The pairwise approach to analysing species co-occurrence
|
|
|
- Todd Adams
- 6 years ago
- Views:
Transcription
1 Journal of Biogeography (J. Biogeogr.) (2014) 41, GUEST EDITORIAL The pairwise approach to analysing species co-occurrence Joseph A. Veech Department of Biology, Texas State University, San Marcos, TX , USA Correspondence: Joseph A. Veech, Department of Biology, Texas State University, San Marcos, TX , USA. ABSTRACT The of species co-occurrence patterns continues to be a main pursuit of ecologists, primarily because the coexistence of species is fundamentally important in evaluating various theories, principles and concepts. Examples include community assembly, equilibrium versus non-equilibrium organization of communities, resource partitioning and ecological character displacement, the local regional species diversity relationship, and the metacommunity concept. Traditionally, co-occurrence has been measured and tested at the level of an entire species presence absence matrix wherein various algorithms are used to randomize matrices and produce statistical null distributions of metrics that quantify structure in the matrix. This approach implicitly recognizes a presence absence matrix as having some real ecological identity (e.g. a set of species exhibiting nestedness among a set of islands) in addition to being a unit of statistical. An emerging alternative is to test for non-random co-occurrence between paired species. The pairwise approach does not analyse matrix-level structure and thus views a species pair as the fundamental unit of co-occurrence. Inferring process from pattern is very difficult in analyses of co-occurrence; however, the pairwise approach may make this task easier by simplifying the and resulting inferences to associations between paired species. Keywords Assembly rules, competition, C-score, data randomization, ecological community, hypothesis testing, Jared Diamond, pattern, presence absence matrix, species coexistence. INTRODUCTION The of species co-occurrence has a long history played out in studies ranging from community ecology to biogeography. Jared Diamond s classic study of the distributions of terrestrial birds on islands of New Guinea was one of the first analyses of species co-occurrence (Diamond, 1973, 1975). His study emphasized diffuse competition the complex situations resulting from the sum of competitive effects from many other somewhat similar species (Diamond, 1975, p. 348) and introduced the idea of assembly rules and forbidden combinations (of species). By definition, testing for diffuse competition requires of co-occurrence among multiple species within an assemblage, not between just two species of a pair. In subsequent years, Diamond s analytical methods were rigorously criticized (Connor & Simberloff, 1979) and confidently defended (Diamond & Gilpin, 1982; Gilpin & Diamond, 1984); see Sanderson (2000) for a review. As a result of these historical beginnings, studies of species co-occurrence have come to be characterized by the methods utilized (and defended) as well as a primary focus on the entire assemblage (i.e. the presence absence matrix) as the unit of. The matrix-level approach is well illustrated by the study of nestedness, the way in which smaller sets of species form orderly subsets of increasingly larger sets (Wright & Reeves, 1992; Almeido- Neto et al., 2008; Ulrich et al., 2009). An emerging alternative to the matrix approach is to examine pairwise co-occurrence (Sanderson, 2000; Sfenthourakis et al., 2004, 2006; Veech, 2006; Sanderson et al., 2009; Gotelli & Ulrich, 2010; Collins et al., 2011; Pitta et al., 2012) with methods that can detect patterns of positive, negative or random association between two species (Veech, 2013; see also Ricklefs, 2011, for a pairwise assessment of abundances). Beyond being a more comprehensive statistical assessment, these new methods are also suggesting (sometimes implicitly) doi: /jbi.12318
2 J. A. Veech that species may not be organized into neat and tidy communities that exist as discrete spatial entities. At the very least, the local community concept (i.e. Clementsian organization of nature into very similar sets of species repeated at different locales) may be insufficient as a framework for understanding the geographical distribution of species and diversity patterns (Ricklefs, 2008). If so, then analyses of cooccurrence patterns are most appropriate when conducted on species pairs instead of on entire species presence absence matrices. Of interest, Pielou (1977) described a pairwise approach several decades ago, although it never caught on. In this essay, I describe more recent pairwise approaches and distinguish them from matrix-level approaches and specifically emphasize benefits of the former. I also argue that the study of species co-occurrence now needs to move beyond the view that nature is organized into presence absence matrices. PAIRWISE AND MATRIX-LEVEL APROACHES Most methods for analysing species co-occurrence can be classified as either pairwise or matrix-level approaches (Table 1). The main difference is that the latter calculate a co-occurrence metric as a property of the entire presence absence matrix whereas the former examine co-occurrence species-by-species (Sfenthourakis et al., 2006; Pitta et al., 2012) to determine whether a particular pair of species is aggregated, segregated or random in occurrence (Gotelli & Ulrich, 2010). A recent approach developed by Arita et al. (2012) is best described as a hybrid given that it requires calculations performed on an entire matrix but the metric (proportional species richness) is obtained for each species separately (Table 1). Examples of matrix-level metrics include matrix temperature (Patterson & Atmar, 1986; Atmar & Patterson, 1993), number of unique species combinations (in the matrix), and the variance ratio (Schluter, 1984). Other metrics (e.g. C-score, number of checkerboards) can be calculated for species pairs but are typically aggregated (summed or averaged) for entire matrices. The most straightforward way to measure co-occurrence between two species is by the observed number of times that the two species co-occur relative to the expected number of times (Sanderson, 2000; Sfenthourakis et al., 2004, 2006; Veech, 2006, 2013; Pitta et al., 2012). Some authors have referred to this as the natural metric (Sfenthourakis et al., 2004, 2006; Pitta et al., 2012). The expected co-occurrence can be obtained through randomization of the matrix (randomizing species occurrences among sites) (Sanderson, 2000; Sfenthourakis et al., 2006) or through basic probability theory as J exp = (N 1 /N) 9 (N 2 /N) 9 N (Bowers & Brown, 1982; Table 1 Analytical methods previously used to study species co-occurrence (broadly defined). More detailed descriptions of each method can be found in the listed references (although these are not intended to represent a complete list). Method of Type of Description and usage Reference(s) Classic null models Nestedness Network Matrixlevel Rangediversity plots Causality Probability model Matrixlevel Matrixlevel Hybrid Pairwise Pairwise Broad class of null models that simultaneously randomize the occurrences of all species among sites. The models are distinguished by different constraints on the randomization algorithms. Used to find statistically significant co-occurrence patterns for entire presence absence matrices. Various algorithms (also sometimes used in classic null models) that randomize species occurrences among sites. Used to detect patterns of nestedness less species-rich sites as orderly compositional subsets of sites with greater species richness. Annealing algorithm is applied to a presence absence matrix (a geographical, taxonomic or guild-based matrix) so as to maximize modularity or the amount of connectedness among nodes (species). Used to find modules (groups of highly connected species) and compartments (spatial clustering of species range boundaries). Calculates and depicts proportional range richness of a species as the mean number of species occurring at the same sites as the focal species. Tests of significance can be conducted through various matrix randomization algorithms. Used to examine co-occurrence in the context of an entire assemblage and its geographical distribution (among other uses). Compares pairwise co-occurrence from two or more types of presence absence matrix. The geographical matrix is the traditional form where species are recorded from sampling sites; the ecological matrices consist of species assigned to units representing habitat types or other ecological factors. A fixed fixed algorithm randomizes species occurrences among units while conserving unit species richness and species incidences. For each matrix, species pairs are classified as positive, negative or no association. Used to examine the causal basis of co-occurrence and to identify the ecological factors that most influence co-occurrence. Applies pure probability-based equations to identify species pairs as having a positive, a negative, a random or an indecisive association. Used to analytically classify (without data randomization) species pairs to the above categories. Gotelli (2000), specifically Table 2 Patterson & Atmar (1986), Wright & Reeves (1992), Ulrich et al. (2009) Carstensen & Olesen (2009), Thebault (2013) Arita et al. (2012) Sfenthourakis et al. (2006) Veech (2013) 1030
3 Pairwise co-occurrence Veech, 2006, 2013; Araujo et al., 2011), where J exp represents the expected number of sites (or samples) that have species 1 and 2, N 1 is the number of sites that have species 1, N 2 is the number of sites that have species 2, and N is the total number of sites. Note that N 1 /N and N 2 /N are the marginal probabilities of occurrence of species 1 and 2, respectively, also known as their incidence rates. When one wishes to test for statistical significance of an observed co-occurrence metric (pairwise or matrix-level) then the presence absence matrix is often randomized to produce a null distribution of the metric as a basis for comparison (i.e. deriving a P-value). There are many algorithms for conducting the randomization and these vary in their Type I and II error rates (Gotelli, 2000). Perhaps, the most common and oft-recommended algorithm is the fixed fixed (F-F) randomization; the column and row sums of the matrix remain fixed at their original values during the randomization (Gotelli, 2000). That is, the randomization does not alter the species incidence rates (row sums) or richness of the sampling sites (column sums). Incidence rates and richness are implicitly assumed to be real ecological properties of the species assemblage that should not be altered when testing for non-random patterns of co-occurrence in the matrix or between species in a pair. The opposite of the F-F algorithm is the equiprobable equiprobable (E-E) randomization in which column and row sums are free to vary. All species are assumed to be equally probable in occurring at a given sampling site (differences in observed incidence rates are ignored during the randomization) and all sampling sites are assumed to be equally probable in having a given species (differences in observed richness among sites are ignored). The logic behind these different randomization algorithms has been thoroughly discussed elsewhere (Connor & Simberloff, 1979; Gotelli & Graves, 1996) and will not be repeated in this essay. However, my main point, relevant to this essay, is that no one randomization algorithm is more logical or ecologically realistic than any other; the algorithms primarily differ in their statistical properties (Gotelli, 2000; Ulrich & Gotelli, 2013). See Ulrich & Gotelli (2013) for a complete review and evaluation of 15 metrics used to measure species co-occurrence in conjunction with matrix randomization. When testing matrix-level co-occurrence patterns, it is obviously necessary to randomize the entire presence absence matrix in order to determine if the co-occurrence metric is significantly different from random. However, this need not be the case when using a pairwise approach. In its most basic form, the pairwise approach is solely concerned with testing pairs of species one-by-one as though there are S p number of 2 9 N matrices (species by sampling site) each examined separately, where S p could represent all possible species pairs, S p = N!/[2 9 (N 2)!], or some pre-selected group of pairs. Any of the randomization algorithms and matrix-level metrics could be applied to these 2 9 N matrices. However, a recently proposed pairwise method circumvents the need for randomization. PROBABILISTIC MODEL OF CO-OCCURRENCE The probabilistic model of species co-occurrence (Veech, 2013) is based on calculating the number of ways that two species can co-occur at exactly j number of sampling sites given that each species occurs at N 1 and N 2 number of sites out of a total of N. This quantity is the product of the following combinations: a = C(N, j), b = C(N j, N 2 j), c = C(N N 2, N 1 j), where a represents the number of unique arrangements of species 1 and 2 co-occurring at j number of sites among all sites, b is the number of ways that species 1 can be placed among the remaining sites that do not already have species 2, and c is the number of ways that species 2 can be placed among the remaining sites that do not already have species 1. The product, abc, is then divided by the number of unique ways that species 1 and 2 can be arranged among all sites without regard for j. This quantity is d 9 e, where d = C(N, N 2 ) and e = C(N, N 1 ). The probability, p j = abc/de, represents the probability that species 1 and 2 co-occur at exactly j number of sites. The probability, p j, is then calculated for all j satisfying the inequality: max{0, N 1 + N 2 N} j min{n 1, N 2 }. Finally, the quantity Σp j for all j < J obs represents the probability that the two species would co-occur at fewer that J obs sites (the number of sites where the two species actually co-occur). Likewise, Σp j for all j > J obs is the probability that the two species would co-occur at greater than J obs sites. Without requiring any randomization of the data, the probabilistic model analytically determines the probability that two species of a pair co-occur at an observed frequency (J obs ) greater than (P gt ) or less than (P lt ) the frequency expected if the two species were distributed randomly of each other (J exp ). Species pairs can be classified as positive, negative or random associations based upon the values of P gt (or P lt ) relative to a pre-defined significance level. The model has several desirable statistical properties including rigorous control of Type I and II error probabilities (Veech, 2013). The probabilistic model is essentially an analytical analogue of an F-E randomization algorithm in which species incidences are fixed but sampling sites are equiprobable in their chances of having a given species. (Granted, the researcher has control over which sites are included in the set of N and it is generally best practice not to include sites outside the species geographical range.) Therefore, the probabilistic model examines cooccurrence without regard for any constraints imposed by variation in species richness among sampling sites or variation in any other site characteristic (e.g. area). In this way, the probabilistic model implicitly recognizes the biological validity of incidence rates (which are fixed in the model) but does not subsume co-occurrence patterns under patterns of species richness. In fact, the contrary may be more realistic: co-occurrence patterns determine differences among sampling sites (ecological communities) in richness and composition and thus there is no reason that sites must be equiprobable in the probabilistic model. That is, site richness can be an unconstrained variable in the model if one views richness as the consequence of co-occurrence, instead of vice versa. Nonetheless, 1031
4 J. A. Veech the F-E randomization algorithm (in null models) is most appropriately applied to data from equal-sized sampling units (sites) of similar ecological conditions. In situations where sites vary in a meaningful environmental property, the equiprobable assumption might not be justified. I am working on a modification of the probabilistic model that will remove the assumption of equiprobable sites (see below). INDEPENDENCE OF SPECIES IN CO-OCCURRENCE ANALYSES Previous authors have correctly pointed out that the co-occurrence of two species in a pair may not be completely independent of a third species (Zaman & Simberloff, 2002; Sfenthourakis et al., 2006; Gotelli & Ulrich, 2010; Pitta et al., 2012). That is, if species x and y tend to positively co-occur while x and z also positively co-occur then species y and z might also be expected to co-occur positively (the same scenario can also be presented with negative co-occurrence). This non-independence could be problematic for interpreting pairwise co-occurrence patterns involving multiple species pairs (Zaman & Simberloff, 2002); for example, x and y may be positively and significantly associated with one another simply because each is significantly associated with z. However, this need not always be the case. Species x and y might co-occur at different sites than do x and z such that y and z never or rarely co-occur (Sfenthourakis et al., 2006). From the perspective of set theory, co-occurrence of x, y and z with each other (in pairs) is non-independent only when the intersection (number of co-occurrence sites) of x and y is large and intersection of x and z is large compared with each set x, y and z. In that case, intersection of y and z would also be large and the three species would co-occur at a large proportion of the total number of sites and the species pairs (xy, xz and yz) would not be independent. Researchers using a pairwise approach should be cognizant of such non-independence when tallying the number of positive and negative associations in a dataset. Of course, non-independence of species pairs might represent ecological reality. For example, positive co-occurrence should exist among two or more competing prey species and a predator species when there is predator-mediated coexistence. This describes the situation where competitively inferior prey species are able to coexist with a competitively dominant prey species whose abundance is limited by the predator (Paine, 1974; Caswell, 1978; Shurin & Allen, 2001). Non-random co-occurrence of three or more species (i.e. non-independence of species pairs) is a characteristic of a dataset more so than the method of. Indeed, some methods (e.g. network ) could be useful in identifying sets of closely linked species (see below). TESTABLE PREDICTIONS 1032 Several authors have recently suggested that the study of species co-occurrence could be improved conceptually and heuristically if researchers would specify a priori the co-occurrence patterns expected (predicted) in the data (Sfenthourakis et al., 2006; Gotelli & Ulrich, 2010; Veech, 2013). Historically, of co-occurrence has consisted primarily of post hoc interpretation; that is, testing for non-random pattern and then proposing an explanation instead of boldly and precisely predicting patterns based upon knowledge of ecological processes and principles. At the very least, researchers should be able to develop predictions based on species traits (e.g. habitat preferences, competitive ability, climatic tolerances) and site properties. The importance of prediction (i.e. formal hypothesis testing) is that it provides a more rigorous test of proposed explanation. A previous study by Sfenthourakis et al. (2006) is a step in the right direction. Although they did not make precise predictions for how specified species should be associated, they did propose and use an insightful framework to distinguish between ecological and geographical factors (processes). This framework involves compiling an ecological matrix in addition to the typical geographical matrix. The geographical matrix is a species 9 sampling site matrix. The ecological matrix replaces actual physical sampling locations with particular environmental variables or characteristics (e.g. low elevation, coniferous vegetation, calcareous substrate) that have been recorded for each site (see Table 4 in Sfenthourakis et al., 2006). For example, if species 1 was found at sampling sites that have coniferous vegetation then the column representing coniferous vegetation and the row representing species 1 is given a value of 1 to denote the species presence for that habitat variable. Applying a pairwise approach on both the geographical and ecological matrices allows a researcher to predict and interpret patterns of co-occurrence (see Table 1 in Sfenthourakis et al., 2006). For example, competitive exclusion should lead to two potentially competing species having a positive pairwise association in the ecological matrix (because they have similar niches) but a negative pairwise association in the geographical matrix (i.e. no syntopy anywhere) (Sfenthourakis et al., 2006). Much of the insight and benefit of the Sfenthourakis et al. (2006) framework is a result of the comprehensive characterization of environmental conditions at each site. They used an ecological matrix that had 41 environmental variables representing all types of factors that could be important to the distribution of the study organisms, terrestrial isopods. They also explicitly used a pairwise approach for analysing co-occurrence instead of a matrix-level approach. This further facilitated their ability to classify each species pair as a positive, negative or neutral association. In a somewhat analogous way, the probabilistic model can also be used to examine the relative roles of different environmental variables on species co-occurrence. The set of sampling sites (N X ) used in the can be specified so as to only include sites that are similar for a given variable, X. The association between two species is then examined for N X and classified as positive, negative or random along with calculating the difference between observed and expected co-occurrence, J obs J exp. The same steps are repeated for the set of remaining sites not in N X. Subsequent comparison of
5 Pairwise co-occurrence the results of the two analyses (and multiple analyses of this type) then may indicate which environmental factors are most responsible for the observed co-occurrence (or lack thereof). The environmental variables could even be species richness and the presence of a particular third species. Predictions about species co-occurrence can also be based on species traits, regardless of whether the uses information about site characteristics. It is relatively common to test for non-random co-occurrence patterns in subsets of species within the same taxonomic units (e.g. genera or families) to determine if evolutionary relatedness (and hence trait similarity) affects co-occurrence. Such tests rely on the notion that evolutionarily closely related species are more likely to be potential competitors (i.e. have relatively greater niche overlap) than are distantly related species (Elton, 1946; Denno et al., 1995; Losos, 2008; Valiente-Banuet & Verdu, 2008; Wiens et al., 2010; Ricklefs, 2011; Violle et al., 2011). However, researchers could go even further by identifying a priori particular traits that might mediate (or prevent) coexistence. Exact predictions could then be made for specific pairs based on their traits. Further, species traits (or at least trait environment interactions) can also be formally incorporated into the co-occurrence (e.g. Sfenthourakis et al., 2006). I am currently developing an extension of the probabilistic model that will allow users to specify a probability that a particular species could occur at a given site. In this way, N (total number of occupiable sites) is adjusted based on either species traits or environmental properties of the sites. The pairwise approach does not rely on the overly simplistic assumption that species affect one another only as pairs isolated from all other species. Processes such as predatormediated coexistence involve more than two species and hence the resultant co-occurrence pattern will reflect that. Pairwise analyses might still be used in those scenarios to test hypotheses. However, new methods such as network (Carstensen & Olesen, 2009; Araujo et al., 2011; Fontaine et al., 2011; Carstensen et al., 2012) that test for non-random clusters of species among sites (i.e. modules and compartments; Thebault, 2013) may be ideal for uncovering the cooccurrence patterns among multiple (more than two) species (Table 1). These methods primarily attempt to identify the clusters (modules) and compartments within a matrix in a way roughly analogous to how pairwise approaches attempt to identify the positive, negative and random species associations. These are essentially tests for structure within a matrix but not tests for any emergent property of a matrix (see next section). As such, network does not carry the hidden assumption that a presence absence matrix is a real ecological entity, unless of course an entire matrix is found to be one large module. PRESENCE ABSENCE MATRICES: NECESSITY OR CONVENIENCE? Is the species presence absence matrix a necessity for analysing co-occurrence and thinking about how species assemblages are geographically structured or is it simply a convenient way to organize and store data? The pairwise approach suggests the latter. If causative mechanisms can be tested and found by analysing species associations one-by-one then there is no need to think that the presence absence matrix has any meaningful ecological property (in and of itself). This argument also covers multispecies associations finding non-random clusters or modules of species within a matrix probably indicates that some ecological process is at work producing the association although it does not mean that the entire matrix itself has any emergent properties. Matrix-level metrics simply summarize the co-occurrence patterns (or lack thereof) that exist among the species; the metrics themselves are not measuring any kind of co-occurrence pattern that exists only at the matrix level. One way to see this is to consider that the value of a matrixlevel metric typically will not change much when a given species or site is removed. If some emergent property of the matrix was being measured, then even a very slight or small change in the matrix should lead to a dramatic change in the metric in which case, a presence absence matrix truly would be more than the sum of its parts. Although the study of cooccurrence and community assembly is historically rooted in the of presence absence matrices, there is no compelling reason to keep searching for nonrandom structure at the level of an entire matrix. CONCLUSIONS Using a pairwise approach, species interactions can be classified as positive, negative or random (neutral). Although, as with any statistical test, some species pairs might not be classifiable simply as a result of the test having low power (Sfenthourakis et al., 2006; Veech, 2013). Together these interactions represent all the possible ways that two species could associate in nature, and presumably each type of association represents the outcome of real ecological and evolutionary processes. Matrix-level metrics do not allow for classifying pairs as positive, negative or random associations. Certainly, in any species assemblage or set of communities, there may be meaningful associations (i.e. modules and compartments) that simultaneously involve multiple species. Nonetheless, this does not detract from using a pairwise approach. Indeed, a recent study using network showed that many species are only weakly linked to most other species in a network (Araujo et al., 2011). This suggests that multispecies interactions (e.g. diffuse competition) may not be relevant (see also Pitta et al., 2012). The main conceptual limitation on the of cooccurrence patterns is that we cannot directly infer the exact process(es) responsible for any particular pattern (Schluter, 1984). This is probably a more severe limitation when analyses are matrix-based than when they are based on species pairs. The pairwise approach is very amenable to hypothesistesting in that predictions about co-occurrence can be made a priori for paired species based on their traits and/or the properties of the sampling sites. 1033
6 J. A. Veech ACKNOWLEDGEMENTS I thank Yoni Gavish, Ivan Castro-Arellano, Dan McGarvey, Michael Huston, Dominique Gravel, Spyros Sfenthourakis, Werner Ulrich and an anonymous referee for providing insightful comments that helped me improve the manuscript. REFERENCES Almeido-Neto, M., Guimar~aes, P., Guimar~aes, P.R., Loyola, R.D. & Ulrich, W. (2008) A consistent metric for nestedness in ecological systems: reconciling concept and measurement. Oikos, 117, Araujo, M.B., Rozenfeld, A., Rahbek, C. & Marquet, P.A. (2011) Using species co-occurrence networks to assess the impacts of climate change. Ecography, 34, Arita, H.T., Christen, A., Rodrıguez, P. & Soberon, J. (2012) The presence absence matrix reloaded: the use and interpretation of range-diversity plots. Global Ecology and Biogeography, 21, Atmar, W. & Patterson, B.D. (1993) The measure of order and disorder in the distribution of species in fragmented habitat. Oecologia, 96, Bowers, M.A. & Brown, J.H. (1982) Body size and coexistence in desert rodents: chance or community structure? Ecology, 63, Carstensen, D.W. & Olesen, J.M. (2009) Wallacea and its nectarivorous birds: nestedness and modules. Journal of Biogeography, 36, Carstensen, D.W., Dalsgaard, B., Svenning, J.-C., Rahbek, C., Fjeldsa, J., Sutherland, W.J. & Olesen, J.M. (2012) Biogeographical modules and island roles: a comparison of Wallacea and the West Indies. Journal of Biogeography, 39, Caswell, H. (1978) Predator-mediated coexistence: non-equilibrium model. The American Naturalist, 112, Collins, M.D., Simberloff, D. & Connor, E.F. (2011) Binary matrices and checkerboard distributions of birds in the Bismarck Archipelago. Journal of Biogeography, 38, Connor, E.F. & Simberloff, D. (1979) The assembly of species communities: chance or competition? Ecology, 60, Denno, R.F., McClure, M.S. & Ott, J.R. (1995) Interspecific interactions in phytophagous insects: competition reexamined and resurrected. Annual Review of Entomology, 40, Diamond, J.M. (1973) Distributional ecology of New Guinea birds. Science, 179, Diamond, J.M. (1975) Assembly of species communities. Ecology and evolution of communities (ed. by M.L. Cody and J.M. Diamond), pp Harvard University Press, Cambridge, MA. Diamond, J.M. & Gilpin, M.E. (1982) Examination of the null model of Connor and Simberloff for species cooccurrences on islands. Oecologia, 52, Elton, C. (1946) Competition and the structure of ecological communities. Journal of Animal Ecology, 15, Fontaine, C., Guimar~aes, P.R., Kefi, S., Loeuille, N., Memmott, J., van der Putten, W.H., van Veen, F.J.F. & Thebault, E. (2011) The ecological and evolutionary implications of merging different types of networks. Ecology Letters, 11, Gilpin, M.E. & Diamond, J.M. (1984) Are species co-occurrences on islands non-random and are null hypotheses useful in community ecology? Ecological communities: conceptual issues and the evidence (ed. by D.R. Strong, D. Simberloff, L.G. Abele and A.B. Thistle), pp Princeton University Press, Princeton, NJ. Gotelli, N.J. (2000) Null model of species co-occurrence patterns. Ecology, 81, Gotelli, N.J. & Graves, G.R. (1996) Null models in ecology. Smithsonian Institution Press, Washington, DC. Gotelli, N.J. & Ulrich, W. (2010) The empirical Bayes approach as a tool to identify non-random species associations. Oecologia, 162, Losos, J.B. (2008) Phylogenetic niche conservatism, phylogenetic signal and the relationship between phylogenetic relatedness and ecological similarity among species. Ecology Letters, 11, Paine, R.T. (1974) Intertidal community structure: experimental studies on relationship between a dominant competitor and its principal predator. Oecologia, 15, Patterson, B.D. & Atmar, W. (1986) Nested subsets and the structure of insular mammalian faunas and archipelagos. Biological Journal of the Linnean Society, 28, Pielou, E.C. (1977) Mathematical ecology. Wiley Interscience, New York. Pitta, E., Giokas, S. & Sfenthourakis, S. (2012) Significant pairwise co-occurrence patterns are not the rule in the majority of biotic communities. Diversity, 4, Ricklefs, R.E. (2008) Disintegration of the ecological community. The American Naturalist, 172, Ricklefs, R.E. (2011) Applying a regional community concept to forest birds of eastern North America. Proceedings of the National Academy of Sciences USA, 108, Sanderson, J.G. (2000) Testing ecological patterns: a well-known algorithm from computer science aids the evaluation of species distributions. American Scientist, 88, Sanderson, J.G., Diamond, J.M. & Pimm, S.L. (2009) Pairwise co-existence of Bismarck and Solomon landbird species. Evolutionary Ecology Research, 11, Schluter, D. (1984) A variance test for detecting species associations with some example applications. Ecology, 65, Sfenthourakis, S., Giokas, S. & Tzanatos, E. (2004) From sampling stations to archipelagos: investigating aspects of the assemblage of insular biota. Global Ecology and Biogeography, 13, Sfenthourakis, S., Tzanatos, E. & Giokas, S. (2006) Species co-occurrence: the case of congeneric species and a causal
7 Pairwise co-occurrence approach to patterns of species association. Global Ecology and Biogeography, 15, Shurin, J.B. & Allen, E.G. (2001) Effects of competition, predation, and dispersal on species richness at local and regional scales. The American Naturalist, 158, Thebault, E. (2013) Identifying compartments in presence absence matrices and bipartite networks: insights into modularity measures. Journal of Biogeography, 40, Ulrich, W. & Gotelli, N.J. (2013) Pattern detection in null model. Oikos, 122, Ulrich, W., Almeida-Neto, M. & Gotelli, N.J. (2009) A consumer s guide to nestedness. Oikos, 118, Valiente-Banuet, A. & Verdu, M. (2008) Temporal shifts from facilitation to competition occur between closely related taxa. Journal of Ecology, 96, Veech, J.A. (2006) A probability-based of temporal and spatial co-occurrence in grassland birds. Journal of Biogeography, 33, Veech, J.A. (2013) A probabilistic model for analysing species co-occurrence. Global Ecology and Biogeography, 22, Violle, C., Nemergut, D.R., Pu, Z. & Jiang, L. (2011) Phylogenetic limiting similarity and competitive exclusion. Ecology Letters, 14, Wiens, J.J., Ackerly, D.D., Allen, A.P., Anacker, B.L., Buckley, L.B., Cornell, H.V., Damschen, E.I., Davies, T.J., Grytnes, J.A., Harrison, S.P., Hawkins, B.A., Holt, R.D., McCain, C.M. & Stephens, P.R. (2010) Niche conservatism as an emerging principle in ecology and conservation biology. Ecology Letters, 13, Wright, D.H. & Reeves, J.H. (1992) On the meaning and measurement of nestedness of species assemblages. Oecologia, 92, Zaman, A. & Simberloff, D. (2002) Random binary matrices in biogeographical ecology instituting a good neighbor policy. Environmental and Ecological Statistics, 9, BIOSKETCH Joseph Veech is broadly interested in the ecological and evolutionary factors that affect species distribution and abundance at a wide range of spatial scales. He also works to develop new statistical methods for the testing of pattern and evaluates previously proposed methods. Most recently, he has taken an interest in promoting the use of probability theory in ecological research. Editor: Miguel Araujo 1035
Pairs a FORTRAN program for studying pair wise species associations in ecological matrices Version 1.0
 Pairs 1 Pairs a FORTRAN program for studying pair wise species associations in ecological matrices Version 1.0 Werner Ulrich Nicolaus Copernicus University in Toruń Department of Animal Ecology Gagarina
Pairs 1 Pairs a FORTRAN program for studying pair wise species associations in ecological matrices Version 1.0 Werner Ulrich Nicolaus Copernicus University in Toruń Department of Animal Ecology Gagarina
Species co-occurrences and neutral models: reassessing J. M. Diamond s assembly rules
 OIKOS 107: 603/609, 2004 Species co-occurrences and neutral models: reassessing J. M. Diamond s assembly rules Werner Ulrich Ulrich, W. 2004. Species co-occurrences and neutral models: reassessing J. M.
OIKOS 107: 603/609, 2004 Species co-occurrences and neutral models: reassessing J. M. Diamond s assembly rules Werner Ulrich Ulrich, W. 2004. Species co-occurrences and neutral models: reassessing J. M.
"PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200 Spring 2014 University of California, Berkeley
 "PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200 Spring 2014 University of California, Berkeley D.D. Ackerly April 16, 2014. Community Ecology and Phylogenetics Readings: Cavender-Bares,
"PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200 Spring 2014 University of California, Berkeley D.D. Ackerly April 16, 2014. Community Ecology and Phylogenetics Readings: Cavender-Bares,
Diversity partitioning without statistical independence of alpha and beta
 1964 Ecology, Vol. 91, No. 7 Ecology, 91(7), 2010, pp. 1964 1969 Ó 2010 by the Ecological Society of America Diversity partitioning without statistical independence of alpha and beta JOSEPH A. VEECH 1,3
1964 Ecology, Vol. 91, No. 7 Ecology, 91(7), 2010, pp. 1964 1969 Ó 2010 by the Ecological Society of America Diversity partitioning without statistical independence of alpha and beta JOSEPH A. VEECH 1,3
EFFECTS OF TAXONOMIC GROUPS AND GEOGRAPHIC SCALE ON PATTERNS OF NESTEDNESS
 EFFECTS OF TAXONOMIC GROUPS AND GEOGRAPHIC SCALE ON PATTERNS OF NESTEDNESS SFENTHOURAKIS Spyros, GIOKAS Sinos & LEGAKIS Anastasios Zoological Museum, Department of Biology, University of Athens, Greece
EFFECTS OF TAXONOMIC GROUPS AND GEOGRAPHIC SCALE ON PATTERNS OF NESTEDNESS SFENTHOURAKIS Spyros, GIOKAS Sinos & LEGAKIS Anastasios Zoological Museum, Department of Biology, University of Athens, Greece
ecological area-network relations: methodology Christopher Moore cs765: complex networks 16 November 2011
 ecological area-network relations: methodology Christopher Moore cs765: complex networks 16 November 2011 ecology: the study of the spatial and temporal patterns of the distribution and abundance of organisms,
ecological area-network relations: methodology Christopher Moore cs765: complex networks 16 November 2011 ecology: the study of the spatial and temporal patterns of the distribution and abundance of organisms,
Species Co-occurrence, an Annotated Bibliography
 I have listed below a set of papers describing research in the area of using species lists from a set of sites (primarily islands in an archipelago). The list is primarily constructed of papers that I
I have listed below a set of papers describing research in the area of using species lists from a set of sites (primarily islands in an archipelago). The list is primarily constructed of papers that I
SPECIES CO-OCCURRENCE: A META-ANALYSIS OF J. M. DIAMOND S ASSEMBLY RULES MODEL
 Ecology, 83(8), 2002, pp. 2091 2096 2002 by the Ecological Society of America SPECIES CO-OCCURRENCE: A META-ANALYSIS OF J. M. DIAMOND S ASSEMBLY RULES MODEL NICHOLAS J. GOTELLI 1 AND DECLAN J. MCCABE 2
Ecology, 83(8), 2002, pp. 2091 2096 2002 by the Ecological Society of America SPECIES CO-OCCURRENCE: A META-ANALYSIS OF J. M. DIAMOND S ASSEMBLY RULES MODEL NICHOLAS J. GOTELLI 1 AND DECLAN J. MCCABE 2
Community phylogenetics review/quiz
 Community phylogenetics review/quiz A. This pattern represents and is a consequent of. Most likely to observe this at phylogenetic scales. B. This pattern represents and is a consequent of. Most likely
Community phylogenetics review/quiz A. This pattern represents and is a consequent of. Most likely to observe this at phylogenetic scales. B. This pattern represents and is a consequent of. Most likely
BIOS 6150: Ecology Dr. Stephen Malcolm, Department of Biological Sciences
 BIOS 6150: Ecology Dr. Stephen Malcolm, Department of Biological Sciences Week 14: Roles of competition, predation & disturbance in community structure. Lecture summary: (A) Competition: Pattern vs process.
BIOS 6150: Ecology Dr. Stephen Malcolm, Department of Biological Sciences Week 14: Roles of competition, predation & disturbance in community structure. Lecture summary: (A) Competition: Pattern vs process.
Rank-abundance. Geometric series: found in very communities such as the
 Rank-abundance Geometric series: found in very communities such as the Log series: group of species that occur _ time are the most frequent. Useful for calculating a diversity metric (Fisher s alpha) Most
Rank-abundance Geometric series: found in very communities such as the Log series: group of species that occur _ time are the most frequent. Useful for calculating a diversity metric (Fisher s alpha) Most
Transitivity a FORTRAN program for the analysis of bivariate competitive interactions Version 1.1
 Transitivity 1 Transitivity a FORTRAN program for the analysis of bivariate competitive interactions Version 1.1 Werner Ulrich Nicolaus Copernicus University in Toruń Chair of Ecology and Biogeography
Transitivity 1 Transitivity a FORTRAN program for the analysis of bivariate competitive interactions Version 1.1 Werner Ulrich Nicolaus Copernicus University in Toruń Chair of Ecology and Biogeography
Gary G. Mittelbach Michigan State University
 Community Ecology Gary G. Mittelbach Michigan State University Sinauer Associates, Inc. Publishers Sunderland, Massachusetts U.S.A. Brief Table of Contents 1 Community Ecology s Roots 1 PART I The Big
Community Ecology Gary G. Mittelbach Michigan State University Sinauer Associates, Inc. Publishers Sunderland, Massachusetts U.S.A. Brief Table of Contents 1 Community Ecology s Roots 1 PART I The Big
The tangled link between b- and c-diversity: a Narcissus effect weakens statistical inferences in null model analyses of diversity patterns
 Global Ecology and Biogeography, (Global Ecol. Biogeogr.) (2017) 26, 1 5 ECOLOGICAL SOUNDINGS The tangled link between b- and c-diversity: a Narcissus effect weakens statistical inferences in null model
Global Ecology and Biogeography, (Global Ecol. Biogeogr.) (2017) 26, 1 5 ECOLOGICAL SOUNDINGS The tangled link between b- and c-diversity: a Narcissus effect weakens statistical inferences in null model
Welcome! Text: Community Ecology by Peter J. Morin, Blackwell Science ISBN (required) Topics covered: Date Topic Reading
 Welcome! Text: Community Ecology by Peter J. Morin, Blackwell Science ISBN 0-86542-350-4 (required) Topics covered: Date Topic Reading 1 Sept Syllabus, project, Ch1, Ch2 Communities 8 Sept Competition
Welcome! Text: Community Ecology by Peter J. Morin, Blackwell Science ISBN 0-86542-350-4 (required) Topics covered: Date Topic Reading 1 Sept Syllabus, project, Ch1, Ch2 Communities 8 Sept Competition
Integrative Biology 200 "PRINCIPLES OF PHYLOGENETICS" Spring 2018 University of California, Berkeley
 Integrative Biology 200 "PRINCIPLES OF PHYLOGENETICS" Spring 2018 University of California, Berkeley B.D. Mishler Feb. 14, 2018. Phylogenetic trees VI: Dating in the 21st century: clocks, & calibrations;
Integrative Biology 200 "PRINCIPLES OF PHYLOGENETICS" Spring 2018 University of California, Berkeley B.D. Mishler Feb. 14, 2018. Phylogenetic trees VI: Dating in the 21st century: clocks, & calibrations;
of a landscape to support biodiversity and ecosystem processes and provide ecosystem services in face of various disturbances.
 L LANDSCAPE ECOLOGY JIANGUO WU Arizona State University Spatial heterogeneity is ubiquitous in all ecological systems, underlining the significance of the pattern process relationship and the scale of
L LANDSCAPE ECOLOGY JIANGUO WU Arizona State University Spatial heterogeneity is ubiquitous in all ecological systems, underlining the significance of the pattern process relationship and the scale of
ENMTools: a toolbox for comparative studies of environmental niche models
 Ecography 33: 607611, 2010 doi: 10.1111/j.1600-0587.2009.06142.x # 2010 The Authors. Journal compilation # 2010 Ecography Subject Editor: Thiago Rangel. Accepted 4 September 2009 ENMTools: a toolbox for
Ecography 33: 607611, 2010 doi: 10.1111/j.1600-0587.2009.06142.x # 2010 The Authors. Journal compilation # 2010 Ecography Subject Editor: Thiago Rangel. Accepted 4 September 2009 ENMTools: a toolbox for
Community Structure. Community An assemblage of all the populations interacting in an area
 Community Structure Community An assemblage of all the populations interacting in an area Community Ecology The ecological community is the set of plant and animal species that occupy an area Questions
Community Structure Community An assemblage of all the populations interacting in an area Community Ecology The ecological community is the set of plant and animal species that occupy an area Questions
Niche The sum of all interactions a species has with biotic/abiotic components of the environment N-dimensional hypervolume
 Niche The sum of all interactions a species has with biotic/abiotic components of the environment N-dimensional hypervolume Each dimension is a biotic or abiotic resource Ecomorphology Ecology (niche)
Niche The sum of all interactions a species has with biotic/abiotic components of the environment N-dimensional hypervolume Each dimension is a biotic or abiotic resource Ecomorphology Ecology (niche)
Non-independence in Statistical Tests for Discrete Cross-species Data
 J. theor. Biol. (1997) 188, 507514 Non-independence in Statistical Tests for Discrete Cross-species Data ALAN GRAFEN* AND MARK RIDLEY * St. John s College, Oxford OX1 3JP, and the Department of Zoology,
J. theor. Biol. (1997) 188, 507514 Non-independence in Statistical Tests for Discrete Cross-species Data ALAN GRAFEN* AND MARK RIDLEY * St. John s College, Oxford OX1 3JP, and the Department of Zoology,
ISLAND BIOGEOGRAPHY Lab 7
 Reminders! Bring memory stick Read papers for Discussion Key Concepts Biogeography/Island biogeography Convergent evolution Dynamic equilibrium Student Learning Outcomes After Lab 7 students will be able
Reminders! Bring memory stick Read papers for Discussion Key Concepts Biogeography/Island biogeography Convergent evolution Dynamic equilibrium Student Learning Outcomes After Lab 7 students will be able
Island biogeography. Key concepts. Introduction. Island biogeography theory. Colonization-extinction balance. Island-biogeography theory
 Island biogeography Key concepts Colonization-extinction balance Island-biogeography theory Introduction At the end of the last chapter, it was suggested that another mechanism for the maintenance of α-diversity
Island biogeography Key concepts Colonization-extinction balance Island-biogeography theory Introduction At the end of the last chapter, it was suggested that another mechanism for the maintenance of α-diversity
Disentangling spatial structure in ecological communities. Dan McGlinn & Allen Hurlbert.
 Disentangling spatial structure in ecological communities Dan McGlinn & Allen Hurlbert http://mcglinn.web.unc.edu daniel.mcglinn@usu.edu The Unified Theories of Biodiversity 6 unified theories of diversity
Disentangling spatial structure in ecological communities Dan McGlinn & Allen Hurlbert http://mcglinn.web.unc.edu daniel.mcglinn@usu.edu The Unified Theories of Biodiversity 6 unified theories of diversity
ANALYSIS OF CHARACTER DIVERGENCE ALONG ENVIRONMENTAL GRADIENTS AND OTHER COVARIATES
 ORIGINAL ARTICLE doi:10.1111/j.1558-5646.2007.00063.x ANALYSIS OF CHARACTER DIVERGENCE ALONG ENVIRONMENTAL GRADIENTS AND OTHER COVARIATES Dean C. Adams 1,2,3 and Michael L. Collyer 1,4 1 Department of
ORIGINAL ARTICLE doi:10.1111/j.1558-5646.2007.00063.x ANALYSIS OF CHARACTER DIVERGENCE ALONG ENVIRONMENTAL GRADIENTS AND OTHER COVARIATES Dean C. Adams 1,2,3 and Michael L. Collyer 1,4 1 Department of
"PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200 Spring 2018 University of California, Berkeley
 "PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200 Spring 2018 University of California, Berkeley D.D. Ackerly Feb. 26, 2018 Maximum Likelihood Principles, and Applications to
"PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200 Spring 2018 University of California, Berkeley D.D. Ackerly Feb. 26, 2018 Maximum Likelihood Principles, and Applications to
Package MBI. February 19, 2015
 Type Package Package MBI February 19, 2015 Title (M)ultiple-site (B)iodiversity (I)ndices Calculator Version 1.0 Date 2012-10-17 Author Youhua Chen Maintainer Over 20 multiple-site diversity indices can
Type Package Package MBI February 19, 2015 Title (M)ultiple-site (B)iodiversity (I)ndices Calculator Version 1.0 Date 2012-10-17 Author Youhua Chen Maintainer Over 20 multiple-site diversity indices can
Name Student ID. Good luck and impress us with your toolkit of ecological knowledge and concepts!
 Page 1 BIOLOGY 150 Final Exam Winter Quarter 2000 Before starting be sure to put your name and student number on the top of each page. MINUS 3 POINTS IF YOU DO NOT WRITE YOUR NAME ON EACH PAGE! You have
Page 1 BIOLOGY 150 Final Exam Winter Quarter 2000 Before starting be sure to put your name and student number on the top of each page. MINUS 3 POINTS IF YOU DO NOT WRITE YOUR NAME ON EACH PAGE! You have
On nestedness analyses: rethinking matrix temperature and anti-nestedness
 Oikos 000: 000 000, 2007 doi: 10.1111/j.2007.0030-1299.15803.x, Copyright # Oikos 2007, ISSN 0030-1299 Subject Editor: Jordi Bascompte, Accepted 27 December 2006 On nestedness analyses: rethinking matrix
Oikos 000: 000 000, 2007 doi: 10.1111/j.2007.0030-1299.15803.x, Copyright # Oikos 2007, ISSN 0030-1299 Subject Editor: Jordi Bascompte, Accepted 27 December 2006 On nestedness analyses: rethinking matrix
BIO S380T Page 1 Summer 2005: Exam 2
 BIO S380T Page 1 Part I: Definitions. [5 points for each term] For each term, provide a brief definition that also indicates why the term is important in ecology or evolutionary biology. Where I ve provided
BIO S380T Page 1 Part I: Definitions. [5 points for each term] For each term, provide a brief definition that also indicates why the term is important in ecology or evolutionary biology. Where I ve provided
The role of stochastic processes in producing nested patterns of species distributions
 OIKOS 114: 159167, 26 The role of stochastic processes in producing nested patterns of species distributions Christopher L. Higgins, Michael R. Willig and Richard E. Strauss Higgins, C. L., Willig, M.
OIKOS 114: 159167, 26 The role of stochastic processes in producing nested patterns of species distributions Christopher L. Higgins, Michael R. Willig and Richard E. Strauss Higgins, C. L., Willig, M.
Parameter Sensitivity In A Lattice Ecosystem With Intraguild Predation
 Parameter Sensitivity In A Lattice Ecosystem With Intraguild Predation N. Nakagiri a, K. Tainaka a, T. Togashi b, T. Miyazaki b and J. Yoshimura a a Department of Systems Engineering, Shizuoka University,
Parameter Sensitivity In A Lattice Ecosystem With Intraguild Predation N. Nakagiri a, K. Tainaka a, T. Togashi b, T. Miyazaki b and J. Yoshimura a a Department of Systems Engineering, Shizuoka University,
POPULATIONS and COMMUNITIES
 POPULATIONS and COMMUNITIES Ecology is the study of organisms and the nonliving world they inhabit. Central to ecology is the complex set of interactions between organisms, both intraspecific (between
POPULATIONS and COMMUNITIES Ecology is the study of organisms and the nonliving world they inhabit. Central to ecology is the complex set of interactions between organisms, both intraspecific (between
Performance Comparison of K-Means and Expectation Maximization with Gaussian Mixture Models for Clustering EE6540 Final Project
 Performance Comparison of K-Means and Expectation Maximization with Gaussian Mixture Models for Clustering EE6540 Final Project Devin Cornell & Sushruth Sastry May 2015 1 Abstract In this article, we explore
Performance Comparison of K-Means and Expectation Maximization with Gaussian Mixture Models for Clustering EE6540 Final Project Devin Cornell & Sushruth Sastry May 2015 1 Abstract In this article, we explore
Interspecific Competition
 Interspecific Competition Intraspecific competition Classic logistic model Interspecific extension of densitydependence Individuals of other species may also have an effect on per capita birth & death
Interspecific Competition Intraspecific competition Classic logistic model Interspecific extension of densitydependence Individuals of other species may also have an effect on per capita birth & death
Current controversies in Marine Ecology with an emphasis on Coral reef systems
 Current controversies in Marine Ecology with an emphasis on Coral reef systems Open vs closed populations (already discussed) The extent and importance of larval dispersal Maintenance of Diversity Equilibrial
Current controversies in Marine Ecology with an emphasis on Coral reef systems Open vs closed populations (already discussed) The extent and importance of larval dispersal Maintenance of Diversity Equilibrial
Multiple regression and inference in ecology and conservation biology: further comments on identifying important predictor variables
 Biodiversity and Conservation 11: 1397 1401, 2002. 2002 Kluwer Academic Publishers. Printed in the Netherlands. Multiple regression and inference in ecology and conservation biology: further comments on
Biodiversity and Conservation 11: 1397 1401, 2002. 2002 Kluwer Academic Publishers. Printed in the Netherlands. Multiple regression and inference in ecology and conservation biology: further comments on
Metacommunities Spatial Ecology of Communities
 Spatial Ecology of Communities Four perspectives for multiple species Patch dynamics principles of metapopulation models (patchy pops, Levins) Mass effects principles of source-sink and rescue effects
Spatial Ecology of Communities Four perspectives for multiple species Patch dynamics principles of metapopulation models (patchy pops, Levins) Mass effects principles of source-sink and rescue effects
Chapter 6 Reading Questions
 Chapter 6 Reading Questions 1. Fill in 5 key events in the re-establishment of the New England forest in the Opening Story: 1. Farmers begin leaving 2. 3. 4. 5. 6. 7. Broadleaf forest reestablished 2.
Chapter 6 Reading Questions 1. Fill in 5 key events in the re-establishment of the New England forest in the Opening Story: 1. Farmers begin leaving 2. 3. 4. 5. 6. 7. Broadleaf forest reestablished 2.
A consumer s guide to nestedness analysis
 Oikos 118: 317, 2009 doi: 10.1111/j.1600-0706.2008.17053.x, # 2008 The Authors. Journal compilation # 2008 Oikos Subject Editor: Jennifer Dunne. Accepted 25 July 2008 A consumer s guide to nestedness analysis
Oikos 118: 317, 2009 doi: 10.1111/j.1600-0706.2008.17053.x, # 2008 The Authors. Journal compilation # 2008 Oikos Subject Editor: Jennifer Dunne. Accepted 25 July 2008 A consumer s guide to nestedness analysis
Development Team. Department of Zoology, University of Delhi. Department of Zoology, University of Delhi
 Paper No. : 12 Module : 18 diversity index, abundance, species richness, vertical and horizontal Development Team Principal Investigator: Co-Principal Investigator: Paper Coordinator: Content Writer: Content
Paper No. : 12 Module : 18 diversity index, abundance, species richness, vertical and horizontal Development Team Principal Investigator: Co-Principal Investigator: Paper Coordinator: Content Writer: Content
"PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200B Spring 2011 University of California, Berkeley
 "PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200B Spring 2011 University of California, Berkeley B.D. Mishler Feb. 1, 2011. Qualitative character evolution (cont.) - comparing
"PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200B Spring 2011 University of California, Berkeley B.D. Mishler Feb. 1, 2011. Qualitative character evolution (cont.) - comparing
Phylogenetic beta diversity: linking ecological and evolutionary processes across space in time
 Ecology Letters, (2008) 11: xxx xxx doi: 10.1111/j.1461-0248.2008.01256.x IDEA AND PERSPECTIVE Phylogenetic beta diversity: linking ecological and evolutionary processes across space in time Catherine
Ecology Letters, (2008) 11: xxx xxx doi: 10.1111/j.1461-0248.2008.01256.x IDEA AND PERSPECTIVE Phylogenetic beta diversity: linking ecological and evolutionary processes across space in time Catherine
Predicting the relationship between local and regional species richness from a patch occupancy dynamics model
 Ecology 2000, 69, Predicting the relationship between local and regional species richness from a patch occupancy dynamics model B. HUGUENY* and H.V. CORNELL{ *ORSTOM, Laboratoire d'ecologie des eaux douces,
Ecology 2000, 69, Predicting the relationship between local and regional species richness from a patch occupancy dynamics model B. HUGUENY* and H.V. CORNELL{ *ORSTOM, Laboratoire d'ecologie des eaux douces,
Current controversies in Marine Ecology with an emphasis on Coral reef systems. Niche Diversification Hypothesis Assumptions:
 Current controversies in Marine Ecology with an emphasis on Coral reef systems Open vs closed populations (already Discussed) The extent and importance of larval dispersal Maintenance of Diversity Equilibrial
Current controversies in Marine Ecology with an emphasis on Coral reef systems Open vs closed populations (already Discussed) The extent and importance of larval dispersal Maintenance of Diversity Equilibrial
Evidence for Competition
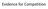 Evidence for Competition Population growth in laboratory experiments carried out by the Russian scientist Gause on growth rates in two different yeast species Each of the species has the same food e.g.,
Evidence for Competition Population growth in laboratory experiments carried out by the Russian scientist Gause on growth rates in two different yeast species Each of the species has the same food e.g.,
NGSS Example Bundles. Page 1 of 23
 High School Conceptual Progressions Model III Bundle 2 Evolution of Life This is the second bundle of the High School Conceptual Progressions Model Course III. Each bundle has connections to the other
High School Conceptual Progressions Model III Bundle 2 Evolution of Life This is the second bundle of the High School Conceptual Progressions Model Course III. Each bundle has connections to the other
Historical contingency, niche conservatism and the tendency for some taxa to be more diverse towards the poles
 Electronic Supplementary Material Historical contingency, niche conservatism and the tendency for some taxa to be more diverse towards the poles Ignacio Morales-Castilla 1,2 *, Jonathan T. Davies 3 and
Electronic Supplementary Material Historical contingency, niche conservatism and the tendency for some taxa to be more diverse towards the poles Ignacio Morales-Castilla 1,2 *, Jonathan T. Davies 3 and
THE CONSEQUENCES OF GENETIC DIVERSITY IN COMPETITIVE COMMUNITIES MARK VELLEND 1
 Ecology, 87(2), 2006, pp. 304 311 2006 by the Ecological Society of America THE CONSEQUENCES OF GENETIC DIVERSITY IN COMPETITIVE COMMUNITIES MARK VELLEND 1 National Center for Ecological Analysis and Synthesis,
Ecology, 87(2), 2006, pp. 304 311 2006 by the Ecological Society of America THE CONSEQUENCES OF GENETIC DIVERSITY IN COMPETITIVE COMMUNITIES MARK VELLEND 1 National Center for Ecological Analysis and Synthesis,
ANOVA approach. Investigates interaction terms. Disadvantages: Requires careful sampling design with replication
 ANOVA approach Advantages: Ideal for evaluating hypotheses Ideal to quantify effect size (e.g., differences between groups) Address multiple factors at once Investigates interaction terms Disadvantages:
ANOVA approach Advantages: Ideal for evaluating hypotheses Ideal to quantify effect size (e.g., differences between groups) Address multiple factors at once Investigates interaction terms Disadvantages:
Aggregations on larger scales. Metapopulation. Definition: A group of interconnected subpopulations Sources and Sinks
 Aggregations on larger scales. Metapopulation Definition: A group of interconnected subpopulations Sources and Sinks Metapopulation - interconnected group of subpopulations sink source McKillup and McKillup
Aggregations on larger scales. Metapopulation Definition: A group of interconnected subpopulations Sources and Sinks Metapopulation - interconnected group of subpopulations sink source McKillup and McKillup
On Likelihoodism and Intelligent Design
 On Likelihoodism and Intelligent Design Sebastian Lutz Draft: 2011 02 14 Abstract Two common and plausible claims in the philosophy of science are that (i) a theory that makes no predictions is not testable
On Likelihoodism and Intelligent Design Sebastian Lutz Draft: 2011 02 14 Abstract Two common and plausible claims in the philosophy of science are that (i) a theory that makes no predictions is not testable
Enduring understanding 1.A: Change in the genetic makeup of a population over time is evolution.
 The AP Biology course is designed to enable you to develop advanced inquiry and reasoning skills, such as designing a plan for collecting data, analyzing data, applying mathematical routines, and connecting
The AP Biology course is designed to enable you to develop advanced inquiry and reasoning skills, such as designing a plan for collecting data, analyzing data, applying mathematical routines, and connecting
A consistent metric for nestedness analysis in ecological systems: reconciling concept and measurement
 Oikos 7: 739, doi:./j..3-99.66.x, # The Authors. Journal compilation # Oikos Subject Editor: Ulrich Brose. Accepted March A consistent metric for nestedness analysis in ecological systems: reconciling
Oikos 7: 739, doi:./j..3-99.66.x, # The Authors. Journal compilation # Oikos Subject Editor: Ulrich Brose. Accepted March A consistent metric for nestedness analysis in ecological systems: reconciling
Effects of trophic similarity on community composition
 Ecology Letters, (2014) 17: 1495 1506 doi: 10.1111/ele.12356 IDEA AND PERSPECTIVE Effects of trophic similarity on community composition Helene Morlon, 1,2 * Sonia Kefi 3 and Neo D. Martinez 4,5 Abstract
Ecology Letters, (2014) 17: 1495 1506 doi: 10.1111/ele.12356 IDEA AND PERSPECTIVE Effects of trophic similarity on community composition Helene Morlon, 1,2 * Sonia Kefi 3 and Neo D. Martinez 4,5 Abstract
Competition: Observations and Experiments. Cedar Creek MN, copyright David Tilman
 Competition: Observations and Experiments Cedar Creek MN, copyright David Tilman Resource-Ratio (R*) Theory Species differ in critical limiting concentration for resources (R* values) R* values differ
Competition: Observations and Experiments Cedar Creek MN, copyright David Tilman Resource-Ratio (R*) Theory Species differ in critical limiting concentration for resources (R* values) R* values differ
Module 4: Community structure and assembly
 Module 4: Community structure and assembly Class Topic Reading(s) Day 1 (Thu Intro, definitions, some history. Messing Nov 2) around with a simple dataset in R. Day 2 (Tue Nov 7) Day 3 (Thu Nov 9) Day
Module 4: Community structure and assembly Class Topic Reading(s) Day 1 (Thu Intro, definitions, some history. Messing Nov 2) around with a simple dataset in R. Day 2 (Tue Nov 7) Day 3 (Thu Nov 9) Day
Ecology 203, Exam III. November 16, Print name:
 Ecology 203, Exam III. November 16, 2005. Print name: Read carefully. Work accurately and efficiently. The exam is worth 100 points (plus 6 extra credit points). Choose four of ten concept-exploring questions
Ecology 203, Exam III. November 16, 2005. Print name: Read carefully. Work accurately and efficiently. The exam is worth 100 points (plus 6 extra credit points). Choose four of ten concept-exploring questions
APPENDIX DR1: METHODOLOGICAL DETAILS AND INFORMATION
 GSA DATA REPOSITORY 2018261 Klompmaker and Finnegan APPENDIX DR1: METHODOLOGICAL DETAILS AND INFORMATION METHODS Fossil datasets (Appendix DR2) were downloaded from the Paleobiology Database (PBDB: https://paleobiodb.org)
GSA DATA REPOSITORY 2018261 Klompmaker and Finnegan APPENDIX DR1: METHODOLOGICAL DETAILS AND INFORMATION METHODS Fossil datasets (Appendix DR2) were downloaded from the Paleobiology Database (PBDB: https://paleobiodb.org)
Age (x) nx lx. Population dynamics Population size through time should be predictable N t+1 = N t + B + I - D - E
 Population dynamics Population size through time should be predictable N t+1 = N t + B + I - D - E Time 1 N = 100 20 births 25 deaths 10 immigrants 15 emmigrants Time 2 100 + 20 +10 25 15 = 90 Life History
Population dynamics Population size through time should be predictable N t+1 = N t + B + I - D - E Time 1 N = 100 20 births 25 deaths 10 immigrants 15 emmigrants Time 2 100 + 20 +10 25 15 = 90 Life History
Oecologia. Competitive exclusion, or species aggregation? An aid in deciding. Oecologia (1992) 91: Springer-Verlag 1992
 Oecologia (1992) 91:419 424 Oecologia 9 Springer-Verlag 1992 Competitive exclusion, or species aggregation? An aid in deciding Lewi Stone 1 and Alan Roberts 2 1 Australian Environmental Studies, Griffith
Oecologia (1992) 91:419 424 Oecologia 9 Springer-Verlag 1992 Competitive exclusion, or species aggregation? An aid in deciding Lewi Stone 1 and Alan Roberts 2 1 Australian Environmental Studies, Griffith
Weather is the day-to-day condition of Earth s atmosphere.
 4.1 Climate Weather and Climate Weather is the day-to-day condition of Earth s atmosphere. Climate refers to average conditions over long periods and is defined by year-after-year patterns of temperature
4.1 Climate Weather and Climate Weather is the day-to-day condition of Earth s atmosphere. Climate refers to average conditions over long periods and is defined by year-after-year patterns of temperature
Community Structure Temporal Patterns
 Community Structure Temporal Patterns Temporal Patterns Seasonality Phenology study of repeated patterns in time and their relationship to physical aspects of the environment Seasonal changes that are
Community Structure Temporal Patterns Temporal Patterns Seasonality Phenology study of repeated patterns in time and their relationship to physical aspects of the environment Seasonal changes that are
Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut
 Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic analysis Phylogenetic Basics: Biological
Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic analysis Phylogenetic Basics: Biological
NICHE BREADTH AND RESOURCE PARTIONING
 22 NICHE BREADTH AND RESOURCE PARTIONING Objectives Compute niche breadth for two organisms coexisting in a community. Compute niche overlap for the two coexisting organisms. Use the Solver function to
22 NICHE BREADTH AND RESOURCE PARTIONING Objectives Compute niche breadth for two organisms coexisting in a community. Compute niche overlap for the two coexisting organisms. Use the Solver function to
What is the range of a taxon? A scaling problem at three levels: Spa9al scale Phylogene9c depth Time
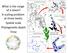 What is the range of a taxon? A scaling problem at three levels: Spa9al scale Phylogene9c depth Time 1 5 0.25 0.15 5 0.05 0.05 0.10 2 0.10 0.10 0.20 4 Reminder of what a range-weighted tree is Actual Tree
What is the range of a taxon? A scaling problem at three levels: Spa9al scale Phylogene9c depth Time 1 5 0.25 0.15 5 0.05 0.05 0.10 2 0.10 0.10 0.20 4 Reminder of what a range-weighted tree is Actual Tree
The Tempo of Macroevolution: Patterns of Diversification and Extinction
 The Tempo of Macroevolution: Patterns of Diversification and Extinction During the semester we have been consider various aspects parameters associated with biodiversity. Current usage stems from 1980's
The Tempo of Macroevolution: Patterns of Diversification and Extinction During the semester we have been consider various aspects parameters associated with biodiversity. Current usage stems from 1980's
Computational Aspects of Aggregation in Biological Systems
 Computational Aspects of Aggregation in Biological Systems Vladik Kreinovich and Max Shpak University of Texas at El Paso, El Paso, TX 79968, USA vladik@utep.edu, mshpak@utep.edu Summary. Many biologically
Computational Aspects of Aggregation in Biological Systems Vladik Kreinovich and Max Shpak University of Texas at El Paso, El Paso, TX 79968, USA vladik@utep.edu, mshpak@utep.edu Summary. Many biologically
BRIEF COMMUNICATIONS
 Evolution, 59(12), 2005, pp. 2705 2710 BRIEF COMMUNICATIONS THE EFFECT OF INTRASPECIFIC SAMPLE SIZE ON TYPE I AND TYPE II ERROR RATES IN COMPARATIVE STUDIES LUKE J. HARMON 1 AND JONATHAN B. LOSOS Department
Evolution, 59(12), 2005, pp. 2705 2710 BRIEF COMMUNICATIONS THE EFFECT OF INTRASPECIFIC SAMPLE SIZE ON TYPE I AND TYPE II ERROR RATES IN COMPARATIVE STUDIES LUKE J. HARMON 1 AND JONATHAN B. LOSOS Department
DETECTING BIOLOGICAL AND ENVIRONMENTAL CHANGES: DESIGN AND ANALYSIS OF MONITORING AND EXPERIMENTS (University of Bologna, 3-14 March 2008)
 Dipartimento di Biologia Evoluzionistica Sperimentale Centro Interdipartimentale di Ricerca per le Scienze Ambientali in Ravenna INTERNATIONAL WINTER SCHOOL UNIVERSITY OF BOLOGNA DETECTING BIOLOGICAL AND
Dipartimento di Biologia Evoluzionistica Sperimentale Centro Interdipartimentale di Ricerca per le Scienze Ambientali in Ravenna INTERNATIONAL WINTER SCHOOL UNIVERSITY OF BOLOGNA DETECTING BIOLOGICAL AND
Computational Tasks and Models
 1 Computational Tasks and Models Overview: We assume that the reader is familiar with computing devices but may associate the notion of computation with specific incarnations of it. Our first goal is to
1 Computational Tasks and Models Overview: We assume that the reader is familiar with computing devices but may associate the notion of computation with specific incarnations of it. Our first goal is to
GENERAL ECOLOGY STUDY NOTES
 1.0 INTRODUCTION GENERAL ECOLOGY STUDY NOTES A community is made up of populations of different organisms living together in a unit environment. The manner in which these organisms relate together for
1.0 INTRODUCTION GENERAL ECOLOGY STUDY NOTES A community is made up of populations of different organisms living together in a unit environment. The manner in which these organisms relate together for
Chapter 4. Probability Theory. 4.1 Probability theory Events
 Chapter 4 Probability Theory Probability theory is a branch of mathematics that is an essential component of statistics. It originally evolved from efforts to understand the odds and probabilities involved
Chapter 4 Probability Theory Probability theory is a branch of mathematics that is an essential component of statistics. It originally evolved from efforts to understand the odds and probabilities involved
Chapter 6 Population and Community Ecology
 Chapter 6 Population and Community Ecology Friedland and Relyea Environmental Science for AP, second edition 2015 W.H. Freeman and Company/BFW AP is a trademark registered and/or owned by the College Board,
Chapter 6 Population and Community Ecology Friedland and Relyea Environmental Science for AP, second edition 2015 W.H. Freeman and Company/BFW AP is a trademark registered and/or owned by the College Board,
BLAST: Target frequencies and information content Dannie Durand
 Computational Genomics and Molecular Biology, Fall 2016 1 BLAST: Target frequencies and information content Dannie Durand BLAST has two components: a fast heuristic for searching for similar sequences
Computational Genomics and Molecular Biology, Fall 2016 1 BLAST: Target frequencies and information content Dannie Durand BLAST has two components: a fast heuristic for searching for similar sequences
Big Idea 1: The process of evolution drives the diversity and unity of life.
 Big Idea 1: The process of evolution drives the diversity and unity of life. understanding 1.A: Change in the genetic makeup of a population over time is evolution. 1.A.1: Natural selection is a major
Big Idea 1: The process of evolution drives the diversity and unity of life. understanding 1.A: Change in the genetic makeup of a population over time is evolution. 1.A.1: Natural selection is a major
Measuring phylogenetic biodiversity
 CHAPTER 14 Measuring phylogenetic biodiversity Mark Vellend, William K. Cornwell, Karen Magnuson-Ford, and Arne Ø. Mooers 14.1 Introduction 14.1.1 Overview Biodiversity has been described as the biology
CHAPTER 14 Measuring phylogenetic biodiversity Mark Vellend, William K. Cornwell, Karen Magnuson-Ford, and Arne Ø. Mooers 14.1 Introduction 14.1.1 Overview Biodiversity has been described as the biology
Effects to Communities & Ecosystems
 Biology 5868 Ecotoxicology Effects to Communities & Ecosystems April 18, 2007 Definitions Ecological Community an assemblage of populations living in a prescribed area or physical habitat [It is] the living
Biology 5868 Ecotoxicology Effects to Communities & Ecosystems April 18, 2007 Definitions Ecological Community an assemblage of populations living in a prescribed area or physical habitat [It is] the living
Affinity analysis: methodologies and statistical inference
 Vegetatio 72: 89-93, 1987 Dr W. Junk Publishers, Dordrecht - Printed in the Netherlands 89 Affinity analysis: methodologies and statistical inference Samuel M. Scheiner 1,2,3 & Conrad A. Istock 1,2 1Department
Vegetatio 72: 89-93, 1987 Dr W. Junk Publishers, Dordrecht - Printed in the Netherlands 89 Affinity analysis: methodologies and statistical inference Samuel M. Scheiner 1,2,3 & Conrad A. Istock 1,2 1Department
Research Note: A more powerful test statistic for reasoning about interference between units
 Research Note: A more powerful test statistic for reasoning about interference between units Jake Bowers Mark Fredrickson Peter M. Aronow August 26, 2015 Abstract Bowers, Fredrickson and Panagopoulos (2012)
Research Note: A more powerful test statistic for reasoning about interference between units Jake Bowers Mark Fredrickson Peter M. Aronow August 26, 2015 Abstract Bowers, Fredrickson and Panagopoulos (2012)
AP Curriculum Framework with Learning Objectives
 Big Ideas Big Idea 1: The process of evolution drives the diversity and unity of life. AP Curriculum Framework with Learning Objectives Understanding 1.A: Change in the genetic makeup of a population over
Big Ideas Big Idea 1: The process of evolution drives the diversity and unity of life. AP Curriculum Framework with Learning Objectives Understanding 1.A: Change in the genetic makeup of a population over
Chapter 54: Community Ecology
 AP Biology Guided Reading Name Chapter 54: Community Ecology Overview 1. What does community ecology explore? Concept 54.1 Community interactions are classified by whether they help, harm, or have no effect
AP Biology Guided Reading Name Chapter 54: Community Ecology Overview 1. What does community ecology explore? Concept 54.1 Community interactions are classified by whether they help, harm, or have no effect
The Ghost of Competition Present
 vol. 173, no. 3 the american naturalist march 2009 The Ghost of Competition Present T. E. Miller, * C. P. terhorst, and J. H. Burns Department of Biological Science, Florida State University, Tallahassee,
vol. 173, no. 3 the american naturalist march 2009 The Ghost of Competition Present T. E. Miller, * C. P. terhorst, and J. H. Burns Department of Biological Science, Florida State University, Tallahassee,
SLOSS debate. reserve design principles. Caribbean Anolis. SLOSS debate- criticisms. Single large or several small Debate over reserve design
 SLOSS debate reserve design principles Single large or several small Debate over reserve design SLOSS debate- criticisms Caribbean Anolis Pattern not always supported Other factors may explain diversity
SLOSS debate reserve design principles Single large or several small Debate over reserve design SLOSS debate- criticisms Caribbean Anolis Pattern not always supported Other factors may explain diversity
Chapter 6 Population and Community Ecology. Thursday, October 19, 17
 Chapter 6 Population and Community Ecology Module 18 The Abundance and Distribution of After reading this module you should be able to explain how nature exists at several levels of complexity. discuss
Chapter 6 Population and Community Ecology Module 18 The Abundance and Distribution of After reading this module you should be able to explain how nature exists at several levels of complexity. discuss
Unit 8: Ecology Guided Reading Questions (60 pts total)
 AP Biology Biology, Campbell and Reece, 10th Edition Adapted from chapter reading guides originally created by Lynn Miriello Name: Unit 8: Ecology Guided Reading Questions (60 pts total) Chapter 51 Animal
AP Biology Biology, Campbell and Reece, 10th Edition Adapted from chapter reading guides originally created by Lynn Miriello Name: Unit 8: Ecology Guided Reading Questions (60 pts total) Chapter 51 Animal
The Royal Entomological Society Journals
 Read the latest Virtual Special Issues from The Royal Entomological Society Journals Click on the buttons below to view the Virtual Special Issues Agricultural and Forest Pests Introduction This virtual
Read the latest Virtual Special Issues from The Royal Entomological Society Journals Click on the buttons below to view the Virtual Special Issues Agricultural and Forest Pests Introduction This virtual
Ecosystem change: an example Ecosystem change: an example
 5/13/13 Community = An assemblage of populations (species) in a particular area or habitat. Here is part of a community in the grassland of the Serengetti. Trophic downgrading of planet Earth: What escapes
5/13/13 Community = An assemblage of populations (species) in a particular area or habitat. Here is part of a community in the grassland of the Serengetti. Trophic downgrading of planet Earth: What escapes
The Structure of Ecological Networks and Consequences for Fragility
 The Structure of Ecological Networks and Consequences for Fragility closely connected clustered Emily I. Jones ECOL 596H Feb. 13, 2008 Why ecological network structure matters 2. 3. the network contains
The Structure of Ecological Networks and Consequences for Fragility closely connected clustered Emily I. Jones ECOL 596H Feb. 13, 2008 Why ecological network structure matters 2. 3. the network contains
Overview. How many species are there? Major patterns of diversity Causes of these patterns Conserving biodiversity
 Overview How many species are there? Major patterns of diversity Causes of these patterns Conserving biodiversity Biodiversity The variability among living organisms from all sources, including, inter
Overview How many species are there? Major patterns of diversity Causes of these patterns Conserving biodiversity Biodiversity The variability among living organisms from all sources, including, inter
Ecology Regulation, Fluctuations and Metapopulations
 Ecology Regulation, Fluctuations and Metapopulations The Influence of Density on Population Growth and Consideration of Geographic Structure in Populations Predictions of Logistic Growth The reality of
Ecology Regulation, Fluctuations and Metapopulations The Influence of Density on Population Growth and Consideration of Geographic Structure in Populations Predictions of Logistic Growth The reality of
Complex Systems Theory and Evolution
 Complex Systems Theory and Evolution Melanie Mitchell and Mark Newman Santa Fe Institute, 1399 Hyde Park Road, Santa Fe, NM 87501 In Encyclopedia of Evolution (M. Pagel, editor), New York: Oxford University
Complex Systems Theory and Evolution Melanie Mitchell and Mark Newman Santa Fe Institute, 1399 Hyde Park Road, Santa Fe, NM 87501 In Encyclopedia of Evolution (M. Pagel, editor), New York: Oxford University
Humans on Planet Earth (HOPE) Long-term impacts on biosphere dynamics
 Humans on Planet Earth (HOPE) Long-term impacts on biosphere dynamics John Birks Department of Biology, University of Bergen ERC Advanced Grant 741413 2018-2022 Some Basic Definitions Patterns what we
Humans on Planet Earth (HOPE) Long-term impacts on biosphere dynamics John Birks Department of Biology, University of Bergen ERC Advanced Grant 741413 2018-2022 Some Basic Definitions Patterns what we
A A A A B B1
 LEARNING OBJECTIVES FOR EACH BIG IDEA WITH ASSOCIATED SCIENCE PRACTICES AND ESSENTIAL KNOWLEDGE Learning Objectives will be the target for AP Biology exam questions Learning Objectives Sci Prac Es Knowl
LEARNING OBJECTIVES FOR EACH BIG IDEA WITH ASSOCIATED SCIENCE PRACTICES AND ESSENTIAL KNOWLEDGE Learning Objectives will be the target for AP Biology exam questions Learning Objectives Sci Prac Es Knowl
A Genetic Algorithm Approach for Doing Misuse Detection in Audit Trail Files
 A Genetic Algorithm Approach for Doing Misuse Detection in Audit Trail Files Pedro A. Diaz-Gomez and Dean F. Hougen Robotics, Evolution, Adaptation, and Learning Laboratory (REAL Lab) School of Computer
A Genetic Algorithm Approach for Doing Misuse Detection in Audit Trail Files Pedro A. Diaz-Gomez and Dean F. Hougen Robotics, Evolution, Adaptation, and Learning Laboratory (REAL Lab) School of Computer
Fundamental ecological principles
 What Important Ideas Will Emerge in Your Study of Ecology? Fundamental ecological principles Application of the scientific method to answer specific ecological questions Ecology is a quantitative science
What Important Ideas Will Emerge in Your Study of Ecology? Fundamental ecological principles Application of the scientific method to answer specific ecological questions Ecology is a quantitative science
Computational Biology, University of Maryland, College Park, MD, USA
 1 Data Sharing in Ecology and Evolution: Why Not? Cynthia S. Parr 1 and Michael P. Cummings 2 1 Institute for Advanced Computer Studies, 2 Center for Bioinformatics and Computational Biology, University
1 Data Sharing in Ecology and Evolution: Why Not? Cynthia S. Parr 1 and Michael P. Cummings 2 1 Institute for Advanced Computer Studies, 2 Center for Bioinformatics and Computational Biology, University
What determines: 1) Species distributions? 2) Species diversity? Patterns and processes
 Species diversity What determines: 1) Species distributions? 2) Species diversity? Patterns and processes At least 120 different (overlapping) hypotheses explaining species richness... We are going to
Species diversity What determines: 1) Species distributions? 2) Species diversity? Patterns and processes At least 120 different (overlapping) hypotheses explaining species richness... We are going to
Yakın Doğu Üniversitesi Mimarlık Fakültesi Peyzaj Mimarlığı Bölümü. PM 317 Human and Environment Assoc. Prof. Dr. Salih GÜCEL
 Yakın Doğu Üniversitesi Mimarlık Fakültesi Peyzaj Mimarlığı Bölümü PM 317 Human and Environment Assoc. Prof. Dr. Salih GÜCEL Ecology & Ecosystems Principles of Ecology Ecology is the study of the interactions
Yakın Doğu Üniversitesi Mimarlık Fakültesi Peyzaj Mimarlığı Bölümü PM 317 Human and Environment Assoc. Prof. Dr. Salih GÜCEL Ecology & Ecosystems Principles of Ecology Ecology is the study of the interactions
Logistic Regression: Regression with a Binary Dependent Variable
 Logistic Regression: Regression with a Binary Dependent Variable LEARNING OBJECTIVES Upon completing this chapter, you should be able to do the following: State the circumstances under which logistic regression
Logistic Regression: Regression with a Binary Dependent Variable LEARNING OBJECTIVES Upon completing this chapter, you should be able to do the following: State the circumstances under which logistic regression
