Trees of Unusual Size: Biased Inference of Early Bursts from Large Molecular Phylogenies
|
|
|
- Alvin Thompson
- 6 years ago
- Views:
Transcription
1 Trees of Unusual Size: Biased Inference of Early Bursts from Large Molecular Phylogenies Matthew W. Pennell 1,2,3 *, Brice A. J. Sarver 1,2,3, Luke J. Harmon 1,2,3 1 Institute for Bioinformatics and Evolutionary Studies (IBEST), University of Idaho, Moscow, Idaho, United States of America, 2 Department of Biological Sciences, University of Idaho, Moscow, Idaho, United States of America, 3 BEACON Center for the Study of Evolution in Action, East Lansing, Michigan, United States of America Abstract An early burst of speciation followed by a subsequent slowdown in the rate of diversification is commonly inferred from molecular phylogenies. This pattern is consistent with some verbal theory of ecological opportunity and adaptive radiations. One often-overlooked source of bias in these studies is that of sampling at the level of whole clades, as researchers tend to choose large, speciose clades to study. In this paper, we investigate the performance of common methods across the distribution of clade sizes that can be generated by a constant-rate birth-death process. Clades which are larger than expected for a given constant-rate branching process tend to show a pattern of an early burst even when both speciation and extinction rates are constant through time. All methods evaluated were susceptible to detecting this false signature when extinction was low. Under moderate extinction, both the c-statistic and diversity-dependent models did not detect such a slowdown but only because the signature of a slowdown was masked by subsequent extinction. Some models which estimate time-varying speciation rates are able to detect early bursts under higher extinction rates, but are extremely prone to sampling bias. We suggest that examining clades in isolation may result in spurious inferences that rates of diversification have changed through time. Citation: Pennell MW, Sarver BAJ, Harmon LJ (2012) Trees of Unusual Size: Biased Inference of Early Bursts from Large Molecular Phylogenies. PLoS ONE 7(9): e doi: /journal.pone Editor: Simon Ho, University of Sydney, Australia Received April 11, 2012; Accepted July 19, 2012; Published September 5, 2012 Copyright: ß 2012 Pennell et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. Funding: LJH and MWP were funded by National Science Foundation grant DEB MWP also received funding from a NSERC-PGS-M award. This material is based in part upon work supported by the National Science Foundation under Cooperative Agreement No. DBI Any opinions, findings, and conclusions or recommendations expressed in this material are those of the authors and do not necessarily reflect the views of the National Science Foundation. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript. Competing Interests: The authors have declared that no competing interests exist. * mwpennell@gmail.com Introduction The branching patterns of reconstructed molecular phylogenies contain information about the tempo and mode of evolution [1], [2]. This insight has been invaluable to our understanding of many evolutionary processes and patterns. Recent studies have identified a common pattern of apparent slowdowns in the rate of diversification through time (reviewed in [3], [4]). Such a pattern is consistent with some theoretical work on adaptive radiations, based on the idea that diversification rates will decrease as species fill available niches [5], [6], [7] (but see [8] for an alternate interpretation). Phylogenies with slowdowns have been inferred for diverse groups of organisms (e.g. lizards [9], birds [10], [11], fish [12], and many groups of plants [13]). The most commonly used metric for detecting shifts in the rate of diversification is the c-statistic introduced by Pybus and Harvey [14]. This statistic quantifies how internode distances (i.e. waiting times to speciation) vary through time compared to what one would expect under a pure-birth model of diversification. Under the pure-birth null expectation, c is distributed according to a standard normal distribution. A c-value less than (onetailed test) represents a statistically significant slowdown in diversification rate. Positive values of c can be caused by either speed-ups in diversification rates or species turnover as recently diverged lineages have have not been around long enough to have been pruned by extinction, creating an overabundance of nodes closer to the present ( pull of the present ; [2]). These two scenarios cannot be distinguished with the c-statistic, so most studies follow the authors [14] original recommendation to disregard significantly positive c-values. There has been considerable controversy surrounding the interpretations of slowdowns using the c-statistic on molecular phylogenies. A preponderance of studies have inferred significantly negative c- values across a wide range of taxonomic groups. McPeek [8] collected a large number of phylogenies from the literature and found that 80% of clades had cv0 and 42% had cv{1:96. However, a number of other studies have pointed out that negative c-values can result from factors other than slowdowns in diversification rates. It has been shown that c can be biased by under-parameterization of the model of sequence evolution [15] and non-random taxon sampling [16] (see [17] and [18] for potential solutions to this problem). Recent speciation events will have a similar effect; if speciation is modelled as a process rather than as a singular event, this can lead to apparent slowdowns in diversification rates [19], [20]. A recent simulation study, conducted by Liow et al. [21], found that c tended to be positively biased when trees were simulated under a birth-death process with variable rates of speciation. They found that, even when present, slowdowns in net diversification rates were very difficult to detect using c, as short branches near the tips of the tree tend to obscure the signal of the slowdown. Under the conditions simulated by Liow et al. [21], it should be PLOS ONE 1 September 2012 Volume 7 Issue 9 e43348
2 only possible to observe a slowdown for a small subset of values of speciation rate (l) and extinction rate (m) and if lineages are sampled at a particular time in the clade s history. It seems unrealistic to propose that these two conditions are met for the majority of phylogenies. Nevertheless slowdowns are commonly inferred from empirical data. The reasons for the discordance between the analysis of empirical data and expectations derived from simulation studies are poorly understood and warrant further investigation. Several new modeling approaches have been developed to detect non-homogeneous diversification. One is to use a model in which speciation and/or extinction vary through time [22], [22], [23], [24], [25]. Diversity-dependence (in which speciation rate varies as a function of the number of taxa at a given time) has also been proposed as an explanation for the patterns of species richness [26]. This has recently been modeled in a number of studies (e.g. [27], [28]) to look for signatures of an adaptive radiation. Diversity-dependence is a useful approach as it is seemingly consistent with theory on adaptive radiation, which posits that diversification should slow as niches become filled [6], [7]. There is also some evidence from the fossil record [29] (but see [30] for an opposing perspective) to support this modeling approach. However, such models from both paleobiology and comparative biology are susceptible to the criticism that the ecological mechanisms by which diversity dependence would operate on the scale of a clade are not entirely clear [31]. While the mathematics of the various methods differ, they all involve fitting non-homogeneous birth-death models to phylogenies rather than using summary statistics such as the c-statistic. The behavior and performance of these methods have not been explored in as much detail as the c-statistic. One important consideration that is often overlooked in studies of clade diversification is that our inferences may be biased by the way we choose clades to investigate. There are several types of sampling bias that are likely to be important. Cusimano and Renner [16], Brock et al. [17] and Cusimano et al. [18] investigated systematists tendency to sample representative taxa, leading to an overabundance of nodes deep in the tree. However, we should also consider that we are only sampling clades that survive to the present day. This means that the observable distribution of surviving trees is only a subset of all possible trees [32],[33]. Another form of bias is that researchers interested in the adaptive radiations are likely to be interested in the speciose clades. These clades likely reside in the tails of the distribution of possible clade sizes and thus give a biased sample for the inference of diversification rates [11]. This effect of sampling clades has seldom been addressed in the literature (but see [32],[33]). Inferences of early bursts are particularly problematic because even under a constant pure-birth or birth-death process, large clades are likely to show patterns of a rapid diversification early in their history. This is simply due to the fact that in order to be large, they are more likely to have undergone a stochastically high rate of speciation early in the process and subsequently regressed to the mean speciation rate [34], [11]. Analyzing large clades in isolation may lead to inferring ecological processes (such as adaptive radiations) attributable solely to these stochastic processes. Phillimore and Price [11] investigated the influence of clade size and age on the c-statistic though their simulations were limited to a small set of parameter space. Specifically, they conditioned their simulations on tree age to investigate the correlation between c and clade size and on clade size to investigate the correlation between c and tree age. As a result, the simulations tended to produce trees where the distribution of the parameter of interest was centered on the expected value. Here, we are specifically interested in investigating the statistical properties of trees that are unexpected, that is, unusually large. By simultaneously conditioning our phylogenies on both tree age and size, we were able to obtain trees from the tails of the distribution. In addition to using the c- statistic, we investigate the effects of the non-random sampling of clades using a model selection approach. We show that the effect of this bias can be severe but it affects different methods in different ways, and we make recommendations on the appropriate use of these methods. Results and Discussion For all phylogenies simulated, regardless of the value of extinction rate used, large phylogenies are disproportionally likely to have undergone an initial burst of speciation events [35], [11]. This can be visualized with a lineages-through-time plot (Figure 1). This result has consequences for the inference of diversification rate patterns; as the clades we choose to examine are not a random sample of clades but often tend to be interesting and speciose and, at least for molecular phylogenies, have survived to the present day. It is also the case that these large clades are the ones most amenable to statistical analysis. We simulated phylogenies under constant-rate diversification, conditioning on both the number of taxa in the clade (N) and the age of the tree (t). We did this in order to investigate the statistical properties of trees drawn from different parts of the distribution of possible tree sizes that can result from a constant-rate process. Consistent with previous work [35], [11], trees which are large at the present day (i.e. those drawn from the right tail of the distribution) are more likely to harbor a signal of a slowdown using the c-statistic even when the phylogeny is generated under a constant-rate process with low extinction. This is evident in Figure 2, where data points corresponding to tree sizes larger than the expectation (denoted by a dashed line) show elevated Type-1 error rates when simulated under low extinction rates. However this is not the case when background extinction rates are higher as extinction alters the distribution of branch lengths on the reconstructed phylogeny. The resulting distribution effectively obscures any signal of a burst (in our simulations, the burst is entirely attributable to stochastic processes) that might have occured deep in the tree [10], [36], [21]. This is especially true for large phylogenies (Figure 2); c is a summary statistic for all nodes in the tree and therefore the few early branching events will have an even smaller influence on the calculation of the c-statistic for large trees with proportionally more nodes. Our results suggest two things about the use of the c-statistic. First, the test has very low power to detect changes in speciation rates when species turnover rates are even modestly high [21]. Second, while there is some concern that significantly negative c- values may potentially be misleading [15], [16], [37], bias in sampling large clades does not tend to create false signatures of a slowdown under modest extinction rates. We know from the fossil record that extinction rates have been moderately high for most groups. This implies that, even if slowdowns in diversification rates are relatively common, significantly negative c-values should be rare. However, this is not the case [8]. The widely reported evidence for slowdowns in studies using c is thus somewhat paradoxical and deserves explanation. Pessimistically, we may attribute this to some yet unstudied artifact such as sampling effort. More intensive sampling of lineages would have the effect of increasing the number of nodes close to the present. This would deteriorate any signal of an early burst detectable by the c-statistic. It should be noted however, that a number of the phylogenies that provide evidence for a slowdown are near complete at the species PLOS ONE 2 September 2012 Volume 7 Issue 9 e43348
3 Figure 1. Exemplar Lineages-Through-Time Plot. An example of a lineages-through-time (LTT) plot for a tree (shown on left) drawn from the far right tail of the distribution of tree sizes (5 percent of surviving trees are expected to be this large or larger) for l~1, m~0:5 and t~5. The dotted line is the expected number of lineages under a constant diversification rate. This LTT plot shows the typical signature of an early burst of speciation yet this signature is not captured by the c-statistic (c~0:602; not significant) as the burst is masked by later extinction events. doi: /journal.pone g001 level (e.g. [11]), though there may still be additional lineages that should actually be recognized as species. It may be the case that if more lineages were included, many of the signatures of early bursts in the literature would no longer be present. This is difficult to evaluate except on a case-by-case basis. We found that the diversity-dependent models are prone to bias due to sampling in a similar manner as the c-statistic (Figure 3); when death rates are low, the diversity-dependent model was preferred to a constant birth-death model more frequently for clades at the large end of the distribution of clade size. But even with moderate extinction, the diversity-dependent model was rarely preferred. In fact, for higher extinction rates, the reverse was true larger clades were less likely to fit a diversity-dependent model than smaller clades (Figure 3). However, it should be noted that the model of diversity-dependence we used here did not explicity model extinction. Consequently the model estimates Figure 2. Type-1 Error Rate for the c-statistic. Results from simulations showing the Type-1 error rate for the c statisitc, signifying a false inference of a slowdown. All trees were generated under a constant-rate birth-death (or pure-birth) process. We recognize a Type-1 error if the value of cv{1:645 (significant at p~0:05; one-tailed test). Extinction rate, e~m=l, varies across the plots (e~0,0:1,0:25,0:5); speciation rate, l, and total tree-depth, t are held constant (l~1 and t~5). All are plotted against the expected number of taxa across the cumulative distribution of probability densities (from 0.99 to 0.01). The dashed vertical line represents the expected value for N under the simulating conditions. Each point represents 1000 simulations. (Results for e~0:75 and e~1 not shown.) doi: /journal.pone g002 PLOS ONE 3 September 2012 Volume 7 Issue 9 e43348
4 diversification rate as a function of contemporaneous lineages in the reconstructed phylogeny rather than as a function of contemporaneous lineages in the true (unobserved) phylogeny [38]. Recently Etienne et al. [28] provided a full likelihood solution for diversity-dependence with extinction which uses a Hidden-Markov approach to fit the model to a phylogeny. We justify the use of a diversity-dependent model which does not estimate extinction in our simulation study due to the fact that it is currently the more commonly employed approach. The diversitydependent model thus has the same drawback as the c-statistic. Extinction will tend to erode the signal of the early diversification events, especially so for large trees. The results from the timevarying speciation rate model (Figures 4 and 5; discussed below) suggest that this result will not hold if both extinction rates and speciation rates are explicitly modeled. However, it should be noted that there is some evidence that estimates of extinction can be inconsistent when there is rate variation across clades [39]. A correction that has been employed to increase the power to detect slowdowns for both the c-statistic and diversity-dependent models is to collapse some nodes close to the present. Species delimitation is a problem that has recently received a great deal of attention from molecular systematists (e.g. [40],[41],[42]). Subdividing lineages into subspecies will necessarily increase the effect of the pull of the present and further obscure evidence of a slowdown. Collapsing these recent nodes will certainly influence our ability to infer slowdowns and this procedure has been done by some researchers (e.g. [11], [43]), by regarding recently-diverged species as not being good species. However, there is currently a lack of theory to guide such decision-making. We therefore recommend that authors should be cautious in collapsing nodes as we do not fully understand how species delimitation affects these methods. The time-varying speciation models [36], [23] were preferred for data generated under a constant-rate process with increasing frequency as tree size increased. We found this to be true even when extinction was present (Figures 4 and 5), though the effect was dampened as extinction increased. As stated above, large trees generated under a constant rate birth-death process are subject to having undergone stochastic bursts of speciation in order to obtain their current size. The time-varying speciation rate models are sensitive to these bursts, which is both a positive and negative; positive because they have more power to detect changes in diversification through time and negative because these models are more prone to inferring spurious results as a consequence of sampling bias. We took two alternative approaches to compare model fits of a constant-rate birth-death model versus a timevarying speciation model: the full-likelihood approach of Rabosky and Lovette [36] and an approximate approach based on the coalescent process, recently derived by Morlon et al. [23] (see Methods for details). Contrary to our expectations, we found that these two approaches differed substantially in terms of which model was prefered when using Akaike Information Criterion (AIC). For trees from the larger end of the distribution, the coalescent approach tended to provide support for a time-varying speciation model over a constant-rate model at much higher frequencies than the full likelihood approach, especially when extinction rates were low (Figure 5). The reasons for these differences are not entirely clear. While many of the large trees do show early bursts due to stochastically high rates early in the process (see Figure 1 for an example), the proneness of the coalescent-based approach to favor a model more complex than the generating model is worrisome as sampling bias appears to very strongly influence model choice. Both approaches are relatively new and their respective statistical properties have not been explored at all beyond the publications in which they were presented [36], [23]. The statistical properties of these and other related models is something that certainly warrants further investigation. Researchers are increasingly fitting more complex models of diversification to study diversity dynamics through time and across clades (e.g. [44], [25], [24]) and it is imperative that we have a good understanding of these if we are to make meaningful inferences. Stochastic bursts, which are more likely to have occured in unusually large trees, may be falsely inferred to have been caused by ecological processes (e.g. adaptive radiations). While the bursts are certainly real, they are not attributable to any biological phenomena. If the background extinction is close to zero, then all methods investigated in this paper are susceptible to this false inference. However, when background extinction is relatively high, these stochastically generated slowdowns are not likely to be detected by the c-statistic as the subsequent extinction removes the signal. As a summary statistic for the whole phylogeny, the power of the c-statistic to detect slowdowns (stochastically or ecologically Figure 3. Proportion of Trees Showing Support for Diversity-Dependent Model. Results from simulations showing the proportion of phylogenies for which a density-dependent (DD) model is preferred over a constant-rate birth-death (BD) model in using AIC to select amongst the models. Only the DD model and a BD model were compared. We used a DAIC cutoff of 4 to favor a DD model when the generating model was a constant-rate process. Extinction rate, e~m=l, varies across the plots (e~0,0:1,0:25,0:5); speciation rate, l, and total tree-depth, t are held constant (l~1 and t~5). All are plotted against the expected number of taxa across the cumulative distribution of probability densities (from 0.99 to 0.01). The dashed vertical line represents the expected value for N under the simulating conditions. Each point represents 1000 simulations. (Results for e~0:75 and e~1 not shown.) doi: /journal.pone g003 PLOS ONE 4 September 2012 Volume 7 Issue 9 e43348
5 Figure 4. Proportion of Trees Showing Support for Temporally-Varying Speciation Model. Results from simulations showing the proportion of phylogenies for which the temporally-varying speciation (TVS) model (the SPVAR model of [36]) is preferred over a constant-rate birthdeath (BD) model using AIC to select amongst the models. Only the TVS and BD models were compared. We used a DAIC cutoff of 4 to favor a TVS model when the generating model was a constant-rate process. Extinction rate, e~m=l, varies across the plots (e~0,0:1,0:25,0:5); speciation rate, l, and total tree-depth, t are held constant (l~1 and t~5). All are plotted against the expected number of taxa across the cumulative distribution of probability densities (from 0.99 to 0.01). The dashed vertical line represents the expected value for N under the simulating conditions. Each point represents 1000 simulations. (Results for e~0:75 and e~1 not shown.) doi: /journal.pone g004 produced) when species turnover is present is very low [21], [45]. It is fair to suggest that while there are some reasons to believe that significantly negative c-values in the empirical literature may result from known statistical artifacts [15], [16], [37], the cause of their ubiquity remains unclear. We suggest that failing to sample cryptic species may contribute to this. Methods that do not deal directly with extinction and extinct lineages, like the diversity-dependent model [27] used here, show similar patterns of performance to the c-statistic. Our findings are also of relevance to studies examining the efficacy of diversification rate models and statistics. There are a multitude of ways to conduct simulations of birth-death models and Stadler [46] has discussed the statistical properties of various conditioning schemes in detail. We caution theoreticians to pay close attention to these differences as conditioning simultaneously on the number of taxa and the age of the tree can produce phylogenies drawn from the tails of the distribution and thus prone to be bottom heavy. Such trees may lead to biased estimation of parameters and misconstrued inferences. We suggest three future directions to address this issue of sampling bias. First, performing diversification analysis on megaphylogenies, including the clade of interest, may be advisable instead of examining clades in isolation. Investigating large clades while ignoring closely related groups that are less speciose may lead to spurious patterns of slowdowns in diversification. This will require further development of methods that relax the assumption of uniformity of the diversity dynamics across the tree (e.g. [44]). We appreciate that for many groups, such an approach may not always be feasible so we urge researchers to at least be cautious in their interpretation of their results. Second, increased attention should be given to how lineages are defined; lineages that are currently denoted as subspecies may soon be considered good species [20]. Whether these lineages are included or not will influence the inference of slowdowns. Third, further investigation Figure 5. Proportion of Trees Showing Support for Temporally-Varying Speciation Model Using Coalescent Approximation. Results from simulations showing the proportion of phylogenies for which a temporally-varying speciation (TVS) model (Model 4a of [23]) is preferred over a constant-rate birth-death model (BD; Model 3 of [23]) using AIC to select amongst the models. Both the models were formulated according to a coalescent-based approximation of the likelihood [23]. We used a DAIC cutoff of 4 to favor the TVS model when the generating model was a constant-rate process. Extinction rate, e~m=l, varies across the plots (e~0,0:1,0:25,0:5); speciation rate, l, and total tree-depth, t are held constant (l~1 and t~5). All are plotted against the expected number of taxa across the cumulative distribution of probability densities (from 0.99 to 0.01). The dashed vertical line represents the expected value for N under the simulating conditions. Each point represents 1000 simulations. (Results for e~0:75 and e~1 not shown.) doi: /journal.pone g005 PLOS ONE 5 September 2012 Volume 7 Issue 9 e43348
6 of the statistical properties of these models may allow researchers to be more confident that the patterns they observe represent truly meaningful variation. Methods All of our simulations were carried out using constant rates of speciation and extinction through time, so that any detection of a slowdown can be considered a Type-1 error. Tree simulations were conducted with Stadler s [46] TreeSim R package, modified slightly for the purposes of computational efficiency. For each simulated phylogeny, we calculated the c-statistic [14] and fit five models of diversification: 1) a constant-rate birth-death model [47]; 2) a diversity-dependent model [27]; 3) a time-varying speciation rate model [36]; 4) a constant-rate birth-death model using a coalescent approach [23]; and 5) a time-varying speciation rate model, also using a coalescent approach [23]. Models 1 3 were fit using the R package laser [48]. Models 4 and 5 were fit using code from Morlon et al. [23], also modified for computational efficiency. We simulated phylogenies conditioned on both the age of the tree t and the number of taxa N (for a discussion on various methods of conditioning simulated phylogenies, see [46]). In order to simulate trees from different regions of the cumulative probability distribution (i.e. 1 st percentile, 2 nd percentile,..., 99 th percentile), we used the equations of Foote et al. [49] for calculating Pr(NDl,m,t) where l is the instantaneous speciation rate per unit time and m is the instantaneous extinction rate per unit time. The probability of observing a phylogenetic tree of age t, containing at least n taxa, is given by the equations: for l~m and and for l=m Pr(N njl,m,t)~1{ m(exp½(l{m)tš{1) l exp½(l{m)tš{m { X n{1 i~1 b~ la m (1{a)(1{b)b i{1 Pr(N ndl,m,t)~1{ lt (1zlt) { Xn{1 (lt) i{1 (1zlt) iz1 Note that in their formulation of these equations, Foote et al. [49] use the paleontological convention of denoting l as p and m as q. We set l~1, t~5 and varied m (m~f0,0:1,0:25,0:5,0:75,1:0g). Conditioning on the survival of at least one lineage to the present day, we used (1) and (2) to calculate the number of taxa associated with the percentiles of the cumulative distribution as discussed above. For each value of m and each percentile (p~0:01,0:02,...,0:99), we simulated 1000 phylogenies under a constant-rate birth-death model (or in the case of m~0, purebirth). We also varied l and t but the results of our analysis did not differ qualitatively. As stated above, the c-statistic [14] quantifies the temporal distribution of internode distances. We calculated c for each phylogeny using the R package ape [50]. Following the author s i~1 ð1þ ð2þ original recommendation [14] and the conventions of the literature, we used a one-tailed test, such that c was considered to be significant if it was v{1:645. Again, as all phylogenies were simulated under a constant-rate process, we considered it to be a Type-1 Error if c was significantly negative. The diversity-dependent (DD) model we used was the exponential declining model of Rabosky and Lovette [27] where speciation rate is modeled as a function of the number of contemporaneous lineages in the reconstructed phylogeny. If N(t) the number of lineages at time t, l(0) is the initial (background) speciation rate and k describes the strength of diversity-dependence, l(t) can be described as the function l(t)~l(0)n(t) {k : ð3þ In the DD model, background extinction rates are assumed to be 0. The decline in diversification rates at higher densities is thus only due to a decrease in speciation rate. To model time-varying speciation without explicitly considering diversity-dependence, we used the SPVAR model of Rabosky and Lovette [36] where speciation rate is an exponential function of time such that l(t)~l(0)e {xt ð4þ where x is a constant describing the relationship. Here, extinction rate m is explicitly modeled but assumed to be constant across the phylogeny. As an alternative to using the full-likelihood equation for a timevarying speciation model [36], we also employed a recently derived approach [23] for fitting birth-death models, based on the coalescent process from population genetics [51]. Morlon et al. [23] model a population of species evolving under the Wright- Fisher process. Following Morlon et al. [23], the likelihood L of observing the internode distances g i between node i and node iz1 can be written as L(g i )~ i(iz1) 2 1 N(u i ) " ð # ui 1 u i {g i N(t) dt exp { i(iz1) 2 where u i is the distance between node i and the present and N(t) is the number of species (population size in population genetics) at time t in the past. Note that this is an approximation of the likelihood as N(t) is approximated by its expected value E½N(t) Š [23]; this was done to make the model analytically tractable. The constant-rate birth-death model we use corresponds to Model 3 from [23]. The time-varying speciation model was specified in the same way in the full-likelihood approach described above, such that l(t)~l(0)e {xt. This corresponds to Model 4a from [23]. For the model-based approaches, we compared a constant-rate birth-death model to each alternative model in our set. We used an AIC approach (sensu [52]) to select the best fit model between a constant-rate birth-death model and a time-varying speciation rate/diversity-dependent model. We emphasize for clarity that only 2 models were considered at a time. We used a DAIC cutoff of 4 for the more parameter-rich model to be favored. While a DAIC of 4 is an arbitrary value, we justify its use as it is conventionally used in phylogenetic comparative methods, following recomendations in [52]. All analyses were conducted in the R programming environment [53]. The code modified from Stadler [46] and Morlon et al. [23] will be available upon acceptance on M.W.P. s personal website wordpress.com/ ð5þ PLOS ONE 6 September 2012 Volume 7 Issue 9 e43348
7 Acknowledgments We thank Dan Wegmann, Graham Slater and Joseph Brown for providing some code to speed up the simulations and Jon Eastman, Tyler Hether, Jamie Voyles, Simon Uribe-Convers, Simon Ho and Lee Hsiang Liow for insightful comments on earlier versions of this manuscript. We also thank the RoHa lab and the Phylogenetics Reading Group (PuRGe) at the University of Idaho for stimulating discussion of these topics. References 1. Raup DM, Gould SJ, Schopf TJM, Simberlo DS (1973) Stochastic-models of phylogeny and evolution of diversity. Journal of Geology 81: Nee S, Mooers AO, Harvey PH (1992) Tempo and mode of evolution revealed from molecular phylogenies. Proceedings of the National Academy of Sciences of the United States of America 89: Gavrilets S, Losos JB (2009) Adaptive radiation: Contrasting theory with data. Science 323: Glor RE (2010) Phylogenetic insights on adaptive radiation. Annual Review of Ecology, Evolution, and Systematics, Vol 41 41: Simpson GG (1953) Major Features of Evolution. Columbia University Press. 6. Schluter D (2000) The Ecology of Adaptive Radiation. Oxford University Press. 7. Yoder JB, Clancey E, Des Roches S, Eastman JM, Gentry L, et al. (2010) Ecological opportunity and the origin of adaptive radiations. Journal of Evolutionary Biology 23: McPeek MA (2008) The ecological dynamics of clade diversification and community assembly. American Naturalist 172: E270 E Harmon LJ, Schulte JA, Larson A, Losos JB (2003) Tempo and mode of evolutionary radiation in iguanian lizards. Science 301: Weir JT (2006) Divergent timing and patterns of species accumulation in lowland and highland neotropical birds. Evolution 60: Phillimore AB, Price TD (2008) Density-dependent cladogenesis in birds. Plos Biology 6: Rüber L, Zardoya R (2005) Rapid cladogenesis in marine fishes revisited. Evolution 59: Davies TJ, Savolainen V, Chase MW, Moat J, Barraclough TG (2004) Environmental energy and evolutionary rates in owering plants. Proceedings of the Royal Society of London Series B-Biological Sciences 271: Pybus OG, Harvey PH (2000) Testing macro-evolutionary models using incomplete molecular phylogenies. Proceedings of the Royal Society of London Series B-Biological Sciences 267: Revell LJ, Harmon LJ, Glor RE (2005) Underparameterized model of sequence evolution leads to bias in the estimation of diversification rates from molecular phylogenies. Systematic Biology 54: Cusimano N, Renner SS (2010) Slowdowns in diversification rates from real phylogenies may not be real. Systematic Biology 59: Brock CD, Harmon LJ, Alfaro ME (2011) Testing for temporal variation in diversification rates when sampling is incomplete and nonrandom. Systematic Biology 60: Cusimano N, Stadler T, Renner SS (2012) A new method for handling missing species in diversifi-cation analysis applicable to randomly or non-randomly sampled phylogenies. Systematic Biology. 19. Etienne RS, Rosindell J (2012) Prolonging the past counteracts the pull of the present: Protracted speciation can explain observed slowdowns in diversification. Systematic Biology 61: Rosenblum EB, Sarver BAJ, Brown JW, Des Roches S, Hardwick KM, et al. (2012) Goldilocks meets santa rosalia: An ephemeral speciation model explains patterns of diversification across time scales. Evolutionary Biology in press: doi /s x. 21. Liow LH, Quental TB, Marshall CR (2010) When can decreasing diversification rates be detected with molecular phylogenies and the fossil record? Systematic Biology 59: Rabosky DL (2006) Likelihood methods for detecting temporal shifts in diversification rates. Evolution 60: Morlon H, Potts MD, Plotkin JB (2010) Inferring the dynamics of diversification: A coalescent approach. Plos Biology Stadler T (2011) Mammalian phylogeny reveals recent diversification rate shifts. Proceedings of the National Academy of Sciences of the United States of America 108: Morlon H, Parsons TL, Plotkin JB (2011) Reconciling molecular phylogenies with the fossil record. Proceedings of the National Academy of Sciences 108: Rabosky DL (2009) Ecological limits and diversification rate: alternative paradigms to explain the variation in species richness among clades and regions. Ecology Letters 12: Rabosky DL, Lovette IJ (2008) Density-dependent diversification in north american wood warblers. Proceedings of the Royal Society B-Biological Sciences 275: Author Contributions Conceived and designed the experiments: MWP BAJS LJH. Performed the experiments: MWP. Analyzed the data: MWP. Contributed reagents/ materials/analysis tools: MWP BAJS. Wrote the paper: MWP BAJS LJH. 28. Etienne RS, Haegeman B, Stadler T, Aze T, Pearson PN, et al. (2012) Diversitydependence brings molecular phylogenies closer to agreement with the fossil record. Proceedings of the Royal Society B: Biological Sciences 279: Alroy J (2008) Dynamics of origination and extinction in the marine fossil record. Proceedings of the National Academy of Sciences 105: Benton MJ, Emerson BC (2007) How did life become so diverse? the dynamics of diversification according to the fossil record and molecular phylogenetics. Palaeontology 50: Wiens JJ (2011) The causes of species richness patterns across space, time, and clades and the role of ecological limits. Quarterly Review of Biology 86: Slowinski JB, Guyer C (1989) Testing the stochasticity of patterns of organismal diversity an improved null model. American Naturalist 134: Slowinski JB, Guyer C (1993) Testing whether certain traits have caused amplified diversification - an improved method based on a model of random speciation and extinction. American Naturalist 142: Price T (2007) Speciation in Birds. Roberts & Co. Publishers. 35. Stadler T (2008) Lineages-through-time plots of neutral models for speciation. Mathematical Biosciences 216: Rabosky DL, Lovette IJ (2008) Explosive evolutionary radiations: Decreasing speciation or increasing extinction through time? Evolution 62: Fordyce JA (2010) Interpreting the gamma statistic in phylogenetic diversification rate studies: A rate decrease does not necessarily indicate an early burst. Plos One Bokma F (2009) Problems detecting density-dependent diversification on phylogenies. Proceedings of the Royal Society B-Biological Sciences 276: Rabosky DL (2010) Extinction rates should not be estimated from molecular phylogenies. Evolution 64: O Meara BC (2010) New heuristic methods for joint species delimitation and species tree inference. Systematic Biology 59: Yang Z, Rannala B (2010) Bayesian species delimitation using multilocus sequence data. Proceedings of the National Academy of Sciences 107: Kubatko LS, Gibbs HL, Bloomquist EW (2011) Inferring species-level phylogenies and taxonomic distinctiveness using multilocus data in sistrurus rattlesnakes. Systematic Biology 60: Slater GJ, Price SA, Santini F, Alfaro ME (2010) Diversity versus disparity and the radiation of modern cetaceans. Proceedings of the Royal Society B- Biological Sciences 277: Alfaro ME, Santini F, Brock C, Alamillo H, Dornburg A, et al. (2009) Nine exceptional radiations plus high turnover explain species diversity in jawed vertebrates. Proceedings of the National Academy of Sciences 106: Quental TB, Marshall CR (2009) Extinction during evolutionary radiations: Reconciling the fossil record with molecular phylogenies. Evolution 63: Stadler T (2011) Simulating trees with a fixed number of extant species. Systematic Biology 60: Nee S, May RM, Harvey PH (1994) The reconstructed evolutionary process. Philosophical Transactions of the Royal Society: Biological Sciences 344: Rabosky DL (2006) Laser: A maximum likelihood toolkit for detecting temporal shifts in diversi-fication rates from molecular phylogenies. Evolutionary Bioinformatics 2: Foote M, Hunter JP, Janis CM, Sepkoski JJ (1999) Evolutionary and preservational constraints on origins of biologic groups: Divergence times of eutherian mammals. Science 283: Paradis E, Claude J, Strimmer K (2004) Ape: Analyses of phylogenetics and evolution in r language. Bioinformatics 20: Griffiths RC, Tavare S (1994) Sampling theory for neutral alleles in a varying environment. Philosophical Transactions of the Royal Society B: Biological Sciences 344: Burnham KP, Anderson DR (2002) Model Selection and Multimodel Inference. Springer. 53. R Development Core Team (2011) R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing, Vienna, Austria. URL ISBN PLOS ONE 7 September 2012 Volume 7 Issue 9 e43348
Inferring the Dynamics of Diversification: A Coalescent Approach
 Inferring the Dynamics of Diversification: A Coalescent Approach Hélène Morlon 1 *, Matthew D. Potts 2, Joshua B. Plotkin 1 * 1 Department of Biology, University of Pennsylvania, Philadelphia, Pennsylvania,
Inferring the Dynamics of Diversification: A Coalescent Approach Hélène Morlon 1 *, Matthew D. Potts 2, Joshua B. Plotkin 1 * 1 Department of Biology, University of Pennsylvania, Philadelphia, Pennsylvania,
Phylogenetic approaches for studying diversification
 Ecology Letters, (2014) doi: 10.1111/ele.12251 REVIEW AND SYNTHESIS Phylogenetic approaches for studying diversification Helene Morlon* Center for Applied Mathematics, Ecole Polytechnique, Palaiseau, Essonne,
Ecology Letters, (2014) doi: 10.1111/ele.12251 REVIEW AND SYNTHESIS Phylogenetic approaches for studying diversification Helene Morlon* Center for Applied Mathematics, Ecole Polytechnique, Palaiseau, Essonne,
Phylogeny-Based Analyses of Evolution with a Paleo-Focus Part 2. Diversification of Lineages (Mar 02, 2012 David W. Bapst
 Phylogeny-Based Analyses of Evolution with a Paleo-Focus Part 2. Diversification of Lineages (Mar 02, 2012 David W. Bapst Recommended Reading: Nee, S. 2006. Birth-Death Models in Macroevolution. Annual
Phylogeny-Based Analyses of Evolution with a Paleo-Focus Part 2. Diversification of Lineages (Mar 02, 2012 David W. Bapst Recommended Reading: Nee, S. 2006. Birth-Death Models in Macroevolution. Annual
EXTINCTION RATES SHOULD NOT BE ESTIMATED FROM MOLECULAR PHYLOGENIES
 ORIGINAL ARTICLE doi:10.1111/j.1558-5646.2009.00926.x EXTINCTION RATES SHOULD NOT BE ESTIMATED FROM MOLECULAR PHYLOGENIES Daniel L. Rabosky 1,2,3,4 1 Department of Ecology and Evolutionary Biology, Cornell
ORIGINAL ARTICLE doi:10.1111/j.1558-5646.2009.00926.x EXTINCTION RATES SHOULD NOT BE ESTIMATED FROM MOLECULAR PHYLOGENIES Daniel L. Rabosky 1,2,3,4 1 Department of Ecology and Evolutionary Biology, Cornell
Ecological Limits on Clade Diversification in Higher Taxa
 vol. 173, no. 5 the american naturalist may 2009 Ecological Limits on Clade Diversification in Higher Taxa Daniel L. Rabosky * Department of Ecology and Evolutionary Biology, Cornell University, Ithaca,
vol. 173, no. 5 the american naturalist may 2009 Ecological Limits on Clade Diversification in Higher Taxa Daniel L. Rabosky * Department of Ecology and Evolutionary Biology, Cornell University, Ithaca,
APPENDIX S1: DESCRIPTION OF THE ESTIMATION OF THE VARIANCES OF OUR MAXIMUM LIKELIHOOD ESTIMATORS
 APPENDIX S1: DESCRIPTION OF THE ESTIMATION OF THE VARIANCES OF OUR MAXIMUM LIKELIHOOD ESTIMATORS For large n, the negative inverse of the second derivative of the likelihood function at the optimum can
APPENDIX S1: DESCRIPTION OF THE ESTIMATION OF THE VARIANCES OF OUR MAXIMUM LIKELIHOOD ESTIMATORS For large n, the negative inverse of the second derivative of the likelihood function at the optimum can
EXPLOSIVE EVOLUTIONARY RADIATIONS: DECREASING SPECIATION OR INCREASING EXTINCTION THROUGH TIME?
 ORIGINAL ARTICLE doi:10.1111/j.1558-5646.2008.00409.x EXPLOSIVE EVOLUTIONARY RADIATIONS: DECREASING SPECIATION OR INCREASING EXTINCTION THROUGH TIME? Daniel L. Rabosky 1,2,3 and Irby J. Lovette 1,3 1 Department
ORIGINAL ARTICLE doi:10.1111/j.1558-5646.2008.00409.x EXPLOSIVE EVOLUTIONARY RADIATIONS: DECREASING SPECIATION OR INCREASING EXTINCTION THROUGH TIME? Daniel L. Rabosky 1,2,3 and Irby J. Lovette 1,3 1 Department
POSITIVE CORRELATION BETWEEN DIVERSIFICATION RATES AND PHENOTYPIC EVOLVABILITY CAN MIMIC PUNCTUATED EQUILIBRIUM ON MOLECULAR PHYLOGENIES
 doi:10.1111/j.1558-5646.2012.01631.x POSITIVE CORRELATION BETWEEN DIVERSIFICATION RATES AND PHENOTYPIC EVOLVABILITY CAN MIMIC PUNCTUATED EQUILIBRIUM ON MOLECULAR PHYLOGENIES Daniel L. Rabosky 1,2,3 1 Department
doi:10.1111/j.1558-5646.2012.01631.x POSITIVE CORRELATION BETWEEN DIVERSIFICATION RATES AND PHENOTYPIC EVOLVABILITY CAN MIMIC PUNCTUATED EQUILIBRIUM ON MOLECULAR PHYLOGENIES Daniel L. Rabosky 1,2,3 1 Department
The Tempo of Macroevolution: Patterns of Diversification and Extinction
 The Tempo of Macroevolution: Patterns of Diversification and Extinction During the semester we have been consider various aspects parameters associated with biodiversity. Current usage stems from 1980's
The Tempo of Macroevolution: Patterns of Diversification and Extinction During the semester we have been consider various aspects parameters associated with biodiversity. Current usage stems from 1980's
Center for Applied Mathematics, École Polytechnique, UMR 7641 CNRS, Route de Saclay,
 1 Opinion Paper for Trends in Ecology and Evolution (TREE) 2 3 Title: Why does diversification slow down? 4 5 Authors: Daniel Moen 1 and Hélène Morlon 6 7 8 9 10 Center for Applied Mathematics, École Polytechnique,
1 Opinion Paper for Trends in Ecology and Evolution (TREE) 2 3 Title: Why does diversification slow down? 4 5 Authors: Daniel Moen 1 and Hélène Morlon 6 7 8 9 10 Center for Applied Mathematics, École Polytechnique,
From Individual-based Population Models to Lineage-based Models of Phylogenies
 From Individual-based Population Models to Lineage-based Models of Phylogenies Amaury Lambert (joint works with G. Achaz, H.K. Alexander, R.S. Etienne, N. Lartillot, H. Morlon, T.L. Parsons, T. Stadler)
From Individual-based Population Models to Lineage-based Models of Phylogenies Amaury Lambert (joint works with G. Achaz, H.K. Alexander, R.S. Etienne, N. Lartillot, H. Morlon, T.L. Parsons, T. Stadler)
paleotree: an R package for paleontological and phylogenetic analyses of evolution
 Methods in Ecology and Evolution 2012, 3, 803 807 doi: 10.1111/j.2041-210X.2012.00223.x APPLICATION paleotree: an R package for paleontological and phylogenetic analyses of evolution David W. Bapst* Department
Methods in Ecology and Evolution 2012, 3, 803 807 doi: 10.1111/j.2041-210X.2012.00223.x APPLICATION paleotree: an R package for paleontological and phylogenetic analyses of evolution David W. Bapst* Department
Three Monte Carlo Models. of Faunal Evolution PUBLISHED BY NATURAL HISTORY THE AMERICAN MUSEUM SYDNEY ANDERSON AND CHARLES S.
 AMERICAN MUSEUM Notltates PUBLISHED BY THE AMERICAN MUSEUM NATURAL HISTORY OF CENTRAL PARK WEST AT 79TH STREET NEW YORK, N.Y. 10024 U.S.A. NUMBER 2563 JANUARY 29, 1975 SYDNEY ANDERSON AND CHARLES S. ANDERSON
AMERICAN MUSEUM Notltates PUBLISHED BY THE AMERICAN MUSEUM NATURAL HISTORY OF CENTRAL PARK WEST AT 79TH STREET NEW YORK, N.Y. 10024 U.S.A. NUMBER 2563 JANUARY 29, 1975 SYDNEY ANDERSON AND CHARLES S. ANDERSON
BAMMtools: an R package for the analysis of evolutionary dynamics on phylogenetic trees
 Methods in Ecology and Evolution 2, 5, 71 77 doi: 1.1111/241-21X.12199 APPLICATION BAMMtools: an R package for the analysis of evolutionary dynamics on phylogenetic trees Daniel L. Rabosky*, Michael Grundler,
Methods in Ecology and Evolution 2, 5, 71 77 doi: 1.1111/241-21X.12199 APPLICATION BAMMtools: an R package for the analysis of evolutionary dynamics on phylogenetic trees Daniel L. Rabosky*, Michael Grundler,
Diversity dynamics: molecular phylogenies need the fossil record
 Opinion Diversity dynamics: molecular phylogenies need the fossil record Tiago B. Quental and Charles R. Marshall Museum of Paleontology and Department of Integrative Biology, University of California
Opinion Diversity dynamics: molecular phylogenies need the fossil record Tiago B. Quental and Charles R. Marshall Museum of Paleontology and Department of Integrative Biology, University of California
Markov chain Monte-Carlo to estimate speciation and extinction rates: making use of the forest hidden behind the (phylogenetic) tree
 Markov chain Monte-Carlo to estimate speciation and extinction rates: making use of the forest hidden behind the (phylogenetic) tree Nicolas Salamin Department of Ecology and Evolution University of Lausanne
Markov chain Monte-Carlo to estimate speciation and extinction rates: making use of the forest hidden behind the (phylogenetic) tree Nicolas Salamin Department of Ecology and Evolution University of Lausanne
Appendix from L. J. Revell, On the Analysis of Evolutionary Change along Single Branches in a Phylogeny
 008 by The University of Chicago. All rights reserved.doi: 10.1086/588078 Appendix from L. J. Revell, On the Analysis of Evolutionary Change along Single Branches in a Phylogeny (Am. Nat., vol. 17, no.
008 by The University of Chicago. All rights reserved.doi: 10.1086/588078 Appendix from L. J. Revell, On the Analysis of Evolutionary Change along Single Branches in a Phylogeny (Am. Nat., vol. 17, no.
Models of Continuous Trait Evolution
 Models of Continuous Trait Evolution By Ricardo Betancur (slides courtesy of Graham Slater, J. Arbour, L. Harmon & M. Alfaro) How fast do animals evolve? C. Darwin G. Simpson J. Haldane How do we model
Models of Continuous Trait Evolution By Ricardo Betancur (slides courtesy of Graham Slater, J. Arbour, L. Harmon & M. Alfaro) How fast do animals evolve? C. Darwin G. Simpson J. Haldane How do we model
Integrating Fossils into Phylogenies. Throughout the 20th century, the relationship between paleontology and evolutionary biology has been strained.
 IB 200B Principals of Phylogenetic Systematics Spring 2011 Integrating Fossils into Phylogenies Throughout the 20th century, the relationship between paleontology and evolutionary biology has been strained.
IB 200B Principals of Phylogenetic Systematics Spring 2011 Integrating Fossils into Phylogenies Throughout the 20th century, the relationship between paleontology and evolutionary biology has been strained.
Testing macro-evolutionary models using incomplete molecular phylogenies
 doi 10.1098/rspb.2000.1278 Testing macro-evolutionary models using incomplete molecular phylogenies Oliver G. Pybus * and Paul H. Harvey Department of Zoology, University of Oxford, South Parks Road, Oxford
doi 10.1098/rspb.2000.1278 Testing macro-evolutionary models using incomplete molecular phylogenies Oliver G. Pybus * and Paul H. Harvey Department of Zoology, University of Oxford, South Parks Road, Oxford
Goldilocks Meets Santa Rosalia: An Ephemeral Speciation Model Explains Patterns of Diversification Across Time Scales
 DOI 10.1007/s11692-012-9171-x SYNTHESIS PAPER Goldilocks Meets Santa Rosalia: An Ephemeral Speciation Model Explains Patterns of Diversification Across Time Scales Erica Bree Rosenblum Brice A. J. Sarver
DOI 10.1007/s11692-012-9171-x SYNTHESIS PAPER Goldilocks Meets Santa Rosalia: An Ephemeral Speciation Model Explains Patterns of Diversification Across Time Scales Erica Bree Rosenblum Brice A. J. Sarver
PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION
 "PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200 Spring 2018 University of California, Berkeley D. Ackerly April 2, 2018 Lineage diversification Reading: Morlon, H. (2014).
"PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200 Spring 2018 University of California, Berkeley D. Ackerly April 2, 2018 Lineage diversification Reading: Morlon, H. (2014).
PO Box 11103, 9700 CC, Groningen, The Netherlands 2 INRIA research team MERE, UMR Systems Analysis and Biometrics, 2 place Pierre Viala,
 Downloaded from http://rspb.royalsocietypublishing.org/ on June 8, 208 Diversity-dependence brings molecular phylogenies closer to agreement with the fossil record Rampal S. Etienne, *, Bart Haegeman 2,
Downloaded from http://rspb.royalsocietypublishing.org/ on June 8, 208 Diversity-dependence brings molecular phylogenies closer to agreement with the fossil record Rampal S. Etienne, *, Bart Haegeman 2,
Does convergent evolution always result from different lineages experiencing similar
 Different evolutionary dynamics led to the convergence of clinging performance in lizard toepads Sarin Tiatragula 1 *, Gopal Murali 2 *, James T. Stroud 3 1 Department of Biological Sciences, Auburn University,
Different evolutionary dynamics led to the convergence of clinging performance in lizard toepads Sarin Tiatragula 1 *, Gopal Murali 2 *, James T. Stroud 3 1 Department of Biological Sciences, Auburn University,
Bayesian Estimation of Speciation and Extinction from Incomplete Fossil Occurrence Data
 Syst. Biol. 63(3):349 367, 2014 The Author(s) 2014. Published by Oxford University Press, on behalf of the Society of Systematic Biologists. All rights reserved. For Permissions, please email: journals.permissions@oup.com
Syst. Biol. 63(3):349 367, 2014 The Author(s) 2014. Published by Oxford University Press, on behalf of the Society of Systematic Biologists. All rights reserved. For Permissions, please email: journals.permissions@oup.com
Comparing the rates of speciation and extinction between phylogenetic trees
 Received: 29 November 2017 Revised: 21 February 2018 Accepted: 8 March 2018 DOI: 10.1002/ece3.4030 ORIGINAL RESEARCH Comparing the rates of speciation and extinction between phylogenetic trees Liam J.
Received: 29 November 2017 Revised: 21 February 2018 Accepted: 8 March 2018 DOI: 10.1002/ece3.4030 ORIGINAL RESEARCH Comparing the rates of speciation and extinction between phylogenetic trees Liam J.
arxiv: v1 [q-bio.pe] 22 Apr 2014 Sacha S.J. Laurent 12, Marc Robinson-Rechavi 12, and Nicolas Salamin 12
![arxiv: v1 [q-bio.pe] 22 Apr 2014 Sacha S.J. Laurent 12, Marc Robinson-Rechavi 12, and Nicolas Salamin 12 arxiv: v1 [q-bio.pe] 22 Apr 2014 Sacha S.J. Laurent 12, Marc Robinson-Rechavi 12, and Nicolas Salamin 12](/thumbs/79/79272132.jpg) Version dated: April 23, 2014 Are we able to detect mass extinction events using phylogenies? arxiv:1404.5441v1 [q-bio.pe] 22 Apr 2014 Sacha S.J. Laurent 12, Marc Robinson-Rechavi 12, and Nicolas Salamin
Version dated: April 23, 2014 Are we able to detect mass extinction events using phylogenies? arxiv:1404.5441v1 [q-bio.pe] 22 Apr 2014 Sacha S.J. Laurent 12, Marc Robinson-Rechavi 12, and Nicolas Salamin
Evolutionary models with ecological interactions. By: Magnus Clarke
 Evolutionary models with ecological interactions By: Magnus Clarke A thesis submitted in partial fulfilment of the requirements for the degree of Doctor of Philosophy The University of Sheffield Faculty
Evolutionary models with ecological interactions By: Magnus Clarke A thesis submitted in partial fulfilment of the requirements for the degree of Doctor of Philosophy The University of Sheffield Faculty
Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut
 Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic analysis Phylogenetic Basics: Biological
Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic analysis Phylogenetic Basics: Biological
MOTMOT: models of trait macroevolution on trees
 Methods in Ecology and Evolution 2012, 3, 145 151 doi: 10.1111/j.2041-210X.2011.00132.x APPLICATION MOTMOT: models of trait macroevolution on trees Gavin H. Thomas 1 * and Robert P. Freckleton 2 1 School
Methods in Ecology and Evolution 2012, 3, 145 151 doi: 10.1111/j.2041-210X.2011.00132.x APPLICATION MOTMOT: models of trait macroevolution on trees Gavin H. Thomas 1 * and Robert P. Freckleton 2 1 School
Estimating Age-Dependent Extinction: Contrasting Evidence from Fossils and Phylogenies
 Syst. Biol. 0(0):1 17, 2017 The Author(s) 2017. Published by Oxford University Press, on behalf of the Society of Systematic Biologists. This is an Open Access article distributed under the terms of the
Syst. Biol. 0(0):1 17, 2017 The Author(s) 2017. Published by Oxford University Press, on behalf of the Society of Systematic Biologists. This is an Open Access article distributed under the terms of the
Integrative Biology 200 "PRINCIPLES OF PHYLOGENETICS" Spring 2018 University of California, Berkeley
 Integrative Biology 200 "PRINCIPLES OF PHYLOGENETICS" Spring 2018 University of California, Berkeley B.D. Mishler Feb. 14, 2018. Phylogenetic trees VI: Dating in the 21st century: clocks, & calibrations;
Integrative Biology 200 "PRINCIPLES OF PHYLOGENETICS" Spring 2018 University of California, Berkeley B.D. Mishler Feb. 14, 2018. Phylogenetic trees VI: Dating in the 21st century: clocks, & calibrations;
Dr. Amira A. AL-Hosary
 Phylogenetic analysis Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic Basics: Biological
Phylogenetic analysis Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic Basics: Biological
Using the Birth-Death Process to Infer Changes in the Pattern of Lineage Gain and Loss. Nathaniel Malachi Hallinan
 Using the Birth-Death Process to Infer Changes in the Pattern of Lineage Gain and Loss by Nathaniel Malachi Hallinan A dissertation submitted in partial satisfaction of the requirements for the degree
Using the Birth-Death Process to Infer Changes in the Pattern of Lineage Gain and Loss by Nathaniel Malachi Hallinan A dissertation submitted in partial satisfaction of the requirements for the degree
Model Inadequacy and Mistaken Inferences of Trait-Dependent Speciation
 1 Model Inadequacy and Mistaken Inferences of Trait-Dependent Speciation Daniel L. Rabosky 1* and Emma E. Goldberg 2* 1 Museum of Zoology and Department of Ecology and Evolutionary Biology, University
1 Model Inadequacy and Mistaken Inferences of Trait-Dependent Speciation Daniel L. Rabosky 1* and Emma E. Goldberg 2* 1 Museum of Zoology and Department of Ecology and Evolutionary Biology, University
A Rough Guide to CladeAge
 A Rough Guide to CladeAge Setting fossil calibrations in BEAST 2 Michael Matschiner Remco Bouckaert michaelmatschiner@mac.com remco@cs.auckland.ac.nz January 17, 2014 Contents 1 Introduction 1 1.1 Windows...............................
A Rough Guide to CladeAge Setting fossil calibrations in BEAST 2 Michael Matschiner Remco Bouckaert michaelmatschiner@mac.com remco@cs.auckland.ac.nz January 17, 2014 Contents 1 Introduction 1 1.1 Windows...............................
Biol 206/306 Advanced Biostatistics Lab 11 Models of Trait Evolution Fall 2016
 Biol 206/306 Advanced Biostatistics Lab 11 Models of Trait Evolution Fall 2016 By Philip J. Bergmann 0. Laboratory Objectives 1. Explore how evolutionary trait modeling can reveal different information
Biol 206/306 Advanced Biostatistics Lab 11 Models of Trait Evolution Fall 2016 By Philip J. Bergmann 0. Laboratory Objectives 1. Explore how evolutionary trait modeling can reveal different information
A Simple Method for Estimating Informative Node Age Priors for the Fossil Calibration of Molecular Divergence Time Analyses
 A Simple Method for Estimating Informative Node Age Priors for the Fossil Calibration of Molecular Divergence Time Analyses Michael D. Nowak 1 *, Andrew B. Smith 2, Carl Simpson 3, Derrick J. Zwickl 4
A Simple Method for Estimating Informative Node Age Priors for the Fossil Calibration of Molecular Divergence Time Analyses Michael D. Nowak 1 *, Andrew B. Smith 2, Carl Simpson 3, Derrick J. Zwickl 4
Taming the Beast Workshop
 Workshop and Chi Zhang June 28, 2016 1 / 19 Species tree Species tree the phylogeny representing the relationships among a group of species Figure adapted from [Rogers and Gibbs, 2014] Gene tree the phylogeny
Workshop and Chi Zhang June 28, 2016 1 / 19 Species tree Species tree the phylogeny representing the relationships among a group of species Figure adapted from [Rogers and Gibbs, 2014] Gene tree the phylogeny
DNA-based species delimitation
 DNA-based species delimitation Phylogenetic species concept based on tree topologies Ø How to set species boundaries? Ø Automatic species delimitation? druhů? DNA barcoding Species boundaries recognized
DNA-based species delimitation Phylogenetic species concept based on tree topologies Ø How to set species boundaries? Ø Automatic species delimitation? druhů? DNA barcoding Species boundaries recognized
Testing the time-for-speciation effect in the assembly of regional biotas
 Methods in Ecology and Evolution 2012, 3, 224 233 Testing the time-for-speciation effect in the assembly of regional biotas Daniel L. Rabosky doi: 10.1111/j.2041-210X.2011.00166.x Department of Integrative
Methods in Ecology and Evolution 2012, 3, 224 233 Testing the time-for-speciation effect in the assembly of regional biotas Daniel L. Rabosky doi: 10.1111/j.2041-210X.2011.00166.x Department of Integrative
Model Inadequacy and Mistaken Inferences of Trait-Dependent Speciation
 Syst. Biol. 64(2):340 355, 2015 The Author(s) 2015. Published by Oxford University Press, on behalf of the Society of Systematic Biologists. All rights reserved. For Permissions, please email: journals.permissions@oup.com
Syst. Biol. 64(2):340 355, 2015 The Author(s) 2015. Published by Oxford University Press, on behalf of the Society of Systematic Biologists. All rights reserved. For Permissions, please email: journals.permissions@oup.com
Using phylogenetics to estimate species divergence times... Basics and basic issues for Bayesian inference of divergence times (plus some digression)
 Using phylogenetics to estimate species divergence times... More accurately... Basics and basic issues for Bayesian inference of divergence times (plus some digression) "A comparison of the structures
Using phylogenetics to estimate species divergence times... More accurately... Basics and basic issues for Bayesian inference of divergence times (plus some digression) "A comparison of the structures
"PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200B Spring 2009 University of California, Berkeley
 "PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200B Spring 2009 University of California, Berkeley B.D. Mishler Jan. 22, 2009. Trees I. Summary of previous lecture: Hennigian
"PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200B Spring 2009 University of California, Berkeley B.D. Mishler Jan. 22, 2009. Trees I. Summary of previous lecture: Hennigian
Postal address: Ruthven Museums Building, Museum of Zoology, University of Michigan, Ann Arbor,
 1 RH: RATE-SHIFT MODELS IN MACROEVOLUTION Is BAMM flawed? Theoretical and practical concerns in the analysis of multi-rate diversification models Daniel L. Rabosky 1,*, Jonathan S. Mitchell 1, and Jonathan
1 RH: RATE-SHIFT MODELS IN MACROEVOLUTION Is BAMM flawed? Theoretical and practical concerns in the analysis of multi-rate diversification models Daniel L. Rabosky 1,*, Jonathan S. Mitchell 1, and Jonathan
FiSSE: A simple nonparametric test for the effects of a binary character on lineage diversification rates
 ORIGINAL ARTICLE doi:0./evo.3227 FiSSE: A simple nonparametric test for the effects of a binary character on lineage diversification rates Daniel L. Rabosky,2, and Emma E. Goldberg 3, Department of Ecology
ORIGINAL ARTICLE doi:0./evo.3227 FiSSE: A simple nonparametric test for the effects of a binary character on lineage diversification rates Daniel L. Rabosky,2, and Emma E. Goldberg 3, Department of Ecology
Non-independence in Statistical Tests for Discrete Cross-species Data
 J. theor. Biol. (1997) 188, 507514 Non-independence in Statistical Tests for Discrete Cross-species Data ALAN GRAFEN* AND MARK RIDLEY * St. John s College, Oxford OX1 3JP, and the Department of Zoology,
J. theor. Biol. (1997) 188, 507514 Non-independence in Statistical Tests for Discrete Cross-species Data ALAN GRAFEN* AND MARK RIDLEY * St. John s College, Oxford OX1 3JP, and the Department of Zoology,
STEM-hy: Species Tree Estimation using Maximum likelihood (with hybridization)
 STEM-hy: Species Tree Estimation using Maximum likelihood (with hybridization) Laura Salter Kubatko Departments of Statistics and Evolution, Ecology, and Organismal Biology The Ohio State University kubatko.2@osu.edu
STEM-hy: Species Tree Estimation using Maximum likelihood (with hybridization) Laura Salter Kubatko Departments of Statistics and Evolution, Ecology, and Organismal Biology The Ohio State University kubatko.2@osu.edu
Phylogenies support out-of-equilibrium models of biodiversity
 Ecology Letters, (2015) doi: 10.1111/ele.12415 LETTER Phylogenies support out-of-equilibrium models of biodiversity Marc Manceau, 1,2 Amaury Lambert, 2,3 and Helene Morlon, 1,2 Abstract There is a long
Ecology Letters, (2015) doi: 10.1111/ele.12415 LETTER Phylogenies support out-of-equilibrium models of biodiversity Marc Manceau, 1,2 Amaury Lambert, 2,3 and Helene Morlon, 1,2 Abstract There is a long
Phylogenetic comparative methods, CEB Spring 2010 [Revised April 22, 2010]
![Phylogenetic comparative methods, CEB Spring 2010 [Revised April 22, 2010] Phylogenetic comparative methods, CEB Spring 2010 [Revised April 22, 2010]](/thumbs/93/112416870.jpg) Phylogenetic comparative methods, CEB 35300 Spring 2010 [Revised April 22, 2010] Instructors: Andrew Hipp, Herbarium Curator, Morton Arboretum. ahipp@mortonarb.org; 630-725-2094 Rick Ree, Associate Curator,
Phylogenetic comparative methods, CEB 35300 Spring 2010 [Revised April 22, 2010] Instructors: Andrew Hipp, Herbarium Curator, Morton Arboretum. ahipp@mortonarb.org; 630-725-2094 Rick Ree, Associate Curator,
"PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200 Spring 2018 University of California, Berkeley
 "PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200 Spring 2018 University of California, Berkeley D.D. Ackerly Feb. 26, 2018 Maximum Likelihood Principles, and Applications to
"PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200 Spring 2018 University of California, Berkeley D.D. Ackerly Feb. 26, 2018 Maximum Likelihood Principles, and Applications to
Greater host breadth still not associated with increased diversification rate in the Nymphalidae A response to Janz et al.
 doi:10.1111/evo.12914 Greater host breadth still not associated with increased diversification rate in the Nymphalidae A response to Janz et al. Christopher A. Hamm 1,2 and James A. Fordyce 3 1 Department
doi:10.1111/evo.12914 Greater host breadth still not associated with increased diversification rate in the Nymphalidae A response to Janz et al. Christopher A. Hamm 1,2 and James A. Fordyce 3 1 Department
Lesson 1 Syllabus Reference
 Lesson 1 Syllabus Reference Outcomes A student Explains how biological understanding has advanced through scientific discoveries, technological developments and the needs of society. Content The theory
Lesson 1 Syllabus Reference Outcomes A student Explains how biological understanding has advanced through scientific discoveries, technological developments and the needs of society. Content The theory
Inferring Ecological Processes from Taxonomic, Phylogenetic and Functional Trait b-diversity
 Inferring Ecological Processes from Taxonomic, Phylogenetic and Functional Trait b-diversity James C. Stegen*, Allen H. Hurlbert Department of Biology, University of North Carolina, Chapel Hill, North
Inferring Ecological Processes from Taxonomic, Phylogenetic and Functional Trait b-diversity James C. Stegen*, Allen H. Hurlbert Department of Biology, University of North Carolina, Chapel Hill, North
Supplementary Figure 1: Distribution of carnivore species within the Jetz et al. phylogeny. The phylogeny presented here was randomly sampled from
 Supplementary Figure 1: Distribution of carnivore species within the Jetz et al. phylogeny. The phylogeny presented here was randomly sampled from the 1 trees from the collection using the Ericson backbone.
Supplementary Figure 1: Distribution of carnivore species within the Jetz et al. phylogeny. The phylogeny presented here was randomly sampled from the 1 trees from the collection using the Ericson backbone.
Phylogenomics. Jeffrey P. Townsend Department of Ecology and Evolutionary Biology Yale University. Tuesday, January 29, 13
 Phylogenomics Jeffrey P. Townsend Department of Ecology and Evolutionary Biology Yale University How may we improve our inferences? How may we improve our inferences? Inferences Data How may we improve
Phylogenomics Jeffrey P. Townsend Department of Ecology and Evolutionary Biology Yale University How may we improve our inferences? How may we improve our inferences? Inferences Data How may we improve
ESTIMATION OF CONSERVATISM OF CHARACTERS BY CONSTANCY WITHIN BIOLOGICAL POPULATIONS
 ESTIMATION OF CONSERVATISM OF CHARACTERS BY CONSTANCY WITHIN BIOLOGICAL POPULATIONS JAMES S. FARRIS Museum of Zoology, The University of Michigan, Ann Arbor Accepted March 30, 1966 The concept of conservatism
ESTIMATION OF CONSERVATISM OF CHARACTERS BY CONSTANCY WITHIN BIOLOGICAL POPULATIONS JAMES S. FARRIS Museum of Zoology, The University of Michigan, Ann Arbor Accepted March 30, 1966 The concept of conservatism
Discrete & continuous characters: The threshold model
 Discrete & continuous characters: The threshold model Discrete & continuous characters: the threshold model So far we have discussed continuous & discrete character models separately for estimating ancestral
Discrete & continuous characters: The threshold model Discrete & continuous characters: the threshold model So far we have discussed continuous & discrete character models separately for estimating ancestral
Integrative Biology 200A "PRINCIPLES OF PHYLOGENETICS" Spring 2008
 Integrative Biology 200A "PRINCIPLES OF PHYLOGENETICS" Spring 2008 University of California, Berkeley B.D. Mishler March 18, 2008. Phylogenetic Trees I: Reconstruction; Models, Algorithms & Assumptions
Integrative Biology 200A "PRINCIPLES OF PHYLOGENETICS" Spring 2008 University of California, Berkeley B.D. Mishler March 18, 2008. Phylogenetic Trees I: Reconstruction; Models, Algorithms & Assumptions
Thursday, March 21, 13. Evolution
 Evolution What is Evolution? Evolution involves inheritable changes in a population of organisms through time Fundamental to biology and paleontology Paleontology is the study of life history as revealed
Evolution What is Evolution? Evolution involves inheritable changes in a population of organisms through time Fundamental to biology and paleontology Paleontology is the study of life history as revealed
A Bayesian approach for detecting the impact of mass-extinction events on molecular phylogenies when rates of lineage diversification may vary
 Methods in Ecology and Evolution 16, 7, 947 99 doi: 1.1111/41-1X.163 A Bayesian approach for detecting the impact of mass-extinction events on molecular phylogenies when rates of lineage diversification
Methods in Ecology and Evolution 16, 7, 947 99 doi: 1.1111/41-1X.163 A Bayesian approach for detecting the impact of mass-extinction events on molecular phylogenies when rates of lineage diversification
History is written by the victors: The effect of the push of the past on the fossil record
 doi:10.1111/evo.13593 History is written by the victors: The effect of the push of the past on the fossil record Graham E. Budd 1,2 and Richard P. Mann 3 1 Department of Earth Sciences, Palaeobiology,
doi:10.1111/evo.13593 History is written by the victors: The effect of the push of the past on the fossil record Graham E. Budd 1,2 and Richard P. Mann 3 1 Department of Earth Sciences, Palaeobiology,
R eports. Incompletely resolved phylogenetic trees inflate estimates of phylogenetic conservatism
 Ecology, 93(2), 2012, pp. 242 247 Ó 2012 by the Ecological Society of America Incompletely resolved phylogenetic trees inflate estimates of phylogenetic conservatism T. JONATHAN DAVIES, 1,6 NATHAN J. B.
Ecology, 93(2), 2012, pp. 242 247 Ó 2012 by the Ecological Society of America Incompletely resolved phylogenetic trees inflate estimates of phylogenetic conservatism T. JONATHAN DAVIES, 1,6 NATHAN J. B.
5/31/17. Week 10; Monday MEMORIAL DAY NO CLASS. Page 88
 Week 10; Monday MEMORIAL DAY NO CLASS Page 88 Week 10; Wednesday Announcements: Family ID final in lab Today Final exam next Tuesday at 8:30 am here Lecture: Species concepts & Speciation. What are species?
Week 10; Monday MEMORIAL DAY NO CLASS Page 88 Week 10; Wednesday Announcements: Family ID final in lab Today Final exam next Tuesday at 8:30 am here Lecture: Species concepts & Speciation. What are species?
New Inferences from Tree Shape: Numbers of Missing Taxa and Population Growth Rates
 Syst. Biol. 51(6):881 888, 2002 DOI: 10.1080/10635150290155881 New Inferences from Tree Shape: Numbers of Missing Taxa and Population Growth Rates O. G. PYBUS,A.RAMBAUT,E.C.HOLMES, AND P. H. HARVEY Department
Syst. Biol. 51(6):881 888, 2002 DOI: 10.1080/10635150290155881 New Inferences from Tree Shape: Numbers of Missing Taxa and Population Growth Rates O. G. PYBUS,A.RAMBAUT,E.C.HOLMES, AND P. H. HARVEY Department
The origin of species richness patterns along environmental gradients: uniting explanations based on time, diversification rate and carrying capacity
 (J. Biogeogr. (2016 SPECIAL PAPER The origin of species richness patterns along environmental gradients: uniting explanations based on time, diversification rate and carrying capacity Mikael Pontarp 1,2,
(J. Biogeogr. (2016 SPECIAL PAPER The origin of species richness patterns along environmental gradients: uniting explanations based on time, diversification rate and carrying capacity Mikael Pontarp 1,2,
Bayesian estimation of multiple clade competition from fossil data
 Evolutionary Ecology Research, 2017, 18: 41 59 Bayesian estimation of multiple clade competition from fossil data Daniele Silvestro 1,2,3, Mathias M. Pires 4, Tiago B. Quental 4 and Nicolas Salamin 2,3
Evolutionary Ecology Research, 2017, 18: 41 59 Bayesian estimation of multiple clade competition from fossil data Daniele Silvestro 1,2,3, Mathias M. Pires 4, Tiago B. Quental 4 and Nicolas Salamin 2,3
Rapid evolution of the cerebellum in humans and other great apes
 Rapid evolution of the cerebellum in humans and other great apes Article Accepted Version Barton, R. A. and Venditti, C. (2014) Rapid evolution of the cerebellum in humans and other great apes. Current
Rapid evolution of the cerebellum in humans and other great apes Article Accepted Version Barton, R. A. and Venditti, C. (2014) Rapid evolution of the cerebellum in humans and other great apes. Current
Estimating a Binary Character s Effect on Speciation and Extinction
 Syst. Biol. 56(5):701 710, 2007 Copyright c Society of Systematic Biologists ISSN: 1063-5157 print / 1076-836X online DOI: 10.1080/10635150701607033 Estimating a Binary Character s Effect on Speciation
Syst. Biol. 56(5):701 710, 2007 Copyright c Society of Systematic Biologists ISSN: 1063-5157 print / 1076-836X online DOI: 10.1080/10635150701607033 Estimating a Binary Character s Effect on Speciation
Biology 211 (2) Week 1 KEY!
 Biology 211 (2) Week 1 KEY Chapter 1 KEY FIGURES: 1.2, 1.3, 1.4, 1.5, 1.6, 1.7 VOCABULARY: Adaptation: a trait that increases the fitness Cells: a developed, system bound with a thin outer layer made of
Biology 211 (2) Week 1 KEY Chapter 1 KEY FIGURES: 1.2, 1.3, 1.4, 1.5, 1.6, 1.7 VOCABULARY: Adaptation: a trait that increases the fitness Cells: a developed, system bound with a thin outer layer made of
Some of these slides have been borrowed from Dr. Paul Lewis, Dr. Joe Felsenstein. Thanks!
 Some of these slides have been borrowed from Dr. Paul Lewis, Dr. Joe Felsenstein. Thanks! Paul has many great tools for teaching phylogenetics at his web site: http://hydrodictyon.eeb.uconn.edu/people/plewis
Some of these slides have been borrowed from Dr. Paul Lewis, Dr. Joe Felsenstein. Thanks! Paul has many great tools for teaching phylogenetics at his web site: http://hydrodictyon.eeb.uconn.edu/people/plewis
3 Hours 18 / 06 / 2012 EXAMS OFFICE USE ONLY University of the Witwatersrand, Johannesburg Course or topic No(s) ANAT 4000
 3 Hours 18 / 06 / 2012 EXAMS OFFICE USE ONLY University of the Witwatersrand, Johannesburg Course or topic No(s) ANAT 4000 Course or topic name(s) Paper Number & title HUMAN BIOLOGY HONOURS: PAPER 1 Examination
3 Hours 18 / 06 / 2012 EXAMS OFFICE USE ONLY University of the Witwatersrand, Johannesburg Course or topic No(s) ANAT 4000 Course or topic name(s) Paper Number & title HUMAN BIOLOGY HONOURS: PAPER 1 Examination
ASYMMETRIES IN PHYLOGENETIC DIVERSIFICATION AND CHARACTER CHANGE CAN BE UNTANGLED
 doi:1.1111/j.1558-5646.27.252.x ASYMMETRIES IN PHYLOGENETIC DIVERSIFICATION AND CHARACTER CHANGE CAN BE UNTANGLED Emmanuel Paradis Institut de Recherche pour le Développement, UR175 CAVIAR, GAMET BP 595,
doi:1.1111/j.1558-5646.27.252.x ASYMMETRIES IN PHYLOGENETIC DIVERSIFICATION AND CHARACTER CHANGE CAN BE UNTANGLED Emmanuel Paradis Institut de Recherche pour le Développement, UR175 CAVIAR, GAMET BP 595,
Modeling continuous trait evolution in the absence of phylogenetic information
 Modeling continuous trait evolution in the absence of phylogenetic information Mariia Koroliuk GU university, EM in complex systems under the supervision of Daniele Silvestro June 29, 2014 Contents 1 Introduction
Modeling continuous trait evolution in the absence of phylogenetic information Mariia Koroliuk GU university, EM in complex systems under the supervision of Daniele Silvestro June 29, 2014 Contents 1 Introduction
Biome evolution and biogeographical change through time
 thesis abstract Biome evolution and biogeographical change through time Christine D. Bacon ISSN 1948-6596 Department of Biological and Environmental Sciences, University of Gothenburg, Gothenburg, Sweden;
thesis abstract Biome evolution and biogeographical change through time Christine D. Bacon ISSN 1948-6596 Department of Biological and Environmental Sciences, University of Gothenburg, Gothenburg, Sweden;
University of Groningen
 University of Groningen Can clade age alone explain the relationship between body size and diversity? Etienne, Rampal S.; de Visser, Sara N.; Janzen, Thijs; Olsen, Jeanine L.; Olff, Han; Rosindell, James
University of Groningen Can clade age alone explain the relationship between body size and diversity? Etienne, Rampal S.; de Visser, Sara N.; Janzen, Thijs; Olsen, Jeanine L.; Olff, Han; Rosindell, James
Integrative Biology 200 "PRINCIPLES OF PHYLOGENETICS" Spring 2016 University of California, Berkeley. Parsimony & Likelihood [draft]
![Integrative Biology 200 PRINCIPLES OF PHYLOGENETICS Spring 2016 University of California, Berkeley. Parsimony & Likelihood [draft] Integrative Biology 200 PRINCIPLES OF PHYLOGENETICS Spring 2016 University of California, Berkeley. Parsimony & Likelihood [draft]](/thumbs/83/87443456.jpg) Integrative Biology 200 "PRINCIPLES OF PHYLOGENETICS" Spring 2016 University of California, Berkeley K.W. Will Parsimony & Likelihood [draft] 1. Hennig and Parsimony: Hennig was not concerned with parsimony
Integrative Biology 200 "PRINCIPLES OF PHYLOGENETICS" Spring 2016 University of California, Berkeley K.W. Will Parsimony & Likelihood [draft] 1. Hennig and Parsimony: Hennig was not concerned with parsimony
Ecological controls of mammalian diversification vary with phylogenetic scale
 Received: 9 November 2016 Revised: 28 July 2017 Accepted: 2 August 2017 DOI: 10.1111/geb.12642 RESEARCH PAPER Ecological controls of mammalian diversification vary with phylogenetic scale Antonin Machac
Received: 9 November 2016 Revised: 28 July 2017 Accepted: 2 August 2017 DOI: 10.1111/geb.12642 RESEARCH PAPER Ecological controls of mammalian diversification vary with phylogenetic scale Antonin Machac
Four aspects of a sampling strategy necessary to make accurate and precise inferences about populations are:
 Why Sample? Often researchers are interested in answering questions about a particular population. They might be interested in the density, species richness, or specific life history parameters such as
Why Sample? Often researchers are interested in answering questions about a particular population. They might be interested in the density, species richness, or specific life history parameters such as
Historical Biogeography. Historical Biogeography. Systematics
 Historical Biogeography I. Definitions II. Fossils: problems with fossil record why fossils are important III. Phylogeny IV. Phenetics VI. Phylogenetic Classification Disjunctions debunked: Examples VII.
Historical Biogeography I. Definitions II. Fossils: problems with fossil record why fossils are important III. Phylogeny IV. Phenetics VI. Phylogenetic Classification Disjunctions debunked: Examples VII.
POPULATION GENETICS Winter 2005 Lecture 17 Molecular phylogenetics
 POPULATION GENETICS Winter 2005 Lecture 17 Molecular phylogenetics - in deriving a phylogeny our goal is simply to reconstruct the historical relationships between a group of taxa. - before we review the
POPULATION GENETICS Winter 2005 Lecture 17 Molecular phylogenetics - in deriving a phylogeny our goal is simply to reconstruct the historical relationships between a group of taxa. - before we review the
PyRate: a new program to estimate speciation and extinction rates from incomplete fossil data
 Methods in Ecology and Evolution 2014, 5, 1126 1131 doi: 10.1111/2041-210X.12263 APPLICATION PyRate: a new program to estimate speciation and extinction rates from incomplete fossil data Daniele Silvestro
Methods in Ecology and Evolution 2014, 5, 1126 1131 doi: 10.1111/2041-210X.12263 APPLICATION PyRate: a new program to estimate speciation and extinction rates from incomplete fossil data Daniele Silvestro
ECOLOGICAL CAUSES OF DECELERATING DIVERSIFICATION IN CARNIVORAN MAMMALS
 ORIGINAL ARTICLE doi:10.1111/evo.12126 ECOLOGICAL CAUSES OF DECELERATING DIVERSIFICATION IN CARNIVORAN MAMMALS Antonin Machac, 1,2,3,4 David Storch, 2,3 and John J. Wiens 1,5 1 Department of Ecology and
ORIGINAL ARTICLE doi:10.1111/evo.12126 ECOLOGICAL CAUSES OF DECELERATING DIVERSIFICATION IN CARNIVORAN MAMMALS Antonin Machac, 1,2,3,4 David Storch, 2,3 and John J. Wiens 1,5 1 Department of Ecology and
Chapter 27: Evolutionary Genetics
 Chapter 27: Evolutionary Genetics Student Learning Objectives Upon completion of this chapter you should be able to: 1. Understand what the term species means to biology. 2. Recognize the various patterns
Chapter 27: Evolutionary Genetics Student Learning Objectives Upon completion of this chapter you should be able to: 1. Understand what the term species means to biology. 2. Recognize the various patterns
ANALYSIS OF CHARACTER DIVERGENCE ALONG ENVIRONMENTAL GRADIENTS AND OTHER COVARIATES
 ORIGINAL ARTICLE doi:10.1111/j.1558-5646.2007.00063.x ANALYSIS OF CHARACTER DIVERGENCE ALONG ENVIRONMENTAL GRADIENTS AND OTHER COVARIATES Dean C. Adams 1,2,3 and Michael L. Collyer 1,4 1 Department of
ORIGINAL ARTICLE doi:10.1111/j.1558-5646.2007.00063.x ANALYSIS OF CHARACTER DIVERGENCE ALONG ENVIRONMENTAL GRADIENTS AND OTHER COVARIATES Dean C. Adams 1,2,3 and Michael L. Collyer 1,4 1 Department of
A (short) introduction to phylogenetics
 A (short) introduction to phylogenetics Thibaut Jombart, Marie-Pauline Beugin MRC Centre for Outbreak Analysis and Modelling Imperial College London Genetic data analysis with PR Statistics, Millport Field
A (short) introduction to phylogenetics Thibaut Jombart, Marie-Pauline Beugin MRC Centre for Outbreak Analysis and Modelling Imperial College London Genetic data analysis with PR Statistics, Millport Field
Chapter 16: Reconstructing and Using Phylogenies
 Chapter Review 1. Use the phylogenetic tree shown at the right to complete the following. a. Explain how many clades are indicated: Three: (1) chimpanzee/human, (2) chimpanzee/ human/gorilla, and (3)chimpanzee/human/
Chapter Review 1. Use the phylogenetic tree shown at the right to complete the following. a. Explain how many clades are indicated: Three: (1) chimpanzee/human, (2) chimpanzee/ human/gorilla, and (3)chimpanzee/human/
AP Biology. Cladistics
 Cladistics Kingdom Summary Review slide Review slide Classification Old 5 Kingdom system Eukaryote Monera, Protists, Plants, Fungi, Animals New 3 Domain system reflects a greater understanding of evolution
Cladistics Kingdom Summary Review slide Review slide Classification Old 5 Kingdom system Eukaryote Monera, Protists, Plants, Fungi, Animals New 3 Domain system reflects a greater understanding of evolution
What is the range of a taxon? A scaling problem at three levels: Spa9al scale Phylogene9c depth Time
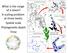 What is the range of a taxon? A scaling problem at three levels: Spa9al scale Phylogene9c depth Time 1 5 0.25 0.15 5 0.05 0.05 0.10 2 0.10 0.10 0.20 4 Reminder of what a range-weighted tree is Actual Tree
What is the range of a taxon? A scaling problem at three levels: Spa9al scale Phylogene9c depth Time 1 5 0.25 0.15 5 0.05 0.05 0.10 2 0.10 0.10 0.20 4 Reminder of what a range-weighted tree is Actual Tree
Lecture V Phylogeny and Systematics Dr. Kopeny
 Delivered 1/30 and 2/1 Lecture V Phylogeny and Systematics Dr. Kopeny Lecture V How to Determine Evolutionary Relationships: Concepts in Phylogeny and Systematics Textbook Reading: pp 425-433, 435-437
Delivered 1/30 and 2/1 Lecture V Phylogeny and Systematics Dr. Kopeny Lecture V How to Determine Evolutionary Relationships: Concepts in Phylogeny and Systematics Textbook Reading: pp 425-433, 435-437
"PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200B Spring 2011 University of California, Berkeley
 "PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200B Spring 2011 University of California, Berkeley B.D. Mishler Feb. 1, 2011. Qualitative character evolution (cont.) - comparing
"PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200B Spring 2011 University of California, Berkeley B.D. Mishler Feb. 1, 2011. Qualitative character evolution (cont.) - comparing
Wright-Fisher Models, Approximations, and Minimum Increments of Evolution
 Wright-Fisher Models, Approximations, and Minimum Increments of Evolution William H. Press The University of Texas at Austin January 10, 2011 1 Introduction Wright-Fisher models [1] are idealized models
Wright-Fisher Models, Approximations, and Minimum Increments of Evolution William H. Press The University of Texas at Austin January 10, 2011 1 Introduction Wright-Fisher models [1] are idealized models
Anatomy of a species tree
 Anatomy of a species tree T 1 Size of current and ancestral Populations (N) N Confidence in branches of species tree t/2n = 1 coalescent unit T 2 Branch lengths and divergence times of species & populations
Anatomy of a species tree T 1 Size of current and ancestral Populations (N) N Confidence in branches of species tree t/2n = 1 coalescent unit T 2 Branch lengths and divergence times of species & populations
How can molecular phylogenies illuminate morphological evolution?
 How can molecular phylogenies illuminate morphological evolution? 27 October 2016. Joe Felsenstein UNAM How can molecular phylogenies illuminate morphological evolution? p.1/81 Where this lecture fits
How can molecular phylogenies illuminate morphological evolution? 27 October 2016. Joe Felsenstein UNAM How can molecular phylogenies illuminate morphological evolution? p.1/81 Where this lecture fits
How should we organize the diversity of animal life?
 How should we organize the diversity of animal life? The difference between Taxonomy Linneaus, and Cladistics Darwin What are phylogenies? How do we read them? How do we estimate them? Classification (Taxonomy)
How should we organize the diversity of animal life? The difference between Taxonomy Linneaus, and Cladistics Darwin What are phylogenies? How do we read them? How do we estimate them? Classification (Taxonomy)
Quantitative Traits and Diversification
 Syst. Biol. 59(6):619 633, 2010 c The Author(s) 2010. Published by Oxford University Press, on behalf of the Society of Systematic Biologists. All rights reserved. For Permissions, please email: journals.permissions@oxfordjournals.org
Syst. Biol. 59(6):619 633, 2010 c The Author(s) 2010. Published by Oxford University Press, on behalf of the Society of Systematic Biologists. All rights reserved. For Permissions, please email: journals.permissions@oxfordjournals.org
Estimating Trait-Dependent Speciation and Extinction Rates from Incompletely Resolved Phylogenies
 Syst. Biol. 58(6):595 611, 2009 c The Author(s) 2009. Published by Oxford University Press, on behalf of the Society of Systematic Biologists. All rights reserved. For Permissions, please email: journals.permissions@oxfordjournals.org
Syst. Biol. 58(6):595 611, 2009 c The Author(s) 2009. Published by Oxford University Press, on behalf of the Society of Systematic Biologists. All rights reserved. For Permissions, please email: journals.permissions@oxfordjournals.org
Historical contingency, niche conservatism and the tendency for some taxa to be more diverse towards the poles
 Electronic Supplementary Material Historical contingency, niche conservatism and the tendency for some taxa to be more diverse towards the poles Ignacio Morales-Castilla 1,2 *, Jonathan T. Davies 3 and
Electronic Supplementary Material Historical contingency, niche conservatism and the tendency for some taxa to be more diverse towards the poles Ignacio Morales-Castilla 1,2 *, Jonathan T. Davies 3 and
SPECIATION. REPRODUCTIVE BARRIERS PREZYGOTIC: Barriers that prevent fertilization. Habitat isolation Populations can t get together
 SPECIATION Origin of new species=speciation -Process by which one species splits into two or more species, accounts for both the unity and diversity of life SPECIES BIOLOGICAL CONCEPT Population or groups
SPECIATION Origin of new species=speciation -Process by which one species splits into two or more species, accounts for both the unity and diversity of life SPECIES BIOLOGICAL CONCEPT Population or groups
An introduction to the picante package
 An introduction to the picante package Steven Kembel (steve.kembel@gmail.com) April 2010 Contents 1 Installing picante 1 2 Data formats in picante 2 2.1 Phylogenies................................ 2 2.2
An introduction to the picante package Steven Kembel (steve.kembel@gmail.com) April 2010 Contents 1 Installing picante 1 2 Data formats in picante 2 2.1 Phylogenies................................ 2 2.2
