Molecular Dynamics. What to choose in an integrator The Verlet algorithm Boundary Conditions in Space and time Reading Assignment: F&S Chapter 4
|
|
|
- Roderick Cannon
- 5 years ago
- Views:
Transcription
1 Molecular Dynamics What to choose in an integrator The Verlet algorithm Boundary Conditions in Space and time Reading Assignment: F&S Chapter 4 MSE485/PHY466/CSE485 1
2 The Molecular Dynamics (MD) method Pick particles, masses and forces (or potential). Initialize positions and momentum (boundary conditions in space and time). Solve F = m a to determine r(t), v(t). (the integrator) We need to make time discrete (t k ) not continuous Compute properties along the trajectory Estimate errors. Try to use the simulation to answer physical questions. Alder & Wainwright, J Chem. Phys. 27, MSE485/PHY466/CSE485 2
3 Characteristics of simulations. Potentials are highly non-linear, with discontinuous higher derivatives either at the origin or at the box edge. Small changes in accuracy lead to totally different trajectories (the mixing or ergodic property). Long-time information is mostly statistical. We need low accuracy because the potentials are not very realistic. Universality saves us: a badly integrated system is probably similar to our original system. Not true in the field of non-linear dynamics e.g. in studying the solar system. CPU time is totally dominated by the calculation of forces. We do not want to compute higher-order derivatives of the potential. They might not exist! What is a good measure of computer/algorithm efficiency? real time/computer time or CPU time/step (for fixed N). Memory limits are not too important. Energy conservation is important; roughly equivalent to time-reversal invariance. rule of thumb: allow 0.01k B T fluctuation in the total energy. MSE485/PHY466/CSE485 3
4 Criteria for an Integrator simplicity (How long does it take to write and debug?) efficiency (How fast to advance a given system?) stability (what is the long-term energy conservation?) reliability (Can it handle a variety of T, densities, potentials?) Compare statistical errors (~ h -1/2 ) with time step errors (~ h p where p=2,3.) to pick the time step! Choose time step so that : statistical error ~ systematic errors. MSE485/PHY466/CSE485 4
5 The Verlet Algorithm Ubiquitous choice for an integrator: Verlet (or leapfrog) r(t+h) = r(t) + v(t) h + 1/2 a(t) h 2 + b(t) h 3 + O(h 4 ) Taylor expand r(t-h) = r(t) - v(t) h + 1/2 a(t) h 2 - b(t) h 3 + O(h 4 ) Reverse time r(t+h) = 2 r(t) - r(t-h) + a(t) h 2 + O(h 4 ) v(t) = (r(t+h) - r(t-h))/(2h) + O(h 2 ) Add estimate velocities Time reversal invariance is built in: the energy does not drift either up or down. Prove this. Historical Point of Interest: Method also suggested by Newton and Newmark! MSE485/PHY466/CSE485 5
6 How to set the time step Adjust to get energy conservation to 1% of kinetic energy. Even if errors are large, you are close to the exact trajectory of a nearby physical system with a different potential. Because we don t really know the potential surface that accurately, this is often satisfactory. Leapfrog algorithm has a problem with round-off error. Use the equivalent velocity-verlet instead: r(t+h) = r(t) + h [ v(t) +(h/2) a(t)] v(t+h/2) = v(t) + (h/2) a(t) v(t+h) = v(t+h/2) + (h/2) a(t+h) MSE485/PHY466/CSE485 6
7 Higher Order Methods? Statistical error for fixed CPU time. Higher-order does not always mean better. Problem is that potentials are not analytic Systematic error Usually one tries to balance all sources of errors Berendsen 86 Gear methods MSE485/PHY466/CSE485 7
8 Quote from Berendsen How accurate a simulation has to be in order to provide reliable statistical results is still a matter of debate. The purpose is never to produce reliable trajectories. Whatever accuracy the forces and the algorithms have, significant deviations from the exact solution will inevitably occur. Also, in the real physical world, individual trajectories have no meaning: in a quantum mechanical sense they do not exist and even classically they become unpredictable in the long run due to the nonisolated character of any real system. So what counts are statistical averages. It seems that very little systematic evaluation of algorithm has been done with this in mind. We have the impression that a noise as high as 10% of the kinetic energy fluctuation is still acceptable, although the accuracy of fluctuations may not be sufficient to obtain thermodynamic data from them. With such a level of inaccuracy the Verlet of leap frog algorithm is always to be preferred. MSE485/PHY466/CSE485 8
9 Long-term stability of Verlet Verlet trajectory and exact trajectory must remain close together. There exists another Hamiltonian, H *, whose energy is exactly conserved by the Verlet trajectory. H * (P,R) H(P,R) ~O(Δt 4 ) The true and pseudo energy must remain close to each other for small time-step (so-called symplectic behavior that conserves phase-space volume). Hence there are no long-term drifts in the (original) energy. Always use sympletic algorithms - higher order ones exist. MSE485/PHY466/CSE485 9
10 Spatial Boundary Conditions Important because spatial scales are limited. What BC can we choose? There are different possibilities: No boundaries; e.g. droplet, protein in vacuum. If droplet has 1 million atoms and surface layer is 5 atoms thick, then 25% of atoms are on the surface. Periodic Boundaries: most popular choice because there are no surfaces (see next slide) but there can still be problems. Simulations on a sphere External potentials Mixed boundaries (e.g. infinite in z, periodic in x and y) MSE485/PHY466/CSE485 10
11 Why use Periodic Boundary Conditions? Macroscopic systems O(10 23 ) particles. We eliminate surfaces from simulation. Allows us to get at bulk properties with few particles. Applied to solids, liquids, gases, and plasmas. Some finite-size effects remain, but can often be removed with scaling. MSE485/PHY466/CSE485 11
12 Periodic Boundary Conditions Is the system periodic or just an infinite array of images? How do you calculate distances in periodic system? MSE485/PHY466/CSE485 12
13 Periodic distances Minimum Image Convention: Take the closest distance r M = min (r+nl) -3L/2 -L -L/2 0 L/2 L 3L/2 How to handle the tail of the potential: Potential cutoff : V(r) = 0 for r > L/2 continuous potential: for r < r c, V c (r) =V(r) V(r c ) continuous V&F: for r < r c. V c (r) =V(r) V(r c ) V(r c )(r r c ) Image potential: V I = V(r i r j + nl) For long-range potential this leads to the Ewald image potential. You need a charge background and convergence factor (more later) MSE485/PHY466/CSE485 13
14 LJ Force Computation! Loop over all pairs of atoms. do i=2,natoms do j=1,i-1!compute distance between i and j. r2 = 0 do k=1,ndim dx(k) = r(k,i) - r(k,j)!periodic boundary conditions. if(dx(k).gt. ell2(k)) dx(k) = dx(k)-ell(k) if(dx(k).lt.-ell2(k)) dx(k) = dx(k)+ell(k) r2 = r2 + dx(k)*dx(k) enddo We can do better with the PBC! dx(k)=dx(k)-ell(k)*nint(dx(k)/ell(k))!only compute for pairs inside radial cutoff. if(r2.lt.rcut2) then r2i=sigma2/r2 r6i=r2i*r2i*r2i!shifted Lennard-Jones potential.!radial force. pot = pot+eps4*r6i*(r6i-1)- potcut rforce = eps24*r6i*r2i*(2*r6i-1) do k = 1, ndim force(k,i)=force(k,i) + rforce*dx(k) force(k,j)=force(k,j) - rforce*dx(k) enddo endif enddo enddo Why? MSE485/PHY466/CSE485 14
15 Complexity of Force Calculations Complexity is defined as how a computer algorithm scales with the number of degrees of freedom (atoms). Number of terms in pair potential is N(N-1)/2 ~ O(N 2 ) For short-range V(r), use neighbor tables to reduce it to O(N). Explicit Neighbor list (Verlet) for systems that move slowly (keep a list and update occassionally.) Bin Sort list (sort particles into bins and sum over neighboring bins) Long-range potentials require more sophisticated treatment: Ewald sums are O(N 3/2 ) Fast Multipole Algorithms are O(N), for very large N. Tree Methods More later MSE485/PHY466/CSE485 15
16 Neighbor Lists: O(N 2 ) to O(N) Pair Potentials: If V c (r) = 0 for r > r c (i.e., short-ranged), then Number of computations of the potential is mn, where m is average number of neighbors inside r c and is independent of system size! So, once we have the neighbor table, the calculation is O(N)! Constructing Neighbor Tables: O(N 2 ), must check if particles fall inside each other s radius. Then, how is table useful? If clever, only have to reconstruct the table occasionally. MSE485/PHY466/CSE485 16
17 A Neighbor Table MSE485/PHY466/CSE485 17
18 Constructing Neighbor Lists: use skin depth Cut off table at r c + Δ (a skin depth). allows a particle to move a while before we new table needed. Keep track of the two largest distances traveled, d 1 and d 2, when d 1 + d 2, Δ get new table. As Δ increases, more neighbors and more force evaluations. As Δ increases, need to calculate neighbor table less often. So, choose Δ to optimize efficiency. MSE485/PHY466/CSE485 18
19 Improvements: the Cell Method When computing table, divide box in cells of size L > r c + Δ. Need only particle s own and neighoring cell. Hence, table construction is O(N)! MSE485/PHY466/CSE485 19
20 CLAMPS input file TYPE argon POTENTIAL argon argon DENSITY 0.35 TEMPERATURE LATTICE SEED 10 WRITE_SCALARS 10 WRITE_COORD 20 Setup system Output intervals e.g., 864 argon with mass 48 e.g., atom-atom potential pot 1, σ=1.0, ε=1.0, r c = 2.5 RUN MD Executes 300 molecular dynamics steps See detailed description on the CLAMPS page. MSE485/PHY466/CSE485 20
21 Potentials for MD See Calendar for HW, etc. Next time MSE485/PHY466/CSE485 21
Molecular Dynamics. The Molecular Dynamics (MD) method for classical systems (not H or He)
 Molecular Dynamics What to choose in an integrator The Verlet algorithm Boundary Conditions in Space and time Reading Assignment: F&S Chapter 4 1 The Molecular Dynamics (MD) method for classical systems
Molecular Dynamics What to choose in an integrator The Verlet algorithm Boundary Conditions in Space and time Reading Assignment: F&S Chapter 4 1 The Molecular Dynamics (MD) method for classical systems
Molecular Dynamics 9/6/16
 Molecular Dynamics What to choose in an integrator The Verlet algorithm Boundary Conditions in Space and time Reading Assignment: Lesar Chpt 6, F&S Chpt 4 1 The Molecular Dynamics (MD) method for classical
Molecular Dynamics What to choose in an integrator The Verlet algorithm Boundary Conditions in Space and time Reading Assignment: Lesar Chpt 6, F&S Chpt 4 1 The Molecular Dynamics (MD) method for classical
What is Classical Molecular Dynamics?
 What is Classical Molecular Dynamics? Simulation of explicit particles (atoms, ions,... ) Particles interact via relatively simple analytical potential functions Newton s equations of motion are integrated
What is Classical Molecular Dynamics? Simulation of explicit particles (atoms, ions,... ) Particles interact via relatively simple analytical potential functions Newton s equations of motion are integrated
 A Nobel Prize for Molecular Dynamics and QM/MM What is Classical Molecular Dynamics? Simulation of explicit particles (atoms, ions,... ) Particles interact via relatively simple analytical potential
A Nobel Prize for Molecular Dynamics and QM/MM What is Classical Molecular Dynamics? Simulation of explicit particles (atoms, ions,... ) Particles interact via relatively simple analytical potential
Interatomic Potentials. The electronic-structure problem
 Interatomic Potentials Before we can start a simulation, we need the model! Interaction between atoms and molecules is determined by quantum mechanics: Schrödinger Equation + Born-Oppenheimer approximation
Interatomic Potentials Before we can start a simulation, we need the model! Interaction between atoms and molecules is determined by quantum mechanics: Schrödinger Equation + Born-Oppenheimer approximation
Molecular Dynamics Simulations
 Molecular Dynamics Simulations Dr. Kasra Momeni www.knanosys.com Outline Long-range Interactions Ewald Sum Fast Multipole Method Spherically Truncated Coulombic Potential Speeding up Calculations SPaSM
Molecular Dynamics Simulations Dr. Kasra Momeni www.knanosys.com Outline Long-range Interactions Ewald Sum Fast Multipole Method Spherically Truncated Coulombic Potential Speeding up Calculations SPaSM
Neighbor Tables Long-Range Potentials
 Neighbor Tables Long-Range Potentials Today we learn how we can handle long range potentials. Neighbor tables Long-range potential Ewald sums MSE485/PHY466/CSE485 1 Periodic distances Minimum Image Convention:
Neighbor Tables Long-Range Potentials Today we learn how we can handle long range potentials. Neighbor tables Long-range potential Ewald sums MSE485/PHY466/CSE485 1 Periodic distances Minimum Image Convention:
Introduction to Simulation - Lectures 17, 18. Molecular Dynamics. Nicolas Hadjiconstantinou
 Introduction to Simulation - Lectures 17, 18 Molecular Dynamics Nicolas Hadjiconstantinou Molecular Dynamics Molecular dynamics is a technique for computing the equilibrium and non-equilibrium properties
Introduction to Simulation - Lectures 17, 18 Molecular Dynamics Nicolas Hadjiconstantinou Molecular Dynamics Molecular dynamics is a technique for computing the equilibrium and non-equilibrium properties
Molecular dynamics simulation of Aquaporin-1. 4 nm
 Molecular dynamics simulation of Aquaporin-1 4 nm Molecular Dynamics Simulations Schrödinger equation i~@ t (r, R) =H (r, R) Born-Oppenheimer approximation H e e(r; R) =E e (R) e(r; R) Nucleic motion described
Molecular dynamics simulation of Aquaporin-1 4 nm Molecular Dynamics Simulations Schrödinger equation i~@ t (r, R) =H (r, R) Born-Oppenheimer approximation H e e(r; R) =E e (R) e(r; R) Nucleic motion described
Advanced Molecular Molecular Dynamics
 Advanced Molecular Molecular Dynamics Technical details May 12, 2014 Integration of harmonic oscillator r m period = 2 k k and the temperature T determine the sampling of x (here T is related with v 0
Advanced Molecular Molecular Dynamics Technical details May 12, 2014 Integration of harmonic oscillator r m period = 2 k k and the temperature T determine the sampling of x (here T is related with v 0
Bioengineering 215. An Introduction to Molecular Dynamics for Biomolecules
 Bioengineering 215 An Introduction to Molecular Dynamics for Biomolecules David Parker May 18, 2007 ntroduction A principal tool to study biological molecules is molecular dynamics simulations (MD). MD
Bioengineering 215 An Introduction to Molecular Dynamics for Biomolecules David Parker May 18, 2007 ntroduction A principal tool to study biological molecules is molecular dynamics simulations (MD). MD
Gear methods I + 1/18
 Gear methods I + 1/18 Predictor-corrector type: knowledge of history is used to predict an approximate solution, which is made more accurate in the following step we do not want (otherwise good) methods
Gear methods I + 1/18 Predictor-corrector type: knowledge of history is used to predict an approximate solution, which is made more accurate in the following step we do not want (otherwise good) methods
Molecular Dynamics. A very brief introduction
 Molecular Dynamics A very brief introduction Sander Pronk Dept. of Theoretical Physics KTH Royal Institute of Technology & Science For Life Laboratory Stockholm, Sweden Why computer simulations? Two primary
Molecular Dynamics A very brief introduction Sander Pronk Dept. of Theoretical Physics KTH Royal Institute of Technology & Science For Life Laboratory Stockholm, Sweden Why computer simulations? Two primary
Scientific Computing II
 Scientific Computing II Molecular Dynamics Numerics Michael Bader SCCS Technical University of Munich Summer 018 Recall: Molecular Dynamics System of ODEs resulting force acting on a molecule: F i = j
Scientific Computing II Molecular Dynamics Numerics Michael Bader SCCS Technical University of Munich Summer 018 Recall: Molecular Dynamics System of ODEs resulting force acting on a molecule: F i = j
Metropolis, 2D Ising model
 Metropolis, 2D Ising model You can get visual understanding from the java applets available, like: http://physics.ucsc.edu/~peter/ising/ising.html Average value of spin is magnetization. Abs of this as
Metropolis, 2D Ising model You can get visual understanding from the java applets available, like: http://physics.ucsc.edu/~peter/ising/ising.html Average value of spin is magnetization. Abs of this as
Scientific Computing II
 Scientific Computing II Molecular Dynamics Simulation Michael Bader SCCS Summer Term 2015 Molecular Dynamics Simulation, Summer Term 2015 1 Continuum Mechanics for Fluid Mechanics? Molecular Dynamics the
Scientific Computing II Molecular Dynamics Simulation Michael Bader SCCS Summer Term 2015 Molecular Dynamics Simulation, Summer Term 2015 1 Continuum Mechanics for Fluid Mechanics? Molecular Dynamics the
Simulations with MM Force Fields. Monte Carlo (MC) and Molecular Dynamics (MD) Video II.vi
 Simulations with MM Force Fields Monte Carlo (MC) and Molecular Dynamics (MD) Video II.vi Some slides taken with permission from Howard R. Mayne Department of Chemistry University of New Hampshire Walking
Simulations with MM Force Fields Monte Carlo (MC) and Molecular Dynamics (MD) Video II.vi Some slides taken with permission from Howard R. Mayne Department of Chemistry University of New Hampshire Walking
André Schleife Department of Materials Science and Engineering
 André Schleife Department of Materials Science and Engineering Length Scales (c) ICAMS: http://www.icams.de/cms/upload/01_home/01_research_at_icams/length_scales_1024x780.png Goals for today: Background
André Schleife Department of Materials Science and Engineering Length Scales (c) ICAMS: http://www.icams.de/cms/upload/01_home/01_research_at_icams/length_scales_1024x780.png Goals for today: Background
Computer simulation methods (2) Dr. Vania Calandrini
 Computer simulation methods (2) Dr. Vania Calandrini in the previous lecture: time average versus ensemble average MC versus MD simulations equipartition theorem (=> computing T) virial theorem (=> computing
Computer simulation methods (2) Dr. Vania Calandrini in the previous lecture: time average versus ensemble average MC versus MD simulations equipartition theorem (=> computing T) virial theorem (=> computing
A MOLECULAR DYNAMICS SIMULATION OF A BUBBLE NUCLEATION ON SOLID SURFACE
 A MOLECULAR DYNAMICS SIMULATION OF A BUBBLE NUCLEATION ON SOLID SURFACE Shigeo Maruyama and Tatsuto Kimura Department of Mechanical Engineering The University of Tokyo 7-- Hongo, Bunkyo-ku, Tokyo -866,
A MOLECULAR DYNAMICS SIMULATION OF A BUBBLE NUCLEATION ON SOLID SURFACE Shigeo Maruyama and Tatsuto Kimura Department of Mechanical Engineering The University of Tokyo 7-- Hongo, Bunkyo-ku, Tokyo -866,
Multiple time step Monte Carlo
 JOURNAL OF CHEMICAL PHYSICS VOLUME 117, NUMBER 18 8 NOVEMBER 2002 Multiple time step Monte Carlo Balázs Hetényi a) Department of Chemistry, Princeton University, Princeton, NJ 08544 and Department of Chemistry
JOURNAL OF CHEMICAL PHYSICS VOLUME 117, NUMBER 18 8 NOVEMBER 2002 Multiple time step Monte Carlo Balázs Hetényi a) Department of Chemistry, Princeton University, Princeton, NJ 08544 and Department of Chemistry
Part 2: Molecular Dynamics. Literature History Practical Issues
 Part 2: Molecular Dynamics Literature History Practical Issues 1 Literature A. K. Hartmann Big Practical Guide to Computer Simulations (World Scientific, 2015) M. P. Allen and D.J. Tildesley Computer Simulations
Part 2: Molecular Dynamics Literature History Practical Issues 1 Literature A. K. Hartmann Big Practical Guide to Computer Simulations (World Scientific, 2015) M. P. Allen and D.J. Tildesley Computer Simulations
Basics of Statistical Mechanics
 Basics of Statistical Mechanics Review of ensembles Microcanonical, canonical, Maxwell-Boltzmann Constant pressure, temperature, volume, Thermodynamic limit Ergodicity (see online notes also) Reading assignment:
Basics of Statistical Mechanics Review of ensembles Microcanonical, canonical, Maxwell-Boltzmann Constant pressure, temperature, volume, Thermodynamic limit Ergodicity (see online notes also) Reading assignment:
Basics of Statistical Mechanics
 Basics of Statistical Mechanics Review of ensembles Microcanonical, canonical, Maxwell-Boltzmann Constant pressure, temperature, volume, Thermodynamic limit Ergodicity (see online notes also) Reading assignment:
Basics of Statistical Mechanics Review of ensembles Microcanonical, canonical, Maxwell-Boltzmann Constant pressure, temperature, volume, Thermodynamic limit Ergodicity (see online notes also) Reading assignment:
Introduction to molecular dynamics
 1 Introduction to molecular dynamics Yves Lansac Université François Rabelais, Tours, France Visiting MSE, GIST for the summer Molecular Simulation 2 Molecular simulation is a computational experiment.
1 Introduction to molecular dynamics Yves Lansac Université François Rabelais, Tours, France Visiting MSE, GIST for the summer Molecular Simulation 2 Molecular simulation is a computational experiment.
Ab initio molecular dynamics. Simone Piccinin CNR-IOM DEMOCRITOS Trieste, Italy. Bangalore, 04 September 2014
 Ab initio molecular dynamics Simone Piccinin CNR-IOM DEMOCRITOS Trieste, Italy Bangalore, 04 September 2014 What is MD? 1) Liquid 4) Dye/TiO2/electrolyte 2) Liquids 3) Solvated protein 5) Solid to liquid
Ab initio molecular dynamics Simone Piccinin CNR-IOM DEMOCRITOS Trieste, Italy Bangalore, 04 September 2014 What is MD? 1) Liquid 4) Dye/TiO2/electrolyte 2) Liquids 3) Solvated protein 5) Solid to liquid
LECTURE 6 : BASICS FOR MOLECULAR SIMULATIONS - Historical perspective - Skimming over Statistical Mechanics - General idea of Molecular Dynamics -
 LECTURE 6 : BASICS FOR MOLECULAR SIMULATIONS - Historical perspective - Skimming over Statistical Mechanics - General idea of Molecular Dynamics - Force calculations, structure of MD, equations of motion
LECTURE 6 : BASICS FOR MOLECULAR SIMULATIONS - Historical perspective - Skimming over Statistical Mechanics - General idea of Molecular Dynamics - Force calculations, structure of MD, equations of motion
Molecular Dynamics Simulation of Argon
 Molecular Dynamics Simulation of Argon The fundamental work on this problem was done by A. Rahman, Phys. Rev. 136, A405 (1964). It was extended in many important ways by L. Verlet, Phys. Rev. 159, 98 (1967),
Molecular Dynamics Simulation of Argon The fundamental work on this problem was done by A. Rahman, Phys. Rev. 136, A405 (1964). It was extended in many important ways by L. Verlet, Phys. Rev. 159, 98 (1967),
MD Thermodynamics. Lecture 12 3/26/18. Harvard SEAS AP 275 Atomistic Modeling of Materials Boris Kozinsky
 MD Thermodynamics Lecture 1 3/6/18 1 Molecular dynamics The force depends on positions only (not velocities) Total energy is conserved (micro canonical evolution) Newton s equations of motion (second order
MD Thermodynamics Lecture 1 3/6/18 1 Molecular dynamics The force depends on positions only (not velocities) Total energy is conserved (micro canonical evolution) Newton s equations of motion (second order
Coarse-Grained Models!
 Coarse-Grained Models! Large and complex molecules (e.g. long polymers) can not be simulated on the all-atom level! Requires coarse-graining of the model! Coarse-grained models are usually also particles
Coarse-Grained Models! Large and complex molecules (e.g. long polymers) can not be simulated on the all-atom level! Requires coarse-graining of the model! Coarse-grained models are usually also particles
Temperature and Pressure Controls
 Ensembles Temperature and Pressure Controls 1. (E, V, N) microcanonical (constant energy) 2. (T, V, N) canonical, constant volume 3. (T, P N) constant pressure 4. (T, V, µ) grand canonical #2, 3 or 4 are
Ensembles Temperature and Pressure Controls 1. (E, V, N) microcanonical (constant energy) 2. (T, V, N) canonical, constant volume 3. (T, P N) constant pressure 4. (T, V, µ) grand canonical #2, 3 or 4 are
Analysis of the simulation
 Analysis of the simulation Marcus Elstner and Tomáš Kubař January 7, 2014 Thermodynamic properties time averages of thermodynamic quantites correspond to ensemble averages (ergodic theorem) some quantities
Analysis of the simulation Marcus Elstner and Tomáš Kubař January 7, 2014 Thermodynamic properties time averages of thermodynamic quantites correspond to ensemble averages (ergodic theorem) some quantities
PHY2053 Lecture 11 Conservation of Energy. Conservation of Energy Kinetic Energy Gravitational Potential Energy
 PHY2053 Lecture 11 Conservation of Energy Conservation of Energy Kinetic Energy Gravitational Potential Energy Symmetries in Physics Symmetry - fundamental / descriptive property of the Universe itself
PHY2053 Lecture 11 Conservation of Energy Conservation of Energy Kinetic Energy Gravitational Potential Energy Symmetries in Physics Symmetry - fundamental / descriptive property of the Universe itself
Temperature and Pressure Controls
 Ensembles Temperature and Pressure Controls 1. (E, V, N) microcanonical (constant energy) 2. (T, V, N) canonical, constant volume 3. (T, P N) constant pressure 4. (T, V, µ) grand canonical #2, 3 or 4 are
Ensembles Temperature and Pressure Controls 1. (E, V, N) microcanonical (constant energy) 2. (T, V, N) canonical, constant volume 3. (T, P N) constant pressure 4. (T, V, µ) grand canonical #2, 3 or 4 are
Classical Equations of Motion
 3 Classical Equations of Motion Several formulations are in use Newtonian Lagrangian Hamiltonian Advantages of non-newtonian formulations more general, no need for fictitious forces better suited for multiparticle
3 Classical Equations of Motion Several formulations are in use Newtonian Lagrangian Hamiltonian Advantages of non-newtonian formulations more general, no need for fictitious forces better suited for multiparticle
Development of a Water Cluster Evaporation Model using Molecular Dynamics
 Development of a Water Cluster Evaporation Model using Molecular Dynamics Arnaud Borner, Zheng Li, Deborah A. Levin. Department of Aerospace Engineering, The Pennsylvania State University, University Park,
Development of a Water Cluster Evaporation Model using Molecular Dynamics Arnaud Borner, Zheng Li, Deborah A. Levin. Department of Aerospace Engineering, The Pennsylvania State University, University Park,
Computation of non-bonded interactions: Part 1
 Computation of non-bonded interactions: Part 1 Computation of non-local (non-bonded) interactions in the potential function E p non-local scales with the number of atoms as N. For the large molecular systems
Computation of non-bonded interactions: Part 1 Computation of non-local (non-bonded) interactions in the potential function E p non-local scales with the number of atoms as N. For the large molecular systems
Example questions for Molecular modelling (Level 4) Dr. Adrian Mulholland
 Example questions for Molecular modelling (Level 4) Dr. Adrian Mulholland 1) Question. Two methods which are widely used for the optimization of molecular geometies are the Steepest descents and Newton-Raphson
Example questions for Molecular modelling (Level 4) Dr. Adrian Mulholland 1) Question. Two methods which are widely used for the optimization of molecular geometies are the Steepest descents and Newton-Raphson
Why Proteins Fold? (Parts of this presentation are based on work of Ashok Kolaskar) CS490B: Introduction to Bioinformatics Mar.
 Why Proteins Fold? (Parts of this presentation are based on work of Ashok Kolaskar) CS490B: Introduction to Bioinformatics Mar. 25, 2002 Molecular Dynamics: Introduction At physiological conditions, the
Why Proteins Fold? (Parts of this presentation are based on work of Ashok Kolaskar) CS490B: Introduction to Bioinformatics Mar. 25, 2002 Molecular Dynamics: Introduction At physiological conditions, the
Molecular Dynamics. Timothy O Sullivan Supervised by Naida Lacevic & John Sader The University of Melbourne
 Molecular Dynamics Timothy O Sullivan Supervised by Naida Lacevic & John Sader The University of Melbourne 1 1 Introduction Molecular Dynamics (MD) is a method of simulating systems of nanoscale volumes,
Molecular Dynamics Timothy O Sullivan Supervised by Naida Lacevic & John Sader The University of Melbourne 1 1 Introduction Molecular Dynamics (MD) is a method of simulating systems of nanoscale volumes,
Molecular computer experiment
 Molecular computer experiment 1/13 Also simulation or pseudoexperiment REAL EXPERIMENT Record everything in a lab notebook Build the experimental apparatus (from parts) Purchase chemicals, synthetise if
Molecular computer experiment 1/13 Also simulation or pseudoexperiment REAL EXPERIMENT Record everything in a lab notebook Build the experimental apparatus (from parts) Purchase chemicals, synthetise if
Molecular Dynamics Simulation of a Nanoconfined Water Film
 Molecular Dynamics Simulation of a Nanoconfined Water Film Kyle Lindquist, Shu-Han Chao May 7, 2013 1 Introduction The behavior of water confined in nano-scale environment is of interest in many applications.
Molecular Dynamics Simulation of a Nanoconfined Water Film Kyle Lindquist, Shu-Han Chao May 7, 2013 1 Introduction The behavior of water confined in nano-scale environment is of interest in many applications.
N-body simulations. Phys 750 Lecture 10
 N-body simulations Phys 750 Lecture 10 N-body simulations Forward integration in time of strongly coupled classical particles or (rigid-body composite objects) dissipation Brownian motion Very general
N-body simulations Phys 750 Lecture 10 N-body simulations Forward integration in time of strongly coupled classical particles or (rigid-body composite objects) dissipation Brownian motion Very general
Scalable, performant, and resilient large-scale applications of molecular process engineering
 Scalable, performant, and resilient large-scale applications of molecular process engineering M. Horsch,1 P. Gralka,2 C. Niethammer,3 N. Tchipev,4 J. Vrabec,5 H. Hasse1 1 University of Kaiserslautern,
Scalable, performant, and resilient large-scale applications of molecular process engineering M. Horsch,1 P. Gralka,2 C. Niethammer,3 N. Tchipev,4 J. Vrabec,5 H. Hasse1 1 University of Kaiserslautern,
Introduction to model potential Molecular Dynamics A3hourcourseatICTP
 Introduction to model potential Molecular Dynamics A3hourcourseatICTP Alessandro Mattoni 1 1 Istituto Officina dei Materiali CNR-IOM Unità di Cagliari SLACS ICTP School on numerical methods for energy,
Introduction to model potential Molecular Dynamics A3hourcourseatICTP Alessandro Mattoni 1 1 Istituto Officina dei Materiali CNR-IOM Unità di Cagliari SLACS ICTP School on numerical methods for energy,
Using Molecular Dynamics to Compute Properties CHEM 430
 Using Molecular Dynamics to Compute Properties CHEM 43 Heat Capacity and Energy Fluctuations Running an MD Simulation Equilibration Phase Before data-collection and results can be analyzed the system
Using Molecular Dynamics to Compute Properties CHEM 43 Heat Capacity and Energy Fluctuations Running an MD Simulation Equilibration Phase Before data-collection and results can be analyzed the system
ON THE INTEGRATION OF EQUATIONS OF MOTION: FEM AND MOLECULAR DYNAMICS PROBLEMS
 8th International Congress on Computational Mechanics, Volos, 1-15 July 015 ON THE INTEGRATION OF EQUATIONS OF MOTION: FEM AND MOLECULAR DYNAMICS PROBLEMS E.G. Kakouris, V.K. Koumousis Institute of Structural
8th International Congress on Computational Mechanics, Volos, 1-15 July 015 ON THE INTEGRATION OF EQUATIONS OF MOTION: FEM AND MOLECULAR DYNAMICS PROBLEMS E.G. Kakouris, V.K. Koumousis Institute of Structural
Department of Chemical Engineering University of California, Santa Barbara Spring Exercise 3. Due: Thursday, 5/3/12
 Department of Chemical Engineering ChE 210D University of California, Santa Barbara Spring 2012 Exercise 3 Due: Thursday, 5/3/12 Objective: To learn how to write & compile Fortran libraries for Python,
Department of Chemical Engineering ChE 210D University of California, Santa Barbara Spring 2012 Exercise 3 Due: Thursday, 5/3/12 Objective: To learn how to write & compile Fortran libraries for Python,
Developing Monovalent Ion Parameters for the Optimal Point Charge (OPC) Water Model. John Dood Hope College
 Developing Monovalent Ion Parameters for the Optimal Point Charge (OPC) Water Model John Dood Hope College What are MD simulations? Model and predict the structure and dynamics of large macromolecules.
Developing Monovalent Ion Parameters for the Optimal Point Charge (OPC) Water Model John Dood Hope College What are MD simulations? Model and predict the structure and dynamics of large macromolecules.
arxiv: v1 [cond-mat.stat-mech] 7 Mar 2019
![arxiv: v1 [cond-mat.stat-mech] 7 Mar 2019 arxiv: v1 [cond-mat.stat-mech] 7 Mar 2019](/thumbs/96/126841930.jpg) Langevin thermostat for robust configurational and kinetic sampling Oded Farago, Department of Chemistry, University of Cambridge, Lensfield Road, Cambridge CB EW, United Kingdom Department of Biomedical
Langevin thermostat for robust configurational and kinetic sampling Oded Farago, Department of Chemistry, University of Cambridge, Lensfield Road, Cambridge CB EW, United Kingdom Department of Biomedical
CE 530 Molecular Simulation
 CE 530 Molecular Simulation Lecture Molecular Dynamics Simulation David A. Kofke Department of Chemical Engineering SUNY Buffalo kofke@eng.buffalo.edu MD of hard disks intuitive Review and Preview collision
CE 530 Molecular Simulation Lecture Molecular Dynamics Simulation David A. Kofke Department of Chemical Engineering SUNY Buffalo kofke@eng.buffalo.edu MD of hard disks intuitive Review and Preview collision
Random Walks A&T and F&S 3.1.2
 Random Walks A&T 110-123 and F&S 3.1.2 As we explained last time, it is very difficult to sample directly a general probability distribution. - If we sample from another distribution, the overlap will
Random Walks A&T 110-123 and F&S 3.1.2 As we explained last time, it is very difficult to sample directly a general probability distribution. - If we sample from another distribution, the overlap will
Brownian Motion and Langevin Equations
 1 Brownian Motion and Langevin Equations 1.1 Langevin Equation and the Fluctuation- Dissipation Theorem The theory of Brownian motion is perhaps the simplest approximate way to treat the dynamics of nonequilibrium
1 Brownian Motion and Langevin Equations 1.1 Langevin Equation and the Fluctuation- Dissipation Theorem The theory of Brownian motion is perhaps the simplest approximate way to treat the dynamics of nonequilibrium
A (short) practical introduction to kinetic theory and thermodynamic properties of gases through molecular dynamics
 A (short) practical introduction to kinetic theory and thermodynamic properties of gases through molecular dynamics Miguel A. Caro mcaroba@gmail.com March 28, 2018 Contents 1 Preface 3 2 Review of thermodynamics
A (short) practical introduction to kinetic theory and thermodynamic properties of gases through molecular dynamics Miguel A. Caro mcaroba@gmail.com March 28, 2018 Contents 1 Preface 3 2 Review of thermodynamics
Ab initio molecular dynamics and nuclear quantum effects
 Ab initio molecular dynamics and nuclear quantum effects Luca M. Ghiringhelli Fritz Haber Institute Hands on workshop density functional theory and beyond: First principles simulations of molecules and
Ab initio molecular dynamics and nuclear quantum effects Luca M. Ghiringhelli Fritz Haber Institute Hands on workshop density functional theory and beyond: First principles simulations of molecules and
Speeding up path integral simulations
 Speeding up path integral simulations Thomas Markland and David Manolopoulos Department of Chemistry University of Oxford Funded by the US Office of Naval Research and the UK EPSRC Outline 1. Ring polymer
Speeding up path integral simulations Thomas Markland and David Manolopoulos Department of Chemistry University of Oxford Funded by the US Office of Naval Research and the UK EPSRC Outline 1. Ring polymer
Energy-stepping integrators in
 Energy-stepping integrators in Lagrangian mechanics M. Gonzalez, B. Schmidt, M. Ortiz California Institute of Technology Technische Universität München Structured Integrators Workshop Caltech, May 8, 2009
Energy-stepping integrators in Lagrangian mechanics M. Gonzalez, B. Schmidt, M. Ortiz California Institute of Technology Technische Universität München Structured Integrators Workshop Caltech, May 8, 2009
the EL equation for the x coordinate is easily seen to be (exercise)
 Physics 6010, Fall 2016 Relevant Sections in Text: 1.3 1.6 Examples After all this formalism it is a good idea to spend some time developing a number of illustrative examples. These examples represent
Physics 6010, Fall 2016 Relevant Sections in Text: 1.3 1.6 Examples After all this formalism it is a good idea to spend some time developing a number of illustrative examples. These examples represent
Computer simulation methods (1) Dr. Vania Calandrini
 Computer simulation methods (1) Dr. Vania Calandrini Why computational methods To understand and predict the properties of complex systems (many degrees of freedom): liquids, solids, adsorption of molecules
Computer simulation methods (1) Dr. Vania Calandrini Why computational methods To understand and predict the properties of complex systems (many degrees of freedom): liquids, solids, adsorption of molecules
Lecture IV: Time Discretization
 Lecture IV: Time Discretization Motivation Kinematics: continuous motion in continuous time Computer simulation: Discrete time steps t Discrete Space (mesh particles) Updating Position Force induces acceleration.
Lecture IV: Time Discretization Motivation Kinematics: continuous motion in continuous time Computer simulation: Discrete time steps t Discrete Space (mesh particles) Updating Position Force induces acceleration.
Non-bonded interactions
 speeding up the number-crunching continued Marcus Elstner and Tomáš Kubař December 3, 2013 why care? number of individual pair-wise interactions bonded interactions proportional to N: O(N) non-bonded interactions
speeding up the number-crunching continued Marcus Elstner and Tomáš Kubař December 3, 2013 why care? number of individual pair-wise interactions bonded interactions proportional to N: O(N) non-bonded interactions
Monte Carlo. Lecture 15 4/9/18. Harvard SEAS AP 275 Atomistic Modeling of Materials Boris Kozinsky
 Monte Carlo Lecture 15 4/9/18 1 Sampling with dynamics In Molecular Dynamics we simulate evolution of a system over time according to Newton s equations, conserving energy Averages (thermodynamic properties)
Monte Carlo Lecture 15 4/9/18 1 Sampling with dynamics In Molecular Dynamics we simulate evolution of a system over time according to Newton s equations, conserving energy Averages (thermodynamic properties)
Computer Simulation of Shock Waves in Condensed Matter. Matthew R. Farrow 2 November 2007
 Computer Simulation of Shock Waves in Condensed Matter Matthew R. Farrow 2 November 2007 Outline of talk Shock wave theory Results Conclusion Computer simulation of shock waves Shock Wave Theory Shock
Computer Simulation of Shock Waves in Condensed Matter Matthew R. Farrow 2 November 2007 Outline of talk Shock wave theory Results Conclusion Computer simulation of shock waves Shock Wave Theory Shock
Project 4/5 - Molecular dynamics part II: advanced study of Lennard-Jones fluids, deadline December 1st (noon)
 Format for delivery of report and programs The format of the project is that of a printed file or hand-written report. The programs should also be included with the report. Write only your candidate number
Format for delivery of report and programs The format of the project is that of a printed file or hand-written report. The programs should also be included with the report. Write only your candidate number
Physics 115/242 The leapfrog method and other symplectic algorithms for integrating Newton s laws of motion
 Physics 115/242 The leapfrog method and other symplectic algorithms for integrating Newton s laws of motion Peter Young (Dated: April 14, 2009) I. INTRODUCTION One frequently obtains detailed dynamical
Physics 115/242 The leapfrog method and other symplectic algorithms for integrating Newton s laws of motion Peter Young (Dated: April 14, 2009) I. INTRODUCTION One frequently obtains detailed dynamical
NUCLEATION IN FINITE SYSTEMS: THEORY AND COMPUTER SIMULATION*t. M. RAO and B. J. BERNE. Chemistry Dept, Columbia University, New York, U.S.A.
 NUCLEATION IN FINITE SYSTEMS: THEORY AND COMPUTER SIMULATION*t M. RAO and B. J. BERNE Chemistry Dept, Columbia University, New York, U.S.A. Abstract. The free energy of formation of a droplet in a finite
NUCLEATION IN FINITE SYSTEMS: THEORY AND COMPUTER SIMULATION*t M. RAO and B. J. BERNE Chemistry Dept, Columbia University, New York, U.S.A. Abstract. The free energy of formation of a droplet in a finite
Introduction. Chapter The Purpose of Statistical Mechanics
 Chapter 1 Introduction 1.1 The Purpose of Statistical Mechanics Statistical Mechanics is the mechanics developed to treat a collection of a large number of atoms or particles. Such a collection is, for
Chapter 1 Introduction 1.1 The Purpose of Statistical Mechanics Statistical Mechanics is the mechanics developed to treat a collection of a large number of atoms or particles. Such a collection is, for
Javier Junquera. Statistical mechanics
 Javier Junquera Statistical mechanics From the microscopic to the macroscopic level: the realm of statistical mechanics Computer simulations Thermodynamic state Generates information at the microscopic
Javier Junquera Statistical mechanics From the microscopic to the macroscopic level: the realm of statistical mechanics Computer simulations Thermodynamic state Generates information at the microscopic
Parallelization of Molecular Dynamics (with focus on Gromacs) SeSE 2014 p.1/29
 Parallelization of Molecular Dynamics (with focus on Gromacs) SeSE 2014 p.1/29 Outline A few words on MD applications and the GROMACS package The main work in an MD simulation Parallelization Stream computing
Parallelization of Molecular Dynamics (with focus on Gromacs) SeSE 2014 p.1/29 Outline A few words on MD applications and the GROMACS package The main work in an MD simulation Parallelization Stream computing
ICCP Project 2 - Advanced Monte Carlo Methods Choose one of the three options below
 ICCP Project 2 - Advanced Monte Carlo Methods Choose one of the three options below Introduction In statistical physics Monte Carlo methods are considered to have started in the Manhattan project (1940
ICCP Project 2 - Advanced Monte Carlo Methods Choose one of the three options below Introduction In statistical physics Monte Carlo methods are considered to have started in the Manhattan project (1940
Kobe-Brown Simulation Summer School 2015 Project: DPD Simulation of a Membrane
 Kobe-Brown Simulation Summer School 2015 Project: DPD Simulation of a Membrane Clark Bowman Karen Larson Yuji Funaki Ross Parker Tae Woo Kim Week 1: Introduction to Molecular Dynamics (MD) Computer simulation
Kobe-Brown Simulation Summer School 2015 Project: DPD Simulation of a Membrane Clark Bowman Karen Larson Yuji Funaki Ross Parker Tae Woo Kim Week 1: Introduction to Molecular Dynamics (MD) Computer simulation
MOLECULAR DYNAMICS SIMULATION OF VAPOR BUBBLE NUCLEATION ON A SOLID SURFACE. Tatsuto Kimura and Shigeo Maruyama
 MOLECULAR DYNAMICS SIMULATION OF VAPOR BUBBLE NUCLEATION ON A SOLID SURFACE Tatsuto Kimura and Shigeo Maruyama * Department of Mechanical Engineering, The University of Tokyo, 7-- Hongo, Bunkyo-ku, Tokyo
MOLECULAR DYNAMICS SIMULATION OF VAPOR BUBBLE NUCLEATION ON A SOLID SURFACE Tatsuto Kimura and Shigeo Maruyama * Department of Mechanical Engineering, The University of Tokyo, 7-- Hongo, Bunkyo-ku, Tokyo
Introduction Statistical Thermodynamics. Monday, January 6, 14
 Introduction Statistical Thermodynamics 1 Molecular Simulations Molecular dynamics: solve equations of motion Monte Carlo: importance sampling r 1 r 2 r n MD MC r 1 r 2 2 r n 2 3 3 4 4 Questions How can
Introduction Statistical Thermodynamics 1 Molecular Simulations Molecular dynamics: solve equations of motion Monte Carlo: importance sampling r 1 r 2 r n MD MC r 1 r 2 2 r n 2 3 3 4 4 Questions How can
Intermolecular Forces and Monte-Carlo Integration 열역학특수연구
 Intermolecular Forces and Monte-Carlo Integration 열역학특수연구 2003.3.28 Source of the lecture note. J.M.Prausnitz and others, Molecular Thermodynamics of Fluid Phase Equiliria Atkins, Physical Chemistry Lecture
Intermolecular Forces and Monte-Carlo Integration 열역학특수연구 2003.3.28 Source of the lecture note. J.M.Prausnitz and others, Molecular Thermodynamics of Fluid Phase Equiliria Atkins, Physical Chemistry Lecture
Ideal Gas Behavior. NC State University
 Chemistry 331 Lecture 6 Ideal Gas Behavior NC State University Macroscopic variables P, T Pressure is a force per unit area (P= F/A) The force arises from the change in momentum as particles hit an object
Chemistry 331 Lecture 6 Ideal Gas Behavior NC State University Macroscopic variables P, T Pressure is a force per unit area (P= F/A) The force arises from the change in momentum as particles hit an object
Error Analysis of the Poisson P 3 MForce Field Scheme for Particle-Based Simulations of Biological Systems
 Journal of Computational Electronics 4: 179 183, 2005 c 2005 Springer Science + Business Media, Inc. Manufactured in The Netherlands. Error Analysis of the Poisson P 3 MForce Field Scheme for Particle-Based
Journal of Computational Electronics 4: 179 183, 2005 c 2005 Springer Science + Business Media, Inc. Manufactured in The Netherlands. Error Analysis of the Poisson P 3 MForce Field Scheme for Particle-Based
Exploring the energy landscape
 Exploring the energy landscape ChE210D Today's lecture: what are general features of the potential energy surface and how can we locate and characterize minima on it Derivatives of the potential energy
Exploring the energy landscape ChE210D Today's lecture: what are general features of the potential energy surface and how can we locate and characterize minima on it Derivatives of the potential energy
Computational Chemistry - MD Simulations
 Computational Chemistry - MD Simulations P. Ojeda-May pedro.ojeda-may@umu.se Department of Chemistry/HPC2N, Umeå University, 901 87, Sweden. May 2, 2017 Table of contents 1 Basics on MD simulations Accelerated
Computational Chemistry - MD Simulations P. Ojeda-May pedro.ojeda-may@umu.se Department of Chemistry/HPC2N, Umeå University, 901 87, Sweden. May 2, 2017 Table of contents 1 Basics on MD simulations Accelerated
Constructing a neighbour list
 Constructing a neighbour list In MD simulations (and actually many other applications) one of the central operations is the calculation of distances between atoms. In MD this is needed in the energy and
Constructing a neighbour list In MD simulations (and actually many other applications) one of the central operations is the calculation of distances between atoms. In MD this is needed in the energy and
Simulation of molecular systems by molecular dynamics
 Simulation of molecular systems by molecular dynamics Yohann Moreau yohann.moreau@ujf-grenoble.fr November 26, 2015 Yohann Moreau (UJF) Molecular Dynamics, Label RFCT 2015 November 26, 2015 1 / 35 Introduction
Simulation of molecular systems by molecular dynamics Yohann Moreau yohann.moreau@ujf-grenoble.fr November 26, 2015 Yohann Moreau (UJF) Molecular Dynamics, Label RFCT 2015 November 26, 2015 1 / 35 Introduction
Charge equilibration
 Charge equilibration Taylor expansion of energy of atom A @E E A (Q) =E A0 + Q A + 1 @Q A 0 2 Q2 A @ 2 E @Q 2 A 0 +... The corresponding energy of cation/anion and neutral atom E A (+1) = E A0 + @E @Q
Charge equilibration Taylor expansion of energy of atom A @E E A (Q) =E A0 + Q A + 1 @Q A 0 2 Q2 A @ 2 E @Q 2 A 0 +... The corresponding energy of cation/anion and neutral atom E A (+1) = E A0 + @E @Q
PH 548 Atomistic Simulation Techniques
 PH 548 Atomistic Simulation Techniques Lectures: Lab: 2-0-2-6 Tuesday 12-1 PM Thursday 12-1 PM Monday 2-5 PM P. K. Padmanabhan PH548: Two Excellent Books How to do it? 1 2 F 12 F 23 3 F i = F ij j i F
PH 548 Atomistic Simulation Techniques Lectures: Lab: 2-0-2-6 Tuesday 12-1 PM Thursday 12-1 PM Monday 2-5 PM P. K. Padmanabhan PH548: Two Excellent Books How to do it? 1 2 F 12 F 23 3 F i = F ij j i F
MOLECULAR DYNAMICS SIMULATION OF HETEROGENEOUS NUCLEATION OF LIQUID DROPLET ON SOLID SURFACE
 MOLECULAR DYNAMICS SIMULATION OF HETEROGENEOUS NUCLEATION OF LIQUID DROPLET ON SOLID SURFACE Tatsuto Kimura* and Shigeo Maruyama** *Department of Mechanical Engineering, The University of Tokyo 7-- Hongo,
MOLECULAR DYNAMICS SIMULATION OF HETEROGENEOUS NUCLEATION OF LIQUID DROPLET ON SOLID SURFACE Tatsuto Kimura* and Shigeo Maruyama** *Department of Mechanical Engineering, The University of Tokyo 7-- Hongo,
An Extended van der Waals Equation of State Based on Molecular Dynamics Simulation
 J. Comput. Chem. Jpn., Vol. 8, o. 3, pp. 97 14 (9) c 9 Society of Computer Chemistry, Japan An Extended van der Waals Equation of State Based on Molecular Dynamics Simulation Yosuke KATAOKA* and Yuri YAMADA
J. Comput. Chem. Jpn., Vol. 8, o. 3, pp. 97 14 (9) c 9 Society of Computer Chemistry, Japan An Extended van der Waals Equation of State Based on Molecular Dynamics Simulation Yosuke KATAOKA* and Yuri YAMADA
COMS 6100 Class Notes
 COMS 6100 Class Notes Daniel Solus September 20, 2016 1 General Remarks The Lecture notes submitted by the class have been very good. Integer division seemed to be a common oversight when working the Fortran
COMS 6100 Class Notes Daniel Solus September 20, 2016 1 General Remarks The Lecture notes submitted by the class have been very good. Integer division seemed to be a common oversight when working the Fortran
Heat Transport in Glass-Forming Liquids
 Heat Transport in Glass-Forming Liquids by VINAY VAIBHAV The Institute of Mathematical Sciences CIT Campus, Taramani, Chennai 600 113, India. A thesis submitted in partial fulfillment of requirements for
Heat Transport in Glass-Forming Liquids by VINAY VAIBHAV The Institute of Mathematical Sciences CIT Campus, Taramani, Chennai 600 113, India. A thesis submitted in partial fulfillment of requirements for
Imperfect Gases. NC State University
 Chemistry 431 Lecture 3 Imperfect Gases NC State University The Compression Factor One way to represent the relationship between ideal and real gases is to plot the deviation from ideality as the gas is
Chemistry 431 Lecture 3 Imperfect Gases NC State University The Compression Factor One way to represent the relationship between ideal and real gases is to plot the deviation from ideality as the gas is
Non-bonded interactions
 speeding up the number-crunching Marcus Elstner and Tomáš Kubař May 8, 2015 why care? key to understand biomolecular structure and function binding of a ligand efficiency of a reaction color of a chromophore
speeding up the number-crunching Marcus Elstner and Tomáš Kubař May 8, 2015 why care? key to understand biomolecular structure and function binding of a ligand efficiency of a reaction color of a chromophore
Advanced Molecular Dynamics
 Advanced Molecular Dynamics Introduction May 2, 2017 Who am I? I am an associate professor at Theoretical Physics Topics I work on: Algorithms for (parallel) molecular simulations including GPU acceleration
Advanced Molecular Dynamics Introduction May 2, 2017 Who am I? I am an associate professor at Theoretical Physics Topics I work on: Algorithms for (parallel) molecular simulations including GPU acceleration
arxiv: v1 [physics.chem-ph] 4 Feb 2014
![arxiv: v1 [physics.chem-ph] 4 Feb 2014 arxiv: v1 [physics.chem-ph] 4 Feb 2014](/thumbs/93/113619303.jpg) Discrete dynamics versus analytic dynamics Søren Toxvaerd DNRF centre Glass and Time, IMFUFA, Department of Sciences, Roskilde University, arxiv:1402.0679v1 [physics.chem-ph] 4 Feb 2014 Postbox 260, DK-4000
Discrete dynamics versus analytic dynamics Søren Toxvaerd DNRF centre Glass and Time, IMFUFA, Department of Sciences, Roskilde University, arxiv:1402.0679v1 [physics.chem-ph] 4 Feb 2014 Postbox 260, DK-4000
JASS Modeling and visualization of molecular dynamic processes
 JASS 2009 Konstantin Shefov Modeling and visualization of molecular dynamic processes St Petersburg State University, Physics faculty, Department of Computational Physics Supervisor PhD Stepanova Margarita
JASS 2009 Konstantin Shefov Modeling and visualization of molecular dynamic processes St Petersburg State University, Physics faculty, Department of Computational Physics Supervisor PhD Stepanova Margarita
Lecture V: The game-engine loop & Time Integration
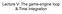 Lecture V: The game-engine loop & Time Integration The Basic Game-Engine Loop Previous state: " #, %(#) ( #, )(#) Forces -(#) Integrate velocities and positions Resolve Interpenetrations Per-body change
Lecture V: The game-engine loop & Time Integration The Basic Game-Engine Loop Previous state: " #, %(#) ( #, )(#) Forces -(#) Integrate velocities and positions Resolve Interpenetrations Per-body change
Molecular Dynamics Simulation Study of Transport Properties of Diatomic Gases
 MD Simulation of Diatomic Gases Bull. Korean Chem. Soc. 14, Vol. 35, No. 1 357 http://dx.doi.org/1.51/bkcs.14.35.1.357 Molecular Dynamics Simulation Study of Transport Properties of Diatomic Gases Song
MD Simulation of Diatomic Gases Bull. Korean Chem. Soc. 14, Vol. 35, No. 1 357 http://dx.doi.org/1.51/bkcs.14.35.1.357 Molecular Dynamics Simulation Study of Transport Properties of Diatomic Gases Song
Monte Carlo Model of Comet Dust
 Monte Carlo Model of Comet Dust Computational Physics Monte Carlo Model of Comet Dust Project 1 About Comets... Coma (Head) Dust Tail Ion Tail Nucleus 10^4-10^5 km 10^5-10^6 km 10^5-10^6 km 1 km All the
Monte Carlo Model of Comet Dust Computational Physics Monte Carlo Model of Comet Dust Project 1 About Comets... Coma (Head) Dust Tail Ion Tail Nucleus 10^4-10^5 km 10^5-10^6 km 10^5-10^6 km 1 km All the
Introduction to Hamiltonian systems
 Introduction to Hamiltonian systems Marlis Hochbruck Heinrich-Heine Universität Düsseldorf Oberwolfach Seminar, November 2008 Examples Mathematical biology: Lotka-Volterra model First numerical methods
Introduction to Hamiltonian systems Marlis Hochbruck Heinrich-Heine Universität Düsseldorf Oberwolfach Seminar, November 2008 Examples Mathematical biology: Lotka-Volterra model First numerical methods
Basics of molecular dynamics
 Basics of molecular dynamics The basic idea of molecular dynamics (MD) simulations is to calculate how a system of particles evolves in time. The method was first used by Alder and Wainwright in 1957 to
Basics of molecular dynamics The basic idea of molecular dynamics (MD) simulations is to calculate how a system of particles evolves in time. The method was first used by Alder and Wainwright in 1957 to
Molecular Dynamics Simulation of Heat Transfer and Phase Change During Laser Material Interaction
 Xinwei Wang Xianfan Xu e-mail: xxu@ecn.purdue.edu School of Mechanical Engineering, Purdue University, West Lafayette, IN 47907 Molecular Dynamics Simulation of Heat Transfer and Phase Change During Laser
Xinwei Wang Xianfan Xu e-mail: xxu@ecn.purdue.edu School of Mechanical Engineering, Purdue University, West Lafayette, IN 47907 Molecular Dynamics Simulation of Heat Transfer and Phase Change During Laser
Molecular Dynamics Simulations. Dr. Noelia Faginas Lago Dipartimento di Chimica,Biologia e Biotecnologie Università di Perugia
 Molecular Dynamics Simulations Dr. Noelia Faginas Lago Dipartimento di Chimica,Biologia e Biotecnologie Università di Perugia 1 An Introduction to Molecular Dynamics Simulations Macroscopic properties
Molecular Dynamics Simulations Dr. Noelia Faginas Lago Dipartimento di Chimica,Biologia e Biotecnologie Università di Perugia 1 An Introduction to Molecular Dynamics Simulations Macroscopic properties
CS115 Computer Simulation Project list
 CS115 Computer Simulation Project list The final project for this class is worth 40% of your grade. Below are your choices. You only need to do one of them. Project MC: Monte Carlo vs. Deterministic Volume
CS115 Computer Simulation Project list The final project for this class is worth 40% of your grade. Below are your choices. You only need to do one of them. Project MC: Monte Carlo vs. Deterministic Volume
