BAT Biodiversity Assessment Tools, an R package for the measurement and estimation of alpha and beta taxon, phylogenetic and functional diversity
|
|
|
- Willa Pitts
- 5 years ago
- Views:
Transcription
1 Methods in Ecology and Evolution 2015, 6, doi: / X APPLICATION BAT Biodiversity Assessment Tools, an R package for the measurement and estimation of alpha and beta taxon, phylogenetic and functional diversity Pedro Cardoso 1,2 *, Francßois Rigal 2 and Jose C. Carvalho 2,3 1 Finnish Museum of Natural History, University of Helsinki, Helsinki, Finland; 2 CE3C - Centre for Ecology, Evolution and Environmental Changes / Azorean Biodiversity Group, University of the Azores, Angra do Heroısmo, Portugal; and 3 CBMA Centre of Molecular and Environmental Biology, Department of Biology, University of Minho, Braga, Portugal Summary 1. Novel algorithms have been recently developed to estimate alpha and partition beta diversity in all their dimensions (taxon, phylogenetic and functional diversity TD, PD and FD), whether communities are completely sampled or not. 2. The R package BAT Biodiversity Assessment Tools performs a number of analyses based on either species identities (TD) or trees depicting species relationships (PD and FD). Functions include building randomized accumulation curves for alpha and beta diversity, alpha diversity estimation from incomplete samples and the partitioning of beta diversity in its replacement and richness difference components. 3. All functions allow the rarefaction of communities. Estimation methods include curve-fitting and nonparametric algorithms. Beta diversity indices include the Jaccard and Sørensen families of measures and deal with both incidence and abundance data. Two auxiliary functions that allow judging the efficiency of the algorithms are also included. 4. Several examples are shown using the data included in the package, which demonstrate the usefulness of the different methods. The BAT package constitutes an open platform for further development of new biodiversity assessment tools. Key-words: accumulation curves, biodiversity estimation, diversity partition, extrapolation, nonparametric estimators, sampling bias Introduction The most studied biodiversity components are alpha diversity (a) that measures the diversity intrinsictoeachcommunityor site and beta diversity (b) that reflects the differences among communities or sites. Biodiversity can also be characterized according to different aspects measured by how species relate. Taxon diversity (TD) is the most common and is quantified based on the number of taxa and, often, on the distribution of abundances. However, TD disregards the fact that communities are composed of species with different evolutionary histories and a diverse array of ecological functions. Thus, the last decade has seen a growing interest in alternative representations of biodiversity, including phylogenetic diversity (PD) and functional diversity (FD) (Stegen & Hurlbert 2011). Studying biodiversity patterns and the respective driving processes in all these dimensions is invariably challenging. As alpha diversity (TD, PD or FD) increases with the number of individuals or sampling units, observed richness almost invariably underestimates true richness (Coddington et al. *Correspondence author. pedro.cardoso@helsinki.fi 2009; Cardoso et al. 2014a). Without complete inventories of communities, comparisons of alpha diversity values cannot be reliably made if one does not consider the sampling effort or completeness attained (Gotelli & Colwell 2001). The same problems have been identified for beta diversity, since underestimation of similarity also occurs because of the failure to account for unseen shared species (Cardoso, Borges & Veech 2009). Some measures are more sensitive than others, and the comparison of randomized accumulation curves for both alpha (Gotelli & Colwell 2001) and beta (Cardoso, Borges & Veech 2009) diversity has been advocated as a previous step in many studies for which undersampling is suspected. Estimating species richness from incomplete samples has long been proposed as a potential way of dealing with undersampling (Colwell & Coddington 1994). Two kinds of data may be used for estimating diversity: (i) abundance data and (ii) incidence data. A number of approaches for estimating true diversity are possible, mainly the fit of asymptotic functions to accumulation curves (Soberon & Llorente 1993) and the use of nonparametric estimators based on the abundance or incidence of rare species. Among the latter are the jackknife (Heltshe & Forrester 1983) and Anne Chao s formulas (Chao 2014 The Authors. Methods in Ecology and Evolution 2014 British Ecological Society
2 BAT Biodiversity assessment tools , 1987). All these approaches were first developed for TD and only recently adapted for other dimensions of biodiversity, namely PD and FD (Cardoso et al. 2014a). The term beta diversity has been used to refer to a variety of phenomena, although all of these encompass compositional heterogeneity between communities. Two distinct processes shape communities and their differences: species replacement and species loss (or gain) (Williams, de Klerk & Crowe 1999; Lennon et al. 2001). How to partition these different components has been a field of much recent development (Baselga 2010, 2012; Podani & Schmera 2011; Carvalho, Cardoso & Gomes 2012; Carvalho et al. 2013; Legendre 2014), including when phylogenetic or functional relations between species are taken into account (Cardoso et al. 2014c). Our objective was to develop code for the measurement and the estimation of alpha and beta diversities in their multiple facets (taxon, phylogenetic and functional) along with the partitioning of beta diversity. The BAT package The R package BAT Biodiversity Assessment Tools (Cardoso, Rigal & Carvalho 2014b) allows performing a number of analyses based on either species identities (TD) or ultrametric trees depicting species relationships (PD and FD). Functions include building randomized accumulation curves for alpha (alpha, alpha.accum) and beta (beta.accum) diversity, alpha diversity estimation from incomplete samples (alpha.estimate) and the partitioning of beta diversity in its replacement and richness differences components (beta), including for multiple sites simultaneously (beta.multi). Beta diversity indices include the Jaccard and Sørensen families of measures and deal with both incidence and abundance data. The package also implements two auxiliary functions that allow judging the efficiency of the algorithms (accuracy, slope). The BAT package is written in R and can be installed from the Comprehensive R Archive Network ( org/web/packages/bat/). The raw data accepted by most functions are (1) a sites/sampling units (rows) by species (columns) matrix, with either abundance or incidence data and (2) a phylo object as implemented in the R package ape (Paradis, Claude & Strimmer 2004) or a hclust object (both used only for PD or FD). All functions allow the rarefaction of communities. The functions provided by the package may be grouped into three categories, according to their scope, as follows: ALPHA DIVERSITY 1. alpha (comm, tree, raref, runs) calculates the observed alpha diversity of multiple sites with possible rarefaction. The argument comm is a sites 9 species matrix, with either abundance or incidence data. As in all functions below, tree is a phylo or hclust object (used only for PD or FD). Calculations of PD and FD are based on Faith (1992) and Petchey & Gaston (2002), respectively, which measure PD and FD of a community as the total branch length of a tree linking all species represented in such community. As in other functions below, raref specifies the number of individuals to be used for individualbased rarefaction. runs is the number of rarefaction procedures to execute. The function returns a matrix of sites 9 diversity values. 2. alpha.accum (comm, tree, func, runs) estimates alpha diversity of a single site with accumulation of sampling units. The argument comm is a sampling units 9 species matrix, with either abundance or incidence data. func is the class of estimators to be used, either curve-fitting, nonparametric or, for PD and FD, based on the completeness of TD (see Cardoso et al. 2014a). runs is the number of random permutations to be made to the sampling units order. The function returns a matrix of sampling units 9 diversity values (individuals, observed and estimated diversity). 3. alpha.estimate (comm, tree, func) estimates alpha diversity of multiple sites simultaneously. The argument comm is a sites 9 species matrix, with either abundance or number of incidences (sampling units where each species occurs) data. As above, func is the class of estimators to be used, but only nonparametric (jackknife and Chao for both incidence and abundance data) or, for PD and FD, based on the completeness of TD, are available (as no sampling unit data are given for curve fitting). The function returns a matrix of sites 9 diversity values (individuals, observed and estimated diversity). BETA DIVERSITY 4. beta (comm, tree, abund, func, raref, runs) calculates beta diversity and its partitioning for multiple sites simultaneously with possible rarefaction. comm is a sites 9 species matrix, with either abundance or incidence data. abund is a Boolean indicating whether abundance data should be used or converted to incidence before analysis. As for all beta diversity functions in the package, func indicates whether the Jaccard or Sørensen family of beta diversity measures should be used. The function returns three distance matrices with beta diversity measured between all pairs of sites (dist object), one per each of the three beta diversity measures (b total overall beta diversity, b repl the replacement component and b rich the richness differences component). 5. beta.accum (comm1, comm2, tree, abund, func, runs) computes an accumulation curve for the beta diversity between two communities (comm1 and comm2). Both comm1 and comm2 are sampling units 9 species matrices, with either abundance or incidence data. The function returns three matrices of sampling units 9 diversity values, one per each of the three beta diversity measures (individuals and observed diversity). 6. beta.multi (comm, tree, abund, func, raref, runs) calculates beta diversity among multiple sites with possible rarefaction. Themultiplesitesmeasureiscalculatedastheaverageorvariance of all pairwise values. The argument comm is a sites 9 species matrix, with either abundance or incidence data. The function returns a matrix of beta measures 9 diversity values (average and variance).
3 234 P. Cardoso, F. Rigal & J. C. Carvalho RELIABILITY 7. accuracy (accum, target) calculates the scaled mean squared error of accumulation curves compared with a known true diversity value (target). The argument accum is a matrix resulting from the alpha.accum or beta.accum functions (sampling units 9 diversity values). The function returns a vector with accuracy values for all observed and estimated curves. 8. slope (accum) calculates the slope between adjacent points along the accumulation curves produced by either alpha.accum or beta.accum (the single argument accum). This value reflects the expected gain in diversity when sampling a new individual. The function returns a matrix of sampling units 9 slope values. DATA SETS The BAT package includes five data sets. Three of them depict the abundance of 338 species of spiders (Araneae) in each of either 192 or 320 sampling units collected in three sites in Portugal (arrabida, geres and guadiana, see details in Cardoso 2009 and references therein). The data set phylotree represents an approximation to the phylogenetic tree for the 338 species. The tree is based on the Linnean hierarchy, with different suborders separated by 1 unit, families by 075, genera by 05and species by 025. Finally, the data set functree represents the functional tree for these same species. Several traits were recorded for each species: average size, type of web, type of hunting, stenophagy, vertical stratification in vegetation and circadian activity (Cardoso et al. 2011). The species 9 traits matrix was subjected to a hierarchical clustering procedure using UPGMA with Gower distances to produce a final dendrogram. EXAMPLES Here, we provide an example of a typical R session using the BAT package. All examples can be reproduced using the included data sets. We start by loading the package and all data. library(bat) data(arrabida) data(geres) data(guadiana) data(phylotree) data(functree) We create a new matrix with species abundance per site. comm <- rbind(colsums(arrabida), colsums(geres), colsums(guadiana)) sites <- c( Arrabida, Geres, Guadiana ) row.names(comm) <- sites and calculate alpha diversity (TD, PD and FD) for all sites. alpha(comm) alpha(comm, phylotree) alpha(comm, functree) Although Ger^es has more species and phylogenetic diversity, it is less diverse functionally than Arrabida. But are these results determined by the sampled abundance? We may rarefy to 1000 individuals and plot the results. raref.td <- alpha(comm, raref = 1000) raref.pd <- alpha(comm, phylotree, raref = 1000) raref.fd <- alpha(comm, functree, raref = 1000) par(mfrow=c(1,3)) boxplot(t(raref.td), names = sites, main= TD, pars = list boxplot(t(raref.pd), names = sites, main= PD, pars = list boxplot(t(raref.fd), names = sites, main= FD, pars = list Fig. 1. Results of the rarefaction procedure, for 1000 individuals and 100 runs, for alpha taxon (TD), phylogenetic (PD) and functional (FD) diversity performed with the function alpha of the R package BAT. Arrabida, Geres and Guadiana are three sites exhaustively sampled for spiders in Portugal and whose data are included in the package (see text for details).
4 BAT Biodiversity assessment tools 235 The three sites present similar FD values when data are rarified (Fig. 1). But results may be better understood if observed diversity is corrected through estimates (Chao1P is used throughout for illustration purposes only). alpha.estimate(comm) alpha.estimate(comm, phylotree) alpha.estimate(comm, functree) Estimates indicate that Guadiana may be more functionally diverse than the other sites. The reliability of any estimate can be judged based on the accumulation curves. acc.arrabida <- alpha.accum(arrabida) par(mfrow=c(1,2)) plot(acc.arrabida[,2], acc.arrabida[,17], xlab= Individuals, ylab = Chao 1P ) The Chao1P estimator values seem to have reached the asymptote (Fig. 2), which can be tested with plot(acc.arrabida[,2], slope(acc.arrabida)[,17], xlab= Individuals, ylab= Slope ) The slope in the last part of the curve is null or very close to it (Fig. 2), an indication of reliability of the estimate (although other indices should be taken into account). If we knew the true diversity of the site (e.g. species richness = 170), the accuracy of the different estimators could be tested as accuracy(acc.arrabida, target = 170) In this case, Chao2P has the smallest value (0017), being considered the most accurate. Beta diversity between each pair of sites can be calculated as: beta(comm) If func is not specified, the Jaccard dissimilarity is used by default. Total beta diversity is larger between Ger^es and Guadiana (b total = 0901), and although dissimilarity between sites is mostly explained by replacement of species (b repl > 0632 in all cases), differences in richness are larger between those two sites (b rich = 0268). Multiple phylogenetic beta with rarefactionmadetothesameabundanceasthesitewithleastindividuals sampled can be calculated as beta.multi(comm, phylotree, raref = 1) Again, b repl is clearly dominant overall (averages of b total = 0731, b repl = 0656, b rich = 0075). Finally, we may want to check the reliability of beta diversity values according to sampling intensity, taking abundances into account acc.beta <- beta.accum(arrabida, geres, abund = TRUE) par(mfrow=c(1,3)) plot(acc.beta[,2], xlab= Sampling units, ylab = expression (beta[total])) plot(acc.beta[,3], xlab= Sampling units, ylab = expression (beta[repl])) plot(acc.beta[,4], xlab= Sampling units, ylab = expression (beta[rich])) In all cases, the diversity values seem to stabilize and be reliable early in the accumulation process (Fig. 3). STRENGTHS The package BAT implements a number of methods that are not available in any other package or software. These include Fig. 2. Accumulation curve for the nonparametric estimator Chao1P provided by the function alpha.accum of the R package BAT and the slopes between every consecutive point along the curve, produced by the function slope. Fig. 3. Accumulation curve for the beta diversity values provided by the function beta.accum of the R package BAT.
5 236 P. Cardoso, F. Rigal & J. C. Carvalho the estimation of phylogenetic and functional diversity from incomplete sampling, the computation of accumulation curves for beta diversity and the partitioning of beta diversity, including with abundance, phylogenetic or functional data, in its replacement and richness difference components. The output of the functions can be easily used as response variables in statistical models using environmental, spatial or temporal data as explanatory variables. ALTERNATIVES Other software packages and R functions exist that calculate parts of the same or alternative methods to those provided by BAT. For species richness estimation, these include the EstimateS software (Colwell 2013), and the estimater and estaccumr functions of the R package vegan (Oksanen et al. 2013). However, none of these is able to deal with phylogenetic or functional data. For beta diversity decomposition, the function beta.div.comp found in Legendre (2014) provides similar results to ours for the coefficients computed by both packages. The package betapart (Baselga & Orme 2012) implements alternative methods, although we disagree with the methodological framework developed by these authors (see Carvalho, Cardoso & Gomes 2012; Carvalho et al. 2013; Cardoso et al. 2014c). CITATION Researchers using BAT in a published paper should cite this article and in addition can also cite the BAT package directly. Updatedcitationinformationcanbeobtainedbytyping:citation ( BAT ) Acknowledgements We thank all the co-authors of past papers that helped us developing the methods and data sets used here. Pierre Legendre and David Nipperess provided very useful comments on this manuscript. Data accessibility All data used in the manuscript are included in the R package: Funding F.R. was partly supported by FCT project PTDC/BIA-BIC/119255/2010. References Baselga, A. (2010) Partitioning the turnover and nestedness components of beta diversity. Global Ecology and Biogeography, 19, Baselga, A. (2012) The relationship between species replacement, dissimilarity derived from nestedness and nestedness. Global Ecology and Biogeography, 21, Baselga, A. & Orme, C.D.L. (2012) betapart: an R package for the study of beta diversity. Methods in Ecology and Evolution, 3, Cardoso, P. (2009) Standardization and optimization of arthropod inventories the case of Iberian spiders. Biodiversity and Conservation, 18, Cardoso, P., Borges, P.A.V. & Veech, J.A. (2009) Testing the performance of beta diversity measures based on incidence data: the robustness to undersampling. Diversity and Distributions, 15, Cardoso, P., Rigal, F. & Carvalho, J.C. (2014b) BAT: Biodiversity Assessment Tools. R package version URL [accessed 22 October 2014] Cardoso, P., Pekar, S., Jocque, R. & Coddington, J.A. (2011) Global patterns of guild composition and functional diversity of spiders. PLoS One, 6, e Cardoso, P., Rigal, F., Borges, P.A.V. & Carvalho, J.C. (2014a) A new frontier in biodiversity inventory: a proposal for estimators of phylogenetic and functional diversity. Methods in Ecology and Evolution, 5, Cardoso,P.,Rigal,F.,Carvalho,J.C.,Fortelius,M.,Borges,P.A.V.,Podani,J. & Schmera, D. (2014c) Partitioning taxon, phylogenetic and functional beta diversity into replacement and richness difference components. Journal of Biogeography, 41, Carvalho, J.C., Cardoso, P. & Gomes, P. (2012) Determining the relative roles of species replacement and species richness differences in generating beta-diversity patterns. Global Ecology and Biogeography, 21, Carvalho, J.C., Cardoso, P., Borges, P.A.V., Schmera, D. & Podani, J. (2013) Measuring fractions of beta diversity and their relationships to nestedness: a theoretical and empirical comparison of novel approaches. Oikos, 122, Chao, A. (1984) Nonparametric estimation of the number of classes in a population. Scandinavian Journal of Statistics, 11, Chao, A. (1987) Estimating the population size for capture-recapture data with unequal catchability. Biometrics, 43, Coddington, J.A., Agnarsson, I., Miller, J.A., Kuntner, M. & Hormiga, G. (2009) Undersampling bias: the null hypothesis for singleton species in tropical arthropod surveys. Journal of Animal Ecology, 78, Colwell, R.K. (2013) EstimateS: Statistical Estimation of Species Richness and Shared Species From Samples. Version 9.1. URL [accessed 01 October 2014] Colwell, R.K. & Coddington, J.A. (1994) Estimating terrestrial biodiversity through extrapolation. Philosophical Transactions of the Royal Society of London Biological Sciences, 345, Faith, D.P. (1992) Conservation evaluation and phylogenetic diversity. Biological Conservation, 61, Gotelli, N.J. & Colwell, R.K. (2001) Quantifying biodiversity: procedures and pitfalls in the measurement and comparison of species richness. Ecology Letters, 4, Heltshe, J. & Forrester, N.E. (1983) Estimating species richness using the jackknife procedure. Biometrics, 39, Legendre, P. (2014) Interpreting the replacement and richness difference components of beta diversity. Global Ecology and Biogeography, 23, Lennon, J.J., Koleff, P., Greenwood, J.J.D. & Gaston, K.J. (2001) The geographical structure of British bird distributions: diversity, spatial turnover and scale. Journal of Animal Ecology, 70, Oksanen, J., Blanchet, F.G., Kindt, R., Legendre, P., Minchin, P.R., O Hara, R.B. et al. (2013) Vegan: Community Ecology Package. R package version URL cran.r-project.org/package=vegan [accessed 01 October 2014]. Paradis, E., Claude, J. & Strimmer, K. (2004) APE: analyses of phylogenetics and evolution in R language. Bioinformatics, 20, Petchey, O.L. & Gaston, K.J. (2002) Functional diversity (FD), species richness and community composition. Ecology Letters, 5, Podani, J. & Schmera, D. (2011) A new conceptual and methodological framework for exploring and explaining pattern in presence-absence data. Oikos, 120, Soberon, M.J. & Llorente, J. (1993) The use of species accumulation functions for the prediction of species richness. Conservation Biology, 7, Stegen, J.C. & Hurlbert, A.H. (2011) Inferring eco-evolutionary processes from taxonomic, phylogenetic and functional trait b-diversity. PLoS One, 6, e Williams, P.H., de Klerk, H.M. & Crowe, T.M. (1999) Interpreting biogeographical boundaries among Afro-tropical birds: spatial patterns in richness gradients and species replacement. Journal of Biogeography, 26, Received 21 August 2014; accepted 11 November 2014 Handling Editor: Steven Kembel
Sampling e ects on beta diversity
 Introduction Methods Results Conclusions Sampling e ects on beta diversity Ben Bolker, Adrian Stier, Craig Osenberg McMaster University, Mathematics & Statistics and Biology UBC, Zoology University of
Introduction Methods Results Conclusions Sampling e ects on beta diversity Ben Bolker, Adrian Stier, Craig Osenberg McMaster University, Mathematics & Statistics and Biology UBC, Zoology University of
A new frontier in biodiversity inventory: a proposal for estimators of phylogenetic and functional diversity
 Methods in Ecology and Evolution 2014, 5, 452 461 doi: 10.1111/2041-210X.12173 A new frontier in biodiversity inventory: a proposal for estimators of phylogenetic and functional diversity Pedro Cardoso
Methods in Ecology and Evolution 2014, 5, 452 461 doi: 10.1111/2041-210X.12173 A new frontier in biodiversity inventory: a proposal for estimators of phylogenetic and functional diversity Pedro Cardoso
Package MBI. February 19, 2015
 Type Package Package MBI February 19, 2015 Title (M)ultiple-site (B)iodiversity (I)ndices Calculator Version 1.0 Date 2012-10-17 Author Youhua Chen Maintainer Over 20 multiple-site diversity indices can
Type Package Package MBI February 19, 2015 Title (M)ultiple-site (B)iodiversity (I)ndices Calculator Version 1.0 Date 2012-10-17 Author Youhua Chen Maintainer Over 20 multiple-site diversity indices can
Appendix S1 Replacement, richness difference and nestedness indices
 Appendix to: Legendre, P. (2014) Interpreting the replacement and richness difference components of beta diversity. Global Ecology and Biogeography, 23, 1324-1334. Appendix S1 Replacement, richness difference
Appendix to: Legendre, P. (2014) Interpreting the replacement and richness difference components of beta diversity. Global Ecology and Biogeography, 23, 1324-1334. Appendix S1 Replacement, richness difference
betapart: an R package for the study of beta diversity Andre s Baselga 1 * and C. David L. Orme 2
 Methods in Ecology and Evolution 2012, 3, 808 812 doi: 10.1111/j.2041-210X.2012.00224.x APPLICATION betapart: an R package for the study of beta diversity Andre s Baselga 1 * and C. David L. Orme 2 1 Departamento
Methods in Ecology and Evolution 2012, 3, 808 812 doi: 10.1111/j.2041-210X.2012.00224.x APPLICATION betapart: an R package for the study of beta diversity Andre s Baselga 1 * and C. David L. Orme 2 1 Departamento
Pedro Cardoso 1,2,3 *, Paulo A. V. Borges 1 and Joseph A. Veech 4. Main conclusions No index was found to perform without bias in all
 Diversity and Distributions, (Diversity Distrib.) (2009) 15, 1081 1090 A Journal of Conservation Biogeography BIODIVERSITY RESEARCH 1 CITA-A (Azorean Biodiversity Group), Dep. Ciências Agrárias, Universidade
Diversity and Distributions, (Diversity Distrib.) (2009) 15, 1081 1090 A Journal of Conservation Biogeography BIODIVERSITY RESEARCH 1 CITA-A (Azorean Biodiversity Group), Dep. Ciências Agrárias, Universidade
CHAO, JACKKNIFE AND BOOTSTRAP ESTIMATORS OF SPECIES RICHNESS
 IJAMAA, Vol. 12, No. 1, (January-June 2017), pp. 7-15 Serials Publications ISSN: 0973-3868 CHAO, JACKKNIFE AND BOOTSTRAP ESTIMATORS OF SPECIES RICHNESS CHAVAN KR. SARMAH ABSTRACT: The species richness
IJAMAA, Vol. 12, No. 1, (January-June 2017), pp. 7-15 Serials Publications ISSN: 0973-3868 CHAO, JACKKNIFE AND BOOTSTRAP ESTIMATORS OF SPECIES RICHNESS CHAVAN KR. SARMAH ABSTRACT: The species richness
Measuring fractions of beta diversity and their relationships to. nestedness: a theoretical and empirical comparison of novel
 1 Oikos 122: 825-834 (2013) 2 3 4 5 Measuring fractions of beta diversity and their relationships to nestedness: a theoretical and empirical comparison of novel approaches 6 7 8 José C. Carvalho 1,2,*,
1 Oikos 122: 825-834 (2013) 2 3 4 5 Measuring fractions of beta diversity and their relationships to nestedness: a theoretical and empirical comparison of novel approaches 6 7 8 José C. Carvalho 1,2,*,
An introduction to the picante package
 An introduction to the picante package Steven Kembel (steve.kembel@gmail.com) April 2010 Contents 1 Installing picante 1 2 Data formats in picante 2 2.1 Phylogenies................................ 2 2.2
An introduction to the picante package Steven Kembel (steve.kembel@gmail.com) April 2010 Contents 1 Installing picante 1 2 Data formats in picante 2 2.1 Phylogenies................................ 2 2.2
Ecological Archives E A2
 Ecological Archives E091-147-A2 Ilyas Siddique, Ima Célia Guimarães Vieira, Susanne Schmidt, David Lamb, Cláudio José Reis Carvalho, Ricardo de Oliveira Figueiredo, Simon Blomberg, Eric A. Davidson. Year.
Ecological Archives E091-147-A2 Ilyas Siddique, Ima Célia Guimarães Vieira, Susanne Schmidt, David Lamb, Cláudio José Reis Carvalho, Ricardo de Oliveira Figueiredo, Simon Blomberg, Eric A. Davidson. Year.
What is the range of a taxon? A scaling problem at three levels: Spa9al scale Phylogene9c depth Time
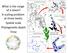 What is the range of a taxon? A scaling problem at three levels: Spa9al scale Phylogene9c depth Time 1 5 0.25 0.15 5 0.05 0.05 0.10 2 0.10 0.10 0.20 4 Reminder of what a range-weighted tree is Actual Tree
What is the range of a taxon? A scaling problem at three levels: Spa9al scale Phylogene9c depth Time 1 5 0.25 0.15 5 0.05 0.05 0.10 2 0.10 0.10 0.20 4 Reminder of what a range-weighted tree is Actual Tree
Comparing methods to separate components of beta diversity
 Methods in Ecology and Evolution 2015, 6, 1069 1079 doi: 10.1111/2041-210X.12388 Comparing methods to separate components of beta diversity Andres Baselga 1 * and Fabien Leprieur 2 1 Departamento de Zoologıa,
Methods in Ecology and Evolution 2015, 6, 1069 1079 doi: 10.1111/2041-210X.12388 Comparing methods to separate components of beta diversity Andres Baselga 1 * and Fabien Leprieur 2 1 Departamento de Zoologıa,
Other resources. Greengenes (bacterial) Silva (bacteria, archaeal and eukarya)
 General QIIME resources http://qiime.org/ Blog (news, updates): http://qiime.wordpress.com/ Support/forum: https://groups.google.com/forum/#!forum/qiimeforum Citing QIIME: Caporaso, J.G. et al., QIIME
General QIIME resources http://qiime.org/ Blog (news, updates): http://qiime.wordpress.com/ Support/forum: https://groups.google.com/forum/#!forum/qiimeforum Citing QIIME: Caporaso, J.G. et al., QIIME
Package SPECIES. R topics documented: April 23, Type Package. Title Statistical package for species richness estimation. Version 1.
 Package SPECIES April 23, 2011 Type Package Title Statistical package for species richness estimation Version 1.0 Date 2010-01-24 Author Ji-Ping Wang, Maintainer Ji-Ping Wang
Package SPECIES April 23, 2011 Type Package Title Statistical package for species richness estimation Version 1.0 Date 2010-01-24 Author Ji-Ping Wang, Maintainer Ji-Ping Wang
Manuscript: Changes in Latitudinal Diversity Gradient during the Great. Björn Kröger, Finnish Museum of Natural History, PO Box 44, Fi Helsinki,
 GSA Data Repository 2018029 1 Supplementary information 2 3 4 Manuscript: Changes in Latitudinal Diversity Gradient during the Great Ordovician Biodiversification Event 5 6 7 Björn Kröger, Finnish Museum
GSA Data Repository 2018029 1 Supplementary information 2 3 4 Manuscript: Changes in Latitudinal Diversity Gradient during the Great Ordovician Biodiversification Event 5 6 7 Björn Kröger, Finnish Museum
Supplementary Material
 Supplementary Material The impact of logging and forest conversion to oil palm on soil bacterial communities in Borneo Larisa Lee-Cruz 1, David P. Edwards 2,3, Binu Tripathi 1, Jonathan M. Adams 1* 1 Department
Supplementary Material The impact of logging and forest conversion to oil palm on soil bacterial communities in Borneo Larisa Lee-Cruz 1, David P. Edwards 2,3, Binu Tripathi 1, Jonathan M. Adams 1* 1 Department
"PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200 Spring 2014 University of California, Berkeley
 "PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200 Spring 2014 University of California, Berkeley D.D. Ackerly April 16, 2014. Community Ecology and Phylogenetics Readings: Cavender-Bares,
"PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200 Spring 2014 University of California, Berkeley D.D. Ackerly April 16, 2014. Community Ecology and Phylogenetics Readings: Cavender-Bares,
Deciphering the Enigma of Undetected Species, Phylogenetic, and Functional Diversity. Based on Good-Turing Theory
 Metadata S1 Deciphering the Enigma of Undetected Species, Phylogenetic, and Functional Diversity Based on Good-Turing Theory Anne Chao, Chun-Huo Chiu, Robert K. Colwell, Luiz Fernando S. Magnago, Robin
Metadata S1 Deciphering the Enigma of Undetected Species, Phylogenetic, and Functional Diversity Based on Good-Turing Theory Anne Chao, Chun-Huo Chiu, Robert K. Colwell, Luiz Fernando S. Magnago, Robin
Diversity partitioning without statistical independence of alpha and beta
 1964 Ecology, Vol. 91, No. 7 Ecology, 91(7), 2010, pp. 1964 1969 Ó 2010 by the Ecological Society of America Diversity partitioning without statistical independence of alpha and beta JOSEPH A. VEECH 1,3
1964 Ecology, Vol. 91, No. 7 Ecology, 91(7), 2010, pp. 1964 1969 Ó 2010 by the Ecological Society of America Diversity partitioning without statistical independence of alpha and beta JOSEPH A. VEECH 1,3
Remote sensing of biodiversity: measuring ecological complexity from space
 Remote sensing of biodiversity: measuring ecological complexity from space Duccio Rocchini Fondazione Edmund Mach Research and Innovation Centre Department of Biodiversity and Molecular Ecology San Michele
Remote sensing of biodiversity: measuring ecological complexity from space Duccio Rocchini Fondazione Edmund Mach Research and Innovation Centre Department of Biodiversity and Molecular Ecology San Michele
Towards a consensus for calculating dendrogram-based functional diversity indices
 Oikos 117: 794800, 2008 doi: 10.1111/j.2008.0030-1299.16594.x, # The Authors. Journal compilation # Oikos 2008 Subject Editor: Nicholas Gotelli, Accepted 1 February 2008 Towards a consensus for calculating
Oikos 117: 794800, 2008 doi: 10.1111/j.2008.0030-1299.16594.x, # The Authors. Journal compilation # Oikos 2008 Subject Editor: Nicholas Gotelli, Accepted 1 February 2008 Towards a consensus for calculating
A (short) introduction to phylogenetics
 A (short) introduction to phylogenetics Thibaut Jombart, Marie-Pauline Beugin MRC Centre for Outbreak Analysis and Modelling Imperial College London Genetic data analysis with PR Statistics, Millport Field
A (short) introduction to phylogenetics Thibaut Jombart, Marie-Pauline Beugin MRC Centre for Outbreak Analysis and Modelling Imperial College London Genetic data analysis with PR Statistics, Millport Field
Lecture 2: Descriptive statistics, normalizations & testing
 Lecture 2: Descriptive statistics, normalizations & testing From sequences to OTU table Sequencing Sample 1 Sample 2... Sample N Abundances of each microbial taxon in each of the N samples 2 1 Normalizing
Lecture 2: Descriptive statistics, normalizations & testing From sequences to OTU table Sequencing Sample 1 Sample 2... Sample N Abundances of each microbial taxon in each of the N samples 2 1 Normalizing
Studying the effect of species dominance on diversity patterns using Hill numbers-based indices
 Studying the effect of species dominance on diversity patterns using Hill numbers-based indices Loïc Chalmandrier Loïc Chalmandrier Diversity pattern analysis November 8th 2017 1 / 14 Introduction Diversity
Studying the effect of species dominance on diversity patterns using Hill numbers-based indices Loïc Chalmandrier Loïc Chalmandrier Diversity pattern analysis November 8th 2017 1 / 14 Introduction Diversity
Phylogenetic diversity and conservation
 Phylogenetic diversity and conservation Dan Faith The Australian Museum Applied ecology and human dimensions in biological conservation Biota Program/ FAPESP Nov. 9-10, 2009 BioGENESIS Providing an evolutionary
Phylogenetic diversity and conservation Dan Faith The Australian Museum Applied ecology and human dimensions in biological conservation Biota Program/ FAPESP Nov. 9-10, 2009 BioGENESIS Providing an evolutionary
Package hierdiversity
 Version 0.1 Date 2015-03-11 Package hierdiversity March 20, 2015 Title Hierarchical Multiplicative Partitioning of Complex Phenotypes Author Zachary Marion, James Fordyce, and Benjamin Fitzpatrick Maintainer
Version 0.1 Date 2015-03-11 Package hierdiversity March 20, 2015 Title Hierarchical Multiplicative Partitioning of Complex Phenotypes Author Zachary Marion, James Fordyce, and Benjamin Fitzpatrick Maintainer
THEORY. Based on sequence Length According to the length of sequence being compared it is of following two types
 Exp 11- THEORY Sequence Alignment is a process of aligning two sequences to achieve maximum levels of identity between them. This help to derive functional, structural and evolutionary relationships between
Exp 11- THEORY Sequence Alignment is a process of aligning two sequences to achieve maximum levels of identity between them. This help to derive functional, structural and evolutionary relationships between
How to quantify biological diversity: taxonomical, functional and evolutionary aspects. Hanna Tuomisto, University of Turku
 How to quantify biological diversity: taxonomical, functional and evolutionary aspects Hanna Tuomisto, University of Turku Why quantify biological diversity? understanding the structure and function of
How to quantify biological diversity: taxonomical, functional and evolutionary aspects Hanna Tuomisto, University of Turku Why quantify biological diversity? understanding the structure and function of
Supplement: Beta Diversity & The End Ordovician Extinctions. Appendix for: Response of beta diversity to pulses of Ordovician-Silurian extinction
 Appendix for: Response of beta diversity to pulses of Ordovician-Silurian extinction Collection- and formation-based sampling biases within the original dataset Both numbers of occurrences and numbers
Appendix for: Response of beta diversity to pulses of Ordovician-Silurian extinction Collection- and formation-based sampling biases within the original dataset Both numbers of occurrences and numbers
Stochastic and deterministic drivers of spatial and temporal turnover in breeding bird communities
 bs_bs_banner Global Ecology and Biogeography, (Global Ecol. Biogeogr.) (2013) 22, 202 212 RESEARCH PAPER Stochastic and deterministic drivers of spatial and temporal turnover in breeding bird communities
bs_bs_banner Global Ecology and Biogeography, (Global Ecol. Biogeogr.) (2013) 22, 202 212 RESEARCH PAPER Stochastic and deterministic drivers of spatial and temporal turnover in breeding bird communities
EFFECTS OF TAXONOMIC GROUPS AND GEOGRAPHIC SCALE ON PATTERNS OF NESTEDNESS
 EFFECTS OF TAXONOMIC GROUPS AND GEOGRAPHIC SCALE ON PATTERNS OF NESTEDNESS SFENTHOURAKIS Spyros, GIOKAS Sinos & LEGAKIS Anastasios Zoological Museum, Department of Biology, University of Athens, Greece
EFFECTS OF TAXONOMIC GROUPS AND GEOGRAPHIC SCALE ON PATTERNS OF NESTEDNESS SFENTHOURAKIS Spyros, GIOKAS Sinos & LEGAKIS Anastasios Zoological Museum, Department of Biology, University of Athens, Greece
Spatial non-stationarity, anisotropy and scale: The interactive visualisation of spatial turnover
 19th International Congress on Modelling and Simulation, Perth, Australia, 12 16 December 2011 http://mssanz.org.au/modsim2011 Spatial non-stationarity, anisotropy and scale: The interactive visualisation
19th International Congress on Modelling and Simulation, Perth, Australia, 12 16 December 2011 http://mssanz.org.au/modsim2011 Spatial non-stationarity, anisotropy and scale: The interactive visualisation
Lecture: Mixture Models for Microbiome data
 Lecture: Mixture Models for Microbiome data Lecture 3: Mixture Models for Microbiome data Outline: - - Sequencing thought experiment Mixture Models (tangent) - (esp. Negative Binomial) - Differential abundance
Lecture: Mixture Models for Microbiome data Lecture 3: Mixture Models for Microbiome data Outline: - - Sequencing thought experiment Mixture Models (tangent) - (esp. Negative Binomial) - Differential abundance
Outline Classes of diversity measures. Species Divergence and the Measurement of Microbial Diversity. How do we describe and compare diversity?
 Species Divergence and the Measurement of Microbial Diversity Cathy Lozupone University of Colorado, Boulder. Washington University, St Louis. Outline Classes of diversity measures α vs β diversity Quantitative
Species Divergence and the Measurement of Microbial Diversity Cathy Lozupone University of Colorado, Boulder. Washington University, St Louis. Outline Classes of diversity measures α vs β diversity Quantitative
Experimental Design and Data Analysis for Biologists
 Experimental Design and Data Analysis for Biologists Gerry P. Quinn Monash University Michael J. Keough University of Melbourne CAMBRIDGE UNIVERSITY PRESS Contents Preface page xv I I Introduction 1 1.1
Experimental Design and Data Analysis for Biologists Gerry P. Quinn Monash University Michael J. Keough University of Melbourne CAMBRIDGE UNIVERSITY PRESS Contents Preface page xv I I Introduction 1 1.1
Package breakaway. R topics documented: March 30, 2016
 Title Species Richness Estimation and Modeling Version 3.0 Date 2016-03-29 Author and John Bunge Maintainer Package breakaway March 30, 2016 Species richness estimation is an important
Title Species Richness Estimation and Modeling Version 3.0 Date 2016-03-29 Author and John Bunge Maintainer Package breakaway March 30, 2016 Species richness estimation is an important
G.S. Gilbert, ENVS291 Transition to R vf13 Class 10 Vegan and Picante. Class 10b - Tools for Phylogenetic Ecology
 Class 10b - Tools for Phylogenetic Ecology This is a very brief introduction to some of the R tools available for phylogenetic community ecology and trait analysis. There are lots of levels to this, with
Class 10b - Tools for Phylogenetic Ecology This is a very brief introduction to some of the R tools available for phylogenetic community ecology and trait analysis. There are lots of levels to this, with
ENMTools: a toolbox for comparative studies of environmental niche models
 Ecography 33: 607611, 2010 doi: 10.1111/j.1600-0587.2009.06142.x # 2010 The Authors. Journal compilation # 2010 Ecography Subject Editor: Thiago Rangel. Accepted 4 September 2009 ENMTools: a toolbox for
Ecography 33: 607611, 2010 doi: 10.1111/j.1600-0587.2009.06142.x # 2010 The Authors. Journal compilation # 2010 Ecography Subject Editor: Thiago Rangel. Accepted 4 September 2009 ENMTools: a toolbox for
Characterizing and predicting cyanobacterial blooms in an 8-year
 1 2 3 4 5 Characterizing and predicting cyanobacterial blooms in an 8-year amplicon sequencing time-course Authors Nicolas Tromas 1*, Nathalie Fortin 2, Larbi Bedrani 1, Yves Terrat 1, Pedro Cardoso 4,
1 2 3 4 5 Characterizing and predicting cyanobacterial blooms in an 8-year amplicon sequencing time-course Authors Nicolas Tromas 1*, Nathalie Fortin 2, Larbi Bedrani 1, Yves Terrat 1, Pedro Cardoso 4,
Analyzing patterns of species diversity as departures from random expectations
 OIKOS 8: 49/55, 25 Analyzing patterns of species diversity as departures from random expectations Joseph A. Veech Veech, J. A. 25. Analyzing patterns of species diversity as departures from random expectations.
OIKOS 8: 49/55, 25 Analyzing patterns of species diversity as departures from random expectations Joseph A. Veech Veech, J. A. 25. Analyzing patterns of species diversity as departures from random expectations.
Thirty years of progeny from Chao s inequality: Estimating and comparing richness with incidence data and incomplete sampling
 Statistics & Operations Research ransactions SOR 41 (1) January-June 2017, 3-54 ISSN: 1696-2281 eissn: 2013-8830 www.idescat.cat/sort/ Statistics & Operations Research Institut d Estadística de Catalunya
Statistics & Operations Research ransactions SOR 41 (1) January-June 2017, 3-54 ISSN: 1696-2281 eissn: 2013-8830 www.idescat.cat/sort/ Statistics & Operations Research Institut d Estadística de Catalunya
Assessing Congruence Among Ultrametric Distance Matrices
 Journal of Classification 26:103-117 (2009) DOI: 10.1007/s00357-009-9028-x Assessing Congruence Among Ultrametric Distance Matrices Véronique Campbell Université de Montréal, Canada Pierre Legendre Université
Journal of Classification 26:103-117 (2009) DOI: 10.1007/s00357-009-9028-x Assessing Congruence Among Ultrametric Distance Matrices Véronique Campbell Université de Montréal, Canada Pierre Legendre Université
Contrasting beta diversity among regions: how do classical and multivariate approaches compare?
 Global Ecologyand and Biogeography, (Global (Global Ecol. Biogeogr.) Ecol. Biogeogr.) (2015) () 25, 368 377 MACROECOLOGICAL METHODS Contrasting beta diversity among regions: how do classical and multivariate
Global Ecologyand and Biogeography, (Global (Global Ecol. Biogeogr.) Ecol. Biogeogr.) (2015) () 25, 368 377 MACROECOLOGICAL METHODS Contrasting beta diversity among regions: how do classical and multivariate
ecological area-network relations: methodology Christopher Moore cs765: complex networks 16 November 2011
 ecological area-network relations: methodology Christopher Moore cs765: complex networks 16 November 2011 ecology: the study of the spatial and temporal patterns of the distribution and abundance of organisms,
ecological area-network relations: methodology Christopher Moore cs765: complex networks 16 November 2011 ecology: the study of the spatial and temporal patterns of the distribution and abundance of organisms,
Dissimilarity and transformations. Pierre Legendre Département de sciences biologiques Université de Montréal
 and transformations Pierre Legendre Département de sciences biologiques Université de Montréal http://www.numericalecology.com/ Pierre Legendre 2017 Definitions An association coefficient is a function
and transformations Pierre Legendre Département de sciences biologiques Université de Montréal http://www.numericalecology.com/ Pierre Legendre 2017 Definitions An association coefficient is a function
Linking species-compositional dissimilarities and environmental data for biodiversity assessment
 Linking species-compositional dissimilarities and environmental data for biodiversity assessment D. P. Faith, S. Ferrier Australian Museum, 6 College St., Sydney, N.S.W. 2010, Australia; N.S.W. National
Linking species-compositional dissimilarities and environmental data for biodiversity assessment D. P. Faith, S. Ferrier Australian Museum, 6 College St., Sydney, N.S.W. 2010, Australia; N.S.W. National
APPENDIX E: Estimating diversity profile based on the proposed RAD estimator (for abundance data).
 Anne Chao, T. C. Hsieh, Robin L. Chazdon, Robert K. Colwell, and Nicholas J. Gotelli. 2015. Unveiling the species-rank abundance distribution by generalizing the Good-Turing sample coverage theory. Ecology
Anne Chao, T. C. Hsieh, Robin L. Chazdon, Robert K. Colwell, and Nicholas J. Gotelli. 2015. Unveiling the species-rank abundance distribution by generalizing the Good-Turing sample coverage theory. Ecology
BIO 111: Biological Diversity and Evolution
 BIO 111: Biological Diversity and Evolution Varsha 2017 Ullasa Kodandaramaiah & Hema Somanathan School of Biology MODULE: BIODIVERSITY AND CONSERVATION BIOLOGY Part I - FUNDAMENTAL CONCEPTS OF BIODIVERSITY
BIO 111: Biological Diversity and Evolution Varsha 2017 Ullasa Kodandaramaiah & Hema Somanathan School of Biology MODULE: BIODIVERSITY AND CONSERVATION BIOLOGY Part I - FUNDAMENTAL CONCEPTS OF BIODIVERSITY
No citation for this material is currently available (but will be once Jarrett Byrnes contributes the lavspatialcorrect package to the semtools
 No citation for this material is currently available (but will be once Jarrett Byrnes contributes the lavspatialcorrect package to the semtools package). In the meantime, references that present the underlying
No citation for this material is currently available (but will be once Jarrett Byrnes contributes the lavspatialcorrect package to the semtools package). In the meantime, references that present the underlying
AUTOMATED DISCOVERY OF RELATIONSHIPS, MODELS AND PRINCIPLES IN ECOLOGY
 AUTOMATED DISCOVERY OF RELATIONSHIPS, MODELS AND PRINCIPLES IN ECOLOGY Pedro Cardoso 1,2,*, Paulo A.V. Borges 2, José Carlos Carvalho 2,3, François Rigal 2, Rosalina Gabriel 2, José Cascalho 4,5, Luís
AUTOMATED DISCOVERY OF RELATIONSHIPS, MODELS AND PRINCIPLES IN ECOLOGY Pedro Cardoso 1,2,*, Paulo A.V. Borges 2, José Carlos Carvalho 2,3, François Rigal 2, Rosalina Gabriel 2, José Cascalho 4,5, Luís
Distance-Based Functional Diversity Measures and Their Decomposition: A Framework Based on Hill Numbers
 and Their Decomposition: A Framework Based on Hill Numbers Chun-Huo Chiu, Anne Chao* Institute of Statistics, National Tsing Hua University, Hsin-Chu, Taiwan Abstract Hill numbers (or the effective number
and Their Decomposition: A Framework Based on Hill Numbers Chun-Huo Chiu, Anne Chao* Institute of Statistics, National Tsing Hua University, Hsin-Chu, Taiwan Abstract Hill numbers (or the effective number
VarCan (version 1): Variation Estimation and Partitioning in Canonical Analysis
 VarCan (version 1): Variation Estimation and Partitioning in Canonical Analysis Pedro R. Peres-Neto March 2005 Department of Biology University of Regina Regina, SK S4S 0A2, Canada E-mail: Pedro.Peres-Neto@uregina.ca
VarCan (version 1): Variation Estimation and Partitioning in Canonical Analysis Pedro R. Peres-Neto March 2005 Department of Biology University of Regina Regina, SK S4S 0A2, Canada E-mail: Pedro.Peres-Neto@uregina.ca
Dr. Amira A. AL-Hosary
 Phylogenetic analysis Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic Basics: Biological
Phylogenetic analysis Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic Basics: Biological
Multi-scale sampling boosts inferences from beta diversity patterns in coastal forests, South Africa. Pieter I. Olivier & Rudi J.
 Article accepted for publication in the Journal of Biogeography, 41, 1428-1439. doi:10.1111/jbi.12303 Multi-scale sampling boosts inferences from beta diversity patterns in coastal forests, South Africa
Article accepted for publication in the Journal of Biogeography, 41, 1428-1439. doi:10.1111/jbi.12303 Multi-scale sampling boosts inferences from beta diversity patterns in coastal forests, South Africa
2/7/2018. Strata. Strata
 The strata option allows you to control how permutations are done. Specifically, to constrain permutations. Why would you want to do this? In this dataset, there are clear differences in area (A vs. B),
The strata option allows you to control how permutations are done. Specifically, to constrain permutations. Why would you want to do this? In this dataset, there are clear differences in area (A vs. B),
entropart: An R Package to Measure and Partition Diversity
 entropart: An R Package to Measure and Partition Diversity Eric Marcon AgroParisTech UMR EcoFoG Bruno Hérault Cirad UMR EcoFoG Abstract entropart is a package for R designed to estimate diversity based
entropart: An R Package to Measure and Partition Diversity Eric Marcon AgroParisTech UMR EcoFoG Bruno Hérault Cirad UMR EcoFoG Abstract entropart is a package for R designed to estimate diversity based
Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut
 Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic analysis Phylogenetic Basics: Biological
Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic analysis Phylogenetic Basics: Biological
TOTAL INERTIA DECOMPOSITION IN TABLES WITH A FACTORIAL STRUCTURE
 Statistica Applicata - Italian Journal of Applied Statistics Vol. 29 (2-3) 233 doi.org/10.26398/ijas.0029-012 TOTAL INERTIA DECOMPOSITION IN TABLES WITH A FACTORIAL STRUCTURE Michael Greenacre 1 Universitat
Statistica Applicata - Italian Journal of Applied Statistics Vol. 29 (2-3) 233 doi.org/10.26398/ijas.0029-012 TOTAL INERTIA DECOMPOSITION IN TABLES WITH A FACTORIAL STRUCTURE Michael Greenacre 1 Universitat
The biogenesis-atbc2012 Training workshop "Evolutionary Approaches to Biodiversity Science" June 2012, Bonito, Brazil
 The biogenesis-atbc2012 Training workshop "Evolutionary Approaches to Biodiversity Science" 16-18 June 2012, Bonito, Brazil Phylogenetic and functional diversity (including PD) and phylogenetic conservation
The biogenesis-atbc2012 Training workshop "Evolutionary Approaches to Biodiversity Science" 16-18 June 2012, Bonito, Brazil Phylogenetic and functional diversity (including PD) and phylogenetic conservation
DETECTING BIOLOGICAL AND ENVIRONMENTAL CHANGES: DESIGN AND ANALYSIS OF MONITORING AND EXPERIMENTS (University of Bologna, 3-14 March 2008)
 Dipartimento di Biologia Evoluzionistica Sperimentale Centro Interdipartimentale di Ricerca per le Scienze Ambientali in Ravenna INTERNATIONAL WINTER SCHOOL UNIVERSITY OF BOLOGNA DETECTING BIOLOGICAL AND
Dipartimento di Biologia Evoluzionistica Sperimentale Centro Interdipartimentale di Ricerca per le Scienze Ambientali in Ravenna INTERNATIONAL WINTER SCHOOL UNIVERSITY OF BOLOGNA DETECTING BIOLOGICAL AND
Using rarefaction to isolate the effects of patch size and sampling effort on beta diversity
 Using rarefaction to isolate the effects of patch size and sampling effort on beta diversity ADRIAN C. STIER, 1, BENJAMIN M. BOLKER, 2 AND CRAIG W. OSENBERG 3 1 Department of Ecology, Evolution and Marine
Using rarefaction to isolate the effects of patch size and sampling effort on beta diversity ADRIAN C. STIER, 1, BENJAMIN M. BOLKER, 2 AND CRAIG W. OSENBERG 3 1 Department of Ecology, Evolution and Marine
Compositional similarity and β (beta) diversity
 CHAPTER 6 Compositional similarity and β (beta) diversity Lou Jost, Anne Chao, and Robin L. Chazdon 6.1 Introduction Spatial variation in species composition is one of the most fundamental and conspicuous
CHAPTER 6 Compositional similarity and β (beta) diversity Lou Jost, Anne Chao, and Robin L. Chazdon 6.1 Introduction Spatial variation in species composition is one of the most fundamental and conspicuous
Island biogeography. Key concepts. Introduction. Island biogeography theory. Colonization-extinction balance. Island-biogeography theory
 Island biogeography Key concepts Colonization-extinction balance Island-biogeography theory Introduction At the end of the last chapter, it was suggested that another mechanism for the maintenance of α-diversity
Island biogeography Key concepts Colonization-extinction balance Island-biogeography theory Introduction At the end of the last chapter, it was suggested that another mechanism for the maintenance of α-diversity
Inferring Ecological Processes from Taxonomic, Phylogenetic and Functional Trait b-diversity
 Inferring Ecological Processes from Taxonomic, Phylogenetic and Functional Trait b-diversity James C. Stegen*, Allen H. Hurlbert Department of Biology, University of North Carolina, Chapel Hill, North
Inferring Ecological Processes from Taxonomic, Phylogenetic and Functional Trait b-diversity James C. Stegen*, Allen H. Hurlbert Department of Biology, University of North Carolina, Chapel Hill, North
Measuring beta diversity for presence absence data
 Ecology 003 7, Measuring beta diversity for presence absence data Blackwell Publishing Ltd. PATRICIA KOLEFF*, KEVIN J. GASTON* and JACK J. LENNON *Biodiversity and Macroecology Group, Department of Animal
Ecology 003 7, Measuring beta diversity for presence absence data Blackwell Publishing Ltd. PATRICIA KOLEFF*, KEVIN J. GASTON* and JACK J. LENNON *Biodiversity and Macroecology Group, Department of Animal
Lecture 3: Mixture Models for Microbiome data. Lecture 3: Mixture Models for Microbiome data
 Lecture 3: Mixture Models for Microbiome data 1 Lecture 3: Mixture Models for Microbiome data Outline: - Mixture Models (Negative Binomial) - DESeq2 / Don t Rarefy. Ever. 2 Hypothesis Tests - reminder
Lecture 3: Mixture Models for Microbiome data 1 Lecture 3: Mixture Models for Microbiome data Outline: - Mixture Models (Negative Binomial) - DESeq2 / Don t Rarefy. Ever. 2 Hypothesis Tests - reminder
Journal of Statistical Software
 JSS Journal of Statistical Software April 2011, Volume 40, Issue 9. http://www.jstatsoft.org/ SPECIES: An R Package for Species Richness Estimation Ji-Ping Wang Northwestern University Abstract We introduce
JSS Journal of Statistical Software April 2011, Volume 40, Issue 9. http://www.jstatsoft.org/ SPECIES: An R Package for Species Richness Estimation Ji-Ping Wang Northwestern University Abstract We introduce
Transitivity a FORTRAN program for the analysis of bivariate competitive interactions Version 1.1
 Transitivity 1 Transitivity a FORTRAN program for the analysis of bivariate competitive interactions Version 1.1 Werner Ulrich Nicolaus Copernicus University in Toruń Chair of Ecology and Biogeography
Transitivity 1 Transitivity a FORTRAN program for the analysis of bivariate competitive interactions Version 1.1 Werner Ulrich Nicolaus Copernicus University in Toruń Chair of Ecology and Biogeography
Development Team. Department of Zoology, University of Delhi. Department of Zoology, University of Delhi
 Paper No. : 12 Module : 18 diversity index, abundance, species richness, vertical and horizontal Development Team Principal Investigator: Co-Principal Investigator: Paper Coordinator: Content Writer: Content
Paper No. : 12 Module : 18 diversity index, abundance, species richness, vertical and horizontal Development Team Principal Investigator: Co-Principal Investigator: Paper Coordinator: Content Writer: Content
Vegan: ecological diversity
 Vegan: ecological diversity Jari Oksanen processed with vegan 2.4-5 in R Under development (unstable) (2017-12-01 r73812) on December 1, 2017 Abstract This document explains diversity related methods in
Vegan: ecological diversity Jari Oksanen processed with vegan 2.4-5 in R Under development (unstable) (2017-12-01 r73812) on December 1, 2017 Abstract This document explains diversity related methods in
Sampling bias is a challenge for quantifying specialization and network structure: lessons from a quantitative niche model
 Oikos : 1 12, 215 doi: 1.1111/oik.2256 215 The Authors. Oikos Nordic Society Oikos Subject Editor: Stefano Allesina. Editor-in-Chief: Dries Bonte. Accepted 7 May 215 Sampling bias is a challenge for quantifying
Oikos : 1 12, 215 doi: 1.1111/oik.2256 215 The Authors. Oikos Nordic Society Oikos Subject Editor: Stefano Allesina. Editor-in-Chief: Dries Bonte. Accepted 7 May 215 Sampling bias is a challenge for quantifying
The tangled link between b- and c-diversity: a Narcissus effect weakens statistical inferences in null model analyses of diversity patterns
 Global Ecology and Biogeography, (Global Ecol. Biogeogr.) (2017) 26, 1 5 ECOLOGICAL SOUNDINGS The tangled link between b- and c-diversity: a Narcissus effect weakens statistical inferences in null model
Global Ecology and Biogeography, (Global Ecol. Biogeogr.) (2017) 26, 1 5 ECOLOGICAL SOUNDINGS The tangled link between b- and c-diversity: a Narcissus effect weakens statistical inferences in null model
ESTIMATION OF CONSERVATISM OF CHARACTERS BY CONSTANCY WITHIN BIOLOGICAL POPULATIONS
 ESTIMATION OF CONSERVATISM OF CHARACTERS BY CONSTANCY WITHIN BIOLOGICAL POPULATIONS JAMES S. FARRIS Museum of Zoology, The University of Michigan, Ann Arbor Accepted March 30, 1966 The concept of conservatism
ESTIMATION OF CONSERVATISM OF CHARACTERS BY CONSTANCY WITHIN BIOLOGICAL POPULATIONS JAMES S. FARRIS Museum of Zoology, The University of Michigan, Ann Arbor Accepted March 30, 1966 The concept of conservatism
Selective extinction drives taxonomic and functional alpha and beta diversities in island bird assemblages
 Journal of Animal Ecology 2016, 85, 409 418 doi: 10.1111/1365-2656.12478 Selective extinction drives taxonomic and functional alpha and beta diversities in island bird assemblages Xingfeng Si 1, Andres
Journal of Animal Ecology 2016, 85, 409 418 doi: 10.1111/1365-2656.12478 Selective extinction drives taxonomic and functional alpha and beta diversities in island bird assemblages Xingfeng Si 1, Andres
Algorithms in Bioinformatics
 Algorithms in Bioinformatics Sami Khuri Department of Computer Science San José State University San José, California, USA khuri@cs.sjsu.edu www.cs.sjsu.edu/faculty/khuri Distance Methods Character Methods
Algorithms in Bioinformatics Sami Khuri Department of Computer Science San José State University San José, California, USA khuri@cs.sjsu.edu www.cs.sjsu.edu/faculty/khuri Distance Methods Character Methods
Supplementary Information
 Supplementary Information For the article"comparable system-level organization of Archaea and ukaryotes" by J. Podani, Z. N. Oltvai, H. Jeong, B. Tombor, A.-L. Barabási, and. Szathmáry (reference numbers
Supplementary Information For the article"comparable system-level organization of Archaea and ukaryotes" by J. Podani, Z. N. Oltvai, H. Jeong, B. Tombor, A.-L. Barabási, and. Szathmáry (reference numbers
"PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200 Spring 2018 University of California, Berkeley
 "PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200 Spring 2018 University of California, Berkeley D.D. Ackerly Feb. 26, 2018 Maximum Likelihood Principles, and Applications to
"PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200 Spring 2018 University of California, Berkeley D.D. Ackerly Feb. 26, 2018 Maximum Likelihood Principles, and Applications to
Biodiversity, temperature, and energy
 Biodiversity, temperature, and energy David Storch Center for Theoretical Study & Department of Ecology, Faculty of Science Charles University, Czech Republic Global diversity gradients energy-related
Biodiversity, temperature, and energy David Storch Center for Theoretical Study & Department of Ecology, Faculty of Science Charles University, Czech Republic Global diversity gradients energy-related
A diversity of beta diversities: straightening up a concept gone awry. Part 2. Quantifying beta diversity and related phenomena
 Ecography 33: 2345, 21 doi: 1.1111/j.16-587.29.6148.x # 21 The Author. Journal compilation # 21 Ecography Subject Editor: Robert K. Colwell. Accepted 18 November 29 A diversity of beta diversities: straightening
Ecography 33: 2345, 21 doi: 1.1111/j.16-587.29.6148.x # 21 The Author. Journal compilation # 21 Ecography Subject Editor: Robert K. Colwell. Accepted 18 November 29 A diversity of beta diversities: straightening
Vegan: ecological diversity
 Vegan: ecological diversity Jari Oksanen Abstract This document explains diversity related methods in vegan. The methods are briefly described, and the equations used them are given often in more detail
Vegan: ecological diversity Jari Oksanen Abstract This document explains diversity related methods in vegan. The methods are briefly described, and the equations used them are given often in more detail
Censusing the Sea in the 21 st Century
 Censusing the Sea in the 21 st Century Nancy Knowlton & Matthieu Leray Photo: Ove Hoegh-Guldberg Smithsonian s National Museum of Natural History Estimates of Marine/Reef Species Numbers (Millions) Marine
Censusing the Sea in the 21 st Century Nancy Knowlton & Matthieu Leray Photo: Ove Hoegh-Guldberg Smithsonian s National Museum of Natural History Estimates of Marine/Reef Species Numbers (Millions) Marine
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/322/5899/258/dc1 Supporting Online Material for Global Warming, Elevational Range Shifts, and Lowland Biotic Attrition in the Wet Tropics Robert K. Colwell,* Gunnar
www.sciencemag.org/cgi/content/full/322/5899/258/dc1 Supporting Online Material for Global Warming, Elevational Range Shifts, and Lowland Biotic Attrition in the Wet Tropics Robert K. Colwell,* Gunnar
Lecture 2: Diversity, Distances, adonis. Lecture 2: Diversity, Distances, adonis. Alpha- Diversity. Alpha diversity definition(s)
 Lecture 2: Diversity, Distances, adonis Lecture 2: Diversity, Distances, adonis Diversity - alpha, beta (, gamma) Beta- Diversity in practice: Ecological Distances Unsupervised Learning: Clustering, etc
Lecture 2: Diversity, Distances, adonis Lecture 2: Diversity, Distances, adonis Diversity - alpha, beta (, gamma) Beta- Diversity in practice: Ecological Distances Unsupervised Learning: Clustering, etc
Phylogenies & Classifying species (AKA Cladistics & Taxonomy) What are phylogenies & cladograms? How do we read them? How do we estimate them?
 Phylogenies & Classifying species (AKA Cladistics & Taxonomy) What are phylogenies & cladograms? How do we read them? How do we estimate them? Carolus Linneaus:Systema Naturae (1735) Swedish botanist &
Phylogenies & Classifying species (AKA Cladistics & Taxonomy) What are phylogenies & cladograms? How do we read them? How do we estimate them? Carolus Linneaus:Systema Naturae (1735) Swedish botanist &
Beta diversity and latitude in North American mammals: testing the hypothesis of covariation
 ECOGRAPHY 27: 547/556, 2004 Beta diversity and latitude in North American mammals: testing the hypothesis of covariation Pilar Rodríguez and Héctor T. Arita Rodríguez, P. and Arita, H. T. 2004. Beta diversity
ECOGRAPHY 27: 547/556, 2004 Beta diversity and latitude in North American mammals: testing the hypothesis of covariation Pilar Rodríguez and Héctor T. Arita Rodríguez, P. and Arita, H. T. 2004. Beta diversity
Metacommunities Spatial Ecology of Communities
 Spatial Ecology of Communities Four perspectives for multiple species Patch dynamics principles of metapopulation models (patchy pops, Levins) Mass effects principles of source-sink and rescue effects
Spatial Ecology of Communities Four perspectives for multiple species Patch dynamics principles of metapopulation models (patchy pops, Levins) Mass effects principles of source-sink and rescue effects
Appendix from L. J. Revell, On the Analysis of Evolutionary Change along Single Branches in a Phylogeny
 008 by The University of Chicago. All rights reserved.doi: 10.1086/588078 Appendix from L. J. Revell, On the Analysis of Evolutionary Change along Single Branches in a Phylogeny (Am. Nat., vol. 17, no.
008 by The University of Chicago. All rights reserved.doi: 10.1086/588078 Appendix from L. J. Revell, On the Analysis of Evolutionary Change along Single Branches in a Phylogeny (Am. Nat., vol. 17, no.
BIO 682 Multivariate Statistics Spring 2008
 BIO 682 Multivariate Statistics Spring 2008 Steve Shuster http://www4.nau.edu/shustercourses/bio682/index.htm Lecture 11 Properties of Community Data Gauch 1982, Causton 1988, Jongman 1995 a. Qualitative:
BIO 682 Multivariate Statistics Spring 2008 Steve Shuster http://www4.nau.edu/shustercourses/bio682/index.htm Lecture 11 Properties of Community Data Gauch 1982, Causton 1988, Jongman 1995 a. Qualitative:
Range-weighted metrics of species and phylogenetic turnover can better resolve biogeographic transition zones
 Methods in Ecology and Evolution 2016, 7, 580 588 doi: 10.1111/2041-210X.12513 Range-weighted metrics of species and phylogenetic turnover can better resolve biogeographic transition zones Shawn W. Laffan
Methods in Ecology and Evolution 2016, 7, 580 588 doi: 10.1111/2041-210X.12513 Range-weighted metrics of species and phylogenetic turnover can better resolve biogeographic transition zones Shawn W. Laffan
Consistency Index (CI)
 Consistency Index (CI) minimum number of changes divided by the number required on the tree. CI=1 if there is no homoplasy negatively correlated with the number of species sampled Retention Index (RI)
Consistency Index (CI) minimum number of changes divided by the number required on the tree. CI=1 if there is no homoplasy negatively correlated with the number of species sampled Retention Index (RI)
Biology 559R: Introduction to Phylogenetic Comparative Methods Topics for this week:
 Biology 559R: Introduction to Phylogenetic Comparative Methods Topics for this week: Course general information About the course Course objectives Comparative methods: An overview R as language: uses and
Biology 559R: Introduction to Phylogenetic Comparative Methods Topics for this week: Course general information About the course Course objectives Comparative methods: An overview R as language: uses and
Package velociraptr. February 15, 2017
 Type Package Title Fossil Analysis Version 1.0 Author Package velociraptr February 15, 2017 Maintainer Andrew A Zaffos Functions for downloading, reshaping, culling, cleaning, and analyzing
Type Package Title Fossil Analysis Version 1.0 Author Package velociraptr February 15, 2017 Maintainer Andrew A Zaffos Functions for downloading, reshaping, culling, cleaning, and analyzing
Bird Species richness per 110x110 km grid square (so, strictly speaking, alpha diversity) -most species live there!
 We "know" there are more species in the tropics Why are the Tropics so biodiverse? And the tropics are special: 1. Oldest known ecological pattern (Humboldt, 1807) 2. Well-known by Darwin and Wallace 3.
We "know" there are more species in the tropics Why are the Tropics so biodiverse? And the tropics are special: 1. Oldest known ecological pattern (Humboldt, 1807) 2. Well-known by Darwin and Wallace 3.
Phylogenetic Diversity and distribution patterns of the Compositae family in the high Andes of South America
 Phylogenetic Diversity and distribution patterns of the Compositae family in the high Andes of South America Scherson, R.A., Naulin,P.I., Albornoz, A., Hagemann, T., Vidal, P.M., Riveros, N., and Arroyo,
Phylogenetic Diversity and distribution patterns of the Compositae family in the high Andes of South America Scherson, R.A., Naulin,P.I., Albornoz, A., Hagemann, T., Vidal, P.M., Riveros, N., and Arroyo,
Webinar Session 1. Introduction to Modern Methods for Analyzing Capture- Recapture Data: Closed Populations 1
 Webinar Session 1. Introduction to Modern Methods for Analyzing Capture- Recapture Data: Closed Populations 1 b y Bryan F.J. Manly Western Ecosystems Technology Inc. Cheyenne, Wyoming bmanly@west-inc.com
Webinar Session 1. Introduction to Modern Methods for Analyzing Capture- Recapture Data: Closed Populations 1 b y Bryan F.J. Manly Western Ecosystems Technology Inc. Cheyenne, Wyoming bmanly@west-inc.com
Vellend, Mark. Do commonly used indices of β-diversity measure species turnover?
 Journal of Vegetation Science 12: 545-552, 2001 IAVS; Opulus Press Uppsala. Printed in Sweden - Do commonly used indices of β-diversity measure species turnover? - 545 Do commonly used indices of β-diversity
Journal of Vegetation Science 12: 545-552, 2001 IAVS; Opulus Press Uppsala. Printed in Sweden - Do commonly used indices of β-diversity measure species turnover? - 545 Do commonly used indices of β-diversity
Discrimination Among Groups. Discrimination Among Groups
 Discrimination Among Groups Id Species Canopy Snag Canopy Cover Density Height 1 A 80 1.2 35 2 A 75 0.5 32 3 A 72 2.8 28..... 31 B 35 3.3 15 32 B 75 4.1 25 60 B 15 5.0 3..... 61 C 5 2.1 5 62 C 8 3.4 2
Discrimination Among Groups Id Species Canopy Snag Canopy Cover Density Height 1 A 80 1.2 35 2 A 75 0.5 32 3 A 72 2.8 28..... 31 B 35 3.3 15 32 B 75 4.1 25 60 B 15 5.0 3..... 61 C 5 2.1 5 62 C 8 3.4 2
Multivariate Analysis of Ecological Data using CANOCO
 Multivariate Analysis of Ecological Data using CANOCO JAN LEPS University of South Bohemia, and Czech Academy of Sciences, Czech Republic Universitats- uric! Lanttesbibiiothek Darmstadt Bibliothek Biologie
Multivariate Analysis of Ecological Data using CANOCO JAN LEPS University of South Bohemia, and Czech Academy of Sciences, Czech Republic Universitats- uric! Lanttesbibiiothek Darmstadt Bibliothek Biologie
Decomposing Phylodiversity
 Decomposing Phylodiversity Eric Marcon, Bruno Hérault To cite this version: Eric Marcon, Bruno Hérault. Decomposing Phylodiversity. 2014. HAL Id: hal-00946177 https://hal-agroparistech.archives-ouvertes.fr/hal-00946177
Decomposing Phylodiversity Eric Marcon, Bruno Hérault To cite this version: Eric Marcon, Bruno Hérault. Decomposing Phylodiversity. 2014. HAL Id: hal-00946177 https://hal-agroparistech.archives-ouvertes.fr/hal-00946177
Package PVR. February 15, 2013
 Package PVR February 15, 2013 Type Package Title Computes phylogenetic eigenvectors regression (PVR) and phylogenetic signal-representation curve (PSR) (with null and Brownian expectations) Version 0.2.1
Package PVR February 15, 2013 Type Package Title Computes phylogenetic eigenvectors regression (PVR) and phylogenetic signal-representation curve (PSR) (with null and Brownian expectations) Version 0.2.1
