Solutions to obligatorisk oppgave 2, STK2100
|
|
|
- Rodger Stokes
- 5 years ago
- Views:
Transcription
1 Solutions to obligatorisk oppgave 2, STK2100 Vinnie Ko May 14, 2018 Disclaimer: This document is made solely for my own personal use and can contain many errors. Oppgave 1 We load packages and read data before we start with the assignment. 1 > # STK > # Obligatorisk oppgave 2 3 > 4 > # Clean up the memory before we start. 5 > rm(list=ls(all=true)) 6 > 7 > # Load packages 8 > library(gam) 9 > 10 > # Read data. 11 > Spam = read.table(" data.txt", header = T) 12 > head(spam) 13 x1 x2 x3 x4 x5 x6 x7 x8 x9 x10 x11 x12 x13 x14 x15 x x17 x18 x19 x20 x21 x22 x23 x24 x25 x26 x27 x28 x29 x30 x31 x32 x33 x x35 x36 x37 x38 x39 x40 x41 x42 x43 x44 x45 x46 x47 x48 x49 x50 x51 x x53 x54 x55 x56 x57 y train TRUE TRUE TRUE FALSE TRUE TRUE TRUE FALSE 1
2 TRUE FALSE TRUE FALSE (a) We fit logistic regression with training data, by using all explanatory variables. 1 > # a) 2 > n.test = sum(spam[,"train"] == 0) 3 > 4 > # Fit logistic regression. 5 > glm.fit = glm(y. train, data = Spam, family = binomial, subset = Spam[,"train"]) 6 Warning message: 7 glm.fit: fitted probabilities numerically 0 or 1 occurred 8 > # Predict from the fitted logistic regression model. 9 > y.hat.test = predict(glm.fit, Spam[(Spam[,"train"] == 0),], type = "response") > > y.test = Spam[(Spam[,"train"] == 0), "y"] 11 > test.rss = var(y.test) (length(y.test) 1) 12 > 13 > # Test error rate 14 > test.error.rate = mean(y.test!= y.hat.test) 15 > test.error.rate 16 [1] > 18 > # Test MSE 19 > test.mse = mean((y.test y.hat.test)ˆ2) 20 > test.mse 21 [1] > 23 > # Test R squared 24 > test.r.squared = 1 n.test test.mse/test.rss 25 > test.r.squared 26 [1] We considered test prediction error rate, test MSE and test R 2 as possible model performance criterion. Note that since y i {0, 1} and ŷ i {0, 1}, test prediction error rate and test MSE are always the same with this data set. Throughout this assignment, we use test prediction error rate as model performance criterion. Warning message: glm.fit: fitted probabilities numerically 0 or 1 occurred This warning message happens most probably because there exists a line that perfectly separates two categories. When p = 0 or 1, we have numerical problem with log-likelihood: L = p y (1 p) 1 y l = y log(p) + (1 y) log(1 p) (b) We first transform the explanatory variables into principal components. Then, we fit logistic regression with first k principal components. We try out k = 1,, 57 and choose the optimal value of k based on the test prediction error rate. 2
3 1 > # b) 2 > # Number of variables 3 > d = ncol(spam) 2 4 > 5 > # Compute principal components 6 > X.PCA = prcomp(spam[,1:d], retx = TRUE, scale = TRUE) 7 > Spam.PCA = data.frame(x.pca$x, y = Spam[,"y"], train = Spam[,"train"]) 8 > test.error.rate.pca.glm = data.frame(n.pc = NA, err = NA) 9 > # Try out different number of principal components. 10 > for(k in 1:d) { 11 + # Fit logistic regression with i principal components glm.fit.pca = glm(y. train, data = Spam.PCA[,c(1:k,58,59)], family = binomial, subset = Spam.PCA[," train"]) 13 + # Predict from the fitted model y.hat.pca.test = predict(glm.fit.pca, Spam.PCA[Spam.PCA[,"train"] == 0,], type = "response") > # Compute test error rate test.error.rate.pca.glm[k,"err"] = mean(spam.pca[(spam.pca[,"train"] == 0),"y"]!= y.hat.pca.test) 17 + test.error.rate.pca.glm[k,"n.pc"] = k 18 + } 19 There were 50 or more warnings (use warnings() to see the first 50) 20 > test.error.rate.pca.glm = test.error.rate.pca.glm[order(test.error.rate.pca.glm[,"err"]),] 21 > rownames(test.error.rate.pca.glm) = NULL 22 > head(test.error.rate.pca.glm) 23 n.pc err > 31 > plot(x = test.error.rate.pca.glm[,"n.pc"], y = test.error.rate.pca.glm[,"err"], xlab = "number of PC", ylab = "test error rate") 32 > 33 > warnings() 34 Warning messages: 35 1: glm.fit: fitted probabilities numerically 0 or 1 occurred 36 2: glm.fit: fitted probabilities numerically 0 or 1 occurred 37 3: glm.fit: fitted probabilities numerically 0 or 1 occurred 38 4: glm.fit: fitted probabilities numerically 0 or 1 occurred 39 5: glm.fit: fitted probabilities numerically 0 or 1 occurred 40 6: glm.fit: fitted probabilities numerically 0 or 1 occurred 41 7: glm.fit: fitted probabilities numerically 0 or 1 occurred 42 8: glm.fit: fitted probabilities numerically 0 or 1 occurred 43 9: glm.fit: fitted probabilities numerically 0 or 1 occurred 44 10: glm.fit: fitted probabilities numerically 0 or 1 occurred 45 11: glm.fit: fitted probabilities numerically 0 or 1 occurred 46 12: glm.fit: fitted probabilities numerically 0 or 1 occurred 47 13: glm.fit: fitted probabilities numerically 0 or 1 occurred 48 14: glm.fit: fitted probabilities numerically 0 or 1 occurred 49 15: glm.fit: fitted probabilities numerically 0 or 1 occurred 50 16: glm.fit: fitted probabilities numerically 0 or 1 occurred 51 17: glm.fit: fitted probabilities numerically 0 or 1 occurred 52 18: glm.fit: fitted probabilities numerically 0 or 1 occurred 53 19: glm.fit: fitted probabilities numerically 0 or 1 occurred 54 20: glm.fit: fitted probabilities numerically 0 or 1 occurred 55 21: glm.fit: fitted probabilities numerically 0 or 1 occurred 56 22: glm.fit: fitted probabilities numerically 0 or 1 occurred 57 23: glm.fit: fitted probabilities numerically 0 or 1 occurred 58 24: glm.fit: fitted probabilities numerically 0 or 1 occurred 59 25: glm.fit: fitted probabilities numerically 0 or 1 occurred 3
4 60 26: glm.fit: fitted probabilities numerically 0 or 1 occurred 61 27: glm.fit: fitted probabilities numerically 0 or 1 occurred 62 28: glm.fit: fitted probabilities numerically 0 or 1 occurred 63 29: glm.fit: fitted probabilities numerically 0 or 1 occurred 64 30: glm.fit: fitted probabilities numerically 0 or 1 occurred 65 31: glm.fit: fitted probabilities numerically 0 or 1 occurred 66 32: glm.fit: fitted probabilities numerically 0 or 1 occurred 67 33: glm.fit: fitted probabilities numerically 0 or 1 occurred 68 34: glm.fit: fitted probabilities numerically 0 or 1 occurred 69 35: glm.fit: fitted probabilities numerically 0 or 1 occurred 70 36: glm.fit: fitted probabilities numerically 0 or 1 occurred 71 37: glm.fit: fitted probabilities numerically 0 or 1 occurred 72 38: glm.fit: fitted probabilities numerically 0 or 1 occurred 73 39: glm.fit: fitted probabilities numerically 0 or 1 occurred 74 40: glm.fit: fitted probabilities numerically 0 or 1 occurred 75 41: glm.fit: fitted probabilities numerically 0 or 1 occurred 76 42: glm.fit: fitted probabilities numerically 0 or 1 occurred 77 43: glm.fit: fitted probabilities numerically 0 or 1 occurred 78 44: glm.fit: fitted probabilities numerically 0 or 1 occurred 79 45: glm.fit: fitted probabilities numerically 0 or 1 occurred 80 46: glm.fit: fitted probabilities numerically 0 or 1 occurred 81 47: glm.fit: fitted probabilities numerically 0 or 1 occurred 82 48: glm.fit: fitted probabilities numerically 0 or 1 occurred 83 49: glm.fit: fitted probabilities numerically 0 or 1 occurred 84 50: glm.fit: fitted probabilities numerically 0 or 1 occurred With k = 45, we get the lowest test error rate. With the given dataset, using principal components (instead of original variables) results in better prediction performance. test error rate number of PC Figure 1: Test error rate against k. (c) We fit logistic regression and GAM by using only the first 3 explanatory variables and compare them. 4
5 1 > # c) 2 > # Logistic regression with first 3 variables. 3 > glm.fit.nv.3 = glm(y x1 + x2 + x3, subset = Spam[,"train"], data = Spam, family = binomial) 4 > y.hat.glm.nv.3 = predict(glm.fit.nv.3, Spam[(Spam[,"train"] == 0), ], type = "response") > > test.error.rate.glm.nv.3 = mean(spam[(spam[,"train"] == 0),"y"]!= y.hat.glm.nv.3) 6 > test.error.rate.glm.nv.3 7 [1] > # GAM with first 3 variables. 9 > gam.fit.nv.3 = gam(y s(x1) + s(x2) + s(x3), subset = Spam[,"train"], data = Spam, family = binomial) 10 > y.hat.gam.nv.3 = predict(gam.fit.nv.3, Spam[(Spam[,"train"] == 0),], type = "response") > > test.error.rate.gam.nv.3 = mean(spam[(spam[,"train"] == 0),"y"]!= y.hat.gam.nv.3) 12 > test.error.rate.gam.nv.3 13 [1] > 15 > # GAM Plot 16 > plot(gam.fit.nv.3, se = TRUE) 17 > 18 > # Histrogram 19 > hist(spam[,"x1"], breaks = 100) 20 > hist(spam[,"x2"], breaks = 100) 21 > hist(spam[,"x3"], breaks = 100) 22 > GAM model has lower test error rate than logistic regression model. So, the non-linear terms improve the model. > summary(gam.fit.nv.3) Call: gam(formula = y ~ s(x1) + s(x2) + s(x3), family = binomial, data = Spam, subset = Spam[, "train"]) Deviance Residuals: Min 1Q Median 3Q Max (Dispersion Parameter for binomial family taken to be 1) Null Deviance: on 1535 degrees of freedom Residual Deviance: on 1523 degrees of freedom AIC: Number of Local Scoring Iterations: 15 Anova for Parametric Effects Df Sum Sq Mean Sq F value Pr(>F) s(x1) e-05 *** s(x2) e-09 *** s(x3) e-07 *** Residuals Signif. codes: 0 *** ** 0.01 * Anova for Nonparametric Effects Npar Df Npar Chisq P(Chi) (Intercept) s(x1) e-06 *** s(x2) e-12 *** s(x3) e-14 *** 5
6 --- Signif. codes: 0 *** ** 0.01 * > s(x1) s(x2) x x2 s(x3) x3 Figure 2: The non-linear terms of GAM model. (gam package was used.) Figure 2 shows the non-linear structures of the fitted GAM model. From visual inspection, we conclude that there is non-linear relationship between response variable and explanatory variables. According to the ANOVA tests from the model summary of gam.fit.nv.3, all three splines are significant. So, we conclude that there is non-linear relationship between response variable and explanatory variables. The plot of X2 looks linear at a first glance, but if one zooms into the left region where the most data points are, one can observe wiggly curves. 6
7 Histogram of Spam[, "x1"] Histogram of Spam[, "x2"] Frequency Frequency Spam[, "x1"] Spam[, "x2"] Histogram of Spam[, "x3"] Frequency Spam[, "x3"] Figure 3: Histogram of the first 3 explanatory variables of the Spam dataset. The variable X 1, X 2, X 3 are word frequencies in and from Figure 3 we can see that most data points have relatively low values of frequency. This means that the (non-linear) patterns on the rightregion of the plots are determined by merely a few data points. (This is also visible by wide 95% CI s on the wide side of the plots.) In this case, one can consider to put restrictions on the smoothness of the right-region. 7
8 s(x1,7.79) s(x2,5.55) x x2 s(x3,4.12) x3 Figure 4: The non-linear terms of GAM model. (mgcv package was used.) (d) We repeat part (c), but this time, we use principal components instead of the original explanatory variables. (i.e. We fit logistic regression and GAM by using only the first 3 principal components and we compare them.) 1 > # d) 2 > # Logistic regression with first 3 principal components. 3 > glm.fit.npc.3 = glm(y PC1 + PC2 + PC3, subset = Spam.PCA[,"train"], data = Spam.PCA, family = binomial) 4 > y.hat.glm.npc.3 = predict(glm.fit.npc.3, Spam.PCA[(Spam.PCA[,"train"] == 0), ], type = "response") > > test.error.rate.glm.npc.3 = mean(spam.pca[(spam.pca[,"train"] == 0),"y"]!= y.hat.glm.npc.3) 6 > test.error.rate.glm.npc.3 7 [1] > # GAM with first 3 principal components. 8
9 9 > gam.fit.npc.3 = gam(y s(pc1) + s(pc2) + s(pc3), subset = Spam.PCA[,"train"], data = Spam.PCA, family = binomial) 10 > y.hat.gam.npc.3 = predict(gam.fit.npc.3, Spam.PCA[(Spam.PCA[,"train"] == 0),], type = "response") > > test.error.rate.gam.npc.3 = mean(spam.pca[(spam.pca[,"train"] == 0),"y"]!= y.hat.gam.npc.3) 12 > test.error.rate.gam.npc.3 13 [1] > 15 > # Plot the composition of principal components. 16 > plot.ts( 17 + X.PCA$rotation[,"PC1"], 18 + xlab = "Variable number", 19 + ylab = "Variable loading", 20 + main = "The composition of principal component 1", 21 + font.main = 1) 22 > plot.ts( 23 + X.PCA$rotation[,"PC2"], 24 + xlab = "Variable number", 25 + ylab = "Variable loading", 26 + main = "The composition of principal component 2", 27 + font.main = 1) 28 > plot.ts( 29 + X.PCA$rotation[,"PC3"], 30 + xlab = "Variable number", 31 + ylab = "Variable loading", 32 + main = "The composition of principal component 3", 33 + font.main = 1) 34 > 35 > # GAM plot 36 > plot(gam.fit.npc.3, se = TRUE) 37 > Adding non-linear terms decreases the test error rate in this case. Figure 5 shows the non-linear structures of the fitted GAM model. From visual inspection, we conclude that there is non-linear relationship between response variable and explanatory variables. > summary(gam.fit.npc.3) Call: gam(formula = y ~ s(pc1) + s(pc2) + s(pc3), family = binomial, data = Spam.PCA, subset = Spam.PCA[, "train"]) Deviance Residuals: Min 1Q Median 3Q Max (Dispersion Parameter for binomial family taken to be 1) Null Deviance: on 1535 degrees of freedom Residual Deviance: on 1523 degrees of freedom AIC: Number of Local Scoring Iterations: 16 Anova for Parametric Effects Df Sum Sq Mean Sq F value Pr(>F) s(pc1) < 2.2e-16 *** s(pc2) < 2.2e-16 *** s(pc3) e-11 *** Residuals
10 --- Signif. codes: 0 *** ** 0.01 * Anova for Nonparametric Effects Npar Df Npar Chisq P(Chi) (Intercept) s(pc1) *** s(pc2) e-16 *** s(pc3) e-05 *** --- Signif. codes: 0 *** ** 0.01 * > According to the ANOVA tests from the model summary of gam.fit.npc.3, all three splines are significant. So, we conclude that there is non-linear relationship between response variable and principal components. Here, we observe similar phenomenon as in question (c). The principal components have less problem that the most of the data points are located in the left side of the plot. However, the non-linearity on the right corners are still determined by only a few data points. In this case, one can consider to put restrictions on the smoothness of the right-region. 10
11 s(pc1) s(pc2) PC PC2 s(pc3) PC3 Figure 5: The non-linear terms of GAM model with 3 principal components. (gam package was used.) 11
12 s(pc1,5.93) s(pc2,3.49) PC1 PC2 s(pc3,3.06) PC3 Figure 6: The non-linear terms of GAM model with 3 principal components. (mgcv package was used.) For both logistic regression and GAM, using 3 principal components instead of original variables decreases test error rate quite a lot. A possible explanation is that PCA merges similar variables into one principal component. Many of the variables are for example word frequency of a certain word. It can happen that word frequency of some words are highly correlated to each other. Those variables will have tendency of being merged into the same PCA. This means that different principal components will try to span the variable space as large as possible. The fact that principal component 1 and 2 are highly orthogonal in Figure 7 supports this. So, with the same number of variables, principal components cover wider variable space and hence result in better model in terms of prediction performance. 12
13 The composition of principal component 1 The composition of principal component 2 Variable loading Variable loading Variable number Variable number The composition of principal component 3 Variable loading Variable number Figure 7: Composition of principal components. (e) 1 > # e) 2 > k = 20 3 > PCA.name.vec = names(spam.pca)[1:57] 4 > # Compose formula 5 > formula.obj = as.formula(paste("y", paste(paste("s(", PCA.name.vec[1:k], ")", sep=""), collapse = "+"), sep=" ")) 6 > # Fit GAM with principal components 7 > gam.fit.npc.20 = gam(formula.obj, subset = Spam.PCA[,"train"], data = Spam.PCA, family = binomial) 8 > # Predict from the fitted model 9 > y.hat.gam.npc.20 = predict(gam.fit.npc.20, Spam.PCA[(Spam.PCA[,"train"] == 0),], type = "response") > > # Compute error rate. 11 > test.error.rate.gam.npc.20 = mean(spam.pca[(spam.pca[,"train"] == 0),"y"]!= y.hat.gam.npc.20) 12 > test.error.rate.gam.npc [1]
14 The GAM model with 20 principal components result in lower test error rate than the GAM model with 3 principal components from part (d). (f) 1 # A function for part f) 2 find.best.k.for.pca.gam.func = function (K, parallel = FALSE, n.core = NULL, silent = FALSE) { 3 4 # tic: global 5 start.time.global = Sys.time() 6 7 # Read data 8 Spam = read.table(" data.txt", header = T) 9 d = ncol(spam) # Compute principal components 12 X.PCA = prcomp(spam[,1:d], retx = TRUE, scale = TRUE) 13 Spam.PCA = data.frame(x.pca$x, y = Spam[,"y"], train = Spam[,"train"]) # Make a frame 16 gam.fit.list = list() 17 test.error.rate.mat = data.frame(n.pc = NA, test.err = NA) 18 PCA.name.vec = names(spam.pca)[1:57] # Option 1: non parallelized for loop 21 if (parallel == FALSE) { 22 # Display message 23 if (silent == FALSE) { 24 cat("fitting process started with K = ", K, " (non-parallelized). ", "\n", sep = "") 25 } 26 # Try out given k values. 27 for (k in 1:K) { 28 # tic: fit.gam 29 start.time.fit.gam = Sys.time() 30 # Display message 31 if (silent == FALSE) { 32 cat("fitting GAM with k = ", k, " PC. ", sep = "") 33 } # Compose formula 36 formula.obj = as.formula(paste("y", paste(paste("s(", PCA.name.vec[1:k], ")", sep=""), collapse = "+"), sep=" ")) 37 # Fit GAM with principal components. 38 gam.fit.list[[k]] = gam(formula.obj, subset = Spam.PCA[,"train"], data = Spam.PCA, family = binomial) 39 names(gam.fit.list)[k] = paste("k.", k, sep="") 40 # Predict from the fitted model. 41 y.hat = predict(gam.fit.list[[k]], Spam.PCA[(Spam.PCA[,"train"] == 0),], type = "response") > # Compute error rate with test set. 43 test.error.rate.mat[k,"n.pc"] = k 44 test.error.rate.mat[k,"test.err"] = mean(spam.pca[(spam.pca[,"train"] == 0),"y"]!= y.hat) 45 # Clean up 46 remove(formula.obj, y.hat) # toc: fit.gam 49 end.time.fit.gam = Sys.time() 50 time.taken.fit.gam = end.time.fit.gam start.time.fit.gam 51 # Display a message. 52 if (silent == FALSE) { 14
15 53 cat("finished (", round(time.taken.fit.gam, 2), " ", units(time.taken.fit.gam), " elapsed).", "\n", sep = "") 54 } 55 } 56 } else if (parallel == TRUE) { 57 # Worker function 58 worker.func = function(i) { 59 # Compose formula. 60 formula.obj = as.formula(paste("y", paste(paste("s(", PCA.name.vec[1:i], ")", sep=""), collapse = "+"), sep=" ")) 61 # Fit GAM with principal components. 62 gam.fit = gam(formula.obj, subset = Spam.PCA[,"train"], data = Spam.PCA, family = binomial) 63 # Predict from the fitted model. 64 y.hat = predict(gam.fit, Spam.PCA[(Spam.PCA[,"train"] == 0),], type = "response") > # Compute error rate with test set. 66 test.error.rate = mean(spam.pca[(spam.pca[,"train"] == 0),"y"]!= y.hat) # Wrap up the result. 69 result = list(k = i, gam.fit = gam.fit, test.error.rate = test.error.rate) 70 return(result) 71 } # Set number of cores to be used. 74 if (is.null(n.core)) { 75 n.workers = detectcores() 76 } else { 77 n.workers = n.core 78 } # tic: cluster setup 81 start.time.cl.setup = Sys.time() 82 # Display a message. 83 if (silent == FALSE) { 84 cat("setting up ", n.workers, " clusters with ", sep = "") 85 } # Set up clusters 88 sys.type = Sys.info()[["sysname"]] 89 if (sys.type == "Windows") { 90 if (silent == FALSE) { 91 cat(" PSOCK. ", sep = "") 92 } 93 cl = makecluster(n.workers, type = "PSOCK") 94 # Load packages to all clusters 95 clustercall(cl, function() { 96 library(mgcv) 97 } 98 ) 99 # Make all variables available to all clusters 100 clusterexport(cl = cl, varlist = objects(), envir = environment()) 101 } else { 102 if (silent == FALSE) { 103 cat(" FORK. ", sep = "") 104 } 105 cl = makecluster(n.workers, type = "FORK") 106 } # toc: cluster set up 109 end.time.cl.setup = Sys.time() 110 time.taken.cl.setup = end.time.cl.setup start.time.cl.setup 111 # Display a message. 15
16 112 if (silent == FALSE) { 113 cat("finished (", round(time.taken.cl.setup, 2), " ", units(time.taken.cl.setup), " elapsed).", "\n", sep = "") 114 } # tic: gam.fit 117 start.time.gam.fit = Sys.time() 118 # Display a message. 119 if (silent == FALSE) { 120 cat("fitting GAM with K = ", K, " (parallelized with ", n.workers, " cores). ", sep = "") 121 } # Perform parallel calculation (parlapply) 124 clusters.result = parlapply(cl, 1:K, worker.func) 125 # Shut down cluster 126 stopcluster(cl) # toc: gam.fit 129 end.time.gam.fit = Sys.time() 130 time.taken.gam.fit = end.time.gam.fit start.time.gam.fit 131 # Display a message. 132 if (silent == FALSE) { 133 cat("finished (", round(time.taken.gam.fit, 2), " ", units(time.taken.gam.fit), " elapsed).", "\n", sep = "") 134 } # Rearrange the result. 137 for (k in 1:K) { 138 gam.fit.list[[k]] = clusters.result[[k]]$gam.fit 139 names(gam.fit.list)[k] = paste("k.", k, sep="") 140 test.error.rate.mat[k,] = c(clusters.result[[k]]$k, clusters.result[[k]]$test.error.rate) 141 } 142 } # Order the result matrix 145 test.error.rate.mat = test.error.rate.mat[order(test.error.rate.mat[,"test.err"]),] 146 rownames(test.error.rate.mat) = NULL # toc: global 149 end.time.global = Sys.time() 150 time.taken.global = end.time.global start.time.global # Wrap up the result. 153 result = list(k = K, gam.fit.list = gam.fit.list, test.error.rate.mat = test.error.rate.mat, time.taken = time. taken.global) 154 return(result) 155 } We try GAM models with k principal components with k {1,, 57}. 1 > source("find.best.k.for.pca.gam.func.r") 2 > 3 > GAM.PCA.result.list = find.best.k.for.pca.gam.func(k = 57, parallel = TRUE) 4 Setting up 8 clusters with FORK. Finished (0.34 secs elapsed). 5 Fitting GAM with K = 57 (parallelized with 8 cores). Finished (2.14 mins elapsed). 6 > 7 > head(gam.pca.result.list$test.error.rate.mat) 8 n.pc test.err
17 > 16 > plot(x = GAM.PCA.result.list$test.error.rate.mat[,"n.PC"], y = GAM.PCA.result.list$test.error.rate.mat[," test.err"], xlab = "k", ylab = "test error rate") 17 > points(gam.pca.result.list$test.error.rate.mat[1,"n.pc"],gam.pca.result.list$test.error.rate.mat[1,"test. err"], pch = 19, col = "red") 18 > The optimal value is k = 50. So, when we use 50 principal components for GAM, we obtain the best test error rate performance. test error rate k Figure 8: Test error rate against k. (g) So, in this assignment, we learned: - We can model binomial response variable with logit link function. - Adding non linear terms can improve the prediction performance model. (We used GAM with splines.) - When we have many variables that measure similar things, condensing them with principal component analysis can improve the model. Possible weaknesses: - GAM can be too wiggly in the region where there are only a few data points. One can consider to put extra restrictions on those regions. - We only tried one training-test set split. If we assign training and test data again, the results can change. One can try different splits of training and data set or can utilize K-fold cross validation. - The explanatory variables are highly skewed. One can consider to log-transform these. 17
Generalized Additive Models
 Generalized Additive Models The Model The GLM is: g( µ) = ß 0 + ß 1 x 1 + ß 2 x 2 +... + ß k x k The generalization to the GAM is: g(µ) = ß 0 + f 1 (x 1 ) + f 2 (x 2 ) +... + f k (x k ) where the functions
Generalized Additive Models The Model The GLM is: g( µ) = ß 0 + ß 1 x 1 + ß 2 x 2 +... + ß k x k The generalization to the GAM is: g(µ) = ß 0 + f 1 (x 1 ) + f 2 (x 2 ) +... + f k (x k ) where the functions
Logistic Regression 21/05
 Logistic Regression 21/05 Recall that we are trying to solve a classification problem in which features x i can be continuous or discrete (coded as 0/1) and the response y is discrete (0/1). Logistic regression
Logistic Regression 21/05 Recall that we are trying to solve a classification problem in which features x i can be continuous or discrete (coded as 0/1) and the response y is discrete (0/1). Logistic regression
Exercise 5.4 Solution
 Exercise 5.4 Solution Niels Richard Hansen University of Copenhagen May 7, 2010 1 5.4(a) > leukemia
Exercise 5.4 Solution Niels Richard Hansen University of Copenhagen May 7, 2010 1 5.4(a) > leukemia
Statistical Prediction
 Statistical Prediction P.R. Hahn Fall 2017 1 Some terminology The goal is to use data to find a pattern that we can exploit. y: response/outcome/dependent/left-hand-side x: predictor/covariate/feature/independent
Statistical Prediction P.R. Hahn Fall 2017 1 Some terminology The goal is to use data to find a pattern that we can exploit. y: response/outcome/dependent/left-hand-side x: predictor/covariate/feature/independent
A Handbook of Statistical Analyses Using R. Brian S. Everitt and Torsten Hothorn
 A Handbook of Statistical Analyses Using R Brian S. Everitt and Torsten Hothorn CHAPTER 6 Logistic Regression and Generalised Linear Models: Blood Screening, Women s Role in Society, and Colonic Polyps
A Handbook of Statistical Analyses Using R Brian S. Everitt and Torsten Hothorn CHAPTER 6 Logistic Regression and Generalised Linear Models: Blood Screening, Women s Role in Society, and Colonic Polyps
Logistic Regression. 0.1 Frogs Dataset
 Logistic Regression We move now to the classification problem from the regression problem and study the technique ot logistic regression. The setting for the classification problem is the same as that
Logistic Regression We move now to the classification problem from the regression problem and study the technique ot logistic regression. The setting for the classification problem is the same as that
Introduction to Statistics and R
 Introduction to Statistics and R Mayo-Illinois Computational Genomics Workshop (2018) Ruoqing Zhu, Ph.D. Department of Statistics, UIUC rqzhu@illinois.edu June 18, 2018 Abstract This document is a supplimentary
Introduction to Statistics and R Mayo-Illinois Computational Genomics Workshop (2018) Ruoqing Zhu, Ph.D. Department of Statistics, UIUC rqzhu@illinois.edu June 18, 2018 Abstract This document is a supplimentary
Linear Regression Models P8111
 Linear Regression Models P8111 Lecture 25 Jeff Goldsmith April 26, 2016 1 of 37 Today s Lecture Logistic regression / GLMs Model framework Interpretation Estimation 2 of 37 Linear regression Course started
Linear Regression Models P8111 Lecture 25 Jeff Goldsmith April 26, 2016 1 of 37 Today s Lecture Logistic regression / GLMs Model framework Interpretation Estimation 2 of 37 Linear regression Course started
cor(dataset$measurement1, dataset$measurement2, method= pearson ) cor.test(datavector1, datavector2, method= pearson )
 Tutorial 7: Correlation and Regression Correlation Used to test whether two variables are linearly associated. A correlation coefficient (r) indicates the strength and direction of the association. A correlation
Tutorial 7: Correlation and Regression Correlation Used to test whether two variables are linearly associated. A correlation coefficient (r) indicates the strength and direction of the association. A correlation
Nature vs. nurture? Lecture 18 - Regression: Inference, Outliers, and Intervals. Regression Output. Conditions for inference.
 Understanding regression output from software Nature vs. nurture? Lecture 18 - Regression: Inference, Outliers, and Intervals In 1966 Cyril Burt published a paper called The genetic determination of differences
Understanding regression output from software Nature vs. nurture? Lecture 18 - Regression: Inference, Outliers, and Intervals In 1966 Cyril Burt published a paper called The genetic determination of differences
How to deal with non-linear count data? Macro-invertebrates in wetlands
 How to deal with non-linear count data? Macro-invertebrates in wetlands In this session we l recognize the advantages of making an effort to better identify the proper error distribution of data and choose
How to deal with non-linear count data? Macro-invertebrates in wetlands In this session we l recognize the advantages of making an effort to better identify the proper error distribution of data and choose
Generalized linear models for binary data. A better graphical exploratory data analysis. The simple linear logistic regression model
 Stat 3302 (Spring 2017) Peter F. Craigmile Simple linear logistic regression (part 1) [Dobson and Barnett, 2008, Sections 7.1 7.3] Generalized linear models for binary data Beetles dose-response example
Stat 3302 (Spring 2017) Peter F. Craigmile Simple linear logistic regression (part 1) [Dobson and Barnett, 2008, Sections 7.1 7.3] Generalized linear models for binary data Beetles dose-response example
R code and output of examples in text. Contents. De Jong and Heller GLMs for Insurance Data R code and output. 1 Poisson regression 2
 R code and output of examples in text Contents 1 Poisson regression 2 2 Negative binomial regression 5 3 Quasi likelihood regression 6 4 Logistic regression 6 5 Ordinal regression 10 6 Nominal regression
R code and output of examples in text Contents 1 Poisson regression 2 2 Negative binomial regression 5 3 Quasi likelihood regression 6 4 Logistic regression 6 5 Ordinal regression 10 6 Nominal regression
Generalized Linear Models in R
 Generalized Linear Models in R NO ORDER Kenneth K. Lopiano, Garvesh Raskutti, Dan Yang last modified 28 4 2013 1 Outline 1. Background and preliminaries 2. Data manipulation and exercises 3. Data structures
Generalized Linear Models in R NO ORDER Kenneth K. Lopiano, Garvesh Raskutti, Dan Yang last modified 28 4 2013 1 Outline 1. Background and preliminaries 2. Data manipulation and exercises 3. Data structures
Variance Decomposition and Goodness of Fit
 Variance Decomposition and Goodness of Fit 1. Example: Monthly Earnings and Years of Education In this tutorial, we will focus on an example that explores the relationship between total monthly earnings
Variance Decomposition and Goodness of Fit 1. Example: Monthly Earnings and Years of Education In this tutorial, we will focus on an example that explores the relationship between total monthly earnings
Logistic Regressions. Stat 430
 Logistic Regressions Stat 430 Final Project Final Project is, again, team based You will decide on a project - only constraint is: you are supposed to use techniques for a solution that are related to
Logistic Regressions Stat 430 Final Project Final Project is, again, team based You will decide on a project - only constraint is: you are supposed to use techniques for a solution that are related to
Classification. Chapter Introduction. 6.2 The Bayes classifier
 Chapter 6 Classification 6.1 Introduction Often encountered in applications is the situation where the response variable Y takes values in a finite set of labels. For example, the response Y could encode
Chapter 6 Classification 6.1 Introduction Often encountered in applications is the situation where the response variable Y takes values in a finite set of labels. For example, the response Y could encode
A Generalized Linear Model for Binomial Response Data. Copyright c 2017 Dan Nettleton (Iowa State University) Statistics / 46
 A Generalized Linear Model for Binomial Response Data Copyright c 2017 Dan Nettleton (Iowa State University) Statistics 510 1 / 46 Now suppose that instead of a Bernoulli response, we have a binomial response
A Generalized Linear Model for Binomial Response Data Copyright c 2017 Dan Nettleton (Iowa State University) Statistics 510 1 / 46 Now suppose that instead of a Bernoulli response, we have a binomial response
Generalized Additive Models (GAMs)
 Generalized Additive Models (GAMs) Israel Borokini Advanced Analysis Methods in Natural Resources and Environmental Science (NRES 746) October 3, 2016 Outline Quick refresher on linear regression Generalized
Generalized Additive Models (GAMs) Israel Borokini Advanced Analysis Methods in Natural Resources and Environmental Science (NRES 746) October 3, 2016 Outline Quick refresher on linear regression Generalized
Stat 4510/7510 Homework 7
 Stat 4510/7510 Due: 1/10. Stat 4510/7510 Homework 7 1. Instructions: Please list your name and student number clearly. In order to receive credit for a problem, your solution must show sufficient details
Stat 4510/7510 Due: 1/10. Stat 4510/7510 Homework 7 1. Instructions: Please list your name and student number clearly. In order to receive credit for a problem, your solution must show sufficient details
R Output for Linear Models using functions lm(), gls() & glm()
 LM 04 lm(), gls() &glm() 1 R Output for Linear Models using functions lm(), gls() & glm() Different kinds of output related to linear models can be obtained in R using function lm() {stats} in the base
LM 04 lm(), gls() &glm() 1 R Output for Linear Models using functions lm(), gls() & glm() Different kinds of output related to linear models can be obtained in R using function lm() {stats} in the base
Stat/F&W Ecol/Hort 572 Review Points Ané, Spring 2010
 1 Linear models Y = Xβ + ɛ with ɛ N (0, σ 2 e) or Y N (Xβ, σ 2 e) where the model matrix X contains the information on predictors and β includes all coefficients (intercept, slope(s) etc.). 1. Number of
1 Linear models Y = Xβ + ɛ with ɛ N (0, σ 2 e) or Y N (Xβ, σ 2 e) where the model matrix X contains the information on predictors and β includes all coefficients (intercept, slope(s) etc.). 1. Number of
Stat 5303 (Oehlert): Randomized Complete Blocks 1
 Stat 5303 (Oehlert): Randomized Complete Blocks 1 > library(stat5303libs);library(cfcdae);library(lme4) > immer Loc Var Y1 Y2 1 UF M 81.0 80.7 2 UF S 105.4 82.3 3 UF V 119.7 80.4 4 UF T 109.7 87.2 5 UF
Stat 5303 (Oehlert): Randomized Complete Blocks 1 > library(stat5303libs);library(cfcdae);library(lme4) > immer Loc Var Y1 Y2 1 UF M 81.0 80.7 2 UF S 105.4 82.3 3 UF V 119.7 80.4 4 UF T 109.7 87.2 5 UF
Week 7 Multiple factors. Ch , Some miscellaneous parts
 Week 7 Multiple factors Ch. 18-19, Some miscellaneous parts Multiple Factors Most experiments will involve multiple factors, some of which will be nuisance variables Dealing with these factors requires
Week 7 Multiple factors Ch. 18-19, Some miscellaneous parts Multiple Factors Most experiments will involve multiple factors, some of which will be nuisance variables Dealing with these factors requires
Hands on cusp package tutorial
 Hands on cusp package tutorial Raoul P. P. P. Grasman July 29, 2015 1 Introduction The cusp package provides routines for fitting a cusp catastrophe model as suggested by (Cobb, 1978). The full documentation
Hands on cusp package tutorial Raoul P. P. P. Grasman July 29, 2015 1 Introduction The cusp package provides routines for fitting a cusp catastrophe model as suggested by (Cobb, 1978). The full documentation
A Handbook of Statistical Analyses Using R 2nd Edition. Brian S. Everitt and Torsten Hothorn
 A Handbook of Statistical Analyses Using R 2nd Edition Brian S. Everitt and Torsten Hothorn CHAPTER 7 Logistic Regression and Generalised Linear Models: Blood Screening, Women s Role in Society, Colonic
A Handbook of Statistical Analyses Using R 2nd Edition Brian S. Everitt and Torsten Hothorn CHAPTER 7 Logistic Regression and Generalised Linear Models: Blood Screening, Women s Role in Society, Colonic
Business Statistics. Lecture 10: Course Review
 Business Statistics Lecture 10: Course Review 1 Descriptive Statistics for Continuous Data Numerical Summaries Location: mean, median Spread or variability: variance, standard deviation, range, percentiles,
Business Statistics Lecture 10: Course Review 1 Descriptive Statistics for Continuous Data Numerical Summaries Location: mean, median Spread or variability: variance, standard deviation, range, percentiles,
Tento projekt je spolufinancován Evropským sociálním fondem a Státním rozpočtem ČR InoBio CZ.1.07/2.2.00/
 Tento projekt je spolufinancován Evropským sociálním fondem a Státním rozpočtem ČR InoBio CZ.1.07/2.2.00/28.0018 Statistical Analysis in Ecology using R Linear Models/GLM Ing. Daniel Volařík, Ph.D. 13.
Tento projekt je spolufinancován Evropským sociálním fondem a Státním rozpočtem ČR InoBio CZ.1.07/2.2.00/28.0018 Statistical Analysis in Ecology using R Linear Models/GLM Ing. Daniel Volařík, Ph.D. 13.
Exam details. Final Review Session. Things to Review
 Exam details Final Review Session Short answer, similar to book problems Formulae and tables will be given You CAN use a calculator Date and Time: Dec. 7, 006, 1-1:30 pm Location: Osborne Centre, Unit
Exam details Final Review Session Short answer, similar to book problems Formulae and tables will be given You CAN use a calculator Date and Time: Dec. 7, 006, 1-1:30 pm Location: Osborne Centre, Unit
Chapter 16: Understanding Relationships Numerical Data
 Chapter 16: Understanding Relationships Numerical Data These notes reflect material from our text, Statistics, Learning from Data, First Edition, by Roxy Peck, published by CENGAGE Learning, 2015. Linear
Chapter 16: Understanding Relationships Numerical Data These notes reflect material from our text, Statistics, Learning from Data, First Edition, by Roxy Peck, published by CENGAGE Learning, 2015. Linear
Using R in 200D Luke Sonnet
 Using R in 200D Luke Sonnet Contents Working with data frames 1 Working with variables........................................... 1 Analyzing data............................................... 3 Random
Using R in 200D Luke Sonnet Contents Working with data frames 1 Working with variables........................................... 1 Analyzing data............................................... 3 Random
Logistic Regression - problem 6.14
 Logistic Regression - problem 6.14 Let x 1, x 2,, x m be given values of an input variable x and let Y 1,, Y m be independent binomial random variables whose distributions depend on the corresponding values
Logistic Regression - problem 6.14 Let x 1, x 2,, x m be given values of an input variable x and let Y 1,, Y m be independent binomial random variables whose distributions depend on the corresponding values
R Hints for Chapter 10
 R Hints for Chapter 10 The multiple logistic regression model assumes that the success probability p for a binomial random variable depends on independent variables or design variables x 1, x 2,, x k.
R Hints for Chapter 10 The multiple logistic regression model assumes that the success probability p for a binomial random variable depends on independent variables or design variables x 1, x 2,, x k.
Interactions in Logistic Regression
 Interactions in Logistic Regression > # UCBAdmissions is a 3-D table: Gender by Dept by Admit > # Same data in another format: > # One col for Yes counts, another for No counts. > Berkeley = read.table("http://www.utstat.toronto.edu/~brunner/312f12/
Interactions in Logistic Regression > # UCBAdmissions is a 3-D table: Gender by Dept by Admit > # Same data in another format: > # One col for Yes counts, another for No counts. > Berkeley = read.table("http://www.utstat.toronto.edu/~brunner/312f12/
7/28/15. Review Homework. Overview. Lecture 6: Logistic Regression Analysis
 Lecture 6: Logistic Regression Analysis Christopher S. Hollenbeak, PhD Jane R. Schubart, PhD The Outcomes Research Toolbox Review Homework 2 Overview Logistic regression model conceptually Logistic regression
Lecture 6: Logistic Regression Analysis Christopher S. Hollenbeak, PhD Jane R. Schubart, PhD The Outcomes Research Toolbox Review Homework 2 Overview Logistic regression model conceptually Logistic regression
mgcv: GAMs in R Simon Wood Mathematical Sciences, University of Bath, U.K.
 mgcv: GAMs in R Simon Wood Mathematical Sciences, University of Bath, U.K. mgcv, gamm4 mgcv is a package supplied with R for generalized additive modelling, including generalized additive mixed models.
mgcv: GAMs in R Simon Wood Mathematical Sciences, University of Bath, U.K. mgcv, gamm4 mgcv is a package supplied with R for generalized additive modelling, including generalized additive mixed models.
BOOtstrapping the Generalized Linear Model. Link to the last RSS article here:factor Analysis with Binary items: A quick review with examples. -- Ed.
 MyUNT EagleConnect Blackboard People & Departments Maps Calendars Giving to UNT Skip to content Benchmarks ABOUT BENCHMARK ONLINE SEARCH ARCHIVE SUBSCRIBE TO BENCHMARKS ONLINE Columns, October 2014 Home»
MyUNT EagleConnect Blackboard People & Departments Maps Calendars Giving to UNT Skip to content Benchmarks ABOUT BENCHMARK ONLINE SEARCH ARCHIVE SUBSCRIBE TO BENCHMARKS ONLINE Columns, October 2014 Home»
Generalised linear models. Response variable can take a number of different formats
 Generalised linear models Response variable can take a number of different formats Structure Limitations of linear models and GLM theory GLM for count data GLM for presence \ absence data GLM for proportion
Generalised linear models Response variable can take a number of different formats Structure Limitations of linear models and GLM theory GLM for count data GLM for presence \ absence data GLM for proportion
STATS216v Introduction to Statistical Learning Stanford University, Summer Midterm Exam (Solutions) Duration: 1 hours
 Instructions: STATS216v Introduction to Statistical Learning Stanford University, Summer 2017 Remember the university honor code. Midterm Exam (Solutions) Duration: 1 hours Write your name and SUNet ID
Instructions: STATS216v Introduction to Statistical Learning Stanford University, Summer 2017 Remember the university honor code. Midterm Exam (Solutions) Duration: 1 hours Write your name and SUNet ID
Multivariate Statistics in Ecology and Quantitative Genetics Summary
 Multivariate Statistics in Ecology and Quantitative Genetics Summary Dirk Metzler & Martin Hutzenthaler http://evol.bio.lmu.de/_statgen 5. August 2011 Contents Linear Models Generalized Linear Models Mixed-effects
Multivariate Statistics in Ecology and Quantitative Genetics Summary Dirk Metzler & Martin Hutzenthaler http://evol.bio.lmu.de/_statgen 5. August 2011 Contents Linear Models Generalized Linear Models Mixed-effects
Consider fitting a model using ordinary least squares (OLS) regression:
 Example 1: Mating Success of African Elephants In this study, 41 male African elephants were followed over a period of 8 years. The age of the elephant at the beginning of the study and the number of successful
Example 1: Mating Success of African Elephants In this study, 41 male African elephants were followed over a period of 8 years. The age of the elephant at the beginning of the study and the number of successful
STAT 510 Final Exam Spring 2015
 STAT 510 Final Exam Spring 2015 Instructions: The is a closed-notes, closed-book exam No calculator or electronic device of any kind may be used Use nothing but a pen or pencil Please write your name and
STAT 510 Final Exam Spring 2015 Instructions: The is a closed-notes, closed-book exam No calculator or electronic device of any kind may be used Use nothing but a pen or pencil Please write your name and
ssh tap sas913, sas https://www.statlab.umd.edu/sasdoc/sashtml/onldoc.htm
 Kedem, STAT 430 SAS Examples: Logistic Regression ==================================== ssh abc@glue.umd.edu, tap sas913, sas https://www.statlab.umd.edu/sasdoc/sashtml/onldoc.htm a. Logistic regression.
Kedem, STAT 430 SAS Examples: Logistic Regression ==================================== ssh abc@glue.umd.edu, tap sas913, sas https://www.statlab.umd.edu/sasdoc/sashtml/onldoc.htm a. Logistic regression.
Non-Gaussian Response Variables
 Non-Gaussian Response Variables What is the Generalized Model Doing? The fixed effects are like the factors in a traditional analysis of variance or linear model The random effects are different A generalized
Non-Gaussian Response Variables What is the Generalized Model Doing? The fixed effects are like the factors in a traditional analysis of variance or linear model The random effects are different A generalized
df=degrees of freedom = n - 1
 One sample t-test test of the mean Assumptions: Independent, random samples Approximately normal distribution (from intro class: σ is unknown, need to calculate and use s (sample standard deviation)) Hypotheses:
One sample t-test test of the mean Assumptions: Independent, random samples Approximately normal distribution (from intro class: σ is unknown, need to calculate and use s (sample standard deviation)) Hypotheses:
Modeling Overdispersion
 James H. Steiger Department of Psychology and Human Development Vanderbilt University Regression Modeling, 2009 1 Introduction 2 Introduction In this lecture we discuss the problem of overdispersion in
James H. Steiger Department of Psychology and Human Development Vanderbilt University Regression Modeling, 2009 1 Introduction 2 Introduction In this lecture we discuss the problem of overdispersion in
Booklet of Code and Output for STAD29/STA 1007 Midterm Exam
 Booklet of Code and Output for STAD29/STA 1007 Midterm Exam List of Figures in this document by page: List of Figures 1 Packages................................ 2 2 Hospital infection risk data (some).................
Booklet of Code and Output for STAD29/STA 1007 Midterm Exam List of Figures in this document by page: List of Figures 1 Packages................................ 2 2 Hospital infection risk data (some).................
Leftovers. Morris. University Farm. University Farm. Morris. yield
 Leftovers SI 544 Lada Adamic 1 Trellis graphics Trebi Wisconsin No. 38 No. 457 Glabron Peatland Velvet No. 475 Manchuria No. 462 Svansota Trebi Wisconsin No. 38 No. 457 Glabron Peatland Velvet No. 475
Leftovers SI 544 Lada Adamic 1 Trellis graphics Trebi Wisconsin No. 38 No. 457 Glabron Peatland Velvet No. 475 Manchuria No. 462 Svansota Trebi Wisconsin No. 38 No. 457 Glabron Peatland Velvet No. 475
Duration of Unemployment - Analysis of Deviance Table for Nested Models
 Duration of Unemployment - Analysis of Deviance Table for Nested Models February 8, 2012 The data unemployment is included as a contingency table. The response is the duration of unemployment, gender and
Duration of Unemployment - Analysis of Deviance Table for Nested Models February 8, 2012 The data unemployment is included as a contingency table. The response is the duration of unemployment, gender and
Variance Decomposition in Regression James M. Murray, Ph.D. University of Wisconsin - La Crosse Updated: October 04, 2017
 Variance Decomposition in Regression James M. Murray, Ph.D. University of Wisconsin - La Crosse Updated: October 04, 2017 PDF file location: http://www.murraylax.org/rtutorials/regression_anovatable.pdf
Variance Decomposition in Regression James M. Murray, Ph.D. University of Wisconsin - La Crosse Updated: October 04, 2017 PDF file location: http://www.murraylax.org/rtutorials/regression_anovatable.pdf
Exam Applied Statistical Regression. Good Luck!
 Dr. M. Dettling Summer 2011 Exam Applied Statistical Regression Approved: Tables: Note: Any written material, calculator (without communication facility). Attached. All tests have to be done at the 5%-level.
Dr. M. Dettling Summer 2011 Exam Applied Statistical Regression Approved: Tables: Note: Any written material, calculator (without communication facility). Attached. All tests have to be done at the 5%-level.
Prediction problems 3: Validation and Model Checking
 Prediction problems 3: Validation and Model Checking Data Science 101 Team May 17, 2018 Outline Validation Why is it important How should we do it? Model checking Checking whether your model is a good
Prediction problems 3: Validation and Model Checking Data Science 101 Team May 17, 2018 Outline Validation Why is it important How should we do it? Model checking Checking whether your model is a good
Chapter 8 Conclusion
 1 Chapter 8 Conclusion Three questions about test scores (score) and student-teacher ratio (str): a) After controlling for differences in economic characteristics of different districts, does the effect
1 Chapter 8 Conclusion Three questions about test scores (score) and student-teacher ratio (str): a) After controlling for differences in economic characteristics of different districts, does the effect
Logistic & Tobit Regression
 Logistic & Tobit Regression Different Types of Regression Binary Regression (D) Logistic transformation + e P( y x) = 1 + e! " x! + " x " P( y x) % ln$ ' = ( + ) x # 1! P( y x) & logit of P(y x){ P(y
Logistic & Tobit Regression Different Types of Regression Binary Regression (D) Logistic transformation + e P( y x) = 1 + e! " x! + " x " P( y x) % ln$ ' = ( + ) x # 1! P( y x) & logit of P(y x){ P(y
PAPER 206 APPLIED STATISTICS
 MATHEMATICAL TRIPOS Part III Thursday, 1 June, 2017 9:00 am to 12:00 pm PAPER 206 APPLIED STATISTICS Attempt no more than FOUR questions. There are SIX questions in total. The questions carry equal weight.
MATHEMATICAL TRIPOS Part III Thursday, 1 June, 2017 9:00 am to 12:00 pm PAPER 206 APPLIED STATISTICS Attempt no more than FOUR questions. There are SIX questions in total. The questions carry equal weight.
Administration. Homework 1 on web page, due Feb 11 NSERC summer undergraduate award applications due Feb 5 Some helpful books
 STA 44/04 Jan 6, 00 / 5 Administration Homework on web page, due Feb NSERC summer undergraduate award applications due Feb 5 Some helpful books STA 44/04 Jan 6, 00... administration / 5 STA 44/04 Jan 6,
STA 44/04 Jan 6, 00 / 5 Administration Homework on web page, due Feb NSERC summer undergraduate award applications due Feb 5 Some helpful books STA 44/04 Jan 6, 00... administration / 5 STA 44/04 Jan 6,
Regression Methods for Survey Data
 Regression Methods for Survey Data Professor Ron Fricker! Naval Postgraduate School! Monterey, California! 3/26/13 Reading:! Lohr chapter 11! 1 Goals for this Lecture! Linear regression! Review of linear
Regression Methods for Survey Data Professor Ron Fricker! Naval Postgraduate School! Monterey, California! 3/26/13 Reading:! Lohr chapter 11! 1 Goals for this Lecture! Linear regression! Review of linear
Stat 401B Exam 3 Fall 2016 (Corrected Version)
 Stat 401B Exam 3 Fall 2016 (Corrected Version) I have neither given nor received unauthorized assistance on this exam. Name Signed Date Name Printed ATTENTION! Incorrect numerical answers unaccompanied
Stat 401B Exam 3 Fall 2016 (Corrected Version) I have neither given nor received unauthorized assistance on this exam. Name Signed Date Name Printed ATTENTION! Incorrect numerical answers unaccompanied
Logistic Regression. 1 Analysis of the budworm moth data 1. 2 Estimates and confidence intervals for the parameters 2
 Logistic Regression Ulrich Halekoh, Jørgen Vinslov Hansen, Søren Højsgaard Biometry Research Unit Danish Institute of Agricultural Sciences March 31, 2006 Contents 1 Analysis of the budworm moth data 1
Logistic Regression Ulrich Halekoh, Jørgen Vinslov Hansen, Søren Højsgaard Biometry Research Unit Danish Institute of Agricultural Sciences March 31, 2006 Contents 1 Analysis of the budworm moth data 1
HW1 Roshena MacPherson Feb 1, 2017
 HW1 Roshena MacPherson Feb 1, 2017 This is an R Markdown Notebook. When you execute code within the notebook, the results appear beneath the code. Question 1: In this question we will consider some real
HW1 Roshena MacPherson Feb 1, 2017 This is an R Markdown Notebook. When you execute code within the notebook, the results appear beneath the code. Question 1: In this question we will consider some real
Introduction to the Generalized Linear Model: Logistic regression and Poisson regression
 Introduction to the Generalized Linear Model: Logistic regression and Poisson regression Statistical modelling: Theory and practice Gilles Guillot gigu@dtu.dk November 4, 2013 Gilles Guillot (gigu@dtu.dk)
Introduction to the Generalized Linear Model: Logistic regression and Poisson regression Statistical modelling: Theory and practice Gilles Guillot gigu@dtu.dk November 4, 2013 Gilles Guillot (gigu@dtu.dk)
Generalized Linear Models 1
 Generalized Linear Models 1 STA 2101/442: Fall 2012 1 See last slide for copyright information. 1 / 24 Suggested Reading: Davison s Statistical models Exponential families of distributions Sec. 5.2 Chapter
Generalized Linear Models 1 STA 2101/442: Fall 2012 1 See last slide for copyright information. 1 / 24 Suggested Reading: Davison s Statistical models Exponential families of distributions Sec. 5.2 Chapter
Class Notes: Week 8. Probit versus Logit Link Functions and Count Data
 Ronald Heck Class Notes: Week 8 1 Class Notes: Week 8 Probit versus Logit Link Functions and Count Data This week we ll take up a couple of issues. The first is working with a probit link function. While
Ronald Heck Class Notes: Week 8 1 Class Notes: Week 8 Probit versus Logit Link Functions and Count Data This week we ll take up a couple of issues. The first is working with a probit link function. While
Two Hours. Mathematical formula books and statistical tables are to be provided THE UNIVERSITY OF MANCHESTER. 26 May :00 16:00
 Two Hours MATH38052 Mathematical formula books and statistical tables are to be provided THE UNIVERSITY OF MANCHESTER GENERALISED LINEAR MODELS 26 May 2016 14:00 16:00 Answer ALL TWO questions in Section
Two Hours MATH38052 Mathematical formula books and statistical tables are to be provided THE UNIVERSITY OF MANCHESTER GENERALISED LINEAR MODELS 26 May 2016 14:00 16:00 Answer ALL TWO questions in Section
UNIVERSITY OF TORONTO. Faculty of Arts and Science APRIL 2010 EXAMINATIONS STA 303 H1S / STA 1002 HS. Duration - 3 hours. Aids Allowed: Calculator
 UNIVERSITY OF TORONTO Faculty of Arts and Science APRIL 2010 EXAMINATIONS STA 303 H1S / STA 1002 HS Duration - 3 hours Aids Allowed: Calculator LAST NAME: FIRST NAME: STUDENT NUMBER: There are 27 pages
UNIVERSITY OF TORONTO Faculty of Arts and Science APRIL 2010 EXAMINATIONS STA 303 H1S / STA 1002 HS Duration - 3 hours Aids Allowed: Calculator LAST NAME: FIRST NAME: STUDENT NUMBER: There are 27 pages
Random Independent Variables
 Random Independent Variables Example: e 1, e 2, e 3 ~ Poisson(5) X 1 = e 1 + e 3 X 2 = e 2 +e 3 X 3 ~ N(10,10) ind. of e 1 e 2 e 3 Y = β 0 + β 1 X 1 + β 2 X 2 +β 3 X 3 + ε ε ~ N(0,σ 2 ) β 0 = 0 β 1 = 1
Random Independent Variables Example: e 1, e 2, e 3 ~ Poisson(5) X 1 = e 1 + e 3 X 2 = e 2 +e 3 X 3 ~ N(10,10) ind. of e 1 e 2 e 3 Y = β 0 + β 1 X 1 + β 2 X 2 +β 3 X 3 + ε ε ~ N(0,σ 2 ) β 0 = 0 β 1 = 1
Clinical Trials. Olli Saarela. September 18, Dalla Lana School of Public Health University of Toronto.
 Introduction to Dalla Lana School of Public Health University of Toronto olli.saarela@utoronto.ca September 18, 2014 38-1 : a review 38-2 Evidence Ideal: to advance the knowledge-base of clinical medicine,
Introduction to Dalla Lana School of Public Health University of Toronto olli.saarela@utoronto.ca September 18, 2014 38-1 : a review 38-2 Evidence Ideal: to advance the knowledge-base of clinical medicine,
Model checking overview. Checking & Selecting GAMs. Residual checking. Distribution checking
 Model checking overview Checking & Selecting GAMs Simon Wood Mathematical Sciences, University of Bath, U.K. Since a GAM is just a penalized GLM, residual plots should be checked exactly as for a GLM.
Model checking overview Checking & Selecting GAMs Simon Wood Mathematical Sciences, University of Bath, U.K. Since a GAM is just a penalized GLM, residual plots should be checked exactly as for a GLM.
Checking, Selecting & Predicting with GAMs. Simon Wood Mathematical Sciences, University of Bath, U.K.
 Checking, Selecting & Predicting with GAMs Simon Wood Mathematical Sciences, University of Bath, U.K. Model checking Since a GAM is just a penalized GLM, residual plots should be checked, exactly as for
Checking, Selecting & Predicting with GAMs Simon Wood Mathematical Sciences, University of Bath, U.K. Model checking Since a GAM is just a penalized GLM, residual plots should be checked, exactly as for
R in Linguistic Analysis. Wassink 2012 University of Washington Week 6
 R in Linguistic Analysis Wassink 2012 University of Washington Week 6 Overview R for phoneticians and lab phonologists Johnson 3 Reading Qs Equivalence of means (t-tests) Multiple Regression Principal
R in Linguistic Analysis Wassink 2012 University of Washington Week 6 Overview R for phoneticians and lab phonologists Johnson 3 Reading Qs Equivalence of means (t-tests) Multiple Regression Principal
How to work correctly statistically about sex ratio
 How to work correctly statistically about sex ratio Marc Girondot Version of 12th April 2014 Contents 1 Load packages 2 2 Introduction 2 3 Confidence interval of a proportion 4 3.1 Pourcentage..............................
How to work correctly statistically about sex ratio Marc Girondot Version of 12th April 2014 Contents 1 Load packages 2 2 Introduction 2 3 Confidence interval of a proportion 4 3.1 Pourcentage..............................
1 Multiple Regression
 1 Multiple Regression In this section, we extend the linear model to the case of several quantitative explanatory variables. There are many issues involved in this problem and this section serves only
1 Multiple Regression In this section, we extend the linear model to the case of several quantitative explanatory variables. There are many issues involved in this problem and this section serves only
BMI 541/699 Lecture 22
 BMI 541/699 Lecture 22 Where we are: 1. Introduction and Experimental Design 2. Exploratory Data Analysis 3. Probability 4. T-based methods for continous variables 5. Power and sample size for t-based
BMI 541/699 Lecture 22 Where we are: 1. Introduction and Experimental Design 2. Exploratory Data Analysis 3. Probability 4. T-based methods for continous variables 5. Power and sample size for t-based
Review of Multiple Regression
 Ronald H. Heck 1 Let s begin with a little review of multiple regression this week. Linear models [e.g., correlation, t-tests, analysis of variance (ANOVA), multiple regression, path analysis, multivariate
Ronald H. Heck 1 Let s begin with a little review of multiple regression this week. Linear models [e.g., correlation, t-tests, analysis of variance (ANOVA), multiple regression, path analysis, multivariate
Reaction Days
 Stat April 03 Week Fitting Individual Trajectories # Straight-line, constant rate of change fit > sdat = subset(sleepstudy, Subject == "37") > sdat Reaction Days Subject > lm.sdat = lm(reaction ~ Days)
Stat April 03 Week Fitting Individual Trajectories # Straight-line, constant rate of change fit > sdat = subset(sleepstudy, Subject == "37") > sdat Reaction Days Subject > lm.sdat = lm(reaction ~ Days)
ESP 178 Applied Research Methods. 2/23: Quantitative Analysis
 ESP 178 Applied Research Methods 2/23: Quantitative Analysis Data Preparation Data coding create codebook that defines each variable, its response scale, how it was coded Data entry for mail surveys and
ESP 178 Applied Research Methods 2/23: Quantitative Analysis Data Preparation Data coding create codebook that defines each variable, its response scale, how it was coded Data entry for mail surveys and
1. Logistic Regression, One Predictor 2. Inference: Estimating the Parameters 3. Multiple Logistic Regression 4. AIC and BIC in Logistic Regression
 Logistic Regression 1. Logistic Regression, One Predictor 2. Inference: Estimating the Parameters 3. Multiple Logistic Regression 4. AIC and BIC in Logistic Regression 5. Target Marketing: Tabloid Data
Logistic Regression 1. Logistic Regression, One Predictor 2. Inference: Estimating the Parameters 3. Multiple Logistic Regression 4. AIC and BIC in Logistic Regression 5. Target Marketing: Tabloid Data
Various Issues in Fitting Contingency Tables
 Various Issues in Fitting Contingency Tables Statistics 149 Spring 2006 Copyright 2006 by Mark E. Irwin Complete Tables with Zero Entries In contingency tables, it is possible to have zero entries in a
Various Issues in Fitting Contingency Tables Statistics 149 Spring 2006 Copyright 2006 by Mark E. Irwin Complete Tables with Zero Entries In contingency tables, it is possible to have zero entries in a
STK 2100 Oblig 1. Zhou Siyu. February 15, 2017
 STK 200 Oblig Zhou Siyu February 5, 207 Question a) Make a scatter box plot for the data set. Answer:Here is the code I used to plot the scatter box in R. library ( MASS ) 2 pairs ( Boston ) Figure : Scatter
STK 200 Oblig Zhou Siyu February 5, 207 Question a) Make a scatter box plot for the data set. Answer:Here is the code I used to plot the scatter box in R. library ( MASS ) 2 pairs ( Boston ) Figure : Scatter
22s:152 Applied Linear Regression. Take random samples from each of m populations.
 22s:152 Applied Linear Regression Chapter 8: ANOVA NOTE: We will meet in the lab on Monday October 10. One-way ANOVA Focuses on testing for differences among group means. Take random samples from each
22s:152 Applied Linear Regression Chapter 8: ANOVA NOTE: We will meet in the lab on Monday October 10. One-way ANOVA Focuses on testing for differences among group means. Take random samples from each
Explanatory variables are: weight, width of shell, color (medium light, medium, medium dark, dark), and condition of spine.
 Horseshoe crab example: There are 173 female crabs for which we wish to model the presence or absence of male satellites dependant upon characteristics of the female horseshoe crabs. 1 satellite present
Horseshoe crab example: There are 173 female crabs for which we wish to model the presence or absence of male satellites dependant upon characteristics of the female horseshoe crabs. 1 satellite present
EPSY 905: Fundamentals of Multivariate Modeling Online Lecture #7
 Introduction to Generalized Univariate Models: Models for Binary Outcomes EPSY 905: Fundamentals of Multivariate Modeling Online Lecture #7 EPSY 905: Intro to Generalized In This Lecture A short review
Introduction to Generalized Univariate Models: Models for Binary Outcomes EPSY 905: Fundamentals of Multivariate Modeling Online Lecture #7 EPSY 905: Intro to Generalized In This Lecture A short review
Statistical Methods III Statistics 212. Problem Set 2 - Answer Key
 Statistical Methods III Statistics 212 Problem Set 2 - Answer Key 1. (Analysis to be turned in and discussed on Tuesday, April 24th) The data for this problem are taken from long-term followup of 1423
Statistical Methods III Statistics 212 Problem Set 2 - Answer Key 1. (Analysis to be turned in and discussed on Tuesday, April 24th) The data for this problem are taken from long-term followup of 1423
STA102 Class Notes Chapter Logistic Regression
 STA0 Class Notes Chapter 0 0. Logistic Regression We continue to study the relationship between a response variable and one or more eplanatory variables. For SLR and MLR (Chapters 8 and 9), our response
STA0 Class Notes Chapter 0 0. Logistic Regression We continue to study the relationship between a response variable and one or more eplanatory variables. For SLR and MLR (Chapters 8 and 9), our response
Sample solutions. Stat 8051 Homework 8
 Sample solutions Stat 8051 Homework 8 Problem 1: Faraway Exercise 3.1 A plot of the time series reveals kind of a fluctuating pattern: Trying to fit poisson regression models yields a quadratic model if
Sample solutions Stat 8051 Homework 8 Problem 1: Faraway Exercise 3.1 A plot of the time series reveals kind of a fluctuating pattern: Trying to fit poisson regression models yields a quadratic model if
22s:152 Applied Linear Regression. There are a couple commonly used models for a one-way ANOVA with m groups. Chapter 8: ANOVA
 22s:152 Applied Linear Regression Chapter 8: ANOVA NOTE: We will meet in the lab on Monday October 10. One-way ANOVA Focuses on testing for differences among group means. Take random samples from each
22s:152 Applied Linear Regression Chapter 8: ANOVA NOTE: We will meet in the lab on Monday October 10. One-way ANOVA Focuses on testing for differences among group means. Take random samples from each
Neural networks (not in book)
 (not in book) Another approach to classification is neural networks. were developed in the 1980s as a way to model how learning occurs in the brain. There was therefore wide interest in neural networks
(not in book) Another approach to classification is neural networks. were developed in the 1980s as a way to model how learning occurs in the brain. There was therefore wide interest in neural networks
Generalized Linear Models
 Generalized Linear Models Lecture 7. Models with binary response II GLM (Spring, 2018) Lecture 7 1 / 13 Existence of estimates Lemma (Claudia Czado, München, 2004) The log-likelihood ln L(β) in logistic
Generalized Linear Models Lecture 7. Models with binary response II GLM (Spring, 2018) Lecture 7 1 / 13 Existence of estimates Lemma (Claudia Czado, München, 2004) The log-likelihood ln L(β) in logistic
Glossary. The ISI glossary of statistical terms provides definitions in a number of different languages:
 Glossary The ISI glossary of statistical terms provides definitions in a number of different languages: http://isi.cbs.nl/glossary/index.htm Adjusted r 2 Adjusted R squared measures the proportion of the
Glossary The ISI glossary of statistical terms provides definitions in a number of different languages: http://isi.cbs.nl/glossary/index.htm Adjusted r 2 Adjusted R squared measures the proportion of the
Package clogitboost. R topics documented: December 21, 2015
 Package December 21, 2015 Type Package Title Boosting Conditional Logit Model Version 1.1 Date 2015-12-09 Author Haolun Shi and Guosheng Yin Maintainer A set of functions to fit a boosting conditional
Package December 21, 2015 Type Package Title Boosting Conditional Logit Model Version 1.1 Date 2015-12-09 Author Haolun Shi and Guosheng Yin Maintainer A set of functions to fit a boosting conditional
β j = coefficient of x j in the model; β = ( β1, β2,
 Regression Modeling of Survival Time Data Why regression models? Groups similar except for the treatment under study use the nonparametric methods discussed earlier. Groups differ in variables (covariates)
Regression Modeling of Survival Time Data Why regression models? Groups similar except for the treatment under study use the nonparametric methods discussed earlier. Groups differ in variables (covariates)
An Introduction to Path Analysis
 An Introduction to Path Analysis PRE 905: Multivariate Analysis Lecture 10: April 15, 2014 PRE 905: Lecture 10 Path Analysis Today s Lecture Path analysis starting with multivariate regression then arriving
An Introduction to Path Analysis PRE 905: Multivariate Analysis Lecture 10: April 15, 2014 PRE 905: Lecture 10 Path Analysis Today s Lecture Path analysis starting with multivariate regression then arriving
Density Temp vs Ratio. temp
 Temp Ratio Density 0.00 0.02 0.04 0.06 0.08 0.10 0.12 Density 0.0 0.2 0.4 0.6 0.8 1.0 1. (a) 170 175 180 185 temp 1.0 1.5 2.0 2.5 3.0 ratio The histogram shows that the temperature measures have two peaks,
Temp Ratio Density 0.00 0.02 0.04 0.06 0.08 0.10 0.12 Density 0.0 0.2 0.4 0.6 0.8 1.0 1. (a) 170 175 180 185 temp 1.0 1.5 2.0 2.5 3.0 ratio The histogram shows that the temperature measures have two peaks,
Lecture 18: Simple Linear Regression
 Lecture 18: Simple Linear Regression BIOS 553 Department of Biostatistics University of Michigan Fall 2004 The Correlation Coefficient: r The correlation coefficient (r) is a number that measures the strength
Lecture 18: Simple Linear Regression BIOS 553 Department of Biostatistics University of Michigan Fall 2004 The Correlation Coefficient: r The correlation coefficient (r) is a number that measures the strength
Handout 4: Simple Linear Regression
 Handout 4: Simple Linear Regression By: Brandon Berman The following problem comes from Kokoska s Introductory Statistics: A Problem-Solving Approach. The data can be read in to R using the following code:
Handout 4: Simple Linear Regression By: Brandon Berman The following problem comes from Kokoska s Introductory Statistics: A Problem-Solving Approach. The data can be read in to R using the following code:
ANOVA (Analysis of Variance) output RLS 11/20/2016
 ANOVA (Analysis of Variance) output RLS 11/20/2016 1. Analysis of Variance (ANOVA) The goal of ANOVA is to see if the variation in the data can explain enough to see if there are differences in the means.
ANOVA (Analysis of Variance) output RLS 11/20/2016 1. Analysis of Variance (ANOVA) The goal of ANOVA is to see if the variation in the data can explain enough to see if there are differences in the means.
Regression and the 2-Sample t
 Regression and the 2-Sample t James H. Steiger Department of Psychology and Human Development Vanderbilt University James H. Steiger (Vanderbilt University) Regression and the 2-Sample t 1 / 44 Regression
Regression and the 2-Sample t James H. Steiger Department of Psychology and Human Development Vanderbilt University James H. Steiger (Vanderbilt University) Regression and the 2-Sample t 1 / 44 Regression
Linear Regression. Data Model. β, σ 2. Process Model. ,V β. ,s 2. s 1. Parameter Model
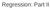 Regression: Part II Linear Regression y~n X, 2 X Y Data Model β, σ 2 Process Model Β 0,V β s 1,s 2 Parameter Model Assumptions of Linear Model Homoskedasticity No error in X variables Error in Y variables
Regression: Part II Linear Regression y~n X, 2 X Y Data Model β, σ 2 Process Model Β 0,V β s 1,s 2 Parameter Model Assumptions of Linear Model Homoskedasticity No error in X variables Error in Y variables
The prediction of house price
 000 001 002 003 004 005 006 007 008 009 010 011 012 013 014 015 016 017 018 019 020 021 022 023 024 025 026 027 028 029 030 031 032 033 034 035 036 037 038 039 040 041 042 043 044 045 046 047 048 049 050
000 001 002 003 004 005 006 007 008 009 010 011 012 013 014 015 016 017 018 019 020 021 022 023 024 025 026 027 028 029 030 031 032 033 034 035 036 037 038 039 040 041 042 043 044 045 046 047 048 049 050
Classification: Logistic Regression and Naive Bayes Book Chapter 4. Carlos M. Carvalho The University of Texas McCombs School of Business
 Classification: Logistic Regression and Naive Bayes Book Chapter 4. Carlos M. Carvalho The University of Texas McCombs School of Business 1 1. Classification 2. Logistic Regression, One Predictor 3. Inference:
Classification: Logistic Regression and Naive Bayes Book Chapter 4. Carlos M. Carvalho The University of Texas McCombs School of Business 1 1. Classification 2. Logistic Regression, One Predictor 3. Inference:
