Supplementary Materials: Probing the Ion Binding Site in a DNA Holliday Junction Using Förster Resonance Energy Transfer (FRET)
|
|
|
- Thomas Parks
- 6 years ago
- Views:
Transcription
1 S1 of S5 Supplementary Materials: Probing the Ion Binding Site in a DNA Holliday Junction Using Förster Resonance Energy Transfer (FRET) Jacob L. Litke, Yan Li, Laura M. Nocka and Ishita Mukerji (a) (b) Figure S1. (a) Interduplex angle determined from FRET measured distances as described in the text. In general we observed for both monovalent and polyvalent ions that the interduplex angle was proportional to the hydrated ion radius. The experimentally determined distances and associated interduplex angles with errors are given in Table S1; (b) Model of a stacked Holliday junction with an IDA of 45. On the left the distance between the C5 atoms of the 17th bp from the junction center of two co-axially stacked arms is used to estimate the distance between dyes in the isoi conformation (113 Å). Right: Distances are shown from the top of the arm to the 17th bp on both strands (53 and 57 Å). Also shown are the distances from the 17th bp of one arm to the 17th bp of the adjacent arm (58 and 66 Å) to estimate the effect of helix position on the distances determined. In both cases, we find that the difference in distance introduced by the position on the helix is smaller than the error in the distance reported from the relative mobility of the dyes. All distances are measured at the C5 atom.
2 S2 of S5 Figure S2. Analysis of Mg 2+ :4WJ binding interaction using a model of two identical, non-interacting binding sites (α = n k[mg 2+ ]/(1 + k[mg 2+ ]) where α = fraction folded, n is the number of binding sites and k is the microscopic association constant). Data shown are taken from Figure 1B in the manuscript. This fitting model gave comparable Ka values to the apparent Ka values obtained from fitting the data to Equation (5) in the manuscript (59,400 M 1 vs. 45,400 M 1 ). We note that in order to obtain the two-site binding fit, the number of binding sites needed to be constrained to 2. (Left): Data and fit are shown plotted on a log scale; (Right): Data and fit are shown plotted on a linear scale. (a) (b) Figure S3. Representative fits of the ITC data to one binding component. (a) Fitting of the ITC data to a one-to-one binding model where all parameters were allowed to vary; (b) Fitting of the ITC data to a one-to-one binding model, where Ka and ΔH were held fixed to the van t Hoff determined parameters of 40,181 M 1 and 14.9 kcal/mol, respectively. Residuals depicted are generated between the actual data and the fit shown. All parameters of the fits are given in Table S2. Generally, χ 2 -values and visual inspection of the fit and the residuals indicated that another component was needed to adequately describe the data.
3 S3 of S C C C C Mg 2+ (μm) Figure S4. Energy transfer efficiencies measured for the Mg 2+ -junction interaction measured at 4, 20, 25, and 30 C. The Ka values were determined as described in the text and used to determine ΔH and ΔS values for the ion binding reaction. All titrations were carried out in a 0.5 mm Na + and 10 mm Tris HCl, ph 7.4 buffer.
4 S4 of S5 Figure S5. Tb 3+ binding to 4WJ DNA and 34 bp duplex DNA measured by DNA-sensitized emission at 543 nm. Experimental parameters are the same as for Figure 4. Both data sets were well described by a simple two-state model for ion binding (Equation (5)). The ion binding affinity to the duplex is weaker than ion binding affinity to the junction. However, it is likely that some non-specific ion binding to the junction arms is reflected in the curve shown above and in Figure 4. Figure S6. Excitation curve of the 2 F0 5 D2 transition of aqueous Eu 3+ at 64 µm, with 6.3 µm junction. Lifetime measurements were performed at the peak wavelength (464.4 nm), and also at two other wavelengths (464.1 and nm). Use of this transition allowed for sufficient intensity for lifetime measurements. Figure S7. Excitation spectrum of Eu 3+ luminescence. Titration of Eu 3+ into a solution of 8.8 µm junction verifies a junction-bound peak at nm. No further changes in Eu 3+ luminescence were observed after a ratio of 16:1, Eu 3+ :4WJ was reached.
5 S5 of S5 Table S1. FRET efficiencies and calculated junction angles for different ions. Ion Kd (app) a R (Å) b IDA ( ) b Ion Radius (pm) c Minimal ion concentration 80 ± ± 20 Na + 29 ± 4 mm a 48 ± 9 45 ± K + 5 ± 1 mm a 59 ± ± Mg ± 8 µm 53 ± ± Ca ± 8 µm 56 ± ± Eu 3+ 3 ± 2 µm 57 ± ± Tb µm 56 ± ± Nd µm 58 ± ± a The Kd (app) values were taken from Vitoc and Mukerji [16]. The other Kd (app) values were determined from the data shown in Figure 1; b Distances and interduplex angles were determined as described in the text. The calculated error on the distances included the experimental errors in measuring the efficiencies and consideration of the minimal and maximal values of κ 2 based on anisotropy measurements as described previously [25]; c Hydrated ion radii were taken from Ohtaki and Radnai [26]. Table S2. Representative fits of the ITC data with one and two components. Binding Site a Binding Site b Binding Site c Binding Sites b Binding Sites a Binding Sites c χ ,000 11, n (sites) 4.9 ± ± ± ± Ka (M 1 ) (1.4 ± 0.5) ,181 19,000 ± , ,000 ± 340, ± 7600 ΔH (kcal/mol) 0.8 ± ± ± ± 13.3 ΔS (cal/( mol)) n2 (sites) 15.9 ± ± ± 5.0 Ka,2 (M 1 ) (2.5 ± 0.3) 10 4 (1.1 ± 2.1) 10 6 (1.4 ± 0.4) 10 4 ΔH2 (kcal/mol) 1.9 ± ± ± 1.6 ΔS2 (cal/( mol)) a All fitting parameters (n, Ka, ΔH) were allowed to vary freely in the fit; b In these fits, the parameters Ka,1 and ΔH1 were held fixed to parameters independently determined from a van t Hoff analysis of temperature dependent FRET data as described in the text. All other fitting parameters (n1, n2, Ka,2, ΔH2) were allowed to vary freely; c In these fits the n1 parameter was held fixed at 1, all other parameters (n2, Ka,1, Ka,2, ΔH1, ΔH2) were allowed to vary freely.
Supplementary Information. The Solution Structural Ensembles of RNA Kink-turn Motifs and Their Protein Complexes
 Supplementary Information The Solution Structural Ensembles of RNA Kink-turn Motifs and Their Protein Complexes Xuesong Shi, a Lin Huang, b David M. J. Lilley, b Pehr B. Harbury a,c and Daniel Herschlag
Supplementary Information The Solution Structural Ensembles of RNA Kink-turn Motifs and Their Protein Complexes Xuesong Shi, a Lin Huang, b David M. J. Lilley, b Pehr B. Harbury a,c and Daniel Herschlag
Supplemental Materials and Methods
 Supplemental Materials and Methods Time-resolved FRET (trfret) to probe for changes in the Box A/A stem upon complex assembly U3 MINI was folded and the decay of Fl fluorescence was measured at 20 ºC (see
Supplemental Materials and Methods Time-resolved FRET (trfret) to probe for changes in the Box A/A stem upon complex assembly U3 MINI was folded and the decay of Fl fluorescence was measured at 20 ºC (see
According to the manufacture s direction (Pierce), RNA and DNA
 Supplementary method Electrophoretic Mobility-shift assay (EMSA) According to the manufacture s direction (Pierce), RNA and DNA oligonuleotides were firstly labeled by biotin. TAVb (1pM) was incubated
Supplementary method Electrophoretic Mobility-shift assay (EMSA) According to the manufacture s direction (Pierce), RNA and DNA oligonuleotides were firstly labeled by biotin. TAVb (1pM) was incubated
A rule of seven in Watson-Crick base-pairing of mismatched sequences
 A rule of seven in Watson-Crick base-pairing of mismatched sequences Ibrahim I. Cisse 1,3, Hajin Kim 1,2, Taekjip Ha 1,2 1 Department of Physics and Center for the Physics of Living Cells, University of
A rule of seven in Watson-Crick base-pairing of mismatched sequences Ibrahim I. Cisse 1,3, Hajin Kim 1,2, Taekjip Ha 1,2 1 Department of Physics and Center for the Physics of Living Cells, University of
Problem solving steps
 Problem solving steps Determine the reaction Write the (balanced) equation ΔG K v Write the equilibrium constant v Find the equilibrium constant using v If necessary, solve for components K K = [ p ] ν
Problem solving steps Determine the reaction Write the (balanced) equation ΔG K v Write the equilibrium constant v Find the equilibrium constant using v If necessary, solve for components K K = [ p ] ν
Simulative and experimental characterization of a ph-dependent
 Simulative and experimental characterization of a ph-dependent clamp-like DNA triple-helix nanoswitch Federico Iacovelli, # Andrea Idili, # Alessandro Benincasa, Davide Mariottini, Alessio Ottaviani, Mattia
Simulative and experimental characterization of a ph-dependent clamp-like DNA triple-helix nanoswitch Federico Iacovelli, # Andrea Idili, # Alessandro Benincasa, Davide Mariottini, Alessio Ottaviani, Mattia
Supplementary Figures
 1 Supplementary Figures Supplementary Figure 1 Type I FGFR1 inhibitors (a) Chemical structures of a pyrazolylaminopyrimidine inhibitor (henceforth referred to as PAPI; PDB-code of the FGFR1-PAPI complex:
1 Supplementary Figures Supplementary Figure 1 Type I FGFR1 inhibitors (a) Chemical structures of a pyrazolylaminopyrimidine inhibitor (henceforth referred to as PAPI; PDB-code of the FGFR1-PAPI complex:
Self-assembled Nanoscale DNA-porphyrin Complex for. Artificial Light-harvesting
 Supporting Information for Self-assembled Nanoscale DNA-porphyrin Complex for Artificial Light-harvesting Jakob G. Woller, Jonas K. Hannestad, and Bo Albinsson Department of Chemical and Biological Engineering/Physical
Supporting Information for Self-assembled Nanoscale DNA-porphyrin Complex for Artificial Light-harvesting Jakob G. Woller, Jonas K. Hannestad, and Bo Albinsson Department of Chemical and Biological Engineering/Physical
The ph-responsive behaviour of aqueous solutions of poly(acrylic acid) is dependent on molar mass
 Electronic Supplementary Material (ESI) for Soft Matter. This journal is The Royal Society of Chemistry 2016 The ph-responsive behaviour of aqueous solutions of poly(acrylic acid) is dependent on molar
Electronic Supplementary Material (ESI) for Soft Matter. This journal is The Royal Society of Chemistry 2016 The ph-responsive behaviour of aqueous solutions of poly(acrylic acid) is dependent on molar
Fluorescence Resonance Energy Transfer (FRET) Microscopy
 Fluorescence Resonance Energy Transfer () Microscopy Mike Lorenz Optical Technology Development mlorenz@mpi-cbg.de -FLM course, May 2009 What is fluorescence? Stoke s shift Fluorescence light is always
Fluorescence Resonance Energy Transfer () Microscopy Mike Lorenz Optical Technology Development mlorenz@mpi-cbg.de -FLM course, May 2009 What is fluorescence? Stoke s shift Fluorescence light is always
Reconfigurable DNA Origami Nanocapsule for ph- Controlled Encapsulation and Display of Cargo
 Supporting Information Reconfigurable DNA Origami Nanocapsule for ph- Controlled Encapsulation and Display of Cargo Heini Ijäs, Iiris Hakaste, Boxuan Shen, Mauri A. Kostiainen, and Veikko Linko* Biohybrid
Supporting Information Reconfigurable DNA Origami Nanocapsule for ph- Controlled Encapsulation and Display of Cargo Heini Ijäs, Iiris Hakaste, Boxuan Shen, Mauri A. Kostiainen, and Veikko Linko* Biohybrid
Analysis of nucleotide binding to p97 reveals the properties of a tandem AAA hexameric ATPase
 SUPPLEMENTARY INFORMATION Analysis of nucleotide binding to p97 reveals the properties of a tandem AAA hexameric ATPase Louise C Briggs, Geoff S Baldwin, Non Miyata, Hisao Kondo, Xiaodong Zhang, Paul S
SUPPLEMENTARY INFORMATION Analysis of nucleotide binding to p97 reveals the properties of a tandem AAA hexameric ATPase Louise C Briggs, Geoff S Baldwin, Non Miyata, Hisao Kondo, Xiaodong Zhang, Paul S
Supporting Information
 Supporting Information To Engineered Holliday Junctions as Single-Molecule Reporters for Protein-DNA Interactions with Application to a MerR-family Regulator Susanta K. Sarkar, a Nesha May Andoy, a Jaime
Supporting Information To Engineered Holliday Junctions as Single-Molecule Reporters for Protein-DNA Interactions with Application to a MerR-family Regulator Susanta K. Sarkar, a Nesha May Andoy, a Jaime
Substrate-dependent switching of the allosteric binding mechanism of a dimeric enzyme
 Supplementary Information: Substrate-dependent switching of the allosteric binding mechanism of a dimeric enzyme Lee Freiburger, 1 Teresa Miletti, 1 Siqi Zhu, 1 Oliver Baettig, Albert Berghuis, Karine
Supplementary Information: Substrate-dependent switching of the allosteric binding mechanism of a dimeric enzyme Lee Freiburger, 1 Teresa Miletti, 1 Siqi Zhu, 1 Oliver Baettig, Albert Berghuis, Karine
Single-Molecule Methods I - in vitro
 Single-Molecule Methods I - in vitro Bo Huang Macromolecules 2014.03.10 F 1 -ATPase: a case study Membrane ADP ATP Rotation of the axle when hydrolyzing ATP Kinosita group, 1997-2005 Single Molecule Methods
Single-Molecule Methods I - in vitro Bo Huang Macromolecules 2014.03.10 F 1 -ATPase: a case study Membrane ADP ATP Rotation of the axle when hydrolyzing ATP Kinosita group, 1997-2005 Single Molecule Methods
Thermodynamics of Ion-Induced RNA Folding in the Hammerhead Ribozyme: An Isothermal Titration Calorimetric Study
 Biochemistry 2001, 40, 1423-1429 1423 Thermodynamics of Ion-Induced RNA Folding in the Hammerhead Ribozyme: An Isothermal Titration Calorimetric Study Christian Hammann, Alan Cooper, and David M. J. Lilley*,
Biochemistry 2001, 40, 1423-1429 1423 Thermodynamics of Ion-Induced RNA Folding in the Hammerhead Ribozyme: An Isothermal Titration Calorimetric Study Christian Hammann, Alan Cooper, and David M. J. Lilley*,
Supporting Information
 Supporting Information Arai et al. 10.1073/pnas.15179911 SI Text Protein Expression and Purification. Myb3 (mouse, residues 84 315) was expressed in Escherichia coli as a fusion with the B1 domain of protein
Supporting Information Arai et al. 10.1073/pnas.15179911 SI Text Protein Expression and Purification. Myb3 (mouse, residues 84 315) was expressed in Escherichia coli as a fusion with the B1 domain of protein
Supporting Information for:
 Supporting Information for: A Simple Molecular Model for Thermophilic Adaptation of Functional Nucleic Acids Joshua M. Blose *, Scott K. Silverman, and Philip C. Bevilacqua * * Department of Chemistry,
Supporting Information for: A Simple Molecular Model for Thermophilic Adaptation of Functional Nucleic Acids Joshua M. Blose *, Scott K. Silverman, and Philip C. Bevilacqua * * Department of Chemistry,
Supplementary Materials: Localization and Spectroscopic Analysis of the Cu(I) Binding Site in Wheat Metallothionein Ec-1
 S1 of S8 Supplementary Materials: Localization and Spectroscopic Analysis of the Cu(I) Binding Site in Wheat Metallothionein Ec-1 Katsiaryna Tarasava, Jens Loebus and Eva Freisinger Figure S1. Deconvoluted
S1 of S8 Supplementary Materials: Localization and Spectroscopic Analysis of the Cu(I) Binding Site in Wheat Metallothionein Ec-1 Katsiaryna Tarasava, Jens Loebus and Eva Freisinger Figure S1. Deconvoluted
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10458 Active Site Remodeling in the Bifunctional Fructose-1,6- bisphosphate aldolase/phosphatase Juan Du, Rafael F. Say, Wei Lü, Georg Fuchs & Oliver Einsle SUPPLEMENTARY FIGURES Figure
doi:10.1038/nature10458 Active Site Remodeling in the Bifunctional Fructose-1,6- bisphosphate aldolase/phosphatase Juan Du, Rafael F. Say, Wei Lü, Georg Fuchs & Oliver Einsle SUPPLEMENTARY FIGURES Figure
A label-free DNA reduced graphene oxide-based fluorescent. sensor for highly sensitive and selective detection of hemin
 A label-free DNA reduced graphene oxide-based fluorescent sensor for highly sensitive and selective detection of hemin Yan Shi, Wei Tao Huang, Hong Qun Luo and Nian Bing Li* Key Laboratory on Luminescence
A label-free DNA reduced graphene oxide-based fluorescent sensor for highly sensitive and selective detection of hemin Yan Shi, Wei Tao Huang, Hong Qun Luo and Nian Bing Li* Key Laboratory on Luminescence
Supplementary Information. Overlap between folding and functional energy landscapes for. adenylate kinase conformational change
 Supplementary Information Overlap between folding and functional energy landscapes for adenylate kinase conformational change by Ulrika Olsson & Magnus Wolf-Watz Contents: 1. Supplementary Note 2. Supplementary
Supplementary Information Overlap between folding and functional energy landscapes for adenylate kinase conformational change by Ulrika Olsson & Magnus Wolf-Watz Contents: 1. Supplementary Note 2. Supplementary
The change of corrin-amides to carboxylates leads to altered structures of the B 12 -responding btub riboswitch
 Gallo et al. ESI 1 The change of corrin-amides to carboxylates leads to altered structures of the B 12 -responding btub riboswitch Electronic Supplementary Information (ESI) Sofia Gallo, Stefan Mundwiler,
Gallo et al. ESI 1 The change of corrin-amides to carboxylates leads to altered structures of the B 12 -responding btub riboswitch Electronic Supplementary Information (ESI) Sofia Gallo, Stefan Mundwiler,
1. Use the Data for RNAse to estimate:
 Chem 78 - - Spr 1 03/14/01 Assignment 4 - Answers Thermodynamic Analysis of RNAseA Denaturation by UV- Vis Difference Absorption Spectroscopy (and Differential Scanning Calorimetry). The accompanying excel
Chem 78 - - Spr 1 03/14/01 Assignment 4 - Answers Thermodynamic Analysis of RNAseA Denaturation by UV- Vis Difference Absorption Spectroscopy (and Differential Scanning Calorimetry). The accompanying excel
τ and τ, which represent the time-averaged single-molecule rates of I II and II I transitions
 Supporting nformation To Single-Molecule Study of Metalloregulator CueR DNA nteractions Using Engineered Holliday Junctions Nesha May Andoy, Susanta K. Sarar, Qi Wang, Debashis Panda, Jaime J. Benítez,
Supporting nformation To Single-Molecule Study of Metalloregulator CueR DNA nteractions Using Engineered Holliday Junctions Nesha May Andoy, Susanta K. Sarar, Qi Wang, Debashis Panda, Jaime J. Benítez,
Supplementary Information
 Electronic Supplementary Information for Dalton Transactions Insights into the Binding Properties of a Cuprous Ion Embedded in the Tren Cap of a Calix[6]arene and Supramolecular Trapping of an Intermediate.
Electronic Supplementary Information for Dalton Transactions Insights into the Binding Properties of a Cuprous Ion Embedded in the Tren Cap of a Calix[6]arene and Supramolecular Trapping of an Intermediate.
Principles of Physical Biochemistry
 Principles of Physical Biochemistry Kensal E. van Hold e W. Curtis Johnso n P. Shing Ho Preface x i PART 1 MACROMOLECULAR STRUCTURE AND DYNAMICS 1 1 Biological Macromolecules 2 1.1 General Principles
Principles of Physical Biochemistry Kensal E. van Hold e W. Curtis Johnso n P. Shing Ho Preface x i PART 1 MACROMOLECULAR STRUCTURE AND DYNAMICS 1 1 Biological Macromolecules 2 1.1 General Principles
Contents. xiii. Preface v
 Contents Preface Chapter 1 Biological Macromolecules 1.1 General PrincipIes 1.1.1 Macrornolecules 1.2 1.1.2 Configuration and Conformation Molecular lnteractions in Macromolecular Structures 1.2.1 Weak
Contents Preface Chapter 1 Biological Macromolecules 1.1 General PrincipIes 1.1.1 Macrornolecules 1.2 1.1.2 Configuration and Conformation Molecular lnteractions in Macromolecular Structures 1.2.1 Weak
Triangulating Nucleic Acid Conformations Using Multicolor Surface Energy Transfer
 Triangulating Nucleic cid Conformations Using Multicolor Surface Energy Transfer Supporting Information Ryan. Riskowski 1, Rachel E. rmstrong 2, Nancy L. Greenbaum 3, Geoffrey F. Strouse 1,2 * 1 Molecular
Triangulating Nucleic cid Conformations Using Multicolor Surface Energy Transfer Supporting Information Ryan. Riskowski 1, Rachel E. rmstrong 2, Nancy L. Greenbaum 3, Geoffrey F. Strouse 1,2 * 1 Molecular
Carbazole Derivatives Binding to c-kit G-quadruplex DNA
 Supplementary Materials Carbazole Derivatives Binding to c-kit G-quadruplex DNA Agata Głuszyńska 1, *, Bernard Juskowiak 1, Martyna Kuta-Siejkowska 2, Marcin Hoffmann 2 and Shozeb Haider 3 1 Laboratory
Supplementary Materials Carbazole Derivatives Binding to c-kit G-quadruplex DNA Agata Głuszyńska 1, *, Bernard Juskowiak 1, Martyna Kuta-Siejkowska 2, Marcin Hoffmann 2 and Shozeb Haider 3 1 Laboratory
CHEMISTRY - BROWN 13E CH.7 - PERIODIC PROPERTIES OF THE ELEMENTS
 !! www.clutchprep.com CONCEPT: EFFECTIVE NUCLEAR CHARGE & SLATER S RULES When looking at any particular electron within an atom it experiences two major forces. A(n) force from the nucleus and a(n) force
!! www.clutchprep.com CONCEPT: EFFECTIVE NUCLEAR CHARGE & SLATER S RULES When looking at any particular electron within an atom it experiences two major forces. A(n) force from the nucleus and a(n) force
4 Single molecule FRET
 4 Single molecule FRET FRET basics Energie Dipole-dipole interaction Teil I SM Fluo, Kap. 4 FRET FRET basics transfer rate (from Fermis Golden Rule) k t = 1 0 1 r 6 apple 2 9 ln(10) n 4 N A 128 5 Z d f
4 Single molecule FRET FRET basics Energie Dipole-dipole interaction Teil I SM Fluo, Kap. 4 FRET FRET basics transfer rate (from Fermis Golden Rule) k t = 1 0 1 r 6 apple 2 9 ln(10) n 4 N A 128 5 Z d f
Awanish Kumar, Anjeeta Rani and Pannuru Venkatesu* Department of Chemistry, University of Delhi, Delhi
 Electronic Supplementary Material (ESI) for New Journal of Chemistry. This journal is The Royal Society of Chemistry and the Centre National de la Recherche Scientifique 2014 Supplimentary Informations
Electronic Supplementary Material (ESI) for New Journal of Chemistry. This journal is The Royal Society of Chemistry and the Centre National de la Recherche Scientifique 2014 Supplimentary Informations
single-molecule fluorescence resonance energy transfer
 single-molecule fluorescence resonance energy transfer (2) determing the Förster radius: quantum yield, donor lifetime, spectral overlap, anisotropy michael börsch 26/05/2004 1 fluorescence (1) absorbance
single-molecule fluorescence resonance energy transfer (2) determing the Förster radius: quantum yield, donor lifetime, spectral overlap, anisotropy michael börsch 26/05/2004 1 fluorescence (1) absorbance
Advanced Fluorescence Microscopy I: Fluorescence (Foster) Resonance Energy Transfer
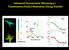 Advanced Fluorescence Microscopy I: Fluorescence (Foster) Resonance Energy Transfer 3.0 Pax 2.5 2.0 1200 800 400 GFP- Pax GFP-Pax + FATmCherry FAT FAT 1.5 Lifetime (ns) 0 1500 2000 2500 3000 Average lifetime
Advanced Fluorescence Microscopy I: Fluorescence (Foster) Resonance Energy Transfer 3.0 Pax 2.5 2.0 1200 800 400 GFP- Pax GFP-Pax + FATmCherry FAT FAT 1.5 Lifetime (ns) 0 1500 2000 2500 3000 Average lifetime
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature11524 Supplementary discussion Functional analysis of the sugar porter family (SP) signature motifs. As seen in Fig. 5c, single point mutation of the conserved
SUPPLEMENTARY INFORMATION doi:10.1038/nature11524 Supplementary discussion Functional analysis of the sugar porter family (SP) signature motifs. As seen in Fig. 5c, single point mutation of the conserved
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Resonance assignment and NMR spectra for hairpin and duplex A 6 constructs. (a) 2D HSQC spectra of hairpin construct (hp-a 6 -RNA) with labeled assignments. (b) 2D HSQC or SOFAST-HMQC
Supplementary Figure 1 Resonance assignment and NMR spectra for hairpin and duplex A 6 constructs. (a) 2D HSQC spectra of hairpin construct (hp-a 6 -RNA) with labeled assignments. (b) 2D HSQC or SOFAST-HMQC
BMB Class 17, November 30, Single Molecule Biophysics (II)
 BMB 178 2018 Class 17, November 30, 2018 15. Single Molecule Biophysics (II) New Advances in Single Molecule Techniques Atomic Force Microscopy Single Molecule Manipulation - optical traps and tweezers
BMB 178 2018 Class 17, November 30, 2018 15. Single Molecule Biophysics (II) New Advances in Single Molecule Techniques Atomic Force Microscopy Single Molecule Manipulation - optical traps and tweezers
CD Basis Set of Spectra that is used is that derived from comparing the spectra of globular proteins whose secondary structures are known from X-ray
 CD Basis Set of Spectra that is used is that derived from comparing the spectra of globular proteins whose secondary structures are known from X-ray crystallography An example of the use of CD Modeling
CD Basis Set of Spectra that is used is that derived from comparing the spectra of globular proteins whose secondary structures are known from X-ray crystallography An example of the use of CD Modeling
Imaging Nucleic Acids with the AFM. W Travis Johnson PhD Agilent Technologies Nanomeasurements Division
 Imaging Nucleic Acids with the AFM W Travis Johnson PhD Agilent Technologies Nanomeasurements Division Structure of DNA A T G C Standard Watson-Crick A-T & G-C base pairs in B-DNA DNA double helix composed
Imaging Nucleic Acids with the AFM W Travis Johnson PhD Agilent Technologies Nanomeasurements Division Structure of DNA A T G C Standard Watson-Crick A-T & G-C base pairs in B-DNA DNA double helix composed
Supporting Information. The Role of Structural Enthalpy in Spherical Nucleic Acid Hybridization
 Supporting Information The Role of Structural Enthalpy in Spherical Nucleic Acid Hybridization Lam-Kiu Fong, 1,6 Ziwei Wang, 2,6 George C. Schatz, 1,6 Erik Luijten,* 3,4,5 and Chad A. Mirkin* 1,3,6 1 Department
Supporting Information The Role of Structural Enthalpy in Spherical Nucleic Acid Hybridization Lam-Kiu Fong, 1,6 Ziwei Wang, 2,6 George C. Schatz, 1,6 Erik Luijten,* 3,4,5 and Chad A. Mirkin* 1,3,6 1 Department
Raman study of magnesium induced conversion of polyu polya duplexes to polyu polya polyu triplexes
 Spectroscopy 24 (2010) 445 448 445 DOI 10.3233/SPE-2010-0473 IOS Press Raman study of magnesium induced conversion of polyu polya duplexes to polyu polya polyu triplexes S.J. Espinoza Herrera and J. Štěpánek
Spectroscopy 24 (2010) 445 448 445 DOI 10.3233/SPE-2010-0473 IOS Press Raman study of magnesium induced conversion of polyu polya duplexes to polyu polya polyu triplexes S.J. Espinoza Herrera and J. Štěpánek
Supplementary Figure 1. Stability constants of metal monohydroxides. The log K values are summarized according to the atomic number of each element
 Supplementary Figure 1. Stability constants of metal monohydroxides. The log K values are summarized according to the atomic number of each element as determined in a previous study 1. The log K value
Supplementary Figure 1. Stability constants of metal monohydroxides. The log K values are summarized according to the atomic number of each element as determined in a previous study 1. The log K value
Lecture 5. Anisotropy decay/data analysis. Enrico Gratton
 Lecture 5. Anisotropy decay/data analysis Enrico Gratton Anisotropy decay Energy-transfer distance distributions Time resolved spectra Excited-state reactions Basic physics concept in polarization The
Lecture 5. Anisotropy decay/data analysis Enrico Gratton Anisotropy decay Energy-transfer distance distributions Time resolved spectra Excited-state reactions Basic physics concept in polarization The
A New Solvatochromic Fluorophore for Exploring Nonpolar Environments Created by Biopolymers
 Electronic Supplementary Information A New Solvatochromic Fluorophore for Exploring Nonpolar Environments Created by Biopolymers Abulfazl Fakhari M. and Steven E. Rokita Contribution from the Department
Electronic Supplementary Information A New Solvatochromic Fluorophore for Exploring Nonpolar Environments Created by Biopolymers Abulfazl Fakhari M. and Steven E. Rokita Contribution from the Department
Determination of Binding Affinities and Molecular Mechanisms case studies with nucleotide binding proteins
 Determination of Binding Affinities and Molecular Mechanisms case studies with nucleotide binding proteins Jochen Reinstein Max Planck-Institute for Medical Research Dept. of Biomolecular Mechanisms Heidelberg,
Determination of Binding Affinities and Molecular Mechanisms case studies with nucleotide binding proteins Jochen Reinstein Max Planck-Institute for Medical Research Dept. of Biomolecular Mechanisms Heidelberg,
Reviewers' comments: Reviewer #1 (Remarks to the Author):
 Reviewers' comments: Reviewer #1 (Remarks to the Author): The manuscript by Malli and coworkers reports the successful development and characterization of the first K+ specific FRET-based sensor protein.
Reviewers' comments: Reviewer #1 (Remarks to the Author): The manuscript by Malli and coworkers reports the successful development and characterization of the first K+ specific FRET-based sensor protein.
Low-Affinity Zinc Sensor Showing Fluorescence
 Supporting Information for Low-Affinity Zinc Sensor Showing Fluorescence Responses with Minimal Artifacts Xinhao Yan, a Jin Ju Kim, b Hey Sun Jeong, c Yu Kyung Moon, b Yoon Kyung Cho, c Soyeon Ahn, a Sang
Supporting Information for Low-Affinity Zinc Sensor Showing Fluorescence Responses with Minimal Artifacts Xinhao Yan, a Jin Ju Kim, b Hey Sun Jeong, c Yu Kyung Moon, b Yoon Kyung Cho, c Soyeon Ahn, a Sang
Biological Thermodynamics
 Biological Thermodynamics Classical thermodynamics is the only physical theory of universal content concerning which I am convinced that, within the framework of applicability of its basic contents, will
Biological Thermodynamics Classical thermodynamics is the only physical theory of universal content concerning which I am convinced that, within the framework of applicability of its basic contents, will
Crystal Violet as a Fluorescent Switch-On Probe for I-Motif: Label-Free DNA-Based Logic Gate
 Crystal Violet as a Fluorescent Switch-On Probe for I-Motif: Label-Free DNA-Based Logic Gate Dik-Lung Ma,* a Maria Hiu-Tung Kwan, a Daniel Shiu-Hin Chan, a Paul Lee, a Hui Yang, a Victor Pui-Yan Ma, a
Crystal Violet as a Fluorescent Switch-On Probe for I-Motif: Label-Free DNA-Based Logic Gate Dik-Lung Ma,* a Maria Hiu-Tung Kwan, a Daniel Shiu-Hin Chan, a Paul Lee, a Hui Yang, a Victor Pui-Yan Ma, a
Bottom-up Optimization of SERS Hot Spots. Supplementary Information
 Bottom-up Optimization of SERS Hot Spots Laura Fabris, * Department of Materials Science and Engineering, Institute for Advanced Materials Devices ad Nanotechnology, Rutgers, The State University of New
Bottom-up Optimization of SERS Hot Spots Laura Fabris, * Department of Materials Science and Engineering, Institute for Advanced Materials Devices ad Nanotechnology, Rutgers, The State University of New
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES Supplementary Figure 1 Protein sequence alignment of Vibrionaceae with either a 40-residue insertion or a 44-residue insertion. Identical residues are indicated by red background.
SUPPLEMENTARY FIGURES Supplementary Figure 1 Protein sequence alignment of Vibrionaceae with either a 40-residue insertion or a 44-residue insertion. Identical residues are indicated by red background.
1. Transition dipole moment
 1. Transition dipole moment You have measured absorption spectra of aqueous (n=1.33) solutions of two different chromophores (A and B). The concentrations of the solutions were the same. The absorption
1. Transition dipole moment You have measured absorption spectra of aqueous (n=1.33) solutions of two different chromophores (A and B). The concentrations of the solutions were the same. The absorption
17. Biomolecular Interaction
 17. Biomolecular Interaction Methods for characterizing biomolecular interactions Sequence-specific DNA binding ligands Molecular mechanisms of drug action and drug resistance In silico compound design
17. Biomolecular Interaction Methods for characterizing biomolecular interactions Sequence-specific DNA binding ligands Molecular mechanisms of drug action and drug resistance In silico compound design
Supplementary figure 1 Application of tmfret in LeuT. (a) To assess the feasibility of using tmfret for distance-dependent measurements in LeuT, a
 Supplementary figure 1 Application of tmfret in LeuT. (a) To assess the feasibility of using tmfret for distance-dependent measurements in LeuT, a series of tmfret-pairs comprised of single cysteine mutants
Supplementary figure 1 Application of tmfret in LeuT. (a) To assess the feasibility of using tmfret for distance-dependent measurements in LeuT, a series of tmfret-pairs comprised of single cysteine mutants
Supplemental Data for: Direct Observation of Translocation in Individual DNA Polymerase Complexes
 Supplemental Data for: Direct Observation of Translocation in Individual DNA Polymerase Complexes Joseph M. Dahl 1, Ai H. Mai 1, Gerald M. Cherf 1, Nahid N. Jetha 4, Daniel R. Garalde 3, Andre Marziali
Supplemental Data for: Direct Observation of Translocation in Individual DNA Polymerase Complexes Joseph M. Dahl 1, Ai H. Mai 1, Gerald M. Cherf 1, Nahid N. Jetha 4, Daniel R. Garalde 3, Andre Marziali
J- vs. H-type assembly: pentamethine cyanine (Cy5) as near IR chiroptical. reporter
 S1 Supporting Information: J- vs. H-type assembly: pentamethine cyanine (Cy5) as near IR chiroptical reporter Larysa I. Markova, a,b Vladimir L. Malinovskii, a Leonid D. Patsenker b and Robert Häner* a
S1 Supporting Information: J- vs. H-type assembly: pentamethine cyanine (Cy5) as near IR chiroptical reporter Larysa I. Markova, a,b Vladimir L. Malinovskii, a Leonid D. Patsenker b and Robert Häner* a
Stopped-Flow Studies of the Kinetics of Single-Stranded DNA Binding and Wrapping around the Escherichia coli SSB Tetramer
 6032 Biochemistry 2002, 41, 6032-6044 Stopped-Flow Studies of the Kinetics of Single-Stranded DNA Binding and Wrapping around the Escherichia coli SSB Tetramer Alexander G. Kozlov and Timothy M. Lohman*
6032 Biochemistry 2002, 41, 6032-6044 Stopped-Flow Studies of the Kinetics of Single-Stranded DNA Binding and Wrapping around the Escherichia coli SSB Tetramer Alexander G. Kozlov and Timothy M. Lohman*
Atomic and molecular interactions. Scanning probe microscopy.
 Atomic and molecular interactions. Scanning probe microscopy. Balázs Kiss Nanobiotechnology and Single Molecule Research Group, Department of Biophysics and Radiation Biology 27. November 2013. 2 Atomic
Atomic and molecular interactions. Scanning probe microscopy. Balázs Kiss Nanobiotechnology and Single Molecule Research Group, Department of Biophysics and Radiation Biology 27. November 2013. 2 Atomic
Free energy, electrostatics, and the hydrophobic effect
 Protein Physics 2016 Lecture 3, January 26 Free energy, electrostatics, and the hydrophobic effect Magnus Andersson magnus.andersson@scilifelab.se Theoretical & Computational Biophysics Recap Protein structure
Protein Physics 2016 Lecture 3, January 26 Free energy, electrostatics, and the hydrophobic effect Magnus Andersson magnus.andersson@scilifelab.se Theoretical & Computational Biophysics Recap Protein structure
Coordinated conformational and compositional dynamics drive. ribosome translocation
 Coordinated conformational and compositional dynamics drive ribosome translocation Supplementary Information Jin Chen,2, Alexey Petrov 2, Albert Tsai,2, Seán E. O Leary 2, & Joseph D. Puglisi 2,3 Department
Coordinated conformational and compositional dynamics drive ribosome translocation Supplementary Information Jin Chen,2, Alexey Petrov 2, Albert Tsai,2, Seán E. O Leary 2, & Joseph D. Puglisi 2,3 Department
Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1.
 Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1. PDZK1 constru cts Amino acids MW [kda] KD [μm] PEPT2-CT- FITC KD [μm] NHE3-CT- FITC KD [μm] PDZK1-CT-
Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1. PDZK1 constru cts Amino acids MW [kda] KD [μm] PEPT2-CT- FITC KD [μm] NHE3-CT- FITC KD [μm] PDZK1-CT-
Presentation Microcalorimetry for Life Science Research
 Presentation Microcalorimetry for Life Science Research MicroCalorimetry The Universal Detector Heat is either generated or absorbed in every chemical process Capable of thermal measurements over a wide
Presentation Microcalorimetry for Life Science Research MicroCalorimetry The Universal Detector Heat is either generated or absorbed in every chemical process Capable of thermal measurements over a wide
LS1a Fall 2014 Problem Set #2 Due Monday 10/6 at 6 pm in the drop boxes on the Science Center 2 nd Floor
 LS1a Fall 2014 Problem Set #2 Due Monday 10/6 at 6 pm in the drop boxes on the Science Center 2 nd Floor Note: Adequate space is given for each answer. Questions that require a brief explanation should
LS1a Fall 2014 Problem Set #2 Due Monday 10/6 at 6 pm in the drop boxes on the Science Center 2 nd Floor Note: Adequate space is given for each answer. Questions that require a brief explanation should
Effect of the Single and Double Chain Surfactant Cobalt(III) Complexes on Their Hydrophobicity, Micelle Formation,
 Electronic Supplementary Material (ESI) for Inorganic Chemistry Frontiers. This journal is the Partner Organisations 2014 Supplementary Information Effect of the Single and Double Chain Surfactant Cobalt(III)
Electronic Supplementary Material (ESI) for Inorganic Chemistry Frontiers. This journal is the Partner Organisations 2014 Supplementary Information Effect of the Single and Double Chain Surfactant Cobalt(III)
Lipid Regulated Intramolecular Conformational Dynamics of SNARE-Protein Ykt6
 Supplementary Information for: Lipid Regulated Intramolecular Conformational Dynamics of SNARE-Protein Ykt6 Yawei Dai 1, 2, Markus Seeger 3, Jingwei Weng 4, Song Song 1, 2, Wenning Wang 4, Yan-Wen 1, 2,
Supplementary Information for: Lipid Regulated Intramolecular Conformational Dynamics of SNARE-Protein Ykt6 Yawei Dai 1, 2, Markus Seeger 3, Jingwei Weng 4, Song Song 1, 2, Wenning Wang 4, Yan-Wen 1, 2,
Circular dichroism spectroscopic studies on structures formed by telomeric DNA sequences in vitro
 Circular dichroism spectroscopic studies on structures formed by telomeric DNA sequences in vitro ZHANG Xiaoyan 1, CAO Enhua 1, SUN Xueguang 1 & BAI Chunli 2 1. Institute of Biophysics, Chinese Academy
Circular dichroism spectroscopic studies on structures formed by telomeric DNA sequences in vitro ZHANG Xiaoyan 1, CAO Enhua 1, SUN Xueguang 1 & BAI Chunli 2 1. Institute of Biophysics, Chinese Academy
Novel fluorescent cationic benzothiazole dye response to G-quadruplex aptamer as a novel K + sensor
 Electronic Supplementary Material (ESI) for Analyst. This journal is The Royal Society of Chemistry 2017 Novel fluorescent cationic benzothiazole dye response to G-quadruplex aptamer as a novel K + sensor
Electronic Supplementary Material (ESI) for Analyst. This journal is The Royal Society of Chemistry 2017 Novel fluorescent cationic benzothiazole dye response to G-quadruplex aptamer as a novel K + sensor
Biology Chemistry & Physics of Biomolecules. Examination #1. Proteins Module. September 29, Answer Key
 Biology 5357 Chemistry & Physics of Biomolecules Examination #1 Proteins Module September 29, 2017 Answer Key Question 1 (A) (5 points) Structure (b) is more common, as it contains the shorter connection
Biology 5357 Chemistry & Physics of Biomolecules Examination #1 Proteins Module September 29, 2017 Answer Key Question 1 (A) (5 points) Structure (b) is more common, as it contains the shorter connection
Supporting Information
 Supporting Information Materials and Methods: Tris-hydroxymethylaminomethane (Tris) was purchased from USB. All the other reagents used in the experiments were purchased from Sigma. All the DNA oligonucleotides
Supporting Information Materials and Methods: Tris-hydroxymethylaminomethane (Tris) was purchased from USB. All the other reagents used in the experiments were purchased from Sigma. All the DNA oligonucleotides
SUPPLEMENTARY INFORMATION. Pistol Ribozyme Adopts a Pseudoknot Fold. Facilitating Site-specific In-line Cleavage
 UPPLEMENTAY INFMATIN Pistol ibozyme Adopts a Pseudoknot Fold Facilitating ite-specific In-line Cleavage Aiming en 1,2,4, Nikola Vušurović 3,4, Jennifer Gebetsberger 3, Pu Gao 2, Michael Juen 3, Christoph
UPPLEMENTAY INFMATIN Pistol ibozyme Adopts a Pseudoknot Fold Facilitating ite-specific In-line Cleavage Aiming en 1,2,4, Nikola Vušurović 3,4, Jennifer Gebetsberger 3, Pu Gao 2, Michael Juen 3, Christoph
Ionic effect on the proton transfer mechanism in aqueous solutions
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2017 Electronic Supplementary Information for Ionic effect on the proton transfer mechanism
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2017 Electronic Supplementary Information for Ionic effect on the proton transfer mechanism
Supplementary Information
 Supplementary Information Single molecule FRET reveals the energy landscape of the full length SAM I riboswitch Christoph Manz, 1,2 Andrei Yu. Kobitski, 1 Ayan Samanta, 3 Bettina G. Keller 4, Andres Jäschke,
Supplementary Information Single molecule FRET reveals the energy landscape of the full length SAM I riboswitch Christoph Manz, 1,2 Andrei Yu. Kobitski, 1 Ayan Samanta, 3 Bettina G. Keller 4, Andres Jäschke,
Entropy-Driven Folding of an RNA Helical Junction: An Isothermal Titration Calorimetric Analysis of the Hammerhead Ribozyme
 5870 Biochemistry 2004, 43, 5870-5881 Entropy-Driven Folding of an RNA Helical Junction: An Isothermal Titration Calorimetric Analysis of the Hammerhead Ribozyme Peter J. Mikulecky, Jennifer C. Takach,
5870 Biochemistry 2004, 43, 5870-5881 Entropy-Driven Folding of an RNA Helical Junction: An Isothermal Titration Calorimetric Analysis of the Hammerhead Ribozyme Peter J. Mikulecky, Jennifer C. Takach,
SUPPLEMENTARY INFORMATION
 Figure S1. Secondary structure of CAP (in the camp 2 -bound state) 10. α-helices are shown as cylinders and β- strands as arrows. Labeling of secondary structure is indicated. CDB, DBD and the hinge are
Figure S1. Secondary structure of CAP (in the camp 2 -bound state) 10. α-helices are shown as cylinders and β- strands as arrows. Labeling of secondary structure is indicated. CDB, DBD and the hinge are
The Fic protein Doc uses an inverted substrate to phosphorylate and. inactivate EF-Tu
 The Fic protein Doc uses an inverted substrate to phosphorylate and inactivate EF-Tu Daniel Castro-Roa 1, Abel Garcia-Pino 2,3 *, Steven De Gieter 2,3, Nico A.J. van Nuland 2,3, Remy Loris 2,3, Nikolay
The Fic protein Doc uses an inverted substrate to phosphorylate and inactivate EF-Tu Daniel Castro-Roa 1, Abel Garcia-Pino 2,3 *, Steven De Gieter 2,3, Nico A.J. van Nuland 2,3, Remy Loris 2,3, Nikolay
Supporting Information
 Supporting Information Figure S1. 2D ( 1 H- 1 H) COSY90 NMR (300 MHz, 3:1 TFA:TFA-d) spectrum of oligo(l-glu-co- 25%L-Cys) synthesized using from 7:3 L-Glu-(Et) 2 :L-Cys-Et, 0.5 M total substrate concentration,
Supporting Information Figure S1. 2D ( 1 H- 1 H) COSY90 NMR (300 MHz, 3:1 TFA:TFA-d) spectrum of oligo(l-glu-co- 25%L-Cys) synthesized using from 7:3 L-Glu-(Et) 2 :L-Cys-Et, 0.5 M total substrate concentration,
Thermodynamic Stability of Hoogsteen and Watson-Crick. Base Pairs in the Presence of Histone H3-Mimicking Peptide
 Supporting information Thermodynamic Stability of Hoogsteen and Watson-Crick Base Pairs in the Presence of Histone H3-Mimicking Peptide Smritimoy Pramanik a, Kaori Nakamura a, Kenji Usui a,b, Shu-ichi
Supporting information Thermodynamic Stability of Hoogsteen and Watson-Crick Base Pairs in the Presence of Histone H3-Mimicking Peptide Smritimoy Pramanik a, Kaori Nakamura a, Kenji Usui a,b, Shu-ichi
Cationic carbosilane dendrimers and oligonucleotide binding: an energetic affair
 Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2014 Supporting information Cationic carbosilane dendrimers and oligonucleotide binding: an energetic
Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2014 Supporting information Cationic carbosilane dendrimers and oligonucleotide binding: an energetic
Gas-Phase DNA Helix Conformations
 Gas-Phase DNA Helix Conformations Erin Shammel Baker, Jennifer Gidden, Alessandra Ferzoco, Thomas Wyttenbach and Michael Bowers utline Experimental Method Theoretical Method Instrumentation DNA Background
Gas-Phase DNA Helix Conformations Erin Shammel Baker, Jennifer Gidden, Alessandra Ferzoco, Thomas Wyttenbach and Michael Bowers utline Experimental Method Theoretical Method Instrumentation DNA Background
Page 2. Define the term electron affinity for chlorine (2)
 Q1.(a) Define the term electron affinity for chlorine. (b) Complete this Born Haber cycle for magnesium chloride by giving the missing species on the dotted lines. Include state symbols where appropriate.
Q1.(a) Define the term electron affinity for chlorine. (b) Complete this Born Haber cycle for magnesium chloride by giving the missing species on the dotted lines. Include state symbols where appropriate.
Supplementary Information
 1 Supplementary Information Figure S1 The V=0.5 Harker section of an anomalous difference Patterson map calculated using diffraction data from the NNQQNY crystal at 1.3 Å resolution. The position of the
1 Supplementary Information Figure S1 The V=0.5 Harker section of an anomalous difference Patterson map calculated using diffraction data from the NNQQNY crystal at 1.3 Å resolution. The position of the
A nano-positioning system for macromolecular structural analysis
 nature methods A nano-positioning system for macromolecular structural analysis Adam Muschielok, Joanna Andrecka, Anass Jawhari, Florian Brückner, Patrick Cramer & Jens Michaelis Supplementary figures
nature methods A nano-positioning system for macromolecular structural analysis Adam Muschielok, Joanna Andrecka, Anass Jawhari, Florian Brückner, Patrick Cramer & Jens Michaelis Supplementary figures
A. K. R. Junker et al 1 Investigating subtle 4f vs. 5f...
 Electronic Supplementary Material (ESI) for Dalton Transactions. This journal is The Royal Society of Chemistry 2018 A. K. R. Junker et al 1 Investigating subtle 4f vs. 5f... Electronic Supplementary Information
Electronic Supplementary Material (ESI) for Dalton Transactions. This journal is The Royal Society of Chemistry 2018 A. K. R. Junker et al 1 Investigating subtle 4f vs. 5f... Electronic Supplementary Information
Temperature ( o C)
 Viscosity (Pa sec) Supplementary Information 10 8 10 6 10 4 10 2 150 200 250 300 Temperature ( o C) Supplementary Figure 1 Viscosity of fibre components (PC cladding blue; As 2 Se 5 red; CPE black) as
Viscosity (Pa sec) Supplementary Information 10 8 10 6 10 4 10 2 150 200 250 300 Temperature ( o C) Supplementary Figure 1 Viscosity of fibre components (PC cladding blue; As 2 Se 5 red; CPE black) as
SUPPLEMENTARY INFORMATION
 Supplementary Results DNA binding property of the SRA domain was examined by an electrophoresis mobility shift assay (EMSA) using synthesized 12-bp oligonucleotide duplexes containing unmodified, hemi-methylated,
Supplementary Results DNA binding property of the SRA domain was examined by an electrophoresis mobility shift assay (EMSA) using synthesized 12-bp oligonucleotide duplexes containing unmodified, hemi-methylated,
DNA Structure. Voet & Voet: Chapter 29 Pages Slide 1
 DNA Structure Voet & Voet: Chapter 29 Pages 1107-1122 Slide 1 Review The four DNA bases and their atom names The four common -D-ribose conformations All B-DNA ribose adopt the C2' endo conformation All
DNA Structure Voet & Voet: Chapter 29 Pages 1107-1122 Slide 1 Review The four DNA bases and their atom names The four common -D-ribose conformations All B-DNA ribose adopt the C2' endo conformation All
Other Cells. Hormones. Viruses. Toxins. Cell. Bacteria
 Other Cells Hormones Viruses Toxins Cell Bacteria ΔH < 0 reaction is exothermic, tells us nothing about the spontaneity of the reaction Δ H > 0 reaction is endothermic, tells us nothing about the spontaneity
Other Cells Hormones Viruses Toxins Cell Bacteria ΔH < 0 reaction is exothermic, tells us nothing about the spontaneity of the reaction Δ H > 0 reaction is endothermic, tells us nothing about the spontaneity
Because light behaves like a wave, we can describe it in one of two ways by its wavelength or by its frequency.
 Light We can use different terms to describe light: Color Wavelength Frequency Light is composed of electromagnetic waves that travel through some medium. The properties of the medium determine how light
Light We can use different terms to describe light: Color Wavelength Frequency Light is composed of electromagnetic waves that travel through some medium. The properties of the medium determine how light
SUPPLEMENTARY MATERIAL FOR
 SUPPLEMENTARY MATERIAL FOR THE LIPID-BINDING DOMAIN OF WILD TYPE AND MUTANT ALPHA- SYNUCLEIN: COMPACTNESS AND INTERCONVERSION BETWEEN THE BROKEN- AND EXTENDED-HELIX FORMS. Elka R. Georgieva 1, Trudy F.
SUPPLEMENTARY MATERIAL FOR THE LIPID-BINDING DOMAIN OF WILD TYPE AND MUTANT ALPHA- SYNUCLEIN: COMPACTNESS AND INTERCONVERSION BETWEEN THE BROKEN- AND EXTENDED-HELIX FORMS. Elka R. Georgieva 1, Trudy F.
Real-time Ratiometric Fluorescent Assay for Alkaline Phosphatase Activity with Stimulus Responsive Infinite Coordination Polymer Nanoparticles
 Supporting information Real-time Ratiometric luorescent Assay for Alkaline Phosphatase Activity with Stimulus Responsive Infinite Coordination Polymer Nanoparticles Jingjing Deng, Ping Yu, Yuexiang Wang,
Supporting information Real-time Ratiometric luorescent Assay for Alkaline Phosphatase Activity with Stimulus Responsive Infinite Coordination Polymer Nanoparticles Jingjing Deng, Ping Yu, Yuexiang Wang,
Free-Probe Fluorescence of Light-up Probes
 J. Am. Chem. Soc. 2001, 123, 803-809 803 Free-Probe Fluorescence of Light-up Probes Nicke Svanvik, Jan Nygren, Gunnar Westman, and Mikael Kubista*, Contribution from the Department of Molecular Biotechnology,
J. Am. Chem. Soc. 2001, 123, 803-809 803 Free-Probe Fluorescence of Light-up Probes Nicke Svanvik, Jan Nygren, Gunnar Westman, and Mikael Kubista*, Contribution from the Department of Molecular Biotechnology,
Simple Method for Determining the Thermodynamic Parameters of Ligand with Nucleic Acids Interaction
 Armenian Journal of Physics, 2017, vol. 10, issue 1, pp. 42-48 Simple Method for Determining the Thermodynamic Parameters of Ligand with Nucleic Acids Interaction M.A. Torosyan National University of Architecture
Armenian Journal of Physics, 2017, vol. 10, issue 1, pp. 42-48 Simple Method for Determining the Thermodynamic Parameters of Ligand with Nucleic Acids Interaction M.A. Torosyan National University of Architecture
Structural basis for catalytically restrictive dynamics of a high-energy enzyme state
 Supplementary Material Structural basis for catalytically restrictive dynamics of a high-energy enzyme state Michael Kovermann, Jörgen Ådén, Christin Grundström, A. Elisabeth Sauer-Eriksson, Uwe H. Sauer
Supplementary Material Structural basis for catalytically restrictive dynamics of a high-energy enzyme state Michael Kovermann, Jörgen Ådén, Christin Grundström, A. Elisabeth Sauer-Eriksson, Uwe H. Sauer
Supporting Information for
 Supporting Information for Direct Fluorescent Detection of Blood Potassium by Ion-selective Formation of Intermolecular G-quadruplex and Ligand Binding Le Yang, Zhihe Qing,, * Changhui Liu, Qiao Tang,
Supporting Information for Direct Fluorescent Detection of Blood Potassium by Ion-selective Formation of Intermolecular G-quadruplex and Ligand Binding Le Yang, Zhihe Qing,, * Changhui Liu, Qiao Tang,
Co-localization, FRET
 Co-localization, FRET Last class FRAP Diffusion This class Co-localization Correlation FRET Co-localization Can you infer function of protein from it s intracellular location How do you measure if two
Co-localization, FRET Last class FRAP Diffusion This class Co-localization Correlation FRET Co-localization Can you infer function of protein from it s intracellular location How do you measure if two
RNA Helix Stability in Mixed Na 1 /Mg 21 Solution
 Biophysical Journal Volume 9 May 007 365 363 365 RNA Helix Stability in Mixed Na /Mg Solution Zhi-Jie Tan and Shi-Jie Chen Department of Physics and Astronomy and Department of Biochemistry, University
Biophysical Journal Volume 9 May 007 365 363 365 RNA Helix Stability in Mixed Na /Mg Solution Zhi-Jie Tan and Shi-Jie Chen Department of Physics and Astronomy and Department of Biochemistry, University
Upconverting Nanoparticle to Quantum Dot Förster Resonance Energy Transfer: Increasing the Efficiency Through Donor Design
 Electronic Supporting Information Upconverting Nanoparticle to Quantum Dot Förster Resonance Energy Transfer: Increasing the Efficiency Through Donor Design R. Marin a,b, L. Labrador-Paéz c, A. Skripka
Electronic Supporting Information Upconverting Nanoparticle to Quantum Dot Förster Resonance Energy Transfer: Increasing the Efficiency Through Donor Design R. Marin a,b, L. Labrador-Paéz c, A. Skripka
Supporting Information
 Supporting Information Cyclodextrin Supramolecular Complex as Water Soluble Ratiometric Sensor for ferric Ion Sensing Meiyun Xu, Shuizhu Wu,* Fang Zeng, Changmin Yu College of Materials Science & Engineering,
Supporting Information Cyclodextrin Supramolecular Complex as Water Soluble Ratiometric Sensor for ferric Ion Sensing Meiyun Xu, Shuizhu Wu,* Fang Zeng, Changmin Yu College of Materials Science & Engineering,
PART 2 Electronic Structure and the Periodic Table. Reference: Chapter 7 8 in textbook
 PART 2 Electronic Structure and the Periodic Table Reference: Chapter 7 8 in textbook 1 Experiment to Discover Atom Structure -particle: He 2+ mass number = 4 Nucleus and Electron Model 2 Atomic Structure
PART 2 Electronic Structure and the Periodic Table Reference: Chapter 7 8 in textbook 1 Experiment to Discover Atom Structure -particle: He 2+ mass number = 4 Nucleus and Electron Model 2 Atomic Structure
