LINEAR MIXED-EFFECT MODELS
|
|
|
- Meryl Fields
- 5 years ago
- Views:
Transcription
1 LINEAR MIXED-EFFECT MODELS Studies / data / models seen previously in 511 assumed a single source of error variation y = Xβ + ɛ. β are fixed constants (in the frequentist approach to inference) ɛ is the only random effect What if there are multiple sources of error variation? c Dept. Statistics () Stat part 3 Spring / 116
2 Examples: Subsampling Seedling weight in 2 genotype study from Aitken model section. Seedling weight measured on each seedling. Two (potential) sources of variation: among flats and among seedlings within a flat. Y ijk = µ + γ i + T ij + ɛ ijk T ij N(0, σ 2 F ) ɛ ijk N(0, σ 2 e) where i indexes genotype, j indexes flat within genotype, and k indexes seedling within flat σf 2 quantifies variation among flats, if have perfect knowledge of the seedlings in them σ 2 e quantifies variation among seedlings c Dept. Statistics () Stat part 3 Spring / 116
3 Examples: Split plot experimental design Influence of two factors: temperature and a catalyst on fermentation of dry distillers grain (byproduct of EtOH production from corn) Response is CO 2 production, measured in a tube Catalyst (+/-) randomly assigned to tubes Temperature randomly assigned to growth chambers 6 tubes per growth chamber, 3 +catalyst, 3 -catalyst all 6 at same temperature Use 6 growth chambers, 2 at each temperature The o.u. is a tube. What is the experimental unit? c Dept. Statistics () Stat part 3 Spring / 116
4 Examples: Split plot experimental design - 2 Two sizes of experimental unit: tube and temperature Y ijkl = µ + α i + γ ik + β j + αβ ij + ɛ ijkl γ ik N(0, σg 2 ) Var among growth chambers ɛ ijkl N(0, σ 2 ) Var among tubes where i indexes temperature, j indexes g.c. within temp., k indexes catalyst, and l indexes tube within temp., g.c., and catalyst σg 2 quantifies variation among g.c., if have perfect knowledge of the tubes in them σ 2 quantifies variation among tubes c Dept. Statistics () Stat part 3 Spring / 116
5 Examples: Gauge R&R study Want to quantify repeatability and reproducibility of a measurement process Repeatability: variation in meas. taken by a single person or instrument on the same item. Reproducibility: variation in meas. made by different people (or different labs), if perfect repeatability One possible design: 10 parts, each measured twice by 10 operators. 200 obs. Y ijk = µ + α i + β j + αβ ij + ɛ ijk α i N(0, σp 2 ) Var among parts β j N(0, σr 2 ) Reproducibility variance αβ ij N(0, σi 2 ) Interaction variance ɛ ijk N(0, σ 2 ) Repeatability variance c Dept. Statistics () Stat part 3 Spring / 116
6 Linear Mixed Effects model y = Xβ + Z u + ɛ X n p matrix of known constants β IR p an unknown parameter vector Z n q matrix of known constants u q 1 random vector ɛ n 1 vector of random errors The elements of β are considered to be non-random and are called fixed effects. The elements of u are called random effects The errors are always a random effect c Dept. Statistics () Stat part 3 Spring / 116
7 Review of crossing and nesting Seedling weight study: seedling effects nested in flats DDG study: temperature effects crossed with catalyst effects tube effects nested within growth chamber effects Gauge R&R study: part effects crossed with operator effects measurement effects nested within part operator fixed effects are usually crossed, rarely nested random effects are commonly nested, sometimes crossed c Dept. Statistics () Stat part 3 Spring / 116
8 Because the model includes both fixed and random effects (in addition to the residual error), it is called a mixed-effects model or, more simply, a mixed model. The model is called a linear mixed-effects model because (as we will soon see) E(y u) = Xβ + Z u, a linear function of fixed and random effects. The usual LME assumes that E(ɛ) = 0 E(u) = 0 Cov(ɛ, u) = 0. Var(ɛ) = R Var(u) = G It follows that: Ey = Xβ Vary = Z GZ + R c Dept. Statistics () Stat part 3 Spring / 116
9 The details: Ey = E(Xβ + Z u + ɛ) = Xβ + Z E(u) + E(ɛ) = Xβ and Vary = Var(Xβ + Z u + ɛ) = Var(Z u + ɛ) = Var(Z u) + Var(ɛ) = Z Var(u)Z + R = Z GZ + R Σ The conditional moments, given the random effects, are: Ey u = Xβ + Z u and Vary u = R c Dept. Statistics () Stat part 3 Spring / 116
10 We usually consider the special case in which [ ] ([ ] [ ]) u 0 G 0 N, ɛ 0 0 R = y N(Xβ, Z GZ + R). Example: Recall the seedling metabolite study. Previously, we looked at metabolite concentration. Each individual seedling was weighed Let y ijk denote the weight of the k th seedling from the j th flat of genotype i (i = 1, 2, j = 1, 2, 3, 4, k = 1,..., n ij ; where n ij = # of seedlings for genotype i flat j.) c Dept. Statistics () Stat part 3 Spring / 116
11 Seedling Weight Tray Genotype A Genotype B c Dept. Statistics () Stat part 3 Spring / 116
12 Consider the model y ijk = µ + γ i + F ij + ɛ ijk F ij s i.i.d. N(0, σ 2 F ) independent of ɛ ijk s i.i.d. N(0, σ 2 e) This model can be written in the form y = Xβ + Z u + ɛ c Dept. Statistics () Stat part 3 Spring / 116
13 y = y 111 y 112 y 113 y 114 y 115 y 121 y y 247 y 248 β = µ γ 1 γ 2 µ = F 11 F 12 F 13 F 14 F 21 F 22 F 23 F 24 ɛ = e 111 e 112 e 113 e 114 e 115 e 121 e e 247 e 248 c Dept. Statistics () Stat part 3 Spring / 116
14 X = Z = c Dept. Statistics () Stat part 3 Spring / 116
15 What is Vary? G = Var(µ) = Var([F 11,..., F 24 ] ) = σ 2 F I 8 8 R = Var(ɛ) = σ 2 e I Var(y) = Z GZ + R = Z σf 2 IZ + σe 2 I = σfz 2 Z + σei 2 1 n11 1 n n12 1 n Z Z n13 1 n 13 = n24 1 n 24 This is a block diagonal matrix with blocks of size n ij n ij, i = 1, 2, j = 1, 2, 3, 4. c Dept. Statistics () Stat part 3 Spring / 116
16 Thus, Var(y) = σf 2Z Z + σei 2 is also block diagonal. The first block is y 111 σf 2 + σ2 e σf 2 σf 2 σf 2 σf 2 y 112 Var y 113 y 114 = σf 2 σf 2 + σ2 e σf 2 σf 2 σf 2 σf 2 σf 2 σf 2 + σ2 e σf 2 σf 2 σf 2 σf 2 σf 2 σf 2 + σ2 e σf 2 y 115 σf 2 σf 2 σf 2 σf 2 σf 2 + σ2 e Other blocks are the same except that the dimensions are (# of seedlings per tray ) 2. A Σ matrix with this form is called Compound Symmetric c Dept. Statistics () Stat part 3 Spring / 116
17 Two different ways to write the same model 1 As a mixed model: i.i.d. Y ijk = µ + γ i + T ij + ɛ ijk, T ij N(0, σf 2 ), 2 As a fixed effects model with correlated errors: Y ijk = µ + γ i + ɛ ijk, ɛ N(0, Σ) In either model: ɛ ijk i.i.d. N(0, σ 2 e) Var(y ijk ) = σf 2 + σ2 e i, j, k. Cov(y ijk, y ijk ) = σf 2 i, j, and k k. Cov(y ijk, y i j k ) = 0 if i i or j j. Any two observations from the same flat have covariance σf 2. Any two observations from different flats are uncorrelated. c Dept. Statistics () Stat part 3 Spring / 116
18 What about the variance of the flat average: Varȳ ij. = Var( 1 n ij 1 (y ij,..., y ijnij ) ) = 1 n 2 ij 1 (σ 2 F11 + σ 2 ei) 1 = 1 n 2 ij (σ 2 F σ 2 e1 I1) = 1 n 2 ij (σ 2 F n ijn ij + σ 2 en ij ) = σ 2 F + σ2 e n ij i, j. If we analyzed seedling dry weight as we did average metabolite concentration, that analysis assumes σ 2 F = 0. Because that analysis models Var(ȳ ij ) as σ2 n ij i, j. c Dept. Statistics () Stat part 3 Spring / 116
19 The variance of the genotype average, using f i flats per genotype: Varȳ i.. = 1 Varȳij. fi 2 = σ2 F + 1 σ 2 f i fi 2 e /n ij i When all flats have same number of seedlings per flat, i.e. n ij = n i, j Varȳ i.. = σ2 F f i + σ2 e f i n i Var among flats divided by # flats + Var among sdl. divided by total # seedlings When balanced, mixed model analysis same as computing flat averages, then using OLS model on ȳ ij. except that mixed model analysis provides estimate of σ 2 F c Dept. Statistics () Stat part 3 Spring / 116
20 Note that Var(y) may be written as σ 2 ev where V is a block diagonal matrix with blocks of the form Thus, if σ2 F σ 2 e 1 + σ2 F σ σ 2 F 2 σ e σ 2 F e σe 2 σf σ2 σ 2 F σ e σ 2 F e σe σf 2 σ 2 σe 2 F σ2 σe 2 F σe 2 were known, we would have the Aitken Model. y = Xβ + Z u + ɛ = Xβ + ɛ, ɛ N(0, σ 2 V ), σ 2 σ 2 e We would use GLS to estimate any estimable Cβ by C ˆβ v = C(X V 1 X) X V 1 y c Dept. Statistics () Stat part 3 Spring / 116
21 However, we seldom know σ2 F or, more generally, Σ or V. σe 2 Thus, our strategy usually involves estimating the unknown parameters in Σ to obtain ˆΣ. Then inference proceeds based on C ˆβ ˆV = C(X ˆV 1 X) X ˆV 1 y or C ˆβˆ = C(X ˆΣ 1 X) X ˆΣ 1 y. Remaining Questions: 1 How do we estimate the unknown parameters in V (or Σ)? 2 What is the distribution of C ˆβˆv = C(X ˆV 1 X) X ˆV 1 y? 3 How should we conduct inference regarding Cβ? 4 Can we test Ho: σ 2 F = 0? 5 Can we use the data to help us choose a model for Σ? c Dept. Statistics () Stat part 3 Spring / 116
22 R code for LME s d <- read.table( SeedlingDryWeight2.txt,as.is=T, header=t) names(d) with(d, table(genotype, Tray)) with(d, table(tray,seedling)) # for demonstration, construct a balanced data set # 5 seedlings for each tray d2 <- d[d$seedling <=5,] # create factors d2$g <- as.factor(d2$genotype) d2$t <- as.factor(d2$tray) d2$s <- as.factor(d2$seedling) c Dept. Statistics () Stat part 3 Spring / 116
23 # fit the ANOVA temp <- lm(seedlingweight g+t, data=d2) # get the ANOVA table anova(temp) # note that all F tests use MSE = seedlings # within tray as denominator # will need to hand-calculate test for genotypes # better to fit LME. 2 functions: # lmer() in lme4 or lme() in nlme # for most models lmer() is easier to use library(lme4) c Dept. Statistics () Stat part 3 Spring / 116
24 temp <- lmer(seedlingweight g+(1 t),data=d2) summary(temp) anova(temp) # various extractor functions: fixef(temp) # estimates of fixed effects vcov(temp) # VC matrix for the fixed effects VarCorr(temp) # estimated variance(-covariance) for r ranef(temp) # predictions of random effects coef(temp) # fixed effects + pred s of random effe fitted(temp) resid(temp) # conditional means for each obs (X bha # conditional residuals (Y - fitted) c Dept. Statistics () Stat part 3 Spring / 116
25 # REML is the default method for estimating # variance components. # if want to use ML, can specify that lmer(seedlingweight g+(1 t),reml=f, data=d2) # but how do we get p-values for tests # or decide what df to use to construct a ci? # we ll talk about inference in R soon c Dept. Statistics () Stat part 3 Spring / 116
26 Experimental Designs and LME s LME models provide one way to model correlations among observations Very useful for experimental designs where there is more than one size of experimental unit Or designs where the observation unit is not the same as the experimental unit. My philosophy (widespread at ISU and elsewhere) is that the way an experiment is conducted specifies the random effect structure for the analysis. Observational studies are very different No randomization scheme, so no clearcut random effect structure Often use model selection methods to help chose the random effect structure NOT SO for a randomized experiment. c Dept. Statistics () Stat part 3 Spring / 116
27 One example: study designed as an RCBD. treatments are randomly assigned within a block analyze data, find out block effects are small should you delete block from the model and reanalyze? above philosophy says tough. Block effects stay in the analysis for the next study, seriously consider: blocking in a different way or using a CRD Following pages work through details of LME s for some common study designs c Dept. Statistics () Stat part 3 Spring / 116
28 Mouse muscle study Mice grouped into blocks by litter. 4 mice per block, 4 blocks. Four treatments randomly assigned within each block Collect two replicate muscle samples from each mouse Measure response on each muscle sample 16 eu., 32 ou. Questions concern differences in mean response among the four treatments The model usually used for a study like this is: y ijk = µ + τ i + l j + m ij + e ijk, where y ijk is the k th measurement of the response for the mouse from litter j that received treatment i, (i = 1, 2, 3, 4; j = 1, 2, 3, 4; k = 1, 2). c Dept. Statistics () Stat part 3 Spring / 116
29 The fixed effects: µ τ 1 β = τ 2 τ 3 IR5 is an unknown vector of fixed parameters τ 4 u = [l 1, l 2, l 3, l 4, m 11, m 21, m 31, m 41, m 12,..., m 34, m 44 ] is a vector of random effects describing litters and mice. ɛ = [e 111, e 112, e 211, e 212,..., e 441, e 442,..., e 441, e 442 ] is a vector of random errors. with y = [y 111, y 112, y 211, y 212,..., y 411, y 412,..., y 441, y 442 ], the model can be written as a linear mixed effects model y = Xβ + Z u + ɛ, where c Dept. Statistics () Stat part 3 Spring / 116
30 X = Z = c Dept. Statistics () Stat part 3 Spring / 116
31 We can write less and be more precise using Kronecker product notation. X = 1 }{{} 4 1 [}{{} 1, }{{} I }{{} 1 ] Z = [}{{} I }{{} 1, I }{{} }{{} 1 ] 2 1 In this experiment, we have two random factors: Litter and Mouse. We can partition our random effects vector u into a vector of litter effects and a vector of mouse effects: u = [ l m ], l = l 1 l 2 l 3 l 4, m = m 11 m 21 m 31 m 41 m m 44 c Dept. Statistics () Stat part 3 Spring / 116
32 We make the usual assumption that [ ] ([ l 0 u = N m 0 ], [ σ 2 l I 0 0 σ 2 mi Where σl 2, σm 2 IR + are unknown parameters. We can partition: Z = [ I }{{} 4 4 }{{} 1, I 8 1 }{{} ]) }{{} 1 ] = [Z l, Z m ] 2 1 We have Z u = [ Z l, Z m ] [ l m ] = Z l l + Z m m and c Dept. Statistics () Stat part 3 Spring / 116
33 Var(Z u) = Z GZ = [ Z l, Z m ] [ σ 2 l I 0 0 σ 2 mi ] [ Z l Z m ] = σ 2 l I }{{} 4 4 = Z l (σ 2 l I)Z l + Z m (σ 2 mi)z m = σ 2 l Z l Z l + σ 2 mz m Z m 11 }{{} 8 8 +σ 2 m I }{{} }{{} 2 2 We usually assume that all random effects and random errors are mutually independent and that the errors (like the effects within each factor) are identically distributed: l m ɛ N 0 0 0, σl 2 I σmi σei 2 c Dept. Statistics () Stat part 3 Spring / 116
34 The unknown variance parameters σ 2 l, σ 2 m, σ 2 e IR + are called variance components. In this case, we have R = Var(ɛ) = σ 2 ei. Thus, Var(y) = Z GZ + R = σ 2 l Z lz l + σ 2 mz m Z m + σ 2 ei. This is a block diagonal matrix. Each litter is one of the following blocks. a + b + c a + b a a a a a a a + b a + b + c a a a a a a a a a + b + c a + b a a a a a a a + b a + b + c a a a a a a a a a + b + c a + b a a a a a a a + b a + b + c a a a a a a a a a + b + c a + b a a a a a a a + b a + b + c (To save space, a = σ 2 l, b = σ 2 m, c = σ 2 e) c Dept. Statistics () Stat part 3 Spring / 116
35 The random effects specify the correlation structure of the observations This is determined (in my philosophy) by the study design Changing the random effects changes the implied study design Delete the mouse random effects: The study is now an RCBD with 8 mice (one per obs.) per block Also delete the litter random effects: The study is now a CRD with 8 mice per treatment c Dept. Statistics () Stat part 3 Spring / 116
36 Should blocks be fixed or random? Some views: The design is called an RCBD, so of course blocks are random. But, R in the name is Randomized (describing treatment assignment). Introductory ANOVA: blocks are fixed, because that s all we know about. Some things that influence my answer If blocks are complete and balanced (all blocks have equal # s of all treatments), inferences about treatment differences (or contrasts) are same either way. If blocks are incomplete, an analysis with fixed blocks evaluates treatment effects within each block, then averages effects over blocks. An analysis with random blocks, recovers the additional information about treatment differences provided by the block means. Will talk about this later, if time. c Dept. Statistics () Stat part 3 Spring / 116
37 However, the biggest and most important difference concerns the relationship between the se s of treatment means and the se s of treatment differences (or contrasts). Simpler version of mouse study: no subsampling, 4 blocks, 4 mice/block, 4 treatments Y ij = µ + τ i + l j + ɛ ij, ɛ ij N(0, σ 2 e) If blocks are random, l j N(0, σ 2 l ) Blocks are: Fixed Random Var ˆτ 1 ˆτ 2 2 σe/4 2 2 σe/4 2 Var ˆµ + ˆτ 1 σe/4 2 (σl 2 + σe)/4 2 When blocks are random, Var trt mean includes the block-block variability. Matches implied inference space: repeating study on new litters and new mice c Dept. Statistics () Stat part 3 Spring / 116
38 Common to add ± se bars to a plot of means If blocking effective, σ 2 l is large, so you get: Random Blocks Fixed Blocks Mean Mean Treatment Treatment The random blocks picture does not support claims about trt. diff. c Dept. Statistics () Stat part 3 Spring / 116
39 So should blocks be Fixed or Random? My approach: When the goal is to summarize difference among treatments, use fixed blocks When the goal is to predict means for new blocks, use random blocks If blocks are incomplete, think carefully about goals of study Many (most) have different views. c Dept. Statistics () Stat part 3 Spring / 116
40 Split plot experimental design Field experiment comparing fertilizer response of 3 corn genotypes Large plots planted with one genotype (A, B, C) Each large plot subdivided into 4 small plots. Small plots fertilized with 0, 50, 100, or 150 lbs Nitrogen / acre. Large plots grouped into blocks; Genotype randomly assigned to large plot, Small plots randomly assigned to N level within each large plot. Key feature: Two sizes of experimental units c Dept. Statistics () Stat part 3 Spring / 116
41 Field Plot Block 1 Block 2 Block 3 Block 4 Genotype C Genotype B Genotype A Genotype B Genotype C Genotype B Genotype A Genotype A Genotype C Genotype B Genotype C Genotype A Split Plot or Sub Plot c Dept. Statistics () Stat part 3 Spring / 116
42 Names: The large plots are called main plots or whole plots Genotype is the main plot factor The small plots are split plots Fertilizer is the split plot factor Two sizes of eu s Main plot is the eu. for genotype Split plot is the eu. for fertilizer Split plots are nested in Main plots Many different variations, this is the most common RCB for main plots; CRD for split plots within main plots Can extend to three or more sizes of eu. split-split plot or split-split-split plot designs c Dept. Statistics () Stat part 3 Spring / 116
43 Split plot studies commonly mis-analyzed as RCBD with one eu. Treat Genotype Fertilizer combination (12 treatments) as if they were randomly assigned to each small plot Field Block 1 B 100 B 0 A 0 C 100 B 150 C 50 A 50 A C B 50 C 0 A 100 Block 2 A 150 A 0 C 50 A 50 B 100 B 50 C 100 C 0 A C B B 0 Block 3 C 0 A 0 A 100 B 100 B 50 B 0 A 150 C 50 A 50 C C B Block 4 B 0 C 150 B 50 A C A 0 B 150 C 50 B 100 C 0 A 100 A 50 c Dept. Statistics () Stat part 3 Spring / 116
44 And if you ignore blocks, you have a very different set of randomizations Field B 50 B 0 A B A C A 50 B 0 A 50 C 100 C 0 C 100 A 50 A 0 C 50 B 50 B 150 B 50 A 0 C 0 A 100 C 50 B 150 B 0 C 0 A 0 A A A 0 B 0 A 150 B 150 A 50 B C A 100 B 50 B B C C 100 C 50 A 150 C 50 C 150 C 0 C B c Dept. Statistics () Stat part 3 Spring / 116
45 Confusion is (somewhat understandable) Same treatment structure (2 way complete factorial) Different experimental design because different way of randomizing treatments to eu.s So different model than the usual RCBD Use random effects to account for the two sizes of eu. Rarely done on purpose. Usually forced by treatment constraints Cleaning out planters is time consuming, so easier to plant large areas Growth chambers can only be set to one temperature, have room for many flats In the engineering literature, split plot designs are often called hard-to-change factor designs c Dept. Statistics () Stat part 3 Spring / 116
46 Diet and Drug effects on mice Split plot designs may not involve a traditional plot Study of diet and drug effect on mice Have 2 mice from each of 8 litters; cages hold 2 mice each. Put the two mice from a litter into a cage. Randomly assign diet (1 or 2) to the cage/litter Randomly assign drug (A or B) to a mouse within a cage Main plots = cage/litter, in a CRD Diet is the main plot treatment factor Split plots = mouse, in a CRD within cage Drug is the split plot treatment factor Let s construct a model for this mouse study, 16 obs. c Dept. Statistics () Stat part 3 Spring / 116
47 y ijk = µ + α i + β j + γ ij + l ik + e ijk (i = 1, 2; j = 1, 2; k = 1,..., 4) y 111 y 121 y 112 e 111 y 122 y µ e y 123 α 1 l 11. y 114 α 2 l 12. y y = 124 β 1 l 13. β = y 211 β 2 µ = l 14. y 221 γ 11 l 21 ɛ =. y 212 γ l γ l 23. y l y 213 γ y 223 e 224 y 214 y c Dept. Statistics () 224 Stat part 3 Spring / 116
48 X = [ 1 }{{} 16 1, I }{{} 2 2 [ u ɛ }{{} 1, }{{} Z = I }{{} 8 8 }{{} I }{{} 2 1, I y = Xβ + Z u + ɛ ] ([ ] 0 N, 0 Var(Z u) = Z GZ = σ 2 l Z Z = σ 2 l [ I }{{} 8 8 = σ 2 l I }{{} }{{} 2 1 }{{} 2 2 [ σ 2 l I 0 0 σ 2 ei 1 }{{} 4 1 ]) }{{} I ] 2 2 }{{} 1 ][}{{} I }{{} ] [ σ = Block Diagonal with blocks 2 σ2 l σ 2 l l σl 2 ] c Dept. Statistics () Stat part 3 Spring / 116
49 Var(y) = σ 2 l Var(ɛ) = R = σ 2 ei I }{{} 8 8 }{{} 11 +σei [ σ 2 = Block Diagonal with blocks l + σe 2 σl 2 σl 2 σl 2 + σe 2 ] c Dept. Statistics () Stat part 3 Spring / 116
50 Thus, the covariance between two observations from the same litter is σ 2 l and the correlation is σl 2. σl 2 +σe 2 This is also easy to compute from the non-matrix expressinon of the model. i, j Var(y ijk ) = Var(µ+α i +β j +γ ij +l ik +e ijk ) = Var(l ik +e ijk ) = σ 2 l +σ 2 e Cov(y i1k, y i2k ) = Cov(µ + α i + β 1 + γ i1 + l ik + e i1k, = Cov(l ik + e i1k, l ik + e i2k ) µ + α i + β 2 + γ i2 + l ik + e i2k ) = Cov(l ik, l ik ) + Cov(l ik + e i2k ) + Cov(e i1k, l ik ) + Cov(e i1k + e i2k ) = Cov(l ik + l ik = Var(l ik ) = σ 2 l Cor(y i1k, y i2k ) = Cov(y i1k, y i2k ) Var(yi1k )Var(y i2k ) = σ2 l σ 2 l + σ 2 e c Dept. Statistics () Stat part 3 Spring / 116
51 THE ANOVA APPROACH TO THE ANALYSIS OF LINEAR MIXED EFFECTS MODELS A model for expt. data with subsampling y ijk = µ + τ i + u ij + e ijk, (i = 1,..., t; j = 1,..., n; k = 1,..., m) β = (µ, τ i,..., τ t ), u = (u 11, u 12,..., u tn ), ɛ = (e 111, e 112,..., e tnm ), β IR t+1, an unknown parameter vector, [ ] ([ ] [ ]) u 0 σ 2 N, u I 0 ɛ 0 0 σei 2 σ 2 u, σ 2 e IR +, unknown variance components c Dept. Statistics () Stat part 3 Spring / 116
52 This is the commonly-used model for a CRD with t treatments, n experimental units per treatment, and m observations per experimental unit. We can write the model as y = Xβ + Z u + ɛ, where X = [ 1 }{{} tnm 1, I }{{} t t 1 }{{} nm 1 ] and Z = [ I }{{} tn tn }{{} 1 ] m 1 Consider the sequence of three column spaces: X 1 = 1 tnm 1, X 2 = [1 tnm 1, I t t 1 nm 1 ], X 3 = [1 tnm 1, I t t 1 nm 1, I tn tn 1 m 1 ] These correspond to the models: X 1 : X 2 : X 3 : Y ijk = µ + ɛ ijk Y ijk = µ + τ i + ɛ ijk Y ijk = µ + τ i + u ij + ɛ ijk c Dept. Statistics () Stat part 3 Spring / 116
53 ANOVA table for subsampling Source SS df df treatments y (P 2 P 1 )y rank(x 2 ) rank(x 1 ) t 1 eu(treatments) y (P 3 P 2 )y rank(x 3 ) rank(x 2 ) t(n 1) ou(eu, treatments) y (I P 3 )y tnm rank(x 3 ) tn(m 1) C.total tnm 1 In terms of squared differences: Source df Sum of Squares trt t 1 t n m i=1 j=1 k=1 (y i.. y...)2 eu(trt) t(n 1) t n m i=1 j=1 k=1 (y ij. y i..)2 ou(eu, trt) tn(m 1) t n m i=1 j=1 k=1 (y ijk y ij.)2 C.total tnm 1 t n m i=1 j=1 k=1 (y ijk y...) 2 c Dept. Statistics () Stat part 3 Spring / 116
54 Which simplifies to: Source df Sum of Squares trt t 1 nm t i=1 (y i.. y...)2 eu(trt) tn t m t n i=1 j=1 (y ij. y i..)2 ou(eu, trt) tnm tn t n m i=1 j=1 k=1 (y ijk y ij...)2 C.total tnm 1 n m j=1 k=1 (y ijk y...) 2 Each line has a MS = SS / df t i=1 MS s are random variables. Their expectations are informative. c Dept. Statistics () Stat part 3 Spring / 116
55 Expected Mean Squares, EMS E(MS trt ) = nm t 1 = nm t 1 = nm t 1 = nm t i=1 E(y i.. y... )2 t i=1 E(µ + τ i + u i. + e i.. µ τ. u.. e...) 2 t i=1 E(τ i τ. + u i u.. + e i.. e...) 2 t t 1 i=1 [(τ i τ.) 2 + E(u i u..) 2 + E(e i.. e...) 2 ] = nm t 1 [ t i=1 (τ i τ) 2 + E( t i=1 (u i u..) 2 ) +E( t i=1 e i. e) 2 )] u 1,...,u t. i.i.d. N(0, σ2 u n ). Thus, E( t i=1 (u i u..) 2 ) = (t 1) σ2 u n e 1...,...,e t.. i.i.d. N(0, σ2 u nm ). Thus, E( t i=1 (e i e..) 2 ) = (t 1) σ2 e mn It follows that E(MS trt )= nm t 1 t i=1 (τ i τ) 2 + mσ 2 u + σ 2 e c Dept. Statistics () Stat part 3 Spring / 116
56 Similar calculations allow us to add Expected Mean Squares (EMS) to our Anova table. Source trt eu(trt) ou(eu, trt) EMS σe 2 + mσu 2 + nm t t 1 i=1 (τ i τ) 2 σe 2 + mσu 2 Mean Squares could also be computed using E(y Ay) = tr(aɛ) + [E(y)] AE(y), σ 2 e Where = Var(y)=ZGZ + R = σ 2 u I }{{} tn tn E(y) = [µ + τ 1, µ + τ 2,..., µ + τ t ] }{{} 11 +σe 2 }{{} I and m m tnm tnm I }{{} nm 1 c Dept. Statistics () Stat part 3 Spring / 116
57 Futhermore, it can be shown that y (P 2 P 1 ) (σ 2 e+mσ 2 u )y χ2 t 1 y (P 3 P 2 ) (σe+mσ 2 u 2)y χ2 tn t y χ 2 tnm tn y (I P 3 ) σ 2 e ( nm [ ] t σe+mσ 2 u 2 i=1 (τ i τ.) 2 /(t 1) and these three χ 2 random variables are independent. It follows that F 1 = MStrt MS eu(trt) F [ nm t σe 2+mσ2 i=1 (τ i τ.) 2 /(t 1)] u t 1, tn t use F 1 to test H 0 : τ 1 =... = τ t F 2 = MS eu(trt) MS ou(eu,trt) ( σ2 e +mσ2 u )F σe 2 tn t, tnm tn use F 2 to test H 0 : σu 2 = 0 ) c Dept. Statistics () Stat part 3 Spring / 116
58 Notice an important difference between the distributions of the F statistics testing Ho: τ 1 = τ 2 =... = τ T and testing Ho: σ 2 u = 0: The F statistic for a test of fixed effects, e.g. F 1, has a non-central F distribution under Ha The F statistic for a test of a variance component, e.g. F 2, has a scaled central F distribution under Ha Both have same central F distribution under Ho If change a factor from fixed to random : critical value for α-level test is same, but power/sample size calculations are not the same! c Dept. Statistics () Stat part 3 Spring / 116
59 Shortcut to identify Ho The EMS tell you what Ho associated with any F test Ho for F = MS A /MS B is whatever is necessary for EMS A /EMS B to equal 1. F statistic EMS ratio Ho σe MS trt /MS 2+mσ2 u + nm t t 1 i=1 (τ i τ) 2 t eu(trt) i=1 (τ i τ) 2 = 0 σ 2 e+mσ 2 u MS eu(trt) /MS ou(eu,trt) MS trt /MS ou(eu,trt) σ 2 e+mσ 2 u σ 2 e σe+mσ 2 u+ 2 nm t t 1 i=1 (τ i τ) 2 σe 2 σ 2 u = 0 σ 2 u = 0, and t i=1 (τ i τ) 2 = 0 c Dept. Statistics () Stat part 3 Spring / 116
60 Estimating estimable functions of β Consider the GLS estimator ˆβ Σ = (X Σ 1 X) X Σ 1 y When m, number of subsamples per eu. constant, Var(y) = σ 2 u I }{{} tn tn }{{} 11 +σe 2 }{{} I m m tnm tnm Σ (1) This is a very special Σ. It turns out that ˆβ Σ = (X Σ 1 X) X Σ 1 y = (X X) X y = ˆβ Thus, when Σ has the form of (1), the OLS estimator of any estimable Cβ is the GLS Estimates of Cβ do not depend on σ 2 u or σ 2 e Estimates of Cβ are BLUE c Dept. Statistics () Stat part 3 Spring / 116
61 ANOVA estimate of a variance component Given data, how can you estimate σ 2 u? E( MS eu(trt) MS ou(eu,trt) m ) = (σ2 e +mσ2 u ) σ2 e m = σu 2 Thus, MS eu(trt) MS ou(eu,trt) m is an unbiased estimator of σu 2 Method of Moments estimator because it is obtained by equating observed statistics with their moments (expected values in this case) and solving the resulting set of equations for unknown parameters in terms of observed statistics. Notice ˆσ u 2 is smaller than Var ȳ ij. = sample var among e.u. s. ȳ ij. includes variability among eu s, σu, 2 and variability among observations, σe. 2 Hence my if perfect knowledge (σe 2 = 0) qualifications when I introduced variance components. Although MS eu(trt) MS ou(eu,trt) m is an unbiased estimator of σu, 2 it can take negative values. σu, 2 the population variance of the u random effects, cannot be negative! c Dept. Statistics () Stat part 3 Spring / 116
62 Why ˆσ 2 u might be < 0 Sampling variation in MS eu(trt) and MS ou(eu,trt) especially if σ 2 u = 0 or 0. Incorrect estimate of σ 2 e One outlier can inflate MS ou(eu,trt) Consequence is ˆσ 2 u might be < 0 Model not correct Var Y ijk u ij may be incorrectly specified e.g. R = σ 2 ei, but Var Y ijk u ij not constant or obs not independent My advice: don t blindly force ˆσ 2 u to be 0. Check data, think carefully about model. c Dept. Statistics () Stat part 3 Spring / 116
63 Two ways to analyze data with subsampling Mixed model with σ 2 u and σ 2 e y ijk = µ + τ i + u ij + e ijk, (i = 1,..., t; j = 1,..., n; k = 1,..., m) Analyses of e.u. averages The average of observations for experimental unit ij is y ij. = µ + τ i + u ij + e ij. y ij = µ + τ i + ɛ ij, where ɛ ij = u ij + e ij. i, j. N(0, σ 2 ) where σ 2 = σu 2 + σ2 e m This is a ngm model for the averages { y ij. : i = 1,..., t; j = 1,..., n } When m ij = m for all e.u. s, inferences about estimable functions of β are exactly the same ɛ ij i.i.d. ˆσ 2 is an estimate of σ 2 u + σ2 e m. We can t separately estimate σ 2 u and σ 2 e, but not an issue if focus is on inference for estimable functions of β. c Dept. Statistics () Stat part 3 Spring / 116
64 Support for these claims What can we estimate? For both models, E(y) = [µ + τ 1, µ + τ 2,..., µ + τ t ] I }{{} nm 1 The only estimable quantities are linear combinations of the treatment means µ + τ 1, µ + τ 2,..., µ + τ t Best Linear Unbiased Estimators are y 1.., y 2..,.., y t..,, respectively. c Dept. Statistics () Stat part 3 Spring / 116
65 What about Var y ij.? Two ways to compute: Var(y i..) = Var(µ + τ i + u i. + e i..) = Var(u i. + e i..) = Var(u i.) + Var(e i..) = σ2 u n + σ2 e nm = 1 n (σ2 u + σ2 e m ) = σ2 n c Dept. Statistics () Stat part 3 Spring / 116
66 Or using matrix representation, Var(y i..) = Var( 1 n n y ij.) j=1 = 1 n Var(y i1. ) = 1 n Var( 1 m }{{} 1 (y i11,..., y i1m ) ) m 1 = 1 1 n m 2 }{{} 1 (σei 2 + σu11 2 )1 = 1 n m 1 1 m 2 (σ2 em + σ 2 um 2 ) = σ2 e + mσ 2 u nm = σ2 n c Dept. Statistics () Stat part 3 Spring / 116
67 Thus, Var y 1.. y y t.. Var C y y t.. = σ2 n I }{{} t t and = σ2 n CC. c Dept. Statistics () Stat part 3 Spring / 116
68 Notice Var Cβ depends only on σ 2 = σu 2 + σe/m 2 Don t need separate estimates of σu 2 and σe 2 to carry out inference for estimable Cβ. Do need to estimate σ 2 = σu 2 + σ2 e m. Mixed model: ˆσ 2 = MS eu(trt) m Analysis of averages: ˆσ 2 = MSE For example: Thus, Var(y 1.. y 2.. ) = Var(y 1.. ) + Var(y 2.. ) = 2 σ2 n = 2(σ2 u n + σ2 e mn ) = 2 mn (σ2 e + mσ 2 u) = 2 mn E(MS eu(trt)) Var(y 1.. y 2.. ) = 2MS eu(trt) mn c Dept. Statistics () Stat part 3 Spring / 116
69 error d.f. are the same in both analyses: Mixed model: MS eu(trt) has t(n 1) df Analysis of averages: MSE has t(n 1) df A 100(1 α)% Confidence interval for τ 1 τ 2 is y 1.. y 2.. ± t α 2 2MSeu(trt) t(n 1) mn. A test of H 0 : τ 1 = τ 2 can be based on t = y 1.. y 2.. t 2MS t(n 1) τ 1 τ 2 eu(trt) 2(σe 2+mσ2 u ) mn mn All assumes same number of observations per experimental unit c Dept. Statistics () Stat part 3 Spring / 116
70 What if the number of observations per experimental unit is not the same for all experimental units? Let s look at two miniature examples. Treatment 1: two eu s, one obs each Treatment 2: one eu. but two obs on that eu y = y 111 y 121 y 211 y 212 Treatment 1 Treatment 2 y 111 y 121 y 211 y 212 X = Z = c Dept. Statistics () Stat part 3 Spring / 116
71 X 1 = 1, X 2 = X, X 3 = Z MS TRT = y (P 2 P 1 )y = 2(y 1.. y... ) 2 + 2(y 2.. y... ) 2 = (y 1.. y 2.. ) 2 MS eu(trt ) = y (P 3 P 2 )y = (y 111 y 1.. ) 2 +(y 121 y 1.. ) 2 = 1 2 (y 111 y 121 ) 2 MS ou(eu.trt ) = y (I P 3 )y = (y 211 y 2.. ) 2 +(y 212 y 2.. ) 2 = 1 2 (y 211 y 212 ) 2 c Dept. Statistics () Stat part 3 Spring / 116
72 E(MS TRT ) = E(y 1.. y 2.. ) 2 = E(τ 1 τ 2 + u 1. u 21 + e 1.. e 2.. ) 2 = (τ 1 τ 2 ) 2 + Var(u 1. ) + Var(u 21 ) + Var(e 1.. ) + Var(e 2.. ) = (τ 1 τ 2 ) 2 + σ2 u 2 + σu 2 + σ2 e 2 + σ2 e 2 = (τ 1 τ 2 ) σ 2 u + σ 2 e c Dept. Statistics () Stat part 3 Spring / 116
73 E(MS eu(trt ) ) = 1 2 E(y 111 y 121 ) 2 = 1 2 E(u 11 u 12 + e 111 e 121 ) 2 = 1 2 (2σ2 u + 2σ 2 e) = σ 2 u + σ 2 e E(MS ou(eu,trt ) ) = 1 2 E(y 211 y 212 ) 2 = 1 2 E(e 211 e 212 ) 2 = σ 2 e c Dept. Statistics () Stat part 3 Spring / 116
74 SOURCE trt eu(trt ) ou(eu, TRT ) F = EMS (τ 1 τ 2 ) σu 2 + σe 2 σu 2 + σe 2 σ 2 e MS TRT 1.5σ 2 u+σ 2 e MS eu(trt ) σ 2 u+σ 2 e F ( ) (τ 1 τ 2 ) 2 1.5σu 2+σ2 e 1,1 c Dept. Statistics () Stat part 3 Spring / 116
75 The test statistic that we used to test H 0 : τ 1 =... = τ t in the balanced case is not F distributed in this unbalanced case: MS TRT MS eu(trt ) 1.5σ2 u + σ 2 e σ 2 u + σ 2 e F ( ) (τ 1 τ 2 ) 2 1.5σu 2+σ2 e 1,1 We d like our denominator to be an unbiased estimator of 1.5σ 2 u + σ 2 e in this case. No MS with this expectation, so construct one Consider 1.5MS eu(trt ) 0.5MS ou(eu,trt ) The expectation is 1.5(σu 2 + σe) 2 0.5σe 2 = 1.5σu 2 + σe 2 Use F = MS TRT 1.5MS eu(trt ) 0.5MS ou(eu,trt ) What is its distribution under Ho? c Dept. Statistics () Stat part 3 Spring / 116
76 COCHRAN-SATTERTHWAITE APPROXIMATION FOR LINEAR COMBINATIONS OF MEAN SQUARES Cochran-Satterthwaite method provides an approximation to the distribution of linear combinations of Chi-square random variables Suppose MS 1,..., MS k are independent random variables and that df i MS i E(MS i ) χ 2 df i i = 1,..., k. Consider the random variable S 2 = a 1 MS 1 + a 2 MS a k MS k, where a 1, a 2,..., a k are known positive constants in IR + C-S says νcss2 E(S 2 ) χ2 ν cs, where d.f., ν cs, can be computed Often used when some a i < 0. Theoretical support in this case much weaker. Support of the random variable should be IR + but S 2 may be < 0. c Dept. Statistics () Stat part 3 Spring / 116
77 Deriving the approximate df for S 2 = a 1 MS 1 + a 2 MS a k MS k : E(S 2 ) = a 1 E(MS 1 ) a k E(MS k ) Var(S 2 ) = a1 2 Var(MS 1) ak 2 Var(MS K ) = a1 2 [E(MS 1) ] 2 2df ak 2 df [E(MS k) ] 2 2df k 1 df k = 2 ai 2[E(MS i)] 2 df i A natural estimator of Var(S 2 ) is Var(S ˆ 2 ) 2 k i=1 a2 i MS2 i /df i Recall that df i MS i E(MS i ) Xdf 2 i i = 1,..., k. If S 2 = a 1 MS a k MS k is distributed like each of the random variables in the linear combination, Xν 2 cs May be a stretch when some a i < 0 νcs S2 E(S 2 ) c Dept. Statistics () Stat part 3 Spring / 116
78 Note that E[ ν css 2 E(S 2 ) ] = ν cs = E(X 2 ν cs ) Var[ ν css 2 E(S 2 ) ] = ν 2 cs [E(S 2 )] 2 Var(S2 ) Equating this expression to Var(Xν 2 cs ) = 2ν cs and solving for ν cs yields ν cs = 2[E(S2 )] 2 Var(S 2 ). Replacing E(S 2 ) with S 2 and Var(S 2 ) with Var(S ˆ 2 ) gives the Cochran-Satterthwaite formula for the approximate degrees of freedom of a linear combination of Mean Squares ˆν cs = (S 2 ) 2 k i=1 a2 i MS2 i /df i = ( k i=1 a ims i ) 2 k i=1 a2 i MS2 i /df i c Dept. Statistics () Stat part 3 Spring / 116
79 1/ν cs is linear combination of 1/df i We want approximate df for 1.5MS eu(trt) 0.5MS ou(eu,trt) That is: ( ) 2 1.5MSeu(trt) 0.5MS ou(eu,trt) ˆν cs = (1.5MS eu(trt) ) 2 /df eu(trt) + (0.5MS ou(eu,trt) ) 2 /df ou(eu,trt) So our F statistic F 1,νcs Examples: eu(trt) ou(eu,trt) 1.5MS MS 2 1.5MS 1 0.5MS 2 MS df MS df ˆν cs ˆν cs c Dept. Statistics () Stat part 3 Spring / 116
80 What does the BLUE of the treatment means look like in this case? [ ] µ1 β = X = Z = µ c Dept. Statistics () Stat part 3 Spring / 116
81 It follows that Var(y) = = ZGZ + R = σuzz 2 + σei = σu σ e ˆβ Σ = (X Σ 1 X) X Σ 1 y [ 1 ] [ ] 1 = y y = 1.. y Fortunately, this is a linear estimator that does not depend on unknown variance components. c Dept. Statistics () Stat part 3 Spring / 116
82 Consider a slightly different scenario: Treatment 1: eu #1 with 2 obs, eu #2 with 1 obs Treatment 2: 1 eu with 1 obs y = y 111 y 112 y 121 y 211 X = Z = c Dept. Statistics () Stat part 3 Spring / 116
83 In this case, it can be shown that ˆβ Σ = (X Σ 1 X) X Σ 1 y [ ] y σ e +σu 2 σe+σ 2 2 = 3σe+4σ 2 u 2 u σe+2σ 2 2 3σe+4σ 2 u 2 u 0 3σe+4σ 2 u 2 y y 121 = [ 2σ 2 e +2σ 2 u 3σ 2 e+4σ 2 u y y 211 σ 2 e +2σ2 u 3σ 2 e+4σ 2 u y 121 y 211 ] c Dept. Statistics () Stat part 3 Spring / 116
84 It is straightforward to show that the weights on y 11. and y 121 are 1 Var(y 11. ) 1 Var(y 11. ) + 1 Var(y 121 ) and 1 Var(y 121 ) 1 Var(y 11. ) + 1 Var(y 121 ) respectively Of course, ˆβ Σ = [ 2σ 2 e +2σ 2 u 3σ 2 e+4σ 2 u y σ 2 e+2σ 2 u 3σ 2 e+4σ 2 u y 121 ] y 211 is not an estimator because it is a function of unknown parameters. Thus we use ˆβ Σ as our estimator (replace σ 2 e and σ 2 u by estimates in the expression above). c Dept. Statistics () Stat part 3 Spring / 116
85 ˆβ Σ is an approximation to the BLUE. ˆβ Σ is not even a linear estimator in this case. Its exact distribution is unknown. When sample sizes are large, it is reasonable to assume that the distribution of ˆβ Σ is approx. the same as the distribution of ˆβ Σ. Var(ˆβ Σ ) = Var[(X Σ 1 X) 1 X Σ 1 y] = (X Σ 1 X) 1 X Σ 1 Var(y)[(X Σ 1 X) 1 X Σ 1 ] = (X Σ 1 X) 1 X Σ 1 )ΣΣ 1 X(X Σ 1 X) 1 = (X Σ 1 X) 1 X Σ 1 X(X Σ 1 X) 1 = (X Σ 1 X) 1 Var(ˆβ Σ) = Var[(X Σ 1 X) 1 X Σ 1 y] =? (X Σ 1 X) 1 c Dept. Statistics () Stat part 3 Spring / 116
86 Summary of Main Points Many of the concepts we have seen by examining special cases hold in greater generality. For many of the linear mixed models commonly used in practice, balanced data are nice because 1. It is relatively easy to determine degrees of freedom, sums of Squares and expected mean squares in an ANOVA table. 2. Ratios of appropriate mean squares can be used to obtain exact F-tests. 3. For estimable Cβ, C ˆβ Σ = C ˆβ.(The OLS estimator equals the GLS estimator). 4. When Var(C ˆβ) = constant E(MS), exact inferences about Cβ can be obtained by constructing t tests or confidence intervals C t = ˆβ Cβ t νcs(ms) constant (MS) 5. Simple analysis based on experimental unit averages gives the same results as those obtained by linear mixed model analysis of the full data set. c Dept. Statistics () Stat part 3 Spring / 116
87 When data are unbalanced, the analysis of linear mixed may be considerably more complicated. 1. Approximate F tests can be obtained by forming linear combinations of Mean Squares to obtain denominators for test statistics. 2. The estimator C ˆβ Σ may be a nonlinear estimator of Cβ whose exact distribution is unknown. 3. Approximate inference for Cβ is often obtained by using the distribution of C ˆβ Σ, with unknowns in that distribution replaced by estimates. Whether data are balanced or unbalanced, unbiased estimators of variance components can be obtained by the method of moments. c Dept. Statistics () Stat part 3 Spring / 116
88 ANOVA ANALYSIS OF A BALANCED SPLIT-PLOT EXPERIMENT Example: the corn genotype and fertilization response study Main plots: genotypes, in blocks Split plots: fertilization 2 way factorial treatment structure split plot variability nested in main plot variability nested in blocks. c Dept. Statistics () Stat part 3 Spring / 116
89 Field Plot Block 1 Block 2 Block 3 Block 4 Genotype C Genotype B Genotype A Genotype B Genotype C Genotype B Genotype A Genotype A Genotype C Genotype B Genotype C Genotype A Split Plot or Sub Plot c Dept. Statistics () Stat part 3 Spring / 116
90 A model: y ijk = µ ij + b k + w ik + e ijk µ ij =Mean for Genotype i, Fertilizer j Genotype i = 1, 2, 3, Fertilizer j = 1, 2, 3, 4, Block k = 1, 2, 3, 4 u = b 1. b 4 w 11 w 21. w 34 = [ u ɛ [ b w ] N ] N ([ 0 0 ([ 0 0 ] [ G 0, 0 σe 2 I ] [ σ 2, b I 0 0 σw 2 I ]) ]) c Dept. Statistics () Stat part 3 Spring / 116
91 Source DF Blocks 4 1 = 3 Genotypes 3 1 = 2 Blocks Geno (4 1)(3 1) = 6 whole plot u Fert 4 1 = 3 Geno Fert (3 1)(4 1) = 6 Block Fert (4 1)(4 1) +Blocks Geno Fert +(4 1)(3 1)(4 1) 27 split plot u C.Total c Dept. Statistics () Stat part 3 Spring / 116
92 Source Sum of Squares Blocks i=1 j=1 k=1 (ȳ..k ȳ... ) 2 Geno i=1 j=1 k=1 (ȳ i.. ȳ... ) 2 Blocks Geno i=1 j=1 k=1 (ȳ i.k ȳ i.. ȳ..k + ȳ... ) 2 Fert i=1 j=1 k=1 (ȳ.j. ȳ... ) 2 Geno Fert i=1 j=1 k=1 (ȳ ij. ȳ i.. ȳ.j. + ȳ... ) 2 error i=1 j=1 k=1 (ȳ ijk ȳ i.k ȳ ij. + ȳ i.. ) 2 C.Total i=1 j=1 k=1 (ȳ ijk ȳ... ) 2 c Dept. Statistics () Stat part 3 Spring / 116
93 E(MS GENO ) = FB G G 1 i=1 E(ȳ i.. ȳ...) 2 = FB G G 1 i=1 E( µ i. µ.. + w i. w.. + ē i. ē... ) 2 G i=1 = FB ( µ i. µ..) 2 = FB G 1 G i=1 + FB E ( w i. w..) 2 G i=1 G 1 + FB E (ē i. ē..) G 1 G i=1 ( µ i. µ..) 2 G 1 + FB σ2 w B + FB σ2 e FB G i=1 = FB ( µ i. µ..) 2 G 1 + Fσw 2 + σe 2 c Dept. Statistics () Stat part 3 Spring / 116
94 = = = E(MS Block GENO ) = F (B 1)(G 1) G i=1 k=1 F G (B 1)(G 1) E[ F (B 1)(G 1) G i=1 k=1 B E(ȳ i.k ȳ i.. ȳ..k ȳ...) 2 B E(w ik w i. w.k + w.. +ē i.k ē i.. +ē..k +ē...) 2 i=1 k=1 + G F G (B 1)(G 1) E[ = B (w ik w i. ) 2 2 i=1 k=1 i=1 k=1 G i=1 k=1 B ( w.k w.. ) 2 + e terms] B (w ik w i. ) 2 G B (w ik w i. )( w.k w.. ) B (w.k w.. ) 2 + e terms] k=1 F (B 1)(G 1) [G(B 1)σ2 w G(B 1)σ 2 w/g + e terms] c Dept. Statistics () = Stat Fσ part + σ3 2 Spring / 116
95 To test for genotype main effects, i.e., H 0 : µ 1. = µ 2. = µ 3. H 0 : MS GENO MS Blcok Geno compare (B 1)(G 1) df. Reject H 0 at level α iff FB G 1 G ( µ i. µ.. ) 2 = 0, i=1 to a central F distribution with G 1 and MS GENO MS Blcok Geno N.B. notation: F α is the 1 α quantile. F α G 1, (B 1)(G 1) c Dept. Statistics () Stat part 3 Spring / 116
96 Contrasts among genotype means: Var(ȳ 1.. ȳ 2.. ) = Var( µ 1. µ 2. + w 1. + w 2. + ē 1.. ē 2.. ) = 2σ2 w B + 2σ2 e FB = 2 FB (Fσ2 w + σ 2 e) = 2 FB E(MS Block GENO) Can use Var(ȳ ˆ 1.. ȳ 2.. ) = 2 FB MS Block GENO t = ȳ1.. ȳ 2.. ( µ 1. µ 2. ) 2 FB MS Block GENO t (B 1)(G 1) to get tests of H 0 : µ 1. = µ 2. or construct confidence intervals for µ 1. µ 2. c Dept. Statistics () Stat part 3 Spring / 116
97 Furthermore, suppose C is a matrix whose rows are contrast vectors so that C1 = 0. Then y 1.. b. + w 1. + ē 1.. Var C. = Var C. y G.. b. + w G. + ē G.. w 1. + ē 1.. w 1. + ē 1.. = Var C1 b. + C. = C Var. C w G. + ē G.. w G. + ē G.. = C( σ2 w B + σ2 e FB )IC = ( σ2 w B + σ2 e FB )CC = E(MS MS ) Block GENO CC FB c Dept. Statistics () Stat part 3 Spring / 116
98 An F statistic for testing F = H 0 : C C µ 1.. µ G. ȳ 1.. ȳ G. = 0, with df = q, (B 1)(G 1), is [ MSBlock GENO FB q ] 1 CC C ȳ 1... ȳ G.. where q is the number of rows of C (which must have full row rank to ensure that the hypothesis is testable) c Dept. Statistics () Stat part 3 Spring / 116
99 Inference on fertilizer main effects, µ.j : E(MS FERT ) = GB F 1 = GB F 1 = GB F 1 FE(ȳ.j. ȳ... ) 2 i=1 FE( µ.j µ.. + ē.j ē.. ) 2 i=1 F( µ.j µ.. ) 2 + σe 2 i=1 To test for fertilizer main effects, i.e. Ho : µ.1 = µ.2 = µ.3 = µ.4 Ho : GB F 1 F(µ.i µ.. ) 2 = 0, compare MS FERT MS E rror to a central F distribution with F 1 and G(B 1)(F 1) df i=1 c Dept. Statistics () Stat part 3 Spring / 116
100 Contrasts among fertilizer means: Note: ȳ.1. ȳ.2. = (µ.1 + b. + w.. + ē.1. ) (µ.2 + b. + w.. + ē.2. ) Var(ȳ.1. ȳ.2. ) = Var( µ.1 µ.2 + ē.1. ē.2. ) = 2σ2 e GB = 2 GB E(MS Error ) ˆ Var(ȳ.1. ȳ.2. ) = 2 GB MS Error Can use t = ȳ.1. ȳ.2. ( µ.1 µ.2 ) 2 GB MS Error t G(B 1)(F 1) to get tests of H 0 : µ.1 = µ.2 or construct confidence intervals for µ.1 µ.2 c Dept. Statistics () Stat part 3 Spring / 116
101 Furthermore, suppose C is a matrix whose rows are contrast vectors so that C1 = 0. Then y.1. b. + w.. + ē.1. Var C. = Var C. y.f. b. + w.. + ē.f. = Var C1 b. + C1 w.. + = C ē.1.. ē.f. = C Var ( ) σ 2 e IC = E(MS Error ) CC GB GB ē.1.. ē.f. C c Dept. Statistics () Stat part 3 Spring / 116
102 An F statistic for testing H 0 : C F = µ.1. µ.f C = 0, with df = q, G(B 1)(F 1), is ȳ.1. ȳ.f [ MSError FB ] 1 CC C q ȳ.1.. ȳ.f. where q is the number of rows of C (which must have full row rank to ensure that the hypothesis is testable) c Dept. Statistics () Stat part 3 Spring / 116
103 Inferences on the interactions: same as fertilizer main effects Use MS Error Inferences on simple effects: Two types, details different Diff. between two fertilizers within a genotype, e.g. µ 11 µ 12 Var(ȳ 11. ȳ 12. ) = Var(µ 11 µ 11 + b. b. + w.. w.. + ē 11. ē 12 = 2σ2 e B ˆ Var(ȳ 11. ȳ 12. ) = 2 B MS Error Diff. between two genotypes within a fertilizer, e.g. µ 11 µ 21 Var(ȳ 11. ȳ 21. ) = Var(µ 11 µ 21 + w 1. w 2. + ē 11. ē 21. ) = 2σ2 w B + 2σ2 e B = 2 B (σ2 w + σ 2 e) c Dept. Statistics () Stat part 3 Spring / 116
104 Need an estimator of σ 2 w + σ 2 e From ANOVA table: E MS Block GENO = F σ 2 w + σ 2 e E MS Error = σ 2 e Use a linear combination of these to estimate σw 2 + σe 2 ( ) MSBlock GENO E F + F 1 F MS ERROR = σw 2 + σ2 e F + (F 1)σ2 e F = σw 2 + σe 2 Var(ȳ ˆ 11. ȳ 21. ) 2 BF MS Block GENO + unbiased estimate of Var(ȳ 11. ȳ 21. ) ȳ 11. ȳ 21. (µ 11 µ 21 ) ˆ Var(ȳ 11. ȳ 21. ) 2(F 1) BF MS error is an t with d.f. determined by the Cochran-Satterthwaite approximation. For simple effects in a split plot, required linear combination is usually a sum of MS. CS works well for sums. c Dept. Statistics () Stat part 3 Spring / 116
105 Inferences on means: E MS for random effect terms in the ANOVA table E MS Block = σ 2 e + Fσ 2 w + GFσ 2 b E MS Block GENO = σ 2 e + Fσ 2 w E MS Error = σ 2 e Cell mean, µ ij Var ȳ ij. = Var(µ ij + b. + w i. + ē ij. ) = σ2 b B + σ2 w B + σ2 e B Need to construct an estimator: Var(ȳ ˆ ij. ) = 1 BGF [MS Block + (G 1) MS Block GENO + F(G 1) MS Error ] with approximate df from Cochran-Satterthwaite c Dept. Statistics () Stat part 3 Spring / 116
106 Main plot marginal mean, µ i. requires MoM estimate If blocks considered fixed: Var ȳ i.. = Var(µ i. + b. + w i. + ē i.. ) = σ2 b B + σ2 w B + σ2 e FB Var ȳ i.. = Var(µ i. + b. + w i. + ē i.. ) = σ2 w = 1 FB B + σ2 e FB ( Fσ 2 w + σ 2 e ) Can estimate this by MS Block GENO FB with (B 1)(G 1) df c Dept. Statistics () Stat part 3 Spring / 116
107 Split plot marginal mean, µ.j If blocks considered fixed: Both require MoM estimates. Var ȳ.i. = Var(µ.j + b. + w.. + ē.j. ) = σ2 b B + σ2 w BG + σ2 e BG Var ȳ.i. = Var(µ.j + b. + w.. + ē.j. ) = σ2 w BG + σ2 e BG c Dept. Statistics () Stat part 3 Spring / 116
108 Summary of ANOVA for a balanced split plot study Use main plot error MS for inferences on main plot marginal means (e.g. genotype) Use split plot error MS for inferences on split plot marginal means (e.g. fertilizer) main*split interactions (e.g. geno fert) a simple effect within a main plot treatment Construct a method of moments estimator for inferences on a simple effect within a split plot treatment a simple effect between two split trt and two main trt, e.g. µ 11 µ 22 most means c Dept. Statistics () Stat part 3 Spring / 116
109 Unbalanced split plots Unequal numbers of observations per treatment Details get complicated fast In general: coefficients for random effect variances in E MS do not match so MS Block GENO is not the appropriate denominator for MS GENO need to construct MoM estimators for all F denominators All inference based on approximate F tests or approximate t tests c Dept. Statistics () Stat part 3 Spring / 116
110 Two approaches for E MS RCBD with random blocks and multiple obs. per block Y ijk = µ + β i + τ j + βτ ij + ɛ ijk where i {1,..., B}, j {1,..., T }, k {1,..., N}. with ANOVA table: Source df R Blocks B-1 Treatments T-1 R Block Trt (B-1)(T-1) R Error BT(N-1) C. total BTN - 1 c Dept. Statistics () Stat part 3 Spring / 116
111 Expected Mean Squares from two different sources Source 1: Searle (1971) 2: Graybill (1976) Blocks Treatments σe 2 + Nσp 2 + NT σb 2 σe 2 + Nσp 2 + Q(τ) ηe 2 + NT ηb 2 ηe 2 + Nηp 2 + Q(τ) Block Trt σe 2 + Nσp 2 ηe 2 + Nηp 2 Error σe 2 ηe 2 1: Searle, S (1971) Linear Models 2: Graybill, F (1976) Theory and Application of the Linear Model. Same except for Blocks Different F tests for Blocks EMS 1: MS Blocks / MS Block*Trt EMS 2: MS Blocks / MS Error c Dept. Statistics () Stat part 3 Spring / 116
THE ANOVA APPROACH TO THE ANALYSIS OF LINEAR MIXED EFFECTS MODELS
 THE ANOVA APPROACH TO THE ANALYSIS OF LINEAR MIXED EFFECTS MODELS We begin with a relatively simple special case. Suppose y ijk = µ + τ i + u ij + e ijk, (i = 1,..., t; j = 1,..., n; k = 1,..., m) β =
THE ANOVA APPROACH TO THE ANALYSIS OF LINEAR MIXED EFFECTS MODELS We begin with a relatively simple special case. Suppose y ijk = µ + τ i + u ij + e ijk, (i = 1,..., t; j = 1,..., n; k = 1,..., m) β =
11. Linear Mixed-Effects Models. Copyright c 2018 Dan Nettleton (Iowa State University) 11. Statistics / 49
 11. Linear Mixed-Effects Models Copyright c 2018 Dan Nettleton (Iowa State University) 11. Statistics 510 1 / 49 The Linear Mixed-Effects Model y = Xβ + Zu + e X is an n p matrix of known constants β R
11. Linear Mixed-Effects Models Copyright c 2018 Dan Nettleton (Iowa State University) 11. Statistics 510 1 / 49 The Linear Mixed-Effects Model y = Xβ + Zu + e X is an n p matrix of known constants β R
Linear Mixed-Effects Models. Copyright c 2012 Dan Nettleton (Iowa State University) Statistics / 34
 Linear Mixed-Effects Models Copyright c 2012 Dan Nettleton (Iowa State University) Statistics 611 1 / 34 The Linear Mixed-Effects Model y = Xβ + Zu + e X is an n p design matrix of known constants β R
Linear Mixed-Effects Models Copyright c 2012 Dan Nettleton (Iowa State University) Statistics 611 1 / 34 The Linear Mixed-Effects Model y = Xβ + Zu + e X is an n p design matrix of known constants β R
REPEATED MEASURES. Copyright c 2012 (Iowa State University) Statistics / 29
 REPEATED MEASURES Copyright c 2012 (Iowa State University) Statistics 511 1 / 29 Repeated Measures Example In an exercise therapy study, subjects were assigned to one of three weightlifting programs i=1:
REPEATED MEASURES Copyright c 2012 (Iowa State University) Statistics 511 1 / 29 Repeated Measures Example In an exercise therapy study, subjects were assigned to one of three weightlifting programs i=1:
20. REML Estimation of Variance Components. Copyright c 2018 (Iowa State University) 20. Statistics / 36
 20. REML Estimation of Variance Components Copyright c 2018 (Iowa State University) 20. Statistics 510 1 / 36 Consider the General Linear Model y = Xβ + ɛ, where ɛ N(0, Σ) and Σ is an n n positive definite
20. REML Estimation of Variance Components Copyright c 2018 (Iowa State University) 20. Statistics 510 1 / 36 Consider the General Linear Model y = Xβ + ɛ, where ɛ N(0, Σ) and Σ is an n n positive definite
ANOVA Variance Component Estimation. Copyright c 2012 Dan Nettleton (Iowa State University) Statistics / 32
 ANOVA Variance Component Estimation Copyright c 2012 Dan Nettleton (Iowa State University) Statistics 611 1 / 32 We now consider the ANOVA approach to variance component estimation. The ANOVA approach
ANOVA Variance Component Estimation Copyright c 2012 Dan Nettleton (Iowa State University) Statistics 611 1 / 32 We now consider the ANOVA approach to variance component estimation. The ANOVA approach
Peter Hoff Linear and multilinear models April 3, GLS for multivariate regression 5. 3 Covariance estimation for the GLM 8
 Contents 1 Linear model 1 2 GLS for multivariate regression 5 3 Covariance estimation for the GLM 8 4 Testing the GLH 11 A reference for some of this material can be found somewhere. 1 Linear model Recall
Contents 1 Linear model 1 2 GLS for multivariate regression 5 3 Covariance estimation for the GLM 8 4 Testing the GLH 11 A reference for some of this material can be found somewhere. 1 Linear model Recall
Lecture 3: Linear Models. Bruce Walsh lecture notes Uppsala EQG course version 28 Jan 2012
 Lecture 3: Linear Models Bruce Walsh lecture notes Uppsala EQG course version 28 Jan 2012 1 Quick Review of the Major Points The general linear model can be written as y = X! + e y = vector of observed
Lecture 3: Linear Models Bruce Walsh lecture notes Uppsala EQG course version 28 Jan 2012 1 Quick Review of the Major Points The general linear model can be written as y = X! + e y = vector of observed
Regression Review. Statistics 149. Spring Copyright c 2006 by Mark E. Irwin
 Regression Review Statistics 149 Spring 2006 Copyright c 2006 by Mark E. Irwin Matrix Approach to Regression Linear Model: Y i = β 0 + β 1 X i1 +... + β p X ip + ɛ i ; ɛ i iid N(0, σ 2 ), i = 1,..., n
Regression Review Statistics 149 Spring 2006 Copyright c 2006 by Mark E. Irwin Matrix Approach to Regression Linear Model: Y i = β 0 + β 1 X i1 +... + β p X ip + ɛ i ; ɛ i iid N(0, σ 2 ), i = 1,..., n
One-way ANOVA (Single-Factor CRD)
 One-way ANOVA (Single-Factor CRD) STAT:5201 Week 3: Lecture 3 1 / 23 One-way ANOVA We have already described a completed randomized design (CRD) where treatments are randomly assigned to EUs. There is
One-way ANOVA (Single-Factor CRD) STAT:5201 Week 3: Lecture 3 1 / 23 One-way ANOVA We have already described a completed randomized design (CRD) where treatments are randomly assigned to EUs. There is
ANOVA Variance Component Estimation. Copyright c 2012 Dan Nettleton (Iowa State University) Statistics / 32
 ANOVA Variance Component Estimation Copyright c 2012 Dan Nettleton (Iowa State University) Statistics 611 1 / 32 We now consider the ANOVA approach to variance component estimation. The ANOVA approach
ANOVA Variance Component Estimation Copyright c 2012 Dan Nettleton (Iowa State University) Statistics 611 1 / 32 We now consider the ANOVA approach to variance component estimation. The ANOVA approach
Stat 217 Final Exam. Name: May 1, 2002
 Stat 217 Final Exam Name: May 1, 2002 Problem 1. Three brands of batteries are under study. It is suspected that the lives (in weeks) of the three brands are different. Five batteries of each brand are
Stat 217 Final Exam Name: May 1, 2002 Problem 1. Three brands of batteries are under study. It is suspected that the lives (in weeks) of the three brands are different. Five batteries of each brand are
Unit 7: Random Effects, Subsampling, Nested and Crossed Factor Designs
 Unit 7: Random Effects, Subsampling, Nested and Crossed Factor Designs STA 643: Advanced Experimental Design Derek S. Young 1 Learning Objectives Understand how to interpret a random effect Know the different
Unit 7: Random Effects, Subsampling, Nested and Crossed Factor Designs STA 643: Advanced Experimental Design Derek S. Young 1 Learning Objectives Understand how to interpret a random effect Know the different
Lecture 2: Linear Models. Bruce Walsh lecture notes Seattle SISG -Mixed Model Course version 23 June 2011
 Lecture 2: Linear Models Bruce Walsh lecture notes Seattle SISG -Mixed Model Course version 23 June 2011 1 Quick Review of the Major Points The general linear model can be written as y = X! + e y = vector
Lecture 2: Linear Models Bruce Walsh lecture notes Seattle SISG -Mixed Model Course version 23 June 2011 1 Quick Review of the Major Points The general linear model can be written as y = X! + e y = vector
2. A Review of Some Key Linear Models Results. Copyright c 2018 Dan Nettleton (Iowa State University) 2. Statistics / 28
 2. A Review of Some Key Linear Models Results Copyright c 2018 Dan Nettleton (Iowa State University) 2. Statistics 510 1 / 28 A General Linear Model (GLM) Suppose y = Xβ + ɛ, where y R n is the response
2. A Review of Some Key Linear Models Results Copyright c 2018 Dan Nettleton (Iowa State University) 2. Statistics 510 1 / 28 A General Linear Model (GLM) Suppose y = Xβ + ɛ, where y R n is the response
Formal Statement of Simple Linear Regression Model
 Formal Statement of Simple Linear Regression Model Y i = β 0 + β 1 X i + ɛ i Y i value of the response variable in the i th trial β 0 and β 1 are parameters X i is a known constant, the value of the predictor
Formal Statement of Simple Linear Regression Model Y i = β 0 + β 1 X i + ɛ i Y i value of the response variable in the i th trial β 0 and β 1 are parameters X i is a known constant, the value of the predictor
Stat 579: Generalized Linear Models and Extensions
 Stat 579: Generalized Linear Models and Extensions Linear Mixed Models for Longitudinal Data Yan Lu April, 2018, week 15 1 / 38 Data structure t1 t2 tn i 1st subject y 11 y 12 y 1n1 Experimental 2nd subject
Stat 579: Generalized Linear Models and Extensions Linear Mixed Models for Longitudinal Data Yan Lu April, 2018, week 15 1 / 38 Data structure t1 t2 tn i 1st subject y 11 y 12 y 1n1 Experimental 2nd subject
Two-Way Factorial Designs
 81-86 Two-Way Factorial Designs Yibi Huang 81-86 Two-Way Factorial Designs Chapter 8A - 1 Problem 81 Sprouting Barley (p166 in Oehlert) Brewer s malt is produced from germinating barley, so brewers like
81-86 Two-Way Factorial Designs Yibi Huang 81-86 Two-Way Factorial Designs Chapter 8A - 1 Problem 81 Sprouting Barley (p166 in Oehlert) Brewer s malt is produced from germinating barley, so brewers like
Linear models and their mathematical foundations: Simple linear regression
 Linear models and their mathematical foundations: Simple linear regression Steffen Unkel Department of Medical Statistics University Medical Center Göttingen, Germany Winter term 2018/19 1/21 Introduction
Linear models and their mathematical foundations: Simple linear regression Steffen Unkel Department of Medical Statistics University Medical Center Göttingen, Germany Winter term 2018/19 1/21 Introduction
STAT22200 Spring 2014 Chapter 8A
 STAT22200 Spring 2014 Chapter 8A Yibi Huang May 13, 2014 81-86 Two-Way Factorial Designs Chapter 8A - 1 Problem 81 Sprouting Barley (p166 in Oehlert) Brewer s malt is produced from germinating barley,
STAT22200 Spring 2014 Chapter 8A Yibi Huang May 13, 2014 81-86 Two-Way Factorial Designs Chapter 8A - 1 Problem 81 Sprouting Barley (p166 in Oehlert) Brewer s malt is produced from germinating barley,
Matrix Approach to Simple Linear Regression: An Overview
 Matrix Approach to Simple Linear Regression: An Overview Aspects of matrices that you should know: Definition of a matrix Addition/subtraction/multiplication of matrices Symmetric/diagonal/identity matrix
Matrix Approach to Simple Linear Regression: An Overview Aspects of matrices that you should know: Definition of a matrix Addition/subtraction/multiplication of matrices Symmetric/diagonal/identity matrix
13. The Cochran-Satterthwaite Approximation for Linear Combinations of Mean Squares
 13. The Cochran-Satterthwaite Approximation for Linear Combinations of Mean Squares opyright c 2018 Dan Nettleton (Iowa State University) 13. Statistics 510 1 / 18 Suppose M 1,..., M k are independent
13. The Cochran-Satterthwaite Approximation for Linear Combinations of Mean Squares opyright c 2018 Dan Nettleton (Iowa State University) 13. Statistics 510 1 / 18 Suppose M 1,..., M k are independent
Topic 25 - One-Way Random Effects Models. Outline. Random Effects vs Fixed Effects. Data for One-way Random Effects Model. One-way Random effects
 Topic 5 - One-Way Random Effects Models One-way Random effects Outline Model Variance component estimation - Fall 013 Confidence intervals Topic 5 Random Effects vs Fixed Effects Consider factor with numerous
Topic 5 - One-Way Random Effects Models One-way Random effects Outline Model Variance component estimation - Fall 013 Confidence intervals Topic 5 Random Effects vs Fixed Effects Consider factor with numerous
Applied Statistics Preliminary Examination Theory of Linear Models August 2017
 Applied Statistics Preliminary Examination Theory of Linear Models August 2017 Instructions: Do all 3 Problems. Neither calculators nor electronic devices of any kind are allowed. Show all your work, clearly
Applied Statistics Preliminary Examination Theory of Linear Models August 2017 Instructions: Do all 3 Problems. Neither calculators nor electronic devices of any kind are allowed. Show all your work, clearly
Chapter 12. Analysis of variance
 Serik Sagitov, Chalmers and GU, January 9, 016 Chapter 1. Analysis of variance Chapter 11: I = samples independent samples paired samples Chapter 1: I 3 samples of equal size J one-way layout two-way layout
Serik Sagitov, Chalmers and GU, January 9, 016 Chapter 1. Analysis of variance Chapter 11: I = samples independent samples paired samples Chapter 1: I 3 samples of equal size J one-way layout two-way layout
Lectures on Simple Linear Regression Stat 431, Summer 2012
 Lectures on Simple Linear Regression Stat 43, Summer 0 Hyunseung Kang July 6-8, 0 Last Updated: July 8, 0 :59PM Introduction Previously, we have been investigating various properties of the population
Lectures on Simple Linear Regression Stat 43, Summer 0 Hyunseung Kang July 6-8, 0 Last Updated: July 8, 0 :59PM Introduction Previously, we have been investigating various properties of the population
Simple Linear Regression
 Simple Linear Regression In simple linear regression we are concerned about the relationship between two variables, X and Y. There are two components to such a relationship. 1. The strength of the relationship.
Simple Linear Regression In simple linear regression we are concerned about the relationship between two variables, X and Y. There are two components to such a relationship. 1. The strength of the relationship.
Introduction to Estimation Methods for Time Series models. Lecture 1
 Introduction to Estimation Methods for Time Series models Lecture 1 Fulvio Corsi SNS Pisa Fulvio Corsi Introduction to Estimation () Methods for Time Series models Lecture 1 SNS Pisa 1 / 19 Estimation
Introduction to Estimation Methods for Time Series models Lecture 1 Fulvio Corsi SNS Pisa Fulvio Corsi Introduction to Estimation () Methods for Time Series models Lecture 1 SNS Pisa 1 / 19 Estimation
STAT 540: Data Analysis and Regression
 STAT 540: Data Analysis and Regression Wen Zhou http://www.stat.colostate.edu/~riczw/ Email: riczw@stat.colostate.edu Department of Statistics Colorado State University Fall 205 W. Zhou (Colorado State
STAT 540: Data Analysis and Regression Wen Zhou http://www.stat.colostate.edu/~riczw/ Email: riczw@stat.colostate.edu Department of Statistics Colorado State University Fall 205 W. Zhou (Colorado State
Ch 2: Simple Linear Regression
 Ch 2: Simple Linear Regression 1. Simple Linear Regression Model A simple regression model with a single regressor x is y = β 0 + β 1 x + ɛ, where we assume that the error ɛ is independent random component
Ch 2: Simple Linear Regression 1. Simple Linear Regression Model A simple regression model with a single regressor x is y = β 0 + β 1 x + ɛ, where we assume that the error ɛ is independent random component
Bias Variance Trade-off
 Bias Variance Trade-off The mean squared error of an estimator MSE(ˆθ) = E([ˆθ θ] 2 ) Can be re-expressed MSE(ˆθ) = Var(ˆθ) + (B(ˆθ) 2 ) MSE = VAR + BIAS 2 Proof MSE(ˆθ) = E((ˆθ θ) 2 ) = E(([ˆθ E(ˆθ)]
Bias Variance Trade-off The mean squared error of an estimator MSE(ˆθ) = E([ˆθ θ] 2 ) Can be re-expressed MSE(ˆθ) = Var(ˆθ) + (B(ˆθ) 2 ) MSE = VAR + BIAS 2 Proof MSE(ˆθ) = E((ˆθ θ) 2 ) = E(([ˆθ E(ˆθ)]
BIOS 2083 Linear Models c Abdus S. Wahed
 Chapter 5 206 Chapter 6 General Linear Model: Statistical Inference 6.1 Introduction So far we have discussed formulation of linear models (Chapter 1), estimability of parameters in a linear model (Chapter
Chapter 5 206 Chapter 6 General Linear Model: Statistical Inference 6.1 Introduction So far we have discussed formulation of linear models (Chapter 1), estimability of parameters in a linear model (Chapter
General Linear Model: Statistical Inference
 Chapter 6 General Linear Model: Statistical Inference 6.1 Introduction So far we have discussed formulation of linear models (Chapter 1), estimability of parameters in a linear model (Chapter 4), least
Chapter 6 General Linear Model: Statistical Inference 6.1 Introduction So far we have discussed formulation of linear models (Chapter 1), estimability of parameters in a linear model (Chapter 4), least
Estimating σ 2. We can do simple prediction of Y and estimation of the mean of Y at any value of X.
 Estimating σ 2 We can do simple prediction of Y and estimation of the mean of Y at any value of X. To perform inferences about our regression line, we must estimate σ 2, the variance of the error term.
Estimating σ 2 We can do simple prediction of Y and estimation of the mean of Y at any value of X. To perform inferences about our regression line, we must estimate σ 2, the variance of the error term.
Spatial analysis of a designed experiment. Uniformity trials. Blocking
 Spatial analysis of a designed experiment RA Fisher s 3 principles of experimental design randomization unbiased estimate of treatment effect replication unbiased estimate of error variance blocking =
Spatial analysis of a designed experiment RA Fisher s 3 principles of experimental design randomization unbiased estimate of treatment effect replication unbiased estimate of error variance blocking =
Stat 579: Generalized Linear Models and Extensions
 Stat 579: Generalized Linear Models and Extensions Mixed models Yan Lu Feb, 2018, week 7 1 / 17 Some commonly used experimental designs related to mixed models Two way or three way random/mixed effects
Stat 579: Generalized Linear Models and Extensions Mixed models Yan Lu Feb, 2018, week 7 1 / 17 Some commonly used experimental designs related to mixed models Two way or three way random/mixed effects
Example 1: Two-Treatment CRD
 Introduction to Mixed Linear Models in Microarray Experiments //0 Copyright 0 Dan Nettleton Statistical Models A statistical model describes a formal mathematical data generation mechanism from which an
Introduction to Mixed Linear Models in Microarray Experiments //0 Copyright 0 Dan Nettleton Statistical Models A statistical model describes a formal mathematical data generation mechanism from which an
Math 423/533: The Main Theoretical Topics
 Math 423/533: The Main Theoretical Topics Notation sample size n, data index i number of predictors, p (p = 2 for simple linear regression) y i : response for individual i x i = (x i1,..., x ip ) (1 p)
Math 423/533: The Main Theoretical Topics Notation sample size n, data index i number of predictors, p (p = 2 for simple linear regression) y i : response for individual i x i = (x i1,..., x ip ) (1 p)
Estimating Estimable Functions of β. Copyright c 2012 Dan Nettleton (Iowa State University) Statistics / 17
 Estimating Estimable Functions of β Copyright c 202 Dan Nettleton (Iowa State University) Statistics 5 / 7 The Response Depends on β Only through Xβ In the Gauss-Markov or Normal Theory Gauss-Markov Linear
Estimating Estimable Functions of β Copyright c 202 Dan Nettleton (Iowa State University) Statistics 5 / 7 The Response Depends on β Only through Xβ In the Gauss-Markov or Normal Theory Gauss-Markov Linear
Analysis of Variance
 Analysis of Variance Math 36b May 7, 2009 Contents 2 ANOVA: Analysis of Variance 16 2.1 Basic ANOVA........................... 16 2.1.1 the model......................... 17 2.1.2 treatment sum of squares.................
Analysis of Variance Math 36b May 7, 2009 Contents 2 ANOVA: Analysis of Variance 16 2.1 Basic ANOVA........................... 16 2.1.1 the model......................... 17 2.1.2 treatment sum of squares.................
Weighted Least Squares
 Weighted Least Squares The standard linear model assumes that Var(ε i ) = σ 2 for i = 1,..., n. As we have seen, however, there are instances where Var(Y X = x i ) = Var(ε i ) = σ2 w i. Here w 1,..., w
Weighted Least Squares The standard linear model assumes that Var(ε i ) = σ 2 for i = 1,..., n. As we have seen, however, there are instances where Var(Y X = x i ) = Var(ε i ) = σ2 w i. Here w 1,..., w
Factorial Treatment Structure: Part I. Lukas Meier, Seminar für Statistik
 Factorial Treatment Structure: Part I Lukas Meier, Seminar für Statistik Factorial Treatment Structure So far (in CRD), the treatments had no structure. So called factorial treatment structure exists if
Factorial Treatment Structure: Part I Lukas Meier, Seminar für Statistik Factorial Treatment Structure So far (in CRD), the treatments had no structure. So called factorial treatment structure exists if
Lecture 11: Nested and Split-Plot Designs
 Lecture 11: Nested and Split-Plot Designs Montgomery, Chapter 14 1 Lecture 11 Page 1 Crossed vs Nested Factors Factors A (a levels)and B (b levels) are considered crossed if Every combinations of A and
Lecture 11: Nested and Split-Plot Designs Montgomery, Chapter 14 1 Lecture 11 Page 1 Crossed vs Nested Factors Factors A (a levels)and B (b levels) are considered crossed if Every combinations of A and
Ordinary Least Squares Regression
 Ordinary Least Squares Regression Goals for this unit More on notation and terminology OLS scalar versus matrix derivation Some Preliminaries In this class we will be learning to analyze Cross Section
Ordinary Least Squares Regression Goals for this unit More on notation and terminology OLS scalar versus matrix derivation Some Preliminaries In this class we will be learning to analyze Cross Section
Mixed-Models. version 30 October 2011
 Mixed-Models version 30 October 2011 Mixed models Mixed models estimate a vector! of fixed effects and one (or more) vectors u of random effects Both fixed and random effects models always include a vector
Mixed-Models version 30 October 2011 Mixed models Mixed models estimate a vector! of fixed effects and one (or more) vectors u of random effects Both fixed and random effects models always include a vector
Sampling Distributions
 Merlise Clyde Duke University September 8, 2016 Outline Topics Normal Theory Chi-squared Distributions Student t Distributions Readings: Christensen Apendix C, Chapter 1-2 Prostate Example > library(lasso2);
Merlise Clyde Duke University September 8, 2016 Outline Topics Normal Theory Chi-squared Distributions Student t Distributions Readings: Christensen Apendix C, Chapter 1-2 Prostate Example > library(lasso2);
Stat 511 Outline (Spring 2009 Revision of 2008 Version)
 Stat 511 Outline Spring 2009 Revision of 2008 Version Steve Vardeman Iowa State University September 29, 2014 Abstract This outline summarizes the main points of lectures based on Ken Koehler s class notes
Stat 511 Outline Spring 2009 Revision of 2008 Version Steve Vardeman Iowa State University September 29, 2014 Abstract This outline summarizes the main points of lectures based on Ken Koehler s class notes
Confidence Intervals, Testing and ANOVA Summary
 Confidence Intervals, Testing and ANOVA Summary 1 One Sample Tests 1.1 One Sample z test: Mean (σ known) Let X 1,, X n a r.s. from N(µ, σ) or n > 30. Let The test statistic is H 0 : µ = µ 0. z = x µ 0
Confidence Intervals, Testing and ANOVA Summary 1 One Sample Tests 1.1 One Sample z test: Mean (σ known) Let X 1,, X n a r.s. from N(µ, σ) or n > 30. Let The test statistic is H 0 : µ = µ 0. z = x µ 0
1 Introduction to Generalized Least Squares
 ECONOMICS 7344, Spring 2017 Bent E. Sørensen April 12, 2017 1 Introduction to Generalized Least Squares Consider the model Y = Xβ + ɛ, where the N K matrix of regressors X is fixed, independent of the
ECONOMICS 7344, Spring 2017 Bent E. Sørensen April 12, 2017 1 Introduction to Generalized Least Squares Consider the model Y = Xβ + ɛ, where the N K matrix of regressors X is fixed, independent of the
Ch 3: Multiple Linear Regression
 Ch 3: Multiple Linear Regression 1. Multiple Linear Regression Model Multiple regression model has more than one regressor. For example, we have one response variable and two regressor variables: 1. delivery
Ch 3: Multiple Linear Regression 1. Multiple Linear Regression Model Multiple regression model has more than one regressor. For example, we have one response variable and two regressor variables: 1. delivery
Reference: Chapter 14 of Montgomery (8e)
 Reference: Chapter 14 of Montgomery (8e) 99 Maghsoodloo The Stage Nested Designs So far emphasis has been placed on factorial experiments where all factors are crossed (i.e., it is possible to study the
Reference: Chapter 14 of Montgomery (8e) 99 Maghsoodloo The Stage Nested Designs So far emphasis has been placed on factorial experiments where all factors are crossed (i.e., it is possible to study the
Analysis of Variance
 Analysis of Variance Blood coagulation time T avg A 62 60 63 59 61 B 63 67 71 64 65 66 66 C 68 66 71 67 68 68 68 D 56 62 60 61 63 64 63 59 61 64 Blood coagulation time A B C D Combined 56 57 58 59 60 61
Analysis of Variance Blood coagulation time T avg A 62 60 63 59 61 B 63 67 71 64 65 66 66 C 68 66 71 67 68 68 68 D 56 62 60 61 63 64 63 59 61 64 Blood coagulation time A B C D Combined 56 57 58 59 60 61
Lecture 10: Experiments with Random Effects
 Lecture 10: Experiments with Random Effects Montgomery, Chapter 13 1 Lecture 10 Page 1 Example 1 A textile company weaves a fabric on a large number of looms. It would like the looms to be homogeneous
Lecture 10: Experiments with Random Effects Montgomery, Chapter 13 1 Lecture 10 Page 1 Example 1 A textile company weaves a fabric on a large number of looms. It would like the looms to be homogeneous
The Pennsylvania State University The Graduate School Eberly College of Science INTRABLOCK, INTERBLOCK AND COMBINED ESTIMATES
 The Pennsylvania State University The Graduate School Eberly College of Science INTRABLOCK, INTERBLOCK AND COMBINED ESTIMATES IN INCOMPLETE BLOCK DESIGNS: A NUMERICAL STUDY A Thesis in Statistics by Yasin
The Pennsylvania State University The Graduate School Eberly College of Science INTRABLOCK, INTERBLOCK AND COMBINED ESTIMATES IN INCOMPLETE BLOCK DESIGNS: A NUMERICAL STUDY A Thesis in Statistics by Yasin
Lec 5: Factorial Experiment
 November 21, 2011 Example Study of the battery life vs the factors temperatures and types of material. A: Types of material, 3 levels. B: Temperatures, 3 levels. Example Study of the battery life vs the
November 21, 2011 Example Study of the battery life vs the factors temperatures and types of material. A: Types of material, 3 levels. B: Temperatures, 3 levels. Example Study of the battery life vs the
ML and REML Variance Component Estimation. Copyright c 2012 Dan Nettleton (Iowa State University) Statistics / 58
 ML and REML Variance Component Estimation Copyright c 2012 Dan Nettleton (Iowa State University) Statistics 611 1 / 58 Suppose y = Xβ + ε, where ε N(0, Σ) for some positive definite, symmetric matrix Σ.
ML and REML Variance Component Estimation Copyright c 2012 Dan Nettleton (Iowa State University) Statistics 611 1 / 58 Suppose y = Xβ + ε, where ε N(0, Σ) for some positive definite, symmetric matrix Σ.
Stat/F&W Ecol/Hort 572 Review Points Ané, Spring 2010
 1 Linear models Y = Xβ + ɛ with ɛ N (0, σ 2 e) or Y N (Xβ, σ 2 e) where the model matrix X contains the information on predictors and β includes all coefficients (intercept, slope(s) etc.). 1. Number of
1 Linear models Y = Xβ + ɛ with ɛ N (0, σ 2 e) or Y N (Xβ, σ 2 e) where the model matrix X contains the information on predictors and β includes all coefficients (intercept, slope(s) etc.). 1. Number of
Lecture 6: Linear models and Gauss-Markov theorem
 Lecture 6: Linear models and Gauss-Markov theorem Linear model setting Results in simple linear regression can be extended to the following general linear model with independently observed response variables
Lecture 6: Linear models and Gauss-Markov theorem Linear model setting Results in simple linear regression can be extended to the following general linear model with independently observed response variables
Reference: Chapter 13 of Montgomery (8e)
 Reference: Chapter 1 of Montgomery (8e) Maghsoodloo 89 Factorial Experiments with Random Factors So far emphasis has been placed on factorial experiments where all factors are at a, b, c,... fixed levels
Reference: Chapter 1 of Montgomery (8e) Maghsoodloo 89 Factorial Experiments with Random Factors So far emphasis has been placed on factorial experiments where all factors are at a, b, c,... fixed levels
Econometrics Multiple Regression Analysis: Heteroskedasticity
 Econometrics Multiple Regression Analysis: João Valle e Azevedo Faculdade de Economia Universidade Nova de Lisboa Spring Semester João Valle e Azevedo (FEUNL) Econometrics Lisbon, April 2011 1 / 19 Properties
Econometrics Multiple Regression Analysis: João Valle e Azevedo Faculdade de Economia Universidade Nova de Lisboa Spring Semester João Valle e Azevedo (FEUNL) Econometrics Lisbon, April 2011 1 / 19 Properties
STAT 135 Lab 10 Two-Way ANOVA, Randomized Block Design and Friedman s Test
 STAT 135 Lab 10 Two-Way ANOVA, Randomized Block Design and Friedman s Test Rebecca Barter April 13, 2015 Let s now imagine a dataset for which our response variable, Y, may be influenced by two factors,
STAT 135 Lab 10 Two-Way ANOVA, Randomized Block Design and Friedman s Test Rebecca Barter April 13, 2015 Let s now imagine a dataset for which our response variable, Y, may be influenced by two factors,
Lecture 14 Topic 10: ANOVA models for random and mixed effects. Fixed and Random Models in One-way Classification Experiments
 Lecture 14 Topic 10: ANOVA models for random and mixed effects To this point, we have considered only the Fixed Model (Model I) ANOVA; now we will extend the method of ANOVA to other experimental objectives.
Lecture 14 Topic 10: ANOVA models for random and mixed effects To this point, we have considered only the Fixed Model (Model I) ANOVA; now we will extend the method of ANOVA to other experimental objectives.
MA 575 Linear Models: Cedric E. Ginestet, Boston University Midterm Review Week 7
 MA 575 Linear Models: Cedric E. Ginestet, Boston University Midterm Review Week 7 1 Random Vectors Let a 0 and y be n 1 vectors, and let A be an n n matrix. Here, a 0 and A are non-random, whereas y is
MA 575 Linear Models: Cedric E. Ginestet, Boston University Midterm Review Week 7 1 Random Vectors Let a 0 and y be n 1 vectors, and let A be an n n matrix. Here, a 0 and A are non-random, whereas y is
Econometrics I KS. Module 2: Multivariate Linear Regression. Alexander Ahammer. This version: April 16, 2018
 Econometrics I KS Module 2: Multivariate Linear Regression Alexander Ahammer Department of Economics Johannes Kepler University of Linz This version: April 16, 2018 Alexander Ahammer (JKU) Module 2: Multivariate
Econometrics I KS Module 2: Multivariate Linear Regression Alexander Ahammer Department of Economics Johannes Kepler University of Linz This version: April 16, 2018 Alexander Ahammer (JKU) Module 2: Multivariate
Heteroskedasticity. Part VII. Heteroskedasticity
 Part VII Heteroskedasticity As of Oct 15, 2015 1 Heteroskedasticity Consequences Heteroskedasticity-robust inference Testing for Heteroskedasticity Weighted Least Squares (WLS) Feasible generalized Least
Part VII Heteroskedasticity As of Oct 15, 2015 1 Heteroskedasticity Consequences Heteroskedasticity-robust inference Testing for Heteroskedasticity Weighted Least Squares (WLS) Feasible generalized Least
General Linear Test of a General Linear Hypothesis. Copyright c 2012 Dan Nettleton (Iowa State University) Statistics / 35
 General Linear Test of a General Linear Hypothesis Copyright c 2012 Dan Nettleton (Iowa State University) Statistics 611 1 / 35 Suppose the NTGMM holds so that y = Xβ + ε, where ε N(0, σ 2 I). opyright
General Linear Test of a General Linear Hypothesis Copyright c 2012 Dan Nettleton (Iowa State University) Statistics 611 1 / 35 Suppose the NTGMM holds so that y = Xβ + ε, where ε N(0, σ 2 I). opyright
Maximum Likelihood Estimation
 Maximum Likelihood Estimation Merlise Clyde STA721 Linear Models Duke University August 31, 2017 Outline Topics Likelihood Function Projections Maximum Likelihood Estimates Readings: Christensen Chapter
Maximum Likelihood Estimation Merlise Clyde STA721 Linear Models Duke University August 31, 2017 Outline Topics Likelihood Function Projections Maximum Likelihood Estimates Readings: Christensen Chapter
Contents. 1 Review of Residuals. 2 Detecting Outliers. 3 Influential Observations. 4 Multicollinearity and its Effects
 Contents 1 Review of Residuals 2 Detecting Outliers 3 Influential Observations 4 Multicollinearity and its Effects W. Zhou (Colorado State University) STAT 540 July 6th, 2015 1 / 32 Model Diagnostics:
Contents 1 Review of Residuals 2 Detecting Outliers 3 Influential Observations 4 Multicollinearity and its Effects W. Zhou (Colorado State University) STAT 540 July 6th, 2015 1 / 32 Model Diagnostics:
Outline. Example and Model ANOVA table F tests Pairwise treatment comparisons with LSD Sample and subsample size determination
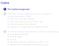 Outline 1 The traditional approach 2 The Mean Squares approach for the Completely randomized design (CRD) CRD and one-way ANOVA Variance components and the F test Inference about the intercept Sample vs.
Outline 1 The traditional approach 2 The Mean Squares approach for the Completely randomized design (CRD) CRD and one-way ANOVA Variance components and the F test Inference about the intercept Sample vs.
MIXED MODELS THE GENERAL MIXED MODEL
 MIXED MODELS This chapter introduces best linear unbiased prediction (BLUP), a general method for predicting random effects, while Chapter 27 is concerned with the estimation of variances by restricted
MIXED MODELS This chapter introduces best linear unbiased prediction (BLUP), a general method for predicting random effects, while Chapter 27 is concerned with the estimation of variances by restricted
Random and Mixed Effects Models - Part III
 Random and Mixed Effects Models - Part III Statistics 149 Spring 2006 Copyright 2006 by Mark E. Irwin Quasi-F Tests When we get to more than two categorical factors, some times there are not nice F tests
Random and Mixed Effects Models - Part III Statistics 149 Spring 2006 Copyright 2006 by Mark E. Irwin Quasi-F Tests When we get to more than two categorical factors, some times there are not nice F tests
Power & Sample Size Calculation
 Chapter 7 Power & Sample Size Calculation Yibi Huang Chapter 7 Section 10.3 Power & Sample Size Calculation for CRDs Power & Sample Size for Factorial Designs Chapter 7-1 Power & Sample Size Calculation
Chapter 7 Power & Sample Size Calculation Yibi Huang Chapter 7 Section 10.3 Power & Sample Size Calculation for CRDs Power & Sample Size for Factorial Designs Chapter 7-1 Power & Sample Size Calculation
Chapter 11 MIVQUE of Variances and Covariances
 Chapter 11 MIVQUE of Variances and Covariances C R Henderson 1984 - Guelph The methods described in Chapter 10 for estimation of variances are quadratic, translation invariant, and unbiased For the balanced
Chapter 11 MIVQUE of Variances and Covariances C R Henderson 1984 - Guelph The methods described in Chapter 10 for estimation of variances are quadratic, translation invariant, and unbiased For the balanced
IDENTIFYING AN APPROPRIATE MODEL
 IDENTIFYING AN APPROPRIATE MODEL Given a description of a study, how do you construct an appropriate model? Context: more than one size of e.u. A made-up example, intended to be complicated (but far from
IDENTIFYING AN APPROPRIATE MODEL Given a description of a study, how do you construct an appropriate model? Context: more than one size of e.u. A made-up example, intended to be complicated (but far from
IDENTIFYING AN APPROPRIATE MODEL
 IDENTIFYING AN APPROPRIATE MODEL Given a description of a study, how do you construct an appropriate model? Context: more than one size of e.u. A made-up example, intended to be complicated (but far from
IDENTIFYING AN APPROPRIATE MODEL Given a description of a study, how do you construct an appropriate model? Context: more than one size of e.u. A made-up example, intended to be complicated (but far from
Spatial inference. Spatial inference. Accounting for spatial correlation. Multivariate normal distributions
 Spatial inference I will start with a simple model, using species diversity data Strong spatial dependence, Î = 0.79 what is the mean diversity? How precise is our estimate? Sampling discussion: The 64
Spatial inference I will start with a simple model, using species diversity data Strong spatial dependence, Î = 0.79 what is the mean diversity? How precise is our estimate? Sampling discussion: The 64
Factorial and Unbalanced Analysis of Variance
 Factorial and Unbalanced Analysis of Variance Nathaniel E. Helwig Assistant Professor of Psychology and Statistics University of Minnesota (Twin Cities) Updated 04-Jan-2017 Nathaniel E. Helwig (U of Minnesota)
Factorial and Unbalanced Analysis of Variance Nathaniel E. Helwig Assistant Professor of Psychology and Statistics University of Minnesota (Twin Cities) Updated 04-Jan-2017 Nathaniel E. Helwig (U of Minnesota)
Simple Linear Regression
 Simple Linear Regression ST 430/514 Recall: A regression model describes how a dependent variable (or response) Y is affected, on average, by one or more independent variables (or factors, or covariates)
Simple Linear Regression ST 430/514 Recall: A regression model describes how a dependent variable (or response) Y is affected, on average, by one or more independent variables (or factors, or covariates)
Linear Regression. In this problem sheet, we consider the problem of linear regression with p predictors and one intercept,
 Linear Regression In this problem sheet, we consider the problem of linear regression with p predictors and one intercept, y = Xβ + ɛ, where y t = (y 1,..., y n ) is the column vector of target values,
Linear Regression In this problem sheet, we consider the problem of linear regression with p predictors and one intercept, y = Xβ + ɛ, where y t = (y 1,..., y n ) is the column vector of target values,
STAT5044: Regression and Anova. Inyoung Kim
 STAT5044: Regression and Anova Inyoung Kim 2 / 51 Outline 1 Matrix Expression 2 Linear and quadratic forms 3 Properties of quadratic form 4 Properties of estimates 5 Distributional properties 3 / 51 Matrix
STAT5044: Regression and Anova Inyoung Kim 2 / 51 Outline 1 Matrix Expression 2 Linear and quadratic forms 3 Properties of quadratic form 4 Properties of estimates 5 Distributional properties 3 / 51 Matrix
Regression with a Single Regressor: Hypothesis Tests and Confidence Intervals
 Regression with a Single Regressor: Hypothesis Tests and Confidence Intervals (SW Chapter 5) Outline. The standard error of ˆ. Hypothesis tests concerning β 3. Confidence intervals for β 4. Regression
Regression with a Single Regressor: Hypothesis Tests and Confidence Intervals (SW Chapter 5) Outline. The standard error of ˆ. Hypothesis tests concerning β 3. Confidence intervals for β 4. Regression
The Random Effects Model Introduction
 The Random Effects Model Introduction Sometimes, treatments included in experiment are randomly chosen from set of all possible treatments. Conclusions from such experiment can then be generalized to other
The Random Effects Model Introduction Sometimes, treatments included in experiment are randomly chosen from set of all possible treatments. Conclusions from such experiment can then be generalized to other
Restricted Maximum Likelihood in Linear Regression and Linear Mixed-Effects Model
 Restricted Maximum Likelihood in Linear Regression and Linear Mixed-Effects Model Xiuming Zhang zhangxiuming@u.nus.edu A*STAR-NUS Clinical Imaging Research Center October, 015 Summary This report derives
Restricted Maximum Likelihood in Linear Regression and Linear Mixed-Effects Model Xiuming Zhang zhangxiuming@u.nus.edu A*STAR-NUS Clinical Imaging Research Center October, 015 Summary This report derives
For more information about how to cite these materials visit
 Author(s): Kerby Shedden, Ph.D., 2010 License: Unless otherwise noted, this material is made available under the terms of the Creative Commons Attribution Share Alike 3.0 License: http://creativecommons.org/licenses/by-sa/3.0/
Author(s): Kerby Shedden, Ph.D., 2010 License: Unless otherwise noted, this material is made available under the terms of the Creative Commons Attribution Share Alike 3.0 License: http://creativecommons.org/licenses/by-sa/3.0/
3. (a) (8 points) There is more than one way to correctly express the null hypothesis in matrix form. One way to state the null hypothesis is
 Stat 501 Solutions and Comments on Exam 1 Spring 005-4 0-4 1. (a) (5 points) Y ~ N, -1-4 34 (b) (5 points) X (X,X ) = (5,8) ~ N ( 11.5, 0.9375 ) 3 1 (c) (10 points, for each part) (i), (ii), and (v) are
Stat 501 Solutions and Comments on Exam 1 Spring 005-4 0-4 1. (a) (5 points) Y ~ N, -1-4 34 (b) (5 points) X (X,X ) = (5,8) ~ N ( 11.5, 0.9375 ) 3 1 (c) (10 points, for each part) (i), (ii), and (v) are
Ph.D. Qualifying Exam Monday Tuesday, January 4 5, 2016
 Ph.D. Qualifying Exam Monday Tuesday, January 4 5, 2016 Put your solution to each problem on a separate sheet of paper. Problem 1. (5106) Find the maximum likelihood estimate of θ where θ is a parameter
Ph.D. Qualifying Exam Monday Tuesday, January 4 5, 2016 Put your solution to each problem on a separate sheet of paper. Problem 1. (5106) Find the maximum likelihood estimate of θ where θ is a parameter
Stat 502 Design and Analysis of Experiments General Linear Model
 1 Stat 502 Design and Analysis of Experiments General Linear Model Fritz Scholz Department of Statistics, University of Washington December 6, 2013 2 General Linear Hypothesis We assume the data vector
1 Stat 502 Design and Analysis of Experiments General Linear Model Fritz Scholz Department of Statistics, University of Washington December 6, 2013 2 General Linear Hypothesis We assume the data vector
Unit 8: 2 k Factorial Designs, Single or Unequal Replications in Factorial Designs, and Incomplete Block Designs
 Unit 8: 2 k Factorial Designs, Single or Unequal Replications in Factorial Designs, and Incomplete Block Designs STA 643: Advanced Experimental Design Derek S. Young 1 Learning Objectives Revisit your
Unit 8: 2 k Factorial Designs, Single or Unequal Replications in Factorial Designs, and Incomplete Block Designs STA 643: Advanced Experimental Design Derek S. Young 1 Learning Objectives Revisit your
18.S096 Problem Set 3 Fall 2013 Regression Analysis Due Date: 10/8/2013
 18.S096 Problem Set 3 Fall 013 Regression Analysis Due Date: 10/8/013 he Projection( Hat ) Matrix and Case Influence/Leverage Recall the setup for a linear regression model y = Xβ + ɛ where y and ɛ are
18.S096 Problem Set 3 Fall 013 Regression Analysis Due Date: 10/8/013 he Projection( Hat ) Matrix and Case Influence/Leverage Recall the setup for a linear regression model y = Xβ + ɛ where y and ɛ are
Mixed models with correlated measurement errors
 Mixed models with correlated measurement errors Rasmus Waagepetersen October 9, 2018 Example from Department of Health Technology 25 subjects where exposed to electric pulses of 11 different durations
Mixed models with correlated measurement errors Rasmus Waagepetersen October 9, 2018 Example from Department of Health Technology 25 subjects where exposed to electric pulses of 11 different durations
COMPLETELY RANDOMIZED DESIGNS (CRD) For now, t unstructured treatments (e.g. no factorial structure)
 STAT 52 Completely Randomized Designs COMPLETELY RANDOMIZED DESIGNS (CRD) For now, t unstructured treatments (e.g. no factorial structure) Completely randomized means no restrictions on the randomization
STAT 52 Completely Randomized Designs COMPLETELY RANDOMIZED DESIGNS (CRD) For now, t unstructured treatments (e.g. no factorial structure) Completely randomized means no restrictions on the randomization
STAT 8200 Design and Analysis of Experiments for Research Workers Lecture Notes
 STAT 8200 Design and Analysis of Experiments for Research Workers Lecture Notes Basics of Experimental Design Terminology Response (Outcome, Dependent) Variable: (y) The variable who s distribution is
STAT 8200 Design and Analysis of Experiments for Research Workers Lecture Notes Basics of Experimental Design Terminology Response (Outcome, Dependent) Variable: (y) The variable who s distribution is
Lecture 4. Random Effects in Completely Randomized Design
 Lecture 4. Random Effects in Completely Randomized Design Montgomery: 3.9, 13.1 and 13.7 1 Lecture 4 Page 1 Random Effects vs Fixed Effects Consider factor with numerous possible levels Want to draw inference
Lecture 4. Random Effects in Completely Randomized Design Montgomery: 3.9, 13.1 and 13.7 1 Lecture 4 Page 1 Random Effects vs Fixed Effects Consider factor with numerous possible levels Want to draw inference
Homework 3 - Solution
 STAT 526 - Spring 2011 Homework 3 - Solution Olga Vitek Each part of the problems 5 points 1. KNNL 25.17 (Note: you can choose either the restricted or the unrestricted version of the model. Please state
STAT 526 - Spring 2011 Homework 3 - Solution Olga Vitek Each part of the problems 5 points 1. KNNL 25.17 (Note: you can choose either the restricted or the unrestricted version of the model. Please state
The Simple Linear Regression Model
 The Simple Linear Regression Model Lesson 3 Ryan Safner 1 1 Department of Economics Hood College ECON 480 - Econometrics Fall 2017 Ryan Safner (Hood College) ECON 480 - Lesson 3 Fall 2017 1 / 77 Bivariate
The Simple Linear Regression Model Lesson 3 Ryan Safner 1 1 Department of Economics Hood College ECON 480 - Econometrics Fall 2017 Ryan Safner (Hood College) ECON 480 - Lesson 3 Fall 2017 1 / 77 Bivariate
Chapter 5 Matrix Approach to Simple Linear Regression
 STAT 525 SPRING 2018 Chapter 5 Matrix Approach to Simple Linear Regression Professor Min Zhang Matrix Collection of elements arranged in rows and columns Elements will be numbers or symbols For example:
STAT 525 SPRING 2018 Chapter 5 Matrix Approach to Simple Linear Regression Professor Min Zhang Matrix Collection of elements arranged in rows and columns Elements will be numbers or symbols For example:
STAT 525 Fall Final exam. Tuesday December 14, 2010
 STAT 525 Fall 2010 Final exam Tuesday December 14, 2010 Time: 2 hours Name (please print): Show all your work and calculations. Partial credit will be given for work that is partially correct. Points will
STAT 525 Fall 2010 Final exam Tuesday December 14, 2010 Time: 2 hours Name (please print): Show all your work and calculations. Partial credit will be given for work that is partially correct. Points will
9 Correlation and Regression
 9 Correlation and Regression SW, Chapter 12. Suppose we select n = 10 persons from the population of college seniors who plan to take the MCAT exam. Each takes the test, is coached, and then retakes the
9 Correlation and Regression SW, Chapter 12. Suppose we select n = 10 persons from the population of college seniors who plan to take the MCAT exam. Each takes the test, is coached, and then retakes the
Lecture 3: Inference in SLR
 Lecture 3: Inference in SLR STAT 51 Spring 011 Background Reading KNNL:.1.6 3-1 Topic Overview This topic will cover: Review of hypothesis testing Inference about 1 Inference about 0 Confidence Intervals
Lecture 3: Inference in SLR STAT 51 Spring 011 Background Reading KNNL:.1.6 3-1 Topic Overview This topic will cover: Review of hypothesis testing Inference about 1 Inference about 0 Confidence Intervals
