Expectation Maximization Machine Learning CSE546 Carlos Guestrin University of Washington November 13, 2014 1 E.M.: The General Case E.M. widely used beyond mixtures of Gaussians The recipe is the same
Sample Size Estimation for Studies of High-Dimensional Data
 Sample Size Estimation for Studies of High-Dimensional Data James J. Chen, Ph.D. National Center for Toxicological Research Food and Drug Administration June 3, 2009 China Medical University Taichung,
Sample Size Estimation for Studies of High-Dimensional Data James J. Chen, Ph.D. National Center for Toxicological Research Food and Drug Administration June 3, 2009 China Medical University Taichung,
Expression Data Exploration: Association, Patterns, Factors & Regression Modelling
 Expression Data Exploration: Association, Patterns, Factors & Regression Modelling Exploring gene expression data Scale factors, median chip correlation on gene subsets for crude data quality investigation
Expression Data Exploration: Association, Patterns, Factors & Regression Modelling Exploring gene expression data Scale factors, median chip correlation on gene subsets for crude data quality investigation
What is Principal Component Analysis?
 What is Principal Component Analysis? Principal component analysis (PCA) Reduce the dimensionality of a data set by finding a new set of variables, smaller than the original set of variables Retains most
What is Principal Component Analysis? Principal component analysis (PCA) Reduce the dimensionality of a data set by finding a new set of variables, smaller than the original set of variables Retains most
Multivariate Analysis of Ecological Data using CANOCO
 Multivariate Analysis of Ecological Data using CANOCO JAN LEPS University of South Bohemia, and Czech Academy of Sciences, Czech Republic Universitats- uric! Lanttesbibiiothek Darmstadt Bibliothek Biologie
Multivariate Analysis of Ecological Data using CANOCO JAN LEPS University of South Bohemia, and Czech Academy of Sciences, Czech Republic Universitats- uric! Lanttesbibiiothek Darmstadt Bibliothek Biologie
ANOVA approach. Investigates interaction terms. Disadvantages: Requires careful sampling design with replication
 ANOVA approach Advantages: Ideal for evaluating hypotheses Ideal to quantify effect size (e.g., differences between groups) Address multiple factors at once Investigates interaction terms Disadvantages:
ANOVA approach Advantages: Ideal for evaluating hypotheses Ideal to quantify effect size (e.g., differences between groups) Address multiple factors at once Investigates interaction terms Disadvantages:
Experimental Design and Data Analysis for Biologists
 Experimental Design and Data Analysis for Biologists Gerry P. Quinn Monash University Michael J. Keough University of Melbourne CAMBRIDGE UNIVERSITY PRESS Contents Preface page xv I I Introduction 1 1.1
Experimental Design and Data Analysis for Biologists Gerry P. Quinn Monash University Michael J. Keough University of Melbourne CAMBRIDGE UNIVERSITY PRESS Contents Preface page xv I I Introduction 1 1.1
1 Interpretation. Contents. Biplots, revisited. Biplots, revisited 2. Biplots, revisited 1
 Biplots, revisited 1 Biplots, revisited 2 1 Interpretation Biplots, revisited Biplots show the following quantities of a data matrix in one display: Slide 1 Ulrich Kohler kohler@wz-berlin.de Slide 3 the
Biplots, revisited 1 Biplots, revisited 2 1 Interpretation Biplots, revisited Biplots show the following quantities of a data matrix in one display: Slide 1 Ulrich Kohler kohler@wz-berlin.de Slide 3 the
The following postestimation commands are of special interest after discrim qda: The following standard postestimation commands are also available:
 Title stata.com discrim qda postestimation Postestimation tools for discrim qda Syntax for predict Menu for predict Options for predict Syntax for estat Menu for estat Options for estat Remarks and examples
Title stata.com discrim qda postestimation Postestimation tools for discrim qda Syntax for predict Menu for predict Options for predict Syntax for estat Menu for estat Options for estat Remarks and examples
Single gene analysis of differential expression
 Single gene analysis of differential expression Giorgio Valentini DSI Dipartimento di Scienze dell Informazione Università degli Studi di Milano valentini@dsi.unimi.it Comparing two conditions Each condition
Single gene analysis of differential expression Giorgio Valentini DSI Dipartimento di Scienze dell Informazione Università degli Studi di Milano valentini@dsi.unimi.it Comparing two conditions Each condition
An Empirical Comparison of Dimensionality Reduction Methods for Classifying Gene and Protein Expression Datasets
 An Empirical Comparison of Dimensionality Reduction Methods for Classifying Gene and Protein Expression Datasets George Lee 1, Carlos Rodriguez 2, and Anant Madabhushi 1 1 Rutgers, The State University
An Empirical Comparison of Dimensionality Reduction Methods for Classifying Gene and Protein Expression Datasets George Lee 1, Carlos Rodriguez 2, and Anant Madabhushi 1 1 Rutgers, The State University
