Lecture 10: The quality of the Estimate
|
|
|
- Meagan Cook
- 5 years ago
- Views:
Transcription
1 Computational Biology Group (CoBI), D-BSSE, ETHZ Lecture 10: The quality of the Estimate Prof Dagmar Iber, PhD DPhil BSc Biotechnology 2015/16
2 May 12, / 102 Contents 1 Review of Previous Lectures 2 Post-regression diagnostics Goodness of fit (GOF) Uncertainty in the Parameters 3 Model Selection 4 Incorporation of prior information 5 Error Propagation
3 May 12, / 102 Literature D.S. Sivia, Data Analysis, OUP Jaqaman & Danuser, Linking data to models: data regression. Nat Rev Mol Cell Biol (2006) 7,
4 May 12, / 102 Parameter Inference 1 Pre-regression Diagnostics: Identifiability 2 Bayesian Inference 3 Post-regression Diagnostics
5 Review of Previous Lectures Computational Biology Group (CoBI), D-BSSE, ETHZ Prof Dagmar Iber, PhD DPhil Lecture 10 BSc Biotechnology May 12, / 102
6 May 12, / 102 Structural Identifiability Structural Identifiability A model with M state variables ( y) and P parameters ( p) is structurally identifiable if its sensitivity matrix S m i p j = y i p j, i = 1,..., M j = 1,..., P. (1) satisfies two conditions: each column has at least one large entry (i.e. each parameter has a large impact on at least one experimental measurement) the matrix has full rank (i.e. all columns must be linearly independent, which means that the effects of the parameters on the measurements must be independent of each other.)
7 May 12, / 102 Structural & Practical Identifiability If parameters are correlated then only relative values can be determined for the correlated parameters since their effects compensate for each other.
8 Bayes Theorem for Parameter Estimation According to Bayes Theorem prob(x D, I ) = prob(d X, I ) prob(x I ) prob(d I ) prob(x D, I ): posterior probability density function (pdf) that we want to determine. prob(d X, I ) : likelihood function prob(x I ): prior probability density function (pdf) that reflects our knowledge about the system prob(d I ): evidence, i.e. the likelihood of the data based on our knowledge. Here one could incorporate knowledge about the quality of different experimental techniques or experimental groups. Computational Biology Group (CoBI), D-BSSE, ETHZ Prof Dagmar Iber, PhD DPhil Lecture 10 BSc Biotechnology May 12, / 102
9 May 12, / 102 Maximum likelihood estimate prob(x D, I ) prob(d X, I ) (2) posterior pdf likelihood function. Maximum likelihood estimate Our best estimate X 0, given by the maximum of the posterior, is equivalent to the solution that yields the greatest value for the probability of the observed data.
10 May 12, / 102 Assume Gaussian Process We can then write prob(d k X, I ) = 1 σ k 2π exp ( (F k D k ) 2 2σ 2 k ). (3) We can rewrite this equation as prob(d X, I ) exp ) ( χ2 2 with χ 2 = k ( ) Fk D 2 k = R 2 k σ k σ 2 k k. Residuals The R k = F k D k are referred to as residuals.
11 May 12, / 102 Least-squares estimate Take the logarithm of the likelihood function: Least-squares estimate L = ln (prob(d X, I )) = const χ2 2. (4) Since the maximum of the posterior will occur when χ 2 is smallest, the corresponding optimal solution X 0 is called least-squares estimate.
12 May 12, / 102 Parameter Estimation for ODE Models Consider a dynamical system with N state variables which we describe by a set of ordinary differential equations: d x(t) dt = f ( x(t), t, k), x(t 0 ) = x 0. (5) x(t): vector with all state variables k: vector with all parameters x 0 : vector for all initial expressions
13 Observables Often, the state variables cannot be directly observed. We specify an observation function g : R N R M which maps the state variables x to a set of M observables, y(t) = g( x(t), s) (6) We require both f ( ) and g( ) to be continuously differentiable functions with respect to their parameters. Note that we may be able to only partially observe the system such that M < N. The parameter vector p now comprises the kinetic parameters, k, the initial conditions x 0, and the parameters of the observation function, s, such that p = { x 0, k, s}. (7) Computational Biology Group (CoBI), D-BSSE, ETHZ Prof Dagmar Iber, PhD DPhil Lecture 10 BSc Biotechnology May 12, / 102
14 Maximum Likelihood The optimal parameter set is the one with the highest probability of observing the data and can be determined by maximizing the likelihood, prob( y p) of the data y ij with respect to the parameter set p prob( y p) = T M i=1 j=1 ( 1 exp 1 σ ij 2π 2 ) (g j ( x(t i, p), p)) y ij ) 2. σ 2 ij Computational Biology Group (CoBI), D-BSSE, ETHZ Prof Dagmar Iber, PhD DPhil Lecture 10 BSc Biotechnology May 12, / 102
15 May 12, / 102 Log-likelihood In practical terms, to find the maximum of the likelihood function the negative log-likelihood, L, is minimized L = log[prob( y p)] = T M i=1 j=1 1 2 R ij( p) 2 + c ij, R ij ( p) = g j( x(t i, p), p)) y ij σ ij, c ij = log[σ ij 2π]. R ij is called residual. The term c ij is independent of p, and can be left out of the minimization.
16 May 12, / 102 The maximum likelihood estimator The maximum likelihood estimator for the model parameters is thus given by log[l( y p)] T M i=1 j=1 1 2 R ij( p) 2. (8) In the background of independent Gaussian measurement errors the parameters p can therefore be determined by least squares minimization.
17 May 12, / 102 Optimization Algorithms 1 Local Methods Gradient-based Methods Newton-Raphson Iterative Algorithm Levenberg-Marquard Direct, derivative-free Methods Simplex Methods Nelder-Mead Method Conjugate Gradient Method 2 Global Methods Simulated Annealing Evolutionary Algorithms
18 May 12, / 102 Gradient of the weighted residuals R ij R ij ( p) = 1 dg j ( x(t i, p l ), p l )) p l σ ij dp l ( = 1 N g j dx n σ ij x n dp l + g ) j ti p l n=1 ti ti (9) g j x n and g j p l are the Jacobians of the differential equation system with respect to the state variables and with respect to the parameters. Sp n l = (dx n )/(dp l ) are the sensitivities of the state variables to changes in the parameter values that we discussed above.
19 May 12, / 102 Calculation of the Sensitivities The sensitivities can be computed by an integration of the sensitivity equations (as discussed above) in parallel with the ODE model. dsp n l = d dx n = d dx n dt dt dp l dp l dt = df (t, x(t), k) = dp l ( ) { Sp n dxq 1 : pl {x l (0) = (0) = 0 } dp l 0 : p l {s, k} N q=1 f n x q dx q dp l + f n p l
20 May 12, / 102 Workflow of gradient based minimization procedures Initialize model system and parameters LOOP Integrate ODE and sensitivity equations Calculate Jacobian of residuals Calculate residuals IF change in norm(residuals) < threshold BREAK ELSE Update parameter values using current parameter values and Jacobian ENDIF ENDLOOP Calculate fit statistics, parameter variances and confidence limits
21 Post-regression diagnostics Computational Biology Group (CoBI), D-BSSE, ETHZ Prof Dagmar Iber, PhD DPhil Lecture 10 BSc Biotechnology May 12, / 102
22 The Workflow Computational Biology Group (CoBI), D-BSSE, ETHZ Prof Dagmar Iber, PhD DPhil Lecture 10 BSc Biotechnology May 12, / 102
23 May 12, / 102 Post-regression diagnostics After fitting a regression model it is important to determine whether all the necessary model assumptions are valid before performing inference. In constructing our regression models we assumed that the errors were independent and normal random variables with mean 0 and constant variance σ 2 ij. Model diagnostic procedures involve both graphical methods and formal statistical tests. These procedures allow us to explore whether the assumptions of the regression model are valid and decide whether we can trust subsequent inference results.
24 Goodness of fit (GOF) Computational Biology Group (CoBI), D-BSSE, ETHZ Prof Dagmar Iber, PhD DPhil Lecture 10 BSc Biotechnology May 12, / 102
25 May 12, / 102 Goodness of fit We have so far assumed that the measurement error is Gaussian distributed. The weighted residuals, R ij ( p) = g j( x(t i, p), p)) y ij σ ij, (10) must then also be Gaussian distributed with unit variance. The sum of squared residuals follows a χ 2 distribution with T M degrees of freedom. T M i=1 j=1 R ij (p) 2 χ 2 df (p), df = T M. (11)
26 May 12, / 102 Chi-square distribution By definition, the Chi-square distribution is the distribution of the sum of the squared values of the observations drawn from the N(0, 1) distribution. It is denoted by the symbol χ 2, which is pronounced Ky square. More precisely, and more formally : Let {X 1, X 2,..., X n } be n independent random variables, all N(0, 1). Then the χ 2 n is defined as the distribution of the sum{x X X 2 n }. So there is not one χ 2 distribution, but a family of distributions, indexed by n. This parameter is called the number of degrees of freedom of the distribution. The Chi-square distribution with n degrees of freedom is therefore the distribution of the sum of n independent squared random variables all N(0, 1).
27 May 12, / 102 What does the χ 2 value tell us? T M i=1 j=1 R ij (p) 2 χ 2 df (p), df = T M Intuitively we expect from a good fit that the deviations of the model from the data should be of the same order as the measurement error, i.e. R ij 1, which means that the sum should be centered around T M. A much larger χ 2 value than T M indicates some variation in the data which is not accounted for by the model.
28 May 12, / 102 Goodness of fit (GOF) test T M i=1 j=1 R ij (p) 2 χ 2 df (p), df = T M. A goodness of fit (GOF) test calculates the probability of observing an as large or larger value than the value of χ 2 (p ) at the minimum. Note however that we have adjusted the parameters in order to minimize χ 2 df (p) and therefore the degrees of freedom in the GOF test are df = T M p. Usually a cut-off value such as Pr[χ 2 df (p )] < 0.05 is used to reject the fit.
29 The GOF test is strictly one sided: a much smaller value of χ 2 df (p ) than expected does not necessarily indicate an overfitting, which would be the result if the model would fit the particular realization of the measurement error. Computational Biology Group (CoBI), D-BSSE, ETHZ Prof Dagmar Iber, PhD DPhil Lecture 10 BSc Biotechnology May 12, / 102 Limitations T M i=1 j=1 R ij (p) 2 χ 2 df (p), df = T M. In some cases, an exceptionally large value of χ 2 df (p ), i.e., a small probability Pr[χ 2 df (p )], does not necessarily result from a bad model fit. It can also result from an underestimation of the measurements errors.
30 Example: TGF-β Model Computational Biology Group (CoBI), D-BSSE, ETHZ Prof Dagmar Iber, PhD DPhil Lecture 10 BSc Biotechnology May 12, / 102
31 May 12, / 102 Example: Time Courses Sensitivity analysis of the TGFβ signaling model. (A) Time course of the stimulation protocol and the system output, nuclear Smad2 /Smad4 complex. (B) Time resolved sensitivities of parameters controlling the dynamics of nuclear Smad2 /Smad4 complex and of the nuclear Smad2 /Smad4 complex. For details see main text. (C) Clustergram of the steady state control coefficients.
32 May 12, / 102 Performance of different optimization procedures. The table provides a comparison of the Matlab algorithm lsqnonlin to alternative minimizers in the Systems Biology Toolbox ( Av. comp. time (cputime 10 3 ) % correct matches Best perf. (χ 2 ) Max iter. NonLinLS tribes Gen. Alg Simplex Chc PSO Hill Cmaes Different optimizers were run on the same data set. The first column show the average computation time of each method. Every algorithm was run 5 times with different initial parameter values. The second column measures the number of times the algorithm produces an acceptable result (χ 2 < 30). The third column is the average χ 2 value at the optimum. The last column is the maximum number of iterations we allowed for each algorithm before it was stopped. Abbreviations: NonLinLS: Matlab lsqnonlin routine. simpex: Downhill Simplex Method in Multidimensions. cmaes: (15,50)-Evolution Strategy with Covariance Matrix Adaptation. chc: Clustering multi-start hill climbing. ga: Standard Genetic Algorithm with elitism. hill: Multi-start hill climbing. pso: Particle Swarm Optimization with constriction. tribes: adaptive Particle Swarm Optimization.
33 Uncertainty in the Parameters Computational Biology Group (CoBI), D-BSSE, ETHZ Prof Dagmar Iber, PhD DPhil Lecture 10 BSc Biotechnology May 12, / 102
34 May 12, / 102 Uncertainty in the Parameters Suppose we have estimated the parameter vector ˆp. How close is this estimate to the true parameter vector p? To this end we define the vector P = ˆp p to specify the difference.
35 May 12, / 102 Confidence Intervals There is no simple and generally valid way of calculating confidence limits of parameters for all problems faced in nonlinear optimization. However, in our context there is an approximate result which is valid in the limit of infinitely many data points and complete parameter identifiability. Specifically, one can relate the variance in the parameters to the curvature of the χ 2 df (p ) function at its minimum in order to derive parameter covariances and asymptotic confidence intervals.
36 May 12, / 102 Covariance Matrix We have previously noted that the covariance matrix Σ for our parameter estimate is related to the inverse of the Hessian of the likelihood function, We will now revisit this idea. Σ = [ L] 1. (12)
37 May 12, / 102 Maximum Likelihood Estimate The log-likelihood function L is given as L = ln (prob(d X, I )) = const χ2 2. (13) For easier readability, we will now write p = prob(d X, I )). (14) At the maximum, i.e. for the optimal parameter vector X, we have L = L ln (p) X = X = 1 p p X = 0. (15)
38 May 12, / 102 Maximum Likelihood Estimate By definition, we have pdd = 1 (16) Differentiating with respect to the parameter vector X and using we have L = 1 p pdd = p X = p p = 0. (17) p LdD = 0. (18)
39 May 12, / 102 Maximum Likelihood Estimate pdd = p LdD = 0. (19) Differentiating again with respect to the parameter vector X, we obtain pdd = 0 = p LdD + p( L) 2 dd + p LdD p LdD (20) We thus have E[( L) 2 ] = E[ L]. (21)
40 May 12, / 102 Fisher-Information Matrix E[( L) 2 ] = E[ L]. (22) For large E[ L] we have large curvature of the likelihood function at its optimum and thus tight confidence intervals and thus more information. Accordingly, we have Information Matrix is called the Information Matrix. FIM = E[ L] = E[( L) 2 ] (23)
41 May 12, / 102 The Cramer-Rao Bound The Cramer-Rao lower bound is a useful, maybe the best statistical indicator for the errors made in estimating the true parameter values. The Cramer-Rao Bound The so-called Cramer-Rao inequality provides a lower bound to the variance of an unbiased estimator, as will be seen in the sequel.
42 May 12, / 102 The Cramer-Rao Bound Let X e (D) be any estimator of the true parameter value X based on the measurements D. We will now drop the vector arrows for better readability. Let X e (D) = E[X e (D)] be the expectation of the estimate. Its variance is given by σ 2 Xe = E[X e(d) X e (D)) 2 ]. (24)
43 May 12, / 102 Bias in an estimator The bias in an estimator is defined as E[X e (D) X ] = X e (D)p(D X, I )dd X = x(x ). (25) If x(x ) = 0, then it is called an unbiased estimator because then on average the expected value of the estimate is the same as the true parameter. Bias in general cannot be determined since it depends on the true value of the parameter, which in practice is unknown. Often the estimates would be biased, if the noise were not zero mean.
44 May 12, / 102 Bias in an estimator We now have X e (D)p(D X, I )dd = X + x(x ). (26) Differentiating on both sides with respect to (w.r.t.) X we get X e (D) p(d X, I )dd = since X e (D) is a function of the data D only. X e (D)( L)pdD = 1 + x(x ). (27)
45 May 12, / 102 Bias in an estimator Multiplying pdd = with X e (D) and adding it to yields X e (D) p(d X, I )dd = p LdD = 0. (28) X e (D)( L)pdD = 1 + x(x ). (29) (X e (D) X e (D))( L)pdD = 1 + x(x ) (X e (D) X e (D)) p( L) pdd = 1 + x(x ). (30)
46 Cauchy-Schwarz inequality Now we are ready to apply the well-known Cauchy-Schwarz inequality [ 2 f (z)g(z)dz] Equality applies if f (z) = kg(z), for a constant k. You may know the form for vectors: Cauchy-Schwarz inequality f (z) 2 dz g(z) 2 dz. (31) The Cauchy-Schwarz inequality states that for all vectors x and y of an inner product space it is true that x, y 2 x, x y, y, (32) where, is the inner product also known as dot product. Computational Biology Group (CoBI), D-BSSE, ETHZ Prof Dagmar Iber, PhD DPhil Lecture 10 BSc Biotechnology May 12, / 102
47 May 12, / 102 Cauchy-Schwarz inequality We thus have (X e (D) X e (D)) p( L) pdd = 1 + x(x ). (33) (1 + x(x )) 2 = ( (X e (D) X e (D)) p( L) ) 2 pdd (X e (D) X e (D)) 2 pdd } {{ } σxe 2 ( L) 2 pdd } {{ } FIM
48 May 12, / 102 Cramer-Rao Inequality σxe 2 (1 + x(x ))2 FIM 1 (34) For an unbiased estimator, x(x ) = 0, and hence σxe 2 FIM 1. (35) The equality sign holds if X e (D) X e (D) = k L. (36) For an unbiased, efficient estimator, we thus have Cramer-Rao Bound σ 2 Xe = FIM 1 = E[ L] 1 = E[( L) 2 ] 1. (37)
49 May 12, / 102 The MLE is efficient For efficiency we have to show that At the optimum we have X e (D) X e (D) = k L. (38) L = 0. (39) In case of an unbiased estimate, x(x ) = 0, we also have at the optimum X e (D) X e (D) = 0. (40) Hence, the equality is established and the ML estimator is proved efficient. This is a very important property of the ML estimator.
50 May 12, / 102 Fisher-Information Matrix The log-likelihood function L is given as L = ln (p(d, X )). (41) Given a probability density function p(d, X ), the Fisher Information Matrix is thus defined as [ ] [ ] log p log p FIM = E = E 2 log p Σ 1. (42) X i X j X i X j It is not a coincidence that the Fisher Information Matrix appears to be the reciprocal of the accuracy with which we can expect to be able to estimate X given an observation data D. As the variance decreases, the amount of information increases.
51 Determination of FIM Recall that for parameters X L = log (p) = T M i=1 j=1 1 2 R ij( X ) 2 + c ij = const χ2, such that with R ij ( X ) = g j( x(t i, X ), X )) y ij σ ij, c ij = log[σ ij 2π]. log (p) X l = χ χ X l = T M i=1 j=1 R ij ( X ) = 1 dg j ( x(t i, X l ), X l )) = 1 X l σ ij dx l σ ij R ij ( X ) R ij X l ( N g j x n n=1 ti dx n dx l + g ) j ti X l Computational Biology Group (CoBI), D-BSSE, ETHZ Prof Dagmar Iber, PhD DPhil Lecture 10 BSc Biotechnology May 12, / 102 ti
52 May 12, / 102 Determination of FIM with log (p) = T i=1 j=1 R ij ( X ) = 1 dg j ( x(t i, X l ), X l )) = 1 X l σ ij dx l σ ij M R ij ( X ) R ij ( X ) ( N g j x n n=1 ti dx n dx l + g ) j ti X l ti We can further approximate ( L) kl = (log (p)) kl T M i=1 j=1 where we neglected the second order derivative. R ij X k T M i=1 j=1 R ij X l
53 May 12, / 102 Fisher Information Matrix (FIM) The Fisher Information Matrix can then be calculated from the residuals during minimization. [ ] FIM ij = E 2 log p S T S (43) X i X j The approximation neglects the second derivative terms but is computationally inexpensive as S l = ij ( R ij/ X l ) is the calculated gradient matrix of the residuals during minimization.
54 May 12, / 102 Confidence Intervals Asymptotic confidence intervals can be calculated by taking into account the distribution of the χ 2 values, which are approximately Gaussian for large degrees of freedom. The 95% confidence intervals given by p ± 1.96 diag(c). (44) Note that this asymptotic result is just a lower bound on the uncertainty of the parameter estimate (the Cramer-Rao bound).
55 May 12, / 102 Observed Fisher-Information Matrix In the following we will approximate [ log p FIM = E X i ] log p S T S. (45) X j Strictly, this definition corresponds to the observed Fisher information. If no expectation is taken we obtain a data-dependent quantity that is called the observed Fisher information. Based on the sensitivity matrix S or rather on the Fisher information matrix (FIM), there are a number of easy-to-compute indicators.
56 The Quality of the Parameter Estimate Computational Biology Group (CoBI), D-BSSE, ETHZ Prof Dagmar Iber, PhD DPhil Lecture 10 BSc Biotechnology May 12, / 102
57 May 12, / 102 Confidence intervals Assuming that all other parameters are exact, a confidence interval for a specific parameter is the intersection of the ellipsoidal region with the parameter axis. This is the dependent confidence interval: D p i = C(α)/ [(S T S) ii ] 1 (46). The independent confidence interval is given by the projection of the ellipsoidal region onto the parameter axis: I p i = C(α)/ ([S T S] 1 ) ii (47).
58 May 12, / 102 Dependent & Independent Confidence Intervals If dependent and independent confidence intervals are similar and small, ˆp i is well-determined. In case of a strong correlation between parameters, the dependent confidence intervals underestimate the confidence region, whereas the independent confidence intervals overestimate it.
59 May 12, / 102 Correlation and Co-Variance Another way to obtain information about the correlations between parameters is to look at the covariance matrix cov = (S T S) 1. The correlation coefficient of the ith and jth parameter is given by:. cor ij = cov ij cov ii cov jj (48)
60 May 12, / 102 Interpretation in terms of eigenvalues & eigenvectors Using the singular value decomposition for S, S = UΦV T, (49) where U is an an unitary matrix (U T U = UU T = I ) and V T is the conjugate transpose of the unitary matrix V, we get S T S = V (ˆp)Φ T U T UΦV (ˆp) T = V (ˆp)Φ T ΦV (ˆp) T (50) where the eigenvectors of S T S are the columns of the matrix of V (ˆp).
61 May 12, / 102 Interpretation in terms of eigenvalues & eigenvectors So, the principal axes of the ellipsoidal confidence region are given by the singular vectors, the column vectors of the matrix V (ˆp), and the length of the principal axes is proportional to the reciprocal of the corresponding singular values, the diagonal elements of Φ(ˆp).
62 May 12, / 102 Interpretation in terms of eigenvalues & eigenvectors Using the transformation (rotation): the equation for the ellipsoid can be rewritten as: z = V T (ˆp)P (51) m σi 2 zi 2 C(α) (52) i=1 Note that C(α) is approximately proportional to the variance in the measurement errors.
63 May 12, / 102 Practical identifiable The precise definition of practical identifiable depends on the level of accuracy, r e, one requires for the parameter estimates. This defines the sphere: m zi 2 = re 2 (53) i=1 To be able to determine z i accurately enough, the radius along the ellipsoid s ith principal axis should not exceed the radius of the sphere, which leads to the following inequality: σ i C(α) r e (54)
64 May 12, / 102 Practical identifiable Suppose that only the first k largest singular values satisfy C(α) σ i r e, then only the first k entries of z are estimated with the required accuracy. If a principal axis of the ellipsoid makes a significant angle with the axis in parameter space (i.e. there exists more than one significant entry in the eigenvector), this corresponds to the presence of correlation among parameters in ˆp. In this case, only a combination of parameters can be determined.
65 May 12, / 102 Practical identifiable To summarize, the level of noise in the data, in combination with the accuracy requirement for the parameter estimates, defines the threshold for significant singular values in the matrix Φ. The number of singular values exceeding this threshold determines the number of parameter relations that can be derived from the experiment. How these relations relate to the individual parameters is described by the corresponding columns in the matrix V.
66 How to improve your estimates? m σi 2 zi 2 C(α) (55) i=1 indicates that having, for example, two times more accurate data so that the standard deviation σ is halved will decrease the radii along the ellipsoid s principal axes by a factor of 2. Therefore, in case of very small singular values φ i (i.e. strongly elongated ellipsoids), more accurate data obtained by the experimentalist will not improve much the quality of the corresponding parameter estimates. In such a case, one certainly needs additional measurements of a different type (e.g. different components, different time points, or in the case of partial differential equations, different spatial points). Computational Biology Group (CoBI), D-BSSE, ETHZ Prof Dagmar Iber, PhD DPhil Lecture 10 BSc Biotechnology May 12, / 102
67 May 12, / 102 Parameter Correlation and Identifiability Frequently, the optimization procedure does not yield a unique optimal parameter set, because there is no unique optimal χ 2 (p ) value given the available data. In this case the value of some or all parameters are non-identifiable. Non-identifiability is the result of a non-unique χ 2 minimum, which can be caused, e.g., by a very flat χ 2 landscape. The latter implies a functional relationship between parameters along which the χ 2 value is unaltered. As a result parameter estimates are highly correlated. There are three common ways to deal with non-identifiability.
68 May 12, / 102 Addressing Parameter Correlations 1 Fix some of the non-identifiable parameter at educated values and only estimate the remaining parameters. These estimates are of course biased since their optimum is in a functional relation to the fixed parameters. 2 Subsequent analyses can be based on all admissible parameter sets and the parameter sets can then be clustered according to the predictions derived from them. 3 Reduce the model such that it does not contain the non-identifiable parameters, e.g. by course graining the model.
69 May 12, / 102 Non-Identifiability & Data Quality It is noteworthy that non-identifiability of parameters does not imply a poor fit to the data, but that parameter values cannot be constrained to a unique value. The predictive power of the model will therefore be limited to model predictions that are not sensitive to non-identifiable parameters.
70 May 12, / 102 Bootstrap Methods In many real world settings the uncertainty in parameter estimates is larger due to a limited amount of data. Here, an alternative but computationally more expensive way to determine parameter uncertainties, called bootstrap, is more appropriate. Bootstrap methods construct the empirical distribution of the parameter estimates by an repeated data resampling and consecutive parameter estimation. In this way, parameter uncertainties can be inferred from the shape of the empirical parameter distributions.
71 Example: TGF-β Model Computational Biology Group (CoBI), D-BSSE, ETHZ Prof Dagmar Iber, PhD DPhil Lecture 10 BSc Biotechnology May 12, / 102
72 May 12, / 102 Example: Time Courses Sensitivity analysis of the TGFβ signaling model. (A) Time course of the stimulation protocol and the system output, nuclear Smad2 /Smad4 complex. (B) Time resolved sensitivities of parameters controlling the dynamics of nuclear Smad2 /Smad4 complex and of the nuclear Smad2 /Smad4 complex. For details see main text. (C) Clustergram of the steady state control coefficients.
73 Example: Sensitivities & Correlations Computational Biology Group (CoBI), D-BSSE, ETHZ Prof Dagmar Iber, PhD DPhil Lecture 10 BSc Biotechnology May 12, / 102
74 Example: Sensitivities & Correlations Computational Biology Group (CoBI), D-BSSE, ETHZ Prof Dagmar Iber, PhD DPhil Lecture 10 BSc Biotechnology May 12, / 102
75 Model Selection Computational Biology Group (CoBI), D-BSSE, ETHZ Prof Dagmar Iber, PhD DPhil Lecture 10 BSc Biotechnology May 12, / 102
76 May 12, / 102 Bayesian Information Criterion (BIC) In case of large datasets, the Bayesian Information Criterion (BIC) is more appropriate than test-based compare-and-rank models (CRM). This method assigns a score based on its likelihood, the number m of estimated free parameters it it and the number n of fitted data points. The BIC for a model M is given by 2 ln p(d M) BIC = 2 ln ˆL + m (ln(n) ln(2π)). (56) where ˆL is the maximised value of the likelihood. For large n, this can be approximated by: BIC = 2 ln ˆL + m ln(n). (57)
77 May 12, / 102 Bayesian Information Criterion (BIC) BIC = 2 ln ˆL + m ln(n). (58) In the case of normally distributed measurements χ 2 min = 2 ln ˆL + C (59) for some constant C, which does not vary between candidate models but depends only upon the data points. The BIC decreases as the likelihood increases, and increases as the number of parameters increases. Among competing models, the model that minimises the BIC is the most suitable to describe the available data.
78 May 12, / 102 Bayesian Information Criterion (BIC) BIC = 2 ln ˆL + m ln(n). (60) Because the first term grows linearly with the number n of fitted data points, while the second term is proportional to ln n, the penalty for having too many parameters is diminished as the data set gets larger.
79 Incorporation of prior information Computational Biology Group (CoBI), D-BSSE, ETHZ Prof Dagmar Iber, PhD DPhil Lecture 10 BSc Biotechnology May 12, / 102
80 May 12, / 102 Incorporation of prior information In most practical cases we have some prior knowledge about plausible parameter ranges. In case of physical constants we know that the parameters are constant, and in case of biochemical binding and reaction rates we know plausible ranges. Further experiments may further restrict these bounds. We will now discuss how such information can be incorporated in the parameter estimation process.
81 May 12, / 102 Incorporation of prior information Recall that prob(x D, I ) = prob(d X, I ) prob(x I ) prob(d I ) (61) So far we have assumed that the prior is constant, i.e. prob(x I ) =const. We will now assume that we know a range in which the parameter value must lie.
82 May 12, / 102 Incorporation of prior information If we already knew that X j = x 0j ± ɛ j, for example, then the assignment of an uncorrelated Gaussian pdf for the prior of X yields [ ] M 1 prob(x I ) = exp (X j x 0j ) 2 ( ɛ j 2π 2ɛ 2 = exp C ) (62) j 2 where j=1 C = M ( ) Xj x 2 0j. (63) ɛ j j=1 For the logarithm of the posterior pdf we then have L = ln [prob(x D, I )] = constant 1 2 [χ2 + C]. (64)
83 Error Propagation Computational Biology Group (CoBI), D-BSSE, ETHZ Prof Dagmar Iber, PhD DPhil Lecture 10 BSc Biotechnology May 12, / 102
84 May 12, / 102 Error Propagation - changing variables The Problem: Suppose that we have the probability distribution function (pdf) for X. What is the pdf for Y if Y = f (X )?
85 May 12, / 102 Error Propagation - changing variables Imagine taking a very small interval δx about some arbitrary point X = X ; the probability that X lies in the range X δx /2 to X + δx /2 is given by ( prob X δx 2 X X + δx 2 ) I prob(x = X I )δx (65) where the equality becomes exact in the limit δx 0.
86 May 12, / 102 Error Propagation - changing variables Now Y = f (X ) will map the point X = X to Y = f (X ) and the interval δx to δy. prob(x = X I )δx = prob(y = Y I )δy. (66) As this must be true for any point in X -space, in the limit of infinitesimally small intervals, we obtain the relationship prob(x I ) = prob(y I ) dy dx. (67) The term dy = df is the Jacobian. dx dx
87 May 12, / 102 Error Propagation in case of several variables If we want to write the pdf for M parameters {X j } in terms of the same quantities {Y j }, then we require prob({x j } I )δx 1 δx 2 δx M = prob({y j } I )δ M Vol({Y j }). (68) where δ M Vol({Y j }) is the M-dimensional volume in Y -space mapped out by the small hypercube region δx 1 δx 2 δx M in X -space. δ M Vol({Y j }) = (Y 1, Y 2,, Y M ) (X 1, X 2,, X M ) δx 1δX 2 δx M (69) where the quantity in the modulus sign is the multivariate Jacobian.
88 May 12, / 102 Error Propagation - several variables Error Propagation - several variables prob({x j } I ) = prob({y j } I ) d(y 1, Y 2,, Y M ) d(x 1, X 2,, X M )
89 May 12, / 102 Error Propagation - an example Consider the transformation of a pdf defined on a two-dimensional Cartesian grid (x, y) to its equivalent form in polar coordinates (R, θ). We have For the determinant of the Jacobian x = R cos θ y = R sin θ (70) (x, y) (R, θ) = cos θ sin θ R sin θ R cos θ = R(cos2 θ + sin 2 θ) = R. (71) Therefore: prob(r, θ I ) = prob(x, y I ) R. (72)
90 May 12, / 102 Error Propagation - an example Thus, if the pdf for x and y was an isotropic, bivariate Gaussian prob(x, y I ) = 1 [ 2πσ 2 exp (x 2 + y 2 ] ) 2σ 2 (73) then the corresponding pdf for R and θ would take the form prob(r, θ I ) = R ] [ 2πσ 2 exp R2 2σ 2. (74)
91 May 12, / 102 Error Propagation - an example Finally, we determine the pdf for the radius R by marginalizing the joint pdf prob(r, θ I ) over θ. prob(r I ) = 2π 0 prob(r, θ I )dθ = R [ ] σ 2 exp R2 2σ 2. (75)
92 May 12, / 102 Error Propagation Consider Z = f (X, Y ) = X + Y : prob(z I ) = = = prob(z X, Y, I )prob(x, Y I )dxdy δ(z f (X, Y ))prob(x, Y I )dxdy δ(z (X + Y ))prob(x, Y I )dxdy. (76) where the Dirac δ-function in the second line is zero unless Z = f (X, Y ).
93 May 12, / 102 Error Propagation Assume further X = x 0 ± σ x, Y = y 0 ± σ y, and that these parameters are uncorrelated. Then: prob(z I ) = dx prob(x I ) δ(z (X + Y ))prob(y I )dy. Since the Dirac δ-function is infinitely sharp (but has unit area), the Y -integrand is zero unless Y = Z X : prob(z I ) = prob(x I )prob(y = Z X I )dx.
94 May 12, / 102 Error Propagation prob(z I ) = prob(x I )prob(y = Z X I )dx. Since X = x 0 ± σ x, Y = y 0 ± σ y use Gaussian pdfs prob(z I ) = 1 2πσ x σ y exp ( (X x 0) 2 ) exp 2σ 2 x ( (Z X y 0) 2 2σ 2 y ) dx.
95 May 12, / 102 Error Propagation prob(z I ) = 1 2πσ x σ y exp ( (X x 0) 2 ) exp 2σ 2 x ( (Z X y 0) 2 2σ 2 y ) dx. After some tedious algebra prob(z I ) = 1 exp ( (Z z 0) 2 ) 2πσz 2σ 2 z where z 0 = x 0 + y 0 and σ 2 z = σ 2 x + σ 2 y.
96 May 12, / 102 Error Propagation prob(z I ) = 1 exp ( (Z z 0) 2 ) 2πσz 2σ 2 z where z 0 = x 0 + y 0 and σ 2 z = σ 2 x + σ 2 y. The pdf for Z = X Y is the same, except that z 0 = x 0 y 0.
97 May 12, / 102 A more intuitive approach Suppose we perturb Z = X Y. Intuitively we might have guessed, that the best estimate for the difference of the two parameters would be x 0 y 0. Let s now have a look at the corresponding error-bar. Z Z 0 = (X X 0 ) (Y Y 0 ) δz = δx δy Recall: σx 2 =< (X X 0) 2 >= (X X 0 ) 2 prob(x, Y {data}, I ) dxdy Thus: < δx 2 >= σx 2, < δy 2 >= σy 2, < δx δy >= 0
98 May 12, / 102 A more intuitive approach Since we have < δz 2 > = < (δx δy ) 2 > = < δ 2 X + δy 2 2δX δy > = < δ 2 X > + < δy 2 > 2 < δx δy > < δx 2 >= σx 2, < δy 2 >= σy 2, < δx δy >= 0 σ Z = < δz 2 > = σ 2 X + σ2 Y
99 May 12, / 102 A more intuitive approach Let s next consider Z = X /Y. Using the quotient rule of differentiation we have This can be rewritten as δz = Y δx X δy Y 2 δz Z = δx X δy Y
100 A more intuitive approach Squaring both sides and taking expectation values, we obtain < δz 2 > z 2 0 = < δx 2 > x < δy 2 > y 2 0 < δx δy > x 0 y 0. Here the X, Y, Z in the denominators were replaced by x 0, y 0, z 0 = x 0 y 0 because we are interested in deviations from the optimal solution. Thus finally, σ z z 0 = σ 2 x x σ2 Y y0 2. Computational Biology Group (CoBI), D-BSSE, ETHZ Prof Dagmar Iber, PhD DPhil Lecture 10 BSc Biotechnology May 12, / 102
101 May 12, / 102 WARNING This intuitive approach may no longer work if the prior cuts the posterior pdf, i.e. because the parameter values must be positive!!
102 May 12, / 102 Thanks!! Thanks for your attention! Slides for this talk will be available at:
Lecture 10: Parameter Estimation for ODE Models
 Computational Biology Group (CoBI), D-BSSE, ETHZ Lecture 10: Parameter Estimation for ODE Models Prof Dagmar Iber, PhD DPhil BSc Biotechnology 2016/17 December 7, 2016 2 / 93 Contents 1 Review of Previous
Computational Biology Group (CoBI), D-BSSE, ETHZ Lecture 10: Parameter Estimation for ODE Models Prof Dagmar Iber, PhD DPhil BSc Biotechnology 2016/17 December 7, 2016 2 / 93 Contents 1 Review of Previous
Physics 403. Segev BenZvi. Propagation of Uncertainties. Department of Physics and Astronomy University of Rochester
 Physics 403 Propagation of Uncertainties Segev BenZvi Department of Physics and Astronomy University of Rochester Table of Contents 1 Maximum Likelihood and Minimum Least Squares Uncertainty Intervals
Physics 403 Propagation of Uncertainties Segev BenZvi Department of Physics and Astronomy University of Rochester Table of Contents 1 Maximum Likelihood and Minimum Least Squares Uncertainty Intervals
Lecture 4: Types of errors. Bayesian regression models. Logistic regression
 Lecture 4: Types of errors. Bayesian regression models. Logistic regression A Bayesian interpretation of regularization Bayesian vs maximum likelihood fitting more generally COMP-652 and ECSE-68, Lecture
Lecture 4: Types of errors. Bayesian regression models. Logistic regression A Bayesian interpretation of regularization Bayesian vs maximum likelihood fitting more generally COMP-652 and ECSE-68, Lecture
Econometrics I, Estimation
 Econometrics I, Estimation Department of Economics Stanford University September, 2008 Part I Parameter, Estimator, Estimate A parametric is a feature of the population. An estimator is a function of the
Econometrics I, Estimation Department of Economics Stanford University September, 2008 Part I Parameter, Estimator, Estimate A parametric is a feature of the population. An estimator is a function of the
Parameter estimation! and! forecasting! Cristiano Porciani! AIfA, Uni-Bonn!
 Parameter estimation! and! forecasting! Cristiano Porciani! AIfA, Uni-Bonn! Questions?! C. Porciani! Estimation & forecasting! 2! Cosmological parameters! A branch of modern cosmological research focuses
Parameter estimation! and! forecasting! Cristiano Porciani! AIfA, Uni-Bonn! Questions?! C. Porciani! Estimation & forecasting! 2! Cosmological parameters! A branch of modern cosmological research focuses
Physics 403. Segev BenZvi. Parameter Estimation, Correlations, and Error Bars. Department of Physics and Astronomy University of Rochester
 Physics 403 Parameter Estimation, Correlations, and Error Bars Segev BenZvi Department of Physics and Astronomy University of Rochester Table of Contents 1 Review of Last Class Best Estimates and Reliability
Physics 403 Parameter Estimation, Correlations, and Error Bars Segev BenZvi Department of Physics and Astronomy University of Rochester Table of Contents 1 Review of Last Class Best Estimates and Reliability
Introduction to Maximum Likelihood Estimation
 Introduction to Maximum Likelihood Estimation Eric Zivot July 26, 2012 The Likelihood Function Let 1 be an iid sample with pdf ( ; ) where is a ( 1) vector of parameters that characterize ( ; ) Example:
Introduction to Maximum Likelihood Estimation Eric Zivot July 26, 2012 The Likelihood Function Let 1 be an iid sample with pdf ( ; ) where is a ( 1) vector of parameters that characterize ( ; ) Example:
Statistics. Lent Term 2015 Prof. Mark Thomson. 2: The Gaussian Limit
 Statistics Lent Term 2015 Prof. Mark Thomson Lecture 2 : The Gaussian Limit Prof. M.A. Thomson Lent Term 2015 29 Lecture Lecture Lecture Lecture 1: Back to basics Introduction, Probability distribution
Statistics Lent Term 2015 Prof. Mark Thomson Lecture 2 : The Gaussian Limit Prof. M.A. Thomson Lent Term 2015 29 Lecture Lecture Lecture Lecture 1: Back to basics Introduction, Probability distribution
Statistics and Data Analysis
 Statistics and Data Analysis The Crash Course Physics 226, Fall 2013 "There are three kinds of lies: lies, damned lies, and statistics. Mark Twain, allegedly after Benjamin Disraeli Statistics and Data
Statistics and Data Analysis The Crash Course Physics 226, Fall 2013 "There are three kinds of lies: lies, damned lies, and statistics. Mark Twain, allegedly after Benjamin Disraeli Statistics and Data
Multivariate Statistics
 Multivariate Statistics Chapter 2: Multivariate distributions and inference Pedro Galeano Departamento de Estadística Universidad Carlos III de Madrid pedro.galeano@uc3m.es Course 2016/2017 Master in Mathematical
Multivariate Statistics Chapter 2: Multivariate distributions and inference Pedro Galeano Departamento de Estadística Universidad Carlos III de Madrid pedro.galeano@uc3m.es Course 2016/2017 Master in Mathematical
Fall 2017 STAT 532 Homework Peter Hoff. 1. Let P be a probability measure on a collection of sets A.
 1. Let P be a probability measure on a collection of sets A. (a) For each n N, let H n be a set in A such that H n H n+1. Show that P (H n ) monotonically converges to P ( k=1 H k) as n. (b) For each n
1. Let P be a probability measure on a collection of sets A. (a) For each n N, let H n be a set in A such that H n H n+1. Show that P (H n ) monotonically converges to P ( k=1 H k) as n. (b) For each n
Physics 403. Segev BenZvi. Numerical Methods, Maximum Likelihood, and Least Squares. Department of Physics and Astronomy University of Rochester
 Physics 403 Numerical Methods, Maximum Likelihood, and Least Squares Segev BenZvi Department of Physics and Astronomy University of Rochester Table of Contents 1 Review of Last Class Quadratic Approximation
Physics 403 Numerical Methods, Maximum Likelihood, and Least Squares Segev BenZvi Department of Physics and Astronomy University of Rochester Table of Contents 1 Review of Last Class Quadratic Approximation
[y i α βx i ] 2 (2) Q = i=1
![[y i α βx i ] 2 (2) Q = i=1 [y i α βx i ] 2 (2) Q = i=1](/thumbs/75/72005858.jpg) Least squares fits This section has no probability in it. There are no random variables. We are given n points (x i, y i ) and want to find the equation of the line that best fits them. We take the equation
Least squares fits This section has no probability in it. There are no random variables. We are given n points (x i, y i ) and want to find the equation of the line that best fits them. We take the equation
Lecture 5: Travelling Waves
 Computational Biology Group (CoBI), D-BSSE, ETHZ Lecture 5: Travelling Waves Prof Dagmar Iber, PhD DPhil MSc Computational Biology 2015 26. Oktober 2016 2 / 68 Contents 1 Introduction to Travelling Waves
Computational Biology Group (CoBI), D-BSSE, ETHZ Lecture 5: Travelling Waves Prof Dagmar Iber, PhD DPhil MSc Computational Biology 2015 26. Oktober 2016 2 / 68 Contents 1 Introduction to Travelling Waves
Gaussian Processes. Le Song. Machine Learning II: Advanced Topics CSE 8803ML, Spring 2012
 Gaussian Processes Le Song Machine Learning II: Advanced Topics CSE 8803ML, Spring 01 Pictorial view of embedding distribution Transform the entire distribution to expected features Feature space Feature
Gaussian Processes Le Song Machine Learning II: Advanced Topics CSE 8803ML, Spring 01 Pictorial view of embedding distribution Transform the entire distribution to expected features Feature space Feature
1 Data Arrays and Decompositions
 1 Data Arrays and Decompositions 1.1 Variance Matrices and Eigenstructure Consider a p p positive definite and symmetric matrix V - a model parameter or a sample variance matrix. The eigenstructure is
1 Data Arrays and Decompositions 1.1 Variance Matrices and Eigenstructure Consider a p p positive definite and symmetric matrix V - a model parameter or a sample variance matrix. The eigenstructure is
Stat 5101 Lecture Notes
 Stat 5101 Lecture Notes Charles J. Geyer Copyright 1998, 1999, 2000, 2001 by Charles J. Geyer May 7, 2001 ii Stat 5101 (Geyer) Course Notes Contents 1 Random Variables and Change of Variables 1 1.1 Random
Stat 5101 Lecture Notes Charles J. Geyer Copyright 1998, 1999, 2000, 2001 by Charles J. Geyer May 7, 2001 ii Stat 5101 (Geyer) Course Notes Contents 1 Random Variables and Change of Variables 1 1.1 Random
University of Cambridge Engineering Part IIB Module 3F3: Signal and Pattern Processing Handout 2:. The Multivariate Gaussian & Decision Boundaries
 University of Cambridge Engineering Part IIB Module 3F3: Signal and Pattern Processing Handout :. The Multivariate Gaussian & Decision Boundaries..15.1.5 1 8 6 6 8 1 Mark Gales mjfg@eng.cam.ac.uk Lent
University of Cambridge Engineering Part IIB Module 3F3: Signal and Pattern Processing Handout :. The Multivariate Gaussian & Decision Boundaries..15.1.5 1 8 6 6 8 1 Mark Gales mjfg@eng.cam.ac.uk Lent
Multiple Random Variables
 Multiple Random Variables Joint Probability Density Let X and Y be two random variables. Their joint distribution function is F ( XY x, y) P X x Y y. F XY ( ) 1, < x
Multiple Random Variables Joint Probability Density Let X and Y be two random variables. Their joint distribution function is F ( XY x, y) P X x Y y. F XY ( ) 1, < x
Tutorial: Statistical distance and Fisher information
 Tutorial: Statistical distance and Fisher information Pieter Kok Department of Materials, Oxford University, Parks Road, Oxford OX1 3PH, UK Statistical distance We wish to construct a space of probability
Tutorial: Statistical distance and Fisher information Pieter Kok Department of Materials, Oxford University, Parks Road, Oxford OX1 3PH, UK Statistical distance We wish to construct a space of probability
MACHINE LEARNING ADVANCED MACHINE LEARNING
 MACHINE LEARNING ADVANCED MACHINE LEARNING Recap of Important Notions on Estimation of Probability Density Functions 2 2 MACHINE LEARNING Overview Definition pdf Definition joint, condition, marginal,
MACHINE LEARNING ADVANCED MACHINE LEARNING Recap of Important Notions on Estimation of Probability Density Functions 2 2 MACHINE LEARNING Overview Definition pdf Definition joint, condition, marginal,
STAT 730 Chapter 4: Estimation
 STAT 730 Chapter 4: Estimation Timothy Hanson Department of Statistics, University of South Carolina Stat 730: Multivariate Analysis 1 / 23 The likelihood We have iid data, at least initially. Each datum
STAT 730 Chapter 4: Estimation Timothy Hanson Department of Statistics, University of South Carolina Stat 730: Multivariate Analysis 1 / 23 The likelihood We have iid data, at least initially. Each datum
The Expectation-Maximization Algorithm
 1/29 EM & Latent Variable Models Gaussian Mixture Models EM Theory The Expectation-Maximization Algorithm Mihaela van der Schaar Department of Engineering Science University of Oxford MLE for Latent Variable
1/29 EM & Latent Variable Models Gaussian Mixture Models EM Theory The Expectation-Maximization Algorithm Mihaela van der Schaar Department of Engineering Science University of Oxford MLE for Latent Variable
01 Probability Theory and Statistics Review
 NAVARCH/EECS 568, ROB 530 - Winter 2018 01 Probability Theory and Statistics Review Maani Ghaffari January 08, 2018 Last Time: Bayes Filters Given: Stream of observations z 1:t and action data u 1:t Sensor/measurement
NAVARCH/EECS 568, ROB 530 - Winter 2018 01 Probability Theory and Statistics Review Maani Ghaffari January 08, 2018 Last Time: Bayes Filters Given: Stream of observations z 1:t and action data u 1:t Sensor/measurement
Density Estimation. Seungjin Choi
 Density Estimation Seungjin Choi Department of Computer Science and Engineering Pohang University of Science and Technology 77 Cheongam-ro, Nam-gu, Pohang 37673, Korea seungjin@postech.ac.kr http://mlg.postech.ac.kr/
Density Estimation Seungjin Choi Department of Computer Science and Engineering Pohang University of Science and Technology 77 Cheongam-ro, Nam-gu, Pohang 37673, Korea seungjin@postech.ac.kr http://mlg.postech.ac.kr/
POLI 8501 Introduction to Maximum Likelihood Estimation
 POLI 8501 Introduction to Maximum Likelihood Estimation Maximum Likelihood Intuition Consider a model that looks like this: Y i N(µ, σ 2 ) So: E(Y ) = µ V ar(y ) = σ 2 Suppose you have some data on Y,
POLI 8501 Introduction to Maximum Likelihood Estimation Maximum Likelihood Intuition Consider a model that looks like this: Y i N(µ, σ 2 ) So: E(Y ) = µ V ar(y ) = σ 2 Suppose you have some data on Y,
Estimation Theory. as Θ = (Θ 1,Θ 2,...,Θ m ) T. An estimator
 Estimation Theory Estimation theory deals with finding numerical values of interesting parameters from given set of data. We start with formulating a family of models that could describe how the data were
Estimation Theory Estimation theory deals with finding numerical values of interesting parameters from given set of data. We start with formulating a family of models that could describe how the data were
2. What are the tradeoffs among different measures of error (e.g. probability of false alarm, probability of miss, etc.)?
 ECE 830 / CS 76 Spring 06 Instructors: R. Willett & R. Nowak Lecture 3: Likelihood ratio tests, Neyman-Pearson detectors, ROC curves, and sufficient statistics Executive summary In the last lecture we
ECE 830 / CS 76 Spring 06 Instructors: R. Willett & R. Nowak Lecture 3: Likelihood ratio tests, Neyman-Pearson detectors, ROC curves, and sufficient statistics Executive summary In the last lecture we
APPENDIX A. Background Mathematics. A.1 Linear Algebra. Vector algebra. Let x denote the n-dimensional column vector with components x 1 x 2.
 APPENDIX A Background Mathematics A. Linear Algebra A.. Vector algebra Let x denote the n-dimensional column vector with components 0 x x 2 B C @. A x n Definition 6 (scalar product). The scalar product
APPENDIX A Background Mathematics A. Linear Algebra A.. Vector algebra Let x denote the n-dimensional column vector with components 0 x x 2 B C @. A x n Definition 6 (scalar product). The scalar product
σ(a) = a N (x; 0, 1 2 ) dx. σ(a) = Φ(a) =
 Until now we have always worked with likelihoods and prior distributions that were conjugate to each other, allowing the computation of the posterior distribution to be done in closed form. Unfortunately,
Until now we have always worked with likelihoods and prior distributions that were conjugate to each other, allowing the computation of the posterior distribution to be done in closed form. Unfortunately,
Machine Learning Basics: Maximum Likelihood Estimation
 Machine Learning Basics: Maximum Likelihood Estimation Sargur N. srihari@cedar.buffalo.edu This is part of lecture slides on Deep Learning: http://www.cedar.buffalo.edu/~srihari/cse676 1 Topics 1. Learning
Machine Learning Basics: Maximum Likelihood Estimation Sargur N. srihari@cedar.buffalo.edu This is part of lecture slides on Deep Learning: http://www.cedar.buffalo.edu/~srihari/cse676 1 Topics 1. Learning
component risk analysis
 273: Urban Systems Modeling Lec. 3 component risk analysis instructor: Matteo Pozzi 273: Urban Systems Modeling Lec. 3 component reliability outline risk analysis for components uncertain demand and uncertain
273: Urban Systems Modeling Lec. 3 component risk analysis instructor: Matteo Pozzi 273: Urban Systems Modeling Lec. 3 component reliability outline risk analysis for components uncertain demand and uncertain
Brandon C. Kelly (Harvard Smithsonian Center for Astrophysics)
 Brandon C. Kelly (Harvard Smithsonian Center for Astrophysics) Probability quantifies randomness and uncertainty How do I estimate the normalization and logarithmic slope of a X ray continuum, assuming
Brandon C. Kelly (Harvard Smithsonian Center for Astrophysics) Probability quantifies randomness and uncertainty How do I estimate the normalization and logarithmic slope of a X ray continuum, assuming
Statistical Data Analysis Stat 3: p-values, parameter estimation
 Statistical Data Analysis Stat 3: p-values, parameter estimation London Postgraduate Lectures on Particle Physics; University of London MSci course PH4515 Glen Cowan Physics Department Royal Holloway,
Statistical Data Analysis Stat 3: p-values, parameter estimation London Postgraduate Lectures on Particle Physics; University of London MSci course PH4515 Glen Cowan Physics Department Royal Holloway,
Gaussian processes. Chuong B. Do (updated by Honglak Lee) November 22, 2008
 Gaussian processes Chuong B Do (updated by Honglak Lee) November 22, 2008 Many of the classical machine learning algorithms that we talked about during the first half of this course fit the following pattern:
Gaussian processes Chuong B Do (updated by Honglak Lee) November 22, 2008 Many of the classical machine learning algorithms that we talked about during the first half of this course fit the following pattern:
Modern Methods of Data Analysis - WS 07/08
 Modern Methods of Data Analysis Lecture VIc (19.11.07) Contents: Maximum Likelihood Fit Maximum Likelihood (I) Assume N measurements of a random variable Assume them to be independent and distributed according
Modern Methods of Data Analysis Lecture VIc (19.11.07) Contents: Maximum Likelihood Fit Maximum Likelihood (I) Assume N measurements of a random variable Assume them to be independent and distributed according
Association studies and regression
 Association studies and regression CM226: Machine Learning for Bioinformatics. Fall 2016 Sriram Sankararaman Acknowledgments: Fei Sha, Ameet Talwalkar Association studies and regression 1 / 104 Administration
Association studies and regression CM226: Machine Learning for Bioinformatics. Fall 2016 Sriram Sankararaman Acknowledgments: Fei Sha, Ameet Talwalkar Association studies and regression 1 / 104 Administration
Statistical Distribution Assumptions of General Linear Models
 Statistical Distribution Assumptions of General Linear Models Applied Multilevel Models for Cross Sectional Data Lecture 4 ICPSR Summer Workshop University of Colorado Boulder Lecture 4: Statistical Distributions
Statistical Distribution Assumptions of General Linear Models Applied Multilevel Models for Cross Sectional Data Lecture 4 ICPSR Summer Workshop University of Colorado Boulder Lecture 4: Statistical Distributions
p(z)
 Chapter Statistics. Introduction This lecture is a quick review of basic statistical concepts; probabilities, mean, variance, covariance, correlation, linear regression, probability density functions and
Chapter Statistics. Introduction This lecture is a quick review of basic statistical concepts; probabilities, mean, variance, covariance, correlation, linear regression, probability density functions and
Fundamentals. CS 281A: Statistical Learning Theory. Yangqing Jia. August, Based on tutorial slides by Lester Mackey and Ariel Kleiner
 Fundamentals CS 281A: Statistical Learning Theory Yangqing Jia Based on tutorial slides by Lester Mackey and Ariel Kleiner August, 2011 Outline 1 Probability 2 Statistics 3 Linear Algebra 4 Optimization
Fundamentals CS 281A: Statistical Learning Theory Yangqing Jia Based on tutorial slides by Lester Mackey and Ariel Kleiner August, 2011 Outline 1 Probability 2 Statistics 3 Linear Algebra 4 Optimization
Likelihood-Based Methods
 Likelihood-Based Methods Handbook of Spatial Statistics, Chapter 4 Susheela Singh September 22, 2016 OVERVIEW INTRODUCTION MAXIMUM LIKELIHOOD ESTIMATION (ML) RESTRICTED MAXIMUM LIKELIHOOD ESTIMATION (REML)
Likelihood-Based Methods Handbook of Spatial Statistics, Chapter 4 Susheela Singh September 22, 2016 OVERVIEW INTRODUCTION MAXIMUM LIKELIHOOD ESTIMATION (ML) RESTRICTED MAXIMUM LIKELIHOOD ESTIMATION (REML)
COMS 4721: Machine Learning for Data Science Lecture 19, 4/6/2017
 COMS 4721: Machine Learning for Data Science Lecture 19, 4/6/2017 Prof. John Paisley Department of Electrical Engineering & Data Science Institute Columbia University PRINCIPAL COMPONENT ANALYSIS DIMENSIONALITY
COMS 4721: Machine Learning for Data Science Lecture 19, 4/6/2017 Prof. John Paisley Department of Electrical Engineering & Data Science Institute Columbia University PRINCIPAL COMPONENT ANALYSIS DIMENSIONALITY
Bayesian Decision Theory
 Bayesian Decision Theory Selim Aksoy Department of Computer Engineering Bilkent University saksoy@cs.bilkent.edu.tr CS 551, Fall 2017 CS 551, Fall 2017 c 2017, Selim Aksoy (Bilkent University) 1 / 46 Bayesian
Bayesian Decision Theory Selim Aksoy Department of Computer Engineering Bilkent University saksoy@cs.bilkent.edu.tr CS 551, Fall 2017 CS 551, Fall 2017 c 2017, Selim Aksoy (Bilkent University) 1 / 46 Bayesian
Estimation theory. Parametric estimation. Properties of estimators. Minimum variance estimator. Cramer-Rao bound. Maximum likelihood estimators
 Estimation theory Parametric estimation Properties of estimators Minimum variance estimator Cramer-Rao bound Maximum likelihood estimators Confidence intervals Bayesian estimation 1 Random Variables Let
Estimation theory Parametric estimation Properties of estimators Minimum variance estimator Cramer-Rao bound Maximum likelihood estimators Confidence intervals Bayesian estimation 1 Random Variables Let
Sparse Linear Models (10/7/13)
 STA56: Probabilistic machine learning Sparse Linear Models (0/7/) Lecturer: Barbara Engelhardt Scribes: Jiaji Huang, Xin Jiang, Albert Oh Sparsity Sparsity has been a hot topic in statistics and machine
STA56: Probabilistic machine learning Sparse Linear Models (0/7/) Lecturer: Barbara Engelhardt Scribes: Jiaji Huang, Xin Jiang, Albert Oh Sparsity Sparsity has been a hot topic in statistics and machine
8 - Continuous random vectors
 8-1 Continuous random vectors S. Lall, Stanford 2011.01.25.01 8 - Continuous random vectors Mean-square deviation Mean-variance decomposition Gaussian random vectors The Gamma function The χ 2 distribution
8-1 Continuous random vectors S. Lall, Stanford 2011.01.25.01 8 - Continuous random vectors Mean-square deviation Mean-variance decomposition Gaussian random vectors The Gamma function The χ 2 distribution
Lecture 3: More on regularization. Bayesian vs maximum likelihood learning
 Lecture 3: More on regularization. Bayesian vs maximum likelihood learning L2 and L1 regularization for linear estimators A Bayesian interpretation of regularization Bayesian vs maximum likelihood fitting
Lecture 3: More on regularization. Bayesian vs maximum likelihood learning L2 and L1 regularization for linear estimators A Bayesian interpretation of regularization Bayesian vs maximum likelihood fitting
Tools for Parameter Estimation and Propagation of Uncertainty
 Tools for Parameter Estimation and Propagation of Uncertainty Brian Borchers Department of Mathematics New Mexico Tech Socorro, NM 87801 borchers@nmt.edu Outline Models, parameters, parameter estimation,
Tools for Parameter Estimation and Propagation of Uncertainty Brian Borchers Department of Mathematics New Mexico Tech Socorro, NM 87801 borchers@nmt.edu Outline Models, parameters, parameter estimation,
Discriminant analysis and supervised classification
 Discriminant analysis and supervised classification Angela Montanari 1 Linear discriminant analysis Linear discriminant analysis (LDA) also known as Fisher s linear discriminant analysis or as Canonical
Discriminant analysis and supervised classification Angela Montanari 1 Linear discriminant analysis Linear discriminant analysis (LDA) also known as Fisher s linear discriminant analysis or as Canonical
MACHINE LEARNING ADVANCED MACHINE LEARNING
 MACHINE LEARNING ADVANCED MACHINE LEARNING Recap of Important Notions on Estimation of Probability Density Functions 22 MACHINE LEARNING Discrete Probabilities Consider two variables and y taking discrete
MACHINE LEARNING ADVANCED MACHINE LEARNING Recap of Important Notions on Estimation of Probability Density Functions 22 MACHINE LEARNING Discrete Probabilities Consider two variables and y taking discrete
Eco517 Fall 2004 C. Sims MIDTERM EXAM
 Eco517 Fall 2004 C. Sims MIDTERM EXAM Answer all four questions. Each is worth 23 points. Do not devote disproportionate time to any one question unless you have answered all the others. (1) We are considering
Eco517 Fall 2004 C. Sims MIDTERM EXAM Answer all four questions. Each is worth 23 points. Do not devote disproportionate time to any one question unless you have answered all the others. (1) We are considering
STA 4273H: Sta-s-cal Machine Learning
 STA 4273H: Sta-s-cal Machine Learning Russ Salakhutdinov Department of Computer Science! Department of Statistical Sciences! rsalakhu@cs.toronto.edu! h0p://www.cs.utoronto.ca/~rsalakhu/ Lecture 2 In our
STA 4273H: Sta-s-cal Machine Learning Russ Salakhutdinov Department of Computer Science! Department of Statistical Sciences! rsalakhu@cs.toronto.edu! h0p://www.cs.utoronto.ca/~rsalakhu/ Lecture 2 In our
Overfitting, Bias / Variance Analysis
 Overfitting, Bias / Variance Analysis Professor Ameet Talwalkar Professor Ameet Talwalkar CS260 Machine Learning Algorithms February 8, 207 / 40 Outline Administration 2 Review of last lecture 3 Basic
Overfitting, Bias / Variance Analysis Professor Ameet Talwalkar Professor Ameet Talwalkar CS260 Machine Learning Algorithms February 8, 207 / 40 Outline Administration 2 Review of last lecture 3 Basic
Lecture Note 1: Probability Theory and Statistics
 Univ. of Michigan - NAME 568/EECS 568/ROB 530 Winter 2018 Lecture Note 1: Probability Theory and Statistics Lecturer: Maani Ghaffari Jadidi Date: April 6, 2018 For this and all future notes, if you would
Univ. of Michigan - NAME 568/EECS 568/ROB 530 Winter 2018 Lecture Note 1: Probability Theory and Statistics Lecturer: Maani Ghaffari Jadidi Date: April 6, 2018 For this and all future notes, if you would
Computer Vision Group Prof. Daniel Cremers. 3. Regression
 Prof. Daniel Cremers 3. Regression Categories of Learning (Rep.) Learnin g Unsupervise d Learning Clustering, density estimation Supervised Learning learning from a training data set, inference on the
Prof. Daniel Cremers 3. Regression Categories of Learning (Rep.) Learnin g Unsupervise d Learning Clustering, density estimation Supervised Learning learning from a training data set, inference on the
Ch 4. Linear Models for Classification
 Ch 4. Linear Models for Classification Pattern Recognition and Machine Learning, C. M. Bishop, 2006. Department of Computer Science and Engineering Pohang University of Science and echnology 77 Cheongam-ro,
Ch 4. Linear Models for Classification Pattern Recognition and Machine Learning, C. M. Bishop, 2006. Department of Computer Science and Engineering Pohang University of Science and echnology 77 Cheongam-ro,
LECTURE NOTES FYS 4550/FYS EXPERIMENTAL HIGH ENERGY PHYSICS AUTUMN 2013 PART I A. STRANDLIE GJØVIK UNIVERSITY COLLEGE AND UNIVERSITY OF OSLO
 LECTURE NOTES FYS 4550/FYS9550 - EXPERIMENTAL HIGH ENERGY PHYSICS AUTUMN 2013 PART I PROBABILITY AND STATISTICS A. STRANDLIE GJØVIK UNIVERSITY COLLEGE AND UNIVERSITY OF OSLO Before embarking on the concept
LECTURE NOTES FYS 4550/FYS9550 - EXPERIMENTAL HIGH ENERGY PHYSICS AUTUMN 2013 PART I PROBABILITY AND STATISTICS A. STRANDLIE GJØVIK UNIVERSITY COLLEGE AND UNIVERSITY OF OSLO Before embarking on the concept
Restricted Maximum Likelihood in Linear Regression and Linear Mixed-Effects Model
 Restricted Maximum Likelihood in Linear Regression and Linear Mixed-Effects Model Xiuming Zhang zhangxiuming@u.nus.edu A*STAR-NUS Clinical Imaging Research Center October, 015 Summary This report derives
Restricted Maximum Likelihood in Linear Regression and Linear Mixed-Effects Model Xiuming Zhang zhangxiuming@u.nus.edu A*STAR-NUS Clinical Imaging Research Center October, 015 Summary This report derives
Basic Sampling Methods
 Basic Sampling Methods Sargur Srihari srihari@cedar.buffalo.edu 1 1. Motivation Topics Intractability in ML How sampling can help 2. Ancestral Sampling Using BNs 3. Transforming a Uniform Distribution
Basic Sampling Methods Sargur Srihari srihari@cedar.buffalo.edu 1 1. Motivation Topics Intractability in ML How sampling can help 2. Ancestral Sampling Using BNs 3. Transforming a Uniform Distribution
Modeling Data with Linear Combinations of Basis Functions. Read Chapter 3 in the text by Bishop
 Modeling Data with Linear Combinations of Basis Functions Read Chapter 3 in the text by Bishop A Type of Supervised Learning Problem We want to model data (x 1, t 1 ),..., (x N, t N ), where x i is a vector
Modeling Data with Linear Combinations of Basis Functions Read Chapter 3 in the text by Bishop A Type of Supervised Learning Problem We want to model data (x 1, t 1 ),..., (x N, t N ), where x i is a vector
Bayesian Regression Linear and Logistic Regression
 When we want more than point estimates Bayesian Regression Linear and Logistic Regression Nicole Beckage Ordinary Least Squares Regression and Lasso Regression return only point estimates But what if we
When we want more than point estimates Bayesian Regression Linear and Logistic Regression Nicole Beckage Ordinary Least Squares Regression and Lasso Regression return only point estimates But what if we
Estimation Tasks. Short Course on Image Quality. Matthew A. Kupinski. Introduction
 Estimation Tasks Short Course on Image Quality Matthew A. Kupinski Introduction Section 13.3 in B&M Keep in mind the similarities between estimation and classification Image-quality is a statistical concept
Estimation Tasks Short Course on Image Quality Matthew A. Kupinski Introduction Section 13.3 in B&M Keep in mind the similarities between estimation and classification Image-quality is a statistical concept
Introduction to Bayesian Statistics and Markov Chain Monte Carlo Estimation. EPSY 905: Multivariate Analysis Spring 2016 Lecture #10: April 6, 2016
 Introduction to Bayesian Statistics and Markov Chain Monte Carlo Estimation EPSY 905: Multivariate Analysis Spring 2016 Lecture #10: April 6, 2016 EPSY 905: Intro to Bayesian and MCMC Today s Class An
Introduction to Bayesian Statistics and Markov Chain Monte Carlo Estimation EPSY 905: Multivariate Analysis Spring 2016 Lecture #10: April 6, 2016 EPSY 905: Intro to Bayesian and MCMC Today s Class An
Generalized Linear Models
 Generalized Linear Models Lecture 3. Hypothesis testing. Goodness of Fit. Model diagnostics GLM (Spring, 2018) Lecture 3 1 / 34 Models Let M(X r ) be a model with design matrix X r (with r columns) r n
Generalized Linear Models Lecture 3. Hypothesis testing. Goodness of Fit. Model diagnostics GLM (Spring, 2018) Lecture 3 1 / 34 Models Let M(X r ) be a model with design matrix X r (with r columns) r n
Statistics: Learning models from data
 DS-GA 1002 Lecture notes 5 October 19, 2015 Statistics: Learning models from data Learning models from data that are assumed to be generated probabilistically from a certain unknown distribution is a crucial
DS-GA 1002 Lecture notes 5 October 19, 2015 Statistics: Learning models from data Learning models from data that are assumed to be generated probabilistically from a certain unknown distribution is a crucial
Chapter 3: Maximum-Likelihood & Bayesian Parameter Estimation (part 1)
 HW 1 due today Parameter Estimation Biometrics CSE 190 Lecture 7 Today s lecture was on the blackboard. These slides are an alternative presentation of the material. CSE190, Winter10 CSE190, Winter10 Chapter
HW 1 due today Parameter Estimation Biometrics CSE 190 Lecture 7 Today s lecture was on the blackboard. These slides are an alternative presentation of the material. CSE190, Winter10 CSE190, Winter10 Chapter
Maximum Likelihood Estimation
 Connexions module: m11446 1 Maximum Likelihood Estimation Clayton Scott Robert Nowak This work is produced by The Connexions Project and licensed under the Creative Commons Attribution License Abstract
Connexions module: m11446 1 Maximum Likelihood Estimation Clayton Scott Robert Nowak This work is produced by The Connexions Project and licensed under the Creative Commons Attribution License Abstract
Regression. Oscar García
 Regression Oscar García Regression methods are fundamental in Forest Mensuration For a more concise and general presentation, we shall first review some matrix concepts 1 Matrices An order n m matrix is
Regression Oscar García Regression methods are fundamental in Forest Mensuration For a more concise and general presentation, we shall first review some matrix concepts 1 Matrices An order n m matrix is
x. Figure 1: Examples of univariate Gaussian pdfs N (x; µ, σ 2 ).
 .8.6 µ =, σ = 1 µ = 1, σ = 1 / µ =, σ =.. 3 1 1 3 x Figure 1: Examples of univariate Gaussian pdfs N (x; µ, σ ). The Gaussian distribution Probably the most-important distribution in all of statistics
.8.6 µ =, σ = 1 µ = 1, σ = 1 / µ =, σ =.. 3 1 1 3 x Figure 1: Examples of univariate Gaussian pdfs N (x; µ, σ ). The Gaussian distribution Probably the most-important distribution in all of statistics
Linear Methods for Prediction
 Chapter 5 Linear Methods for Prediction 5.1 Introduction We now revisit the classification problem and focus on linear methods. Since our prediction Ĝ(x) will always take values in the discrete set G we
Chapter 5 Linear Methods for Prediction 5.1 Introduction We now revisit the classification problem and focus on linear methods. Since our prediction Ĝ(x) will always take values in the discrete set G we
Variational Principal Components
 Variational Principal Components Christopher M. Bishop Microsoft Research 7 J. J. Thomson Avenue, Cambridge, CB3 0FB, U.K. cmbishop@microsoft.com http://research.microsoft.com/ cmbishop In Proceedings
Variational Principal Components Christopher M. Bishop Microsoft Research 7 J. J. Thomson Avenue, Cambridge, CB3 0FB, U.K. cmbishop@microsoft.com http://research.microsoft.com/ cmbishop In Proceedings
Parametric Models. Dr. Shuang LIANG. School of Software Engineering TongJi University Fall, 2012
 Parametric Models Dr. Shuang LIANG School of Software Engineering TongJi University Fall, 2012 Today s Topics Maximum Likelihood Estimation Bayesian Density Estimation Today s Topics Maximum Likelihood
Parametric Models Dr. Shuang LIANG School of Software Engineering TongJi University Fall, 2012 Today s Topics Maximum Likelihood Estimation Bayesian Density Estimation Today s Topics Maximum Likelihood
A Few Notes on Fisher Information (WIP)
 A Few Notes on Fisher Information (WIP) David Meyer dmm@{-4-5.net,uoregon.edu} Last update: April 30, 208 Definitions There are so many interesting things about Fisher Information and its theoretical properties
A Few Notes on Fisher Information (WIP) David Meyer dmm@{-4-5.net,uoregon.edu} Last update: April 30, 208 Definitions There are so many interesting things about Fisher Information and its theoretical properties
Review (Probability & Linear Algebra)
 Review (Probability & Linear Algebra) CE-725 : Statistical Pattern Recognition Sharif University of Technology Spring 2013 M. Soleymani Outline Axioms of probability theory Conditional probability, Joint
Review (Probability & Linear Algebra) CE-725 : Statistical Pattern Recognition Sharif University of Technology Spring 2013 M. Soleymani Outline Axioms of probability theory Conditional probability, Joint
MS&E 226: Small Data. Lecture 11: Maximum likelihood (v2) Ramesh Johari
 MS&E 226: Small Data Lecture 11: Maximum likelihood (v2) Ramesh Johari ramesh.johari@stanford.edu 1 / 18 The likelihood function 2 / 18 Estimating the parameter This lecture develops the methodology behind
MS&E 226: Small Data Lecture 11: Maximum likelihood (v2) Ramesh Johari ramesh.johari@stanford.edu 1 / 18 The likelihood function 2 / 18 Estimating the parameter This lecture develops the methodology behind
Unsupervised Learning
 2018 EE448, Big Data Mining, Lecture 7 Unsupervised Learning Weinan Zhang Shanghai Jiao Tong University http://wnzhang.net http://wnzhang.net/teaching/ee448/index.html ML Problem Setting First build and
2018 EE448, Big Data Mining, Lecture 7 Unsupervised Learning Weinan Zhang Shanghai Jiao Tong University http://wnzhang.net http://wnzhang.net/teaching/ee448/index.html ML Problem Setting First build and
Theory of Maximum Likelihood Estimation. Konstantin Kashin
 Gov 2001 Section 5: Theory of Maximum Likelihood Estimation Konstantin Kashin February 28, 2013 Outline Introduction Likelihood Examples of MLE Variance of MLE Asymptotic Properties What is Statistical
Gov 2001 Section 5: Theory of Maximum Likelihood Estimation Konstantin Kashin February 28, 2013 Outline Introduction Likelihood Examples of MLE Variance of MLE Asymptotic Properties What is Statistical
Perhaps the simplest way of modeling two (discrete) random variables is by means of a joint PMF, defined as follows.
 Chapter 5 Two Random Variables In a practical engineering problem, there is almost always causal relationship between different events. Some relationships are determined by physical laws, e.g., voltage
Chapter 5 Two Random Variables In a practical engineering problem, there is almost always causal relationship between different events. Some relationships are determined by physical laws, e.g., voltage
A6523 Modeling, Inference, and Mining Jim Cordes, Cornell University
 A6523 Modeling, Inference, and Mining Jim Cordes, Cornell University Lecture 19 Modeling Topics plan: Modeling (linear/non- linear least squares) Bayesian inference Bayesian approaches to spectral esbmabon;
A6523 Modeling, Inference, and Mining Jim Cordes, Cornell University Lecture 19 Modeling Topics plan: Modeling (linear/non- linear least squares) Bayesian inference Bayesian approaches to spectral esbmabon;
STAT215: Solutions for Homework 2
 STAT25: Solutions for Homework 2 Due: Wednesday, Feb 4. (0 pt) Suppose we take one observation, X, from the discrete distribution, x 2 0 2 Pr(X x θ) ( θ)/4 θ/2 /2 (3 θ)/2 θ/4, 0 θ Find an unbiased estimator
STAT25: Solutions for Homework 2 Due: Wednesday, Feb 4. (0 pt) Suppose we take one observation, X, from the discrete distribution, x 2 0 2 Pr(X x θ) ( θ)/4 θ/2 /2 (3 θ)/2 θ/4, 0 θ Find an unbiased estimator
Lecture 3: Pattern Classification
 EE E6820: Speech & Audio Processing & Recognition Lecture 3: Pattern Classification 1 2 3 4 5 The problem of classification Linear and nonlinear classifiers Probabilistic classification Gaussians, mixtures
EE E6820: Speech & Audio Processing & Recognition Lecture 3: Pattern Classification 1 2 3 4 5 The problem of classification Linear and nonlinear classifiers Probabilistic classification Gaussians, mixtures
Introduction to Simple Linear Regression
 Introduction to Simple Linear Regression Yang Feng http://www.stat.columbia.edu/~yangfeng Yang Feng (Columbia University) Introduction to Simple Linear Regression 1 / 68 About me Faculty in the Department
Introduction to Simple Linear Regression Yang Feng http://www.stat.columbia.edu/~yangfeng Yang Feng (Columbia University) Introduction to Simple Linear Regression 1 / 68 About me Faculty in the Department
Introduction to gradient descent
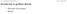 6-1: Introduction to gradient descent Prof. J.C. Kao, UCLA Introduction to gradient descent Derivation and intuitions Hessian 6-2: Introduction to gradient descent Prof. J.C. Kao, UCLA Introduction Our
6-1: Introduction to gradient descent Prof. J.C. Kao, UCLA Introduction to gradient descent Derivation and intuitions Hessian 6-2: Introduction to gradient descent Prof. J.C. Kao, UCLA Introduction Our
Lecture 5: Linear models for classification. Logistic regression. Gradient Descent. Second-order methods.
 Lecture 5: Linear models for classification. Logistic regression. Gradient Descent. Second-order methods. Linear models for classification Logistic regression Gradient descent and second-order methods
Lecture 5: Linear models for classification. Logistic regression. Gradient Descent. Second-order methods. Linear models for classification Logistic regression Gradient descent and second-order methods
COM336: Neural Computing
 COM336: Neural Computing http://www.dcs.shef.ac.uk/ sjr/com336/ Lecture 2: Density Estimation Steve Renals Department of Computer Science University of Sheffield Sheffield S1 4DP UK email: s.renals@dcs.shef.ac.uk
COM336: Neural Computing http://www.dcs.shef.ac.uk/ sjr/com336/ Lecture 2: Density Estimation Steve Renals Department of Computer Science University of Sheffield Sheffield S1 4DP UK email: s.renals@dcs.shef.ac.uk
ECE531 Lecture 10b: Maximum Likelihood Estimation
 ECE531 Lecture 10b: Maximum Likelihood Estimation D. Richard Brown III Worcester Polytechnic Institute 05-Apr-2011 Worcester Polytechnic Institute D. Richard Brown III 05-Apr-2011 1 / 23 Introduction So
ECE531 Lecture 10b: Maximum Likelihood Estimation D. Richard Brown III Worcester Polytechnic Institute 05-Apr-2011 Worcester Polytechnic Institute D. Richard Brown III 05-Apr-2011 1 / 23 Introduction So
L11: Pattern recognition principles
 L11: Pattern recognition principles Bayesian decision theory Statistical classifiers Dimensionality reduction Clustering This lecture is partly based on [Huang, Acero and Hon, 2001, ch. 4] Introduction
L11: Pattern recognition principles Bayesian decision theory Statistical classifiers Dimensionality reduction Clustering This lecture is partly based on [Huang, Acero and Hon, 2001, ch. 4] Introduction
Parametric Techniques Lecture 3
 Parametric Techniques Lecture 3 Jason Corso SUNY at Buffalo 22 January 2009 J. Corso (SUNY at Buffalo) Parametric Techniques Lecture 3 22 January 2009 1 / 39 Introduction In Lecture 2, we learned how to
Parametric Techniques Lecture 3 Jason Corso SUNY at Buffalo 22 January 2009 J. Corso (SUNY at Buffalo) Parametric Techniques Lecture 3 22 January 2009 1 / 39 Introduction In Lecture 2, we learned how to
Pattern Recognition and Machine Learning. Bishop Chapter 2: Probability Distributions
 Pattern Recognition and Machine Learning Chapter 2: Probability Distributions Cécile Amblard Alex Kläser Jakob Verbeek October 11, 27 Probability Distributions: General Density Estimation: given a finite
Pattern Recognition and Machine Learning Chapter 2: Probability Distributions Cécile Amblard Alex Kläser Jakob Verbeek October 11, 27 Probability Distributions: General Density Estimation: given a finite
SGN Advanced Signal Processing: Lecture 8 Parameter estimation for AR and MA models. Model order selection
 SG 21006 Advanced Signal Processing: Lecture 8 Parameter estimation for AR and MA models. Model order selection Ioan Tabus Department of Signal Processing Tampere University of Technology Finland 1 / 28
SG 21006 Advanced Signal Processing: Lecture 8 Parameter estimation for AR and MA models. Model order selection Ioan Tabus Department of Signal Processing Tampere University of Technology Finland 1 / 28
Variations. ECE 6540, Lecture 10 Maximum Likelihood Estimation
 Variations ECE 6540, Lecture 10 Last Time BLUE (Best Linear Unbiased Estimator) Formulation Advantages Disadvantages 2 The BLUE A simplification Assume the estimator is a linear system For a single parameter
Variations ECE 6540, Lecture 10 Last Time BLUE (Best Linear Unbiased Estimator) Formulation Advantages Disadvantages 2 The BLUE A simplification Assume the estimator is a linear system For a single parameter
A Very Brief Summary of Statistical Inference, and Examples
 A Very Brief Summary of Statistical Inference, and Examples Trinity Term 2008 Prof. Gesine Reinert 1 Data x = x 1, x 2,..., x n, realisations of random variables X 1, X 2,..., X n with distribution (model)
A Very Brief Summary of Statistical Inference, and Examples Trinity Term 2008 Prof. Gesine Reinert 1 Data x = x 1, x 2,..., x n, realisations of random variables X 1, X 2,..., X n with distribution (model)
Lecture 6. Regression
 Lecture 6. Regression Prof. Alan Yuille Summer 2014 Outline 1. Introduction to Regression 2. Binary Regression 3. Linear Regression; Polynomial Regression 4. Non-linear Regression; Multilayer Perceptron
Lecture 6. Regression Prof. Alan Yuille Summer 2014 Outline 1. Introduction to Regression 2. Binary Regression 3. Linear Regression; Polynomial Regression 4. Non-linear Regression; Multilayer Perceptron
Statistics 3858 : Maximum Likelihood Estimators
 Statistics 3858 : Maximum Likelihood Estimators 1 Method of Maximum Likelihood In this method we construct the so called likelihood function, that is L(θ) = L(θ; X 1, X 2,..., X n ) = f n (X 1, X 2,...,
Statistics 3858 : Maximum Likelihood Estimators 1 Method of Maximum Likelihood In this method we construct the so called likelihood function, that is L(θ) = L(θ; X 1, X 2,..., X n ) = f n (X 1, X 2,...,
Probabilistic & Unsupervised Learning
 Probabilistic & Unsupervised Learning Week 2: Latent Variable Models Maneesh Sahani maneesh@gatsby.ucl.ac.uk Gatsby Computational Neuroscience Unit, and MSc ML/CSML, Dept Computer Science University College
Probabilistic & Unsupervised Learning Week 2: Latent Variable Models Maneesh Sahani maneesh@gatsby.ucl.ac.uk Gatsby Computational Neuroscience Unit, and MSc ML/CSML, Dept Computer Science University College
Parameter estimation and forecasting. Cristiano Porciani AIfA, Uni-Bonn
 Parameter estimation and forecasting Cristiano Porciani AIfA, Uni-Bonn Questions? C. Porciani Estimation & forecasting 2 Temperature fluctuations Variance at multipole l (angle ~180o/l) C. Porciani Estimation
Parameter estimation and forecasting Cristiano Porciani AIfA, Uni-Bonn Questions? C. Porciani Estimation & forecasting 2 Temperature fluctuations Variance at multipole l (angle ~180o/l) C. Porciani Estimation
Parameter estimation and forecasting. Cristiano Porciani AIfA, Uni-Bonn
 Parameter estimation and forecasting Cristiano Porciani AIfA, Uni-Bonn Questions? C. Porciani Estimation & forecasting 2 Cosmological parameters A branch of modern cosmological research focuses on measuring
Parameter estimation and forecasting Cristiano Porciani AIfA, Uni-Bonn Questions? C. Porciani Estimation & forecasting 2 Cosmological parameters A branch of modern cosmological research focuses on measuring
DS-GA 1002 Lecture notes 0 Fall Linear Algebra. These notes provide a review of basic concepts in linear algebra.
 DS-GA 1002 Lecture notes 0 Fall 2016 Linear Algebra These notes provide a review of basic concepts in linear algebra. 1 Vector spaces You are no doubt familiar with vectors in R 2 or R 3, i.e. [ ] 1.1
DS-GA 1002 Lecture notes 0 Fall 2016 Linear Algebra These notes provide a review of basic concepts in linear algebra. 1 Vector spaces You are no doubt familiar with vectors in R 2 or R 3, i.e. [ ] 1.1
Some Curiosities Arising in Objective Bayesian Analysis
 . Some Curiosities Arising in Objective Bayesian Analysis Jim Berger Duke University Statistical and Applied Mathematical Institute Yale University May 15, 2009 1 Three vignettes related to John s work
. Some Curiosities Arising in Objective Bayesian Analysis Jim Berger Duke University Statistical and Applied Mathematical Institute Yale University May 15, 2009 1 Three vignettes related to John s work
University of Cambridge Engineering Part IIB Module 4F10: Statistical Pattern Processing Handout 2: Multivariate Gaussians
 Engineering Part IIB: Module F Statistical Pattern Processing University of Cambridge Engineering Part IIB Module F: Statistical Pattern Processing Handout : Multivariate Gaussians. Generative Model Decision
Engineering Part IIB: Module F Statistical Pattern Processing University of Cambridge Engineering Part IIB Module F: Statistical Pattern Processing Handout : Multivariate Gaussians. Generative Model Decision
