Effects of interaction between nanopore and polymer on translocation time
|
|
|
- Aubrie Patrick
- 5 years ago
- Views:
Transcription
1 Effects of interaction between nanopore and polymer on translocation time Mohammadreza Niknam Hamidabad and Rouhollah Haji Abdolvahab Physics Department, Iran University of Science and Technology (IUST), , Tehran, Iran Abstract: Here using LAMMPS molecular dynamics (MD) software, we simulate polymer translocation in 2 dimensions. We do the simulations for weak and moderate forces and for different pore diameters. Our results show that in both non-equilibrium and equilibrium initial conditions, translocation time will always increase by increasing binding energy and or increasing pore diameter. Moreover, scaling exponent of time versus force is in accordance to our predecessors. The comparison between equilibrium and non-equilibrium initial condition shows that the translocation time is very sensitive to the initial condition. Translocation time of the relaxed polymers for interaction energy of 8k B T is smaller from the non-equilibrium case even in the small energy of 1k B T. Keywords: Biopolymer, Translocation, Nanopore, Molecular Dynamic, Binding energy 1-Introduction Biopolymer s translocation through nanopores is a ubiquitous process in both biology and biotechnology. Protein translocation through membranes, RNA translocation through nucleopores and RNA injection through host cell by a virus are some instances [1, 2]. Moreover, it has technological applications such as drug delivery [3] and DNA sequencing [4]. Consequently, polymer translocation through the nanopore is one of the most active fields in biophysics and soft matter [5, 6]. In vivo Chaperone-assisted polymer translocation is one of the important deriving mechanism. In this method, which is used in cells [7], as soon as the polymer went through the Trans side, a protein, called chaperone, bound to it and prevent the polymer from backsliding [7-12]. However, in vitro, among the methods of polymer translocation through a nanopore, translocation deriving by external force is one of the most common experimental and computational approaches [6, 13-19]. In translocation deriving by an external force, different parameters like, external force, nanopore length, polymer length, crowding etc. on translocation time are investigated. One of the important parameters here is binding energy between channel wall and polymer, which we consider in this work. In our simulation, we investigate polymer translocation in two different initial conditions, namely, equilibrium and non-equilibrium in at least 8 different interaction energies. Moreover, we change the pore diameter and consider 3 different
2 amount of it. Simulation results indicate an increase in translocation time by increasing interaction energy and/or pore diameter in both cases. In what follows, we first outline our theoretical model. Then we report and analyze the results from our simulations. We finished our work by a conclusion based on the simulation results. 2-Theoretical model We model the polymer using a chain of masses and springs so that there is a finitely extensible nonlinear elastic (FENE) potential (Eq. 1) between adjacent monomers. There is also a Leonard-Jones potential (Eq. 2) between all monomers. U FENE = kr 0 2 ln ( 1- r2 R 0 2) (1) U LJ = 4ε [( σ 12 r ) - ( σ 6] r ) +ε r r cut 0 r > r cut (2) R 0 is the maximum allowed distance between adjacent monomers and k is the spring constant. σ, shows the monomer size, ε is the potential depth and r cut is the effective radius of the Leonard-Jones potential. The Leonard-Jones potential is also used for pore walls with a different r cut. We use of Langevin dynamics for the simulation. In this method for each monomer, one can write: mr i = F i C + F i F + F i R (3) in which m is the monomer mass, F i C calls for conservative forces, F i F is the friction and F i R shows stochastic force on the monomer. F i R is related to monomer s velocity as: F F i = ξ Vi (4) where ξ is the friction constant. For conservative force: F i C = (U LJ + U FENE ) + F external (5)
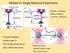
3 F external is the external force on the polymer through nanopore which we define: F external = Fx (6) where the direction of the force coincides with the axis of the nanopore through the Trans side. 2-1-Initial configuration and simulation parameters Nanopore has a length of 6σ, where σ is the size of a monomer and we consider pores of diameters 3, 4 and 5σ. We put the first monomer at the end of the nanopore. We situate the other monomers of the polymer in the equilibrium state with respect to each other but in a straight line in front of the pore. In non-equilibrium initial condition, we just start the simulation while in equilibrium initial condition the polymer have enough time to equilibrate and then the translocation starts. In equilibration of the second initial state, we fix the monomers through the channel. This part of simthe ulation, takes of around 20% (for the slowest polymers) to 40% (for the fastest polymer) of the whole simulation time. In order to decrease the error of the mean translocation time, we repeat the process for at least 1000 times. We use from gyration radius to find the equilibration. We see the gyration radius change over time and continue the simulation until it reaches to its saturation value with small fluctuations. Figure 1: Initial configuration for a non-equilibrium sample with a pore diameter of 5σ and N = 50.
4 Figure 2: Initial configuration for an equilibrium sample with a pore diameter of 5σ and N = 50. Time unit of the simulation is obtained from: t LJ =( mσ2 ε )1 2 This time unit is of the order of 10fs. Here we use two different external forces of weak and moderate for simulation. The moderate force satisfies the following relation [15]: (7) k B T σn ν F k BT σ (8) in which, ν is the Flory exponent and N is the total number of monomers. External forces that we used as weak and moderate forces are 3.5pN and 6.5pN, respectively. Effective interaction radius for inthe teraction of polymer and nanopore take as 2σ and for other interactions 2 1/6 σ. By setting the cut off distance r cut = 2 1/6 σ, to exclude attractive interactions, we models the polymer to be immersed in a good solvent [10]. Moreover, we take the parameter of energy (apart from polymer and nanopore) as ε = k B T. The length is of order angstrom. Friction coefficient equals ξ=0.7 m t LJ. Parameters of FENE potential taken to be k = 40 ε σ 2, R 0 = 1.5σ and the mass of each monomer m = 70amu [20].
5 3-Results and discussion Due to two states of different initial conditions of equilibrium and non-equilibrium, we present our results accordingly. 3-1-Non-equilibrium state Increasing binding energy will always increase the translocation time in nonequilibrium initial condition. Moreover, our simulation results reveal that increasing pore diameter from 3 to 4 and then 5σ will always increase the mean translocation time of the polymer in both weak and moderate forces (figures 3, 4). Figure 4: Mean translocation time of the polymer in the non-equilibrium state against interaction energy in 3 different pore size of 3, 4 and 5σ and for the weak force of 3.5pN. The mean is taken over 1000 sample.
6 Figure 3: Mean translocation time of the polymer in the non-equilibrium state against interaction energy in 3 different pore size of 3, 4 and 5σ and for the moderate force of 6.5pN. The mean is taken over 1000 sample. Comparison of figure 3 and figure 4 shows that the mean translocation time of the polymer in weak force is significantly larger than translocation time in the moderate force. This increase is expected due to decreasing the driving force. Moreover, the difference between translocation time of the polymer in case of pore diameter 5σ and to smaller pores increase by increasing driving force from weak to moderate. Polymer translocation time will decrease by increasing the external force. This change in moderate forces helps us to calculate a scaling exponent. We plot the translocation time versus force for moderate forces in the log-log axis in figure 5. We measure the scaling exponent of translocation time versus forces using the process of curve fitting as In these simulations the interaction energy ε = 6, pore diameter was 4σ and polymer length N = 50. It has been calculated in literature only for equilibrium initial state [14, 16-18].
7 Figure 5: Polymer translocation time against moderate forces in a log-log plot. The measured scaling exponent is equilibrium state In equilibrium initial configuration, the results for weak forces are similar to that of non-equilibrium state. Increasing both the binding energy and pore diameter, will increase the translocation time and slow down the translocation process (figure 6). An interesting result here is the slope of the curves. The slope of the curve for the diameter of size 5σ is significantly greater than that of pore diameters of 3 and 4σ.
8 Figure 6: Mean translocation time of the polymer in equilibrium state against interaction energy in 3 different pore size of 3, 4 and 5σ and for the weak force of 3.5pN. The mean is taken over 1000 sample. Figure 7: Mean translocation time of the polymer in equilibrium state against interaction energy in 3 different pore size of 3, 4 and 5σ and for the weak force of 6.5pN. The mean is taken over 1000 sample. We change the external force in the simulation and calculate the translocation time. Log-log plot of translocation time against forces is sketched in figure 8. Curve fitting on the figure shows that the scaling exponent of translocation time versus forces is
9 equal to Here the interaction energy is taken to be ε = 6, pore diameter was 4σ and polymer length N = 50. Luo et al. in 2009 report the exponent 1 for weak and moderate forces[18]. Huopaniemi et al. in 2007 calculate the exponent for weak, moderate and strong forces as -0.94[14], which are in good agreement with our calculations. Figure 8: Polymer translocation time against force in a log-log plot. The measured scaling exponent is
10 Figure 9: Plots of waiting times for both initial conditions and in 2 different binding energies 1 and 8, for pore of diameter size 5σ. Figure 10: Plots of cumulative waiting times for both initial conditions and in 2 different binding energies 1 and 8, for pore of diameter size 5σ.
11 Waiting time of the polymer translocation for pore diameter 5σ, in both initial conditions and for to binding energies of 1 and 8 is plotted in figure 9. It shows 3 stages of polymer translocation. Due to the empty space in the right the translocation of the first monomers are fast. Afterward, the translocation becomes slower and slower until, nearly, the middle of the polymer. Then the translocation s speed increase until the end. Cumulative of the above waiting time is shown in the figure 10. As expected from the waiting times, most of the separation of translocation times take place in the middle of the translocation. Conclusion We studied polymer translocation through a nanopore using a series of MD simulations. An external force derives the translocation. We investigated especially the effects of interaction between polymer and pore wall in our coarse-grained simulation. Our simulation results show that in both initial conditions, the translocation time will be increased by increasing the interaction energy and/or pore diameter. However, the initial condition is very important. As an example the translocation time for the unrelaxed polymer in binding energy of 1 is larger than the translocation time for the relaxed polymer with energy 8. The results also show that by increasing the pore diameter from 3σ to 5σ the difference between translocation times will be increased. This result may be used to separate different polymers using their translocation time. Acknowledgement: We use from LAMMPS molecular dynamic software for the simulation [21]. We also used from VMD for graphic representation [22] and Gnu plot for plotting the figures [23]. References 1. B. Alberts, et al., Molecular Biology of the Cell 2002: Garland Publishing. 2. A. Guyton, a.j.h., Textbook of Medical Physiology. 7th ed. 2005: Saunders. 3. Chang, D.C., Guide to Electroporation and Electrofusion. 1992, New York Academic. 4. Meller, A., Dynamics of polynucleotide transport through nanometer-scale pores. J. Phys. Condens. Matt., : p. R581-R Muthukumar, M., Polymer translocation through a hole. Journal of Chemical Physics, (22): p V.V. Palyulin, T.A.-N., and R. Metzler, Polymer translocation: the first two decades and the recent diversification. Soft Matter, (45): p
12 7. Danuta Tomkiewicz, N.N., and Arnold J.M. Driessen, Pushing, pulling and trapping Modes of motor protein supported protein translocation. Fed. Eur. Biochem. Soc. Lett., (15): p Abdolvahab, R.H., M.R. Ejtehadi, and R. Metzler, Sequence dependence of the binding energy in chaperone-driven polymer translocation through a nanopore. Physical Review E, (1): p Abdolvahab, R.H., Chaperone-driven polymer translocation through nanopore: Spatial distribution and binding energy. European Physical Journal E, (4): p Suhonen, P. and R. Linna, Chaperone-assisted translocation of flexible polymers in three dimensions. Physical Review E, (1): p Yu, W. and K. Luo, Polymer translocation through a nanopore driven by binding particles: Influence of chain rigidity. Physical Review E, (4): p Yu, W. and K. Luo, Chaperone-assisted translocation of a polymer through a nanopore. Journal of the American Chemical Society, (34): p Huopaniemi, I., K. Luo, and T. Ala-Nissila, Langevin dynamics simulations of polymer translocation through nanopores. Journal of Chemical Physics, (12): p Huopaniemi, I., et al., Polymer translocation through a nanopore under a pulling force. Physical Review E, (6): p M Magill, C.F., Ed Waller and H W de Haan, Translocation Time through a Nanopore with an Internal Cavity Is Minimal for Polymers of Intermediate Length. Phys. Rev. Lett., (24): p V Lehtola, R.L., and K Kaski, Critical evaluation of the computational methods used in the forced polymer translocation. Phys. Rev. E, : p Bhattacharya, A., et al., Scaling exponents of forced polymer translocation through a nanopore. European Physical Journal E, (4): p K Luo, T.A.-N., S Yingand, and R Metzler, Driven polymer translocation through nanopores: Slow-vs.-fast dynamics. EPPL, : p Luo, K., et al., Driven polymer translocation through nanopores: Slow-vs.-fast dynamics. Epl, (6): p Junfang Wang, Y.W., and Kaifu Luo, Dynamics of polymer translocation through kinked nanopores. J. Chem. Phys., (8): p Plimpton, S., Fast parallel algorithms for short-range molecular dynamics. J. Comp. Phys., (1): p Humphrey, W., A. Dalke, and K. Schulten, VMD: visual molecular dynamics. Journal of molecular graphics, (1): p T Williams, C.K.a.m.o., Gnuplot 4.4: an interactive plotting program
Scaling exponents of Forced Polymer Translocation through a nano-pore
 Scaling exponents of Forced Polymer Translocation through a nano-pore Aniket Bhattacharya, 1, William H. Morrison, 1 Kaifu Luo, 2 Tapio Ala-Nissila, 2, See-Chen Ying, Andrey Milchev, 4 and Kurt Binder
Scaling exponents of Forced Polymer Translocation through a nano-pore Aniket Bhattacharya, 1, William H. Morrison, 1 Kaifu Luo, 2 Tapio Ala-Nissila, 2, See-Chen Ying, Andrey Milchev, 4 and Kurt Binder
Huopaniemi, I.; Luo, K.; Ala-Nissilä, Tapio; Ying, S.C. Langevin Dynamics Simulations of Polymer Translocation through Nanopores
 Powered by TCPDF (www.tcpdf.org) This is an electronic reprint of the original article. This reprint may differ from the original in pagination and typographic detail. Huopaniemi, I.; Luo, K.; Ala-Nissilä,
Powered by TCPDF (www.tcpdf.org) This is an electronic reprint of the original article. This reprint may differ from the original in pagination and typographic detail. Huopaniemi, I.; Luo, K.; Ala-Nissilä,
Deconvoluting chain heterogeneity from driven translocation through a nano-pore
 epl draft Deconvoluting chain heterogeneity from driven translocation through a nano-pore Department of Physics, University of Central Florida, Orlando, Florida 32826-238, USA PACS 87.1.ap First pacs description
epl draft Deconvoluting chain heterogeneity from driven translocation through a nano-pore Department of Physics, University of Central Florida, Orlando, Florida 32826-238, USA PACS 87.1.ap First pacs description
arxiv: v3 [cond-mat.soft] 10 Nov 2008
![arxiv: v3 [cond-mat.soft] 10 Nov 2008 arxiv: v3 [cond-mat.soft] 10 Nov 2008](/thumbs/92/109756414.jpg) Scaling exponents of Forced Polymer Translocation through a nano-pore arxiv:88.1868v3 [cond-mat.soft] 1 Nov 28 Aniket Bhattacharya, 1, William H. Morrison, 1 Kaifu Luo, 2 Tapio Ala-Nissila, 2,3 See-Chen
Scaling exponents of Forced Polymer Translocation through a nano-pore arxiv:88.1868v3 [cond-mat.soft] 1 Nov 28 Aniket Bhattacharya, 1, William H. Morrison, 1 Kaifu Luo, 2 Tapio Ala-Nissila, 2,3 See-Chen
This is an electronic reprint of the original article. This reprint may differ from the original in pagination and typographic detail.
 Powered by TCPDF (www.tcpdf.org) This is an electronic reprint of the original article. This reprint may differ from the original in pagination and typographic detail. Luo, Kaifu; Ollila, Santtu T.T.;
Powered by TCPDF (www.tcpdf.org) This is an electronic reprint of the original article. This reprint may differ from the original in pagination and typographic detail. Luo, Kaifu; Ollila, Santtu T.T.;
Translocation Dynamics of a Semi-flexible Chain Under a Bias: Comparison with Tension Propagation Theory
 Translocation Dynamics of a Semi-flexible Chain Under a Bias: Comparison with Tension Propagation Theory Aniket Bhattacharya Department of Physics, University of Central Florida, Orlando, Flor ida 32816-2385,
Translocation Dynamics of a Semi-flexible Chain Under a Bias: Comparison with Tension Propagation Theory Aniket Bhattacharya Department of Physics, University of Central Florida, Orlando, Flor ida 32816-2385,
Out-of-equilibrium characteristics of a forced translocating chain through a nanopore
 PHYSICAL REVIEW E 8, 8 Out-of-equilibrium characteristics of a forced locating chain through a nanopore Aniket Bhattacharya, * and Kurt Binder Department of Physics, University of Central Florida, Orlando,
PHYSICAL REVIEW E 8, 8 Out-of-equilibrium characteristics of a forced locating chain through a nanopore Aniket Bhattacharya, * and Kurt Binder Department of Physics, University of Central Florida, Orlando,
Deconvoluting chain heterogeneity from driven translocation through a nanopore
 February 201 EPL, 9 (201) 38001 doi:.1209/029-07/9/38001 www.epljournal.org Deconvoluting chain heterogeneity from driven translocation through a nanopore Ramesh Adhikari and Aniket Bhattacharya Department
February 201 EPL, 9 (201) 38001 doi:.1209/029-07/9/38001 www.epljournal.org Deconvoluting chain heterogeneity from driven translocation through a nanopore Ramesh Adhikari and Aniket Bhattacharya Department
Heteropolymer translocation through nanopores
 THE JOURNAL OF CHEMICAL PHYSICS 126, 145101 2007 Heteropolymer translocation through nanopores Kaifu Luo a Laboratory of Physics, Helsinki University of Technology, P.O. Box 1100, FIN-02015 TKK Espoo,
THE JOURNAL OF CHEMICAL PHYSICS 126, 145101 2007 Heteropolymer translocation through nanopores Kaifu Luo a Laboratory of Physics, Helsinki University of Technology, P.O. Box 1100, FIN-02015 TKK Espoo,
ABSTRACT We investigate the translocation dynamics of heteropolymers driven through a
 Heteropolymer translocation through nanopores Kaifu Luo 1*, Tapio Ala-Nissila 1,2, See-Chen Ying 2 and Aniket Bhattacharya 3 1 Laboratory of Physics, Helsinki University of Technology, P.O. Box 1100, FIN-02015
Heteropolymer translocation through nanopores Kaifu Luo 1*, Tapio Ala-Nissila 1,2, See-Chen Ying 2 and Aniket Bhattacharya 3 1 Laboratory of Physics, Helsinki University of Technology, P.O. Box 1100, FIN-02015
arxiv: v1 [physics.bio-ph] 7 May 2009
![arxiv: v1 [physics.bio-ph] 7 May 2009 arxiv: v1 [physics.bio-ph] 7 May 2009](/thumbs/78/76737080.jpg) epl draft Dynamics of forced biopolymer translocation V.V. LEHTOLA, R.P. LINNA and K. KASKI arxiv:95.114v1 [physics.bio-ph] 7 May 29 Department of Biomedical Engineering and Computational Science, Helsinki
epl draft Dynamics of forced biopolymer translocation V.V. LEHTOLA, R.P. LINNA and K. KASKI arxiv:95.114v1 [physics.bio-ph] 7 May 29 Department of Biomedical Engineering and Computational Science, Helsinki
Dynamics of DNA translocation through an attractive nanopore
 Dynamics of DNA translocation through an attractive nanopore Kaifu Luo, 1,2, * Tapio Ala-Nissila, 1,3 See-Chen Ying, 3 and Aniket Bhattacharya 4 1 Department of Applied Physics, Helsinki University of
Dynamics of DNA translocation through an attractive nanopore Kaifu Luo, 1,2, * Tapio Ala-Nissila, 1,3 See-Chen Ying, 3 and Aniket Bhattacharya 4 1 Department of Applied Physics, Helsinki University of
Coarse-Grained Models!
 Coarse-Grained Models! Large and complex molecules (e.g. long polymers) can not be simulated on the all-atom level! Requires coarse-graining of the model! Coarse-grained models are usually also particles
Coarse-Grained Models! Large and complex molecules (e.g. long polymers) can not be simulated on the all-atom level! Requires coarse-graining of the model! Coarse-grained models are usually also particles
Confinement of polymer chains and gels
 Confinement of polymer chains and gels Nefeli Georgoulia - Student number: 70732831 1 Introduction Confinement of polymer chains is significant in industrial as well as biological applications. For this
Confinement of polymer chains and gels Nefeli Georgoulia - Student number: 70732831 1 Introduction Confinement of polymer chains is significant in industrial as well as biological applications. For this
(Entropic) Stochastic Resonance in Biological Systems at Mesoscale
 (Entropic) Stochastic Resonance in Biological Systems at Mesoscale Wokyung Sung Department of Physics, POSTECH, IBS center for Self-assembly and Complexity, Pohang, 790-784, South Korea As interconnected,
(Entropic) Stochastic Resonance in Biological Systems at Mesoscale Wokyung Sung Department of Physics, POSTECH, IBS center for Self-assembly and Complexity, Pohang, 790-784, South Korea As interconnected,
STRUCTURE OF IONS AND WATER AROUND A POLYELECTROLYTE IN A POLARIZABLE NANOPORE
 International Journal of Modern Physics C Vol. 2, No. 9 (29) 1485 1492 c World Scientific Publishing Company STRUCTURE OF IONS AND WATER AROUND A POLYELECTROLYTE IN A POLARIZABLE NANOPORE LEI GUO and ERIK
International Journal of Modern Physics C Vol. 2, No. 9 (29) 1485 1492 c World Scientific Publishing Company STRUCTURE OF IONS AND WATER AROUND A POLYELECTROLYTE IN A POLARIZABLE NANOPORE LEI GUO and ERIK
On the Dynamics and Disentanglement in Thin and Two-Dimensional Polymer Films
 J. Phys. IV France 1 (006) Pr1-1 c EDP Sciences, Les Ulis On the Dynamics and Disentanglement in Thin and Two-Dimensional Polymer Films H. Meyer, T. Kreer, A. Cavallo, J. P. Wittmer and J. Baschnagel 1
J. Phys. IV France 1 (006) Pr1-1 c EDP Sciences, Les Ulis On the Dynamics and Disentanglement in Thin and Two-Dimensional Polymer Films H. Meyer, T. Kreer, A. Cavallo, J. P. Wittmer and J. Baschnagel 1
Advantages of a Finite Extensible Nonlinear Elastic Potential in Lattice Boltzmann Simulations
 The Hilltop Review Volume 7 Issue 1 Winter 2014 Article 10 December 2014 Advantages of a Finite Extensible Nonlinear Elastic Potential in Lattice Boltzmann Simulations Tai-Hsien Wu Western Michigan University
The Hilltop Review Volume 7 Issue 1 Winter 2014 Article 10 December 2014 Advantages of a Finite Extensible Nonlinear Elastic Potential in Lattice Boltzmann Simulations Tai-Hsien Wu Western Michigan University
Anatoly B. Kolomeisky Department of Chemistry Center for Theoretical Biological Physics How to Understand Molecular Transport through Channels: The
 Anatoly B. Kolomeisy Department of Chemistry Center for Theoretical Biological Physics How to Understand Molecular Transport through Channels: The Role of Interactions Transport Through Channels Oil pumping
Anatoly B. Kolomeisy Department of Chemistry Center for Theoretical Biological Physics How to Understand Molecular Transport through Channels: The Role of Interactions Transport Through Channels Oil pumping
Proteins polymer molecules, folded in complex structures. Konstantin Popov Department of Biochemistry and Biophysics
 Proteins polymer molecules, folded in complex structures Konstantin Popov Department of Biochemistry and Biophysics Outline General aspects of polymer theory Size and persistent length of ideal linear
Proteins polymer molecules, folded in complex structures Konstantin Popov Department of Biochemistry and Biophysics Outline General aspects of polymer theory Size and persistent length of ideal linear
Driven translocation of a semiflexible polymer through a. nanopore OPEN.
 www.nature.com/scientificreports OPEN eceived: March 07 Accepted: 6 June 07 Published: xx xx xxxx Driven translocation of a semiflexible polymer through a nanopore Jalal Sarabadani, Timo Ikonen, Harri
www.nature.com/scientificreports OPEN eceived: March 07 Accepted: 6 June 07 Published: xx xx xxxx Driven translocation of a semiflexible polymer through a nanopore Jalal Sarabadani, Timo Ikonen, Harri
Biopolymer Entering into and Escaping from a Nanometer Pore
 CHEM. RES. CHINESE UNIVERSITIES 29, 25(6), 945 949 iopolymer Entering into and Escaping from a Nanometer Pore DING Ke-ian 1, ZHENG Yan-peng 1, GUAN Wei-un 2* and MA Yue-hui 2* 1. College of Life Sciences
CHEM. RES. CHINESE UNIVERSITIES 29, 25(6), 945 949 iopolymer Entering into and Escaping from a Nanometer Pore DING Ke-ian 1, ZHENG Yan-peng 1, GUAN Wei-un 2* and MA Yue-hui 2* 1. College of Life Sciences
Polyelectrolyte Threading through a Nanopore
 polymers Article Polyelectrolyte Threading through a Nanopore Pai-Yi Hsiao Department of Engineering and System Science, National Tsing Hua University, Hsinchu 313, Taiwan; pyhsiao@ess.nthu.edu.tw; Tel.:
polymers Article Polyelectrolyte Threading through a Nanopore Pai-Yi Hsiao Department of Engineering and System Science, National Tsing Hua University, Hsinchu 313, Taiwan; pyhsiao@ess.nthu.edu.tw; Tel.:
Diffusion of Water and Diatomic Oxygen in Poly(3-hexylthiophene) Melt: A Molecular Dynamics Simulation Study
 Diffusion of Water and Diatomic Oxygen in Poly(3-hexylthiophene) Melt: A Molecular Dynamics Simulation Study Julia Deitz, Yeneneh Yimer, and Mesfin Tsige Department of Polymer Science University of Akron
Diffusion of Water and Diatomic Oxygen in Poly(3-hexylthiophene) Melt: A Molecular Dynamics Simulation Study Julia Deitz, Yeneneh Yimer, and Mesfin Tsige Department of Polymer Science University of Akron
Semiflexible macromolecules in quasi-one-dimensional confinement: Discrete versus continuous bond angles
 THE JOURNAL OF CHEMICAL PHYSICS 143, 243102 (2015) Semiflexible macromolecules in quasi-one-dimensional confinement: Discrete versus continuous bond angles Aiqun Huang, 1,a) Hsiao-Ping Hsu, 2,3,b) Aniket
THE JOURNAL OF CHEMICAL PHYSICS 143, 243102 (2015) Semiflexible macromolecules in quasi-one-dimensional confinement: Discrete versus continuous bond angles Aiqun Huang, 1,a) Hsiao-Ping Hsu, 2,3,b) Aniket
Simulation of Coarse-Grained Equilibrium Polymers
 Simulation of Coarse-Grained Equilibrium Polymers J. P. Wittmer, Institut Charles Sadron, CNRS, Strasbourg, France Collaboration with: M.E. Cates (Edinburgh), P. van der Schoot (Eindhoven) A. Milchev (Sofia),
Simulation of Coarse-Grained Equilibrium Polymers J. P. Wittmer, Institut Charles Sadron, CNRS, Strasbourg, France Collaboration with: M.E. Cates (Edinburgh), P. van der Schoot (Eindhoven) A. Milchev (Sofia),
Soft Matter and Biological Physics
 Dr. Ulrich F. Keyser - ufk20 (at) cam.ac.uk Soft Matter and Biological Physics Question Sheet Michaelmas 2011 Version: November 2, 2011 Question 0: Sedimentation Initially consider identical small particles
Dr. Ulrich F. Keyser - ufk20 (at) cam.ac.uk Soft Matter and Biological Physics Question Sheet Michaelmas 2011 Version: November 2, 2011 Question 0: Sedimentation Initially consider identical small particles
arxiv: v1 [cond-mat.soft] 16 Nov 2014
![arxiv: v1 [cond-mat.soft] 16 Nov 2014 arxiv: v1 [cond-mat.soft] 16 Nov 2014](/thumbs/91/106587590.jpg) Polyelectrolytes polarization in non-uniform electric fields Farnoush Farahpour 1, Fathollah Varnik 2, and Mohammad Reza Ejtehadi 1 1 Sharif University of Technology, Department of Physics, P.O. Box 11155-9161,
Polyelectrolytes polarization in non-uniform electric fields Farnoush Farahpour 1, Fathollah Varnik 2, and Mohammad Reza Ejtehadi 1 1 Sharif University of Technology, Department of Physics, P.O. Box 11155-9161,
Transport of single molecules along the periodic parallel lattices with coupling
 THE JOURNAL OF CHEMICAL PHYSICS 124 204901 2006 Transport of single molecules along the periodic parallel lattices with coupling Evgeny B. Stukalin The James Franck Institute The University of Chicago
THE JOURNAL OF CHEMICAL PHYSICS 124 204901 2006 Transport of single molecules along the periodic parallel lattices with coupling Evgeny B. Stukalin The James Franck Institute The University of Chicago
Polymer Translocation By M. Muthukumar
 Polymer Translocation By M. Muthukumar If you are searching for a book by M. Muthukumar Polymer Translocation in pdf format, then you've come to the right website. We present full variation of this book
Polymer Translocation By M. Muthukumar If you are searching for a book by M. Muthukumar Polymer Translocation in pdf format, then you've come to the right website. We present full variation of this book
The Molecular Dynamics Method
 The Molecular Dynamics Method Thermal motion of a lipid bilayer Water permeation through channels Selective sugar transport Potential Energy (hyper)surface What is Force? Energy U(x) F = d dx U(x) Conformation
The Molecular Dynamics Method Thermal motion of a lipid bilayer Water permeation through channels Selective sugar transport Potential Energy (hyper)surface What is Force? Energy U(x) F = d dx U(x) Conformation
Anatoly B. Kolomeisky. Department of Chemistry CAN WE UNDERSTAND THE COMPLEX DYNAMICS OF MOTOR PROTEINS USING SIMPLE STOCHASTIC MODELS?
 Anatoly B. Kolomeisky Department of Chemistry CAN WE UNDERSTAND THE COMPLEX DYNAMICS OF MOTOR PROTEINS USING SIMPLE STOCHASTIC MODELS? Motor Proteins Enzymes that convert the chemical energy into mechanical
Anatoly B. Kolomeisky Department of Chemistry CAN WE UNDERSTAND THE COMPLEX DYNAMICS OF MOTOR PROTEINS USING SIMPLE STOCHASTIC MODELS? Motor Proteins Enzymes that convert the chemical energy into mechanical
Distribution of chains in polymer brushes produced by a grafting from mechanism
 SUPPLEMENTARY INFORMATION Distribution of chains in polymer brushes produced by a grafting from mechanism Andre Martinez, Jan-Michael Y. Carrillo, Andrey V. Dobrynin,, * and Douglas H. Adamson, * Department
SUPPLEMENTARY INFORMATION Distribution of chains in polymer brushes produced by a grafting from mechanism Andre Martinez, Jan-Michael Y. Carrillo, Andrey V. Dobrynin,, * and Douglas H. Adamson, * Department
Untangling the Mechanics of Entangled Biopolymers
 Untangling the Mechanics of Entangled Biopolymers Rae M. Robertson-Anderson Physics Department University of San Diego students/postdocs: Cole Chapman, PhD Tobias Falzone, PhD Stephanie Gorczyca, USD 16
Untangling the Mechanics of Entangled Biopolymers Rae M. Robertson-Anderson Physics Department University of San Diego students/postdocs: Cole Chapman, PhD Tobias Falzone, PhD Stephanie Gorczyca, USD 16
2.1 Traditional and modern applications of polymers. Soft and light materials good heat and electrical insulators
 . Polymers.1. Traditional and modern applications.. From chemistry to statistical description.3. Polymer solutions and polymer blends.4. Amorphous polymers.5. The glass transition.6. Crystalline polymers.7.
. Polymers.1. Traditional and modern applications.. From chemistry to statistical description.3. Polymer solutions and polymer blends.4. Amorphous polymers.5. The glass transition.6. Crystalline polymers.7.
Analyzing Ion channel Simulations
 Analyzing Ion channel Simulations (Neher and Sakmann, Scientific American 1992) Single channel current (Heurteaux et al, EMBO 2004) Computational Patch Clamp (Molecular Dynamics) Atoms move according to
Analyzing Ion channel Simulations (Neher and Sakmann, Scientific American 1992) Single channel current (Heurteaux et al, EMBO 2004) Computational Patch Clamp (Molecular Dynamics) Atoms move according to
(Crystal) Nucleation: The language
 Why crystallization requires supercooling (Crystal) Nucleation: The language 2r 1. Transferring N particles from liquid to crystal yields energy. Crystal nucleus Δµ: thermodynamic driving force N is proportional
Why crystallization requires supercooling (Crystal) Nucleation: The language 2r 1. Transferring N particles from liquid to crystal yields energy. Crystal nucleus Δµ: thermodynamic driving force N is proportional
Universal Repulsive Contribution to the. Solvent-Induced Interaction Between Sizable, Curved Hydrophobes: Supporting Information
 Universal Repulsive Contribution to the Solvent-Induced Interaction Between Sizable, Curved Hydrophobes: Supporting Information B. Shadrack Jabes, Dusan Bratko, and Alenka Luzar Department of Chemistry,
Universal Repulsive Contribution to the Solvent-Induced Interaction Between Sizable, Curved Hydrophobes: Supporting Information B. Shadrack Jabes, Dusan Bratko, and Alenka Luzar Department of Chemistry,
Phys 450 Spring 2011 Solution set 6. A bimolecular reaction in which A and B combine to form the product P may be written as:
 Problem Phys 45 Spring Solution set 6 A bimolecular reaction in which A and combine to form the product P may be written as: k d A + A P k d k a where k d is a diffusion-limited, bimolecular rate constant
Problem Phys 45 Spring Solution set 6 A bimolecular reaction in which A and combine to form the product P may be written as: k d A + A P k d k a where k d is a diffusion-limited, bimolecular rate constant
Swelling and Collapse of Single Polymer Molecules and Gels.
 Swelling and Collapse of Single Polymer Molecules and Gels. Coil-Globule Transition in Single Polymer Molecules. the coil-globule transition If polymer chains are not ideal, interactions of non-neighboring
Swelling and Collapse of Single Polymer Molecules and Gels. Coil-Globule Transition in Single Polymer Molecules. the coil-globule transition If polymer chains are not ideal, interactions of non-neighboring
arxiv: v1 [cond-mat.soft] 3 Jul 2012
![arxiv: v1 [cond-mat.soft] 3 Jul 2012 arxiv: v1 [cond-mat.soft] 3 Jul 2012](/thumbs/73/68495873.jpg) Topological jamming of spontaneously knotted polyelectrolyte chains driven through a nanopore A. Rosa, M. Di Ventra 2, and C. Micheletti,3 SISSA - Scuola Internazionale Superiore di Studi Avanzati, Via
Topological jamming of spontaneously knotted polyelectrolyte chains driven through a nanopore A. Rosa, M. Di Ventra 2, and C. Micheletti,3 SISSA - Scuola Internazionale Superiore di Studi Avanzati, Via
arxiv: v1 [cond-mat.soft] 24 Feb 2014
![arxiv: v1 [cond-mat.soft] 24 Feb 2014 arxiv: v1 [cond-mat.soft] 24 Feb 2014](/thumbs/91/105443009.jpg) Biopolymer filtration in corrugated nanochannels arxiv:1402.597v1 [cond-mat.soft] 24 Feb 2014 Santtu T. T. Ollila, 1, 2, Colin Denniston, 2, Mikko Karttunen, 2,, 1, 4, and Tapio Ala-Nissila 1 COMP Centre
Biopolymer filtration in corrugated nanochannels arxiv:1402.597v1 [cond-mat.soft] 24 Feb 2014 Santtu T. T. Ollila, 1, 2, Colin Denniston, 2, Mikko Karttunen, 2,, 1, 4, and Tapio Ala-Nissila 1 COMP Centre
ICCP Project 2 - Advanced Monte Carlo Methods Choose one of the three options below
 ICCP Project 2 - Advanced Monte Carlo Methods Choose one of the three options below Introduction In statistical physics Monte Carlo methods are considered to have started in the Manhattan project (1940
ICCP Project 2 - Advanced Monte Carlo Methods Choose one of the three options below Introduction In statistical physics Monte Carlo methods are considered to have started in the Manhattan project (1940
Single molecule investigations of the phdependent interaction between nanoparticles and an a-hemolysin protein pore
 Single molecule investigations of the phdependent interaction between nanoparticles and an a-hemolysin protein pore Dr. Alina ASANDEI The Science Department of Alexandru Ioan Cuza University Iasi 2012
Single molecule investigations of the phdependent interaction between nanoparticles and an a-hemolysin protein pore Dr. Alina ASANDEI The Science Department of Alexandru Ioan Cuza University Iasi 2012
Improved Resolution of Tertiary Structure Elasticity in Muscle Protein
 Improved Resolution of Tertiary Structure Elasticity in Muscle Protein Jen Hsin and Klaus Schulten* Department of Physics and Beckman Institute, University of Illinois at Urbana-Champaign, Urbana, Illinois
Improved Resolution of Tertiary Structure Elasticity in Muscle Protein Jen Hsin and Klaus Schulten* Department of Physics and Beckman Institute, University of Illinois at Urbana-Champaign, Urbana, Illinois
JASS Modeling and visualization of molecular dynamic processes
 JASS 2009 Konstantin Shefov Modeling and visualization of molecular dynamic processes St Petersburg State University, Physics faculty, Department of Computational Physics Supervisor PhD Stepanova Margarita
JASS 2009 Konstantin Shefov Modeling and visualization of molecular dynamic processes St Petersburg State University, Physics faculty, Department of Computational Physics Supervisor PhD Stepanova Margarita
3 Biopolymers Uncorrelated chains - Freely jointed chain model
 3.4 Entropy When we talk about biopolymers, it is important to realize that the free energy of a biopolymer in thermal equilibrium is not constant. Unlike solids, biopolymers are characterized through
3.4 Entropy When we talk about biopolymers, it is important to realize that the free energy of a biopolymer in thermal equilibrium is not constant. Unlike solids, biopolymers are characterized through
V = 2ze 2 n. . a. i=1
 IITS: Statistical Physics in Biology Assignment # 3 KU Leuven 5/29/2013 Coulomb Interactions & Polymers 1. Flory Theory: The Coulomb energy of a ball of charge Q and dimension R in d spacial dimensions
IITS: Statistical Physics in Biology Assignment # 3 KU Leuven 5/29/2013 Coulomb Interactions & Polymers 1. Flory Theory: The Coulomb energy of a ball of charge Q and dimension R in d spacial dimensions
Polymer escape from a confining potential
 THE JOURAL OF CHEMICAL PHYSICS 140,054907(2014) Polymer escape from a confining potential Harri Mökkönen, 1,2,a) Timo Ikonen, 1,3 Hannes Jónsson, 1,2,4 and Tapio Ala-issila 1,4 1 Department of Applied
THE JOURAL OF CHEMICAL PHYSICS 140,054907(2014) Polymer escape from a confining potential Harri Mökkönen, 1,2,a) Timo Ikonen, 1,3 Hannes Jónsson, 1,2,4 and Tapio Ala-issila 1,4 1 Department of Applied
Françoise Brochard. Université Paris 6, Inst. Curie. March 13,
 Polymers in confined geometries Françoise Brochard Université Paris 6, Inst. Curie March 13, 2008 1 Three classes of polymers Flexible Rigid: proteins, colloids Semi-flexible: DNA Transfer of T5 genome
Polymers in confined geometries Françoise Brochard Université Paris 6, Inst. Curie March 13, 2008 1 Three classes of polymers Flexible Rigid: proteins, colloids Semi-flexible: DNA Transfer of T5 genome
CECAM Tutorial Stuttgart, Oct. 2012
 CECAM Tutorial Stuttgart, Oct. 2012 09.10.2012 ESPResSo++ CECAM Tutorial 1 / 40 ESPResSo++ ESPResSo++ is an open-source simulation package designed to perform molecular dynamics and monte carlo simulations
CECAM Tutorial Stuttgart, Oct. 2012 09.10.2012 ESPResSo++ CECAM Tutorial 1 / 40 ESPResSo++ ESPResSo++ is an open-source simulation package designed to perform molecular dynamics and monte carlo simulations
8.592J HST.452J: Statistical Physics in Biology
 Assignment # 4 8.592J HST.452J: Statistical Physics in Biology Coulomb Interactions 1. Flory Theory: The Coulomb energy of a ball of charge Q and dimension R in d spacial dimensions scales as Q 2 E c.
Assignment # 4 8.592J HST.452J: Statistical Physics in Biology Coulomb Interactions 1. Flory Theory: The Coulomb energy of a ball of charge Q and dimension R in d spacial dimensions scales as Q 2 E c.
Electrophoretic Mobility within a Confining Well
 Electrophoretic Mobility within a Confining Well Tyler N. Shendruk Martin Bertrand Gary W. Slater University of Ottawa December 5, 2013 Free-Solution Electrophoresis: free-draining polyelectrolytes Free-Draining
Electrophoretic Mobility within a Confining Well Tyler N. Shendruk Martin Bertrand Gary W. Slater University of Ottawa December 5, 2013 Free-Solution Electrophoresis: free-draining polyelectrolytes Free-Draining
ATP hydrolysis 1 1 1
 ATP hydrolysis 1 1 1 ATP hydrolysis 2 2 2 The binding zipper 1 3 3 ATP hydrolysis/synthesis is coupled to a torque Yasuda, R., et al (1998). Cell 93:1117 1124. Abrahams, et al (1994). Nature 370:621-628.
ATP hydrolysis 1 1 1 ATP hydrolysis 2 2 2 The binding zipper 1 3 3 ATP hydrolysis/synthesis is coupled to a torque Yasuda, R., et al (1998). Cell 93:1117 1124. Abrahams, et al (1994). Nature 370:621-628.
Dynamics of two coupled particles: comparison of Lennard-Jones and spring forces
 Dynamics of two coupled particles: comparison of Lennard-Jones and spring forces Renata Retkute a and James P. Gleeson a a Department of Applied Mathematics, UCC, Cork, Ireland; ABSTRACT The motion of
Dynamics of two coupled particles: comparison of Lennard-Jones and spring forces Renata Retkute a and James P. Gleeson a a Department of Applied Mathematics, UCC, Cork, Ireland; ABSTRACT The motion of
Experimental Soft Matter (M. Durand, G. Foffi)
 Master 2 PCS/PTSC 2016-2017 10/01/2017 Experimental Soft Matter (M. Durand, G. Foffi) Nota Bene Exam duration : 3H ecture notes are not allowed. Electronic devices (including cell phones) are prohibited,
Master 2 PCS/PTSC 2016-2017 10/01/2017 Experimental Soft Matter (M. Durand, G. Foffi) Nota Bene Exam duration : 3H ecture notes are not allowed. Electronic devices (including cell phones) are prohibited,
Origin of the Electrophoretic Force on DNA in Nanopores. Biological and Soft Systems - Cavendish Laboratory
 Origin of the Electrophoretic Force on DNA in Nanopores Ulrich F. Keyser Biological and Soft Systems - Cavendish Laboratory Acknowledgements Delft Cees Dekker, Nynke H. Dekker, Serge G. Lemay R. Smeets,
Origin of the Electrophoretic Force on DNA in Nanopores Ulrich F. Keyser Biological and Soft Systems - Cavendish Laboratory Acknowledgements Delft Cees Dekker, Nynke H. Dekker, Serge G. Lemay R. Smeets,
arxiv:q-bio/ v1 [q-bio.bm] 21 Aug 2005
![arxiv:q-bio/ v1 [q-bio.bm] 21 Aug 2005 arxiv:q-bio/ v1 [q-bio.bm] 21 Aug 2005](/thumbs/80/81642473.jpg) arxiv:q-bio/0508029v1 [q-bio.bm] 21 Aug 2005 Directed motion emerging from two coupled random processes: Translocation of a chain through a membrane nanopore driven by binding proteins Tobias Ambjörnsson,
arxiv:q-bio/0508029v1 [q-bio.bm] 21 Aug 2005 Directed motion emerging from two coupled random processes: Translocation of a chain through a membrane nanopore driven by binding proteins Tobias Ambjörnsson,
Dissipative Particle Dynamics: Foundation, Evolution and Applications
 Dissipative Particle Dynamics: Foundation, Evolution and Applications Lecture 4: DPD in soft matter and polymeric applications George Em Karniadakis Division of Applied Mathematics, Brown University &
Dissipative Particle Dynamics: Foundation, Evolution and Applications Lecture 4: DPD in soft matter and polymeric applications George Em Karniadakis Division of Applied Mathematics, Brown University &
Models for Single Molecule Experiments
 Models for Single Molecule Experiments Binding Unbinding Folding - Unfolding -> Kramers Bell theory 1 Polymer elasticity: - Hooke s law (?) - FJC (Freely joined chain) - WLC (Worm-like chain) 3 Molecular
Models for Single Molecule Experiments Binding Unbinding Folding - Unfolding -> Kramers Bell theory 1 Polymer elasticity: - Hooke s law (?) - FJC (Freely joined chain) - WLC (Worm-like chain) 3 Molecular
arxiv: v1 [cond-mat.soft] 13 May 2016
![arxiv: v1 [cond-mat.soft] 13 May 2016 arxiv: v1 [cond-mat.soft] 13 May 2016](/thumbs/71/65742964.jpg) Efficient dynamical correction of the transition state theory rate estimate for a flat energy barrier arxiv:1605.04158v1 [cond-mat.soft] 13 May 2016 Harri Mökkönen 1,2,3, Tapio Ala-Nissila 1,2,4, and Hannes
Efficient dynamical correction of the transition state theory rate estimate for a flat energy barrier arxiv:1605.04158v1 [cond-mat.soft] 13 May 2016 Harri Mökkönen 1,2,3, Tapio Ala-Nissila 1,2,4, and Hannes
Overview & Applications. T. Lezon Hands-on Workshop in Computational Biophysics Pittsburgh Supercomputing Center 04 June, 2015
 Overview & Applications T. Lezon Hands-on Workshop in Computational Biophysics Pittsburgh Supercomputing Center 4 June, 215 Simulations still take time Bakan et al. Bioinformatics 211. Coarse-grained Elastic
Overview & Applications T. Lezon Hands-on Workshop in Computational Biophysics Pittsburgh Supercomputing Center 4 June, 215 Simulations still take time Bakan et al. Bioinformatics 211. Coarse-grained Elastic
PHASE TRANSITIONS IN SOFT MATTER SYSTEMS
 OUTLINE: Topic D. PHASE TRANSITIONS IN SOFT MATTER SYSTEMS Definition of a phase Classification of phase transitions Thermodynamics of mixing (gases, polymers, etc.) Mean-field approaches in the spirit
OUTLINE: Topic D. PHASE TRANSITIONS IN SOFT MATTER SYSTEMS Definition of a phase Classification of phase transitions Thermodynamics of mixing (gases, polymers, etc.) Mean-field approaches in the spirit
Supplementary Information for: Controlling Cellular Uptake of Nanoparticles with ph-sensitive Polymers
 Supplementary Information for: Controlling Cellular Uptake of Nanoparticles with ph-sensitive Polymers Hong-ming Ding 1 & Yu-qiang Ma 1,2, 1 National Laboratory of Solid State Microstructures and Department
Supplementary Information for: Controlling Cellular Uptake of Nanoparticles with ph-sensitive Polymers Hong-ming Ding 1 & Yu-qiang Ma 1,2, 1 National Laboratory of Solid State Microstructures and Department
Macromolecular Crowding
 Macromolecular Crowding Keng-Hwee Chiam Mathematical and Theoretical Biology Group Goodsell (1994) Macromolecular Crowding, Oct. 15, 2003 p.1/33 Outline What: introduction, definition Why: implications
Macromolecular Crowding Keng-Hwee Chiam Mathematical and Theoretical Biology Group Goodsell (1994) Macromolecular Crowding, Oct. 15, 2003 p.1/33 Outline What: introduction, definition Why: implications
MARTINI simulation details
 S1 Appendix MARTINI simulation details MARTINI simulation initialization and equilibration In this section, we describe the initialization of simulations from Main Text section Residue-based coarsegrained
S1 Appendix MARTINI simulation details MARTINI simulation initialization and equilibration In this section, we describe the initialization of simulations from Main Text section Residue-based coarsegrained
Molecular Machines and Enzymes
 Molecular Machines and Enzymes Principles of functioning of molecular machines Enzymes and catalysis Molecular motors: kinesin 1 NB Queste diapositive sono state preparate per il corso di Biofisica tenuto
Molecular Machines and Enzymes Principles of functioning of molecular machines Enzymes and catalysis Molecular motors: kinesin 1 NB Queste diapositive sono state preparate per il corso di Biofisica tenuto
simulations Extracting coarse-grained elastic behavior of capsid proteins from molecular dynamics Outline Feb. 2009
 Extracting coarse-grained elastic behavior of capsid proteins from molecular dynamics simulations Stephen D. Hicks Christopher L. Henley Cornell University Gordon Conf. on Physical Virology Feb. 2009 Outline
Extracting coarse-grained elastic behavior of capsid proteins from molecular dynamics simulations Stephen D. Hicks Christopher L. Henley Cornell University Gordon Conf. on Physical Virology Feb. 2009 Outline
IPLS Retreat Bath, Oct 17, 2017 Novel Physics arising from phase transitions in biology
 IPLS Retreat Bath, Oct 17, 2017 Novel Physics arising from phase transitions in biology Chiu Fan Lee Department of Bioengineering, Imperial College London, UK Biology inspires new physics Biology Physics
IPLS Retreat Bath, Oct 17, 2017 Novel Physics arising from phase transitions in biology Chiu Fan Lee Department of Bioengineering, Imperial College London, UK Biology inspires new physics Biology Physics
Self-avoiding polymer trapped inside a cylindrical pore: Flory free energy and unexpected dynamics
 PHYSICAL REVIEW E 79, 6191 9 Self-avoiding polymer trapped inside a cylindrical pore: Flory free energy and unexpected dynamics Youngkyun Jung* Supercomputing Center, Korea Institute of Science and Technology
PHYSICAL REVIEW E 79, 6191 9 Self-avoiding polymer trapped inside a cylindrical pore: Flory free energy and unexpected dynamics Youngkyun Jung* Supercomputing Center, Korea Institute of Science and Technology
Mechanics of Motor Proteins and the Cytoskeleton Jonathon Howard Chapter 10 Force generation 2 nd part. Andrea and Yinyun April 4 th,2012
 Mechanics of Motor Proteins and the Cytoskeleton Jonathon Howard Chapter 10 Force generation 2 nd part Andrea and Yinyun April 4 th,2012 I. Equilibrium Force Reminder: http://www.youtube.com/watch?v=yt59kx_z6xm
Mechanics of Motor Proteins and the Cytoskeleton Jonathon Howard Chapter 10 Force generation 2 nd part Andrea and Yinyun April 4 th,2012 I. Equilibrium Force Reminder: http://www.youtube.com/watch?v=yt59kx_z6xm
Kinetics of Loop Formation in Polymer Chains
 6094 J. Phys. Chem. B 2008, 112, 6094-6106 Kinetics of Loop Formation in Polymer Chains Ngo Minh Toan,, Greg Morrison,, Changbong Hyeon, and D. Thirumalai*,,# Biophysics Program, Institute for Physical
6094 J. Phys. Chem. B 2008, 112, 6094-6106 Kinetics of Loop Formation in Polymer Chains Ngo Minh Toan,, Greg Morrison,, Changbong Hyeon, and D. Thirumalai*,,# Biophysics Program, Institute for Physical
Statistical Mechanics and Thermodynamics of Small Systems
 Statistical Mechanics and Thermodynamics of Small Systems Luca Cerino Advisors: A. Puglisi and A. Vulpiani Final Seminar of PhD course in Physics Cycle XXIX Rome, October, 26 2016 Outline of the talk 1.
Statistical Mechanics and Thermodynamics of Small Systems Luca Cerino Advisors: A. Puglisi and A. Vulpiani Final Seminar of PhD course in Physics Cycle XXIX Rome, October, 26 2016 Outline of the talk 1.
Modeling the combined effect of surface roughness and shear rate on slip flow of simple fluids
 Modeling the combined effect of surface roughness and shear rate on slip flow of simple fluids Anoosheh Niavarani and Nikolai Priezjev www.egr.msu.edu/~niavaran November 2009 A. Niavarani and N.V. Priezjev,
Modeling the combined effect of surface roughness and shear rate on slip flow of simple fluids Anoosheh Niavarani and Nikolai Priezjev www.egr.msu.edu/~niavaran November 2009 A. Niavarani and N.V. Priezjev,
Lecture 1: Introduction. Keywords: Mathematics as a language, Need of learning mathematics, Applications of mathematics in BIology
 NPTEL Syllabus Biomathematics - Video course COURSE OUTLINE Graphs and functions, Derivative of a function, Techniques of differentiation Differentiation and its application in Biology, Finding maxima,
NPTEL Syllabus Biomathematics - Video course COURSE OUTLINE Graphs and functions, Derivative of a function, Techniques of differentiation Differentiation and its application in Biology, Finding maxima,
Polymer Physics MSE 458 / CHEM 482 Spring 2018
 Polymer Physics MSE 458 / CHEM 482 Spring 2018 Instructor: Prof. A.L. Ferguson 204 MSEB (217) 300-2354 alf@illinois.edu Grader: Class: Location: 4101 MSEB Time: 2:00 3:20 pm Days: T, Th Sections: A3 (CRN-38260)
Polymer Physics MSE 458 / CHEM 482 Spring 2018 Instructor: Prof. A.L. Ferguson 204 MSEB (217) 300-2354 alf@illinois.edu Grader: Class: Location: 4101 MSEB Time: 2:00 3:20 pm Days: T, Th Sections: A3 (CRN-38260)
Supporting Information for Solid-liquid Thermal Transport and its Relationship with Wettability and the Interfacial Liquid Structure
 Supporting Information for Solid-liquid Thermal Transport and its Relationship with Wettability and the Interfacial Liquid Structure Bladimir Ramos-Alvarado, Satish Kumar, and G. P. Peterson The George
Supporting Information for Solid-liquid Thermal Transport and its Relationship with Wettability and the Interfacial Liquid Structure Bladimir Ramos-Alvarado, Satish Kumar, and G. P. Peterson The George
Polymerization and force generation
 Polymerization and force generation by Eric Cytrynbaum April 8, 2008 Equilibrium polymer in a box An equilibrium polymer is a polymer has no source of extraneous energy available to it. This does not mean
Polymerization and force generation by Eric Cytrynbaum April 8, 2008 Equilibrium polymer in a box An equilibrium polymer is a polymer has no source of extraneous energy available to it. This does not mean
Physical Biochemistry. Kwan Hee Lee, Ph.D. Handong Global University
 Physical Biochemistry Kwan Hee Lee, Ph.D. Handong Global University Week 9 CHAPTER 4 Physical Equilibria Basic Concepts Biological organisms are highly inhomogeneous. Different regions of each cell have
Physical Biochemistry Kwan Hee Lee, Ph.D. Handong Global University Week 9 CHAPTER 4 Physical Equilibria Basic Concepts Biological organisms are highly inhomogeneous. Different regions of each cell have
Structure Analysis of Bottle-Brush Polymers: Simulation and Experiment
 Jacob Klein Science 323, 47, 2009 Manfred Schmidt (Mainz) Structure Analysis of Bottle-Brush Polymers: Simulation and Experiment http://www.blueberryforest.com Hsiao-Ping Hsu Institut für Physik, Johannes
Jacob Klein Science 323, 47, 2009 Manfred Schmidt (Mainz) Structure Analysis of Bottle-Brush Polymers: Simulation and Experiment http://www.blueberryforest.com Hsiao-Ping Hsu Institut für Physik, Johannes
Modeling endothelial glycocalyx and its interaction with blood flow
 Modeling endothelial glycocalyx and its interaction with blood flow Sofia Biagi PhD under the joint supervision of DdR Chaouqi Misbah LIPhy, Grenoble Prof. Francesco Sciortino Università Sapienza, Roma
Modeling endothelial glycocalyx and its interaction with blood flow Sofia Biagi PhD under the joint supervision of DdR Chaouqi Misbah LIPhy, Grenoble Prof. Francesco Sciortino Università Sapienza, Roma
Final Morphology of Complex Materials
 120314 Final Morphology of Complex Materials 1) Proteins are the prototypical model for hierarchy. a) Give the generic chemical structure for an amino acid and a protein molecule (a tripeptide). b) Label
120314 Final Morphology of Complex Materials 1) Proteins are the prototypical model for hierarchy. a) Give the generic chemical structure for an amino acid and a protein molecule (a tripeptide). b) Label
Anomalous diffusion of a tagged monomer with absorbing boundaries
 Anomalous diffusion of a tagged monomer with absorbing boundaries Thesis submitted as part of the requirements for the degree of Master of Science (M.Sc.) in Physics at Tel Aviv University School of Physics
Anomalous diffusion of a tagged monomer with absorbing boundaries Thesis submitted as part of the requirements for the degree of Master of Science (M.Sc.) in Physics at Tel Aviv University School of Physics
Potential energy and conservation of energy
 Chapter 8 Potential energy and conservation of energy Copyright 8.1_2 Potential Energy and Work Potential energy U is energy that can be associated with the configuration (arrangement) of a system of objects
Chapter 8 Potential energy and conservation of energy Copyright 8.1_2 Potential Energy and Work Potential energy U is energy that can be associated with the configuration (arrangement) of a system of objects
Liquid-vapour oscillations of water in hydrophobic nanopores
 Alicante 2003 4th European Biophysics Congress Liquid-vapour oscillations of water in hydrophobic nanopores 7 th July 2003 Oliver Beckstein and Mark S. P. Sansom Department of Biochemistry, Laboratory
Alicante 2003 4th European Biophysics Congress Liquid-vapour oscillations of water in hydrophobic nanopores 7 th July 2003 Oliver Beckstein and Mark S. P. Sansom Department of Biochemistry, Laboratory
Introduction to Computer Simulations of Soft Matter Methodologies and Applications Boulder July, 19-20, 2012
 Introduction to Computer Simulations of Soft Matter Methodologies and Applications Boulder July, 19-20, 2012 K. Kremer Max Planck Institute for Polymer Research, Mainz Overview Simulations, general considerations
Introduction to Computer Simulations of Soft Matter Methodologies and Applications Boulder July, 19-20, 2012 K. Kremer Max Planck Institute for Polymer Research, Mainz Overview Simulations, general considerations
MAE 545: Lecture 4 (9/29) Statistical mechanics of proteins
 MAE 545: Lecture 4 (9/29) Statistical mechanics of proteins Ideal freely jointed chain N identical unstretchable links (Kuhn segments) of length a with freely rotating joints ~r 2 a ~r ~R ~r N In first
MAE 545: Lecture 4 (9/29) Statistical mechanics of proteins Ideal freely jointed chain N identical unstretchable links (Kuhn segments) of length a with freely rotating joints ~r 2 a ~r ~R ~r N In first
Supplemental Information - Glassy Dynamics in Composite Biopolymer Networks
 Electronic Supplementary Material (ESI) for Soft Matter. This journal is The Royal Society of Chemistry 2018 Supplemental Information - Glassy Dynamics in Composite Biopolymer Networks Tom Golde, 1 Constantin
Electronic Supplementary Material (ESI) for Soft Matter. This journal is The Royal Society of Chemistry 2018 Supplemental Information - Glassy Dynamics in Composite Biopolymer Networks Tom Golde, 1 Constantin
Are DNA Transcription Factor Proteins Maxwellian Demons?
 Are DNA Transcription Factor Proteins Maxwellian Demons? * physics as a design constraint on molecular evolution. Alexander Grosberg, Longhua Hu, U. Minnesota, NYU U. Minnesota Biophysical Journal 2008
Are DNA Transcription Factor Proteins Maxwellian Demons? * physics as a design constraint on molecular evolution. Alexander Grosberg, Longhua Hu, U. Minnesota, NYU U. Minnesota Biophysical Journal 2008
arxiv: v2 [cond-mat.stat-mech] 6 Jun 2010
![arxiv: v2 [cond-mat.stat-mech] 6 Jun 2010 arxiv: v2 [cond-mat.stat-mech] 6 Jun 2010](/thumbs/81/83040079.jpg) Chaos in Sandpile Models Saman Moghimi-Araghi and Ali Mollabashi Physics department, Sharif University of Technology, P.O. Box 55-96, Tehran, Iran We have investigated the weak chaos exponent to see if
Chaos in Sandpile Models Saman Moghimi-Araghi and Ali Mollabashi Physics department, Sharif University of Technology, P.O. Box 55-96, Tehran, Iran We have investigated the weak chaos exponent to see if
Polymers Dynamics by Dielectric Spectroscopy
 Polymers Dynamics by Dielectric Spectroscopy What s a polymer bulk? A condensed matter system where the structural units are macromolecules Polymers Shape of a Macromolecule in the Bulk Flory's prediction
Polymers Dynamics by Dielectric Spectroscopy What s a polymer bulk? A condensed matter system where the structural units are macromolecules Polymers Shape of a Macromolecule in the Bulk Flory's prediction
Introduction to polymer physics Lecture 1
 Introduction to polymer physics Lecture 1 Boulder Summer School July August 3, 2012 A.Grosberg New York University Lecture 1: Ideal chains Course outline: Ideal chains (Grosberg) Real chains (Rubinstein)
Introduction to polymer physics Lecture 1 Boulder Summer School July August 3, 2012 A.Grosberg New York University Lecture 1: Ideal chains Course outline: Ideal chains (Grosberg) Real chains (Rubinstein)
Non equilibrium thermodynamics: foundations, scope, and extension to the meso scale. Miguel Rubi
 Non equilibrium thermodynamics: foundations, scope, and extension to the meso scale Miguel Rubi References S.R. de Groot and P. Mazur, Non equilibrium Thermodynamics, Dover, New York, 1984 J.M. Vilar and
Non equilibrium thermodynamics: foundations, scope, and extension to the meso scale Miguel Rubi References S.R. de Groot and P. Mazur, Non equilibrium Thermodynamics, Dover, New York, 1984 J.M. Vilar and
MONTE CARLO DYNAMICS OF DIAMOND-LATTICE MULTICHAIN SYSTEMS
 241 MONTE CARLO DYNAMICS OF DIAMOND-LATTICE MULTICHAIN SYSTEMS Andrzej Kolinski,* Jeffrey Skolnick t and Robert Yaris Department of Chemistry, Washington University, St. Louis, MO 63130 ABSTRACT We present
241 MONTE CARLO DYNAMICS OF DIAMOND-LATTICE MULTICHAIN SYSTEMS Andrzej Kolinski,* Jeffrey Skolnick t and Robert Yaris Department of Chemistry, Washington University, St. Louis, MO 63130 ABSTRACT We present
Referee Report Bell et al., Asymmetric dynamics of DNA entering and exiting a strongly confining nanopore - MS NCOMMS
 Reviewers' comments: Reviewer #1 (Remarks to the Author): Referee Report Bell et al., Asymmetric dynamics of DNA entering and exiting a strongly confining nanopore - MS NCOMMS-16-30539 I have read the
Reviewers' comments: Reviewer #1 (Remarks to the Author): Referee Report Bell et al., Asymmetric dynamics of DNA entering and exiting a strongly confining nanopore - MS NCOMMS-16-30539 I have read the
arxiv: v1 [cond-mat.stat-mech] 15 Sep 2007
![arxiv: v1 [cond-mat.stat-mech] 15 Sep 2007 arxiv: v1 [cond-mat.stat-mech] 15 Sep 2007](/thumbs/94/119661638.jpg) Current in a three-dimensional periodic tube with unbiased forces Bao-quan Ai a and Liang-gang Liu b a School of Physics and Telecommunication Engineering, South China Normal University, 56 GuangZhou,
Current in a three-dimensional periodic tube with unbiased forces Bao-quan Ai a and Liang-gang Liu b a School of Physics and Telecommunication Engineering, South China Normal University, 56 GuangZhou,
hydrogels Lecture 7 Spring
 hydrogels Lecture 7 Spring 2006 2 Thermodynamics of hydrogel swelling polymerize Move to a new, larger aqueous bath V r swelling V s Lecture 7 Spring 2006 3 Thermodynamics of hydrogel swelling Competing
hydrogels Lecture 7 Spring 2006 2 Thermodynamics of hydrogel swelling polymerize Move to a new, larger aqueous bath V r swelling V s Lecture 7 Spring 2006 3 Thermodynamics of hydrogel swelling Competing
Origins of Mechanical and Rheological Properties of Polymer Nanocomposites. Venkat Ganesan
 Department of Chemical Engineering University of Texas@Austin Origins of Mechanical and Rheological Properties of Polymer Nanocomposites Venkat Ganesan $$$: NSF DMR, Welch Foundation Megha Surve, Victor
Department of Chemical Engineering University of Texas@Austin Origins of Mechanical and Rheological Properties of Polymer Nanocomposites Venkat Ganesan $$$: NSF DMR, Welch Foundation Megha Surve, Victor
Effect of nanostructures on evaporation and explosive boiling of thin liquid films: a molecular dynamics study
 Appl Phys A DOI 10.1007/s00339-011-6577-8 Effect of nanostructures on evaporation and explosive boiling of thin liquid films: a molecular dynamics study A.K.M.M. Morshed Taitan C. Paul Jamil A. Khan Received:
Appl Phys A DOI 10.1007/s00339-011-6577-8 Effect of nanostructures on evaporation and explosive boiling of thin liquid films: a molecular dynamics study A.K.M.M. Morshed Taitan C. Paul Jamil A. Khan Received:
Weak Ergodicity Breaking. Manchester 2016
 Weak Ergodicity Breaking Eli Barkai Bar-Ilan University Burov, Froemberg, Garini, Metzler PCCP 16 (44), 24128 (2014) Akimoto, Saito Manchester 2016 Outline Experiments: anomalous diffusion of single molecules
Weak Ergodicity Breaking Eli Barkai Bar-Ilan University Burov, Froemberg, Garini, Metzler PCCP 16 (44), 24128 (2014) Akimoto, Saito Manchester 2016 Outline Experiments: anomalous diffusion of single molecules
