Use of Agilent Feature Extraction Software (v8.1) QC Report to Evaluate Microarray Performance
|
|
|
- Beverley Wheeler
- 5 years ago
- Views:
Transcription
1 Use of Agilent Feature Extraction Software (v8.1) QC Report to Evaluate Microarray Performance Anthea Dokidis Glenda Delenstarr Abstract The performance of the Agilent microarray system can now be evaluated using statistical results generated in the QC report, a new Feature Extraction (FE) output file (see FE User's Manual). There are three classes of metrics reported in the QC Report: (1) those that are independent of the platform; (2) those that rely on replicated probes; and (3) those that require both the coamplification and co-labeling of E1a spike-in targets, and the presence of E1a probes on the microarray. In this study, microarrays processed with Agilent's Two-Color Microarray- Based Gene Expression Analysis, version 4.0 protocol were experimentally manipulated to produce several images that can be used to examine the trends in the QC Report metrics. These conditions were wash artifacts, ozone exposure, and hybridization of crna signal generated from a degraded total RNA.
2 Introduction Figure 1 The performance of microarray target labeling, hybridization, and washes can now be evaluated and monitored using control E1a RNA targets supplied in the Agilent RNA Spike-In kit (P/N ) in combination with Agilent FE v8.1.these E1a control targets are in vitro synthesized, polyadenylated transcripts premixed at specific concentrations. In this way, when the E1a targets are amplified and labeled in the same reaction as the sample target RNA, they can be used to monitor the entire Agilent Two-Color Microarray-Based Gene Expression Analysis system for uniformity, sensitivity, and accuracy of log ratio signal across microarrays and between microarray experiments. In addition, the E1a targets are an invaluable tool in optimizing and troubleshooting experiments. To accommodate and make this E1a metric information easily available, the new version of the FE software (v8.1) produces an optional output file, the QC Report, which includes statistical results for both the E1a controls, for the non-control probes, and for the calculated backgrounds. These metrics are also available in the FE Stats table output, making it easy for the user to generate tracking charts (e.g. run charts ) of various metrics over time. This report shows examples of a few of these QC metrics, tracking the perturbations to the microarray system caused by manipulation of the experimental conditions. These conditions, described in the Experimental Methods, are: microarray wash artifacts, ozone exposure, and hybridization of crna generated from a degraded total RNA. Experimental Design Total RNA (30 arrays) Ozone (10 arrays) No wash 3 Exposed to ozone crna Targets Degraded RNA (10 arrays) Arrays Hybridizations (40 arrays) Array Washes (40 arrays) (10 arrays) Generate wash artifacts with wash3+10% wash2 Normal Wash (Control) (20 arrays) 10 Control arrays 10 Degraded RNA microarrays Total RNA from human adult and fetal heart were used to generate labeled crna. In addition, degraded total RNA (see Experimental Methods) from both adult and fetal heart were also used to generate labeled crna targets. Forty microarrays were hybridized the same day (Figure 1). Ten of these microarrays were hybridized with crna generated from the degraded total RNA for both adult and fetal heart. After hybridization all microarrays were washed with Gene Expression Wash buffer 1 and 2. Thirty microarrays including the normal wash control and the degraded total RNA microarrays were also washed a third time with the Agilent Stabilization and Drying solution and scanned for fluorescence signal. The remaining ten microarrays were washed with Gene Expression Wash Buffer 1 and 2 but not with the Agilent Stabilization and Drying solution. These microarrays were scanned and then removed from the scanner and were exposed to an ozone dose (see Experimental Methods). These microarrays were scanned, removed from the scanner, exposed to a second ozone dose and then scanned again. Scan Microarrays Microarrays: Whole Human Genome Oligo Microarray -G4112A Wash 1: Gene Expression Wash Buffer 1 (P/N ) at room temperature for 1 minute Wash 2: Gene Expression Wash Buffer 2 (P/N ) at 37ºC for 1 minute Wash 3: Stabilization and Drying Solution (P/N ), at room temperature for 30 seconds 2
3 Experimental Methods Degraded total RNA: Degraded total RNA was prepared by addding 0.001X RNase I 'A' from the Agilent Low RNA Input Fluorescent Linear Amplification Kit and leaving for 15 minutes at room temperature. After phenol chloroform purification and ethanol precipitation the degraded RNA was purified and spectrophotometrically quantitated while the level of degradation was determined using the RNA 6000 Nanoassay on the Agilent 2100 BioAnalyzer. (Figure 2) Fluorescent crna Targets 200 ng of total RNA from human adult and fetal heart and 12.5 pg of target E1a spike-in RNA were used to generate labeled Cyanine3- and Cyanine5-cRNA targets respectively using the Agilent Low RNA Input Fluorescent Linear Amplification Kit. Hybridization and washes 0.75ng of Cyanine3- and Cyanine5- labeled crna targets and 1X control targets in 1X hybridization buffer were incubated with the human microarrays G4112A, in a rotating oven at 65ºC for 17 hours. The microarrays were washed with Gene Expression Wash Buffer 1 at room temperature for 1 minute followed by a second wash of Gene Expression Wash Buffer 2 at 37ºC for 1 minute. The third wash consisted of Agilent Stabilization and Drying solution for 30 seconds at room temperature. Figure 2: Total RNA from Adult Heart: RNAse Treatment Bioanalyzer Electropherogram Adult Heart Total RNA (No RNAse) Adult Heart Total RNA (treated with 0.001X RNAse I 'A') Microarrays with wash artifacts were generated by washing hybridized microarrays with the Agilent Stabilization and Drying solution which has a 10% added volume of the Gene Expression Wash Buffer 2 for 30 seconds at room temperature. Ozone Exposure Hybridized Microarrays were washed with the first and second wash as described above but not with the stabilization and drying solution. The microarrays were then scanned on the Agilent microarray scanner to obtain a signal baseline. The microarrays were removed from the scanner and were exposed to 50 ppb of ozone for 1 minute while they were still in the scanner B-type slide holder. After this first dose of ozone the microarrays were scanned to obtain a microarray image for the fist ozone dose. The microarrays were removed and then exposed to a second ozone dose of 50 ppb for 1 minute and scanned again to produce microarray images for the second ozone dose. 4,000 2,000 1,000 RNA Ladder No RNase RNase 0.001x Adult Heart Total RNA Treatments Bioanalyzer Gel Image 3
4 Microarray images Figure 3: Log Scale Control, normal microarray Microarray with wash artifacts Microarray with degraded total RNA 4
5 Figure 4: Log Scale Microarray before ozone dose Same microarray after first ozone dose Same microarray after third ozone dose 5
6 Figure 5: Linear Scale Microarray before ozone doses Microarray after first ozone dose Microarray after third dose of ozone 6
7 Trends Observed in QC Report Metric Results To examine and visualize these trends, the QC metric in question is plotted on a graph for each microarray for all of the experimental conditions tested. Signal Dynamic range Net signal statistics can be a good indication of the signal dynamic range for a specific hybridized target. It should be noted that the net signal is independent of the Agilent scanner version. It is calculated as the (MeanSignal - scanner offset) and is also shown as a Features data column in the text output of the Feature Extraction v8.1. In Figures 6 and 7, the green and red net signal intensity at the 99th percentile of all non-control probes are plotted against the microarray experimental condition. From these figures, it is apparent that the microarrays hybridized with the crnas from the degraded total RNA (yellow symbols) show a dramatic decline in their net signal for both channels. A decrease in signal is also seen in the microarrays with the wash artifacts (light blue symbols). Microarrays with exposure to ozone clearly exhibit a significant effect in the red channel but not in the green channel, for both ozone doses (green and black symbols). Background Signal Figures 8 and 9 show the standard deviation for the net signal of the microarray negative controls for both channels. The microarrays with wash artifacts show a higher and wider range of standard deviations in their net signal distribution, as expected, as compared to the control microarrays. These negative control statistics are not shown on the QC Report, but are available in the Stats table output. The standard deviations of the local background inliers are shown on the QC Report. These SD's also increased on the microarrays with wash artifacts compared to control microarrays (data not shown). In comparison, the background standard deviations from the microarrays from the other conditions are similar to the range seen with control microarrays. Microarray Performance Uniformity The average of the signal to noise ratios (S/N) of the hybridized E1a probes can be used as one of the metrics for the microarray uniformity and indicate the amount of up or down regulated differential expression. Each of the ten different E1a probes is present with 30 replicate features on the microarray used for these experiments. An average and standard deviation (SD) of the log ratio for each of the ten E1a probes is calculated. Then, for each E1a probe, the S/N is calculated as the absolute (Avg_LogRatio/ SD_LogRatio). The average S/N across these ten S/N measurements is reported as a QC metric. A high S/N represents differential expression that is reproducible (e.g. a S/N > 3 is commonly used as a significance threshold). In Figure 10, the average of the S/N of the E1a probes is plotted for each microarray in each of the experimental conditions. The E1a Figure 6 Figure 7 Green net Signal At 99 th Percentile for Non-Control Probes Red Net Signal At 99 th Percentile for Non-Control Probes Legend Control Degraded Total RNA Before Ozone Doses Ozone First Dose Ozone Second Dose 7
8 hybridized to the microarrays with the wash artifacts and to the ozone treated microarrays are considerably closer to zero (less differential expression and/or poorly reproducible log ratios) than the control normal microarrays. This indicates that this metric can be used to show relative changes in the microarray uniformity within microarrays. Note that the microarrays that were scanned before exposure to ozone were not washed with the stabilization and Drying solution and they yield a wider variation of S/N ratios as compared to the control microarrays. Additionally, the E1a S/N's are unaffected by the degradation of the crna, as expected. Thus, this metric can be used to differentiate between problems that affect only the sample RNA from other types of problems that affect the entire sample, E1a amplification, and labeling reactions. Figure 8 Std Dev for Green Net Signal of Negative Controls Microarray Reproducibility The median percent coefficient of variation (%CV) of the background subtracted signals for the replicate features is used as a measure of the intra-array signal reproducibility. The %CV is calculated for each group of replicated probes, as (SD_Signals/Avg_Signals)*100%. The median of this group of %CV's is reported in the QC Report. A lower %CV indicates better reproducibility of signal intensity and hybridization uniformity. In Figures 11-14, the %CV of the background subtracted signal in both channels is shown for the non-control probes and the control E1a probes. Clearly, the %CV of the background subtracted signals for the non-control probes in the microarrays with the wash artifacts is increased in both channels compared to all the other microarrays (Figures 11 and 12). The microarrays with the wash artifacts are not expected to show a reproducible signal across the replicate probes within the microarrays, which is observed with this metric as expected. Figure 9 Std Dev for Red Net Signal of Negative Controls This microarray trend in the slides with wash artifacts is also observed in the %CV of the background subtracted signal for the E1a probes (Figures 13 and 14). Note that the %CV's of the E1a probes in the red channel are also sensitive to ozone (Figure 14). The %CV's seen with the E1a probes are higher, in this experiment, than the %CV's with the non-control probes. The E1a probes are designed to have a wide range of signal intensities across the 9 different probe sequences. The spread observed in the %CV of the background subtracted signal for the E1a replicate probes reflects this signal range; that is, higher %CV's are generally obtained from the E1a probes with the lower signal ranges. This relation is also shown as a plot on the QC Report: E1a %CV's vs. average background subtracted signal (not shown). Legend Control Degraded Total RNA Before Ozone Doses Ozone First Dose Ozone Second Dose 8
9 Figure 10 E1A S/N Log Ratio Legend Control Degraded Total RNA Before Ozone Doses Ozone First Dose Ozone Second Dose Figure 11 Green Non-Control Median %CV Background Subtracted Signal Figure 12 Red Non-Control Median %CV Background Subtracted Signal 9
10 Figure 13 Green E1A Median %CV Background Subtracted Signal Figure 14 Red E1A Median %CV Background Subtracted Signal Discussion From the results of this study, it is seen that significant changes in some of the metrics calculated in the FE output QC report appear to be well correlated with certain microarray process defects. For example, in the Ozone-treated hybridized microarrays, the net signal intensity at the 99th percentile of all non-control probes is greatly affected in the red channel (Figure 7) but not affected in the green channel (Figure 6). Similarly, the log ratio for the E1a probes on these microarrays is seen to be closer to zero, compressed and/or non-reproducible, (Figure 10), compared to the control microarrays indicating a change in their signal intensities and an increase in noise. Furthermore, the ozone results also show an increase in the average %CV for the E1a probes in the red channel (and not green channel), indicating a decrease in reproducibility due to the effect of ozone on the Cyanine5-labeled targets (Figure 14). Microarrays exhibiting wash artifacts have the highest number of non feature uniform outliers (graph not shown) and a lower signal dynamic range (Figures 6 and 7). This decrease in performance trend continues in the microarray uniformity metric for E1a controls (Figure 10). We expect the microarrays with wash artifacts to show less signal and log ratio uniformity within microarrays and also across microarrays of this experimental condition (Figure 11 and 12). Additionally, the negative controls on the microarrays with wash artifacts have an increased net signal standard deviation that shows a dramatically wider spread than the control microarrays (Figures 8 and 9). This is particularly evident in the green channel. Interestingly, the microarrays hybridized with crna generated from a degraded sample total RNA (degraded human adult and fetal heart RNA) appear to have greatly decreased signal dynamic ranges in both the red and the green channels, as compared to microarrays from all other conditions (Figure 6). In contrast, the reproducibility of signal and log ratios with the E1a controls is unaffected by the crna degradation condition, as can be with the E1a signal %CV metrics (Figures 13 and 14) and the E1a S/N of log ratio metric (Figures 10). This is expected since the E1a spike-ins were added post sample RNA degradation. Conclusion Legend Control Degraded Total RNA Before Ozone Doses Ozone First Dose Ozone Second Dose In this study, it is demonstrated that the QC report generated using FE 8.1 and the E1A targets is capable of identifying microarray images which exhibit visible hybridized microarray problems such as wash artifacts as well as microarrays with non-visible problems such as exposure to Ozone or hybridization with crna from degraded Total RNA. The ability to detect deviations from normal trends reported in the QC Report enable researchers to identify results which may require closer inspection. The trends of the QC Report metrics observed in this study were as expected for each particular microarray artifact condition proving that it is numerically possible to track the performance of the microarray processes, such as target preparation, microarray hybridization, and washes using the QC Report statistics. 10
11 Agilent Technologies, Inc Printed in the U.S.A. Agilent Technologies Bioresearch Solutions Unit 3500 Deer Creek Road Palo Alto, CA Agilent Gene Expression Microarrays Website Agilent Lab-on-a-Chip Website Information, descriptions and specifications are subject to change without notice. Please register online with Agilent to recieve product updates at: EN July 20, 2005
Performance of the RNA and the High Sensitivity RNA ScreenTape Assay for the Agilent 4200 TapeStation System
 Performance of the RNA and the High Sensitivity RNA ScreenTape Assay for the Agilent TapeStation System Technical Overview Introduction The Agilent TapeStation system provides an automated system for fast
Performance of the RNA and the High Sensitivity RNA ScreenTape Assay for the Agilent TapeStation System Technical Overview Introduction The Agilent TapeStation system provides an automated system for fast
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
U.S. Patent No. 9,051,563 and other pending patents. Ver
 INSTRUCTION MANUAL Direct-zol 96 RNA Catalog Nos. R2054, R2055, R2056 & R2057 Highlights Quick, 96-well purification of high-quality (DNA-free) total RNA directly from TRIzol, TRI Reagent and all other
INSTRUCTION MANUAL Direct-zol 96 RNA Catalog Nos. R2054, R2055, R2056 & R2057 Highlights Quick, 96-well purification of high-quality (DNA-free) total RNA directly from TRIzol, TRI Reagent and all other
RNA Transport. R preps R preps
 RNA Transport R0527-00 5 preps R0527-01 50 preps July 2014 RNA Transport Table of Contents Introduction...2 Kit Contents/Storage and Stability...3 Protocol...4 Storage Procedure...4 Recovery Procedure...5
RNA Transport R0527-00 5 preps R0527-01 50 preps July 2014 RNA Transport Table of Contents Introduction...2 Kit Contents/Storage and Stability...3 Protocol...4 Storage Procedure...4 Recovery Procedure...5
RayBio Human IDUA ELISA Kit
 RayBio Human IDUA ELISA Kit Catalog #: ELH-IDUA User Manual Last revised April 15, 2016 Caution: Extraordinarily useful information enclosed ISO 13485 Certified 3607 Parkway Lane, Suite 100 Norcross, GA
RayBio Human IDUA ELISA Kit Catalog #: ELH-IDUA User Manual Last revised April 15, 2016 Caution: Extraordinarily useful information enclosed ISO 13485 Certified 3607 Parkway Lane, Suite 100 Norcross, GA
Easy Illumina Nextera DNA FLEX Library Preparation using the epmotion 5075t automated liquid handler
 WHITE PAPER No. 13 Easy Illumina Nextera DNA FLEX Library Preparation using the epmotion 5075t automated liquid handler Executive Summary Library preparation steps, including DNA extraction, quantification,
WHITE PAPER No. 13 Easy Illumina Nextera DNA FLEX Library Preparation using the epmotion 5075t automated liquid handler Executive Summary Library preparation steps, including DNA extraction, quantification,
SPOTTED cdna MICROARRAYS
 SPOTTED cdna MICROARRAYS Spot size: 50um - 150um SPOTTED cdna MICROARRAYS Compare the genetic expression in two samples of cells PRINT cdna from one gene on each spot SAMPLES cdna labelled red/green e.g.
SPOTTED cdna MICROARRAYS Spot size: 50um - 150um SPOTTED cdna MICROARRAYS Compare the genetic expression in two samples of cells PRINT cdna from one gene on each spot SAMPLES cdna labelled red/green e.g.
RayBio Human Thyroid Peroxidase ELISA Kit
 RayBio Human Thyroid Peroxidase ELISA Kit Catalog #: ELH-TPX User Manual Last revised July 6, 2017 Caution: Extraordinarily useful information enclosed ISO 13485 Certified 3607 Parkway Lane, Suite 100
RayBio Human Thyroid Peroxidase ELISA Kit Catalog #: ELH-TPX User Manual Last revised July 6, 2017 Caution: Extraordinarily useful information enclosed ISO 13485 Certified 3607 Parkway Lane, Suite 100
Quantification Strategies Using the High Sensitivity Protein 250 Assay for the Agilent 2100 Bioanalyzer. Technical Note. Abstract
 Quantification Strategies Using the High Sensitivity Protein 25 Assay for the Agilent 21 Bioanalyzer Technical Note.. 1. 1. 1..1.1 1. 1. 1... Abstract The Agilent High Sensitivity Protein 25 assay for
Quantification Strategies Using the High Sensitivity Protein 25 Assay for the Agilent 21 Bioanalyzer Technical Note.. 1. 1. 1..1.1 1. 1. 1... Abstract The Agilent High Sensitivity Protein 25 assay for
DEVELOP YOUR LAB, YOUR WAY
 Technical brochure DEVELOP YOUR LAB, YOUR WAY The FLOW Solution: Expand your potential today Redesign your lab with the FLOW Solution The FLOW Solution is a highly flexible modular, semi-automated data
Technical brochure DEVELOP YOUR LAB, YOUR WAY The FLOW Solution: Expand your potential today Redesign your lab with the FLOW Solution The FLOW Solution is a highly flexible modular, semi-automated data
RayBio Rat TNF-alpha ELISA Kit (For Lysates)
 RayBio Rat TNF-alpha ELISA Kit (For Lysates) Catalog #: ELR-TNFa-CL User Manual Last revised April 15, 2016 Caution: Extraordinarily useful information enclosed ISO 13485 Certified 3607 Parkway Lane, Suite
RayBio Rat TNF-alpha ELISA Kit (For Lysates) Catalog #: ELR-TNFa-CL User Manual Last revised April 15, 2016 Caution: Extraordinarily useful information enclosed ISO 13485 Certified 3607 Parkway Lane, Suite
cdna Microarray Analysis
 cdna Microarray Analysis with BioConductor packages Nolwenn Le Meur Copyright 2007 Outline Data acquisition Pre-processing Quality assessment Pre-processing background correction normalization summarization
cdna Microarray Analysis with BioConductor packages Nolwenn Le Meur Copyright 2007 Outline Data acquisition Pre-processing Quality assessment Pre-processing background correction normalization summarization
Data Sheet. Azide Cy5 RNA T7 Transcription Kit
 Cat. No. Size 1. Description PP-501-Cy5 10 reactions à 40 µl For in vitro use only Quality guaranteed for 12 months Store all components at -20 C. Avoid freeze and thaw cycles. DBCO-Sulfo-Cy5 must be stored
Cat. No. Size 1. Description PP-501-Cy5 10 reactions à 40 µl For in vitro use only Quality guaranteed for 12 months Store all components at -20 C. Avoid freeze and thaw cycles. DBCO-Sulfo-Cy5 must be stored
U.S. Patent No. 9,051,563 and other pending patents. Ver
 INSTRUCTION MANUAL Direct-zol RNA MiniPrep Catalog Nos. R050, R05, R05, & R053 Highlights Quick, spin column purification of high-quality (DNA-free) total RNA directly from TRIzol, TRI Reagent and other
INSTRUCTION MANUAL Direct-zol RNA MiniPrep Catalog Nos. R050, R05, R05, & R053 Highlights Quick, spin column purification of high-quality (DNA-free) total RNA directly from TRIzol, TRI Reagent and other
Amersham Cy3/HyPer5 reactive dye protocols for post labeling amino allyl-cdna and amino allyl-arna
 GE Healthcare Amersham Cy3/HyPer5 reactive dye protocols for post labeling amino allyl-cdna and amino allyl-arna For microarray applications Product booklet Codes: 28-9224-18 28-9224-19 Page finder 1.
GE Healthcare Amersham Cy3/HyPer5 reactive dye protocols for post labeling amino allyl-cdna and amino allyl-arna For microarray applications Product booklet Codes: 28-9224-18 28-9224-19 Page finder 1.
Mag-Bind Soil DNA Kit. M preps M preps M preps
 Mag-Bind Soil DNA Kit M5635-00 5 preps M5635-01 50 preps M5635-02 200 preps January 2013 Mag-Bind Soil DNA Kit Table of Contents Introduction and Overview...2 Kit Contents/Storage and Stability...3 Preparing
Mag-Bind Soil DNA Kit M5635-00 5 preps M5635-01 50 preps M5635-02 200 preps January 2013 Mag-Bind Soil DNA Kit Table of Contents Introduction and Overview...2 Kit Contents/Storage and Stability...3 Preparing
Globin-Zero Gold Kit
 Cat. No. GZG1206 (Contains 1 box of Cat. No. GZRR1306 and 1 box of Cat. No. MRZ116C) Cat. No. GZG1224 (Contains 1 box of Cat. No. GZRR1324 and 1 box of Cat. No. MRZ11124C) Connect with Epicentre on our
Cat. No. GZG1206 (Contains 1 box of Cat. No. GZRR1306 and 1 box of Cat. No. MRZ116C) Cat. No. GZG1224 (Contains 1 box of Cat. No. GZRR1324 and 1 box of Cat. No. MRZ11124C) Connect with Epicentre on our
Fast and simple fluorescent RNA quality assessment
 Fast and simple fluorescent RNA quality assessment Chris Vonnegut Presented at ASHG Laboratory CoLab Session October, The world leader in serving science RNA Quality and Quantity RNA Sample Isolation Cells
Fast and simple fluorescent RNA quality assessment Chris Vonnegut Presented at ASHG Laboratory CoLab Session October, The world leader in serving science RNA Quality and Quantity RNA Sample Isolation Cells
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Bradley Broom Department of Bioinformatics and Computational Biology UT M. D. Anderson Cancer Center kabagg@mdanderson.org bmbroom@mdanderson.org
GS01 0163 Analysis of Microarray Data Keith Baggerly and Bradley Broom Department of Bioinformatics and Computational Biology UT M. D. Anderson Cancer Center kabagg@mdanderson.org bmbroom@mdanderson.org
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
Instructor s Advance Preparation
 INSTRUCTOR'S MANUAL Instructor s Advance Preparation This protocol is designed for 80 workstations of 4 students. Each group will prepare a set of standards, a blank, and 2 milk samples (can be a blind
INSTRUCTOR'S MANUAL Instructor s Advance Preparation This protocol is designed for 80 workstations of 4 students. Each group will prepare a set of standards, a blank, and 2 milk samples (can be a blind
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
Product Name Quantity* Product No.
 Page 1 of 6 Lit.# ML002 Rev. 11/15/06 Label IT Nucleic Acid Labeling Kits Product Name Quantity* Product No. Label IT CX-Rhodamine Labeling Kit Full Size MIR 3100 Trial Size MIR 3125 Label IT Fluorescein
Page 1 of 6 Lit.# ML002 Rev. 11/15/06 Label IT Nucleic Acid Labeling Kits Product Name Quantity* Product No. Label IT CX-Rhodamine Labeling Kit Full Size MIR 3100 Trial Size MIR 3125 Label IT Fluorescein
RayBio Human VEGFR1 ELISA Kit
 RayBio Human VEGFR1 ELISA Kit Catalog #: ELH-VEGFR1 User Manual Last revised July 6, 2017 Caution: Extraordinarily useful information enclosed ISO 13485 Certified 3607 Parkway Lane, Suite 100 Norcross,
RayBio Human VEGFR1 ELISA Kit Catalog #: ELH-VEGFR1 User Manual Last revised July 6, 2017 Caution: Extraordinarily useful information enclosed ISO 13485 Certified 3607 Parkway Lane, Suite 100 Norcross,
RayBio Human LBP ELISA Kit
 RayBio Human LBP ELISA Kit Catalog #: ELH-LBP User Manual Last revised April 15, 2016 Caution: Extraordinarily useful information enclosed ISO 13485 Certified 3607 Parkway Lane, Suite 100 Norcross, GA
RayBio Human LBP ELISA Kit Catalog #: ELH-LBP User Manual Last revised April 15, 2016 Caution: Extraordinarily useful information enclosed ISO 13485 Certified 3607 Parkway Lane, Suite 100 Norcross, GA
Ribo-Zero Magnetic Gold Kit*
 Ribo-Zero Magnetic Gold Kit* (Epidemiology) Cat. No. MRZE706 (Contains 1 box of Cat. No. RZE1206 and 1 box of Cat. No. MRZ116C) Cat. No. MRZE724 (Contains 1 box of Cat. No. RZE1224 and 1 box of Cat. No.
Ribo-Zero Magnetic Gold Kit* (Epidemiology) Cat. No. MRZE706 (Contains 1 box of Cat. No. RZE1206 and 1 box of Cat. No. MRZ116C) Cat. No. MRZE724 (Contains 1 box of Cat. No. RZE1224 and 1 box of Cat. No.
RayBio Human CA-125 ELISA Kit
 RayBio Human CA-125 ELISA Kit Catalog #: ELH-CA125 User Manual Last revised April 15, 2016 Caution: Extraordinarily useful information enclosed ISO 13485 Certified 3607 Parkway Lane, Suite 100 Norcross,
RayBio Human CA-125 ELISA Kit Catalog #: ELH-CA125 User Manual Last revised April 15, 2016 Caution: Extraordinarily useful information enclosed ISO 13485 Certified 3607 Parkway Lane, Suite 100 Norcross,
RayBio Human Semaphorin 6D ELISA Kit
 RayBio Human Semaphorin 6D ELISA Kit Catalog #: ELH-SEMA6D User Manual Last revised February 9, 2017 Caution: Extraordinarily useful information enclosed ISO 13485 Certified 3607 Parkway Lane, Suite 100
RayBio Human Semaphorin 6D ELISA Kit Catalog #: ELH-SEMA6D User Manual Last revised February 9, 2017 Caution: Extraordinarily useful information enclosed ISO 13485 Certified 3607 Parkway Lane, Suite 100
Ribo-Zero Magnetic Kit*
 Ribo-Zero Magnetic Kit* (Bacteria) Cat. No. MRZMB126 (Contains 1 box of Cat. No. RZMB11086 and 1 box of Cat. No. MRZ116C) Cat. No. MRZB12424 24 Reactions (Contains 1 box of Cat. No. RZMB12324 and 1 box
Ribo-Zero Magnetic Kit* (Bacteria) Cat. No. MRZMB126 (Contains 1 box of Cat. No. RZMB11086 and 1 box of Cat. No. MRZ116C) Cat. No. MRZB12424 24 Reactions (Contains 1 box of Cat. No. RZMB12324 and 1 box
Biochip informatics-(i)
 Biochip informatics-(i) : biochip normalization & differential expression Ju Han Kim, M.D., Ph.D. SNUBI: SNUBiomedical Informatics http://www.snubi snubi.org/ Biochip Informatics - (I) Biochip basics Preprocessing
Biochip informatics-(i) : biochip normalization & differential expression Ju Han Kim, M.D., Ph.D. SNUBI: SNUBiomedical Informatics http://www.snubi snubi.org/ Biochip Informatics - (I) Biochip basics Preprocessing
RayBio Human B7H1/PD-L1 ELISA Kit
 RayBio Human B7H1/PD-L1 ELISA Kit Catalog #: ELH-B7H1 User Manual Last revised April 15, 2016 Caution: Extraordinarily useful information enclosed ISO 13485 Certified 3607 Parkway Lane, Suite 100 Norcross,
RayBio Human B7H1/PD-L1 ELISA Kit Catalog #: ELH-B7H1 User Manual Last revised April 15, 2016 Caution: Extraordinarily useful information enclosed ISO 13485 Certified 3607 Parkway Lane, Suite 100 Norcross,
Discovering Correlation in Data. Vinh Nguyen Research Fellow in Data Science Computing and Information Systems DMD 7.
 Discovering Correlation in Data Vinh Nguyen (vinh.nguyen@unimelb.edu.au) Research Fellow in Data Science Computing and Information Systems DMD 7.14 Discovering Correlation Why is correlation important?
Discovering Correlation in Data Vinh Nguyen (vinh.nguyen@unimelb.edu.au) Research Fellow in Data Science Computing and Information Systems DMD 7.14 Discovering Correlation Why is correlation important?
Evaluation of the performance of Amersham HyPer5 dye in comparative genomic hybridization microarray applications
 GE Healthcare Application note 28-9304-76 AA Nucleic acid labeling and detection Evaluation of the performance of Amersham HyPer5 dye in comparative genomic hybridization microarray applications Key words:
GE Healthcare Application note 28-9304-76 AA Nucleic acid labeling and detection Evaluation of the performance of Amersham HyPer5 dye in comparative genomic hybridization microarray applications Key words:
WELCOME!! LABORATORY MATH PERCENT CONCENTRATION. Things to do ASAP: Concepts to deal with:
 WELCOME!! Things to do ASAP: Read the course syllabus; information regarding testing, homework, lecture schedules, expectations and course objectives are all there Read the weekly overview; lecture objectives
WELCOME!! Things to do ASAP: Read the course syllabus; information regarding testing, homework, lecture schedules, expectations and course objectives are all there Read the weekly overview; lecture objectives
RNA Labeling Kit. User Manual
 RNA Labeling Kit User Manual RNA Labeling Kit The RNA Labeling Kit contains reagents to perform 10 transcription reactions (50 µl each) and 12 independent labeling reactions. Introduction and product description:
RNA Labeling Kit User Manual RNA Labeling Kit The RNA Labeling Kit contains reagents to perform 10 transcription reactions (50 µl each) and 12 independent labeling reactions. Introduction and product description:
Automated Illumina TruSeq Stranded mrna library construction with the epmotion 5075t/TMX
 SHORT PROTOCOL No. 02 I November 2014 Automated Illumina TruSeq Stranded mrna library construction with the epmotion 5075t/TMX Introduction For the MiSeq and HiSeq next generation sequencing (NGS) systems,
SHORT PROTOCOL No. 02 I November 2014 Automated Illumina TruSeq Stranded mrna library construction with the epmotion 5075t/TMX Introduction For the MiSeq and HiSeq next generation sequencing (NGS) systems,
E.Z.N.A. MicroElute Clean-up Kits Table of Contents
 E.Z.N.A. MicroElute Clean-up Kits Table of Contents Introduction... 2 Kit Contents... 3 Preparing Reagents/Storage and Stability... 4 Guideline for Vacuum Manifold... 5 MicroElute Cycle-Pure - Spin Protocol...
E.Z.N.A. MicroElute Clean-up Kits Table of Contents Introduction... 2 Kit Contents... 3 Preparing Reagents/Storage and Stability... 4 Guideline for Vacuum Manifold... 5 MicroElute Cycle-Pure - Spin Protocol...
Lesson 11. Functional Genomics I: Microarray Analysis
 Lesson 11 Functional Genomics I: Microarray Analysis Transcription of DNA and translation of RNA vary with biological conditions 3 kinds of microarray platforms Spotted Array - 2 color - Pat Brown (Stanford)
Lesson 11 Functional Genomics I: Microarray Analysis Transcription of DNA and translation of RNA vary with biological conditions 3 kinds of microarray platforms Spotted Array - 2 color - Pat Brown (Stanford)
ELISA QUALITY ASSURANCE Analytical Phase
 LOGO ELISA QUALITY ASSURANCE Analytical Phase - DR. ALI MIRJALILI QUALITY ASSURANCE Dept. PISHTAZ TEB DIAGNOSTICS 01/02/92 Quality improvement congress Definitions: Quality Assurance Quality Control:-
LOGO ELISA QUALITY ASSURANCE Analytical Phase - DR. ALI MIRJALILI QUALITY ASSURANCE Dept. PISHTAZ TEB DIAGNOSTICS 01/02/92 Quality improvement congress Definitions: Quality Assurance Quality Control:-
Low-volume, High Throughput Workflow for Analysis of Nucleic Acid Samples for Biobanking
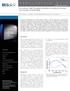 A p p l i c a t i o n N o t e Low-volume, High Throughput Workflow for Analysis of Nucleic Acid Samples for Biobanking Peter J. Brescia, Jr., Chris Wilson, and Peter Banks, BioTek Instruments, Inc., Winooski,
A p p l i c a t i o n N o t e Low-volume, High Throughput Workflow for Analysis of Nucleic Acid Samples for Biobanking Peter J. Brescia, Jr., Chris Wilson, and Peter Banks, BioTek Instruments, Inc., Winooski,
RayBio Mouse TNF-alpha ELISA Kit (For Lysates)
 RayBio Mouse TNF-alpha ELISA Kit (For Lysates) Catalog #: ELM-TNFa-CL User Manual Last revised September 15, 2017 Caution: Extraordinarily useful information enclosed ISO 13485 Certified 3607 Parkway Lane,
RayBio Mouse TNF-alpha ELISA Kit (For Lysates) Catalog #: ELM-TNFa-CL User Manual Last revised September 15, 2017 Caution: Extraordinarily useful information enclosed ISO 13485 Certified 3607 Parkway Lane,
Human Syndecan-3 ELISA Kit
 BioAim Scientific Inc Human Syndecan-3 ELISA Kit Cat.No: 3010574 Instruction Manual (Last revised May 01, 2017) For research use only TABLE OF CONTENTS I. INTRODUCTION.......3 II. MATERIALS SUPPLIED 4
BioAim Scientific Inc Human Syndecan-3 ELISA Kit Cat.No: 3010574 Instruction Manual (Last revised May 01, 2017) For research use only TABLE OF CONTENTS I. INTRODUCTION.......3 II. MATERIALS SUPPLIED 4
ncounter PlexSet Data Analysis Guidelines
 ncounter PlexSet Data Analysis Guidelines NanoString Technologies, Inc. 530 airview Ave North Seattle, Washington 98109 USA Telephone: 206.378.6266 888.358.6266 E-mail: info@nanostring.com Molecules That
ncounter PlexSet Data Analysis Guidelines NanoString Technologies, Inc. 530 airview Ave North Seattle, Washington 98109 USA Telephone: 206.378.6266 888.358.6266 E-mail: info@nanostring.com Molecules That
Quant-iT dsdna High-Sensitivity Assay Kit
 Quant-iT dsdna High-Sensitivity Assay Kit Table 1. Contents and storage information. Material Amount Concentration Storage Stability Quant-iT dsdna HS reagent (Component A) λ dsdna HS standards (Component
Quant-iT dsdna High-Sensitivity Assay Kit Table 1. Contents and storage information. Material Amount Concentration Storage Stability Quant-iT dsdna HS reagent (Component A) λ dsdna HS standards (Component
ActiPix SDI300 Surface Dissolution Imaging System
 Product Note: PN002B Figure 1. SDI300 Surface Dissolution Imaging System Introduction The ActiPix SDI300 is a powerful UV area imaging system which enables quantitative imaging of surface phenomena for
Product Note: PN002B Figure 1. SDI300 Surface Dissolution Imaging System Introduction The ActiPix SDI300 is a powerful UV area imaging system which enables quantitative imaging of surface phenomena for
Ion Sphere Assay on the Qubit 3.0 Fluorometer
 USER GUIDE Ion Sphere Assay on the Qubit 3.0 Fluorometer for use with: Ion Sphere Quality Control Kit (Cat. No. 4468656) Publication Number MAN0016388 Revision A.0 Ion Sphere Assay overview... 2 Materials
USER GUIDE Ion Sphere Assay on the Qubit 3.0 Fluorometer for use with: Ion Sphere Quality Control Kit (Cat. No. 4468656) Publication Number MAN0016388 Revision A.0 Ion Sphere Assay overview... 2 Materials
Mouse TNF-alpha ELISAKit
 BioAim Scientific Inc Mouse TNF-alpha ELISAKit Cat.No: 3020014 Instruction Manual (Last revised Dec 24, 2015) For research use only TABLE OF CONTENTS I. INTRODUCTION.......3 II. MATERIALS SUPPLIED 4 III.
BioAim Scientific Inc Mouse TNF-alpha ELISAKit Cat.No: 3020014 Instruction Manual (Last revised Dec 24, 2015) For research use only TABLE OF CONTENTS I. INTRODUCTION.......3 II. MATERIALS SUPPLIED 4 III.
TruSight Cancer Workflow on the MiniSeq System
 TruSight Cancer Workflow on the MiniSeq System Prepare Library Sequence Analyze Data TruSight Cancer 1.5 days ~ 24 hours < 2 hours TruSight Cancer Library Prep MiniSeq System Local Run Manager Enrichment
TruSight Cancer Workflow on the MiniSeq System Prepare Library Sequence Analyze Data TruSight Cancer 1.5 days ~ 24 hours < 2 hours TruSight Cancer Library Prep MiniSeq System Local Run Manager Enrichment
Automated Illumina TruSight HLA v2 Sequencing Panel Library Preparation with the epmotion 5075t
 SHORT PROTOCOL No. 41 Automated Illumina TruSight HLA v2 Sequencing Panel Library Preparation with the epmotion 5075t Introduction This protocol describes the workstation configuration and pre-programmed
SHORT PROTOCOL No. 41 Automated Illumina TruSight HLA v2 Sequencing Panel Library Preparation with the epmotion 5075t Introduction This protocol describes the workstation configuration and pre-programmed
Globin-Zero Gold rrna Removal Kit Reference Guide
 Globin-Zero Gold rrna Removal Kit Reference Guide For Research Use Only. Not for use in diagnostic procedures. ILLUMINA PROPRIETARY Document # 15066013 v02 August 2016 Customize a short end-to-end workflow
Globin-Zero Gold rrna Removal Kit Reference Guide For Research Use Only. Not for use in diagnostic procedures. ILLUMINA PROPRIETARY Document # 15066013 v02 August 2016 Customize a short end-to-end workflow
SUPPLEMENTAL DATA: ROBUST ESTIMATORS FOR EXPRESSION ANALYSIS EARL HUBBELL, WEI-MIN LIU, AND RUI MEI
 SUPPLEMENTAL DATA: ROBUST ESTIMATORS FOR EXPRESSION ANALYSIS EARL HUBBELL, WEI-MIN LIU, AND RUI MEI ABSTRACT. This is supplemental data extracted from the paper Robust Estimators for Expression Analysis
SUPPLEMENTAL DATA: ROBUST ESTIMATORS FOR EXPRESSION ANALYSIS EARL HUBBELL, WEI-MIN LIU, AND RUI MEI ABSTRACT. This is supplemental data extracted from the paper Robust Estimators for Expression Analysis
PSA (Human) ELISA Kit
 PSA (Human) ELISA Kit Catalog Number KA0208 96 assays Version: 04 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use... 3 Principle of the Assay... 3 General
PSA (Human) ELISA Kit Catalog Number KA0208 96 assays Version: 04 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use... 3 Principle of the Assay... 3 General
RayBio Human CD86/B7-2 ELISA Kit
 RayBio Human CD86/B7-2 ELISA Kit Catalog #: ELH-CD86 User Manual Last revised April 15, 2016 Caution: Extraordinarily useful information enclosed ISO 13485 Certified 3607 Parkway Lane, Suite 100 Norcross,
RayBio Human CD86/B7-2 ELISA Kit Catalog #: ELH-CD86 User Manual Last revised April 15, 2016 Caution: Extraordinarily useful information enclosed ISO 13485 Certified 3607 Parkway Lane, Suite 100 Norcross,
Axiom TM 2.0 Target Prep 384 Samples
 Quick Reference Card Axiom TM 2.0 Target Prep 384 Samples Stage 1 DNA Amplification Introduction Running the Axiom 2.0 Assay for 384 Samples Assay requires the following sets of steps: 1. Genomic DNA Prep,
Quick Reference Card Axiom TM 2.0 Target Prep 384 Samples Stage 1 DNA Amplification Introduction Running the Axiom 2.0 Assay for 384 Samples Assay requires the following sets of steps: 1. Genomic DNA Prep,
Isolation of Total RNA and mrna from Plant Tissues
 Promega Notes Magazine Number 54, 1995, p.02 Isolation of Total RNA and mrna from Plant Tissues By: Isabel Murillo, Dora Raventos, Estelle Jaeck, Blanca San Segundo* Centro de Investigacion y Desarrollo
Promega Notes Magazine Number 54, 1995, p.02 Isolation of Total RNA and mrna from Plant Tissues By: Isabel Murillo, Dora Raventos, Estelle Jaeck, Blanca San Segundo* Centro de Investigacion y Desarrollo
TrioMol Isolation Reagent
 TrioMol Isolation Reagent Technical Manual No. 0242 Version 06142007 I Description... 1 II Key Features... 1 III Storage..... 1 IV General Protocol Using Triomol Isolation Reagent 1 V Troubleshooting.
TrioMol Isolation Reagent Technical Manual No. 0242 Version 06142007 I Description... 1 II Key Features... 1 III Storage..... 1 IV General Protocol Using Triomol Isolation Reagent 1 V Troubleshooting.
Human IL-1 beta ELISA Kit
 For detection and measurement of human interleukin 1 beta Catalog #02004 Catalog #02005 2 Plates 10 Plates Product Description The Human Interleukin 1 beta (IL-1β) ELISA Kit is designed for the quantitative
For detection and measurement of human interleukin 1 beta Catalog #02004 Catalog #02005 2 Plates 10 Plates Product Description The Human Interleukin 1 beta (IL-1β) ELISA Kit is designed for the quantitative
SUPPLEMENTARY INFORMATION
 High-density integration of carbon nanotubes by chemical self-assembly Hongsik Park, Ali Afzali, Shu-Jen Han, George S. Tulevski, Aaron D. Franklin, Jerry Tersoff, James B. Hannon and Wilfried Haensch
High-density integration of carbon nanotubes by chemical self-assembly Hongsik Park, Ali Afzali, Shu-Jen Han, George S. Tulevski, Aaron D. Franklin, Jerry Tersoff, James B. Hannon and Wilfried Haensch
FOR RESEARCH USE ONLY.
 MAN-10039-04 Vantage 3D DNA SNV Qualification Kit Vantage 3D DNA SNV Qualification Kit The ncounter Vantage 3D DNA SNV Qualification Kit is designed to assess whether the ncounter MAX, FLEX, or SPRINT
MAN-10039-04 Vantage 3D DNA SNV Qualification Kit Vantage 3D DNA SNV Qualification Kit The ncounter Vantage 3D DNA SNV Qualification Kit is designed to assess whether the ncounter MAX, FLEX, or SPRINT
Low-Level Analysis of High- Density Oligonucleotide Microarray Data
 Low-Level Analysis of High- Density Oligonucleotide Microarray Data Ben Bolstad http://www.stat.berkeley.edu/~bolstad Biostatistics, University of California, Berkeley UC Berkeley Feb 23, 2004 Outline
Low-Level Analysis of High- Density Oligonucleotide Microarray Data Ben Bolstad http://www.stat.berkeley.edu/~bolstad Biostatistics, University of California, Berkeley UC Berkeley Feb 23, 2004 Outline
In fact, we are going to be sneaky and use Hess s Law to determine the heat of magnesium combustion indirectly. Go to the website:
 Chemistry 161 Please prepare your notebook though the data tables before class on Wednesday, October 27. Write an abstract and place it at the front of your individual report. The abstract and the report
Chemistry 161 Please prepare your notebook though the data tables before class on Wednesday, October 27. Write an abstract and place it at the front of your individual report. The abstract and the report
New Approaches to the Development of GC/MS Selected Ion Monitoring Acquisition and Quantitation Methods Technique/Technology
 New Approaches to the Development of GC/MS Selected Ion Monitoring Acquisition and Quantitation Methods Technique/Technology Gas Chromatography/Mass Spectrometry Author Harry Prest 1601 California Avenue
New Approaches to the Development of GC/MS Selected Ion Monitoring Acquisition and Quantitation Methods Technique/Technology Gas Chromatography/Mass Spectrometry Author Harry Prest 1601 California Avenue
TrioMol Isolation Reagent
 TrioMol Isolation Reagent Technical Manual No. 0242 Version 06142007 I Description... 1 II Key Features... 1 III Storage..... 1 IV General Protocol Using Triomol Isolation Reagent 1 V Troubleshooting.
TrioMol Isolation Reagent Technical Manual No. 0242 Version 06142007 I Description... 1 II Key Features... 1 III Storage..... 1 IV General Protocol Using Triomol Isolation Reagent 1 V Troubleshooting.
CSA: Simian Panel E. (Product Number ) User Protocol. 1.0 Introduction Kit Contents Required Materials...
 L757 28Jun2018 1 of 9 Table of Contents 1.0 Introduction... 2 2.0 Kit Contents... 3 3.0 Required Materials... 3 4.0 Storage... 3 5.0 Safety and Handling... 4 6.0 Protocol Overview... 4 7.0 Procedure...
L757 28Jun2018 1 of 9 Table of Contents 1.0 Introduction... 2 2.0 Kit Contents... 3 3.0 Required Materials... 3 4.0 Storage... 3 5.0 Safety and Handling... 4 6.0 Protocol Overview... 4 7.0 Procedure...
Qubit RNA IQ Assay Kits
 USER GUIDE Qubit RNA IQ s Catalog No. Q33221, Q33222 Pub. No. MAN0017405 Rev. B.0 Product information The Qubit RNA IQ provides a fast, simple method to check whether an RNA sample has degraded using the
USER GUIDE Qubit RNA IQ s Catalog No. Q33221, Q33222 Pub. No. MAN0017405 Rev. B.0 Product information The Qubit RNA IQ provides a fast, simple method to check whether an RNA sample has degraded using the
And how to do them. Denise L Seman City of Youngstown
 And how to do them Denise L Seman City of Youngstown Quality Control (QC) is defined as the process of detecting analytical errors to ensure both reliability and accuracy of the data generated QC can be
And how to do them Denise L Seman City of Youngstown Quality Control (QC) is defined as the process of detecting analytical errors to ensure both reliability and accuracy of the data generated QC can be
Normalization. Example of Replicate Data. Biostatistics Rafael A. Irizarry
 This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike License. Your use of this material constitutes acceptance of that license and the conditions of use of materials on this
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike License. Your use of this material constitutes acceptance of that license and the conditions of use of materials on this
Highly automated protein formulation development: a case study
 Application Note Highly automated protein formulation development: a case study Introduction and background Therapeutic proteins can be inherently prone to degradation and instability, and they often pose
Application Note Highly automated protein formulation development: a case study Introduction and background Therapeutic proteins can be inherently prone to degradation and instability, and they often pose
ab MDA Assay Kit (competitive ELISA)
 Version 1 Last updated 30 August 2018 ab238537 MDA Assay Kit (competitive ELISA) For the quantitative measurement of MDA in protein samples such as purified protein, plasma, serum and cell lysate. This
Version 1 Last updated 30 August 2018 ab238537 MDA Assay Kit (competitive ELISA) For the quantitative measurement of MDA in protein samples such as purified protein, plasma, serum and cell lysate. This
Human Coagulation Factor XII Total Antigen ELISA Kit
 Human Coagulation Factor XII Total Antigen ELISA Kit Catalog No: IHFXIIKT-TOT Lot No: SAMPLE INTENDED USE This human coagulation Factor XII antigen assay is intended for the quantitative determination
Human Coagulation Factor XII Total Antigen ELISA Kit Catalog No: IHFXIIKT-TOT Lot No: SAMPLE INTENDED USE This human coagulation Factor XII antigen assay is intended for the quantitative determination
11,12-EET/DHET ELISA kit
 11,12-EET/DHET ELISA kit Catalog Number: DH5/DH15/DH25 Store at -20 C. FOR RESEARCH USE ONLY V.0522010 Introduction It is well known that arachidonic acid (AA) will be converted to EET by P 450 arachidonic
11,12-EET/DHET ELISA kit Catalog Number: DH5/DH15/DH25 Store at -20 C. FOR RESEARCH USE ONLY V.0522010 Introduction It is well known that arachidonic acid (AA) will be converted to EET by P 450 arachidonic
Ribo-Zero rrna Removal Kit Reference Guide
 Ribo-Zero rrna Removal Kit Reference Guide For Research Use Only. Not for use in diagnostic procedures. ILLUMINA PROPRIETARY Document # 15066012 v02 August 2016 Customize a short end-to-end workflow guide
Ribo-Zero rrna Removal Kit Reference Guide For Research Use Only. Not for use in diagnostic procedures. ILLUMINA PROPRIETARY Document # 15066012 v02 August 2016 Customize a short end-to-end workflow guide
RayBio Mouse VEGF R1 ELISA Kit
 RayBio Mouse VEGF R1 ELISA Kit Catalog #: ELM-VEGFR1 User Manual Last revised April 15, 2016 Caution: Extraordinarily useful information enclosed ISO 13485 Certified 3607 Parkway Lane, Suite 100 Norcross,
RayBio Mouse VEGF R1 ELISA Kit Catalog #: ELM-VEGFR1 User Manual Last revised April 15, 2016 Caution: Extraordinarily useful information enclosed ISO 13485 Certified 3607 Parkway Lane, Suite 100 Norcross,
Probe-Level Analysis of Affymetrix GeneChip Microarray Data
 Probe-Level Analysis of Affymetrix GeneChip Microarray Data Ben Bolstad http://www.stat.berkeley.edu/~bolstad Biostatistics, University of California, Berkeley University of Minnesota Mar 30, 2004 Outline
Probe-Level Analysis of Affymetrix GeneChip Microarray Data Ben Bolstad http://www.stat.berkeley.edu/~bolstad Biostatistics, University of California, Berkeley University of Minnesota Mar 30, 2004 Outline
mrna Isolation Kit for Blood/Bone Marrow For isolation mrna from blood or bone marrow lysates Cat. No
 For isolation mrna from blood or bone marrow lysates Cat. No. 1 934 333 Principle Starting material Application Time required Results Key advantages The purification of mrna requires two steps: 1. Cells
For isolation mrna from blood or bone marrow lysates Cat. No. 1 934 333 Principle Starting material Application Time required Results Key advantages The purification of mrna requires two steps: 1. Cells
Optimal normalization of DNA-microarray data
 Optimal normalization of DNA-microarray data Daniel Faller 1, HD Dr. J. Timmer 1, Dr. H. U. Voss 1, Prof. Dr. Honerkamp 1 and Dr. U. Hobohm 2 1 Freiburg Center for Data Analysis and Modeling 1 F. Hoffman-La
Optimal normalization of DNA-microarray data Daniel Faller 1, HD Dr. J. Timmer 1, Dr. H. U. Voss 1, Prof. Dr. Honerkamp 1 and Dr. U. Hobohm 2 1 Freiburg Center for Data Analysis and Modeling 1 F. Hoffman-La
THE UNIVERSITY OF NEWCASTLE- DISCIPLINE OF MEDICAL BIOCHEMISTRY. STANDARD OPERATING PROCEDURE PROCEDURE NO: GLP 019 MOD: 1st Issue Page: 1 of 6
 Page: 1 of 6 1. Risk Assessment:. This Risk Assessment is to be used as a general guide and as such, cannot accommodate all the varying factors that may be encountered when using this equipment. Therefore,
Page: 1 of 6 1. Risk Assessment:. This Risk Assessment is to be used as a general guide and as such, cannot accommodate all the varying factors that may be encountered when using this equipment. Therefore,
Spread Spectrum Watermarking. Information Technologies for IPR Protection
 Spread Spectrum Watermarking Information Technologies for IPR Protection Spread Spectrum Communication A Narrow-band signal is transmitted over a much larger bandwidth such that the signal energy presented
Spread Spectrum Watermarking Information Technologies for IPR Protection Spread Spectrum Communication A Narrow-band signal is transmitted over a much larger bandwidth such that the signal energy presented
MyBioSource.com. This package insert must be read in its entirety before using this product.
 Tetracyclines ELISA Kit Catalog Number. MBS940077 This immunoassay kit allows for the in vitro quantitative determination of Tetracyclines concentrations in honey, tissue(chicken, pork). This package insert
Tetracyclines ELISA Kit Catalog Number. MBS940077 This immunoassay kit allows for the in vitro quantitative determination of Tetracyclines concentrations in honey, tissue(chicken, pork). This package insert
Ion Channel Discovery Services
 www.essenbioscience.com Ion Channel Discovery Services Customer: Customer PO: Essen contact: Date of assay: Customer contact: Essen Quote No: Srini Kanumilli Lab Notebook Ref: Date of Report: LNB-2650
www.essenbioscience.com Ion Channel Discovery Services Customer: Customer PO: Essen contact: Date of assay: Customer contact: Essen Quote No: Srini Kanumilli Lab Notebook Ref: Date of Report: LNB-2650
Data Analysis II. CU- Boulder CHEM-4181 Instrumental Analysis Laboratory. Prof. Jose-Luis Jimenez Spring 2007
 Data Analysis II CU- Boulder CHEM-48 Instrumental Analysis Laboratory Prof. Jose-Luis Jimenez Spring 007 Lecture will be posted on course web page based on lab manual, Skoog, web links Summary of Last
Data Analysis II CU- Boulder CHEM-48 Instrumental Analysis Laboratory Prof. Jose-Luis Jimenez Spring 007 Lecture will be posted on course web page based on lab manual, Skoog, web links Summary of Last
Name Date Period. 1. If drops of ACID are added to a ph buffer, then the ph of the buffer will [increase / decrease / stay the same].
![Name Date Period. 1. If drops of ACID are added to a ph buffer, then the ph of the buffer will [increase / decrease / stay the same]. Name Date Period. 1. If drops of ACID are added to a ph buffer, then the ph of the buffer will [increase / decrease / stay the same].](/thumbs/89/98268119.jpg) Name Date Period ACIDS AND BASES Organisms are often very sensitive to the effect of s and s in their environment. They need to maintain a stable internal ph in order to survive even in the event of environmental
Name Date Period ACIDS AND BASES Organisms are often very sensitive to the effect of s and s in their environment. They need to maintain a stable internal ph in order to survive even in the event of environmental
RFI Mitigation for the Parkes Galactic All-Sky Survey (GASS)
 RFI Mitigation for the Parkes Galactic All-Sky Survey (GASS) Peter Kalberla Argelander-Institut für Astronomie, Auf dem Hügel 71, D-53121 Bonn, Germany E-mail: pkalberla@astro.uni-bonn.de The GASS is a
RFI Mitigation for the Parkes Galactic All-Sky Survey (GASS) Peter Kalberla Argelander-Institut für Astronomie, Auf dem Hügel 71, D-53121 Bonn, Germany E-mail: pkalberla@astro.uni-bonn.de The GASS is a
user manual blotting Hoefer PR648 Slot blot manifold um PR648-IM/Rev.F0/08-12
 user manual blotting Hoefer PR648 Slot blot manifold um PR648-IM/Rev.F0/08-12 Page finder Introduction...1 Specifications...2 Procedure for standard use of the Hoefer PR648...3 Setting up the Hoefer PR648...3
user manual blotting Hoefer PR648 Slot blot manifold um PR648-IM/Rev.F0/08-12 Page finder Introduction...1 Specifications...2 Procedure for standard use of the Hoefer PR648...3 Setting up the Hoefer PR648...3
Virtual Beach Building a GBM Model
 Virtual Beach 3.0.6 Building a GBM Model Building, Evaluating and Validating Anytime Nowcast Models In this module you will learn how to: A. Build and evaluate an anytime GBM model B. Optimize a GBM model
Virtual Beach 3.0.6 Building a GBM Model Building, Evaluating and Validating Anytime Nowcast Models In this module you will learn how to: A. Build and evaluate an anytime GBM model B. Optimize a GBM model
Color Compensation Set 1
 Color Compensation Set 1 Version 2, March 2012 Solutions for the generation of color compensation objects for the LightCycler System. Order No. A 500 08 For 5 calibration runs. Store in dark at -15 to
Color Compensation Set 1 Version 2, March 2012 Solutions for the generation of color compensation objects for the LightCycler System. Order No. A 500 08 For 5 calibration runs. Store in dark at -15 to
Objective A: Mean, Median and Mode Three measures of central of tendency: the mean, the median, and the mode.
 Chapter 3 Numerically Summarizing Data Chapter 3.1 Measures of Central Tendency Objective A: Mean, Median and Mode Three measures of central of tendency: the mean, the median, and the mode. A1. Mean The
Chapter 3 Numerically Summarizing Data Chapter 3.1 Measures of Central Tendency Objective A: Mean, Median and Mode Three measures of central of tendency: the mean, the median, and the mode. A1. Mean The
Aflatoxin M1 (AFM1) ELISA Kit
 Aflatoxin M1 (AFM1) ELISA Kit Catalog Number. CSB-EL027236 This immunoassay kit allows for the in vitro quantitative determination of Aflatoxin M1 concentrations in milk, milk power. This package insert
Aflatoxin M1 (AFM1) ELISA Kit Catalog Number. CSB-EL027236 This immunoassay kit allows for the in vitro quantitative determination of Aflatoxin M1 concentrations in milk, milk power. This package insert
mag nanogram kit Description Kit uses
 mag nanogram kit Catalogue number 40701 and 40710 (For research use only. Not for use in diagnostic procedures.) Description mag kits use magnetic separation for the preparation of nucleic acids. Superparamagnetic
mag nanogram kit Catalogue number 40701 and 40710 (For research use only. Not for use in diagnostic procedures.) Description mag kits use magnetic separation for the preparation of nucleic acids. Superparamagnetic
Error models and normalization. Wolfgang Huber DKFZ Heidelberg
 Error models and normalization Wolfgang Huber DKFZ Heidelberg Acknowledgements Anja von Heydebreck, Martin Vingron Andreas Buness, Markus Ruschhaupt, Klaus Steiner, Jörg Schneider, Katharina Finis, Anke
Error models and normalization Wolfgang Huber DKFZ Heidelberg Acknowledgements Anja von Heydebreck, Martin Vingron Andreas Buness, Markus Ruschhaupt, Klaus Steiner, Jörg Schneider, Katharina Finis, Anke
For the quantitative measurement of PEGylated molecules in plasma, serum and cell culture media
 ab138914 Polyethylene Glycol (PEG) ELISA Kit Instructions for Use For the quantitative measurement of PEGylated molecules in plasma, serum and cell culture media This product is for research use only and
ab138914 Polyethylene Glycol (PEG) ELISA Kit Instructions for Use For the quantitative measurement of PEGylated molecules in plasma, serum and cell culture media This product is for research use only and
NOVABEADS FOOD 1 DNA KIT
 NOVABEADS FOOD 1 DNA KIT NOVABEADS FOOD DNA KIT is the new generation tool in molecular biology techniques and allows DNA isolations from highly processed food products. The method is based on the use
NOVABEADS FOOD 1 DNA KIT NOVABEADS FOOD DNA KIT is the new generation tool in molecular biology techniques and allows DNA isolations from highly processed food products. The method is based on the use
Automated Illumina TruSeq DNA PCR-Free library construction with the epmotion 5075t/TMX
 SHORT PROTOCOL No. 15 I August 2016 Automated Illumina TruSeq DNA PCR-Free library construction with the epmotion 5075t/TMX Introduction This protocol describes the configuration and preprogrammed methods
SHORT PROTOCOL No. 15 I August 2016 Automated Illumina TruSeq DNA PCR-Free library construction with the epmotion 5075t/TMX Introduction This protocol describes the configuration and preprogrammed methods
Carcino embryonic Antigen Human ELISA Kit
 ab108635 Carcino embryonic Antigen Human ELISA Kit Instructions for Use For the quantitative measurement of Human Carcino embryonic Antigen (CEA) concentrations in serum. This product is for research use
ab108635 Carcino embryonic Antigen Human ELISA Kit Instructions for Use For the quantitative measurement of Human Carcino embryonic Antigen (CEA) concentrations in serum. This product is for research use
The recent biological and therapeutic discoveries of RNA have led to numerous alterations in the chemical synthesis
 Application Note: TN-16 RNA Sample Preparation & High-Throughput Purification for TBDMS & TOM Chemistries Using Clarity QSP Author: Greg Scott Introduction The recent biological and therapeutic discoveries
Application Note: TN-16 RNA Sample Preparation & High-Throughput Purification for TBDMS & TOM Chemistries Using Clarity QSP Author: Greg Scott Introduction The recent biological and therapeutic discoveries
THE UNIVERSITY OF NEWCASTLE- DISCIPLINE OF MEDICAL BIOCHEMISTRY. STANDARD OPERATING PROCEDURE PROCEDURE NO: GLP 021 MOD: 1st Issue Page: 1 of 6
 Page: 1 of 6 1. Risk Assessment: This Risk Assessment is to be used as a general guide and as such, cannot accommodate all the varying factors that may be encountered when using this equipment. Therefore,
Page: 1 of 6 1. Risk Assessment: This Risk Assessment is to be used as a general guide and as such, cannot accommodate all the varying factors that may be encountered when using this equipment. Therefore,
Design of Microarray Experiments. Xiangqin Cui
 Design of Microarray Experiments Xiangqin Cui Experimental design Experimental design: is a term used about efficient methods for planning the collection of data, in order to obtain the maximum amount
Design of Microarray Experiments Xiangqin Cui Experimental design Experimental design: is a term used about efficient methods for planning the collection of data, in order to obtain the maximum amount
Microarray Preprocessing
 Microarray Preprocessing Normaliza$on Normaliza$on is needed to ensure that differences in intensi$es are indeed due to differen$al expression, and not some prin$ng, hybridiza$on, or scanning ar$fact.
Microarray Preprocessing Normaliza$on Normaliza$on is needed to ensure that differences in intensi$es are indeed due to differen$al expression, and not some prin$ng, hybridiza$on, or scanning ar$fact.
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Bradley Broom Department of Bioinformatics and Computational Biology UT M. D. Anderson Cancer Center kabagg@mdanderson.org bmbroom@mdanderson.org
GS01 0163 Analysis of Microarray Data Keith Baggerly and Bradley Broom Department of Bioinformatics and Computational Biology UT M. D. Anderson Cancer Center kabagg@mdanderson.org bmbroom@mdanderson.org
