When you download HuLiS you are requested to fill a form that tells us where HuLiS is used. It can be skipped, but we appreciate if you do so.
|
|
|
- Annice West
- 6 years ago
- Views:
Transcription
1 User s manual Version 3.3 February S. umbel, ism2, Aix-Marseille University France For ulis 3.3 Native english teachers (or students): if you like the program but not the manual, send me corrections (use the file for corrections) Overview ulis is a Java applet that targets at the calculation of electronic structures within the uckel framework. In addition to the standard simple ückel method, ulis implements the calculation of mesomerism (i.e the user can build Lewis structures, and evaluate the weights of Lewis structures). Mesomerism is written as a linear combination of Lewis structures, and coefficients are also calculated. ulis is freely available a standalone applet (to download from the web site : We also propose an on-line version that runs from the web site Last, an TML5 exists for mobiles and tablets (see m.hulis.free.fr). This manual describes the standalone applet. Some features are removed in the online version because the required access to the user s computer is blocked for security reasons (undo/redo, a robotized tutorial, file handling). When you download ulis you are requested to fill a form that tells us where ulis is used. It can be skipped, but we appreciate if you do so. In the same spirit, when you start ulis the first time, our privacy policy is explained, and you can let ulis send us some informations such as how many times you use ulis, which version of ulis you are using, which country you are using it from, etc. This choice can be modified later with the preference panel. Nota : ulis interface is in English, but the for some langages (Italian, French Arabic) depending of the OS. This choice can be changed in the appropriate menu. User s manual ulis 3.3 1
2 Areas Nota: Flying over a button with the mouse display an help. The taskbar also gives indications on how to use ulis. The ulis window is divided into 5 areas (Figure 1). ückel tools Menu bar (& langages) Mesomerism Lewis structures Main panel (& orbitals) Taskbar Figure 1 : Areas is ulis (in grey are buttons that are not relevant in the context) 1. Taskbar Instant help (depends on the button in use). This area targets mainly at beginners. 2. Menu Bar <File> Open/save molecules & mesomeric structures (if any). The file format is that of Gaussian + NBO (see on the right here and examples in the library available for download from the website). Preferences are here also: privacy, number of digits in outputs can be adjusted here. Molecular Orbitals scale factor for drawings, molecule drawing preferences, Lewis preferences, etc. <Edition> undo/redo + paste paste can use several format Gaussian type of file ( here opposite)==> Or simply XYZ coordinates of atoms (without the number of atoms) Format xyz C From gaussian output in Å From Gamess output in au (Bohr) C The molecule is automatically rotated along the xy plan. The initial orientation can be retrieved (see the results panel for Lewis). Gaussian type %mem=256mb #P F 6-31G(d) pop=nboread! File generated by the ulis Formamide C N S $nbo nrt $end $NRTSTR! Trust factor = 90.90% (LP) STR! ulis Wgt = 65.40% (LP) S1 LONE END BOND D 1 6 S 1 2 S 1 3 S 2 4 S 2 5 END END $END User s manual ulis 3.3 2
3 <Display> just zooms for the main panel. <Compute> expert mode and choice of the Lewis calculation method : L-CI or L-P (default). The expert mode gives the opportunity to define a singlet coupling of two electrons in principle at long range (a green bond is drawn). <Tutorial> One robot is defined, it builds a molecule, and shows the orbitals <elp> events log (for debug), and the «About» <Langage - flags> Nota: The menu bar has fewer options in the on-line version to match security requirements. 3. Main panel ere are thumbnails for the molecule (ψ tot ) and the Lewis structures (ψ 1, ψ 2, ). For each, the Molecular Orbital (MO) energetic diagram is drawn. When the mouse flies over an atom or a bond, the corresponding parameters are displayed. The MO are drawn by a clic on the energetic level. In the standalone version, the content of this panel can be saved as a vectorial picture (.svg) for a better quality: A second clic erases the MO (bascule). When a MO is on the screen coefficients are displayed if the mouse flies over atoms. The scale factor for the MO drawing can be changed in Preferences (Zoom MO - slider). C C This image comes from a conversion ffrom.svg to.pdf (using Inkscape). The pdf is then inserted in word, and saved as a pdf to put on the web site. C User s manual ulis 3.3 3
4 4. ückel tools In the left part (blue) are the tools to build the molecule for simple ückel calculations. Results display panel is as neat as we could. Nota : As uckel calculations have delocalized π bonds ( delocalized structure ), these are not drawn for ückel calculations. Bonds are rather drawn with dashes with a dash density that is proportional to π bond indices. Build, Change, Delete : tools to build the molecule s connectivity. "Build" creates a sp 2 (1 électron) Carbon. The appropriate valence is completed by atoms. "Change" modifies sp 2 Carbons into other atoms. When this button is selected, a clic on an atom (or a bond) open a small panel with compatible choices. Default parameters are proposed (built-in), with a reasonable choice of atoms (these parameters are from Van-Catledge s paper [1] ). All values can be modified. A Xx generic atom is also available. Atomic integral, h x, number of neighbours (n neighb ), number of π electron (n e- ) and symbol Xx can be changed. "Delete" delete an object atom or bond and correct the valency if necessary. (a) [ ] & [ + ] increase/decrease the total charge. (b) Figure 2 : Change the parameters: for an atom (a) or for a bond (b). Move, Rotate, Optimize, Center to correct the display of the molecule. Sort sort the atoms. The applet waits for either all atoms are selected or that the button is deselected). Atoms are sorted in the order they have been selected. Three small buttons ("radio") Numbering Parameters Charges To display (or not) the corresponding stuff. Results open a pop up panel (light blue background) with ückel results. Erase all erase the molecule and reinitialize ulis. Quit (standalone only) Quit ulis. User s manual ulis 3.3 4
5 5. Mesomery / Lewis Structures The right hand-side part (orange) provides tools to build Lewis structures and see the corresponding results. As elsewhere in ulis, when the mouse flies over a button, an explanation is displayed (yellow little panel). When selected a larger explanation is written in the taskbar. Nota: consistent with the principle of localization (in Lewis structures), and a contrario to the delocalized electronic structures, π bonds and lone pairs are explicitly displayed in Lewis structures. The interaction between these structures gives rise to π delocalization (mesomerism). ulis computes the coefficients of this interaction, hence the weights of each structure. Nota bis: ulis targets at a good matching between the delocalized envision (ückel) and the localized one (Lewis). Two strategies are available: (1) Wave function matching, or ückel-lewis-projection (L-P); it is the most accurate approach and the default one. owever, it can require many computations. Large systems and/or systems with many Lewis structures might heat the cpu of your computer. [2] (2) Energetic matching, or ückel-lewis-configuration-interaction (L-CI); faster but less accurate, it is a physically interesting approach where the interaction between localized states (Lewis structures) is accounted, for via an effective amiltonian. [3] Generate all The corresponding papers can be asked (and obtained) upon request. Creates all the Lewis structures that have 0 or 1 charge separation (1 +/ but not 1 st neighbour), and biradicals if requested in the pref panel. Biradicals are singlet coupled shown as a green bond ( ab + ba ). A structure can thus be both zwitterionic (+/ ) and a biradical:!!! Create to define a new 1 Lewis structure (new thumbnail) It copies the skeleton of the molecule in ψ i (a new thumbnail). Charges displayed correspond to the π effective charge (Z eff ) for each atom as defined in the ückel framework. The Lewis structure is to be completed with bonds, and odd or pairs of electron - use the Bond/Electr button. Bond/Electr to define bonds and electron dots of the Lewis structure On an atom, it modifies the number of electron as 1 > 2 > 0 > 1 etc. On a bond, its π bonding indice is modified as single > double > single > etc (there are no triple bond on ulis). The charge is updated at each clic. ulis checks for redundancy in Lewis structures. In L-CI the weight is shared among the redundant structures. In L-P, the weights are not computed when redundancy is encountered. A flag is put on the redundant structure(s). User s manual ulis 3.3 5
6 Results open a pop up panel (light orange background) with mesomeric results. Example for acrolein <ψ tot ψ 2 > <ψ tot ψ Mesomery > Lewis Structures ψ Mesomery ψ Mesomery Weights of the Structures adds NBO strings & produces gaussian inputs (a) (b) (c) Delocalized (a) and mesomeric (b) (c) structures. Figure 3 : Popup of Lewis results for acrolein. ere, 2 mesomeric structures are active among the set. Only L-P is active. When L-CI also activated, The effective amiltonian of the Configuration Interaction is also written. Nota : the trust factor, (trust rank) (τ) corresponds to the overlap between the mesomeric wavefunction and ückel s (τ = <Ψ ückel Ψ mesom. >). It is written as %. Slider In version before ulis 3.2, τ was calculated differently. To select part of the Lewis structures. Structures are ordered vertically using their overlap with the ückel wave function, from smallest overlap (top) to the largest (bottom). The overlap value is indicated in black. The red number in parenthesis indicates how many structures have the same overlap with the ückel wave function. The blue arrow indicates the threshold. When overlaps are not computed we use the energy of the structures from the highest in energy (top) to the lowest (bottom). (a) (b) Figure 4 : with the slider one selects structures (a) per overlap (see text) or (b) per energies (L-CI only). Nota : The energy difference between the delocalized wavefunction (ückel level) and the lowest Lewis give the resonance energy (at the ückel level, in terms of β). Erase 1 Delete the resonance structure of the current thumbnail. Erase mes. Delete all resonance structures. Returns to ückel calculations. User s manual ulis 3.3 6
7 Remarks 1. To read ückel results of a specific Lewis structure: Select its thumbnail Clic on Results of the ückel (blue) area. This opens a blue-background popup. Note that the amiltonien matrix has zeros along single bonds. This is the way electrons are localized in ulis. Charges & indices are also here. Localized orbitales can be viewed with a clik on the orbital level (main panel). 2. Biradical structures are singlet coupled. A green bond is drawn. Only 1 (one) spin coupling can be defined. So a max of 3 unpaired electrons are allowed. The expert mode, adds a button labelled "Spin coupling" to define which 2 electrons are coupled. Only useful for 3 electrons cases. 3. Chemical bond, Valence Bond covalent vs ionics. The chemical bond between A B describes how two electrons are paired in a bonding orbital. This holds for π bonds. If the orbital π =1/ 2 (p A +p B ) is bioccupied, the standard π bond (a black line) includes : - Covalent bonding (A B) - AND Ionics o A( ) (+)B when p A si bioccupied o A(+) ( )B when p B si bioccupied Adding ionics structures to such a standard π bond would introduce redundancy. Redundancy should be avoided. It is a good practice to remove structures with a negative weight. The green bond only introduces the covalent part. Ionics can (must) be added there. Only one green bond can be set in ulis. 1 F.A. Van-Catledge A Pariser-Parr-Pople-based set of ueckel molecular orbital parameters, J. Org. Chem. 1980, 45, ( 2 Y. Carissan, D. agebaum-reignier, N. Goudard, S. umbel ückel-lewis-projection Method: A Weights Watcher for Mesomeric Structures: J. Phys. Chem. A. 2008, 112 (50), S. umbel Getting the Weights of Lewis Structures out of ückel Theory: ückel Lewis Configuration Interaction (L-CI) J. Chem. Educ , User s manual ulis 3.3 7
NMR Predictor. Introduction
 NMR Predictor This manual gives a walk-through on how to use the NMR Predictor: Introduction NMR Predictor QuickHelp NMR Predictor Overview Chemical features GUI features Usage Menu system File menu Edit
NMR Predictor This manual gives a walk-through on how to use the NMR Predictor: Introduction NMR Predictor QuickHelp NMR Predictor Overview Chemical features GUI features Usage Menu system File menu Edit
HuLiS, a program to teach mesomerism and more
 Journal of Physics: Conference Series PAPER OPEN ACCESS HuLiS, a program to teach mesomerism and more To cite this article: Yannick Carissan et al 2016 J. Phys.: Conf. Ser. 738 012015 View the article
Journal of Physics: Conference Series PAPER OPEN ACCESS HuLiS, a program to teach mesomerism and more To cite this article: Yannick Carissan et al 2016 J. Phys.: Conf. Ser. 738 012015 View the article
Introduction to Hartree-Fock calculations in Spartan
 EE5 in 2008 Hannes Jónsson Introduction to Hartree-Fock calculations in Spartan In this exercise, you will get to use state of the art software for carrying out calculations of wavefunctions for molecues,
EE5 in 2008 Hannes Jónsson Introduction to Hartree-Fock calculations in Spartan In this exercise, you will get to use state of the art software for carrying out calculations of wavefunctions for molecues,
Exercises for Windows
 Exercises for Windows CAChe User Interface for Windows Select tool Application window Document window (workspace) Style bar Tool palette Select entire molecule Select Similar Group Select Atom tool Rotate
Exercises for Windows CAChe User Interface for Windows Select tool Application window Document window (workspace) Style bar Tool palette Select entire molecule Select Similar Group Select Atom tool Rotate
Molecular Modeling and Conformational Analysis with PC Spartan
 Molecular Modeling and Conformational Analysis with PC Spartan Introduction Molecular modeling can be done in a variety of ways, from using simple hand-held models to doing sophisticated calculations on
Molecular Modeling and Conformational Analysis with PC Spartan Introduction Molecular modeling can be done in a variety of ways, from using simple hand-held models to doing sophisticated calculations on
Instructions for Using Spartan 14
 Instructions for Using Spartan 14 Log in to the computer with your Colby ID and password. Click on the Spartan 14 icon in the dock at the bottom of your screen. I. Building Molecules Spartan has one main
Instructions for Using Spartan 14 Log in to the computer with your Colby ID and password. Click on the Spartan 14 icon in the dock at the bottom of your screen. I. Building Molecules Spartan has one main
Calculating Bond Enthalpies of the Hydrides
 Proposed Exercise for the General Chemistry Section of the Teaching with Cache Workbook: Calculating Bond Enthalpies of the Hydrides Contributed by James Foresman, Rachel Fogle, and Jeremy Beck, York College
Proposed Exercise for the General Chemistry Section of the Teaching with Cache Workbook: Calculating Bond Enthalpies of the Hydrides Contributed by James Foresman, Rachel Fogle, and Jeremy Beck, York College
The Hückel Approximation
 The ückel Approximation 1 In this exercise you will use a program called ückel to look at the π molecular orbitals in conjugated molecules. The program calculates the energies and shapes of π (pi) molecular
The ückel Approximation 1 In this exercise you will use a program called ückel to look at the π molecular orbitals in conjugated molecules. The program calculates the energies and shapes of π (pi) molecular
ISIS/Draw "Quick Start"
 ISIS/Draw "Quick Start" Click to print, or click Drawing Molecules * Basic Strategy 5.1 * Drawing Structures with Template tools and template pages 5.2 * Drawing bonds and chains 5.3 * Drawing atoms 5.4
ISIS/Draw "Quick Start" Click to print, or click Drawing Molecules * Basic Strategy 5.1 * Drawing Structures with Template tools and template pages 5.2 * Drawing bonds and chains 5.3 * Drawing atoms 5.4
After obtaining access to the Jmol program you should see an image on your computer screen that looks similar to the following figure. Figure 1.
 A C T I V I T I E S Molecular Modeling Introduction to Molecular Modeling In the following laboratory activities you will examine three-dimensional models of molecules using the computer-based molecular
A C T I V I T I E S Molecular Modeling Introduction to Molecular Modeling In the following laboratory activities you will examine three-dimensional models of molecules using the computer-based molecular
Application Note. U. Heat of Formation of Ethyl Alcohol and Dimethyl Ether. Introduction
 Application Note U. Introduction The molecular builder (Molecular Builder) is part of the MEDEA standard suite of building tools. This tutorial provides an overview of the Molecular Builder s basic functionality.
Application Note U. Introduction The molecular builder (Molecular Builder) is part of the MEDEA standard suite of building tools. This tutorial provides an overview of the Molecular Builder s basic functionality.
Learning to Use Scigress Wagner, Eugene P. (revised May 15, 2018)
 Learning to Use Scigress Wagner, Eugene P. (revised May 15, 2018) Abstract Students are introduced to basic features of Scigress by building molecules and performing calculations on them using semi-empirical
Learning to Use Scigress Wagner, Eugene P. (revised May 15, 2018) Abstract Students are introduced to basic features of Scigress by building molecules and performing calculations on them using semi-empirical
Chemistry 112 Laboratory Experiment 5: Visualizing Molecular Orbitals: A MacSpartan Pro Experience (This experiment will be conducted in OR341)
 Chemistry 112 Laboratory Experiment 5: Visualizing Molecular Orbitals: A MacSpartan Pro Experience (This experiment will be conducted in OR341) Introduction In class we have discussed Lewis structures,
Chemistry 112 Laboratory Experiment 5: Visualizing Molecular Orbitals: A MacSpartan Pro Experience (This experiment will be conducted in OR341) Introduction In class we have discussed Lewis structures,
Synteny Portal Documentation
 Synteny Portal Documentation Synteny Portal is a web application portal for visualizing, browsing, searching and building synteny blocks. Synteny Portal provides four main web applications: SynCircos,
Synteny Portal Documentation Synteny Portal is a web application portal for visualizing, browsing, searching and building synteny blocks. Synteny Portal provides four main web applications: SynCircos,
Project 3: Molecular Orbital Calculations of Diatomic Molecules. This project is worth 30 points and is due on Wednesday, May 2, 2018.
 Chemistry 362 Spring 2018 Dr. Jean M. Standard April 20, 2018 Project 3: Molecular Orbital Calculations of Diatomic Molecules In this project, you will investigate the molecular orbitals and molecular
Chemistry 362 Spring 2018 Dr. Jean M. Standard April 20, 2018 Project 3: Molecular Orbital Calculations of Diatomic Molecules In this project, you will investigate the molecular orbitals and molecular
This experiment is a continuation of the earlier experiment on molecular
 Molecular Modeling: Experiment 2 Page 115 Bonding and Molecular Structure Experiment 2 This experiment is a continuation of the earlier experiment on molecular structure. In that experiment you used a
Molecular Modeling: Experiment 2 Page 115 Bonding and Molecular Structure Experiment 2 This experiment is a continuation of the earlier experiment on molecular structure. In that experiment you used a
How to Create a Substance Answer Set
 How to Create a Substance Answer Set Select among five search techniques to find substances Since substances can be described by multiple names or other characteristics, SciFinder gives you the flexibility
How to Create a Substance Answer Set Select among five search techniques to find substances Since substances can be described by multiple names or other characteristics, SciFinder gives you the flexibility
Chemistry 14CL. Worksheet for the Molecular Modeling Workshop. (Revised FULL Version 2012 J.W. Pang) (Modified A. A. Russell)
 Chemistry 14CL Worksheet for the Molecular Modeling Workshop (Revised FULL Version 2012 J.W. Pang) (Modified A. A. Russell) Structure of the Molecular Modeling Assignment The molecular modeling assignment
Chemistry 14CL Worksheet for the Molecular Modeling Workshop (Revised FULL Version 2012 J.W. Pang) (Modified A. A. Russell) Structure of the Molecular Modeling Assignment The molecular modeling assignment
Technical Note Calculations of Orbital Overlap Range Function EDR( r ; d) and Overlap Distance D(r )using Multiwfn
 Technical Note Calculations of Orbital Overlap Range Function EDR( r ; d) and Overlap Distance D(r )using Multiwfn Abstract The orbital overlap range function EDR( r; d) (J. Chem. Phys. 2014, 141, 144104)
Technical Note Calculations of Orbital Overlap Range Function EDR( r ; d) and Overlap Distance D(r )using Multiwfn Abstract The orbital overlap range function EDR( r; d) (J. Chem. Phys. 2014, 141, 144104)
Computer simulation of radioactive decay
 Computer simulation of radioactive decay y now you should have worked your way through the introduction to Maple, as well as the introduction to data analysis using Excel Now we will explore radioactive
Computer simulation of radioactive decay y now you should have worked your way through the introduction to Maple, as well as the introduction to data analysis using Excel Now we will explore radioactive
The Hückel Approximation Consider a conjugated molecule i.e. a molecule with alternating double and single bonds, as shown in Figure 1.
 The Hückel Approximation In this exercise you will use a program called Hückel to look at the p molecular orbitals in conjugated molecules. The program calculates the energies and shapes of p (pi) molecular
The Hückel Approximation In this exercise you will use a program called Hückel to look at the p molecular orbitals in conjugated molecules. The program calculates the energies and shapes of p (pi) molecular
Introduc)on to IQmol: Part I.!!! Shirin Faraji, Ilya Kaliman, and Anna Krylov
 Introduc)on to IQmol: Part I!!! Shirin Faraji, Ilya Kaliman, and Anna Krylov! 1 Resources! Written by Dr. Andrew Gilbert Keep yourself up to date with IQmol website: http://iqmol.org! IQmol Youtube channel:
Introduc)on to IQmol: Part I!!! Shirin Faraji, Ilya Kaliman, and Anna Krylov! 1 Resources! Written by Dr. Andrew Gilbert Keep yourself up to date with IQmol website: http://iqmol.org! IQmol Youtube channel:
Computational Chemistry Lab Module: Conformational Analysis of Alkanes
 Introduction Computational Chemistry Lab Module: Conformational Analysis of Alkanes In this experiment, we will use CAChe software package to model the conformations of butane, 2-methylbutane, and substituted
Introduction Computational Chemistry Lab Module: Conformational Analysis of Alkanes In this experiment, we will use CAChe software package to model the conformations of butane, 2-methylbutane, and substituted
Molecular Shape and Molecular Polarity. Molecular Shape and Molecular Polarity. Molecular Shape and Molecular Polarity
 Molecular Shape and Molecular Polarity When there is a difference in electronegativity between two atoms, then the bond between them is polar. It is possible for a molecule to contain polar bonds, but
Molecular Shape and Molecular Polarity When there is a difference in electronegativity between two atoms, then the bond between them is polar. It is possible for a molecule to contain polar bonds, but
Structure and Bonding of Organic Molecules
 Chem 220 Notes Page 1 Structure and Bonding of Organic Molecules I. Types of Chemical Bonds A. Why do atoms forms bonds? Atoms want to have the same number of electrons as the nearest noble gas atom (noble
Chem 220 Notes Page 1 Structure and Bonding of Organic Molecules I. Types of Chemical Bonds A. Why do atoms forms bonds? Atoms want to have the same number of electrons as the nearest noble gas atom (noble
Valence electrons octet rule. Lewis structure Lewis structures
 Lewis Dot Diagrams Valence electrons are the electrons in the outermost energy level of an atom. An element with a full octet of valence electrons has a stable configuration. The tendency of bonded atoms
Lewis Dot Diagrams Valence electrons are the electrons in the outermost energy level of an atom. An element with a full octet of valence electrons has a stable configuration. The tendency of bonded atoms
Chapter 8 Covalent Bonding Test B Answers Cordlessore
 We have made it easy for you to find a PDF Ebooks without any digging. And by having access to our ebooks online or by storing it on your computer, you have convenient answers with chapter 8 covalent bonding
We have made it easy for you to find a PDF Ebooks without any digging. And by having access to our ebooks online or by storing it on your computer, you have convenient answers with chapter 8 covalent bonding
Mesomerism, Ring & Substituent Effects, A Computational Chemistry Experiments
 Journal of Laboratory hemical Education 2016, 4(2): 25-34 DI: 10.5923/j.jlce.20160402.01 Mesomerism, Ring & Substituent Effects, A omputational hemistry Experiments Djameleddine Khatmi 1,2, Yannick arissan
Journal of Laboratory hemical Education 2016, 4(2): 25-34 DI: 10.5923/j.jlce.20160402.01 Mesomerism, Ring & Substituent Effects, A omputational hemistry Experiments Djameleddine Khatmi 1,2, Yannick arissan
Dock Ligands from a 2D Molecule Sketch
 Dock Ligands from a 2D Molecule Sketch March 31, 2016 Sample to Insight CLC bio, a QIAGEN Company Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.clcbio.com support-clcbio@qiagen.com
Dock Ligands from a 2D Molecule Sketch March 31, 2016 Sample to Insight CLC bio, a QIAGEN Company Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.clcbio.com support-clcbio@qiagen.com
Molecular Orbitals. Based on Inorganic Chemistry, Miessler and Tarr, 4 th edition, 2011, Pearson Prentice Hall
 Molecular Orbitals Based on Inorganic Chemistry, Miessler and Tarr, 4 th edition, 2011, Pearson Prentice Hall Images from Miessler and Tarr Inorganic Chemistry 2011 obtained from Pearson Education, Inc.
Molecular Orbitals Based on Inorganic Chemistry, Miessler and Tarr, 4 th edition, 2011, Pearson Prentice Hall Images from Miessler and Tarr Inorganic Chemistry 2011 obtained from Pearson Education, Inc.
Using Web-Based Computations in Organic Chemistry
 10/30/2017 1 Using Web-Based Computations in Organic Chemistry John Keller UAF Department of Chemistry & Biochemistry The UAF WebMO site Practical aspects of computational chemistry theory and nomenclature
10/30/2017 1 Using Web-Based Computations in Organic Chemistry John Keller UAF Department of Chemistry & Biochemistry The UAF WebMO site Practical aspects of computational chemistry theory and nomenclature
Comparing whole genomes
 BioNumerics Tutorial: Comparing whole genomes 1 Aim The Chromosome Comparison window in BioNumerics has been designed for large-scale comparison of sequences of unlimited length. In this tutorial you will
BioNumerics Tutorial: Comparing whole genomes 1 Aim The Chromosome Comparison window in BioNumerics has been designed for large-scale comparison of sequences of unlimited length. In this tutorial you will
Curriculum Support Maps for the Study of Indiana Coal (Student Handout)
 Curriculum Support Maps for the Study of Indiana Coal (Student Handout) Introduction In this lesson you will learn how to use a geographic information system (GIS) program (IndianaMap) to investigate coal
Curriculum Support Maps for the Study of Indiana Coal (Student Handout) Introduction In this lesson you will learn how to use a geographic information system (GIS) program (IndianaMap) to investigate coal
OneStop Map Viewer Navigation
 OneStop Map Viewer Navigation» Intended User: Industry Map Viewer users Overview The OneStop Map Viewer is an interactive map tool that helps you find and view information associated with energy development,
OneStop Map Viewer Navigation» Intended User: Industry Map Viewer users Overview The OneStop Map Viewer is an interactive map tool that helps you find and view information associated with energy development,
ST-Links. SpatialKit. Version 3.0.x. For ArcMap. ArcMap Extension for Directly Connecting to Spatial Databases. ST-Links Corporation.
 ST-Links SpatialKit For ArcMap Version 3.0.x ArcMap Extension for Directly Connecting to Spatial Databases ST-Links Corporation www.st-links.com 2012 Contents Introduction... 3 Installation... 3 Database
ST-Links SpatialKit For ArcMap Version 3.0.x ArcMap Extension for Directly Connecting to Spatial Databases ST-Links Corporation www.st-links.com 2012 Contents Introduction... 3 Installation... 3 Database
EXPERIMENT 15: MOLECULAR MODELS
 EXPERIMENT 15: MLEULAR MDELS Introduction: Given formulas of some molecules and ions, you will use the periodic table, valence electron count, and electronegativities to deduce their geometry and polarities.
EXPERIMENT 15: MLEULAR MDELS Introduction: Given formulas of some molecules and ions, you will use the periodic table, valence electron count, and electronegativities to deduce their geometry and polarities.
VSEPR Model. Valence-Shell Electron-Pair Repulsion Bonds (single or multiple) and lone pairs are thought of as charge clouds
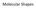 Molecular Shapes VSEPR Model Valence-Shell Electron-Pair Repulsion Bonds (single or multiple) and lone pairs are thought of as charge clouds They repel each other and stay as far away from each other as
Molecular Shapes VSEPR Model Valence-Shell Electron-Pair Repulsion Bonds (single or multiple) and lone pairs are thought of as charge clouds They repel each other and stay as far away from each other as
Experiment 15: Atomic Orbitals, Bond Length, and Molecular Orbitals
 Experiment 15: Atomic Orbitals, Bond Length, and Molecular Orbitals Introduction Molecular orbitals result from the mixing of atomic orbitals that overlap during the bonding process allowing the delocalization
Experiment 15: Atomic Orbitals, Bond Length, and Molecular Orbitals Introduction Molecular orbitals result from the mixing of atomic orbitals that overlap during the bonding process allowing the delocalization
1 An Experimental and Computational Investigation of the Dehydration of 2-Butanol
 1 An Experimental and Computational Investigation of the Dehydration of 2-Butanol Summary. 2-Butanol will be dehydrated to a mixture of 1-butene and cis- and trans-2-butene using the method described in
1 An Experimental and Computational Investigation of the Dehydration of 2-Butanol Summary. 2-Butanol will be dehydrated to a mixture of 1-butene and cis- and trans-2-butene using the method described in
Carbon and Its Compounds
 Chapter 1 Carbon and Its Compounds Copyright 2018 by Nelson Education Limited 1 1.2 Organic Molecules from the Inside Out I: The Modelling of Atoms Copyright 2018 by Nelson Education Limited 2 s orbitals:
Chapter 1 Carbon and Its Compounds Copyright 2018 by Nelson Education Limited 1 1.2 Organic Molecules from the Inside Out I: The Modelling of Atoms Copyright 2018 by Nelson Education Limited 2 s orbitals:
Chapter 10: Modern Atomic Theory and the Periodic Table. How does atomic structure relate to the periodic table? 10.1 Electromagnetic Radiation
 Chapter 10: Modern Atomic Theory and the Periodic Table How does atomic structure relate to the periodic table? 10.1 Electromagnetic Radiation Electromagnetic (EM) radiation is a form of energy that exhibits
Chapter 10: Modern Atomic Theory and the Periodic Table How does atomic structure relate to the periodic table? 10.1 Electromagnetic Radiation Electromagnetic (EM) radiation is a form of energy that exhibits
Moving into the information age: From records to Google Earth
 Moving into the information age: From records to Google Earth David R. R. Smith Psychology, School of Life Sciences, University of Hull e-mail: davidsmith.butterflies@gmail.com Introduction Many of us
Moving into the information age: From records to Google Earth David R. R. Smith Psychology, School of Life Sciences, University of Hull e-mail: davidsmith.butterflies@gmail.com Introduction Many of us
Experiment 1 Scientific Writing Tools
 Experiment 1 Scientific Writing Tools OUTCOMES After completing this experiment, the student should be able to: insert a variety of mathematical equations into a Word document. draw line structures of
Experiment 1 Scientific Writing Tools OUTCOMES After completing this experiment, the student should be able to: insert a variety of mathematical equations into a Word document. draw line structures of
Molecular Shapes and VSEPR (Valence Shell Electron Pair Repulsion Theory)
 AP Chemistry Ms. Ye Name Date Block Molecular Shapes and VSEPR (Valence Shell Electron Pair Repulsion Theory) Go to bit.ly/vseprshapes Introduction Atoms bond to satisfy their need for more electrons.
AP Chemistry Ms. Ye Name Date Block Molecular Shapes and VSEPR (Valence Shell Electron Pair Repulsion Theory) Go to bit.ly/vseprshapes Introduction Atoms bond to satisfy their need for more electrons.
Ligand Scout Tutorials
 Ligand Scout Tutorials Step : Creating a pharmacophore from a protein-ligand complex. Type ke6 in the upper right area of the screen and press the button Download *+. The protein will be downloaded and
Ligand Scout Tutorials Step : Creating a pharmacophore from a protein-ligand complex. Type ke6 in the upper right area of the screen and press the button Download *+. The protein will be downloaded and
Calculating NMR Chemical Shifts for beta-ionone O
 Calculating NMR Chemical Shifts for beta-ionone O Molecular orbital calculations can be used to get good estimates for chemical shifts. In this exercise we will calculate the chemical shifts for beta-ionone.
Calculating NMR Chemical Shifts for beta-ionone O Molecular orbital calculations can be used to get good estimates for chemical shifts. In this exercise we will calculate the chemical shifts for beta-ionone.
Section 6.2 1/13/2014. Most Chemical Compounds. Molecular (or Covalent) Compound. Covalent Bonding and Molecular Compounds
 Section 6.2 Covalent Bonding and Molecular Compounds Most Chemical Compounds Are molecules, a neutral group of atoms that are held together by covalent bonds. It is a single unit capable of existing on
Section 6.2 Covalent Bonding and Molecular Compounds Most Chemical Compounds Are molecules, a neutral group of atoms that are held together by covalent bonds. It is a single unit capable of existing on
Basic chromatographic parameters and optimization in LC
 AM0925 Assignment Basic chromatographic parameters and optimization in LC Introduction This is a computer exercise where you will apply a simulator of reversed phase LC to study the influence of chromatographic
AM0925 Assignment Basic chromatographic parameters and optimization in LC Introduction This is a computer exercise where you will apply a simulator of reversed phase LC to study the influence of chromatographic
IFM Chemistry Computational Chemistry 2010, 7.5 hp LAB2. Computer laboratory exercise 1 (LAB2): Quantum chemical calculations
 Computer laboratory exercise 1 (LAB2): Quantum chemical calculations Introduction: The objective of the second computer laboratory exercise is to get acquainted with a program for performing quantum chemical
Computer laboratory exercise 1 (LAB2): Quantum chemical calculations Introduction: The objective of the second computer laboratory exercise is to get acquainted with a program for performing quantum chemical
Lab Activity 4: Structures and Properties of Aliphatic Compounds
 Chemistry 2202 Structures and Properties of Aliphatic Compounds Page 1 of 6 Lab Activity 4: Structures and Properties of Aliphatic Compounds Introduction Aliphatic compounds are hydrocarbons that have
Chemistry 2202 Structures and Properties of Aliphatic Compounds Page 1 of 6 Lab Activity 4: Structures and Properties of Aliphatic Compounds Introduction Aliphatic compounds are hydrocarbons that have
Manual Seatrack Web Brofjorden
 December 2011 Manual Seatrack Web Brofjorden A user-friendly system for forecasts and backtracking of drift and spreading of oil, chemicals and substances in water 1. Introduction and Background... 3 1.1
December 2011 Manual Seatrack Web Brofjorden A user-friendly system for forecasts and backtracking of drift and spreading of oil, chemicals and substances in water 1. Introduction and Background... 3 1.1
To visualize the three-dimensional structures of some common molecules. To obtain bond angle, bond length, and hybridization data for molecules.
 Molecular Geometry PURPOSE A B C To explore some simple molecular structures. To explore the relationship between bond order and bond length. To explore resonance structures. GOALS To compare Lewis structures
Molecular Geometry PURPOSE A B C To explore some simple molecular structures. To explore the relationship between bond order and bond length. To explore resonance structures. GOALS To compare Lewis structures
GIS Workshop UCLS_Fall Forum 2014 Sowmya Selvarajan, PhD TABLE OF CONTENTS
 TABLE OF CONTENTS TITLE PAGE NO. 1. ArcGIS Basics I 2 a. Open and Save a Map Document 2 b. Work with Map Layers 2 c. Navigate in a Map Document 4 d. Measure Distances 4 2. ArcGIS Basics II 5 a. Work with
TABLE OF CONTENTS TITLE PAGE NO. 1. ArcGIS Basics I 2 a. Open and Save a Map Document 2 b. Work with Map Layers 2 c. Navigate in a Map Document 4 d. Measure Distances 4 2. ArcGIS Basics II 5 a. Work with
Marvin. Sketching, viewing and predicting properties with Marvin - features, tips and tricks. Gyorgy Pirok. Solutions for Cheminformatics
 Marvin Sketching, viewing and predicting properties with Marvin - features, tips and tricks Gyorgy Pirok Solutions for Cheminformatics The Marvin family The Marvin toolkit provides web-enabled components
Marvin Sketching, viewing and predicting properties with Marvin - features, tips and tricks Gyorgy Pirok Solutions for Cheminformatics The Marvin family The Marvin toolkit provides web-enabled components
Chapter 8 Covalent Bonding Study Guide Answers Pearson
 Chapter 8 Covalent Bonding Study Guide Answers Pearson We have made it easy for you to find a PDF Ebooks without any digging. And by having access to our ebooks online or by storing it on your computer,
Chapter 8 Covalent Bonding Study Guide Answers Pearson We have made it easy for you to find a PDF Ebooks without any digging. And by having access to our ebooks online or by storing it on your computer,
Lesson Plan 2 - Middle and High School Land Use and Land Cover Introduction. Understanding Land Use and Land Cover using Google Earth
 Understanding Land Use and Land Cover using Google Earth Image an image is a representation of reality. It can be a sketch, a painting, a photograph, or some other graphic representation such as satellite
Understanding Land Use and Land Cover using Google Earth Image an image is a representation of reality. It can be a sketch, a painting, a photograph, or some other graphic representation such as satellite
Chapter 7 Ionic And Metallic Bonding Answer Key Pearson Education
 Chapter 7 Ionic And Metallic Bonding Answer Key Pearson Education CHAPTER 7 IONIC AND METALLIC BONDING ANSWER KEY PEARSON EDUCATION PDF - Are you looking for chapter 7 ionic and metallic bonding answer
Chapter 7 Ionic And Metallic Bonding Answer Key Pearson Education CHAPTER 7 IONIC AND METALLIC BONDING ANSWER KEY PEARSON EDUCATION PDF - Are you looking for chapter 7 ionic and metallic bonding answer
CHEM 463: Advanced Inorganic Chemistry Modeling Metalloproteins for Structural Analysis
 CHEM 463: Advanced Inorganic Chemistry Modeling Metalloproteins for Structural Analysis Purpose: The purpose of this laboratory is to introduce some of the basic visualization and modeling tools for viewing
CHEM 463: Advanced Inorganic Chemistry Modeling Metalloproteins for Structural Analysis Purpose: The purpose of this laboratory is to introduce some of the basic visualization and modeling tools for viewing
Chapter 8 Covalent Bonding Study Guide Answers Pearson
 Chapter 8 Covalent Bonding Study Guide Answers Pearson We have made it easy for you to find a PDF Ebooks without any digging. And by having access to our ebooks online or by storing it on your computer,
Chapter 8 Covalent Bonding Study Guide Answers Pearson We have made it easy for you to find a PDF Ebooks without any digging. And by having access to our ebooks online or by storing it on your computer,
Be H. Delocalized Bonding. Localized Bonding. σ 2. σ 1. Two (sp-1s) Be-H σ bonds. The two σ bonding MO s in BeH 2. MO diagram for BeH 2
 The Delocalized Approach to Bonding: The localized models for bonding we have examined (Lewis and VBT) assume that all electrons are restricted to specific bonds between atoms or in lone pairs. In contrast,
The Delocalized Approach to Bonding: The localized models for bonding we have examined (Lewis and VBT) assume that all electrons are restricted to specific bonds between atoms or in lone pairs. In contrast,
Orange Visualization Tool (OVT) Manual
 Orange Visualization Tool (OVT) Manual This manual describes the features of the tool and how to use it. 1. Contents of the OVT Once the OVT is open (the first time it may take some seconds), it should
Orange Visualization Tool (OVT) Manual This manual describes the features of the tool and how to use it. 1. Contents of the OVT Once the OVT is open (the first time it may take some seconds), it should
Refine & Validate. In the *.res file, be sure to add the following four commands after the UNIT instruction and before any atoms: ACTA CONF WPDB -2
 Refine & Validate Refinement is simply a way to improve the fit between the measured intensities and the intensities calculated from the model. The peaks in the difference map and the list of worst fitting
Refine & Validate Refinement is simply a way to improve the fit between the measured intensities and the intensities calculated from the model. The peaks in the difference map and the list of worst fitting
Worksheet: Coulomb Force Simulation
 Screen shot of simulation showing force between two charged particles. Worksheet: Coulomb Force Simulation Key Topic/Concept: Coulomb s Law and Newton s 3 rd Law Materials: One guide sheet for each student
Screen shot of simulation showing force between two charged particles. Worksheet: Coulomb Force Simulation Key Topic/Concept: Coulomb s Law and Newton s 3 rd Law Materials: One guide sheet for each student
Chapter 12 CHEMICAL BONDING
 Chapter 12 CHEMICAL BONDING Sharing electrons is fun! H F Do you smell what the Rock is cooking? I. Types of Chemical Bonds A. Formation of Covalent Bonds B. Lewis Symbols and Covalent Bonding C. Other
Chapter 12 CHEMICAL BONDING Sharing electrons is fun! H F Do you smell what the Rock is cooking? I. Types of Chemical Bonds A. Formation of Covalent Bonds B. Lewis Symbols and Covalent Bonding C. Other
Ethene. Introduction. The ethene molecule is planar (i.e. all the six atoms lie in the same plane) and has a high degree of symmetry:
 FY1006 Innføring i kvantefysikk og TFY4215 Kjemisk fysikk og kvantemekanikk Spring 2012 Chemical Physics Exercise 1 To be delivered by Friday 27.04.12 Introduction Ethene. Ethylene, C 2 H 4, or ethene,
FY1006 Innføring i kvantefysikk og TFY4215 Kjemisk fysikk og kvantemekanikk Spring 2012 Chemical Physics Exercise 1 To be delivered by Friday 27.04.12 Introduction Ethene. Ethylene, C 2 H 4, or ethene,
Molecular Orbitals in Inorganic Chemistry. Dr. P. Hunt Rm 167 (Chemistry)
 Molecular Orbitals in Inorganic Chemistry Dr. P. Hunt p.hunt@imperial.ac.uk Rm 167 (Chemistry http://www.ch.ic.ac.uk/hunt/ Outline LCAO theory orbital size matters where to put the FO energy levels the
Molecular Orbitals in Inorganic Chemistry Dr. P. Hunt p.hunt@imperial.ac.uk Rm 167 (Chemistry http://www.ch.ic.ac.uk/hunt/ Outline LCAO theory orbital size matters where to put the FO energy levels the
Tutorial. Getting started. Sample to Insight. March 31, 2016
 Getting started March 31, 2016 Sample to Insight CLC bio, a QIAGEN Company Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.clcbio.com support-clcbio@qiagen.com Getting started
Getting started March 31, 2016 Sample to Insight CLC bio, a QIAGEN Company Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.clcbio.com support-clcbio@qiagen.com Getting started
MDSS Functional Prototype Display System Preview April 2002
 MDSS Functional Prototype Display System Preview April 2002 Bill Mahoney National Center for Atmospheric Research Images shown are valid as of 15 April 2002 NCAR MDSS Display System - Overview The MDSS
MDSS Functional Prototype Display System Preview April 2002 Bill Mahoney National Center for Atmospheric Research Images shown are valid as of 15 April 2002 NCAR MDSS Display System - Overview The MDSS
EXPERIMENT 12: MOLECULAR ARCHITECTURE
 Name Section EXPERIMENT 12: MLECULAR ARCITECTURE PRE-LABRATRY QUESTINS The following preparatory questions should be answered before coming to lab. They are intended to introduce you to several ideas important
Name Section EXPERIMENT 12: MLECULAR ARCITECTURE PRE-LABRATRY QUESTINS The following preparatory questions should be answered before coming to lab. They are intended to introduce you to several ideas important
Watershed Modeling Orange County Hydrology Using GIS Data
 v. 10.0 WMS 10.0 Tutorial Watershed Modeling Orange County Hydrology Using GIS Data Learn how to delineate sub-basins and compute soil losses for Orange County (California) hydrologic modeling Objectives
v. 10.0 WMS 10.0 Tutorial Watershed Modeling Orange County Hydrology Using GIS Data Learn how to delineate sub-basins and compute soil losses for Orange County (California) hydrologic modeling Objectives
Chapter 6. Table of Contents. Section 1 Covalent Bonds. Section 2 Drawing and Naming Molecules. Section 3 Molecular Shapes. Covalent Compounds
 Covalent Compounds Table of Contents Section 1 Covalent Bonds Section 2 Drawing and Naming Molecules Section 3 Molecular Shapes Section 1 Covalent Bonds Bellringer Make a list of the elements that form
Covalent Compounds Table of Contents Section 1 Covalent Bonds Section 2 Drawing and Naming Molecules Section 3 Molecular Shapes Section 1 Covalent Bonds Bellringer Make a list of the elements that form
Introduction Molecular Structure Script Console External resources Advanced topics. JMol tutorial. Giovanni Morelli.
 Gen 19th, 2017 1 2 Create and edit Display and view Mesurament and labelling Surface and Orbitals 3 4 from Database Protein Enzyme Crystal Structure and Unit Cell 5 Symmetry Animation General information
Gen 19th, 2017 1 2 Create and edit Display and view Mesurament and labelling Surface and Orbitals 3 4 from Database Protein Enzyme Crystal Structure and Unit Cell 5 Symmetry Animation General information
OCEAN/ESS 410 Lab 4. Earthquake location
 Lab 4. Earthquake location To complete this exercise you will need to (a) Complete the table on page 2. (b) Identify phases on the seismograms on pages 3-6 as requested on page 11. (c) Locate the earthquake
Lab 4. Earthquake location To complete this exercise you will need to (a) Complete the table on page 2. (b) Identify phases on the seismograms on pages 3-6 as requested on page 11. (c) Locate the earthquake
E-BOOK // OF CHEMICAL FORMULAS OWNERS MANUAL EBOOK
 02 January, 2018 E-BOOK // OF CHEMICAL FORMULAS OWNERS MANUAL EBOOK Document Filetype: PDF 389.32 KB 0 E-BOOK // OF CHEMICAL FORMULAS OWNERS MANUAL EBOOK This is a list of common chemical compounds with
02 January, 2018 E-BOOK // OF CHEMICAL FORMULAS OWNERS MANUAL EBOOK Document Filetype: PDF 389.32 KB 0 E-BOOK // OF CHEMICAL FORMULAS OWNERS MANUAL EBOOK This is a list of common chemical compounds with
Essential Organic Chemistry. Paula Y. Bruice Second Edition
 Essential rganic hemistry Paula Y. Bruice Second Edition Pearson Education Limited Edinburgh Gate arlow Essex M20 2JE England and Associated ompanies throughout the world Visit us on the World Wide Web
Essential rganic hemistry Paula Y. Bruice Second Edition Pearson Education Limited Edinburgh Gate arlow Essex M20 2JE England and Associated ompanies throughout the world Visit us on the World Wide Web
Lab 1 Uniform Motion - Graphing and Analyzing Motion
 Lab 1 Uniform Motion - Graphing and Analyzing Motion Objectives: < To observe the distance-time relation for motion at constant velocity. < To make a straight line fit to the distance-time data. < To interpret
Lab 1 Uniform Motion - Graphing and Analyzing Motion Objectives: < To observe the distance-time relation for motion at constant velocity. < To make a straight line fit to the distance-time data. < To interpret
General Chemistry Lab Molecular Modeling
 PURPOSE The objectives of this experiment are PROCEDURE General Chemistry Lab Molecular Modeling To learn how to use molecular modeling software, a commonly used tool in chemical research and industry.
PURPOSE The objectives of this experiment are PROCEDURE General Chemistry Lab Molecular Modeling To learn how to use molecular modeling software, a commonly used tool in chemical research and industry.
NPA/NBO-Analysis. Examples POP =
 NPA/NBO-Analysis Examples POP = NBO Requests a full Natural Bond Orbital analysis, using NBO version 3 NPA Requests just the Natural Population Analysis phase of NBO. NBORead Requests a full NBO analysis,
NPA/NBO-Analysis Examples POP = NBO Requests a full Natural Bond Orbital analysis, using NBO version 3 NPA Requests just the Natural Population Analysis phase of NBO. NBORead Requests a full NBO analysis,
Chapter 6 Molecular Structure
 hapter 6 Molecular Structure 1. Draw the Lewis structure of each of the following ions, showing all nonzero formal charges. Indicate whether each ion is linear or bent. If the ion is bent, what is the
hapter 6 Molecular Structure 1. Draw the Lewis structure of each of the following ions, showing all nonzero formal charges. Indicate whether each ion is linear or bent. If the ion is bent, what is the
Mnova Software for Analyzing Reaction Monitoring NMR Spectra
 Mnova Software for Analyzing Reaction Monitoring NMR Spectra Version 10 Chen Peng, PhD, VP of Business Development, US & China Mestrelab Research SL San Diego, CA, USA chen.peng@mestrelab.com 858.736.4563
Mnova Software for Analyzing Reaction Monitoring NMR Spectra Version 10 Chen Peng, PhD, VP of Business Development, US & China Mestrelab Research SL San Diego, CA, USA chen.peng@mestrelab.com 858.736.4563
The Nature of Covalent Bonding
 Chemistry 1 of 50 The colors in this map indicate the concentrations of ozone in various parts of Earth s atmosphere. Oxygen atoms can join in pairs to form the oxygen you breathe and can also join in
Chemistry 1 of 50 The colors in this map indicate the concentrations of ozone in various parts of Earth s atmosphere. Oxygen atoms can join in pairs to form the oxygen you breathe and can also join in
Molecular Orbitals. Chapter 9. Sigma bonding orbitals. Sigma bonding orbitals. Pi bonding orbitals. Sigma and pi bonds
 Molecular Orbitals Chapter 9 Orbitals and Covalent Bond The overlap of atomic orbitals from separate atoms makes molecular orbitals Each molecular orbital has room for two electrons Two types of MO Sigma
Molecular Orbitals Chapter 9 Orbitals and Covalent Bond The overlap of atomic orbitals from separate atoms makes molecular orbitals Each molecular orbital has room for two electrons Two types of MO Sigma
BD1. Reciprocal space and Fourier transforms. IB Mineral Sciences Module B: Reciprocal Space, Symmetry and Crystallography
 Reciprocal space and Fourier transforms The aim of this practical is to give you a chance to develop your intuition about the relationship between real space and reciprocal space. There are two halves
Reciprocal space and Fourier transforms The aim of this practical is to give you a chance to develop your intuition about the relationship between real space and reciprocal space. There are two halves
Patrick: An Introduction to Medicinal Chemistry 5e MOLECULAR MODELLING EXERCISES CHAPTER 17
 MOLECULAR MODELLING EXERCISES CHAPTER 17 Exercise 17.6 Conformational analysis of n-butane Introduction Figure 1 Butane Me Me In this exercise, we will consider the possible stable conformations of butane
MOLECULAR MODELLING EXERCISES CHAPTER 17 Exercise 17.6 Conformational analysis of n-butane Introduction Figure 1 Butane Me Me In this exercise, we will consider the possible stable conformations of butane
Valence Bond Theory - Description
 Bonding and Molecular Structure - PART 2 - Valence Bond Theory and Hybridization 1. Understand and be able to describe the Valence Bond Theory description of covalent bond formation. 2. Understand and
Bonding and Molecular Structure - PART 2 - Valence Bond Theory and Hybridization 1. Understand and be able to describe the Valence Bond Theory description of covalent bond formation. 2. Understand and
Tutorial 2: Analysis of DIA data in Skyline
 Tutorial 2: Analysis of DIA data in Skyline In this tutorial we will learn how to use Skyline to perform targeted post-acquisition analysis for peptide and inferred protein detection and quantitation using
Tutorial 2: Analysis of DIA data in Skyline In this tutorial we will learn how to use Skyline to perform targeted post-acquisition analysis for peptide and inferred protein detection and quantitation using
Molecular Orbitals for Ozone
 Molecular Orbitals for Ozone Purpose: In this exercise you will do semi-empirical molecular orbital calculations on ozone with the goal of understanding the molecular orbital print out provided by Spartan
Molecular Orbitals for Ozone Purpose: In this exercise you will do semi-empirical molecular orbital calculations on ozone with the goal of understanding the molecular orbital print out provided by Spartan
Gas Pressure and Temperature Relationships *
 Gas Pressure and Temperature Relationships * MoLE Activities To begin this assignment you must be able to log on to the Internet (the software requires OSX for mac users). Type the following address into
Gas Pressure and Temperature Relationships * MoLE Activities To begin this assignment you must be able to log on to the Internet (the software requires OSX for mac users). Type the following address into
Review Package #3 Atomic Models and Subatomic Particles The Periodic Table Chemical Bonding
 Chemistry 11 Review Package #3 Atomic Models and Subatomic Particles The Periodic Table Chemical Bonding 1. Atomic Models and Subatomic Particles: A. Subatomic Particles and Average Atomic Mass: - Subatomic
Chemistry 11 Review Package #3 Atomic Models and Subatomic Particles The Periodic Table Chemical Bonding 1. Atomic Models and Subatomic Particles: A. Subatomic Particles and Average Atomic Mass: - Subatomic
We can model covalent bonding in molecules in essentially two ways:
 CHEM 2060 Lecture 22: VB Theory L22-1 PART FIVE: The Covalent Bond We can model covalent bonding in molecules in essentially two ways: 1. Localized Bonds (retains electron pair concept of Lewis Structures)
CHEM 2060 Lecture 22: VB Theory L22-1 PART FIVE: The Covalent Bond We can model covalent bonding in molecules in essentially two ways: 1. Localized Bonds (retains electron pair concept of Lewis Structures)
Lewis Structures (The Localized Electron Model)
 Lewis Structures (The Localized Electron Model) G. N. Lewis 1875-1946 Using electron-dot symbols, G. N. Lewis developed the Localized Electron Model of chemical bonding (1916) in which valence electrons
Lewis Structures (The Localized Electron Model) G. N. Lewis 1875-1946 Using electron-dot symbols, G. N. Lewis developed the Localized Electron Model of chemical bonding (1916) in which valence electrons
CHAPTER TEN MOLECULAR GEOMETRY MOLECULAR GEOMETRY V S E P R CHEMICAL BONDING II: MOLECULAR GEOMETRY AND HYBRIDIZATION OF ATOMIC ORBITALS
 CHAPTER TEN CHEMICAL BONDING II: AND HYBRIDIZATION O ATOMIC ORBITALS V S E P R VSEPR Theory In VSEPR theory, multiple bonds behave like a single electron pair Valence shell electron pair repulsion (VSEPR)
CHAPTER TEN CHEMICAL BONDING II: AND HYBRIDIZATION O ATOMIC ORBITALS V S E P R VSEPR Theory In VSEPR theory, multiple bonds behave like a single electron pair Valence shell electron pair repulsion (VSEPR)
CHEMISTRY 110 EXAM 2 Feb 25, 2013 FORM A
 EMISTRY 110 EXAM 2 Feb 25, 2013 FORM A 1. ow many valence electrons and lone pairs are in the structure of the ammonium ion? # valence electrons # lone pairs A. 8 0 B. 10 1. 8 1 D. 10 2 E. 12 3 2. Which
EMISTRY 110 EXAM 2 Feb 25, 2013 FORM A 1. ow many valence electrons and lone pairs are in the structure of the ammonium ion? # valence electrons # lone pairs A. 8 0 B. 10 1. 8 1 D. 10 2 E. 12 3 2. Which
The CSC Interface to Sky in Google Earth
 The CSC Interface to Sky in Google Earth CSC Threads The CSC Interface to Sky in Google Earth 1 Table of Contents The CSC Interface to Sky in Google Earth - CSC Introduction How to access CSC data with
The CSC Interface to Sky in Google Earth CSC Threads The CSC Interface to Sky in Google Earth 1 Table of Contents The CSC Interface to Sky in Google Earth - CSC Introduction How to access CSC data with
VCell Tutorial. Building a Rule-Based Model
 VCell Tutorial Building a Rule-Based Model We will demonstrate how to create a rule-based model of EGFR receptor interaction with two adapter proteins Grb2 and Shc. A Receptor-monomer reversibly binds
VCell Tutorial Building a Rule-Based Model We will demonstrate how to create a rule-based model of EGFR receptor interaction with two adapter proteins Grb2 and Shc. A Receptor-monomer reversibly binds
CHEMICAL BONDING IN MATTER Text p ) Chemical Bonds are attractive electrostatic forces that hold atoms or ions together in a substance.
 CHEMICAL BONDING IN MATTER Text p. 162 164) Chemical Bonds are attractive electrostatic forces that hold atoms or ions together in a substance. Bohr's Model of the Atom protons and neutrons are located
CHEMICAL BONDING IN MATTER Text p. 162 164) Chemical Bonds are attractive electrostatic forces that hold atoms or ions together in a substance. Bohr's Model of the Atom protons and neutrons are located
Computational Chemistry Using the University of Alaska WebMO Site
 2/7/2017 1 Computational Chemistry Using the University of Alaska WebMO Site John Keller Department of Chemistry & Biochemistry University of Alaska Fairbanks Intro and operation of WebMO and MOPAC Basic
2/7/2017 1 Computational Chemistry Using the University of Alaska WebMO Site John Keller Department of Chemistry & Biochemistry University of Alaska Fairbanks Intro and operation of WebMO and MOPAC Basic
Chemical Bonds, Lewis Structures, Bond Order, and Formal Charge
 Chemical Bonds, Lewis Structures, Bond Order, and Formal Charge PRELAB ASSIGNMENT Read the entire laboratory write up. Write an objective, any hazards associated with this lab, and answer the following
Chemical Bonds, Lewis Structures, Bond Order, and Formal Charge PRELAB ASSIGNMENT Read the entire laboratory write up. Write an objective, any hazards associated with this lab, and answer the following
Lewis Structures and Molecular Shapes
 Lewis Structures and Molecular Shapes Rules for Writing Lewis Structures 1. Determine the correct skeleton structure (connectivity of atoms). Usually, the most electronegative atoms go around the edges
Lewis Structures and Molecular Shapes Rules for Writing Lewis Structures 1. Determine the correct skeleton structure (connectivity of atoms). Usually, the most electronegative atoms go around the edges
Tutorial: Structural Analysis of a Protein-Protein Complex
 Molecular Modeling Section (MMS) Department of Pharmaceutical and Pharmacological Sciences University of Padova Via Marzolo 5-35131 Padova (IT) @contact: stefano.moro@unipd.it Tutorial: Structural Analysis
Molecular Modeling Section (MMS) Department of Pharmaceutical and Pharmacological Sciences University of Padova Via Marzolo 5-35131 Padova (IT) @contact: stefano.moro@unipd.it Tutorial: Structural Analysis
