Molecular Simulation I
|
|
|
- Ashlie Hawkins
- 6 years ago
- Views:
Transcription
1 Molecular Simulation I Quantum Chemistry Classical Mechanics E = Ψ H Ψ ΨΨ U = E bond +E angle +E torsion +E non-bond Jeffry D. Madura Department of Chemistry & Biochemistry Center for Computational Sciences Duquesne University
2 Quantum Chemistry Molecular Orbital Theory Based on a wave function approach Schrödinger equation Density Functional Theory Based on the total electron density Hohenberg Kohn theorem Semi-empirical Some to most integrals parameterized MNDO, AM1, EHT Empirical All integrals are parameterized Huckel method
3 The Beginning... Schrödinger equation H Ψ = Hamiltonian operator EΨ h h H = + + V r V r = + 2m x y z 2m Wave function ( ) Ψ characterizes the particles motion ( ) 2 ( ) various properties of the particle can be derived
4 Quantum Chemistry Start with Schrödinger s equation HΨ = EΨ Make some assumptions Born-Oppenheimer approximation Linear combination of atomic orbitals Ψ = c ϕ + cϕ a a b b Apply the variational method E Ψ * HΨdτ * ΨΨ dτ
5 LCAO A practical and common approach to solving the Hartree-Fock equations is to write each spin orbital as a linear combination of single electron orbitals (LCAO) K ψ i = ν = 1 c ν i the φ v are commonly called basis functions and often correspond to atomic orbitals K basis functions lead to K molecular orbitals the point at which the energy is not reduced by the addition of basis functions is known as the Hartree- Fock limit φ ν
6 Basis Sets Slater type orbitals (STO) Gaussian type orbitals (GTO) functional form n+ 1/2 1/2 n 1 r R ( ) ( 2 ) ( 2 ) nl r = ζ n! r e ζ x yze a a b c - r zeroth-order Gaussian function 2 3/4 2α r gs ( α, r) = e α π 2
7 Property of Gaussian functions is that the product of two Gaussians can be expressed as a single Gaussian, located along the line joining the centers of the two Gaussians a a -a - a = a + a -a m n rmn 2 mrm nrn m n rc e e e e fx ( ) 0.6 g( x) fx ( ). g( x) x 6
8 STO vs. GTO
9 Gaussian expansion the coefficient L φ ( ) the exponent μ = diμφi αiμ i= 1 uncontratcted or primitive and contracted s and p exponents in the same shell are equal Minimal basis set STO-NG Double zeta basis set linear combination of a contracted function and a diffuse function. Split valence 3-21G, 4-31G, 6-31G
10 Polarization to solve the problem of non-isotropic charge distribution. 6-31G*, 6-31G** Diffuse functions fulfill as deficiency of the basis sets to describe significant amounts of electron density away from the nuclear centers. (e.g. anions, lone pairs, etc.) 3-21+G, G
11 RHF vs. UHF Restricted Hartree-Fock (RHF) closed-shell molecules Restricted Open-shell Hartree-Fock (ROHF) combination of singly and doubly occupied molecular orbitals. Unrestricted Hartree-Fock (UHF) open-shell molecules Pople and Nesbet: one set of molecular orbitals for α spin and another for the β spin.
12 UHF and RHF Dissociation Curves for H E(H 2 )-2E(H) a.u R (a.u.)
13 Electron Correlation The most significant drawback to HF theory is that it fails to adequately represent electron correlation. NR E = E E corr Configuration Interactions HF excited states are included in the description of an electronic state Many Body Perturbation Theory based upon Rayleigh-Schrödinger perturbation theory
14 Configuration Interaction The CI wavefunction is written as Ψ= c0ψ 0 + c1ψ 1+ c2ψ 2 +L where Ψ 0 is the HF single determinant where Ψ 1 is the configuration derived by replacing one of the occupied spin orbitals by a virtual spin orbital where Ψ 2 is the configuration derived by replacing one of the occupied spin orbitals by a virtual spin orbital The system energy is minimized in order to determine the coefficients, c 0, c 1, etc., using a linear variational approach
15 Many Body Peturbation Theory Based upon perturbation concepts The correction to the energies are ( 0) ( 0) ( 0) E = Ψ H Ψ dτ i i 0 i ( 1) ( 0) ( 0) E = Ψ VΨ dτ i i i ( 2) ( 0) ( 1) E = Ψ VΨ dτ i i i ( 3) ( 0) ( 2) E = Ψ VΨ dτ i i i Perturbation methods are size independent these methods are not variational H = H0 + V
16 Theoretical Model Theoretical Model = Level of Theory + Basis Set Level of Theory = HF, MP2, DFT, CI, CCSD, etc Basis Set = STO-3G, 3-21G, 6-31G*, G(d,p), etc
17 Geometry Optimization Derivatives of the energy ( ) ( ) ( ) ( ) ( ) 2 E x 1 E x ( )( ) L E xf = E x + xif xi + xif xi xj xj + x 2 x x i i i j i j the first term is set to zero the second term can be shown to be equivalent to a force the third term can be shown to be equivalent to a force constant
18 Internal coordinate, Cartesian coordinate, and redundant coordinate optimization choice of coordinate set can determine whether a structure reaches a minimum/maximum and the speed of this convergence. Internal coordinates are defined as bond lengths, bond angles, and torsions. There are 3N-6 (3N-5) such degrees of freedom for each molecule. Chemists work in this world. Z-matrix... Cartesian coordinates are the standard x, y, z coordinates. Programs often work in this world. Redundant coordinates are defined as the number of coordinates larger than 3N-6.
19 Potential Energy Surfaces f k dv = dr = 2 dv dr 2
20 Frequency Calculation The second derivatives of the energy with respect to the displacement of coordinate yields the force constants. These force constants in turn can be used to calculate frequencies. All real frequencies (positive force constants): local minimum One imaginary frequency (one negative force constant): saddle point, a.k.a. transition state. From vibrational analysis can compute thermodynamic data
21 Charges Molecular Properties Mulliken Löwdin electrostatic fitted (ESP) Bond orders Bonding Natural Bond Analysis Bader s AIM method Molecular orbitals and total electron density Dipole Moment Energies ionization and electron affinity
22 Energies Koopman s theorem equating the energy of an electron in an orbital to the energy required to remove the electron to the corresponding ion. frozen orbitals lack of electron correlation effects
23 Dipole Moments The electric multipole moments of a molecule reflect the distribution of charge. Simplest is the dipole moment nuclear component electronic μ M = Z R nuclear A A A= 1 K K μ= 1 ν= 1 μ = qr ( ) μelectronic = P dτφ r φ μν μ ν i i i
24 Molecular Orbitals and Total Electron Density Electron density at a point r 2 ( r) = 2 ( r) = P ( r) ( r) + 2 P ( r) ( r) Number of electrons is Molecular orbitals HOMO LUMO N /2 K K K ρ ψ φ φ φ φ i μμ μ μ μν μ ν i= 1 μ= 1 μ= 1ν= μ+ 1 N /2 2 ( ) K K K ψ i μμ μν μν i= 1 μ= 1 μ= 1ν= μ+ 1 N = 2 dr r = P + 2 P S
25 Natural Bond Analysis Bonding a way to describe N-electron wave functions in terms of localized orbitals that are closely tied to chemical concepts. Bader F. W. Bader s theory of atoms in molecules. This method provides an alternative way to partition the electrons among the atoms in a molecule. Gradient vector path bond critical points charges are relatively invariant to the basis set
26 Bond Orders Wiberg Mayer W AB AB = μon Aνon B μon Aνon B P μν B PS PS = Bond orders can be computed for intermediate structures which can be useful way to describe similarity of the TS to the reactants or to the products. 2 ( ) ( ) μν νμ
27 Mulliken A Löwdin A Charges K K K q = Z P P S μμ μν μν μ= 1; on A μ= 1; on Aν= 1; ν μ atomic orbitals are transformed to an orthogonal set, along with the mo coefficients φ K ' 1/2 μ = ν = 1 A A ( S ) K νμ φ ( 1/2 1/2 ) q = Z S P ν μ= 1; μon A μμ
28 Summary of Methods
29 Summary of Results
30 Summary of Results
31 Summary of Results
32 Summary of Results
33 Summary of Results
34 Electrostatic potentials the electrostatic potential at a point r, φ(r), is defined as the work done to bring a unit positive charge from infinity to the point. the electrostatic interaction energy between a point charge q located at r and the molecule equals qφ(r). there is a nuclear part and electronic part φ nucl r ( ) = Z M A ( ) elec r A= 1 r RA φ = dr r ' ρ ' r ( ) r φ r = φ r + φ r ( ) ( ) ( ) nucl elec
35 Electrostatic Potential
36 G03W - Water Example
37 Water Example Geometry Z-MATRIX (ANGSTROMS AND DEGREES) CD Cent Atom N1 Length/X N2 Alpha/Y N3 Beta/Z J H 2 2 O ( 1) Energy 3 3 H ( 2) ( 3) E(RHF) = A.U. Population Analysis ********************************************************************** Population analysis using the SCF density. ********************************************************************** Alpha occ. eigenvalues Alpha virt. eigenvalues Condensed to atoms (all electrons): H O H Total atomic charges: 1 1 H O H
38 GaussView Water Example
39 GaussView Water Example
40 Limitations, Strengths & Reliability Limitations Requires more CPU time Can treat smaller molecules Calculations are more complex Have to worry about electronic configuration Strengths No experimental bias Can improve a calculation in a logical manner (e.g. basis set, level of theory, ) Provides information on intermediate species, including spectroscopic data Can calculate novel structures Can calculate any electronic state Reliability The mean deviation between experiment and theory for heavy-atom bond lengths in twoheavy-atom hydrides drops from A for the RHF/STO-3G level of theory to just A for MP2/6-31G(d). Heats of hydrogenation of a range of saturated and unsaturated systems are calculated sufficiently well at the Hartree-Fock level of theory with a moderate basis set (increasing the basis set from 6-31G(d) to 6-31G(d,p) has little effect on the accuracy of these numbers). Inclusion of electron correlation is mandatory in order to get good agreement between experiment and theory for bond dissociation energies (MP2/6-31G(d,p) does very well for the one-heavy-atom hydrides).
41 Summary What can you do with electronic structure methods? Geometry optimizations (minima and transition states) Energies of minima and transition states Chemical reactivity IR, UV, NMR spectra Physical properties of molecules Interaction energy between two or more molecules
42 Molecular Simulation II Quantum Chemistry Classical Mechanics E = Ψ H Ψ ΨΨ U = E bond +E angle +E torsion +E non-bond Jeffry D. Madura Department of Chemistry & Biochemistry Center for Computational Sciences Duquesne University
43 Classical Mechanical Treatment I. Classical Mechanics II. a. Implicit treatment of electrons b. Use simple analytical functions (i.e., harmonic springs) c. Use Cartesian coordinates, not the z-matrix Force Fields a. Have evolved over time b. Use different analytical terms and parameters c. Are specific for classes of molecules (proteins, carbohydrates, nucleic acids, organic molecules, etc.)
44 What is a force field? Force Field A mathematical expression that describes the dependence of the energy of a molecule on the coordinates of the atoms in the molecule Also this sometimes used as another term for potential energy function. What are force fields used for? Structure determination Conformational energies Rotational and Pyramidal inversion barriers Vibrational frequencies Molecular dynamics
45 Force Field History Pre-1970 Harmonic 1970 For molecules with less than 100 atoms one class of force fields went for high accuracy to match experimental results The other class of force fields was for macromolecules. Present There are highly accurate force fields designed for small molecules and there are force fields for studying protein and other large molecules
46 Force Field First force fields developed from experimental data X-ray NMR Microwave Current force fields have made use of quantum mechanical calculations CFF MMFF94 There is no single best force field
47 Force Fields MM2/3/4: Molecular Mechanic Force field for small moelcules CHARMM: Chemistry at Harvard Macromolecular Mechanics AMBER: Assisted Model Building with Energy Refinement OPLS: Optimized Parameters for Liquid Simulation CFF: Consistent Force Field CVFF: Valence Consistent Force Field MMFF94: Merck Molecular Force Field 94 UFF: Universal Force Field
48 Comparison and Evaluation Engler, E. M., J. D. Andose and P. v. R. Schleyer (1973) "Critical Evaluation of Molecular Mechanics," J. Am. Chem. Soc. 95, Gundertofte, K., J. Palm, I. Pettersson and A. Stamvik (1991) "A Comparison of Conformational Energies Calculated by Molecular Mechanics (MM2(85), Sybyl 5.1, Sybyl 5.21, ChemX) and Semiempirical (AM1 and PM3) Methods," J. Comp. Chem. 12, Gundertofte, K., T. Liljefors, P-O Norrby, I. Pattesson (1996) "Comparison of Conformational Energies Calculated by Several Molecular Mechanics Methods," J. Comp. Chem. 17, (1996). Hall, D., and N. Pavitt (1984) "A Appraisal of Molecular Force Fields for the Representation of Polypeptides," J. Comp. Chem. 5, Hobza, P., M. Kabelac, J. Sponer, P. Mejzlik and J. Vondrasek (1997) "Performance of Empirical Potentials (AMBER, CFF95, CVFF, CHARMM, OPS, POLTEV), Semiemprical Quantum Chemical Methods (AM1, MNDO/M, PM3) and ab initio Hartree-Fock Method for Interaction of DNA Bases: Comparison of Nonempirical Beyond Hartree-Fock Results," J. Comp. Chem. 18, Kini, R. M., and H. J. Evans (1992) "Comparison of Protein Models Minimized by the All-Atom and United Atom Models in the AMBER force Field," J. Biomol. Structure and Dynamics 10, Roterman, I. K., Gibson, K. D., and Scheraga, H. A. (1989) "A Comparison of the CHARMM, AMBER, and ECEPP/2 Potential for Peptides I." J. Biomol. Struct. and Dynamics 7, Roterman, I. K., Lambert, M. H., Gibson, K. D., and Scheraga, H. A. (1989) "A Comparison of the CHARMM, AMBER, and ECEPP/2 Potential for Peptides II." J. Biomol. Struct. and Dynamics 7, Whitlow, M., and M. M. Teeter (1986) "A Empirical Examination of Potential Energy Minimization using the Well-Determined Structure of the Protein Crambin," J. Am. Chem. Soc. 108,
49 Potential Energy Function The potential energy function is a mathematical model which describes the various interactions between the atoms of a molecule or system of molecules. In general, the function is composed of intramolecular terms (1st three terms) and intermolecular terms (last two terms). Ur 1 2 () = ( b b ) + o ( θ θ ) + o 2 1 j+ 1 Vj[ 1+ ( 1) cos( n jφ δ) ] + 2 j ε[( σ 12 4 ) ( σ 6 ) ] + rij rij qq i j 4εr i j
50 Bond Stretch E = k l l l l ( ) 2 0 Harmonic approximation is used k b is the force constant l 0 is the reference bond length Higher order terms ( ) 2 '( ) 3 ''( ) 4 l = l 0 + l 0 + l 0 E k l l k l l k l l
51 Angle Bending ( 0 ) 2 Eθ = kθ θ θ Harmonic approximation k θ is the bending force constant θ 0 is the reference angle Other forms include Eθ = kθ 1+ cosθ ( )
52 Torsion Interactions ( 1 cos ) ( 1 cos 2 ) ( 1 cos 3 ) E V V V ϕ = 1 + ϕ + 2 ϕ ϕ Represented as a Fourier series This term accounts for the energetics of twisting the 1-4 atoms First term: important for describing the conformational energies (cis-trans) Second term: important for determining the relatively large barrier to rotation about conjugated bonds Third term: allows for accurate of the energy barrier for rotation about bonds where one or both of the atoms in the bond have sp 3 hybridization
53 Out-of-plane Bending Eω = kω ω ω ( ) 2 0 Harmonic approximation k ω is the oop force constant ω 0 is the reference value Diffferent methods in which to calculate ω MMF: angle between a bond i-j and a plane formed by j-k-l and j is the central atom MM3: angle between a bond i-j and a point located in the place formed by i-k-j.
54 Van Der Waals Interactions E vdw R = ε 2 R ij R ij 12 6 * * ij Rij Lennard-Jones 12-6 potential ε is the well depth R ij* is the sum of the van der Waals radii (of atoms i and j) R ij is the distance between interacting atoms
55 Electrostatics E elec = qq i DR j ij Coulomb s law q i and q j are the charges on atom i and j respectively D is the dielectric constant R ij is the distance between atoms i and j Bond increment model (used in CFF and MMFF) q q δ = + 0 i i ij j
56 Charge Classes Class I Calculated directly from experiment Class II Extracted from a quantum mechanical wave function (Mulliken analysis) Class III Extracted from a wave function by analyzing a physical observable predicted from the wave function. (Electrostatic fitting) Class IV A parameterization procedure to improve class II and III charges by mapping them to reproduce charge-dependent observables obtained from experiment
57 Cross Terms bb bθ bb bθ ( 0)( 0) E = k b b b b E θθ ( 0)( 0 θ ) 1 θ1 θ2 θ2 = k θθ ( 0)( 0 θ θ ) E = k b b Bond/bond Needed to get the correct splitting in the vibrational frequencies of the symmetric and asymmetric C-H bond stretching modes Angle/angle Needed to determine correctly the extent of splitting in angular deformation modes for the cases in which the bending modes are centered on a single atom Bond/angle Needed to predict the observed bond lengthening that often occurs when a bond angle is reduced
58 Molecular Mechanics In the molecular mechanics model, a molecule is described as a series of point charges (atoms) linked by springs (bonds). A mathematical function (the force-field) describes the freedom of bond lengths, bond angles, and torsions to change. The force-field also contains a description of the van der Waals and electrostatic interactions between atoms that are not directly bonded. The force-field is used to describe the potential energy of the molecule or system of interest. Molecular mechanics is a mathematical procedure used to explore the potential energy surface of a molecule or system of interest. Force F = U Potential Energy
59 Potential Energy Minimizations Potential Energy Surface: Has minima (stable structures) and saddle points (transition states). Below: 2 minima & 1 Saddle Point.
60 Energy Minimization Given a function f which depends on one or more independent variables, x 1, x 2,, find the values of those variables where f has a minimum value. Potential Energy Surface f = x i 0 2 f > 0 2 x
61 Energy Minimization Methods Taylor series expansion about point x k 2 2 U( xk) ( x xk) U( xk) U( x) = U( xk) + ( x xk) + + L xk 2 xk xj the second term is known as the gradient (force) the third term is known as the Hessian (force constant) Algorithms are classified by order, or the highest derivative used in the Taylor series. Common algorithms (1 st Order): Steepest Descent (SD), Conjugated Gradients (CONJ) Non-derivative Simplex Sequential univariate method
62 Energy Minimization Methods Derivative Steepest descents Moves are made in the direction parallel to the net force Conjugate gradient The gradients and the direction of successive steps are orthogonal Newton-Raphson Second-order method; both first and second derivatives are used BFGS Quasi-Newton method (a.k.a. variable metric methods) build up the inverse Hessian matrix in successive iterations
63 Energy Minimization Methods Truncated Newton-Raphson Initially follow a descent direction and near the solution solve more accurately using a Newton method. Different from QN in that the Hessian is sparse allowing for a faster evaluation
64 Comparing 1 st Order Algorithms BOTH: iterate over the following equation in order to perform the minimization: R k = R k-1 + l k S k Where R k is the new position at step k, R k-1 in the position at the previous step k-1, lk is the size of the step to be taken at step k and S k is the direction. SD: At each step the gradient of the potential g k (i.e., the first derivative in multi-dimensions) is calculated and a displacement is added to all the coordinates in a direction opposite to the gradient. S k = -g k CONJ: In each step, weighs in the previous gradients to compensate for the lack of curvature information. For all steps k > 1 the direction of the step is a weighted average of the current gradient and the previous step direction, i.e., S k = -g k + b k S k-1
65 SD versus CONJ SD versus CONJ. Starting from point A, SD will follow a path A-B-C. CONJ will follow A-B-O because it modifies the second direction to take into account the previous gradient along A-B and the current gradient along B-C.
66 Convergence Comparison of Methods Small change in energy Small norm of the gradient RMS gradient Number of steps vs. time grad RMS du = i dxi grad Steepest descents: 500 steps in secs (not converged) Conjugate gradient: 72 steps in seconds Newton-Raphson: 15 steps in seconds = n
67 Which Method Should I Use? Must consider Storage: Steepest descents little memory needed while Newton- Raphson methods require lots. Availability of derivatives: Simplex, none are needed, steepest descents, only first derivatives, Newton-Raphson needs first and second derivatives. The following is common practice SD or CG for the initial rough minimization followed by a few steps of NR. SD is superior to CG when starting structure is far from the minimum TN method after a few SD and/or CG appears to give the best overall and fastest convergence
68 Conformational Analysis Molecular conformations term used to describe molecular structures that interconvert under ambient conditions. this implies several conformations may be present, in differing conc., under ambient conditions. a proper description of the molecular structure, the molecular energy, or the spectrum for a molecule with several conformations must comprise a proper weighting of all of the conformations.
69 Boltzmann s equation P i = j fe i Ei RT fe j E j RT f i is the number of states or conf. of energy E i R is 1.98 cal/mol-k (the ideal gas constant) T is the absolute temperature (K) j is the summation over all the conformations
70 Butane Conformational Analysis
71 Conformational Analysis Example Using Boltzmann s equation We have a population of 89.74% at -180 and 10.26% at +/- 60 assuming a relative energy difference of 1.7 kcal/mol.
72 Conformational Analysis: A Cautionary Note MM2 Dreiding Term Trans Gauche ΔE Trans Gauche ΔE Stretch Stretch-Bend Bend Torsion VDW Total Even though the energetic difference given by the two models is similar, different contributions give rise to those differences.
73 Molecular Mechanics Energetics Steric energy the energy reported by most molecular mechanics programs energy of structure at 0 K. correct for vibrational motion by adding the zero point energy 1 ZPE = ν i MM energy is NOT equal to free energy!! MM energy can be equivalent to enthalpy if one assumes the PV term can be ignored i
74 Molecular Mechanics Energy Strain energy Differences in steric energy are oly valid for different conformations or configurations. Strain energy permits the comparison between different molecules. A strainless reference point must be determined. A particular reference point might be the all trans conformations of the straight-chain alkanes from methane to hexane (Allinger definition). These compounds can be used to derive a set of strainless energy parameters for constituent parts of molecules. Subtracting the strainless energy from the steric energy Allinger and co-workers concluded that the chair cyclohexane has an inherent strain energy due to the presence of 1,4 van der Waals interactions between the carbon atom within the ring.
75 Molecular Mechanics Energy Interaction energy This is the difference between the energies of two isolated species and the energy of the intermolecular complex Mathematically this is represented as ( ) E = E E + E ie ab a b
76 Steric Energies Using steric energies to predict the thermodynamics of simple tautomerization O OH The experimental heat of formation difference is approximately 8 kcal/mol MM2 steric energy difference is 2.3 kcal/mol The large error is due to error in bond energy terms, I.e. the number/types of bonds broken and made are not precisely balanced.
77 Positional Isomers In this case the number and precise types of bonds are retianed. Consider hydracrlic acid vs. lactic acid. HO O OH HO O OH MM2 energy for hydracrylic acid is kcal/mol while that for lactic acid is 2.37 kcal/mol yielding an energy difference of kcal/mol in favor of hydracrylic acid. Experimentally the heat of formation difference is 4.4 kcal/mol in favor of lactic acid.
78 Conformers and Configurational Isomers Molecular mechanics can be employed to predict relative energies of conformers and configurational isomers. It should not be used on positional isomers or the relative energies of different molecules. Molecular mechanics can be used to study the binding between molecules is intermolecular interactions have been appropriately parameterized.
79 Ideal Gas Statistical Thermodyanmics The free energy can be written as G =- kt ln Q+ PV where Q = i fe i E i RT The difference in free energy can be written as Δ G = RTln P P 1 2
Molecular Simulation II. Classical Mechanical Treatment
 Molecular Simulation II Quantum Chemistry Classical Mechanics E = Ψ H Ψ ΨΨ U = E bond +E angle +E torsion +E non-bond Jeffry D. Madura Department of Chemistry & Biochemistry Center for Computational Sciences
Molecular Simulation II Quantum Chemistry Classical Mechanics E = Ψ H Ψ ΨΨ U = E bond +E angle +E torsion +E non-bond Jeffry D. Madura Department of Chemistry & Biochemistry Center for Computational Sciences
Molecular Simulation I
 Molecular Simulation I Quantum Chemistry Classical Mechanics E = Ψ H Ψ ΨΨ U = E bond +E angle +E torsion +E non-bond Jeffry D. Madura Department of Chemistry & Biochemistry Center for Computational Sciences
Molecular Simulation I Quantum Chemistry Classical Mechanics E = Ψ H Ψ ΨΨ U = E bond +E angle +E torsion +E non-bond Jeffry D. Madura Department of Chemistry & Biochemistry Center for Computational Sciences
Molecular Simulation II
 Molecular Simulation II Quantum Chemistry Classical Mechanics E = Ψ H Ψ ΨΨ U = E bond +E angle +E torsion +E non-bond Jeffry D. Madura Department of Chemistry & Biochemistry Center for Computational Sciences
Molecular Simulation II Quantum Chemistry Classical Mechanics E = Ψ H Ψ ΨΨ U = E bond +E angle +E torsion +E non-bond Jeffry D. Madura Department of Chemistry & Biochemistry Center for Computational Sciences
Computational Methods. Chem 561
 Computational Methods Chem 561 Lecture Outline 1. Ab initio methods a) HF SCF b) Post-HF methods 2. Density Functional Theory 3. Semiempirical methods 4. Molecular Mechanics Computational Chemistry " Computational
Computational Methods Chem 561 Lecture Outline 1. Ab initio methods a) HF SCF b) Post-HF methods 2. Density Functional Theory 3. Semiempirical methods 4. Molecular Mechanics Computational Chemistry " Computational
Example questions for Molecular modelling (Level 4) Dr. Adrian Mulholland
 Example questions for Molecular modelling (Level 4) Dr. Adrian Mulholland 1) Question. Two methods which are widely used for the optimization of molecular geometies are the Steepest descents and Newton-Raphson
Example questions for Molecular modelling (Level 4) Dr. Adrian Mulholland 1) Question. Two methods which are widely used for the optimization of molecular geometies are the Steepest descents and Newton-Raphson
Molecular Simulation III
 Molecular Simulation III Quantum Chemistry Classical Mechanics E = Ψ H Ψ ΨΨ U = E bond +E angle +E torsion +E non-bond Jeffry D. Madura Department of Chemistry & Biochemistry Center for Computational Sciences
Molecular Simulation III Quantum Chemistry Classical Mechanics E = Ψ H Ψ ΨΨ U = E bond +E angle +E torsion +E non-bond Jeffry D. Madura Department of Chemistry & Biochemistry Center for Computational Sciences
Computational Chemistry. An Introduction to Molecular Dynamic Simulations
 Computational Chemistry An Introduction to Molecular Dynamic Simulations Computational chemistry simulates chemical structures and reactions numerically, based in full or in part on the fundamental laws
Computational Chemistry An Introduction to Molecular Dynamic Simulations Computational chemistry simulates chemical structures and reactions numerically, based in full or in part on the fundamental laws
Session 1. Introduction to Computational Chemistry. Computational (chemistry education) and/or (Computational chemistry) education
 Session 1 Introduction to Computational Chemistry 1 Introduction to Computational Chemistry Computational (chemistry education) and/or (Computational chemistry) education First one: Use computational tools
Session 1 Introduction to Computational Chemistry 1 Introduction to Computational Chemistry Computational (chemistry education) and/or (Computational chemistry) education First one: Use computational tools
T6.2 Molecular Mechanics
 T6.2 Molecular Mechanics We have seen that Benson group additivities are capable of giving heats of formation of molecules with accuracies comparable to those of the best ab initio procedures. However,
T6.2 Molecular Mechanics We have seen that Benson group additivities are capable of giving heats of formation of molecules with accuracies comparable to those of the best ab initio procedures. However,
AN INTRODUCTION TO QUANTUM CHEMISTRY. Mark S. Gordon Iowa State University
 AN INTRODUCTION TO QUANTUM CHEMISTRY Mark S. Gordon Iowa State University 1 OUTLINE Theoretical Background in Quantum Chemistry Overview of GAMESS Program Applications 2 QUANTUM CHEMISTRY In principle,
AN INTRODUCTION TO QUANTUM CHEMISTRY Mark S. Gordon Iowa State University 1 OUTLINE Theoretical Background in Quantum Chemistry Overview of GAMESS Program Applications 2 QUANTUM CHEMISTRY In principle,
Basis Set for Molecular Orbital Theory
 Basis Set for Molecular Orbital Theory! Different Types of Basis Functions! Different Types of Atom Center Basis Functions! Classifications of Gaussian Basis Sets! Pseudopotentials! Molecular Properties
Basis Set for Molecular Orbital Theory! Different Types of Basis Functions! Different Types of Atom Center Basis Functions! Classifications of Gaussian Basis Sets! Pseudopotentials! Molecular Properties
Lecture 11: Potential Energy Functions
 Lecture 11: Potential Energy Functions Dr. Ronald M. Levy ronlevy@temple.edu Originally contributed by Lauren Wickstrom (2011) Microscopic/Macroscopic Connection The connection between microscopic interactions
Lecture 11: Potential Energy Functions Dr. Ronald M. Levy ronlevy@temple.edu Originally contributed by Lauren Wickstrom (2011) Microscopic/Macroscopic Connection The connection between microscopic interactions
Chemistry 4560/5560 Molecular Modeling Fall 2014
 Final Exam Name:. User s guide: 1. Read questions carefully and make sure you understand them before answering (if not, ask). 2. Answer only the question that is asked, not a different question. 3. Unless
Final Exam Name:. User s guide: 1. Read questions carefully and make sure you understand them before answering (if not, ask). 2. Answer only the question that is asked, not a different question. 3. Unless
Beyond the Hartree-Fock Approximation: Configuration Interaction
 Beyond the Hartree-Fock Approximation: Configuration Interaction The Hartree-Fock (HF) method uses a single determinant (single electronic configuration) description of the electronic wavefunction. For
Beyond the Hartree-Fock Approximation: Configuration Interaction The Hartree-Fock (HF) method uses a single determinant (single electronic configuration) description of the electronic wavefunction. For
Quantum Chemical Simulations and Descriptors. Dr. Antonio Chana, Dr. Mosè Casalegno
 Quantum Chemical Simulations and Descriptors Dr. Antonio Chana, Dr. Mosè Casalegno Classical Mechanics: basics It models real-world objects as point particles, objects with negligible size. The motion
Quantum Chemical Simulations and Descriptors Dr. Antonio Chana, Dr. Mosè Casalegno Classical Mechanics: basics It models real-world objects as point particles, objects with negligible size. The motion
Computational Material Science Part II. Ito Chao ( ) Institute of Chemistry Academia Sinica
 Computational Material Science Part II Ito Chao ( ) Institute of Chemistry Academia Sinica Ab Initio Implementations of Hartree-Fock Molecular Orbital Theory Fundamental assumption of HF theory: each electron
Computational Material Science Part II Ito Chao ( ) Institute of Chemistry Academia Sinica Ab Initio Implementations of Hartree-Fock Molecular Orbital Theory Fundamental assumption of HF theory: each electron
Close agreement between the orientation dependence of hydrogen bonds observed in protein structures and quantum mechanical calculations
 Close agreement between the orientation dependence of hydrogen bonds observed in protein structures and quantum mechanical calculations Alexandre V. Morozov, Tanja Kortemme, Kiril Tsemekhman, David Baker
Close agreement between the orientation dependence of hydrogen bonds observed in protein structures and quantum mechanical calculations Alexandre V. Morozov, Tanja Kortemme, Kiril Tsemekhman, David Baker
CE 530 Molecular Simulation
 1 CE 530 Molecular Simulation Lecture 14 Molecular Models David A. Kofke Department of Chemical Engineering SUNY Buffalo kofke@eng.buffalo.edu 2 Review Monte Carlo ensemble averaging, no dynamics easy
1 CE 530 Molecular Simulation Lecture 14 Molecular Models David A. Kofke Department of Chemical Engineering SUNY Buffalo kofke@eng.buffalo.edu 2 Review Monte Carlo ensemble averaging, no dynamics easy
Electronic structure calculations: fundamentals George C. Schatz Northwestern University
 Electronic structure calculations: fundamentals George C. Schatz Northwestern University Electronic Structure (often called Quantum Chemistry) calculations use quantum mechanics to determine the wavefunctions
Electronic structure calculations: fundamentals George C. Schatz Northwestern University Electronic Structure (often called Quantum Chemistry) calculations use quantum mechanics to determine the wavefunctions
The successful wavefunction can be written as a determinant: # 1 (2) # 2 (2) Electrons. This can be generalized to our 2N-electron wavefunction:
 T2. CNDO to AM1: The Semiempirical Molecular Orbital Models The discussion in sections T2.1 T2.3 applies also to ab initio molecular orbital calculations. T2.1 Slater Determinants Consider the general
T2. CNDO to AM1: The Semiempirical Molecular Orbital Models The discussion in sections T2.1 T2.3 applies also to ab initio molecular orbital calculations. T2.1 Slater Determinants Consider the general
( ) + ( ) + ( ) = 0.00
 Chem 3502/4502 Physical Chemistry II (Quantum Mechanics) 3 Credits Spring Semester 2006 Christopher J. Cramer Lecture 32, April 14, 2006 (Some material in this lecture has been adapted from Cramer, C.
Chem 3502/4502 Physical Chemistry II (Quantum Mechanics) 3 Credits Spring Semester 2006 Christopher J. Cramer Lecture 32, April 14, 2006 (Some material in this lecture has been adapted from Cramer, C.
Introduction to Computational Chemistry Computational (chemistry education) and/or. (Computational chemistry) education
 Introduction to Computational Chemistry Computational (chemistry education) and/or (Computational chemistry) education First one: Use computational tools to help increase student understanding of material
Introduction to Computational Chemistry Computational (chemistry education) and/or (Computational chemistry) education First one: Use computational tools to help increase student understanding of material
Exercise 1: Structure and dipole moment of a small molecule
 Introduction to computational chemistry Exercise 1: Structure and dipole moment of a small molecule Vesa Hänninen 1 Introduction In this exercise the equilibrium structure and the dipole moment of a small
Introduction to computational chemistry Exercise 1: Structure and dipole moment of a small molecule Vesa Hänninen 1 Introduction In this exercise the equilibrium structure and the dipole moment of a small
Chem 1140; Molecular Modeling
 P. Wipf 1 Chem 1140 $E = -! (q a + q b )" ab S ab + ab!k
P. Wipf 1 Chem 1140 $E = -! (q a + q b )" ab S ab + ab!k
Chemistry 334 Part 2: Computational Quantum Chemistry
 Chemistry 334 Part 2: Computational Quantum Chemistry 1. Definition Louis Scudiero, Ben Shepler and Kirk Peterson Washington State University January 2006 Computational chemistry is an area of theoretical
Chemistry 334 Part 2: Computational Quantum Chemistry 1. Definition Louis Scudiero, Ben Shepler and Kirk Peterson Washington State University January 2006 Computational chemistry is an area of theoretical
Reactivity and Organocatalysis. (Adalgisa Sinicropi and Massimo Olivucci)
 Reactivity and Organocatalysis (Adalgisa Sinicropi and Massimo Olivucci) The Aldol Reaction - O R 1 O R 1 + - O O OH * * H R 2 R 1 R 2 The (1957) Zimmerman-Traxler Transition State Counterion (e.g. Li
Reactivity and Organocatalysis (Adalgisa Sinicropi and Massimo Olivucci) The Aldol Reaction - O R 1 O R 1 + - O O OH * * H R 2 R 1 R 2 The (1957) Zimmerman-Traxler Transition State Counterion (e.g. Li
An Introduction to Quantum Chemistry and Potential Energy Surfaces. Benjamin G. Levine
 An Introduction to Quantum Chemistry and Potential Energy Surfaces Benjamin G. Levine This Week s Lecture Potential energy surfaces What are they? What are they good for? How do we use them to solve chemical
An Introduction to Quantum Chemistry and Potential Energy Surfaces Benjamin G. Levine This Week s Lecture Potential energy surfaces What are they? What are they good for? How do we use them to solve chemical
This is a very succinct primer intended as supplementary material for an undergraduate course in physical chemistry.
 1 Computational Chemistry (Quantum Chemistry) Primer This is a very succinct primer intended as supplementary material for an undergraduate course in physical chemistry. TABLE OF CONTENTS Methods...1 Basis
1 Computational Chemistry (Quantum Chemistry) Primer This is a very succinct primer intended as supplementary material for an undergraduate course in physical chemistry. TABLE OF CONTENTS Methods...1 Basis
Session 7 Overview: Part A I. Prediction of Vibrational Frequencies (IR) Part B III. Prediction of Electronic Transitions (UV-Vis) IV.
 Session 7 Overview: Part A I. Prediction of Vibrational Frequencies (IR) II. Thermochemistry Part B III. Prediction of Electronic Transitions (UV-Vis) IV. NMR Predictions 1 I. Prediction of Vibrational
Session 7 Overview: Part A I. Prediction of Vibrational Frequencies (IR) II. Thermochemistry Part B III. Prediction of Electronic Transitions (UV-Vis) IV. NMR Predictions 1 I. Prediction of Vibrational
Handbook of Computational Quantum Chemistry. DAVID B. COOK The Department of Chemistry, University of Sheffield
 Handbook of Computational Quantum Chemistry DAVID B. COOK The Department of Chemistry, University of Sheffield Oxford New York Tokyo OXFORD UNIVERSITY PRESS 1998 CONTENTS 1 Mechanics and molecules 1 1.1
Handbook of Computational Quantum Chemistry DAVID B. COOK The Department of Chemistry, University of Sheffield Oxford New York Tokyo OXFORD UNIVERSITY PRESS 1998 CONTENTS 1 Mechanics and molecules 1 1.1
Chemistry 3502/4502. Final Exam Part I. May 14, 2005
 Advocacy chit Chemistry 350/450 Final Exam Part I May 4, 005. For which of the below systems is = where H is the Hamiltonian operator and T is the kinetic-energy operator? (a) The free particle
Advocacy chit Chemistry 350/450 Final Exam Part I May 4, 005. For which of the below systems is = where H is the Hamiltonian operator and T is the kinetic-energy operator? (a) The free particle
Semi-Empirical MO Methods
 Semi-Empirical MO Methods the high cost of ab initio MO calculations is largely due to the many integrals that need to be calculated (esp. two electron integrals) semi-empirical MO methods start with the
Semi-Empirical MO Methods the high cost of ab initio MO calculations is largely due to the many integrals that need to be calculated (esp. two electron integrals) semi-empirical MO methods start with the
Structural Bioinformatics (C3210) Molecular Mechanics
 Structural Bioinformatics (C3210) Molecular Mechanics How to Calculate Energies Calculation of molecular energies is of key importance in protein folding, molecular modelling etc. There are two main computational
Structural Bioinformatics (C3210) Molecular Mechanics How to Calculate Energies Calculation of molecular energies is of key importance in protein folding, molecular modelling etc. There are two main computational
Introduction to Vibrational Spectroscopy
 Introduction to Vibrational Spectroscopy Harmonic oscillators The classical harmonic oscillator The uantum mechanical harmonic oscillator Harmonic approximations in molecular vibrations Vibrational spectroscopy
Introduction to Vibrational Spectroscopy Harmonic oscillators The classical harmonic oscillator The uantum mechanical harmonic oscillator Harmonic approximations in molecular vibrations Vibrational spectroscopy
Chemistry 3502/4502. Final Exam Part I. May 14, 2005
 Chemistry 3502/4502 Final Exam Part I May 14, 2005 1. For which of the below systems is = where H is the Hamiltonian operator and T is the kinetic-energy operator? (a) The free particle (e) The
Chemistry 3502/4502 Final Exam Part I May 14, 2005 1. For which of the below systems is = where H is the Hamiltonian operator and T is the kinetic-energy operator? (a) The free particle (e) The
CHEMISTRY 4021/8021 MIDTERM EXAM 1 SPRING 2014
 CHEMISTRY 4021/8021 Q1) Propose a simple, united-atom molecular mechanics force-field needed to generate a potential energy surface for an isolated molecule of acetone (Me 2 CO). I.e., provide an energy
CHEMISTRY 4021/8021 Q1) Propose a simple, united-atom molecular mechanics force-field needed to generate a potential energy surface for an isolated molecule of acetone (Me 2 CO). I.e., provide an energy
Introduction to computational chemistry Exercise I: Structure and electronic energy of a small molecule. Vesa Hänninen
 Introduction to computational chemistry Exercise I: Structure and electronic energy of a small molecule Vesa Hänninen 1 Introduction In this exercise the equilibrium structure and the electronic energy
Introduction to computational chemistry Exercise I: Structure and electronic energy of a small molecule Vesa Hänninen 1 Introduction In this exercise the equilibrium structure and the electronic energy
Why Is Molecular Interaction Important in Our Life
 Why Is Molecular Interaction Important in ur Life QuLiS and Graduate School of Science iroshima University http://www.nabit.hiroshima-u.ac.jp/iwatasue/indexe.htm Suehiro Iwata Sept. 29, 2007 Department
Why Is Molecular Interaction Important in ur Life QuLiS and Graduate School of Science iroshima University http://www.nabit.hiroshima-u.ac.jp/iwatasue/indexe.htm Suehiro Iwata Sept. 29, 2007 Department
Lecture 4: methods and terminology, part II
 So theory guys have got it made in rooms free of pollution. Instead of problems with the reflux, they have only solutions... In other words, experimentalists will likely die of cancer From working hard,
So theory guys have got it made in rooms free of pollution. Instead of problems with the reflux, they have only solutions... In other words, experimentalists will likely die of cancer From working hard,
Molecular Mechanics. C. David Sherrill School of Chemistry and Biochemistry Georgia Institute of Technology. January 2001
 Molecular Mechanics C. David Sherrill School of Chemistry and Biochemistry Georgia Institute of Technology January 2001 Introduction Molecular Mechanics uses classical type models to predict the energy
Molecular Mechanics C. David Sherrill School of Chemistry and Biochemistry Georgia Institute of Technology January 2001 Introduction Molecular Mechanics uses classical type models to predict the energy
Scuola di Chimica Computazionale
 Societa Chimica Italiana Gruppo Interdivisionale di Chimica Computazionale Scuola di Chimica Computazionale Introduzione, per Esercizi, all Uso del Calcolatore in Chimica Organica e Biologica Modellistica
Societa Chimica Italiana Gruppo Interdivisionale di Chimica Computazionale Scuola di Chimica Computazionale Introduzione, per Esercizi, all Uso del Calcolatore in Chimica Organica e Biologica Modellistica
Introduction to Computational Quantum Chemistry: Theory
 Introduction to Computational Quantum Chemistry: Theory Dr Andrew Gilbert Rm 118, Craig Building, RSC 3108 Course Lectures 2007 Introduction Hartree Fock Theory Configuration Interaction Lectures 1 Introduction
Introduction to Computational Quantum Chemistry: Theory Dr Andrew Gilbert Rm 118, Craig Building, RSC 3108 Course Lectures 2007 Introduction Hartree Fock Theory Configuration Interaction Lectures 1 Introduction
Introduction to Computational Chemistry: Theory
 Introduction to Computational Chemistry: Theory Dr Andrew Gilbert Rm 118, Craig Building, RSC andrew.gilbert@anu.edu.au 3023 Course Lectures Introduction Hartree Fock Theory Basis Sets Lecture 1 1 Introduction
Introduction to Computational Chemistry: Theory Dr Andrew Gilbert Rm 118, Craig Building, RSC andrew.gilbert@anu.edu.au 3023 Course Lectures Introduction Hartree Fock Theory Basis Sets Lecture 1 1 Introduction
Same idea for polyatomics, keep track of identical atom e.g. NH 3 consider only valence electrons F(2s,2p) H(1s)
 XIII 63 Polyatomic bonding -09 -mod, Notes (13) Engel 16-17 Balance: nuclear repulsion, positive e-n attraction, neg. united atom AO ε i applies to all bonding, just more nuclei repulsion biggest at low
XIII 63 Polyatomic bonding -09 -mod, Notes (13) Engel 16-17 Balance: nuclear repulsion, positive e-n attraction, neg. united atom AO ε i applies to all bonding, just more nuclei repulsion biggest at low
Yingwei Wang Computational Quantum Chemistry 1 Hartree energy 2. 2 Many-body system 2. 3 Born-Oppenheimer approximation 2
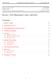 Purdue University CHM 67300 Computational Quantum Chemistry REVIEW Yingwei Wang October 10, 2013 Review: Prof Slipchenko s class, Fall 2013 Contents 1 Hartree energy 2 2 Many-body system 2 3 Born-Oppenheimer
Purdue University CHM 67300 Computational Quantum Chemistry REVIEW Yingwei Wang October 10, 2013 Review: Prof Slipchenko s class, Fall 2013 Contents 1 Hartree energy 2 2 Many-body system 2 3 Born-Oppenheimer
Introduction to DFTB. Marcus Elstner. July 28, 2006
 Introduction to DFTB Marcus Elstner July 28, 2006 I. Non-selfconsistent solution of the KS equations DFT can treat up to 100 atoms in routine applications, sometimes even more and about several ps in MD
Introduction to DFTB Marcus Elstner July 28, 2006 I. Non-selfconsistent solution of the KS equations DFT can treat up to 100 atoms in routine applications, sometimes even more and about several ps in MD
All-atom Molecular Mechanics. Trent E. Balius AMS 535 / CHE /27/2010
 All-atom Molecular Mechanics Trent E. Balius AMS 535 / CHE 535 09/27/2010 Outline Molecular models Molecular mechanics Force Fields Potential energy function functional form parameters and parameterization
All-atom Molecular Mechanics Trent E. Balius AMS 535 / CHE 535 09/27/2010 Outline Molecular models Molecular mechanics Force Fields Potential energy function functional form parameters and parameterization
Quantum mechanics can be used to calculate any property of a molecule. The energy E of a wavefunction Ψ evaluated for the Hamiltonian H is,
 Chapter : Molecules Quantum mechanics can be used to calculate any property of a molecule The energy E of a wavefunction Ψ evaluated for the Hamiltonian H is, E = Ψ H Ψ Ψ Ψ 1) At first this seems like
Chapter : Molecules Quantum mechanics can be used to calculate any property of a molecule The energy E of a wavefunction Ψ evaluated for the Hamiltonian H is, E = Ψ H Ψ Ψ Ψ 1) At first this seems like
Density Functional Theory
 Chemistry 380.37 Fall 2015 Dr. Jean M. Standard October 28, 2015 Density Functional Theory What is a Functional? A functional is a general mathematical quantity that represents a rule to convert a function
Chemistry 380.37 Fall 2015 Dr. Jean M. Standard October 28, 2015 Density Functional Theory What is a Functional? A functional is a general mathematical quantity that represents a rule to convert a function
Ab initio calculations for potential energy surfaces. D. Talbi GRAAL- Montpellier
 Ab initio calculations for potential energy surfaces D. Talbi GRAAL- Montpellier A theoretical study of a reaction is a two step process I-Electronic calculations : techniques of quantum chemistry potential
Ab initio calculations for potential energy surfaces D. Talbi GRAAL- Montpellier A theoretical study of a reaction is a two step process I-Electronic calculations : techniques of quantum chemistry potential
Molecular Mechanics, Dynamics & Docking
 Molecular Mechanics, Dynamics & Docking Lawrence Hunter, Ph.D. Director, Computational Bioscience Program University of Colorado School of Medicine Larry.Hunter@uchsc.edu http://compbio.uchsc.edu/hunter
Molecular Mechanics, Dynamics & Docking Lawrence Hunter, Ph.D. Director, Computational Bioscience Program University of Colorado School of Medicine Larry.Hunter@uchsc.edu http://compbio.uchsc.edu/hunter
Atomic and molecular interaction forces in biology
 Atomic and molecular interaction forces in biology 1 Outline Types of interactions relevant to biology Van der Waals interactions H-bond interactions Some properties of water Hydrophobic effect 2 Types
Atomic and molecular interaction forces in biology 1 Outline Types of interactions relevant to biology Van der Waals interactions H-bond interactions Some properties of water Hydrophobic effect 2 Types
Lecture 3: methods and terminology, part I
 Eritis sicut Deus, scientes bonum et malum. Lecture 3: methods and terminology, part I Molecular dynamics, internal coordinates, Z-matrices, force fields, periodic boundary conditions, Hartree-Fock theory,
Eritis sicut Deus, scientes bonum et malum. Lecture 3: methods and terminology, part I Molecular dynamics, internal coordinates, Z-matrices, force fields, periodic boundary conditions, Hartree-Fock theory,
MO Calculation for a Diatomic Molecule. /4 0 ) i=1 j>i (1/r ij )
 MO Calculation for a Diatomic Molecule Introduction The properties of any molecular system can in principle be found by looking at the solutions to the corresponding time independent Schrodinger equation
MO Calculation for a Diatomic Molecule Introduction The properties of any molecular system can in principle be found by looking at the solutions to the corresponding time independent Schrodinger equation
Density Functional Theory
 Density Functional Theory March 26, 2009 ? DENSITY FUNCTIONAL THEORY is a method to successfully describe the behavior of atomic and molecular systems and is used for instance for: structural prediction
Density Functional Theory March 26, 2009 ? DENSITY FUNCTIONAL THEORY is a method to successfully describe the behavior of atomic and molecular systems and is used for instance for: structural prediction
( R)Ψ el ( r;r) = E el ( R)Ψ el ( r;r)
 Born Oppenheimer Approximation: Ĥ el ( R)Ψ el ( r;r) = E el ( R)Ψ el ( r;r) For a molecule with N electrons and M nuclei: Ĥ el What is E el (R)? s* potential surface Reaction Barrier Unstable intermediate
Born Oppenheimer Approximation: Ĥ el ( R)Ψ el ( r;r) = E el ( R)Ψ el ( r;r) For a molecule with N electrons and M nuclei: Ĥ el What is E el (R)? s* potential surface Reaction Barrier Unstable intermediate
Molecular Mechanics. Yohann Moreau. November 26, 2015
 Molecular Mechanics Yohann Moreau yohann.moreau@ujf-grenoble.fr November 26, 2015 Yohann Moreau (UJF) Molecular Mechanics, Label RFCT 2015 November 26, 2015 1 / 29 Introduction A so-called Force-Field
Molecular Mechanics Yohann Moreau yohann.moreau@ujf-grenoble.fr November 26, 2015 Yohann Moreau (UJF) Molecular Mechanics, Label RFCT 2015 November 26, 2015 1 / 29 Introduction A so-called Force-Field
Lec20 Fri 3mar17
 564-17 Lec20 Fri 3mar17 [PDF]GAUSSIAN 09W TUTORIAL www.molcalx.com.cn/wp-content/uploads/2015/01/gaussian09w_tutorial.pdf by A Tomberg - Cited by 8 - Related articles GAUSSIAN 09W TUTORIAL. AN INTRODUCTION
564-17 Lec20 Fri 3mar17 [PDF]GAUSSIAN 09W TUTORIAL www.molcalx.com.cn/wp-content/uploads/2015/01/gaussian09w_tutorial.pdf by A Tomberg - Cited by 8 - Related articles GAUSSIAN 09W TUTORIAL. AN INTRODUCTION
Jack Simons, Henry Eyring Scientist and Professor Chemistry Department University of Utah
 1. Born-Oppenheimer approx.- energy surfaces 2. Mean-field (Hartree-Fock) theory- orbitals 3. Pros and cons of HF- RHF, UHF 4. Beyond HF- why? 5. First, one usually does HF-how? 6. Basis sets and notations
1. Born-Oppenheimer approx.- energy surfaces 2. Mean-field (Hartree-Fock) theory- orbitals 3. Pros and cons of HF- RHF, UHF 4. Beyond HF- why? 5. First, one usually does HF-how? 6. Basis sets and notations
Molecular Mechanics. I. Quantum mechanical treatment of molecular systems
 Molecular Mechanics I. Quantum mechanical treatment of molecular systems The first principle approach for describing the properties of molecules, including proteins, involves quantum mechanics. For example,
Molecular Mechanics I. Quantum mechanical treatment of molecular systems The first principle approach for describing the properties of molecules, including proteins, involves quantum mechanics. For example,
Gaussian: Basic Tutorial
 Input file: # hf sto-g pop=full Water - Single Point Energy 0 H.0 H.0 H 04.5 Route Section Start with # Contains the keywords Gaussian: Basic Tutorial Route Section Title Section Charge-Multiplicity Molecule
Input file: # hf sto-g pop=full Water - Single Point Energy 0 H.0 H.0 H 04.5 Route Section Start with # Contains the keywords Gaussian: Basic Tutorial Route Section Title Section Charge-Multiplicity Molecule
Chemistry 4681 Module: Electronic Structure of Small Molecules Background Handout
 Chemistry 4681 Module: Electronic Structure of Small Molecules Background Handout C. David Sherrill Last Revised: 6 January 2000 1 Computational Chemistry The term computational chemistry is used to mean
Chemistry 4681 Module: Electronic Structure of Small Molecules Background Handout C. David Sherrill Last Revised: 6 January 2000 1 Computational Chemistry The term computational chemistry is used to mean
Bioengineering 215. An Introduction to Molecular Dynamics for Biomolecules
 Bioengineering 215 An Introduction to Molecular Dynamics for Biomolecules David Parker May 18, 2007 ntroduction A principal tool to study biological molecules is molecular dynamics simulations (MD). MD
Bioengineering 215 An Introduction to Molecular Dynamics for Biomolecules David Parker May 18, 2007 ntroduction A principal tool to study biological molecules is molecular dynamics simulations (MD). MD
Molecular mechanics. classical description of molecules. Marcus Elstner and Tomáš Kubař. April 29, 2016
 classical description of molecules April 29, 2016 Chemical bond Conceptual and chemical basis quantum effect solution of the SR numerically expensive (only small molecules can be treated) approximations
classical description of molecules April 29, 2016 Chemical bond Conceptual and chemical basis quantum effect solution of the SR numerically expensive (only small molecules can be treated) approximations
DFT calculations of NMR indirect spin spin coupling constants
 DFT calculations of NMR indirect spin spin coupling constants Dalton program system Program capabilities Density functional theory Kohn Sham theory LDA, GGA and hybrid theories Indirect NMR spin spin coupling
DFT calculations of NMR indirect spin spin coupling constants Dalton program system Program capabilities Density functional theory Kohn Sham theory LDA, GGA and hybrid theories Indirect NMR spin spin coupling
Walter Kohn was awarded with the Nobel Prize in Chemistry in 1998 for his development of the density functional theory.
 Walter Kohn was awarded with the Nobel Prize in Chemistry in 1998 for his development of the density functional theory. Walter Kohn receiving his Nobel Prize from His Majesty the King at the Stockholm
Walter Kohn was awarded with the Nobel Prize in Chemistry in 1998 for his development of the density functional theory. Walter Kohn receiving his Nobel Prize from His Majesty the King at the Stockholm
Chem 442 Review for Exam 2. Exact separation of the Hamiltonian of a hydrogenic atom into center-of-mass (3D) and relative (3D) components.
 Chem 44 Review for Exam Hydrogenic atoms: The Coulomb energy between two point charges Ze and e: V r Ze r Exact separation of the Hamiltonian of a hydrogenic atom into center-of-mass (3D) and relative
Chem 44 Review for Exam Hydrogenic atoms: The Coulomb energy between two point charges Ze and e: V r Ze r Exact separation of the Hamiltonian of a hydrogenic atom into center-of-mass (3D) and relative
Figure 1: Transition State, Saddle Point, Reaction Pathway
 Computational Chemistry Workshops West Ridge Research Building-UAF Campus 9:00am-4:00pm, Room 009 Electronic Structure - July 19-21, 2016 Molecular Dynamics - July 26-28, 2016 Potential Energy Surfaces
Computational Chemistry Workshops West Ridge Research Building-UAF Campus 9:00am-4:00pm, Room 009 Electronic Structure - July 19-21, 2016 Molecular Dynamics - July 26-28, 2016 Potential Energy Surfaces
Lecture 10. Born-Oppenheimer approximation LCAO-MO application to H + The potential energy surface MOs for diatomic molecules. NC State University
 Chemistry 431 Lecture 10 Diatomic molecules Born-Oppenheimer approximation LCAO-MO application to H + 2 The potential energy surface MOs for diatomic molecules NC State University Born-Oppenheimer approximation
Chemistry 431 Lecture 10 Diatomic molecules Born-Oppenheimer approximation LCAO-MO application to H + 2 The potential energy surface MOs for diatomic molecules NC State University Born-Oppenheimer approximation
Semi-Empirical Methods CHEM 430
 Semi-Empirical Methods CHEM 430 Cost, Hartree%Fock, scales,as,n 4,(N=#, basis,funcfons), Due,to,two% electron, integrals, within,fock, matrix, Semi%empirical,cut, cost,by,reducing, number,of, integrals,
Semi-Empirical Methods CHEM 430 Cost, Hartree%Fock, scales,as,n 4,(N=#, basis,funcfons), Due,to,two% electron, integrals, within,fock, matrix, Semi%empirical,cut, cost,by,reducing, number,of, integrals,
k θ (θ θ 0 ) 2 angles r i j r i j
 1 Force fields 1.1 Introduction The term force field is slightly misleading, since it refers to the parameters of the potential used to calculate the forces (via gradient) in molecular dynamics simulations.
1 Force fields 1.1 Introduction The term force field is slightly misleading, since it refers to the parameters of the potential used to calculate the forces (via gradient) in molecular dynamics simulations.
Handbook of Computational Quantum Chemistry
 Handbook of Computational Quantum Chemistry David B. Cook Dept. of Chemistry University of Sheffield DOVER PUBLICATIONS, INC. Mineola, New York F Contents 1 Mechanics and molecules 1 1.1 1.2 1.3 1.4 1.5
Handbook of Computational Quantum Chemistry David B. Cook Dept. of Chemistry University of Sheffield DOVER PUBLICATIONS, INC. Mineola, New York F Contents 1 Mechanics and molecules 1 1.1 1.2 1.3 1.4 1.5
one ν im: transition state saddle point
 Hypothetical Potential Energy Surface Ethane conformations Hartree-Fock theory, basis set stationary points all ν s >0: minimum eclipsed one ν im: transition state saddle point multiple ν im: hilltop 1
Hypothetical Potential Energy Surface Ethane conformations Hartree-Fock theory, basis set stationary points all ν s >0: minimum eclipsed one ν im: transition state saddle point multiple ν im: hilltop 1
Lecture 9. Hartree Fock Method and Koopman s Theorem
 Lecture 9 Hartree Fock Method and Koopman s Theorem Ψ(N) is approximated as a single slater determinant Φ of N orthogonal One electron spin-orbitals. One electron orbital φ i = φ i (r) χ i (σ) χ i (σ)
Lecture 9 Hartree Fock Method and Koopman s Theorem Ψ(N) is approximated as a single slater determinant Φ of N orthogonal One electron spin-orbitals. One electron orbital φ i = φ i (r) χ i (σ) χ i (σ)
Electronic structure theory: Fundamentals to frontiers. 1. Hartree-Fock theory
 Electronic structure theory: Fundamentals to frontiers. 1. Hartree-Fock theory MARTIN HEAD-GORDON, Department of Chemistry, University of California, and Chemical Sciences Division, Lawrence Berkeley National
Electronic structure theory: Fundamentals to frontiers. 1. Hartree-Fock theory MARTIN HEAD-GORDON, Department of Chemistry, University of California, and Chemical Sciences Division, Lawrence Berkeley National
Electron Correlation
 Electron Correlation Levels of QM Theory HΨ=EΨ Born-Oppenheimer approximation Nuclear equation: H n Ψ n =E n Ψ n Electronic equation: H e Ψ e =E e Ψ e Single determinant SCF Semi-empirical methods Correlation
Electron Correlation Levels of QM Theory HΨ=EΨ Born-Oppenheimer approximation Nuclear equation: H n Ψ n =E n Ψ n Electronic equation: H e Ψ e =E e Ψ e Single determinant SCF Semi-empirical methods Correlation
The Potential Energy Surface (PES) And the Basic Force Field Chem 4021/8021 Video II.iii
 The Potential Energy Surface (PES) And the Basic Force Field Chem 4021/8021 Video II.iii Fundamental Points About Which to Be Thinking It s clear the PES is useful, so how can I construct it for an arbitrary
The Potential Energy Surface (PES) And the Basic Force Field Chem 4021/8021 Video II.iii Fundamental Points About Which to Be Thinking It s clear the PES is useful, so how can I construct it for an arbitrary
LUMO + 1 LUMO. Tómas Arnar Guðmundsson Report 2 Reikniefnafræði G
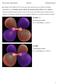 Q1: Display all the MOs for N2 in your report and classify each one of them as bonding, antibonding or non-bonding, and say whether the symmetry of the orbital is σ or π. Sketch a molecular orbital diagram
Q1: Display all the MOs for N2 in your report and classify each one of them as bonding, antibonding or non-bonding, and say whether the symmetry of the orbital is σ or π. Sketch a molecular orbital diagram
Introduction to Computational Chemistry for Experimental Chemists... (Part 2/2)
 12 th PhD seminar, Garching, October 31 st 2008 Introduction to Computational Chemistry for Experimental Chemists... (Part 2/2) Dr. Markus Drees, TU München Universität Regensburg Universität Augsburg
12 th PhD seminar, Garching, October 31 st 2008 Introduction to Computational Chemistry for Experimental Chemists... (Part 2/2) Dr. Markus Drees, TU München Universität Regensburg Universität Augsburg
SCF calculation on HeH +
 SCF calculation on HeH + Markus Meuwly Department of Chemistry, University of Basel, Basel, Switzerland Abstract This document describes the main steps involved in carrying out a SCF calculation on the
SCF calculation on HeH + Markus Meuwly Department of Chemistry, University of Basel, Basel, Switzerland Abstract This document describes the main steps involved in carrying out a SCF calculation on the
Advanced Electronic Structure Theory Density functional theory. Dr Fred Manby
 Advanced Electronic Structure Theory Density functional theory Dr Fred Manby fred.manby@bris.ac.uk http://www.chm.bris.ac.uk/pt/manby/ 6 Strengths of DFT DFT is one of many theories used by (computational)
Advanced Electronic Structure Theory Density functional theory Dr Fred Manby fred.manby@bris.ac.uk http://www.chm.bris.ac.uk/pt/manby/ 6 Strengths of DFT DFT is one of many theories used by (computational)
Project 1 Report File: Chem4PB3_project_1_2017-solutions last changed: 02-Feb-2017
 Project 1 Report File: Chem4PB3_project_1_2017-solutions last changed: 02-Feb-2017 1. Formaldehyde comparison of results from different methods Yellow shaded boxes are closest to experimental Method #e-
Project 1 Report File: Chem4PB3_project_1_2017-solutions last changed: 02-Feb-2017 1. Formaldehyde comparison of results from different methods Yellow shaded boxes are closest to experimental Method #e-
Homework Problem Set 5 Solutions. E e + H corr (a.u.) a.) Determine the bond dissociation enthalpy of ethane in kcal/mol (1 a.u. = kcal/mol).
 Chemistry 380.37 Dr. Jean M. Standard Homework Problem Set 5 Solutions 1. Given below are the sum of electronic and thermal enthalpies, E e + H corr, from Hartree-Fock calculations using a 6-31G(d) basis
Chemistry 380.37 Dr. Jean M. Standard Homework Problem Set 5 Solutions 1. Given below are the sum of electronic and thermal enthalpies, E e + H corr, from Hartree-Fock calculations using a 6-31G(d) basis
Structure of Cement Phases from ab initio Modeling Crystalline C-S-HC
 Structure of Cement Phases from ab initio Modeling Crystalline C-S-HC Sergey V. Churakov sergey.churakov@psi.ch Paul Scherrer Institute Switzerland Cement Phase Composition C-S-H H Solid Solution Model
Structure of Cement Phases from ab initio Modeling Crystalline C-S-HC Sergey V. Churakov sergey.churakov@psi.ch Paul Scherrer Institute Switzerland Cement Phase Composition C-S-H H Solid Solution Model
Practical 1: Structure and electronic properties of organic molecules. B/ Structure, electronic and vibrational properties of the water molecule
 D1CH9116 - MMCO Molecular Modelling applied to organic chemistry Practical 1: tructure and electronic properties of organic molecules B/ tructure, electronic and vibrational properties of the water molecule
D1CH9116 - MMCO Molecular Modelling applied to organic chemistry Practical 1: tructure and electronic properties of organic molecules B/ tructure, electronic and vibrational properties of the water molecule
TDDFT in Chemistry and Biochemistry III
 TDDFT in Chemistry and Biochemistry III Dmitrij Rappoport Department of Chemistry and Chemical Biology Harvard University TDDFT Winter School Benasque, January 2010 Dmitrij Rappoport (Harvard U.) TDDFT
TDDFT in Chemistry and Biochemistry III Dmitrij Rappoport Department of Chemistry and Chemical Biology Harvard University TDDFT Winter School Benasque, January 2010 Dmitrij Rappoport (Harvard U.) TDDFT
OVERVIEW OF QUANTUM CHEMISTRY METHODS
 OVERVIEW OF QUANTUM CHEMISTRY METHODS Outline I Generalities Correlation, basis sets Spin II Wavefunction methods Hartree-Fock Configuration interaction Coupled cluster Perturbative methods III Density
OVERVIEW OF QUANTUM CHEMISTRY METHODS Outline I Generalities Correlation, basis sets Spin II Wavefunction methods Hartree-Fock Configuration interaction Coupled cluster Perturbative methods III Density
Electron Correlation Methods
 Electron Correlation Methods HF method: electron-electron interaction is replaced by an average interaction E HF c = E 0 E HF E 0 exact ground state energy E HF HF energy for a given basis set HF E c
Electron Correlation Methods HF method: electron-electron interaction is replaced by an average interaction E HF c = E 0 E HF E 0 exact ground state energy E HF HF energy for a given basis set HF E c
Jack Simons, Henry Eyring Scientist and Professor Chemistry Department University of Utah
 1. Born-Oppenheimer approx.- energy surfaces 2. Mean-field (Hartree-Fock) theory- orbitals 3. Pros and cons of HF- RHF, UHF 4. Beyond HF- why? 5. First, one usually does HF-how? 6. Basis sets and notations
1. Born-Oppenheimer approx.- energy surfaces 2. Mean-field (Hartree-Fock) theory- orbitals 3. Pros and cons of HF- RHF, UHF 4. Beyond HF- why? 5. First, one usually does HF-how? 6. Basis sets and notations
GEM4 Summer School OpenCourseWare
 GEM4 Summer School OpenCourseWare http://gem4.educommons.net/ http://www.gem4.org/ Lecture: Molecular Mechanics by Ju Li. Given August 9, 2006 during the GEM4 session at MIT in Cambridge, MA. Please use
GEM4 Summer School OpenCourseWare http://gem4.educommons.net/ http://www.gem4.org/ Lecture: Molecular Mechanics by Ju Li. Given August 9, 2006 during the GEM4 session at MIT in Cambridge, MA. Please use
Semi Empirical Force Fields and Their Limitations. Potential Energy Surface (PES)
 Semi Empirical Force Fields and Their Limitations Ioan Kosztin Beckman Institute University of Illinois at Urbana-Champaign Potential Energy Surface (PES) Schrödinger equation: H T Ψ( r, = E Ψ( r, H =
Semi Empirical Force Fields and Their Limitations Ioan Kosztin Beckman Institute University of Illinois at Urbana-Champaign Potential Energy Surface (PES) Schrödinger equation: H T Ψ( r, = E Ψ( r, H =
with the larger dimerization energy also exhibits the larger structural changes.
 A7. Looking at the image and table provided below, it is apparent that the monomer and dimer are structurally almost identical. Although angular and dihedral data were not included, these data are also
A7. Looking at the image and table provided below, it is apparent that the monomer and dimer are structurally almost identical. Although angular and dihedral data were not included, these data are also
NMR and IR spectra & vibrational analysis
 Lab 5: NMR and IR spectra & vibrational analysis A brief theoretical background 1 Some of the available chemical quantum methods for calculating NMR chemical shifts are based on the Hartree-Fock self-consistent
Lab 5: NMR and IR spectra & vibrational analysis A brief theoretical background 1 Some of the available chemical quantum methods for calculating NMR chemical shifts are based on the Hartree-Fock self-consistent
Density Functional Theory. Martin Lüders Daresbury Laboratory
 Density Functional Theory Martin Lüders Daresbury Laboratory Ab initio Calculations Hamiltonian: (without external fields, non-relativistic) impossible to solve exactly!! Electrons Nuclei Electron-Nuclei
Density Functional Theory Martin Lüders Daresbury Laboratory Ab initio Calculations Hamiltonian: (without external fields, non-relativistic) impossible to solve exactly!! Electrons Nuclei Electron-Nuclei
Subject of the Lecture:
 Subject of the Lecture: Conceptual basis for the development of force fields. Implementation/validation Water - a worked example Extensions - combining molecular mechanics and quantum mechanics (QM/MM)
Subject of the Lecture: Conceptual basis for the development of force fields. Implementation/validation Water - a worked example Extensions - combining molecular mechanics and quantum mechanics (QM/MM)
Chem 4502 Introduction to Quantum Mechanics and Spectroscopy 3 Credits Fall Semester 2014 Laura Gagliardi. Lecture 28, December 08, 2014
 Chem 4502 Introduction to Quantum Mechanics and Spectroscopy 3 Credits Fall Semester 2014 Laura Gagliardi Lecture 28, December 08, 2014 Solved Homework Water, H 2 O, involves 2 hydrogen atoms and an oxygen
Chem 4502 Introduction to Quantum Mechanics and Spectroscopy 3 Credits Fall Semester 2014 Laura Gagliardi Lecture 28, December 08, 2014 Solved Homework Water, H 2 O, involves 2 hydrogen atoms and an oxygen
Transition states and reaction paths
 Transition states and reaction paths Lab 4 Theoretical background Transition state A transition structure is the molecular configuration that separates reactants and products. In a system with a single
Transition states and reaction paths Lab 4 Theoretical background Transition state A transition structure is the molecular configuration that separates reactants and products. In a system with a single
Introduction to Geometry Optimization. Computational Chemistry lab 2009
 Introduction to Geometry Optimization Computational Chemistry lab 9 Determination of the molecule configuration H H Diatomic molecule determine the interatomic distance H O H Triatomic molecule determine
Introduction to Geometry Optimization Computational Chemistry lab 9 Determination of the molecule configuration H H Diatomic molecule determine the interatomic distance H O H Triatomic molecule determine
Computational Chemistry Using the University of Alaska WebMO Site
 2/7/2017 1 Computational Chemistry Using the University of Alaska WebMO Site John Keller Department of Chemistry & Biochemistry University of Alaska Fairbanks Intro and operation of WebMO and MOPAC Basic
2/7/2017 1 Computational Chemistry Using the University of Alaska WebMO Site John Keller Department of Chemistry & Biochemistry University of Alaska Fairbanks Intro and operation of WebMO and MOPAC Basic
The digital copy of this thesis is protected by the Copyright Act 1994 (New Zealand).
 http://waikato.researchgateway.ac.nz/ Research Commons at the University of Waikato Copyright Statement: The digital copy of this thesis is protected by the Copyright Act 1994 (New Zealand). The thesis
http://waikato.researchgateway.ac.nz/ Research Commons at the University of Waikato Copyright Statement: The digital copy of this thesis is protected by the Copyright Act 1994 (New Zealand). The thesis
