Solvation and Macromolecular Structure. The structure and dynamics of biological macromolecules are strongly influenced by water:
|
|
|
- Sharlene Davidson
- 5 years ago
- Views:
Transcription
1 Overview Solvation and Macromolecular Structure The structure and dynamics of biological macromolecules are strongly influenced by water: Electrostatic effects: charges are screened by water molecules and counterions. Hydrophobic effect: Entropic forces favor conformations that sequester non-polar hydrophobic domains within the interior of the molecule. Hydrodynamic effects: collision with solvent molecules imparts kinetic energy to macromolecules. Jay Taylor (ASU) APM Lecture 6 Fall / 32
2 Overview Solvation of Nucleic Acids Solvent effects on DNA and RNA are especially pronounced: Screening of the negatively-charged phosphates by water and counterions stabilizes helical and other tertiary structural motifs. 76% of the charge is reduced by water; 24% is reduced by divalent counterions. The transition between A-form and B-form DNA is partly controlled by hydration: B-DNA has waters per nucleotide. A-DNA has waters per nucleotide. DNA bending is promoted by hydration of the phosphates. Jay Taylor (ASU) APM Lecture 6 Fall / 32
3 Overview DNA is surrounded by multiple layers of water. Water density is increased up to six-fold in the first layer, especially around the phosphates and in the minor and major groove. A less stable second hydration shell extends out to 10 Å. A spine of hydration runs through the minor groove. Jay Taylor (ASU) APM Lecture 6 Fall / 32
4 Overview Explicit Solvation Models Solvent effects must also be included in molecular simulations. One approach is to explicitly include a large number of solvent molecules surrounding the macromolecule. Boundary conditions are usually periodic. Counterions can also be explicitly modeled. A large number of solvent molecules is generally required: Approximately 3000 waters are needed in a simulation of a 10 bp DNA duplex. Solvent viscosity leads to slower sampling of conformational space. Jay Taylor (ASU) APM Lecture 6 Fall / 32
5 Overview Implicit Solvation Models Implicit solvent models represent the solvent and counterions as a continuous medium. Implicit simulations can usually be run more quickly than explicit simulations. We are usually not interested in the distribution of individual water molecules in the solvent-solute interface. Residence times of water in DNA vary from hundreds of picoseconds to nanoseconds in the minor groove of A-tracts. Several methods are available: Solvent accessible surface area models Poisson-Boltzmann equation Generalized Born models Jay Taylor (ASU) APM Lecture 6 Fall / 32
6 SASA models Solvent Accessible Surface Area SASA models express the free energy of solvation as a sum G solv = i σ i ASA i where the sum is over all atoms in the macromolecule; ASA i is the surface area of atom i accessible to the solvent; σ i is an atom-specific surface tension parameter. The parameters σ i have been estimated from empirical hydration free energies for various organic compounds in water. Jay Taylor (ASU) APM Lecture 6 Fall / 32
7 SASA models Accessible Surface Area Calculations Atoms and solvent molecules are both modeled as spheres. The ASA of an atom is the area of those points on the surface of a sphere of radius R which can contact the center of a spherical solvent molecule that intersects no other atoms. R is the sum of the van der Walls radius of the atom and the radius of the solvent molecule (1.4 Å for water). Jay Taylor (ASU) APM Lecture 6 Fall / 32
8 SASA models Shrake-Rupley Algorithm The Shrake-Rupley algorithm (1973) uses a discretization of the molecular surface to estimate ASA: 92 points are distributed uniformly along a sphere centered at each atom with radius R (defined on the previous slide). The ASA is estimated by calculating the proportion of points that can be contacted by a solvent molecule that intersects no other atoms. Related methods approximate the surface using polyhedra. Gradients of discretized ASA-estimates must be evaluated numerically. Jay Taylor (ASU) APM Lecture 6 Fall / 32
9 SASA models Linear Combination of Pairwise Overlaps Weiser et al. (1999) proposed an approximate analytical expression for the ASA based on an inclusion-exclusion-like formula: A i P 1 S i + P 2 j N(i) A ij + S i is the surface area of atom i. j,k N(i) k j (P 3 + P 4 A ij )A jk A ij is the surface area of sphere i buried inside sphere j. N(i) is the neighbor list of atom i. P 1 - P 4 were calculated using least squares regression. The relative error is in the range %. The resulting formula can be differentiated. Jay Taylor (ASU) APM Lecture 6 Fall / 32
10 SASA models Limitations of SASA Models SASA models have several important limitations: Solvation free energies are not linearly related to surface area, e.g., SASA overestimates hydration free energies of cyclic alkanes. The solvation free energy calculated using SASA ignores the electrostatic effects of the solvent. SASA does not account for interactions between the solvent and polar atoms that are buried in the interior of the macromolecule. Jay Taylor (ASU) APM Lecture 6 Fall / 32
11 Poisson-Boltzmann equation Continuum Electrostatics Continuum electrostatic models account for the effects of the solvent on the electrostatics of the macromolecule. The simplest approach is to introduce a distance-dependent dielectric function: ɛ(r) = D s + D ke κ(ds+d 0) r D 0. ɛ(r) mimics electrostatic screening by the solvent. This approach ignores the fact that the solute itself influences the distribution of solvent molecules: Solvent molecules are often excluded from the interior of the solute. Counterions and polar solvent molecules condense around charged atoms. Jay Taylor (ASU) APM Lecture 6 Fall / 32
12 Poisson-Boltzmann equation Potential of Mean Force More accurate results can be obtained by modeling the distribution of the solvent around the macromolecule. At equilibrium, the total charge density of the solvent and any dissolved salts is described by the Boltzmann distribution ρ solvent (r) = i q i c i e Ṽi (r)/k B T where q i and c i are the charge and concentration of the i th ion; The potential of mean force for ion i is approximately related to the bulk electrostatic field by Ṽ i (r) q i Φ(r). Jay Taylor (ASU) APM Lecture 6 Fall / 32
13 Poisson-Boltzmann equation The Poisson-Boltzmann Equation We also know that the bulk potential Φ is related to the charge distribution of the solvent and the solute via Poisson s equation [ɛ(r) Φ(r)] = 4πρ solute (r) 4πρ solvent (r). Then, by combining this result with the expression for ρ solvent from the previous slide, we obtain the Poisson-Boltzmann equation: [ɛ(r) Φ(r)] = 4πρ solute (r) 4π i q i c i e q i Φ(r)/k B T. Jay Taylor (ASU) APM Lecture 6 Fall / 32
14 Poisson-Boltzmann equation The Poisson-Boltzmann Equation When the solvent consists of two oppositely charged ions with the same concentration c, the Poisson-Boltzmann equation can be written in the form: [ɛ(r) Φ(r)] = 4πρ solute (r) 4πqc (e ) qφ(r)/kbt e qφ(r)/k BT = 4πρ solute (r) 8πqc sinh(qφ(r)/k B T ). Jay Taylor (ASU) APM Lecture 6 Fall / 32
15 Poisson-Boltzmann equation The Linearized Poisson-Boltzmann Equation In practice, the Poisson-Boltzmann equation is usually linearized about 0: [ɛ(r) Φ(r)] = 4πρ solute (r) 4π i q i c i (1 q i Φ(r)/k B T ) where we have assumed/defined = 4πρ solute (r) + γ 2 Φ(r), 0 = i c i q i and γ 2 = 8π k B T c i qi 2. i This linearized version is appropriate when the electrostatic potential is much smaller than the thermal energy: q i Φ(r) k B T. Jay Taylor (ASU) APM Lecture 6 Fall / 32
16 Poisson-Boltzmann equation Debye-Hückel Theory Analytical solutions to the linearized P-B equation are available for some special cases. The Debye-Hückel theory considers a single spherical ion in an electrolyte with radius a and uniform surface charge q; constant interior and solvent permittivities ɛ i and ɛ s. In this case, the LPB can be written as (ɛ s /ɛ i )κ 2 Φ(r) if r < a 2 Φ(r) = κ 2 Φ(r) if r > a, where κ 2 = γ 2 /ɛ s. Jay Taylor (ASU) APM Lecture 6 Fall / 32
17 Poisson-Boltzmann equation Debye-Hückel Theory For r a, this equation coincides with the Helmholtz equation and has a unique solution in C 1 : Φ(r) = [ ] q 1 4π ɛ i r 1 ɛ i a + 1 ɛ sa(1+κa) if r < a ( ) exp(κa) q exp( κ r ) 1+κa ɛ r if r a. The second part of the solution illustrates the exponential screening of the ion by counterions in the solvent. κ 1 is called the Debye screening length and is inversely proportional to the square root of the ionic strength. Jay Taylor (ASU) APM Lecture 6 Fall / 32
18 Poisson-Boltzmann equation Numerical Solutions to the Poisson-Boltzmann Equation In practice, both the PB and the LPB equations must be solved numerically. Several difficulties need to be addressed: Repetitive solution of the PBE is needed for energy minimization and MD. Solutions need to be smooth in space and time to be used in MD. Boundary conditions need to be accurately specified. The dielectric function changes rapidly at the solvent-solute interface. Finite-difference, finite-element and boundary element approaches have all been developed for this problem. Jay Taylor (ASU) APM Lecture 6 Fall / 32
19 Poisson-Boltzmann equation Finite Difference Methods for the LPBE Luo et al. (2002) describe an efficient implementation of the finite difference approach. A 1 Å cubic mesh is used for high-accuracy solutions. The atomic charges and dielectric function are interpolated onto the grid points. The linearized PBE is replaced by a finite-difference equation Ax = b where A is a 7-banded symmetric positive-definite matrix; b is the interpolated charge distribution. The FDPB equation is solved iteratively using the Modified Incomplete Cholsky Conjugate Gradient method. Jay Taylor (ASU) APM Lecture 6 Fall / 32
20 Poisson-Boltzmann equation Preconditioning The preconditioning matrix used in MICCG is M = (L + D)D 1 (D + U) where U and L are the strictly upper and lower parts of A and D is the diagonal matrix with diagonal entries d 1 i = A i,1 A 2 i 1,2(A 2 i 1,2 + αa 2 i 1,3 + αa 2 i 1,4)d i 1 A 2 i nx,3(αa 2 i nx,2 + A 2 i nx,3 + αa 2 i nx,4)d i nx A 2 i nxny,4(αa 2 i nxny,2 + αa 2 i nxny,3 + A 2 i nxny,4)d i nxny. Here A has dimension n x n y n z with diagonal element A i,1 and non-zero off-diagonal elements A i,2, A i,3, A i,4 at row i. Jay Taylor (ASU) APM Lecture 6 Fall / 32
21 Poisson-Boltzmann equation Acceleration of the FDBE Iterative solution of the FDBE can be accelerated by the following practices: The convergence criterion for the conjugate gradient algorithm can be taken to be 10 2 rather than This leads to a 6-fold speedup with correlation coefficient of 0.99 compared to the 10 6 calculation. The current value the potential can be used as an initial guess for the subsequent conjugate gradient iteration. The PB potential can be updated intermittently (every 2-10 steps). There is a tradeoff between the frequency of updating and the number of iterations required per update. Jay Taylor (ASU) APM Lecture 6 Fall / 32
22 Poisson-Boltzmann equation Electrostatic Focusing Electrostatic focusing can be used to obtain accurate solutions to elliptic problems with non-periodic boundary conditions: A low-accuracy solution is first obtained on a coarse mesh spanning the entire space. This solution is then used to define boundary conditions for a higher accuracy calculation on a subdomain using a finer mesh. Example: Electrostatic focusing accelerates the FDBE core routine by a factor of 3-4 when applied to HIV-1 protease with an initial 2 Å grid of 100 Å 3 followed by a final 1 Å grid of Å. Jay Taylor (ASU) APM Lecture 6 Fall / 32
23 Poisson-Boltzmann equation Electrostatic Focusing and Parallelization Baker et al. (2001) showed that electrostatic focusing can also be used to parallelize the FDBE algorithm. Each processor obtains a coarse solution over the whole domain. This coarse solution is then used to assign boundary conditions to each of a collection of overlapping subdomains. Each processor then solves the PBE on a finer mesh on each subdomain. This method keeps communication between processors to a minimum. Jay Taylor (ASU) APM Lecture 6 Fall / 32
24 Poisson-Boltzmann equation Application to Microtubule Electrostatics By combining a multigrid algorithm with parallel focusing, the authors are able to solve the linearized PBE for a microtubule containing 90 dimers and over 1.25 million atoms. The microtubule structure was inferred from X-ray data. Both the coarse global mesh and the finer subdomain meshes contained 97 3 grid points, with a final resolution of 0.54 Å. The LPBE was solved in less than 1 hour using 686 processors. Jay Taylor (ASU) APM Lecture 6 Fall / 32
25 Poisson-Boltzmann equation Applications to Nucleic Acids Because of their high charge density, continuum electrostatic models of nucleic acids usually resort to the full non-linear PBE. Misra & Draper (2000) showed that the PBE quantitatively predicts the free energy of Mg 2+ binding to a yeast trna (right). Similar results have been obtained for salt effects on DNA-ligand binding and peptide-rna docking (Misra et al., 1994; Garcia-Garcia & Draper, 2003). Jay Taylor (ASU) APM Lecture 6 Fall / 32
26 Generalized Born models Generalized Born Models Generalized Born models approximate the solvation free energy by the formula: where ( G GB = 1 1 ) ɛ w i qi 2 1 ( 1 1 ) 2α i 2 ɛ w i j α i is the effective Born radius of atom i; ɛ w is the dielectric constant of water; f GB (r ij, α i, α j ) = rij 2 + α iα j exp( rij 2/4α iα j ). q i q j f GB (r ij, α i, α j ) Jay Taylor (ASU) APM Lecture 6 Fall / 32
27 Generalized Born models Solvation Free Energy of Isolated Charges The effective Born radius α i is chosen so that the quantity ( 1 1 ) q 2 i ɛ w 2α i is the electrostatic solvation energy of an isolated charge surrounded by a spherical shell that excludes the solvent. α i can be interpreted as the average distance of atom i from the solvent accessible surface of the molecule; This is an approximation since most atoms will not be surrounded by spherical cavities. Jay Taylor (ASU) APM Lecture 6 Fall / 32
28 Generalized Born models Calculation of the Effective Born Radii The effective Born radii can be estimated numerically but more often are evaluated using an approximate analytical expression (Hawkins et al. 1996): α 1 i ρ 1 i 1 j i ρ i r 2 H ij(r, ρ j )dr ρ 1 i j g(r ij, ρ i, ρ j ) where ρ i is the intrinsic radius of atom i (i.e., in isolation). H ij (r, ρ j ) is the fraction of the area of a sphere of radius r centered at atom i that is shielded by atom j. Jay Taylor (ASU) APM Lecture 6 Fall / 32
29 Generalized Born models Interaction Solvation Energy The second term that appears in the generalized Born model 1 2 ( 1 1 ) q i q j ɛ w f GB (r ij, α i, α j ) i j is the pairwise contribution of the charges in the molecule to the solvation energy. The interpolation function f GB has been chosen so that: lim r ij 0 f GB (r ij, α i, α j ) = α i α j lim r 1 r ij ij f GB (r ij, α i, α j ) = 1. Jay Taylor (ASU) APM Lecture 6 Fall / 32
30 Generalized Born models Generalized Born Models in Practice The Generalized Born model is often used in combination with the SASA estimate of the nonpolar solvation energy (GBSA models). Because the GBSA solvation energy can be calculated relatively quickly and can also be differentiated analytically, these models have seen more widespread use in MD than the Poisson-Boltzmann equation. On the other hand, GBSA is considered to be a cruder approximation for the solvation energy than PBE. Jay Taylor (ASU) APM Lecture 6 Fall / 32
31 Generalized Born models Generalized Born Simulations of DNA Tsui & Case (2001) compared the results of explicit solvent and generalized Born simulations of a 10 base-pair DNA helix for a 12 ns period. Both methods produce stable structures over the 12 ns trajectory. Hydrogen bond distances are shorter by Å in the explicit solvent simulation. The atomic fluctuations of the inner base pairs are highly correlated between the two methods (right). Jay Taylor (ASU) APM Lecture 6 Fall / 32
32 References References Luo, R., David, L. and Gilson, M. K. (2002) Accelerated Poisson-Boltzmann Calculations for Static and Dynamic Systems. J. Comput. Chem. 23: Schlick, T. (2006) Molecular Modeling and Simulation. Springer. Tsui, V. and Case, D.A. (2000) Theory and applications of the generalized Born solvation model in macromolecular simulations. Biopolymers 56: Zacharias, M. (2006) Continuum Solvent Models to Study the Structure and Dynamics of Nucleic Acids and Complexes with Ligands. In Computational Studies of RNA and DNA, ed. by Jiri Sponer and F. Lankas. Springer. Jay Taylor (ASU) APM Lecture 6 Fall / 32
Can a continuum solvent model reproduce the free energy landscape of a β-hairpin folding in water?
 Can a continuum solvent model reproduce the free energy landscape of a β-hairpin folding in water? Ruhong Zhou 1 and Bruce J. Berne 2 1 IBM Thomas J. Watson Research Center; and 2 Department of Chemistry,
Can a continuum solvent model reproduce the free energy landscape of a β-hairpin folding in water? Ruhong Zhou 1 and Bruce J. Berne 2 1 IBM Thomas J. Watson Research Center; and 2 Department of Chemistry,
ONETEP PB/SA: Application to G-Quadruplex DNA Stability. Danny Cole
 ONETEP PB/SA: Application to G-Quadruplex DNA Stability Danny Cole Introduction Historical overview of structure and free energy calculation of complex molecules using molecular mechanics and continuum
ONETEP PB/SA: Application to G-Quadruplex DNA Stability Danny Cole Introduction Historical overview of structure and free energy calculation of complex molecules using molecular mechanics and continuum
Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015,
 Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015, Course,Informa5on, BIOC%530% GraduateAlevel,discussion,of,the,structure,,func5on,,and,chemistry,of,proteins,and, nucleic,acids,,control,of,enzyma5c,reac5ons.,please,see,the,course,syllabus,and,
Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015, Course,Informa5on, BIOC%530% GraduateAlevel,discussion,of,the,structure,,func5on,,and,chemistry,of,proteins,and, nucleic,acids,,control,of,enzyma5c,reac5ons.,please,see,the,course,syllabus,and,
Bchem 675 Lecture 9 Electrostatics-Lecture 2 Debye-Hückel: Continued Counter ion condensation
 Bchem 675 Lecture 9 Electrostatics-Lecture 2 Debye-Hückel: Continued Counter ion condensation Ion:ion interactions What is the free energy of ion:ion interactions ΔG i-i? Consider an ion in a solution
Bchem 675 Lecture 9 Electrostatics-Lecture 2 Debye-Hückel: Continued Counter ion condensation Ion:ion interactions What is the free energy of ion:ion interactions ΔG i-i? Consider an ion in a solution
Aqueous solutions. Solubility of different compounds in water
 Aqueous solutions Solubility of different compounds in water The dissolution of molecules into water (in any solvent actually) causes a volume change of the solution; the size of this volume change is
Aqueous solutions Solubility of different compounds in water The dissolution of molecules into water (in any solvent actually) causes a volume change of the solution; the size of this volume change is
INTRODUCTION TO FINITE ELEMENT METHODS ON ELLIPTIC EQUATIONS LONG CHEN
 INTROUCTION TO FINITE ELEMENT METHOS ON ELLIPTIC EQUATIONS LONG CHEN CONTENTS 1. Poisson Equation 1 2. Outline of Topics 3 2.1. Finite ifference Method 3 2.2. Finite Element Method 3 2.3. Finite Volume
INTROUCTION TO FINITE ELEMENT METHOS ON ELLIPTIC EQUATIONS LONG CHEN CONTENTS 1. Poisson Equation 1 2. Outline of Topics 3 2.1. Finite ifference Method 3 2.2. Finite Element Method 3 2.3. Finite Volume
GENERALIZED BORN MODELS OF MACROMOLECULARSOLVATION EFFECTS
 Annu. Rev. Phys. Chem. 2000. 51:129 52 Copyright c 2000 by Annual Reviews. All rights reserved GENERALIZED BORN MODELS OF MACROMOLECULARSOLVATION EFFECTS Donald Bashford and David A. Case Department of
Annu. Rev. Phys. Chem. 2000. 51:129 52 Copyright c 2000 by Annual Reviews. All rights reserved GENERALIZED BORN MODELS OF MACROMOLECULARSOLVATION EFFECTS Donald Bashford and David A. Case Department of
Lecture 12: Solvation Models: Molecular Mechanics Modeling of Hydration Effects
 Statistical Thermodynamics Lecture 12: Solvation Models: Molecular Mechanics Modeling of Hydration Effects Dr. Ronald M. Levy ronlevy@temple.edu Bare Molecular Mechanics Atomistic Force Fields: torsion
Statistical Thermodynamics Lecture 12: Solvation Models: Molecular Mechanics Modeling of Hydration Effects Dr. Ronald M. Levy ronlevy@temple.edu Bare Molecular Mechanics Atomistic Force Fields: torsion
Electrostatic correlations and fluctuations for ion binding to a finite length polyelectrolyte
 THE JOURNAL OF CHEMICAL PHYSICS 122, 044903 2005 Electrostatic correlations and fluctuations for ion binding to a finite length polyelectrolyte Zhi-Jie Tan and Shi-Jie Chen a) Department of Physics and
THE JOURNAL OF CHEMICAL PHYSICS 122, 044903 2005 Electrostatic correlations and fluctuations for ion binding to a finite length polyelectrolyte Zhi-Jie Tan and Shi-Jie Chen a) Department of Physics and
Villa et al. (2005) Structural dynamics of the lac repressor-dna complex revealed by a multiscale simulation. PNAS 102:
 Villa et al. (2005) Structural dynamics of the lac repressor-dna complex revealed by a multiscale simulation. PNAS 102: 6783-6788. Background: The lac operon is a cluster of genes in the E. coli genome
Villa et al. (2005) Structural dynamics of the lac repressor-dna complex revealed by a multiscale simulation. PNAS 102: 6783-6788. Background: The lac operon is a cluster of genes in the E. coli genome
Molecular Modeling -- Lecture 15 Surfaces and electrostatics
 Molecular Modeling -- Lecture 15 Surfaces and electrostatics Molecular surfaces The Hydrophobic Effect Electrostatics Poisson-Boltzmann Equation Electrostatic maps Electrostatic surfaces in MOE 15.1 The
Molecular Modeling -- Lecture 15 Surfaces and electrostatics Molecular surfaces The Hydrophobic Effect Electrostatics Poisson-Boltzmann Equation Electrostatic maps Electrostatic surfaces in MOE 15.1 The
Molecular Simulation III
 Molecular Simulation III Quantum Chemistry Classical Mechanics E = Ψ H Ψ ΨΨ U = E bond +E angle +E torsion +E non-bond Molecular Dynamics Jeffry D. Madura Department of Chemistry & Biochemistry Center
Molecular Simulation III Quantum Chemistry Classical Mechanics E = Ψ H Ψ ΨΨ U = E bond +E angle +E torsion +E non-bond Molecular Dynamics Jeffry D. Madura Department of Chemistry & Biochemistry Center
Continuum Electrostatics for Ionic Solutions with Nonuniform Ionic Sizes
 Continuum Electrostatics for Ionic Solutions with Nonuniform Ionic Sizes Bo Li Department of Mathematics and Center for Theoretical Biological Physics (CTBP) University of California, San Diego, USA Supported
Continuum Electrostatics for Ionic Solutions with Nonuniform Ionic Sizes Bo Li Department of Mathematics and Center for Theoretical Biological Physics (CTBP) University of California, San Diego, USA Supported
Peptide folding in non-aqueous environments investigated with molecular dynamics simulations Soto Becerra, Patricia
 University of Groningen Peptide folding in non-aqueous environments investigated with molecular dynamics simulations Soto Becerra, Patricia IMPORTANT NOTE: You are advised to consult the publisher's version
University of Groningen Peptide folding in non-aqueous environments investigated with molecular dynamics simulations Soto Becerra, Patricia IMPORTANT NOTE: You are advised to consult the publisher's version
Lecture 2-3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability
 Lecture 2-3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability Part I. Review of forces Covalent bonds Non-covalent Interactions Van der Waals Interactions
Lecture 2-3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability Part I. Review of forces Covalent bonds Non-covalent Interactions Van der Waals Interactions
Lecture 2 and 3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability
 Lecture 2 and 3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability Part I. Review of forces Covalent bonds Non-covalent Interactions: Van der Waals Interactions
Lecture 2 and 3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability Part I. Review of forces Covalent bonds Non-covalent Interactions: Van der Waals Interactions
The generalized Born model: its foundation, applications, and limitations
 The generalized Born model: its foundation, applications, and limitations Alexey Onufriev September 8, 2010 Abstract The generalized Born (GB) approximation is introduced in the context of the implicit
The generalized Born model: its foundation, applications, and limitations Alexey Onufriev September 8, 2010 Abstract The generalized Born (GB) approximation is introduced in the context of the implicit
Applications of Molecular Dynamics
 June 4, 0 Molecular Modeling and Simulation Applications of Molecular Dynamics Agricultural Bioinformatics Research Unit, Graduate School of Agricultural and Life Sciences, The University of Tokyo Tohru
June 4, 0 Molecular Modeling and Simulation Applications of Molecular Dynamics Agricultural Bioinformatics Research Unit, Graduate School of Agricultural and Life Sciences, The University of Tokyo Tohru
2 Structure. 2.1 Coulomb interactions
 2 Structure 2.1 Coulomb interactions While the information needed for reproduction of living systems is chiefly maintained in the sequence of macromolecules, any practical use of this information must
2 Structure 2.1 Coulomb interactions While the information needed for reproduction of living systems is chiefly maintained in the sequence of macromolecules, any practical use of this information must
Multigrid and Domain Decomposition Methods for Electrostatics Problems
 Multigrid and Domain Decomposition Methods for Electrostatics Problems Michael Holst and Faisal Saied Abstract. We consider multigrid and domain decomposition methods for the numerical solution of electrostatics
Multigrid and Domain Decomposition Methods for Electrostatics Problems Michael Holst and Faisal Saied Abstract. We consider multigrid and domain decomposition methods for the numerical solution of electrostatics
A SHORT NOTE COMPARING MULTIGRID AND DOMAIN DECOMPOSITION FOR PROTEIN MODELING EQUATIONS
 A SHORT NOTE COMPARING MULTIGRID AND DOMAIN DECOMPOSITION FOR PROTEIN MODELING EQUATIONS MICHAEL HOLST AND FAISAL SAIED Abstract. We consider multigrid and domain decomposition methods for the numerical
A SHORT NOTE COMPARING MULTIGRID AND DOMAIN DECOMPOSITION FOR PROTEIN MODELING EQUATIONS MICHAEL HOLST AND FAISAL SAIED Abstract. We consider multigrid and domain decomposition methods for the numerical
Phase Equilibria and Molecular Solutions Jan G. Korvink and Evgenii Rudnyi IMTEK Albert Ludwig University Freiburg, Germany
 Phase Equilibria and Molecular Solutions Jan G. Korvink and Evgenii Rudnyi IMTEK Albert Ludwig University Freiburg, Germany Preliminaries Learning Goals Phase Equilibria Phase diagrams and classical thermodynamics
Phase Equilibria and Molecular Solutions Jan G. Korvink and Evgenii Rudnyi IMTEK Albert Ludwig University Freiburg, Germany Preliminaries Learning Goals Phase Equilibria Phase diagrams and classical thermodynamics
BIOC : Homework 1 Due 10/10
 Contact information: Name: Student # BIOC530 2012: Homework 1 Due 10/10 Department Email address The following problems are based on David Baker s lectures of forces and protein folding. When numerical
Contact information: Name: Student # BIOC530 2012: Homework 1 Due 10/10 Department Email address The following problems are based on David Baker s lectures of forces and protein folding. When numerical
Biochemistry 530: Introduction to Structural Biology. Autumn Quarter 2014 BIOC 530
 Biochemistry 530: Introduction to Structural Biology Autumn Quarter 2014 Course Information Course Description Graduate-level discussion of the structure, function, and chemistry of proteins and nucleic
Biochemistry 530: Introduction to Structural Biology Autumn Quarter 2014 Course Information Course Description Graduate-level discussion of the structure, function, and chemistry of proteins and nucleic
All-atom Molecular Mechanics. Trent E. Balius AMS 535 / CHE /27/2010
 All-atom Molecular Mechanics Trent E. Balius AMS 535 / CHE 535 09/27/2010 Outline Molecular models Molecular mechanics Force Fields Potential energy function functional form parameters and parameterization
All-atom Molecular Mechanics Trent E. Balius AMS 535 / CHE 535 09/27/2010 Outline Molecular models Molecular mechanics Force Fields Potential energy function functional form parameters and parameterization
Level-Set Variational Solvation Coupling Solute Molecular Mechanics with Continuum Solvent
 Level-Set Variational Solvation Coupling Solute Molecular Mechanics with Continuum Solvent Bo Li Department of Mathematics and Center for Theoretical Biological Physics (CTBP) University of California,
Level-Set Variational Solvation Coupling Solute Molecular Mechanics with Continuum Solvent Bo Li Department of Mathematics and Center for Theoretical Biological Physics (CTBP) University of California,
SOLVATION ENERGY OF BIOMOLECULAR STRUCTURES: A STUDY OF THE EFFECT OF SALT ON BIOMOLECULES THROUGH IMPLICIT AS WELL AS EXPLICIT SOLVATION METHODS
 Clemson University TigerPrints All Dissertations Dissertations 12-2012 SOLVATION ENERGY OF BIOMOLECULAR STRUCTURES: A STUDY OF THE EFFECT OF SALT ON BIOMOLECULES THROUGH IMPLICIT AS WELL AS EXPLICIT SOLVATION
Clemson University TigerPrints All Dissertations Dissertations 12-2012 SOLVATION ENERGY OF BIOMOLECULAR STRUCTURES: A STUDY OF THE EFFECT OF SALT ON BIOMOLECULES THROUGH IMPLICIT AS WELL AS EXPLICIT SOLVATION
Electrolytes. Chapter Basics = = 131 2[ ]. (c) From both of the above = = 120 8[
![Electrolytes. Chapter Basics = = 131 2[ ]. (c) From both of the above = = 120 8[ Electrolytes. Chapter Basics = = 131 2[ ]. (c) From both of the above = = 120 8[](/thumbs/83/88791608.jpg) Chapter 1 Electrolytes 1.1 Basics Here we consider species that dissociate into positively and negatively charged species in solution. 1. Consider: 1 H (g) + 1 Cl (g) + ()+ () = { } = (+ )+ ( ) = 167[
Chapter 1 Electrolytes 1.1 Basics Here we consider species that dissociate into positively and negatively charged species in solution. 1. Consider: 1 H (g) + 1 Cl (g) + ()+ () = { } = (+ )+ ( ) = 167[
Lectures 11-13: Electrostatics of Salty Solutions
 Lectures 11-13: Electrostatics of Salty Solutions Lecturer: Brigita Urbanc Office: 1-909 (E-mail: brigita@drexel.edu) Course website: www.physics.drexel.edu/~brigita/courses/biophys_011-01/ 1 Water as
Lectures 11-13: Electrostatics of Salty Solutions Lecturer: Brigita Urbanc Office: 1-909 (E-mail: brigita@drexel.edu) Course website: www.physics.drexel.edu/~brigita/courses/biophys_011-01/ 1 Water as
Molecular Mechanics, Dynamics & Docking
 Molecular Mechanics, Dynamics & Docking Lawrence Hunter, Ph.D. Director, Computational Bioscience Program University of Colorado School of Medicine Larry.Hunter@uchsc.edu http://compbio.uchsc.edu/hunter
Molecular Mechanics, Dynamics & Docking Lawrence Hunter, Ph.D. Director, Computational Bioscience Program University of Colorado School of Medicine Larry.Hunter@uchsc.edu http://compbio.uchsc.edu/hunter
INTERMOLECULAR AND SURFACE FORCES
 INTERMOLECULAR AND SURFACE FORCES SECOND EDITION JACOB N. ISRAELACHVILI Department of Chemical & Nuclear Engineering and Materials Department University of California, Santa Barbara California, USA ACADEMIC
INTERMOLECULAR AND SURFACE FORCES SECOND EDITION JACOB N. ISRAELACHVILI Department of Chemical & Nuclear Engineering and Materials Department University of California, Santa Barbara California, USA ACADEMIC
Explicit Ion, Implicit Water Solvation for Molecular Dynamics of Nucleic Acids and Highly Charged Molecules
 Explicit Ion, Implicit Water Solvation for Molecular Dynamics of Nucleic Acids and Highly Charged Molecules NINAD V. PRABHU, 1 MANORANJAN PANDA, 2 QINGYI YANG, 3 KIM A. SHARP 3 1 Schering-Plough, Kenilworth,
Explicit Ion, Implicit Water Solvation for Molecular Dynamics of Nucleic Acids and Highly Charged Molecules NINAD V. PRABHU, 1 MANORANJAN PANDA, 2 QINGYI YANG, 3 KIM A. SHARP 3 1 Schering-Plough, Kenilworth,
Proteins polymer molecules, folded in complex structures. Konstantin Popov Department of Biochemistry and Biophysics
 Proteins polymer molecules, folded in complex structures Konstantin Popov Department of Biochemistry and Biophysics Outline General aspects of polymer theory Size and persistent length of ideal linear
Proteins polymer molecules, folded in complex structures Konstantin Popov Department of Biochemistry and Biophysics Outline General aspects of polymer theory Size and persistent length of ideal linear
Biophysics II. Hydrophobic Bio-molecules. Key points to be covered. Molecular Interactions in Bio-molecular Structures - van der Waals Interaction
 Biophysics II Key points to be covered By A/Prof. Xiang Yang Liu Biophysics & Micro/nanostructures Lab Department of Physics, NUS 1. van der Waals Interaction 2. Hydrogen bond 3. Hydrophilic vs hydrophobic
Biophysics II Key points to be covered By A/Prof. Xiang Yang Liu Biophysics & Micro/nanostructures Lab Department of Physics, NUS 1. van der Waals Interaction 2. Hydrogen bond 3. Hydrophilic vs hydrophobic
Where do ions solvate?
 PRAMANA c Indian Academy of Sciences Vol. 64, No. 6 journal of June 25 physics pp. 957 961 YAN LEVIN Instituto de Física, Universidade Federal do Rio Grande do Sul, Caixa Postal 1551, CEP 9151-97, Porto
PRAMANA c Indian Academy of Sciences Vol. 64, No. 6 journal of June 25 physics pp. 957 961 YAN LEVIN Instituto de Física, Universidade Federal do Rio Grande do Sul, Caixa Postal 1551, CEP 9151-97, Porto
Research Statement. Shenggao Zhou. November 3, 2014
 Shenggao Zhou November 3, My research focuses on: () Scientific computing and numerical analysis (numerical PDEs, numerical optimization, computational fluid dynamics, and level-set method for interface
Shenggao Zhou November 3, My research focuses on: () Scientific computing and numerical analysis (numerical PDEs, numerical optimization, computational fluid dynamics, and level-set method for interface
A generalized Born formalism for heterogeneous dielectric environments: Application to the implicit modeling of biological membranes
 THE JOURNAL OF CHEMICAL PHYSICS 122, 124706 2005 A generalized Born formalism for heterogeneous dielectric environments: Application to the implicit modeling of biological membranes Seiichiro Tanizaki
THE JOURNAL OF CHEMICAL PHYSICS 122, 124706 2005 A generalized Born formalism for heterogeneous dielectric environments: Application to the implicit modeling of biological membranes Seiichiro Tanizaki
Langevin Dynamics of a Single Particle
 Overview Langevin Dynamics of a Single Particle We consider a spherical particle of radius r immersed in a viscous fluid and suppose that the dynamics of the particle depend on two forces: A drag force
Overview Langevin Dynamics of a Single Particle We consider a spherical particle of radius r immersed in a viscous fluid and suppose that the dynamics of the particle depend on two forces: A drag force
Density Functional Theory for the Nonspecific Binding of Salt to Polyelectrolytes: Thermodynamic Properties
 Biophysical Journal Volume 78 February 2000 699 706 699 Density Functional Theory for the Nonspecific Binding of Salt to Polyelectrolytes: Thermodynamic Properties Chandra N. Patra and Arun Yethiraj Department
Biophysical Journal Volume 78 February 2000 699 706 699 Density Functional Theory for the Nonspecific Binding of Salt to Polyelectrolytes: Thermodynamic Properties Chandra N. Patra and Arun Yethiraj Department
Biochemistry 675, Lecture 8 Electrostatic Interactions
 Biochemistry 675, Lecture 8 Electrostatic Interactions Previous Classes Hydrophobic Effect Hydrogen Bonds Today: Electrostatic Interactions Reading: Handout:P.R. Bergethon & E.R. Simons, Biophysical Chemistry,
Biochemistry 675, Lecture 8 Electrostatic Interactions Previous Classes Hydrophobic Effect Hydrogen Bonds Today: Electrostatic Interactions Reading: Handout:P.R. Bergethon & E.R. Simons, Biophysical Chemistry,
Bioengineering 215. An Introduction to Molecular Dynamics for Biomolecules
 Bioengineering 215 An Introduction to Molecular Dynamics for Biomolecules David Parker May 18, 2007 ntroduction A principal tool to study biological molecules is molecular dynamics simulations (MD). MD
Bioengineering 215 An Introduction to Molecular Dynamics for Biomolecules David Parker May 18, 2007 ntroduction A principal tool to study biological molecules is molecular dynamics simulations (MD). MD
Polypeptide Folding Using Monte Carlo Sampling, Concerted Rotation, and Continuum Solvation
 Polypeptide Folding Using Monte Carlo Sampling, Concerted Rotation, and Continuum Solvation Jakob P. Ulmschneider and William L. Jorgensen J.A.C.S. 2004, 126, 1849-1857 Presented by Laura L. Thomas and
Polypeptide Folding Using Monte Carlo Sampling, Concerted Rotation, and Continuum Solvation Jakob P. Ulmschneider and William L. Jorgensen J.A.C.S. 2004, 126, 1849-1857 Presented by Laura L. Thomas and
Dielectric Boundary in Biomolecular Solvation Bo Li Department of Mathematics and Center for Theoretical Biological Physics UC San Diego
 Dielectric Boundary in Biomolecular Solvation Bo Li Department of Mathematics and Center for Theoretical Biological Physics UC San Diego International Conference on Free Boundary Problems Newton Institute,
Dielectric Boundary in Biomolecular Solvation Bo Li Department of Mathematics and Center for Theoretical Biological Physics UC San Diego International Conference on Free Boundary Problems Newton Institute,
Lecture 5: Electrostatic Interactions & Screening
 Lecture 5: Electrostatic Interactions & Screening Lecturer: Prof. Brigita Urbanc (brigita@drexel.edu) PHYS 461 & 561, Fall 2009-2010 1 A charged particle (q=+1) in water, at the interface between water
Lecture 5: Electrostatic Interactions & Screening Lecturer: Prof. Brigita Urbanc (brigita@drexel.edu) PHYS 461 & 561, Fall 2009-2010 1 A charged particle (q=+1) in water, at the interface between water
MM-GBSA for Calculating Binding Affinity A rank-ordering study for the lead optimization of Fxa and COX-2 inhibitors
 MM-GBSA for Calculating Binding Affinity A rank-ordering study for the lead optimization of Fxa and COX-2 inhibitors Thomas Steinbrecher Senior Application Scientist Typical Docking Workflow Databases
MM-GBSA for Calculating Binding Affinity A rank-ordering study for the lead optimization of Fxa and COX-2 inhibitors Thomas Steinbrecher Senior Application Scientist Typical Docking Workflow Databases
Numerical Solution of Nonlinear Poisson Boltzmann Equation
 Numerical Solution of Nonlinear Poisson Boltzmann Equation Student: Jingzhen Hu, Southern Methodist University Advisor: Professor Robert Krasny A list of work done extended the numerical solution of nonlinear
Numerical Solution of Nonlinear Poisson Boltzmann Equation Student: Jingzhen Hu, Southern Methodist University Advisor: Professor Robert Krasny A list of work done extended the numerical solution of nonlinear
Modeling in Solution. Important Observations. Energy Concepts
 Modeling in Solution 1 Important Observations Various electrostatic effects for a solvated molecule are often less important than for an isolated gaseous molecule when the molecule is dissolved in a solvent
Modeling in Solution 1 Important Observations Various electrostatic effects for a solvated molecule are often less important than for an isolated gaseous molecule when the molecule is dissolved in a solvent
Biophysical Journal, Volume 99. Supporting Material. Chromatin ionic atmosphere analyzed by a mesoscale electrostatic approach
 Biophysical Journal, Volume 99 Supporting Material Chromatin ionic atmosphere analyzed by a mesoscale electrostatic approach Hin Hark Gan and Tamar Schlick Supporting Material Chromatin ionic atmosphere
Biophysical Journal, Volume 99 Supporting Material Chromatin ionic atmosphere analyzed by a mesoscale electrostatic approach Hin Hark Gan and Tamar Schlick Supporting Material Chromatin ionic atmosphere
Mirror Charges, Hydrogen Bonds, And the Water-Dielectric Interface
 Mirror Charges, ydrogen Bonds, And the Water-Dielectric Interface Kevin Cahill cahill@unm.edu http://dna.phys.unm.edu/ 1 Field Energy The energy E of an electric field E( x, t) is the space integral of
Mirror Charges, ydrogen Bonds, And the Water-Dielectric Interface Kevin Cahill cahill@unm.edu http://dna.phys.unm.edu/ 1 Field Energy The energy E of an electric field E( x, t) is the space integral of
The Molecular Dynamics Method
 The Molecular Dynamics Method Thermal motion of a lipid bilayer Water permeation through channels Selective sugar transport Potential Energy (hyper)surface What is Force? Energy U(x) F = d dx U(x) Conformation
The Molecular Dynamics Method Thermal motion of a lipid bilayer Water permeation through channels Selective sugar transport Potential Energy (hyper)surface What is Force? Energy U(x) F = d dx U(x) Conformation
Supporting Information. The hinge region strengthens the nonspecific interaction. between lac-repressor and DNA: a computer simulation.
 Supporting Information The hinge region strengthens the nonspecific interaction between lac-repressor and DNA: a computer simulation study Lili Sun 1, Marcin Tabaka 1, Sen Hou 1, Lin Li 2, Krzysztof Burdzy
Supporting Information The hinge region strengthens the nonspecific interaction between lac-repressor and DNA: a computer simulation study Lili Sun 1, Marcin Tabaka 1, Sen Hou 1, Lin Li 2, Krzysztof Burdzy
Multimedia : Boundary Lubrication Podcast, Briscoe, et al. Nature , ( )
 3.05 Nanomechanics of Materials and Biomaterials Thursday 04/05/07 Prof. C. Ortiz, MITDMSE I LECTURE 14: TE ELECTRICAL DOUBLE LAYER (EDL) Outline : REVIEW LECTURE #11 : INTRODUCTION TO TE ELECTRICAL DOUBLE
3.05 Nanomechanics of Materials and Biomaterials Thursday 04/05/07 Prof. C. Ortiz, MITDMSE I LECTURE 14: TE ELECTRICAL DOUBLE LAYER (EDL) Outline : REVIEW LECTURE #11 : INTRODUCTION TO TE ELECTRICAL DOUBLE
Colloid Chemistry. La chimica moderna e la sua comunicazione Silvia Gross.
 Colloid Chemistry La chimica moderna e la sua comunicazione Silvia Gross Istituto Dipartimento di Scienze di e Scienze Tecnologie Chimiche Molecolari ISTM-CNR, Università Università degli Studi degli Studi
Colloid Chemistry La chimica moderna e la sua comunicazione Silvia Gross Istituto Dipartimento di Scienze di e Scienze Tecnologie Chimiche Molecolari ISTM-CNR, Università Università degli Studi degli Studi
Homology modeling. Dinesh Gupta ICGEB, New Delhi 1/27/2010 5:59 PM
 Homology modeling Dinesh Gupta ICGEB, New Delhi Protein structure prediction Methods: Homology (comparative) modelling Threading Ab-initio Protein Homology modeling Homology modeling is an extrapolation
Homology modeling Dinesh Gupta ICGEB, New Delhi Protein structure prediction Methods: Homology (comparative) modelling Threading Ab-initio Protein Homology modeling Homology modeling is an extrapolation
Structural Bioinformatics (C3210) Molecular Mechanics
 Structural Bioinformatics (C3210) Molecular Mechanics How to Calculate Energies Calculation of molecular energies is of key importance in protein folding, molecular modelling etc. There are two main computational
Structural Bioinformatics (C3210) Molecular Mechanics How to Calculate Energies Calculation of molecular energies is of key importance in protein folding, molecular modelling etc. There are two main computational
BIOC 530 Fall, 2011 BIOC 530
 Fall, 2011 Course Information Course Description Graduate-level discussion of the structure, function, and chemistry of proteins and nucleic acids, control of enzymatic reactions. Please see the course
Fall, 2011 Course Information Course Description Graduate-level discussion of the structure, function, and chemistry of proteins and nucleic acids, control of enzymatic reactions. Please see the course
Stabilization and Acceleration of Algebraic Multigrid Method
 Stabilization and Acceleration of Algebraic Multigrid Method Recursive Projection Algorithm A. Jemcov J.P. Maruszewski Fluent Inc. October 24, 2006 Outline 1 Need for Algorithm Stabilization and Acceleration
Stabilization and Acceleration of Algebraic Multigrid Method Recursive Projection Algorithm A. Jemcov J.P. Maruszewski Fluent Inc. October 24, 2006 Outline 1 Need for Algorithm Stabilization and Acceleration
An electrokinetic LB based model for ion transport and macromolecular electrophoresis
 An electrokinetic LB based model for ion transport and macromolecular electrophoresis Raffael Pecoroni Supervisor: Michael Kuron July 8, 2016 1 Introduction & Motivation So far an mesoscopic coarse-grained
An electrokinetic LB based model for ion transport and macromolecular electrophoresis Raffael Pecoroni Supervisor: Michael Kuron July 8, 2016 1 Introduction & Motivation So far an mesoscopic coarse-grained
1. Poisson-Boltzmann 1.1. Poisson equation. We consider the Laplacian. which is given in spherical coordinates by (2)
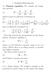 1. Poisson-Boltzmann 1.1. Poisson equation. We consider the Laplacian operator (1) 2 = 2 x + 2 2 y + 2 2 z 2 which is given in spherical coordinates by (2) 2 = 1 ( r 2 ) + 1 r 2 r r r 2 sin θ θ and in
1. Poisson-Boltzmann 1.1. Poisson equation. We consider the Laplacian operator (1) 2 = 2 x + 2 2 y + 2 2 z 2 which is given in spherical coordinates by (2) 2 = 1 ( r 2 ) + 1 r 2 r r r 2 sin θ θ and in
SECOND PUBLIC EXAMINATION. Honour School of Physics Part C: 4 Year Course. Honour School of Physics and Philosophy Part C C7: BIOLOGICAL PHYSICS
 2757 SECOND PUBLIC EXAMINATION Honour School of Physics Part C: 4 Year Course Honour School of Physics and Philosophy Part C C7: BIOLOGICAL PHYSICS TRINITY TERM 2013 Monday, 17 June, 2.30 pm 5.45 pm 15
2757 SECOND PUBLIC EXAMINATION Honour School of Physics Part C: 4 Year Course Honour School of Physics and Philosophy Part C C7: BIOLOGICAL PHYSICS TRINITY TERM 2013 Monday, 17 June, 2.30 pm 5.45 pm 15
Ranjit P. Bahadur Assistant Professor Department of Biotechnology Indian Institute of Technology Kharagpur, India. 1 st November, 2013
 Hydration of protein-rna recognition sites Ranjit P. Bahadur Assistant Professor Department of Biotechnology Indian Institute of Technology Kharagpur, India 1 st November, 2013 Central Dogma of life DNA
Hydration of protein-rna recognition sites Ranjit P. Bahadur Assistant Professor Department of Biotechnology Indian Institute of Technology Kharagpur, India 1 st November, 2013 Central Dogma of life DNA
Improved Solvation Models using Boundary Integral Equations
 Improved Solvation Models using Boundary Integral Equations Matthew Knepley and Jaydeep Bardhan Computational and Applied Mathematics Rice University SIAM Conference on the Life Sciences Minneapolis, MN
Improved Solvation Models using Boundary Integral Equations Matthew Knepley and Jaydeep Bardhan Computational and Applied Mathematics Rice University SIAM Conference on the Life Sciences Minneapolis, MN
Molecular dynamics simulation. CS/CME/BioE/Biophys/BMI 279 Oct. 5 and 10, 2017 Ron Dror
 Molecular dynamics simulation CS/CME/BioE/Biophys/BMI 279 Oct. 5 and 10, 2017 Ron Dror 1 Outline Molecular dynamics (MD): The basic idea Equations of motion Key properties of MD simulations Sample applications
Molecular dynamics simulation CS/CME/BioE/Biophys/BMI 279 Oct. 5 and 10, 2017 Ron Dror 1 Outline Molecular dynamics (MD): The basic idea Equations of motion Key properties of MD simulations Sample applications
The AGBNP2 Implicit Solvation Model
 2544 J. Chem. Theory Comput. 2009, 5, 2544 2564 The AGBNP2 Implicit Solvation Model Emilio Gallicchio,* Kristina Paris, and Ronald M. Levy Department of Chemistry and Chemical Biology and BioMaPS Institute
2544 J. Chem. Theory Comput. 2009, 5, 2544 2564 The AGBNP2 Implicit Solvation Model Emilio Gallicchio,* Kristina Paris, and Ronald M. Levy Department of Chemistry and Chemical Biology and BioMaPS Institute
Solutions and Non-Covalent Binding Forces
 Chapter 3 Solutions and Non-Covalent Binding Forces 3.1 Solvent and solution properties Molecules stick together using the following forces: dipole-dipole, dipole-induced dipole, hydrogen bond, van der
Chapter 3 Solutions and Non-Covalent Binding Forces 3.1 Solvent and solution properties Molecules stick together using the following forces: dipole-dipole, dipole-induced dipole, hydrogen bond, van der
Error Analysis of the Poisson P 3 MForce Field Scheme for Particle-Based Simulations of Biological Systems
 Journal of Computational Electronics 4: 179 183, 2005 c 2005 Springer Science + Business Media, Inc. Manufactured in The Netherlands. Error Analysis of the Poisson P 3 MForce Field Scheme for Particle-Based
Journal of Computational Electronics 4: 179 183, 2005 c 2005 Springer Science + Business Media, Inc. Manufactured in The Netherlands. Error Analysis of the Poisson P 3 MForce Field Scheme for Particle-Based
8.592J HST.452J: Statistical Physics in Biology
 Assignment # 4 8.592J HST.452J: Statistical Physics in Biology Coulomb Interactions 1. Flory Theory: The Coulomb energy of a ball of charge Q and dimension R in d spacial dimensions scales as Q 2 E c.
Assignment # 4 8.592J HST.452J: Statistical Physics in Biology Coulomb Interactions 1. Flory Theory: The Coulomb energy of a ball of charge Q and dimension R in d spacial dimensions scales as Q 2 E c.
arxiv: v1 [q-bio.bm] 6 Apr 2016
![arxiv: v1 [q-bio.bm] 6 Apr 2016 arxiv: v1 [q-bio.bm] 6 Apr 2016](/thumbs/88/115645706.jpg) Multi-shell model of ion-induced nucleic acid condensation Igor S. Tolokh Department of Computer Science, Virginia Tech, Blacksburg, VA 24061, USA Aleksander Drozdetski Department of Physics, Virginia
Multi-shell model of ion-induced nucleic acid condensation Igor S. Tolokh Department of Computer Science, Virginia Tech, Blacksburg, VA 24061, USA Aleksander Drozdetski Department of Physics, Virginia
AD-A jlmainpg M o 7408
 OTIC FILE COPY Q AD-A218 125 jlmainpg M o 7408 A D A2 8 lb RESTRICTIVE MARKINGS '1 IT NA 2a. SECURITY CLASSIFICATION AU 3. DISTRIBUTION /AVAILABILITY OF REPORT NA Form Approved 2b. DECLASSIFICATION/DOWNGRI
OTIC FILE COPY Q AD-A218 125 jlmainpg M o 7408 A D A2 8 lb RESTRICTIVE MARKINGS '1 IT NA 2a. SECURITY CLASSIFICATION AU 3. DISTRIBUTION /AVAILABILITY OF REPORT NA Form Approved 2b. DECLASSIFICATION/DOWNGRI
F. Piazza Center for Molecular Biophysics and University of Orléans, France. Selected topic in Physical Biology. Lecture 1
 Zhou Pei-Yuan Centre for Applied Mathematics, Tsinghua University November 2013 F. Piazza Center for Molecular Biophysics and University of Orléans, France Selected topic in Physical Biology Lecture 1
Zhou Pei-Yuan Centre for Applied Mathematics, Tsinghua University November 2013 F. Piazza Center for Molecular Biophysics and University of Orléans, France Selected topic in Physical Biology Lecture 1
Effective Born Radii in the Generalized Born Approximation: The Importance of Being Perfect
 Effective Born Radii in the Generalized Born Approximation: The Importance of Being Perfect ALEXEY ONUFRIEV, DAVID A. CASE, DONALD BASHFORD Department of Molecular Biology, The Scripps Research Institute,
Effective Born Radii in the Generalized Born Approximation: The Importance of Being Perfect ALEXEY ONUFRIEV, DAVID A. CASE, DONALD BASHFORD Department of Molecular Biology, The Scripps Research Institute,
Chapter 1. Topic: Overview of basic principles
 Chapter 1 Topic: Overview of basic principles Four major themes of biochemistry I. What are living organism made from? II. How do organism acquire and use energy? III. How does an organism maintain its
Chapter 1 Topic: Overview of basic principles Four major themes of biochemistry I. What are living organism made from? II. How do organism acquire and use energy? III. How does an organism maintain its
Coupling the Level-Set Method with Variational Implicit Solvent Modeling of Molecular Solvation
 Coupling the Level-Set Method with Variational Implicit Solvent Modeling of Molecular Solvation Bo Li Math Dept & CTBP, UCSD Li-Tien Cheng (Math, UCSD) Zhongming Wang (Math & Biochem, UCSD) Yang Xie (MAE,
Coupling the Level-Set Method with Variational Implicit Solvent Modeling of Molecular Solvation Bo Li Math Dept & CTBP, UCSD Li-Tien Cheng (Math, UCSD) Zhongming Wang (Math & Biochem, UCSD) Yang Xie (MAE,
Classical Models of the Interface between an Electrode and Electrolyte. M.Sc. Ekaterina Gongadze
 Presented at the COMSOL Conference 009 Milan Classical Models of the Interface between an Electrode and Electrolyte M.Sc. Ekaterina Gongadze Faculty of Informatics and Electrical Engineering Comsol Conference
Presented at the COMSOL Conference 009 Milan Classical Models of the Interface between an Electrode and Electrolyte M.Sc. Ekaterina Gongadze Faculty of Informatics and Electrical Engineering Comsol Conference
RESEARCH ARTICLES. Effective Energy Function for Proteins in Solution
 PROTEINS: Structure, Function, and Genetics 35:133 152 (1999) RESEARCH ARTICLES Effective Energy Function for Proteins in Solution Themis Lazaridis 1 and Martin Karplus 1,2 * 1 Department of Chemistry
PROTEINS: Structure, Function, and Genetics 35:133 152 (1999) RESEARCH ARTICLES Effective Energy Function for Proteins in Solution Themis Lazaridis 1 and Martin Karplus 1,2 * 1 Department of Chemistry
ProteinMolecularDynamicsWithElectrostaticForce EntirelyDeterminedbyaSinglePoisson-Boltzmann Calculation
 PROTEINS: Structure, Function, and Genetics 48:497 504 (2002) ProteinMolecularDynamicsWithElectrostaticForce EntirelyDeterminedbyaSinglePoisson-Boltzmann Calculation BenZhuoLu, 1,2 WeiZuChen, 2 CunXinWang,
PROTEINS: Structure, Function, and Genetics 48:497 504 (2002) ProteinMolecularDynamicsWithElectrostaticForce EntirelyDeterminedbyaSinglePoisson-Boltzmann Calculation BenZhuoLu, 1,2 WeiZuChen, 2 CunXinWang,
Exploring the Sequence Dependent Structure and Dynamics of DNA with Molecular Dynamics Simulation
 Exploring the Sequence Dependent Structure and Dynamics of DNA with Molecular Dynamics Simulation Sarah Harris School of Physics and Astronomy University of Leeds Introduction Calculations of the charge
Exploring the Sequence Dependent Structure and Dynamics of DNA with Molecular Dynamics Simulation Sarah Harris School of Physics and Astronomy University of Leeds Introduction Calculations of the charge
Principles of Physical Biochemistry
 Principles of Physical Biochemistry Kensal E. van Hold e W. Curtis Johnso n P. Shing Ho Preface x i PART 1 MACROMOLECULAR STRUCTURE AND DYNAMICS 1 1 Biological Macromolecules 2 1.1 General Principles
Principles of Physical Biochemistry Kensal E. van Hold e W. Curtis Johnso n P. Shing Ho Preface x i PART 1 MACROMOLECULAR STRUCTURE AND DYNAMICS 1 1 Biological Macromolecules 2 1.1 General Principles
Why Proteins Fold? (Parts of this presentation are based on work of Ashok Kolaskar) CS490B: Introduction to Bioinformatics Mar.
 Why Proteins Fold? (Parts of this presentation are based on work of Ashok Kolaskar) CS490B: Introduction to Bioinformatics Mar. 25, 2002 Molecular Dynamics: Introduction At physiological conditions, the
Why Proteins Fold? (Parts of this presentation are based on work of Ashok Kolaskar) CS490B: Introduction to Bioinformatics Mar. 25, 2002 Molecular Dynamics: Introduction At physiological conditions, the
Non-bonded interactions
 speeding up the number-crunching continued Marcus Elstner and Tomáš Kubař December 3, 2013 why care? number of individual pair-wise interactions bonded interactions proportional to N: O(N) non-bonded interactions
speeding up the number-crunching continued Marcus Elstner and Tomáš Kubař December 3, 2013 why care? number of individual pair-wise interactions bonded interactions proportional to N: O(N) non-bonded interactions
Module 8: "Stability of Colloids" Lecture 38: "" The Lecture Contains: Calculation for CCC (n c )
 The Lecture Contains: Calculation for CCC (n c ) Relation between surface charge and electrostatic potential Extensions to DLVO theory file:///e /courses/colloid_interface_science/lecture38/38_1.htm[6/16/2012
The Lecture Contains: Calculation for CCC (n c ) Relation between surface charge and electrostatic potential Extensions to DLVO theory file:///e /courses/colloid_interface_science/lecture38/38_1.htm[6/16/2012
Macroscopic modeling and simulations of supercoiled DNA with bound proteins
 JOURNAL OF CHEMICAL PHYSICS VOLUME 117, NUMBER 18 8 NOVEMBER 2002 Macroscopic modeling and simulations of supercoiled DNA with bound proteins Jing Huang a) and Tamar Schlick b) Department of Chemistry
JOURNAL OF CHEMICAL PHYSICS VOLUME 117, NUMBER 18 8 NOVEMBER 2002 Macroscopic modeling and simulations of supercoiled DNA with bound proteins Jing Huang a) and Tamar Schlick b) Department of Chemistry
G pol = 1 2 where ρ( x) is the charge density at position x and the reaction electrostatic potential
 EFFICIENT AND ACCURATE HIGHER-ORDER FAST MULTIPOLE BOUNDARY ELEMENT METHOD FOR POISSON BOLTZMANN ELECTROSTATICS CHANDRAJIT BAJAJ SHUN-CHUAN CHEN Abstract. The Poisson-Boltzmann equation is a partial differential
EFFICIENT AND ACCURATE HIGHER-ORDER FAST MULTIPOLE BOUNDARY ELEMENT METHOD FOR POISSON BOLTZMANN ELECTROSTATICS CHANDRAJIT BAJAJ SHUN-CHUAN CHEN Abstract. The Poisson-Boltzmann equation is a partial differential
The change in free energy on transferring an ion from a medium of low dielectric constantε1 to one of high dielectric constant ε2:
 The Born Energy of an Ion The free energy density of an electric field E arising from a charge is ½(ε 0 ε E 2 ) per unit volume Integrating the energy density of an ion over all of space = Born energy:
The Born Energy of an Ion The free energy density of an electric field E arising from a charge is ½(ε 0 ε E 2 ) per unit volume Integrating the energy density of an ion over all of space = Born energy:
Molecular Interactions F14NMI. Lecture 4: worked answers to practice questions
 Molecular Interactions F14NMI Lecture 4: worked answers to practice questions http://comp.chem.nottingham.ac.uk/teaching/f14nmi jonathan.hirst@nottingham.ac.uk (1) (a) Describe the Monte Carlo algorithm
Molecular Interactions F14NMI Lecture 4: worked answers to practice questions http://comp.chem.nottingham.ac.uk/teaching/f14nmi jonathan.hirst@nottingham.ac.uk (1) (a) Describe the Monte Carlo algorithm
Variational Implicit Solvation of Biomolecules: From Theory to Numerical Computations
 Variational Implicit Solvation of Biomolecules: From Theory to Numerical Computations Bo Li Department of Mathematics and Center for Theoretical Biological Physics UC San Diego CECAM Workshop: New Perspectives
Variational Implicit Solvation of Biomolecules: From Theory to Numerical Computations Bo Li Department of Mathematics and Center for Theoretical Biological Physics UC San Diego CECAM Workshop: New Perspectives
Computer simulation methods (2) Dr. Vania Calandrini
 Computer simulation methods (2) Dr. Vania Calandrini in the previous lecture: time average versus ensemble average MC versus MD simulations equipartition theorem (=> computing T) virial theorem (=> computing
Computer simulation methods (2) Dr. Vania Calandrini in the previous lecture: time average versus ensemble average MC versus MD simulations equipartition theorem (=> computing T) virial theorem (=> computing
Laplacian-Centered Poisson Solvers and Multilevel Summation Algorithms
 Laplacian-Centered Poisson Solvers and Multilevel Summation Algorithms Dmitry Yershov 1 Stephen Bond 1 Robert Skeel 2 1 University of Illinois at Urbana-Champaign 2 Purdue University 2009 SIAM Annual Meeting
Laplacian-Centered Poisson Solvers and Multilevel Summation Algorithms Dmitry Yershov 1 Stephen Bond 1 Robert Skeel 2 1 University of Illinois at Urbana-Champaign 2 Purdue University 2009 SIAM Annual Meeting
Molecular Forces in Biological Systems - Electrostatic Interactions; - Shielding of charged objects in solution
 Molecular Forces in Biological Systems - Electrostatic Interactions; - Shielding of charged objects in solution Electrostatic self-energy, effects of size and dielectric constant q q r r ε ε 1 ε 2? δq
Molecular Forces in Biological Systems - Electrostatic Interactions; - Shielding of charged objects in solution Electrostatic self-energy, effects of size and dielectric constant q q r r ε ε 1 ε 2? δq
Analysis of the Stability of Looped-Out and Stacked-In Conformations of an Adenine Bulge in DNA Using a Continuum Model for Solvent and Ions
 2990 Biophysical Journal Volume 73 December 1997 2990-3003 Analysis of the Stability of Looped-Out and Stacked-In Conformations of an Adenine Bulge in DNA Using a Continuum Model for Solvent and Ions Martin
2990 Biophysical Journal Volume 73 December 1997 2990-3003 Analysis of the Stability of Looped-Out and Stacked-In Conformations of an Adenine Bulge in DNA Using a Continuum Model for Solvent and Ions Martin
Roles of Boundary Conditions in DNA Simulations: Analysis of Ion Distributions with the Finite-Difference Poisson-Boltzmann Method
 554 Biophysical Journal Volume 97 July 2009 554 562 Roles of Boundary Conditions in DNA Simulations: Analysis of Ion Distributions with the Finite-Difference Poisson-Boltzmann Method Xiang Ye, Qin Cai,
554 Biophysical Journal Volume 97 July 2009 554 562 Roles of Boundary Conditions in DNA Simulations: Analysis of Ion Distributions with the Finite-Difference Poisson-Boltzmann Method Xiang Ye, Qin Cai,
Electrolyte Solutions
 Chapter 8 Electrolyte Solutions In the last few chapters of this book, we will deal with several specific types of chemical systems. The first one is solutions and equilibria involving electrolytes, which
Chapter 8 Electrolyte Solutions In the last few chapters of this book, we will deal with several specific types of chemical systems. The first one is solutions and equilibria involving electrolytes, which
1044 Lecture #14 of 18
 Lecture #14 of 18 1044 1045 Q: What s in this set of lectures? A: B&F Chapter 13 main concepts: Section 1.2.3: Diffuse double layer structure Sections 13.1 & 13.2: Gibbs adsorption isotherm; Electrocapillary
Lecture #14 of 18 1044 1045 Q: What s in this set of lectures? A: B&F Chapter 13 main concepts: Section 1.2.3: Diffuse double layer structure Sections 13.1 & 13.2: Gibbs adsorption isotherm; Electrocapillary
Alek Marenich Cramer -Truhlar Group. University of Minnesota
 and Computational Electrochemistry Alek Marenich Cramer -Truhlar Group University of Minnesota Outline Solvation and charge models Energetics and dynamics in solution Computational electrochemistry Future
and Computational Electrochemistry Alek Marenich Cramer -Truhlar Group University of Minnesota Outline Solvation and charge models Energetics and dynamics in solution Computational electrochemistry Future
Introduction to Polymer Physics
 Introduction to Polymer Physics Enrico Carlon, KU Leuven, Belgium February-May, 2016 Enrico Carlon, KU Leuven, Belgium Introduction to Polymer Physics February-May, 2016 1 / 28 Polymers in Chemistry and
Introduction to Polymer Physics Enrico Carlon, KU Leuven, Belgium February-May, 2016 Enrico Carlon, KU Leuven, Belgium Introduction to Polymer Physics February-May, 2016 1 / 28 Polymers in Chemistry and
Lecture 3 Charged interfaces
 Lecture 3 Charged interfaces rigin of Surface Charge Immersion of some materials in an electrolyte solution. Two mechanisms can operate. (1) Dissociation of surface sites. H H H H H M M M +H () Adsorption
Lecture 3 Charged interfaces rigin of Surface Charge Immersion of some materials in an electrolyte solution. Two mechanisms can operate. (1) Dissociation of surface sites. H H H H H M M M +H () Adsorption
Statistical Theory and Learning from Molecular Simulations
 Statistical Theory and Learning from Molecular Simulations Lawrence R. Pratt 1 and Susan B. Rempe 2 1 Department of Chemical & Biomolecular Engineering 2 Center for Biological and Material Sciences, Sandia
Statistical Theory and Learning from Molecular Simulations Lawrence R. Pratt 1 and Susan B. Rempe 2 1 Department of Chemical & Biomolecular Engineering 2 Center for Biological and Material Sciences, Sandia
Structuring of hydrophobic and hydrophilic polymers at interfaces Stephen Donaldson ChE 210D Final Project Abstract
 Structuring of hydrophobic and hydrophilic polymers at interfaces Stephen Donaldson ChE 210D Final Project Abstract In this work, a simplified Lennard-Jones (LJ) sphere model is used to simulate the aggregation,
Structuring of hydrophobic and hydrophilic polymers at interfaces Stephen Donaldson ChE 210D Final Project Abstract In this work, a simplified Lennard-Jones (LJ) sphere model is used to simulate the aggregation,
Molecular dynamics simulation of Aquaporin-1. 4 nm
 Molecular dynamics simulation of Aquaporin-1 4 nm Molecular Dynamics Simulations Schrödinger equation i~@ t (r, R) =H (r, R) Born-Oppenheimer approximation H e e(r; R) =E e (R) e(r; R) Nucleic motion described
Molecular dynamics simulation of Aquaporin-1 4 nm Molecular Dynamics Simulations Schrödinger equation i~@ t (r, R) =H (r, R) Born-Oppenheimer approximation H e e(r; R) =E e (R) e(r; R) Nucleic motion described
Introduction to molecular dynamics
 1 Introduction to molecular dynamics Yves Lansac Université François Rabelais, Tours, France Visiting MSE, GIST for the summer Molecular Simulation 2 Molecular simulation is a computational experiment.
1 Introduction to molecular dynamics Yves Lansac Université François Rabelais, Tours, France Visiting MSE, GIST for the summer Molecular Simulation 2 Molecular simulation is a computational experiment.
