Shortest DNA cyclic cover in compressed space
|
|
|
- Aubrey Jasmin French
- 5 years ago
- Views:
Transcription
1 26 Data Compression Conference Shortest DNA cyclic cover in compressed space Bastien Cazaux, Rodrigo Cánovas and Eric Rivals L.I.R.M.M. & Institut Biologie Computationnelle Université de Montpellier, CNRS F Montpellier Cedex 5, France Abstract For a set of input words, finding a superstring (a string containing each word of the set as a substring) of minimal length is hard. Most approximation algorithms solve the Shortest Cyclic Cover problem before merging the cyclic strings into a linear superstring. A cyclic cover is a set of cyclic strings in which the input words occur as a substring. We investigate a variant of the Shortest Cyclic Cover problem for the case of DNA. Because the two strands that compose DNA have a reverse complementary sequence, and because the sequencing process often overlooks the strand of a read, each read or its reverse complement must occur as a substring in a cyclic cover. We exhibit a linear time algorithm based on graphs for solving the Shortest DNA Cyclic Cover problem and propose compressed data structures for storing the underlying graphs. All results and algorithms can be adapted to the case where strings are simply reversed but not complemented (e.g. in pattern recognition). Introduction Computing a shortest superstring for a set of input words is a way to compress a language. Indeed, overlaps between the words allow to save characters. The Shortest Superstring problem is crucial in text compression. Unfortunately, its hardness prevents us from finding, not only an optimal, but even an approximate solution that is arbitrarily near to optimal [, 2]. Rather than for a single linear superstring, it is thus better to opt for a compressed version that consists in a set of cyclic superstrings, whose total length is minimal. This representation is called a shortest cyclic cover. Indeed, the length of a shortest cyclic cover is smaller than that of a shortest linear superstring. These problems are related to DNA or genome assembly in bioinformatics: A highly redundant set of sequencing reads is given as input and one wishes to reconstruct the DNA sequence. As DNA is made of two complementary and reverse strands, a read can come from any strand, and thus either word or its reverse complement should occur in the superstring/genome. Moreover, chromosomes can be linear or cyclic molecules, and a sequenced sample may contain a mixture of cyclic genomes. We investigate the Shortest DNA Cyclic Cover problem and present a linear time algorithm for it. We also propose a compressed data structure for storing the underlying graphs used to solve the problem. All results described here can be adapted to the case where strings are simply reversed, and applied for instance to pattern matching of 2D shapes [3]. Outline First, we define the problem of finding a shortest DNA cyclic cover of a set strings and show the optimality of a greedy algorithm (Section ). Then, we introduce a graph, the Hierarchical Overlap Graph (HOG), which encodes all the information of the Overlap Graph using linear space, and also present a variation of the HOG adapted to 68-34/6 $3. 26 IEEE DOI.9/DCC
2 the DNA case. In the latter, nodes representing a word and its the reverse complementary sequence are glued together in a single node (Section 2). We then show that the Shortest DNA cyclic cover problem can be solved in linear time with this graph. Finally, we provide a construction algorithm for the HOG and a compressed representation that reduces the memory requirement (Section 3) before concluding.. Notation and problem definition Shortest DNA Cyclic Cover of Strings We introduce a notation for linear and cyclic strings/words. Let Σ be a finite alphabet. Let Σ denote the set of finite (linear) strings over Σ, and ε denote the empty string. Let u,v,w,x be linear strings of Σ. We denote by u the length of u. If x = uvw then u,v and w are all substrings of x, u is a prefix of x, and w a suffix of x. Furthermore, we say that v is a proper substring of x if v x (the definitions of a proper prefix/suffix are similar). An overlap from u over v is a linear string that is a proper suffix of u and a proper prefix of v. We denote by ov(u,v) the longest overlap from u to v (also termed maximal overlap). Overlaps are not symmetrical. The prefix from u to v, denoted by pr(u,v) is the string satisfying u = pr(u,v)ov(u,v), while the suffix from u to v, denoted su(u,v), is the substring of v satisfying v = ov(u,v)su(u,v). A DNA sequence is viewed as a string over Σ = {A,T,C,G}. As DNA is a double stranded molecule, any sequence also occurs as its reverse complement on the other strand, i.e. read from right to left and complemented (A T, C G). We denote the reverse complement of a string w by w. This notion and the notation are extended to any finite set of strings. The operation of reverse complementing a string is the composition of an inversion with an order 2 permutation over Σ. The following key property links the notions of maximal overlap and reverse complement. Property Let u and v be two linear strings. Then, ov( u, v ) = ov(v,u). Throughout the text, let P := {s,...,s n } be a finite set of linear words over Σ. We term the norm of P the quantity si P s i, and denote it by P. Without loss of generality, we assume that P is substring free: no input word is a substring of another. Computing a superstring for a set words, which is a string that includes each word as a substring, is a key question in stringology and data compression, but also in bioinformatics where it relates to DNA assembly [4]. DNA assembly, when the initial words are unstranded, consists in finding a superstring that includes either the word or its reverse complement for each input word. Therefore, it is interesting to design algorithms for finding superstrings and generalisation thereof for the DNA case (even without considering sequencing errors or mutations). Superstrings obtained by merging words of P using their maximal overlaps can be described by a permutation on {,...,n} that for any s i gives its successor in the superstring. Conversely, given a permutation one can easily compute the superstring induced by this Superstrings built using non maximal overlaps are disregarded because of their non optimality. 537
3 P {AT CA, AGT A, CT GA} + ( 3, 2 ) σ AT CA T CAG AGT A 3 2 pr(, 3) pr( 3, ) pr(2, 2) { P (σ) = A T CAG, AGT } Figure : Example of permutation and cyclic cover for the input set P := {ATCA,AGTA,CT GA}. An instance of a DNA cyclic cover of P obtained with a permutation σ. We obtain the cyclic cover P(σ) = {ATCAG,AGT }, where ATCAG and AGT are cyclic words. Note that this cyclic cover is not optimal. permutation. Similarly, a cyclic superstring or a cyclic cover can also be described by a permutation. Finding a shortest superstring of a set P is known to be difficult [, 4], and approximation algorithms often use the slightly more general concept of cyclic cover. A cyclic cover is a set C of cyclic strings such that each word of P is a substring of a cyclic string of C. We introduce the variant for the DNA case, the DNA Cyclic Cover: it is a cyclic cover C such that for each s of P, there exists c C such that s or s is a substring of c. Let DCC(P) be the set of DNA cyclic covers for the strings of P. In the DNA case, the substring freeness hypothesis is extended to P P. We introduce a variation of the Shortest Cyclic Cover problem adapted to the DNA case. Problem (Shortest DNA Cyclic Cover of Strings (SDCCS)) Let P be a set of linear strings. Find C, a DNA cyclic cover of strings in P, minimising C. For the input P, we denote by OPT(P), the set of optimal solutions for SDCCS..2 Permutations for DNA cyclic covers To describe a DNA cyclic cover by a permutation, we need a permutation on the set { P,...,,,..., P }, where, a positive element i stands for s i, while i represents si. As a cyclic string for DNA can be read in both directions, the permutation describes the cyclic cover in both directions; a constraint enforces that in each direction either s i or si is used in the induced DNA cyclic cover. Figure shows an example for a cyclic cover (see definition below). Example Consider P := {ATCA,AGTA,CT GA} := {s,s 2,s 3 }. One can form the DNA cyclic cover {TCAGTA} (with a single cyclic string) as the induced DNA cyclic cover of 538
4 P of the permutation σ := ( 3 2, 2 3 ). This means merging s with s 3 with s 2, or equivalently merging s 2 with s 3 with s. The permutation σ can also be written as (2,3, 2, 3,, ), where the numbers for the cycle 3 2 appear in bold, and the others in italic. Clearly for an input P, the set of DNA cyclic covers that are induced by permutations (using maximal overlaps), which we denote by PDCC(P), is a subset of all the DNA cyclic covers of P. Surely, an optimal DNA cyclic cover can be induced by a permutation. Hence, we get OPT(P) PDCC(P) DCC(P)..3 Optimal and Greedy DNA cyclic covers The classical Shortest Cyclic Cover of Strings (SCCS) has been investigated because of its central role in nearly all approximation algorithms for the Shortest Superstring Problem, starting with [2]. It can be solved in polynomial time by computing a minimum assignment on the distance graph in O( P + P 3 ) [5]. However, a greedy algorithm also yields an optimal solution [6], for which a linear time version was recently proposed [7]. Algorithm below gives the greedy algorithm for the SDCCS problem. Algorithm : The greedy algorithm for SDCCS Input: P a set of linear words Output: C a DNA cyclic cover of strings of P; C := / while P is not empty do u and v two elements of P P which have the longest overlap (u can be equal to v but not to v ) w is the merge of u and v P := P \ {u, u,v, v } if u = v (i.e. w is a cyclic string) then C := C {w} else P := P {w} return C We denote by GREEDY(P), the set of solutions of the greedy for SDCCS (see Algorithm ). The greedy algorithm for SCCS and SDCCS are similar. Since the line of proof applied to SCCS remains valid for SDCCS, we obtain that GREEDY(P) OPT(P). This proof relies on the properties of subset systems and on the Monge condition for string overlaps [6]. In summary, we obtain the following theorem, which implies that greedy DNA cyclic covers are optimal. Theorem (Figure 2) For any set of linear strings P, we have: GREEDY(P) OPT(P) PDCC(P) DCC(P) 2 Hierarchical Overlap Graph In the sequel, we use graph and data structures in which each node is associated with a string. For simplicity, we confound the node and the string it represents. 539
5 Figure 2: Inclusions of sets of solutions for SDCCS into the set of DNA Cyclic Covers. The Shortest Superstring problem can be solved by computing a maximum weight Hamiltonian path in the Overlap Graph [2]. Definition (Overlap Graph) The Overlap Graph (OG) of P is a complete, oriented graph, weighted on its arcs, whose nodes are the strings of P, and in which the weight of an arc (u,v) equals the length of the maximal overlap from string u to string v. The overlap graph for our running example is shown on Figure 4. The number of arcs in the OG is quadratic in the number of words. However, Gusfield et al. have exhibited an algorithm for computing the maximal overlaps in O( P ) time, plus O(n 2 ) time for outputting the weights [8]. However, the OG does not take advantage from the fact that several words can share the same or similar overlaps. A clear improvement would be a graph summarising the same information than OG, but would take O( P ) space. For this sake we propose the Hierarchical Overlap Graph. 2. Hierarchical Overlap Graph Consider a Generalised Suffix Tree (ST) for P equipped with Suffix Links (SL) (see [4] for the definition and properties of Suffix Trees, including their generalised version). For practical reasons, we extend the notion of SL to leaves of the ST. To ease the distinction, we say that the suffix links are red, while arcs of the ST are blue. Consider two leaves representing say s i and s j for some i, j n. The maximal overlaps of s i over s j is a suffix of s i and a prefix of s j : it is a repeated string within P and thus an explicit node that is an ancestor of s j in the ST. Hence, there exists a chain of suffix links linking the leaf s i to the node representing ov(s i,s j ), while there exists a blue chain from ov(s i,s j ) to s j. Note that this property remains valid for any two nodes in the ST. We call this path the Red-Blue Path between two nodes. Let Ov(P) be the set of maximum overlaps from a string of P to another string or the same string of P. Let us define the Hierarchical Overlap Graph as follows (see an illustration for our running example in Figure 4). Definition 2 The Hierarchical Overlap Graph of P, denoted by HOG(P), is the oriented graph (P Ov(P),R B) where: R = {(x,y) (P Ov(P)) 2 y longest suffix of x in V }, B = {(y,x) (P Ov(P)) 2 y longest prefix of x in V }. The HOG is reminiscent of the Hierarchical Graph of [9], where the arcs also encodes inclusion between substrings of the input words. However, in the Hierarchical Graph, each 54
6 arc extends or shortens the string by a single symbol, making it larger than the HOG. The HOG is in fact embedded in the ST of P, but successive blue arcs may be contracted in the HOG (the same for red arcs). We will see in Theorem 3 on page 8 that HOG(P) takes linear space in P. Property 2 Let u and v be two strings of P. In the HOG, there exists a Red-Blue path from u to v such that the largest node that is an ancestor of u and v is the overlap from u to v. The length of this path is at most pr(u,v) + su(u,v). Since, the HOG is embedded in the Generalised Suffix Tree of P, we obtain the following space bound. Corollary The total length of Red-Blue paths on the HOG is linear in P. The total length of the Red-Blue cycles used in a cyclic cover is linear in the norm of the cyclic cover. 2.2 Glued Hierarchical Overlap Graph Now, we want to find a solution of SDCCS and we are considering the graph HOG(P P ) (not simply HOG(P)). Because we considered for each word its normal sequence and its reverse complementary sequence, a sought superstring must include at least one of these. For each input word, we take into account the two forms of the word, and the corresponding nodes in the OG, forming a class. The OG is partitioned into n classes, where n is the number of words [6]. Hence, a DNA superstring corresponds to a partitioned path in the OG, while a DNA cyclic cover is a partitioned cyclic cover in the OG. Let X be a set such that X X = (P P ) Ov(P P ). X can be seen as a quotient, i.e. the set of representatives, of the equivalence relation of the reverse complement: Two different elements are in the same equivalence class, if the first is the reverse complement of the second. In the sequel, we take X as the set of minimal lexicographically strings between w and w for all strings of (P P ) Ov(P P ). Note that the construction can be adapted to the case where X is another quotient of the equivalence relation of the reverse complement. For a string w of (P P ) Ov(P P ), we set a(w) as the representative element of X of the string w, and d(w) as the element of {,,} such that, d(w) = if w = w, d(w) = if w X, or d(w) = if w / X. Definition 3 (Glued Hierarchical Overlap Graph) The Glued Hierarchical Overlap Graph of P, denoted by GHOG(P), is the undirected graph (V,E,l) labeled { on its edges, such that a(u) d(u) V = X and for all (u,v) B, (a(u),a(v)) E and l((a(u),a(v))) : a(v) d(v) where B is the set of blue arcs of HOG(P P ). The Glued Hierarchical Overlap Graph for our example is shown on Figure 4. By Property, there exists a bijection from B onto R, i.e. between the set of blue arcs and the set of red arcs of HOG(P P ). Hence, in Definition 3, we only consider arcs of B to 54
7 define the edges of the GHOG(P). The function l applies to edges of GHOG(P), and a(u) tells whether or not one needs to complement the string u. In other words, d(u) records of the original string we had when entering the edge: if d(u) = then u represents a(u), if it equals then u is equal to a(u). We define the RF-path between two elements of (P P ) X) as the smallest path p = (e = (u,u 2 ),...,e m = (u m,u m+ )) of the GHOG(P) such that there exists i between 2 and m such that: for all j < i, u j u j+ and l((u j,u j ))(u j ) = l((u j,u j+ ))(u j ), for all j i, u j u j+ and l((u j,u j ))(u j ) = l((u j,u j+ ))(u j ), l((u i,u i ))(u i ) = l((u i,u i+ ))(u i ). An example of a RB-path in the HOG and a RF-path in the GHOG are shown on Figure 3. ATCA TCA A AG AGTA TGAT TGA T CT TACT ATCA TGAT - - TCA - - A AG TGA T CT AGTA TACT Figure 3: An example of RB-path in the HOG and a RF-path in the GHOG. Theorem 2 There exists an algorithm that exactly solves SDCCS in linear time in P. Proof Theorem states that the greedy algorithm solves exactly SDCCS. The idea is to build a greedy solution for SDCCS in linear time in P using the GHOG(P). To achieve that we adapt the Superstring Graph designed for solving the SCCS problem to the DNA case [7]. In fact the Superstring graph is a subgraph of the HOG in which each arc has a multiplicity that counts the number of traversal for that arc. The multiplicity are set to make the Superstring Graph Eulerian. Its main property is to encode all greedy solutions in linear space. For SCCS, paths in the Superstring Graph are Red-Blue Paths between the leaves representing the input words (i.e. the s i for i n). For SDCCS, the RF paths replace the Red-Blue paths in the Superstring Graph built on the GHOG. 3 Data structures and construction algorithms for the GHOG 3. Representing HOG(P) using a compressed Suffix tree Consider a Generalised Suffix Tree (ST) for P equipped with Suffix Links (SL) [4]. As previously explained, for maximal overlaps between two words of P, each element of Ov(P) is an explicit, internal node of the ST of P. Hence, P Ov(P) takes a space that is linear in P. Now, as both (P Ov(P),B) and (P Ov(P),R) are trees, we get that HOG(P) is also linear in P. Gusfield et al. gave a characterisation of overlap nodes in the ST of P, with which they solved the all pairs suffix-prefix problem [8]. With this characterisation, one can mark in the ST of P the nodes that belong to HOG(P) in linear time in P. 542
8 ATCA 3 TCAG 2 TGAT 2 2 AGTA ATCA TCA TCAG 3 CTGA 2 TACT AGTA TACT TGAT AT T ε AG A TA AGTA ATCA TGAT TCA TGA AT AT A T ε TA TA AG CT TGA CT TACT CTGA CTGA TCAG Figure 4: Running example with P := {ATCA,AGTA,CT GA} := {s,s 2,s 3 }. We have P = {T GAT,TACT,TCAG}. The figure shows for P P i.e. in DNA case (Top) the Overlap Graph, (bottom left) the Hierarchical Overlap Graph, and (right) the Glued Hierarchical Overlap Graph. In the HOG and GHOG, input words appear on the outer circle, and their substrings are ordered on inner circles according to their length, with the empty word ε at the centre. In the HOG, one sees the Red-Blue paths, which become RF paths in the GHOG. In the GHOG, the, or labels store whether the traversal implies a change in the DNA strand or not. Theorem 3 The graph HOG(P) can be built in linear time in P. The algorithm uses O( P ) space, and finally HOG(P) occupies a space that is linear in min( P 2, P ). The Suffix Tree occupies large space in practice. Currently, several implementations of Compressed Suffix Tree (CST) data structures exist that support all classical operations including Suffix Links, and occupy between 5 and 2 bits per symbol [, ]. These implementations offer an advantageous trade off between storage and efficacy of operations. Using a CST and an additional membership table that marks which nodes belongs to the HOG, it is possible to emulate the traversal of blue and red arcs between nodes of the HOG. Remind that a blue arc is path of one or more arcs of the ST dictated by the labels, while a red arc is a chain of one or more SL starting from a node. 543
9 Let v be a node of CST(P) that belongs to HOG(P). To emulate a red arc starting from v, we iteratively traverse the chain of SL until we reach a marked node. At most this ends up at the root, taking in the worst case, v complexity(sl) time. Now to traverse a blue arc of the HOG using a label, we iteratively use the operation Child of v with the given label. If the node reached belongs to HOG(P) then the path is valid, taking in the worse case, v complexity(child) time. To obtain all possible outgoing blue arcs from v, we first check whether there exist marked nodes in its subtree and interrupt the search if there are none (we assume that checking the existence of marked nodes within a sub-tree can be done in constant time). Otherwise, we compute the Children of v in the ST and output those children that belong to the HOG. For the remaining children of v, we iterate the process in their subtree until all branches have been considered. This procedure stops at most when it reaches a leaf representing a word of P. It takes ((max i n s i ) v ) complexity(children) time. Alternative solution To encode HOG(P), one could use either a bi-directional wavelet index [2, 3], which is a compressed representation of the affix array [4]. This would almost double the space compared to a single CST, since the bi-directional wavelet index stores the trees for P and for the reverse of P. However, it would simplify the traversal of a red arc, since we could use the parent function on the reverse of P. Note that because the Superstring graph is a subgraph of the HOG, it can also be emulated on top of the compressed data structure used for HOG(P). 3.2 Representing GHOG(P) To represent GHOG(P) we need to build the HOG for the set P P and to glue pairs of substrings that are the respective reverse complement of each other. Each node encodes the lexicographically smallest substring of the pair, but represents both of them. Above, we detail how to build and emulate the HOG using a CST. To emulate GHOG(P) with the same data structure, we propose to add a new operation called Dlink (for DNA link), which links a node/string to the node representing its reverse complementary string. A Dlink is needed only for the nodes of GHOG(P) (not for all the ST nodes); this can be stored either using a permutation of the nodes, or by traversing the CST. Using the CST structure, Dlink and the marked nodes, one can emulate all the edges going out of a node of GHOG(P). Determining whether a string is the representative of its node is done by comparing it once with its reverse complement using Dlink, and the result is recorded in an array. This procedure also yields the edge s label (i.e. in {,,}). One traversal of the GHOG is enough to compute the representative string for each node and all labels. Theorem 4 The graph GHOG(P) can be built in linear time in P using a Compressed Suffix Tree data structure. 4 Conclusion and perspectives DNA cyclic covers are a natural generalisation of cyclic covers to handle the double strandedness of DNA. We propose a linear time algorithm for finding the Shortest DNA Cyclic Cover of a set of words using compressed data structures. In the general case, shortest 544
10 cyclic covers are used to compute approximate shortest superstrings. Hence, our work opens the way to approximation algorithms for linear or cyclic shortest superstrings in the DNA case. Currently for the DNA case, computing the Overlap Graph takes too much space. The HOG and GHOG reduce the memory needed and are thus valuable alternatives. Our algorithm to build these graphs with compressed data structures is another step further in term of improved memory requirement. Our proposal is flexible in several aspects. The graph used to encode the greedy solutions to the problem can be adapted to the cases where one imposes thresholds on the overlap lengths. With real sequencing data, an overlap must be long enough compared to the word size to be considered as reliable. Moreover, the graph could incorporate information on the multiplicity of input words, i.e. the minimal number of occurrences each word should have in the cyclic cover. Of course, extensions of our algorithms to approximate overlaps is a challenging perspective. Both graphs, the HOG and GHOG, which we introduced to compute cyclic covers, can be updated dynamically in linear time. Conceiving re-optimisation algorithms, which can recompute a solution once the instance has been modified, for example after adding or deleting a word is motivating line of research. Acknowledgements: This work is supported by ANR Colib read (ANR-2-BS2-8) and Défi MASTODONS SePhHaDe from CNRS. References [] J. Gallant, D. Maier, and J. A. Storer, On finding minimal length superstrings, Journal of Computer and System Sciences, vol. 2, pp. 5 58, 98. [2] A. Blum, T. Jiang, M. Li, J. Tromp, and M. Yannakakis, Linear approximation of shortest superstrings, in ACM Symposium on the Theory of Computing, 99, pp [3] H. Bunke and U. Buehler, Applications of approximate string matching to 2D shape recognition, Pattern Recognition, vol. 26, no. 2, pp , 993. [4] D. Gusfield, Algorithms on Strings, Trees and Sequences. Cambridge Univ. Press, 997. [5] C. H. Papadimitriou and K. Steiglitz, Combinatorial optimization : algorithms and complexity, 2nd ed. Dover Publications, Inc., 998, 496 p. [6] B. Cazaux and E. Rivals, The power of greedy algorithms for approximating Max-ATSP, Cyclic Cover, and superstrings, Discrete Applied Mathematics, p. doi:.6/j.dam , 25. [7], A linear time algorithm for Shortest Cyclic Cover of Strings, LIRMM Research report, p. 2, Feb. 25, hal.lirmm.fr. [8] D. Gusfield, G. M. Landau, and B. Schieber, An efficient algorithm for the all pairs suffixprefix problem, Inf. Proc. Letters, vol. 4, no. 4, pp. 8 85, 992. [9] A. Golovnev, A. S. Kulikov, and I. Mihajlin, Solving SCS for bounded length strings in fewer than 2 n steps, Inf. Proc. Letters, vol. 4, no. 8, pp , 24. [] K. Sadakane, Compressed suffix trees with full functionality, Theory Comput. Syst., vol. 4, no. 4, pp , 27. [] G. Navarro and L. M. S. Russo, Fast fully-compressed suffix trees, in Data Compression Conference, DCC 24, 24, pp [2] T. Schnattinger, E. Ohlebusch, and S. Gog, Bidirectional search in a string with wavelet trees and bidirectional matching statistics, Inf. Comput., vol. 23, pp. 3 22, 22. [3] T. W. Lam, R. Li, A. Tam, S. C. K. Wong, E. Wu, and S. Yiu, High throughput short read alignment via bi-directional BWT, in Proc. IEEE BIBM, 29, pp [4] D. Strothmann, The affix array data structure and its applications to RNA secondary structure analysis, Theor. Comput. Sci., vol. 389, no. -2, pp ,
Shortest DNA cyclic cover in compressed space
 Shortest DNA cyclic cover in compressed space Bastien Cazaux, Rodrigo Cánovas, Eric Rivals To cite this version: Bastien Cazaux, Rodrigo Cánovas, Eric Rivals. Shortest DNA cyclic cover in compressed space.
Shortest DNA cyclic cover in compressed space Bastien Cazaux, Rodrigo Cánovas, Eric Rivals To cite this version: Bastien Cazaux, Rodrigo Cánovas, Eric Rivals. Shortest DNA cyclic cover in compressed space.
Hierarchical Overlap Graph
 Hierarchical Overlap Graph B. Cazaux and E. Rivals LIRMM & IBC, Montpellier 8. Feb. 2018 arxiv:1802.04632 2018 B. Cazaux & E. Rivals 1 / 29 Overlap Graph for a set of words Consider the set P := {abaa,
Hierarchical Overlap Graph B. Cazaux and E. Rivals LIRMM & IBC, Montpellier 8. Feb. 2018 arxiv:1802.04632 2018 B. Cazaux & E. Rivals 1 / 29 Overlap Graph for a set of words Consider the set P := {abaa,
Approximating Shortest Superstring Problem Using de Bruijn Graphs
 Approximating Shortest Superstring Problem Using de Bruijn Advanced s Course Presentation Farshad Barahimi Department of Computer Science University of Lethbridge November 19, 2013 This presentation is
Approximating Shortest Superstring Problem Using de Bruijn Advanced s Course Presentation Farshad Barahimi Department of Computer Science University of Lethbridge November 19, 2013 This presentation is
The power of greedy algorithms for approximating Max-ATSP, Cyclic Cover, and Superstrings
 The power of greedy algorithms for approximating Max-ATSP, Cyclic Cover, and Superstrings Bastien Cazaux and Eric Rivals L.I.R.M.M. & Institut Biologie Computationnelle Université de Montpellier, CNRS
The power of greedy algorithms for approximating Max-ATSP, Cyclic Cover, and Superstrings Bastien Cazaux and Eric Rivals L.I.R.M.M. & Institut Biologie Computationnelle Université de Montpellier, CNRS
Lecture 18 April 26, 2012
 6.851: Advanced Data Structures Spring 2012 Prof. Erik Demaine Lecture 18 April 26, 2012 1 Overview In the last lecture we introduced the concept of implicit, succinct, and compact data structures, and
6.851: Advanced Data Structures Spring 2012 Prof. Erik Demaine Lecture 18 April 26, 2012 1 Overview In the last lecture we introduced the concept of implicit, succinct, and compact data structures, and
Compressed Index for Dynamic Text
 Compressed Index for Dynamic Text Wing-Kai Hon Tak-Wah Lam Kunihiko Sadakane Wing-Kin Sung Siu-Ming Yiu Abstract This paper investigates how to index a text which is subject to updates. The best solution
Compressed Index for Dynamic Text Wing-Kai Hon Tak-Wah Lam Kunihiko Sadakane Wing-Kin Sung Siu-Ming Yiu Abstract This paper investigates how to index a text which is subject to updates. The best solution
Greedy Conjecture for Strings of Length 4
 Greedy Conjecture for Strings of Length 4 Alexander S. Kulikov 1, Sergey Savinov 2, and Evgeniy Sluzhaev 1,2 1 St. Petersburg Department of Steklov Institute of Mathematics 2 St. Petersburg Academic University
Greedy Conjecture for Strings of Length 4 Alexander S. Kulikov 1, Sergey Savinov 2, and Evgeniy Sluzhaev 1,2 1 St. Petersburg Department of Steklov Institute of Mathematics 2 St. Petersburg Academic University
A dynamic programming approach to generation of strong traces
 A dynamic programming approach to generation of strong traces Nino Bašić Faculty of Mathematics, Natural Sciences and Information Technologies University of Primorska 32 nd TBI Winterseminar Bled, 16 February
A dynamic programming approach to generation of strong traces Nino Bašić Faculty of Mathematics, Natural Sciences and Information Technologies University of Primorska 32 nd TBI Winterseminar Bled, 16 February
Alphabet Friendly FM Index
 Alphabet Friendly FM Index Author: Rodrigo González Santiago, November 8 th, 2005 Departamento de Ciencias de la Computación Universidad de Chile Outline Motivations Basics Burrows Wheeler Transform FM
Alphabet Friendly FM Index Author: Rodrigo González Santiago, November 8 th, 2005 Departamento de Ciencias de la Computación Universidad de Chile Outline Motivations Basics Burrows Wheeler Transform FM
Least Random Suffix/Prefix Matches in Output-Sensitive Time
 Least Random Suffix/Prefix Matches in Output-Sensitive Time Niko Välimäki Department of Computer Science University of Helsinki nvalimak@cs.helsinki.fi 23rd Annual Symposium on Combinatorial Pattern Matching
Least Random Suffix/Prefix Matches in Output-Sensitive Time Niko Välimäki Department of Computer Science University of Helsinki nvalimak@cs.helsinki.fi 23rd Annual Symposium on Combinatorial Pattern Matching
arxiv: v1 [cs.ds] 9 Apr 2018
![arxiv: v1 [cs.ds] 9 Apr 2018 arxiv: v1 [cs.ds] 9 Apr 2018](/thumbs/91/107706762.jpg) From Regular Expression Matching to Parsing Philip Bille Technical University of Denmark phbi@dtu.dk Inge Li Gørtz Technical University of Denmark inge@dtu.dk arxiv:1804.02906v1 [cs.ds] 9 Apr 2018 Abstract
From Regular Expression Matching to Parsing Philip Bille Technical University of Denmark phbi@dtu.dk Inge Li Gørtz Technical University of Denmark inge@dtu.dk arxiv:1804.02906v1 [cs.ds] 9 Apr 2018 Abstract
A GREEDY APPROXIMATION ALGORITHM FOR CONSTRUCTING SHORTEST COMMON SUPERSTRINGS *
 A GREEDY APPROXIMATION ALGORITHM FOR CONSTRUCTING SHORTEST COMMON SUPERSTRINGS * 1 Jorma Tarhio and Esko Ukkonen Department of Computer Science, University of Helsinki Tukholmankatu 2, SF-00250 Helsinki,
A GREEDY APPROXIMATION ALGORITHM FOR CONSTRUCTING SHORTEST COMMON SUPERSTRINGS * 1 Jorma Tarhio and Esko Ukkonen Department of Computer Science, University of Helsinki Tukholmankatu 2, SF-00250 Helsinki,
Smaller and Faster Lempel-Ziv Indices
 Smaller and Faster Lempel-Ziv Indices Diego Arroyuelo and Gonzalo Navarro Dept. of Computer Science, Universidad de Chile, Chile. {darroyue,gnavarro}@dcc.uchile.cl Abstract. Given a text T[1..u] over an
Smaller and Faster Lempel-Ziv Indices Diego Arroyuelo and Gonzalo Navarro Dept. of Computer Science, Universidad de Chile, Chile. {darroyue,gnavarro}@dcc.uchile.cl Abstract. Given a text T[1..u] over an
Structure-Based Comparison of Biomolecules
 Structure-Based Comparison of Biomolecules Benedikt Christoph Wolters Seminar Bioinformatics Algorithms RWTH AACHEN 07/17/2015 Outline 1 Introduction and Motivation Protein Structure Hierarchy Protein
Structure-Based Comparison of Biomolecules Benedikt Christoph Wolters Seminar Bioinformatics Algorithms RWTH AACHEN 07/17/2015 Outline 1 Introduction and Motivation Protein Structure Hierarchy Protein
Analysis of tree edit distance algorithms
 Analysis of tree edit distance algorithms Serge Dulucq 1 and Hélène Touzet 1 LaBRI - Universit Bordeaux I 33 0 Talence cedex, France Serge.Dulucq@labri.fr LIFL - Universit Lille 1 9 6 Villeneuve d Ascq
Analysis of tree edit distance algorithms Serge Dulucq 1 and Hélène Touzet 1 LaBRI - Universit Bordeaux I 33 0 Talence cedex, France Serge.Dulucq@labri.fr LIFL - Universit Lille 1 9 6 Villeneuve d Ascq
Proofs, Strings, and Finite Automata. CS154 Chris Pollett Feb 5, 2007.
 Proofs, Strings, and Finite Automata CS154 Chris Pollett Feb 5, 2007. Outline Proofs and Proof Strategies Strings Finding proofs Example: For every graph G, the sum of the degrees of all the nodes in G
Proofs, Strings, and Finite Automata CS154 Chris Pollett Feb 5, 2007. Outline Proofs and Proof Strategies Strings Finding proofs Example: For every graph G, the sum of the degrees of all the nodes in G
Small-Space Dictionary Matching (Dissertation Proposal)
 Small-Space Dictionary Matching (Dissertation Proposal) Graduate Center of CUNY 1/24/2012 Problem Definition Dictionary Matching Input: Dictionary D = P 1,P 2,...,P d containing d patterns. Text T of length
Small-Space Dictionary Matching (Dissertation Proposal) Graduate Center of CUNY 1/24/2012 Problem Definition Dictionary Matching Input: Dictionary D = P 1,P 2,...,P d containing d patterns. Text T of length
Pattern Matching (Exact Matching) Overview
 CSI/BINF 5330 Pattern Matching (Exact Matching) Young-Rae Cho Associate Professor Department of Computer Science Baylor University Overview Pattern Matching Exhaustive Search DFA Algorithm KMP Algorithm
CSI/BINF 5330 Pattern Matching (Exact Matching) Young-Rae Cho Associate Professor Department of Computer Science Baylor University Overview Pattern Matching Exhaustive Search DFA Algorithm KMP Algorithm
arxiv: v1 [cs.dm] 18 May 2016
![arxiv: v1 [cs.dm] 18 May 2016 arxiv: v1 [cs.dm] 18 May 2016](/thumbs/75/72625353.jpg) A note on the shortest common superstring of NGS reads arxiv:1605.05542v1 [cs.dm] 18 May 2016 Tristan Braquelaire Marie Gasparoux Mathieu Raffinot Raluca Uricaru May 19, 2016 Abstract The Shortest Superstring
A note on the shortest common superstring of NGS reads arxiv:1605.05542v1 [cs.dm] 18 May 2016 Tristan Braquelaire Marie Gasparoux Mathieu Raffinot Raluca Uricaru May 19, 2016 Abstract The Shortest Superstring
Theoretical Computer Science. Why greed works for shortest common superstring problem
 Theoretical Computer Science 410 (2009) 5374 5381 Contents lists available at ScienceDirect Theoretical Computer Science journal homepage: www.elsevier.com/locate/tcs Why greed works for shortest common
Theoretical Computer Science 410 (2009) 5374 5381 Contents lists available at ScienceDirect Theoretical Computer Science journal homepage: www.elsevier.com/locate/tcs Why greed works for shortest common
An O(N) Semi-Predictive Universal Encoder via the BWT
 An O(N) Semi-Predictive Universal Encoder via the BWT Dror Baron and Yoram Bresler Abstract We provide an O(N) algorithm for a non-sequential semi-predictive encoder whose pointwise redundancy with respect
An O(N) Semi-Predictive Universal Encoder via the BWT Dror Baron and Yoram Bresler Abstract We provide an O(N) algorithm for a non-sequential semi-predictive encoder whose pointwise redundancy with respect
Lecture 4 : Adaptive source coding algorithms
 Lecture 4 : Adaptive source coding algorithms February 2, 28 Information Theory Outline 1. Motivation ; 2. adaptive Huffman encoding ; 3. Gallager and Knuth s method ; 4. Dictionary methods : Lempel-Ziv
Lecture 4 : Adaptive source coding algorithms February 2, 28 Information Theory Outline 1. Motivation ; 2. adaptive Huffman encoding ; 3. Gallager and Knuth s method ; 4. Dictionary methods : Lempel-Ziv
A PARSIMONY APPROACH TO ANALYSIS OF HUMAN SEGMENTAL DUPLICATIONS
 A PARSIMONY APPROACH TO ANALYSIS OF HUMAN SEGMENTAL DUPLICATIONS CRYSTAL L. KAHN and BENJAMIN J. RAPHAEL Box 1910, Brown University Department of Computer Science & Center for Computational Molecular Biology
A PARSIMONY APPROACH TO ANALYSIS OF HUMAN SEGMENTAL DUPLICATIONS CRYSTAL L. KAHN and BENJAMIN J. RAPHAEL Box 1910, Brown University Department of Computer Science & Center for Computational Molecular Biology
SUFFIX TREE. SYNONYMS Compact suffix trie
 SUFFIX TREE Maxime Crochemore King s College London and Université Paris-Est, http://www.dcs.kcl.ac.uk/staff/mac/ Thierry Lecroq Université de Rouen, http://monge.univ-mlv.fr/~lecroq SYNONYMS Compact suffix
SUFFIX TREE Maxime Crochemore King s College London and Université Paris-Est, http://www.dcs.kcl.ac.uk/staff/mac/ Thierry Lecroq Université de Rouen, http://monge.univ-mlv.fr/~lecroq SYNONYMS Compact suffix
Combinatorial Optimization
 Combinatorial Optimization Problem set 8: solutions 1. Fix constants a R and b > 1. For n N, let f(n) = n a and g(n) = b n. Prove that f(n) = o ( g(n) ). Solution. First we observe that g(n) 0 for all
Combinatorial Optimization Problem set 8: solutions 1. Fix constants a R and b > 1. For n N, let f(n) = n a and g(n) = b n. Prove that f(n) = o ( g(n) ). Solution. First we observe that g(n) 0 for all
Implementing Approximate Regularities
 Implementing Approximate Regularities Manolis Christodoulakis Costas S. Iliopoulos Department of Computer Science King s College London Kunsoo Park School of Computer Science and Engineering, Seoul National
Implementing Approximate Regularities Manolis Christodoulakis Costas S. Iliopoulos Department of Computer Science King s College London Kunsoo Park School of Computer Science and Engineering, Seoul National
Bio nformatics. Lecture 16. Saad Mneimneh
 Bio nformatics Lecture 16 DNA sequencing To sequence a DNA is to obtain the string of bases that it contains. It is impossible to sequence the whole DNA molecule directly. We may however obtain a piece
Bio nformatics Lecture 16 DNA sequencing To sequence a DNA is to obtain the string of bases that it contains. It is impossible to sequence the whole DNA molecule directly. We may however obtain a piece
UNIT I INFORMATION THEORY. I k log 2
 UNIT I INFORMATION THEORY Claude Shannon 1916-2001 Creator of Information Theory, lays the foundation for implementing logic in digital circuits as part of his Masters Thesis! (1939) and published a paper
UNIT I INFORMATION THEORY Claude Shannon 1916-2001 Creator of Information Theory, lays the foundation for implementing logic in digital circuits as part of his Masters Thesis! (1939) and published a paper
Hardness of Optimal Spaced Seed Design. François Nicolas, Eric Rivals,1
 Hardness of Optimal Spaced Seed Design François Nicolas, Eric Rivals,1 L.I.R.M.M., U.M.R. 5506 C.N.R.S. Université de Montpellier II 161, rue Ada 34392 Montpellier Cedex 5 France Correspondence address:
Hardness of Optimal Spaced Seed Design François Nicolas, Eric Rivals,1 L.I.R.M.M., U.M.R. 5506 C.N.R.S. Université de Montpellier II 161, rue Ada 34392 Montpellier Cedex 5 France Correspondence address:
Superstring Graph: A New Approach for Genome Assembly
 Superstring Graph: A New Approach for Genome Assembly Bastien Cazaux, Gustavo Sacomoto, Eric Rivals To cite this version: Bastien Cazaux, Gustavo Sacomoto, Eric Rivals. Superstring Graph: A New Approach
Superstring Graph: A New Approach for Genome Assembly Bastien Cazaux, Gustavo Sacomoto, Eric Rivals To cite this version: Bastien Cazaux, Gustavo Sacomoto, Eric Rivals. Superstring Graph: A New Approach
COMPRESSED INDEXING DATA STRUCTURES FOR BIOLOGICAL SEQUENCES
 COMPRESSED INDEXING DATA STRUCTURES FOR BIOLOGICAL SEQUENCES DO HUY HOANG (B.C.S. (Hons), NUS) A THESIS SUBMITTED FOR THE DEGREE OF DOCTOR OF PHILOSOPHY IN COMPUTER SCIENCE SCHOOL OF COMPUTING NATIONAL
COMPRESSED INDEXING DATA STRUCTURES FOR BIOLOGICAL SEQUENCES DO HUY HOANG (B.C.S. (Hons), NUS) A THESIS SUBMITTED FOR THE DEGREE OF DOCTOR OF PHILOSOPHY IN COMPUTER SCIENCE SCHOOL OF COMPUTING NATIONAL
Indexing LZ77: The Next Step in Self-Indexing. Gonzalo Navarro Department of Computer Science, University of Chile
 Indexing LZ77: The Next Step in Self-Indexing Gonzalo Navarro Department of Computer Science, University of Chile gnavarro@dcc.uchile.cl Part I: Why Jumping off the Cliff The Past Century Self-Indexing:
Indexing LZ77: The Next Step in Self-Indexing Gonzalo Navarro Department of Computer Science, University of Chile gnavarro@dcc.uchile.cl Part I: Why Jumping off the Cliff The Past Century Self-Indexing:
Transducers for bidirectional decoding of prefix codes
 Transducers for bidirectional decoding of prefix codes Laura Giambruno a,1, Sabrina Mantaci a,1 a Dipartimento di Matematica ed Applicazioni - Università di Palermo - Italy Abstract We construct a transducer
Transducers for bidirectional decoding of prefix codes Laura Giambruno a,1, Sabrina Mantaci a,1 a Dipartimento di Matematica ed Applicazioni - Università di Palermo - Italy Abstract We construct a transducer
Covering Linear Orders with Posets
 Covering Linear Orders with Posets Proceso L. Fernandez, Lenwood S. Heath, Naren Ramakrishnan, and John Paul C. Vergara Department of Information Systems and Computer Science, Ateneo de Manila University,
Covering Linear Orders with Posets Proceso L. Fernandez, Lenwood S. Heath, Naren Ramakrishnan, and John Paul C. Vergara Department of Information Systems and Computer Science, Ateneo de Manila University,
CSE 549: Computational Biology. Computer Science for Biologists Biology
 CSE 549: Computational Biology Computer Science for Biologists Biology What is Computer Science? http://people.cs.pitt.edu/~kirk/cs2110/computer_science_major.png What is Computer Science? Not actually
CSE 549: Computational Biology Computer Science for Biologists Biology What is Computer Science? http://people.cs.pitt.edu/~kirk/cs2110/computer_science_major.png What is Computer Science? Not actually
Motif Extraction from Weighted Sequences
 Motif Extraction from Weighted Sequences C. Iliopoulos 1, K. Perdikuri 2,3, E. Theodoridis 2,3,, A. Tsakalidis 2,3 and K. Tsichlas 1 1 Department of Computer Science, King s College London, London WC2R
Motif Extraction from Weighted Sequences C. Iliopoulos 1, K. Perdikuri 2,3, E. Theodoridis 2,3,, A. Tsakalidis 2,3 and K. Tsichlas 1 1 Department of Computer Science, King s College London, London WC2R
Theory of Computation
 Theory of Computation (Feodor F. Dragan) Department of Computer Science Kent State University Spring, 2018 Theory of Computation, Feodor F. Dragan, Kent State University 1 Before we go into details, what
Theory of Computation (Feodor F. Dragan) Department of Computer Science Kent State University Spring, 2018 Theory of Computation, Feodor F. Dragan, Kent State University 1 Before we go into details, what
Run-length & Entropy Coding. Redundancy Removal. Sampling. Quantization. Perform inverse operations at the receiver EEE
 General e Image Coder Structure Motion Video x(s 1,s 2,t) or x(s 1,s 2 ) Natural Image Sampling A form of data compression; usually lossless, but can be lossy Redundancy Removal Lossless compression: predictive
General e Image Coder Structure Motion Video x(s 1,s 2,t) or x(s 1,s 2 ) Natural Image Sampling A form of data compression; usually lossless, but can be lossy Redundancy Removal Lossless compression: predictive
arxiv: v1 [cs.ds] 15 Feb 2012
![arxiv: v1 [cs.ds] 15 Feb 2012 arxiv: v1 [cs.ds] 15 Feb 2012](/thumbs/92/110725364.jpg) Linear-Space Substring Range Counting over Polylogarithmic Alphabets Travis Gagie 1 and Pawe l Gawrychowski 2 1 Aalto University, Finland travis.gagie@aalto.fi 2 Max Planck Institute, Germany gawry@cs.uni.wroc.pl
Linear-Space Substring Range Counting over Polylogarithmic Alphabets Travis Gagie 1 and Pawe l Gawrychowski 2 1 Aalto University, Finland travis.gagie@aalto.fi 2 Max Planck Institute, Germany gawry@cs.uni.wroc.pl
arxiv: v1 [cs.ds] 21 May 2013
![arxiv: v1 [cs.ds] 21 May 2013 arxiv: v1 [cs.ds] 21 May 2013](/thumbs/79/80107468.jpg) Easy identification of generalized common nested intervals Fabien de Montgolfier 1, Mathieu Raffinot 1, and Irena Rusu 2 arxiv:1305.4747v1 [cs.ds] 21 May 2013 1 LIAFA, Univ. Paris Diderot - Paris 7, 75205
Easy identification of generalized common nested intervals Fabien de Montgolfier 1, Mathieu Raffinot 1, and Irena Rusu 2 arxiv:1305.4747v1 [cs.ds] 21 May 2013 1 LIAFA, Univ. Paris Diderot - Paris 7, 75205
4.8 Huffman Codes. These lecture slides are supplied by Mathijs de Weerd
 4.8 Huffman Codes These lecture slides are supplied by Mathijs de Weerd Data Compression Q. Given a text that uses 32 symbols (26 different letters, space, and some punctuation characters), how can we
4.8 Huffman Codes These lecture slides are supplied by Mathijs de Weerd Data Compression Q. Given a text that uses 32 symbols (26 different letters, space, and some punctuation characters), how can we
Online Computation of Abelian Runs
 Online Computation of Abelian Runs Gabriele Fici 1, Thierry Lecroq 2, Arnaud Lefebvre 2, and Élise Prieur-Gaston2 1 Dipartimento di Matematica e Informatica, Università di Palermo, Italy Gabriele.Fici@unipa.it
Online Computation of Abelian Runs Gabriele Fici 1, Thierry Lecroq 2, Arnaud Lefebvre 2, and Élise Prieur-Gaston2 1 Dipartimento di Matematica e Informatica, Università di Palermo, Italy Gabriele.Fici@unipa.it
On the Sizes of Decision Diagrams Representing the Set of All Parse Trees of a Context-free Grammar
 Proceedings of Machine Learning Research vol 73:153-164, 2017 AMBN 2017 On the Sizes of Decision Diagrams Representing the Set of All Parse Trees of a Context-free Grammar Kei Amii Kyoto University Kyoto
Proceedings of Machine Learning Research vol 73:153-164, 2017 AMBN 2017 On the Sizes of Decision Diagrams Representing the Set of All Parse Trees of a Context-free Grammar Kei Amii Kyoto University Kyoto
Lecture 1 : Data Compression and Entropy
 CPS290: Algorithmic Foundations of Data Science January 8, 207 Lecture : Data Compression and Entropy Lecturer: Kamesh Munagala Scribe: Kamesh Munagala In this lecture, we will study a simple model for
CPS290: Algorithmic Foundations of Data Science January 8, 207 Lecture : Data Compression and Entropy Lecturer: Kamesh Munagala Scribe: Kamesh Munagala In this lecture, we will study a simple model for
Fast String Kernels. Alexander J. Smola Machine Learning Group, RSISE The Australian National University Canberra, ACT 0200
 Fast String Kernels Alexander J. Smola Machine Learning Group, RSISE The Australian National University Canberra, ACT 0200 Alex.Smola@anu.edu.au joint work with S.V.N. Vishwanathan Slides (soon) available
Fast String Kernels Alexander J. Smola Machine Learning Group, RSISE The Australian National University Canberra, ACT 0200 Alex.Smola@anu.edu.au joint work with S.V.N. Vishwanathan Slides (soon) available
CSE 549: Computational Biology. Computer Science for Biologists Biology
 CSE 549: Computational Biology Computer Science for Biologists Biology What is Computer Science? http://people.cs.pitt.edu/~kirk/cs2110/computer_science_major.png What is Computer Science? Not actually
CSE 549: Computational Biology Computer Science for Biologists Biology What is Computer Science? http://people.cs.pitt.edu/~kirk/cs2110/computer_science_major.png What is Computer Science? Not actually
Complexity Theory VU , SS The Polynomial Hierarchy. Reinhard Pichler
 Complexity Theory Complexity Theory VU 181.142, SS 2018 6. The Polynomial Hierarchy Reinhard Pichler Institut für Informationssysteme Arbeitsbereich DBAI Technische Universität Wien 15 May, 2018 Reinhard
Complexity Theory Complexity Theory VU 181.142, SS 2018 6. The Polynomial Hierarchy Reinhard Pichler Institut für Informationssysteme Arbeitsbereich DBAI Technische Universität Wien 15 May, 2018 Reinhard
Outline. Complexity Theory EXACT TSP. The Class DP. Definition. Problem EXACT TSP. Complexity of EXACT TSP. Proposition VU 181.
 Complexity Theory Complexity Theory Outline Complexity Theory VU 181.142, SS 2018 6. The Polynomial Hierarchy Reinhard Pichler Institut für Informationssysteme Arbeitsbereich DBAI Technische Universität
Complexity Theory Complexity Theory Outline Complexity Theory VU 181.142, SS 2018 6. The Polynomial Hierarchy Reinhard Pichler Institut für Informationssysteme Arbeitsbereich DBAI Technische Universität
Efficient Polynomial-Time Algorithms for Variants of the Multiple Constrained LCS Problem
 Efficient Polynomial-Time Algorithms for Variants of the Multiple Constrained LCS Problem Hsing-Yen Ann National Center for High-Performance Computing Tainan 74147, Taiwan Chang-Biau Yang and Chiou-Ting
Efficient Polynomial-Time Algorithms for Variants of the Multiple Constrained LCS Problem Hsing-Yen Ann National Center for High-Performance Computing Tainan 74147, Taiwan Chang-Biau Yang and Chiou-Ting
Branch-and-Bound for the Travelling Salesman Problem
 Branch-and-Bound for the Travelling Salesman Problem Leo Liberti LIX, École Polytechnique, F-91128 Palaiseau, France Email:liberti@lix.polytechnique.fr March 15, 2011 Contents 1 The setting 1 1.1 Graphs...............................................
Branch-and-Bound for the Travelling Salesman Problem Leo Liberti LIX, École Polytechnique, F-91128 Palaiseau, France Email:liberti@lix.polytechnique.fr March 15, 2011 Contents 1 The setting 1 1.1 Graphs...............................................
arxiv: v3 [cs.fl] 2 Jul 2018
![arxiv: v3 [cs.fl] 2 Jul 2018 arxiv: v3 [cs.fl] 2 Jul 2018](/thumbs/93/111950553.jpg) COMPLEXITY OF PREIMAGE PROBLEMS FOR DETERMINISTIC FINITE AUTOMATA MIKHAIL V. BERLINKOV arxiv:1704.08233v3 [cs.fl] 2 Jul 2018 Institute of Natural Sciences and Mathematics, Ural Federal University, Ekaterinburg,
COMPLEXITY OF PREIMAGE PROBLEMS FOR DETERMINISTIC FINITE AUTOMATA MIKHAIL V. BERLINKOV arxiv:1704.08233v3 [cs.fl] 2 Jul 2018 Institute of Natural Sciences and Mathematics, Ural Federal University, Ekaterinburg,
Finding all covers of an indeterminate string in O(n) time on average
 Finding all covers of an indeterminate string in O(n) time on average Md. Faizul Bari, M. Sohel Rahman, and Rifat Shahriyar Department of Computer Science and Engineering Bangladesh University of Engineering
Finding all covers of an indeterminate string in O(n) time on average Md. Faizul Bari, M. Sohel Rahman, and Rifat Shahriyar Department of Computer Science and Engineering Bangladesh University of Engineering
arxiv: v2 [cs.ds] 3 Oct 2017
![arxiv: v2 [cs.ds] 3 Oct 2017 arxiv: v2 [cs.ds] 3 Oct 2017](/thumbs/84/90870651.jpg) Orthogonal Vectors Indexing Isaac Goldstein 1, Moshe Lewenstein 1, and Ely Porat 1 1 Bar-Ilan University, Ramat Gan, Israel {goldshi,moshe,porately}@cs.biu.ac.il arxiv:1710.00586v2 [cs.ds] 3 Oct 2017 Abstract
Orthogonal Vectors Indexing Isaac Goldstein 1, Moshe Lewenstein 1, and Ely Porat 1 1 Bar-Ilan University, Ramat Gan, Israel {goldshi,moshe,porately}@cs.biu.ac.il arxiv:1710.00586v2 [cs.ds] 3 Oct 2017 Abstract
Efficient Reassembling of Graphs, Part 1: The Linear Case
 Efficient Reassembling of Graphs, Part 1: The Linear Case Assaf Kfoury Boston University Saber Mirzaei Boston University Abstract The reassembling of a simple connected graph G = (V, E) is an abstraction
Efficient Reassembling of Graphs, Part 1: The Linear Case Assaf Kfoury Boston University Saber Mirzaei Boston University Abstract The reassembling of a simple connected graph G = (V, E) is an abstraction
Alphabet-Independent Compressed Text Indexing
 Alphabet-Independent Compressed Text Indexing DJAMAL BELAZZOUGUI Université Paris Diderot GONZALO NAVARRO University of Chile Self-indexes are able to represent a text within asymptotically the information-theoretic
Alphabet-Independent Compressed Text Indexing DJAMAL BELAZZOUGUI Université Paris Diderot GONZALO NAVARRO University of Chile Self-indexes are able to represent a text within asymptotically the information-theoretic
1 Introduction to information theory
 1 Introduction to information theory 1.1 Introduction In this chapter we present some of the basic concepts of information theory. The situations we have in mind involve the exchange of information through
1 Introduction to information theory 1.1 Introduction In this chapter we present some of the basic concepts of information theory. The situations we have in mind involve the exchange of information through
Automata-based Verification - III
 COMP30172: Advanced Algorithms Automata-based Verification - III Howard Barringer Room KB2.20: email: howard.barringer@manchester.ac.uk March 2009 Third Topic Infinite Word Automata Motivation Büchi Automata
COMP30172: Advanced Algorithms Automata-based Verification - III Howard Barringer Room KB2.20: email: howard.barringer@manchester.ac.uk March 2009 Third Topic Infinite Word Automata Motivation Büchi Automata
arxiv: v2 [cs.ds] 16 Mar 2015
![arxiv: v2 [cs.ds] 16 Mar 2015 arxiv: v2 [cs.ds] 16 Mar 2015](/thumbs/95/125127202.jpg) Longest common substrings with k mismatches Tomas Flouri 1, Emanuele Giaquinta 2, Kassian Kobert 1, and Esko Ukkonen 3 arxiv:1409.1694v2 [cs.ds] 16 Mar 2015 1 Heidelberg Institute for Theoretical Studies,
Longest common substrings with k mismatches Tomas Flouri 1, Emanuele Giaquinta 2, Kassian Kobert 1, and Esko Ukkonen 3 arxiv:1409.1694v2 [cs.ds] 16 Mar 2015 1 Heidelberg Institute for Theoretical Studies,
An Algebraic View of the Relation between Largest Common Subtrees and Smallest Common Supertrees
 An Algebraic View of the Relation between Largest Common Subtrees and Smallest Common Supertrees Francesc Rosselló 1, Gabriel Valiente 2 1 Department of Mathematics and Computer Science, Research Institute
An Algebraic View of the Relation between Largest Common Subtrees and Smallest Common Supertrees Francesc Rosselló 1, Gabriel Valiente 2 1 Department of Mathematics and Computer Science, Research Institute
Computation Theory Finite Automata
 Computation Theory Dept. of Computing ITT Dublin October 14, 2010 Computation Theory I 1 We would like a model that captures the general nature of computation Consider two simple problems: 2 Design a program
Computation Theory Dept. of Computing ITT Dublin October 14, 2010 Computation Theory I 1 We would like a model that captures the general nature of computation Consider two simple problems: 2 Design a program
Dominating Set Counting in Graph Classes
 Dominating Set Counting in Graph Classes Shuji Kijima 1, Yoshio Okamoto 2, and Takeaki Uno 3 1 Graduate School of Information Science and Electrical Engineering, Kyushu University, Japan kijima@inf.kyushu-u.ac.jp
Dominating Set Counting in Graph Classes Shuji Kijima 1, Yoshio Okamoto 2, and Takeaki Uno 3 1 Graduate School of Information Science and Electrical Engineering, Kyushu University, Japan kijima@inf.kyushu-u.ac.jp
CSE 202 Homework 4 Matthias Springer, A
 CSE 202 Homework 4 Matthias Springer, A99500782 1 Problem 2 Basic Idea PERFECT ASSEMBLY N P: a permutation P of s i S is a certificate that can be checked in polynomial time by ensuring that P = S, and
CSE 202 Homework 4 Matthias Springer, A99500782 1 Problem 2 Basic Idea PERFECT ASSEMBLY N P: a permutation P of s i S is a certificate that can be checked in polynomial time by ensuring that P = S, and
Theoretical Computer Science
 Theoretical Computer Science 410 (2009) 2759 2766 Contents lists available at ScienceDirect Theoretical Computer Science journal homepage: www.elsevier.com/locate/tcs Note Computing the longest topological
Theoretical Computer Science 410 (2009) 2759 2766 Contents lists available at ScienceDirect Theoretical Computer Science journal homepage: www.elsevier.com/locate/tcs Note Computing the longest topological
On Pattern Matching With Swaps
 On Pattern Matching With Swaps Fouad B. Chedid Dhofar University, Salalah, Oman Notre Dame University - Louaize, Lebanon P.O.Box: 2509, Postal Code 211 Salalah, Oman Tel: +968 23237200 Fax: +968 23237720
On Pattern Matching With Swaps Fouad B. Chedid Dhofar University, Salalah, Oman Notre Dame University - Louaize, Lebanon P.O.Box: 2509, Postal Code 211 Salalah, Oman Tel: +968 23237200 Fax: +968 23237720
An Approximation Algorithm for Constructing Error Detecting Prefix Codes
 An Approximation Algorithm for Constructing Error Detecting Prefix Codes Artur Alves Pessoa artur@producao.uff.br Production Engineering Department Universidade Federal Fluminense, Brazil September 2,
An Approximation Algorithm for Constructing Error Detecting Prefix Codes Artur Alves Pessoa artur@producao.uff.br Production Engineering Department Universidade Federal Fluminense, Brazil September 2,
p 3 p 2 p 4 q 2 q 7 q 1 q 3 q 6 q 5
 Discrete Fréchet distance Consider Professor Bille going for a walk with his personal dog. The professor follows a path of points p 1,..., p n and the dog follows a path of points q 1,..., q m. We assume
Discrete Fréchet distance Consider Professor Bille going for a walk with his personal dog. The professor follows a path of points p 1,..., p n and the dog follows a path of points q 1,..., q m. We assume
Enhancing Active Automata Learning by a User Log Based Metric
 Master Thesis Computing Science Radboud University Enhancing Active Automata Learning by a User Log Based Metric Author Petra van den Bos First Supervisor prof. dr. Frits W. Vaandrager Second Supervisor
Master Thesis Computing Science Radboud University Enhancing Active Automata Learning by a User Log Based Metric Author Petra van den Bos First Supervisor prof. dr. Frits W. Vaandrager Second Supervisor
Parikh s theorem. Håkan Lindqvist
 Parikh s theorem Håkan Lindqvist Abstract This chapter will discuss Parikh s theorem and provide a proof for it. The proof is done by induction over a set of derivation trees, and using the Parikh mappings
Parikh s theorem Håkan Lindqvist Abstract This chapter will discuss Parikh s theorem and provide a proof for it. The proof is done by induction over a set of derivation trees, and using the Parikh mappings
More Dynamic Programming
 CS 374: Algorithms & Models of Computation, Spring 2017 More Dynamic Programming Lecture 14 March 9, 2017 Chandra Chekuri (UIUC) CS374 1 Spring 2017 1 / 42 What is the running time of the following? Consider
CS 374: Algorithms & Models of Computation, Spring 2017 More Dynamic Programming Lecture 14 March 9, 2017 Chandra Chekuri (UIUC) CS374 1 Spring 2017 1 / 42 What is the running time of the following? Consider
More Dynamic Programming
 Algorithms & Models of Computation CS/ECE 374, Fall 2017 More Dynamic Programming Lecture 14 Tuesday, October 17, 2017 Sariel Har-Peled (UIUC) CS374 1 Fall 2017 1 / 48 What is the running time of the following?
Algorithms & Models of Computation CS/ECE 374, Fall 2017 More Dynamic Programming Lecture 14 Tuesday, October 17, 2017 Sariel Har-Peled (UIUC) CS374 1 Fall 2017 1 / 48 What is the running time of the following?
On-line String Matching in Highly Similar DNA Sequences
 On-line String Matching in Highly Similar DNA Sequences Nadia Ben Nsira 1,2,ThierryLecroq 1,,MouradElloumi 2 1 LITIS EA 4108, Normastic FR3638, University of Rouen, France 2 LaTICE, University of Tunis
On-line String Matching in Highly Similar DNA Sequences Nadia Ben Nsira 1,2,ThierryLecroq 1,,MouradElloumi 2 1 LITIS EA 4108, Normastic FR3638, University of Rouen, France 2 LaTICE, University of Tunis
New successor rules for constructing de Bruijn sequences
 New successor rules for constructing de Bruijn sequences Dennis Wong Northwest Missouri State University Daniel Gabric and Joe Sawada University of Guelph, CAN Aaron Williams Simon s Rock, USA Southeastern
New successor rules for constructing de Bruijn sequences Dennis Wong Northwest Missouri State University Daniel Gabric and Joe Sawada University of Guelph, CAN Aaron Williams Simon s Rock, USA Southeastern
The maximum edge-disjoint paths problem in complete graphs
 Theoretical Computer Science 399 (2008) 128 140 www.elsevier.com/locate/tcs The maximum edge-disjoint paths problem in complete graphs Adrian Kosowski Department of Algorithms and System Modeling, Gdańsk
Theoretical Computer Science 399 (2008) 128 140 www.elsevier.com/locate/tcs The maximum edge-disjoint paths problem in complete graphs Adrian Kosowski Department of Algorithms and System Modeling, Gdańsk
Streaming Algorithms for Optimal Generation of Random Bits
 Streaming Algorithms for Optimal Generation of Random Bits ongchao Zhou Electrical Engineering Department California Institute of echnology Pasadena, CA 925 Email: hzhou@caltech.edu Jehoshua Bruck Electrical
Streaming Algorithms for Optimal Generation of Random Bits ongchao Zhou Electrical Engineering Department California Institute of echnology Pasadena, CA 925 Email: hzhou@caltech.edu Jehoshua Bruck Electrical
Lecture 15. Saad Mneimneh
 Computat onal Biology Lecture 15 DNA sequencing Shortest common superstring SCS An elegant theoretical abstraction, but fundamentally flawed R. Karp Given a set of fragments F, Find the shortest string
Computat onal Biology Lecture 15 DNA sequencing Shortest common superstring SCS An elegant theoretical abstraction, but fundamentally flawed R. Karp Given a set of fragments F, Find the shortest string
DNA sequencing. Bad example (repeats) Lecture 15. Shortest common superstring SCS. Given a set of fragments F,
 Computat onal Biology Lecture 15 DNA sequencing Shortest common superstring SCS An elegant theoretical abstraction, but fundamentally flawed R. Karp Given a set of fragments F, Find the shortest string
Computat onal Biology Lecture 15 DNA sequencing Shortest common superstring SCS An elegant theoretical abstraction, but fundamentally flawed R. Karp Given a set of fragments F, Find the shortest string
Alternative Algorithms for Lyndon Factorization
 Alternative Algorithms for Lyndon Factorization Suhpal Singh Ghuman 1, Emanuele Giaquinta 2, and Jorma Tarhio 1 1 Department of Computer Science and Engineering, Aalto University P.O.B. 15400, FI-00076
Alternative Algorithms for Lyndon Factorization Suhpal Singh Ghuman 1, Emanuele Giaquinta 2, and Jorma Tarhio 1 1 Department of Computer Science and Engineering, Aalto University P.O.B. 15400, FI-00076
Discrete Wiskunde II. Lecture 5: Shortest Paths & Spanning Trees
 , 2009 Lecture 5: Shortest Paths & Spanning Trees University of Twente m.uetz@utwente.nl wwwhome.math.utwente.nl/~uetzm/dw/ Shortest Path Problem "#$%&'%()*%"()$#+,&- Given directed "#$%&'()*+,%+('-*.#/'01234564'.*,'7+"-%/8',&'5"4'84%#3
, 2009 Lecture 5: Shortest Paths & Spanning Trees University of Twente m.uetz@utwente.nl wwwhome.math.utwente.nl/~uetzm/dw/ Shortest Path Problem "#$%&'%()*%"()$#+,&- Given directed "#$%&'()*+,%+('-*.#/'01234564'.*,'7+"-%/8',&'5"4'84%#3
CMPSCI 311: Introduction to Algorithms Second Midterm Exam
 CMPSCI 311: Introduction to Algorithms Second Midterm Exam April 11, 2018. Name: ID: Instructions: Answer the questions directly on the exam pages. Show all your work for each question. Providing more
CMPSCI 311: Introduction to Algorithms Second Midterm Exam April 11, 2018. Name: ID: Instructions: Answer the questions directly on the exam pages. Show all your work for each question. Providing more
Foreword. Grammatical inference. Examples of sequences. Sources. Example of problems expressed by sequences Switching the light
 Foreword Vincent Claveau IRISA - CNRS Rennes, France In the course of the course supervised symbolic machine learning technique concept learning (i.e. 2 classes) INSA 4 Sources s of sequences Slides and
Foreword Vincent Claveau IRISA - CNRS Rennes, France In the course of the course supervised symbolic machine learning technique concept learning (i.e. 2 classes) INSA 4 Sources s of sequences Slides and
CSE 431/531: Analysis of Algorithms. Dynamic Programming. Lecturer: Shi Li. Department of Computer Science and Engineering University at Buffalo
 CSE 431/531: Analysis of Algorithms Dynamic Programming Lecturer: Shi Li Department of Computer Science and Engineering University at Buffalo Paradigms for Designing Algorithms Greedy algorithm Make a
CSE 431/531: Analysis of Algorithms Dynamic Programming Lecturer: Shi Li Department of Computer Science and Engineering University at Buffalo Paradigms for Designing Algorithms Greedy algorithm Make a
On the Average Complexity of Brzozowski s Algorithm for Deterministic Automata with a Small Number of Final States
 On the Average Complexity of Brzozowski s Algorithm for Deterministic Automata with a Small Number of Final States Sven De Felice 1 and Cyril Nicaud 2 1 LIAFA, Université Paris Diderot - Paris 7 & CNRS
On the Average Complexity of Brzozowski s Algorithm for Deterministic Automata with a Small Number of Final States Sven De Felice 1 and Cyril Nicaud 2 1 LIAFA, Université Paris Diderot - Paris 7 & CNRS
Optimal prefix codes for pairs of geometrically-distributed random variables
 Author manuscript, published in IEEE International Symposium on Information Theory (ISIT'06), United States (2006) Optimal prefix codes for pairs of geometrically-distributed random variables Frédériue
Author manuscript, published in IEEE International Symposium on Information Theory (ISIT'06), United States (2006) Optimal prefix codes for pairs of geometrically-distributed random variables Frédériue
arxiv: v1 [cs.dm] 26 Apr 2010
![arxiv: v1 [cs.dm] 26 Apr 2010 arxiv: v1 [cs.dm] 26 Apr 2010](/thumbs/73/68325063.jpg) A Simple Polynomial Algorithm for the Longest Path Problem on Cocomparability Graphs George B. Mertzios Derek G. Corneil arxiv:1004.4560v1 [cs.dm] 26 Apr 2010 Abstract Given a graph G, the longest path
A Simple Polynomial Algorithm for the Longest Path Problem on Cocomparability Graphs George B. Mertzios Derek G. Corneil arxiv:1004.4560v1 [cs.dm] 26 Apr 2010 Abstract Given a graph G, the longest path
Automata-based Verification - III
 CS3172: Advanced Algorithms Automata-based Verification - III Howard Barringer Room KB2.20/22: email: howard.barringer@manchester.ac.uk March 2005 Third Topic Infinite Word Automata Motivation Büchi Automata
CS3172: Advanced Algorithms Automata-based Verification - III Howard Barringer Room KB2.20/22: email: howard.barringer@manchester.ac.uk March 2005 Third Topic Infinite Word Automata Motivation Büchi Automata
Sorting suffixes of two-pattern strings
 Sorting suffixes of two-pattern strings Frantisek Franek W. F. Smyth Algorithms Research Group Department of Computing & Software McMaster University Hamilton, Ontario Canada L8S 4L7 April 19, 2004 Abstract
Sorting suffixes of two-pattern strings Frantisek Franek W. F. Smyth Algorithms Research Group Department of Computing & Software McMaster University Hamilton, Ontario Canada L8S 4L7 April 19, 2004 Abstract
Fragment Assembly of DNA
 Wright State University CORE Scholar Computer Science and Engineering Faculty Publications Computer Science and Engineering 2003 Fragment Assembly of DNA Dan E. Krane Wright State University - Main Campus,
Wright State University CORE Scholar Computer Science and Engineering Faculty Publications Computer Science and Engineering 2003 Fragment Assembly of DNA Dan E. Krane Wright State University - Main Campus,
Computing Techniques for Parallel and Distributed Systems with an Application to Data Compression. Sergio De Agostino Sapienza University di Rome
 Computing Techniques for Parallel and Distributed Systems with an Application to Data Compression Sergio De Agostino Sapienza University di Rome Parallel Systems A parallel random access machine (PRAM)
Computing Techniques for Parallel and Distributed Systems with an Application to Data Compression Sergio De Agostino Sapienza University di Rome Parallel Systems A parallel random access machine (PRAM)
Bandwidth: Communicate large complex & highly detailed 3D models through lowbandwidth connection (e.g. VRML over the Internet)
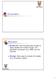 Compression Motivation Bandwidth: Communicate large complex & highly detailed 3D models through lowbandwidth connection (e.g. VRML over the Internet) Storage: Store large & complex 3D models (e.g. 3D scanner
Compression Motivation Bandwidth: Communicate large complex & highly detailed 3D models through lowbandwidth connection (e.g. VRML over the Internet) Storage: Store large & complex 3D models (e.g. 3D scanner
Module 5 EMBEDDED WAVELET CODING. Version 2 ECE IIT, Kharagpur
 Module 5 EMBEDDED WAVELET CODING Lesson 13 Zerotree Approach. Instructional Objectives At the end of this lesson, the students should be able to: 1. Explain the principle of embedded coding. 2. Show the
Module 5 EMBEDDED WAVELET CODING Lesson 13 Zerotree Approach. Instructional Objectives At the end of this lesson, the students should be able to: 1. Explain the principle of embedded coding. 2. Show the
String Regularities and Degenerate Strings
 M. Sc. Thesis Defense Md. Faizul Bari (100705050P) Supervisor: Dr. M. Sohel Rahman String Regularities and Degenerate Strings Department of Computer Science and Engineering Bangladesh University of Engineering
M. Sc. Thesis Defense Md. Faizul Bari (100705050P) Supervisor: Dr. M. Sohel Rahman String Regularities and Degenerate Strings Department of Computer Science and Engineering Bangladesh University of Engineering
Theory of Computation Time Complexity
 Theory of Computation Time Complexity Bow-Yaw Wang Academia Sinica Spring 2012 Bow-Yaw Wang (Academia Sinica) Time Complexity Spring 2012 1 / 59 Time for Deciding a Language Let us consider A = {0 n 1
Theory of Computation Time Complexity Bow-Yaw Wang Academia Sinica Spring 2012 Bow-Yaw Wang (Academia Sinica) Time Complexity Spring 2012 1 / 59 Time for Deciding a Language Let us consider A = {0 n 1
Even More on Dynamic Programming
 Algorithms & Models of Computation CS/ECE 374, Fall 2017 Even More on Dynamic Programming Lecture 15 Thursday, October 19, 2017 Sariel Har-Peled (UIUC) CS374 1 Fall 2017 1 / 26 Part I Longest Common Subsequence
Algorithms & Models of Computation CS/ECE 374, Fall 2017 Even More on Dynamic Programming Lecture 15 Thursday, October 19, 2017 Sariel Har-Peled (UIUC) CS374 1 Fall 2017 1 / 26 Part I Longest Common Subsequence
A BIJECTION BETWEEN WELL-LABELLED POSITIVE PATHS AND MATCHINGS
 Séminaire Lotharingien de Combinatoire 63 (0), Article B63e A BJECTON BETWEEN WELL-LABELLED POSTVE PATHS AND MATCHNGS OLVER BERNARD, BERTRAND DUPLANTER, AND PHLPPE NADEAU Abstract. A well-labelled positive
Séminaire Lotharingien de Combinatoire 63 (0), Article B63e A BJECTON BETWEEN WELL-LABELLED POSTVE PATHS AND MATCHNGS OLVER BERNARD, BERTRAND DUPLANTER, AND PHLPPE NADEAU Abstract. A well-labelled positive
SIGNAL COMPRESSION Lecture 7. Variable to Fix Encoding
 SIGNAL COMPRESSION Lecture 7 Variable to Fix Encoding 1. Tunstall codes 2. Petry codes 3. Generalized Tunstall codes for Markov sources (a presentation of the paper by I. Tabus, G. Korodi, J. Rissanen.
SIGNAL COMPRESSION Lecture 7 Variable to Fix Encoding 1. Tunstall codes 2. Petry codes 3. Generalized Tunstall codes for Markov sources (a presentation of the paper by I. Tabus, G. Korodi, J. Rissanen.
The Lefthanded Local Lemma characterizes chordal dependency graphs
 The Lefthanded Local Lemma characterizes chordal dependency graphs Wesley Pegden March 30, 2012 Abstract Shearer gave a general theorem characterizing the family L of dependency graphs labeled with probabilities
The Lefthanded Local Lemma characterizes chordal dependency graphs Wesley Pegden March 30, 2012 Abstract Shearer gave a general theorem characterizing the family L of dependency graphs labeled with probabilities
Algorithms: Lecture 12. Chalmers University of Technology
 Algorithms: Lecture 1 Chalmers University of Technology Today s Topics Shortest Paths Network Flow Algorithms Shortest Path in a Graph Shortest Path Problem Shortest path network. Directed graph G = (V,
Algorithms: Lecture 1 Chalmers University of Technology Today s Topics Shortest Paths Network Flow Algorithms Shortest Path in a Graph Shortest Path Problem Shortest path network. Directed graph G = (V,
Minmax Tree Cover in the Euclidean Space
 Journal of Graph Algorithms and Applications http://jgaa.info/ vol. 15, no. 3, pp. 345 371 (2011) Minmax Tree Cover in the Euclidean Space Seigo Karakawa 1 Ehab Morsy 1 Hiroshi Nagamochi 1 1 Department
Journal of Graph Algorithms and Applications http://jgaa.info/ vol. 15, no. 3, pp. 345 371 (2011) Minmax Tree Cover in the Euclidean Space Seigo Karakawa 1 Ehab Morsy 1 Hiroshi Nagamochi 1 1 Department
Theoretical Computer Science. Dynamic rank/select structures with applications to run-length encoded texts
 Theoretical Computer Science 410 (2009) 4402 4413 Contents lists available at ScienceDirect Theoretical Computer Science journal homepage: www.elsevier.com/locate/tcs Dynamic rank/select structures with
Theoretical Computer Science 410 (2009) 4402 4413 Contents lists available at ScienceDirect Theoretical Computer Science journal homepage: www.elsevier.com/locate/tcs Dynamic rank/select structures with
1 Alphabets and Languages
 1 Alphabets and Languages Look at handout 1 (inference rules for sets) and use the rules on some examples like {a} {{a}} {a} {a, b}, {a} {{a}}, {a} {{a}}, {a} {a, b}, a {{a}}, a {a, b}, a {{a}}, a {a,
1 Alphabets and Languages Look at handout 1 (inference rules for sets) and use the rules on some examples like {a} {{a}} {a} {a, b}, {a} {{a}}, {a} {{a}}, {a} {a, b}, a {{a}}, a {a, b}, a {{a}}, a {a,
