Mestrelab s Users Meeting. Santi Dominguez MBA CEO Dr Manuel Pérez Senior VP October 22 nd 2015
|
|
|
- Dortha Arleen Moody
- 5 years ago
- Views:
Transcription
1 Mestrelab s Users Meeting Santi Dominguez MBA CEO Dr Manuel Pérez Senior VP October 22 nd 2015
2 Layout Layout 13:00 13:10: Introduction 13:10 13:50: MestreNova(Mnova) NMR plugin: a wealth of functionality 13:50 14:20: Mnova Verification: predictions and understanding results 14:20 14:40: Coffee Break 14:20 14:50: MnovaSMA and qnmr: analysis and quantification of single and multiple compounds 14:50 15:20: MnovaDB: using databases to capture, share and create knowledge 15:20 15:40: Mnova MS plugin: main characteristics 16:00 17:00: Q&A
3 10 years of Mestrelab!!! From1st Dec2004 today From2 FTE to 30 FTE From 10 customers to 85,000 1/12/ /09/2015 Intro & What s New 3
4 The development team: Nikolay Larin(Head of Software Development) Isaac Iglesias (Developer) Felipe Seoane(Developer) Pablo Monje (Apps Chemist) Santi Fraga (Developer) Michael Goebel(Developer) Esther Vaz (QA PhD Chemist) Santi López (Developer) José Antonio García (Developer) Oleg Ovchinnikov (Developer) Agustin Barba (Developer) José Antonio García Pulido (Developer) The scientific team: Carlos Cobas Javier Sardina Mike Bernstein Manuel Perez Chen Peng Brian Marquez The business team: Santi Dominguez Chen Peng Cristina Geada Dani Fraga Olalla Lema Manuel Perez Enrique Sánchez Izabella Kroll The Team We are over> in thefamily! The scientific partners(collaborations and contractors) Modgraph Consultants Ltd Rashmi Mistry Jeff Seymour Mike Wainwright Wolfgang Robien Ray Abraham Erno Pretsch Portanova GmbH Pius Portmann USC Manuel Martin Pastor Origenis Michael Thormann ebyte Stan Sykora Nucleomatica Giuseppe Balacco Sierra Analytics David Stranz Scott Campbell George Maydwell Mspin Armando Navarro R4R Silvia Mari Intro & What s New 5
5 The customers Continued market expansion led by NMR 34 of top Worlduniversitieshavecampus licensesof Mnova(9 of top 10) Latest additions are Stanford, Oxford, Nottingham and Strasbourg Several pharmaceutical and chemical companies have gone for Company wide Mnova installations: Pfizer Novartis BASF LGChem Boehringer Ingelheim Leo Pharma Intro & What s New 6
6 The customers Japan Japanese market starting to grow Very important market, huge potential and high level of expertise Hand in hand with SystemPlus, who are our partners for supporting the Japanese market Thisisourfirsttripto Japan, buttherewillbe manymore Full training session in Spring 2016 Intro & What s New 6
7 Areas y Methodologies Manufacturing Development Research Mnova MS Databases SMA qnmr Verification and Prediction NMR: Functionality
8 What most of us want i
9 It is easy
10 Introduction The Trinity of NMR (or a combination of) Integra tion Chemic al Shift Coupli ng
11 Phasing, Baseline correction (and lies) Will influence: Integration Chemical Shift Need to: Learn to appreciate their impact Know your algorithms
12 Pre-Acquisiton Considerations O O Expansion of aromatic region d1 = 0 sec d1 = 1 sec d1 = 3 sec d1 = 10 sec OH 37 % error (on signal at 7.8 ppm) 29 % error (Standard 1H) 17 % error 3 % error (Proton_Quantitative )
13 Phasing Strategies No perfect automatic solution But. Play!
14 Phase correction: Let s look at some integrals ph0 +3
15 Baseline Correction Considerations Influence over quantitation Multiple Choices Smooth: Polynomial Methods Aggressive: Smoothers, Splines Manual: slower but more precise
16 Baseline correction Whittaker: Influence on integrals
17 Referencing: Absolute Referencing Influence over chemical shift Consistency IUPAC recommendation: unified scale Added benefit: Alignment across all experiments
18 Absolute Referencing in Mnova For all nuclei: Manage References:
19 Peak Picking Different Approaches Standard Global Spectral Deconvolution Effect over: Chemical Shift Coupling
20 Standard Peak Picking Simple algorithm: Look for peak maxima Good enough for certain cases But. Peak overlap Insufficient peak resolution Not accurate coupling constants
21 GSD Intro Multiplication of the FID by a resolution-enhancement function But: poorer SNR and peak shape distortions
22 GSD Intro Resolution Booster Original Integral part of GSD
23 Why GSD (overlap) Overlap
24 Why GSD (Auto-classification of peaks)
25 Why GSD - Couplings
26 But why?! Because we can reconstruct the data
27 To the report! Now that some of you are scared Creation of reporting templates, including: Exact look and feel All the analysis operations (phase and baseline correction, peak picking, autoclassification, reports of multiplets and parameters )
28
29 Tracking Data COSY TOCSY NOESY HSQC HMBC 1 H- 15 N HMBC
30 Tracking assignments Correlations Assignment Tables
31 Data Analysis T1 calculation
32 Reaction Monitoring by NMR
33 Verification and Prediction Areas y Methodologies Manufacturing Development Research Mnova MS Databases SMA qnmr Verification and Prediction NMR: Functionality
34 Verification and Prediction Approaches 1 H NMR Increments (Prof Erno Prest) Conformers (Prof Ray Abraham) Access through:
35 Verification and Prediction Approaches 13 C NMR HOSE Code Wolfgang Robien Neural Networks Wolfgang Robien
36 1 H NMR Prediction Increments Similar (but different to HOSE) Uses substructures Very accurate Easy to anonymize Over 10,000 structurally different molecules in the DB Data organised into multiple databases
37 1 H NMR Prediction Conformers Semi empirical approach Takes into consideration the 3D structure of the compound Thousands of compounds used to create the models Effective when dealing with flexible systems Visualise the conformers that contribute to the predictions
38 13 C Prediction 13 C HOSE Concept of Shells : how similar are the substructures as we go further away from the Carbon atom predicted: 5 shells: perfectly represented 1 shell: not represented Over compounds in the prediction database Constantly updated
39 13 C Prediction 13 C Neural Networks Fall-back approach More accurate than any other when no representative data can be found in the database
40 Method Selection Best Approach and Error Estimation Algorithm to decide which model produces the best predictions Errors calculated following considerable atom by atom type analysis
41 Solvents Solvents For 1 H NMR three solvents are considered by default: CDCl 3 DMSO D 2 O For 13 C NMR the solvent is automatically detected and the predictions are produced from relevant data.
42 Error interpretation Error Interpretation It is possible for any given prediction to display the estimated errors of the prediction graphically or in a table format:
43 Results Result Overview The inclusion of user data has a marked effect: Total refers to the effect over the entire of the dataset, Partial only for those atom that showed a change Total Partial Total Partial 13C 13C 1H 1H Improvement %
44 Structural Determination Need workflows to Track information across multiple Record these effectively But need the killer experiment
45 Verification What is verification Is my Structure compatible with my data?
46 Verification What is Mnova Verify? Candidate Structures(one or several) Spectra 1H or/& 13C or/& HSQC or/& LC/GC/MS NMRPrediction (1H, 13C, 19F, 31P, 17º, 19N, 29Si) -CHARGE - Increments -HOSE -CSEARCH MS Prediction -m/z value - Isotopic cluster pattern -MSMS NMR Analysis - Autoprocess - GSD Enhance resolution Quantitate overlapped peaks - Autoedit Mark solvent lines Identify impurities Label labiles MS Analysis -Autoprocess - Integration/Peak Pick - Clustering Comparison Test 1 Global metrics Test 2 Nuclide count Test 3 Predicted multiplets Test 4 Js and assignme nts Test 5 HSQC XCOR Test 6 Isotope cluster distance Test 7 MSMS Peaks Scoring System(Scores and Significances) a novel scoring system able to apply fuzzy logic to combining many tests of different reliability in each specific case
47 Applications Structure Verification Foundation: GSD Auto classification of peaks Assignment Classification
48 Verification Reporting and result visualization Meaningful feedback for chemists on verification results Assignments on molecule Percentage or score result NMR Purity
49 Reporting and result visualization Meaningful feedback for chemists on verification results Results explained Reasons for specific verification scores
50 Integration with concentration calculation Quantification and Verification We can now run concentration and purity calculations as part of the verification workflow generate extra value for verification users and get more information out of your data with quantification for free
51 Reporting and result visualization Latest Developments in Mnova Verify Meaningful feedback for chemists on verification results Acquisition issues which may affect verification or quantification accuracy
52 Latest Developments in Mnova Verify Ranking between possible reaction outcomes Confidence for chemists on the outcome of their reactions
53 Latest Developments in Mnova Verify Ranking between possible reaction outcomes Meaningful feedback for chemists on verification results
54 Automation in Verify Automation - Batch There is now a batch verification plugin available, which allows users to verify any number of samples in automation
55 Automation - Listener Latest Developments in Mnova Verify Verification can now be run as a listener launched by the arrival of new data in a specifically preselected location
56 Latest Developments in Mnova Verify Automation Verification listener
57 Result visualisation Multiple verification review Verification viewer allows the fast review of any number of verification analysis Results for specific well Detailed Results for specific well Wellplate Concentration information
58 Integration with MyData / Spectral DB Databasing of Verify results Storage in Spectral DB integrated as part of the verification workflow
59 Reporting and result visualization Latest Developments in Mnova Verify Integration of verification with layout templates for automated report generation including reporting results
60 Verification and Prediction Areas y Methodologies Manufacturing Development Research Mnova MS Databases SMA qnmr Verification and Prediction NMR: Functionality
61 Methodologies Internal Reference External reference Artificial Signal Conversion Factor 2D Methods Relative Response Factors
62 qnmr qnmr: General aspects of data processing Sample preparation, shimming, temperature stability Interpulse delay Signal-to-noise-ratio (SNR)(guideline: 150) Number of data points/peak Apodisation Phase correction Baseline correction Integration method Integration regions Blind regions
63 qnmr Basics of qnmr spectroscopy Fundamental relationship of qnmr: the signal intensity in the NMR spectrum is directly proportional to the number of nuclei responsible for that particular resonance (1) so I I X X = N K X S N X where K S is the spectrometer constant and remains the same for all the resonances in a NMR spectrum Relative quantitation method The easiest method where the molar ratio between two compounds (M x /M y ) can be calculated M M X X ( 2) = Y I N X N I Y Y Absolute quantitation method and Purity By determining the absolute concentration of an analyte it is possible to derive its purity: I. If all the impurities (or other components) present in the NMR spectrum can be measured quantitatively, then the assay is just a difference from the 100% value II. Otherwise purity of a compound X is derived by the comparison with a standard according to the formula: I N MW X STD X ( 3) PX = ISTD NX MWSTD W W STD P STD where, I, N, M, W and P are integral area, number of nuclei, molar mass, gravimetric weight and purity of analyte (X) and standard (STD), respectively 63
64 Potency Potency (Purity) Purity/Potency
65 Concentration Determination Concentration Response factor calculation
66 qnmr Setup: load Spectrum, Compound Library and General Settings W-qNMR directory Open in Mnova 66
67 Adding DMSO2 as a reference to the table of compounds qnmr Finally: Save Compound Library 67
68 qnmr Check that Nuclides are correct 68
69 qnmr Preparing to Calculate Do this after Multiplets Ananlysis using Sum integration! 69
70 qnmr Expected result 70
71 qnmr Save experiment dialogue Check Analysis with replicates if >1 spectrum of a sample 71
72 qnmr Running an experiment 72
73 Verification and Prediction Areas y Methodologies Manufacturing Development Research Mnova MS Databases SMA qnmr Verification and Prediction NMR: Functionality
74 SMA Mnova capabilities for mixtures analysis Unsupervised methods Targeted analysis(=quantitation of defined substances) qnmr Simple Mixtures Analysis(SMA) 74 74
75 SMA is useful for Drug substances for release (CoA) Compound "Purity" determinations Quantitate the components of biological fluids, fermentation broths Wines and beverages Oligo-/polysaccharide hydrolysis sugar analysis Petroleum analysis PUFAs Metabolomics Other Low-field, high-field, 1D, 2D data 75
76 SMA You can design an SMA experiment to: Search for a multiplet or integrate a spectral region Use of GSD in areas of peak overlap Peak pattern recognition SMA will automatically determine these areas for each spectrum Specify an equationthat converts that area into meaningful quantification If necessary, use an internal quantitation standard / 76
77 SMA basics Other data Quantitation of C 1 Equation for SMA I1 I2 I3 77
78 SMA basics Equations Automatic and manual Measurement Equation SMA equation Mole ratio (I1*NNR1)/(IR1*NN1) Concentration Purity Ref: RC * (NN1/I1) Compound: CCF * (I1/NN1) (I1*NNR1*RW*MW) /(IR1*NN1*SW*MWR) Statistics,e.g. average )/2 0.5*(C_A+ C_B) 78
79 SMA basics Further flexibility in equation usage Factorise partly hidden, or semi-saturated signals 1D or 2D Specify the NMR experiment for a sample mix and match Use a previous result in a later equation 79
80 Extending targeted analysis Multiple spectral information for a sample Extends the scope of targeted analysis 80
81 Extending targeted analysis Helping to deal with overlap AI AI t = AI* 2 AI AI t = AI-AI 81 81
82 SMA workflow for complex mixtures The SMA analysis workflow: analyse-alert-refinereanalyse Search for multiplets Alert the user to possible false peak identification (J, relative integral, etc.) or review each region Allow the user to manually fix a problem Reanalyse Reporting 82
83 SMA worked example Example: concentrations of Aspirin C components Acetyl salicylic acid Salicylic acid Ascorbic acid Citric acid Acetic acid 83
84 Verification and Prediction Areas y Methodologies Manufacturing Development Research Mnova MS Databases SMA qnmr Verification and Prediction NMR: Functionality
85 Verification and Prediction Areas y Methodologies Manufacturing Development Research Mnova MS Databases SMA qnmr Verification and Prediction NMR: Functionality
86 LC/MS data formats Mnova MS can open Formats supported Vendor Windows Mac Linux Agilent ChemStation, MassHunter, Ion Trap ChemStation ChemStation Bruker 1 XMass, Compass XMass XMass Waters MassLynx MassLynx MassLynx Thermo Scientific Xcalibur, Exactive, Q-Exactive JEOL MSQ 1000, FastFlight AB SCIEX Analyst Shimadzu 2 LabSolutions mzdata, mzxml mzdata, mzxml mzdata, mzxml mzdata, mzxml NetCDF ANDI-MS NetCDF ANDI-MS
87 To open your LC/MS data Opening data Choose File Page Setup Orientation and change the page orientation to portrait, if you prefer Choose File Opento open any file in the folder containing the raw data, or drag/dropthe folder from Windows Explorer to Mnova Mnovaautomatically converts your data and does peak picking TIC Drag & drop Mass spectrum (at highest TIC)
88 To browse the mass spectra Browsing Data Click to switch to crosshair cursor, and click on the TIC to display the mass spectrum at that retention time, or click-and-drag to display co-added spectra Click to change to appending mode if you want to display multiple mass spectra Choose the Spectrum Selection Mode to display mass spectra conveniently: Manual mode: You click to display a single MS, or click-and-drag to co-add multiple MS Peak mode: You click on a peak to display the co-added MS within the peak range Peak (Background subtraction) mode: You click on a peak to display the co-added MS within the peak range with the background subtracted
89 To browse the UV traces MS browser Click to show the MS Browser dialog Highlight the Total UV Absorbance under Traces, and click to display it Repeat the above step to display the other traces if any
90 Adding a chromatogram To extract a UV trace at selected wavenumber Highlight the DAD in the MS Browser Click to display it In the New Chromatogram dialog, choose Wavelength, and enter a wave length and a tolerance to display the extracted UV trace
91 To display a UV spectrum at selected retention time If available, highlight the DAD Trace in the MS Browser and click to display it Click for Crosshair cursor, press and hold Alt key, click on the DAD trace to display the UV spectrum at that retention time. DAD In crosshair cursor mode, click on the PDA curve to display the UV spectrum at that retention time UV spectrum
92 To edit and report peak integration results Peaks are automatically integrated when you open a chromatogram Use the Peak Detection tool menu to re-detect peaks, add, delete or clear peaks Hover your cursor on the wedges, click and drag the green boxes to change the range of a peak Or press Shift, click and drag the green boxes to change the baseline of a peak Choose View Tables Mass Peaksto display or report the Mass Peaks Table SHIFT+ Drag
93 To display extracted ion chromatogram from an m/zvalue Click (or choose Mass Analysis New Mass Chromatogram Manually) In the New Chromatogram dialog, enter the m/z value that you are interested in, and a suitable Tolerance Press OK to display the EIC TIC EIC at /-0.25 Da
94 To display extracted ion chromatogram for an MS peak TIC First display the MS trace and zoom into the molecular ion peak that you are interested. Next select Click (or choose Mass Analysis New Mass Chromatogram Graphically), clickand-drag around the peak to define a mass range An EIC will be displayed within the mass range MS at 3.08 min EIC at Da
95 To confirm proposed structures using Molecule Match Molecular Match TIC Import one or several structures by copy/pasting from ChemDraw, Isis/Draw or ChemSketch, or by opening.mol or.sdf files. Click (or choose Mass Analysis Molecule Match Calculate). In the Molecule Match Table, click on a molecule to see the matching results Matched Isotope Cluster Chromat. Matched Isotope Cluster Mol Match Results
96 Molecular Match You can choose Mass Analysis Molecule Match Settings to change the settings for Molecule Match. The default settings are for low-resolution MS. Change Tolerance to 5-10 ppm if you are using high-resolution MS. Edit the Adducts or Losses and other parameters if you want. Click to run the Molecule Match again To confirm proposed structures using Molecule Match (2) Tip: Click the + buttons to add a new adduct. Enter + for a radical cation. Highlight one and click the x button to remove it. Click Restore to reset to the default or previously saved settings.
97 To confirm proposed MF using Molecule Match Molecular Match If you don t have a structure but only a MF, choose the Calculate From Molecular Formula tool Enter one or more molecular formulas The results are displayed in Molecule Match Table
98 To predict isotope clusters from a mol. formula High Resolution data Choose Mass Analysis Spectrum Prediction Predict Enter one or more molecular formulae to predict the isotope clusters with all predefined adducts and losses Click on TIC to display a mass spec (or a range of co-added mass spec), and highlight a row in the Mass Prediction Table. Mnovadisplays the predicted isotope cluster with matched peaks in the experimental peaks, if any Choose an adduct/loss Click on TIC to choose a mass spec Predicted (blue) and matched experimental peaks (red)
99 To enumerate elemental compositions Elemental composition Zoom into the molecular ion peak of a MS trace Click or choose Mass Analysis Elemental Composition Calculate. Click on the molecule ion peak. An Elemental Composition Table is displayed Click on a row to see the match of observed and predicted isotope peaks Choose Mass Analysis Elemental Composition Settings to change the settings if necessary. Then click on the ion peak to recalculate. Possible Elemental Compositions Match of observed and predicted isotope peaks
100 Acknowledgements Mestrelab Team Questions?
Mnova Software Tools for Fragment-Based Drug Discovery
 Mnova Software Tools for Fragment-Based Drug Discovery Chen Peng, PhD, VP of Business Development, US & China Mestrelab Research SL San Diego, CA chen.peng@mestrelab.com 858.736.4563 Agenda Brief intro
Mnova Software Tools for Fragment-Based Drug Discovery Chen Peng, PhD, VP of Business Development, US & China Mestrelab Research SL San Diego, CA chen.peng@mestrelab.com 858.736.4563 Agenda Brief intro
Mnova Software for Analyzing Reaction Monitoring NMR Spectra
 Mnova Software for Analyzing Reaction Monitoring NMR Spectra Version 10 Chen Peng, PhD, VP of Business Development, US & China Mestrelab Research SL San Diego, CA, USA chen.peng@mestrelab.com 858.736.4563
Mnova Software for Analyzing Reaction Monitoring NMR Spectra Version 10 Chen Peng, PhD, VP of Business Development, US & China Mestrelab Research SL San Diego, CA, USA chen.peng@mestrelab.com 858.736.4563
Using MnovaScreen to Process, Analyze and Report Ligand-Protein Binding Spectra for Fragment-based Lead Design
 September 2014 Using MnovaScreen to Process, Analyze and Report Ligand-Protein Binding Spectra for Fragment-based Lead Design Dr Manuel Perez Senior VP - Mestrelab 1996: A research project in University
September 2014 Using MnovaScreen to Process, Analyze and Report Ligand-Protein Binding Spectra for Fragment-based Lead Design Dr Manuel Perez Senior VP - Mestrelab 1996: A research project in University
MassHunter Software Overview
 MassHunter Software Overview 1 Qualitative Analysis Workflows Workflows in Qualitative Analysis allow the user to only see and work with the areas and dialog boxes they need for their specific tasks A
MassHunter Software Overview 1 Qualitative Analysis Workflows Workflows in Qualitative Analysis allow the user to only see and work with the areas and dialog boxes they need for their specific tasks A
MassHunter TOF/QTOF Users Meeting
 MassHunter TOF/QTOF Users Meeting 1 Qualitative Analysis Workflows Workflows in Qualitative Analysis allow the user to only see and work with the areas and dialog boxes they need for their specific tasks
MassHunter TOF/QTOF Users Meeting 1 Qualitative Analysis Workflows Workflows in Qualitative Analysis allow the user to only see and work with the areas and dialog boxes they need for their specific tasks
ProMass Deconvolution User Training. Novatia LLC January, 2013
 ProMass Deconvolution User Training Novatia LLC January, 2013 Overview General info about ProMass Features Basics of how ProMass Deconvolution works Example Spectra Manual Deconvolution with ProMass Deconvolution
ProMass Deconvolution User Training Novatia LLC January, 2013 Overview General info about ProMass Features Basics of how ProMass Deconvolution works Example Spectra Manual Deconvolution with ProMass Deconvolution
Agilent MassHunter Quantitative Data Analysis
 Agilent MassHunter Quantitative Data Analysis Presenters: Howard Sanford Stephen Harnos MassHunter Quantitation: Batch Table, Compound Information Setup, Calibration Curve and Globals Settings 1 MassHunter
Agilent MassHunter Quantitative Data Analysis Presenters: Howard Sanford Stephen Harnos MassHunter Quantitation: Batch Table, Compound Information Setup, Calibration Curve and Globals Settings 1 MassHunter
Agilent MassHunter Quantitative Data Analysis
 Agilent MassHunter Quantitative Data Analysis Presenters: Howard Sanford Stephen Harnos MassHunter Quantitation: Batch and Method Setup Outliers, Data Review, Reporting 1 MassHunter Quantitative Analysis
Agilent MassHunter Quantitative Data Analysis Presenters: Howard Sanford Stephen Harnos MassHunter Quantitation: Batch and Method Setup Outliers, Data Review, Reporting 1 MassHunter Quantitative Analysis
NMR Spectra Processing, Verification and Elucidation: challenges and current progress
 NMR Spectra Processing, Verification and Elucidation: challenges and current progress Stanislav Sykora, Extra Byte, Italy www.ebyte.it Juan Carlos Cobas Gómez, Mestrelab, Spain www.mestrelab.com The abstract
NMR Spectra Processing, Verification and Elucidation: challenges and current progress Stanislav Sykora, Extra Byte, Italy www.ebyte.it Juan Carlos Cobas Gómez, Mestrelab, Spain www.mestrelab.com The abstract
Agilent MassHunter Profinder: Solving the Challenge of Isotopologue Extraction for Qualitative Flux Analysis
 Agilent MassHunter Profinder: Solving the Challenge of Isotopologue Extraction for Qualitative Flux Analysis Technical Overview Introduction Metabolomics studies measure the relative abundance of metabolites
Agilent MassHunter Profinder: Solving the Challenge of Isotopologue Extraction for Qualitative Flux Analysis Technical Overview Introduction Metabolomics studies measure the relative abundance of metabolites
2D NMR: HMBC Assignments and Publishing NMR Data Using MNova
 Homework 10 Chem 636, Fall 2014 due at the beginning of lab Nov 18-20 updated 10 Nov 2014 (cgf) 2D NMR: HMBC Assignments and Publishing NMR Data Using MNova Use Artemis (Av-400) or Callisto (Av-500) for
Homework 10 Chem 636, Fall 2014 due at the beginning of lab Nov 18-20 updated 10 Nov 2014 (cgf) 2D NMR: HMBC Assignments and Publishing NMR Data Using MNova Use Artemis (Av-400) or Callisto (Av-500) for
Making Sense of Differences in LCMS Data: Integrated Tools
 Making Sense of Differences in LCMS Data: Integrated Tools David A. Weil Agilent Technologies MassHunter Overview Page 1 March 2008 How Clean is our Water?... Page 2 Chemical Residue Analysis.... From
Making Sense of Differences in LCMS Data: Integrated Tools David A. Weil Agilent Technologies MassHunter Overview Page 1 March 2008 How Clean is our Water?... Page 2 Chemical Residue Analysis.... From
Intelligent NMR productivity tools
 Intelligent NMR productivity tools Till Kühn VP Applications Development Pittsburgh April 2016 Innovation with Integrity A week in the life of Brian Brian Works in a hypothetical pharma company / university
Intelligent NMR productivity tools Till Kühn VP Applications Development Pittsburgh April 2016 Innovation with Integrity A week in the life of Brian Brian Works in a hypothetical pharma company / university
New Approaches to the Development of GC/MS Selected Ion Monitoring Acquisition and Quantitation Methods Technique/Technology
 New Approaches to the Development of GC/MS Selected Ion Monitoring Acquisition and Quantitation Methods Technique/Technology Gas Chromatography/Mass Spectrometry Author Harry Prest 1601 California Avenue
New Approaches to the Development of GC/MS Selected Ion Monitoring Acquisition and Quantitation Methods Technique/Technology Gas Chromatography/Mass Spectrometry Author Harry Prest 1601 California Avenue
NMR Predictor. Introduction
 NMR Predictor This manual gives a walk-through on how to use the NMR Predictor: Introduction NMR Predictor QuickHelp NMR Predictor Overview Chemical features GUI features Usage Menu system File menu Edit
NMR Predictor This manual gives a walk-through on how to use the NMR Predictor: Introduction NMR Predictor QuickHelp NMR Predictor Overview Chemical features GUI features Usage Menu system File menu Edit
Automated, High- Throughput Data Processing & Quantification: Illustrated by a series of Non-Steroidal Anti-Inflammatory Drugs (NSAIDs)
 Automated, High- Throughput Data Processing & Quantification: Illustrated by a series of Non-Steroidal Anti-Inflammatory Drugs (NSAIDs) Automatic Data Processing Mnova offers a number of listeners that
Automated, High- Throughput Data Processing & Quantification: Illustrated by a series of Non-Steroidal Anti-Inflammatory Drugs (NSAIDs) Automatic Data Processing Mnova offers a number of listeners that
profileanalysis Innovation with Integrity Quickly pinpointing and identifying potential biomarkers in Proteomics and Metabolomics research
 profileanalysis Quickly pinpointing and identifying potential biomarkers in Proteomics and Metabolomics research Innovation with Integrity Omics Research Biomarker Discovery Made Easy by ProfileAnalysis
profileanalysis Quickly pinpointing and identifying potential biomarkers in Proteomics and Metabolomics research Innovation with Integrity Omics Research Biomarker Discovery Made Easy by ProfileAnalysis
Application Note 12: Fully Automated Compound Screening and Verification Using Spinsolve and MestReNova
 Application Note : Fully Automated Compound Screening and Verification Using Spinsolve and MestReNova Paul Bowyer, Magritek, Inc. and Mark Dixon, Mestrelab Sample screening to verify the identity or integrity
Application Note : Fully Automated Compound Screening and Verification Using Spinsolve and MestReNova Paul Bowyer, Magritek, Inc. and Mark Dixon, Mestrelab Sample screening to verify the identity or integrity
Compounding insights Thermo Scientific Compound Discoverer Software
 Compounding insights Thermo Scientific Compound Discoverer Software Integrated, complete, toolset solves small-molecule analysis challenges Thermo Scientific Orbitrap mass spectrometers produce information-rich
Compounding insights Thermo Scientific Compound Discoverer Software Integrated, complete, toolset solves small-molecule analysis challenges Thermo Scientific Orbitrap mass spectrometers produce information-rich
Using the ACD/MS Manager Software with Agilent 1100 Series LC/MS Systems. Application Note
 Using the ACD/MS Manager Software with Agilent 1100 Series LC/MS Systems Application Note Bryan Miller, Christine Miller, Paul Goodley, Masahiko Takino Introduction ACD/MS Manager software from Advanced
Using the ACD/MS Manager Software with Agilent 1100 Series LC/MS Systems Application Note Bryan Miller, Christine Miller, Paul Goodley, Masahiko Takino Introduction ACD/MS Manager software from Advanced
Benchtop NMR Combined with GC/MS Confirms Identity of Forensic Case Sample
 APPLICATION NOTE Benchtop NMR Combined with GC/MS Confirms Identity of Forensic Case Sample No. AN52889 Authors: Dean Antic, Ph.D., Thermo Fisher Scientific, San Jose, CA, USA WanLi Wei, Senior Engineer,
APPLICATION NOTE Benchtop NMR Combined with GC/MS Confirms Identity of Forensic Case Sample No. AN52889 Authors: Dean Antic, Ph.D., Thermo Fisher Scientific, San Jose, CA, USA WanLi Wei, Senior Engineer,
NMRPredict Functional Block Diagram
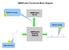 NMRPredict Functional Block Diagram 2 Submit to server. NMRPredict Server NMRPredict DeskTop Software Review results. 3 1 Input structure. Starting the NMRPredict Interface Click on the NMRPredict icon.
NMRPredict Functional Block Diagram 2 Submit to server. NMRPredict Server NMRPredict DeskTop Software Review results. 3 1 Input structure. Starting the NMRPredict Interface Click on the NMRPredict icon.
North Carolina State University Department of Chemistry Varian NMR Training Manual
 North Carolina State University Department of Chemistry Varian NMR Training Manual by J.B. Clark IV & Dr. S. Sankar 1 st Edition 05/15/2009 Section 3: Glide Program Operations for Advanced 1D & 2D Spectra
North Carolina State University Department of Chemistry Varian NMR Training Manual by J.B. Clark IV & Dr. S. Sankar 1 st Edition 05/15/2009 Section 3: Glide Program Operations for Advanced 1D & 2D Spectra
All Ions MS/MS: Targeted Screening and Quantitation Using Agilent TOF and Q-TOF LC/MS Systems
 All Ions MS/MS: Targeted Screening and Quantitation Using Agilent TOF and Q-TOF LC/MS Systems Technical Overview Introduction All Ions MS/MS is a technique that is available for Agilent high resolution
All Ions MS/MS: Targeted Screening and Quantitation Using Agilent TOF and Q-TOF LC/MS Systems Technical Overview Introduction All Ions MS/MS is a technique that is available for Agilent high resolution
Tutorial 2: Analysis of DIA data in Skyline
 Tutorial 2: Analysis of DIA data in Skyline In this tutorial we will learn how to use Skyline to perform targeted post-acquisition analysis for peptide and inferred protein detection and quantitation using
Tutorial 2: Analysis of DIA data in Skyline In this tutorial we will learn how to use Skyline to perform targeted post-acquisition analysis for peptide and inferred protein detection and quantitation using
Cerno Application Note Extending the Limits of Mass Spectrometry
 Creation of Accurate Mass Library for NIST Database Search Novel MS calibration has been shown to enable accurate mass and elemental composition determination on quadrupole GC/MS systems for either molecular
Creation of Accurate Mass Library for NIST Database Search Novel MS calibration has been shown to enable accurate mass and elemental composition determination on quadrupole GC/MS systems for either molecular
Agilent TOF Screening & Impurity Profiling Julie Cichelli, PhD LC/MS Small Molecule Workshop Dec 6, 2012
 1 Agilent TOF Screening & Impurity Profiling Julie Cichelli, PhD LC/MS Small Molecule Workshop Dec 6, 2012 Review: Technology for Accurate Mass Analysis: TOF LC/MS Mass measurements accurate to several
1 Agilent TOF Screening & Impurity Profiling Julie Cichelli, PhD LC/MS Small Molecule Workshop Dec 6, 2012 Review: Technology for Accurate Mass Analysis: TOF LC/MS Mass measurements accurate to several
Agilent All Ions MS/MS
 Agilent All Ions MS/MS Workflow Overview A Determine fragment ions for LC/MS Quant method B Develop final Quant method Develop LC/MS Qualitative Analysis method Process data with Find by Formula Build
Agilent All Ions MS/MS Workflow Overview A Determine fragment ions for LC/MS Quant method B Develop final Quant method Develop LC/MS Qualitative Analysis method Process data with Find by Formula Build
Cerno Bioscience MassWorks: Acquiring Calibration Data on Agilent GC/MSDs
 Cerno Bioscience MassWorks: Acquiring Calibration Data on Agilent GC/MSDs Application Note Author Harry Prest, Ph.D. Senior Chemist in GC-MS Agilent Technologies, Inc. 5301 Stevens Creek Blvd Santa Clara,
Cerno Bioscience MassWorks: Acquiring Calibration Data on Agilent GC/MSDs Application Note Author Harry Prest, Ph.D. Senior Chemist in GC-MS Agilent Technologies, Inc. 5301 Stevens Creek Blvd Santa Clara,
Skyline Small Molecule Targets
 Skyline Small Molecule Targets The Skyline Targeted Proteomics Environment provides informative visual displays of the raw mass spectrometer data you import into your Skyline documents. Originally developed
Skyline Small Molecule Targets The Skyline Targeted Proteomics Environment provides informative visual displays of the raw mass spectrometer data you import into your Skyline documents. Originally developed
Implementation of novel tools to facilitate fragment-based drug discovery by NMR:
 Implementation of novel tools to facilitate fragment-based drug discovery by NMR: Automated analysis of large sets of ligand-observed NMR binding data and 19 F methods Andreas Lingel Global Discovery Chemistry
Implementation of novel tools to facilitate fragment-based drug discovery by NMR: Automated analysis of large sets of ligand-observed NMR binding data and 19 F methods Andreas Lingel Global Discovery Chemistry
Quantification of JEOL XPS Spectra from SpecSurf
 Quantification of JEOL XPS Spectra from SpecSurf The quantification procedure used by the JEOL SpecSurf software involves modifying the Scofield cross-sections to account for both an energy dependency
Quantification of JEOL XPS Spectra from SpecSurf The quantification procedure used by the JEOL SpecSurf software involves modifying the Scofield cross-sections to account for both an energy dependency
La RMN quantitative appliquée aux petites molécules
 La RMN quantitative appliquée aux petites molécules Fabrice Moriaud - Applications Development - Fällanden 30ème Réunion d Utilisateurs RMN Bruker December 9, 2016 1 Covered in this presentation Quantification
La RMN quantitative appliquée aux petites molécules Fabrice Moriaud - Applications Development - Fällanden 30ème Réunion d Utilisateurs RMN Bruker December 9, 2016 1 Covered in this presentation Quantification
A Workflow Approach for the Identification and Structural Elucidation of Impurities of Quetiapine Hemifumarate Drug Substance
 A Workflow Approach for the Identification and Structural Elucidation of Impurities of Quetiapine Hemifumarate Drug Substance Michael D. Jones, Marian Twohig, Karen Haas, and Robert S. Plumb Waters Corporation,
A Workflow Approach for the Identification and Structural Elucidation of Impurities of Quetiapine Hemifumarate Drug Substance Michael D. Jones, Marian Twohig, Karen Haas, and Robert S. Plumb Waters Corporation,
WALKUP LC/MS FOR PHARMACEUTICAL R&D
 Pharmaceutical Workflow Solutions WALKUP LC/MS FOR PHARMACEUTICAL R&D Chemists, Peptide/Protein Chemists, Biologists, and Beyond MASSHUNTER WALKUP A Single User Interface for Robust and Reliable LC/MS
Pharmaceutical Workflow Solutions WALKUP LC/MS FOR PHARMACEUTICAL R&D Chemists, Peptide/Protein Chemists, Biologists, and Beyond MASSHUNTER WALKUP A Single User Interface for Robust and Reliable LC/MS
Protein Deconvolution Version 2.0
 Thermo Protein Deconvolution Version 2.0 User Guide XCALI-97414 Revision A August 2012 2012 Thermo Fisher Scientific Inc. All rights reserved. ReSpect is a trademark of Positive Probability Ltd. Xcalibur
Thermo Protein Deconvolution Version 2.0 User Guide XCALI-97414 Revision A August 2012 2012 Thermo Fisher Scientific Inc. All rights reserved. ReSpect is a trademark of Positive Probability Ltd. Xcalibur
SEAMLESS INTEGRATION OF MASS DETECTION INTO THE UV CHROMATOGRAPHIC WORKFLOW
 SEAMLESS INTEGRATION OF MASS DETECTION INTO THE UV CHROMATOGRAPHIC WORKFLOW Paula Hong, John Van Antwerp, and Patricia McConville Waters Corporation, Milford, MA, USA Historically UV detection has been
SEAMLESS INTEGRATION OF MASS DETECTION INTO THE UV CHROMATOGRAPHIC WORKFLOW Paula Hong, John Van Antwerp, and Patricia McConville Waters Corporation, Milford, MA, USA Historically UV detection has been
MetWorks Metabolite Identification Software
 m a s s s p e c t r o m e t r y MetWorks Metabolite Identification Software Enabling Confident Analysis of Metabolism Data Part of Thermo Fisher Scientific MetWorks Software for the Confident Analysis
m a s s s p e c t r o m e t r y MetWorks Metabolite Identification Software Enabling Confident Analysis of Metabolism Data Part of Thermo Fisher Scientific MetWorks Software for the Confident Analysis
THE UNBEARABLE FUZZINES OF NMR DATA?!?
 THE UNBEARABLE FUZZINES OF NMR DATA?!?... a brief reflection by Stan Sykora (ebyte.it)... Given the limited time, I will present only a few NMR-based illustrations of a broader question that is presently
THE UNBEARABLE FUZZINES OF NMR DATA?!?... a brief reflection by Stan Sykora (ebyte.it)... Given the limited time, I will present only a few NMR-based illustrations of a broader question that is presently
Tools for Structure Elucidation
 Innovation with Integrity Tools for Structure Elucidation Sandra Groscurth Workflow in Natural Product Research Organism Natural Products Extraction Activity Assay Characterization of Unknown Structures
Innovation with Integrity Tools for Structure Elucidation Sandra Groscurth Workflow in Natural Product Research Organism Natural Products Extraction Activity Assay Characterization of Unknown Structures
BASIC NMR HANDBOOK Written by M. A. Eastman Copyright 1997, 2001, 2013, 2015, 2018
 BASIC NMR HANDBOOK Written by M. A. Eastman Copyright 1997, 2001, 2013, 2015, 2018 Basic NMR Handbook Table of Contents: Preface ii viii PART 1 Chapter 1: Introduction to NMR 1 Why Study NMR? 1 The Magnetic
BASIC NMR HANDBOOK Written by M. A. Eastman Copyright 1997, 2001, 2013, 2015, 2018 Basic NMR Handbook Table of Contents: Preface ii viii PART 1 Chapter 1: Introduction to NMR 1 Why Study NMR? 1 The Magnetic
mzmatch Excel Template Tutorial
 mzmatch Excel Template Tutorial Installation & Requirements Installation The template may be used to process mzmatch output text files without additional installations or add-ins. Microsoft Excel 2007
mzmatch Excel Template Tutorial Installation & Requirements Installation The template may be used to process mzmatch output text files without additional installations or add-ins. Microsoft Excel 2007
Understanding Your Spectra Module. Agilent OpenLAB CDS ChemStation Edition
 Understanding Your Spectra Module Agilent OpenLAB CDS ChemStation Edition Notices Agilent Technologies, Inc. 1994-2012, 2013 No part of this manual may be reproduced in any form or by any means (including
Understanding Your Spectra Module Agilent OpenLAB CDS ChemStation Edition Notices Agilent Technologies, Inc. 1994-2012, 2013 No part of this manual may be reproduced in any form or by any means (including
 NanoDrop One Viewer software NanoDrop One Website. NanoDrop One Website NanoDrop One Viewer software NanoDrop One Website Software System Update Update Update Software, Update Note OK Language Measure
NanoDrop One Viewer software NanoDrop One Website. NanoDrop One Website NanoDrop One Viewer software NanoDrop One Website Software System Update Update Update Software, Update Note OK Language Measure
NMR Data workup using NUTS
 omework 1 Chem 636, Fall 2008 due at the beginning of the 2 nd week lab (week of Sept 9) NMR Data workup using NUTS This laboratory and homework introduces the basic processing of one dimensional NMR data
omework 1 Chem 636, Fall 2008 due at the beginning of the 2 nd week lab (week of Sept 9) NMR Data workup using NUTS This laboratory and homework introduces the basic processing of one dimensional NMR data
Last updated: Copyright
 Last updated: 2012-08-20 Copyright 2004-2012 plabel (v2.4) User s Manual by Bioinformatics Group, Institute of Computing Technology, Chinese Academy of Sciences Tel: 86-10-62601016 Email: zhangkun01@ict.ac.cn,
Last updated: 2012-08-20 Copyright 2004-2012 plabel (v2.4) User s Manual by Bioinformatics Group, Institute of Computing Technology, Chinese Academy of Sciences Tel: 86-10-62601016 Email: zhangkun01@ict.ac.cn,
The new Water Screening PCDL
 The new Water Screening PCDL Content and integration in suspect and non-target screening Dr. Thomas Glauner Senior LC/MS Applications Scientist EMEA Market Development Team 1 Accurate mass screening and
The new Water Screening PCDL Content and integration in suspect and non-target screening Dr. Thomas Glauner Senior LC/MS Applications Scientist EMEA Market Development Team 1 Accurate mass screening and
Extend Your Metabolomics Insight!
 Extend Your Metabolomics Insight! Introducing MassHunter VistaFlux April 2016 Agilent THE Leader in Metabolomics! Fiehn EI Library METLIN MS/MS Library Mass Profiler Professional with Pathway Architect
Extend Your Metabolomics Insight! Introducing MassHunter VistaFlux April 2016 Agilent THE Leader in Metabolomics! Fiehn EI Library METLIN MS/MS Library Mass Profiler Professional with Pathway Architect
New happenings in gamma spectroscopy data analysis Sylvie Ward Sales Support Specialist
 New happenings in gamma spectroscopy data analysis Sylvie Ward Sales Support Specialist Genie-2000 Latest Features Cascade Summing Correction LACE (Line Activity Consistency Evaluator) Interactive Nuclide
New happenings in gamma spectroscopy data analysis Sylvie Ward Sales Support Specialist Genie-2000 Latest Features Cascade Summing Correction LACE (Line Activity Consistency Evaluator) Interactive Nuclide
Varian Galaxie Chromatography Data System for Preparative HPLC
 Varian Galaxie Chromatography Data System for Preparative HPLC By Gary Burce Varian, Inc. 2700 Mitchell Drive, Walnut Creek, CA 95498 USA Abstract Galaxie is an ideal chromatography data system for the
Varian Galaxie Chromatography Data System for Preparative HPLC By Gary Burce Varian, Inc. 2700 Mitchell Drive, Walnut Creek, CA 95498 USA Abstract Galaxie is an ideal chromatography data system for the
QuantumMCA QuantumNaI QuantumGe QuantumGold
 QuantumMCA QuantumNaI QuantumGe QuantumGold Berkeley Nucleonics Corporation (San Rafael, CA) and Princeton Gamma Tech (Princeton, NJ) have partnered to offer gamma spectroscopy with either germanium or
QuantumMCA QuantumNaI QuantumGe QuantumGold Berkeley Nucleonics Corporation (San Rafael, CA) and Princeton Gamma Tech (Princeton, NJ) have partnered to offer gamma spectroscopy with either germanium or
Handling Human Interpreted Analytical Data. Workflows for Pharmaceutical R&D. Presented by Peter Russell
 Handling Human Interpreted Analytical Data Workflows for Pharmaceutical R&D Presented by Peter Russell 2011 Survey 88% of R&D organizations lack adequate systems to automatically collect data for reporting,
Handling Human Interpreted Analytical Data Workflows for Pharmaceutical R&D Presented by Peter Russell 2011 Survey 88% of R&D organizations lack adequate systems to automatically collect data for reporting,
Jasco V-670 absorption spectrometer
 Laser Spectroscopy Labs Jasco V-670 absorption spectrometer Operation instructions 1. Turn ON the power switch on the right side of the spectrophotometer. It takes about 5 minutes for the light source
Laser Spectroscopy Labs Jasco V-670 absorption spectrometer Operation instructions 1. Turn ON the power switch on the right side of the spectrophotometer. It takes about 5 minutes for the light source
Information Dependent Acquisition (IDA) 1
 Information Dependent Acquisition (IDA) Information Dependent Acquisition (IDA) enables on the fly acquisition of MS/MS spectra during a chromatographic run. Analyst Software IDA is optimized to generate
Information Dependent Acquisition (IDA) Information Dependent Acquisition (IDA) enables on the fly acquisition of MS/MS spectra during a chromatographic run. Analyst Software IDA is optimized to generate
SIERRA ANALYTICS, INC. Version Polymerix Software User Manual
 SIERRA ANALYTICS, INC. Version 3.0.0 Polymerix Software User Manual V E R S I O N 3.0.0 M A R C H 2013 Polymerix Software User Manual Copyright 2010 to 2013 Sierra Analytics, Inc. 5815 Stoddard Road, Suite
SIERRA ANALYTICS, INC. Version 3.0.0 Polymerix Software User Manual V E R S I O N 3.0.0 M A R C H 2013 Polymerix Software User Manual Copyright 2010 to 2013 Sierra Analytics, Inc. 5815 Stoddard Road, Suite
Operation of the Bruker 400 JB Stothers NMR Facility Department of Chemistry Western University
 Operation of the Bruker 400 JB Stothers NMR Facility Department of Chemistry Western University 1. INTRODUCTION...3 1.1. Overview of the Bruker 400 NMR Spectrometer...3 1.2. Overview of Software... 3 1.2.1.
Operation of the Bruker 400 JB Stothers NMR Facility Department of Chemistry Western University 1. INTRODUCTION...3 1.1. Overview of the Bruker 400 NMR Spectrometer...3 1.2. Overview of Software... 3 1.2.1.
An Effective Workflow for Impurity Analysis Incorporating High Quality HRAM LCMS & MSMS with Intelligent Automated Data Mining
 An Effective Workflow for Impurity Analysis Incorporating High Quality HRAM LCMS & MSMS with Intelligent Automated Data Mining Dave Weil, Ph.D. and Jim Lau, Ph.D. Typical Method Conditions: 1260 UHPLC
An Effective Workflow for Impurity Analysis Incorporating High Quality HRAM LCMS & MSMS with Intelligent Automated Data Mining Dave Weil, Ph.D. and Jim Lau, Ph.D. Typical Method Conditions: 1260 UHPLC
Die Nadel im Heuhaufen
 Die Nadel im Heuhaufen Workflow zur Identifizierung unerwarteter Komponenten in LC Q-Tof Daten Umwelt & Lebensmittel Seminar Tour Andreas Reimann Produktspezialist LC-MS Agilent Technologies Instrumentation
Die Nadel im Heuhaufen Workflow zur Identifizierung unerwarteter Komponenten in LC Q-Tof Daten Umwelt & Lebensmittel Seminar Tour Andreas Reimann Produktspezialist LC-MS Agilent Technologies Instrumentation
Non Uniform Sampling in Routine 2D Correlation Experiments
 Analytical and Imaging Solutions for Advanced R&D Non Uniform Sampling in Routine 2D Correlation Experiments NUCLEAR MAGNETIC RESONANCE INTRODUCTION Data obtained from two-dimensional NMR experiments is
Analytical and Imaging Solutions for Advanced R&D Non Uniform Sampling in Routine 2D Correlation Experiments NUCLEAR MAGNETIC RESONANCE INTRODUCTION Data obtained from two-dimensional NMR experiments is
Tutorial. Getting started. Sample to Insight. March 31, 2016
 Getting started March 31, 2016 Sample to Insight CLC bio, a QIAGEN Company Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.clcbio.com support-clcbio@qiagen.com Getting started
Getting started March 31, 2016 Sample to Insight CLC bio, a QIAGEN Company Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.clcbio.com support-clcbio@qiagen.com Getting started
Application Note. Authors. Abstract. Introduction. Environmental
 Using a Chlorine Filter for Accurate-Mass Data Analysis of Environmental Samples Application Note Environmental Authors Imma Ferrer and E. Michael Thurman Center for Environmental Mass Spectrometry University
Using a Chlorine Filter for Accurate-Mass Data Analysis of Environmental Samples Application Note Environmental Authors Imma Ferrer and E. Michael Thurman Center for Environmental Mass Spectrometry University
SPEED AND ACCURACY TO DRUG DEVELOPMENT
 HOW qnmr BRINGS SPEED ND CCURCY TO DRUG DEVELOPMENT INTRODUCTION The total cost of bringing a drug to market is now estimated at about $2.5 billion 1, and the pressure to advance new drugs faster is greater
HOW qnmr BRINGS SPEED ND CCURCY TO DRUG DEVELOPMENT INTRODUCTION The total cost of bringing a drug to market is now estimated at about $2.5 billion 1, and the pressure to advance new drugs faster is greater
Overview. Introduction. André Schreiber AB SCIEX Concord, Ontario (Canada)
 Quantitation and Identification of Pharmaceuticals and Personal Care Products (PPCP) in Environmental Samples using Advanced TripleTOF MS/MS Technology André Schreiber AB SCIEX Concord, Ontario (Canada)
Quantitation and Identification of Pharmaceuticals and Personal Care Products (PPCP) in Environmental Samples using Advanced TripleTOF MS/MS Technology André Schreiber AB SCIEX Concord, Ontario (Canada)
OECD QSAR Toolbox v.4.1. Tutorial on how to predict Skin sensitization potential taking into account alert performance
 OECD QSAR Toolbox v.4.1 Tutorial on how to predict Skin sensitization potential taking into account alert performance Outlook Background Objectives Specific Aims Read across and analogue approach The exercise
OECD QSAR Toolbox v.4.1 Tutorial on how to predict Skin sensitization potential taking into account alert performance Outlook Background Objectives Specific Aims Read across and analogue approach The exercise
Rapid and Accurate Forensics Analysis using High Resolution All Ions MS/MS
 Rapid and Accurate Forensics Analysis using High Resolution All Ions MS/MS Application Note Forensic Toxicology Authors Martin Josefsson, and Markus Roman National Board of Forensic Medicine Linköping,
Rapid and Accurate Forensics Analysis using High Resolution All Ions MS/MS Application Note Forensic Toxicology Authors Martin Josefsson, and Markus Roman National Board of Forensic Medicine Linköping,
Login -the operator screen should be in view when you first sit down at the spectrometer console:
 Lab #2 1D 1 H Double Resonance (Selective Decoupling) operation of the 400 MHz instrument using automated sample insertion (robot) and automated locking and shimming collection of 1D 1 H spectra retrieving
Lab #2 1D 1 H Double Resonance (Selective Decoupling) operation of the 400 MHz instrument using automated sample insertion (robot) and automated locking and shimming collection of 1D 1 H spectra retrieving
OECD QSAR Toolbox v.4.0. Tutorial on how to predict Skin sensitization potential taking into account alert performance
 OECD QSAR Toolbox v.4.0 Tutorial on how to predict Skin sensitization potential taking into account alert performance Outlook Background Objectives Specific Aims Read across and analogue approach The exercise
OECD QSAR Toolbox v.4.0 Tutorial on how to predict Skin sensitization potential taking into account alert performance Outlook Background Objectives Specific Aims Read across and analogue approach The exercise
Chemistry Department
 Chemistry Department NMR/Instrumentation Facility Users Guide - VNMRJ Prepared by Leila Maurmann The following procedures should be used to acquire one-dimensional proton and carbon NMR data on the 400MHz
Chemistry Department NMR/Instrumentation Facility Users Guide - VNMRJ Prepared by Leila Maurmann The following procedures should be used to acquire one-dimensional proton and carbon NMR data on the 400MHz
Produktivitätswerkzeuge für die NMR
 Produktivitätswerkzeuge für die NMR Till Kühn VP Applications Development Benutzertagung Karlsruhe November 2016 Innovation with Integrity A week in the life of Brian Brian Works in a hypothetical pharma
Produktivitätswerkzeuge für die NMR Till Kühn VP Applications Development Benutzertagung Karlsruhe November 2016 Innovation with Integrity A week in the life of Brian Brian Works in a hypothetical pharma
Agilent 6400 Series Triple Quadrupole LC/MS/MS Users Session
 Agilent 6400 Series Triple Quadrupole LC/MS/MS Users Session QQQ Method Development and Optimization MassHunter Quant: Method setup Peak detection optimization Quant troubleshooting David Presser Application
Agilent 6400 Series Triple Quadrupole LC/MS/MS Users Session QQQ Method Development and Optimization MassHunter Quant: Method setup Peak detection optimization Quant troubleshooting David Presser Application
Pulsar. Delivering NMR to your benchtop
 Pulsar NMR Delivering NMR to your benchtop Pulsar TM NMR for your laboratory The Pulsar TM NMR spectrometer from Oxford Instruments delivers affordable, high performance NMR spectroscopy into the laboratory
Pulsar NMR Delivering NMR to your benchtop Pulsar TM NMR for your laboratory The Pulsar TM NMR spectrometer from Oxford Instruments delivers affordable, high performance NMR spectroscopy into the laboratory
The Theory of HPLC. Quantitative and Qualitative HPLC
 The Theory of HPLC Quantitative and Qualitative HPLC i Wherever you see this symbol, it is important to access the on-line course as there is interactive material that cannot be fully shown in this reference
The Theory of HPLC Quantitative and Qualitative HPLC i Wherever you see this symbol, it is important to access the on-line course as there is interactive material that cannot be fully shown in this reference
for XPS surface analysis
 Thermo Scientific Avantage XPS Software Powerful instrument operation and data processing for XPS surface analysis Avantage Software Atomic Concentration (%) 100 The premier software for surface analysis
Thermo Scientific Avantage XPS Software Powerful instrument operation and data processing for XPS surface analysis Avantage Software Atomic Concentration (%) 100 The premier software for surface analysis
GAS CHROMATOGRAPHY MASS SPECTROMETRY. Pre-Lab Questions
 GAS CHROMATOGRAPHY MASS SPECTROMETRY Pre-Lab Questions Questions to be answered before doing the experiment. The answers are due at the beginning of each experiment without exception (the questions are
GAS CHROMATOGRAPHY MASS SPECTROMETRY Pre-Lab Questions Questions to be answered before doing the experiment. The answers are due at the beginning of each experiment without exception (the questions are
BIOLIGHT STUDIO IN ROUTINE UV/VIS SPECTROSCOPY
 BIOLIGHT STUDIO IN ROUTINE UV/VIS SPECTROSCOPY UV/Vis Spectroscopy is a technique that is widely used to characterize, identify and quantify chemical compounds in all fields of analytical chemistry. The
BIOLIGHT STUDIO IN ROUTINE UV/VIS SPECTROSCOPY UV/Vis Spectroscopy is a technique that is widely used to characterize, identify and quantify chemical compounds in all fields of analytical chemistry. The
HMQC HSQC and HMBC. Gradient HMQC, HMBC on the Bruker400 and 500
 1 Gradient HMQC, HMBC on the Bruker400 and 500 HMQC, HSQC - Heteronuclear Multiple Quantum Correlation. These experiments correlate the chemical shift of proton with the chemical shift of the directly
1 Gradient HMQC, HMBC on the Bruker400 and 500 HMQC, HSQC - Heteronuclear Multiple Quantum Correlation. These experiments correlate the chemical shift of proton with the chemical shift of the directly
How to Create a Substance Answer Set
 How to Create a Substance Answer Set Select among five search techniques to find substances Since substances can be described by multiple names or other characteristics, SciFinder gives you the flexibility
How to Create a Substance Answer Set Select among five search techniques to find substances Since substances can be described by multiple names or other characteristics, SciFinder gives you the flexibility
NEW TOOLS FOR FINDING AND IDENTIFYING METABOLITES IN A METABOLOMICS WORKFLOW
 NEW TOOLS FOR FINDING AND IDENTIFYING METABOLITES IN A METABOLOMICS WORKFLOW Julia E. Wingate 1 ; Elliott Jones 2 ; Armin Graber 3 ; Klaus Weinberger 3 1Applied Biosystems, Toronto, Canada; 2Applied Biosystems,
NEW TOOLS FOR FINDING AND IDENTIFYING METABOLITES IN A METABOLOMICS WORKFLOW Julia E. Wingate 1 ; Elliott Jones 2 ; Armin Graber 3 ; Klaus Weinberger 3 1Applied Biosystems, Toronto, Canada; 2Applied Biosystems,
Experience the most powerful benchtop NMR spectrometer
 Experience the most powerful benchtop NMR spectrometer Outstanding Features Highest field strength (80 MHz) Largest chemical shift spread Highest sensitivity Superb Resolution (0.5 Hz/20 Hz) Multi-nuclear
Experience the most powerful benchtop NMR spectrometer Outstanding Features Highest field strength (80 MHz) Largest chemical shift spread Highest sensitivity Superb Resolution (0.5 Hz/20 Hz) Multi-nuclear
Quantitation of High Resolution MS Data Using UNIFI: Acquiring and Processing Full Scan or Tof-MRM (Targeted HRMS) Datasets for Quantitative Assays
 : Acquiring and Processing Full Scan or Tof-MRM (Targeted HRMS) Datasets for Quantitative Assays Mark Wrona, Jayne Kirk, and Yun Alelyunas Waters Corporation, Milford, MA, USA APPLICATION BENEFITS Ability
: Acquiring and Processing Full Scan or Tof-MRM (Targeted HRMS) Datasets for Quantitative Assays Mark Wrona, Jayne Kirk, and Yun Alelyunas Waters Corporation, Milford, MA, USA APPLICATION BENEFITS Ability
This tutorial is intended to familiarize you with the Geomatica Toolbar and describe the basics of viewing data using Geomatica Focus.
 PCI GEOMATICS GEOMATICA QUICKSTART 1. Introduction This tutorial is intended to familiarize you with the Geomatica Toolbar and describe the basics of viewing data using Geomatica Focus. All data used in
PCI GEOMATICS GEOMATICA QUICKSTART 1. Introduction This tutorial is intended to familiarize you with the Geomatica Toolbar and describe the basics of viewing data using Geomatica Focus. All data used in
Multi-residue analysis of pesticides by GC-HRMS
 An Executive Summary Multi-residue analysis of pesticides by GC-HRMS Dr. Hans Mol is senior scientist at RIKILT- Wageningen UR Introduction Regulatory authorities throughout the world set and enforce strict
An Executive Summary Multi-residue analysis of pesticides by GC-HRMS Dr. Hans Mol is senior scientist at RIKILT- Wageningen UR Introduction Regulatory authorities throughout the world set and enforce strict
Identification and Characterization of an Isolated Impurity Fraction: Analysis of an Unknown Degradant Found in Quetiapine Fumarate
 Identification and Characterization of an Isolated Impurity Fraction: Analysis of an Unknown Degradant Found in Quetiapine Fumarate Michael D. Jones, Xiang Jin Song, Robert S. Plumb, Peter J. Lee, and
Identification and Characterization of an Isolated Impurity Fraction: Analysis of an Unknown Degradant Found in Quetiapine Fumarate Michael D. Jones, Xiang Jin Song, Robert S. Plumb, Peter J. Lee, and
Application Note LCMS-116 What are we eating? MetaboScape Software; Enabling the De-replication and Identification of Unknowns in Food Metabolomics
 Application Note LCMS-116 What are we eating? MetaboScape Software; Enabling the De-replication and Identification of Unknowns in Food Metabolomics Introduction Determining the structure of secondary metabolites
Application Note LCMS-116 What are we eating? MetaboScape Software; Enabling the De-replication and Identification of Unknowns in Food Metabolomics Introduction Determining the structure of secondary metabolites
What s New in NIST11 (April 3, 2011)
 What s New in NIST11 (April 3, 2011) NIST11 consists of the 2011 release of the NIST/EPA/NIH Electron Ionization (EI) Mass Spectral Database, the NIST MS/MS Database, and the NIST GC Methods and Retention
What s New in NIST11 (April 3, 2011) NIST11 consists of the 2011 release of the NIST/EPA/NIH Electron Ionization (EI) Mass Spectral Database, the NIST MS/MS Database, and the NIST GC Methods and Retention
Analytical Roche Benefits and Hurdles of ACD Software. Josef Schneider ACD/Labs Symposium on Laboratory Intelligence Jun 19-20, 2013
 Analytical Chemistry @ Roche Benefits and Hurdles of ACD Software Josef Schneider ACD/Labs Symposium on Laboratory Intelligence Jun 19-20, 2013 Drug Discovery "Making discoveries that matter" MedChem,
Analytical Chemistry @ Roche Benefits and Hurdles of ACD Software Josef Schneider ACD/Labs Symposium on Laboratory Intelligence Jun 19-20, 2013 Drug Discovery "Making discoveries that matter" MedChem,
Become a Microprobe Power User Part 2: Qualitative & Quantitative Analysis
 Become a Microprobe Power User Part 2: Qualitative & Quantitative Analysis Mike Spilde Spring IOM Seminar February 5, 2008 Qualitative Analysis Why use qualitative scans? Elemental ID (especially trace
Become a Microprobe Power User Part 2: Qualitative & Quantitative Analysis Mike Spilde Spring IOM Seminar February 5, 2008 Qualitative Analysis Why use qualitative scans? Elemental ID (especially trace
OECD QSAR Toolbox v.3.4. Step-by-step example of how to build and evaluate a category based on mechanism of action with protein and DNA binding
 OECD QSAR Toolbox v.3.4 Step-by-step example of how to build and evaluate a category based on mechanism of action with protein and DNA binding Outlook Background Objectives Specific Aims The exercise Workflow
OECD QSAR Toolbox v.3.4 Step-by-step example of how to build and evaluate a category based on mechanism of action with protein and DNA binding Outlook Background Objectives Specific Aims The exercise Workflow
Fragment-based drug discovery
 Fragment-based drug discovery Dr. Till Kühn VP Applications Development MRS, Bruker BioSpion User s meeting, Brussels, November 2016 Innovation with Integrity The principle of Fragment Based Screening
Fragment-based drug discovery Dr. Till Kühn VP Applications Development MRS, Bruker BioSpion User s meeting, Brussels, November 2016 Innovation with Integrity The principle of Fragment Based Screening
OECD QSAR Toolbox v.3.3. Step-by-step example of how to build and evaluate a category based on mechanism of action with protein and DNA binding
 OECD QSAR Toolbox v.3.3 Step-by-step example of how to build and evaluate a category based on mechanism of action with protein and DNA binding Outlook Background Objectives Specific Aims The exercise Workflow
OECD QSAR Toolbox v.3.3 Step-by-step example of how to build and evaluate a category based on mechanism of action with protein and DNA binding Outlook Background Objectives Specific Aims The exercise Workflow
Natural Products. Innovation with Integrity. High Performance NMR Solutions for Analysis NMR
 Natural Products High Performance NMR Solutions for Analysis Innovation with Integrity NMR NMR Spectroscopy Continuous advancement in Bruker s NMR technology allows researchers to push the boundaries for
Natural Products High Performance NMR Solutions for Analysis Innovation with Integrity NMR NMR Spectroscopy Continuous advancement in Bruker s NMR technology allows researchers to push the boundaries for
Building Inflation Tables and CER Libraries
 Building Inflation Tables and CER Libraries January 2007 Presented by James K. Johnson Tecolote Research, Inc. Copyright Tecolote Research, Inc. September 2006 Abstract Building Inflation Tables and CER
Building Inflation Tables and CER Libraries January 2007 Presented by James K. Johnson Tecolote Research, Inc. Copyright Tecolote Research, Inc. September 2006 Abstract Building Inflation Tables and CER
Application Note LCMS-112 A Fully Automated Two-Step Procedure for Quality Control of Synthetic Peptides
 Application Note LCMS-112 A Fully Automated Two-Step Procedure for Quality Control of Synthetic Peptides Abstract Here we describe a two-step QC procedure for synthetic peptides. In the first step, the
Application Note LCMS-112 A Fully Automated Two-Step Procedure for Quality Control of Synthetic Peptides Abstract Here we describe a two-step QC procedure for synthetic peptides. In the first step, the
Achieve confident synthesis control with the Thermo Scientific ISQ EC single quadrupole mass spectrometer
 APPLICATION NOTE 72385 Achieve confident synthesis control with the Thermo Scientific ISQ EC single quadrupole mass spectrometer Authors Stephan Meding, Katherine Lovejoy, Martin Ruehl Thermo Fisher Scientific,
APPLICATION NOTE 72385 Achieve confident synthesis control with the Thermo Scientific ISQ EC single quadrupole mass spectrometer Authors Stephan Meding, Katherine Lovejoy, Martin Ruehl Thermo Fisher Scientific,
Structural Analysis by In-Depth Impurity Search Using MetID Solution and High Accuracy MS/MS
 C146-E118 Structural Analysis by In-Depth Impurity Search Using MetID Solution and High Accuracy MS/MS Technical Report vol.16 1. Introduction MetID Solution is a software application that was developed
C146-E118 Structural Analysis by In-Depth Impurity Search Using MetID Solution and High Accuracy MS/MS Technical Report vol.16 1. Introduction MetID Solution is a software application that was developed
Band-Selective Homonuclear 2D Correlation Experiments
 Band-Selective Homonuclear 2D Correlation Experiments Application Note Authors Péter Sándor Agilent Technologies GmbH D76337 Waldbronn Germany Abstract This application note demonstrates the utility of
Band-Selective Homonuclear 2D Correlation Experiments Application Note Authors Péter Sándor Agilent Technologies GmbH D76337 Waldbronn Germany Abstract This application note demonstrates the utility of
Tutorial 1: Setting up your Skyline document
 Tutorial 1: Setting up your Skyline document Caution! For using Skyline the number formats of your computer have to be set to English (United States). Open the Control Panel Clock, Language, and Region
Tutorial 1: Setting up your Skyline document Caution! For using Skyline the number formats of your computer have to be set to English (United States). Open the Control Panel Clock, Language, and Region
OECD QSAR Toolbox v.3.2. Step-by-step example of how to build and evaluate a category based on mechanism of action with protein and DNA binding
 OECD QSAR Toolbox v.3.2 Step-by-step example of how to build and evaluate a category based on mechanism of action with protein and DNA binding Outlook Background Objectives Specific Aims The exercise Workflow
OECD QSAR Toolbox v.3.2 Step-by-step example of how to build and evaluate a category based on mechanism of action with protein and DNA binding Outlook Background Objectives Specific Aims The exercise Workflow
Standards-Based Quantification in DTSA-II Part II
 Standards-Based Quantification in DTSA-II Part II Nicholas W.M. Ritchie National Institute of Standards and Technology, Gaithersburg, MD 20899-8371 nicholas.ritchie@nist.gov Introduction This article is
Standards-Based Quantification in DTSA-II Part II Nicholas W.M. Ritchie National Institute of Standards and Technology, Gaithersburg, MD 20899-8371 nicholas.ritchie@nist.gov Introduction This article is
OECD QSAR Toolbox v.3.3. Step-by-step example of how to build a userdefined
 OECD QSAR Toolbox v.3.3 Step-by-step example of how to build a userdefined QSAR Background Objectives The exercise Workflow of the exercise Outlook 2 Background This is a step-by-step presentation designed
OECD QSAR Toolbox v.3.3 Step-by-step example of how to build a userdefined QSAR Background Objectives The exercise Workflow of the exercise Outlook 2 Background This is a step-by-step presentation designed
