Efficient Polynomial-Time Algorithms for Variants of the Multiple Constrained LCS Problem
|
|
|
- Elinor Pope
- 5 years ago
- Views:
Transcription
1 Efficient Polynomial-Time Algorithms for Variants of the Multiple Constrained LCS Problem Hsing-Yen Ann National Center for High-Performance Computing Tainan 74147, Taiwan Chang-Biau Yang and Chiou-Ting Tseng Department of Computer Science and Engineering National Sun Yat-sen University, Kaohsiung 80424, Taiwan Abstract In this paper, we revisit a recent variant of the longest common subsequence problem, the string-excluding constrained LCS (STR-EC-LCS) problem, which was first addressed by Chen and Chao [Journal of Combinatorial Optimization, 21(3), 2011]. Given two sequences X and Y of lengths n and m, respectively, and a constraint string P of length r, we are to find a common subsequence Z of X and Y which excludes P as a substring and the length of Z is maximized. This problem cannot be correctly solved by the previously proposed algorithm. Thus, we give a correct algorithm with O(nmr) time to solve it. Keywords design of algorithms; longest common subsequence; constrained LCS; NP-hardness; biological application I. INTRODUCTION Computing the similarity of two sequences is one of the most important fundamentals in computer field. Given a sequence X of length n over some fixed alphabet Σ, a subsequence of X is formed by deleting zero or more symbols from X arbitrarily. We use X = x 1 x 2 x n to denote the list of symbols in X. A substring X i..j = x i x i+1 x j of X consists of successive symbols in X, where 1 i j n. A substring X i..j is called a prefix or a suffix of X if i = 1 or j = n, respectively. A sequence is called a common subsequence of two sequences X and Y if it is a subsequence of both X and Y. Given two sequences X and Y, the longest common subsequence (LCS) problem is to find out a subsequence of X and Y where its length is longest among all common subsequences of the two given sequences. The LCS problem is a well-known measurement for computing the similarity of two strings. It can be widely applied in diverse areas, such as file comparison, pattern matching and computational biology. The most referred algorithm was proposed by Wagner and Fischer [21] which solves the LCS problem in quadratic time with the dynamic programming (DP) technique. Other advanced algorithms have also been proposed for the past decades [1, 2, 11, 12, 22]. When the number of input sequences is not fixed, finding the LCS of the multiple sequences has been proved as NP-hard [15], and therefore some approximate and heuristic algorithms have been proposed [5, 17]. Applying the constraints to the LCS problem is meaningful for some biological applications [18]. Therefore, the constrained LCS (CLCS) problem, a recent variant of the LCS problem which was first addressed by Tsai [19], has gained much attention. Given two input sequences X and Y of lengths n and m, respectively, and a constrained sequence P of length r, the CLCS problem is to find the common subsequences Z of X and Y such that P is a subsequence of Z and the length of Z is maximized. In the following, without loss of generality, we may assume that r m n. The most referred algorithms were presented independently [4, 7] which solve the CLCS problem based on the DP technique in O(nmr) time and space. Some improved algorithms have also been proposed. Iliopoulos and Rahman [13] consider the matched pairs and employed a special data structure, van Emde Boas tree [20], to solve this problem. Ann et al. [3] considered the sequences which are in run-length encoding (RLE) format, a lineartime compressing technique, and greatly reduce the number of elements required to be evaluated in the DP lattice. Moreover, Peng et al. [16] extended the problem to the one with weighted constraints, a more generalized problem. Referring to the variant of multiple constraints, the multiple CLCS problem was also proven to be NP-hard [10]. Recently, Gotthilf et al. [9] proposed a variant of the CLCS problem, the restricted LCS problem, which excludes the given constraint as a subsequence of the answer. Furthermore, they proved that the restricted LCS problem also becomes NP-hard when the number of constraints is not fixed. Independently, Chen and Chao [6] addressed the more generalized forms of the CLCS problem, the generalized constrained longest common subsequence (GC-LCS) problem. Given two input sequences X and Y of lengths n and m, respectively, and a constraint string P of length r, the GC-LCS problem is a set of four problems which are to find the longest common subsequence of X and Y including (or excluding) P as a subsequence (or substring), respectively. Table I shows the four problems. Evidently, the original CLCS problem addressed by Tsai [19] is specified as SEQ-IC-LCS, and the restricted LCS problem addressed by Gotthilf et al. [9] is specified as SEQ- EC-LCS. For the problems in Table I except SEQ-IC-LCS, Chen and Chao proposed the corresponding algorithms which are all in O(nmr) time. However, as we will show soon, their algorithm for STR-EC-LCS cannot work correctly. In this paper, we focus on the STR-EC-LCS problem and its variants. In Section II, we review Chen and Chao s algorithm and give a counter example which breaks their algorithm. In Section III, a correct straightforward backtracking algorithm is given, though it runs in exponential time. Then, we propose an efficient algorithm in Section IV. And finally, the conclusion goes in Section V.
2 TABLE I THE FOUR GC-LCS PROBLEMS DEFINED BY CHEN AND CHAO [6] AND THE TIME COMPLEXITIES OF THEIR PROPOSED METHODS. As subsequence As substring Including P SEQ-IC-LCS STR-IC-LCS O(nmr) Excluding P SEQ-EC-LCS O(nmr) STR-EC-LCS O(nmr) II. A COUNTER EXAMPLE TO CHEN AND CHAO S ALGORITHM Definition 1. (STR-EC-LCS problem [6]). Given two input sequences X and Y of lengths n and m, respectively, and a constraint string P of length r, the STR-EC-LCS (string excluding) problem is to find the longest common subsequence Z of X and Y excluding P as a substring. For the STR-EC-LCS problem, Chen and Chao [6] proposed an dynamic programming algorithm with O(nmr) time to solve it. Their recurrence formula is shown in Figure 1, where L Chen (i, j, k) denotes the length of the LCS of X 1..i and Y 1..j excluding P 1..k as a substring. Now we provide a simple counter example, X = axbc, Y = abyc and P = ac, which breaks Chen and Chao s algorithm for the STR-EC-LCS problem. Evidently, the LCS of X and Y, abc, does not contain ac as a substring. That is to say, abc excludes the constraint P. However, Chen and Chao s algorithm would return the answer L Chen (4, 4, 2) = 2, where the corresponding result sequence is ab or bc. Part of the recursive process of calling L Chen (4, 4, 2) is illustrated in Figure 2. One can see that their answer is not correct and the length is shorter than abc = 3. It concludes that Chen and Chao s algorithm fails to solve the STR-EC-LCS problem. III. A STRAIGHTFORWARD BACKTRACKING ALGORITHM In this section, we give a simple straightforward backtracking algorithm which correctly solves the STR-EC-LCS problem. Although it runs in exponential time, we will show that it can be reduced to an efficient polynomial-time algorithm with similar ideas in the next section. Algorithm 1 shows a backtracking algorithm for solving the STR-EC-LCS problem. We denote L 1 (i, j, Tail) as the longest length of the common subsequence of X 1..i and Y 1..j such that the concatenation of the common subsequence and string Tail excludes P as a substring. For each recursive step, string Tail denotes the tail of the possible answer sequence obtained by the previous step. The answer of STR-EC-LCS is obtained by invoking L 1 (n, m, ϕ) recursively, where ϕ denotes the empty string. We consider the example, X = axbc, Y = abyc and P = ac, which breaks Chen and Chao s algorithm for the STR-EC-LCS problem in the previous section. Part of the recursive process of invoking L 1 (4, 4, ϕ) is illustrated in Figure 3. The bold edges indicate the path corresponding to the answer, abc. We also note that L 1 (0, 0, ac ) = since the constraint is violated, which means that node L 1 (0, 0, ac ) is never considered as a part of the answer. It is not difficult to examine the correctness of Algorithm 1. The boundary conditions are handled in Lines 1-4, where Lines 3-4 deal with the normal terminations. Lines 1-2 deal with the case that the constraint P has been contained in Tail as a substring, or more precisely, a prefix. The constraint has been violated in this case, thus is returned which means that we would never consider this state as a part of the answer. Lines 5-7 consider the case that x i and y j are matched. In this case, we call L 1 recursively to determine whether the matched symbol x i should be chosen as a part of the answer or not. If we choose x i as a part of the answer, x i will be inserted in front of Tail to form a new string New Tail, where denotes the string concatenation. Lines 8-9 consider the case of mismatch in which either x i or y j will be dropped. It is evident that even if we use the dynamic programming technique to reduce the total number of states in the recursive process, there are still O(nm Σ m ) states in the worst case. Theorem 1. Algorithm 1 can correctly solve the STR-EC-LCS problem in exponential time. IV. OUR ALGORITHM FOR THE STR-EC-LCS PROBLEM Although Algorithm 1 requires exponential time, in this section, we will show that the required time can be reduced and thus an efficient algorithm with polynomial time is proposed. After investigating the relationship between the string Tail and the constraint P in Algorithm 1, we find that a property holds when solving STR-EC-LCS with the recursive function L 1, as shown in Lemma 1. Definition 2. (Longest prefix-suffix). Given two strings S 1 and S 2, we denote LPS(S 1, S 2 ) as the longest prefix of S 1 that matches a suffix of S 2. Lemma 1. For a given string S excluding P as a substring, L 1 (i, j, S) = L 1 (i, j, LPS(S, P )). Proof. We assume L 1 (i, j, S) L 1 (i, j, LPS(S, P )) and prove it by contradiction. For ease of presentation, we denote S = V V where V = LPS(S, P ). Besides, we denote the corresponding strings of L 1 (i, j, S) and L 1 (i, j, V ) as U 1 and U 2, respectively. In other words, U 1 = L 1 (i, j, S) and U 2 = L 1 (i, j, V ). If L 1 (i, j, S) L 1 (i, j, V ), then two possible cases would be considered as follows. 1) L 1 (i, j, S) > L 1 (i, j, V ). It means that U 1 S excludes P as a substring and U 1 V contains P as a substring, otherwise U 1 can be concatenated with V and gains a better length U 1 > L 1 (i, j, V ). However, U 1 V is a prefix of U 1 S. If U 1 V contains P as a substring, U 1 S would also contain P as a substring, which is a contradiction. 2) L 1 (i, j, S) < L 1 (i, j, V ). It means that U 2 S = U 2 V V contains P as a substring and U 2 V excludes P as a substring. By definition, S = V V excludes P, the occurrence of P in U 2 V V should start inside U 2 and end inside V. Thus, there would exist a suffix of P that is also a prefix of V V and its length is longer than V.
3 L Chen (i, j, k) = L Chen {(i 1, j 1, k) } if k = 1 and x i = y j = p k, 1 + LChen (i 1, j 1, k 1), max if k 2 and x L Chen (i 1, j 1, k) i = y j = p k, if x i = y j and (k = 0, or k > 0 and x i p k ), 1 + L Chen (i 1, j 1, k) { } LChen (i 1, j, k 1), max L Chen (i, j 1, k 1) if x i y j. L Chen (i, 0, k) = L Chen (0, j, k) = 0 for any 0 i n, 0 j m, 0 k r. Fig. 1. Chen and Chao s recurrence formula [6] for the STR-EC-LCS problem. L Chen (4,4,2) +1 (c) L Chen (3,3,2) L Chen (3,3,1) L Chen (2,3,2) L Chen (3,2,2) L Chen (2,3,1) L Chen (3,2,1) +1 (b) +1 (b) L Chen (2,1,2) L Chen (2,1,1) L Chen (1,1,2) L Chen (2,0,2) L Chen (1,1,1) L Chen (2,0,1) +1 (a) L Chen (0,0,2) L Chen (0,0,1) Fig. 2. Part of the process of Chen and Chao s recurrence formula, where X = axbc, Y = abyc and P = ac. The result sequences are ab and bc. Algorithm 1 L 1 (i, j, Tail) 1: if P is a prefix of Tail then { boundary condition: violating the constraint} 2: return 3: else if i = 0 or j = 0 then { normal boundary condition} 4: return 0 5: else if x i = y j then { matched} 6: New Tail x i Tail { new tail is formed by concatenation of x i and Tail} 7: return max{l 1 (i 1, j 1, New Tail) + 1, L 1 (i 1, j, Tail), L 1 (i, j 1, Tail)} 8: else { x i y j, mismatched} 9: return max{l 1 (i 1, j, Tail), L 1 (i, j 1, Tail)} 10: end if However, V = LPS(S, P ), which is a contradiction. For example, if P = abc, it is clear that L 1 (i, j, bca ) = L 1 (i, j, bcdddee ) = L 1 (i, j, bc ) and L 1 (i, j, aaaaa ) = L 1 (i, j, ϕ). According to Lemma 1, before each call of function L 1 in Algorithm 1, we may shorten the length of the third parameter Tail by replacing it with LPS(Tail, P ). Since LPS(Tail, P ) is always a suffix of P, it becomes feasible to record each state of the recursive process in Algorithm 1 with a 3-tuple (i, j, l), where l denotes the length of LPS(Tail, P ). With this trick, we present an improved version of Algorithm 1 as shown in Algorithm 2. The answer of STR-EC-LCS is got by calling L 2 (n, m, 0) recursively, that is, L 2 (n, m, 0) = L 1 (n, m, ϕ). The most notable difference between Algorithm 1 and Algorithm 2 locates in Lines 5-7. In Line 6, P (r l+1)..r represents the longest prefix of Tail which matches a suffix of P. After x i is inserted in front of P (r l+1)..r, we will compute the new length l of the longest prefix of x i P (r l+1)..r which matches a suffix of P. Obviously, there are at most O(nmr) states in the recursive process when calling L 2 (n, m, 0), which are much less than O(nm Σ m ) states when calling L 1 (n, m, ϕ). Given two strings S 1 and S 2, LPS(S 1, S 2 ) can be computed in O( S 1 S 2 ) time by a straightforward algorithm. By utilizing such a straightforward algorithm, Algorithm 2 achieves O(nmr) O(r 2 ) = O(nmr 3 ) running time. To improve the efficiency of Algorithm 2, we have to find a more efficient way to compute the value LPS(x i
4 L 1(4,4,ψ) = 3 +1 (c) L 1(3,3, c ) = 2 L 1(3,4,ψ) L 1(4,3,ψ) L 1 (2,3, c ) L 1 (3,2, c ) = 2 +1 (b) L 1(2,1, bc ) = 1 L 1(2,2, c ) L 1(3,1, c ) L 1 (1,1, bc ) = 1 L 1 (2,0, bc ) L 1 (2,1, c ) L 1 (3,0, c ) +1 (a) L 1(0,0, abc ) = 0 L 1(0,1, bc ) L 1(1,0, bc ) L 1(1,1, c ) = 0 L 1(2,0, c ) +1 (a) L 1 (0,0, ac ) = - L 1 (0,1, c ) = 0 L 1 (1,0, c ) = 0 Fig. 3. Part of the process of invoking L 1 (4, 4, ϕ), where X = axbc, Y = abyc and P = ac. The result shows that L 1 (4, 4, ϕ) = 3. Algorithm 2 L 2 (i, j, l) 1: if l = P then { boundary condition: violating the constraint} 2: return 3: else if i = 0 or j = 0 then { normal boundary condition} 4: return 0 5: else if x i = y j then { matched} 6: l LPS(x i P (r l+1)..r, P ) 7: return max{l 2 (i 1, j 1, l ) + 1, L 2 (i 1, j, l), L 2 (i, j 1, l)} 8: else { x i y j, mismatched} 9: return max{l 2 (i 1, j, l), L 2 (i, j 1, l)} 10: end if P (r l+1)..r, P ) in Line 6. By observing the paths in the recursive process of calling L 2, the behaviors of the third parameter l can be described as follows. When x i is equal to p (r l), we set the new prefix-suffix length l with l + 1, which means that the length of matched suffix of P increases by one. On the other hand, when x i is not equal to p (r l), the length of matched suffix of P has to be shortened to the largest l such that P (r l+1)..(r l+l 1) = P (r l +2)..r and x i = p (r l +1). One can see that this process is similar to slide a pattern when executing Knuth-Morris-Pratt (KMP) algorithm [8, 14], a famous string matching algorithm. KMP algorithm searches for a pattern in a text efficiently by precomputing the prefix function of the given pattern [8] and then sliding the pattern efficiently when the matching failure occurs. Definition 3. (Prefix function [8]). Given a string S, the prefix function f(i) denotes the length of the longest prefix of S 1..i 1 that matches a suffix of S 1..i. Figure 4(a) shows the table of the pattern S = and its corresponding prefix function. For example, f(9) = 4, f(4) = 2 and f(2) = 0 illustrate the sliding process of KMP algorithm [8, 14] when the matching failure occurs after S 1..9 was matched, as shown in Figure 4(b). We should notice that the constraint P is backward matched when L 1 and L 2 are called recursively. Therefore, we will precompute the prefix function of the reversed string P of P. We should also notice that the amortized analysis of the linear-time matching, or more precisely, constant-time sliding, of KMP algorithm would be violated since there are more than one path in the recursive process of calling L 2. To ensure the sliding of the pattern to be executed efficiently, we define the next function as follows, which achieves constant query time by precomputing a two-dimensional table. Definition 4. (Next function). Given a string S and a symbol σ Σ, the next function π(i, σ) denotes the length of the longest prefix of S 1..i+1 that matches a suffix of S 1..i σ.
5 S a b a b b a b a b c f S 1..9 S 1..4 S 1..2 (a) (b) Fig. 4. An example for illustrating the prefix function. (a) The pattern S = and its corresponding prefix function. (b) The sliding process of S 1..9 when the matching failure occurs a b a b b a b a b c a b c S S 1..9 S 1..9 a S 1..9 b S 1..9 c (a) ababbabab ababbababa ababbababb (b) Fig. 5. An example for illustrating the next function. (a) The precomputed table for the next function π of the pattern S =. (b) The sliding results corresponding to π(9, a ), π(9, b ) and π(9, c ). Figure 5(a) shows the precomputed table for the next function of S =. Figure 5(b) shows the sliding result after appending each symbol in alphabet Σ to S For example, after appending symbol c to S 1..9, the matched prefix of pattern S extends to S However, after appending symbol a or b to S 1..9, the length of the matched prefix is shortened to π(9, a ) = 3 or π(9, b ) = 5, respectively. Algorithm 3 shows how to compute the table of next function π when a pattern S and its precomputed prefix function f are given. Clearly, each element can be computed in constant time and the whole table can be computed in O( S Σ ) time. Now go back to our aim, to improve the efficiency of Algorithm 2. One can easily see that the value LPS(x i P (r l+1)..r, P ) in Line 6 of Algorithm 2 can be answered in constant time after precomputing the next function π of the reversed string P. That is, LPS(x i P (r l+1)..r, P ) = π(l, x i ). We summarize the result in the following theorem. Theorem 2. Algorithm 2 solves the STR-EC-LCS problem in O(nmr) time. Proof. The correctness of Algorithm 2 has been discussed above. Now we consider its time complexity. Precompute the next function π of P takes O(r Σ ) time, and it is evident Algorithm 3 Compute-Next-Function 1: for σ Σ and σ s 1 do 2: π(0, σ) 0 3: end for 4: π(0, s 1 ) 1 5: for i = 1 to S 1 do 6: for σ Σ do 7: if σ = s i+1 then 8: π(i, σ) i + 1 9: else 10: π(i, σ) π(f(i), σ) 11: end if 12: end for 13: end for that Σ = O(n + m + r). Remind that we have assumed r m n in the beginning of this paper. Therefore, the overall time complexity is simplified from O(nmr + r Σ ) to O(nmr). V. CONCLUDING REMARKS In this paper, we study the STR-EC-LCS problem with single constraint. We give an algorithm which corrects Chen and Chao s algorithm with the same time complexity. For the STR-EC-LCS problem with multiple constraints, Chen and Chao [6] claimed that the problem is NP-hard. However, the claim may not be correct, since we are developing an efficient algorithm, which requires only polynomial time. The new algorithm is somewhat complicated, so we omit its description here. Another similar variant was also addressed by Chen and Chao [6], the multiple STR-IC-LCS problem, in which the resulting sequence has to contain all constraints as its substrings. In fact, it is not difficult to see that this problem is complementary to the multiple STR-EC-LCS problem. We will also try to solve this problem in the future. REFERENCES [1] H. Y. Ann, C. B. Yang, Y. H. Peng, and B. C. Liaw, Efficient algorithms for the block edit problems, Information and Computation, Vol. 208, No. 3, pp , [2] H. Y. Ann, C. B. Yang, C. T. Tseng, and C. Y. Hor, A fast and simple algorithm for computing the longest common subsequence of run-length encoded strings, Information Processing Letters, Vol. 108, pp , [3] H. Y. Ann, C. B. Yang, C. T. Tseng, and C. Y. Hor, Fast algorithms for computing the constrained LCS of run-length encoded strings, Proc. of the 2009 International Conference on Bioinformatics and Computational Biology (BIOCOMP 09), Las Vegas, USA, pp , [4] A. N. Arslan and Ö. Eğecioğlu, Algorithms for the constrained longest common subsequence problems, In-
6 ternational Journal of Foundations Computer Science, Vol. 16, No. 6, pp , [5] C. Blum, M. J. Blesa, and M. López-Ibáñez, Beam search for the longest common subsequence problem, Computers and Operations Research, Vol. 36, No. 12, pp , [6] Y. C. Chen and K. M. Chao, On the generalized constrained longest common subsequence problems, Journal of Combinatorial Optimization, Vol. 21, No. 3, pp , [7] F. Y. L. Chin, A. D. Santis, A. L. Ferrara, N. L. Ho, and S. K. Kim, A simple algorithm for the constrained sequence problems, Information Processing Letters, Vol. 90, No. 4, pp , [8] T. H. Cormen, C. E. Leiserson, R. L. Rivest, and C. Stein, Introduction to Algorithms. MIT Press, second ed., [9] Z. Gotthilf, D. Hermelin, G. M. Landau, and M. Lewenstein, Restricted LCS, String Processing and Information Retrieval (SPIRE), Lecture Notes in Computer Science, Vol. 6393, pp , [10] Z. Gotthilf, D. Hermelin, and M. Lewenstein, Constrained LCS: hardness and approximation, Combinatorial Pattern Matching, 19th Annual Symposium (CPM2008), Pisa, Italy, pp , [11] D. S. Hirschberg, Algorithms for the longest common subsequence problem, Journal of the ACM, Vol. 24, No. 4, pp , [12] J. W. Hunt and T. G. Szymanski, A fast algorithm for computing longest common subsequences, Communications of the ACM, Vol. 20, No. 5, pp , [13] C. S. Iliopoulos and M. S. Rahman, New efficient algorithms for the LCS and constrained LCS problems, Information Processing Letters, Vol. 106, No. 1, pp , [14] D. E. Knuth, J. H. Morris Jr., and V. Pratt, Fast pattern matching in strings, SIAM Journal on Computing, Vol. 6, No. 2, pp , [15] D. Maier, The complexity of some problems on subsequences and supersequences, Journal of the ACM, Vol. 25, pp , [16] Y. H. Peng, C. B. Yang, K. S. Huang, and K. T. Tseng, An algorithm and applications to sequence alignment with weighted constraints, International Journal of Foundations of Computer Science, Vol. 21, No. 1, pp , [17] S. J. Shyu and C. Y. Tsai, Finding the longest common subsequence for multiple biological sequences by ant colony optimization, Computers and Operations Research, Vol. 36, No. 1, pp , [18] C. Y. Tang, C. L. Lu, M. D. T. Chang, Y. T. Tsai, Y. J. Sun, K. M. Chao, J. M. Chang, Y. H. Chiou, C. M. Wu, H. T. Chang, and W. I. Chou, Constrained multiple sequence alignment tool development and its application to RNase family alignment, Journal of Bioinformatics and Computational Biology, Vol. 1, pp , [19] Y. T. Tsai, The constrained longest common subsequence problem, Information Processing Letters, Vol. 88, No. 4, pp , [20] P. van Emde Boas, Preserving order in a forest in less than logarithmic time and linear space, Information Processing Letters, Vol. 6, No. 3, pp , [21] R. Wagner and M. Fischer, The string-to-string correction problem, Journal of the ACM, Vol. 21, No. 1, pp , [22] C. B. Yang and R. C. T. Lee, Systolic algorithm for the longest common subsequence problem, Journal of the Chinese Institute of Engineers, Vol. 10, No. 6, pp , 1987.
Efficient Algorithms for the Longest Common Subsequence Problem with Sequential Substring Constraints
 2011 11th IEEE International Conference on Bioinformatics and Bioengineering Efficient Algorithms for the Longest Common Subsequence Problem with Sequential Substring Constraints Chiou-Ting Tseng, Chang-Biau
2011 11th IEEE International Conference on Bioinformatics and Bioengineering Efficient Algorithms for the Longest Common Subsequence Problem with Sequential Substring Constraints Chiou-Ting Tseng, Chang-Biau
Information Processing Letters. A fast and simple algorithm for computing the longest common subsequence of run-length encoded strings
 Information Processing Letters 108 (2008) 360 364 Contents lists available at ScienceDirect Information Processing Letters www.vier.com/locate/ipl A fast and simple algorithm for computing the longest
Information Processing Letters 108 (2008) 360 364 Contents lists available at ScienceDirect Information Processing Letters www.vier.com/locate/ipl A fast and simple algorithm for computing the longest
Minimal Height and Sequence Constrained Longest Increasing Subsequence
 Minimal Height and Sequence Constrained Longest Increasing Subsequence Chiou-Ting Tseng, Chang-Biau Yang and Hsing-Yen Ann Department of Computer Science and Engineering National Sun Yat-sen University,
Minimal Height and Sequence Constrained Longest Increasing Subsequence Chiou-Ting Tseng, Chang-Biau Yang and Hsing-Yen Ann Department of Computer Science and Engineering National Sun Yat-sen University,
The NP-hardness and APX-hardness of the Two-Dimensional Largest Common Substructure Problems
 The NP-hardness and APX-hardness of the Two-Dimensional Largest Common Substructure Problems Syuan-Zong Chiou a, Chang-Biau Yang a and Yung-Hsing Peng b a Department of Computer Science and Engineering
The NP-hardness and APX-hardness of the Two-Dimensional Largest Common Substructure Problems Syuan-Zong Chiou a, Chang-Biau Yang a and Yung-Hsing Peng b a Department of Computer Science and Engineering
arxiv: v1 [cs.ds] 2 Dec 2009
![arxiv: v1 [cs.ds] 2 Dec 2009 arxiv: v1 [cs.ds] 2 Dec 2009](/thumbs/71/66068077.jpg) Variants of Constrained Longest Common Subsequence arxiv:0912.0368v1 [cs.ds] 2 Dec 2009 Paola Bonizzoni Gianluca Della Vedova Riccardo Dondi Yuri Pirola Abstract In this work, we consider a variant of
Variants of Constrained Longest Common Subsequence arxiv:0912.0368v1 [cs.ds] 2 Dec 2009 Paola Bonizzoni Gianluca Della Vedova Riccardo Dondi Yuri Pirola Abstract In this work, we consider a variant of
The Longest Common Subsequence Problem
 Advanced Studies in Biology, Vol. 4, 2012, no. 8, 373-380 The Longest Common Subsequence Problem Anna Gorbenko Department of Intelligent Systems and Robotics Ural Federal University 620083 Ekaterinburg,
Advanced Studies in Biology, Vol. 4, 2012, no. 8, 373-380 The Longest Common Subsequence Problem Anna Gorbenko Department of Intelligent Systems and Robotics Ural Federal University 620083 Ekaterinburg,
Pattern Matching. a b a c a a b. a b a c a b. a b a c a b. Pattern Matching 1
 Pattern Matching a b a c a a b 1 4 3 2 Pattern Matching 1 Outline and Reading Strings ( 9.1.1) Pattern matching algorithms Brute-force algorithm ( 9.1.2) Boyer-Moore algorithm ( 9.1.3) Knuth-Morris-Pratt
Pattern Matching a b a c a a b 1 4 3 2 Pattern Matching 1 Outline and Reading Strings ( 9.1.1) Pattern matching algorithms Brute-force algorithm ( 9.1.2) Boyer-Moore algorithm ( 9.1.3) Knuth-Morris-Pratt
A Simple Linear Space Algorithm for Computing a Longest Common Increasing Subsequence
 A Simple Linear Space Algorithm for Computing a Longest Common Increasing Subsequence Danlin Cai, Daxin Zhu, Lei Wang, and Xiaodong Wang Abstract This paper presents a linear space algorithm for finding
A Simple Linear Space Algorithm for Computing a Longest Common Increasing Subsequence Danlin Cai, Daxin Zhu, Lei Wang, and Xiaodong Wang Abstract This paper presents a linear space algorithm for finding
Efficient Algorithms forregular Expression Constrained Sequence Alignment p. 1/35
 Efficient Algorithms for Regular Expression Constrained Sequence Alignment Yun-Sheng Chung, Chin Lung Lu, and Chuan Yi Tang Department of Computer Science National Tsing Hua University, Taiwan Department
Efficient Algorithms for Regular Expression Constrained Sequence Alignment Yun-Sheng Chung, Chin Lung Lu, and Chuan Yi Tang Department of Computer Science National Tsing Hua University, Taiwan Department
Module 9: Tries and String Matching
 Module 9: Tries and String Matching CS 240 - Data Structures and Data Management Sajed Haque Veronika Irvine Taylor Smith Based on lecture notes by many previous cs240 instructors David R. Cheriton School
Module 9: Tries and String Matching CS 240 - Data Structures and Data Management Sajed Haque Veronika Irvine Taylor Smith Based on lecture notes by many previous cs240 instructors David R. Cheriton School
Theoretical Computer Science
 Theoretical Computer Science 410 (2009) 2759 2766 Contents lists available at ScienceDirect Theoretical Computer Science journal homepage: www.elsevier.com/locate/tcs Note Computing the longest topological
Theoretical Computer Science 410 (2009) 2759 2766 Contents lists available at ScienceDirect Theoretical Computer Science journal homepage: www.elsevier.com/locate/tcs Note Computing the longest topological
Pattern Matching. a b a c a a b. a b a c a b. a b a c a b. Pattern Matching Goodrich, Tamassia
 Pattern Matching a b a c a a b 1 4 3 2 Pattern Matching 1 Brute-Force Pattern Matching ( 11.2.1) The brute-force pattern matching algorithm compares the pattern P with the text T for each possible shift
Pattern Matching a b a c a a b 1 4 3 2 Pattern Matching 1 Brute-Force Pattern Matching ( 11.2.1) The brute-force pattern matching algorithm compares the pattern P with the text T for each possible shift
On Pattern Matching With Swaps
 On Pattern Matching With Swaps Fouad B. Chedid Dhofar University, Salalah, Oman Notre Dame University - Louaize, Lebanon P.O.Box: 2509, Postal Code 211 Salalah, Oman Tel: +968 23237200 Fax: +968 23237720
On Pattern Matching With Swaps Fouad B. Chedid Dhofar University, Salalah, Oman Notre Dame University - Louaize, Lebanon P.O.Box: 2509, Postal Code 211 Salalah, Oman Tel: +968 23237200 Fax: +968 23237720
Overview. Knuth-Morris-Pratt & Boyer-Moore Algorithms. Notation Review (2) Notation Review (1) The Kunth-Morris-Pratt (KMP) Algorithm
 Knuth-Morris-Pratt & s by Robert C. St.Pierre Overview Notation review Knuth-Morris-Pratt algorithm Discussion of the Algorithm Example Boyer-Moore algorithm Discussion of the Algorithm Example Applications
Knuth-Morris-Pratt & s by Robert C. St.Pierre Overview Notation review Knuth-Morris-Pratt algorithm Discussion of the Algorithm Example Boyer-Moore algorithm Discussion of the Algorithm Example Applications
/633 Introduction to Algorithms Lecturer: Michael Dinitz Topic: Dynamic Programming II Date: 10/12/17
 601.433/633 Introduction to Algorithms Lecturer: Michael Dinitz Topic: Dynamic Programming II Date: 10/12/17 12.1 Introduction Today we re going to do a couple more examples of dynamic programming. While
601.433/633 Introduction to Algorithms Lecturer: Michael Dinitz Topic: Dynamic Programming II Date: 10/12/17 12.1 Introduction Today we re going to do a couple more examples of dynamic programming. While
String Range Matching
 String Range Matching Juha Kärkkäinen, Dominik Kempa, and Simon J. Puglisi Department of Computer Science, University of Helsinki Helsinki, Finland firstname.lastname@cs.helsinki.fi Abstract. Given strings
String Range Matching Juha Kärkkäinen, Dominik Kempa, and Simon J. Puglisi Department of Computer Science, University of Helsinki Helsinki, Finland firstname.lastname@cs.helsinki.fi Abstract. Given strings
2. Exact String Matching
 2. Exact String Matching Let T = T [0..n) be the text and P = P [0..m) the pattern. We say that P occurs in T at position j if T [j..j + m) = P. Example: P = aine occurs at position 6 in T = karjalainen.
2. Exact String Matching Let T = T [0..n) be the text and P = P [0..m) the pattern. We say that P occurs in T at position j if T [j..j + m) = P. Example: P = aine occurs at position 6 in T = karjalainen.
Implementing Approximate Regularities
 Implementing Approximate Regularities Manolis Christodoulakis Costas S. Iliopoulos Department of Computer Science King s College London Kunsoo Park School of Computer Science and Engineering, Seoul National
Implementing Approximate Regularities Manolis Christodoulakis Costas S. Iliopoulos Department of Computer Science King s College London Kunsoo Park School of Computer Science and Engineering, Seoul National
Quadratic-backtracking Algorithm for String Reconstruction from Substring Compositions
 Quadratic-backtracking Algorithm for String Reconstruction from Substring Compositions Jayadev Acharya UCSD jacharya@ucsdedu Hirakendu Das Yahoo Labs hdas@yahoo-inccom Olgica Milenkovic UIUC milenkov@uiucedu
Quadratic-backtracking Algorithm for String Reconstruction from Substring Compositions Jayadev Acharya UCSD jacharya@ucsdedu Hirakendu Das Yahoo Labs hdas@yahoo-inccom Olgica Milenkovic UIUC milenkov@uiucedu
Analysis of Algorithms Prof. Karen Daniels
 UMass Lowell Computer Science 91.503 Analysis of Algorithms Prof. Karen Daniels Spring, 2012 Tuesday, 4/24/2012 String Matching Algorithms Chapter 32* * Pseudocode uses 2 nd edition conventions 1 Chapter
UMass Lowell Computer Science 91.503 Analysis of Algorithms Prof. Karen Daniels Spring, 2012 Tuesday, 4/24/2012 String Matching Algorithms Chapter 32* * Pseudocode uses 2 nd edition conventions 1 Chapter
A GREEDY APPROXIMATION ALGORITHM FOR CONSTRUCTING SHORTEST COMMON SUPERSTRINGS *
 A GREEDY APPROXIMATION ALGORITHM FOR CONSTRUCTING SHORTEST COMMON SUPERSTRINGS * 1 Jorma Tarhio and Esko Ukkonen Department of Computer Science, University of Helsinki Tukholmankatu 2, SF-00250 Helsinki,
A GREEDY APPROXIMATION ALGORITHM FOR CONSTRUCTING SHORTEST COMMON SUPERSTRINGS * 1 Jorma Tarhio and Esko Ukkonen Department of Computer Science, University of Helsinki Tukholmankatu 2, SF-00250 Helsinki,
String Search. 6th September 2018
 String Search 6th September 2018 Search for a given (short) string in a long string Search problems have become more important lately The amount of stored digital information grows steadily (rapidly?)
String Search 6th September 2018 Search for a given (short) string in a long string Search problems have become more important lately The amount of stored digital information grows steadily (rapidly?)
15 Text search. P.D. Dr. Alexander Souza. Winter term 11/12
 Algorithms Theory 15 Text search P.D. Dr. Alexander Souza Text search Various scenarios: Dynamic texts Text editors Symbol manipulators Static texts Literature databases Library systems Gene databases
Algorithms Theory 15 Text search P.D. Dr. Alexander Souza Text search Various scenarios: Dynamic texts Text editors Symbol manipulators Static texts Literature databases Library systems Gene databases
Finding all covers of an indeterminate string in O(n) time on average
 Finding all covers of an indeterminate string in O(n) time on average Md. Faizul Bari, M. Sohel Rahman, and Rifat Shahriyar Department of Computer Science and Engineering Bangladesh University of Engineering
Finding all covers of an indeterminate string in O(n) time on average Md. Faizul Bari, M. Sohel Rahman, and Rifat Shahriyar Department of Computer Science and Engineering Bangladesh University of Engineering
Computing a Longest Common Palindromic Subsequence
 Fundamenta Informaticae 129 (2014) 1 12 1 DOI 10.3233/FI-2014-860 IOS Press Computing a Longest Common Palindromic Subsequence Shihabur Rahman Chowdhury, Md. Mahbubul Hasan, Sumaiya Iqbal, M. Sohel Rahman
Fundamenta Informaticae 129 (2014) 1 12 1 DOI 10.3233/FI-2014-860 IOS Press Computing a Longest Common Palindromic Subsequence Shihabur Rahman Chowdhury, Md. Mahbubul Hasan, Sumaiya Iqbal, M. Sohel Rahman
INF 4130 / /8-2017
 INF 4130 / 9135 28/8-2017 Algorithms, efficiency, and complexity Problem classes Problems can be divided into sets (classes). Problem classes are defined by the type of algorithm that can (or cannot) solve
INF 4130 / 9135 28/8-2017 Algorithms, efficiency, and complexity Problem classes Problems can be divided into sets (classes). Problem classes are defined by the type of algorithm that can (or cannot) solve
Maximal Unbordered Factors of Random Strings arxiv: v1 [cs.ds] 14 Apr 2017
![Maximal Unbordered Factors of Random Strings arxiv: v1 [cs.ds] 14 Apr 2017 Maximal Unbordered Factors of Random Strings arxiv: v1 [cs.ds] 14 Apr 2017](/thumbs/74/70164248.jpg) Maximal Unbordered Factors of Random Strings arxiv:1704.04472v1 [cs.ds] 14 Apr 2017 Patrick Hagge Cording 1 and Mathias Bæk Tejs Knudsen 2 1 DTU Compute, Technical University of Denmark, phaco@dtu.dk 2
Maximal Unbordered Factors of Random Strings arxiv:1704.04472v1 [cs.ds] 14 Apr 2017 Patrick Hagge Cording 1 and Mathias Bæk Tejs Knudsen 2 1 DTU Compute, Technical University of Denmark, phaco@dtu.dk 2
String Matching with Variable Length Gaps
 String Matching with Variable Length Gaps Philip Bille, Inge Li Gørtz, Hjalte Wedel Vildhøj, and David Kofoed Wind Technical University of Denmark Abstract. We consider string matching with variable length
String Matching with Variable Length Gaps Philip Bille, Inge Li Gørtz, Hjalte Wedel Vildhøj, and David Kofoed Wind Technical University of Denmark Abstract. We consider string matching with variable length
A Survey of the Longest Common Subsequence Problem and Its. Related Problems
 Survey of the Longest Common Subsequence Problem and Its Related Problems Survey of the Longest Common Subsequence Problem and Its Related Problems Thesis Submitted to the Faculty of Department of Computer
Survey of the Longest Common Subsequence Problem and Its Related Problems Survey of the Longest Common Subsequence Problem and Its Related Problems Thesis Submitted to the Faculty of Department of Computer
Algorithm Theory. 13 Text Search - Knuth, Morris, Pratt, Boyer, Moore. Christian Schindelhauer
 Algorithm Theory 13 Text Search - Knuth, Morris, Pratt, Boyer, Moore Institut für Informatik Wintersemester 2007/08 Text Search Scenarios Static texts Literature databases Library systems Gene databases
Algorithm Theory 13 Text Search - Knuth, Morris, Pratt, Boyer, Moore Institut für Informatik Wintersemester 2007/08 Text Search Scenarios Static texts Literature databases Library systems Gene databases
Lecture 3: String Matching
 COMP36111: Advanced Algorithms I Lecture 3: String Matching Ian Pratt-Hartmann Room KB2.38: email: ipratt@cs.man.ac.uk 2017 18 Outline The string matching problem The Rabin-Karp algorithm The Knuth-Morris-Pratt
COMP36111: Advanced Algorithms I Lecture 3: String Matching Ian Pratt-Hartmann Room KB2.38: email: ipratt@cs.man.ac.uk 2017 18 Outline The string matching problem The Rabin-Karp algorithm The Knuth-Morris-Pratt
Structure-Based Comparison of Biomolecules
 Structure-Based Comparison of Biomolecules Benedikt Christoph Wolters Seminar Bioinformatics Algorithms RWTH AACHEN 07/17/2015 Outline 1 Introduction and Motivation Protein Structure Hierarchy Protein
Structure-Based Comparison of Biomolecules Benedikt Christoph Wolters Seminar Bioinformatics Algorithms RWTH AACHEN 07/17/2015 Outline 1 Introduction and Motivation Protein Structure Hierarchy Protein
Pattern Matching (Exact Matching) Overview
 CSI/BINF 5330 Pattern Matching (Exact Matching) Young-Rae Cho Associate Professor Department of Computer Science Baylor University Overview Pattern Matching Exhaustive Search DFA Algorithm KMP Algorithm
CSI/BINF 5330 Pattern Matching (Exact Matching) Young-Rae Cho Associate Professor Department of Computer Science Baylor University Overview Pattern Matching Exhaustive Search DFA Algorithm KMP Algorithm
PATTERN MATCHING WITH SWAPS IN PRACTICE
 International Journal of Foundations of Computer Science c World Scientific Publishing Company PATTERN MATCHING WITH SWAPS IN PRACTICE MATTEO CAMPANELLI Università di Catania, Scuola Superiore di Catania
International Journal of Foundations of Computer Science c World Scientific Publishing Company PATTERN MATCHING WITH SWAPS IN PRACTICE MATTEO CAMPANELLI Università di Catania, Scuola Superiore di Catania
Quantum pattern matching fast on average
 Quantum pattern matching fast on average Ashley Montanaro Department of Computer Science, University of Bristol, UK 12 January 2015 Pattern matching In the traditional pattern matching problem, we seek
Quantum pattern matching fast on average Ashley Montanaro Department of Computer Science, University of Bristol, UK 12 January 2015 Pattern matching In the traditional pattern matching problem, we seek
Efficient Sequential Algorithms, Comp309
 Efficient Sequential Algorithms, Comp309 University of Liverpool 2010 2011 Module Organiser, Igor Potapov Part 2: Pattern Matching References: T. H. Cormen, C. E. Leiserson, R. L. Rivest Introduction to
Efficient Sequential Algorithms, Comp309 University of Liverpool 2010 2011 Module Organiser, Igor Potapov Part 2: Pattern Matching References: T. H. Cormen, C. E. Leiserson, R. L. Rivest Introduction to
arxiv: v1 [cs.ds] 15 Feb 2012
![arxiv: v1 [cs.ds] 15 Feb 2012 arxiv: v1 [cs.ds] 15 Feb 2012](/thumbs/92/110725364.jpg) Linear-Space Substring Range Counting over Polylogarithmic Alphabets Travis Gagie 1 and Pawe l Gawrychowski 2 1 Aalto University, Finland travis.gagie@aalto.fi 2 Max Planck Institute, Germany gawry@cs.uni.wroc.pl
Linear-Space Substring Range Counting over Polylogarithmic Alphabets Travis Gagie 1 and Pawe l Gawrychowski 2 1 Aalto University, Finland travis.gagie@aalto.fi 2 Max Planck Institute, Germany gawry@cs.uni.wroc.pl
INF 4130 / /8-2014
 INF 4130 / 9135 26/8-2014 Mandatory assignments («Oblig-1», «-2», and «-3»): All three must be approved Deadlines around: 25. sept, 25. oct, and 15. nov Other courses on similar themes: INF-MAT 3370 INF-MAT
INF 4130 / 9135 26/8-2014 Mandatory assignments («Oblig-1», «-2», and «-3»): All three must be approved Deadlines around: 25. sept, 25. oct, and 15. nov Other courses on similar themes: INF-MAT 3370 INF-MAT
Lecture 1 : Data Compression and Entropy
 CPS290: Algorithmic Foundations of Data Science January 8, 207 Lecture : Data Compression and Entropy Lecturer: Kamesh Munagala Scribe: Kamesh Munagala In this lecture, we will study a simple model for
CPS290: Algorithmic Foundations of Data Science January 8, 207 Lecture : Data Compression and Entropy Lecturer: Kamesh Munagala Scribe: Kamesh Munagala In this lecture, we will study a simple model for
arxiv: v2 [cs.ds] 5 Mar 2014
![arxiv: v2 [cs.ds] 5 Mar 2014 arxiv: v2 [cs.ds] 5 Mar 2014](/thumbs/93/113153836.jpg) Order-preserving pattern matching with k mismatches Pawe l Gawrychowski 1 and Przemys law Uznański 2 1 Max-Planck-Institut für Informatik, Saarbrücken, Germany 2 LIF, CNRS and Aix-Marseille Université,
Order-preserving pattern matching with k mismatches Pawe l Gawrychowski 1 and Przemys law Uznański 2 1 Max-Planck-Institut für Informatik, Saarbrücken, Germany 2 LIF, CNRS and Aix-Marseille Université,
Longest Common Subsequence Problem for Run-Length-Encoded Strings
 JOURNAL OF COMPUTERS, VOL. 9, NO. 8, AUGUST 2014 1769 Longest Common Subsequence Problem for Run-Length-Encoded Strings Shegufta Bakht Ahsan a, Syeda Persia Aziz a, M. Sohel Rahman a a AlEDA Group Department
JOURNAL OF COMPUTERS, VOL. 9, NO. 8, AUGUST 2014 1769 Longest Common Subsequence Problem for Run-Length-Encoded Strings Shegufta Bakht Ahsan a, Syeda Persia Aziz a, M. Sohel Rahman a a AlEDA Group Department
Optimal spaced seeds for faster approximate string matching
 Optimal spaced seeds for faster approximate string matching Martin Farach-Colton Gad M. Landau S. Cenk Sahinalp Dekel Tsur Abstract Filtering is a standard technique for fast approximate string matching
Optimal spaced seeds for faster approximate string matching Martin Farach-Colton Gad M. Landau S. Cenk Sahinalp Dekel Tsur Abstract Filtering is a standard technique for fast approximate string matching
Optimal spaced seeds for faster approximate string matching
 Optimal spaced seeds for faster approximate string matching Martin Farach-Colton Gad M. Landau S. Cenk Sahinalp Dekel Tsur Abstract Filtering is a standard technique for fast approximate string matching
Optimal spaced seeds for faster approximate string matching Martin Farach-Colton Gad M. Landau S. Cenk Sahinalp Dekel Tsur Abstract Filtering is a standard technique for fast approximate string matching
Knuth-Morris-Pratt Algorithm
 Knuth-Morris-Pratt Algorithm Jayadev Misra June 5, 2017 The Knuth-Morris-Pratt string matching algorithm (KMP) locates all occurrences of a pattern string in a text string in linear time (in the combined
Knuth-Morris-Pratt Algorithm Jayadev Misra June 5, 2017 The Knuth-Morris-Pratt string matching algorithm (KMP) locates all occurrences of a pattern string in a text string in linear time (in the combined
Average Complexity of Exact and Approximate Multiple String Matching
 Average Complexity of Exact and Approximate Multiple String Matching Gonzalo Navarro Department of Computer Science University of Chile gnavarro@dcc.uchile.cl Kimmo Fredriksson Department of Computer Science
Average Complexity of Exact and Approximate Multiple String Matching Gonzalo Navarro Department of Computer Science University of Chile gnavarro@dcc.uchile.cl Kimmo Fredriksson Department of Computer Science
On-line String Matching in Highly Similar DNA Sequences
 On-line String Matching in Highly Similar DNA Sequences Nadia Ben Nsira 1,2,ThierryLecroq 1,,MouradElloumi 2 1 LITIS EA 4108, Normastic FR3638, University of Rouen, France 2 LaTICE, University of Tunis
On-line String Matching in Highly Similar DNA Sequences Nadia Ben Nsira 1,2,ThierryLecroq 1,,MouradElloumi 2 1 LITIS EA 4108, Normastic FR3638, University of Rouen, France 2 LaTICE, University of Tunis
Computing the Longest Common Subsequence of Multiple RLE Strings
 The 29th Workshop on Combinatorial Mathematics an Computation Theory Computing the Longest Common Subsequence of Multiple RLE Strings Ling-Chih Yao an Kuan-Yu Chen Grauate Institute of Networking an Multimeia
The 29th Workshop on Combinatorial Mathematics an Computation Theory Computing the Longest Common Subsequence of Multiple RLE Strings Ling-Chih Yao an Kuan-Yu Chen Grauate Institute of Networking an Multimeia
String Regularities and Degenerate Strings
 M. Sc. Thesis Defense Md. Faizul Bari (100705050P) Supervisor: Dr. M. Sohel Rahman String Regularities and Degenerate Strings Department of Computer Science and Engineering Bangladesh University of Engineering
M. Sc. Thesis Defense Md. Faizul Bari (100705050P) Supervisor: Dr. M. Sohel Rahman String Regularities and Degenerate Strings Department of Computer Science and Engineering Bangladesh University of Engineering
Pushdown Automata. Notes on Automata and Theory of Computation. Chia-Ping Chen
 Pushdown Automata Notes on Automata and Theory of Computation Chia-Ping Chen Department of Computer Science and Engineering National Sun Yat-Sen University Kaohsiung, Taiwan ROC Pushdown Automata p. 1
Pushdown Automata Notes on Automata and Theory of Computation Chia-Ping Chen Department of Computer Science and Engineering National Sun Yat-Sen University Kaohsiung, Taiwan ROC Pushdown Automata p. 1
Complete words and superpatterns
 Honors Theses Mathematics Spring 2015 Complete words and superpatterns Yonah Biers-Ariel Penrose Library, Whitman College Permanent URL: http://hdl.handle.net/10349/20151078 This thesis has been deposited
Honors Theses Mathematics Spring 2015 Complete words and superpatterns Yonah Biers-Ariel Penrose Library, Whitman College Permanent URL: http://hdl.handle.net/10349/20151078 This thesis has been deposited
Consensus Optimizing Both Distance Sum and Radius
 Consensus Optimizing Both Distance Sum and Radius Amihood Amir 1, Gad M. Landau 2, Joong Chae Na 3, Heejin Park 4, Kunsoo Park 5, and Jeong Seop Sim 6 1 Bar-Ilan University, 52900 Ramat-Gan, Israel 2 University
Consensus Optimizing Both Distance Sum and Radius Amihood Amir 1, Gad M. Landau 2, Joong Chae Na 3, Heejin Park 4, Kunsoo Park 5, and Jeong Seop Sim 6 1 Bar-Ilan University, 52900 Ramat-Gan, Israel 2 University
1 Alphabets and Languages
 1 Alphabets and Languages Look at handout 1 (inference rules for sets) and use the rules on some examples like {a} {{a}} {a} {a, b}, {a} {{a}}, {a} {{a}}, {a} {a, b}, a {{a}}, a {a, b}, a {{a}}, a {a,
1 Alphabets and Languages Look at handout 1 (inference rules for sets) and use the rules on some examples like {a} {{a}} {a} {a, b}, {a} {{a}}, {a} {{a}}, {a} {a, b}, a {{a}}, a {a, b}, a {{a}}, a {a,
A Multiobjective Approach to the Weighted Longest Common Subsequence Problem
 A Multiobjective Approach to the Weighted Longest Common Subsequence Problem David Becerra, Juan Mendivelso, and Yoan Pinzón Universidad Nacional de Colombia Facultad de Ingeniería Department of Computer
A Multiobjective Approach to the Weighted Longest Common Subsequence Problem David Becerra, Juan Mendivelso, and Yoan Pinzón Universidad Nacional de Colombia Facultad de Ingeniería Department of Computer
Finding a longest common subsequence between a run-length-encoded string and an uncompressed string
 Journal of Complexity 24 (2008) 173 184 www.elsevier.com/locate/jco Finding a longest common subsequence between a run-length-encoded string and an uncompressed string J.J. Liu a, Y.L. Wang b,, R.C.T.
Journal of Complexity 24 (2008) 173 184 www.elsevier.com/locate/jco Finding a longest common subsequence between a run-length-encoded string and an uncompressed string J.J. Liu a, Y.L. Wang b,, R.C.T.
Lecture 2: Pairwise Alignment. CG Ron Shamir
 Lecture 2: Pairwise Alignment 1 Main source 2 Why compare sequences? Human hexosaminidase A vs Mouse hexosaminidase A 3 www.mathworks.com/.../jan04/bio_genome.html Sequence Alignment עימוד רצפים The problem:
Lecture 2: Pairwise Alignment 1 Main source 2 Why compare sequences? Human hexosaminidase A vs Mouse hexosaminidase A 3 www.mathworks.com/.../jan04/bio_genome.html Sequence Alignment עימוד רצפים The problem:
Introduction. An Introduction to Algorithms and Data Structures
 Introduction An Introduction to Algorithms and Data Structures Overview Aims This course is an introduction to the design, analysis and wide variety of algorithms (a topic often called Algorithmics ).
Introduction An Introduction to Algorithms and Data Structures Overview Aims This course is an introduction to the design, analysis and wide variety of algorithms (a topic often called Algorithmics ).
Lecture 9: The Splitting Method for SAT
 Lecture 9: The Splitting Method for SAT 1 Importance of SAT Cook-Levin Theorem: SAT is NP-complete. The reason why SAT is an important problem can be summarized as below: 1. A natural NP-Complete problem.
Lecture 9: The Splitting Method for SAT 1 Importance of SAT Cook-Levin Theorem: SAT is NP-complete. The reason why SAT is an important problem can be summarized as below: 1. A natural NP-Complete problem.
10-704: Information Processing and Learning Fall Lecture 10: Oct 3
 0-704: Information Processing and Learning Fall 206 Lecturer: Aarti Singh Lecture 0: Oct 3 Note: These notes are based on scribed notes from Spring5 offering of this course. LaTeX template courtesy of
0-704: Information Processing and Learning Fall 206 Lecturer: Aarti Singh Lecture 0: Oct 3 Note: These notes are based on scribed notes from Spring5 offering of this course. LaTeX template courtesy of
About the impossibility to prove P NP or P = NP and the pseudo-randomness in NP
 About the impossibility to prove P NP or P = NP and the pseudo-randomness in NP Prof. Marcel Rémon 1 arxiv:0904.0698v3 [cs.cc] 24 Mar 2016 Abstract The relationship between the complexity classes P and
About the impossibility to prove P NP or P = NP and the pseudo-randomness in NP Prof. Marcel Rémon 1 arxiv:0904.0698v3 [cs.cc] 24 Mar 2016 Abstract The relationship between the complexity classes P and
Chapter 3 Deterministic planning
 Chapter 3 Deterministic planning In this chapter we describe a number of algorithms for solving the historically most important and most basic type of planning problem. Two rather strong simplifying assumptions
Chapter 3 Deterministic planning In this chapter we describe a number of algorithms for solving the historically most important and most basic type of planning problem. Two rather strong simplifying assumptions
Chapter 3 Source Coding. 3.1 An Introduction to Source Coding 3.2 Optimal Source Codes 3.3 Shannon-Fano Code 3.4 Huffman Code
 Chapter 3 Source Coding 3. An Introduction to Source Coding 3.2 Optimal Source Codes 3.3 Shannon-Fano Code 3.4 Huffman Code 3. An Introduction to Source Coding Entropy (in bits per symbol) implies in average
Chapter 3 Source Coding 3. An Introduction to Source Coding 3.2 Optimal Source Codes 3.3 Shannon-Fano Code 3.4 Huffman Code 3. An Introduction to Source Coding Entropy (in bits per symbol) implies in average
arxiv: v1 [cs.cc] 4 Feb 2008
![arxiv: v1 [cs.cc] 4 Feb 2008 arxiv: v1 [cs.cc] 4 Feb 2008](/thumbs/72/66975294.jpg) On the complexity of finding gapped motifs Morris Michael a,1, François Nicolas a, and Esko Ukkonen a arxiv:0802.0314v1 [cs.cc] 4 Feb 2008 Abstract a Department of Computer Science, P. O. Box 68 (Gustaf
On the complexity of finding gapped motifs Morris Michael a,1, François Nicolas a, and Esko Ukkonen a arxiv:0802.0314v1 [cs.cc] 4 Feb 2008 Abstract a Department of Computer Science, P. O. Box 68 (Gustaf
Multiple Pattern Matching
 Multiple Pattern Matching Stephen Fulwider and Amar Mukherjee College of Engineering and Computer Science University of Central Florida Orlando, FL USA Email: {stephen,amar}@cs.ucf.edu Abstract In this
Multiple Pattern Matching Stephen Fulwider and Amar Mukherjee College of Engineering and Computer Science University of Central Florida Orlando, FL USA Email: {stephen,amar}@cs.ucf.edu Abstract In this
arxiv: v2 [cs.ds] 28 Jan 2009
![arxiv: v2 [cs.ds] 28 Jan 2009 arxiv: v2 [cs.ds] 28 Jan 2009](/thumbs/87/95711442.jpg) Minimax Trees in Linear Time Pawe l Gawrychowski 1 and Travis Gagie 2, arxiv:0812.2868v2 [cs.ds] 28 Jan 2009 1 Institute of Computer Science University of Wroclaw, Poland gawry1@gmail.com 2 Research Group
Minimax Trees in Linear Time Pawe l Gawrychowski 1 and Travis Gagie 2, arxiv:0812.2868v2 [cs.ds] 28 Jan 2009 1 Institute of Computer Science University of Wroclaw, Poland gawry1@gmail.com 2 Research Group
Dynamic Programming: Shortest Paths and DFA to Reg Exps
 CS 374: Algorithms & Models of Computation, Spring 207 Dynamic Programming: Shortest Paths and DFA to Reg Exps Lecture 8 March 28, 207 Chandra Chekuri (UIUC) CS374 Spring 207 / 56 Part I Shortest Paths
CS 374: Algorithms & Models of Computation, Spring 207 Dynamic Programming: Shortest Paths and DFA to Reg Exps Lecture 8 March 28, 207 Chandra Chekuri (UIUC) CS374 Spring 207 / 56 Part I Shortest Paths
Lecture 4 : Adaptive source coding algorithms
 Lecture 4 : Adaptive source coding algorithms February 2, 28 Information Theory Outline 1. Motivation ; 2. adaptive Huffman encoding ; 3. Gallager and Knuth s method ; 4. Dictionary methods : Lempel-Ziv
Lecture 4 : Adaptive source coding algorithms February 2, 28 Information Theory Outline 1. Motivation ; 2. adaptive Huffman encoding ; 3. Gallager and Knuth s method ; 4. Dictionary methods : Lempel-Ziv
Average Case Analysis. October 11, 2011
 Average Case Analysis October 11, 2011 Worst-case analysis Worst-case analysis gives an upper bound for the running time of a single execution of an algorithm with a worst-case input and worst-case random
Average Case Analysis October 11, 2011 Worst-case analysis Worst-case analysis gives an upper bound for the running time of a single execution of an algorithm with a worst-case input and worst-case random
Optimal Superprimitivity Testing for Strings
 Optimal Superprimitivity Testing for Strings Alberto Apostolico Martin Farach Costas S. Iliopoulos Fibonacci Report 90.7 August 1990 - Revised: March 1991 Abstract A string w covers another string z if
Optimal Superprimitivity Testing for Strings Alberto Apostolico Martin Farach Costas S. Iliopoulos Fibonacci Report 90.7 August 1990 - Revised: March 1991 Abstract A string w covers another string z if
Compressed Index for Dynamic Text
 Compressed Index for Dynamic Text Wing-Kai Hon Tak-Wah Lam Kunihiko Sadakane Wing-Kin Sung Siu-Ming Yiu Abstract This paper investigates how to index a text which is subject to updates. The best solution
Compressed Index for Dynamic Text Wing-Kai Hon Tak-Wah Lam Kunihiko Sadakane Wing-Kin Sung Siu-Ming Yiu Abstract This paper investigates how to index a text which is subject to updates. The best solution
Improving the KMP Algorithm by Using Properties of Fibonacci String
 Improving the KMP Algorithm by Using Properties of Fibonacci String Yi-Kung Shieh and R. C. T. Lee Department of Computer Science National Tsing Hua University d9762814@oz.nthu.edu.tw and rctlee@ncnu.edu.tw
Improving the KMP Algorithm by Using Properties of Fibonacci String Yi-Kung Shieh and R. C. T. Lee Department of Computer Science National Tsing Hua University d9762814@oz.nthu.edu.tw and rctlee@ncnu.edu.tw
(In)approximability Results for Pattern Matching Problems
 (In)approximability Results for Pattern Matching Problems Raphaël Clifford and Alexandru Popa Department of Computer Science University of Bristol Merchant Venturer s Building Woodland Road, Bristol, BS8
(In)approximability Results for Pattern Matching Problems Raphaël Clifford and Alexandru Popa Department of Computer Science University of Bristol Merchant Venturer s Building Woodland Road, Bristol, BS8
EE5139R: Problem Set 4 Assigned: 31/08/16, Due: 07/09/16
 EE539R: Problem Set 4 Assigned: 3/08/6, Due: 07/09/6. Cover and Thomas: Problem 3.5 Sets defined by probabilities: Define the set C n (t = {x n : P X n(x n 2 nt } (a We have = P X n(x n P X n(x n 2 nt
EE539R: Problem Set 4 Assigned: 3/08/6, Due: 07/09/6. Cover and Thomas: Problem 3.5 Sets defined by probabilities: Define the set C n (t = {x n : P X n(x n 2 nt } (a We have = P X n(x n P X n(x n 2 nt
Graduate Algorithms CS F-20 String Matching
 Graduate Algorithms CS673-2016F-20 String Matching David Galles Department of Computer Science University of San Francisco 20-0: String Matching Given a source text, and a string to match, where does the
Graduate Algorithms CS673-2016F-20 String Matching David Galles Department of Computer Science University of San Francisco 20-0: String Matching Given a source text, and a string to match, where does the
Aside: Golden Ratio. Golden Ratio: A universal law. Golden ratio φ = lim n = 1+ b n = a n 1. a n+1 = a n + b n, a n+b n a n
 Aside: Golden Ratio Golden Ratio: A universal law. Golden ratio φ = lim n a n+b n a n = 1+ 5 2 a n+1 = a n + b n, b n = a n 1 Ruta (UIUC) CS473 1 Spring 2018 1 / 41 CS 473: Algorithms, Spring 2018 Dynamic
Aside: Golden Ratio Golden Ratio: A universal law. Golden ratio φ = lim n a n+b n a n = 1+ 5 2 a n+1 = a n + b n, b n = a n 1 Ruta (UIUC) CS473 1 Spring 2018 1 / 41 CS 473: Algorithms, Spring 2018 Dynamic
Efficient (δ, γ)-pattern-matching with Don t Cares
 fficient (δ, γ)-pattern-matching with Don t Cares Yoan José Pinzón Ardila Costas S. Iliopoulos Manolis Christodoulakis Manal Mohamed King s College London, Department of Computer Science, London WC2R 2LS,
fficient (δ, γ)-pattern-matching with Don t Cares Yoan José Pinzón Ardila Costas S. Iliopoulos Manolis Christodoulakis Manal Mohamed King s College London, Department of Computer Science, London WC2R 2LS,
Finding Consensus Strings With Small Length Difference Between Input and Solution Strings
 Finding Consensus Strings With Small Length Difference Between Input and Solution Strings Markus L. Schmid Trier University, Fachbereich IV Abteilung Informatikwissenschaften, D-54286 Trier, Germany, MSchmid@uni-trier.de
Finding Consensus Strings With Small Length Difference Between Input and Solution Strings Markus L. Schmid Trier University, Fachbereich IV Abteilung Informatikwissenschaften, D-54286 Trier, Germany, MSchmid@uni-trier.de
Algorithm Design and Analysis
 Algorithm Design and Analysis LECTURE 8 Greedy Algorithms V Huffman Codes Adam Smith Review Questions Let G be a connected undirected graph with distinct edge weights. Answer true or false: Let e be the
Algorithm Design and Analysis LECTURE 8 Greedy Algorithms V Huffman Codes Adam Smith Review Questions Let G be a connected undirected graph with distinct edge weights. Answer true or false: Let e be the
CS481F01 Prelim 2 Solutions
 CS481F01 Prelim 2 Solutions A. Demers 7 Nov 2001 1 (30 pts = 4 pts each part + 2 free points). For this question we use the following notation: x y means x is a prefix of y m k n means m n k For each of
CS481F01 Prelim 2 Solutions A. Demers 7 Nov 2001 1 (30 pts = 4 pts each part + 2 free points). For this question we use the following notation: x y means x is a prefix of y m k n means m n k For each of
Alternative Algorithms for Lyndon Factorization
 Alternative Algorithms for Lyndon Factorization Suhpal Singh Ghuman 1, Emanuele Giaquinta 2, and Jorma Tarhio 1 1 Department of Computer Science and Engineering, Aalto University P.O.B. 15400, FI-00076
Alternative Algorithms for Lyndon Factorization Suhpal Singh Ghuman 1, Emanuele Giaquinta 2, and Jorma Tarhio 1 1 Department of Computer Science and Engineering, Aalto University P.O.B. 15400, FI-00076
Two Algorithms for LCS Consecutive Suffix Alignment
 Two Algorithms for LCS Consecutive Suffix Alignment Gad M. Landau Eugene Myers Michal Ziv-Ukelson Abstract The problem of aligning two sequences A and to determine their similarity is one of the fundamental
Two Algorithms for LCS Consecutive Suffix Alignment Gad M. Landau Eugene Myers Michal Ziv-Ukelson Abstract The problem of aligning two sequences A and to determine their similarity is one of the fundamental
Sequence comparison by compression
 Sequence comparison by compression Motivation similarity as a marker for homology. And homology is used to infer function. Sometimes, we are only interested in a numerical distance between two sequences.
Sequence comparison by compression Motivation similarity as a marker for homology. And homology is used to infer function. Sometimes, we are only interested in a numerical distance between two sequences.
Dynamic Programming: Shortest Paths and DFA to Reg Exps
 CS 374: Algorithms & Models of Computation, Fall 205 Dynamic Programming: Shortest Paths and DFA to Reg Exps Lecture 7 October 22, 205 Chandra & Manoj (UIUC) CS374 Fall 205 / 54 Part I Shortest Paths with
CS 374: Algorithms & Models of Computation, Fall 205 Dynamic Programming: Shortest Paths and DFA to Reg Exps Lecture 7 October 22, 205 Chandra & Manoj (UIUC) CS374 Fall 205 / 54 Part I Shortest Paths with
Dynamic Programming. Shuang Zhao. Microsoft Research Asia September 5, Dynamic Programming. Shuang Zhao. Outline. Introduction.
 Microsoft Research Asia September 5, 2005 1 2 3 4 Section I What is? Definition is a technique for efficiently recurrence computing by storing partial results. In this slides, I will NOT use too many formal
Microsoft Research Asia September 5, 2005 1 2 3 4 Section I What is? Definition is a technique for efficiently recurrence computing by storing partial results. In this slides, I will NOT use too many formal
Perfect Sorting by Reversals and Deletions/Insertions
 The Ninth International Symposium on Operations Research and Its Applications (ISORA 10) Chengdu-Jiuzhaigou, China, August 19 23, 2010 Copyright 2010 ORSC & APORC, pp. 512 518 Perfect Sorting by Reversals
The Ninth International Symposium on Operations Research and Its Applications (ISORA 10) Chengdu-Jiuzhaigou, China, August 19 23, 2010 Copyright 2010 ORSC & APORC, pp. 512 518 Perfect Sorting by Reversals
Lecture 4: September 19
 CSCI1810: Computational Molecular Biology Fall 2017 Lecture 4: September 19 Lecturer: Sorin Istrail Scribe: Cyrus Cousins Note: LaTeX template courtesy of UC Berkeley EECS dept. Disclaimer: These notes
CSCI1810: Computational Molecular Biology Fall 2017 Lecture 4: September 19 Lecturer: Sorin Istrail Scribe: Cyrus Cousins Note: LaTeX template courtesy of UC Berkeley EECS dept. Disclaimer: These notes
The Generalization of Quicksort
 The Generalization of Quicksort Shyue-Horng Shiau Chang-Biau Yang Ý Department of Computer Science and Engineering National Sun Yat-Sen University Kaohsiung, Taiwan 804, Republic of China shiaush@cse.nsysu.edu.tw
The Generalization of Quicksort Shyue-Horng Shiau Chang-Biau Yang Ý Department of Computer Science and Engineering National Sun Yat-Sen University Kaohsiung, Taiwan 804, Republic of China shiaush@cse.nsysu.edu.tw
De Bruijn Sequences for the Binary Strings with Maximum Density
 De Bruijn Sequences for the Binary Strings with Maximum Density Joe Sawada 1, Brett Stevens 2, and Aaron Williams 2 1 jsawada@uoguelph.ca School of Computer Science, University of Guelph, CANADA 2 brett@math.carleton.ca
De Bruijn Sequences for the Binary Strings with Maximum Density Joe Sawada 1, Brett Stevens 2, and Aaron Williams 2 1 jsawada@uoguelph.ca School of Computer Science, University of Guelph, CANADA 2 brett@math.carleton.ca
Each internal node v with d(v) children stores d 1 keys. k i 1 < key in i-th sub-tree k i, where we use k 0 = and k d =.
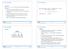 7.5 (a, b)-trees 7.5 (a, b)-trees Definition For b a an (a, b)-tree is a search tree with the following properties. all leaves have the same distance to the root. every internal non-root vertex v has at
7.5 (a, b)-trees 7.5 (a, b)-trees Definition For b a an (a, b)-tree is a search tree with the following properties. all leaves have the same distance to the root. every internal non-root vertex v has at
arxiv: v2 [cs.ds] 3 Oct 2017
![arxiv: v2 [cs.ds] 3 Oct 2017 arxiv: v2 [cs.ds] 3 Oct 2017](/thumbs/84/90870651.jpg) Orthogonal Vectors Indexing Isaac Goldstein 1, Moshe Lewenstein 1, and Ely Porat 1 1 Bar-Ilan University, Ramat Gan, Israel {goldshi,moshe,porately}@cs.biu.ac.il arxiv:1710.00586v2 [cs.ds] 3 Oct 2017 Abstract
Orthogonal Vectors Indexing Isaac Goldstein 1, Moshe Lewenstein 1, and Ely Porat 1 1 Bar-Ilan University, Ramat Gan, Israel {goldshi,moshe,porately}@cs.biu.ac.il arxiv:1710.00586v2 [cs.ds] 3 Oct 2017 Abstract
Guess & Check Codes for Deletions, Insertions, and Synchronization
 Guess & Check Codes for Deletions, Insertions, and Synchronization Serge Kas Hanna, Salim El Rouayheb ECE Department, Rutgers University sergekhanna@rutgersedu, salimelrouayheb@rutgersedu arxiv:759569v3
Guess & Check Codes for Deletions, Insertions, and Synchronization Serge Kas Hanna, Salim El Rouayheb ECE Department, Rutgers University sergekhanna@rutgersedu, salimelrouayheb@rutgersedu arxiv:759569v3
BLAST: Basic Local Alignment Search Tool
 .. CSC 448 Bioinformatics Algorithms Alexander Dekhtyar.. (Rapid) Local Sequence Alignment BLAST BLAST: Basic Local Alignment Search Tool BLAST is a family of rapid approximate local alignment algorithms[2].
.. CSC 448 Bioinformatics Algorithms Alexander Dekhtyar.. (Rapid) Local Sequence Alignment BLAST BLAST: Basic Local Alignment Search Tool BLAST is a family of rapid approximate local alignment algorithms[2].
Average Case Analysis of the Boyer-Moore Algorithm
 Average Case Analysis of the Boyer-Moore Algorithm TSUNG-HSI TSAI Institute of Statistical Science Academia Sinica Taipei 115 Taiwan e-mail: chonghi@stat.sinica.edu.tw URL: http://www.stat.sinica.edu.tw/chonghi/stat.htm
Average Case Analysis of the Boyer-Moore Algorithm TSUNG-HSI TSAI Institute of Statistical Science Academia Sinica Taipei 115 Taiwan e-mail: chonghi@stat.sinica.edu.tw URL: http://www.stat.sinica.edu.tw/chonghi/stat.htm
Computing repetitive structures in indeterminate strings
 Computing repetitive structures in indeterminate strings Pavlos Antoniou 1, Costas S. Iliopoulos 1, Inuka Jayasekera 1, Wojciech Rytter 2,3 1 Dept. of Computer Science, King s College London, London WC2R
Computing repetitive structures in indeterminate strings Pavlos Antoniou 1, Costas S. Iliopoulos 1, Inuka Jayasekera 1, Wojciech Rytter 2,3 1 Dept. of Computer Science, King s College London, London WC2R
arxiv: v3 [cs.fl] 2 Jul 2018
![arxiv: v3 [cs.fl] 2 Jul 2018 arxiv: v3 [cs.fl] 2 Jul 2018](/thumbs/93/111950553.jpg) COMPLEXITY OF PREIMAGE PROBLEMS FOR DETERMINISTIC FINITE AUTOMATA MIKHAIL V. BERLINKOV arxiv:1704.08233v3 [cs.fl] 2 Jul 2018 Institute of Natural Sciences and Mathematics, Ural Federal University, Ekaterinburg,
COMPLEXITY OF PREIMAGE PROBLEMS FOR DETERMINISTIC FINITE AUTOMATA MIKHAIL V. BERLINKOV arxiv:1704.08233v3 [cs.fl] 2 Jul 2018 Institute of Natural Sciences and Mathematics, Ural Federal University, Ekaterinburg,
Sorting suffixes of two-pattern strings
 Sorting suffixes of two-pattern strings Frantisek Franek W. F. Smyth Algorithms Research Group Department of Computing & Software McMaster University Hamilton, Ontario Canada L8S 4L7 April 19, 2004 Abstract
Sorting suffixes of two-pattern strings Frantisek Franek W. F. Smyth Algorithms Research Group Department of Computing & Software McMaster University Hamilton, Ontario Canada L8S 4L7 April 19, 2004 Abstract
COS597D: Information Theory in Computer Science October 19, Lecture 10
 COS597D: Information Theory in Computer Science October 9, 20 Lecture 0 Lecturer: Mark Braverman Scribe: Andrej Risteski Kolmogorov Complexity In the previous lectures, we became acquainted with the concept
COS597D: Information Theory in Computer Science October 9, 20 Lecture 0 Lecturer: Mark Braverman Scribe: Andrej Risteski Kolmogorov Complexity In the previous lectures, we became acquainted with the concept
Efficient High-Similarity String Comparison: The Waterfall Algorithm
 Efficient High-Similarity String Comparison: The Waterfall Algorithm Alexander Tiskin Department of Computer Science University of Warwick http://go.warwick.ac.uk/alextiskin Alexander Tiskin (Warwick)
Efficient High-Similarity String Comparison: The Waterfall Algorithm Alexander Tiskin Department of Computer Science University of Warwick http://go.warwick.ac.uk/alextiskin Alexander Tiskin (Warwick)
Final exam study sheet for CS3719 Turing machines and decidability.
 Final exam study sheet for CS3719 Turing machines and decidability. A Turing machine is a finite automaton with an infinite memory (tape). Formally, a Turing machine is a 6-tuple M = (Q, Σ, Γ, δ, q 0,
Final exam study sheet for CS3719 Turing machines and decidability. A Turing machine is a finite automaton with an infinite memory (tape). Formally, a Turing machine is a 6-tuple M = (Q, Σ, Γ, δ, q 0,
Efficient Approximation of Large LCS in Strings Over Not Small Alphabet
 Efficient Approximation of Large LCS in Strings Over Not Small Alphabet Gad M. Landau 1, Avivit Levy 2,3, and Ilan Newman 1 1 Department of Computer Science, Haifa University, Haifa 31905, Israel. E-mail:
Efficient Approximation of Large LCS in Strings Over Not Small Alphabet Gad M. Landau 1, Avivit Levy 2,3, and Ilan Newman 1 1 Department of Computer Science, Haifa University, Haifa 31905, Israel. E-mail:
A New Dominant Point-Based Parallel Algorithm for Multiple Longest Common Subsequence Problem
 A New Dominant Point-Based Parallel Algorithm for Multiple Longest Common Subsequence Problem Dmitry Korkin This work introduces a new parallel algorithm for computing a multiple longest common subsequence
A New Dominant Point-Based Parallel Algorithm for Multiple Longest Common Subsequence Problem Dmitry Korkin This work introduces a new parallel algorithm for computing a multiple longest common subsequence
