Genomic Arrangement of Regulons in Bacterial Genomes
|
|
|
- Susan Bishop
- 5 years ago
- Views:
Transcription
1 l Genomes Han Zhang 1,2., Yanbin Yin 1,3., Victor Olman 1, Ying Xu 1,3,4 * 1 Computational Systems Biology Laboratory, Department of Biochemistry and Molecular Biology and Institute of Bioinformatics, University of Georgia, Athens, Georgia, United States of America, 2 Department of Automation and Intelligent Science, College of Information Technical Science, Nankai University, Tianjin, China, 3 BioEnergy Science Center, United States of America, 4 College of Computer Science and Technology, Jilin University, Changchun, Jilin, China Abstract Regulons, as groups of transcriptionally co-regulated operons, are the basic units of cellular response systems in bacterial cells. While the concept has been long and widely used in bacterial studies since it was first proposed in 1964, very little is known about how its component operons are arranged in a bacterial genome. We present a computational study to elucidate of the organizational principles of regulons in a bacterial genome, based on the experimentally validated regulons of E. coli and B. subtilis. Our results indicate that (1) genomic locations of transcriptional factors (TFs) are under stronger evolutionary constraints than those of the operons they regulate so changing a TF s genomic location will have larger impact to the bacterium than changing the genomic position of any of its target operons; (2) operons of regulons are generally not uniformly distributed in the genome but tend to form a few closely located clusters, which generally consist of genes working in the same metabolic pathways; and (3) the global arrangement of the component operons of all the regulons in a genome tends to minimize a simple scoring function, indicating that the global arrangement of regulons follows simple organizational principles. Citation: Zhang H, Yin Y, Olman V, Xu Y (2012) Genomic Arrangement of Regulons in Bacterial Genomes. PLoS ONE 7(1): e doi: / journal.pone Editor: Jonathan H. Badger, J. Craig Venter Institute, United States of America Received October 5, 2011; Accepted November 29, 2011; Published January 3, 2012 Copyright: ß 2012 Zhang et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. Funding: DOE of the US, BioEnergy Science Center grant (DE-PS02-06ER64304) NSF of the US, DEB The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript. Competing Interests: The authors have declared that no competing interests exist. * xyn@bmb.uga.edu. These authors contributed equally to this work. Introduction Regulons are the basic units of cellular response systems in bacterial cells, and represent a most basic concept in bacterial studies. A bacterial regulon is a group of operons that are transcriptionally co-regulated by the same regulatory machinery, consisting of trans regulators (transcription factors or simply TFs) and cis regulatory binding elements in the promoters of the operons they regulate. Operationally, a regulon contains operons regulated by one same transcription factor. Since the term regulon was first proposed in 1964 [1], 173 regulons have been fully or partially identified in E. coli K12 [2] and numerous more in other bacteria e.g. B. subtilis. Loosely speaking, regulons can be categorized into two classes: local and global regulons, with the former corresponding to regulons consisting of only a few component operons and the latter having a relatively large number of operons [3]. While the functionalities of the known regulons have been well studied, very little is known about how regulons are organized in a bacterial genome. The only published related work is by Janga et al. [4], which found that for small regulons, TFs tend to be located close to their targets (TGs). We present a computational study here to elucidate the organizational principles of regulons in a bacterial genome. We have carried out our study on E. coli K12 and also on B. subtilis str. 168 to demonstrate the generality of the results. Our key findings are (1) operons of each regulon tend to form a few closely located clusters along with genome; (2) TFs are under stronger evolutionary constraints than their TGs; and (3) the global arrangement of the component operons of all the (known) regulons in a genome tend to minimize a simple scoring function. Results and Discussion We have examined all the 3,684 regulatory relationships between TFs and their TGs in RegulonDB [2], involving 173 TFs and 729 TGs forming 173 regulons. We assigned genes to operons based on the operon information in the DOOR database (14) ( of E. coli K12. Among these regulatory relationships, 105 TFs are self-regulated; 123 (71%) of the 173 TFs regulate more than one TG; 411 (56%) of the 729 TGs are regulated by more than one TF; and 131 (18%) TGs are also TFs so they are regulated by upper-stream TFs while regulating downstream targets in the global transcription regulation network. Operons in a regulon tend to form clusters in terms of their genomic locations Intuitively we would expect that operons in a regulon should stay close to each other in a genome to facilitate efficient coregulation, which was used earlier to explain the formation of operons [5]. To test if this is indeed the case, we examined the distribution of the distances between two neighboring operons within a regulon, measured as the (smallest) number of operons between the two operons (we do not consider the orientations of operons). We noted that 523 (32%) of the 1,624 such distances in the 173 regulons are less than two (and 47% less than 10), as PLoS ONE 1 January 2012 Volume 7 Issue 1 e29496
2 shown in Figure 1A, suggesting that the component operons in a regulons tend to cluster together, although they may form multiple clusters. This remains to be true for all large regulons, which are defined as regulons with more than 5 component operons in this study. For example, crp, the largest regulon in E. coli, consists of 230 operons, 86 of which (37%) have distances less than two. As a control, we have checked a similar distance distribution calculated over 173 artificial regulons by randomly grouping E. coli K12 operons. Figure 1B shows the distance distribution, which is clearly very different from the one in Figure 1A. Similar observations were made when studying the regulons of B. subtilis (Figure 1C and 1D). We hypothesize that operons in each cluster of each regulon encode enzymes participating in the same metabolic pathway. To check for this, we consider a (maximal) list of operons of a regulon forms an operon cluster if the maximum distance between each pair of neighboring operons in this list is less than five. Using this definition, we obtained 353 operon clusters (85% from large regulons with size.5 as defined above), 242 of which each has at least two operons mapped to some SEED metabolic pathways ( [6]. Among them, 191 (79%) clusters have at least two operons participating in the same SEED pathway. Interestingly among these 191 clusters, 158 (83%) have all their mapped operons participating in the same SEED pathways. We noted that all these results do not change substantially when we adjust the distance cutoff from 5 to any integer between 2 and 7 when defining an operon cluster (Table S1). This suggests that each operon cluster generally consists of genes working in the same metabolic pathways. Similar observation (Table S2) was made when studying the B. subtilis regulons. Genomic locations of TFs are under stronger constraints than those of TGs It has been observed that small regulons tend to have their TFs located close to their TGs [4], suggesting that the genomic location of a TF is under strong constraints from its TGs (meaning TG s locations). A natural question is if a TG is under strong constraints from its TFs. We note that 56% TGs are regulated by more than one TF in E. coli K12. From Figure 2A, we conclude that there is no significant difference (Wilcoxon test P-value = 0.31) between Figure 1. Component operons of regulons tend to be clustered. The distance between two neighboring operons within a regulon is defined as the number of operons between the two operons. The bin width of the histogram is 1. (C) and (D) are similar to (A) and (B), respectively, but are for B. subtilis. doi: /journal.pone g001 PLoS ONE 2 January 2012 Volume 7 Issue 1 e29496
3 Figure 2. TFs are under stronger constraints than TGs. (A) All TGs are categorized into two groups, TGs regulated by one TF and TGs regulated by multiple TFs. The distributions of the distances (y axis) from TGs to their TFs are shown as box-plots. (B) All TGs are categorized into two groups, TGs that regulate other TGs, and TGs that do not regulate other TGs. The distributions of the distances (y axis) from TGs to their upstream TFs are shown as box-plots. Throughout this paper, the distance between a TF-TG pair is defined as the number of operons between the two operons. (C) and (D) are similar to (A) and (B), respectively, but are for B. subtilis. P-values of Wilcoxon tests are shown between two neighboring boxes. doi: /journal.pone g002 TGs regulated by one regulator and those regulated by multiple regulators in terms of the distances to their TFs, meaning that an operon does not have to stay close to its regulator even if it is only controlled by one regulator. This is not surprising because the regulator may control many targets, some of which might be close to the regulator while others may not. This finding is opposed to what we found for TFs controlling a small number of operons (Figure S1), suggesting that the genomic locations of TGs are generally under less constraints than those of TFs. To further study this, we divided all TGs into two groups, TGs that are also TFs regulating downstream TGs and TGs that are not TFs. Figure 2B shows that the first group of TGs tends to locate significantly closer (Wilcoxon test P-value = 1e23) to their regulators than the second group, directly suggesting that TGs are under stronger constraints than ordinary TGs from their upstream regulators if they are TFs themselves controlling further downstream targets. This is possibly due to the need for such TGs to have faster reaction time to send the regulatory signal down across the regulatory network. We have also performed all the analyses for B. subtilis str. 168 using data from DBTBS [7] database. The results are shown in Figure 2C and 2D and remain as significant as observed in E. coli, strongly suggesting that the observations made above are independent of which bacteria we use and hence may apply to all bacterial organisms in general. Global genomic arrangement of regulons All these observations led to our main hypothesis that the genomic locations of the component operons of all the regulons encoded in a genome are determined by some global organizational principle. Specifically we hypothesize that the global genomic arrangement of regulons tends to minimize the following function based on our preliminary study: D~ XN d i i~1 PLoS ONE 3 January 2012 Volume 7 Issue 1 e29496
4 where N is the total number of regulons encoded in a genome and d i represents the total distance between the genomic location of the TF and all the TGs of the i th regulon. Note that a similar formula has been used in our recent study on the genomic arrangements of metabolic pathways [8]. We have used the following procedure to demonstrate that the D value of all the known regulons encoded in the E. coli genome is significantly smaller than those of the vast majority of alternatively arranged genomes. Specifically, we have considered one million permutations of the genomic locations of X% of operons (both TFs and TGs) of E. coli K-12, for X = 10, 20,, 100 (see Methods). Figure 3A shows the D value distributions for different percentages of reshuffled locations of operons for the E. coli genome. We can clearly see that the current genomic arrangement of operons of E. coli K12 has a lower D value (the vertical dash line) than the vast majority of the D values of the reshuffled genomes, which is also supported by statistical tests (all P-values,0.05, see Table S3). It is also interesting to see that reshuffling TFs increases D values considerably more than reshuffling the same amount of TGs (Figure 3B, P-value,0.05), consistent with our observation made based on Figure 2 that TFs are under stronger constraints than TGs in terms of their genomic locations. For each known regulon in E. coli K12, we have also arbitrarily selected the same number of operons from the pool of all operons covered by the known regulons to form an artificial regulon, and do this for every known regulon. Again, we see the D value of the Figure 3. The distributions of D values calculated for the actual and reshuffled genomes. The x-axis represents the D values), and the y- axis is the frequency (density). In (A) each curve is calculated using one million permutations of the current arrangement of the operons in a genome under a specified constraint. Ten D distributions are calculated, with each distribution calculated allowing X% of operons randomly selected among all the operons under consideration and being randomly permutated, with X = 10, 20,, 100, respectively, where the ten curves from left to right are consistent with the order of X. The vertical dash line shows the D value for the current arrangement of the operons in the E. coli K12 genome. We also conducted permutations using a second manner, i.e. artificially forming regulons and calculating D values for permutated genomes. The result is shown as a dotted curve. (B) A comparison between the D distributions when randomly permuting 100 TGs (curve on the left) versus randomly permuting 100 TFs (curve on the right) in the genome of E. coli K-12. (C) and (D) are similar to (A) and (B), respectively, but are for B. subtilis. doi: /journal.pone g003 PLoS ONE 4 January 2012 Volume 7 Issue 1 e29496
5 real genome (vertical dashed line) is significantly smaller than those of genomes with artificially formed regulons (the dotted curve in Figure 3A). To ensure that the our observations hold for other bacterial genomes in general, we have checked the observation made in this section on all the 160 known regulons of B. subtilis, using the same procedure on E. coli genome and the results are as shown in Figure 3C and 3D (see also Table S3), which are clearly highly similar to those shown in Figure 3A and 3B. This work presents a systematic study of the genomic arrangement of regulons in terms of their organization in a bacterial genome. We made a number of interesting observations related to the organizational principles of regulons in a bacterial genome, namely (1) transcription factors of regulons are under strong constraints from their regulatory targets while TGs do not seem to be under strong constraints from their TFs; (2) regulons tend to form operon clusters, each of which tend to consist of operons encoding the same metabolic pathway; and (3) the genome tends to minimize the overall distance between the TFs and their TGs across all regulons encoded in the genome. We believe that all the observations are mostly due to the need by the cell to efficiently transcribe the relevant genes. Janga et al suggested that TFs of large regulons usually have high expression levels and presumably get to their targets through diffusion, and this might be the reason that they do not need to locate close to their targets. For small regulons, TFs are simply located closely to their targets which should be evolutionarily favored. For larger regulons consisting of multiply clustered operons, the three dimensional packing of the chromosome needs to be considered. It is likely that these organizational principles, along with a few others including genomic organization of metabolic pathways [8], the selfish operon model [9] and the nucleoid compaction [10,11,12,13,14], collectively determine the local and the global organization of all bacterial genes in a genome [15]. Materials and Methods Date sources The genome of E. coli K-12 MG1655 was downloaded from ftp://ftp.ncbi.nih.gov as of 01/14/2009. All the predicted operons for the organism were downloaded from the DOOR [16] database at All regulons data of E. coli K-12 MG1655 and of B. subtilis str. 168 were downloaded from the RegulonDB [17] and from the DBTBS [17] database, respectively, as of 03/2010. Operon shuffling For each reshuffled genome, the D value defined in the formula was calculated for X = 10, 20,, 100; X is the percentage of References 1. Maas WK (1964) Studies on the Mechanism of Repression of Arginine Biosynthesis in Escherichia Coli. Ii. Dominance of Repressibility in Diploids. J Mol Biol 8: Gama-Castro S, Jimenez-Jacinto V, Peralta-Gil M, Santos-Zavaleta A, Penaloza-Spinola MI, et al. (2008) RegulonDB (version 6.0): gene regulation model of Escherichia coli K-12 beyond transcription, active (experimental) annotated promoters and Textpresso navigation. Nucleic Acids Res 36: D Martinez-Antonio A, Collado-Vides J (2003) Identifying global regulators in transcriptional regulatory networks in bacteria. Curr Opin Microbiol 6: Janga SC, Salgado H, Martinez-Antonio A (2009) Transcriptional regulation shapes the organization of genes on bacterial chromosomes. Nucleic Acids Res 37: Jacob F, Monod J (1961) Genetic regulatory mechanisms in the synthesis of proteins. J Mol Biol 3: operons to be reshuffled (i.e. their genomic locations are permutated). The following two-step procedure was conducted to randomly shuffle a specified fraction (X%) of operons. We first randomly select operons among all operons of the E. coli genome for 10,000 times and then randomly permute their locations 100 times for each specific selection of the 10,000. So we do a total of one million permutations and calculate the D value distribution over the million rearranged genomes. Supporting Information Figure S1 Box plots of the distance distribution of operons (TGs) to their regulators (TFs). (A) is for E. coli and (B) is for B. subtilis. P-values of Wilcoxon tests are shown between two neighboring boxes. (EPS) Table S1 The number of operon clusters participating in the same SEED pathway under different distance cutoffs (E. coli). The first column represents the distance cutoff used to define a cluster. The second column is the number of clusters having at least two operons mapped to some SEED metabolic pathways. The third column is the number of clusters having at least two operons participating in the same SEED pathway. The fourth column is the number of clusters having all their mapped operons participating in the same SEED pathways. Regulons with at least two operons are considered. (DOC) Table S2 The number of operon clusters participating in the same SEED pathway under different distance cutoffs (B. subtilis). See Table S1 legend for details. Note the there are significantly less operons in B. subtilis than in E. coli that are mapped to the SEED pathways. This makes the numbers in Table S2 are much smaller than those in Table S1. (DOC) Table S3 Statistical tests of curves in Figure 3. The skewness and kurtosis columns are calculated to test if the curves in Figure 3 are normal distribution. skewness closer to 0 and kurtosis closer to 3 indicates close to normal distribution. The Pvalue column is calculated to test if the curves are significantly larger than the vertical dash line, indicating that the permutated genomes have significant larger D values than the actual genomes. (DOC) Author Contributions Conceived and designed the experiments: YY YX HZ. Performed the experiments: HZ YY. Analyzed the data: YY HZ. Contributed reagents/ materials/analysis tools: HZ YY VO. Wrote the paper: YY YX HZ. 6. Overbeek R, Begley T, Butler RM, Choudhuri JV, Chuang HY, et al. (2005) The subsystems approach to genome annotation and its use in the project to annotate 1000 genomes. Nucleic Acids Res 33: Sierro N, Makita Y, de Hoon M, Nakai K (2008) DBTBS: a database of transcriptional regulation in Bacillus subtilis containing upstream intergenic conservation information. Nucleic Acids Res 36: D Yin Y, Zhang H, Olman V, Xu Y (2010) Genomic arrangement of bacterial operons is constrained by biological pathways encoded in the genome. Proc Natl Acad Sci U S A 107: Lawrence JG, Roth JR (1996) Selfish operons: horizontal transfer may drive the evolution of gene clusters. Genetics 143: Carpentier AS, Torresani B, Grossmann A, Henaut A (2005) Decoding the nucleoid organisation of Bacillus subtilis and Escherichia coli through gene expression data. BMC Genomics 6: Postow L, Hardy CD, Arsuaga J, Cozzarelli NR (2004) Topological domain structure of the Escherichia coli chromosome. Genes Dev 18: PLoS ONE 5 January 2012 Volume 7 Issue 1 e29496
6 12. Wright MA, Kharchenko P, Church GM, Segre D (2007) Chromosomal periodicity of evolutionarily conserved gene pairs. Proc Natl Acad Sci U S A 104: Jeong KS, Ahn J, Khodursky AB (2004) Spatial patterns of transcriptional activity in the chromosome of Escherichia coli. Genome Biol 5: R Allen TE, Price ND, Joyce AR, Palsson BO (2006) Long-range periodic patterns in microbial genomes indicate significant multi-scale chromosomal organization. PLoS Comput Biol 2: e Rocha EP (2008) The organization of the bacterial genome. Annu Rev Genet 42: Mao F, Dam P, Chou J, Olman V, Xu Y (2009) DOOR: a database for prokaryotic operons. Nucleic Acids Res 37: D Salgado H, Gama-Castro S, Peralta-Gil M, Diaz-Peredo E, Sanchez-Solano F, et al. (2006) RegulonDB (version 5.0): Escherichia coli K-12 transcriptional regulatory network, operon organization, and growth conditions. Nucleic Acids Res 34: D PLoS ONE 6 January 2012 Volume 7 Issue 1 e29496
Computational methods for the analysis of bacterial gene regulation Brouwer, Rutger Wubbe Willem
 University of Groningen Computational methods for the analysis of bacterial gene regulation Brouwer, Rutger Wubbe Willem IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's
University of Groningen Computational methods for the analysis of bacterial gene regulation Brouwer, Rutger Wubbe Willem IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's
Thiago M Venancio and L Aravind
 Minireview Reconstructing prokaryotic transcriptional regulatory networks: lessons from actinobacteria Thiago M Venancio and L Aravind Address: National Center for Biotechnology Information, National Library
Minireview Reconstructing prokaryotic transcriptional regulatory networks: lessons from actinobacteria Thiago M Venancio and L Aravind Address: National Center for Biotechnology Information, National Library
COMPARATIVE PATHWAY ANNOTATION WITH PROTEIN-DNA INTERACTION AND OPERON INFORMATION VIA GRAPH TREE DECOMPOSITION
 COMPARATIVE PATHWAY ANNOTATION WITH PROTEIN-DNA INTERACTION AND OPERON INFORMATION VIA GRAPH TREE DECOMPOSITION JIZHEN ZHAO, DONGSHENG CHE AND LIMING CAI Department of Computer Science, University of Georgia,
COMPARATIVE PATHWAY ANNOTATION WITH PROTEIN-DNA INTERACTION AND OPERON INFORMATION VIA GRAPH TREE DECOMPOSITION JIZHEN ZHAO, DONGSHENG CHE AND LIMING CAI Department of Computer Science, University of Georgia,
Identification and annotation of promoter regions in microbial genome sequences on the basis of DNA stability
 Annotation of promoter regions in microbial genomes 851 Identification and annotation of promoter regions in microbial genome sequences on the basis of DNA stability VETRISELVI RANGANNAN and MANJU BANSAL*
Annotation of promoter regions in microbial genomes 851 Identification and annotation of promoter regions in microbial genome sequences on the basis of DNA stability VETRISELVI RANGANNAN and MANJU BANSAL*
A genomic-scale search for regulatory binding sites in the integration host factor regulon of Escherichia coli K12
 The integration host factor regulon of E. coli K12 genome 783 A genomic-scale search for regulatory binding sites in the integration host factor regulon of Escherichia coli K12 M. Trindade dos Santos and
The integration host factor regulon of E. coli K12 genome 783 A genomic-scale search for regulatory binding sites in the integration host factor regulon of Escherichia coli K12 M. Trindade dos Santos and
The percentage of bacterial genes on leading versus lagging strands is influenced by multiple balancing forces
 The percentage of bacterial genes on leading versus lagging strands is influenced by multiple balancing forces Xizeng Mao 1, Han Zhang 1,4, Yanbin Yin 1, 2 1, 2, 3, Ying Xu 1 Computational Systems Biology
The percentage of bacterial genes on leading versus lagging strands is influenced by multiple balancing forces Xizeng Mao 1, Han Zhang 1,4, Yanbin Yin 1, 2 1, 2, 3, Ying Xu 1 Computational Systems Biology
The EcoCyc Database. January 25, de Nitrógeno, UNAM,Cuernavaca, A.P. 565-A, Morelos, 62100, Mexico;
 The EcoCyc Database Peter D. Karp, Monica Riley, Milton Saier,IanT.Paulsen +, Julio Collado-Vides + Suzanne M. Paley, Alida Pellegrini-Toole,César Bonavides ++, and Socorro Gama-Castro ++ January 25, 2002
The EcoCyc Database Peter D. Karp, Monica Riley, Milton Saier,IanT.Paulsen +, Julio Collado-Vides + Suzanne M. Paley, Alida Pellegrini-Toole,César Bonavides ++, and Socorro Gama-Castro ++ January 25, 2002
Dynamic optimisation identifies optimal programs for pathway regulation in prokaryotes. - Supplementary Information -
 Dynamic optimisation identifies optimal programs for pathway regulation in prokaryotes - Supplementary Information - Martin Bartl a, Martin Kötzing a,b, Stefan Schuster c, Pu Li a, Christoph Kaleta b a
Dynamic optimisation identifies optimal programs for pathway regulation in prokaryotes - Supplementary Information - Martin Bartl a, Martin Kötzing a,b, Stefan Schuster c, Pu Li a, Christoph Kaleta b a
Microbial computational genomics of gene regulation*
 Pure Appl. Chem., Vol. 74, No. 6, pp. 899 905, 2002. 2002 IUPAC Microbial computational genomics of gene regulation* Julio Collado-Vides, Gabriel Moreno-Hagelsieb, and Arturo Medrano-Soto Program of Computational
Pure Appl. Chem., Vol. 74, No. 6, pp. 899 905, 2002. 2002 IUPAC Microbial computational genomics of gene regulation* Julio Collado-Vides, Gabriel Moreno-Hagelsieb, and Arturo Medrano-Soto Program of Computational
This document describes the process by which operons are predicted for genes within the BioHealthBase database.
 1. Purpose This document describes the process by which operons are predicted for genes within the BioHealthBase database. 2. Methods Description An operon is a coexpressed set of genes, transcribed onto
1. Purpose This document describes the process by which operons are predicted for genes within the BioHealthBase database. 2. Methods Description An operon is a coexpressed set of genes, transcribed onto
Hiromi Nishida. 1. Introduction. 2. Materials and Methods
 Evolutionary Biology Volume 212, Article ID 342482, 5 pages doi:1.1155/212/342482 Research Article Comparative Analyses of Base Compositions, DNA Sizes, and Dinucleotide Frequency Profiles in Archaeal
Evolutionary Biology Volume 212, Article ID 342482, 5 pages doi:1.1155/212/342482 Research Article Comparative Analyses of Base Compositions, DNA Sizes, and Dinucleotide Frequency Profiles in Archaeal
Conservation of Gene Co-Regulation between Two Prokaryotes: Bacillus subtilis and Escherichia coli
 116 Genome Informatics 16(1): 116 124 (2005) Conservation of Gene Co-Regulation between Two Prokaryotes: Bacillus subtilis and Escherichia coli Shujiro Okuda 1 Shuichi Kawashima 2 okuda@kuicr.kyoto-u.ac.jp
116 Genome Informatics 16(1): 116 124 (2005) Conservation of Gene Co-Regulation between Two Prokaryotes: Bacillus subtilis and Escherichia coli Shujiro Okuda 1 Shuichi Kawashima 2 okuda@kuicr.kyoto-u.ac.jp
Chapter 15 Active Reading Guide Regulation of Gene Expression
 Name: AP Biology Mr. Croft Chapter 15 Active Reading Guide Regulation of Gene Expression The overview for Chapter 15 introduces the idea that while all cells of an organism have all genes in the genome,
Name: AP Biology Mr. Croft Chapter 15 Active Reading Guide Regulation of Gene Expression The overview for Chapter 15 introduces the idea that while all cells of an organism have all genes in the genome,
DOOR: a prokaryotic operon database for genome analyses and functional inference
 Briefings in Bioinformatics, 2017, 1 8 doi: 10.1093/bib/bbx088 Software Review DOOR: a prokaryotic operon database for genome analyses and functional inference Huansheng Cao*, Qin Ma*, Xin Chen and Ying
Briefings in Bioinformatics, 2017, 1 8 doi: 10.1093/bib/bbx088 Software Review DOOR: a prokaryotic operon database for genome analyses and functional inference Huansheng Cao*, Qin Ma*, Xin Chen and Ying
56:198:582 Biological Networks Lecture 8
 56:198:582 Biological Networks Lecture 8 Course organization Two complementary approaches to modeling and understanding biological networks Constraint-based modeling (Palsson) System-wide Metabolism Steady-state
56:198:582 Biological Networks Lecture 8 Course organization Two complementary approaches to modeling and understanding biological networks Constraint-based modeling (Palsson) System-wide Metabolism Steady-state
BMC Systems Biology. Open Access. Abstract. BioMed Central
 BMC Systems Biology BioMed Central Research article Comparative analysis of the transcription-factor gene regulatory networks of and Lev Guzmán-Vargas* and Moisés Santillán 2,3 Open Access Address: Unidad
BMC Systems Biology BioMed Central Research article Comparative analysis of the transcription-factor gene regulatory networks of and Lev Guzmán-Vargas* and Moisés Santillán 2,3 Open Access Address: Unidad
INTERACTIVE CLUSTERING FOR EXPLORATION OF GENOMIC DATA
 INTERACTIVE CLUSTERING FOR EXPLORATION OF GENOMIC DATA XIUFENG WAN xw6@cs.msstate.edu Department of Computer Science Box 9637 JOHN A. BOYLE jab@ra.msstate.edu Department of Biochemistry and Molecular Biology
INTERACTIVE CLUSTERING FOR EXPLORATION OF GENOMIC DATA XIUFENG WAN xw6@cs.msstate.edu Department of Computer Science Box 9637 JOHN A. BOYLE jab@ra.msstate.edu Department of Biochemistry and Molecular Biology
SUPPLEMENTARY METHODS
 SUPPLEMENTARY METHODS M1: ALGORITHM TO RECONSTRUCT TRANSCRIPTIONAL NETWORKS M-2 Figure 1: Procedure to reconstruct transcriptional regulatory networks M-2 M2: PROCEDURE TO IDENTIFY ORTHOLOGOUS PROTEINSM-3
SUPPLEMENTARY METHODS M1: ALGORITHM TO RECONSTRUCT TRANSCRIPTIONAL NETWORKS M-2 Figure 1: Procedure to reconstruct transcriptional regulatory networks M-2 M2: PROCEDURE TO IDENTIFY ORTHOLOGOUS PROTEINSM-3
Evaluation of the relative contribution of each STRING feature in the overall accuracy operon classification
 Evaluation of the relative contribution of each STRING feature in the overall accuracy operon classification B. Taboada *, E. Merino 2, C. Verde 3 blanca.taboada@ccadet.unam.mx Centro de Ciencias Aplicadas
Evaluation of the relative contribution of each STRING feature in the overall accuracy operon classification B. Taboada *, E. Merino 2, C. Verde 3 blanca.taboada@ccadet.unam.mx Centro de Ciencias Aplicadas
Chapter 16 Lecture. Concepts Of Genetics. Tenth Edition. Regulation of Gene Expression in Prokaryotes
 Chapter 16 Lecture Concepts Of Genetics Tenth Edition Regulation of Gene Expression in Prokaryotes Chapter Contents 16.1 Prokaryotes Regulate Gene Expression in Response to Environmental Conditions 16.2
Chapter 16 Lecture Concepts Of Genetics Tenth Edition Regulation of Gene Expression in Prokaryotes Chapter Contents 16.1 Prokaryotes Regulate Gene Expression in Response to Environmental Conditions 16.2
APGRU6L2. Control of Prokaryotic (Bacterial) Genes
 APGRU6L2 Control of Prokaryotic (Bacterial) Genes 2007-2008 Bacterial metabolism Bacteria need to respond quickly to changes in their environment STOP u if they have enough of a product, need to stop production
APGRU6L2 Control of Prokaryotic (Bacterial) Genes 2007-2008 Bacterial metabolism Bacteria need to respond quickly to changes in their environment STOP u if they have enough of a product, need to stop production
Integration of Omics Data to Investigate Common Intervals
 2011 International Conference on Bioscience, Biochemistry and Bioinformatics IPCBEE vol.5 (2011) (2011) IACSIT Press, Singapore Integration of Omics Data to Investigate Common Intervals Sébastien Angibaud,
2011 International Conference on Bioscience, Biochemistry and Bioinformatics IPCBEE vol.5 (2011) (2011) IACSIT Press, Singapore Integration of Omics Data to Investigate Common Intervals Sébastien Angibaud,
CHAPTER 13 PROKARYOTE GENES: E. COLI LAC OPERON
 PROKARYOTE GENES: E. COLI LAC OPERON CHAPTER 13 CHAPTER 13 PROKARYOTE GENES: E. COLI LAC OPERON Figure 1. Electron micrograph of growing E. coli. Some show the constriction at the location where daughter
PROKARYOTE GENES: E. COLI LAC OPERON CHAPTER 13 CHAPTER 13 PROKARYOTE GENES: E. COLI LAC OPERON Figure 1. Electron micrograph of growing E. coli. Some show the constriction at the location where daughter
Division Ave. High School AP Biology
 Control of Prokaryotic (Bacterial) Genes 20072008 Bacterial metabolism n Bacteria need to respond quickly to changes in their environment u if they have enough of a product, need to stop production n why?
Control of Prokaryotic (Bacterial) Genes 20072008 Bacterial metabolism n Bacteria need to respond quickly to changes in their environment u if they have enough of a product, need to stop production n why?
Bacterial Genetics & Operons
 Bacterial Genetics & Operons The Bacterial Genome Because bacteria have simple genomes, they are used most often in molecular genetics studies Most of what we know about bacterial genetics comes from the
Bacterial Genetics & Operons The Bacterial Genome Because bacteria have simple genomes, they are used most often in molecular genetics studies Most of what we know about bacterial genetics comes from the
Prokaryotic Gene Expression (Learning Objectives)
 Prokaryotic Gene Expression (Learning Objectives) 1. Learn how bacteria respond to changes of metabolites in their environment: short-term and longer-term. 2. Compare and contrast transcriptional control
Prokaryotic Gene Expression (Learning Objectives) 1. Learn how bacteria respond to changes of metabolites in their environment: short-term and longer-term. 2. Compare and contrast transcriptional control
Name Period The Control of Gene Expression in Prokaryotes Notes
 Bacterial DNA contains genes that encode for many different proteins (enzymes) so that many processes have the ability to occur -not all processes are carried out at any one time -what allows expression
Bacterial DNA contains genes that encode for many different proteins (enzymes) so that many processes have the ability to occur -not all processes are carried out at any one time -what allows expression
L3.1: Circuits: Introduction to Transcription Networks. Cellular Design Principles Prof. Jenna Rickus
 L3.1: Circuits: Introduction to Transcription Networks Cellular Design Principles Prof. Jenna Rickus In this lecture Cognitive problem of the Cell Introduce transcription networks Key processing network
L3.1: Circuits: Introduction to Transcription Networks Cellular Design Principles Prof. Jenna Rickus In this lecture Cognitive problem of the Cell Introduce transcription networks Key processing network
Elucidation of the sequential transcriptional activity in Escherichia coli using time-series RNA-seq data
 www.bioinformation.net Volume 13(1) Hypothesis Elucidation of the sequential transcriptional activity in Escherichia coli using time-series RNA-seq data Pui Shan Wong 1, Kosuke Tashiro 2, Satoru Kuhara
www.bioinformation.net Volume 13(1) Hypothesis Elucidation of the sequential transcriptional activity in Escherichia coli using time-series RNA-seq data Pui Shan Wong 1, Kosuke Tashiro 2, Satoru Kuhara
CHAPTER : Prokaryotic Genetics
 CHAPTER 13.3 13.5: Prokaryotic Genetics 1. Most bacteria are not pathogenic. Identify several important roles they play in the ecosystem and human culture. 2. How do variations arise in bacteria considering
CHAPTER 13.3 13.5: Prokaryotic Genetics 1. Most bacteria are not pathogenic. Identify several important roles they play in the ecosystem and human culture. 2. How do variations arise in bacteria considering
56:198:582 Biological Networks Lecture 10
 56:198:582 Biological Networks Lecture 10 Temporal Programs and the Global Structure The single-input module (SIM) network motif The network motifs we have studied so far all had a defined number of nodes.
56:198:582 Biological Networks Lecture 10 Temporal Programs and the Global Structure The single-input module (SIM) network motif The network motifs we have studied so far all had a defined number of nodes.
Introduction. Gene expression is the combined process of :
 1 To know and explain: Regulation of Bacterial Gene Expression Constitutive ( house keeping) vs. Controllable genes OPERON structure and its role in gene regulation Regulation of Eukaryotic Gene Expression
1 To know and explain: Regulation of Bacterial Gene Expression Constitutive ( house keeping) vs. Controllable genes OPERON structure and its role in gene regulation Regulation of Eukaryotic Gene Expression
REVIEW SESSION. Wednesday, September 15 5:30 PM SHANTZ 242 E
 REVIEW SESSION Wednesday, September 15 5:30 PM SHANTZ 242 E Gene Regulation Gene Regulation Gene expression can be turned on, turned off, turned up or turned down! For example, as test time approaches,
REVIEW SESSION Wednesday, September 15 5:30 PM SHANTZ 242 E Gene Regulation Gene Regulation Gene expression can be turned on, turned off, turned up or turned down! For example, as test time approaches,
3.B.1 Gene Regulation. Gene regulation results in differential gene expression, leading to cell specialization.
 3.B.1 Gene Regulation Gene regulation results in differential gene expression, leading to cell specialization. We will focus on gene regulation in prokaryotes first. Gene regulation accounts for some of
3.B.1 Gene Regulation Gene regulation results in differential gene expression, leading to cell specialization. We will focus on gene regulation in prokaryotes first. Gene regulation accounts for some of
University of Groningen. The relative value of operon predictions Brouwer, Rutger W. W.; Kuipers, Oscar; van Hijum, Sacha A. F. T.
 University of Groningen The relative value of operon predictions Brouwer, Rutger W. W.; Kuipers, Oscar; van Hijum, Sacha A. F. T. Published in: Briefings in Bioinformatics DOI: 10.1093/bib/bbn019 IMPORTANT
University of Groningen The relative value of operon predictions Brouwer, Rutger W. W.; Kuipers, Oscar; van Hijum, Sacha A. F. T. Published in: Briefings in Bioinformatics DOI: 10.1093/bib/bbn019 IMPORTANT
AP Bio Module 16: Bacterial Genetics and Operons, Student Learning Guide
 Name: Period: Date: AP Bio Module 6: Bacterial Genetics and Operons, Student Learning Guide Getting started. Work in pairs (share a computer). Make sure that you log in for the first quiz so that you get
Name: Period: Date: AP Bio Module 6: Bacterial Genetics and Operons, Student Learning Guide Getting started. Work in pairs (share a computer). Make sure that you log in for the first quiz so that you get
Is Thermosensing Property of RNA Thermometers Unique?
 Is Thermosensing Property of RNA Thermometers Unique? Premal Shah 1,2 *, Michael A. Gilchrist 1,2 1 Department of Ecology and Evolutionary Biology, University of Tennessee, Knoxville, Tennessee, United
Is Thermosensing Property of RNA Thermometers Unique? Premal Shah 1,2 *, Michael A. Gilchrist 1,2 1 Department of Ecology and Evolutionary Biology, University of Tennessee, Knoxville, Tennessee, United
Lecture 8: Temporal programs and the global structure of transcription networks. Chap 5 of Alon. 5.1 Introduction
 Lecture 8: Temporal programs and the global structure of transcription networks Chap 5 of Alon 5. Introduction We will see in this chapter that sensory transcription networks are largely made of just four
Lecture 8: Temporal programs and the global structure of transcription networks Chap 5 of Alon 5. Introduction We will see in this chapter that sensory transcription networks are largely made of just four
Lecture 4: Transcription networks basic concepts
 Lecture 4: Transcription networks basic concepts - Activators and repressors - Input functions; Logic input functions; Multidimensional input functions - Dynamics and response time 2.1 Introduction The
Lecture 4: Transcription networks basic concepts - Activators and repressors - Input functions; Logic input functions; Multidimensional input functions - Dynamics and response time 2.1 Introduction The
Regulation of Gene Expression at the level of Transcription
 Regulation of Gene Expression at the level of Transcription (examples are mostly bacterial) Diarmaid Hughes ICM/Microbiology VT2009 Regulation of Gene Expression at the level of Transcription (examples
Regulation of Gene Expression at the level of Transcription (examples are mostly bacterial) Diarmaid Hughes ICM/Microbiology VT2009 Regulation of Gene Expression at the level of Transcription (examples
Introduction to Bioinformatics
 CSCI8980: Applied Machine Learning in Computational Biology Introduction to Bioinformatics Rui Kuang Department of Computer Science and Engineering University of Minnesota kuang@cs.umn.edu History of Bioinformatics
CSCI8980: Applied Machine Learning in Computational Biology Introduction to Bioinformatics Rui Kuang Department of Computer Science and Engineering University of Minnesota kuang@cs.umn.edu History of Bioinformatics
Biological Systems: Open Access
 Biological Systems: Open Access Biological Systems: Open Access Liu and Zheng, 2016, 5:1 http://dx.doi.org/10.4172/2329-6577.1000153 ISSN: 2329-6577 Research Article ariant Maps to Identify Coding and
Biological Systems: Open Access Biological Systems: Open Access Liu and Zheng, 2016, 5:1 http://dx.doi.org/10.4172/2329-6577.1000153 ISSN: 2329-6577 Research Article ariant Maps to Identify Coding and
Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression Edited by Shawn Lester PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley
Chapter 18 Regulation of Gene Expression Edited by Shawn Lester PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley
Bio 119 Bacterial Genomics 6/26/10
 BACTERIAL GENOMICS Reading in BOM-12: Sec. 11.1 Genetic Map of the E. coli Chromosome p. 279 Sec. 13.2 Prokaryotic Genomes: Sizes and ORF Contents p. 344 Sec. 13.3 Prokaryotic Genomes: Bioinformatic Analysis
BACTERIAL GENOMICS Reading in BOM-12: Sec. 11.1 Genetic Map of the E. coli Chromosome p. 279 Sec. 13.2 Prokaryotic Genomes: Sizes and ORF Contents p. 344 Sec. 13.3 Prokaryotic Genomes: Bioinformatic Analysis
Genetic Variation: The genetic substrate for natural selection. Horizontal Gene Transfer. General Principles 10/2/17.
 Genetic Variation: The genetic substrate for natural selection What about organisms that do not have sexual reproduction? Horizontal Gene Transfer Dr. Carol E. Lee, University of Wisconsin In prokaryotes:
Genetic Variation: The genetic substrate for natural selection What about organisms that do not have sexual reproduction? Horizontal Gene Transfer Dr. Carol E. Lee, University of Wisconsin In prokaryotes:
Proteomics Systems Biology
 Dr. Sanjeeva Srivastava IIT Bombay Proteomics Systems Biology IIT Bombay 2 1 DNA Genomics RNA Transcriptomics Global Cellular Protein Proteomics Global Cellular Metabolite Metabolomics Global Cellular
Dr. Sanjeeva Srivastava IIT Bombay Proteomics Systems Biology IIT Bombay 2 1 DNA Genomics RNA Transcriptomics Global Cellular Protein Proteomics Global Cellular Metabolite Metabolomics Global Cellular
Control of Prokaryotic (Bacterial) Gene Expression. AP Biology
 Control of Prokaryotic (Bacterial) Gene Expression Figure 18.1 How can this fish s eyes see equally well in both air and water? Aka. Quatro ojas Regulation of Gene Expression: Prokaryotes and eukaryotes
Control of Prokaryotic (Bacterial) Gene Expression Figure 18.1 How can this fish s eyes see equally well in both air and water? Aka. Quatro ojas Regulation of Gene Expression: Prokaryotes and eukaryotes
Supplemental Materials
 JOURNAL OF MICROBIOLOGY & BIOLOGY EDUCATION, May 2013, p. 107-109 DOI: http://dx.doi.org/10.1128/jmbe.v14i1.496 Supplemental Materials for Engaging Students in a Bioinformatics Activity to Introduce Gene
JOURNAL OF MICROBIOLOGY & BIOLOGY EDUCATION, May 2013, p. 107-109 DOI: http://dx.doi.org/10.1128/jmbe.v14i1.496 Supplemental Materials for Engaging Students in a Bioinformatics Activity to Introduce Gene
Gene regulation II Biochemistry 302. Bob Kelm February 28, 2005
 Gene regulation II Biochemistry 302 Bob Kelm February 28, 2005 Catabolic operons: Regulation by multiple signals targeting different TFs Catabolite repression: Activity of lac operon is restricted when
Gene regulation II Biochemistry 302 Bob Kelm February 28, 2005 Catabolic operons: Regulation by multiple signals targeting different TFs Catabolite repression: Activity of lac operon is restricted when
Development Team. Regulation of gene expression in Prokaryotes: Lac Operon. Molecular Cell Biology. Department of Zoology, University of Delhi
 Paper Module : 15 : 23 Development Team Principal Investigator : Prof. Neeta Sehgal Department of Zoology, University of Delhi Co-Principal Investigator : Prof. D.K. Singh Department of Zoology, University
Paper Module : 15 : 23 Development Team Principal Investigator : Prof. Neeta Sehgal Department of Zoology, University of Delhi Co-Principal Investigator : Prof. D.K. Singh Department of Zoology, University
Biology. Biology. Slide 1 of 26. End Show. Copyright Pearson Prentice Hall
 Biology Biology 1 of 26 Fruit fly chromosome 12-5 Gene Regulation Mouse chromosomes Fruit fly embryo Mouse embryo Adult fruit fly Adult mouse 2 of 26 Gene Regulation: An Example Gene Regulation: An Example
Biology Biology 1 of 26 Fruit fly chromosome 12-5 Gene Regulation Mouse chromosomes Fruit fly embryo Mouse embryo Adult fruit fly Adult mouse 2 of 26 Gene Regulation: An Example Gene Regulation: An Example
4. Why not make all enzymes all the time (even if not needed)? Enzyme synthesis uses a lot of energy.
 1 C2005/F2401 '10-- Lecture 15 -- Last Edited: 11/02/10 01:58 PM Copyright 2010 Deborah Mowshowitz and Lawrence Chasin Department of Biological Sciences Columbia University New York, NY. Handouts: 15A
1 C2005/F2401 '10-- Lecture 15 -- Last Edited: 11/02/10 01:58 PM Copyright 2010 Deborah Mowshowitz and Lawrence Chasin Department of Biological Sciences Columbia University New York, NY. Handouts: 15A
Topic 4 - #14 The Lactose Operon
 Topic 4 - #14 The Lactose Operon The Lactose Operon The lactose operon is an operon which is responsible for the transport and metabolism of the sugar lactose in E. coli. - Lactose is one of many organic
Topic 4 - #14 The Lactose Operon The Lactose Operon The lactose operon is an operon which is responsible for the transport and metabolism of the sugar lactose in E. coli. - Lactose is one of many organic
Outline. Genome Evolution. Genome. Genome Architecture. Constraints on Genome Evolution. New Evolutionary Synthesis 11/1/18
 Genome Evolution Outline 1. What: Patterns of Genome Evolution Carol Eunmi Lee Evolution 410 University of Wisconsin 2. Why? Evolution of Genome Complexity and the interaction between Natural Selection
Genome Evolution Outline 1. What: Patterns of Genome Evolution Carol Eunmi Lee Evolution 410 University of Wisconsin 2. Why? Evolution of Genome Complexity and the interaction between Natural Selection
CISC 636 Computational Biology & Bioinformatics (Fall 2016)
 CISC 636 Computational Biology & Bioinformatics (Fall 2016) Predicting Protein-Protein Interactions CISC636, F16, Lec22, Liao 1 Background Proteins do not function as isolated entities. Protein-Protein
CISC 636 Computational Biology & Bioinformatics (Fall 2016) Predicting Protein-Protein Interactions CISC636, F16, Lec22, Liao 1 Background Proteins do not function as isolated entities. Protein-Protein
Evaluation of the Number of Different Genomes on Medium and Identification of Known Genomes Using Composition Spectra Approach.
 Evaluation of the Number of Different Genomes on Medium and Identification of Known Genomes Using Composition Spectra Approach Valery Kirzhner *1 & Zeev Volkovich 2 1 Institute of Evolution, University
Evaluation of the Number of Different Genomes on Medium and Identification of Known Genomes Using Composition Spectra Approach Valery Kirzhner *1 & Zeev Volkovich 2 1 Institute of Evolution, University
Effects of Signalling on the Evolution of Gene Regulatory Networks
 Effects of Signalling on the Evolution of Gene Regulatory Networks Dafyd J. Jenkins and Dov J. Stekel Centre for Systems Biology, School of Biosciences, University of Birmingham, U.K., B15 2TT djj134@bham.ac.uk
Effects of Signalling on the Evolution of Gene Regulatory Networks Dafyd J. Jenkins and Dov J. Stekel Centre for Systems Biology, School of Biosciences, University of Birmingham, U.K., B15 2TT djj134@bham.ac.uk
15.2 Prokaryotic Transcription *
 OpenStax-CNX module: m52697 1 15.2 Prokaryotic Transcription * Shannon McDermott Based on Prokaryotic Transcription by OpenStax This work is produced by OpenStax-CNX and licensed under the Creative Commons
OpenStax-CNX module: m52697 1 15.2 Prokaryotic Transcription * Shannon McDermott Based on Prokaryotic Transcription by OpenStax This work is produced by OpenStax-CNX and licensed under the Creative Commons
Slide 1 / 7. Free Response
 Slide 1 / 7 Free Response Slide 2 / 7 Slide 3 / 7 1 The above diagrams illustrate the experiments carried out by Griffith and Hershey and Chaserespectively. Describe the hypothesis or conclusion that each
Slide 1 / 7 Free Response Slide 2 / 7 Slide 3 / 7 1 The above diagrams illustrate the experiments carried out by Griffith and Hershey and Chaserespectively. Describe the hypothesis or conclusion that each
Complete all warm up questions Focus on operon functioning we will be creating operon models on Monday
 Complete all warm up questions Focus on operon functioning we will be creating operon models on Monday 1. What is the Central Dogma? 2. How does prokaryotic DNA compare to eukaryotic DNA? 3. How is DNA
Complete all warm up questions Focus on operon functioning we will be creating operon models on Monday 1. What is the Central Dogma? 2. How does prokaryotic DNA compare to eukaryotic DNA? 3. How is DNA
BioControl - Week 6, Lecture 1
 BioControl - Week 6, Lecture 1 Goals of this lecture Large metabolic networks organization Design principles for small genetic modules - Rules based on gene demand - Rules based on error minimization Suggested
BioControl - Week 6, Lecture 1 Goals of this lecture Large metabolic networks organization Design principles for small genetic modules - Rules based on gene demand - Rules based on error minimization Suggested
Supplementary Information
 Supplementary Information 1 List of Figures 1 Models of circular chromosomes. 2 Distribution of distances between core genes in Escherichia coli K12, arc based model. 3 Distribution of distances between
Supplementary Information 1 List of Figures 1 Models of circular chromosomes. 2 Distribution of distances between core genes in Escherichia coli K12, arc based model. 3 Distribution of distances between
Computational Cell Biology Lecture 4
 Computational Cell Biology Lecture 4 Case Study: Basic Modeling in Gene Expression Yang Cao Department of Computer Science DNA Structure and Base Pair Gene Expression Gene is just a small part of DNA.
Computational Cell Biology Lecture 4 Case Study: Basic Modeling in Gene Expression Yang Cao Department of Computer Science DNA Structure and Base Pair Gene Expression Gene is just a small part of DNA.
Essentiality in B. subtilis
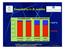 Essentiality in B. subtilis 100% 75% Essential genes Non-essential genes Lagging 50% 25% Leading 0% non-highly expressed highly expressed non-highly expressed highly expressed 1 http://www.pasteur.fr/recherche/unites/reg/
Essentiality in B. subtilis 100% 75% Essential genes Non-essential genes Lagging 50% 25% Leading 0% non-highly expressed highly expressed non-highly expressed highly expressed 1 http://www.pasteur.fr/recherche/unites/reg/
Gene regulation I Biochemistry 302. Bob Kelm February 25, 2005
 Gene regulation I Biochemistry 302 Bob Kelm February 25, 2005 Principles of gene regulation (cellular versus molecular level) Extracellular signals Chemical (e.g. hormones, growth factors) Environmental
Gene regulation I Biochemistry 302 Bob Kelm February 25, 2005 Principles of gene regulation (cellular versus molecular level) Extracellular signals Chemical (e.g. hormones, growth factors) Environmental
Introduction to Molecular and Cell Biology
 Introduction to Molecular and Cell Biology Molecular biology seeks to understand the physical and chemical basis of life. and helps us answer the following? What is the molecular basis of disease? What
Introduction to Molecular and Cell Biology Molecular biology seeks to understand the physical and chemical basis of life. and helps us answer the following? What is the molecular basis of disease? What
Outline. Genome Evolution. Genome. Genome Architecture. Constraints on Genome Evolution. New Evolutionary Synthesis 11/8/16
 Genome Evolution Outline 1. What: Patterns of Genome Evolution Carol Eunmi Lee Evolution 410 University of Wisconsin 2. Why? Evolution of Genome Complexity and the interaction between Natural Selection
Genome Evolution Outline 1. What: Patterns of Genome Evolution Carol Eunmi Lee Evolution 410 University of Wisconsin 2. Why? Evolution of Genome Complexity and the interaction between Natural Selection
Regulation of Gene Expression in Bacteria and Their Viruses
 11 Regulation of Gene Expression in Bacteria and Their Viruses WORKING WITH THE FIGURES 1. Compare the structure of IPTG shown in Figure 11-7 with the structure of galactose shown in Figure 11-5. Why is
11 Regulation of Gene Expression in Bacteria and Their Viruses WORKING WITH THE FIGURES 1. Compare the structure of IPTG shown in Figure 11-7 with the structure of galactose shown in Figure 11-5. Why is
CSCE555 Bioinformatics. Protein Function Annotation
 CSCE555 Bioinformatics Protein Function Annotation Why we need to do function annotation? Fig from: Network-based prediction of protein function. Molecular Systems Biology 3:88. 2007 What s function? The
CSCE555 Bioinformatics Protein Function Annotation Why we need to do function annotation? Fig from: Network-based prediction of protein function. Molecular Systems Biology 3:88. 2007 What s function? The
Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Tuesday, December 27, 16
 Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Enduring understanding 3.B: Expression of genetic information involves cellular and molecular
Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Enduring understanding 3.B: Expression of genetic information involves cellular and molecular
Organization of Genes Differs in Prokaryotic and Eukaryotic DNA Chapter 10 p
 Organization of Genes Differs in Prokaryotic and Eukaryotic DNA Chapter 10 p.110-114 Arrangement of information in DNA----- requirements for RNA Common arrangement of protein-coding genes in prokaryotes=
Organization of Genes Differs in Prokaryotic and Eukaryotic DNA Chapter 10 p.110-114 Arrangement of information in DNA----- requirements for RNA Common arrangement of protein-coding genes in prokaryotes=
2012 Univ Aguilera Lecture. Introduction to Molecular and Cell Biology
 2012 Univ. 1301 Aguilera Lecture Introduction to Molecular and Cell Biology Molecular biology seeks to understand the physical and chemical basis of life. and helps us answer the following? What is the
2012 Univ. 1301 Aguilera Lecture Introduction to Molecular and Cell Biology Molecular biology seeks to understand the physical and chemical basis of life. and helps us answer the following? What is the
V14 extreme pathways
 V14 extreme pathways A torch is directed at an open door and shines into a dark room... What area is lighted? Instead of marking all lighted points individually, it would be sufficient to characterize
V14 extreme pathways A torch is directed at an open door and shines into a dark room... What area is lighted? Instead of marking all lighted points individually, it would be sufficient to characterize
Computational approaches for functional genomics
 Computational approaches for functional genomics Kalin Vetsigian October 31, 2001 The rapidly increasing number of completely sequenced genomes have stimulated the development of new methods for finding
Computational approaches for functional genomics Kalin Vetsigian October 31, 2001 The rapidly increasing number of completely sequenced genomes have stimulated the development of new methods for finding
Network motifs in the transcriptional regulation network (of Escherichia coli):
 Network motifs in the transcriptional regulation network (of Escherichia coli): Janne.Ravantti@Helsinki.Fi (disclaimer: IANASB) Contents: Transcription Networks (aka. The Very Boring Biology Part ) Network
Network motifs in the transcriptional regulation network (of Escherichia coli): Janne.Ravantti@Helsinki.Fi (disclaimer: IANASB) Contents: Transcription Networks (aka. The Very Boring Biology Part ) Network
Gene regulation II Biochemistry 302. February 27, 2006
 Gene regulation II Biochemistry 302 February 27, 2006 Molecular basis of inhibition of RNAP by Lac repressor 35 promoter site 10 promoter site CRP/DNA complex 60 Lewis, M. et al. (1996) Science 271:1247
Gene regulation II Biochemistry 302 February 27, 2006 Molecular basis of inhibition of RNAP by Lac repressor 35 promoter site 10 promoter site CRP/DNA complex 60 Lewis, M. et al. (1996) Science 271:1247
the noisy gene Biology of the Universidad Autónoma de Madrid Jan 2008 Juan F. Poyatos Spanish National Biotechnology Centre (CNB)
 Biology of the the noisy gene Universidad Autónoma de Madrid Jan 2008 Juan F. Poyatos Spanish National Biotechnology Centre (CNB) day III: noisy bacteria - Regulation of noise (B. subtilis) - Intrinsic/Extrinsic
Biology of the the noisy gene Universidad Autónoma de Madrid Jan 2008 Juan F. Poyatos Spanish National Biotechnology Centre (CNB) day III: noisy bacteria - Regulation of noise (B. subtilis) - Intrinsic/Extrinsic
Translation - Prokaryotes
 1 Translation - Prokaryotes Shine-Dalgarno (SD) Sequence rrna 3 -GAUACCAUCCUCCUUA-5 mrna...ggagg..(5-7bp)...aug Influences: Secondary structure!! SD and AUG in unstructured region Start AUG 91% GUG 8 UUG
1 Translation - Prokaryotes Shine-Dalgarno (SD) Sequence rrna 3 -GAUACCAUCCUCCUUA-5 mrna...ggagg..(5-7bp)...aug Influences: Secondary structure!! SD and AUG in unstructured region Start AUG 91% GUG 8 UUG
Bacterial regulon modeling and prediction based on systematic cis regulatory motif analyses
 University of Nebraska - Lincoln DigitalCommons@University of Nebraska - Lincoln CSE Journal Articles Computer Science and Engineering, Department of 2016 Bacterial regulon modeling and prediction based
University of Nebraska - Lincoln DigitalCommons@University of Nebraska - Lincoln CSE Journal Articles Computer Science and Engineering, Department of 2016 Bacterial regulon modeling and prediction based
THE EVOLUTIONARY DYNAMICS OF TRANSCRIPTION FACTORS, OPERATORS, AND THEIR TARGET GENES ACROSS PROKARYOTES
 Wilfrid Laurier University Scholars Commons @ Laurier Theses and Dissertations (Comprehensive) 2014 THE EVOLUTIONARY DYNAMICS OF TRANSCRIPTION FACTORS, OPERATORS, AND THEIR TARGET GENES ACROSS PROKARYOTES
Wilfrid Laurier University Scholars Commons @ Laurier Theses and Dissertations (Comprehensive) 2014 THE EVOLUTIONARY DYNAMICS OF TRANSCRIPTION FACTORS, OPERATORS, AND THEIR TARGET GENES ACROSS PROKARYOTES
Welcome to Class 21!
 Welcome to Class 21! Introductory Biochemistry! Lecture 21: Outline and Objectives l Regulation of Gene Expression in Prokaryotes! l transcriptional regulation! l principles! l lac operon! l trp attenuation!
Welcome to Class 21! Introductory Biochemistry! Lecture 21: Outline and Objectives l Regulation of Gene Expression in Prokaryotes! l transcriptional regulation! l principles! l lac operon! l trp attenuation!
SUPPLEMENTARY INFORMATION
 Supplementary information S1 (box). Supplementary Methods description. Prokaryotic Genome Database Archaeal and bacterial genome sequences were downloaded from the NCBI FTP site (ftp://ftp.ncbi.nlm.nih.gov/genomes/all/)
Supplementary information S1 (box). Supplementary Methods description. Prokaryotic Genome Database Archaeal and bacterial genome sequences were downloaded from the NCBI FTP site (ftp://ftp.ncbi.nlm.nih.gov/genomes/all/)
Computational methods for predicting protein-protein interactions
 Computational methods for predicting protein-protein interactions Tomi Peltola T-61.6070 Special course in bioinformatics I 3.4.2008 Outline Biological background Protein-protein interactions Computational
Computational methods for predicting protein-protein interactions Tomi Peltola T-61.6070 Special course in bioinformatics I 3.4.2008 Outline Biological background Protein-protein interactions Computational
THE EDIBLE OPERON David O. Freier Lynchburg College [BIOL 220W Cellular Diversity]
![THE EDIBLE OPERON David O. Freier Lynchburg College [BIOL 220W Cellular Diversity] THE EDIBLE OPERON David O. Freier Lynchburg College [BIOL 220W Cellular Diversity]](/thumbs/90/102203867.jpg) THE EDIBLE OPERON David O. Freier Lynchburg College [BIOL 220W Cellular Diversity] You have the following resources available to you: Short bread cookies = DNA / genetic elements Fudge Mint cookies = RNA
THE EDIBLE OPERON David O. Freier Lynchburg College [BIOL 220W Cellular Diversity] You have the following resources available to you: Short bread cookies = DNA / genetic elements Fudge Mint cookies = RNA
Initiation of translation in eukaryotic cells:connecting the head and tail
 Initiation of translation in eukaryotic cells:connecting the head and tail GCCRCCAUGG 1: Multiple initiation factors with distinct biochemical roles (linking, tethering, recruiting, and scanning) 2: 5
Initiation of translation in eukaryotic cells:connecting the head and tail GCCRCCAUGG 1: Multiple initiation factors with distinct biochemical roles (linking, tethering, recruiting, and scanning) 2: 5
REGULATION OF GENE EXPRESSION. Bacterial Genetics Lac and Trp Operon
 REGULATION OF GENE EXPRESSION Bacterial Genetics Lac and Trp Operon Levels of Metabolic Control The amount of cellular products can be controlled by regulating: Enzyme activity: alters protein function
REGULATION OF GENE EXPRESSION Bacterial Genetics Lac and Trp Operon Levels of Metabolic Control The amount of cellular products can be controlled by regulating: Enzyme activity: alters protein function
GACE Biology Assessment Test I (026) Curriculum Crosswalk
 Subarea I. Cell Biology: Cell Structure and Function (50%) Objective 1: Understands the basic biochemistry and metabolism of living organisms A. Understands the chemical structures and properties of biologically
Subarea I. Cell Biology: Cell Structure and Function (50%) Objective 1: Understands the basic biochemistry and metabolism of living organisms A. Understands the chemical structures and properties of biologically
V19 Metabolic Networks - Overview
 V19 Metabolic Networks - Overview There exist different levels of computational methods for describing metabolic networks: - stoichiometry/kinetics of classical biochemical pathways (glycolysis, TCA cycle,...
V19 Metabolic Networks - Overview There exist different levels of computational methods for describing metabolic networks: - stoichiometry/kinetics of classical biochemical pathways (glycolysis, TCA cycle,...
Biology 105/Summer Bacterial Genetics 8/12/ Bacterial Genomes p Gene Transfer Mechanisms in Bacteria p.
 READING: 14.2 Bacterial Genomes p. 481 14.3 Gene Transfer Mechanisms in Bacteria p. 486 Suggested Problems: 1, 7, 13, 14, 15, 20, 22 BACTERIAL GENETICS AND GENOMICS We still consider the E. coli genome
READING: 14.2 Bacterial Genomes p. 481 14.3 Gene Transfer Mechanisms in Bacteria p. 486 Suggested Problems: 1, 7, 13, 14, 15, 20, 22 BACTERIAL GENETICS AND GENOMICS We still consider the E. coli genome
Measuring TF-DNA interactions
 Measuring TF-DNA interactions How is Biological Complexity Achieved? Mediated by Transcription Factors (TFs) 2 Regulation of Gene Expression by Transcription Factors TF trans-acting factors TF TF TF TF
Measuring TF-DNA interactions How is Biological Complexity Achieved? Mediated by Transcription Factors (TFs) 2 Regulation of Gene Expression by Transcription Factors TF trans-acting factors TF TF TF TF
Regulation of gene expression. Premedical - Biology
 Regulation of gene expression Premedical - Biology Regulation of gene expression in prokaryotic cell Operon units system of negative feedback positive and negative regulation in eukaryotic cell - at any
Regulation of gene expression Premedical - Biology Regulation of gene expression in prokaryotic cell Operon units system of negative feedback positive and negative regulation in eukaryotic cell - at any
Honors Biology Reading Guide Chapter 11
 Honors Biology Reading Guide Chapter 11 v Promoter a specific nucleotide sequence in DNA located near the start of a gene that is the binding site for RNA polymerase and the place where transcription begins
Honors Biology Reading Guide Chapter 11 v Promoter a specific nucleotide sequence in DNA located near the start of a gene that is the binding site for RNA polymerase and the place where transcription begins
Prokaryotic Gene Expression (Learning Objectives)
 Prokaryotic Gene Expression (Learning Objectives) 1. Learn how bacteria respond to changes of metabolites in their environment: short-term and longer-term. 2. Compare and contrast transcriptional control
Prokaryotic Gene Expression (Learning Objectives) 1. Learn how bacteria respond to changes of metabolites in their environment: short-term and longer-term. 2. Compare and contrast transcriptional control
Introduction to Bioinformatics Integrated Science, 11/9/05
 1 Introduction to Bioinformatics Integrated Science, 11/9/05 Morris Levy Biological Sciences Research: Evolutionary Ecology, Plant- Fungal Pathogen Interactions Coordinator: BIOL 495S/CS490B/STAT490B Introduction
1 Introduction to Bioinformatics Integrated Science, 11/9/05 Morris Levy Biological Sciences Research: Evolutionary Ecology, Plant- Fungal Pathogen Interactions Coordinator: BIOL 495S/CS490B/STAT490B Introduction
Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Chapter 18 Regulation of Gene Expression PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Methods to reconstruct and compare transcriptional regulatory networks
 Methods to reconstruct and compare transcriptional regulatory networks M. Madan Babu 1*, Benjamin Lang 1 & L. Aravind 2 1 MRC Laboratory of Molecular Biology, Hills Road, Cambridge CB22QH, U.K. 2 National
Methods to reconstruct and compare transcriptional regulatory networks M. Madan Babu 1*, Benjamin Lang 1 & L. Aravind 2 1 MRC Laboratory of Molecular Biology, Hills Road, Cambridge CB22QH, U.K. 2 National
Cell biology traditionally identifies proteins based on their individual actions as catalysts, signaling
 Lethality and centrality in protein networks Cell biology traditionally identifies proteins based on their individual actions as catalysts, signaling molecules, or building blocks of cells and microorganisms.
Lethality and centrality in protein networks Cell biology traditionally identifies proteins based on their individual actions as catalysts, signaling molecules, or building blocks of cells and microorganisms.
The binding of transcription factors (TFs) to specific sites is a
 Evolutionary comparisons suggest many novel camp response protein binding sites in Escherichia coli C. T. Brown* and C. G. Callan, Jr. *Division of Biology, California Institute of Technology, Pasadena,
Evolutionary comparisons suggest many novel camp response protein binding sites in Escherichia coli C. T. Brown* and C. G. Callan, Jr. *Division of Biology, California Institute of Technology, Pasadena,
BMC Evolutionary Biology 2008, 8:23
 BMC Evolutionary Biology This Provisional PDF corresponds to the article as it appeared upon acceptance. Fully formatted PDF and full text (HTML) versions will be made available soon. The impact of horizontal
BMC Evolutionary Biology This Provisional PDF corresponds to the article as it appeared upon acceptance. Fully formatted PDF and full text (HTML) versions will be made available soon. The impact of horizontal
UNIVERSITY OF YORK. BA, BSc, and MSc Degree Examinations Department : BIOLOGY. Title of Exam: Molecular microbiology
 Examination Candidate Number: Desk Number: UNIVERSITY OF YORK BA, BSc, and MSc Degree Examinations 2017-8 Department : BIOLOGY Title of Exam: Molecular microbiology Time Allowed: 1 hour 30 minutes Marking
Examination Candidate Number: Desk Number: UNIVERSITY OF YORK BA, BSc, and MSc Degree Examinations 2017-8 Department : BIOLOGY Title of Exam: Molecular microbiology Time Allowed: 1 hour 30 minutes Marking
