ID14-EH3. Adam Round
|
|
|
- Edgar Gervais Alexander
- 5 years ago
- Views:
Transcription
1 ID14-EH3 Adam Round
2 Contents What can be obtained from Bio-SAXS Measurable parameters Modelling strategies How to collect data at Bio-SAXS Procedure Data collection tests Data Verification and quality control Modelling and analysis Future Improvements for Bio-SAXS
3 Crystals versus solution B A Protein molecule X Lattice = Crystal Fourier Transformation 3-dim information about: Length of the vector AB Orientation of vector AB 3-dim protein structure with atomic resolution B X = A 2Θ Protein Solution Random molecule Radial integration Scattering Pair distance function distribution function I(s) [a.u.] Fourier Transformation s [Å] Maximal dimension of the particle Mean value Radius of gyration p(r) r [nm]
4 Based on existing high resolution protein structures from crystallography or NMR the corresponding SAXS profile can be calculated. I(s) = Crystals Structure Validation A( s) 2 Ω = A a ( s) ρ A s s ( s) + δρ A This allows one to verify high resolution structures b b ( s) especially in the case of multimeric protein and larger protein complexes. 2 Ω Example: The muscle protein titin was found in different conformations in the crystal. CRYSOL (X-rays): Svergun et al. (1995). J. Appl. Cryst. 28, 768 CRYSON (neutrons): Svergun et al. (1998) P.N.A.S. USA, 95, 2267
5 Refinement of Rigid Domains Rigid Body Refinement: Moving protein sub-parts (called domains) as rigid bodies to fit the scattering data Example: Structural changes upon ligand binding PX-structure with ligand SAXS Shape obtained by GASBOR - unliganded state Ligand Rigid Body refinement using MASSHA
6 Adding Missing Linkers Remodeling of proteins from high resolution fragments/constructs (. J Program BUNCH (M. Petoukhov; Biophysical A Linkers?!? B Domains A+B Domains B+C ( ABC ) Entire Protein C High resolution protein fragments from X-ray crystallography + sequence data ( TrEMBL/Swissprot ) SAXS Data of constructs AB BC ABC Model for the entire protein including not resolved linker components
7 Ab-initio Modelling A sphere with diameter D max is filled by densely packed beads of radius r 0 << D max. A configuration vector X indicates whether the j-th atom belongs to the particle or to the solvent. Vector of model parameters: Position ( j ) = x( j ) = ( assignments (phase 1 0 if particle if solvent Solvent Particle The number of model parameters M (D max / r 0 ) is too large for conventional minimization methods. D max 2r 0 A Monte-Carlo type search starting from a random X can be employed to find a configuration that yields the calculated scattering curve fitting the experimental data Chacón, P. et al. (1998) Biophys. J. 74, Svergun, D.I. (1999) Biophys. J. 76,
8 Ab-initio Modelling DAMMIN modelling penalties Using simulated annealing, finds a compact dummy atoms configuration X that fits the scattering data by minimizing 2 f ( X ) = χ [ Iexp( s), I( s, X )] + αp( X ) where χ is the discrepancy between the experimental and calculated curves, P(X) is the penalty to ensure compactness and connectivity, α>0 its weight. compact loose disconnected
9 Ab-initio Can it be Trusted? Ab initio bead models compared to high resolution X-ray structures
10 Summary What can we learn from solution SAXS Complete high resolution structure known: validation of crystal structure in solution under physiological conditions Ligand binding reactions: Internal structure of multicomponent particles and large macromolecular complexes High resolution structure of domains/subunits known: quaternary structure using docking/rigid body refinement Incomplete high resolution structure known: probable configuration of missing portions Nothing known: ab initio low resolution structure
11 Solution scattering data collection ( I(s Log sample buffer sample ( subtracted ) buffer s, nm-1 X-ray Beam X-ray Scattering Sample X-ray detector
12 ID14-3 Experimental Hutch Setup
13 ID14-3 Sample Exposure System
14 ID14-3 Beamline Control Interface
15 ID14-3 Beamline Control Interface
16 Primary Data Processing PRIMUS
17 Quality Control Tests Multiple time frames used to check for radiation damage! ( I(s Log sample same sample again RADIATION DAMAGE! s, nm -1 Thanks to A. Kikhney for this slide
18 Merging Data Low and High Concentration ( I(s Log c = 2 mg/ml c = 5 mg/ml s, nm -1 Thanks to A. Kikhney for this slide
19 Merging Data Low and High Concentration ( I(s Log c = 2 mg/ml c = 5 mg/ml s, nm -1 Thanks to A. Kikhney for this slide
20 Merging Data Low and High Concentration ( I(s Log c = 2 mg/ml c = 5 mg/ml s, nm -1 Thanks to A. Kikhney for this slide
21 Rg and I(0) Radius of Gyration and Zero Angle Intensity
22 Rg and I(0) Radius of Gyration and Zero Angle Intensity
23 Using calibration data BSA as a standard... Comparison with unknown proteins The difference at low angles < 1 nm -1 is due to the different molecular weights (MW). The scattering at low angles is proportional the product of c protein MW. With known protein concentration the molecular weight of the sample can be estimated using the parameters estimated from the Guinierplot.
24 Using calibration data BSA as a standard... Comparison with unknown proteins I( s) I lni 0 e Rg 3 2 () s ln I s 0 2 s 2 Rg 3 2 Radius of Gyration Forward scattering I 0 MW protein 66kDa = BSA standard 3.07 nm 185 units Sample protein 5.58 nm 867 units MW protein = 307kDa Thanks to M. Roessle for this slide
25 Porod Analysis in Primus gives excluded volume
26 Porod Analysis in Primus gives excluded volume
27 Distance Distribution P(r) Function Calculated by GNOM give the Dmax and the input file required for Ab-initio modeling
28 Distance Distribution P(r) Function Calculated by GNOM give the Dmax and the input file required for Ab-initio modeling Indirect Fourier Transform!
29 Distance Distribution P(r) Function Calculated by GNOM give the Dmax and the input file required for Ab-initio modeling
30 Distance Distribution P(r) Function Calculated by GNOM give the Dmax and the input file required for Ab-initio modeling
31 Distance Distribution P(r) Function Calculated by GNOM give the Dmax and the input file required for Ab-initio modeling
32 Summary what should be done while at the beamline Data collection Radial averaging 1D Normalization Background subtraction Checks for effects of Radiation Concentration Log plot ( MM Guinier plot (R g, ( volume ) Porod plot P(r) plot ( P(r ( I(s Log s Lo-res 3D model r ( I(s Ln I(s)*s 4 s s 2
33 Automation! Making life easier! The future of ID14-3
34 ID14-3 BioSAXS Data collection to be Automated
35 Automated Data Processing Pipeline Data AUTOSUB AUTOPilatus Automatic data transformation of 2D raw data into 1D scattering curves Automatic samplebuffer subtraction with automatic estimation of radius of ( AUTORG ) gyration AUTOGNOM Automatic estimation of distance distribution function Results: Sample checks for radiation damage and concentration effects Sample Size: Rg, Dmax, Volume and Mw Low Resolution particle shape DAMMIF Automaticly start the abinitio shape determination program and fitting with simple geometrical bodies
36 Closing Remarks SAXS is a powerful tool for structural biology Structure validation Structure of multi domain proteins With addition of missing linkers if needed Ab-initio modeling Optimized data collection at ID14-3 Standard experimental setup greatly improves efficiency Easy access using rolling application procedure The future is Automation Data collection through collaboration between ESRF and the EMBL Grenoble and Hamburg outstations Data Processing based on the Hamburg SAXS data processing pipeline
37 Acknowledgments EMBL-Hamburg SAXS Group D. Svergun M. Roessle M. Petoukhov D. Franke A. Kikhney P. Konarev EMBL-Grenoble S. Cusack A. Mc Carthy F. Cipriani Synchrotron Instrumentation Team ESRF P. Pernot S. McSweeney G. Leonard D. Nurizzo D. Spruce R. Fernandez P. Theveneau J. Surr T. Giraud D. Davison D. Flot S. Larson
Data reduction and processing tutorial
 Data reduction and processing tutorial Petr V. Konarev European Molecular Biology Laboratory, Hamburg Outstation BioSAXS group EMBL BioSAXS beamline X33, 2012 Optics Vacuum cell Completely redesigned 2005-2012
Data reduction and processing tutorial Petr V. Konarev European Molecular Biology Laboratory, Hamburg Outstation BioSAXS group EMBL BioSAXS beamline X33, 2012 Optics Vacuum cell Completely redesigned 2005-2012
SAXS/SANS data processing and overall parameters
 EMBO Global Exchange Lecture Course 30 November 2012 Hyderabad India SAXS/SANS data processing and overall parameters Petr V. Konarev European Molecular Biology Laboratory, Hamburg Outstation BioSAXS group
EMBO Global Exchange Lecture Course 30 November 2012 Hyderabad India SAXS/SANS data processing and overall parameters Petr V. Konarev European Molecular Biology Laboratory, Hamburg Outstation BioSAXS group
Small-Angle Scattering Atomic Structure Based Modeling
 Small-Angle Scattering Atomic Structure Based Modeling Alejandro Panjkovich EMBL Hamburg 07.12.2017 A. Panjkovich (EMBL) BioSAS atomic modeling 07.12.2017 1 / 49 From the forest to the particle accelerator
Small-Angle Scattering Atomic Structure Based Modeling Alejandro Panjkovich EMBL Hamburg 07.12.2017 A. Panjkovich (EMBL) BioSAS atomic modeling 07.12.2017 1 / 49 From the forest to the particle accelerator
Small-Angle Scattering from Biomolecular Solutions
 T H E U N I V E R S I T Y of T E X A S S C H O O L O F H E A L T H I N F O R M A T I O N S C I E N C E S A T H O U S T O N Small-Angle Scattering from Biomolecular Solutions For students of HI 6001-125
T H E U N I V E R S I T Y of T E X A S S C H O O L O F H E A L T H I N F O R M A T I O N S C I E N C E S A T H O U S T O N Small-Angle Scattering from Biomolecular Solutions For students of HI 6001-125
Modelling against small angle scattering data. Al Kikhney EMBL Hamburg, Germany
 Modelling against small angle scattering data Al Kikhney EMBL Hamburg, Germany Validation of atomic models CRYSOL Rigid body modelling SASREF BUNCH CORAL Oligomeric mixtures OLIGOMER Flexible systems EOM
Modelling against small angle scattering data Al Kikhney EMBL Hamburg, Germany Validation of atomic models CRYSOL Rigid body modelling SASREF BUNCH CORAL Oligomeric mixtures OLIGOMER Flexible systems EOM
BM29 biosaxs data processing tutorial
 HERCULES 2014 BM29 biosaxs data processing tutorial Page 2 OUTLINE Sample Changer Primary Data Processing Model Validation HPLC-SAXS Primary Data Processing Model Validation Ab Initio Model Software in
HERCULES 2014 BM29 biosaxs data processing tutorial Page 2 OUTLINE Sample Changer Primary Data Processing Model Validation HPLC-SAXS Primary Data Processing Model Validation Ab Initio Model Software in
Introduction to Biological Small Angle Scattering
 Introduction to Biological Small Angle Scattering Tom Grant, Ph.D. Staff Scientist BioXFEL Science and Technology Center Hauptman-Woodward Institute Buffalo, New York, USA tgrant@hwi.buffalo.edu SAXS Literature
Introduction to Biological Small Angle Scattering Tom Grant, Ph.D. Staff Scientist BioXFEL Science and Technology Center Hauptman-Woodward Institute Buffalo, New York, USA tgrant@hwi.buffalo.edu SAXS Literature
How to judge data quality
 SSRL Workshop: Small-Angle X-ray Scattering and Diffraction Studies, March 28-30, 2016 How to judge data quality Tsutomu Matsui SSRL Lab / Dept. of Chemistry Stanford University Subject of this session
SSRL Workshop: Small-Angle X-ray Scattering and Diffraction Studies, March 28-30, 2016 How to judge data quality Tsutomu Matsui SSRL Lab / Dept. of Chemistry Stanford University Subject of this session
Biological Small Angle X-ray Scattering (SAXS) Dec 2, 2013
 Biological Small Angle X-ray Scattering (SAXS) Dec 2, 2013 Structural Biology Shape Dynamic Light Scattering Electron Microscopy Small Angle X-ray Scattering Cryo-Electron Microscopy Wide Angle X- ray
Biological Small Angle X-ray Scattering (SAXS) Dec 2, 2013 Structural Biology Shape Dynamic Light Scattering Electron Microscopy Small Angle X-ray Scattering Cryo-Electron Microscopy Wide Angle X- ray
Characterizing Biological Macromolecules by SAXS Detlef Beckers, Jörg Bolze, Bram Schierbeek, PANalytical B.V., Almelo, The Netherlands
 Characterizing Biological Macromolecules by SAXS Detlef Beckers, Jörg Bolze, Bram Schierbeek, PANalytical B.V., Almelo, The Netherlands This document was presented at PPXRD - Pharmaceutical Powder X-ray
Characterizing Biological Macromolecules by SAXS Detlef Beckers, Jörg Bolze, Bram Schierbeek, PANalytical B.V., Almelo, The Netherlands This document was presented at PPXRD - Pharmaceutical Powder X-ray
Small-Angle X-ray Scattering (SAXS) SPring-8/JASRI Naoto Yagi
 Small-Angle X-ray Scattering (SAXS) SPring-8/JASRI Naoto Yagi 1 Wikipedia Small-angle X-ray scattering (SAXS) is a small-angle scattering (SAS) technique where the elastic scattering of X-rays (wavelength
Small-Angle X-ray Scattering (SAXS) SPring-8/JASRI Naoto Yagi 1 Wikipedia Small-angle X-ray scattering (SAXS) is a small-angle scattering (SAS) technique where the elastic scattering of X-rays (wavelength
Supplemental Information. Structural and Mechanistic Paradigm. of Leptin Receptor Activation Revealed
 Structure, Volume 22 Supplemental Information Structural and Mechanistic Paradigm of Leptin Receptor Activation Revealed by Complexes with Wild-Type and Antagonist Leptins Kedar Moharana, Lennart Zabeau,
Structure, Volume 22 Supplemental Information Structural and Mechanistic Paradigm of Leptin Receptor Activation Revealed by Complexes with Wild-Type and Antagonist Leptins Kedar Moharana, Lennart Zabeau,
Robust, high-throughput solution structural analyses by small angle X-ray scattering (SAXS)
 nature methods Robust, high-throughput solution structural analyses by small angle X-ray scattering (SAXS) Greg L Hura, Angeli L Menon, Michal Hammel, Robert P Rambo, Farris L Poole II, Susan E Tsutakawa,
nature methods Robust, high-throughput solution structural analyses by small angle X-ray scattering (SAXS) Greg L Hura, Angeli L Menon, Michal Hammel, Robert P Rambo, Farris L Poole II, Susan E Tsutakawa,
Introduction to biological small angle scattering
 Introduction to biological small angle scattering Frank Gabel (IBS/ILL) EMBO Practical Course (May 6th 013) F. Gabel (May 6th 013) EMBO Practical Course Length-scales and tools in structural biology small
Introduction to biological small angle scattering Frank Gabel (IBS/ILL) EMBO Practical Course (May 6th 013) F. Gabel (May 6th 013) EMBO Practical Course Length-scales and tools in structural biology small
SI Text S1 Solution Scattering Data Collection and Analysis. SI references
 SI Text S1 Solution Scattering Data Collection and Analysis. The X-ray photon energy was set to 8 kev. The PILATUS hybrid pixel array detector (RIGAKU) was positioned at a distance of 606 mm from the sample.
SI Text S1 Solution Scattering Data Collection and Analysis. The X-ray photon energy was set to 8 kev. The PILATUS hybrid pixel array detector (RIGAKU) was positioned at a distance of 606 mm from the sample.
SAXS Basics for BioSAXS. Robert P. Rambo Diamond Light Source B21
 SAXS Basics for BioSAXS Robert P. Rambo Diamond Light Source B21 Scattering and Size SAXS 0.11 nm (1.1 Å) smaller than C-C bond 6000x range 690 nm (6900 Å) 3.5x larger than E.coli 10x larger than a ribosome
SAXS Basics for BioSAXS Robert P. Rambo Diamond Light Source B21 Scattering and Size SAXS 0.11 nm (1.1 Å) smaller than C-C bond 6000x range 690 nm (6900 Å) 3.5x larger than E.coli 10x larger than a ribosome
Supplementary figure 1. Comparison of unbound ogm-csf and ogm-csf as captured in the GIF:GM-CSF complex. Alignment of two copies of unbound ovine
 Supplementary figure 1. Comparison of unbound and as captured in the GIF:GM-CSF complex. Alignment of two copies of unbound ovine GM-CSF (slate) with bound GM-CSF in the GIF:GM-CSF complex (GIF: green,
Supplementary figure 1. Comparison of unbound and as captured in the GIF:GM-CSF complex. Alignment of two copies of unbound ovine GM-CSF (slate) with bound GM-CSF in the GIF:GM-CSF complex (GIF: green,
Small Angle X-Ray Solution Scattering of Biological Macromolecules
 Small Angle X-Ray Solution Scattering of Biological Macromolecules Emre Brookes UltraScan Workshop 15 June 2014 Overview Experimental method Sample preparation Experimental data analysis Experimental method
Small Angle X-Ray Solution Scattering of Biological Macromolecules Emre Brookes UltraScan Workshop 15 June 2014 Overview Experimental method Sample preparation Experimental data analysis Experimental method
P. Vachette IBBMC (CNRS-Université Paris-Sud), Orsay, France
 Scattering of X-rays P. Vachette IBBMC (CNRS-Université Paris-Sud), Orsay, France SAXS measurement Sample SAXS measuring cell SAXS measurement Scattering experiment X-ray beam?? Detector SAXS measurement
Scattering of X-rays P. Vachette IBBMC (CNRS-Université Paris-Sud), Orsay, France SAXS measurement Sample SAXS measuring cell SAXS measurement Scattering experiment X-ray beam?? Detector SAXS measurement
SOCS3 binds specific receptor JAK complexes to control cytokine signaling by direct kinase inhibition SUPPLEMENTARY INFORMATION
 SOCS3 binds specific receptor JAK complexes to control cytokine signaling by direct kinase inhibition Nadia J. Kershaw 1,2, James M. Murphy 1,2, Nicholas P.D. Liau 1,2, Leila N. Varghese 1,2, Artem Laktyushin
SOCS3 binds specific receptor JAK complexes to control cytokine signaling by direct kinase inhibition Nadia J. Kershaw 1,2, James M. Murphy 1,2, Nicholas P.D. Liau 1,2, Leila N. Varghese 1,2, Artem Laktyushin
BCM Protein crystallography - I. Crystal symmetry X-ray diffraction Protein crystallization X-ray sources SAXS
 BCM 6200 - Protein crystallography - I Crystal symmetry X-ray diffraction Protein crystallization X-ray sources SAXS What SAXS can do Small-angle X-ray scattering (SAXS) yields information on biological
BCM 6200 - Protein crystallography - I Crystal symmetry X-ray diffraction Protein crystallization X-ray sources SAXS What SAXS can do Small-angle X-ray scattering (SAXS) yields information on biological
Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1.
 Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1. PDZK1 constru cts Amino acids MW [kda] KD [μm] PEPT2-CT- FITC KD [μm] NHE3-CT- FITC KD [μm] PDZK1-CT-
Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1. PDZK1 constru cts Amino acids MW [kda] KD [μm] PEPT2-CT- FITC KD [μm] NHE3-CT- FITC KD [μm] PDZK1-CT-
Development of Novel Small- Angle X-ray Scattering Data Analysis Methods for Study of Flexible Proteins. Michael Kachala EMBL-Hamburg, Germany
 Development of Novel Small- Angle X-ray Scattering Data Analysis Methods for Study of Flexible Proteins Michael Kachala EMBL-Hamburg, Germany 60 mkl >1 mg/ml Monocromatic X-ray beam Sample Mono- or polydisperse
Development of Novel Small- Angle X-ray Scattering Data Analysis Methods for Study of Flexible Proteins Michael Kachala EMBL-Hamburg, Germany 60 mkl >1 mg/ml Monocromatic X-ray beam Sample Mono- or polydisperse
SAS Data Analysis Colloids. Dr Karen Edler
 SAS Data Analysis Colloids Dr Karen Edler Size Range Comparisons 10 1 0.1 0.01 0.001 proteins viruses nanoparticles micelles polymers Q = 2π/d (Å -1 ) bacteria molecules nanotubes precipitates grain boundaries
SAS Data Analysis Colloids Dr Karen Edler Size Range Comparisons 10 1 0.1 0.01 0.001 proteins viruses nanoparticles micelles polymers Q = 2π/d (Å -1 ) bacteria molecules nanotubes precipitates grain boundaries
Small-angle scattering studies of biological macromolecules in solution
 INSTITUTE OF PHYSICS PUBLISHING Rep. Prog. Phys. 66 (2003) 1735 1782 REPORTS ON PROGRESS IN PHYSICS PII: S0034-4885(03)12688-7 Small-angle scattering studies of biological macromolecules in solution Dmitri
INSTITUTE OF PHYSICS PUBLISHING Rep. Prog. Phys. 66 (2003) 1735 1782 REPORTS ON PROGRESS IN PHYSICS PII: S0034-4885(03)12688-7 Small-angle scattering studies of biological macromolecules in solution Dmitri
Crystals, X-rays and Proteins
 Crystals, X-rays and Proteins Comprehensive Protein Crystallography Dennis Sherwood MA (Hons), MPhil, PhD Jon Cooper BA (Hons), PhD OXFORD UNIVERSITY PRESS Contents List of symbols xiv PART I FUNDAMENTALS
Crystals, X-rays and Proteins Comprehensive Protein Crystallography Dennis Sherwood MA (Hons), MPhil, PhD Jon Cooper BA (Hons), PhD OXFORD UNIVERSITY PRESS Contents List of symbols xiv PART I FUNDAMENTALS
Overview - Macromolecular Crystallography
 Overview - Macromolecular Crystallography 1. Overexpression and crystallization 2. Crystal characterization and data collection 3. The diffraction experiment 4. Phase problem 1. MIR (Multiple Isomorphous
Overview - Macromolecular Crystallography 1. Overexpression and crystallization 2. Crystal characterization and data collection 3. The diffraction experiment 4. Phase problem 1. MIR (Multiple Isomorphous
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature11539 Supplementary Figure 1 Schematic representation of plant (A) and mammalian (B) P 2B -ATPase domain organization. Actuator (A-), nucleotide binding (N-),
SUPPLEMENTARY INFORMATION doi:10.1038/nature11539 Supplementary Figure 1 Schematic representation of plant (A) and mammalian (B) P 2B -ATPase domain organization. Actuator (A-), nucleotide binding (N-),
Acta Cryst. (2017). D73, , doi: /s
 Supporting information Volume 73 (2017) Supporting information for article: 2017 publication guidelines for structural modelling of small-angle scattering data from biomolecules in solution: an update
Supporting information Volume 73 (2017) Supporting information for article: 2017 publication guidelines for structural modelling of small-angle scattering data from biomolecules in solution: an update
High Brilliance SAXS on Synchrotrons
 High Brilliance SAXS on Synchrotrons Manfred Roessle Luebeck University of Applied Science Lab of X-ray Engineering X-ray generation Synchrotron Sources Shanghai Synchrotron Radiation Facility China Deutsches
High Brilliance SAXS on Synchrotrons Manfred Roessle Luebeck University of Applied Science Lab of X-ray Engineering X-ray generation Synchrotron Sources Shanghai Synchrotron Radiation Facility China Deutsches
Studying conformational dynamics and molecular recognition using integrated structural biology in solution Michael Sattler
 Studying conformational dynamics and molecular recognition using integrated structural biology in solution Michael Sattler http://www.nmr.ch.tum.de http://www.helmholtz-muenchen.de/stb/ Outline Dynamics
Studying conformational dynamics and molecular recognition using integrated structural biology in solution Michael Sattler http://www.nmr.ch.tum.de http://www.helmholtz-muenchen.de/stb/ Outline Dynamics
SAXS and SANS facilities and experimental practice. Clement Blanchet
 SAXS and SANS facilities and experimental practice Clement Blanchet SAS experiment Detector X-ray or neutron Beam Sample 2 s Buffer X-rays Roengten, 1895 Electromagnetic wave The electromagnetic spectrum
SAXS and SANS facilities and experimental practice Clement Blanchet SAS experiment Detector X-ray or neutron Beam Sample 2 s Buffer X-rays Roengten, 1895 Electromagnetic wave The electromagnetic spectrum
Scattering of X-rays. EMBO Practical Course on Solution Scattering from Biological Macromolecules Hamburg 4 11 September 2001
 Scattering of X-rays Books on SAS - " The origins" (no recent edition) : Small Angle Scattering of X-rays A. Guinier and A. Fournet, (1955), in English, ed. Wiley, NY - Small-Angle X-ray Scattering O.
Scattering of X-rays Books on SAS - " The origins" (no recent edition) : Small Angle Scattering of X-rays A. Guinier and A. Fournet, (1955), in English, ed. Wiley, NY - Small-Angle X-ray Scattering O.
The basics of structural biology. And Why we use synchrotron sources Sean McSweeney ESRF Structural Biology Group
 The basics of structural biology And Why we use synchrotron sources Sean McSweeney ESRF Structural Biology Group The rise and rise of structural biology. 2 The aim of the game 3 What information does structure
The basics of structural biology And Why we use synchrotron sources Sean McSweeney ESRF Structural Biology Group The rise and rise of structural biology. 2 The aim of the game 3 What information does structure
Light scattering Small and large particles
 Scattering by macromolecules E B Incident light Scattered Light particle Oscillating E field from light makes electronic cloud oscillate surrounding the particle Intensity: I E Accelerating charges means
Scattering by macromolecules E B Incident light Scattered Light particle Oscillating E field from light makes electronic cloud oscillate surrounding the particle Intensity: I E Accelerating charges means
X-ray Crystallography
 2009/11/25 [ 1 ] X-ray Crystallography Andrew Torda, wintersemester 2009 / 2010 X-ray numerically most important more than 4/5 structures Goal a set of x, y, z coordinates different properties to NMR History
2009/11/25 [ 1 ] X-ray Crystallography Andrew Torda, wintersemester 2009 / 2010 X-ray numerically most important more than 4/5 structures Goal a set of x, y, z coordinates different properties to NMR History
Protein Crystallography
 Protein Crystallography Part II Tim Grüne Dept. of Structural Chemistry Prof. G. Sheldrick University of Göttingen http://shelx.uni-ac.gwdg.de tg@shelx.uni-ac.gwdg.de Overview The Reciprocal Lattice The
Protein Crystallography Part II Tim Grüne Dept. of Structural Chemistry Prof. G. Sheldrick University of Göttingen http://shelx.uni-ac.gwdg.de tg@shelx.uni-ac.gwdg.de Overview The Reciprocal Lattice The
Nature Structural and Molecular Biology: doi: /nsmb Supplementary Figure 1. Definition and assessment of ciap1 constructs.
 Supplementary Figure 1 Definition and assessment of ciap1 constructs. (a) ciap1 constructs used in this study are shown as primary structure schematics with domains colored as in the main text. Mutations
Supplementary Figure 1 Definition and assessment of ciap1 constructs. (a) ciap1 constructs used in this study are shown as primary structure schematics with domains colored as in the main text. Mutations
Supplemental Information for:
 Supplemental Information for: New Insight into the Structure of RNA in Red clover necrotic mosaic virus and the Role of Divalent Cations Revealed by Small-Angle Neutron Scattering Stanton L. Martin a,
Supplemental Information for: New Insight into the Structure of RNA in Red clover necrotic mosaic virus and the Role of Divalent Cations Revealed by Small-Angle Neutron Scattering Stanton L. Martin a,
Neutron Scattering (Basics).
 Neutron Scattering (Basics). Cy M. Jeffries Introduction One of the new frontiers of structural biology is the study of the interactome: There are approximately 30 000 structural genes in the human genome.
Neutron Scattering (Basics). Cy M. Jeffries Introduction One of the new frontiers of structural biology is the study of the interactome: There are approximately 30 000 structural genes in the human genome.
Small-Angle X-Ray Scattering Reveals Compact Domain-Domain Interactions in the N-Terminal Region of Filamin C
 Small-Angle X-Ray Scattering Reveals Compact Domain-Domain Interactions in the N-Terminal Region of Filamin C Ritika Sethi*, Jari Ylänne Department of Biological and Environmental Science and Nanoscience
Small-Angle X-Ray Scattering Reveals Compact Domain-Domain Interactions in the N-Terminal Region of Filamin C Ritika Sethi*, Jari Ylänne Department of Biological and Environmental Science and Nanoscience
Methoden Moderner Röntgenphysik II - Vorlesung im Haupt-/Masterstudiengang, Universität Hamburg, SoSe 2016, S. Roth
 > 31.05. : Small-Angle X-ray Scattering (SAXS) > 0.06. : Applications & A short excursion into Polymeric materials > 04.06. : Grazing incidence SAXS (GISAXS) Methoden Moderner Röntgenphysik II - Vorlesung
> 31.05. : Small-Angle X-ray Scattering (SAXS) > 0.06. : Applications & A short excursion into Polymeric materials > 04.06. : Grazing incidence SAXS (GISAXS) Methoden Moderner Röntgenphysik II - Vorlesung
Sample preparation and characterization around SAXS
 Sample preparation and characterization around SAXS Experimental verification and validation? Rob Meijers EMBL Hamburg Garbage in? The right stuff Molecular weight Oligomerization state Monodispersity
Sample preparation and characterization around SAXS Experimental verification and validation? Rob Meijers EMBL Hamburg Garbage in? The right stuff Molecular weight Oligomerization state Monodispersity
Introduction to biological small angle scattering
 Introduction to biological small angle scattering Frank Gabel (IBS/ILL) HERCULES Specialized Course 16 (September 15 th 014) Length-scales and tools in structural biology small angle scattering in solution
Introduction to biological small angle scattering Frank Gabel (IBS/ILL) HERCULES Specialized Course 16 (September 15 th 014) Length-scales and tools in structural biology small angle scattering in solution
Proteins in solution: charge-tuning, cluster formation, liquid-liquid phase separation, and crystallization
 HERCULES Specialized Course: Non-atomic resolution scattering in biology and soft matter Grenoble, September 14-19, 2014 Proteins in solution: charge-tuning, cluster formation, liquid-liquid phase separation,
HERCULES Specialized Course: Non-atomic resolution scattering in biology and soft matter Grenoble, September 14-19, 2014 Proteins in solution: charge-tuning, cluster formation, liquid-liquid phase separation,
Joana Pereira Lamzin Group EMBL Hamburg, Germany. Small molecules How to identify and build them (with ARP/wARP)
 Joana Pereira Lamzin Group EMBL Hamburg, Germany Small molecules How to identify and build them (with ARP/wARP) The task at hand To find ligand density and build it! Fitting a ligand We have: electron
Joana Pereira Lamzin Group EMBL Hamburg, Germany Small molecules How to identify and build them (with ARP/wARP) The task at hand To find ligand density and build it! Fitting a ligand We have: electron
Introduction to X-ray and neutron scattering
 UNESCO/IUPAC Postgraduate Course in Polymer Science Lecture: Introduction to X-ray and neutron scattering Zhigunov Alexander Institute of Macromolecular Chemistry ASCR, Heyrovsky sq., Prague -16 06 http://www.imc.cas.cz/unesco/index.html
UNESCO/IUPAC Postgraduate Course in Polymer Science Lecture: Introduction to X-ray and neutron scattering Zhigunov Alexander Institute of Macromolecular Chemistry ASCR, Heyrovsky sq., Prague -16 06 http://www.imc.cas.cz/unesco/index.html
Synchrotron Methods in Nanomaterials Research
 Synchrotron Methods in Nanomaterials Research Marcel MiGLiERiNi Slovak University of Technology in Bratislava and Centre for Nanomaterials Research, Olomouc marcel.miglierini@stuba.sk www.nuc.elf.stuba.sk/bruno
Synchrotron Methods in Nanomaterials Research Marcel MiGLiERiNi Slovak University of Technology in Bratislava and Centre for Nanomaterials Research, Olomouc marcel.miglierini@stuba.sk www.nuc.elf.stuba.sk/bruno
Complementary use of SAXS and SANS. Jill Trewhella University of Sydney
 Complementary use of SAXS and SANS Jill Trewhella University of Sydney Conceptual diagram of the small-angle scattering experiment The conceptual experiment and theory is the same for X-rays and neutrons,
Complementary use of SAXS and SANS Jill Trewhella University of Sydney Conceptual diagram of the small-angle scattering experiment The conceptual experiment and theory is the same for X-rays and neutrons,
SUPPLEMENTARY INFORMATION
 Supplementary Information DNA-Programmable Nanoparticle Crystallization Sung Yong Park,* 1 Abigail K. R. Lytton-Jean,* 1 Byeongdu Lee 2, Steven Weigand 3, George C. Schatz 1 and Chad A. Mirkin 1 1 Department
Supplementary Information DNA-Programmable Nanoparticle Crystallization Sung Yong Park,* 1 Abigail K. R. Lytton-Jean,* 1 Byeongdu Lee 2, Steven Weigand 3, George C. Schatz 1 and Chad A. Mirkin 1 1 Department
Growing Biological crystals for Neutrons
 European Molecular Biology Laboratory Grenoble Outstation Growing Biological crystals for Neutrons Monika Budayova-Spano UJF EMBL CNRS / ILL Why neutron protein crystallography? Providing evidence on the
European Molecular Biology Laboratory Grenoble Outstation Growing Biological crystals for Neutrons Monika Budayova-Spano UJF EMBL CNRS / ILL Why neutron protein crystallography? Providing evidence on the
Experimental Techniques in Protein Structure Determination
 Experimental Techniques in Protein Structure Determination Homayoun Valafar Department of Computer Science and Engineering, USC Two Main Experimental Methods X-Ray crystallography Nuclear Magnetic Resonance
Experimental Techniques in Protein Structure Determination Homayoun Valafar Department of Computer Science and Engineering, USC Two Main Experimental Methods X-Ray crystallography Nuclear Magnetic Resonance
The Small Angle X-ray Scattering Technique: An Overview
 The Small Angle X-ray Scattering Technique: An Overview Dr. Gianluca Croce, Ph.D DISTA - Univ. Piemonte Orientale Via T. Michel 11,15121 Alessandria (Italy) gianluca.croce@mfn.unipmn.it Dr. Gianluca Croce
The Small Angle X-ray Scattering Technique: An Overview Dr. Gianluca Croce, Ph.D DISTA - Univ. Piemonte Orientale Via T. Michel 11,15121 Alessandria (Italy) gianluca.croce@mfn.unipmn.it Dr. Gianluca Croce
FRAGMENT SCREENING IN LEAD DISCOVERY BY WEAK AFFINITY CHROMATOGRAPHY (WAC )
 FRAGMENT SCREENING IN LEAD DISCOVERY BY WEAK AFFINITY CHROMATOGRAPHY (WAC ) SARomics Biostructures AB & Red Glead Discovery AB Medicon Village, Lund, Sweden Fragment-based lead discovery The basic idea:
FRAGMENT SCREENING IN LEAD DISCOVERY BY WEAK AFFINITY CHROMATOGRAPHY (WAC ) SARomics Biostructures AB & Red Glead Discovery AB Medicon Village, Lund, Sweden Fragment-based lead discovery The basic idea:
different subdomains of the lectin domain. W282A 10 mins 10 mins 20 mins 20 mins 40 mins 40 mins 1 hour 1 hour 2 hours 2 hours 4 hours 4 hours
 a +0 WT +1 +2 +3 GalNAc residues 1672.940 1875.891 20 mins 20 mins 40 mins 40 mins Intensity Intensity 1 hour 2079.133 2282.226 0 1000 1800 m/z 2200 1672.940 10 mins W282A 1 hour 2 hours 2 hours 4 hours
a +0 WT +1 +2 +3 GalNAc residues 1672.940 1875.891 20 mins 20 mins 40 mins 40 mins Intensity Intensity 1 hour 2079.133 2282.226 0 1000 1800 m/z 2200 1672.940 10 mins W282A 1 hour 2 hours 2 hours 4 hours
The Effect of Linker DNA on the Structure and Interaction of Nucleosome Core Particles
 Electronic Supplementary Material (ESI) for Soft Matter. This journal is The Royal Society of Chemistry 2018 SUPPLEMENTARY INFORMATION The Effect of Linker DNA on the Structure and Interaction of Nucleosome
Electronic Supplementary Material (ESI) for Soft Matter. This journal is The Royal Society of Chemistry 2018 SUPPLEMENTARY INFORMATION The Effect of Linker DNA on the Structure and Interaction of Nucleosome
Introduction to SAXS at SSRL
 Everything You Ever Wanted to Know About Introduction to SAXS at SSRL SAXS But Were Afraid to Ask John A Pople Stanford Synchrotron Radiation Laboratory, Stanford Linear Accelerator Center, Stanford CA
Everything You Ever Wanted to Know About Introduction to SAXS at SSRL SAXS But Were Afraid to Ask John A Pople Stanford Synchrotron Radiation Laboratory, Stanford Linear Accelerator Center, Stanford CA
Nuclear targeting by Nuclear Localization Signals (NLS) Richardson and Laskey (1988)
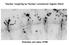 Nuclear targeting by Nuclear Localization Signals (NLS) Richardson and Laskey (1988) The nuclear import pathway of proteins containing a classical Nuclear Localization Signal (NLS) Uptake of NLS-containing
Nuclear targeting by Nuclear Localization Signals (NLS) Richardson and Laskey (1988) The nuclear import pathway of proteins containing a classical Nuclear Localization Signal (NLS) Uptake of NLS-containing
Schematic representation of relation between disorder and scattering
 Crystal lattice Reciprocal lattice FT Schematic representation of relation between disorder and scattering ρ = Δρ + Occupational disorder Diffuse scattering Bragg scattering ρ = Δρ + Positional
Crystal lattice Reciprocal lattice FT Schematic representation of relation between disorder and scattering ρ = Δρ + Occupational disorder Diffuse scattering Bragg scattering ρ = Δρ + Positional
Web-based Auto-Rickshaw for validation of the X-ray experiment at the synchrotron beamline
 Web-based Auto-Rickshaw for validation of the X-ray experiment at the synchrotron beamline Auto-Rickshaw http://www.embl-hamburg.de/auto-rickshaw A platform for automated crystal structure determination
Web-based Auto-Rickshaw for validation of the X-ray experiment at the synchrotron beamline Auto-Rickshaw http://www.embl-hamburg.de/auto-rickshaw A platform for automated crystal structure determination
Rietveld Structure Refinement of Protein Powder Diffraction Data using GSAS
 Rietveld Structure Refinement of Protein Powder Diffraction Data using GSAS Jon Wright ESRF, Grenoble, France Plan This is a users perspective Cover the protein specific aspects (assuming knowledge of
Rietveld Structure Refinement of Protein Powder Diffraction Data using GSAS Jon Wright ESRF, Grenoble, France Plan This is a users perspective Cover the protein specific aspects (assuming knowledge of
Protein crystallography. Garry Taylor
 Protein crystallography Garry Taylor X-ray Crystallography - the Basics Grow crystals Collect X-ray data Determine phases Calculate ρ-map Interpret map Refine coordinates Do the biology. Nitrogen at -180
Protein crystallography Garry Taylor X-ray Crystallography - the Basics Grow crystals Collect X-ray data Determine phases Calculate ρ-map Interpret map Refine coordinates Do the biology. Nitrogen at -180
X-Ray Damage to Biological Crystalline Samples
 X-Ray Damage to Biological Crystalline Samples Gerd Rosenbaum Structural Biology Center, ANL and Dept. of Biochemistry, UGA ACA Summer School IIT, 19 July 2007 A U.S. Department of Energy laboratory managed
X-Ray Damage to Biological Crystalline Samples Gerd Rosenbaum Structural Biology Center, ANL and Dept. of Biochemistry, UGA ACA Summer School IIT, 19 July 2007 A U.S. Department of Energy laboratory managed
Molecular shapes from small-angle X-ray scattering: extension of the theory to higher scattering angles
 Acta Crystallographica Section A Foundations of Crystallography ISSN 0108-7673 Editor: D. Schwarzenbach Molecular shapes from small-angle X-ray scattering: extension of the theory to higher scattering
Acta Crystallographica Section A Foundations of Crystallography ISSN 0108-7673 Editor: D. Schwarzenbach Molecular shapes from small-angle X-ray scattering: extension of the theory to higher scattering
Small Angle X-ray Scattering (SAXS)
 Small Angle X-ray Scattering (SAXS) We have considered that Bragg's Law, d = λ/(2 sinθ), supports a minimum size of measurement of λ/2 in a diffraction experiment (limiting sphere of inverse space) but
Small Angle X-ray Scattering (SAXS) We have considered that Bragg's Law, d = λ/(2 sinθ), supports a minimum size of measurement of λ/2 in a diffraction experiment (limiting sphere of inverse space) but
A program for SAXS data processing and analysis *
 A program for SAXS data processing and analysis * LI Zhi-Hong( ) 1) Beijing Synchrotron Radiation Facility, Institute of High Energy Physics, Chinese Academy of Sciences, Beijing 149, China Abstract: A
A program for SAXS data processing and analysis * LI Zhi-Hong( ) 1) Beijing Synchrotron Radiation Facility, Institute of High Energy Physics, Chinese Academy of Sciences, Beijing 149, China Abstract: A
X-ray Crystallography. Kalyan Das
 X-ray Crystallography Kalyan Das Electromagnetic Spectrum NMR 10 um - 10 mm 700 to 10 4 nm 400 to 700 nm 10 to 400 nm 10-1 to 10 nm 10-4 to 10-1 nm X-ray radiation was discovered by Roentgen in 1895. X-rays
X-ray Crystallography Kalyan Das Electromagnetic Spectrum NMR 10 um - 10 mm 700 to 10 4 nm 400 to 700 nm 10 to 400 nm 10-1 to 10 nm 10-4 to 10-1 nm X-ray radiation was discovered by Roentgen in 1895. X-rays
Methoden moderner Röntgenphysik II Streuung und Abbildung
 Methoden moderner Röntgenphysik II Streuung und Abbildung Stephan V. Roth DESY 1.5.15 Outline > 1.5. : Small-Angle X-ray Scattering (SAXS) > 19.5. : Applications & A short excursion into Polymeric materials
Methoden moderner Röntgenphysik II Streuung und Abbildung Stephan V. Roth DESY 1.5.15 Outline > 1.5. : Small-Angle X-ray Scattering (SAXS) > 19.5. : Applications & A short excursion into Polymeric materials
Structural characterization of NiV N 0 P in solution and in crystal.
 Supplementary Figure 1 Structural characterization of NiV N 0 P in solution and in crystal. (a) SAXS analysis of the N 32-383 0 -P 50 complex. The Guinier plot for complex concentrations of 0.55, 1.1,
Supplementary Figure 1 Structural characterization of NiV N 0 P in solution and in crystal. (a) SAXS analysis of the N 32-383 0 -P 50 complex. The Guinier plot for complex concentrations of 0.55, 1.1,
Internal structure of 15 nm 3-helix micelle revealed by small-angle neutron scattering and coarse-grained MD simulation
 Internal structure of 15 nm 3-helix micelle revealed by small-angle neutron scattering and coarse-grained MD simulation JooChuan Ang 1, Dan Ma, Reidar Lund 3, Sinan Keten, and Ting Xu *1,4,5 1 Department
Internal structure of 15 nm 3-helix micelle revealed by small-angle neutron scattering and coarse-grained MD simulation JooChuan Ang 1, Dan Ma, Reidar Lund 3, Sinan Keten, and Ting Xu *1,4,5 1 Department
THE SOLUTION CONFORMATION OF NOVEL ANTIBODY FRAGMENTS STUDIED USING THE PROTEOMELAB XL-A ANALYTICAL ULTRACENTRIFUGE
 APPLICATION INFORMATION Peter J. Morgan, Olwyn D. Byron, Stephen E. Harding Department of Applied Biochemistry and Food Science University of Nottingham Sutton Bonington, U. K. Introduction One of the
APPLICATION INFORMATION Peter J. Morgan, Olwyn D. Byron, Stephen E. Harding Department of Applied Biochemistry and Food Science University of Nottingham Sutton Bonington, U. K. Introduction One of the
Structural characterization. Part 2
 Structural characterization Part Determining partial pair distribution functions X-ray absorption spectroscopy (XAS). Atoms of different elements have absorption edges at different energies. Structure
Structural characterization Part Determining partial pair distribution functions X-ray absorption spectroscopy (XAS). Atoms of different elements have absorption edges at different energies. Structure
Dilute-solution properties of biomacromolecules as indicators of macromolecular structure and interactions
 Dilute-solution properties of biomacromolecules as indicators of macromolecular structure and interactions José García de la Torre, Departament of Physical Chemistry University of Murcia, Spain jgt@um.es
Dilute-solution properties of biomacromolecules as indicators of macromolecular structure and interactions José García de la Torre, Departament of Physical Chemistry University of Murcia, Spain jgt@um.es
X-ray Data Collection. Bio5325 Spring 2006
 X-ray Data Collection Bio535 Spring 006 Obtaining I hkl and α (Ihkl) from Frame Images Braggs Law -predicts conditions for in-phase scattering by equivalent atoms lying in planes that transect a crystal.
X-ray Data Collection Bio535 Spring 006 Obtaining I hkl and α (Ihkl) from Frame Images Braggs Law -predicts conditions for in-phase scattering by equivalent atoms lying in planes that transect a crystal.
CRYSTALLOGRAPHY AND STORYTELLING WITH DATA. President, Association of Women in Science, Bethesda Chapter STEM Consultant
 CRYSTALLOGRAPHY AND STORYTELLING WITH DATA President, Association of Women in Science, Bethesda Chapter STEM Consultant MY STORY Passion for Science BS Biology Major MS Biotechnology & Project in Bioinformatics
CRYSTALLOGRAPHY AND STORYTELLING WITH DATA President, Association of Women in Science, Bethesda Chapter STEM Consultant MY STORY Passion for Science BS Biology Major MS Biotechnology & Project in Bioinformatics
Scattering Lecture. February 24, 2014
 Scattering Lecture February 24, 2014 Structure Determination by Scattering Waves of radiation scattered by different objects interfere to give rise to an observable pattern! The wavelength needs to close
Scattering Lecture February 24, 2014 Structure Determination by Scattering Waves of radiation scattered by different objects interfere to give rise to an observable pattern! The wavelength needs to close
Direct Method. Very few protein diffraction data meet the 2nd condition
 Direct Method Two conditions: -atoms in the structure are equal-weighted -resolution of data are higher than the distance between the atoms in the structure Very few protein diffraction data meet the 2nd
Direct Method Two conditions: -atoms in the structure are equal-weighted -resolution of data are higher than the distance between the atoms in the structure Very few protein diffraction data meet the 2nd
Diffraction Geometry
 Diffraction Geometry Diffraction from a crystal - Laue equations Reciprocal lattice Ewald construction Data collection strategy Phil Evans LMB May 2013 MRC Laboratory of Molecular Biology Cambridge UK
Diffraction Geometry Diffraction from a crystal - Laue equations Reciprocal lattice Ewald construction Data collection strategy Phil Evans LMB May 2013 MRC Laboratory of Molecular Biology Cambridge UK
radiation damage Development of tools to automate quantitative analysis of radiation damage in SAXS experiments 1. Introduction
 Development of tools to automate quantitative analysis of radiation damage in SAXS experiments ISSN 1600-5775 Jonathan C. Brooks-Bartlett, a Rebecca A. Batters, a Charles S. Bury, a Edward D. Lowe, a Helen
Development of tools to automate quantitative analysis of radiation damage in SAXS experiments ISSN 1600-5775 Jonathan C. Brooks-Bartlett, a Rebecca A. Batters, a Charles S. Bury, a Edward D. Lowe, a Helen
Structure Analysis by Small-Angle X-Ray and Neutron Scattering
 Structure Analysis by Small-Angle X-Ray and Neutron Scattering L. A. Feigin and D. I. Svergun Institute of Crystallography Academy of Sciences of the USSR Moscow, USSR Edited by George W. Taylor Princeton
Structure Analysis by Small-Angle X-Ray and Neutron Scattering L. A. Feigin and D. I. Svergun Institute of Crystallography Academy of Sciences of the USSR Moscow, USSR Edited by George W. Taylor Princeton
research papers Online ion-exchange chromatography for small-angle X-ray scattering
 Online ion-exchange chromatography for small-angle X-ray scattering ISSN 2059-7983 Stephanie Hutin, a Martha Brennich, b * Benoit Maillot b,c and Adam Round d,e Received 20 November 2015 Accepted 9 August
Online ion-exchange chromatography for small-angle X-ray scattering ISSN 2059-7983 Stephanie Hutin, a Martha Brennich, b * Benoit Maillot b,c and Adam Round d,e Received 20 November 2015 Accepted 9 August
Part 8. Special Topic: Light Scattering
 Part 8. Special Topic: Light Scattering Light scattering occurs when polarizable particles in a sample are placed in the oscillating electric field of a beam of light. The varying field induces oscillating
Part 8. Special Topic: Light Scattering Light scattering occurs when polarizable particles in a sample are placed in the oscillating electric field of a beam of light. The varying field induces oscillating
Molecular Biology Course 2006 Protein Crystallography Part I
 Molecular Biology Course 2006 Protein Crystallography Part I Tim Grüne University of Göttingen Dept. of Structural Chemistry November 2006 http://shelx.uni-ac.gwdg.de tg@shelx.uni-ac.gwdg.de Overview Overview
Molecular Biology Course 2006 Protein Crystallography Part I Tim Grüne University of Göttingen Dept. of Structural Chemistry November 2006 http://shelx.uni-ac.gwdg.de tg@shelx.uni-ac.gwdg.de Overview Overview
Handout 7 Reciprocal Space
 Handout 7 Reciprocal Space Useful concepts for the analysis of diffraction data http://homepages.utoledo.edu/clind/ Concepts versus reality Reflection from lattice planes is just a concept that helps us
Handout 7 Reciprocal Space Useful concepts for the analysis of diffraction data http://homepages.utoledo.edu/clind/ Concepts versus reality Reflection from lattice planes is just a concept that helps us
Small-angle X-ray scattering a (mostly) theoretical introduction to the basics
 Small-angle X-ray scattering a (mostly) theoretical introduction to the basics András Wacha Research Centre for Natural Sciences, Hungarian Academy of Sciences Contents Introduction A bit of history The
Small-angle X-ray scattering a (mostly) theoretical introduction to the basics András Wacha Research Centre for Natural Sciences, Hungarian Academy of Sciences Contents Introduction A bit of history The
High Pressure Freezing. Philippe Carpentier,
 High Pressure Freezing Philippe Carpentier, - The system - The method - Some typical examples ESRF Users Meeting 2015: Meeting of MX BAG Representatives and Beamline Staff, 9 th February 2015 Page 1 INTRODUCTION
High Pressure Freezing Philippe Carpentier, - The system - The method - Some typical examples ESRF Users Meeting 2015: Meeting of MX BAG Representatives and Beamline Staff, 9 th February 2015 Page 1 INTRODUCTION
Biology III: Crystallographic phases
 Haupt/Masterstudiengang Physik Methoden moderner Röntgenphysik II: Streuung und Abbildung SS 2013 Biology III: Crystallographic phases Thomas R. Schneider, EMBL Hamburg 25/6/2013 thomas.schneider@embl-hamburg.de
Haupt/Masterstudiengang Physik Methoden moderner Röntgenphysik II: Streuung und Abbildung SS 2013 Biology III: Crystallographic phases Thomas R. Schneider, EMBL Hamburg 25/6/2013 thomas.schneider@embl-hamburg.de
Small Angle X-Ray Scattering What information can you get from this technique?
 Small Angle X-Ray Scattering 1 What information can you get from this technique? Small Angle X-Ray Scattering 2 A wide range of fields: Medicine Biology Chemistry Physics Archaeology Environmental and
Small Angle X-Ray Scattering 1 What information can you get from this technique? Small Angle X-Ray Scattering 2 A wide range of fields: Medicine Biology Chemistry Physics Archaeology Environmental and
Application of Bayesian analysis to indirect Fourier transformation in small-angle scattering
 Journal of Applied Crystallography ISSN 0021-8898 Editor: Gernot Kostorz Application of Bayesian analysis to indirect Fourier transformation in small-angle scattering Bente Vestergaard and Steen Hansen
Journal of Applied Crystallography ISSN 0021-8898 Editor: Gernot Kostorz Application of Bayesian analysis to indirect Fourier transformation in small-angle scattering Bente Vestergaard and Steen Hansen
USPAS course on Recirculated and Energy Recovered Linacs Ivan Bazarov, Cornell University Geoff Krafft, JLAB. ERL as a X-ray Light Source
 USPAS course on Recirculated and Energy Recovered Linacs Ivan Bazarov, Cornell University Geoff Krafft, JLAB ERL as a X-ray Light Source Contents Introduction Light sources landscape General motivation
USPAS course on Recirculated and Energy Recovered Linacs Ivan Bazarov, Cornell University Geoff Krafft, JLAB ERL as a X-ray Light Source Contents Introduction Light sources landscape General motivation
Chapter 11 Protein shape and assembly studied with X ray solution scattering: Fundaments and practice
 Chapter 11 Protein shape and assembly studied with X ray solution scattering: Fundaments and practice Rubén M Buey, Pablo Chacón, José Manuel Andreu and José Fernando Díaz Abstract Small Angle X ray Scattering
Chapter 11 Protein shape and assembly studied with X ray solution scattering: Fundaments and practice Rubén M Buey, Pablo Chacón, José Manuel Andreu and José Fernando Díaz Abstract Small Angle X ray Scattering
A Key Structural Role for Active Site Type 3 Copper Ions in Human Ceruloplasmin*
 THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 277, No. 43, Issue of October 25, pp. 40823 40831, 2002 2002 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. A Key Structural
THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 277, No. 43, Issue of October 25, pp. 40823 40831, 2002 2002 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. A Key Structural
research papers 1. Introduction Thomas C. Terwilliger a * and Joel Berendzen b
 Acta Crystallographica Section D Biological Crystallography ISSN 0907-4449 Discrimination of solvent from protein regions in native Fouriers as a means of evaluating heavy-atom solutions in the MIR and
Acta Crystallographica Section D Biological Crystallography ISSN 0907-4449 Discrimination of solvent from protein regions in native Fouriers as a means of evaluating heavy-atom solutions in the MIR and
SHELXC/D/E. Andrea Thorn
 SHELXC/D/E Andrea Thorn What is experimental phasing? Experimental phasing is what you do if MR doesn t work. What is experimental phasing? Experimental phasing methods depend on intensity differences.
SHELXC/D/E Andrea Thorn What is experimental phasing? Experimental phasing is what you do if MR doesn t work. What is experimental phasing? Experimental phasing methods depend on intensity differences.
The Reciprocal Lattice
 59-553 The Reciprocal Lattice 61 Because of the reciprocal nature of d spacings and θ from Bragg s Law, the pattern of the diffraction we observe can be related to the crystal lattice by a mathematical
59-553 The Reciprocal Lattice 61 Because of the reciprocal nature of d spacings and θ from Bragg s Law, the pattern of the diffraction we observe can be related to the crystal lattice by a mathematical
2) Measure sample and empties. 3) Measure transmissions of sample and empties. 4) Normalize to exposure time and transmission. 5) Subtract the empties
 1) Measure calibrants and direct beam to do Angular Integrations (transforms 2D into 1D) Need the distance SD Need center of beam Need to calculate q-vector 2) Measure sample and empties 3) Measure transmissions
1) Measure calibrants and direct beam to do Angular Integrations (transforms 2D into 1D) Need the distance SD Need center of beam Need to calculate q-vector 2) Measure sample and empties 3) Measure transmissions
How DLS Works: Interference of Light
 Static light scattering vs. Dynamic light scattering Static light scattering measures time-average intensities (mean square fluctuations) molecular weight radius of gyration second virial coefficient Dynamic
Static light scattering vs. Dynamic light scattering Static light scattering measures time-average intensities (mean square fluctuations) molecular weight radius of gyration second virial coefficient Dynamic
Supporting information for: Norovirus capsid proteins self-assemble through. biphasic kinetics via long-lived stave-like.
 Supporting information for: Norovirus capsid proteins self-assemble through biphasic kinetics via long-lived stave-like intermediates Guillaume Tresset,, Clémence Le Cœur, Jean-François Bryche, Mouna Tatou,
Supporting information for: Norovirus capsid proteins self-assemble through biphasic kinetics via long-lived stave-like intermediates Guillaume Tresset,, Clémence Le Cœur, Jean-François Bryche, Mouna Tatou,
shelxl: Refinement of Macromolecular Structures from Neutron Data
 ESS Neutron Protein Crystallography 2013 Aarhus, Denmark shelxl: Refinement of Macromolecular Structures from Neutron Data Tim Grüne University of Göttingen Dept. of Structural Chemistry http://shelx.uni-ac.gwdg.de
ESS Neutron Protein Crystallography 2013 Aarhus, Denmark shelxl: Refinement of Macromolecular Structures from Neutron Data Tim Grüne University of Göttingen Dept. of Structural Chemistry http://shelx.uni-ac.gwdg.de
X- ray crystallography. CS/CME/Biophys/BMI 279 Nov. 12, 2015 Ron Dror
 X- ray crystallography CS/CME/Biophys/BMI 279 Nov. 12, 2015 Ron Dror 1 Outline Overview of x-ray crystallography Crystals Electron density Diffraction patterns The computational problem: determining structure
X- ray crystallography CS/CME/Biophys/BMI 279 Nov. 12, 2015 Ron Dror 1 Outline Overview of x-ray crystallography Crystals Electron density Diffraction patterns The computational problem: determining structure
