mvmorph:an R package for fitting multivariate evolutionary models to morphometric data
|
|
|
- Naomi McCormick
- 5 years ago
- Views:
Transcription
1 Methods in Ecology and Evolution 215, 6, doi: /241-21X.1242 APPLICATION mvmorph:an R package for fitting multivariate evolutionary models to morphometric data Julien Clavel 1,2 *, Gilles Escarguel 2 and Gildas Merceron 3 1 Ecole Normale Superieure, IBENS UMR 8197, CNRS, 46 rue d Ulm, 755 Paris, France; 2 Laboratoire de Geologie de Lyon, UMR 5276, CNRS, UCB Lyon 1, ENS Lyon, Campus de la Doua, 2 rue Rapha el Dubois, Villeurbanne Cedex, France; and 3 IPHEP, UMR 7262, CNRS, Universite de Poitiers, Bat. B35 TSA , rue M. Brunet, 8673 Poitiers Cedex 9, France Summary 1. We present mvmorph, a package of multivariate phylogenetic comparative methods for the R statistical environment. mvmorph is freely available on the CRAN package repository ( mvmorph/). 2. mvmorph allows fitting a range of multivariate evolutionary models under a maximum-likelihood criterion. Initially developed in the context of phylogenetic analysis of multiple morphometric traits, its use can be extended to any biological data set with one or multiple covarying continuous traits. All the fitting models include the possibility to use SIMMAP-like mapping,which may be useful for fitting changes along lineages at a given point in time. All models provide diagnostic metrics for convergence and reliability of estimates, as well as the possibility to include trait measurement errors in model estimates. 3. New features provided by the mvmorph package include the possibility of fitting models with changes in the mode of evolution along the phylogeny, which will be particularly meaningful in comparative analyses that include extinct taxa, for example when testing changes in evolutionary mode associated with global biotic/abiotic events. 4. We briefly describe the models already included in mvmorph and provide some demonstration of the use of the package with two simulated worked examples. Key-words: Brownian motion, Early Burst, multivariate comparative methods, Ornstein Uhlenbeck, stochastic mapping Introduction Comparative methods for fitting evolutionary models to continuous data are becoming increasingly popular (e.g. Hansen 1997; Pagel 1999; Butler & King 24; O Meara et al. 26; Thomas, Freckleton & Szekely 26; Hansen, Pienaar & Orzack 28; Beaulieu et al. 212; Thomas & Freckleton 212; Adams 213; Ingram & Mahler 213). Over the last years, these methods used to study evolutionary patterns of continuous traits within phylogenies have been widely applied on a trait-by-trait basis for estimating shifts in evolutionary rates associated with environmental global changes, the timing of acquisition of key innovations, or the release of biotic interactions (Collar et al. 29; Price et al. 211, 213; Slater 213), as well as changes in modes of evolution and adaptation towards different optima (Scales, King & Butler 29; Collar, Schulte & Losos 211; Frederich et al. 213; Knope & Scales 213; Mahler et al. 213). However, phenomic evolution is *Correspondence author. s: clavel@biologie.ens.fr, julien.clavel@hotmail.fr multivariate by nature, as natural selection operates on functionally related organism s traits (Walsh 27; Walsh & Blows 29), and because traits may covary due to pleiotropic effects, genetic linkages or different sources of developmental constraints (Lande & Arnold 1983; Felsenstein 1988; Armbruster & Schwaegerle 1996; Armbruster et al. 214). Thus, multivariate analyses should logically be favoured over univariate ones (Fig. 1) for describing the complex structure of multiple covarying traits and measurements as routinely studied in morphometrics. Moreover, by considering the evolutionary history of phylogenetically related species, we can further assess how and when phenotypic covariations evolved, either in relation or not with biotic/abiotic events that potentially drive evolution (Fig. 2). Here, we present a new R package, mvmorph, dedicated to the phylogenetic comparative analysis of biological data sets with multiple covarying continuous traits. Phenotypic multivariate evolution has been thoroughly studied over the last decades, particularly in quantitative genetics in relation to the evolution of genetic covariances (Lande 1979; Lande & Arnold 1983; Cheverud 1984; Felsenstein 1988; 215 The Authors. Methods in Ecology and Evolution 215 British Ecological Society
2 1312 J. Clavel, G. Escarguel, & G. Merceron (a1) (b1) (c1) (d) (a2) (b2) (c2) (e) Fig. 1. Different aspects of trait variances and covariances (as recorded in the r² matrices) described by multivariate analysis as the result of distinct selective, functional and/or genetic constraints. Bivariate cases are illustrated here for the sake of graphical simplicity, but same reasoning applies to any multivariate case. In (a1) and (a2), the correlations between traits 1 and 2 are similar (same covariance/variances ratio), but trait variances differ. Conversely, in (b1) and (b2), the variances are identical, but the correlations between traits 1 and 2 differ (higher in b2 than in b1). Finally, in (c1) and (c2) trait relationships differ in both trait variances and correlations. Contrasting with these multivariate cases, univariate analysis where only differences in trait variances can be assessed and interactions (i.e. covariances) between traits are simply ignored is illustrated in (d). Last, traits mayalso differ in their means rather than in their variance covariance structures. When trait means differ along axes of low variance as in (e), trait-by-trait (univariate) analyses often fail at identifying differences between taxa (blue and red ellipses) due to lack of statistical power(see example 1 in Appendix S2). Zeng 1988; Endler 1995; Arnold, Pfrender & Jones 21; Steppan, Phillips & Houle 22) and has led to particular expectations over macroevolutionary times (e.g. Schluter 1996; Steppan 1997; Arnold, Pfrender & Jones 21; Baker & Wilkinson 23; Polly 24; Hohenlohe & Arnold 28; Revell & Harmon 28; Shoval et al. 212; Goswami et al. 214). Importantly, modelling multivariate phenotypic evolution through trait variances and pairwise covariances allows describing several aspects (e.g. correlated evolution, correlated selection) that separate univariate analyses cannot provide (Fig. 2). Indeed, univariate analyses are merely limited for describing complex structures and generally show less statistical power than multivariate counterparts (Zheng et al. 29). Moreover, while complex multivariate data sets are often first reduced through ordination methods such as PCA and a subset of the first principal components is then treated as independent univariate traits, evolutionary model inferences from these transformed spaces can be severely biased (Revell 29; Monteiro 213; Polly et al. 213; Uyeda, Caetano & Pennell 215). Finally, it is a rather common approach in comparative analysis to regressing traits on each other by assuming, often arbitrarily, that one trait is the dependent variable and the other the independent variable. Nevertheless, such an approach implicitly assumes that one trait is evolutionarily responding to the other (Hansen, Pienaar & Orzack 28; Revell 21; Hansen & Bartoszek 212), a critical assumption of biological causality which is actually relaxed in a multivariate setting focused on mere correlations among response variables. The multivariate models provided by the mvmorph package will be also of particular interest for evolutionary biologists integrating fossil specimens and taxa in their studies, as the package allows estimating models of changes in evolutionary modes and/or rates at a given point in time (e.g. Slater 213). Indeed, integration of extinct clades in comparative studies is an important step forward in the study of morphological diversification (Slater, Harmon & Alfaro 212; Slater 213): while some studies have pointed towards the need to consider phylogenies in among-group comparisons of morphological rates of evolution (O Meara et al. 26; Ricklefs 26; Adams et al. 29; Revell & Collar 29), it appears also necessary to consider underlying processes which may affect the realized variances (Hunt 212). Actually, some processes such as those leading to shifts from the ancestral state to the optima can be estimated only when including fossil taxa in the analysis (Hansen 1997; Hunt, Bell & Travis 28). Moreover, although
3 mvmorph 1313 (a) (b) Fig. 2. Examples of two phenotypic traits evolving under a bivariate Brownian motion (BM) process within a 1 species tree [inset in (a)]. In (a), a positive (s12 = 1 6) evolutionary covariance is accounted for by the BM process, making phenotypic changes in traits 1 and 2 correlated throughout the species evolutionary history (i.e. interspecific divergences in phenotype are more likely to occur along the correlation axis between traits 1 and 2). Conversely, in (b), traits evolve independently in the bivariate space (i.e. there is a null evolutionary covariance between traits 1 and 2). Importantly, the covariations between traits of phylogenetically related species may vary through time (e.g. when species are diversifying functionally during an adaptive radiation) or among different clades (e.g. release of competitive pressures) and may lead to particular expectations about changes in evolutionary correlations under various macroevolutionary scenarios that can be explicitly tested with mvmorph by using various models and mapped histories (Table 1). the use of non-ultrametric trees (whether including fossil taxa or fast evolving organisms such as viruses) enhances the accuracy and identifiability of parameter estimates (Slater, Harmon & Alfaro 212; Ho & An e 213), it remains computationally limited to some of the current software implementations (Slater 214). Description THE mvmorph PACKAGE The mvmorph package provides in a unified environment useful tools to deal with ultrametric or non-ultrametric trees over
4 1314 J. Clavel, G. Escarguel, & G. Merceron Table 1. Summary of evolutionary models and main functions currently available in the mvmorph package (version 1..5) Function Model Number of parameters* Short description of the associated evolutionary model mvbm BM1 (p(p + 1)/2) + p BM on the whole tree (unique rate matrix) BMM (p(p + 1)/2)k + p BM with multiple rates matrix (multiple selective regimes) mvou OU1 2(p(p + 1)/2) + p OU process with a unique adaptive optimum per trait OUM 2(p(p + 1)/2) + kp OU process with multiple adaptive optima per trait mveb EB ACDC (p(p + 1)/2) + p + 1 EB model or decelerating model of evolutionary rates. mvshift BMEB/EBBM (p(p + 1)/2) + p + 1 Shift of a BM to/from EB process at a given point in time BMEBi/EBBMi 2(p(p + 1)/2) + p + 1 Same as BMEB/EBBM with independent rates on each time slice BMOU/OUBM 2(p(p + 1)/2) + p Shift of a BM to/from OU process at a given point in time BMOUi/OUBMi 3(p(p + 1)/2) + p Same as BMOU/OUBM with independent rates on each time slice EBOU/OUEB 2(p(p + 1)/2) + p + 1 Shift of a EB to/from OU process at a given point in time EBOUi/OUEBi 3(p(p + 1)/2) + p + 1 Same as EBOU/OUEB with independent rates on each time slice mvsim all Simulate the evolution of traits under the models represented in mvmorph mvll Compute the log-likelihood of an user provided tree or variance covariance matrix half-life OU Compute the phylogenetic half-life for a multivariate OU process stationary OU Compute the multivariate stationary distribution for a multivariate OU process LRT Compute the Log-ratio test for nested models in mvmorph p, number of traits; k, number of distinct selective regimes. Model abbreviations: BM, Brownian motion; OU, Ornstein Uhlenbeck; EB, Early Burst; ACDC, exponentially accelerating or decelerating. *The number of parameters can be actually lower, depending on matrix constraints used. a range of models of continuous trait evolution (Table 1). mvmorph provides fast (see drawback below) multivariate implementations of comparative approaches allowing the incorporation of measurement errors (Ives, Midford & Garland 27; Felsenstein 28; Revell & Reynolds 212; Silvestro et al. 215), missing values and various parameterizations; it allows the user to handle trees in the SIMMAP-like format (Huelsenbeck, Nielsen & Bollback 23; Bollback 26) which provide a flexible way to map evolutionary hypothesis on a phylogeny thanks to existing functions from the phytools package (Revell 212). Last, mvmorph also allows simulating the multivariate evolution of traits (mvsim function) and provides tools for users and developers with a general wrapper mvll for computing log-likelihood of userspecified and customized models of trait evolution with various fast methods (see Appendix S1 for details, Supporting information). MULTIVARIATE MODELS IMPLEMENTED IN mvmorph The multivariate models for continuous trait evolution implemented in mvmorph are summarized in Table 1. Most of these models have been already widely described in an univariate context (Hansen 1997; Blomberg, Garland & Ives 23; Butler & King 24; O Meara et al. 26; Thomas, Freckleton & Szekely 26; Harmon et al. 21; Slater 213). Below we give a short presentation of their multivariate extension as provided in the package (see Appendix S1 for detailed descriptions). LOG-LIKELIHOOD OF MULTIVARIATE MODELS Most phylogenetic comparative analyses are done by fitting a generalized least square (GLS) model of the form: Y ¼ bx þ e eqn 1 For a multivariate model with m traits and n species, Y is a vector of length mn and e is a vector of mn residuals normally distributed in Nð; VÞ,withV a mn 9 mn phylogenetic multivariate variance covariance matrix. If we fit an intercept-only model (i.e. a correlational model where we are estimating the phylogenetic signal in Y), then X is a mn 9 m design matrix, and b the (notional) ancestral state expectation is a vector of parameters to estimate of length m (assuming a unique ancestral state for each trait). As for the univariate case, the fit of multivariate models of continuous trait evolution is assessed by maximization of the log-likelihood eqn 2 (e.g. Martins & Hansen 1997; Blomberg, Garland & Ives 23; O Meara et al. 26). logðlþ ¼ 1 h i mnlog 2p 2 ð Þþlog j V jþ ð Y bx ÞT V 1 ðy bxþ eqn 2 Evolutionary parameters for the various models of phenotypic evolution [Brownian motion (BM), Ornstein Uhlenbeck (OU), etc.] enter this equation through the phylogenetic variance covariance matrix V, while b is given by the GLS solution (eqn 3).
5 mvmorph 1315 b ¼ðX T V 1 XÞ 1 X T V 1 Y eqn 3 Brownian motion The BM model is the most widespread evolutionary model used in comparative methods (e.g. Felsenstein 1985). Its multivariate extension is straightforward and has been already thoroughly described (Felsenstein 24; Revell & Harmon 28; Revell & Collar 29; Freckleton 212). The variance covariance matrix related to a multivariate BM process (eqn 4) is the Kronecker product of the phylogenetic variance covariance matrix C (for which each element C ij is the height above the root for the common ancestor of species i and j; seerohlf 21; Revell & Collar 29) with the maximum-likelihood estimate of the evolutionary rate matrix R (Revell & Harmon 28; Revell & Collar 29; see also Appendix S1). Cov Y ik ; Y jl ¼ Cij R kl eqn 4 Ornstein Uhlenbeck The OU model is an appealing alternative to the BM model as it describes processes where the traits variance is constrained around one or several optima often referred as selective regime or adaptive zone optima at the macroevolutionary scale (Hansen 1997, 212; Butler & King 24). In comparative method studies, this process has often been misleadingly interpreted in terms of Lande s (1976) model of population undergoing stabilizing selection in a static landscape (see also discussion in Harmon et al. 21 and Pennell & Harmon 213). The OU model for a multivariate trait matrix Y along a lineage is described by the differential equation: dyðþ¼ t Aðb Yt ðþþdtþrdwðþ t eqn 5 where A and R are square matrices of the strength of selection and the random drift, respectively, b is a vector of mean (optimum) values, and W is the diffusion parameter (Butler & King 29; Bartoszek et al. 212; Monteiro 213). The eigenvalues and eigenvectors of the matrix A describe the path towards the optimum, while the matrix R describes the covariations of the random perturbations. The variance covariance matrix computed between the m traits at tips i and j canbeobtainedintwo different ways, depending on whether the value at the root is estimated or assumed at the stationary point (see Ho & Ane 213 for the univariate counterpart). Particularly, when A is symmetric positive definite (i.e. A has only real positive eigenvalues), a stationary solution exists (Gardiner 24) for which the variance covariance matrix can be computed as follows: Cov Y i ; Y j ¼ Z Cij A Cii t e ð Þ Re AT ðc jj tþ dt eqn 6 This is the implementation found in the R package OUCH (Butler & King 29). Alternatively, if the covariance matrix is conditioned on the root, a more general form which also allows non-symmetric matrices A may be used (Eq. B.3 in Bartoszek et al. 212; see also Reitan, Schweder & Henderiks 212 and Appendix S1 for details): 1 Cov Y i ; Y j ¼ e AðC ii C ij Z Cij e At Re ATt C dt ð Ae AT C jj C ij Þ eqn 7 This second approach is particularly useful for fitting multivariate models to non-ultrametric trees (e.g. including fossil species) for which the estimation of the root state is feasible. On ultrametric trees, both approaches should converge to the same estimates if the process is stationary. As for the univariate case, the expectation of the OU process (eqn 8) can be computed as the weighted average of the optima and the common ancestral state at the root of the tree (Hansen 1997). E½Y i ðþ t Š ¼ e At E½Y i ðþšþ Z t e Aðt tþ Ab i ðtþdt eqn 8 Thisexpectationisusedtoestimatetherootandtheoptima by entering the coefficients of their relative contribution in the design matrix X of the GLS equation (eqn 3). The various parameterizations implemented for the root state are described and discussed in the Appendix S1. Early Burst The mvmorph package allows computing a joint estimate for a decrease or increase in evolutionary rates in a multivariate data set, following Blomberg, Garland & Ives (23) exponentially accelerating or decelerating (ACDC) model or Harmon et al. s (21) Early Burst (EB) model. These models have been used for studying adaptive radiations where evolutionary theory predicts a burst of evolutionary rates and an increase in morphological disparity, followed by a decrease in rates of morphological evolution as ecological niches are filled (Cooper & Purvis 21; Harmon et al. 21; Slater et al. 21; Ingram, Harmon & Shurin 212; Slater 213). Here, the phylogenetic matrix C is transformed by the exponential transformation (g parameter) of the ACDC model (Blomberg, Garland & Ives 23), which is jointly estimated during likelihood maximization of the evolutionary rate matrix (eqn 9). e gcij 1 Cov Y ik ; Y jl ¼ Rkl g Shifts models eqn 9 Slater (213) recently introduced a heterogeneous model allowing for the shift from one evolutionary process to another in the same tree. Dealing with heterogeneous models was also
6 1316 J. Clavel, G. Escarguel, & G. Merceron previously proposed by using tree-stretching approaches (O Meara 212; but see Slater 214). Such approaches are particularly appealing when mixing fossil and extant data to assess changes in evolutionary modes in addition to temporal changes in evolutionary tempo (Hunt, Bell & Travis 28; Slater & Harmon 213; see Appendix S1). A combination of the various models implemented in mvmorph canbefittedtotwo successive time slices or mapped era using the mvshift function (Table 1). For instance, the variance covariance for the ecological release model proposed by Slater (213), combining an OU process then followed by a BM process, is computed as follows: 1 Y i ; Y j ¼ A Couii Couij Cbmij R þ e ð Þ Z Couij e At Re ATt C dta ð e AT Cou jj Cou ij Þ eqn 1 where Cbm ij represents the history between the taxa i and j shared during the time period where the BM process took place; Cou ij represents the shared history between both taxa accrued during the OU process; and Cou jj and Cou ii represent the evolutionary histories (variances) accrued in each lineage during the OU process. Constraining the parameter spaces Estimating parameters of multivariate models is a complex task because matrices of parameters must always fulfil specific conditions, for example positive definiteness for variance covariance matrices. Generally, one of the efficient ways to solve this issue is to use an unconstrained optimization of the likelihood with a parameterization that enforces the matrices to a particular structure (Pinheiro & Bates 1996). Various parameterizations are used in mvmorph (see Appendix S1) for ensuring positive definiteness of rate matrices (Pinheiro & Bates 1996), conditioning matrices on their eigenvalues (e.g. Sy, Taylor & Cumberland 1997; Jaffrezic, Thompson & Pletcher 24; Bartoszek et al. 212), or testing significance of interactions between traits by constraining the parameter space and taking advantage of joint likelihood optimization (e.g. Revell & Collar 29; Adams 213). Alternatives Other R implementations of multivariate comparative approaches already exist that calculate parts of the same or alternative models to those provided in mvmorph. Multivariate extensions of Pagel (1999) branch-length transformations are proposed in MOTMOT (Thomas & Freckleton 212; no more on the CRAN repository but accessible from the archive); BM with multiple evolutionary rate matrices can be fitted with evol.vcv as well as a phylogenetic canonical correlation analysis with phyl.cca in phytools (Revell 212), and geomorph allows computing BM rates for high-dimensional data sets (Adams & Otarola-Castillo 213). Multivariate OU is available from the OUCH package (Butler & King 29) as well as mvslouch that allows fitting multivariate OU models with a randomly evolving predictor variable (Bartoszek et al. 212). Drawbacks COMPUTATIONAL PERFORMANCES One of the main drawback of working with multivariate dimensions is that computational time grows up exponentially with variance covariance matrix construction, inversion and computation of the determinant (Felsenstein 1973; Henderson 1976; Hadfield & Nakagawa 21; Freckleton 212). Recent alternative algorithms have been published to cope with the problem of growing number of species and dimensions in comparative analyses (Hadfield & Nakagawa 21; FitzJohn 212; Thomas & Freckleton 212; Ho & Ane 214; Lartillot 214). Nevertheless, no general solution emerged so far, and most of these fast algorithms remain actually limited with multivariate data. For instance, the algorithm proposed by Ho & Ane (214) can be easily extended to multivariate case when phylogenetic variance covariance submatrices are proportional and that the multivariate variance covariance matrix maintains a three-point structure, which is unlikely for non-brownian processes such as the multivariate OU. Similarly, the pruning algorithm used by Thomas & Freckleton (212) in the MOTMOT package allows estimating a limited range of multivariate models such as a single optimum OU where selection acts independently on each traits. Facing this lack of general solution, particularly for complex models such as multivariate OU processes, mvmorph implements various approaches to speed up the computations (see Appendix S1 for details). The main implementations consist in different methods that avoid the explicit computation of the inverse and determinant of the variance covariance matrix to solve the linear model in eqn 1 through Cholesky factorization and independent contrasts. Two efficient algorithms are used for computing the Cholesky factors. The first one (method = rpf ) uses the rectangular full-packed format algorithm proposed by Gustavson et al. (21). This algorithm uses half memory for the storage of symmetric matrices and reorganizes the upper triangular matrix in a rectangular format allowing the use of fast BLAS 3 matrix multiplication routines. The second method(method = sparse ) takes advantage of the sparsity structure of phylogenetic variance covariance matrices to use faster sparse methods and updating of the Cholesky factor (Furrer & Sain 21; see Appendix S1). Finally, an extremely fast implementation of the likelihood computation using contrasts (Freckleton 212; see Appendix S1) is also provided in mvmorph for BM-based models (method = pic ). Overall, the computational solutions implemented in mvmorph allow realistic computation times even for very large-sized data sets, with relative speeds several times faster than other available packages (see Appendix S1, Table S1).
7 mvmorph 1317 HIGH-DIMENSIONALITY With increased number of traits and parameters (Table 1), over-parameterized multivariate models are often highly penalized by model selection criteria (Burnham & Anderson 22; Revell & Collar 29). Moreover, when the number of analysed traits reaches the number of studied species, the matrix computations involved in the likelihood calculation cannot be completed and parameter estimates become less accurate (Adams 214a,b,c). Accordingly, Adams (214a,b,c) has proposed methods based on matrix distances to allow the use of large m/small n data sets. Although these nonparametric approaches are appealing for comparing high-dimensional multivariate data sets, they do not fully address all the multivariate questions of trait evolution as they summarize the whole covariation information through a single metric, they assume a common phylogenetic structure for each trait, and they are limited to the BM model. Indeed, high-dimensional methods in phylogenetic comparative analysis should deserve more attention in the future as common approaches used to reduce the dimensionality of such data sets (e.g. principal component analysis) are prone to several biases (Monteiro 213; Polly et al. 213; Uyeda, Caetano & Pennell 215). Worked examples To illustrate how mvmorph works, we describe and comment two simulated examples in the Appendix S2 (also included as vignettes in the mvmorph package). In the first example, we show how multivariate models can out-compete separate univariate analyses for a phylogeny involving two distinct selective regimes corresponding to two separate species groups. In the second worked example, we illustrate how to assess significances of differences in evolutionary covariations by testing a hierarchy of nested models. In addition, the help pages embedded within the mvmorph package also provide several simulated examples illustrating how the various functions work. Conclusion Elaboration of the mvmorph package was initially motivated by the need for a unified R package for fitting multivariate models of phenotypic evolution incorporating measurement errors and using flexible ways to map evolutionary scenarios for morphometric data. Obviously, such objectives can be extended to any biological data set made of covarying continuous traits, as well as univariate traits. mvmorph also implements models that were not previously designed for the multivariate case, as well as new ones. Last, but not least, the fast likelihood implementations provided by the mvmorph package may be particularly appealing when dealing with large data sets and assessing the statistical power of comparative methods through modelling approaches (Boettiger, Coop & Ralph 212). The mvmorph package is in continuous development and can be followed and contributed from the github website ( mvmorph). mvmorph (current version: 1..5) is an opensource R package available for download on the CRAN repository ( Acknowledgements J.C. warmly thanks Aaron A. King and Krzysztof Bartoszek for their patience and help in understanding how the multivariate OU model works. We thank Emmanuel Paradis for sharing codes and Graham J. Slater for sharing an unpublished manuscript as well as for precisions on his models. We thank Jonathan Drury for his advices and help for correcting the package, and Fabien Dubuffet for his help with some computational procedures and access to the cluster under his care for the package testing. Thanks to Helene Morlon and members of her laboratory (Jonathan Drury, Eric Lewitus, Marc Manceau and Olivier Missa) for helpful discussions. We finally thank Vincent Bonhomme, Matthew W. Pennell, Thimothee Poisot, Liam J. Revell, Graham J. Slater, two anonymous reviewers and the Journal editor Robert B. O Hara for comments that greatly improved a first version of this manuscript. Data accessibility This manuscript does not include any data. References Adams, D.C. (213) Comparing evolutionary rates for different phenotypic traits on a phylogeny using likelihood. Systematic Biology, 62, Adams, D.C. (214a) A generalized K statistic for estimating phylogenetic signal from shape and other high-dimensional multivariate data. Systematic Biology, 63, Adams, D.C. (214b) A method for assessing phylogenetic least squares models for shape and other high-dimensional multivariate data. Evolution, 68, Adams, D.C. (214c) Quantifying and comparing phylogenetic evolutionary rates for shape and other high-dimensional phenotypic data. Systematic Biology, 63, Adams, D.C. & Otarola-Castillo, E. (213) geomorph: an R package for the collection and analysis of geometric morphometric shape data. Methods in Ecology and Evolution, 4, Adams, D.C., Berns, C.M., Kozak, K.H. & Wiens, J.J. (29) Are rates of species diversification correlated with rates of morphological evolution? Proceedings of the Royal Society B, 276, Armbruster, W.S. & Schwaegerle, K.E. (1996) Causes of covariation of phenotypic traits among populations. Journal of Evolutionary Biology, 9, Armbruster, W.S., Pelabon, C., Bolstad, G.H. & Hansen, T.F. (214) Integrated phenotypes: understanding trait covariation in plants and animals. Philosophical Transactions of the Royal Society of London B: Biological Sciences, 369, Arnold, S.J., Pfrender, M.E. & Jones, A.G. (21) The adaptive landscape as a conceptual bridge between micro and macroevolution. Genetica, , Baker, R.H. & Wilkinson, G.S. (23) Phylogenetic analysis of correlation structure in stalk-eyed flies (Diasemopsis, Diopsidae). Evolution, 57, Bartoszek, K., Pienaar, J., Mostad, P., Andersson, S. & Hansen, T.F. (212) A phylogenetic comparative method for studying multivariate adaptation. Journal of Theoretical Biology, 314, Beaulieu, J.M., Jhwueng, D.-C., Boettiger, C. & O Meara, B.C. (212) Modeling stabilizing selection: expanding the Ornstein-Uhlenbeck model of adaptive evolution. Evolution, 66, Blomberg, S.P., Garland, T.J. & Ives, A.R. (23) Testing for phylogenetic signal in comparative data: behavioral traits are more labile. Evolution, 57, Boettiger, C., Coop, G. & Ralph, P. (212) Is your phylogeny informative? Measuring the power of comparative methods. Evolution, 66, Bollback, J.B. (26) SIMMAP: Stochastic character mapping of discrete traits on phylogenies. BMC Bioinformatics, 7, 1 7. Burnham, K.P. & Anderson, D.R. (22) Model Selection and Multi-Model Inference: A Practical Information-Theoretic Approach. Springer-Verlag, New York, USA. Butler, M.A. & King, A.A. (24) Phylogenetic comparative analysis: a modeling approach for adaptive evolution. The American Naturalist, 164,
8 1318 J. Clavel, G. Escarguel, & G. Merceron Butler, M.A. & King, A.A. (29) Multivariate comparative analysis using OUCH. Integrative Comparative Biology, 57.5, e24. Cheverud, J. (1984) Quantitative genetics and developmental constraints on evolution by selection. Journal of Theoretical Biology, 11, Collar, D.C., Schulte, J.A. II & Losos, J.B. (211) Evolution of extreme body size disparity in monitor lizards (varanus). Evolution, 65, Collar, D.C., O Meara, B.C., Wainwright, P.C. & Near, T.J. (29) Piscivory limits diversification of feeding morphology in centrarchid fishes. Evolution, 63, Cooper, N. & Purvis, A. (21) Body size evolution in mammals: complexity in tempo and mode. The American Naturalist, 175, Endler, J.A. (1995) Multiple-trait coevolution and environmental gradients in guppies. Trends in Ecology & Evolution, 1, Felsenstein, J. (1973) Maximum-likelihood estimation of evolutionary trees from continuous characters. American Journal of Human Genetics, 25, Felsenstein, J. (1985) Phylogenies and the comparative method. The American Naturalist, 125, Felsenstein, J. (1988) Phylogenies and quantitative characters. Annual Review of Ecology, Evolution and Systematics, 19, Felsenstein, J. (24) Inferring Phylogenies. Sinauer Associates, Sunderland, Massachusetts, USA. Felsenstein, J. (28) Comparative methods with sampling error and within-species variation : contrasts revisited and revised. The American Naturalist, 171, FitzJohn, R.G. (212) Diversitree: comparative phylogenetic analyses of diversification in R. Methods in Ecology and Evolution, 3, Freckleton, R.P. (212) Fast likelihood calculations for comparative analyses. Methods in Ecology and Evolution, 3, Frederich, B., Sorenson, L., Santini, F., Slater, G.J. & Alfaro, M.E. (213) Iterative ecological radiation and convergence during the evolutionary history of damselfishes (Pomacentridae). The American Naturalist, 181, Furrer, R. & Sain, S.R. (21) Spam: A sparse matrix R package with emphasis on MCMC methods for Gaussian Markov random fields. Journal of Statistical Software, 36,1 25. Gardiner, C.W. (24) Handbook of Stochastic Methods for Physics, Chemistry and the Natural Sciences, 3rd edn. Springer, Berlin. Goswami, A., Smaers, J.B., Soligo, C. & Polly, P.D. (214) The macroevolutionary consequences of phenotypic integration: from development to deep time. Philosophical Transactions of the Royal Society of London B: Biological Sciences, 369, Gustavson, F.G., Wasniewski, J., Dongarra, J.J. & Langou, J. (21) Rectangular full packed format for Cholesky s algorithm: factorization, solution and inversion. ACM Transactions on Mathematical Software, 37, Hadfield, J.D. & Nakagawa, S. (21) General quantitative genetic methods for comparative biology: phylogenies, taxonomies and multi-trait models for continuous and categorical characters. Journal of Evolutionary Biology, 23, Hansen, T.F. (1997) Stabilizing selection and the comparative analysis of adaptation. Evolution, 51, Hansen, T.F. (212) Adaptive landscape and macroevolutionary dynamics. The Adaptive Landscape in Evolutionary Biology (eds E. Svensson & R. Calsbeek), pp Oxford University Press, Oxford. Hansen, T.F. & Bartoszek, K. (212) Interpreting the evolutionary regression : the interplay between observational and biological errors in phylogenetic comparative studies. Systematic Biology, 61, Hansen, T.F., Pienaar, J. & Orzack, S.H. (28) A comparative method for studying adaptation to a randomly evolving environment. Evolution, 62, Harmon, L.J., Losos, J.B., Davies, J.T., Gillespie, R.G., Gittleman, J.L., Jennings, B.W. et al. (21) Early bursts of body size and shape evolution are rare in comparative data. Evolution, 64, Henderson, C.R. (1976) A simple method for computing the inverse of a numerator relationship matrix used in prediction of breeding values. Biometrics, 32, Ho, L.S.T. & Ane, C. (213) Asymptotic theory with hierarchical autocorrelation: Ornstein-Uhlenbeck tree models. The Annals of Statistics, 41, Ho, L.S.T.& Ane, C. (214) A linear-time algorithm for Gaussian and non- Gaussian trait evolution models. Systematic Biology, 63, Hohenlohe, P.A. & Arnold, S.J. (28) MIPoD: A hypothesis-testing framework for microevolutionary inference from patterns of divergence. The American Naturalist, 171, Huelsenbeck, J.P., Nielsen, R. & Bollback, J.B. (23) Stochastic mapping of morphological characters. Systematic Biology, 52, Hunt, G. (212) Measuring rates of phenotypic evolution and the inseparability of tempo and mode. Paleobiology, 38, Hunt, G., Bell, M.A. & Travis, M.P. (28) Evolution toward a new adaptive optimum: phenotypic evolution in a fossil stickleback lineage. Evolution, 62, Ingram, T., Harmon, L.J. & Shurin, J.B. (212) When should we expect early bursts of trait evolution in comparative data? Predictions from an evolutionary food web model. Journal of Evolutionary Biology, 25, Ingram, T. & Mahler, D.L. (213) SURFACE: detecting convergent evolution from comparative data by fitting Ornstein-Uhlenbeck models with stepwise Akaike Information Criterion. Methods in Ecology and Evolution, 4, Ives, A.R., Midford, P.E. & Garland, T.J. (27) Within-species variation and measurement error in phylogenetic comparative methods. Systematic Biology, 56, Jaffrezic, F., Thompson, R. & Pletcher, S.D. (24) Multivariate character process models for the analysis of two or more correlated function-valued traits. Genetics, 168, Knope, M.L. & Scales, J.A. (213) Adaptive morphological shifts to novel habitats in marine sculpin fishes. Journal of Evolutionary Biology, 26, Lande, R. (1976) Natural selection and random genetic drift in phenotypic evolution. Evolution, 3, Lande, R. (1979) Quantitative genetic analysis of multivariate evolution, applied to brain-body size allometry. Evolution, 33, Lande, R. & Arnold, S.J. (1983) The measurement of selection on correlated characters. Evolution, 37, Lartillot, N. (214) A phylogenetic Kalman filter for ancestral trait reconstruction using molecular data. Bioinformatics, 3, Mahler, D.L., Ingram, T., Revell, L.J. & Losos, J.B. (213) Exceptional convergence on the macroevolutionary landscape in island lizard radiations. Science, 341, Martins, E.P. & Hansen, T.F. (1997) Phylogenies and the comparative method: a general approach to incorporating phylogenetic information into the analysis of interspecific data. The American Naturalist, 149, Monteiro, L.R. (213) Morphometrics and the comparative method: studying the evolution of biological shape. Hystrix, the Italian Journal of Mammalogy, 24,1 8. O Meara, B.C. (212) Evolutionary inferences from phylogenies: a review of methods. Annual Review of Ecology, Evolution and Systematics, 43, O Meara, B.C., Ane, C., Sanderson, M.J. & Wainwright, P.C. (26) Testing for different rates of continuous trait evolution. Evolution, 6, Pagel, M.D. (1999) Inferring the historical patterns of biological evolution. Nature, 41, Pennell, M.W. & Harmon, L.J. (213) An integrative view of phylogenetic comparative methods: connections to population genetics, community ecology, and paleobiology. Annals of the New York Academy of Sciences, 1289, Pinheiro, J.C. & Bates, D.M. (1996) Unconstrained parameterizations for variance-covariance matrices. Statistics and Computing, 6, Polly, P.D. (24) On the simulation of the evolution of morphological shape: multivariate shape under selection and drift. Palaeontologia Electronica, 7.2.7A,1 28. Polly, P.D., Lawing, M.A., Fabre, A.-C. & Goswami, A. (213) Phylogenetic principal components analysis and geometric morphometrics. Hystrix, 24, 1 9. Price,S.A.,Holzman,R.,Near,T.J.&Wainwright,P.C.(211)Coralreefspromote the evolution of morphological diversity and ecological novelty in labrid fishes. Ecology Letters, 14, Price, S.A., Tavera, J.J., Near, T.J. & Wainwright, P.C. (213) Elevated rates of morphological and functional diversification in reef-dwelling haemulid fishes. Evolution, 67, Reitan, T., Schweder, T. & Henderiks, J. (212) Phenotypic evolution studied by layered stochastic differential equations. The Annals of Applied Statistics, 6, Revell, L.J. (29) Size-correction and principal components for interspecific comparative studies. Evolution, 63, Revell, L.J. (21) Phylogenetic signal and linear regression on species data. Methods in Ecology and Evolution, 1, Revell, L.J. (212) phytools: An R package for phylogenetic comparative biology (and other things). Methods in Ecology and Evolution, 3, Revell, L.J. & Collar, D.C. (29) Phylogenetic analysis of the evolutionary correlation using likelihood. Evolution, 63, Revell, L.J. & Harmon, L.J. (28) Testing quantitative genetic hypotheses about the evolutionary rate matrix for continuous characters. Evolutionary Ecology Research, 1,
9 mvmorph 1319 Revell, L.J. & Reynolds, R.G. (212) A new bayesian method for fitting evolutionary models to comparative data with intraspecific variation. Evolution, 66, Ricklefs,R.E.(26)Time,species,andthe generation of trait variance in clades. Systematic Biology, 55, Rohlf, F.J. (21) Comparative methods for the analysis of continuous variables : geometric interpretations. Evolution, 55, Scales, J.A., King, A.A. & Butler, M.A. (29) Running for your life or running for your dinner: what drives fiber-type evolution in lizard locomotor muscles? The American Naturalist, 173, Schluter, D. (1996) Adaptive radiation along genetic lines of least resistance. Evolution, 5, Shoval, O., Sheftel, H., Shinar, G., Hart, Y., Ramote, O., Mayo, A., Dekel, E., Kavanagh, K. & Alon, U. (212) Evolutionary trade-offs, pareto optimality, and the geometry of phenotype space. Science, 336, Silvestro, D., Kostikova, A., Litsios, G., Pearman, P.B. & Salamin, N. (215) Measurement errors should always be incorporated in phylogenetic comparative analysis. Methods in Ecology and Evolution, 6, Slater, G.J. (213) Phylogenetic evidence for a shift in the mode of mammalian body size evolution at the Cretaceous-Palaeogene boundary. Methods in Ecology and Evolution, 4, Slater, G.J. (214) Correction to Phylogenetic evidence for a shift in the mode of mammalian body size evolution at the Cretaceous-Palaeogene boundary, and a note on fitting macroevolutionary models to comparative paleontological data sets. Methods in Ecology and Evolution, 5, Slater, G.J. & Harmon, L.J. (213) Unifying fossils and phylogenies for comparative analyses of diversification and trait evolution. Methods in Ecology and Evolution, 4, Slater, G.J., Harmon, L.J. & Alfaro, M.E. (212) Integrating fossils with molecular phylogenies improves inference of trait evolution. Evolution, 66, Slater, G.J., Price, S.A., Santini, F. & Alfaro, M.E. (21) Diversity versus disparity and the radiation of modern cetaceans. Proceedings of the Royal Society B, 277, Steppan, S.J. (1997) Phylogenetic analysis of phenotypic covariance structure. II. Reconstructing matrix evolution. Evolution, 51, Steppan, S.J., Phillips, P.C. & Houle, D. (22) Comparative quantitative genetics: evolution of the G matrix. Trends in Ecology and Evolution, 17, Sy, J.P., Taylor, J.M.G. & Cumberland, W.G. (1997) A stochastic model for the analysis of bivariate longitudinal AIDS data. Biometrics, 53, Thomas, G.H. & Freckleton, R.P. (212) MOTMOT: models of trait macroevolution on trees. Methods in Ecology and Evolution, 3, Thomas, G.H., Freckleton, R.P. & Szekely, T. (26) Comparative analyses of the influence of developmental mode on phenotypic diversification rates in shorebirds. Proceedings of the Royal Society B: Biological Sciences, 273, Uyeda, J.C., Caetano, D.S. & Pennell, M.W. (215) Comparative analysis of principal components can be misleading. Systematic Biology, 64, Walsh, B. (27) Escape from flatland. Journal of Evolutionary Biology, 2, Walsh, B. & Blows, M.W. (29) Abundant genetic variation + strong selection = multivariate genetic constraints: a geometric view of adaptation. Annual Review of Ecology, Evolution and Systematics, 4, Zeng, Z.-B. (1988) Long-term correlated response, interpopulation covariation, and interspecific allometry. Evolution, 42, Zheng,L.,Ives,A.R.,Garland,T.J.,Larget,B.R.,Yu,Y.&Cao,K.(29)New multivariate tests for phylogenetic signal and trait correlations applied to ecophysiological phenotypes of nine Manglietia species. Functional Ecology, 23, Received 3 April 215; accepted 4 June 215 Handling Editor: Timothee Poisot Supporting Information Additional Supporting Information may be found in the online version of this article. Appendix S1. Details on computational methods used by mvmorph. Appendix S2. Two worked examples of evolutionary model fitting with mvmorph.
Models of Continuous Trait Evolution
 Models of Continuous Trait Evolution By Ricardo Betancur (slides courtesy of Graham Slater, J. Arbour, L. Harmon & M. Alfaro) How fast do animals evolve? C. Darwin G. Simpson J. Haldane How do we model
Models of Continuous Trait Evolution By Ricardo Betancur (slides courtesy of Graham Slater, J. Arbour, L. Harmon & M. Alfaro) How fast do animals evolve? C. Darwin G. Simpson J. Haldane How do we model
Quantitative evolution of morphology
 Quantitative evolution of morphology Properties of Brownian motion evolution of a single quantitative trait Most likely outcome = starting value Variance of the outcomes = number of step * (rate parameter)
Quantitative evolution of morphology Properties of Brownian motion evolution of a single quantitative trait Most likely outcome = starting value Variance of the outcomes = number of step * (rate parameter)
Package OUwie. August 29, 2013
 Package OUwie August 29, 2013 Version 1.34 Date 2013-5-21 Title Analysis of evolutionary rates in an OU framework Author Jeremy M. Beaulieu , Brian O Meara Maintainer
Package OUwie August 29, 2013 Version 1.34 Date 2013-5-21 Title Analysis of evolutionary rates in an OU framework Author Jeremy M. Beaulieu , Brian O Meara Maintainer
MOTMOT: models of trait macroevolution on trees
 Methods in Ecology and Evolution 2012, 3, 145 151 doi: 10.1111/j.2041-210X.2011.00132.x APPLICATION MOTMOT: models of trait macroevolution on trees Gavin H. Thomas 1 * and Robert P. Freckleton 2 1 School
Methods in Ecology and Evolution 2012, 3, 145 151 doi: 10.1111/j.2041-210X.2011.00132.x APPLICATION MOTMOT: models of trait macroevolution on trees Gavin H. Thomas 1 * and Robert P. Freckleton 2 1 School
BRIEF COMMUNICATIONS
 Evolution, 59(12), 2005, pp. 2705 2710 BRIEF COMMUNICATIONS THE EFFECT OF INTRASPECIFIC SAMPLE SIZE ON TYPE I AND TYPE II ERROR RATES IN COMPARATIVE STUDIES LUKE J. HARMON 1 AND JONATHAN B. LOSOS Department
Evolution, 59(12), 2005, pp. 2705 2710 BRIEF COMMUNICATIONS THE EFFECT OF INTRASPECIFIC SAMPLE SIZE ON TYPE I AND TYPE II ERROR RATES IN COMPARATIVE STUDIES LUKE J. HARMON 1 AND JONATHAN B. LOSOS Department
Chapter 6: Beyond Brownian Motion Section 6.1: Introduction
 Chapter 6: Beyond Brownian Motion Section 6.1: Introduction Detailed studies of contemporary evolution have revealed a rich variety of processes that influence how traits evolve through time. Consider
Chapter 6: Beyond Brownian Motion Section 6.1: Introduction Detailed studies of contemporary evolution have revealed a rich variety of processes that influence how traits evolve through time. Consider
Evolutionary models with ecological interactions. By: Magnus Clarke
 Evolutionary models with ecological interactions By: Magnus Clarke A thesis submitted in partial fulfilment of the requirements for the degree of Doctor of Philosophy The University of Sheffield Faculty
Evolutionary models with ecological interactions By: Magnus Clarke A thesis submitted in partial fulfilment of the requirements for the degree of Doctor of Philosophy The University of Sheffield Faculty
Does convergent evolution always result from different lineages experiencing similar
 Different evolutionary dynamics led to the convergence of clinging performance in lizard toepads Sarin Tiatragula 1 *, Gopal Murali 2 *, James T. Stroud 3 1 Department of Biological Sciences, Auburn University,
Different evolutionary dynamics led to the convergence of clinging performance in lizard toepads Sarin Tiatragula 1 *, Gopal Murali 2 *, James T. Stroud 3 1 Department of Biological Sciences, Auburn University,
Phylogenetic comparative methods, CEB Spring 2010 [Revised April 22, 2010]
![Phylogenetic comparative methods, CEB Spring 2010 [Revised April 22, 2010] Phylogenetic comparative methods, CEB Spring 2010 [Revised April 22, 2010]](/thumbs/93/112416870.jpg) Phylogenetic comparative methods, CEB 35300 Spring 2010 [Revised April 22, 2010] Instructors: Andrew Hipp, Herbarium Curator, Morton Arboretum. ahipp@mortonarb.org; 630-725-2094 Rick Ree, Associate Curator,
Phylogenetic comparative methods, CEB 35300 Spring 2010 [Revised April 22, 2010] Instructors: Andrew Hipp, Herbarium Curator, Morton Arboretum. ahipp@mortonarb.org; 630-725-2094 Rick Ree, Associate Curator,
Testing quantitative genetic hypotheses about the evolutionary rate matrix for continuous characters
 Evolutionary Ecology Research, 2008, 10: 311 331 Testing quantitative genetic hypotheses about the evolutionary rate matrix for continuous characters Liam J. Revell 1 * and Luke J. Harmon 2 1 Department
Evolutionary Ecology Research, 2008, 10: 311 331 Testing quantitative genetic hypotheses about the evolutionary rate matrix for continuous characters Liam J. Revell 1 * and Luke J. Harmon 2 1 Department
Measurement errors should always be incorporated in phylogenetic comparative analysis
 Methods in Ecology and Evolution 25, 6, 34 346 doi:./24-2x.2337 s should always be incorporated in phylogenetic comparative analysis Daniele Silvestro,2,3, Anna Kostikova 2,3, Glenn Litsios 2,3, Peter
Methods in Ecology and Evolution 25, 6, 34 346 doi:./24-2x.2337 s should always be incorporated in phylogenetic comparative analysis Daniele Silvestro,2,3, Anna Kostikova 2,3, Glenn Litsios 2,3, Peter
Intrinsic inference difficulties for trait evolution with Ornstein-Uhlenbeck models
 Methods in Ecology and Evolution 204, 5, 33 46 doi: 0./204-20X.2285 Intrinsic inference difficulties for trait evolution with Ornstein-Uhlenbeck models Lam Si Tung Ho, 2* and Cecile Ane Department of Statistics,
Methods in Ecology and Evolution 204, 5, 33 46 doi: 0./204-20X.2285 Intrinsic inference difficulties for trait evolution with Ornstein-Uhlenbeck models Lam Si Tung Ho, 2* and Cecile Ane Department of Statistics,
Phenotypic Evolution. and phylogenetic comparative methods. G562 Geometric Morphometrics. Department of Geological Sciences Indiana University
 Phenotypic Evolution and phylogenetic comparative methods Phenotypic Evolution Change in the mean phenotype from generation to generation... Evolution = Mean(genetic variation * selection) + Mean(genetic
Phenotypic Evolution and phylogenetic comparative methods Phenotypic Evolution Change in the mean phenotype from generation to generation... Evolution = Mean(genetic variation * selection) + Mean(genetic
How can molecular phylogenies illuminate morphological evolution?
 How can molecular phylogenies illuminate morphological evolution? 27 October 2016. Joe Felsenstein UNAM How can molecular phylogenies illuminate morphological evolution? p.1/81 Where this lecture fits
How can molecular phylogenies illuminate morphological evolution? 27 October 2016. Joe Felsenstein UNAM How can molecular phylogenies illuminate morphological evolution? p.1/81 Where this lecture fits
Biol 206/306 Advanced Biostatistics Lab 11 Models of Trait Evolution Fall 2016
 Biol 206/306 Advanced Biostatistics Lab 11 Models of Trait Evolution Fall 2016 By Philip J. Bergmann 0. Laboratory Objectives 1. Explore how evolutionary trait modeling can reveal different information
Biol 206/306 Advanced Biostatistics Lab 11 Models of Trait Evolution Fall 2016 By Philip J. Bergmann 0. Laboratory Objectives 1. Explore how evolutionary trait modeling can reveal different information
Rapid evolution of the cerebellum in humans and other great apes
 Rapid evolution of the cerebellum in humans and other great apes Article Accepted Version Barton, R. A. and Venditti, C. (2014) Rapid evolution of the cerebellum in humans and other great apes. Current
Rapid evolution of the cerebellum in humans and other great apes Article Accepted Version Barton, R. A. and Venditti, C. (2014) Rapid evolution of the cerebellum in humans and other great apes. Current
The phylogenetic effective sample size and jumps
 arxiv:1809.06672v1 [q-bio.pe] 18 Sep 2018 The phylogenetic effective sample size and jumps Krzysztof Bartoszek September 19, 2018 Abstract The phylogenetic effective sample size is a parameter that has
arxiv:1809.06672v1 [q-bio.pe] 18 Sep 2018 The phylogenetic effective sample size and jumps Krzysztof Bartoszek September 19, 2018 Abstract The phylogenetic effective sample size is a parameter that has
BAMMtools: an R package for the analysis of evolutionary dynamics on phylogenetic trees
 Methods in Ecology and Evolution 2, 5, 71 77 doi: 1.1111/241-21X.12199 APPLICATION BAMMtools: an R package for the analysis of evolutionary dynamics on phylogenetic trees Daniel L. Rabosky*, Michael Grundler,
Methods in Ecology and Evolution 2, 5, 71 77 doi: 1.1111/241-21X.12199 APPLICATION BAMMtools: an R package for the analysis of evolutionary dynamics on phylogenetic trees Daniel L. Rabosky*, Michael Grundler,
POSITIVE CORRELATION BETWEEN DIVERSIFICATION RATES AND PHENOTYPIC EVOLVABILITY CAN MIMIC PUNCTUATED EQUILIBRIUM ON MOLECULAR PHYLOGENIES
 doi:10.1111/j.1558-5646.2012.01631.x POSITIVE CORRELATION BETWEEN DIVERSIFICATION RATES AND PHENOTYPIC EVOLVABILITY CAN MIMIC PUNCTUATED EQUILIBRIUM ON MOLECULAR PHYLOGENIES Daniel L. Rabosky 1,2,3 1 Department
doi:10.1111/j.1558-5646.2012.01631.x POSITIVE CORRELATION BETWEEN DIVERSIFICATION RATES AND PHENOTYPIC EVOLVABILITY CAN MIMIC PUNCTUATED EQUILIBRIUM ON MOLECULAR PHYLOGENIES Daniel L. Rabosky 1,2,3 1 Department
Integrative Biology 200 "PRINCIPLES OF PHYLOGENETICS" Spring 2018 University of California, Berkeley
 Integrative Biology 200 "PRINCIPLES OF PHYLOGENETICS" Spring 2018 University of California, Berkeley B.D. Mishler Feb. 14, 2018. Phylogenetic trees VI: Dating in the 21st century: clocks, & calibrations;
Integrative Biology 200 "PRINCIPLES OF PHYLOGENETICS" Spring 2018 University of California, Berkeley B.D. Mishler Feb. 14, 2018. Phylogenetic trees VI: Dating in the 21st century: clocks, & calibrations;
Appendix from L. J. Revell, On the Analysis of Evolutionary Change along Single Branches in a Phylogeny
 008 by The University of Chicago. All rights reserved.doi: 10.1086/588078 Appendix from L. J. Revell, On the Analysis of Evolutionary Change along Single Branches in a Phylogeny (Am. Nat., vol. 17, no.
008 by The University of Chicago. All rights reserved.doi: 10.1086/588078 Appendix from L. J. Revell, On the Analysis of Evolutionary Change along Single Branches in a Phylogeny (Am. Nat., vol. 17, no.
ANALYSIS OF CHARACTER DIVERGENCE ALONG ENVIRONMENTAL GRADIENTS AND OTHER COVARIATES
 ORIGINAL ARTICLE doi:10.1111/j.1558-5646.2007.00063.x ANALYSIS OF CHARACTER DIVERGENCE ALONG ENVIRONMENTAL GRADIENTS AND OTHER COVARIATES Dean C. Adams 1,2,3 and Michael L. Collyer 1,4 1 Department of
ORIGINAL ARTICLE doi:10.1111/j.1558-5646.2007.00063.x ANALYSIS OF CHARACTER DIVERGENCE ALONG ENVIRONMENTAL GRADIENTS AND OTHER COVARIATES Dean C. Adams 1,2,3 and Michael L. Collyer 1,4 1 Department of
Evolutionary Physiology
 Evolutionary Physiology Yves Desdevises Observatoire Océanologique de Banyuls 04 68 88 73 13 desdevises@obs-banyuls.fr http://desdevises.free.fr 1 2 Physiology Physiology: how organisms work Lot of diversity
Evolutionary Physiology Yves Desdevises Observatoire Océanologique de Banyuls 04 68 88 73 13 desdevises@obs-banyuls.fr http://desdevises.free.fr 1 2 Physiology Physiology: how organisms work Lot of diversity
Optimum selection and OU processes
 Optimum selection and OU processes 29 November 2016. Joe Felsenstein Biology 550D Optimum selection and OU processes p.1/15 With selection... life is harder There is the Breeder s Equation of Wright and
Optimum selection and OU processes 29 November 2016. Joe Felsenstein Biology 550D Optimum selection and OU processes p.1/15 With selection... life is harder There is the Breeder s Equation of Wright and
Statistical nonmolecular phylogenetics: can molecular phylogenies illuminate morphological evolution?
 Statistical nonmolecular phylogenetics: can molecular phylogenies illuminate morphological evolution? 30 July 2011. Joe Felsenstein Workshop on Molecular Evolution, MBL, Woods Hole Statistical nonmolecular
Statistical nonmolecular phylogenetics: can molecular phylogenies illuminate morphological evolution? 30 July 2011. Joe Felsenstein Workshop on Molecular Evolution, MBL, Woods Hole Statistical nonmolecular
Phylogenetic Comparative Analysis: A Modeling Approach for Adaptive Evolution
 vol. 164, no. 6 the american naturalist december 2004 Phylogenetic Comparative Analysis: A Modeling Approach for Adaptive Evolution Marguerite A. Butler * and Aaron A. King Department of Ecology and Evolutionary
vol. 164, no. 6 the american naturalist december 2004 Phylogenetic Comparative Analysis: A Modeling Approach for Adaptive Evolution Marguerite A. Butler * and Aaron A. King Department of Ecology and Evolutionary
GEOMETRIC MORPHOMETRICS. Adrian Castellanos, Michelle Chrpa, & Pedro Afonso Leite
 GEOMETRIC MORPHOMETRICS Adrian Castellanos, Michelle Chrpa, & Pedro Afonso Leite WHAT IS MORPHOMETRICS? Quantitative analysis of form, a concept that encompasses size and shape. Analyses performed on live
GEOMETRIC MORPHOMETRICS Adrian Castellanos, Michelle Chrpa, & Pedro Afonso Leite WHAT IS MORPHOMETRICS? Quantitative analysis of form, a concept that encompasses size and shape. Analyses performed on live
20 Grenoble, France ; CNRS, Laboratoire d Écologie Alpine (LECA), F Grenoble, France
 1/23 1 Improving phylogenetic regression under complex evolutionary models 2 Florent Mazel, T. Jonathan Davies, Damien Georges, Sébastien Lavergne, Wilfried Thuiller and 3 Pedro R. Peres-Neto 4 Running
1/23 1 Improving phylogenetic regression under complex evolutionary models 2 Florent Mazel, T. Jonathan Davies, Damien Georges, Sébastien Lavergne, Wilfried Thuiller and 3 Pedro R. Peres-Neto 4 Running
Concepts and Methods in Molecular Divergence Time Estimation
 Concepts and Methods in Molecular Divergence Time Estimation 26 November 2012 Prashant P. Sharma American Museum of Natural History Overview 1. Why do we date trees? 2. The molecular clock 3. Local clocks
Concepts and Methods in Molecular Divergence Time Estimation 26 November 2012 Prashant P. Sharma American Museum of Natural History Overview 1. Why do we date trees? 2. The molecular clock 3. Local clocks
Phylogenetic Signal, Evolutionary Process, and Rate
 Syst. Biol. 57(4):591 601, 2008 Copyright c Society of Systematic Biologists ISSN: 1063-5157 print / 1076-836X online DOI: 10.1080/10635150802302427 Phylogenetic Signal, Evolutionary Process, and Rate
Syst. Biol. 57(4):591 601, 2008 Copyright c Society of Systematic Biologists ISSN: 1063-5157 print / 1076-836X online DOI: 10.1080/10635150802302427 Phylogenetic Signal, Evolutionary Process, and Rate
Laboratoire d Écologie Alpine (LECA), University of Grenoble Alpes, F-38000, Grenoble, France 2
 Reports Ecology, 97(2), 2016, pp. 286 293 2016 by the Ecological Society of America Improving phylogenetic regression under complex evolutionary models Florent Mazel, 1,2,6 T. Jonathan Davies, 3,4 Damien
Reports Ecology, 97(2), 2016, pp. 286 293 2016 by the Ecological Society of America Improving phylogenetic regression under complex evolutionary models Florent Mazel, 1,2,6 T. Jonathan Davies, 3,4 Damien
"PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200 Spring 2018 University of California, Berkeley
 "PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200 Spring 2018 University of California, Berkeley D.D. Ackerly Feb. 26, 2018 Maximum Likelihood Principles, and Applications to
"PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200 Spring 2018 University of California, Berkeley D.D. Ackerly Feb. 26, 2018 Maximum Likelihood Principles, and Applications to
Contrasts for a within-species comparative method
 Contrasts for a within-species comparative method Joseph Felsenstein, Department of Genetics, University of Washington, Box 357360, Seattle, Washington 98195-7360, USA email address: joe@genetics.washington.edu
Contrasts for a within-species comparative method Joseph Felsenstein, Department of Genetics, University of Washington, Box 357360, Seattle, Washington 98195-7360, USA email address: joe@genetics.washington.edu
Nonlinear relationships and phylogenetically independent contrasts
 doi:10.1111/j.1420-9101.2004.00697.x SHORT COMMUNICATION Nonlinear relationships and phylogenetically independent contrasts S. QUADER,* K. ISVARAN,* R. E. HALE,* B. G. MINER* & N. E. SEAVY* *Department
doi:10.1111/j.1420-9101.2004.00697.x SHORT COMMUNICATION Nonlinear relationships and phylogenetically independent contrasts S. QUADER,* K. ISVARAN,* R. E. HALE,* B. G. MINER* & N. E. SEAVY* *Department
Quantifying and Comparing Phylogenetic Evolutionary Rates for Shape and Other High-Dimensional Phenotypic Data
 Syst. Biol. 63(2):166 177, 2014 The Author(s) 2013. Published by Oxford University Press, on behalf of the Society of Systematic Biologists. All rights reserved. For Permissions, please email: journals.permissions@oup.com
Syst. Biol. 63(2):166 177, 2014 The Author(s) 2013. Published by Oxford University Press, on behalf of the Society of Systematic Biologists. All rights reserved. For Permissions, please email: journals.permissions@oup.com
An integrative view of phylogenetic comparative methods: connections to population genetics, community ecology, and paleobiology
 Ann. N.Y. Acad. Sci. ISSN 0077-8923 ANNALS OF THE NEW YORK ACADEMY OF SCIENCES Issue: The Year in Evolutionary Biology An integrative view of phylogenetic comparative methods: connections to population
Ann. N.Y. Acad. Sci. ISSN 0077-8923 ANNALS OF THE NEW YORK ACADEMY OF SCIENCES Issue: The Year in Evolutionary Biology An integrative view of phylogenetic comparative methods: connections to population
Lineage-dependent rates of evolutionary diversification: analysis of bivariate ellipses
 Functional Ecology 1998 ORIGINAL ARTICLE OA 000 EN Lineage-dependent rates of evolutionary diversification: analysis of bivariate ellipses R. E. RICKLEFS* and P. NEALEN *Department of Biology, University
Functional Ecology 1998 ORIGINAL ARTICLE OA 000 EN Lineage-dependent rates of evolutionary diversification: analysis of bivariate ellipses R. E. RICKLEFS* and P. NEALEN *Department of Biology, University
Phylogeny, trees and morphospace
 G562 Geometric Morphometrics Phylogeny, trees and morphospace Hierarchical patterns in morphometric data WALLABY HUMAN Node 0 LEOPARD Node 1 Node 3 FOSSA Node 2 DOG Node 4 OTTER 0 20 40 60 80 Cottonwood
G562 Geometric Morphometrics Phylogeny, trees and morphospace Hierarchical patterns in morphometric data WALLABY HUMAN Node 0 LEOPARD Node 1 Node 3 FOSSA Node 2 DOG Node 4 OTTER 0 20 40 60 80 Cottonwood
Some of these slides have been borrowed from Dr. Paul Lewis, Dr. Joe Felsenstein. Thanks!
 Some of these slides have been borrowed from Dr. Paul Lewis, Dr. Joe Felsenstein. Thanks! Paul has many great tools for teaching phylogenetics at his web site: http://hydrodictyon.eeb.uconn.edu/people/plewis
Some of these slides have been borrowed from Dr. Paul Lewis, Dr. Joe Felsenstein. Thanks! Paul has many great tools for teaching phylogenetics at his web site: http://hydrodictyon.eeb.uconn.edu/people/plewis
OVERVIEW. L5. Quantitative population genetics
 L5. Quantitative population genetics OVERVIEW. L1. Approaches to ecological modelling. L2. Model parameterization and validation. L3. Stochastic models of population dynamics (math). L4. Animal movement
L5. Quantitative population genetics OVERVIEW. L1. Approaches to ecological modelling. L2. Model parameterization and validation. L3. Stochastic models of population dynamics (math). L4. Animal movement
paleotree: an R package for paleontological and phylogenetic analyses of evolution
 Methods in Ecology and Evolution 2012, 3, 803 807 doi: 10.1111/j.2041-210X.2012.00223.x APPLICATION paleotree: an R package for paleontological and phylogenetic analyses of evolution David W. Bapst* Department
Methods in Ecology and Evolution 2012, 3, 803 807 doi: 10.1111/j.2041-210X.2012.00223.x APPLICATION paleotree: an R package for paleontological and phylogenetic analyses of evolution David W. Bapst* Department
Euclidean Nature of Phylogenetic Distance Matrices
 Syst. Biol. 60(6):86 83, 011 c The Author(s) 011. Published by Oxford University Press, on behalf of the Society of Systematic Biologists. All rights reserved. For Permissions, please email: journals.permissions@oup.com
Syst. Biol. 60(6):86 83, 011 c The Author(s) 011. Published by Oxford University Press, on behalf of the Society of Systematic Biologists. All rights reserved. For Permissions, please email: journals.permissions@oup.com
Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut
 Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic analysis Phylogenetic Basics: Biological
Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic analysis Phylogenetic Basics: Biological
APPENDIX S1: DESCRIPTION OF THE ESTIMATION OF THE VARIANCES OF OUR MAXIMUM LIKELIHOOD ESTIMATORS
 APPENDIX S1: DESCRIPTION OF THE ESTIMATION OF THE VARIANCES OF OUR MAXIMUM LIKELIHOOD ESTIMATORS For large n, the negative inverse of the second derivative of the likelihood function at the optimum can
APPENDIX S1: DESCRIPTION OF THE ESTIMATION OF THE VARIANCES OF OUR MAXIMUM LIKELIHOOD ESTIMATORS For large n, the negative inverse of the second derivative of the likelihood function at the optimum can
ADAPTIVE CONSTRAINTS AND THE PHYLOGENETIC COMPARATIVE METHOD: A COMPUTER SIMULATION TEST
 Evolution, 56(1), 2002, pp. 1 13 ADAPTIVE CONSTRAINTS AND THE PHYLOGENETIC COMPARATIVE METHOD: A COMPUTER SIMULATION TEST EMÍLIA P. MARTINS, 1,2 JOSÉ ALEXANDRE F. DINIZ-FILHO, 3,4 AND ELIZABETH A. HOUSWORTH
Evolution, 56(1), 2002, pp. 1 13 ADAPTIVE CONSTRAINTS AND THE PHYLOGENETIC COMPARATIVE METHOD: A COMPUTER SIMULATION TEST EMÍLIA P. MARTINS, 1,2 JOSÉ ALEXANDRE F. DINIZ-FILHO, 3,4 AND ELIZABETH A. HOUSWORTH
MIPoD: A Hypothesis-Testing Framework for Microevolutionary Inference from Patterns of Divergence
 vol. 171, no. 3 the american naturalist march 2008 MIPoD: A Hypothesis-Testing Framework for Microevolutionary Inference from Patterns of Divergence Paul A. Hohenlohe 1,2,* and Stevan J. Arnold 1, 1. Department
vol. 171, no. 3 the american naturalist march 2008 MIPoD: A Hypothesis-Testing Framework for Microevolutionary Inference from Patterns of Divergence Paul A. Hohenlohe 1,2,* and Stevan J. Arnold 1, 1. Department
Systematics - Bio 615
 Bayesian Phylogenetic Inference 1. Introduction, history 2. Advantages over ML 3. Bayes Rule 4. The Priors 5. Marginal vs Joint estimation 6. MCMC Derek S. Sikes University of Alaska 7. Posteriors vs Bootstrap
Bayesian Phylogenetic Inference 1. Introduction, history 2. Advantages over ML 3. Bayes Rule 4. The Priors 5. Marginal vs Joint estimation 6. MCMC Derek S. Sikes University of Alaska 7. Posteriors vs Bootstrap
New multivariate tests for phylogenetic signal and trait correlations applied to ecophysiological phenotypes of nine Manglietia species
 Functional Ecology 2009, 23, 1059 1069 doi: 10.1111/j.1365-2435.2009.01596.x New multivariate tests for phylogenetic signal and trait correlations applied to ecophysiological phenotypes of nine Manglietia
Functional Ecology 2009, 23, 1059 1069 doi: 10.1111/j.1365-2435.2009.01596.x New multivariate tests for phylogenetic signal and trait correlations applied to ecophysiological phenotypes of nine Manglietia
Multiple regression and inference in ecology and conservation biology: further comments on identifying important predictor variables
 Biodiversity and Conservation 11: 1397 1401, 2002. 2002 Kluwer Academic Publishers. Printed in the Netherlands. Multiple regression and inference in ecology and conservation biology: further comments on
Biodiversity and Conservation 11: 1397 1401, 2002. 2002 Kluwer Academic Publishers. Printed in the Netherlands. Multiple regression and inference in ecology and conservation biology: further comments on
ASYMMETRIES IN PHYLOGENETIC DIVERSIFICATION AND CHARACTER CHANGE CAN BE UNTANGLED
 doi:1.1111/j.1558-5646.27.252.x ASYMMETRIES IN PHYLOGENETIC DIVERSIFICATION AND CHARACTER CHANGE CAN BE UNTANGLED Emmanuel Paradis Institut de Recherche pour le Développement, UR175 CAVIAR, GAMET BP 595,
doi:1.1111/j.1558-5646.27.252.x ASYMMETRIES IN PHYLOGENETIC DIVERSIFICATION AND CHARACTER CHANGE CAN BE UNTANGLED Emmanuel Paradis Institut de Recherche pour le Développement, UR175 CAVIAR, GAMET BP 595,
Integrating Fossils into Phylogenies. Throughout the 20th century, the relationship between paleontology and evolutionary biology has been strained.
 IB 200B Principals of Phylogenetic Systematics Spring 2011 Integrating Fossils into Phylogenies Throughout the 20th century, the relationship between paleontology and evolutionary biology has been strained.
IB 200B Principals of Phylogenetic Systematics Spring 2011 Integrating Fossils into Phylogenies Throughout the 20th century, the relationship between paleontology and evolutionary biology has been strained.
Greater host breadth still not associated with increased diversification rate in the Nymphalidae A response to Janz et al.
 doi:10.1111/evo.12914 Greater host breadth still not associated with increased diversification rate in the Nymphalidae A response to Janz et al. Christopher A. Hamm 1,2 and James A. Fordyce 3 1 Department
doi:10.1111/evo.12914 Greater host breadth still not associated with increased diversification rate in the Nymphalidae A response to Janz et al. Christopher A. Hamm 1,2 and James A. Fordyce 3 1 Department
Generalized Linear Models (GLZ)
 Generalized Linear Models (GLZ) Generalized Linear Models (GLZ) are an extension of the linear modeling process that allows models to be fit to data that follow probability distributions other than the
Generalized Linear Models (GLZ) Generalized Linear Models (GLZ) are an extension of the linear modeling process that allows models to be fit to data that follow probability distributions other than the
Discrete & continuous characters: The threshold model
 Discrete & continuous characters: The threshold model Discrete & continuous characters: the threshold model So far we have discussed continuous & discrete character models separately for estimating ancestral
Discrete & continuous characters: The threshold model Discrete & continuous characters: the threshold model So far we have discussed continuous & discrete character models separately for estimating ancestral
A NEW BAYESIAN METHOD FOR FITTING EVOLUTIONARY MODELS TO COMPARATIVE DATA WITH INTRASPECIFIC VARIATION
 ORIGINAL ARTICLE doi:10.1111/j.1558-5646.01.01645.x A NEW BAYESIAN METHOD FOR FITTING EVOLUTIONARY MODELS TO COMPARATIVE DATA WITH INTRASPECIFIC VARIATION Liam J. Revell 1, and R. Graham Reynolds 1 1 Department
ORIGINAL ARTICLE doi:10.1111/j.1558-5646.01.01645.x A NEW BAYESIAN METHOD FOR FITTING EVOLUTIONARY MODELS TO COMPARATIVE DATA WITH INTRASPECIFIC VARIATION Liam J. Revell 1, and R. Graham Reynolds 1 1 Department
Chapter 7: Models of discrete character evolution
 Chapter 7: Models of discrete character evolution pdf version R markdown to recreate analyses Biological motivation: Limblessness as a discrete trait Squamates, the clade that includes all living species
Chapter 7: Models of discrete character evolution pdf version R markdown to recreate analyses Biological motivation: Limblessness as a discrete trait Squamates, the clade that includes all living species
Likelihood Ratio Tests for Detecting Positive Selection and Application to Primate Lysozyme Evolution
 Likelihood Ratio Tests for Detecting Positive Selection and Application to Primate Lysozyme Evolution Ziheng Yang Department of Biology, University College, London An excess of nonsynonymous substitutions
Likelihood Ratio Tests for Detecting Positive Selection and Application to Primate Lysozyme Evolution Ziheng Yang Department of Biology, University College, London An excess of nonsynonymous substitutions
Experimental Design and Data Analysis for Biologists
 Experimental Design and Data Analysis for Biologists Gerry P. Quinn Monash University Michael J. Keough University of Melbourne CAMBRIDGE UNIVERSITY PRESS Contents Preface page xv I I Introduction 1 1.1
Experimental Design and Data Analysis for Biologists Gerry P. Quinn Monash University Michael J. Keough University of Melbourne CAMBRIDGE UNIVERSITY PRESS Contents Preface page xv I I Introduction 1 1.1
Biology 559R: Introduction to Phylogenetic Comparative Methods Topics for this week:
 Biology 559R: Introduction to Phylogenetic Comparative Methods Topics for this week: Course general information About the course Course objectives Comparative methods: An overview R as language: uses and
Biology 559R: Introduction to Phylogenetic Comparative Methods Topics for this week: Course general information About the course Course objectives Comparative methods: An overview R as language: uses and
"PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200B Spring 2011 University of California, Berkeley
 "PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200B Spring 2011 University of California, Berkeley B.D. Mishler Feb. 1, 2011. Qualitative character evolution (cont.) - comparing
"PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200B Spring 2011 University of California, Berkeley B.D. Mishler Feb. 1, 2011. Qualitative character evolution (cont.) - comparing
Supplemental Information Likelihood-based inference in isolation-by-distance models using the spatial distribution of low-frequency alleles
 Supplemental Information Likelihood-based inference in isolation-by-distance models using the spatial distribution of low-frequency alleles John Novembre and Montgomery Slatkin Supplementary Methods To
Supplemental Information Likelihood-based inference in isolation-by-distance models using the spatial distribution of low-frequency alleles John Novembre and Montgomery Slatkin Supplementary Methods To
Dr. Amira A. AL-Hosary
 Phylogenetic analysis Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic Basics: Biological
Phylogenetic analysis Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic Basics: Biological
Selection on Multiple Traits
 Selection on Multiple Traits Bruce Walsh lecture notes Uppsala EQG 2012 course version 7 Feb 2012 Detailed reading: Chapter 30 Genetic vs. Phenotypic correlations Within an individual, trait values can
Selection on Multiple Traits Bruce Walsh lecture notes Uppsala EQG 2012 course version 7 Feb 2012 Detailed reading: Chapter 30 Genetic vs. Phenotypic correlations Within an individual, trait values can
Elements of Multivariate Time Series Analysis
 Gregory C. Reinsel Elements of Multivariate Time Series Analysis Second Edition With 14 Figures Springer Contents Preface to the Second Edition Preface to the First Edition vii ix 1. Vector Time Series
Gregory C. Reinsel Elements of Multivariate Time Series Analysis Second Edition With 14 Figures Springer Contents Preface to the Second Edition Preface to the First Edition vii ix 1. Vector Time Series
25 : Graphical induced structured input/output models
 10-708: Probabilistic Graphical Models 10-708, Spring 2016 25 : Graphical induced structured input/output models Lecturer: Eric P. Xing Scribes: Raied Aljadaany, Shi Zong, Chenchen Zhu Disclaimer: A large
10-708: Probabilistic Graphical Models 10-708, Spring 2016 25 : Graphical induced structured input/output models Lecturer: Eric P. Xing Scribes: Raied Aljadaany, Shi Zong, Chenchen Zhu Disclaimer: A large
Modeling continuous trait evolution in the absence of phylogenetic information
 Modeling continuous trait evolution in the absence of phylogenetic information Mariia Koroliuk GU university, EM in complex systems under the supervision of Daniele Silvestro June 29, 2014 Contents 1 Introduction
Modeling continuous trait evolution in the absence of phylogenetic information Mariia Koroliuk GU university, EM in complex systems under the supervision of Daniele Silvestro June 29, 2014 Contents 1 Introduction
DNA-based species delimitation
 DNA-based species delimitation Phylogenetic species concept based on tree topologies Ø How to set species boundaries? Ø Automatic species delimitation? druhů? DNA barcoding Species boundaries recognized
DNA-based species delimitation Phylogenetic species concept based on tree topologies Ø How to set species boundaries? Ø Automatic species delimitation? druhů? DNA barcoding Species boundaries recognized
FACTOR ANALYSIS AND MULTIDIMENSIONAL SCALING
 FACTOR ANALYSIS AND MULTIDIMENSIONAL SCALING Vishwanath Mantha Department for Electrical and Computer Engineering Mississippi State University, Mississippi State, MS 39762 mantha@isip.msstate.edu ABSTRACT
FACTOR ANALYSIS AND MULTIDIMENSIONAL SCALING Vishwanath Mantha Department for Electrical and Computer Engineering Mississippi State University, Mississippi State, MS 39762 mantha@isip.msstate.edu ABSTRACT
How to measure and test phylogenetic signal
 Methods in Ecology and Evolution 2012, 3, 743 756 doi: 10.1111/j.2041-210X.2012.00196.x How to measure and test phylogenetic signal Tamara Mu nkemu ller 1 *, Se bastien Lavergne 1, Bruno Bzeznik 2, Ste
Methods in Ecology and Evolution 2012, 3, 743 756 doi: 10.1111/j.2041-210X.2012.00196.x How to measure and test phylogenetic signal Tamara Mu nkemu ller 1 *, Se bastien Lavergne 1, Bruno Bzeznik 2, Ste
From Individual-based Population Models to Lineage-based Models of Phylogenies
 From Individual-based Population Models to Lineage-based Models of Phylogenies Amaury Lambert (joint works with G. Achaz, H.K. Alexander, R.S. Etienne, N. Lartillot, H. Morlon, T.L. Parsons, T. Stadler)
From Individual-based Population Models to Lineage-based Models of Phylogenies Amaury Lambert (joint works with G. Achaz, H.K. Alexander, R.S. Etienne, N. Lartillot, H. Morlon, T.L. Parsons, T. Stadler)
Measuring rates of phenotypic evolution and the inseparability of tempo and mode
 Measuring rates of phenotypic evolution and the inseparability of tempo and mode Gene Hunt This pdf file is licensed for distribution in the form of electronic reprints and by way of personal or institutional
Measuring rates of phenotypic evolution and the inseparability of tempo and mode Gene Hunt This pdf file is licensed for distribution in the form of electronic reprints and by way of personal or institutional
Ronald Christensen. University of New Mexico. Albuquerque, New Mexico. Wesley Johnson. University of California, Irvine. Irvine, California
 Texts in Statistical Science Bayesian Ideas and Data Analysis An Introduction for Scientists and Statisticians Ronald Christensen University of New Mexico Albuquerque, New Mexico Wesley Johnson University
Texts in Statistical Science Bayesian Ideas and Data Analysis An Introduction for Scientists and Statisticians Ronald Christensen University of New Mexico Albuquerque, New Mexico Wesley Johnson University
ANCESTRAL CHARACTER ESTIMATION UNDER THE THRESHOLD MODEL FROM QUANTITATIVE GENETICS
 ORIGINAL ARTICLE doi:10.1111/evo.12300 ANCESTRAL CHARACTER ESTIMATION UNDER THE THRESHOLD MODEL FROM QUANTITATIVE GENETICS Liam J. Revell 1,2 1 Department of Biology, University of Massachusetts Boston,
ORIGINAL ARTICLE doi:10.1111/evo.12300 ANCESTRAL CHARACTER ESTIMATION UNDER THE THRESHOLD MODEL FROM QUANTITATIVE GENETICS Liam J. Revell 1,2 1 Department of Biology, University of Massachusetts Boston,
Phylogenetic signal and linear regression on species data
 Methods in Ecology and Evolution 2010, 1, 319 329 doi: 10.1111/j.2041-210X.2010.00044.x Phylogenetic signal and linear regression on species data Liam J. Revell* National Evolutionary Synthesis Center,
Methods in Ecology and Evolution 2010, 1, 319 329 doi: 10.1111/j.2041-210X.2010.00044.x Phylogenetic signal and linear regression on species data Liam J. Revell* National Evolutionary Synthesis Center,
DETECTING BIOLOGICAL AND ENVIRONMENTAL CHANGES: DESIGN AND ANALYSIS OF MONITORING AND EXPERIMENTS (University of Bologna, 3-14 March 2008)
 Dipartimento di Biologia Evoluzionistica Sperimentale Centro Interdipartimentale di Ricerca per le Scienze Ambientali in Ravenna INTERNATIONAL WINTER SCHOOL UNIVERSITY OF BOLOGNA DETECTING BIOLOGICAL AND
Dipartimento di Biologia Evoluzionistica Sperimentale Centro Interdipartimentale di Ricerca per le Scienze Ambientali in Ravenna INTERNATIONAL WINTER SCHOOL UNIVERSITY OF BOLOGNA DETECTING BIOLOGICAL AND
The Tempo of Macroevolution: Patterns of Diversification and Extinction
 The Tempo of Macroevolution: Patterns of Diversification and Extinction During the semester we have been consider various aspects parameters associated with biodiversity. Current usage stems from 1980's
The Tempo of Macroevolution: Patterns of Diversification and Extinction During the semester we have been consider various aspects parameters associated with biodiversity. Current usage stems from 1980's
Integrative Biology 200 "PRINCIPLES OF PHYLOGENETICS" Spring 2016 University of California, Berkeley. Parsimony & Likelihood [draft]
![Integrative Biology 200 PRINCIPLES OF PHYLOGENETICS Spring 2016 University of California, Berkeley. Parsimony & Likelihood [draft] Integrative Biology 200 PRINCIPLES OF PHYLOGENETICS Spring 2016 University of California, Berkeley. Parsimony & Likelihood [draft]](/thumbs/83/87443456.jpg) Integrative Biology 200 "PRINCIPLES OF PHYLOGENETICS" Spring 2016 University of California, Berkeley K.W. Will Parsimony & Likelihood [draft] 1. Hennig and Parsimony: Hennig was not concerned with parsimony
Integrative Biology 200 "PRINCIPLES OF PHYLOGENETICS" Spring 2016 University of California, Berkeley K.W. Will Parsimony & Likelihood [draft] 1. Hennig and Parsimony: Hennig was not concerned with parsimony
Four aspects of a sampling strategy necessary to make accurate and precise inferences about populations are:
 Why Sample? Often researchers are interested in answering questions about a particular population. They might be interested in the density, species richness, or specific life history parameters such as
Why Sample? Often researchers are interested in answering questions about a particular population. They might be interested in the density, species richness, or specific life history parameters such as
Stat 5101 Lecture Notes
 Stat 5101 Lecture Notes Charles J. Geyer Copyright 1998, 1999, 2000, 2001 by Charles J. Geyer May 7, 2001 ii Stat 5101 (Geyer) Course Notes Contents 1 Random Variables and Change of Variables 1 1.1 Random
Stat 5101 Lecture Notes Charles J. Geyer Copyright 1998, 1999, 2000, 2001 by Charles J. Geyer May 7, 2001 ii Stat 5101 (Geyer) Course Notes Contents 1 Random Variables and Change of Variables 1 1.1 Random
Phylogenetic ANOVA: Group-clade aggregation, biological challenges, and a refined permutation procedure
 ORIGINAL ARTICLE doi:10.1111/evo.13492 Phylogenetic ANOVA: Group-clade aggregation, biological challenges, and a refined permutation procedure Dean C. Adams 1,2,3 and Michael L. Collyer 4 1 Department
ORIGINAL ARTICLE doi:10.1111/evo.13492 Phylogenetic ANOVA: Group-clade aggregation, biological challenges, and a refined permutation procedure Dean C. Adams 1,2,3 and Michael L. Collyer 4 1 Department
Historical contingency, niche conservatism and the tendency for some taxa to be more diverse towards the poles
 Electronic Supplementary Material Historical contingency, niche conservatism and the tendency for some taxa to be more diverse towards the poles Ignacio Morales-Castilla 1,2 *, Jonathan T. Davies 3 and
Electronic Supplementary Material Historical contingency, niche conservatism and the tendency for some taxa to be more diverse towards the poles Ignacio Morales-Castilla 1,2 *, Jonathan T. Davies 3 and
an interspecific test of the van Noordwijk and de Jong model
 Functional Ecology 2000 Trade-offs between egg size and number in waterfowl: Blackwell Science, Ltd an interspecific test of the van Noordwijk and de Jong model J. K. CHRISTIANS Department of Biological
Functional Ecology 2000 Trade-offs between egg size and number in waterfowl: Blackwell Science, Ltd an interspecific test of the van Noordwijk and de Jong model J. K. CHRISTIANS Department of Biological
Non-independence in Statistical Tests for Discrete Cross-species Data
 J. theor. Biol. (1997) 188, 507514 Non-independence in Statistical Tests for Discrete Cross-species Data ALAN GRAFEN* AND MARK RIDLEY * St. John s College, Oxford OX1 3JP, and the Department of Zoology,
J. theor. Biol. (1997) 188, 507514 Non-independence in Statistical Tests for Discrete Cross-species Data ALAN GRAFEN* AND MARK RIDLEY * St. John s College, Oxford OX1 3JP, and the Department of Zoology,
25 : Graphical induced structured input/output models
 10-708: Probabilistic Graphical Models 10-708, Spring 2013 25 : Graphical induced structured input/output models Lecturer: Eric P. Xing Scribes: Meghana Kshirsagar (mkshirsa), Yiwen Chen (yiwenche) 1 Graph
10-708: Probabilistic Graphical Models 10-708, Spring 2013 25 : Graphical induced structured input/output models Lecturer: Eric P. Xing Scribes: Meghana Kshirsagar (mkshirsa), Yiwen Chen (yiwenche) 1 Graph
The Origin of Species
 The Origin of Species Introduction A species can be defined as a group of organisms whose members can breed and produce fertile offspring, but who do not produce fertile offspring with members of other
The Origin of Species Introduction A species can be defined as a group of organisms whose members can breed and produce fertile offspring, but who do not produce fertile offspring with members of other
Trees, branches and (square) roots: why evolutionary relatedness is not linearly related to functional distance
 Methods in Ecology and Evolution 2015, 6, 439 444 doi: 10.1111/2041-210X.12237 SPECIAL FEATURE NEW OPPORTUNITIES AT THE INTERFACE BETWEEN ECOLOGY AND STATISTICS Trees, branches and (square) roots: why
Methods in Ecology and Evolution 2015, 6, 439 444 doi: 10.1111/2041-210X.12237 SPECIAL FEATURE NEW OPPORTUNITIES AT THE INTERFACE BETWEEN ECOLOGY AND STATISTICS Trees, branches and (square) roots: why
Chapter 9: Beyond the Mk Model
 Chapter 9: Beyond the Mk Model Section 9.1: The Evolution of Frog Life History Strategies Frog reproduction is one of the most bizarrely interesting topics in all of biology. Across the nearly 6,000 species
Chapter 9: Beyond the Mk Model Section 9.1: The Evolution of Frog Life History Strategies Frog reproduction is one of the most bizarrely interesting topics in all of biology. Across the nearly 6,000 species
CONVERGENT EVOLUTION OF PHENOTYPIC INTEGRATION AND ITS ALIGNMENT WITH MORPHOLOGICAL DIVERSIFICATION IN CARIBBEAN ANOLIS ECOMORPHS
 ORIGINAL ARTICLE doi:10.1111/j.1558-5646.2011.01416.x CONVERGENT EVOLUTION OF PHENOTYPIC INTEGRATION AND ITS ALIGNMENT WITH MORPHOLOGICAL DIVERSIFICATION IN CARIBBEAN ANOLIS ECOMORPHS Jason J. Kolbe, 1,2
ORIGINAL ARTICLE doi:10.1111/j.1558-5646.2011.01416.x CONVERGENT EVOLUTION OF PHENOTYPIC INTEGRATION AND ITS ALIGNMENT WITH MORPHOLOGICAL DIVERSIFICATION IN CARIBBEAN ANOLIS ECOMORPHS Jason J. Kolbe, 1,2
Estimating Variances and Covariances in a Non-stationary Multivariate Time Series Using the K-matrix
 Estimating Variances and Covariances in a Non-stationary Multivariate ime Series Using the K-matrix Stephen P Smith, January 019 Abstract. A second order time series model is described, and generalized
Estimating Variances and Covariances in a Non-stationary Multivariate ime Series Using the K-matrix Stephen P Smith, January 019 Abstract. A second order time series model is described, and generalized
Historical Biogeography. Historical Biogeography. Systematics
 Historical Biogeography I. Definitions II. Fossils: problems with fossil record why fossils are important III. Phylogeny IV. Phenetics VI. Phylogenetic Classification Disjunctions debunked: Examples VII.
Historical Biogeography I. Definitions II. Fossils: problems with fossil record why fossils are important III. Phylogeny IV. Phenetics VI. Phylogenetic Classification Disjunctions debunked: Examples VII.
A (short) introduction to phylogenetics
 A (short) introduction to phylogenetics Thibaut Jombart, Marie-Pauline Beugin MRC Centre for Outbreak Analysis and Modelling Imperial College London Genetic data analysis with PR Statistics, Millport Field
A (short) introduction to phylogenetics Thibaut Jombart, Marie-Pauline Beugin MRC Centre for Outbreak Analysis and Modelling Imperial College London Genetic data analysis with PR Statistics, Millport Field
Regression. Oscar García
 Regression Oscar García Regression methods are fundamental in Forest Mensuration For a more concise and general presentation, we shall first review some matrix concepts 1 Matrices An order n m matrix is
Regression Oscar García Regression methods are fundamental in Forest Mensuration For a more concise and general presentation, we shall first review some matrix concepts 1 Matrices An order n m matrix is
Modeling Phylogenetic Comparative Methods with Hybridization
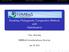 Modeling Phylogenetic Comparative Methods with Hybridization Tony Jhwueng NIMBioS Interdisciplinary Seminar Jan 25 2011 Outline: 1 Introduction: Phylognetic Comparative Methods (PCMs). 2 Modeling PCMs
Modeling Phylogenetic Comparative Methods with Hybridization Tony Jhwueng NIMBioS Interdisciplinary Seminar Jan 25 2011 Outline: 1 Introduction: Phylognetic Comparative Methods (PCMs). 2 Modeling PCMs
arxiv: v2 [q-bio.pe] 22 Sep 2017
![arxiv: v2 [q-bio.pe] 22 Sep 2017 arxiv: v2 [q-bio.pe] 22 Sep 2017](/thumbs/81/83178910.jpg) arxiv:1709.04702v2 [q-bio.pe] 22 Sep 2017 Jugowice, 11 th 15 th September 2017 TRAIT EVOLUTION WITH JUMPS: ILLUSIONARY NORMALITY Krzysztof Bartoszek 1 1 Department of Computer and Information Science,
arxiv:1709.04702v2 [q-bio.pe] 22 Sep 2017 Jugowice, 11 th 15 th September 2017 TRAIT EVOLUTION WITH JUMPS: ILLUSIONARY NORMALITY Krzysztof Bartoszek 1 1 Department of Computer and Information Science,
Plant of the Day Isoetes andicola
 Plant of the Day Isoetes andicola Endemic to central and southern Peru Found in scattered populations above 4000 m Restricted to the edges of bogs and lakes Leaves lack stomata and so CO 2 is obtained,
Plant of the Day Isoetes andicola Endemic to central and southern Peru Found in scattered populations above 4000 m Restricted to the edges of bogs and lakes Leaves lack stomata and so CO 2 is obtained,
Evolutionary Theory. Sinauer Associates, Inc. Publishers Sunderland, Massachusetts U.S.A.
 Evolutionary Theory Mathematical and Conceptual Foundations Sean H. Rice Sinauer Associates, Inc. Publishers Sunderland, Massachusetts U.S.A. Contents Preface ix Introduction 1 CHAPTER 1 Selection on One
Evolutionary Theory Mathematical and Conceptual Foundations Sean H. Rice Sinauer Associates, Inc. Publishers Sunderland, Massachusetts U.S.A. Contents Preface ix Introduction 1 CHAPTER 1 Selection on One
InDel 3-5. InDel 8-9. InDel 3-5. InDel 8-9. InDel InDel 8-9
 Lecture 5 Alignment I. Introduction. For sequence data, the process of generating an alignment establishes positional homologies; that is, alignment provides the identification of homologous phylogenetic
Lecture 5 Alignment I. Introduction. For sequence data, the process of generating an alignment establishes positional homologies; that is, alignment provides the identification of homologous phylogenetic
5/31/17. Week 10; Monday MEMORIAL DAY NO CLASS. Page 88
 Week 10; Monday MEMORIAL DAY NO CLASS Page 88 Week 10; Wednesday Announcements: Family ID final in lab Today Final exam next Tuesday at 8:30 am here Lecture: Species concepts & Speciation. What are species?
Week 10; Monday MEMORIAL DAY NO CLASS Page 88 Week 10; Wednesday Announcements: Family ID final in lab Today Final exam next Tuesday at 8:30 am here Lecture: Species concepts & Speciation. What are species?
PHYLOGENY & THE TREE OF LIFE
 PHYLOGENY & THE TREE OF LIFE PREFACE In this powerpoint we learn how biologists distinguish and categorize the millions of species on earth. Early we looked at the process of evolution here we look at
PHYLOGENY & THE TREE OF LIFE PREFACE In this powerpoint we learn how biologists distinguish and categorize the millions of species on earth. Early we looked at the process of evolution here we look at
PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION
 "PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200 Spring 2018 University of California, Berkeley D. Ackerly April 2, 2018 Lineage diversification Reading: Morlon, H. (2014).
"PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200 Spring 2018 University of California, Berkeley D. Ackerly April 2, 2018 Lineage diversification Reading: Morlon, H. (2014).
