Supplementary Information
|
|
|
- Octavia Heath
- 5 years ago
- Views:
Transcription
1 Supplementary Information For the article"comparable system-level organization of Archaea and ukaryotes" by J. Podani, Z. N. Oltvai, H. Jeong, B. Tombor, A.-L. Barabási, and. Szathmáry (reference numbers are the same as in the paper) 1
2 Web Note A Connectivity distribution of information transfer networks To determine the large-scale structural organization of information transfer networks, we have examined the topologic properties of the information transfer pathways of 43 different organisms based on "Information transfer" portions of data deposited in the WIT database. Similar to that of metabolic networks 11, we have first established a graph theoretic representation of the biochemical reactions taking place in a given information transfer network. In this representation, a network is built up of nodes, which are the substrates that are connected to one another through links, which are the actual biochemical reactions. Note, that beside core metabolic reactions, the WIT DB includes biochemical reactions contributing to (1) information pathways, (2) electron transport, (3) transmembrane transport, (4) signal transduction, and (5) structure and function of the cell. However, with the exception of information transfer pathways, the data deposited for these parts are so incomplete that comparison of e.g., signal transduction pathways is completely impossible. In addition, no information for any metazoan, including for those of the just recently sequenced human genome, is available in the WIT DB. To establish if the topology of information transfer networks is best described by the inherently random and uniform exponential- or the highly heterogeneous scale-free model 11 the connectivity distribution of all 43 networks were analyzed as described for metabolic networks in ref. 11. As illustrated in Web Fig. A, our results convincingly indicate that the probability that a given substrate participates in k reactions follows a power-law distribution, i.e., information transfer networks belong to the class of scale-free networks. Since under physiological conditions a large number of biochemical reactions (links) in a network are preferentially catalyzed in one direction (i.e., the links are directed), for each node we distinguish between incoming and outgoing links. For instance, in scherichia coli the probability that a substrate participates as an educt in k information transfer pathways follows P(k) ~ k -γin, with γ in = 2.1, and the probability that a given substrate is produced by k different information transfer reactions follows a similar distribution, with γ out = 2.2 (Web Fig. Ab). We find that scale-free networks describe the information transfer reaction networks in all organisms in all three domains of life (Web Fig. Aa-c), indicating the generic nature of this structural organization (Web Fig. Ad). 2
3 0 In Out 0 In Out -2-2 P(k) P(k) (a) k -2 In Out P(k) P(k) (b) k In Out -4-6 (c) k -4-6 (d) k Fig. A Connectivity distribution P(k) for the substrates in (a) A. fulgidus (Archaea) (b). coli (Bacteria) (c) C. elegans (ukarya), shown on a log-log plot, counting separately the incoming (IN) and outgoing links (OUT) for each substrate, k in (k out ) corresponding to the number of reactions in which a substrate participates as a product (educt). (d) The connectivity distribution averaged over all 43 organisms. 3
4 Web Note B Database analysis Data types and resemblance coefficients Although the data matrices comprise quantitative information, we decided to rely only upon the qualitative part of the data, for several reasons. First of all, traditionally, qualitative data have been almost exclusively used in molecular systematics. Second, qualitative data are less sensitive to the different intensity by which the organisms are studied, meaning that there is more noise in quantitative data attributable to 'sampling error'. The presence or absence (P/A) of a variable in a given organism is the simplest type of all data. We thus compared each pair of organisms using a simple ratio in which the number of variables present in both was divided by the number of variables present in at least one of them (widely known as the Jaccard index). The coefficient is expressed here as its complement to measure the dissimilarity of the two species being compared. In addition to the P/A scores, we also utilized the ordinal information (i.e., the rank order) in the data. We have previously demonstrated 11 that ordering relationships for each organism are also of great importance when describing metabolic pathways. Consequently, we also compared each pair of species using the ordinal measure suggested by Goodman and Kruskal (their γ function). In this coefficient, the number of variable pairs ordered similarly by both taxa is divided by the total number of pairs of variables that are ordered at all. For taxa j and k it is written as γ jk = (a - b) / (a + b) where a is the number of pairs of variables ordered for objects j and k identically, b is the number of pairs of variables that are reversely ordered in j and k. The coefficient ranges from -1 to 1, indicating complete disagreement and full agreement, respectively. To simplify calculations, we transformed the values to measure dissimilarity in the range of 0 to 2. For details of the methodology used in this paper see refs Ordination A major group of exploratory data analysis tools aims to reduce the dimensionality of the system, more precisely, to show a multidimensional picture in a few, preferably two, dimensions as efficiently as possible. These procedures are commonly referred to as ordination. For the present study, we have chosen the non-metric multidimensional scaling (NMDS) method which is best suited to input dissimilarities that are ordinal anyway. For 4
5 consistency, we applied the very same procedure to the otherwise metric Jaccard coefficient, as well. A fundamental feature of NMDS is that the user defines the final dimensionality of the result, which is generally selected to be two. The procedure then attempts to arrange the points in the space such that the ordering relationships of input dissimilarities are as faithfully preserved in the ordination space as possible. The procedure is iterative, and therefore requires several runs from which we retain the best result. Hierarchical clustering The most widely known hierarchical clustering method is group average clustering (UPGMA, ref. 13). Despite strong criticisms expressed by cladists, the method applies even to phylogenetic problems especially if the molecular clock (i.e., constant rate of evolutionary change on all lineages) is assumed. Nevertheless, UPGMA has been most commonly used on data other than DNA or RNA sequences, thus its properties and interpretability of its results are sufficiently known. In the present study, UPGMA was applied to matrices of the Jaccard index. The method produces an ultrametric tree, the hierarchical levels potentially interpretable as mean dissimilarities between groups fused at that level. Since the matrix calculated by Goodman-Kruskal's γ comprises non-metric information, they were processed by a different hierarchical classification algorithm, namely ordinal hierarchical clustering (OC, see ref. 14 for its underlying criterion). As OC is a method recently developed by one of us (JP); here we provide a more detailed description of this technique. This procedure considers only the ordering information present in the dissimilarity matrix, and then builds a tree in which each level is labeled only by the number of the iteration step in which the actual merger took place. The algorithm of agglomerative ordinal clustering first orders the m(m-1)/2 dissimilarity coefficients (m is the number of taxa), so that each γ jk is replaced by its rank, r jk. Then, the following clustering criterion is considered C = (R w - R min ) / (R max - R min ) where R w is the sum of ranks of within-cluster dissimilarities, R min is the possible minimum of such ranks for the given number of clusters and for the given numbers of objects in each cluster, and R max is the possible maximum. A similar criterion using R exp instead of R max was proposed by ref. 14 for determining the importance of variables in classification. The value of C ranges from 0 to 1, 0 indicating that all within-cluster 5
6 dissimilarities are smaller than the between-cluster dissimilarities, whereas 1 indicating that all between-cluster dissimilarities are smaller than the others. In the dendrogram resulted from agglomerative ordinal clustering we used the ranks of fusions (values from 1 to m-1), rather than the C values themselves, because this criterion does not change monotonically. That is, the result is an ordered dendrogram, rather than a weighted dendrogram, being consistent with the ordinality of the previous steps of the analysis. Neighbor joining (NJ) The method originally developed by Saitou and Nei (ref. 12) has been the most widely applied distance-based tree-building method in phylogenetic reconstruction. Contrary to UPGMA, the tree is not ultrametric, but the additivity of branch lengths holds. In other words, the sum of branch lengths along the path between two taxa in the tree is an approximation to the input distances. The NJ tree is unrooted by definition, although some external criterion (outgroup, longest path) may be used to position a root (a practice not followed here). 6
7 Web Note C Analysis of the effect of database errors Of the 43 organisms whose metabolic- and information transfer pathways we have analyzed the genome of 25 has been completely sequenced (5 Archaea, 18 Bacteria, 2 ukaryotes), while the remaining 18 are only partially sequenced. Therefore, two major sources of possible errors in the database could potentially affect our analysis: (a) the erroneous annotation of enzymes and consequently, biochemical reactions; for the organisms with completely sequenced genomes this is the likely source of error. (b) reactions and pathways missing from the database; for organisms with incompletely sequenced genomes both (a) and (b) are of potential source of error. To determine if these limitations affect our analysis we have performed repeated analyses according to the type of errors to demonstrate if database bias critically influences the validity of our findings. These analyses were restricted to the UPGMA classifications of the P/A case, assuming that ordinations and unrooted trees are similarly influenced by data changes. In the first series of simulations, the four UPGMA classifications (Fig. 1f,h, Fig. 2f,h) were recomputed based on a reduced set of input data such that only the randomly chosen 50% of original variables were retained. The INFO based dendrograms (compare original Fig. 1f,h to the corresponding ones in Web Fig. B) suffered little changes owing to data reduction, the only discrepancy being that Archaea and ukaryotes form three, rather than two small clusters, while Bacteria remain intact as a very tight group. For MTAB data, removal of 50% of the variables from the substrate data set caused only one critical modification of the tree (compare original Fig. 2f to the corresponding one in Web Fig. B); Aeropyrum pernix was moved to the B2 group which was otherwise split into two. On the other hand, the main groupings in the dendrogram of Fig. 2h (NZ data) did not change at all. In the second series of simulations, the incompletely sequenced organisms were simply removed from the analyses, so that clustering was confined to 25 taxa in case of all the four input data sets. Reduction of sample size did not alter the main groupings at all (compare original Fig. 1f,h and 2f,h to the corresponding trees in Web Fig. C), suggesting robustness of the data upon the removal of incompletely sequenced organisms. 7
8 1f 1h A+ B A+ B 2f 2h A B1 B2 A B1 B2 Fig. B UPGMA classifications based on randomly reduced data sets to evaluate potential effects of missing reactions and pathways. Captions refer to comparable dendrograms obtained from complete data. 8
9 1f 1h A B A+ 2f 2h B A B1 B2 A B1 B2 Fig. C UPGMA classifications restricted to completely sequenced species to evaluate the effect of missing information arising from incomplete sequencing. Captions refer to comparable dendrograms obtained for the full set of species. 9
Phylogenetics: Building Phylogenetic Trees
 1 Phylogenetics: Building Phylogenetic Trees COMP 571 Luay Nakhleh, Rice University 2 Four Questions Need to be Answered What data should we use? Which method should we use? Which evolutionary model should
1 Phylogenetics: Building Phylogenetic Trees COMP 571 Luay Nakhleh, Rice University 2 Four Questions Need to be Answered What data should we use? Which method should we use? Which evolutionary model should
Bioinformatics 2. Yeast two hybrid. Proteomics. Proteomics
 GENOME Bioinformatics 2 Proteomics protein-gene PROTEOME protein-protein METABOLISM Slide from http://www.nd.edu/~networks/ Citrate Cycle Bio-chemical reactions What is it? Proteomics Reveal protein Protein
GENOME Bioinformatics 2 Proteomics protein-gene PROTEOME protein-protein METABOLISM Slide from http://www.nd.edu/~networks/ Citrate Cycle Bio-chemical reactions What is it? Proteomics Reveal protein Protein
Phylogenetics: Building Phylogenetic Trees. COMP Fall 2010 Luay Nakhleh, Rice University
 Phylogenetics: Building Phylogenetic Trees COMP 571 - Fall 2010 Luay Nakhleh, Rice University Four Questions Need to be Answered What data should we use? Which method should we use? Which evolutionary
Phylogenetics: Building Phylogenetic Trees COMP 571 - Fall 2010 Luay Nakhleh, Rice University Four Questions Need to be Answered What data should we use? Which method should we use? Which evolutionary
Department of Pathology, Northwestern University Medical School, Chicago, IL 60611
 The large-scale organization of metabolic networks H. Jeong 1, B. Tombor 2, R. Albert 1, Z. N. Oltvai 2 and A.-L. Barabási 1 1 Department of Physics, University of Notre Dame, Notre Dame, IN 46556, and
The large-scale organization of metabolic networks H. Jeong 1, B. Tombor 2, R. Albert 1, Z. N. Oltvai 2 and A.-L. Barabási 1 1 Department of Physics, University of Notre Dame, Notre Dame, IN 46556, and
SUPPLEMENTARY INFORMATION
 Supplementary information S3 (box) Methods Methods Genome weighting The currently available collection of archaeal and bacterial genomes has a highly biased distribution of isolates across taxa. For example,
Supplementary information S3 (box) Methods Methods Genome weighting The currently available collection of archaeal and bacterial genomes has a highly biased distribution of isolates across taxa. For example,
Proteomics. Yeast two hybrid. Proteomics - PAGE techniques. Data obtained. What is it?
 Proteomics What is it? Reveal protein interactions Protein profiling in a sample Yeast two hybrid screening High throughput 2D PAGE Automatic analysis of 2D Page Yeast two hybrid Use two mating strains
Proteomics What is it? Reveal protein interactions Protein profiling in a sample Yeast two hybrid screening High throughput 2D PAGE Automatic analysis of 2D Page Yeast two hybrid Use two mating strains
C3020 Molecular Evolution. Exercises #3: Phylogenetics
 C3020 Molecular Evolution Exercises #3: Phylogenetics Consider the following sequences for five taxa 1-5 and the known outgroup O, which has the ancestral states (note that sequence 3 has changed from
C3020 Molecular Evolution Exercises #3: Phylogenetics Consider the following sequences for five taxa 1-5 and the known outgroup O, which has the ancestral states (note that sequence 3 has changed from
Biological Networks: Comparison, Conservation, and Evolution via Relative Description Length By: Tamir Tuller & Benny Chor
 Biological Networks:,, and via Relative Description Length By: Tamir Tuller & Benny Chor Presented by: Noga Grebla Content of the presentation Presenting the goals of the research Reviewing basic terms
Biological Networks:,, and via Relative Description Length By: Tamir Tuller & Benny Chor Presented by: Noga Grebla Content of the presentation Presenting the goals of the research Reviewing basic terms
Algorithms in Bioinformatics
 Algorithms in Bioinformatics Sami Khuri Department of Computer Science San José State University San José, California, USA khuri@cs.sjsu.edu www.cs.sjsu.edu/faculty/khuri Distance Methods Character Methods
Algorithms in Bioinformatics Sami Khuri Department of Computer Science San José State University San José, California, USA khuri@cs.sjsu.edu www.cs.sjsu.edu/faculty/khuri Distance Methods Character Methods
Phylogenetic Tree Reconstruction
 I519 Introduction to Bioinformatics, 2011 Phylogenetic Tree Reconstruction Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Evolution theory Speciation Evolution of new organisms is driven
I519 Introduction to Bioinformatics, 2011 Phylogenetic Tree Reconstruction Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Evolution theory Speciation Evolution of new organisms is driven
Theory of Evolution Charles Darwin
 Theory of Evolution Charles arwin 858-59: Origin of Species 5 year voyage of H.M.S. eagle (83-36) Populations have variations. Natural Selection & Survival of the fittest: nature selects best adapted varieties
Theory of Evolution Charles arwin 858-59: Origin of Species 5 year voyage of H.M.S. eagle (83-36) Populations have variations. Natural Selection & Survival of the fittest: nature selects best adapted varieties
Dr. Amira A. AL-Hosary
 Phylogenetic analysis Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic Basics: Biological
Phylogenetic analysis Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic Basics: Biological
Cell biology traditionally identifies proteins based on their individual actions as catalysts, signaling
 Lethality and centrality in protein networks Cell biology traditionally identifies proteins based on their individual actions as catalysts, signaling molecules, or building blocks of cells and microorganisms.
Lethality and centrality in protein networks Cell biology traditionally identifies proteins based on their individual actions as catalysts, signaling molecules, or building blocks of cells and microorganisms.
Types of biological networks. I. Intra-cellurar networks
 Types of biological networks I. Intra-cellurar networks 1 Some intra-cellular networks: 1. Metabolic networks 2. Transcriptional regulation networks 3. Cell signalling networks 4. Protein-protein interaction
Types of biological networks I. Intra-cellurar networks 1 Some intra-cellular networks: 1. Metabolic networks 2. Transcriptional regulation networks 3. Cell signalling networks 4. Protein-protein interaction
BINF6201/8201. Molecular phylogenetic methods
 BINF60/80 Molecular phylogenetic methods 0-7-06 Phylogenetics Ø According to the evolutionary theory, all life forms on this planet are related to one another by descent. Ø Traditionally, phylogenetics
BINF60/80 Molecular phylogenetic methods 0-7-06 Phylogenetics Ø According to the evolutionary theory, all life forms on this planet are related to one another by descent. Ø Traditionally, phylogenetics
POPULATION GENETICS Winter 2005 Lecture 17 Molecular phylogenetics
 POPULATION GENETICS Winter 2005 Lecture 17 Molecular phylogenetics - in deriving a phylogeny our goal is simply to reconstruct the historical relationships between a group of taxa. - before we review the
POPULATION GENETICS Winter 2005 Lecture 17 Molecular phylogenetics - in deriving a phylogeny our goal is simply to reconstruct the historical relationships between a group of taxa. - before we review the
Chapter 26 Phylogeny and the Tree of Life
 Chapter 26 Phylogeny and the Tree of Life Chapter focus Shifting from the process of how evolution works to the pattern evolution produces over time. Phylogeny Phylon = tribe, geny = genesis or origin
Chapter 26 Phylogeny and the Tree of Life Chapter focus Shifting from the process of how evolution works to the pattern evolution produces over time. Phylogeny Phylon = tribe, geny = genesis or origin
A Phylogenetic Network Construction due to Constrained Recombination
 A Phylogenetic Network Construction due to Constrained Recombination Mohd. Abdul Hai Zahid Research Scholar Research Supervisors: Dr. R.C. Joshi Dr. Ankush Mittal Department of Electronics and Computer
A Phylogenetic Network Construction due to Constrained Recombination Mohd. Abdul Hai Zahid Research Scholar Research Supervisors: Dr. R.C. Joshi Dr. Ankush Mittal Department of Electronics and Computer
Multiple Sequence Alignment. Sequences
 Multiple Sequence Alignment Sequences > YOR020c mstllksaksivplmdrvlvqrikaqaktasglylpe knveklnqaevvavgpgftdangnkvvpqvkvgdqvl ipqfggstiklgnddevilfrdaeilakiakd > crassa mattvrsvksliplldrvlvqrvkaeaktasgiflpe
Multiple Sequence Alignment Sequences > YOR020c mstllksaksivplmdrvlvqrikaqaktasglylpe knveklnqaevvavgpgftdangnkvvpqvkvgdqvl ipqfggstiklgnddevilfrdaeilakiakd > crassa mattvrsvksliplldrvlvqrvkaeaktasgiflpe
Constructing Evolutionary/Phylogenetic Trees
 Constructing Evolutionary/Phylogenetic Trees 2 broad categories: istance-based methods Ultrametric Additive: UPGMA Transformed istance Neighbor-Joining Character-based Maximum Parsimony Maximum Likelihood
Constructing Evolutionary/Phylogenetic Trees 2 broad categories: istance-based methods Ultrametric Additive: UPGMA Transformed istance Neighbor-Joining Character-based Maximum Parsimony Maximum Likelihood
CS5238 Combinatorial methods in bioinformatics 2003/2004 Semester 1. Lecture 8: Phylogenetic Tree Reconstruction: Distance Based - October 10, 2003
 CS5238 Combinatorial methods in bioinformatics 2003/2004 Semester 1 Lecture 8: Phylogenetic Tree Reconstruction: Distance Based - October 10, 2003 Lecturer: Wing-Kin Sung Scribe: Ning K., Shan T., Xiang
CS5238 Combinatorial methods in bioinformatics 2003/2004 Semester 1 Lecture 8: Phylogenetic Tree Reconstruction: Distance Based - October 10, 2003 Lecturer: Wing-Kin Sung Scribe: Ning K., Shan T., Xiang
BioControl - Week 6, Lecture 1
 BioControl - Week 6, Lecture 1 Goals of this lecture Large metabolic networks organization Design principles for small genetic modules - Rules based on gene demand - Rules based on error minimization Suggested
BioControl - Week 6, Lecture 1 Goals of this lecture Large metabolic networks organization Design principles for small genetic modules - Rules based on gene demand - Rules based on error minimization Suggested
EVOLUTIONARY DISTANCES
 EVOLUTIONARY DISTANCES FROM STRINGS TO TREES Luca Bortolussi 1 1 Dipartimento di Matematica ed Informatica Università degli studi di Trieste luca@dmi.units.it Trieste, 14 th November 2007 OUTLINE 1 STRINGS:
EVOLUTIONARY DISTANCES FROM STRINGS TO TREES Luca Bortolussi 1 1 Dipartimento di Matematica ed Informatica Università degli studi di Trieste luca@dmi.units.it Trieste, 14 th November 2007 OUTLINE 1 STRINGS:
A (short) introduction to phylogenetics
 A (short) introduction to phylogenetics Thibaut Jombart, Marie-Pauline Beugin MRC Centre for Outbreak Analysis and Modelling Imperial College London Genetic data analysis with PR Statistics, Millport Field
A (short) introduction to phylogenetics Thibaut Jombart, Marie-Pauline Beugin MRC Centre for Outbreak Analysis and Modelling Imperial College London Genetic data analysis with PR Statistics, Millport Field
Computational approaches for functional genomics
 Computational approaches for functional genomics Kalin Vetsigian October 31, 2001 The rapidly increasing number of completely sequenced genomes have stimulated the development of new methods for finding
Computational approaches for functional genomics Kalin Vetsigian October 31, 2001 The rapidly increasing number of completely sequenced genomes have stimulated the development of new methods for finding
Phylogenetic trees 07/10/13
 Phylogenetic trees 07/10/13 A tree is the only figure to occur in On the Origin of Species by Charles Darwin. It is a graphical representation of the evolutionary relationships among entities that share
Phylogenetic trees 07/10/13 A tree is the only figure to occur in On the Origin of Species by Charles Darwin. It is a graphical representation of the evolutionary relationships among entities that share
Phylogenetic inference
 Phylogenetic inference Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, March 7 th 016 After this lecture, you can discuss (dis-) advantages of different information types
Phylogenetic inference Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, March 7 th 016 After this lecture, you can discuss (dis-) advantages of different information types
Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut
 Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic analysis Phylogenetic Basics: Biological
Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic analysis Phylogenetic Basics: Biological
V14 Graph connectivity Metabolic networks
 V14 Graph connectivity Metabolic networks In the first half of this lecture section, we use the theory of network flows to give constructive proofs of Menger s theorem. These proofs lead directly to algorithms
V14 Graph connectivity Metabolic networks In the first half of this lecture section, we use the theory of network flows to give constructive proofs of Menger s theorem. These proofs lead directly to algorithms
THEORY. Based on sequence Length According to the length of sequence being compared it is of following two types
 Exp 11- THEORY Sequence Alignment is a process of aligning two sequences to achieve maximum levels of identity between them. This help to derive functional, structural and evolutionary relationships between
Exp 11- THEORY Sequence Alignment is a process of aligning two sequences to achieve maximum levels of identity between them. This help to derive functional, structural and evolutionary relationships between
9/30/11. Evolution theory. Phylogenetic Tree Reconstruction. Phylogenetic trees (binary trees) Phylogeny (phylogenetic tree)
 I9 Introduction to Bioinformatics, 0 Phylogenetic ree Reconstruction Yuzhen Ye (yye@indiana.edu) School of Informatics & omputing, IUB Evolution theory Speciation Evolution of new organisms is driven by
I9 Introduction to Bioinformatics, 0 Phylogenetic ree Reconstruction Yuzhen Ye (yye@indiana.edu) School of Informatics & omputing, IUB Evolution theory Speciation Evolution of new organisms is driven by
A bioinformatics approach to the structural and functional analysis of the glycogen phosphorylase protein family
 A bioinformatics approach to the structural and functional analysis of the glycogen phosphorylase protein family Jieming Shen 1,2 and Hugh B. Nicholas, Jr. 3 1 Bioengineering and Bioinformatics Summer
A bioinformatics approach to the structural and functional analysis of the glycogen phosphorylase protein family Jieming Shen 1,2 and Hugh B. Nicholas, Jr. 3 1 Bioengineering and Bioinformatics Summer
NJMerge: A generic technique for scaling phylogeny estimation methods and its application to species trees
 NJMerge: A generic technique for scaling phylogeny estimation methods and its application to species trees Erin Molloy and Tandy Warnow {emolloy2, warnow}@illinois.edu University of Illinois at Urbana
NJMerge: A generic technique for scaling phylogeny estimation methods and its application to species trees Erin Molloy and Tandy Warnow {emolloy2, warnow}@illinois.edu University of Illinois at Urbana
The architecture of complexity: the structure and dynamics of complex networks.
 SMR.1656-36 School and Workshop on Structure and Function of Complex Networks 16-28 May 2005 ------------------------------------------------------------------------------------------------------------------------
SMR.1656-36 School and Workshop on Structure and Function of Complex Networks 16-28 May 2005 ------------------------------------------------------------------------------------------------------------------------
Theory of Evolution. Charles Darwin
 Theory of Evolution harles arwin 858-59: Origin of Species 5 year voyage of H.M.S. eagle (8-6) Populations have variations. Natural Selection & Survival of the fittest: nature selects best adapted varieties
Theory of Evolution harles arwin 858-59: Origin of Species 5 year voyage of H.M.S. eagle (8-6) Populations have variations. Natural Selection & Survival of the fittest: nature selects best adapted varieties
Chapter 19 Organizing Information About Species: Taxonomy and Cladistics
 Chapter 19 Organizing Information About Species: Taxonomy and Cladistics An unexpected family tree. What are the evolutionary relationships among a human, a mushroom, and a tulip? Molecular systematics
Chapter 19 Organizing Information About Species: Taxonomy and Cladistics An unexpected family tree. What are the evolutionary relationships among a human, a mushroom, and a tulip? Molecular systematics
Network Biology: Understanding the cell s functional organization. Albert-László Barabási Zoltán N. Oltvai
 Network Biology: Understanding the cell s functional organization Albert-László Barabási Zoltán N. Oltvai Outline: Evolutionary origin of scale-free networks Motifs, modules and hierarchical networks Network
Network Biology: Understanding the cell s functional organization Albert-László Barabási Zoltán N. Oltvai Outline: Evolutionary origin of scale-free networks Motifs, modules and hierarchical networks Network
 Tree of Life iological Sequence nalysis Chapter http://tolweb.org/tree/ Phylogenetic Prediction ll organisms on Earth have a common ancestor. ll species are related. The relationship is called a phylogeny
Tree of Life iological Sequence nalysis Chapter http://tolweb.org/tree/ Phylogenetic Prediction ll organisms on Earth have a common ancestor. ll species are related. The relationship is called a phylogeny
Evolutionary trees. Describe the relationship between objects, e.g. species or genes
 Evolutionary trees Bonobo Chimpanzee Human Neanderthal Gorilla Orangutan Describe the relationship between objects, e.g. species or genes Early evolutionary studies The evolutionary relationships between
Evolutionary trees Bonobo Chimpanzee Human Neanderthal Gorilla Orangutan Describe the relationship between objects, e.g. species or genes Early evolutionary studies The evolutionary relationships between
Dynamic optimisation identifies optimal programs for pathway regulation in prokaryotes. - Supplementary Information -
 Dynamic optimisation identifies optimal programs for pathway regulation in prokaryotes - Supplementary Information - Martin Bartl a, Martin Kötzing a,b, Stefan Schuster c, Pu Li a, Christoph Kaleta b a
Dynamic optimisation identifies optimal programs for pathway regulation in prokaryotes - Supplementary Information - Martin Bartl a, Martin Kötzing a,b, Stefan Schuster c, Pu Li a, Christoph Kaleta b a
Microbial Taxonomy. Microbes usually have few distinguishing properties that relate them, so a hierarchical taxonomy mainly has not been possible.
 Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
"Nothing in biology makes sense except in the light of evolution Theodosius Dobzhansky
 MOLECULAR PHYLOGENY "Nothing in biology makes sense except in the light of evolution Theodosius Dobzhansky EVOLUTION - theory that groups of organisms change over time so that descendeants differ structurally
MOLECULAR PHYLOGENY "Nothing in biology makes sense except in the light of evolution Theodosius Dobzhansky EVOLUTION - theory that groups of organisms change over time so that descendeants differ structurally
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/1/8/e1500527/dc1 Supplementary Materials for A phylogenomic data-driven exploration of viral origins and evolution The PDF file includes: Arshan Nasir and Gustavo
advances.sciencemag.org/cgi/content/full/1/8/e1500527/dc1 Supplementary Materials for A phylogenomic data-driven exploration of viral origins and evolution The PDF file includes: Arshan Nasir and Gustavo
Phylogeny Tree Algorithms
 Phylogeny Tree lgorithms Jianlin heng, PhD School of Electrical Engineering and omputer Science University of entral Florida 2006 Free for academic use. opyright @ Jianlin heng & original sources for some
Phylogeny Tree lgorithms Jianlin heng, PhD School of Electrical Engineering and omputer Science University of entral Florida 2006 Free for academic use. opyright @ Jianlin heng & original sources for some
Evolutionary trees. Describe the relationship between objects, e.g. species or genes
 Evolutionary trees Bonobo Chimpanzee Human Neanderthal Gorilla Orangutan Describe the relationship between objects, e.g. species or genes Early evolutionary studies Anatomical features were the dominant
Evolutionary trees Bonobo Chimpanzee Human Neanderthal Gorilla Orangutan Describe the relationship between objects, e.g. species or genes Early evolutionary studies Anatomical features were the dominant
CS224W: Social and Information Network Analysis
 CS224W: Social and Information Network Analysis Reaction Paper Adithya Rao, Gautam Kumar Parai, Sandeep Sripada Keywords: Self-similar networks, fractality, scale invariance, modularity, Kronecker graphs.
CS224W: Social and Information Network Analysis Reaction Paper Adithya Rao, Gautam Kumar Parai, Sandeep Sripada Keywords: Self-similar networks, fractality, scale invariance, modularity, Kronecker graphs.
Phylogenetics: Distance Methods. COMP Spring 2015 Luay Nakhleh, Rice University
 Phylogenetics: Distance Methods COMP 571 - Spring 2015 Luay Nakhleh, Rice University Outline Evolutionary models and distance corrections Distance-based methods Evolutionary Models and Distance Correction
Phylogenetics: Distance Methods COMP 571 - Spring 2015 Luay Nakhleh, Rice University Outline Evolutionary models and distance corrections Distance-based methods Evolutionary Models and Distance Correction
Phylogenetic Trees. What They Are Why We Do It & How To Do It. Presented by Amy Harris Dr Brad Morantz
 Phylogenetic Trees What They Are Why We Do It & How To Do It Presented by Amy Harris Dr Brad Morantz Overview What is a phylogenetic tree Why do we do it How do we do it Methods and programs Parallels
Phylogenetic Trees What They Are Why We Do It & How To Do It Presented by Amy Harris Dr Brad Morantz Overview What is a phylogenetic tree Why do we do it How do we do it Methods and programs Parallels
Microbial Diversity and Assessment (II) Spring, 2007 Guangyi Wang, Ph.D. POST103B
 Microbial Diversity and Assessment (II) Spring, 007 Guangyi Wang, Ph.D. POST03B guangyi@hawaii.edu http://www.soest.hawaii.edu/marinefungi/ocn403webpage.htm General introduction and overview Taxonomy [Greek
Microbial Diversity and Assessment (II) Spring, 007 Guangyi Wang, Ph.D. POST03B guangyi@hawaii.edu http://www.soest.hawaii.edu/marinefungi/ocn403webpage.htm General introduction and overview Taxonomy [Greek
Phylogenetic inference: from sequences to trees
 W ESTFÄLISCHE W ESTFÄLISCHE W ILHELMS -U NIVERSITÄT NIVERSITÄT WILHELMS-U ÜNSTER MM ÜNSTER VOLUTIONARY FUNCTIONAL UNCTIONAL GENOMICS ENOMICS EVOLUTIONARY Bioinformatics 1 Phylogenetic inference: from sequences
W ESTFÄLISCHE W ESTFÄLISCHE W ILHELMS -U NIVERSITÄT NIVERSITÄT WILHELMS-U ÜNSTER MM ÜNSTER VOLUTIONARY FUNCTIONAL UNCTIONAL GENOMICS ENOMICS EVOLUTIONARY Bioinformatics 1 Phylogenetic inference: from sequences
Classification and Phylogeny
 Classification and Phylogeny The diversity of life is great. To communicate about it, there must be a scheme for organization. There are many species that would be difficult to organize without a scheme
Classification and Phylogeny The diversity of life is great. To communicate about it, there must be a scheme for organization. There are many species that would be difficult to organize without a scheme
Biological Networks. Gavin Conant 163B ASRC
 Biological Networks Gavin Conant 163B ASRC conantg@missouri.edu 882-2931 Types of Network Regulatory Protein-interaction Metabolic Signaling Co-expressing General principle Relationship between genes Gene/protein/enzyme
Biological Networks Gavin Conant 163B ASRC conantg@missouri.edu 882-2931 Types of Network Regulatory Protein-interaction Metabolic Signaling Co-expressing General principle Relationship between genes Gene/protein/enzyme
DNA Phylogeny. Signals and Systems in Biology Kushal EE, IIT Delhi
 DNA Phylogeny Signals and Systems in Biology Kushal Shah @ EE, IIT Delhi Phylogenetics Grouping and Division of organisms Keeps changing with time Splitting, hybridization and termination Cladistics :
DNA Phylogeny Signals and Systems in Biology Kushal Shah @ EE, IIT Delhi Phylogenetics Grouping and Division of organisms Keeps changing with time Splitting, hybridization and termination Cladistics :
CSCI1950 Z Computa4onal Methods for Biology Lecture 5
 CSCI1950 Z Computa4onal Methods for Biology Lecture 5 Ben Raphael February 6, 2009 hip://cs.brown.edu/courses/csci1950 z/ Alignment vs. Distance Matrix Mouse: ACAGTGACGCCACACACGT Gorilla: CCTGCGACGTAACAAACGC
CSCI1950 Z Computa4onal Methods for Biology Lecture 5 Ben Raphael February 6, 2009 hip://cs.brown.edu/courses/csci1950 z/ Alignment vs. Distance Matrix Mouse: ACAGTGACGCCACACACGT Gorilla: CCTGCGACGTAACAAACGC
Additive distances. w(e), where P ij is the path in T from i to j. Then the matrix [D ij ] is said to be additive.
![Additive distances. w(e), where P ij is the path in T from i to j. Then the matrix [D ij ] is said to be additive. Additive distances. w(e), where P ij is the path in T from i to j. Then the matrix [D ij ] is said to be additive.](/thumbs/92/110439366.jpg) Additive distances Let T be a tree on leaf set S and let w : E R + be an edge-weighting of T, and assume T has no nodes of degree two. Let D ij = e P ij w(e), where P ij is the path in T from i to j. Then
Additive distances Let T be a tree on leaf set S and let w : E R + be an edge-weighting of T, and assume T has no nodes of degree two. Let D ij = e P ij w(e), where P ij is the path in T from i to j. Then
Phylogenetics. Applications of phylogenetics. Unrooted networks vs. rooted trees. Outline
 Phylogenetics Todd Vision iology 522 March 26, 2007 pplications of phylogenetics Studying organismal or biogeographic history Systematics ating events in the fossil record onservation biology Studying
Phylogenetics Todd Vision iology 522 March 26, 2007 pplications of phylogenetics Studying organismal or biogeographic history Systematics ating events in the fossil record onservation biology Studying
Classification and Phylogeny
 Classification and Phylogeny The diversity it of life is great. To communicate about it, there must be a scheme for organization. There are many species that would be difficult to organize without a scheme
Classification and Phylogeny The diversity it of life is great. To communicate about it, there must be a scheme for organization. There are many species that would be difficult to organize without a scheme
What is Phylogenetics
 What is Phylogenetics Phylogenetics is the area of research concerned with finding the genetic connections and relationships between species. The basic idea is to compare specific characters (features)
What is Phylogenetics Phylogenetics is the area of research concerned with finding the genetic connections and relationships between species. The basic idea is to compare specific characters (features)
Phylogenetic Trees. Phylogenetic Trees Five. Phylogeny: Inference Tool. Phylogeny Terminology. Picture of Last Quagga. Importance of Phylogeny 5.
 Five Sami Khuri Department of Computer Science San José State University San José, California, USA sami.khuri@sjsu.edu v Distance Methods v Character Methods v Molecular Clock v UPGMA v Maximum Parsimony
Five Sami Khuri Department of Computer Science San José State University San José, California, USA sami.khuri@sjsu.edu v Distance Methods v Character Methods v Molecular Clock v UPGMA v Maximum Parsimony
86 Part 4 SUMMARY INTRODUCTION
 86 Part 4 Chapter # AN INTEGRATION OF THE DESCRIPTIONS OF GENE NETWORKS AND THEIR MODELS PRESENTED IN SIGMOID (CELLERATOR) AND GENENET Podkolodny N.L. *1, 2, Podkolodnaya N.N. 1, Miginsky D.S. 1, Poplavsky
86 Part 4 Chapter # AN INTEGRATION OF THE DESCRIPTIONS OF GENE NETWORKS AND THEIR MODELS PRESENTED IN SIGMOID (CELLERATOR) AND GENENET Podkolodny N.L. *1, 2, Podkolodnaya N.N. 1, Miginsky D.S. 1, Poplavsky
Phylogeny: building the tree of life
 Phylogeny: building the tree of life Dr. Fayyaz ul Amir Afsar Minhas Department of Computer and Information Sciences Pakistan Institute of Engineering & Applied Sciences PO Nilore, Islamabad, Pakistan
Phylogeny: building the tree of life Dr. Fayyaz ul Amir Afsar Minhas Department of Computer and Information Sciences Pakistan Institute of Engineering & Applied Sciences PO Nilore, Islamabad, Pakistan
Overview of clustering analysis. Yuehua Cui
 Overview of clustering analysis Yuehua Cui Email: cuiy@msu.edu http://www.stt.msu.edu/~cui A data set with clear cluster structure How would you design an algorithm for finding the three clusters in this
Overview of clustering analysis Yuehua Cui Email: cuiy@msu.edu http://www.stt.msu.edu/~cui A data set with clear cluster structure How would you design an algorithm for finding the three clusters in this
Macroevolution Part I: Phylogenies
 Macroevolution Part I: Phylogenies Taxonomy Classification originated with Carolus Linnaeus in the 18 th century. Based on structural (outward and inward) similarities Hierarchal scheme, the largest most
Macroevolution Part I: Phylogenies Taxonomy Classification originated with Carolus Linnaeus in the 18 th century. Based on structural (outward and inward) similarities Hierarchal scheme, the largest most
Sequence Analysis '17- lecture 8. Multiple sequence alignment
 Sequence Analysis '17- lecture 8 Multiple sequence alignment Ex5 explanation How many random database search scores have e-values 10? (Answer: 10!) Why? e-value of x = m*p(s x), where m is the database
Sequence Analysis '17- lecture 8 Multiple sequence alignment Ex5 explanation How many random database search scores have e-values 10? (Answer: 10!) Why? e-value of x = m*p(s x), where m is the database
Chapter 26: Phylogeny and the Tree of Life Phylogenies Show Evolutionary Relationships
 Chapter 26: Phylogeny and the Tree of Life You Must Know The taxonomic categories and how they indicate relatedness. How systematics is used to develop phylogenetic trees. How to construct a phylogenetic
Chapter 26: Phylogeny and the Tree of Life You Must Know The taxonomic categories and how they indicate relatedness. How systematics is used to develop phylogenetic trees. How to construct a phylogenetic
Molecular Evolution and Phylogenetic Tree Reconstruction
 1 4 Molecular Evolution and Phylogenetic Tree Reconstruction 3 2 5 1 4 2 3 5 Orthology, Paralogy, Inparalogs, Outparalogs Phylogenetic Trees Nodes: species Edges: time of independent evolution Edge length
1 4 Molecular Evolution and Phylogenetic Tree Reconstruction 3 2 5 1 4 2 3 5 Orthology, Paralogy, Inparalogs, Outparalogs Phylogenetic Trees Nodes: species Edges: time of independent evolution Edge length
Workshop: Biosystematics
 Workshop: Biosystematics by Julian Lee (revised by D. Krempels) Biosystematics (sometimes called simply "systematics") is that biological sub-discipline that is concerned with the theory and practice of
Workshop: Biosystematics by Julian Lee (revised by D. Krempels) Biosystematics (sometimes called simply "systematics") is that biological sub-discipline that is concerned with the theory and practice of
Phylogenetics. BIOL 7711 Computational Bioscience
 Consortium for Comparative Genomics! University of Colorado School of Medicine Phylogenetics BIOL 7711 Computational Bioscience Biochemistry and Molecular Genetics Computational Bioscience Program Consortium
Consortium for Comparative Genomics! University of Colorado School of Medicine Phylogenetics BIOL 7711 Computational Bioscience Biochemistry and Molecular Genetics Computational Bioscience Program Consortium
BioSystems 107 (2012) Contents lists available at SciVerse ScienceDirect. BioSystems
 BioSystems 17 (212) 186 196 Contents lists available at SciVerse ScienceDirect BioSystems jo ur nal homep age : www.elsevier.com/locate /biosystems Structural distance and evolutionary relationship of
BioSystems 17 (212) 186 196 Contents lists available at SciVerse ScienceDirect BioSystems jo ur nal homep age : www.elsevier.com/locate /biosystems Structural distance and evolutionary relationship of
Genomic Comparison of Bacterial Species Based on Metabolic Characteristics
 Genomic Comparison of Bacterial Species Based on Metabolic Characteristics Gaurav Jain 1, Haozhu Wang 1, Li Liao 1*, E. Fidelma Boyd 2* 1 Department of Computer and Information Sciences University of Delaware
Genomic Comparison of Bacterial Species Based on Metabolic Characteristics Gaurav Jain 1, Haozhu Wang 1, Li Liao 1*, E. Fidelma Boyd 2* 1 Department of Computer and Information Sciences University of Delaware
Honor pledge: I have neither given nor received unauthorized aid on this test. Name :
 Midterm Exam #1 MB 451 : Microbial Diversity Honor pledge: I have neither given nor received unauthorized aid on this test. Signed : Date : Name : 1. What are the three primary evolutionary branches of
Midterm Exam #1 MB 451 : Microbial Diversity Honor pledge: I have neither given nor received unauthorized aid on this test. Signed : Date : Name : 1. What are the three primary evolutionary branches of
Workshop III: Evolutionary Genomics
 Identifying Species Trees from Gene Trees Elizabeth S. Allman University of Alaska IPAM Los Angeles, CA November 17, 2011 Workshop III: Evolutionary Genomics Collaborators The work in today s talk is joint
Identifying Species Trees from Gene Trees Elizabeth S. Allman University of Alaska IPAM Los Angeles, CA November 17, 2011 Workshop III: Evolutionary Genomics Collaborators The work in today s talk is joint
Network Alignment 858L
 Network Alignment 858L Terms & Questions A homologous h Interolog = B h Species 1 Species 2 Are there conserved pathways? What is the minimum set of pathways required for life? Can we compare networks
Network Alignment 858L Terms & Questions A homologous h Interolog = B h Species 1 Species 2 Are there conserved pathways? What is the minimum set of pathways required for life? Can we compare networks
Non-independence in Statistical Tests for Discrete Cross-species Data
 J. theor. Biol. (1997) 188, 507514 Non-independence in Statistical Tests for Discrete Cross-species Data ALAN GRAFEN* AND MARK RIDLEY * St. John s College, Oxford OX1 3JP, and the Department of Zoology,
J. theor. Biol. (1997) 188, 507514 Non-independence in Statistical Tests for Discrete Cross-species Data ALAN GRAFEN* AND MARK RIDLEY * St. John s College, Oxford OX1 3JP, and the Department of Zoology,
Weighted gene co-expression analysis. Yuehua Cui June 7, 2013
 Weighted gene co-expression analysis Yuehua Cui June 7, 2013 Weighted gene co-expression network (WGCNA) A type of scale-free network: A scale-free network is a network whose degree distribution follows
Weighted gene co-expression analysis Yuehua Cui June 7, 2013 Weighted gene co-expression network (WGCNA) A type of scale-free network: A scale-free network is a network whose degree distribution follows
How to read and make phylogenetic trees Zuzana Starostová
 How to read and make phylogenetic trees Zuzana Starostová How to make phylogenetic trees? Workflow: obtain DNA sequence quality check sequence alignment calculating genetic distances phylogeny estimation
How to read and make phylogenetic trees Zuzana Starostová How to make phylogenetic trees? Workflow: obtain DNA sequence quality check sequence alignment calculating genetic distances phylogeny estimation
Clustering. CSL465/603 - Fall 2016 Narayanan C Krishnan
 Clustering CSL465/603 - Fall 2016 Narayanan C Krishnan ckn@iitrpr.ac.in Supervised vs Unsupervised Learning Supervised learning Given x ", y " "%& ', learn a function f: X Y Categorical output classification
Clustering CSL465/603 - Fall 2016 Narayanan C Krishnan ckn@iitrpr.ac.in Supervised vs Unsupervised Learning Supervised learning Given x ", y " "%& ', learn a function f: X Y Categorical output classification
Evolutionary Tree Analysis. Overview
 CSI/BINF 5330 Evolutionary Tree Analysis Young-Rae Cho Associate Professor Department of Computer Science Baylor University Overview Backgrounds Distance-Based Evolutionary Tree Reconstruction Character-Based
CSI/BINF 5330 Evolutionary Tree Analysis Young-Rae Cho Associate Professor Department of Computer Science Baylor University Overview Backgrounds Distance-Based Evolutionary Tree Reconstruction Character-Based
Consistency Index (CI)
 Consistency Index (CI) minimum number of changes divided by the number required on the tree. CI=1 if there is no homoplasy negatively correlated with the number of species sampled Retention Index (RI)
Consistency Index (CI) minimum number of changes divided by the number required on the tree. CI=1 if there is no homoplasy negatively correlated with the number of species sampled Retention Index (RI)
Name: Class: Date: ID: A
 Class: _ Date: _ Ch 17 Practice test 1. A segment of DNA that stores genetic information is called a(n) a. amino acid. b. gene. c. protein. d. intron. 2. In which of the following processes does change
Class: _ Date: _ Ch 17 Practice test 1. A segment of DNA that stores genetic information is called a(n) a. amino acid. b. gene. c. protein. d. intron. 2. In which of the following processes does change
networks in molecular biology Wolfgang Huber
 networks in molecular biology Wolfgang Huber networks in molecular biology Regulatory networks: components = gene products interactions = regulation of transcription, translation, phosphorylation... Metabolic
networks in molecular biology Wolfgang Huber networks in molecular biology Regulatory networks: components = gene products interactions = regulation of transcription, translation, phosphorylation... Metabolic
Phylogenetic Analysis of Molecular Interaction Networks 1
 Phylogenetic Analysis of Molecular Interaction Networks 1 Mehmet Koyutürk Case Western Reserve University Electrical Engineering & Computer Science 1 Joint work with Sinan Erten, Xin Li, Gurkan Bebek,
Phylogenetic Analysis of Molecular Interaction Networks 1 Mehmet Koyutürk Case Western Reserve University Electrical Engineering & Computer Science 1 Joint work with Sinan Erten, Xin Li, Gurkan Bebek,
1 ATGGGTCTC 2 ATGAGTCTC
 We need an optimality criterion to choose a best estimate (tree) Other optimality criteria used to choose a best estimate (tree) Parsimony: begins with the assumption that the simplest hypothesis that
We need an optimality criterion to choose a best estimate (tree) Other optimality criteria used to choose a best estimate (tree) Parsimony: begins with the assumption that the simplest hypothesis that
Taxonomy. Content. How to determine & classify a species. Phylogeny and evolution
 Taxonomy Content Why Taxonomy? How to determine & classify a species Domains versus Kingdoms Phylogeny and evolution Why Taxonomy? Classification Arrangement in groups or taxa (taxon = group) Nomenclature
Taxonomy Content Why Taxonomy? How to determine & classify a species Domains versus Kingdoms Phylogeny and evolution Why Taxonomy? Classification Arrangement in groups or taxa (taxon = group) Nomenclature
SPATIAL DATA MINING. Ms. S. Malathi, Lecturer in Computer Applications, KGiSL - IIM
 SPATIAL DATA MINING Ms. S. Malathi, Lecturer in Computer Applications, KGiSL - IIM INTRODUCTION The main difference between data mining in relational DBS and in spatial DBS is that attributes of the neighbors
SPATIAL DATA MINING Ms. S. Malathi, Lecturer in Computer Applications, KGiSL - IIM INTRODUCTION The main difference between data mining in relational DBS and in spatial DBS is that attributes of the neighbors
Phylogeny 9/8/2014. Evolutionary Relationships. Data Supporting Phylogeny. Chapter 26
 Phylogeny Chapter 26 Taxonomy Taxonomy: ordered division of organisms into categories based on a set of characteristics used to assess similarities and differences Carolus Linnaeus developed binomial nomenclature,
Phylogeny Chapter 26 Taxonomy Taxonomy: ordered division of organisms into categories based on a set of characteristics used to assess similarities and differences Carolus Linnaeus developed binomial nomenclature,
METABOLIC PATHWAY PREDICTION/ALIGNMENT
 COMPUTATIONAL SYSTEMIC BIOLOGY METABOLIC PATHWAY PREDICTION/ALIGNMENT Hofestaedt R*, Chen M Bioinformatics / Medical Informatics, Technische Fakultaet, Universitaet Bielefeld Postfach 10 01 31, D-33501
COMPUTATIONAL SYSTEMIC BIOLOGY METABOLIC PATHWAY PREDICTION/ALIGNMENT Hofestaedt R*, Chen M Bioinformatics / Medical Informatics, Technische Fakultaet, Universitaet Bielefeld Postfach 10 01 31, D-33501
University of Florida CISE department Gator Engineering. Clustering Part 1
 Clustering Part 1 Dr. Sanjay Ranka Professor Computer and Information Science and Engineering University of Florida, Gainesville What is Cluster Analysis? Finding groups of objects such that the objects
Clustering Part 1 Dr. Sanjay Ranka Professor Computer and Information Science and Engineering University of Florida, Gainesville What is Cluster Analysis? Finding groups of objects such that the objects
Gene Ontology and Functional Enrichment. Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein
 Gene Ontology and Functional Enrichment Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein The parsimony principle: A quick review Find the tree that requires the fewest
Gene Ontology and Functional Enrichment Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein The parsimony principle: A quick review Find the tree that requires the fewest
Fig. 26.7a. Biodiversity. 1. Course Outline Outcomes Instructors Text Grading. 2. Course Syllabus. Fig. 26.7b Table
 Fig. 26.7a Biodiversity 1. Course Outline Outcomes Instructors Text Grading 2. Course Syllabus Fig. 26.7b Table 26.2-1 1 Table 26.2-2 Outline: Systematics and the Phylogenetic Revolution I. Naming and
Fig. 26.7a Biodiversity 1. Course Outline Outcomes Instructors Text Grading 2. Course Syllabus Fig. 26.7b Table 26.2-1 1 Table 26.2-2 Outline: Systematics and the Phylogenetic Revolution I. Naming and
RECOVERING NORMAL NETWORKS FROM SHORTEST INTER-TAXA DISTANCE INFORMATION
 RECOVERING NORMAL NETWORKS FROM SHORTEST INTER-TAXA DISTANCE INFORMATION MAGNUS BORDEWICH, KATHARINA T. HUBER, VINCENT MOULTON, AND CHARLES SEMPLE Abstract. Phylogenetic networks are a type of leaf-labelled,
RECOVERING NORMAL NETWORKS FROM SHORTEST INTER-TAXA DISTANCE INFORMATION MAGNUS BORDEWICH, KATHARINA T. HUBER, VINCENT MOULTON, AND CHARLES SEMPLE Abstract. Phylogenetic networks are a type of leaf-labelled,
ELE4120 Bioinformatics Tutorial 8
 ELE4120 ioinformatics Tutorial 8 ontent lassifying Organisms Systematics and Speciation Taxonomy and phylogenetics Phenetics versus cladistics Phylogenetic trees iological classification Goal: To develop
ELE4120 ioinformatics Tutorial 8 ontent lassifying Organisms Systematics and Speciation Taxonomy and phylogenetics Phenetics versus cladistics Phylogenetic trees iological classification Goal: To develop
Principles of Pattern Recognition. C. A. Murthy Machine Intelligence Unit Indian Statistical Institute Kolkata
 Principles of Pattern Recognition C. A. Murthy Machine Intelligence Unit Indian Statistical Institute Kolkata e-mail: murthy@isical.ac.in Pattern Recognition Measurement Space > Feature Space >Decision
Principles of Pattern Recognition C. A. Murthy Machine Intelligence Unit Indian Statistical Institute Kolkata e-mail: murthy@isical.ac.in Pattern Recognition Measurement Space > Feature Space >Decision
What is the range of a taxon? A scaling problem at three levels: Spa9al scale Phylogene9c depth Time
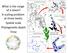 What is the range of a taxon? A scaling problem at three levels: Spa9al scale Phylogene9c depth Time 1 5 0.25 0.15 5 0.05 0.05 0.10 2 0.10 0.10 0.20 4 Reminder of what a range-weighted tree is Actual Tree
What is the range of a taxon? A scaling problem at three levels: Spa9al scale Phylogene9c depth Time 1 5 0.25 0.15 5 0.05 0.05 0.10 2 0.10 0.10 0.20 4 Reminder of what a range-weighted tree is Actual Tree
On the Uniqueness of the Selection Criterion in Neighbor-Joining
 Journal of Classification 22:3-15 (2005) DOI: 10.1007/s00357-005-0003-x On the Uniqueness of the Selection Criterion in Neighbor-Joining David Bryant McGill University, Montreal Abstract: The Neighbor-Joining
Journal of Classification 22:3-15 (2005) DOI: 10.1007/s00357-005-0003-x On the Uniqueness of the Selection Criterion in Neighbor-Joining David Bryant McGill University, Montreal Abstract: The Neighbor-Joining
Lecture 6 Phylogenetic Inference
 Lecture 6 Phylogenetic Inference From Darwin s notebook in 1837 Charles Darwin Willi Hennig From The Origin in 1859 Cladistics Phylogenetic inference Willi Hennig, Cladistics 1. Clade, Monophyletic group,
Lecture 6 Phylogenetic Inference From Darwin s notebook in 1837 Charles Darwin Willi Hennig From The Origin in 1859 Cladistics Phylogenetic inference Willi Hennig, Cladistics 1. Clade, Monophyletic group,
Neighbor Joining Algorithms for Inferring Phylogenies via LCA-Distances
 Neighbor Joining Algorithms for Inferring Phylogenies via LCA-Distances Ilan Gronau Shlomo Moran September 6, 2006 Abstract Reconstructing phylogenetic trees efficiently and accurately from distance estimates
Neighbor Joining Algorithms for Inferring Phylogenies via LCA-Distances Ilan Gronau Shlomo Moran September 6, 2006 Abstract Reconstructing phylogenetic trees efficiently and accurately from distance estimates
Chapter 19. Microbial Taxonomy
 Chapter 19 Microbial Taxonomy 12-17-2008 Taxonomy science of biological classification consists of three separate but interrelated parts classification arrangement of organisms into groups (taxa; s.,taxon)
Chapter 19 Microbial Taxonomy 12-17-2008 Taxonomy science of biological classification consists of three separate but interrelated parts classification arrangement of organisms into groups (taxa; s.,taxon)
Biology 211 (2) Week 1 KEY!
 Biology 211 (2) Week 1 KEY Chapter 1 KEY FIGURES: 1.2, 1.3, 1.4, 1.5, 1.6, 1.7 VOCABULARY: Adaptation: a trait that increases the fitness Cells: a developed, system bound with a thin outer layer made of
Biology 211 (2) Week 1 KEY Chapter 1 KEY FIGURES: 1.2, 1.3, 1.4, 1.5, 1.6, 1.7 VOCABULARY: Adaptation: a trait that increases the fitness Cells: a developed, system bound with a thin outer layer made of
PHYLOGENY AND SYSTEMATICS
 AP BIOLOGY EVOLUTION/HEREDITY UNIT Unit 1 Part 11 Chapter 26 Activity #15 NAME DATE PERIOD PHYLOGENY AND SYSTEMATICS PHYLOGENY Evolutionary history of species or group of related species SYSTEMATICS Study
AP BIOLOGY EVOLUTION/HEREDITY UNIT Unit 1 Part 11 Chapter 26 Activity #15 NAME DATE PERIOD PHYLOGENY AND SYSTEMATICS PHYLOGENY Evolutionary history of species or group of related species SYSTEMATICS Study
