Apparent persistence length renormalization of bent DNA
|
|
|
- Clyde Terry
- 5 years ago
- Views:
Transcription
1 PHYSICAL REVIEW E 7, Apparent persistence length renormalization of bent DNA Igor M. Kulić,* Hervé Mohrbach, Vladimir Lobaskin, Rochish Thaokar, and Helmut Schiessel Max-Planck-Institute for Polymer Research, Theory Group, PO Box 3148, D 5501 Mainz, Germany Received 6 January 005; published 6 October 005 We derive the single molecule equation of state force-extension relation for DNA molecules bearing sliding loops and deflection defects. Analytical results are obtained in the large force limit by employing an analogy with instantons in quantum mechanical tunneling problems. The results reveal a remarkable feature of sliding loops an apparent strong reduction of the persistence length. We generalize these results to several other experimentally interesting situations ranging from rigid DNA-protein loops to the problem of anchoring deflections in atomic force microscopy stretching of semiflexible polymers. Expressions relating the forceextension measurements to the underlying loop or boundary deflection geometry are provided and applied to the case of the gal repressor dimer protein. The theoreticaredictions are complemented and quantitatively confirmed by molecular dynamics simulations. DOI: /PhysRevE PACS numbers: La, a, 8.37.Rs In nature, DNA is rarely found in its straight naked state as usually depicted in the introductory pages of elementary textbooks. In most in vivo situations, an overwhelming fraction of DNA is rather strongly configurationally constrained by binding proteins causing loops, bends, and deflections. The advent of single molecule stretching techniques 1 has opened the possibility of measuring the equation of state of single tethered DNA molecules in a variety of different conditions. While the statistical mechanics of unconstrained DNA under tension is theoretically well understood in the framework of the wormlike chain WLC model 3 the presence of topological constraints such as supercoiling 4,5 or geometrical constraints such as protein-induced kinks and bends 6 9 renders analytical results more difficult. In this paper we expand the repertoire of analytically solvable equations of state by deriving the force-extension relation for a DNA molecule featuring loops and large deflections in the limit of strong stretching forces for the small forces case see 7. The computation is performed by evaluating quadratic fluctuations around the looped solution a nonconstant saddle point of the DNA elastic energy. The method is essentially analogous to the semiclassical treatment of tunneling amplitudes in quantum mechanics and instantons in quantum field theory 10. After deriving the general result that we accompany with molecular dynamics MD simulations we focus on two interesting experimental applications: the stretching of the gal repressor GalR loop complex 11, and of tangentially anchored semiflexible polymers from a surface in atomic force microscopy AFM experiments. Stretching a sliding loop. In the following we neglect the DNA twist degree of freedom for it is not constrained from outside and no external torsional torques are acting on it. In *Present address: Department of Physics and Astronomy, University of Pennsylvania, Philadelphia, PA 19104, USA. Present address: Instituut-Lorentz, Universiteit Leiden, Postbus 9506, 300 RA Leiden, The Netherlands. Present address: Department of Chemical Engineering, Indian Institute of Technology, Bombay, Mumbai, , India. this case, DNA of length L is described in the continuum limit by specifying only the unit vector tangent ts to the chain. Here s is the contour length along the DNA with L/sL/. The chain is submitted to an external constant force so that the kinetic plus potential energy of such a chain reads L/ E 0 = L/ ds A dt F tds with the bending stiffness A=l P k B T where l P is the orientational persistence length and k B T the thermal energy; for DNA at room temperature l P 50 nm 1. In the following we parametrize the tangent as t =cos cos,sin cos,sin and put the force along the x axis so that the potential energy part writes F cos cos. Note that the angle is measured with respect to the equatorial plane as on a globe. This parametrization is necessary to take properly into account the inextensibility of the chain imposed by the condition t =1. In the following it is convenient to introduce the dimensionless contour length t=s/ with the deflection length = A/F 6,13. The latter becomes the relevant length scale characterizing the loss of orientational correlation in the case of DNA under large tension replacing the usual tension-free persistence length l P =A/k B T. In these coordinates the chain energy writes L/ E 0 = 1 AF L/ cos + cos cos dt. 1 The relevant saddle point the loop in the x-y plane shown in Fig. 1a satisfies E 0 =0 with the boundary conditions loop ±L/=0, loop ±L/=0, and. In the limit of large lengths L/ the loop solution is the wellknown kink solution from theory of quantum tunneling 10 cos loop =1 cosh t and loop =0 with the loop energy given by E loop =8 FA LF. For large values of AF we can expand E 0 in terms of quadratic fluctuations, around the looped saddle point loop, loop /005/74/ /$ The American Physical Society
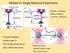
2 KULIĆ et al. FIG. 1. Color online Examples for large deflections in a stretched DNA molecules. a A freely sliding linker protein stabilizes a DNA loop. b A rigid ligand with opening angle causes a kink in the DNA. c Tangentially anchored DNA stretched from a surface by an AFM tip. The tilting angle as well as anchoring angles and can strongly affect the elastic response. E 0 = E loop + AF Tˆ dt + AF Tˆ dt, with the in- and out-of-plane fluctuation operators Tˆ = /t +1 cosh t and Tˆ = /t +1 6 cosh t, respectively. A closer inspection of the discrete spectrum of the two operators reveals the physics behind. Tˆ has a zero eigenvalue resulting from the translational shift invariance of the loop along the chain that costs no energy for L/. The absence of negative eigenvalues is in agreement with intuition, as in two dimensions, the loop is a topologically stable saddle point. The out-of-plane fluctuation operator Tˆ shows a richer behavior. Again it possesses a zero mode, this time resulting from rotational invariance of the problem. More remarkably, in contrast to Tˆ the out-ofplane fluctuation operator Tˆ has a negative eigenvalue 3. The latter underlines the intrinsic instability of the loop in three dimensions as described by the elastic energy expressions 1 and. This brings us to the question how to describe the obviously stable physical situation shown in Fig. 1a. In order to model the action of a sliding linker 7 we introduce a term accounting for the DNA self-interaction. In lowest order it can be written as a function of the perpendicular distance z c = c t t tc sin tdt c tc tdt of the two overcrossing DNA arms at the contact point t c resulting from the crossing condition t c = tanh t c. The total energy of the chain can now be written as t c E tot = E 0 + tdt, 3 V t c with a short-ranged, but otherwise arbitrary, attractive interaction potential V acting at the crossing point. Note that if Vz c has a minimum at z c =0 the loop saddle point stays unaffected by the self-interaction. DNA is known to effectively attract itself in many solvents despite its strong negative bare charge. Typical situations inducing DNA selfattraction are poor solvents such as alcohol, and small neutraolymers such as polyethylene glycol 14, the presence of multivalent counterions like hexaammine cobalt 111 and Spermidine or small cationic proteins acting as linkers between two DNA surfaces. Single molecule stretching experiments on DNA condensed with multivalent counterions performed by several groups 15 might bear loops or related structures such as DNA toroids 16. The partition function of the system in Fig. 1a can generally be written as Q loop = loop t 1D 3 te E tott where loop represents the path integral over the functional neighborhood of the loop solution and the function enforces the chain inextensibility. This partition function is nothing but the Euclidean path integral of a quantum particle moving on an unit sphere under the influence of an external constant force. Using the, parametrization, the partition function can be rewritten as Q loop =DDe E tot, ln C m with a metric term C m =exp0ds lncos resulting from the inextensibility constraint. It can be shown that this term does not contribute at the quadratic level of the approximation because we expand around loop =0. By virtue of the decomposition of the in- and out-of-plane fluctuations at the quadratic level Eqs. and 3 the partition function can be conveniently factorized, with PHYSICAL REVIEW E 7, Q loop = e E loopq Q V, Q V = dz c e Vz c Q z c, and with the in- and out-of-plane plane partition functions Q = * De AF/Tˆ dt, Q = D * z t c c dt e t AF/Tˆ dt. c 7 The notation * reminds us that the translational and rotational zero modes of Tˆ and Tˆ, respectively, have to be handled with care. A naive Gaussian integration would lead to a formal divergence in both cases. After dealing with this problem by introducing collective coordinates, a method well known from tunneling theory 10, and applying the Gelfand-Yaglom method for the computation of the fluctuation determinant we obtain the in-plane partition function Q =4/LFe L/ F/A. The computation of Q which follows similar lines of reasoning, requires rewriting the function in Eq. 7 in its Fourier representation. After introducing c, the characteristic function of the interval t c,t c i.e., c t=1 for t t t c,t c and =0 otherwise, we rewrite c tc dt = c ttdt as a scalar product. The path integral in Eq. 7 becomes a Gaussian with a source term ik c and can be solved by constructing the Green s function for Tˆ. This leads to the result
3 APPARENT PERSISTENCE LENGTH RENORMALIZATION Q =8 l P 3/ e L/ e l P /3 z, with =/9t c 3 30t c 0.35 a numeric constant. Combining those results we obtain Q V = 8 l P 3/ e L/ e l P /3 z Vz dz. The resulting force extension relation x =k B T/Fln Q loop writes in leading order x L =1 1 lp1+8 l P L L F 4LF z e A 1/ F 3/ z Vz dz e A 1/ F 3/ z Vz dz 8. 9 In the limiting case of a very deep potential Vz strongly localized around z=0, the last term is negligible and the force-extension relation becomes independent of the detailed nature of the contact interaction. If we in addition consider large forces, the O1/LF term can be neglected and the relation 9 can be cast in a more illuminating form x L app lp F that resembles the well-known loop-free WLC response x/l1 lp F 1 3, but with a strongly renormalized apparent persistence length l P app, l P app = 1+8 L. 11 Equations 10 and 11 show that one has to be very cautious when interpreting experimental stretching data in terms of persistence length and stiffness. The presence of a loop modifies the elastic response of the chain in such a manner that the persistence length can appear effectively reduced as stated in Eq. 11. For a single loop this is obviously a finitesize effect involving the scaled total length L/. But the effect remains significant over a large range of parameters: for l P /L=0.1 one finds l P app 0.31l P whereas for l P /L=0.0 there is still a remarkable effect, namely l P app 0.74l P.We mention that an apparent reduction of stiffness has also been found in Ref. 17 for a WLC with folding elements but there only in the low force regime. To check the prediction of Eqs. 10 and 11 we performed MD simulations 18. As seen from Fig. a the theoreticaredictions quantitatively agree with the simulation results. Stretching kinked DNA. The same approach can be used to calculate the elastic response of DNA with deflections as depicted in Figs. 1b and 1c 19. However, having solved the sliding loop problem we can obtain the results much more directly. To see this we rewrite Eq. 10 in terms of the three relevant lengths L, l P, and, x L PHYSICAL REVIEW E 7, FIG.. Color online a Stretching DNA looped with a sliding linker: Comparison of MD simulation results data points with theoretical prediction solid lines for the force-extension relation as given by Eq. 10. The force is measured in units of F P =k B T/l P. The different slopes correspond to different values of l P /L. The dashed line shows free chain no loop behavior. b The stretching of a GalR loop complex: MD simulation dots vs theory, Eq. 13 solid lines for various values of l P /L l P L 1 The first term in the brackets is the length loss by thermal fluctuations of a straight DNA chain-with the fluctuation contribution of the loop being negligible in the large F limit L. The second term in the bracket, 4/L, describes the loss of length due to the loop-induced elastic deflection. This asymptotic decomposition of the two contributions in 1 leads to immediate generalizations. After short inspection it is easy to see that any strongly localized 0 DNA deflection as in Figs. 1b and 1c can be appropriately continued and mapped piecewise onto fractions of the full loop solution Fig. 1a. By this reasoning, any length loss induced by a given deflection angle between 0 and is given by s0 1 cos loop sds. As a first example consider the stretching of rigid DNAprotein complexes that come as large kinks or fixed angle loops cf. Fig. 1b. A prominent example is the GalR re
4 KULIĆ et al. pressor complex that was studied in single molecule experiments 11. Following the upper reasoning one obtains the same force-extension relation as in Eq. 10 but with an apparent persistence length given as a function of the opening angle of the complex, l P app = 1+8 cos L We note that a previous numerical study of a discretised version of the WLC with local kinks showed a rescaling of the force-extension curve that can be described by our formula cf. Fig. 5 in Ref. 8 for =90. For the GalR complex the opening angle is not known but there are some indications for the antiparallel configuration, i.e., 0 11,1. With Eqs. 10 and 13 at hand, one can predict the loss of length due to the presence of a GalR complex with the conjectured angle =0. For the force F=0.88 pn applied in the experiment 11 with a loop size of 38 nm 11, Eqs. 10 and 13 predict a loss of length of 56 nm with the DNA hidden inside the loop included. Remarkably, the experimental Lia et al. 11 is 55 nm±5 nm. The latter result together with 10 and 13 gives convincing evidence for the antiparallel loop model. An independent check of 10 and 13 is provided by MD simulations shown in Fig. b, which were performed for various ratios of /L. In conclusion, Eq. 10 enables one to directly measure the angle an important feature of the DNA-protein complex geometry. A second application concerns AFM stretching experiments of semiflexible polymers. A stable anchoring can be achieved when the polymer is tangentially attached at its two ends as in Fig. 1c. Force extension data in such a setup have to be interpreted with care. The boundary anchoring angles at the AFM tip as well as at the surface and, respectively can significantly alter the measured apparent persistence length, a fact that has been overlooked before. A more trivial effect of a tilting angle in Fig. 1c between the line of contact points and the force direction can alter the result in addition 3. A simple calculation for large forces F again gives the same functional relation as in 10 with an apparent persistence length given by l P app = PHYSICAL REVIEW E 7, cos L1 cos + cos. 14 While the angle can be completely canceled by shifting the tip in the lateral direction, the influence of and cannot be fully eliminated by this procedure. This means that for shortto intermediate-sized semiflexible polymers l P /L1 1/30 the influence of boundary conditions has to be taken into account through Eq. 14. In conclusion, the force-extension relation of looped DNA that we derived in the limit of a strong stretching force can be generalized to DNA featuring large deflections. In particular our approach provides an analytical expression for the force-extension response of DNA-bending proteins that can give important information with regard to the DNA-protein complex geometry. Large deflections due to the anchoring at the AFM tip and at the substrate also affect the measured persistence length for not too long molecules. Finally these effects might be related to the strong reduction of the persistence length found in condensed DNA stretching experiments by Baumann et al. 15. The authors thank Nikhil Gunari, Andreas Janshoff and Phil Nelson for useful discussions. 1 T. R. Strick et al., Rep. Prog. Phys. 66, S. B. Smith, L. Finzi, and C. Bustamante, Science 58, ; T. T. Perkins et al., ibid. 64, ; C.G. Baumann et al., Proc. Natl. Acad. Sci. U.S.A. 94, C. Bustamante et al., Science 65, ; J. F. Marko and E. D. Siggia, Macromolecules 8, ; T. Odijk, ibid. 8, ; C. Bouchiat et al., Biophys. J. 76, T. R. Strick et al., Science 71, ; Biophys. J. 74, C. Bouchiat and M. Mezard, Phys. Rev. Lett. 80, ; Eur. Phys. J. E, ; J. D. Moroz and P. Nelson, Proc. Natl. Acad. Sci. U.S.A. 94, ; Macromolecules 31, R. Bruinsma and J. Rudnick, Biophys. J. 76, R. Metzler, Y. Kantor, and M. Kardar, Phys. Rev. E 66, J. Yan and J. F. Marko, Phys. Rev. E 68, I. M. Kulić, H. Mohrbach, R. Thaokar, and H. Schiessel, eprint q-bio.bm/ ; I. M. Kulić, Ph.D. thesis, University of Mainz L. S. Schulman, Techniques and Applications of Path Integration Wiley Classics Library, New York, 1996; H. Kleinert, Path Integrals World Scientific, Singapore, 00; J. Zinn- Justin, Quantum Field Theory and Critical Phenomena Clarendon Press, Oxford, G. Lia et al., Proc. Natl. Acad. Sci. U.S.A. 100, P. J. Hagerman, Annu. Rev. Biophys. Biophys. Chem. 17, T. Odijk, Macromolecules 19, V. A. Bloomfield, Curr. Opin. Struct. Biol. 6, C. G. Baumann et al., Biophys. J. 78, H. Noguchi et al., Chem. Phys. Lett. 61, ; B. Schnurr, F. Gittes, and F. C. MacKintosh, Phys. Rev. E 65, S. Cocco et al., Eur. Phys. J. E 10, Langevin MD simulation of a semiflexible bead-spring chain
5 APPARENT PERSISTENCE LENGTH RENORMALIZATION with up to 00 beads; the stretching force applied at the end beads; the contact potential was realized by a flexible sliding linker, i.e., closed bead-spring chain The exact force-extension relations satisfying boundary conditions at finite intervals involving elliptic integrals are omitted here. The contributions of the boundary conditions and the breaking of translational symmetry both involve O1/LF PHYSICAL REVIEW E 7, terms that are negligible in the large force limit leading immediately to Eqs. 13 and By strongly localized we mean a deflection that decays to zero on a length scale much shorter than the whole molecule. 1 M. Geanacopoulos et al., Nat. Struct. Biol. 8, A. Janshoff et al., Angew. Chem., Int. Ed. 39, M. Rief et al., Science 76,
Single molecule force spectroscopy reveals a highly compliant helical
 Supplementary Information Single molecule force spectroscopy reveals a highly compliant helical folding for the 30 nm chromatin fiber Maarten Kruithof, Fan-Tso Chien, Andrew Routh, Colin Logie, Daniela
Supplementary Information Single molecule force spectroscopy reveals a highly compliant helical folding for the 30 nm chromatin fiber Maarten Kruithof, Fan-Tso Chien, Andrew Routh, Colin Logie, Daniela
New Measurements of DNA Twist Elasticity
 University of Pennsylvania ScholarlyCommons Department of Physics Papers Department of Physics 5-1998 New Measurements of DNA Twist Elasticity Philip C. Nelson University of Pennsylvania, nelson@physics.upenn.edu
University of Pennsylvania ScholarlyCommons Department of Physics Papers Department of Physics 5-1998 New Measurements of DNA Twist Elasticity Philip C. Nelson University of Pennsylvania, nelson@physics.upenn.edu
Magnetic tweezers and its application to DNA mechanics
 Biotechnological Center Research group DNA motors (Seidel group) Handout for Practical Course Magnetic tweezers and its application to DNA mechanics When: 9.00 am Where: Biotec, 3 rd Level, Room 317 Tutors:
Biotechnological Center Research group DNA motors (Seidel group) Handout for Practical Course Magnetic tweezers and its application to DNA mechanics When: 9.00 am Where: Biotec, 3 rd Level, Room 317 Tutors:
Curvature Distribution of Worm-like Chains in Two and Three Dimensions
 Curvature Distribution of Worm-like Chains in Two and Three Dimensions Shay M. Rappaport, Shlomi Medalion and Yitzhak Rabin Department of Physics, Bar-Ilan University, Ramat-Gan 59, Israel arxiv:8.383v
Curvature Distribution of Worm-like Chains in Two and Three Dimensions Shay M. Rappaport, Shlomi Medalion and Yitzhak Rabin Department of Physics, Bar-Ilan University, Ramat-Gan 59, Israel arxiv:8.383v
MAE 545: Lecture 4 (9/29) Statistical mechanics of proteins
 MAE 545: Lecture 4 (9/29) Statistical mechanics of proteins Ideal freely jointed chain N identical unstretchable links (Kuhn segments) of length a with freely rotating joints ~r 2 a ~r ~R ~r N In first
MAE 545: Lecture 4 (9/29) Statistical mechanics of proteins Ideal freely jointed chain N identical unstretchable links (Kuhn segments) of length a with freely rotating joints ~r 2 a ~r ~R ~r N In first
Models for Single Molecule Experiments
 Models for Single Molecule Experiments Binding Unbinding Folding - Unfolding -> Kramers Bell theory 1 Polymer elasticity: - Hooke s law (?) - FJC (Freely joined chain) - WLC (Worm-like chain) 3 Molecular
Models for Single Molecule Experiments Binding Unbinding Folding - Unfolding -> Kramers Bell theory 1 Polymer elasticity: - Hooke s law (?) - FJC (Freely joined chain) - WLC (Worm-like chain) 3 Molecular
GEM4 Summer School OpenCourseWare
 GEM4 Summer School OpenCourseWare http://gem4.educommons.net/ http://www.gem4.org/ Lecture: Polymer Chains by Ju Li. Given August 16, 2006 during the GEM4 session at MIT in Cambridge, MA. Please use the
GEM4 Summer School OpenCourseWare http://gem4.educommons.net/ http://www.gem4.org/ Lecture: Polymer Chains by Ju Li. Given August 16, 2006 during the GEM4 session at MIT in Cambridge, MA. Please use the
Effects of kinks on DNA elasticity
 PHYSICAL REVIEW E 71, 051905 2005 Effects of kinks on DNA elasticity Yuri O. Popov* and Alexei V. Tkachenko Department of Physics, University of Michigan, 450 Church Street, Ann Arbor, Michigan 48109,
PHYSICAL REVIEW E 71, 051905 2005 Effects of kinks on DNA elasticity Yuri O. Popov* and Alexei V. Tkachenko Department of Physics, University of Michigan, 450 Church Street, Ann Arbor, Michigan 48109,
Faculty of math and natural science. Universty of Split. Kratky-Porod model. Nataša Vučemilovid-Alagid. February, 2013.
 Faculty of math and natural science Universty of Split Kratky-Porod model Nataša Vučemilovid-Alagid February, 2013. Table of contents 1.Introduction:... 3 1. DNA:... 4 2. Models of polymer elasticity:...
Faculty of math and natural science Universty of Split Kratky-Porod model Nataša Vučemilovid-Alagid February, 2013. Table of contents 1.Introduction:... 3 1. DNA:... 4 2. Models of polymer elasticity:...
Buckling of stiff polymers: Influence of thermal fluctuations
 PHYSICAL REVIEW E 76, 06907 2007 Buckling of stiff polymers: Influence of thermal fluctuations Marc Emanuel, Hervé Mohrbach, 2 Mehmet Sayar, 3 Helmut Schiessel, and Igor M. Kulić 4 Instituut-Lorentz, Universiteit
PHYSICAL REVIEW E 76, 06907 2007 Buckling of stiff polymers: Influence of thermal fluctuations Marc Emanuel, Hervé Mohrbach, 2 Mehmet Sayar, 3 Helmut Schiessel, and Igor M. Kulić 4 Instituut-Lorentz, Universiteit
Exact Theory of Kinkable Elastic Polymers
 University of Pennsylvania ScholarlyCommons Department of Physics Papers Department of Physics 2-2005 Exact Theory of Kinkable Elastic Polymers Paul A. Wiggins Rob Phillips Philip C. Nelson University
University of Pennsylvania ScholarlyCommons Department of Physics Papers Department of Physics 2-2005 Exact Theory of Kinkable Elastic Polymers Paul A. Wiggins Rob Phillips Philip C. Nelson University
Exact theory of kinkable elastic polymers
 Exact theory of kinkable elastic polymers Paul A. Wiggins* Division of Physics, Mathematics, and Astronomy, California Institute of Technology, Pasadena, California 91125, USA Rob Phillips Division of
Exact theory of kinkable elastic polymers Paul A. Wiggins* Division of Physics, Mathematics, and Astronomy, California Institute of Technology, Pasadena, California 91125, USA Rob Phillips Division of
3 Biopolymers Uncorrelated chains - Freely jointed chain model
 3.4 Entropy When we talk about biopolymers, it is important to realize that the free energy of a biopolymer in thermal equilibrium is not constant. Unlike solids, biopolymers are characterized through
3.4 Entropy When we talk about biopolymers, it is important to realize that the free energy of a biopolymer in thermal equilibrium is not constant. Unlike solids, biopolymers are characterized through
Effect of protein shape on multibody interactions between membrane inclusions
 PHYSICAL REVIEW E VOLUME 61, NUMBER 4 APRIL 000 Effect of protein shape on multibody interactions between membrane inclusions K. S. Kim, 1, * John Neu, and George Oster 3, 1 Department of Physics, Graduate
PHYSICAL REVIEW E VOLUME 61, NUMBER 4 APRIL 000 Effect of protein shape on multibody interactions between membrane inclusions K. S. Kim, 1, * John Neu, and George Oster 3, 1 Department of Physics, Graduate
Exact theory of kinkable elastic polymers
 Exact theory of kinkable elastic polymers Paul A. Wiggins, Rob Phillips, and Philip C. Nelson (Dated: August 31st, 2004) The importance of nonlinearities in material constitutive relations has long been
Exact theory of kinkable elastic polymers Paul A. Wiggins, Rob Phillips, and Philip C. Nelson (Dated: August 31st, 2004) The importance of nonlinearities in material constitutive relations has long been
Lines of Renormalization Group Fixed Points for Fluid and Crystalline Membranes.
 EUROPHYSICS LETTERS 1 October 1988 Europhys. Lett., 7 (3), pp. 255-261 (1988) Lines of Renormalization Group Fixed Points for Fluid and Crystalline Membranes. R. LIPOWSKY Institut für Festkörperforschung
EUROPHYSICS LETTERS 1 October 1988 Europhys. Lett., 7 (3), pp. 255-261 (1988) Lines of Renormalization Group Fixed Points for Fluid and Crystalline Membranes. R. LIPOWSKY Institut für Festkörperforschung
Multimedia : Fibronectin and Titin unfolding simulation movies.
 I LECTURE 21: SINGLE CHAIN ELASTICITY OF BIOMACROMOLECULES: THE GIANT PROTEIN TITIN AND DNA Outline : REVIEW LECTURE #2 : EXTENSIBLE FJC AND WLC... 2 STRUCTURE OF MUSCLE AND TITIN... 3 SINGLE MOLECULE
I LECTURE 21: SINGLE CHAIN ELASTICITY OF BIOMACROMOLECULES: THE GIANT PROTEIN TITIN AND DNA Outline : REVIEW LECTURE #2 : EXTENSIBLE FJC AND WLC... 2 STRUCTURE OF MUSCLE AND TITIN... 3 SINGLE MOLECULE
arxiv:cond-mat/ v1 [cond-mat.soft] 29 May 2002
![arxiv:cond-mat/ v1 [cond-mat.soft] 29 May 2002 arxiv:cond-mat/ v1 [cond-mat.soft] 29 May 2002](/thumbs/90/102416155.jpg) Stretching Instability of Helical Springs David A. Kessler and Yitzhak Rabin Dept. of Physics, Bar-Ilan University, Ramat-Gan, Israel (Dated: October 31, 18) arxiv:cond-mat/05612v1 [cond-mat.soft] 29 May
Stretching Instability of Helical Springs David A. Kessler and Yitzhak Rabin Dept. of Physics, Bar-Ilan University, Ramat-Gan, Israel (Dated: October 31, 18) arxiv:cond-mat/05612v1 [cond-mat.soft] 29 May
Excluded-Volume Effects in Tethered-Particle Experiments: Bead Size Matters
 University of Pennsylvania ScholarlyCommons Department of Physics Papers Department of Physics 3-2006 Excluded-Volume Effects in Tethered-Particle Experiments: Bead Size Matters Darren E. Segall Philip
University of Pennsylvania ScholarlyCommons Department of Physics Papers Department of Physics 3-2006 Excluded-Volume Effects in Tethered-Particle Experiments: Bead Size Matters Darren E. Segall Philip
Supporting Information
 Supporting Information A: Calculation of radial distribution functions To get an effective propagator in one dimension, we first transform 1) into spherical coordinates: x a = ρ sin θ cos φ, y = ρ sin
Supporting Information A: Calculation of radial distribution functions To get an effective propagator in one dimension, we first transform 1) into spherical coordinates: x a = ρ sin θ cos φ, y = ρ sin
The elastic theory of a single DNA molecule
 PRAMANA cfl Indian Academy of Sciences Vol. 61, No. 2 journal of August 2003 physics pp. 353 360 The elastic theory of a single DNA molecule HAIJUN HOU a;b, YANG HANG a and HANG-CAN OU-YANG a;c; * a Institute
PRAMANA cfl Indian Academy of Sciences Vol. 61, No. 2 journal of August 2003 physics pp. 353 360 The elastic theory of a single DNA molecule HAIJUN HOU a;b, YANG HANG a and HANG-CAN OU-YANG a;c; * a Institute
Chapter 1 Introduction
 Chapter 1 Introduction This thesis is concerned with the behaviour of polymers in flow. Both polymers in solutions and polymer melts will be discussed. The field of research that studies the flow behaviour
Chapter 1 Introduction This thesis is concerned with the behaviour of polymers in flow. Both polymers in solutions and polymer melts will be discussed. The field of research that studies the flow behaviour
arxiv:cond-mat/ v2 [cond-mat.soft] 16 Jan 2005
![arxiv:cond-mat/ v2 [cond-mat.soft] 16 Jan 2005 arxiv:cond-mat/ v2 [cond-mat.soft] 16 Jan 2005](/thumbs/94/121793971.jpg) arxiv:cond-mat/412617v2 [cond-mat.soft] 16 Jan 25 On the behaviour of short Kratky-Porod chain Semjon Stepanow Martin-Luther-Universität Halle, Fachbereich Physik, D-699 Halle, Germany PACS numbers: 5.4.-a,
arxiv:cond-mat/412617v2 [cond-mat.soft] 16 Jan 25 On the behaviour of short Kratky-Porod chain Semjon Stepanow Martin-Luther-Universität Halle, Fachbereich Physik, D-699 Halle, Germany PACS numbers: 5.4.-a,
Torsional directed walks, entropic elasticity, and DNA twist stiffness
 Proc. Natl. Acad. Sci. USA Vol. 94, pp. 14418 14422, December 1997 Biophysics Torsional directed walks, entropic elasticity, and DNA twist stiffness J. DAVID MOROZ AND PHILIP NELSON* Department of Physics
Proc. Natl. Acad. Sci. USA Vol. 94, pp. 14418 14422, December 1997 Biophysics Torsional directed walks, entropic elasticity, and DNA twist stiffness J. DAVID MOROZ AND PHILIP NELSON* Department of Physics
Announcements. Homework 3 (Klaus Schulten s Lecture): Due Wednesday at noon. Next homework assigned. Due Wednesday March 1.
 Announcements Homework 3 (Klaus Schulten s Lecture): Due Wednesday at noon. Next homework assigned. Due Wednesday March 1. No lecture next Monday, Feb. 27 th! (Homework is a bit longer.) Marco will have
Announcements Homework 3 (Klaus Schulten s Lecture): Due Wednesday at noon. Next homework assigned. Due Wednesday March 1. No lecture next Monday, Feb. 27 th! (Homework is a bit longer.) Marco will have
Appendix C - Persistence length 183. Consider an ideal chain with N segments each of length a, such that the contour length L c is
 Appendix C - Persistence length 183 APPENDIX C - PERSISTENCE LENGTH Consider an ideal chain with N segments each of length a, such that the contour length L c is L c = Na. (C.1) If the orientation of each
Appendix C - Persistence length 183 APPENDIX C - PERSISTENCE LENGTH Consider an ideal chain with N segments each of length a, such that the contour length L c is L c = Na. (C.1) If the orientation of each
Writhe distribution of stiff biopolymers
 Writhe distribution of stiff biopolymers Abhijit Ghosh Raman Research Institute, Bangalore, India 560080 Received 5 November 2007; published 30 April 2008 Motivated by the interest in the role of stiff
Writhe distribution of stiff biopolymers Abhijit Ghosh Raman Research Institute, Bangalore, India 560080 Received 5 November 2007; published 30 April 2008 Motivated by the interest in the role of stiff
Investigation of dsdna Stretching Meso-Mechanics Using LS-DYNA
 8 th International LS-DYNA Users Conference Simulation echnology (3) Investigation of dsdna Stretching Meso-Mechanics Using LS-DYNA C. A. Yuan and K. N. Chiang Department of Power Mechanical Engineering,
8 th International LS-DYNA Users Conference Simulation echnology (3) Investigation of dsdna Stretching Meso-Mechanics Using LS-DYNA C. A. Yuan and K. N. Chiang Department of Power Mechanical Engineering,
arxiv:cond-mat/ v2 [cond-mat.soft] 6 Aug 2002
![arxiv:cond-mat/ v2 [cond-mat.soft] 6 Aug 2002 arxiv:cond-mat/ v2 [cond-mat.soft] 6 Aug 2002](/thumbs/73/68156181.jpg) Radial Distribution Function of Rod-like Polyelelctrolytes Roya Zandi Department of Chemistry and Biochemistry, UCLA, Box 95569, Los Angeles, California, 995-569 arxiv:cond-mat/7v [cond-mat.soft] 6 Aug
Radial Distribution Function of Rod-like Polyelelctrolytes Roya Zandi Department of Chemistry and Biochemistry, UCLA, Box 95569, Los Angeles, California, 995-569 arxiv:cond-mat/7v [cond-mat.soft] 6 Aug
Effective theory of quadratic degeneracies
 Effective theory of quadratic degeneracies Y. D. Chong,* Xiao-Gang Wen, and Marin Soljačić Department of Physics, Massachusetts Institute of Technology, Cambridge, Massachusetts 02139, USA Received 28
Effective theory of quadratic degeneracies Y. D. Chong,* Xiao-Gang Wen, and Marin Soljačić Department of Physics, Massachusetts Institute of Technology, Cambridge, Massachusetts 02139, USA Received 28
Obtaining the Bending Modulus from a Buckled Lipid Membrane
 Obtaining the Bending Modulus from a Buckled Lipid Membrane Patrick Diggins, Mingyang Hu, Markus Deserno Department of Physics, Carnegie Mellon University, Pittsburgh PA, USA October 11, 2013 Outline Lipid
Obtaining the Bending Modulus from a Buckled Lipid Membrane Patrick Diggins, Mingyang Hu, Markus Deserno Department of Physics, Carnegie Mellon University, Pittsburgh PA, USA October 11, 2013 Outline Lipid
PHYSICAL REVIEW LETTERS
 PHYSICAL REVIEW LETTERS VOLUME 76 4 MARCH 1996 NUMBER 10 Finite-Size Scaling and Universality above the Upper Critical Dimensionality Erik Luijten* and Henk W. J. Blöte Faculty of Applied Physics, Delft
PHYSICAL REVIEW LETTERS VOLUME 76 4 MARCH 1996 NUMBER 10 Finite-Size Scaling and Universality above the Upper Critical Dimensionality Erik Luijten* and Henk W. J. Blöte Faculty of Applied Physics, Delft
DNA supercoiling: plectonemes or curls?
 DNA supercoiling: plectonemes or curls? Sébastien Neukirch CNRS & UPMC Univ. Paris 6 (France) John F. Marko Northwestern University (IL. USA) Why study DNA mechanical properties? mechanical properties
DNA supercoiling: plectonemes or curls? Sébastien Neukirch CNRS & UPMC Univ. Paris 6 (France) John F. Marko Northwestern University (IL. USA) Why study DNA mechanical properties? mechanical properties
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/314/5801/1001/dc1 Supporting Online Material for Direct Measurement of the Full, Sequence-Dependent Folding Landscape of a Nucleic Acid Michael T. Woodside, Peter C.
www.sciencemag.org/cgi/content/full/314/5801/1001/dc1 Supporting Online Material for Direct Measurement of the Full, Sequence-Dependent Folding Landscape of a Nucleic Acid Michael T. Woodside, Peter C.
arxiv: v1 [quant-ph] 7 Mar 2012
![arxiv: v1 [quant-ph] 7 Mar 2012 arxiv: v1 [quant-ph] 7 Mar 2012](/thumbs/73/68782511.jpg) Global Level Number Variance in Integrable Systems Tao Ma, R.A. Serota Department of Physics University of Cincinnati Cincinnati, OH 5-11 (Dated: October, 13) arxiv:3.1v1 [quant-ph] 7 Mar 1 We study previously
Global Level Number Variance in Integrable Systems Tao Ma, R.A. Serota Department of Physics University of Cincinnati Cincinnati, OH 5-11 (Dated: October, 13) arxiv:3.1v1 [quant-ph] 7 Mar 1 We study previously
Classical differential geometry of two-dimensional surfaces
 Classical differential geometry of two-dimensional surfaces 1 Basic definitions This section gives an overview of the basic notions of differential geometry for twodimensional surfaces. It follows mainly
Classical differential geometry of two-dimensional surfaces 1 Basic definitions This section gives an overview of the basic notions of differential geometry for twodimensional surfaces. It follows mainly
Giant Enhancement of Quantum Decoherence by Frustrated Environments
 ISSN 0021-3640, JETP Letters, 2006, Vol. 84, No. 2, pp. 99 103. Pleiades Publishing, Inc., 2006.. Giant Enhancement of Quantum Decoherence by Frustrated Environments S. Yuan a, M. I. Katsnelson b, and
ISSN 0021-3640, JETP Letters, 2006, Vol. 84, No. 2, pp. 99 103. Pleiades Publishing, Inc., 2006.. Giant Enhancement of Quantum Decoherence by Frustrated Environments S. Yuan a, M. I. Katsnelson b, and
Finite Element Analysis Prof. Dr. B. N. Rao Department of Civil Engineering Indian Institute of Technology, Madras. Lecture - 06
 Finite Element Analysis Prof. Dr. B. N. Rao Department of Civil Engineering Indian Institute of Technology, Madras Lecture - 06 In the last lecture, we have seen a boundary value problem, using the formal
Finite Element Analysis Prof. Dr. B. N. Rao Department of Civil Engineering Indian Institute of Technology, Madras Lecture - 06 In the last lecture, we have seen a boundary value problem, using the formal
Theory and Practice of Rotor Dynamics Prof. Dr. Rajiv Tiwari Department of Mechanical Engineering Indian Institute of Technology Guwahati
 Theory and Practice of Rotor Dynamics Prof. Dr. Rajiv Tiwari Department of Mechanical Engineering Indian Institute of Technology Guwahati Module - 2 Simpul Rotors Lecture - 2 Jeffcott Rotor Model In the
Theory and Practice of Rotor Dynamics Prof. Dr. Rajiv Tiwari Department of Mechanical Engineering Indian Institute of Technology Guwahati Module - 2 Simpul Rotors Lecture - 2 Jeffcott Rotor Model In the
Supplementary Material
 1 2 3 Topological defects in confined populations of spindle-shaped cells by G. Duclos et al. Supplementary Material 4 5 6 7 8 9 10 11 12 13 Supplementary Note 1: Characteristic time associated with the
1 2 3 Topological defects in confined populations of spindle-shaped cells by G. Duclos et al. Supplementary Material 4 5 6 7 8 9 10 11 12 13 Supplementary Note 1: Characteristic time associated with the
arxiv: v2 [cond-mat.soft] 10 Sep 2015
![arxiv: v2 [cond-mat.soft] 10 Sep 2015 arxiv: v2 [cond-mat.soft] 10 Sep 2015](/thumbs/82/86252089.jpg) Efficient simulation of semiflexible polymers Debabrata Panja Institute for Theoretical Physics, Universiteit Utrecht, Leuvenlaan 4, 3584 CE Utrecht, The Netherlands and Institute of Physics, Universiteit
Efficient simulation of semiflexible polymers Debabrata Panja Institute for Theoretical Physics, Universiteit Utrecht, Leuvenlaan 4, 3584 CE Utrecht, The Netherlands and Institute of Physics, Universiteit
Nature Methods: doi: /nmeth Supplementary Figure 1. Principle of force-distance (FD) curve based AFM.
 Supplementary Figure 1 Principle of force-distance (FD) curve based AFM. (a) FD-based AFM contours the sample surface while oscillating the AFM tip with a sine wave at a frequency of 0.25 khz. Pixel-by-pixel
Supplementary Figure 1 Principle of force-distance (FD) curve based AFM. (a) FD-based AFM contours the sample surface while oscillating the AFM tip with a sine wave at a frequency of 0.25 khz. Pixel-by-pixel
Dynamics of an active magnetic particle in a rotating magnetic field
 PHYSICAL REVIEW E 73, 021505 2006 Dynamics of an active magnetic particle in a rotating magnetic field A. Cēbers* Institute of Physics, University of Latvia, Salaspils-1, LV-2169, Latvia M. Ozols University
PHYSICAL REVIEW E 73, 021505 2006 Dynamics of an active magnetic particle in a rotating magnetic field A. Cēbers* Institute of Physics, University of Latvia, Salaspils-1, LV-2169, Latvia M. Ozols University
Supplementary Information
 Supplementary Information Mechanical unzipping and rezipping of a single SNARE complex reveals hysteresis as a force-generating mechanism Duyoung Min, Kipom Kim, Changbong Hyeon, Yong Hoon Cho, Yeon-Kyun
Supplementary Information Mechanical unzipping and rezipping of a single SNARE complex reveals hysteresis as a force-generating mechanism Duyoung Min, Kipom Kim, Changbong Hyeon, Yong Hoon Cho, Yeon-Kyun
Distinct regimes of elastic response and deformation modes of cross-linked cytoskeletal and semiflexible polymer networks
 Distinct regimes of elastic response and deformation modes of cross-linked cytoskeletal and semiflexible polymer networks D. A. Head, 1, A. J. Levine,,3 and F. C. MacKintosh 1, 1 Department of Physics
Distinct regimes of elastic response and deformation modes of cross-linked cytoskeletal and semiflexible polymer networks D. A. Head, 1, A. J. Levine,,3 and F. C. MacKintosh 1, 1 Department of Physics
Triangular Lattice Foldings-a Transfer Matrix Study.
 EUROPHYSICS LETTERS Europhys. Lett., 11 (2)) pp. 157-161 (1990) 15 January 1990 Triangular Lattice Foldings-a Transfer Matrix Study. Y. KANT~R(*) and M. V. JARIC(**) (*) School of Physics and Astronomy,
EUROPHYSICS LETTERS Europhys. Lett., 11 (2)) pp. 157-161 (1990) 15 January 1990 Triangular Lattice Foldings-a Transfer Matrix Study. Y. KANT~R(*) and M. V. JARIC(**) (*) School of Physics and Astronomy,
Supporting Information for. DNA elasticity from short DNA to nucleosomal DNA
 Supporting Information for DNA elasticity from short DNA to nucleosomal DNA Ashok Garai, 1 Suman Saurabh, 1 Yves Lansac, 2 and Prabal K. Maiti* 1 1 Centre for Condensed Matter Theory, Department of Physics,
Supporting Information for DNA elasticity from short DNA to nucleosomal DNA Ashok Garai, 1 Suman Saurabh, 1 Yves Lansac, 2 and Prabal K. Maiti* 1 1 Centre for Condensed Matter Theory, Department of Physics,
Representations of Sp(6,R) and SU(3) carried by homogeneous polynomials
 Representations of Sp(6,R) and SU(3) carried by homogeneous polynomials Govindan Rangarajan a) Department of Mathematics and Centre for Theoretical Studies, Indian Institute of Science, Bangalore 560 012,
Representations of Sp(6,R) and SU(3) carried by homogeneous polynomials Govindan Rangarajan a) Department of Mathematics and Centre for Theoretical Studies, Indian Institute of Science, Bangalore 560 012,
Structural investigation of single biomolecules
 Structural investigation of single biomolecules NMR spectroscopy and X-ray crystallography are currently the most common techniques capable of determining the structures of biological macromolecules like
Structural investigation of single biomolecules NMR spectroscopy and X-ray crystallography are currently the most common techniques capable of determining the structures of biological macromolecules like
arxiv:cond-mat/ v1 4 Aug 2003
 Conductivity of thermally fluctuating superconductors in two dimensions Subir Sachdev arxiv:cond-mat/0308063 v1 4 Aug 2003 Abstract Department of Physics, Yale University, P.O. Box 208120, New Haven CT
Conductivity of thermally fluctuating superconductors in two dimensions Subir Sachdev arxiv:cond-mat/0308063 v1 4 Aug 2003 Abstract Department of Physics, Yale University, P.O. Box 208120, New Haven CT
Potts And XY, Together At Last
 Potts And XY, Together At Last Daniel Kolodrubetz Massachusetts Institute of Technology, Center for Theoretical Physics (Dated: May 16, 212) We investigate the behavior of an XY model coupled multiplicatively
Potts And XY, Together At Last Daniel Kolodrubetz Massachusetts Institute of Technology, Center for Theoretical Physics (Dated: May 16, 212) We investigate the behavior of an XY model coupled multiplicatively
4 4 and perturbation theory
 and perturbation theory We now consider interacting theories. A simple case is the interacting scalar field, with a interaction. This corresponds to a -body contact repulsive interaction between scalar
and perturbation theory We now consider interacting theories. A simple case is the interacting scalar field, with a interaction. This corresponds to a -body contact repulsive interaction between scalar
arxiv:cond-mat/ v1 [cond-mat.stat-mech] 12 Jan 2006
![arxiv:cond-mat/ v1 [cond-mat.stat-mech] 12 Jan 2006 arxiv:cond-mat/ v1 [cond-mat.stat-mech] 12 Jan 2006](/thumbs/92/109595830.jpg) arxiv:cond-mat/6263v [cond-mat.stat-mech] 2 Jan 26 A one-dimensional model for theoretical analysis of single molecule experiments. Introduction Erik Van der Straeten and Jan Naudts Departement Fysica,
arxiv:cond-mat/6263v [cond-mat.stat-mech] 2 Jan 26 A one-dimensional model for theoretical analysis of single molecule experiments. Introduction Erik Van der Straeten and Jan Naudts Departement Fysica,
1 Fundamentals. 1.1 Overview. 1.2 Units: Physics 704 Spring 2018
 Physics 704 Spring 2018 1 Fundamentals 1.1 Overview The objective of this course is: to determine and fields in various physical systems and the forces and/or torques resulting from them. The domain of
Physics 704 Spring 2018 1 Fundamentals 1.1 Overview The objective of this course is: to determine and fields in various physical systems and the forces and/or torques resulting from them. The domain of
Engineering Mechanics Prof. U. S. Dixit Department of Mechanical Engineering Indian Institute of Technology, Guwahati Introduction to vibration
 Engineering Mechanics Prof. U. S. Dixit Department of Mechanical Engineering Indian Institute of Technology, Guwahati Introduction to vibration Module 15 Lecture 38 Vibration of Rigid Bodies Part-1 Today,
Engineering Mechanics Prof. U. S. Dixit Department of Mechanical Engineering Indian Institute of Technology, Guwahati Introduction to vibration Module 15 Lecture 38 Vibration of Rigid Bodies Part-1 Today,
Foundations of Chemical Kinetics. Lecture 12: Transition-state theory: The thermodynamic formalism
 Foundations of Chemical Kinetics Lecture 12: Transition-state theory: The thermodynamic formalism Marc R. Roussel Department of Chemistry and Biochemistry Breaking it down We can break down an elementary
Foundations of Chemical Kinetics Lecture 12: Transition-state theory: The thermodynamic formalism Marc R. Roussel Department of Chemistry and Biochemistry Breaking it down We can break down an elementary
, follows from these assumptions.
 Untangling the mechanics and topology in the frictional response of long overhand elastic knots Supplemental Information M.K. Jawed, P. Dieleman, B. Audoly, P. M. Reis S1 Experimental details In this section,
Untangling the mechanics and topology in the frictional response of long overhand elastic knots Supplemental Information M.K. Jawed, P. Dieleman, B. Audoly, P. M. Reis S1 Experimental details In this section,
arxiv: v4 [quant-ph] 9 Jun 2016
![arxiv: v4 [quant-ph] 9 Jun 2016 arxiv: v4 [quant-ph] 9 Jun 2016](/thumbs/73/68830115.jpg) Applying Classical Geometry Intuition to Quantum arxiv:101.030v4 [quant-ph] 9 Jun 016 Spin Dallin S. Durfee and James L. Archibald Department of Physics and Astronomy, Brigham Young University, Provo,
Applying Classical Geometry Intuition to Quantum arxiv:101.030v4 [quant-ph] 9 Jun 016 Spin Dallin S. Durfee and James L. Archibald Department of Physics and Astronomy, Brigham Young University, Provo,
Supplementary Information. The Solution Structural Ensembles of RNA Kink-turn Motifs and Their Protein Complexes
 Supplementary Information The Solution Structural Ensembles of RNA Kink-turn Motifs and Their Protein Complexes Xuesong Shi, a Lin Huang, b David M. J. Lilley, b Pehr B. Harbury a,c and Daniel Herschlag
Supplementary Information The Solution Structural Ensembles of RNA Kink-turn Motifs and Their Protein Complexes Xuesong Shi, a Lin Huang, b David M. J. Lilley, b Pehr B. Harbury a,c and Daniel Herschlag
V = 2ze 2 n. . a. i=1
 IITS: Statistical Physics in Biology Assignment # 3 KU Leuven 5/29/2013 Coulomb Interactions & Polymers 1. Flory Theory: The Coulomb energy of a ball of charge Q and dimension R in d spacial dimensions
IITS: Statistical Physics in Biology Assignment # 3 KU Leuven 5/29/2013 Coulomb Interactions & Polymers 1. Flory Theory: The Coulomb energy of a ball of charge Q and dimension R in d spacial dimensions
Magnetic Torque Tweezers: measuring torsional stiffness in DNA and RecA DNA filaments
 Magnetic Torque Tweezers: measuring torsional stiffness in DNA and RecA DNA filaments Lipfert, J., Kerssemakers, J. W., Jager, T. & Dekker, N. H. Magnetic torque tweezers: measuring torsional stiffness
Magnetic Torque Tweezers: measuring torsional stiffness in DNA and RecA DNA filaments Lipfert, J., Kerssemakers, J. W., Jager, T. & Dekker, N. H. Magnetic torque tweezers: measuring torsional stiffness
Fluctuating Elastic Filaments Under Distributed Loads
 Fluctuating Elastic Filaments Under Distributed Loads Tianxiang Su, Prashant K. Purohit Department of Mechanical Engineering and Applied Mechanics, University of Pennsylvania, Philadelphia, PA 94, USA.
Fluctuating Elastic Filaments Under Distributed Loads Tianxiang Su, Prashant K. Purohit Department of Mechanical Engineering and Applied Mechanics, University of Pennsylvania, Philadelphia, PA 94, USA.
Pulling hairpinned polynucleotide chains: Does base-pair stacking interaction matter?
 JOURNAL OF CHEMICAL PHYSICS VOLUME 114, NUMBER 19 15 MAY 2001 Pulling hairpinned polynucleotide chains: Does base-pair stacking interaction matter? Haijun Zhou a) Max-Planck-Institute of Colloids and Interfaces,
JOURNAL OF CHEMICAL PHYSICS VOLUME 114, NUMBER 19 15 MAY 2001 Pulling hairpinned polynucleotide chains: Does base-pair stacking interaction matter? Haijun Zhou a) Max-Planck-Institute of Colloids and Interfaces,
Towards a manifestly diffeomorphism invariant Exact Renormalization Group
 Towards a manifestly diffeomorphism invariant Exact Renormalization Group Anthony W. H. Preston University of Southampton Supervised by Prof. Tim R. Morris Talk prepared for UK QFT-V, University of Nottingham,
Towards a manifestly diffeomorphism invariant Exact Renormalization Group Anthony W. H. Preston University of Southampton Supervised by Prof. Tim R. Morris Talk prepared for UK QFT-V, University of Nottingham,
Exchange of Counterions in DNA Condensation. Abstract
 Exchange of Counterions in DNA Condensation Yoshihiro Murayama and Masaki Sano Department of Physics, University of Tokyo, Tokyo 113-0033, Japan Abstract We measured the fluorescence intensity of DNA-bound
Exchange of Counterions in DNA Condensation Yoshihiro Murayama and Masaki Sano Department of Physics, University of Tokyo, Tokyo 113-0033, Japan Abstract We measured the fluorescence intensity of DNA-bound
LATERAL STABILITY OF BEAMS WITH ELASTIC END RESTRAINTS
 LATERAL STABILITY OF BEAMS WITH ELASTIC END RESTRAINTS By John J. Zahn, 1 M. ASCE ABSTRACT: In the analysis of the lateral buckling of simply supported beams, the ends are assumed to be rigidly restrained
LATERAL STABILITY OF BEAMS WITH ELASTIC END RESTRAINTS By John J. Zahn, 1 M. ASCE ABSTRACT: In the analysis of the lateral buckling of simply supported beams, the ends are assumed to be rigidly restrained
Elastic property of single double-stranded DNA molecules: Theoretical study and comparison with experiments
 PHYSICA REVIEW E VOUME 62, NUMBER 1 JUY 2 Elastic property of single double-stranded DNA molecules: Theoretical study and comparison with experiments Haijun Zhou, 1,2, * Yang Zhang, 1 and Zhong-can Ou-Yang
PHYSICA REVIEW E VOUME 62, NUMBER 1 JUY 2 Elastic property of single double-stranded DNA molecules: Theoretical study and comparison with experiments Haijun Zhou, 1,2, * Yang Zhang, 1 and Zhong-can Ou-Yang
Chain-configuration and rate dependent rheological properties in transient networks
 Electronic Supplementary Material (ESI) for Soft Matter. This journal is The Royal Society of Chemistry 204 Supplementary Information Chain-configuration and rate dependent rheological properties in transient
Electronic Supplementary Material (ESI) for Soft Matter. This journal is The Royal Society of Chemistry 204 Supplementary Information Chain-configuration and rate dependent rheological properties in transient
Math Methods for Polymer Physics Lecture 1: Series Representations of Functions
 Math Methods for Polymer Physics ecture 1: Series Representations of Functions Series analysis is an essential tool in polymer physics and physical sciences, in general. Though other broadly speaking,
Math Methods for Polymer Physics ecture 1: Series Representations of Functions Series analysis is an essential tool in polymer physics and physical sciences, in general. Though other broadly speaking,
Minimal encounter time and separation determine ligand-receptor binding in cell adhesion SUPPLEMENTARY INFORMATION
 Minimal encounter time and separation determine ligand-receptor binding in cell adhesion SUPPLEMENTARY INFORMATION Philippe Robert Alice Nicolas Said Aranda-Espinoza Pierre Bongrand Laurent Limozin Molecular
Minimal encounter time and separation determine ligand-receptor binding in cell adhesion SUPPLEMENTARY INFORMATION Philippe Robert Alice Nicolas Said Aranda-Espinoza Pierre Bongrand Laurent Limozin Molecular
Randomly Triangulated Surfaces as Models for Fluid and Crystalline Membranes. G. Gompper Institut für Festkörperforschung, Forschungszentrum Jülich
 Randomly Triangulated Surfaces as Models for Fluid and Crystalline Membranes G. Gompper Institut für Festkörperforschung, Forschungszentrum Jülich Motivation: Endo- and Exocytosis Membrane transport of
Randomly Triangulated Surfaces as Models for Fluid and Crystalline Membranes G. Gompper Institut für Festkörperforschung, Forschungszentrum Jülich Motivation: Endo- and Exocytosis Membrane transport of
Liquid crystal in confined environment
 Liquid crystal in confined environment Adviser: Prof. Rudi Podgornik Supervisor: Prof. Igor Muševič By Maryam Nikkhou September 2011 Contents Abstract.................................................................
Liquid crystal in confined environment Adviser: Prof. Rudi Podgornik Supervisor: Prof. Igor Muševič By Maryam Nikkhou September 2011 Contents Abstract.................................................................
Exploring the energy landscape
 Exploring the energy landscape ChE210D Today's lecture: what are general features of the potential energy surface and how can we locate and characterize minima on it Derivatives of the potential energy
Exploring the energy landscape ChE210D Today's lecture: what are general features of the potential energy surface and how can we locate and characterize minima on it Derivatives of the potential energy
Metrics and Curvature
 Metrics and Curvature How to measure curvature? Metrics Euclidian/Minkowski Curved spaces General 4 dimensional space Cosmological principle Homogeneity and isotropy: evidence Robertson-Walker metrics
Metrics and Curvature How to measure curvature? Metrics Euclidian/Minkowski Curved spaces General 4 dimensional space Cosmological principle Homogeneity and isotropy: evidence Robertson-Walker metrics
Transport of single molecules along the periodic parallel lattices with coupling
 THE JOURNAL OF CHEMICAL PHYSICS 124 204901 2006 Transport of single molecules along the periodic parallel lattices with coupling Evgeny B. Stukalin The James Franck Institute The University of Chicago
THE JOURNAL OF CHEMICAL PHYSICS 124 204901 2006 Transport of single molecules along the periodic parallel lattices with coupling Evgeny B. Stukalin The James Franck Institute The University of Chicago
Elasticity of biological gels
 Seminar II Elasticity of biological gels Author: Gašper Gregorič Mentor: assoc. prof. Primož Ziherl Ljubljana, February 2014 Abstract In the seminar we discuss the elastic behavior of biological gels,
Seminar II Elasticity of biological gels Author: Gašper Gregorič Mentor: assoc. prof. Primož Ziherl Ljubljana, February 2014 Abstract In the seminar we discuss the elastic behavior of biological gels,
Supplementary Information
 Supplementary Information Switching of myosin-v motion between the lever-arm swing and Brownian search-and-catch Keisuke Fujita 1*, Mitsuhiro Iwaki 2,3*, Atsuko H. Iwane 1, Lorenzo Marcucci 1 & Toshio
Supplementary Information Switching of myosin-v motion between the lever-arm swing and Brownian search-and-catch Keisuke Fujita 1*, Mitsuhiro Iwaki 2,3*, Atsuko H. Iwane 1, Lorenzo Marcucci 1 & Toshio
Degenerate ground-state lattices of membrane inclusions
 PHYSICAL REVIEW E 78, 011401 008 Degenerate ground-state lattices of membrane inclusions K. S. Kim, 1 Tom Chou,, * and Joseph Rudnick 3 1 Lawrence Livermore National Laboratory, Livermore, California 94550,
PHYSICAL REVIEW E 78, 011401 008 Degenerate ground-state lattices of membrane inclusions K. S. Kim, 1 Tom Chou,, * and Joseph Rudnick 3 1 Lawrence Livermore National Laboratory, Livermore, California 94550,
Monte Carlo simulation calculation of the critical coupling constant for two-dimensional continuum 4 theory
 Monte Carlo simulation calculation of the critical coupling constant for two-dimensional continuum 4 theory Will Loinaz * Institute for Particle Physics and Astrophysics, Physics Department, Virginia Tech,
Monte Carlo simulation calculation of the critical coupling constant for two-dimensional continuum 4 theory Will Loinaz * Institute for Particle Physics and Astrophysics, Physics Department, Virginia Tech,
The Euclidean Propagator in Quantum Models with Non-Equivalent Instantons
 Proceedings of Institute of Mathematics of NAS of Ukraine 2002, Vol. 43, Part 2, 609 615 The Euclidean Propagator in Quantum Models with Non-Equivalent Instantons Javier CASAHORRAN Departamento de Física
Proceedings of Institute of Mathematics of NAS of Ukraine 2002, Vol. 43, Part 2, 609 615 The Euclidean Propagator in Quantum Models with Non-Equivalent Instantons Javier CASAHORRAN Departamento de Física
Elasticity of Short DNA Molecules: Theory and Experiment for Contour Lengths of mm
 4360 Biophysical Journal Volume 93 December 2007 4360 4373 Elasticity of Short DNA Molecules: Theory and Experiment for Contour Lengths of 0.6 7 mm Yeonee Seol,* Jinyu Li, y Philip C. Nelson, { Thomas
4360 Biophysical Journal Volume 93 December 2007 4360 4373 Elasticity of Short DNA Molecules: Theory and Experiment for Contour Lengths of 0.6 7 mm Yeonee Seol,* Jinyu Li, y Philip C. Nelson, { Thomas
Dynamics of a mass-spring-pendulum system with vastly different frequencies
 Dynamics of a mass-spring-pendulum system with vastly different frequencies Hiba Sheheitli, hs497@cornell.edu Richard H. Rand, rhr2@cornell.edu Cornell University, Ithaca, NY, USA Abstract. We investigate
Dynamics of a mass-spring-pendulum system with vastly different frequencies Hiba Sheheitli, hs497@cornell.edu Richard H. Rand, rhr2@cornell.edu Cornell University, Ithaca, NY, USA Abstract. We investigate
arxiv: v3 [cond-mat.stat-mech] 1 Dec 2016
![arxiv: v3 [cond-mat.stat-mech] 1 Dec 2016 arxiv: v3 [cond-mat.stat-mech] 1 Dec 2016](/thumbs/93/112509456.jpg) Effects of Landau-Lifshitz-Gilbert damping on domain growth Kazue Kudo Department of Computer Science, Ochanomizu University, 2-1-1 Ohtsuka, Bunkyo-ku, Tokyo 112-861, Japan (Dated: December 2, 216) arxiv:162.6673v3
Effects of Landau-Lifshitz-Gilbert damping on domain growth Kazue Kudo Department of Computer Science, Ochanomizu University, 2-1-1 Ohtsuka, Bunkyo-ku, Tokyo 112-861, Japan (Dated: December 2, 216) arxiv:162.6673v3
Energy functions and their relationship to molecular conformation. CS/CME/BioE/Biophys/BMI 279 Oct. 3 and 5, 2017 Ron Dror
 Energy functions and their relationship to molecular conformation CS/CME/BioE/Biophys/BMI 279 Oct. 3 and 5, 2017 Ron Dror Yesterday s Nobel Prize: single-particle cryoelectron microscopy 2 Outline Energy
Energy functions and their relationship to molecular conformation CS/CME/BioE/Biophys/BMI 279 Oct. 3 and 5, 2017 Ron Dror Yesterday s Nobel Prize: single-particle cryoelectron microscopy 2 Outline Energy
5 Topological defects and textures in ordered media
 5 Topological defects and textures in ordered media In this chapter we consider how to classify topological defects and textures in ordered media. We give here only a very short account of the method following
5 Topological defects and textures in ordered media In this chapter we consider how to classify topological defects and textures in ordered media. We give here only a very short account of the method following
Laplacian modes for calorons and as a filter
 Instituut-Lorentz for Theoretical Physics, University of Leiden, 2300 RA Leiden, The Netherlands E-mail: bruckmann@lorentz.leidenuniv.nl Ernst-Michael Ilgenfritz Institut für Physik, Humboldt Universität
Instituut-Lorentz for Theoretical Physics, University of Leiden, 2300 RA Leiden, The Netherlands E-mail: bruckmann@lorentz.leidenuniv.nl Ernst-Michael Ilgenfritz Institut für Physik, Humboldt Universität
Vortex stretching in incompressible and compressible fluids
 Vortex stretching in incompressible and compressible fluids Esteban G. Tabak, Fluid Dynamics II, Spring 00 1 Introduction The primitive form of the incompressible Euler equations is given by ( ) du P =
Vortex stretching in incompressible and compressible fluids Esteban G. Tabak, Fluid Dynamics II, Spring 00 1 Introduction The primitive form of the incompressible Euler equations is given by ( ) du P =
Simple Simulations of DNA Condensation
 130 Biophysical Journal Volume 80 January 2001 130 139 Simple Simulations of DNA Condensation Mark J. Stevens Sandia National Laboratory, P.O. Box 5800, MS 1111, Albuquerque, New Mexico 87185 USA ABSTRACT
130 Biophysical Journal Volume 80 January 2001 130 139 Simple Simulations of DNA Condensation Mark J. Stevens Sandia National Laboratory, P.O. Box 5800, MS 1111, Albuquerque, New Mexico 87185 USA ABSTRACT
Multimedia : Podcast : Sacrificial Bonds in Biological Materials; Fantner, et al. Biophys. J , 1411
 3.52 Nanomechanics o Materials and Biomaterials Tuesday 5/1/7 Pro. C. Ortiz, MIT-DMSE I LECTURE 2: THEORETICAL ASPECTS O SINGLE MOLECULE ORCE SPECTROSCOPY 2 : EXTENSIBILITY AND THE WORM LIKE CHAIN (WLC)
3.52 Nanomechanics o Materials and Biomaterials Tuesday 5/1/7 Pro. C. Ortiz, MIT-DMSE I LECTURE 2: THEORETICAL ASPECTS O SINGLE MOLECULE ORCE SPECTROSCOPY 2 : EXTENSIBILITY AND THE WORM LIKE CHAIN (WLC)
Asymptotic series in quantum mechanics: anharmonic oscillator
 Asymptotic series in quantum mechanics: anharmonic oscillator Facultat de Física, Universitat de Barcelona, Diagonal 645, 0808 Barcelona, Spain Thesis supervisors: Dr Bartomeu Fiol Núñez, Dr Alejandro
Asymptotic series in quantum mechanics: anharmonic oscillator Facultat de Física, Universitat de Barcelona, Diagonal 645, 0808 Barcelona, Spain Thesis supervisors: Dr Bartomeu Fiol Núñez, Dr Alejandro
On the Symmetric Molecular Conjectures
 On the Symmetric Molecular Conjectures Josep M. Porta, Lluis Ros, Bernd Schulze, Adnan Sljoka, and Walter Whiteley Abstract A molecular linkage consists of a set of rigid bodies pairwise connected by revolute
On the Symmetric Molecular Conjectures Josep M. Porta, Lluis Ros, Bernd Schulze, Adnan Sljoka, and Walter Whiteley Abstract A molecular linkage consists of a set of rigid bodies pairwise connected by revolute
arxiv: v1 [cond-mat.stat-mech] 4 Feb 2008
![arxiv: v1 [cond-mat.stat-mech] 4 Feb 2008 arxiv: v1 [cond-mat.stat-mech] 4 Feb 2008](/thumbs/88/115727112.jpg) Exact Solution to Ideal Chain with Fixed Angular Momentum J. M. Deutsch Department of Physics, University of California, Santa Cruz, CA 9564. arxiv:8.486v1 [cond-mat.stat-mech] 4 Feb 8 (Dated: February
Exact Solution to Ideal Chain with Fixed Angular Momentum J. M. Deutsch Department of Physics, University of California, Santa Cruz, CA 9564. arxiv:8.486v1 [cond-mat.stat-mech] 4 Feb 8 (Dated: February
Macroscopic modeling and simulations of supercoiled DNA with bound proteins
 JOURNAL OF CHEMICAL PHYSICS VOLUME 117, NUMBER 18 8 NOVEMBER 2002 Macroscopic modeling and simulations of supercoiled DNA with bound proteins Jing Huang a) and Tamar Schlick b) Department of Chemistry
JOURNAL OF CHEMICAL PHYSICS VOLUME 117, NUMBER 18 8 NOVEMBER 2002 Macroscopic modeling and simulations of supercoiled DNA with bound proteins Jing Huang a) and Tamar Schlick b) Department of Chemistry
The Photon Counting Histogram (PCH)
 The Photon Counting Histogram (PCH) While Fluorescence Correlation Spectroscopy (FCS) can measure the diffusion coefficient and concentration of fluorophores, a complementary method, a photon counting
The Photon Counting Histogram (PCH) While Fluorescence Correlation Spectroscopy (FCS) can measure the diffusion coefficient and concentration of fluorophores, a complementary method, a photon counting
Uncoupling shear and uniaxial elastic moduli of semiflexible biopolymer networks:
 Supplemental document for: Uncoupling shear and uniaxial elastic moduli of semiflexible biopolymer networks: compression-softening and stretch-stiffening Anne S. G. van Oosten 1, Mahsa Vahabi 2, Albert
Supplemental document for: Uncoupling shear and uniaxial elastic moduli of semiflexible biopolymer networks: compression-softening and stretch-stiffening Anne S. G. van Oosten 1, Mahsa Vahabi 2, Albert
Light-Cone Quantization of Electrodynamics
 Light-Cone Quantization of Electrodynamics David G. Robertson Department of Physics, The Ohio State University Columbus, OH 43210 Abstract Light-cone quantization of (3+1)-dimensional electrodynamics is
Light-Cone Quantization of Electrodynamics David G. Robertson Department of Physics, The Ohio State University Columbus, OH 43210 Abstract Light-cone quantization of (3+1)-dimensional electrodynamics is
Intensity (a.u.) Intensity (a.u.) Raman Shift (cm -1 ) Oxygen plasma. 6 cm. 9 cm. 1mm. Single-layer graphene sheet. 10mm. 14 cm
 Intensity (a.u.) Intensity (a.u.) a Oxygen plasma b 6 cm 1mm 10mm Single-layer graphene sheet 14 cm 9 cm Flipped Si/SiO 2 Patterned chip Plasma-cleaned glass slides c d After 1 sec normal Oxygen plasma
Intensity (a.u.) Intensity (a.u.) a Oxygen plasma b 6 cm 1mm 10mm Single-layer graphene sheet 14 cm 9 cm Flipped Si/SiO 2 Patterned chip Plasma-cleaned glass slides c d After 1 sec normal Oxygen plasma
A Renormalization Group Primer
 A Renormalization Group Primer Physics 295 2010. Independent Study. Topics in Quantum Field Theory Michael Dine Department of Physics University of California, Santa Cruz May 2010 Introduction: Some Simple
A Renormalization Group Primer Physics 295 2010. Independent Study. Topics in Quantum Field Theory Michael Dine Department of Physics University of California, Santa Cruz May 2010 Introduction: Some Simple
Semiflexible macromolecules in quasi-one-dimensional confinement: Discrete versus continuous bond angles
 THE JOURNAL OF CHEMICAL PHYSICS 143, 243102 (2015) Semiflexible macromolecules in quasi-one-dimensional confinement: Discrete versus continuous bond angles Aiqun Huang, 1,a) Hsiao-Ping Hsu, 2,3,b) Aniket
THE JOURNAL OF CHEMICAL PHYSICS 143, 243102 (2015) Semiflexible macromolecules in quasi-one-dimensional confinement: Discrete versus continuous bond angles Aiqun Huang, 1,a) Hsiao-Ping Hsu, 2,3,b) Aniket
Elasticity of Short DNA Molecules: Theory and Experiment for Contour Lengths of µm
 University of Pennsylvania ScholarlyCommons Department of Physics Papers Department of Physics 12-2007 Elasticity of Short DNA Molecules: Theory and Experiment for Contour Lengths of 0.6 7 µm Yeonee Seol
University of Pennsylvania ScholarlyCommons Department of Physics Papers Department of Physics 12-2007 Elasticity of Short DNA Molecules: Theory and Experiment for Contour Lengths of 0.6 7 µm Yeonee Seol
