supplementary information
|
|
|
- Anissa Stokes
- 5 years ago
- Views:
Transcription
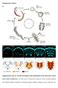
1 DOI: /ncb1825 Figure S1 Venus::UNC-6 expression prior to and during AC invasion. (a-c) Nomarski images overlayed with Venus::UNC-6 ( SP) expression shown in yellow (left), and Venus::UNC-6 ( SP) fluorescence alone (right). Venus::UNC-6 ( SP) lacks a signal sequence and was thus retained in the cells in which unc-6 is expressed. Venus::UNC-6 was retained within the neurons of the ventral nerve cord (VNC, small bracket) under the AC (arrow) prior to invasion at the P6.p 1- and 2-cell stages (large brackets, a and b), and during the time of invasion at constant levels (c). Scale bar in (a) is 5µm in this and all subsequent supplementary figures. 1
2 Figure S2 Venus::UNC-6 expression and localization is not dependent on the vulval cells. Nomarski image (left), and corresponding fluorescence image (right) of an animal expressing Venus::UNC-6 at the early L3 stage after laser-mediated ablation of the vulval cells in the early L2 larval stages. Venus::UNC-6 was expressed normally in the VNC and localized to the basement membrane (arrowhead) under the AC (arrow) as in wild-type animals (n = 24/24 animals). Inset highlights the basement membrane localization of Venus::UNC-6 (arrowhead) under the AC. 2
3 Figure S3 AC-specific expression of UNC-40::GFP. (a) Nomarski image overlayed with UNC-40::GFP fluorescence (green). The unc-40::gfp construct is driven by the AC-specific regulatory element of the cdh-3 gene. (b) The corresponding fluorescence image of UNC-40::GFP fluorescence alone. The cdh-3 AC-specific promoter drives expression solely in the AC (arrow), and not in neighboring cells, including vulval cells (brackets). The asterisk marks autofluorescence from the gut granules. 3
4 Figure S4 The Rac orthologs GFP::MIG-2 and GFP::CED-10 are polarized along the invasive cell membrane of the AC. (a-e) Nomarski images (left), corresponding fluorescence images (right). Time is post-hatching at 20ºC. (a) GFP::MIG-2 expression driven by its endogenous promoter was first expressed in the AC during the late L2 (9.5 hours before invasion) and was not polarized (16/16 animals). (b) MIG-2 was first polarized at the L2 molt (arrowhead; 7/11 animals). (c-e). Polarized MIG-2 was maintained along the invasive cell membrane from the early L3 stage through the time of invasion (20/20 animals for each stage). (f) Projected confocal z-stack showing uterine and vulval expression of GFP::CED-10 driven by its endogenous promoter (left) and the corresponding spectral representation of the fluorescence intensity (right). CED-10 was expressed in all uterine and vulval cells (bracket outlines the 1º VPCs), and was polarized along the basal (invasive) membrane of the AC (arrow) but not in neighboring uterine cells (arrowheads). 4
5 Figure S5 PAR-3::GFP and apical AJM-1::GFP are localized normally in unc-6 mutants. (a, b) Nomarski images (left), corresponding fluorescence images (right). (a) PAR-3::GFP was localized to apical and lateral membranes of wild-type ACs prior to invasion (arrowheads), and this polarity was not altered in unc-6 mutants (b, arrowheads; n = 20/20 animals). (c) In wild-type animals AJM-1::GFP (green) localized to nascent junctions (arrowhead) in the apical region of the AC (expressing zmp-1>mcherry in magenta). (d) In unc-6 mutants apical AJM-1::GFP junctions were formed and positioned normally compared to wild-type animals (n 60 wild-type and unc-6 animals examined). 5
6 Figure S6 Pan-uterine expression of mcherry:: PLCδ PH reveals a unique PI(4,5)P 2 rich invasive membrane domain in the AC. Fluorescence (left) overlaid on corresponding Normarski image (right). (a) Expression of the PI(4,5)P 2 sensor mcherry::plcδ PH in the AC revealed strong polarization at the basal (invasive) cell membrane (arrowhead). (b) In contrast, a section lateral to the AC through the neighboring ventral and dorsal uterine cells (VU/DU) expressing PLCδ PH ::mcherry, revealed that adjacent uterine cells did not have PI(4,5)P 2 concentrated in basal membranes (arrowheads). 6
7 Figure S7 GFP::MIG-2 is localized normally in the AC of fos-1(ar105) animals. Nomarski images (left), corresponding fluorescence images (right). In (a) wild-type animals and (b) fos-1(ar105) mutants, MIG-2::GFP was strongly polarized to the invasive cell membrane of the AC (arrowhead; n = 20/20 fos-1a mutants examined). (c) In unc-6 animals, MIG-2::GFP polarity was perturbed. 7
8 Supplementary Movie Legends Movie S1 Hemicentin::GFP deposition under the AC in wild-type animals. Hemicentin::GFP was localized at low levels along gonadal and ventral epidermal basement membranes, but was deposited at high levels (green, orange arrows) under the AC s invasive cell membrane (expressing cdh-3>mcherry::moeabd in magenta) during AC invasion. Little hemicentin::gfp was deposited along apical or lateral domains of the AC (note small deposit at white arrowhead). Movie S2 Hemicentin::GFP deposition is perturbed unc-6 mutants. In unc-6 mutants there was a dramatic decrease in hemicentin deposited under the AC (expressing cdh-3>mcherry::moeabd in magenta) at the site of basement membrane contact (green, orange arrows), while apical and lateral accumulations of hemicentin increased (white arrowheads). 8
9 8 (Sherwood) Table S1. AC invasion in mutant strains with roles in cell motility and axon outgrowth Invasion at Invasion at P6.p 4-cell P6.p 8-cell stage C. elegans gene Encoded Product stage (number of ACs (number of ACs invaded/number observed) a invaded/number observed) arf-1.2(ok796) ADP-ribosylation factor 1 homolog 10/10 10/10 arf-1.2(ok1322) ADP-ribosylation factor 1 homolog 9/9 4/4 bar-1 (ga80) Armadillo/beta-Catenin/plakoglobin 11/11 9/9 protein tyrosine kinase homolog that is also homologous to C25F6.4(ok874) 15/15 16/16 human RS1 receptor tyrosine kinase of the immunoglobulin superfamily cam-1(gm122) 7/9 28/35 that is orthologous to human ROR1 uncloned locus that affects migration of canal associated cam-2(gm124) 6/6 5/5 neurons cdh-4(ok1323) member of the cadherin superfamily 10/10 8/8 cdh-5(hc181) member of the cadherin superfamily 10/10 7/7 cdh-7(ok428) contains a cadherin domain 10/10 not observed ced-2(e1752) Src homology (SH) 2 and 3-containing adaptor protein 10/10 11/11 ced-5(n1812) homolog of the human protein DOCK180 10/10 13/13 ced-12(k149) Regulator of Rac1, required for phagocytosis and cell migration 11/11 12/12 ceh-10(ct78) Paired-like class of homeodomain proteins 10/10 8/8 C2H2-type zinc finger transcription factor that is a member of ces-1(n1414) 11/11 11/11 the Snail family of proteins C2H2-type zinc finger transcription factor that is a member of ces-1(n703) 16/16 14/14 the Snail family of proteins cle-1(cg120) collagen protein with endostatin domain 9/9 13/13 crb-1(ok931) homolog of Drosophila CRUMBS 9/9 9/9 daf-1(m40) TGF-beta type I receptor homolog 8/9 6/6 daf-4(e1364) type II transforming growth factor-beta (TGF-b) receptors 11/11 14/14 member of the transforming growth factor beta (TGFbeta) daf-7(e1372) 12/12 14/14 superfamily dgn-2(ok209) dystroglycan 27/27 20/20 member of the transforming growth factor beta (TGFbeta) dbl-1(wk70) 21/21 6/6 superfamily dpy-19(e1295) novel transmembrane protein 7/7 7/7 efn-4(bx80) member of the ephrin family of ligands 10/10 17/17 egl-15(mt3324) FGF-like receptor tyrosine kinase 13/13 7/7 egl-17(e1313) fibroblast growth factor-like protein 13/13 8/8 a functional ortholog of human ADP-ribosylation factor-like evl-20(ar103) not observed 4/4 protein 2 F11D5.3(ok574) a putative tyrosine kinase homologous to human RS1 11/11 22/22 flt-1(ok722) a putative homolog of flectin, an extracellular matrix protein 10/10 8/8 a functional metalloprotease that defines a new sub-family of gon-1(e1254)/edf18 secreted proteases known as MPT (metalloprotease with 8/8 11/11 thrombospondin type 1 repeats) glypican, a heparan sulfate proteoglycan anchored to the cell gpn-1(ok377) 13/13 14/14 membrane by a GPI linkage helix-loop-helix protein required for normal muscle hlh-8(nr2061) 10/10 18/18 development lad-1(ok1244)/sax-7 ortholog of human L1CAM 10/10 13/13 let-756(s2613) fibroblast growth factor (FGF)-like ligand 9/11 21/21 member of the glypican family of heparan sulfate lon-2(e678) 13/13 19/19 proteoglycans mab-20 (ev574) semaphorin 9/10 13/13 mab-20(bx24) semaphorin 10/10 10/10 max-1(ju39) conserved PH/MyTH4/FERM domain-containing protein 15/15 10/10 mig-1(e1787) Frizzled-like receptor 10/10 13/13 mig-2(gm103) Rho family of GTP-binding proteins, similar to Rac 12/15 36/37 mig-6(e1931) uncloned locus involved in cell migration 9/10 15/15 mig-14(ga62) homologous to drosophila Wntless 10/10 15/15 mig-15(rh326) Nck-interacting kinase (NIK) 8/9 10/10 secreted metalloprotease that is a member of the ADAM (A mig-17(k113) 10/10 13/13 Disintegrin And Metalloprotease) protein family
10 9 (Sherwood) mom-2(or42) member of the Wnt family of secreted signaling glycoproteins 6/6 20/20 mom-5(or57) Frizzled-like receptor 2/5 7/8 pkc-3(rnai) Serine/threonine protein kinase, atypical Protien Kinase C 15/15 no data pld-1(ok986) phospholipase D1 10/10 2/2 plx-1(nc37) plexin ortholog, semaphorin receptor 10/10 10/10 ptp-3(op147) receptor-like tyrosine phosphatase 10/10 15/15 qid-7(mu533) uncloned locus that affects Q neuroblast polarization and migration 11/11 10/10 qid-8(mu342) uncloned locus that affects Q neuroblast polarization and migration 10/10 13/15 rac-2(ok326) Rho family GTPase that is one of three C. elegans Rac-related proteins 10/10 7/7 rig-4(ok1160) Immunoglobulin C-2 Type/fibronectin type III domains 10/10 5/5 sax-3(ky123) homologous to Drosophila roundabout 11/11 10/10 sdn-1(ok449) a type I transmembrane heparan sulfate proteoglycan, syndecan 13/13 5/5 slt-1(eh15) homolog of Drosophila slit 16/16 11/11 slt-1(ok255) homolog of Drosophila slit 80/81 59/59 sma-6(wk7) serine/threonine protein kinase that is orthologous to type I TGF-beta receptors 10/10 15/15 smp-1 (ev715) semaphorin 10/11 12/12 smp-2 (ev709) semaphorin 10/10 10/10 syg-1(ky652) a novel transmembrane protein 9/9 12/12 syg-2(ky671) transmembrane protein, immunoglobulin superfamily 10/10 4/4 tag-150(gk261) Guanine nucleotide exchange factor for Rho and Rac GTPases 10/10 11/11 unc-3(e151) a protein with homology to immunoglobulin (Ig) domaincontaining transcription factors 10/10 10/10 unc-5(e53) a netrin receptor required for dorsal cell and axon migration 10/10 22/22 unc-6(ev400) netrin ortholog, secreted guidance molecule that regulates pioneer axons and mesodermal cells 4/20 14/18 unc-14(e57) an activity required for both axonogenesis and sex myoblast migration 8/10 11/11 unc-18(e81) an ortholog of Saccharomyces cervisiae SEC1 10/11 12/12 unc-30(e191) Pitx homeodomain transcription factor family member 10/10 10/10 unc-33(mn407) CRMP/TOAD/Ulip/DRP homologue 10/10 10/11 unc-34(e315) EVH1 domain-containing protein that is the sole C. elegans Enabled/VASP homolog 6/10 12/12 unc-34(e566) EVH1 domain-containing protein that is the sole C. elegans Enabled/VASP homolog 5/8 5/5 unc-39(e257) homeodomain transcription factor that belongs to the Six4/5 family 10/10 11/11 unc-40(e271) A netrin receptor required for guiding dorsal and ventral cell and axon migrations 6/21 13/18 unc-44(e1197) ankyrin-like protein 10/10 10/10 unc-44(e362) ankyrin-like protein 10/10 2/2 unc-51 (e369) serine/threonine kinase involved in autophagy 9/10 13/13 unc-51(e1189) serine/threonine kinase involved in autophagy 10/12 10/10 unc-53(e404) orthologous to human NAV1, NAV2/RAINB1, and NAV3 11/11 7/7 unc-53(n152) orthologous to human NAV1, NAV2/RAINB1, and NAV3 10/10 13/13 unc-60(e723) orthologs of actin depolymerizing factor/cofilin 15/15 16/16 unc-60(su158) orthologs of actin depolymerizing factor/cofilin 37/39 30/30 unc-71 (e541) ADAM, a disintegrin and metalloprotease-containing transmembrane protein 10/10 10/10 unc-73 (e936) guanine nucleotide exchange factor (GNEF) similar to the trio protein not observed 39/39 unc-73 (gm40) guanine nucleotide exchange factor (GNEF) similar to the trio protein 6/11 22/22 unc-76(e911) coiled-coil protein that belongs to the FEZ (fasciculation and elongation protein; zygin/zeta-1) family of proteins 9/11 10/10 unc-78(gk27) homolog of actin-interacting protein 1 26/26 27/27 unc-97(su10) LIM domain-containing protein of the PINCH family 10/10 14/14 unc-115(e2225) Actin-binding LIM Zn-finger protein Limatin involved in axon guidance 10/10 15/15 unc-115(ky275) Actin-binding LIM Zn-finger protein Limatin involved in axon guidance 67/67 68/68 unc-115(mn481) Actin-binding LIM Zn-finger protein Limatin 8/11 24/24 unc-129 (ev554) member of the transforming growth factor beta (TGFbeta) superfamily 5/5 11/11
11 10 (Sherwood) vab-1(e2) ephrin receptor 10/10 10/10 vab-8(e1017) novel protein containing an atypical kinesin-like motor domain 10/10 10/10 vab-9(e1744) claudin homolog 8/9 13/13 ver-1(ok859) Fibroblast/platelet-derived growth factor receptor and related receptor tyrosine kinases 10/10 4/4 ver-2(ok897) Fibroblast/platelet-derived growth factor receptor and related receptor tyrosine kinases 10/10 8/8 ver-3(gk227) Fibroblast/platelet-derived growth factor receptor and related receptor tyrosine kinases 11/11 13/13 ver-4(ok1079) Fibroblast/platelet-derived growth factor receptor and related receptor tyrosine kinases 10/10 5/5 zig-1(ok784) secreted 2-immunoglobulin-domain protein 9/9 5/5 zig-2(ok696) secreted 2-immunoglobulin-domain protein 9/9 11/11 zig-3(gk33) secreted 2-immunoglobulin-domain protein 10/10 4/4 zig-4(gk4) secreted 2-immunoglobulin-domain protein 10/10 4/4 zig-5(ok1065) secreted 2-immunoglobulin-domain protein 11/11 10/10 zig-6(ok723) secreted 2-immunoglobulin-domain protein 10/10 3/3 zig-8(ok561) secreted 2-immunoglobulin-domain protein 9/9 4/4 a ACs showing any degree of invasion were scored as invaded and invasion was scored over the entire range of the 4-cell stage, including the L3 molt. In contrast, Table S2 4-cell stage animals were only scored at beginning of the 4-cell stage at the mid-to-late L3 stage, and partial invasions were scored.
12 11 (Sherwood) Table S2. Timing and degree of AC invasion into the vulval epithelium Genotype/Treatment ACs showing full, partial or no invasion P6.p 4-cell stage (mid-to-late L3 stage) P6.p 8-cell stage (early L4 stage) % Full Invasion % Partial Invasion % No Invasion n = % Full Invasion % Partial Invasion % No Invasion n = wild-type (N2) > >100 unc-6/site of action unc-6(ev400) h 54 ghis8[unc-6>venus::unc-6]; unc-6(ev400) a unc-6(ev400); ghex15[glr-1p>venus::unc-6; tph-1p>gfp] b unc-6(ev400);ghex13[egl-17>venus::unc-6] c rde-1(ne219);ghex11[egl-17>rde1::mrfp]; unc-6(rnai) d lin-3(n1059)/lin-3(n378) (Vul) e i 52 lin-3(n1059)/lin-3(n378);unc-6(ev400) j 50 laser killed P3.p-P8.p(Vul); unc-6(ev400) k 20 kyis299 [hs>unc-6::ha] f nd nd nd nd N2(mock heat shock) nd nd nd nd Netrin receptors/unc-40/site of action unc-40(e271) l 53 unc-40(e271); unc-6(ev400) m 70 unc-40(e271); qyis66[cdh-3>unc-40::gfp] unc-5(e51) Intracellular effectors mig-10(ct41) ced-10(n1993) mig-2(mu28) ced-10(n1993); mig-2(mu28) unc-34(gm104) qyis23(cdh-3>mcherry::plcδ PH ) g qyis23(cdh-3>mcherry::plcδ PH ); mig (mu28) qyis23(cdh-3>mcherry::plcδ PH ); unc- 34(gm104) fos-1 pathway interaction unc-40(e271); rrf-3(pk1426) unc-40(e271); rrf-3(pk1426); fos-1(rnai) rrf-3(pk1426); fos-1(rnai) a Venus::unc-6 driven by its own promoter. 11 b Ventral nerve cord specific expression of Venus::unc-6 driven by glr-1 promoter. 11 c 1º VPC specific expression of Venus::unc-6 driven by the egl-17 promoter. 11 d Targeted RNAi mediated knockdown of unc-6 in the 1º VPCs. 11 e A similar percentage of ACs have been shown to invade when vulval cells are removed by ablation. 5 f 2h heat shock at 30ºC followed by 4h recovery at 15ºC. g qyis23(cdh-3> mcherry::plcδ PH ) binds and sequesters PI(4,5)P 2 in the AC, thus reducing its levels there. h-m The number of AC s that detached from the basement membrane at the L4 stage was as follows (number of detached/number that failed to invade): h = 2/12; i =11/42; j = 47/50; k = 18/20 l = 0/9; m= 2/15. nd = not determined
13 12 (Sherwood) Table S3. Polarized marker localization in wild-type and mutant ACs Genotype Marker Polarized (%±SE) Classification of marker localization patterns in wild-type and various mutant ACs P6.p 1-cell stage P6.p 2-cell stage P6.p 4-cell stage Apicolateral No Apicolateral No Apicolateral No Polarized Polarized Accumulation Polarity N= Accumulation Polarity N= Accumulation Polarity (%±SE) (%±SE) (%±SE) (%±SE) (%±SE) (%±SE) (%±SE) (%±SE) wild-type mcherry::plcδ PH 85±8 15± ±8 14± ±5 0 5±5 20 wild-type GFP::MIG-2 86±7 13± wild-type mcherry::moeabd ±5 5± wild-type UNC-40::GFP 90±7 5±5 5± ±6 10± unc-6(ev400) mcherry::plcδ PH 22±10 38±12 38± ±9 53±12 32± ±12 31±12 16 unc-6(ev400) mcherry::moeabd 29±11 53±12 18± ±11 71± ±10 76±11 6±6 17 unc-6(ev400) UNC-40::GFP 21±10 32±11 47± ±5 42±12 53± ±10 75±10 20 unc-6(ev400) GFP::MIG-2 33±11 38±11 29± ±8 43±11 43± ±10 40±11 35±11 20 Vul a mcherry::plcδ PH 84±9 16± ±7 10± ±12 27± Vul a UNC-40::GFP 94±6 6± ±8 14± ±6 6± Vul a GFP::MIG-2 95±5 5± ±7 13± ±10 24± Vul a mcherry::moeabd 88±9 6±6 6± ±8 14± ±7 10± hs> unc-6::ha b UNC-40::GFP nd nd nd nd nd nd nd nd 33±13 33±13 33±13 15 wild-type (mock) c UNC-40::GFP nd nd nd nd nd nd nd nd 93±7 7± a Vul (vulvaless) animals were of the genotype lin-3(n1059)/lin-3(n378). b Heat shock directed expression of unc-6::ha from the integrated transgene kyis299, 2h heat shock at 30ºC followed by 4h recovery at 15ºC. c wild-type animals expressing UNC-40::GFP in the AC were subjected to an identical experimental regimen to confirm that heat shock alone does not cause defects in UNC-40::GFP localization. N=
14 13 (Sherwood) Table S4. Primer sequences and templates used for PCR fusions Primer Sequence Primer Type Amplicon Template 5 TAA TgT gag TTA gct CAC TCA TTA gg 3 forward cdh-3/zmp-1> promoters cdh-3>, ppd107.94/mk62-63; zmp-1>, ppd107.94/mk AAC gat gga TAC gct AAC AAC TTg g 3 forward nested cdh-3/zmp-1> promoters cdh-3>, ppd107.94/mk62-63; zmp-1>, ppd107.94/mk TTT CTg AgC TCg gta CCC TCC AAg 3 reverse cdh-3/zmp-1> promoters cdh-3>, ppd107.94/mk62-63; zmp-1>, ppd107.94/mk TAg gct TTT CCg TAT AgC ATC CTC 3 forward fos-1a> promoter cosmid F29G9 5 gcc CAA CTC TAg TCA TTT CTA gc 3 forward nested fos-1a> promoter cosmid F29G9 5 TCC ACT CTC TTA TAT AgC AgA ggt gc 3 reverse fos-1a> promoter cosmid F29G9 5 CTT gga ggg TAC CgA gct Cag AAA ggt ACC Atg AgT AAA gga gaa g 3 cdh-3/zmp-1 extension, forward GFP::unc-34 punc-86>gfp::unc-34 5 Cgg gaa gct AgA gta AgT AgT TCg CC 3 reverse GFP::unc-34 punc-86>gfp::unc-34 5 CTC TCA Agg ATC TTA CCg CTg TTg 3 reverse nested GFP::unc-34 punc-86>gfp::unc-34 5 CTT gga ggg TAC CgA gct Cag AAA ATg ATT TTg CgA CAT TTC gg 3 cdh-3/zmp-1 extension, forward unc-40::gfp punc-86>unc-40::gfp 5 gtg CCA CCT gac gtc TAA g 3 reverse unc-40::gfp punc-86>unc-40::gfp 5 gta Cgg CCg ACT AgT Agg AAA CAg T 3 reverse nested unc-40::gfp punc-86>unc-40::gfp 5 CTT gga ggg TAC CgA gct CAg AAA ATg gtc TCA AAG ggt gaa g 3 cdh-3/zmp-1 extension, forward mcherry::moeabd pjwz6 5 CAg gaa ACA gct ATg ACC ATg 3 reverse mcherry::moeabd pjwz6 5 gcc gct CTA gaa TCA TCg TTC 3 reverse nested mcherry::moeabd pjwz6 5 CTT gga ggg TAC CgA gct CAg AAA ATg gct CAA ACA AAg CCg ATT gcc 3 cdh-3/zmp-1 extension, forward mcherry::plcδ PH paa173 5 TTC gag CgA Agg TCg CTT TTT ggt C 3 reverse mcherry::plcδ PH paa173 5 TTg AAA TCg AgT TgC AAg CgC gct CC 3 reverse nested mcherry::plcδ PH paa173 5 CTT gga ggg TAC CgA gct Cag AAA Atg gct CAA ACA AAg CCg ATT gcc 3 cdh-3/zmp-1 extension, forward mcherry paa64 5 TTC gag CgA Agg TCg CTT TTT ggt C 3 reverse mcherry paa64 5 TTg AAA TCg AgT TgC AAg CgC gct CC 3 reverse nested mcherry paa64 5 gca CCT CTg CTA TAT AAg AgA gtg gaa Tgg fos-1a extension, forward mcherry::plcδ PH paa173 CTC AAA CAA AgC CgA TTg CC 3 5 TTC gag CgA Agg TCg CTT TTT ggt C 3 reverse mcherry::plcδ PH paa173 5 TTg AAA TCg AgT TgC AAg CgC gct CC 3 reverse nested mcherry::plcδ PH paa173 5 TgT AAA ACg ACg gcc AgT 3 forward unc-6> promoter pvns-unc-6 5 TgT AAA ACg ACg gcc AgT 3 forward nested unc-6> promoter pvns-unc-6 5 gtt CTT CTC CTT TAC TgT TTg TgT gaa Agg gtg reverse unc-6> promoter pvns-unc-6 Taa AgT gga3 5 TCC ACT TTA CAC CCT TTC ACA CAA ACA TgA unc-6 promoter extension, venus::unc-6(δsp) pvns-unc-6 gta AAg gag AAg gag AAg AACTTT TCA CTg g 3 forward 5 CAg gaa ACA gct ATg ACC ATg 3 reverse venus::unc-6(δsp) pvns-unc-6 5 ATg ACC ATg ATT ACg CCA AgC gc 3 reverse nested venus::unc-6(δsp) pvns-unc-6
15 14 (Sherwood) Table S5. Extrachromosomal array and integrated strain generation Ex Designation Is Designation PCR Fusion created Injected Concentration Co-Injection Marker(s) qyex27 qyis23, qyis24, qyis25 cdh-3>mcherry::plcδ PH 0.01ng/µl unc-119 qyex39 qyis50 cdh-3>mcherry::moeabd 2ng/µl unc-119 qyex40 qyis66, qyis67 cdh-3>unc-40::gfp 2ng/µl unc myo-2>yfp qyex30 qyis37 zmp-1>unc-40::gfp 2ng/µl unc-119 qyex42 qyis61 cdh-3>unc-34::gfp 1ng/µl unc-119 qyex60 not integrated fos-1a>mcherry::plcδ PH 0.25ng/µl pha-1 qyex68 qyis7 zmp-1>mcherry 1.0ng/µl unc-119 qyex3 qyis17 pgk41(lam-1::gfp) 10ng/µl unc-119 qyex19 qyis27 ppr80(gfp::ced-10) 75ng/µl unc-119 qyex78 not integrated Venus::unc-6(ΔSP) 15ng/µl unc-119
Practical Bioinformatics
 5/2/2017 Dictionaries d i c t i o n a r y = { A : T, T : A, G : C, C : G } d i c t i o n a r y [ G ] d i c t i o n a r y [ N ] = N d i c t i o n a r y. h a s k e y ( C ) Dictionaries g e n e t i c C o
5/2/2017 Dictionaries d i c t i o n a r y = { A : T, T : A, G : C, C : G } d i c t i o n a r y [ G ] d i c t i o n a r y [ N ] = N d i c t i o n a r y. h a s k e y ( C ) Dictionaries g e n e t i c C o
SUPPORTING INFORMATION FOR. SEquence-Enabled Reassembly of β-lactamase (SEER-LAC): a Sensitive Method for the Detection of Double-Stranded DNA
 SUPPORTING INFORMATION FOR SEquence-Enabled Reassembly of β-lactamase (SEER-LAC): a Sensitive Method for the Detection of Double-Stranded DNA Aik T. Ooi, Cliff I. Stains, Indraneel Ghosh *, David J. Segal
SUPPORTING INFORMATION FOR SEquence-Enabled Reassembly of β-lactamase (SEER-LAC): a Sensitive Method for the Detection of Double-Stranded DNA Aik T. Ooi, Cliff I. Stains, Indraneel Ghosh *, David J. Segal
High throughput near infrared screening discovers DNA-templated silver clusters with peak fluorescence beyond 950 nm
 Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2018 High throughput near infrared screening discovers DNA-templated silver clusters with peak fluorescence
Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2018 High throughput near infrared screening discovers DNA-templated silver clusters with peak fluorescence
Supplemental data. Pommerrenig et al. (2011). Plant Cell /tpc
 Supplemental Figure 1. Prediction of phloem-specific MTK1 expression in Arabidopsis shoots and roots. The images and the corresponding numbers showing absolute (A) or relative expression levels (B) of
Supplemental Figure 1. Prediction of phloem-specific MTK1 expression in Arabidopsis shoots and roots. The images and the corresponding numbers showing absolute (A) or relative expression levels (B) of
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Zn 2+ -binding sites in USP18. (a) The two molecules of USP18 present in the asymmetric unit are shown. Chain A is shown in blue, chain B in green. Bound Zn 2+ ions are shown as
Supplementary Figure 1 Zn 2+ -binding sites in USP18. (a) The two molecules of USP18 present in the asymmetric unit are shown. Chain A is shown in blue, chain B in green. Bound Zn 2+ ions are shown as
SEQUENCE ALIGNMENT BACKGROUND: BIOINFORMATICS. Prokaryotes and Eukaryotes. DNA and RNA
 SEQUENCE ALIGNMENT BACKGROUND: BIOINFORMATICS 1 Prokaryotes and Eukaryotes 2 DNA and RNA 3 4 Double helix structure Codons Codons are triplets of bases from the RNA sequence. Each triplet defines an amino-acid.
SEQUENCE ALIGNMENT BACKGROUND: BIOINFORMATICS 1 Prokaryotes and Eukaryotes 2 DNA and RNA 3 4 Double helix structure Codons Codons are triplets of bases from the RNA sequence. Each triplet defines an amino-acid.
Clay Carter. Department of Biology. QuickTime and a TIFF (Uncompressed) decompressor are needed to see this picture.
 QuickTime and a TIFF (Uncompressed) decompressor are needed to see this picture. Clay Carter Department of Biology QuickTime and a TIFF (LZW) decompressor are needed to see this picture. Ornamental tobacco
QuickTime and a TIFF (Uncompressed) decompressor are needed to see this picture. Clay Carter Department of Biology QuickTime and a TIFF (LZW) decompressor are needed to see this picture. Ornamental tobacco
Electronic supplementary material
 Applied Microbiology and Biotechnology Electronic supplementary material A family of AA9 lytic polysaccharide monooxygenases in Aspergillus nidulans is differentially regulated by multiple substrates and
Applied Microbiology and Biotechnology Electronic supplementary material A family of AA9 lytic polysaccharide monooxygenases in Aspergillus nidulans is differentially regulated by multiple substrates and
SSR ( ) Vol. 48 No ( Microsatellite marker) ( Simple sequence repeat,ssr),
 48 3 () Vol. 48 No. 3 2009 5 Journal of Xiamen University (Nat ural Science) May 2009 SSR,,,, 3 (, 361005) : SSR. 21 516,410. 60 %96. 7 %. (),(Between2groups linkage method),.,, 11 (),. 12,. (, ), : 0.
48 3 () Vol. 48 No. 3 2009 5 Journal of Xiamen University (Nat ural Science) May 2009 SSR,,,, 3 (, 361005) : SSR. 21 516,410. 60 %96. 7 %. (),(Between2groups linkage method),.,, 11 (),. 12,. (, ), : 0.
Supplementary Information for
 Supplementary Information for Evolutionary conservation of codon optimality reveals hidden signatures of co-translational folding Sebastian Pechmann & Judith Frydman Department of Biology and BioX, Stanford
Supplementary Information for Evolutionary conservation of codon optimality reveals hidden signatures of co-translational folding Sebastian Pechmann & Judith Frydman Department of Biology and BioX, Stanford
TM1 TM2 TM3 TM4 TM5 TM6 TM bp
 a 467 bp 1 482 2 93 3 321 4 7 281 6 21 7 66 8 176 19 12 13 212 113 16 8 b ATG TCA GGA CAT GTA ATG GAG GAA TGT GTA GTT CAC GGT ACG TTA GCG GCA GTA TTG CGT TTA ATG GGC GTA GTG M S G H V M E E C V V H G T
a 467 bp 1 482 2 93 3 321 4 7 281 6 21 7 66 8 176 19 12 13 212 113 16 8 b ATG TCA GGA CAT GTA ATG GAG GAA TGT GTA GTT CAC GGT ACG TTA GCG GCA GTA TTG CGT TTA ATG GGC GTA GTG M S G H V M E E C V V H G T
Advanced topics in bioinformatics
 Feinberg Graduate School of the Weizmann Institute of Science Advanced topics in bioinformatics Shmuel Pietrokovski & Eitan Rubin Spring 2003 Course WWW site: http://bioinformatics.weizmann.ac.il/courses/atib
Feinberg Graduate School of the Weizmann Institute of Science Advanced topics in bioinformatics Shmuel Pietrokovski & Eitan Rubin Spring 2003 Course WWW site: http://bioinformatics.weizmann.ac.il/courses/atib
Crick s early Hypothesis Revisited
 Crick s early Hypothesis Revisited Or The Existence of a Universal Coding Frame Ryan Rossi, Jean-Louis Lassez and Axel Bernal UPenn Center for Bioinformatics BIOINFORMATICS The application of computer
Crick s early Hypothesis Revisited Or The Existence of a Universal Coding Frame Ryan Rossi, Jean-Louis Lassez and Axel Bernal UPenn Center for Bioinformatics BIOINFORMATICS The application of computer
SUPPLEMENTARY DATA - 1 -
 - 1 - SUPPLEMENTARY DATA Construction of B. subtilis rnpb complementation plasmids For complementation, the B. subtilis rnpb wild-type gene (rnpbwt) under control of its native rnpb promoter and terminator
- 1 - SUPPLEMENTARY DATA Construction of B. subtilis rnpb complementation plasmids For complementation, the B. subtilis rnpb wild-type gene (rnpbwt) under control of its native rnpb promoter and terminator
Supplemental Figure 1.
 A wt spoiiiaδ spoiiiahδ bofaδ B C D E spoiiiaδ, bofaδ Supplemental Figure 1. GFP-SpoIVFA is more mislocalized in the absence of both BofA and SpoIIIAH. Sporulation was induced by resuspension in wild-type
A wt spoiiiaδ spoiiiahδ bofaδ B C D E spoiiiaδ, bofaδ Supplemental Figure 1. GFP-SpoIVFA is more mislocalized in the absence of both BofA and SpoIIIAH. Sporulation was induced by resuspension in wild-type
Number-controlled spatial arrangement of gold nanoparticles with
 Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2016 Number-controlled spatial arrangement of gold nanoparticles with DNA dendrimers Ping Chen,*
Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2016 Number-controlled spatial arrangement of gold nanoparticles with DNA dendrimers Ping Chen,*
Characterization of Pathogenic Genes through Condensed Matrix Method, Case Study through Bacterial Zeta Toxin
 International Journal of Genetic Engineering and Biotechnology. ISSN 0974-3073 Volume 2, Number 1 (2011), pp. 109-114 International Research Publication House http://www.irphouse.com Characterization of
International Journal of Genetic Engineering and Biotechnology. ISSN 0974-3073 Volume 2, Number 1 (2011), pp. 109-114 International Research Publication House http://www.irphouse.com Characterization of
Supplementary Information
 Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2014 Directed self-assembly of genomic sequences into monomeric and polymeric branched DNA structures
Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2014 Directed self-assembly of genomic sequences into monomeric and polymeric branched DNA structures
NSCI Basic Properties of Life and The Biochemistry of Life on Earth
 NSCI 314 LIFE IN THE COSMOS 4 Basic Properties of Life and The Biochemistry of Life on Earth Dr. Karen Kolehmainen Department of Physics CSUSB http://physics.csusb.edu/~karen/ WHAT IS LIFE? HARD TO DEFINE,
NSCI 314 LIFE IN THE COSMOS 4 Basic Properties of Life and The Biochemistry of Life on Earth Dr. Karen Kolehmainen Department of Physics CSUSB http://physics.csusb.edu/~karen/ WHAT IS LIFE? HARD TO DEFINE,
Supplemental Table 1. Primers used for cloning and PCR amplification in this study
 Supplemental Table 1. Primers used for cloning and PCR amplification in this study Target Gene Primer sequence NATA1 (At2g393) forward GGG GAC AAG TTT GTA CAA AAA AGC AGG CTT CAT GGC GCC TCC AAC CGC AGC
Supplemental Table 1. Primers used for cloning and PCR amplification in this study Target Gene Primer sequence NATA1 (At2g393) forward GGG GAC AAG TTT GTA CAA AAA AGC AGG CTT CAT GGC GCC TCC AAC CGC AGC
Supporting Information
 Supporting Information T. Pellegrino 1,2,3,#, R. A. Sperling 1,#, A. P. Alivisatos 2, W. J. Parak 1,2,* 1 Center for Nanoscience, Ludwig Maximilians Universität München, München, Germany 2 Department of
Supporting Information T. Pellegrino 1,2,3,#, R. A. Sperling 1,#, A. P. Alivisatos 2, W. J. Parak 1,2,* 1 Center for Nanoscience, Ludwig Maximilians Universität München, München, Germany 2 Department of
Table S1. Primers and PCR conditions used in this paper Primers Sequence (5 3 ) Thermal conditions Reference Rhizobacteria 27F 1492R
 Table S1. Primers and PCR conditions used in this paper Primers Sequence (5 3 ) Thermal conditions Reference Rhizobacteria 27F 1492R AAC MGG ATT AGA TAC CCK G GGY TAC CTT GTT ACG ACT T Detection of Candidatus
Table S1. Primers and PCR conditions used in this paper Primers Sequence (5 3 ) Thermal conditions Reference Rhizobacteria 27F 1492R AAC MGG ATT AGA TAC CCK G GGY TAC CTT GTT ACG ACT T Detection of Candidatus
Regulatory Sequence Analysis. Sequence models (Bernoulli and Markov models)
 Regulatory Sequence Analysis Sequence models (Bernoulli and Markov models) 1 Why do we need random models? Any pattern discovery relies on an underlying model to estimate the random expectation. This model
Regulatory Sequence Analysis Sequence models (Bernoulli and Markov models) 1 Why do we need random models? Any pattern discovery relies on an underlying model to estimate the random expectation. This model
Supporting Information for. Initial Biochemical and Functional Evaluation of Murine Calprotectin Reveals Ca(II)-
 Supporting Information for Initial Biochemical and Functional Evaluation of Murine Calprotectin Reveals Ca(II)- Dependence and Its Ability to Chelate Multiple Nutrient Transition Metal Ions Rose C. Hadley,
Supporting Information for Initial Biochemical and Functional Evaluation of Murine Calprotectin Reveals Ca(II)- Dependence and Its Ability to Chelate Multiple Nutrient Transition Metal Ions Rose C. Hadley,
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION DOI:.38/NCHEM.246 Optimizing the specificity of nucleic acid hyridization David Yu Zhang, Sherry Xi Chen, and Peng Yin. Analytic framework and proe design 3.. Concentration-adjusted
SUPPLEMENTARY INFORMATION DOI:.38/NCHEM.246 Optimizing the specificity of nucleic acid hyridization David Yu Zhang, Sherry Xi Chen, and Peng Yin. Analytic framework and proe design 3.. Concentration-adjusted
Building a Multifunctional Aptamer-Based DNA Nanoassembly for Targeted Cancer Therapy
 Supporting Information Building a Multifunctional Aptamer-Based DNA Nanoassembly for Targeted Cancer Therapy Cuichen Wu,, Da Han,, Tao Chen,, Lu Peng, Guizhi Zhu,, Mingxu You,, Liping Qiu,, Kwame Sefah,
Supporting Information Building a Multifunctional Aptamer-Based DNA Nanoassembly for Targeted Cancer Therapy Cuichen Wu,, Da Han,, Tao Chen,, Lu Peng, Guizhi Zhu,, Mingxu You,, Liping Qiu,, Kwame Sefah,
6.047 / Computational Biology: Genomes, Networks, Evolution Fall 2008
 MIT OpenCourseWare http://ocw.mit.edu 6.047 / 6.878 Computational Biology: Genomes, Networks, Evolution Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu 6.047 / 6.878 Computational Biology: Genomes, Networks, Evolution Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
Protein Threading. Combinatorial optimization approach. Stefan Balev.
 Protein Threading Combinatorial optimization approach Stefan Balev Stefan.Balev@univ-lehavre.fr Laboratoire d informatique du Havre Université du Havre Stefan Balev Cours DEA 30/01/2004 p.1/42 Outline
Protein Threading Combinatorial optimization approach Stefan Balev Stefan.Balev@univ-lehavre.fr Laboratoire d informatique du Havre Université du Havre Stefan Balev Cours DEA 30/01/2004 p.1/42 Outline
SUPPLEMENTARY INFORMATION
 DOI:.8/NCHEM. Conditionally Fluorescent Molecular Probes for Detecting Single Base Changes in Double-stranded DNA Sherry Xi Chen, David Yu Zhang, Georg Seelig. Analytic framework and probe design.. Design
DOI:.8/NCHEM. Conditionally Fluorescent Molecular Probes for Detecting Single Base Changes in Double-stranded DNA Sherry Xi Chen, David Yu Zhang, Georg Seelig. Analytic framework and probe design.. Design
The role of the FliD C-terminal domain in pentamer formation and
 The role of the FliD C-terminal domain in pentamer formation and interaction with FliT Hee Jung Kim 1,2,*, Woongjae Yoo 3,*, Kyeong Sik Jin 4, Sangryeol Ryu 3,5 & Hyung Ho Lee 1, 1 Department of Chemistry,
The role of the FliD C-terminal domain in pentamer formation and interaction with FliT Hee Jung Kim 1,2,*, Woongjae Yoo 3,*, Kyeong Sik Jin 4, Sangryeol Ryu 3,5 & Hyung Ho Lee 1, 1 Department of Chemistry,
3. Evolution makes sense of homologies. 3. Evolution makes sense of homologies. 3. Evolution makes sense of homologies
 Richard Owen (1848) introduced the term Homology to refer to structural similarities among organisms. To Owen, these similarities indicated that organisms were created following a common plan or archetype.
Richard Owen (1848) introduced the term Homology to refer to structural similarities among organisms. To Owen, these similarities indicated that organisms were created following a common plan or archetype.
Evolvable Neural Networks for Time Series Prediction with Adaptive Learning Interval
 Evolvable Neural Networs for Time Series Prediction with Adaptive Learning Interval Dong-Woo Lee *, Seong G. Kong *, and Kwee-Bo Sim ** *Department of Electrical and Computer Engineering, The University
Evolvable Neural Networs for Time Series Prediction with Adaptive Learning Interval Dong-Woo Lee *, Seong G. Kong *, and Kwee-Bo Sim ** *Department of Electrical and Computer Engineering, The University
evoglow - express N kit distributed by Cat.#: FP product information broad host range vectors - gram negative bacteria
 evoglow - express N kit broad host range vectors - gram negative bacteria product information distributed by Cat.#: FP-21020 Content: Product Overview... 3 evoglow express N -kit... 3 The evoglow -Fluorescent
evoglow - express N kit broad host range vectors - gram negative bacteria product information distributed by Cat.#: FP-21020 Content: Product Overview... 3 evoglow express N -kit... 3 The evoglow -Fluorescent
AtTIL-P91V. AtTIL-P92V. AtTIL-P95V. AtTIL-P98V YFP-HPR
 Online Resource 1. Primers used to generate constructs AtTIL-P91V, AtTIL-P92V, AtTIL-P95V and AtTIL-P98V and YFP(HPR) using overlapping PCR. pentr/d- TOPO-AtTIL was used as template to generate the constructs
Online Resource 1. Primers used to generate constructs AtTIL-P91V, AtTIL-P92V, AtTIL-P95V and AtTIL-P98V and YFP(HPR) using overlapping PCR. pentr/d- TOPO-AtTIL was used as template to generate the constructs
Sex-Linked Inheritance in Macaque Monkeys: Implications for Effective Population Size and Dispersal to Sulawesi
 Supporting Information http://www.genetics.org/cgi/content/full/genetics.110.116228/dc1 Sex-Linked Inheritance in Macaque Monkeys: Implications for Effective Population Size and Dispersal to Sulawesi Ben
Supporting Information http://www.genetics.org/cgi/content/full/genetics.110.116228/dc1 Sex-Linked Inheritance in Macaque Monkeys: Implications for Effective Population Size and Dispersal to Sulawesi Ben
evoglow - express N kit Cat. No.: product information broad host range vectors - gram negative bacteria
 evoglow - express N kit broad host range vectors - gram negative bacteria product information Cat. No.: 2.1.020 evocatal GmbH 2 Content: Product Overview... 4 evoglow express N kit... 4 The evoglow Fluorescent
evoglow - express N kit broad host range vectors - gram negative bacteria product information Cat. No.: 2.1.020 evocatal GmbH 2 Content: Product Overview... 4 evoglow express N kit... 4 The evoglow Fluorescent
The Trigram and other Fundamental Philosophies
 The Trigram and other Fundamental Philosophies by Weimin Kwauk July 2012 The following offers a minimal introduction to the trigram and other Chinese fundamental philosophies. A trigram consists of three
The Trigram and other Fundamental Philosophies by Weimin Kwauk July 2012 The following offers a minimal introduction to the trigram and other Chinese fundamental philosophies. A trigram consists of three
ChemiScreen CaS Calcium Sensor Receptor Stable Cell Line
 PRODUCT DATASHEET ChemiScreen CaS Calcium Sensor Receptor Stable Cell Line CATALOG NUMBER: HTS137C CONTENTS: 2 vials of mycoplasma-free cells, 1 ml per vial. STORAGE: Vials are to be stored in liquid N
PRODUCT DATASHEET ChemiScreen CaS Calcium Sensor Receptor Stable Cell Line CATALOG NUMBER: HTS137C CONTENTS: 2 vials of mycoplasma-free cells, 1 ml per vial. STORAGE: Vials are to be stored in liquid N
Modelling and Analysis in Bioinformatics. Lecture 1: Genomic k-mer Statistics
 582746 Modelling and Analysis in Bioinformatics Lecture 1: Genomic k-mer Statistics Juha Kärkkäinen 06.09.2016 Outline Course introduction Genomic k-mers 1-Mers 2-Mers 3-Mers k-mers for Larger k Outline
582746 Modelling and Analysis in Bioinformatics Lecture 1: Genomic k-mer Statistics Juha Kärkkäinen 06.09.2016 Outline Course introduction Genomic k-mers 1-Mers 2-Mers 3-Mers k-mers for Larger k Outline
Re- engineering cellular physiology by rewiring high- level global regulatory genes
 Re- engineering cellular physiology by rewiring high- level global regulatory genes Stephen Fitzgerald 1,2,, Shane C Dillon 1, Tzu- Chiao Chao 2, Heather L Wiencko 3, Karsten Hokamp 3, Andrew DS Cameron
Re- engineering cellular physiology by rewiring high- level global regulatory genes Stephen Fitzgerald 1,2,, Shane C Dillon 1, Tzu- Chiao Chao 2, Heather L Wiencko 3, Karsten Hokamp 3, Andrew DS Cameron
part 3: analysis of natural selection pressure
 part 3: analysis of natural selection pressure markov models are good phenomenological codon models do have many benefits: o principled framework for statistical inference o avoiding ad hoc corrections
part 3: analysis of natural selection pressure markov models are good phenomenological codon models do have many benefits: o principled framework for statistical inference o avoiding ad hoc corrections
Evolutionary dynamics of abundant stop codon readthrough in Anopheles and Drosophila
 biorxiv preprint first posted online May. 3, 2016; doi: http://dx.doi.org/10.1101/051557. The copyright holder for this preprint (which was not peer-reviewed) is the author/funder. All rights reserved.
biorxiv preprint first posted online May. 3, 2016; doi: http://dx.doi.org/10.1101/051557. The copyright holder for this preprint (which was not peer-reviewed) is the author/funder. All rights reserved.
Near-instant surface-selective fluorogenic protein quantification using sulfonated
 Electronic Supplementary Material (ESI) for rganic & Biomolecular Chemistry. This journal is The Royal Society of Chemistry 2014 Supplemental nline Materials for ear-instant surface-selective fluorogenic
Electronic Supplementary Material (ESI) for rganic & Biomolecular Chemistry. This journal is The Royal Society of Chemistry 2014 Supplemental nline Materials for ear-instant surface-selective fluorogenic
Supplemental Figure 1. Phenotype of ProRGA:RGAd17 plants under long day
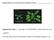 Supplemental Figure 1. Phenotype of ProRGA:RGAd17 plants under long day conditions. Photo was taken when the wild type plant started to bolt. Scale bar represents 1 cm. Supplemental Figure 2. Flowering
Supplemental Figure 1. Phenotype of ProRGA:RGAd17 plants under long day conditions. Photo was taken when the wild type plant started to bolt. Scale bar represents 1 cm. Supplemental Figure 2. Flowering
The 3 Genomic Numbers Discovery: How Our Genome Single-Stranded DNA Sequence Is Self-Designed as a Numerical Whole
 Applied Mathematics, 2013, 4, 37-53 http://dx.doi.org/10.4236/am.2013.410a2004 Published Online October 2013 (http://www.scirp.org/journal/am) The 3 Genomic Numbers Discovery: How Our Genome Single-Stranded
Applied Mathematics, 2013, 4, 37-53 http://dx.doi.org/10.4236/am.2013.410a2004 Published Online October 2013 (http://www.scirp.org/journal/am) The 3 Genomic Numbers Discovery: How Our Genome Single-Stranded
Codon Distribution in Error-Detecting Circular Codes
 life Article Codon Distribution in Error-Detecting Circular Codes Elena Fimmel, * and Lutz Strüngmann Institute for Mathematical Biology, Faculty of Computer Science, Mannheim University of Applied Sciences,
life Article Codon Distribution in Error-Detecting Circular Codes Elena Fimmel, * and Lutz Strüngmann Institute for Mathematical Biology, Faculty of Computer Science, Mannheim University of Applied Sciences,
Timing molecular motion and production with a synthetic transcriptional clock
 Timing molecular motion and production with a synthetic transcriptional clock Elisa Franco,1, Eike Friedrichs 2, Jongmin Kim 3, Ralf Jungmann 2, Richard Murray 1, Erik Winfree 3,4,5, and Friedrich C. Simmel
Timing molecular motion and production with a synthetic transcriptional clock Elisa Franco,1, Eike Friedrichs 2, Jongmin Kim 3, Ralf Jungmann 2, Richard Murray 1, Erik Winfree 3,4,5, and Friedrich C. Simmel
Roles for UNC-6/Netrin Signaling During Cell Invasion in C. Elegans. Joshua Willem Ziel. Department of Biology Duke University.
 Roles for UNC-6/Netrin Signaling During Cell Invasion in C. Elegans by Joshua Willem Ziel Department of Biology Duke University Date: Approved: David Sherwood, Supervisor Amy Bejsovec Daniel Kiehart Daniel
Roles for UNC-6/Netrin Signaling During Cell Invasion in C. Elegans by Joshua Willem Ziel Department of Biology Duke University Date: Approved: David Sherwood, Supervisor Amy Bejsovec Daniel Kiehart Daniel
Encoding of Amino Acids and Proteins from a Communications and Information Theoretic Perspective
 Jacobs University Bremen Encoding of Amino Acids and Proteins from a Communications and Information Theoretic Perspective Semester Project II By: Dawit Nigatu Supervisor: Prof. Dr. Werner Henkel Transmission
Jacobs University Bremen Encoding of Amino Acids and Proteins from a Communications and Information Theoretic Perspective Semester Project II By: Dawit Nigatu Supervisor: Prof. Dr. Werner Henkel Transmission
Chain-like assembly of gold nanoparticles on artificial DNA templates via Click Chemistry
 Electronic Supporting Information: Chain-like assembly of gold nanoparticles on artificial DNA templates via Click Chemistry Monika Fischler, Alla Sologubenko, Joachim Mayer, Guido Clever, Glenn Burley,
Electronic Supporting Information: Chain-like assembly of gold nanoparticles on artificial DNA templates via Click Chemistry Monika Fischler, Alla Sologubenko, Joachim Mayer, Guido Clever, Glenn Burley,
Why do more divergent sequences produce smaller nonsynonymous/synonymous
 Genetics: Early Online, published on June 21, 2013 as 10.1534/genetics.113.152025 Why do more divergent sequences produce smaller nonsynonymous/synonymous rate ratios in pairwise sequence comparisons?
Genetics: Early Online, published on June 21, 2013 as 10.1534/genetics.113.152025 Why do more divergent sequences produce smaller nonsynonymous/synonymous rate ratios in pairwise sequence comparisons?
Supplementary Information
 Supplementary Information Arginine-rhamnosylation as new strategy to activate translation elongation factor P Jürgen Lassak 1,2,*, Eva Keilhauer 3, Max Fürst 1,2, Kristin Wuichet 4, Julia Gödeke 5, Agata
Supplementary Information Arginine-rhamnosylation as new strategy to activate translation elongation factor P Jürgen Lassak 1,2,*, Eva Keilhauer 3, Max Fürst 1,2, Kristin Wuichet 4, Julia Gödeke 5, Agata
FliZ Is a Posttranslational Activator of FlhD 4 C 2 -Dependent Flagellar Gene Expression
 JOURNAL OF BACTERIOLOGY, July 2008, p. 4979 4988 Vol. 190, No. 14 0021-9193/08/$08.00 0 doi:10.1128/jb.01996-07 Copyright 2008, American Society for Microbiology. All Rights Reserved. FliZ Is a Posttranslational
JOURNAL OF BACTERIOLOGY, July 2008, p. 4979 4988 Vol. 190, No. 14 0021-9193/08/$08.00 0 doi:10.1128/jb.01996-07 Copyright 2008, American Society for Microbiology. All Rights Reserved. FliZ Is a Posttranslational
5- Semaphorin-Plexin-Neuropilin
 5- Semaphorin-Plexin-Neuropilin 1 SEMAPHORINS-PLEXINS-NEUROPILINS ligands receptors co-receptors semaphorins and their receptors are known signals for: -axon guidance -cell migration -morphogenesis -immune
5- Semaphorin-Plexin-Neuropilin 1 SEMAPHORINS-PLEXINS-NEUROPILINS ligands receptors co-receptors semaphorins and their receptors are known signals for: -axon guidance -cell migration -morphogenesis -immune
Pathways and Controls of N 2 O Production in Nitritation Anammox Biomass
 Supporting Information for Pathways and Controls of N 2 O Production in Nitritation Anammox Biomass Chun Ma, Marlene Mark Jensen, Barth F. Smets, Bo Thamdrup, Department of Biology, University of Southern
Supporting Information for Pathways and Controls of N 2 O Production in Nitritation Anammox Biomass Chun Ma, Marlene Mark Jensen, Barth F. Smets, Bo Thamdrup, Department of Biology, University of Southern
Insects act as vectors for a number of important diseases of
 pubs.acs.org/synthbio Novel Synthetic Medea Selfish Genetic Elements Drive Population Replacement in Drosophila; a Theoretical Exploration of Medea- Dependent Population Suppression Omar S. Abari,,# Chun-Hong
pubs.acs.org/synthbio Novel Synthetic Medea Selfish Genetic Elements Drive Population Replacement in Drosophila; a Theoretical Exploration of Medea- Dependent Population Suppression Omar S. Abari,,# Chun-Hong
Introduction to Molecular Phylogeny
 Introduction to Molecular Phylogeny Starting point: a set of homologous, aligned DNA or protein sequences Result of the process: a tree describing evolutionary relationships between studied sequences =
Introduction to Molecular Phylogeny Starting point: a set of homologous, aligned DNA or protein sequences Result of the process: a tree describing evolutionary relationships between studied sequences =
Integrin Acts Upstream of Netrin Signaling to Regulate Formation of the Anchor Cell s Invasive Membrane in C. elegans
 Article Integrin Acts Upstream of Netrin Signaling to Regulate Formation of the Anchor Cell s Invasive Membrane in C. elegans Elliott J. Hagedorn, 1 Hanako Yashiro, 2 Joshua W. Ziel, 1 Shinji Ihara, 1
Article Integrin Acts Upstream of Netrin Signaling to Regulate Formation of the Anchor Cell s Invasive Membrane in C. elegans Elliott J. Hagedorn, 1 Hanako Yashiro, 2 Joshua W. Ziel, 1 Shinji Ihara, 1
Supplementary Figure 1. Schematic of split-merger microfluidic device used to add transposase to template drops for fragmentation.
 Supplementary Figure 1. Schematic of split-merger microfluidic device used to add transposase to template drops for fragmentation. Inlets are labelled in blue, outlets are labelled in red, and static channels
Supplementary Figure 1. Schematic of split-merger microfluidic device used to add transposase to template drops for fragmentation. Inlets are labelled in blue, outlets are labelled in red, and static channels
Supplementary information. Porphyrin-Assisted Docking of a Thermophage Portal Protein into Lipid Bilayers: Nanopore Engineering and Characterization.
 Supplementary information Porphyrin-Assisted Docking of a Thermophage Portal Protein into Lipid Bilayers: Nanopore Engineering and Characterization. Benjamin Cressiot #, Sandra J. Greive #, Wei Si ^#,
Supplementary information Porphyrin-Assisted Docking of a Thermophage Portal Protein into Lipid Bilayers: Nanopore Engineering and Characterization. Benjamin Cressiot #, Sandra J. Greive #, Wei Si ^#,
Identification of a Locus Involved in the Utilization of Iron by Haemophilus influenzae
 INFECrION AND IMMUNITY, OCt. 1994, p. 4515-4525 0019-9567/94/$04.00+0 Copyright 1994, American Society for Microbiology Vol. 62, No. 10 Identification of a Locus Involved in the Utilization of Iron by
INFECrION AND IMMUNITY, OCt. 1994, p. 4515-4525 0019-9567/94/$04.00+0 Copyright 1994, American Society for Microbiology Vol. 62, No. 10 Identification of a Locus Involved in the Utilization of Iron by
ydci GTC TGT TTG AAC GCG GGC GAC TGG GCG CGC AAT TAA CGG TGT GTA GGC TGG AGC TGC TTC
 Table S1. DNA primers used in this study. Name ydci P1ydcIkd3 Sequence GTC TGT TTG AAC GCG GGC GAC TGG GCG CGC AAT TAA CGG TGT GTA GGC TGG AGC TGC TTC Kd3ydcIp2 lacz fusion YdcIendP1 YdcItrgP2 GAC AGC
Table S1. DNA primers used in this study. Name ydci P1ydcIkd3 Sequence GTC TGT TTG AAC GCG GGC GAC TGG GCG CGC AAT TAA CGG TGT GTA GGC TGG AGC TGC TTC Kd3ydcIp2 lacz fusion YdcIendP1 YdcItrgP2 GAC AGC
Supporting Information. An Electric Single-Molecule Hybridisation Detector for short DNA Fragments
 Supporting Information An Electric Single-Molecule Hybridisation Detector for short DNA Fragments A.Y.Y. Loh, 1 C.H. Burgess, 2 D.A. Tanase, 1 G. Ferrari, 3 M.A. Maclachlan, 2 A.E.G. Cass, 1 T. Albrecht*
Supporting Information An Electric Single-Molecule Hybridisation Detector for short DNA Fragments A.Y.Y. Loh, 1 C.H. Burgess, 2 D.A. Tanase, 1 G. Ferrari, 3 M.A. Maclachlan, 2 A.E.G. Cass, 1 T. Albrecht*
Axon Guidance. Multiple decision points along a growing axon s trajectory Different types of axon guidance cues:
 Axon Guidance Multiple decision points along a growing axon s trajectory Different types of axon guidance cues: Contact mediated - requires direct contact by growth cone Long range - growth cone responds
Axon Guidance Multiple decision points along a growing axon s trajectory Different types of axon guidance cues: Contact mediated - requires direct contact by growth cone Long range - growth cone responds
Nature Neuroscience: doi: /nn.2717
 Supplementary Fig. 1. Dendrite length is not secondary to body length. Dendrite growth proceeds independently of the rate of body growth and decreases in rate in adults. n 20 on dendrite measurement, n
Supplementary Fig. 1. Dendrite length is not secondary to body length. Dendrite growth proceeds independently of the rate of body growth and decreases in rate in adults. n 20 on dendrite measurement, n
Symmetry Studies. Marlos A. G. Viana
 Symmetry Studies Marlos A. G. Viana aaa aac aag aat caa cac cag cat aca acc acg act cca ccc ccg cct aga agc agg agt cga cgc cgg cgt ata atc atg att cta ctc ctg ctt gaa gac gag gat taa tac tag tat gca gcc
Symmetry Studies Marlos A. G. Viana aaa aac aag aat caa cac cag cat aca acc acg act cca ccc ccg cct aga agc agg agt cga cgc cgg cgt ata atc atg att cta ctc ctg ctt gaa gac gag gat taa tac tag tat gca gcc
The Cell Cycle & Cell Division. Cell Function Cell Cycle. What does the cell do = cell physiology:
 Cell Function 2404 What does the cell do = cell physiology: 1. Cell Cycle & Cell Division 2. Membrane Transport 3. Secretion 4. Membrane Potential 5. Metabolism 6. Cellular Interactions 7. Cellular Control:
Cell Function 2404 What does the cell do = cell physiology: 1. Cell Cycle & Cell Division 2. Membrane Transport 3. Secretion 4. Membrane Potential 5. Metabolism 6. Cellular Interactions 7. Cellular Control:
Biosynthesis of Bacterial Glycogen: Primary Structure of Salmonella typhimurium ADPglucose Synthetase as Deduced from the
 JOURNAL OF BACTERIOLOGY, Sept. 1987, p. 4355-4360 0021-9193/87/094355-06$02.00/0 Copyright X) 1987, American Society for Microbiology Vol. 169, No. 9 Biosynthesis of Bacterial Glycogen: Primary Structure
JOURNAL OF BACTERIOLOGY, Sept. 1987, p. 4355-4360 0021-9193/87/094355-06$02.00/0 Copyright X) 1987, American Society for Microbiology Vol. 169, No. 9 Biosynthesis of Bacterial Glycogen: Primary Structure
THE MATHEMATICAL STRUCTURE OF THE GENETIC CODE: A TOOL FOR INQUIRING ON THE ORIGIN OF LIFE
 STATISTICA, anno LXIX, n. 2 3, 2009 THE MATHEMATICAL STRUCTURE OF THE GENETIC CODE: A TOOL FOR INQUIRING ON THE ORIGIN OF LIFE Diego Luis Gonzalez CNR-IMM, Bologna Section, Via Gobetti 101, I-40129, Bologna,
STATISTICA, anno LXIX, n. 2 3, 2009 THE MATHEMATICAL STRUCTURE OF THE GENETIC CODE: A TOOL FOR INQUIRING ON THE ORIGIN OF LIFE Diego Luis Gonzalez CNR-IMM, Bologna Section, Via Gobetti 101, I-40129, Bologna,
160, and 220 bases, respectively, shorter than pbr322/hag93. (data not shown). The DNA sequence of approximately 100 bases of each
 JOURNAL OF BACTEROLOGY, JUlY 1988, p. 3305-3309 0021-9193/88/073305-05$02.00/0 Copyright 1988, American Society for Microbiology Vol. 170, No. 7 Construction of a Minimum-Size Functional Flagellin of Escherichia
JOURNAL OF BACTEROLOGY, JUlY 1988, p. 3305-3309 0021-9193/88/073305-05$02.00/0 Copyright 1988, American Society for Microbiology Vol. 170, No. 7 Construction of a Minimum-Size Functional Flagellin of Escherichia
A functional homologue of goosecoid in Drosophila
 Development 122, 1641-1650 (1996) Printed in Great Britain The Company of Biologists Limited 1996 DEV1069 1641 A functional homologue of goosecoid in Drosophila Anne Goriely 1, Michael Stella 1, Catherine
Development 122, 1641-1650 (1996) Printed in Great Britain The Company of Biologists Limited 1996 DEV1069 1641 A functional homologue of goosecoid in Drosophila Anne Goriely 1, Michael Stella 1, Catherine
Nature Methods: doi: /nmeth Supplementary Figure 1
 Supplementary Figure 1 Two strategies for application of LOVTRAP. The LOV domain is never entirely open or closed, but rather is in equilibria that favor the open or closed forms in the light and dark,
Supplementary Figure 1 Two strategies for application of LOVTRAP. The LOV domain is never entirely open or closed, but rather is in equilibria that favor the open or closed forms in the light and dark,
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3267 Supplementary Figure 1 A group of genes required for formation or orientation of annular F-actin bundles and aecm ridges: RNAi phenotypes and their validation by standard mutations.
DOI: 10.1038/ncb3267 Supplementary Figure 1 A group of genes required for formation or orientation of annular F-actin bundles and aecm ridges: RNAi phenotypes and their validation by standard mutations.
Using algebraic geometry for phylogenetic reconstruction
 Using algebraic geometry for phylogenetic reconstruction Marta Casanellas i Rius (joint work with Jesús Fernández-Sánchez) Departament de Matemàtica Aplicada I Universitat Politècnica de Catalunya IMA
Using algebraic geometry for phylogenetic reconstruction Marta Casanellas i Rius (joint work with Jesús Fernández-Sánchez) Departament de Matemàtica Aplicada I Universitat Politècnica de Catalunya IMA
Motif Finding Algorithms. Sudarsan Padhy IIIT Bhubaneswar
 Motif Finding Algorithms Sudarsan Padhy IIIT Bhubaneswar Outline Gene Regulation Regulatory Motifs The Motif Finding Problem Brute Force Motif Finding Consensus and Pattern Branching: Greedy Motif Search
Motif Finding Algorithms Sudarsan Padhy IIIT Bhubaneswar Outline Gene Regulation Regulatory Motifs The Motif Finding Problem Brute Force Motif Finding Consensus and Pattern Branching: Greedy Motif Search
Control of Growth Cone Polarity, Microtubule Accumulation, and Protrusion by UNC- 6/Netrin and its Receptors in C. elegans
 Genetics: Early Online, published on July 25, 2018 as 10.1534/genetics.118.301234 1 2 3 4 5 6 7 8 9 10 11 12 Control of Growth Cone Polarity, Microtubule Accumulation, and Protrusion by UNC- 6/Netrin and
Genetics: Early Online, published on July 25, 2018 as 10.1534/genetics.118.301234 1 2 3 4 5 6 7 8 9 10 11 12 Control of Growth Cone Polarity, Microtubule Accumulation, and Protrusion by UNC- 6/Netrin and
It is the author's version of the article accepted for publication in the journal "Biosystems" on 03/10/2015.
 It is the author's version of the article accepted for publication in the journal "Biosystems" on 03/10/2015. The system-resonance approach in modeling genetic structures Sergey V. Petoukhov Institute
It is the author's version of the article accepted for publication in the journal "Biosystems" on 03/10/2015. The system-resonance approach in modeling genetic structures Sergey V. Petoukhov Institute
Supplemental Figure 1. Differences in amino acid composition between the paralogous copies Os MADS17 and Os MADS6.
 Supplemental Data. Reinheimer and Kellogg (2009). Evolution of AGL6-like MADSbox genes in grasses (Poaceae): ovule expression is ancient and palea expression is new Supplemental Figure 1. Differences in
Supplemental Data. Reinheimer and Kellogg (2009). Evolution of AGL6-like MADSbox genes in grasses (Poaceae): ovule expression is ancient and palea expression is new Supplemental Figure 1. Differences in
How DNA barcoding can be more effective in microalgae. identification: a case of cryptic diversity revelation in Scenedesmus
 How DNA barcoding can be more effective in microalgae identification: a case of cryptic diversity revelation in Scenedesmus (Chlorophyceae) Shanmei Zou, Cong Fei, Chun Wang, Zhan Gao, Yachao Bao, Meilin
How DNA barcoding can be more effective in microalgae identification: a case of cryptic diversity revelation in Scenedesmus (Chlorophyceae) Shanmei Zou, Cong Fei, Chun Wang, Zhan Gao, Yachao Bao, Meilin
Chemical Biology on Genomic DNA: minimizing PCR bias. Electronic Supplementary Information (ESI) for Chemical Communications
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 Chemical Biology on Genomic DA: minimizing PCR bias Gordon R. McInroy, Eun-Ang Raiber, & Shankar
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 Chemical Biology on Genomic DA: minimizing PCR bias Gordon R. McInroy, Eun-Ang Raiber, & Shankar
Adam J. Schindler, L. Ryan Baugh, David R. Sherwood* Abstract. Introduction
 Identification of Late Larval Stage Developmental Checkpoints in Caenorhabditis elegans Regulated by Insulin/IGF and Steroid Hormone Signaling Pathways Adam J. Schindler, L. Ryan Baugh, David R. Sherwood*
Identification of Late Larval Stage Developmental Checkpoints in Caenorhabditis elegans Regulated by Insulin/IGF and Steroid Hormone Signaling Pathways Adam J. Schindler, L. Ryan Baugh, David R. Sherwood*
Evidence for RNA editing in mitochondria of all major groups of
 Proc. Natl. Acad. Sci. USA Vol. 91, pp. 629-633, January 1994 Plant Biology Evidence for RNA editing in mitochondria of all major groups of land plants except the Bryophyta RUDOLF HIESEL, BRUNO COMBETTES*,
Proc. Natl. Acad. Sci. USA Vol. 91, pp. 629-633, January 1994 Plant Biology Evidence for RNA editing in mitochondria of all major groups of land plants except the Bryophyta RUDOLF HIESEL, BRUNO COMBETTES*,
Glucosylglycerate phosphorylase, a novel enzyme specificity involved in compatible solute metabolism
 Supplementary Information for Glucosylglycerate phosphorylase, a novel enzyme specificity involved in compatible solute metabolism Jorick Franceus, Denise Pinel, Tom Desmet Corresponding author: Tom Desmet,
Supplementary Information for Glucosylglycerate phosphorylase, a novel enzyme specificity involved in compatible solute metabolism Jorick Franceus, Denise Pinel, Tom Desmet Corresponding author: Tom Desmet,
DNA sequence analysis of the imp UV protection and mutation operon of the plasmid TP110: identification of a third gene
 QD) 1990 Oxford University Press Nucleic Acids Research, Vol. 18, No. 17 5045 DNA sequence analysis of the imp UV protection and mutation operon of the plasmid TP110: identification of a third gene David
QD) 1990 Oxford University Press Nucleic Acids Research, Vol. 18, No. 17 5045 DNA sequence analysis of the imp UV protection and mutation operon of the plasmid TP110: identification of a third gene David
From DNA to protein, i.e. the central dogma
 From DNA to protein, i.e. the central dogma DNA RNA Protein Biochemistry, chapters1 5 and Chapters 29 31. Chapters 2 5 and 29 31 will be covered more in detail in other lectures. ph, chapter 1, will be
From DNA to protein, i.e. the central dogma DNA RNA Protein Biochemistry, chapters1 5 and Chapters 29 31. Chapters 2 5 and 29 31 will be covered more in detail in other lectures. ph, chapter 1, will be
Supplementary Table 3. Membrane/Signaling/Neural Genes of the DmSP. FBgn CG5265 acetyltransferase amino acid metabolism
 Supplementary Table 3 Membrane/Signaling/Neural Genes of the DmSP FlyBase ID Gene Name Molecular function summary Membrane Biological process summary FBgn0038486 CG5265 acetyltransferase amino acid metabolism
Supplementary Table 3 Membrane/Signaling/Neural Genes of the DmSP FlyBase ID Gene Name Molecular function summary Membrane Biological process summary FBgn0038486 CG5265 acetyltransferase amino acid metabolism
Supplemental data. Vos et al. (2008). The plant TPX2 protein regulates pro-spindle assembly before nuclear envelope breakdown.
 Supplemental data. Vos et al. (2008). The plant TPX2 protein regulates pro-spindle assembly before nuclear envelope breakdown. SUPPLEMENTAL FIGURE 1 ONLINE Xenopus laevis! Xenopus tropicalis! Danio rerio!
Supplemental data. Vos et al. (2008). The plant TPX2 protein regulates pro-spindle assembly before nuclear envelope breakdown. SUPPLEMENTAL FIGURE 1 ONLINE Xenopus laevis! Xenopus tropicalis! Danio rerio!
SPO71 encodes a developmental stage-specific partner for VPS13 in Saccharomcyes cerevisiae
 EC Accepts, published online ahead of print on 13 September 2013 Eukaryotic Cell doi:10.1128/ec.00239-13 Copyright 2013, American Society for Microbiology. All Rights Reserved. 1 2 3 4 5 6 7 8 9 10 11
EC Accepts, published online ahead of print on 13 September 2013 Eukaryotic Cell doi:10.1128/ec.00239-13 Copyright 2013, American Society for Microbiology. All Rights Reserved. 1 2 3 4 5 6 7 8 9 10 11
Axon guidance I. Paul Garrity March 15, /9.013
 Axon guidance I Paul Garrity March 15, 2004 7.68/9.013 Neuronal Wiring: Functional Framework of the Nervous System Stretch reflex circuit Early theories of axonogenesis Schwann: many neurons link to form
Axon guidance I Paul Garrity March 15, 2004 7.68/9.013 Neuronal Wiring: Functional Framework of the Nervous System Stretch reflex circuit Early theories of axonogenesis Schwann: many neurons link to form
Regulation of intracellular trafficking. by UNC-50 and the GARP complex. in C. elegans
 Regulation of intracellular trafficking by UNC-50 and the GARP complex in C. elegans PhD Thesis In partial fulfilment of the requirements for the degree Doctor of Philosophy (PhD) in the Neuroscience Program
Regulation of intracellular trafficking by UNC-50 and the GARP complex in C. elegans PhD Thesis In partial fulfilment of the requirements for the degree Doctor of Philosophy (PhD) in the Neuroscience Program
Supplementary Figure 1. Real time in vivo imaging of SG secretion. (a) SGs from Drosophila third instar larvae that express Sgs3-GFP (green) and
 Supplementary Figure 1. Real time in vivo imaging of SG secretion. (a) SGs from Drosophila third instar larvae that express Sgs3-GFP (green) and Lifeact-Ruby (red) were imaged in vivo to visualize secretion
Supplementary Figure 1. Real time in vivo imaging of SG secretion. (a) SGs from Drosophila third instar larvae that express Sgs3-GFP (green) and Lifeact-Ruby (red) were imaged in vivo to visualize secretion
Evidence for Evolution: Change Over Time (Make Up Assignment)
 Lesson 7.2 Evidence for Evolution: Change Over Time (Make Up Assignment) Name Date Period Key Terms Adaptive radiation Molecular Record Vestigial organ Homologous structure Strata Divergent evolution Evolution
Lesson 7.2 Evidence for Evolution: Change Over Time (Make Up Assignment) Name Date Period Key Terms Adaptive radiation Molecular Record Vestigial organ Homologous structure Strata Divergent evolution Evolution
Supplementary Figures
 Supplementary Figures Supplementary Fig. S1: Normal development and organization of the embryonic ventral nerve cord in Platynereis. (A) Life cycle of Platynereis dumerilii. (B-F) Axonal scaffolds and
Supplementary Figures Supplementary Fig. S1: Normal development and organization of the embryonic ventral nerve cord in Platynereis. (A) Life cycle of Platynereis dumerilii. (B-F) Axonal scaffolds and
Reduce, Reuse, Recycle: The tale of two Wnts and the lone C. elegans Syndecan, SDN-1
 Reduce, Reuse, Recycle: The tale of two Wnts and the lone C. elegans Syndecan, SDN-1 BY Copyright 2015 Samantha N. Hartin Submitted to the graduate degree program in Molecular Biosciences and the Graduate
Reduce, Reuse, Recycle: The tale of two Wnts and the lone C. elegans Syndecan, SDN-1 BY Copyright 2015 Samantha N. Hartin Submitted to the graduate degree program in Molecular Biosciences and the Graduate
RACK-1 Acts with Rac GTPase Signaling and UNC-115/ ablim in Caenorhabditis elegans Axon Pathfinding and Cell Migration
 RACK-1 Acts with Rac GTPase Signaling and UNC-115/ ablim in Caenorhabditis elegans Axon Pathfinding and Cell Migration Rafael S. Demarco, Erik A. Lundquist* Programs in Genetics and Molecular, Cellular,
RACK-1 Acts with Rac GTPase Signaling and UNC-115/ ablim in Caenorhabditis elegans Axon Pathfinding and Cell Migration Rafael S. Demarco, Erik A. Lundquist* Programs in Genetics and Molecular, Cellular,
DNA Barcoding Fishery Resources:
 DNA Barcoding Fishery Resources: A case study in Shandong Costal Water Shufang Liu Laboratory of Molecular ecology of fishery resources Yellow Sea Fisheries Research Institute (YSFRI) Chinese Academy of
DNA Barcoding Fishery Resources: A case study in Shandong Costal Water Shufang Liu Laboratory of Molecular ecology of fishery resources Yellow Sea Fisheries Research Institute (YSFRI) Chinese Academy of
C. elegans L1 cell adhesion molecule functions in axon guidance
 C. elegans L1 cell adhesion molecule functions in axon guidance Biorad Lihsia Chen Dept. of Genetics, Cell Biology & Development Developmental Biology Center C. elegans embryogenesis Goldstein lab, UNC-Chapel
C. elegans L1 cell adhesion molecule functions in axon guidance Biorad Lihsia Chen Dept. of Genetics, Cell Biology & Development Developmental Biology Center C. elegans embryogenesis Goldstein lab, UNC-Chapel
Three C. elegans Rac proteins and several alternative Rac regulators control axon guidance, cell migration and apoptotic cell phagocytosis
 Development 128, 4475-4488 (2001) Printed in Great Britain The Company of Biologists Limited 2001 DEV8810 4475 Three C. elegans Rac proteins and several alternative Rac regulators control axon guidance,
Development 128, 4475-4488 (2001) Printed in Great Britain The Company of Biologists Limited 2001 DEV8810 4475 Three C. elegans Rac proteins and several alternative Rac regulators control axon guidance,
Supporting Information
 Supporting Information Mixed DNA-functionalized Nanoparticle Probes for Surface Enhanced Raman Scattering-based Multiplex DNA Detection Zhiliang Zhang, a, b Yongqiang Wen,* a Ying Ma, a Jia Luo, a Lei
Supporting Information Mixed DNA-functionalized Nanoparticle Probes for Surface Enhanced Raman Scattering-based Multiplex DNA Detection Zhiliang Zhang, a, b Yongqiang Wen,* a Ying Ma, a Jia Luo, a Lei
Characterization of Multiple-Antimicrobial-Resistant Salmonella Serovars Isolated from Retail Meats
 APPLIED AND ENVIRONMENTAL MICROBIOLOGY, Jan. 2004, p. 1 7 Vol. 70, No. 1 0099-2240/04/$08.00 0 DOI: 10.1128/AEM.70.1.1 7.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved. Characterization
APPLIED AND ENVIRONMENTAL MICROBIOLOGY, Jan. 2004, p. 1 7 Vol. 70, No. 1 0099-2240/04/$08.00 0 DOI: 10.1128/AEM.70.1.1 7.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved. Characterization
