How To Use CORREP to Estimate Multivariate Correlation and Statistical Inference Procedures
|
|
|
- Valentine Barber
- 5 years ago
- Views:
Transcription
1 How To Use CORREP to Estimate Multivariate Correlation and Statistical Inference Procedures Dongxiao Zhu June 13, Introduction OMICS data are increasingly available to biomedical researchers, and (biological) replications are more and more affordable for gene microarray experiments or proteomics experiments. The functional relationship between a pair of genes or proteins are often inferred by calculating correlation coefficient between their expression profiles. Classical correlation estimation techniques, such as Pearson correlation coefficient, do not explicitly take replicated data into account. As a result, biological replicates are often averaged before correlations are calculated. The averaging is not justified if there is poor concordance between samples and the variance in each sample is not similar. Based on our recently proposed multivariate correlation estimator, CORREP implements functions for estimating multivariate correlation for replicated OMICS data and statistical inference procedures. In this vignette I demo an non-trivial task accomplished using COR- REP. First let s look at examples of replicated OMICS data and non-replicated OMICS data. x 11, x 12,..., x 1n x 21, x 22,..., x 2n... x m11, x m12,..., x m1n y 11, y 12,..., y 1n y 21, y 22,..., y 2n... versus y m21, y m22,..., y m2n x 1, x 2,..., x n y 1, y 2,..., y n In this toy example, X and Y are a pair of genes or proteins of interest. Using microarray or mass spectrum we were able to profile their expression over n biological conditions such as knockouts, overexpression or SiRNA treatments either with replication (upper example) or without replication (lower example). There are m 1 and m 2 replicates for X and Y respectively. It is often biological 1
2 interest how strong is the correlation between X and Y over n conditions and how significant the correlation is. Significant correlation between X and Y often indicates potential functional relevancy. In addition, 1-correlation are usually used to calculate distance matrix of a number of genes or proteins to perform hierarchical clustering. Therefore, correlation estimation is an important problem in functional genomics research. For non-replicated OMICS data, estimating correlation is relatively trivial since there are plenty of established methods such as Pearson correlation coefficient and its non-parametric alternatives: Spearman s ρ and Kendall s τ. However it is not straightforward to estimate correlation from replicated OMICS data. A naive approach may be to average over replicates to transform the replicated data into non-replicated data. This approach might work for relative clean data in which the within-replicate correlation is relatively high. Unfortunately most OMICS data are noisy reflected partially by poor within-replicate correlation. The averaging for those noisy data is not justified. In order to properly estimate correlation of X and Y from replicated data, we must explicitly model the within-replicate correlation together with the between-replicate correlation. The next section briefly describes the models and methods for deriving the multivariate correlation estimator. For interested reader, please refer to our manuscript for more technical details [Zhu and Li, 2007]. Interested reader also refer to [Medvedovic et al., 2004] for related Bayesian mixture model methods. 2 Methods 2.1 Pearson correlation estimator Instead of averaging, we exploit all the replicated observations by assuming the data are i.i.d. samples from a multivariate normal distribution with a specified correlation matrix and a mean vector, i.e., Z j = (x j1,..., x jp, y j1,..., y jq ) T, j = [ 1, 2,..., n, ] follows a p + q-variate normal distribution N(µ, Σ), where µx e µ = m, e µ y e m = (1,..., 1) T is a m 1 vector, the correlation matrix m Σ is a (p + q) (p + q) matrix with structure: 1... ρ x ρ... ρ Σ = ρ x... 1 ρ... ρ ρ... ρ 1... ρ y = ρ... ρ ρ y... 1 [ ] Σx Σ xy Σ T, (1) xy Σ y where the inter-molecule correlation ρ is the parameter of interest, and the intramolecule correlation ρ x or ρ y are nuisance parameters. The ρ x and ρ y indicate the quality of replicates that high quality replicates tend to have high value, and vice versa. We employ three parameters: ρ, ρ x and ρ y to model the correlation structure of replicated omics data. 2
3 2.2 Multivariate correlation estimator Assuming a multivariate normal model, the Maximum Likelihood Estimate (MLE) of ρ can be derived as follows (see Manuscript for more details): Similarly, ˆµ x = 1 1 n m i=1 m x ij (2) [ ] ˆµx e therefore, ˆµ = m ˆµ y e m The MLE of Σ is ˆµ y = 1 1 n m m y ij (3) i=1 ˆΣ = 1 n (Z j ˆµ)(Z j ˆµ) T (4) To derive the MLE of ρ, the ideal method would be obtaining the likelihood explicitly as a function of ρ. However, this proved to be intractable in practice (see manscript for detailed discussion). Our approach is to use the average of the elements of ˆΣ xy estimated via Eq. 4: ˆρ = Avg(ˆΣ xy ) (5) The sample Pearson correlation coefficient can be also written into the following form: cor(x, Y ) = ( x j x)(ȳ j ȳ), (6) (n 1)S X S Y where S X and S Y are standard deviations of X and Y respectively. When there is no replicate (m 1 = m 2 = 1), the correlation matrix Σ is reduced to a 2 by 2 matrix with diagonal elements equal to 1 and off-diagonal elements equal to ρ. It is easy to show from Eq. 4 that ˆρ = ( x j x)(ȳ j ȳ) (7) ns X S Y Hence we derive the connection between the two estimators when there is no replicate as follows: ˆρ = n 1 cor. (8) n 2.3 Statistical inference procedures For very small sample data, eg, n < 4, we recommend using all permutations to approximate the null distribution (see Manuscript for detail). For larger sample data, we provide a Likelihood Ratio (LR) test. For moderate to large sample data, we provide a LRT procedure for testing the hypothesis that the 3
4 multivariate correlation ρ vanishes. Under the multivariate normal distribution assumption, Z j N(µ, Σ), and we test the following hypothesis: H 0 : Z N(µ, Σ 0 ) versus H α : Z N(µ, Σ 1 ). (9) Here, both Σ 0 and Σ 1 are (m 1 + m 2 ) (m 1 + m 2 ) matrices, where ( m 1 and m) 2 Σx 0 are number of replicates for biomolecule X and Y and Σ 0 = m1 0 T, m 2 Σ ( ) y Σx Σ Σ 1 = xy Σ T, where Σ xy Σ x and Σ y, with diagonal elements identity and y all the other entries being ρ x and ρ y respectively. 0 m1 is a m 1 m 1 zero matrix and 0 m2 is a m 2 m 2 zero matrix, that is, under the null hypothesis, the intermolecule correlation ρ vanishes. Σ xy is a m 1 m 1 matrix with all entries equal to ρ. Likewise Σ T xy is a m 2 m 2 matrix with all entries equal to ρ. The Likelihood Ratio (LR) statistic for testing the two different correlation structures can be derived as follows: = ˆΣ 0 n/2 e 1 2 ˆΣ 1 n/2 e 1 2 (Zj ˆµ) (ˆΣ 0) 1 (Z j ˆµ) (Zj ˆµ) (ˆΣ 1) 1 (Z j ˆµ). (10) Note that for the test to be a true LRT, all the estimated quantities ( ) ˆ in the above formula should be MLE s. In Section 2.2, Eqs. 2, 3 and 4 give the formula of MLE s of the mean vector and the correlation matrix under H( α. The MLE ) of the correlation matrix under H 0 can be determined as ˆΣ ˆΣx O 0 = O ˆΣ, y where ˆΣ x = 1 n ˆΣ y = 1 n (X j ˆµ x )(X j ˆµ x ), (11) (Y j ˆµ y )(Y j ˆµ y ). (12) The LR statistic, denoted by G 2, is therefore: G 2 = 2log = n[trm log M (m 1 + m 2 )], (13) where M = (ˆΣ 0 ) 1 ˆΣ1. The LR statistic is asymptotically chi-square distributed with 2(m 1 m 2 ) degrees of freedom under H 0. 3 Data Analysis Examples In this example, we will analyze a subset of 205 genes whose expression were profiled using 4 replicates under 20 physiological/genetic conditions [Medvedovic et al., 2004]. The whole data was initially reported in [Ideker et al., 2000] We first estimate all pairwise correlation and then we use 1-correlation as distance measure to cluster these genes. Medvedovic et al, 2004 [Medvedovic et al., 2004] has already classified the 205 genes into 4 functional groups according to their GO annotations. The four classes are: 4
5 ˆ Biosynthesis; protein metabolism and modification ˆ Energy pathways; carbohydrate metabolism; catabolism ˆ Nucleobase, nucleoside, nucleotide and nucleic acid metabolism ˆ Transport The membership of 205 genes were stored in the internal data true.member. We were able to compare the performance of our multivariate correlation estimator versus Pearson correlation estimator through hierarchical clustering by comparing clustering results to the external knowledge above. > library(correp) > data(gal_all) > ##Take average > gal_avg <- apply(gal_all, 1, function(x) c(mean(x[1:4]), mean(x[5:8]), mean(x[9:12]), mea + mean(x[49:52]), mean(x[53:56]), mean(x[57:60]), mean(x[61:64]), mean(x[65:68]), mean(x[69 > ##Estimate routine correlation based on average profile > M1 <- cor(gal_avg) The above code is to calculate a 205 by 205 correlation matrix using Pearson correlation coefficient implemented in R function cor. Note that we have to average over replicates before the straight-forward application of Pearson correlation coefficient can be done. As we mentioned in the paper, the data used in this example is relatively clean data (see the boxplot below for distribution of within-replicate correlation), which is not favorable condition to apply our estimator, however, the following clustering results show that it still significantly outperforms Pearson correlation coefficient. > ##Take the first row and format > x <- gal_all[1,] > x <- cbind(t(x[1:4]),t(x[5:8]),t(x[9:12]),t(x[13:16]),t(x[17:20]),t(x[21:24]),t(x[25:28]) > ##Loop through to format each row > for(j in 2:205) + { + y <- gal_all[j,] + y <- cbind(t(y[1:4]),t(y[5:8]),t(y[9:12]),t(y[13:16]),t(y[17:20]),t(y[21:24]),t(y[25:28]) + x <- rbind(x,y) + } > boxplot(cor(x)) > rawdata <- x > stddata <- apply(rawdata, 1, function(x) x/sd(x)) > stddata <- t(stddata) The above code is to reshape the data to be compatible with functions implemented in this package. This step is not absolutely necessary if your data is already in the right format, for example, columns correspond to conditions and rows correspond to (replicated) variables. For example, a 820 by 20 matrix for this data. Moreover, data has to be standardized by making variance of each row (gene) equals to 1. The standardization is VERY important and must be followed. 5
6 > M <- cor.balance(stddata, m = 4, G= 205) The above code is to calculate 205 by 205 correlation matrix using the new multivariate correlation estimator implemented in R function cor.balance. Note that there are equivalent number of replicates for this data (balanced). For unbalanced data, R function cor.unbalance should be used. > row.names(m) <- row.names(m1) > colnames(m) <- colnames(m1) We next use 1-correlation as distance measure to cluster the genes into 4 groups, and compare the consistency between these 4 groups and pre-defined 4 groups. > M.rep <- 1-M > M.avg <- 1-M1 > d.rep <- as.dist(m.rep) > d.avg <- as.dist(m.avg) The above code calculates distance matrix for hierarchical clustering using both Pearson correlation coefficient and multivariate correlation estimator. > library(e1071) > library(cluster) > data(true.member) > g.rep <- diana(d.rep) > g.avg <- diana(d.avg) The above code does hierarchical clustering in a top-down fashion ie, Diana, and classifies all 205 genes into a number of clusters. We recommend using top-down method in this case since we are interested in identifying a few (4) large clusters. For other data, if you are interested in identifying a lot of small clusters, we recommend using agglomerative hierarchical clustering methods. See below for examples. > h.rep <- hclust(d.rep, method = "complete") > h.avg <- hclust(d.avg, method = "complete") The R function hclust can be applied to perform bottom-up hierarchical clustering. In addition to choosing a proper distance measure, we will need to determine a method to measure distance between objects such as are complete, average, single. Option average seems to be robust in most of cases. > member.rep.k4 <- cutree(g.rep, k = 4) > member.avg.k4 <- cutree(g.avg, k = 4) > classagreement(table(member.avg.k4, as.matrix(true.member))) $diag [1] 0 $kappa [1] $rand [1]
7 $crand [1] > classagreement(table(member.rep.k4, as.matrix(true.member))) $diag [1] $kappa [1] $rand [1] $crand [1] The above code retrieves the membership of 205 genes as determined by divisive clustering, and accesses consistency of the memberships with the true cluster membership (external knowledge) using adjusted RAND index. The function classagreement that was used to calculate adjusted RAND index is from e1071 package. It is obvious that clustering results based on multivariate correlation estimator are more consistent to the external knowledge (higher adjusted RAND index). We can also examine the quality of clusters using, for example, silhouette plot. Here quality roughly means that ratio between within-cluster deviation and between-cluster deviation. Although it is statistically true that good quality clusters indicate better clustering performance, it is not necessarily true biologically. Good quality clusters are only a sufficient condition for good clustering performance but not a necessary condition. > sil.rep4 <- silhouette(member.rep.k4, d.rep) > sil.avg4 <- silhouette(member.avg.k4, d.avg) > plot(sil.rep4, nmax = 80, cex.names = 0.5) > plot(sil.avg4, nmax = 80, cex.names = 0.5) References [Medvedovic et al., 2004] Medvedovic, M., Yeung, K.Y., Bumgarner, R.E. (2004) Bayesian mixtures for clustering replicated microarray data. Bioinformatics, 20, [Ideker et al., 2000] Ideker, T., Thorsson, V., Siegel, A.F. and Hood, L.E Testing for differentially-expressed genes by maximum-likelihood analysis of microarray data. J. Comput. Biol., 7, [Zhu and Li, 2007] Zhu, D and Li, Y Multivariate Correlation Estimator for Inferring Functional Relationships from Replicated OMICS Data. Submitted. 7
Bivariate Relationships Between Variables
 Bivariate Relationships Between Variables BUS 735: Business Decision Making and Research 1 Goals Specific goals: Detect relationships between variables. Be able to prescribe appropriate statistical methods
Bivariate Relationships Between Variables BUS 735: Business Decision Making and Research 1 Goals Specific goals: Detect relationships between variables. Be able to prescribe appropriate statistical methods
Spatial inference. Spatial inference. Accounting for spatial correlation. Multivariate normal distributions
 Spatial inference I will start with a simple model, using species diversity data Strong spatial dependence, Î = 0.79 what is the mean diversity? How precise is our estimate? Sampling discussion: The 64
Spatial inference I will start with a simple model, using species diversity data Strong spatial dependence, Î = 0.79 what is the mean diversity? How precise is our estimate? Sampling discussion: The 64
Unsupervised machine learning
 Chapter 9 Unsupervised machine learning Unsupervised machine learning (a.k.a. cluster analysis) is a set of methods to assign objects into clusters under a predefined distance measure when class labels
Chapter 9 Unsupervised machine learning Unsupervised machine learning (a.k.a. cluster analysis) is a set of methods to assign objects into clusters under a predefined distance measure when class labels
Hypothesis Testing for Var-Cov Components
 Hypothesis Testing for Var-Cov Components When the specification of coefficients as fixed, random or non-randomly varying is considered, a null hypothesis of the form is considered, where Additional output
Hypothesis Testing for Var-Cov Components When the specification of coefficients as fixed, random or non-randomly varying is considered, a null hypothesis of the form is considered, where Additional output
Optimal exact tests for complex alternative hypotheses on cross tabulated data
 Optimal exact tests for complex alternative hypotheses on cross tabulated data Daniel Yekutieli Statistics and OR Tel Aviv University CDA course 29 July 2017 Yekutieli (TAU) Optimal exact tests for complex
Optimal exact tests for complex alternative hypotheses on cross tabulated data Daniel Yekutieli Statistics and OR Tel Aviv University CDA course 29 July 2017 Yekutieli (TAU) Optimal exact tests for complex
Statistical Inference: Estimation and Confidence Intervals Hypothesis Testing
 Statistical Inference: Estimation and Confidence Intervals Hypothesis Testing 1 In most statistics problems, we assume that the data have been generated from some unknown probability distribution. We desire
Statistical Inference: Estimation and Confidence Intervals Hypothesis Testing 1 In most statistics problems, we assume that the data have been generated from some unknown probability distribution. We desire
Statistical Inference On the High-dimensional Gaussian Covarianc
 Statistical Inference On the High-dimensional Gaussian Covariance Matrix Department of Mathematical Sciences, Clemson University June 6, 2011 Outline Introduction Problem Setup Statistical Inference High-Dimensional
Statistical Inference On the High-dimensional Gaussian Covariance Matrix Department of Mathematical Sciences, Clemson University June 6, 2011 Outline Introduction Problem Setup Statistical Inference High-Dimensional
University of Cambridge. MPhil in Computer Speech Text & Internet Technology. Module: Speech Processing II. Lecture 2: Hidden Markov Models I
 University of Cambridge MPhil in Computer Speech Text & Internet Technology Module: Speech Processing II Lecture 2: Hidden Markov Models I o o o o o 1 2 3 4 T 1 b 2 () a 12 2 a 3 a 4 5 34 a 23 b () b ()
University of Cambridge MPhil in Computer Speech Text & Internet Technology Module: Speech Processing II Lecture 2: Hidden Markov Models I o o o o o 1 2 3 4 T 1 b 2 () a 12 2 a 3 a 4 5 34 a 23 b () b ()
BMD645. Integration of Omics
 BMD645 Integration of Omics Shu-Jen Chen, Chang Gung University Dec. 11, 2009 1 Traditional Biology vs. Systems Biology Traditional biology : Single genes or proteins Systems biology: Simultaneously study
BMD645 Integration of Omics Shu-Jen Chen, Chang Gung University Dec. 11, 2009 1 Traditional Biology vs. Systems Biology Traditional biology : Single genes or proteins Systems biology: Simultaneously study
Lecture 15. Hypothesis testing in the linear model
 14. Lecture 15. Hypothesis testing in the linear model Lecture 15. Hypothesis testing in the linear model 1 (1 1) Preliminary lemma 15. Hypothesis testing in the linear model 15.1. Preliminary lemma Lemma
14. Lecture 15. Hypothesis testing in the linear model Lecture 15. Hypothesis testing in the linear model 1 (1 1) Preliminary lemma 15. Hypothesis testing in the linear model 15.1. Preliminary lemma Lemma
MISCELLANEOUS TOPICS RELATED TO LIKELIHOOD. Copyright c 2012 (Iowa State University) Statistics / 30
 MISCELLANEOUS TOPICS RELATED TO LIKELIHOOD Copyright c 2012 (Iowa State University) Statistics 511 1 / 30 INFORMATION CRITERIA Akaike s Information criterion is given by AIC = 2l(ˆθ) + 2k, where l(ˆθ)
MISCELLANEOUS TOPICS RELATED TO LIKELIHOOD Copyright c 2012 (Iowa State University) Statistics 511 1 / 30 INFORMATION CRITERIA Akaike s Information criterion is given by AIC = 2l(ˆθ) + 2k, where l(ˆθ)
Matrices and vectors A matrix is a rectangular array of numbers. Here s an example: A =
 Matrices and vectors A matrix is a rectangular array of numbers Here s an example: 23 14 17 A = 225 0 2 This matrix has dimensions 2 3 The number of rows is first, then the number of columns We can write
Matrices and vectors A matrix is a rectangular array of numbers Here s an example: 23 14 17 A = 225 0 2 This matrix has dimensions 2 3 The number of rows is first, then the number of columns We can write
Bivariate Paired Numerical Data
 Bivariate Paired Numerical Data Pearson s correlation, Spearman s ρ and Kendall s τ, tests of independence University of California, San Diego Instructor: Ery Arias-Castro http://math.ucsd.edu/~eariasca/teaching.html
Bivariate Paired Numerical Data Pearson s correlation, Spearman s ρ and Kendall s τ, tests of independence University of California, San Diego Instructor: Ery Arias-Castro http://math.ucsd.edu/~eariasca/teaching.html
Overview. and data transformations of gene expression data. Toy 2-d Clustering Example. K-Means. Motivation. Model-based clustering
 Model-based clustering and data transformations of gene expression data Walter L. Ruzzo University of Washington UW CSE Computational Biology Group 2 Toy 2-d Clustering Example K-Means? 3 4 Hierarchical
Model-based clustering and data transformations of gene expression data Walter L. Ruzzo University of Washington UW CSE Computational Biology Group 2 Toy 2-d Clustering Example K-Means? 3 4 Hierarchical
Lecture 5: Clustering, Linear Regression
 Lecture 5: Clustering, Linear Regression Reading: Chapter 10, Sections 3.1-3.2 STATS 202: Data mining and analysis October 4, 2017 1 / 22 .0.0 5 5 1.0 7 5 X2 X2 7 1.5 1.0 0.5 3 1 2 Hierarchical clustering
Lecture 5: Clustering, Linear Regression Reading: Chapter 10, Sections 3.1-3.2 STATS 202: Data mining and analysis October 4, 2017 1 / 22 .0.0 5 5 1.0 7 5 X2 X2 7 1.5 1.0 0.5 3 1 2 Hierarchical clustering
Parametric Empirical Bayes Methods for Microarrays
 Parametric Empirical Bayes Methods for Microarrays Ming Yuan, Deepayan Sarkar, Michael Newton and Christina Kendziorski April 30, 2018 Contents 1 Introduction 1 2 General Model Structure: Two Conditions
Parametric Empirical Bayes Methods for Microarrays Ming Yuan, Deepayan Sarkar, Michael Newton and Christina Kendziorski April 30, 2018 Contents 1 Introduction 1 2 General Model Structure: Two Conditions
10-810: Advanced Algorithms and Models for Computational Biology. Optimal leaf ordering and classification
 10-810: Advanced Algorithms and Models for Computational Biology Optimal leaf ordering and classification Hierarchical clustering As we mentioned, its one of the most popular methods for clustering gene
10-810: Advanced Algorithms and Models for Computational Biology Optimal leaf ordering and classification Hierarchical clustering As we mentioned, its one of the most popular methods for clustering gene
Lecture 5: Clustering, Linear Regression
 Lecture 5: Clustering, Linear Regression Reading: Chapter 10, Sections 3.1-3.2 STATS 202: Data mining and analysis October 4, 2017 1 / 22 Hierarchical clustering Most algorithms for hierarchical clustering
Lecture 5: Clustering, Linear Regression Reading: Chapter 10, Sections 3.1-3.2 STATS 202: Data mining and analysis October 4, 2017 1 / 22 Hierarchical clustering Most algorithms for hierarchical clustering
COM336: Neural Computing
 COM336: Neural Computing http://www.dcs.shef.ac.uk/ sjr/com336/ Lecture 2: Density Estimation Steve Renals Department of Computer Science University of Sheffield Sheffield S1 4DP UK email: s.renals@dcs.shef.ac.uk
COM336: Neural Computing http://www.dcs.shef.ac.uk/ sjr/com336/ Lecture 2: Density Estimation Steve Renals Department of Computer Science University of Sheffield Sheffield S1 4DP UK email: s.renals@dcs.shef.ac.uk
Fall 2017 STAT 532 Homework Peter Hoff. 1. Let P be a probability measure on a collection of sets A.
 1. Let P be a probability measure on a collection of sets A. (a) For each n N, let H n be a set in A such that H n H n+1. Show that P (H n ) monotonically converges to P ( k=1 H k) as n. (b) For each n
1. Let P be a probability measure on a collection of sets A. (a) For each n N, let H n be a set in A such that H n H n+1. Show that P (H n ) monotonically converges to P ( k=1 H k) as n. (b) For each n
DEGseq: an R package for identifying differentially expressed genes from RNA-seq data
 DEGseq: an R package for identifying differentially expressed genes from RNA-seq data Likun Wang Zhixing Feng i Wang iaowo Wang * and uegong Zhang * MOE Key Laboratory of Bioinformatics and Bioinformatics
DEGseq: an R package for identifying differentially expressed genes from RNA-seq data Likun Wang Zhixing Feng i Wang iaowo Wang * and uegong Zhang * MOE Key Laboratory of Bioinformatics and Bioinformatics
Simultaneous Inference for Multiple Testing and Clustering via Dirichlet Process Mixture Models
 Simultaneous Inference for Multiple Testing and Clustering via Dirichlet Process Mixture Models David B. Dahl Department of Statistics Texas A&M University Marina Vannucci, Michael Newton, & Qianxing Mo
Simultaneous Inference for Multiple Testing and Clustering via Dirichlet Process Mixture Models David B. Dahl Department of Statistics Texas A&M University Marina Vannucci, Michael Newton, & Qianxing Mo
Discovering molecular pathways from protein interaction and ge
 Discovering molecular pathways from protein interaction and gene expression data 9-4-2008 Aim To have a mechanism for inferring pathways from gene expression and protein interaction data. Motivation Why
Discovering molecular pathways from protein interaction and gene expression data 9-4-2008 Aim To have a mechanism for inferring pathways from gene expression and protein interaction data. Motivation Why
Composite Hypotheses and Generalized Likelihood Ratio Tests
 Composite Hypotheses and Generalized Likelihood Ratio Tests Rebecca Willett, 06 In many real world problems, it is difficult to precisely specify probability distributions. Our models for data may involve
Composite Hypotheses and Generalized Likelihood Ratio Tests Rebecca Willett, 06 In many real world problems, it is difficult to precisely specify probability distributions. Our models for data may involve
the long tau-path for detecting monotone association in an unspecified subpopulation
 the long tau-path for detecting monotone association in an unspecified subpopulation Joe Verducci Current Challenges in Statistical Learning Workshop Banff International Research Station Tuesday, December
the long tau-path for detecting monotone association in an unspecified subpopulation Joe Verducci Current Challenges in Statistical Learning Workshop Banff International Research Station Tuesday, December
UNIT 4 RANK CORRELATION (Rho AND KENDALL RANK CORRELATION
 UNIT 4 RANK CORRELATION (Rho AND KENDALL RANK CORRELATION Structure 4.0 Introduction 4.1 Objectives 4. Rank-Order s 4..1 Rank-order data 4.. Assumptions Underlying Pearson s r are Not Satisfied 4.3 Spearman
UNIT 4 RANK CORRELATION (Rho AND KENDALL RANK CORRELATION Structure 4.0 Introduction 4.1 Objectives 4. Rank-Order s 4..1 Rank-order data 4.. Assumptions Underlying Pearson s r are Not Satisfied 4.3 Spearman
Data Mining Chapter 4: Data Analysis and Uncertainty Fall 2011 Ming Li Department of Computer Science and Technology Nanjing University
 Data Mining Chapter 4: Data Analysis and Uncertainty Fall 2011 Ming Li Department of Computer Science and Technology Nanjing University Why uncertainty? Why should data mining care about uncertainty? We
Data Mining Chapter 4: Data Analysis and Uncertainty Fall 2011 Ming Li Department of Computer Science and Technology Nanjing University Why uncertainty? Why should data mining care about uncertainty? We
Comparative Network Analysis
 Comparative Network Analysis BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2016 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under CC BY-NC 4.0 by
Comparative Network Analysis BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2016 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under CC BY-NC 4.0 by
Canonical Correlation Analysis of Longitudinal Data
 Biometrics Section JSM 2008 Canonical Correlation Analysis of Longitudinal Data Jayesh Srivastava Dayanand N Naik Abstract Studying the relationship between two sets of variables is an important multivariate
Biometrics Section JSM 2008 Canonical Correlation Analysis of Longitudinal Data Jayesh Srivastava Dayanand N Naik Abstract Studying the relationship between two sets of variables is an important multivariate
I i=1 1 I(J 1) j=1 (Y ij Ȳi ) 2. j=1 (Y j Ȳ )2 ] = 2n( is the two-sample t-test statistic.
![I i=1 1 I(J 1) j=1 (Y ij Ȳi ) 2. j=1 (Y j Ȳ )2 ] = 2n( is the two-sample t-test statistic. I i=1 1 I(J 1) j=1 (Y ij Ȳi ) 2. j=1 (Y j Ȳ )2 ] = 2n( is the two-sample t-test statistic.](/thumbs/85/92729432.jpg) Serik Sagitov, Chalmers and GU, February, 08 Solutions chapter Matlab commands: x = data matrix boxplot(x) anova(x) anova(x) Problem.3 Consider one-way ANOVA test statistic For I = and = n, put F = MS
Serik Sagitov, Chalmers and GU, February, 08 Solutions chapter Matlab commands: x = data matrix boxplot(x) anova(x) anova(x) Problem.3 Consider one-way ANOVA test statistic For I = and = n, put F = MS
A Practical Approach to Inferring Large Graphical Models from Sparse Microarray Data
 A Practical Approach to Inferring Large Graphical Models from Sparse Microarray Data Juliane Schäfer Department of Statistics, University of Munich Workshop: Practical Analysis of Gene Expression Data
A Practical Approach to Inferring Large Graphical Models from Sparse Microarray Data Juliane Schäfer Department of Statistics, University of Munich Workshop: Practical Analysis of Gene Expression Data
Introduction to Normal Distribution
 Introduction to Normal Distribution Nathaniel E. Helwig Assistant Professor of Psychology and Statistics University of Minnesota (Twin Cities) Updated 17-Jan-2017 Nathaniel E. Helwig (U of Minnesota) Introduction
Introduction to Normal Distribution Nathaniel E. Helwig Assistant Professor of Psychology and Statistics University of Minnesota (Twin Cities) Updated 17-Jan-2017 Nathaniel E. Helwig (U of Minnesota) Introduction
Bootstrap Procedures for Testing Homogeneity Hypotheses
 Journal of Statistical Theory and Applications Volume 11, Number 2, 2012, pp. 183-195 ISSN 1538-7887 Bootstrap Procedures for Testing Homogeneity Hypotheses Bimal Sinha 1, Arvind Shah 2, Dihua Xu 1, Jianxin
Journal of Statistical Theory and Applications Volume 11, Number 2, 2012, pp. 183-195 ISSN 1538-7887 Bootstrap Procedures for Testing Homogeneity Hypotheses Bimal Sinha 1, Arvind Shah 2, Dihua Xu 1, Jianxin
Extending the Robust Means Modeling Framework. Alyssa Counsell, Phil Chalmers, Matt Sigal, Rob Cribbie
 Extending the Robust Means Modeling Framework Alyssa Counsell, Phil Chalmers, Matt Sigal, Rob Cribbie One-way Independent Subjects Design Model: Y ij = µ + τ j + ε ij, j = 1,, J Y ij = score of the ith
Extending the Robust Means Modeling Framework Alyssa Counsell, Phil Chalmers, Matt Sigal, Rob Cribbie One-way Independent Subjects Design Model: Y ij = µ + τ j + ε ij, j = 1,, J Y ij = score of the ith
4.2 Estimation on the boundary of the parameter space
 Chapter 4 Non-standard inference As we mentioned in Chapter the the log-likelihood ratio statistic is useful in the context of statistical testing because typically it is pivotal (does not depend on any
Chapter 4 Non-standard inference As we mentioned in Chapter the the log-likelihood ratio statistic is useful in the context of statistical testing because typically it is pivotal (does not depend on any
Minimum Hellinger Distance Estimation in a. Semiparametric Mixture Model
 Minimum Hellinger Distance Estimation in a Semiparametric Mixture Model Sijia Xiang 1, Weixin Yao 1, and Jingjing Wu 2 1 Department of Statistics, Kansas State University, Manhattan, Kansas, USA 66506-0802.
Minimum Hellinger Distance Estimation in a Semiparametric Mixture Model Sijia Xiang 1, Weixin Yao 1, and Jingjing Wu 2 1 Department of Statistics, Kansas State University, Manhattan, Kansas, USA 66506-0802.
Simultaneous inference for multiple testing and clustering via a Dirichlet process mixture model
 Simultaneous inference for multiple testing and clustering via a Dirichlet process mixture model David B Dahl 1, Qianxing Mo 2 and Marina Vannucci 3 1 Texas A&M University, US 2 Memorial Sloan-Kettering
Simultaneous inference for multiple testing and clustering via a Dirichlet process mixture model David B Dahl 1, Qianxing Mo 2 and Marina Vannucci 3 1 Texas A&M University, US 2 Memorial Sloan-Kettering
BTRY 4830/6830: Quantitative Genomics and Genetics
 BTRY 4830/6830: Quantitative Genomics and Genetics Lecture 23: Alternative tests in GWAS / (Brief) Introduction to Bayesian Inference Jason Mezey jgm45@cornell.edu Nov. 13, 2014 (Th) 8:40-9:55 Announcements
BTRY 4830/6830: Quantitative Genomics and Genetics Lecture 23: Alternative tests in GWAS / (Brief) Introduction to Bayesian Inference Jason Mezey jgm45@cornell.edu Nov. 13, 2014 (Th) 8:40-9:55 Announcements
Mixture models for analysing transcriptome and ChIP-chip data
 Mixture models for analysing transcriptome and ChIP-chip data Marie-Laure Martin-Magniette French National Institute for agricultural research (INRA) Unit of Applied Mathematics and Informatics at AgroParisTech,
Mixture models for analysing transcriptome and ChIP-chip data Marie-Laure Martin-Magniette French National Institute for agricultural research (INRA) Unit of Applied Mathematics and Informatics at AgroParisTech,
Summary of Chapter 7 (Sections ) and Chapter 8 (Section 8.1)
 Summary of Chapter 7 (Sections 7.2-7.5) and Chapter 8 (Section 8.1) Chapter 7. Tests of Statistical Hypotheses 7.2. Tests about One Mean (1) Test about One Mean Case 1: σ is known. Assume that X N(µ, σ
Summary of Chapter 7 (Sections 7.2-7.5) and Chapter 8 (Section 8.1) Chapter 7. Tests of Statistical Hypotheses 7.2. Tests about One Mean (1) Test about One Mean Case 1: σ is known. Assume that X N(µ, σ
STAT 730 Chapter 5: Hypothesis Testing
 STAT 730 Chapter 5: Hypothesis Testing Timothy Hanson Department of Statistics, University of South Carolina Stat 730: Multivariate Analysis 1 / 28 Likelihood ratio test def n: Data X depend on θ. The
STAT 730 Chapter 5: Hypothesis Testing Timothy Hanson Department of Statistics, University of South Carolina Stat 730: Multivariate Analysis 1 / 28 Likelihood ratio test def n: Data X depend on θ. The
Lecture 5: Clustering, Linear Regression
 Lecture 5: Clustering, Linear Regression Reading: Chapter 10, Sections 3.1-2 STATS 202: Data mining and analysis Sergio Bacallado September 19, 2018 1 / 23 Announcements Starting next week, Julia Fukuyama
Lecture 5: Clustering, Linear Regression Reading: Chapter 10, Sections 3.1-2 STATS 202: Data mining and analysis Sergio Bacallado September 19, 2018 1 / 23 Announcements Starting next week, Julia Fukuyama
Confidence Intervals, Testing and ANOVA Summary
 Confidence Intervals, Testing and ANOVA Summary 1 One Sample Tests 1.1 One Sample z test: Mean (σ known) Let X 1,, X n a r.s. from N(µ, σ) or n > 30. Let The test statistic is H 0 : µ = µ 0. z = x µ 0
Confidence Intervals, Testing and ANOVA Summary 1 One Sample Tests 1.1 One Sample z test: Mean (σ known) Let X 1,, X n a r.s. from N(µ, σ) or n > 30. Let The test statistic is H 0 : µ = µ 0. z = x µ 0
Probability and Statistics Notes
 Probability and Statistics Notes Chapter Seven Jesse Crawford Department of Mathematics Tarleton State University Spring 2011 (Tarleton State University) Chapter Seven Notes Spring 2011 1 / 42 Outline
Probability and Statistics Notes Chapter Seven Jesse Crawford Department of Mathematics Tarleton State University Spring 2011 (Tarleton State University) Chapter Seven Notes Spring 2011 1 / 42 Outline
STAT 135 Lab 5 Bootstrapping and Hypothesis Testing
 STAT 135 Lab 5 Bootstrapping and Hypothesis Testing Rebecca Barter March 2, 2015 The Bootstrap Bootstrap Suppose that we are interested in estimating a parameter θ from some population with members x 1,...,
STAT 135 Lab 5 Bootstrapping and Hypothesis Testing Rebecca Barter March 2, 2015 The Bootstrap Bootstrap Suppose that we are interested in estimating a parameter θ from some population with members x 1,...,
Sparse Covariance Selection using Semidefinite Programming
 Sparse Covariance Selection using Semidefinite Programming A. d Aspremont ORFE, Princeton University Joint work with O. Banerjee, L. El Ghaoui & G. Natsoulis, U.C. Berkeley & Iconix Pharmaceuticals Support
Sparse Covariance Selection using Semidefinite Programming A. d Aspremont ORFE, Princeton University Joint work with O. Banerjee, L. El Ghaoui & G. Natsoulis, U.C. Berkeley & Iconix Pharmaceuticals Support
Correlation and the Analysis of Variance Approach to Simple Linear Regression
 Correlation and the Analysis of Variance Approach to Simple Linear Regression Biometry 755 Spring 2009 Correlation and the Analysis of Variance Approach to Simple Linear Regression p. 1/35 Correlation
Correlation and the Analysis of Variance Approach to Simple Linear Regression Biometry 755 Spring 2009 Correlation and the Analysis of Variance Approach to Simple Linear Regression p. 1/35 Correlation
O 3 O 4 O 5. q 3. q 4. Transition
 Hidden Markov Models Hidden Markov models (HMM) were developed in the early part of the 1970 s and at that time mostly applied in the area of computerized speech recognition. They are first described in
Hidden Markov Models Hidden Markov models (HMM) were developed in the early part of the 1970 s and at that time mostly applied in the area of computerized speech recognition. They are first described in
Package SHIP. February 19, 2015
 Type Package Package SHIP February 19, 2015 Title SHrinkage covariance Incorporating Prior knowledge Version 1.0.2 Author Maintainer Vincent Guillemot The SHIP-package allows
Type Package Package SHIP February 19, 2015 Title SHrinkage covariance Incorporating Prior knowledge Version 1.0.2 Author Maintainer Vincent Guillemot The SHIP-package allows
1 Data Arrays and Decompositions
 1 Data Arrays and Decompositions 1.1 Variance Matrices and Eigenstructure Consider a p p positive definite and symmetric matrix V - a model parameter or a sample variance matrix. The eigenstructure is
1 Data Arrays and Decompositions 1.1 Variance Matrices and Eigenstructure Consider a p p positive definite and symmetric matrix V - a model parameter or a sample variance matrix. The eigenstructure is
MA 575 Linear Models: Cedric E. Ginestet, Boston University Mixed Effects Estimation, Residuals Diagnostics Week 11, Lecture 1
 MA 575 Linear Models: Cedric E Ginestet, Boston University Mixed Effects Estimation, Residuals Diagnostics Week 11, Lecture 1 1 Within-group Correlation Let us recall the simple two-level hierarchical
MA 575 Linear Models: Cedric E Ginestet, Boston University Mixed Effects Estimation, Residuals Diagnostics Week 11, Lecture 1 1 Within-group Correlation Let us recall the simple two-level hierarchical
Inferences on a Normal Covariance Matrix and Generalized Variance with Monotone Missing Data
 Journal of Multivariate Analysis 78, 6282 (2001) doi:10.1006jmva.2000.1939, available online at http:www.idealibrary.com on Inferences on a Normal Covariance Matrix and Generalized Variance with Monotone
Journal of Multivariate Analysis 78, 6282 (2001) doi:10.1006jmva.2000.1939, available online at http:www.idealibrary.com on Inferences on a Normal Covariance Matrix and Generalized Variance with Monotone
Machine Learning, Midterm Exam: Spring 2009 SOLUTION
 10-601 Machine Learning, Midterm Exam: Spring 2009 SOLUTION March 4, 2009 Please put your name at the top of the table below. If you need more room to work out your answer to a question, use the back of
10-601 Machine Learning, Midterm Exam: Spring 2009 SOLUTION March 4, 2009 Please put your name at the top of the table below. If you need more room to work out your answer to a question, use the back of
1. (Rao example 11.15) A study measures oxygen demand (y) (on a log scale) and five explanatory variables (see below). Data are available as
 ST 51, Summer, Dr. Jason A. Osborne Homework assignment # - Solutions 1. (Rao example 11.15) A study measures oxygen demand (y) (on a log scale) and five explanatory variables (see below). Data are available
ST 51, Summer, Dr. Jason A. Osborne Homework assignment # - Solutions 1. (Rao example 11.15) A study measures oxygen demand (y) (on a log scale) and five explanatory variables (see below). Data are available
Test Code: STA/STB (Short Answer Type) 2013 Junior Research Fellowship for Research Course in Statistics
 Test Code: STA/STB (Short Answer Type) 2013 Junior Research Fellowship for Research Course in Statistics The candidates for the research course in Statistics will have to take two shortanswer type tests
Test Code: STA/STB (Short Answer Type) 2013 Junior Research Fellowship for Research Course in Statistics The candidates for the research course in Statistics will have to take two shortanswer type tests
Parametric Modelling of Over-dispersed Count Data. Part III / MMath (Applied Statistics) 1
 Parametric Modelling of Over-dispersed Count Data Part III / MMath (Applied Statistics) 1 Introduction Poisson regression is the de facto approach for handling count data What happens then when Poisson
Parametric Modelling of Over-dispersed Count Data Part III / MMath (Applied Statistics) 1 Introduction Poisson regression is the de facto approach for handling count data What happens then when Poisson
Notes on the Multivariate Normal and Related Topics
 Version: July 10, 2013 Notes on the Multivariate Normal and Related Topics Let me refresh your memory about the distinctions between population and sample; parameters and statistics; population distributions
Version: July 10, 2013 Notes on the Multivariate Normal and Related Topics Let me refresh your memory about the distinctions between population and sample; parameters and statistics; population distributions
Dr. Maddah ENMG 617 EM Statistics 11/28/12. Multiple Regression (3) (Chapter 15, Hines)
 Dr. Maddah ENMG 617 EM Statistics 11/28/12 Multiple Regression (3) (Chapter 15, Hines) Problems in multiple regression: Multicollinearity This arises when the independent variables x 1, x 2,, x k, are
Dr. Maddah ENMG 617 EM Statistics 11/28/12 Multiple Regression (3) (Chapter 15, Hines) Problems in multiple regression: Multicollinearity This arises when the independent variables x 1, x 2,, x k, are
Experimental Design and Data Analysis for Biologists
 Experimental Design and Data Analysis for Biologists Gerry P. Quinn Monash University Michael J. Keough University of Melbourne CAMBRIDGE UNIVERSITY PRESS Contents Preface page xv I I Introduction 1 1.1
Experimental Design and Data Analysis for Biologists Gerry P. Quinn Monash University Michael J. Keough University of Melbourne CAMBRIDGE UNIVERSITY PRESS Contents Preface page xv I I Introduction 1 1.1
Co-expression analysis of RNA-seq data
 Co-expression analysis of RNA-seq data Etienne Delannoy & Marie-Laure Martin-Magniette & Andrea Rau Plant Science Institut of Paris-Saclay (IPS2) Applied Mathematics and Informatics Unit (MIA-Paris) Genetique
Co-expression analysis of RNA-seq data Etienne Delannoy & Marie-Laure Martin-Magniette & Andrea Rau Plant Science Institut of Paris-Saclay (IPS2) Applied Mathematics and Informatics Unit (MIA-Paris) Genetique
Areal data models. Spatial smoothers. Brook s Lemma and Gibbs distribution. CAR models Gaussian case Non-Gaussian case
 Areal data models Spatial smoothers Brook s Lemma and Gibbs distribution CAR models Gaussian case Non-Gaussian case SAR models Gaussian case Non-Gaussian case CAR vs. SAR STAR models Inference for areal
Areal data models Spatial smoothers Brook s Lemma and Gibbs distribution CAR models Gaussian case Non-Gaussian case SAR models Gaussian case Non-Gaussian case CAR vs. SAR STAR models Inference for areal
Quantitative Genomics and Genetics BTRY 4830/6830; PBSB
 Quantitative Genomics and Genetics BTRY 4830/6830; PBSB.5201.01 Lecture16: Population structure and logistic regression I Jason Mezey jgm45@cornell.edu April 11, 2017 (T) 8:40-9:55 Announcements I April
Quantitative Genomics and Genetics BTRY 4830/6830; PBSB.5201.01 Lecture16: Population structure and logistic regression I Jason Mezey jgm45@cornell.edu April 11, 2017 (T) 8:40-9:55 Announcements I April
GLOBEX Bioinformatics (Summer 2015) Genetic networks and gene expression data
 GLOBEX Bioinformatics (Summer 2015) Genetic networks and gene expression data 1 Gene Networks Definition: A gene network is a set of molecular components, such as genes and proteins, and interactions between
GLOBEX Bioinformatics (Summer 2015) Genetic networks and gene expression data 1 Gene Networks Definition: A gene network is a set of molecular components, such as genes and proteins, and interactions between
THE UNIVERSITY OF CHICAGO Graduate School of Business Business 41912, Spring Quarter 2008, Mr. Ruey S. Tsay. Solutions to Final Exam
 THE UNIVERSITY OF CHICAGO Graduate School of Business Business 41912, Spring Quarter 2008, Mr. Ruey S. Tsay Solutions to Final Exam 1. (13 pts) Consider the monthly log returns, in percentages, of five
THE UNIVERSITY OF CHICAGO Graduate School of Business Business 41912, Spring Quarter 2008, Mr. Ruey S. Tsay Solutions to Final Exam 1. (13 pts) Consider the monthly log returns, in percentages, of five
Reports of the Institute of Biostatistics
 Reports of the Institute of Biostatistics No 01 / 2010 Leibniz University of Hannover Natural Sciences Faculty Titel: Multiple contrast tests for multiple endpoints Author: Mario Hasler 1 1 Lehrfach Variationsstatistik,
Reports of the Institute of Biostatistics No 01 / 2010 Leibniz University of Hannover Natural Sciences Faculty Titel: Multiple contrast tests for multiple endpoints Author: Mario Hasler 1 1 Lehrfach Variationsstatistik,
Scatter Plot Quadrants. Setting. Data pairs of two attributes X & Y, measured at N sampling units:
 Geog 20C: Phaedon C Kriakidis Setting Data pairs of two attributes X & Y, measured at sampling units: ṇ and ṇ there are pairs of attribute values {( n, n ),,,} Scatter plot: graph of - versus -values in
Geog 20C: Phaedon C Kriakidis Setting Data pairs of two attributes X & Y, measured at sampling units: ṇ and ṇ there are pairs of attribute values {( n, n ),,,} Scatter plot: graph of - versus -values in
Testing Statistical Hypotheses
 E.L. Lehmann Joseph P. Romano Testing Statistical Hypotheses Third Edition 4y Springer Preface vii I Small-Sample Theory 1 1 The General Decision Problem 3 1.1 Statistical Inference and Statistical Decisions
E.L. Lehmann Joseph P. Romano Testing Statistical Hypotheses Third Edition 4y Springer Preface vii I Small-Sample Theory 1 1 The General Decision Problem 3 1.1 Statistical Inference and Statistical Decisions
CS 195-5: Machine Learning Problem Set 1
 CS 95-5: Machine Learning Problem Set Douglas Lanman dlanman@brown.edu 7 September Regression Problem Show that the prediction errors y f(x; ŵ) are necessarily uncorrelated with any linear function of
CS 95-5: Machine Learning Problem Set Douglas Lanman dlanman@brown.edu 7 September Regression Problem Show that the prediction errors y f(x; ŵ) are necessarily uncorrelated with any linear function of
STATISTICS SYLLABUS UNIT I
 STATISTICS SYLLABUS UNIT I (Probability Theory) Definition Classical and axiomatic approaches.laws of total and compound probability, conditional probability, Bayes Theorem. Random variable and its distribution
STATISTICS SYLLABUS UNIT I (Probability Theory) Definition Classical and axiomatic approaches.laws of total and compound probability, conditional probability, Bayes Theorem. Random variable and its distribution
Permutation-invariant regularization of large covariance matrices. Liza Levina
 Liza Levina Permutation-invariant covariance regularization 1/42 Permutation-invariant regularization of large covariance matrices Liza Levina Department of Statistics University of Michigan Joint work
Liza Levina Permutation-invariant covariance regularization 1/42 Permutation-invariant regularization of large covariance matrices Liza Levina Department of Statistics University of Michigan Joint work
Chapter 3: Statistical methods for estimation and testing. Key reference: Statistical methods in bioinformatics by Ewens & Grant (2001).
 Chapter 3: Statistical methods for estimation and testing Key reference: Statistical methods in bioinformatics by Ewens & Grant (2001). Chapter 3: Statistical methods for estimation and testing Key reference:
Chapter 3: Statistical methods for estimation and testing Key reference: Statistical methods in bioinformatics by Ewens & Grant (2001). Chapter 3: Statistical methods for estimation and testing Key reference:
Session 3 The proportional odds model and the Mann-Whitney test
 Session 3 The proportional odds model and the Mann-Whitney test 3.1 A unified approach to inference 3.2 Analysis via dichotomisation 3.3 Proportional odds 3.4 Relationship with the Mann-Whitney test Session
Session 3 The proportional odds model and the Mann-Whitney test 3.1 A unified approach to inference 3.2 Analysis via dichotomisation 3.3 Proportional odds 3.4 Relationship with the Mann-Whitney test Session
BTRY 4090: Spring 2009 Theory of Statistics
 BTRY 4090: Spring 2009 Theory of Statistics Guozhang Wang September 25, 2010 1 Review of Probability We begin with a real example of using probability to solve computationally intensive (or infeasible)
BTRY 4090: Spring 2009 Theory of Statistics Guozhang Wang September 25, 2010 1 Review of Probability We begin with a real example of using probability to solve computationally intensive (or infeasible)
The optimal discovery procedure: a new approach to simultaneous significance testing
 J. R. Statist. Soc. B (2007) 69, Part 3, pp. 347 368 The optimal discovery procedure: a new approach to simultaneous significance testing John D. Storey University of Washington, Seattle, USA [Received
J. R. Statist. Soc. B (2007) 69, Part 3, pp. 347 368 The optimal discovery procedure: a new approach to simultaneous significance testing John D. Storey University of Washington, Seattle, USA [Received
Computational functional genomics
 Computational functional genomics (Spring 2005: Lecture 8) David K. Gifford (Adapted from a lecture by Tommi S. Jaakkola) MIT CSAIL Basic clustering methods hierarchical k means mixture models Multi variate
Computational functional genomics (Spring 2005: Lecture 8) David K. Gifford (Adapted from a lecture by Tommi S. Jaakkola) MIT CSAIL Basic clustering methods hierarchical k means mixture models Multi variate
Statistical Inference
 Statistical Inference Classical and Bayesian Methods Revision Class for Midterm Exam AMS-UCSC Th Feb 9, 2012 Winter 2012. Session 1 (Revision Class) AMS-132/206 Th Feb 9, 2012 1 / 23 Topics Topics We will
Statistical Inference Classical and Bayesian Methods Revision Class for Midterm Exam AMS-UCSC Th Feb 9, 2012 Winter 2012. Session 1 (Revision Class) AMS-132/206 Th Feb 9, 2012 1 / 23 Topics Topics We will
Chapter 4: Asymptotic Properties of the MLE (Part 2)
 Chapter 4: Asymptotic Properties of the MLE (Part 2) Daniel O. Scharfstein 09/24/13 1 / 1 Example Let {(R i, X i ) : i = 1,..., n} be an i.i.d. sample of n random vectors (R, X ). Here R is a response
Chapter 4: Asymptotic Properties of the MLE (Part 2) Daniel O. Scharfstein 09/24/13 1 / 1 Example Let {(R i, X i ) : i = 1,..., n} be an i.i.d. sample of n random vectors (R, X ). Here R is a response
Quantitative Genomics and Genetics BTRY 4830/6830; PBSB
 Quantitative Genomics and Genetics BTRY 4830/6830; PBSB.5201.01 Lecture 20: Epistasis and Alternative Tests in GWAS Jason Mezey jgm45@cornell.edu April 16, 2016 (Th) 8:40-9:55 None Announcements Summary
Quantitative Genomics and Genetics BTRY 4830/6830; PBSB.5201.01 Lecture 20: Epistasis and Alternative Tests in GWAS Jason Mezey jgm45@cornell.edu April 16, 2016 (Th) 8:40-9:55 None Announcements Summary
Econ 583 Homework 7 Suggested Solutions: Wald, LM and LR based on GMM and MLE
 Econ 583 Homework 7 Suggested Solutions: Wald, LM and LR based on GMM and MLE Eric Zivot Winter 013 1 Wald, LR and LM statistics based on generalized method of moments estimation Let 1 be an iid sample
Econ 583 Homework 7 Suggested Solutions: Wald, LM and LR based on GMM and MLE Eric Zivot Winter 013 1 Wald, LR and LM statistics based on generalized method of moments estimation Let 1 be an iid sample
Lecture 3. Inference about multivariate normal distribution
 Lecture 3. Inference about multivariate normal distribution 3.1 Point and Interval Estimation Let X 1,..., X n be i.i.d. N p (µ, Σ). We are interested in evaluation of the maximum likelihood estimates
Lecture 3. Inference about multivariate normal distribution 3.1 Point and Interval Estimation Let X 1,..., X n be i.i.d. N p (µ, Σ). We are interested in evaluation of the maximum likelihood estimates
Statistical Significance of Ranking Paradoxes
 Statistical Significance of Ranking Paradoxes Anna E. Bargagliotti and Raymond N. Greenwell 1 February 28, 2009 1 Anna E. Bargagliotti is an Assistant Professor in the Department of Mathematical Sciences
Statistical Significance of Ranking Paradoxes Anna E. Bargagliotti and Raymond N. Greenwell 1 February 28, 2009 1 Anna E. Bargagliotti is an Assistant Professor in the Department of Mathematical Sciences
Statistical Inference of Covariate-Adjusted Randomized Experiments
 1 Statistical Inference of Covariate-Adjusted Randomized Experiments Feifang Hu Department of Statistics George Washington University Joint research with Wei Ma, Yichen Qin and Yang Li Email: feifang@gwu.edu
1 Statistical Inference of Covariate-Adjusted Randomized Experiments Feifang Hu Department of Statistics George Washington University Joint research with Wei Ma, Yichen Qin and Yang Li Email: feifang@gwu.edu
Inferring Gene Regulatory Networks from Time-Ordered Gene Expression Data of Bacillus Subtilis Using Differential Equations
 Inferring Gene Regulatory Networks from Time-Ordered Gene Expression Data of Bacillus Subtilis Using Differential Equations M.J.L. de Hoon, S. Imoto, K. Kobayashi, N. Ogasawara, S. Miyano Pacific Symposium
Inferring Gene Regulatory Networks from Time-Ordered Gene Expression Data of Bacillus Subtilis Using Differential Equations M.J.L. de Hoon, S. Imoto, K. Kobayashi, N. Ogasawara, S. Miyano Pacific Symposium
2. What are the tradeoffs among different measures of error (e.g. probability of false alarm, probability of miss, etc.)?
 ECE 830 / CS 76 Spring 06 Instructors: R. Willett & R. Nowak Lecture 3: Likelihood ratio tests, Neyman-Pearson detectors, ROC curves, and sufficient statistics Executive summary In the last lecture we
ECE 830 / CS 76 Spring 06 Instructors: R. Willett & R. Nowak Lecture 3: Likelihood ratio tests, Neyman-Pearson detectors, ROC curves, and sufficient statistics Executive summary In the last lecture we
Stat 206: Sampling theory, sample moments, mahalanobis
 Stat 206: Sampling theory, sample moments, mahalanobis topology James Johndrow (adapted from Iain Johnstone s notes) 2016-11-02 Notation My notation is different from the book s. This is partly because
Stat 206: Sampling theory, sample moments, mahalanobis topology James Johndrow (adapted from Iain Johnstone s notes) 2016-11-02 Notation My notation is different from the book s. This is partly because
Discrete Mathematics and Probability Theory Fall 2015 Lecture 21
 CS 70 Discrete Mathematics and Probability Theory Fall 205 Lecture 2 Inference In this note we revisit the problem of inference: Given some data or observations from the world, what can we infer about
CS 70 Discrete Mathematics and Probability Theory Fall 205 Lecture 2 Inference In this note we revisit the problem of inference: Given some data or observations from the world, what can we infer about
Statistical Applications in Genetics and Molecular Biology
 Statistical Applications in Genetics and Molecular Biology Volume 6, Issue 1 2007 Article 28 A Comparison of Methods to Control Type I Errors in Microarray Studies Jinsong Chen Mark J. van der Laan Martyn
Statistical Applications in Genetics and Molecular Biology Volume 6, Issue 1 2007 Article 28 A Comparison of Methods to Control Type I Errors in Microarray Studies Jinsong Chen Mark J. van der Laan Martyn
Lecture 3: A basic statistical concept
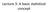 Lecture 3: A basic statistical concept P value In statistical hypothesis testing, the p value is the probability of obtaining a result at least as extreme as the one that was actually observed, assuming
Lecture 3: A basic statistical concept P value In statistical hypothesis testing, the p value is the probability of obtaining a result at least as extreme as the one that was actually observed, assuming
Binary choice 3.3 Maximum likelihood estimation
 Binary choice 3.3 Maximum likelihood estimation Michel Bierlaire Output of the estimation We explain here the various outputs from the maximum likelihood estimation procedure. Solution of the maximum likelihood
Binary choice 3.3 Maximum likelihood estimation Michel Bierlaire Output of the estimation We explain here the various outputs from the maximum likelihood estimation procedure. Solution of the maximum likelihood
A Bayesian Nonparametric Model for Predicting Disease Status Using Longitudinal Profiles
 A Bayesian Nonparametric Model for Predicting Disease Status Using Longitudinal Profiles Jeremy Gaskins Department of Bioinformatics & Biostatistics University of Louisville Joint work with Claudio Fuentes
A Bayesian Nonparametric Model for Predicting Disease Status Using Longitudinal Profiles Jeremy Gaskins Department of Bioinformatics & Biostatistics University of Louisville Joint work with Claudio Fuentes
Learning Multiple Tasks with a Sparse Matrix-Normal Penalty
 Learning Multiple Tasks with a Sparse Matrix-Normal Penalty Yi Zhang and Jeff Schneider NIPS 2010 Presented by Esther Salazar Duke University March 25, 2011 E. Salazar (Reading group) March 25, 2011 1
Learning Multiple Tasks with a Sparse Matrix-Normal Penalty Yi Zhang and Jeff Schneider NIPS 2010 Presented by Esther Salazar Duke University March 25, 2011 E. Salazar (Reading group) March 25, 2011 1
Bayesian Inference for Normal Mean
 Al Nosedal. University of Toronto. November 18, 2015 Likelihood of Single Observation The conditional observation distribution of y µ is Normal with mean µ and variance σ 2, which is known. Its density
Al Nosedal. University of Toronto. November 18, 2015 Likelihood of Single Observation The conditional observation distribution of y µ is Normal with mean µ and variance σ 2, which is known. Its density
High-Throughput Sequencing Course
 High-Throughput Sequencing Course DESeq Model for RNA-Seq Biostatistics and Bioinformatics Summer 2017 Outline Review: Standard linear regression model (e.g., to model gene expression as function of an
High-Throughput Sequencing Course DESeq Model for RNA-Seq Biostatistics and Bioinformatics Summer 2017 Outline Review: Standard linear regression model (e.g., to model gene expression as function of an
Multivariate Time Series
 Multivariate Time Series Notation: I do not use boldface (or anything else) to distinguish vectors from scalars. Tsay (and many other writers) do. I denote a multivariate stochastic process in the form
Multivariate Time Series Notation: I do not use boldface (or anything else) to distinguish vectors from scalars. Tsay (and many other writers) do. I denote a multivariate stochastic process in the form
A Squared Correlation Coefficient of the Correlation Matrix
 A Squared Correlation Coefficient of the Correlation Matrix Rong Fan Southern Illinois University August 25, 2016 Abstract Multivariate linear correlation analysis is important in statistical analysis
A Squared Correlation Coefficient of the Correlation Matrix Rong Fan Southern Illinois University August 25, 2016 Abstract Multivariate linear correlation analysis is important in statistical analysis
Data Mining Stat 588
 Data Mining Stat 588 Lecture 02: Linear Methods for Regression Department of Statistics & Biostatistics Rutgers University September 13 2011 Regression Problem Quantitative generic output variable Y. Generic
Data Mining Stat 588 Lecture 02: Linear Methods for Regression Department of Statistics & Biostatistics Rutgers University September 13 2011 Regression Problem Quantitative generic output variable Y. Generic
Logistic Regression: Regression with a Binary Dependent Variable
 Logistic Regression: Regression with a Binary Dependent Variable LEARNING OBJECTIVES Upon completing this chapter, you should be able to do the following: State the circumstances under which logistic regression
Logistic Regression: Regression with a Binary Dependent Variable LEARNING OBJECTIVES Upon completing this chapter, you should be able to do the following: State the circumstances under which logistic regression
Math 494: Mathematical Statistics
 Math 494: Mathematical Statistics Instructor: Jimin Ding jmding@wustl.edu Department of Mathematics Washington University in St. Louis Class materials are available on course website (www.math.wustl.edu/
Math 494: Mathematical Statistics Instructor: Jimin Ding jmding@wustl.edu Department of Mathematics Washington University in St. Louis Class materials are available on course website (www.math.wustl.edu/
Nesting and Equivalence Testing
 Nesting and Equivalence Testing Tihomir Asparouhov and Bengt Muthén August 13, 2018 Abstract In this note, we discuss the nesting and equivalence testing (NET) methodology developed in Bentler and Satorra
Nesting and Equivalence Testing Tihomir Asparouhov and Bengt Muthén August 13, 2018 Abstract In this note, we discuss the nesting and equivalence testing (NET) methodology developed in Bentler and Satorra
[y i α βx i ] 2 (2) Q = i=1
![[y i α βx i ] 2 (2) Q = i=1 [y i α βx i ] 2 (2) Q = i=1](/thumbs/75/72005858.jpg) Least squares fits This section has no probability in it. There are no random variables. We are given n points (x i, y i ) and want to find the equation of the line that best fits them. We take the equation
Least squares fits This section has no probability in it. There are no random variables. We are given n points (x i, y i ) and want to find the equation of the line that best fits them. We take the equation
