Procedure to Create NCBI KOGS
|
|
|
- May Anthony
- 5 years ago
- Views:
Transcription
1 Procedure to Create NCBI KOGS full details in: Tatusov et al (2003) BMC Bioinformatics 4: Detect and mask typical repetitive domains Reason: masking prevents spurious lumping of non-orthologs based only on common shared domains o Examples PPR Repeat 35 amino acids long up to 18 copiers per protein generally copy number is expanded in plants C2H2 classic zinc finger domain 25 amino acids long binds to major groove of DNA protein can function as a transcription regulator 2. All-against-all BLASTP analysis Comparison of amino acid sequences All sequences used as the database Each sequence used as a query against this database P values can be rather high Ubiquitin cluster
2 3. Mutually consistent triangles of genome specific best hits 4. Merge triangles with common sides 5. Manual analysis of each cluster to eliminate false positives 6. Assignment of masked proteins (step 1) to clusters 7. Perform phylogenetic clustering on KOGs with proteins from multiple species Species Arabidopsis thaliana (thale cress) Caenorhabditis elegans (worm) Drosophila melanogaster (fruit fly) Homo sapiens (human) Saccharomyces cerevisiae (baker yeast) Schizosaccharomyces pombe (fission yeast) Encephalitozoon cuniculi (Microsporidia) Species Designation A C D H Y P E Species Groupings with Largest Members Genomes Members CDH 1147 ACDHYP 928 ACDHYPE 860 ACDH 484 CDHYP 152
3 Examples of KOGs KOG containing all species 860 clusters KOG 0001: Ubiquitin and ubiquitin-like proteins Species Number of members Arabidopsis thaliana (thale cress) 29 Caenorhabditis elegans (worm) 12 Drosophila melanogaster (fruit fly) 3 Homo sapiens (human) 17 Saccharomyces cerevisiae (baker yeast) 2 Schizosaccharomyces pombe (fission yeast) 1 Encephalitozoon cuniculi (Microsporidia) 1
4 Arabidopsis At5g20620 vs. Baker Yeast YLL039c >YLL039c Length = 381 Score = 723 bits (1866), Expect = 0.0 Identities = 370/381 (97%), Positives = 381/381 (99%) Query: 1 MQIFVKTLTGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADYN 60 MQIFVKTLTGKTITLEVESSDTIDNVK+KIQDKEGIPPDQQRLIFAGKQLEDGRTL+DYN Sbjct: 1 MQIFVKTLTGKTITLEVESSDTIDNVKSKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN 60 Query: 61 IQKESTLHLVLRLRGGMQIFVKTLTGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLI 120 IQKESTLHLVLRLRGGMQIFVKTLTGKTITLEVESSDTIDNVK+KIQDKEGIPPDQQRLI Sbjct: 61 IQKESTLHLVLRLRGGMQIFVKTLTGKTITLEVESSDTIDNVKSKIQDKEGIPPDQQRLI 120 Query: 121 FAGKQLEDGRTLADYNIQKESTLHLVLRLRGGMQIFVKTLTGKTITLEVESSDTIDNVKA 180 FAGKQLEDGRTL+DYNIQKESTLHLVLRLRGGMQIFVKTLTGKTITLEVESSDTIDNVK+ Sbjct: 121 FAGKQLEDGRTLSDYNIQKESTLHLVLRLRGGMQIFVKTLTGKTITLEVESSDTIDNVKS 180 Query: 181 KIQDKEGIPPDQQRLIFAGKQLEDGRTLADYNIQKESTLHLVLRLRGGMQIFVKTLTGKT 240 KIQDKEGIPPDQQRLIFAGKQLEDGRTL+DYNIQKESTLHLVLRLRGGMQIFVKTLTGKT Sbjct: 181 KIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYNIQKESTLHLVLRLRGGMQIFVKTLTGKT 240 Query: 241 ITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADYNIQKESTLHLVLR 300 ITLEVESSDTIDNVK+KIQDKEGIPPDQQRLIFAGKQLEDGRTL+DYNIQKESTLHLVLR Sbjct: 241 ITLEVESSDTIDNVKSKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYNIQKESTLHLVLR 300 Query: 301 LRGGMQIFVKTLTGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTL 360 LRGGMQIFVKTLTGKTITLEVESSDTIDNVK+KIQDKEGIPPDQQRLIFAGKQLEDGRTL Sbjct: 301 LRGGMQIFVKTLTGKTITLEVESSDTIDNVKSKIQDKEGIPPDQQRLIFAGKQLEDGRTL 360 Query: 361 ADYNIQKESTLHLVLRLRGGS 381 +DYNIQKESTLHLVLRLRGG+ Sbjct: 361 SDYNIQKESTLHLVLRLRGGN 381 Arabidopsis At5g20620 vs. Arabidopsis At1g14650 >At1g14650 Length = 785 Score = 38.5 bits (88), Expect = Identities = 24/68 (35%), Positives = 39/68 (57%), Gaps = 1/68 (1%) Query: 314 GKTITLEVES-SDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADYNIQKESTLH 372 G+ + + V+S S+ + ++K KI + IP ++Q+L L+D +LA YN+ L Sbjct: 715 GQFMEITVQSLSENVGSLKEKIAGEIQIPANKQKLSGKAGFLKDNMSLAHYNVGAGEILT 774 Query: 373 LVLRLRGG 380 L LR RGG Sbjct: 775 LSLRERGG 782
5 KOG 0001: 40S Ribosomal Protein S17 Species Number of members Arabidopsis thaliana (thale cress) 4 Caenorhabditis elegans (worm) 1 Drosophila melanogaster (fruit fly) 1 Homo sapiens (human) 8 Saccharomyces cerevisiae (baker yeast) 2 Schizosaccharomyces pombe (fission yeast) 2 Encephalitozoon cuniculi (Microsporidia) 1
6
Quantitative Measurement of Genome-wide Protein Domain Co-occurrence of Transcription Factors
 Quantitative Measurement of Genome-wide Protein Domain Co-occurrence of Transcription Factors Arli Parikesit, Peter F. Stadler, Sonja J. Prohaska Bioinformatics Group Institute of Computer Science University
Quantitative Measurement of Genome-wide Protein Domain Co-occurrence of Transcription Factors Arli Parikesit, Peter F. Stadler, Sonja J. Prohaska Bioinformatics Group Institute of Computer Science University
Giri Narasimhan. CAP 5510: Introduction to Bioinformatics CGS 5166: Bioinformatics Tools. Evaluation. Course Homepage.
 CAP 5510: Introduction to Bioinformatics CGS 5166: Bioinformatics Tools Giri Narasimhan ECS 389; Phone: x3748 giri@cis.fiu.edu www.cis.fiu.edu/~giri/teach/bioinfs06.html 1/12/06 CAP5510/CGS5166 1 Evaluation
CAP 5510: Introduction to Bioinformatics CGS 5166: Bioinformatics Tools Giri Narasimhan ECS 389; Phone: x3748 giri@cis.fiu.edu www.cis.fiu.edu/~giri/teach/bioinfs06.html 1/12/06 CAP5510/CGS5166 1 Evaluation
Advanced Algorithms and Models for Computational Biology
 Advanced Algorithms and Models for Computational Biology Introduction to cell biology genomics development and probability Eric ing Lecture January 3 006 Reading: Chap. DTM book Introduction to cell biology
Advanced Algorithms and Models for Computational Biology Introduction to cell biology genomics development and probability Eric ing Lecture January 3 006 Reading: Chap. DTM book Introduction to cell biology
Graph Alignment and Biological Networks
 Graph Alignment and Biological Networks Johannes Berg http://www.uni-koeln.de/ berg Institute for Theoretical Physics University of Cologne Germany p.1/12 Networks in molecular biology New large-scale
Graph Alignment and Biological Networks Johannes Berg http://www.uni-koeln.de/ berg Institute for Theoretical Physics University of Cologne Germany p.1/12 Networks in molecular biology New large-scale
ECOL/MCB 320 and 320H Genetics
 ECOL/MCB 320 and 320H Genetics Instructors Dr. C. William Birky, Jr. Dept. of Ecology and Evolutionary Biology Lecturing on Molecular genetics Transmission genetics Population and evolutionary genetics
ECOL/MCB 320 and 320H Genetics Instructors Dr. C. William Birky, Jr. Dept. of Ecology and Evolutionary Biology Lecturing on Molecular genetics Transmission genetics Population and evolutionary genetics
BIOINFORMATICS LAB AP BIOLOGY
 BIOINFORMATICS LAB AP BIOLOGY Bioinformatics is the science of collecting and analyzing complex biological data. Bioinformatics combines computer science, statistics and biology to allow scientists to
BIOINFORMATICS LAB AP BIOLOGY Bioinformatics is the science of collecting and analyzing complex biological data. Bioinformatics combines computer science, statistics and biology to allow scientists to
CGS 5991 (2 Credits) Bioinformatics Tools
 CAP 5991 (3 Credits) Introduction to Bioinformatics CGS 5991 (2 Credits) Bioinformatics Tools Giri Narasimhan 8/26/03 CAP/CGS 5991: Lecture 1 1 Course Schedules CAP 5991 (3 credit) will meet every Tue
CAP 5991 (3 Credits) Introduction to Bioinformatics CGS 5991 (2 Credits) Bioinformatics Tools Giri Narasimhan 8/26/03 CAP/CGS 5991: Lecture 1 1 Course Schedules CAP 5991 (3 credit) will meet every Tue
Genome Annotation. Bioinformatics and Computational Biology. Genome sequencing Assembly. Gene prediction. Protein targeting.
 Genome Annotation Bioinformatics and Computational Biology Genome Annotation Frank Oliver Glöckner 1 Genome Analysis Roadmap Genome sequencing Assembly Gene prediction Protein targeting trna prediction
Genome Annotation Bioinformatics and Computational Biology Genome Annotation Frank Oliver Glöckner 1 Genome Analysis Roadmap Genome sequencing Assembly Gene prediction Protein targeting trna prediction
Welcome to BIOL 572: Recombinant DNA techniques
 Lecture 1: 1 Welcome to BIOL 572: Recombinant DNA techniques Agenda 1: Introduce yourselves Agenda 2: Course introduction Agenda 3: Some logistics for BIOL 572 Agenda 4: Q&A section Agenda 1: Introduce
Lecture 1: 1 Welcome to BIOL 572: Recombinant DNA techniques Agenda 1: Introduce yourselves Agenda 2: Course introduction Agenda 3: Some logistics for BIOL 572 Agenda 4: Q&A section Agenda 1: Introduce
Genomics and bioinformatics summary. Finding genes -- computer searches
 Genomics and bioinformatics summary 1. Gene finding: computer searches, cdnas, ESTs, 2. Microarrays 3. Use BLAST to find homologous sequences 4. Multiple sequence alignments (MSAs) 5. Trees quantify sequence
Genomics and bioinformatics summary 1. Gene finding: computer searches, cdnas, ESTs, 2. Microarrays 3. Use BLAST to find homologous sequences 4. Multiple sequence alignments (MSAs) 5. Trees quantify sequence
10-810: Advanced Algorithms and Models for Computational Biology. microrna and Whole Genome Comparison
 10-810: Advanced Algorithms and Models for Computational Biology microrna and Whole Genome Comparison Central Dogma: 90s Transcription factors DNA transcription mrna translation Proteins Central Dogma:
10-810: Advanced Algorithms and Models for Computational Biology microrna and Whole Genome Comparison Central Dogma: 90s Transcription factors DNA transcription mrna translation Proteins Central Dogma:
Computational Structural Bioinformatics
 Computational Structural Bioinformatics ECS129 Instructor: Patrice Koehl http://koehllab.genomecenter.ucdavis.edu/teaching/ecs129 koehl@cs.ucdavis.edu Learning curve Math / CS Biology/ Chemistry Pre-requisite
Computational Structural Bioinformatics ECS129 Instructor: Patrice Koehl http://koehllab.genomecenter.ucdavis.edu/teaching/ecs129 koehl@cs.ucdavis.edu Learning curve Math / CS Biology/ Chemistry Pre-requisite
BSC 4934: QʼBIC Capstone Workshop" Giri Narasimhan. ECS 254A; Phone: x3748
 BSC 4934: QʼBIC Capstone Workshop" Giri Narasimhan ECS 254A; Phone: x3748 giri@cs.fiu.edu http://www.cs.fiu.edu/~giri/teach/bsc4934_su10.html July 2010 7/12/10 Q'BIC Bioinformatics 1 Overview of Course"
BSC 4934: QʼBIC Capstone Workshop" Giri Narasimhan ECS 254A; Phone: x3748 giri@cs.fiu.edu http://www.cs.fiu.edu/~giri/teach/bsc4934_su10.html July 2010 7/12/10 Q'BIC Bioinformatics 1 Overview of Course"
Supplementary text for the section Interactions conserved across species: can one select the conserved interactions?
 1 Supporting Information: What Evidence is There for the Homology of Protein-Protein Interactions? Anna C. F. Lewis, Nick S. Jones, Mason A. Porter, Charlotte M. Deane Supplementary text for the section
1 Supporting Information: What Evidence is There for the Homology of Protein-Protein Interactions? Anna C. F. Lewis, Nick S. Jones, Mason A. Porter, Charlotte M. Deane Supplementary text for the section
Introduction to protein alignments
 Introduction to protein alignments Comparative Analysis of Proteins Experimental evidence from one or more proteins can be used to infer function of related protein(s). Gene A Gene X Protein A compare
Introduction to protein alignments Comparative Analysis of Proteins Experimental evidence from one or more proteins can be used to infer function of related protein(s). Gene A Gene X Protein A compare
Lecture Materials are available on the 321 web site
 MENDEL AND MODELS Explore the Course Web Site http://fire.biol.wwu.edu/trent/trent/biol321index.html Lecture Materials are available on the 321 web site http://fire.biol.wwu.edu/trent/trent/321lectureindex.html
MENDEL AND MODELS Explore the Course Web Site http://fire.biol.wwu.edu/trent/trent/biol321index.html Lecture Materials are available on the 321 web site http://fire.biol.wwu.edu/trent/trent/321lectureindex.html
11/24/13. Science, then, and now. Computational Structural Bioinformatics. Learning curve. ECS129 Instructor: Patrice Koehl
 Computational Structural Bioinformatics ECS129 Instructor: Patrice Koehl http://www.cs.ucdavis.edu/~koehl/teaching/ecs129/index.html koehl@cs.ucdavis.edu Learning curve Math / CS Biology/ Chemistry Pre-requisite
Computational Structural Bioinformatics ECS129 Instructor: Patrice Koehl http://www.cs.ucdavis.edu/~koehl/teaching/ecs129/index.html koehl@cs.ucdavis.edu Learning curve Math / CS Biology/ Chemistry Pre-requisite
Science of Information Initiative
 Science of Information Initiative Wojciech Szpankowski and Ananth Grama... to integrate research and teaching activities aimed at investigating the role of information from various viewpoints: from fundamental
Science of Information Initiative Wojciech Szpankowski and Ananth Grama... to integrate research and teaching activities aimed at investigating the role of information from various viewpoints: from fundamental
BLAST Database Searching. BME 110: CompBio Tools Todd Lowe April 8, 2010
 BLAST Database Searching BME 110: CompBio Tools Todd Lowe April 8, 2010 Admin Reading: Read chapter 7, and the NCBI Blast Guide and tutorial http://www.ncbi.nlm.nih.gov/blast/why.shtml Read Chapter 8 for
BLAST Database Searching BME 110: CompBio Tools Todd Lowe April 8, 2010 Admin Reading: Read chapter 7, and the NCBI Blast Guide and tutorial http://www.ncbi.nlm.nih.gov/blast/why.shtml Read Chapter 8 for
Related Courses He who asks is a fool for five minutes, but he who does not ask remains a fool forever.
 CSE 527 Computational Biology http://www.cs.washington.edu/527 Lecture 1: Overview & Bio Review Autumn 2004 Larry Ruzzo Related Courses He who asks is a fool for five minutes, but he who does not ask remains
CSE 527 Computational Biology http://www.cs.washington.edu/527 Lecture 1: Overview & Bio Review Autumn 2004 Larry Ruzzo Related Courses He who asks is a fool for five minutes, but he who does not ask remains
Gene mention normalization in full texts using GNAT and LINNAEUS
 Gene mention normalization in full texts using GNAT and LINNAEUS Illés Solt 1,2, Martin Gerner 3, Philippe Thomas 2, Goran Nenadic 4, Casey M. Bergman 3, Ulf Leser 2, Jörg Hakenberg 5 1 Department of Telecommunications
Gene mention normalization in full texts using GNAT and LINNAEUS Illés Solt 1,2, Martin Gerner 3, Philippe Thomas 2, Goran Nenadic 4, Casey M. Bergman 3, Ulf Leser 2, Jörg Hakenberg 5 1 Department of Telecommunications
SUPPLEMENTARY INFORMATION
 Fig. 1 Influences of crystal lattice contacts on Pol η structures. a. The dominant lattice contact between two hpol η molecules (silver and gold) in the type 1 crystals. b. A close-up view of the hydrophobic
Fig. 1 Influences of crystal lattice contacts on Pol η structures. a. The dominant lattice contact between two hpol η molecules (silver and gold) in the type 1 crystals. b. A close-up view of the hydrophobic
Small RNA in rice genome
 Vol. 45 No. 5 SCIENCE IN CHINA (Series C) October 2002 Small RNA in rice genome WANG Kai ( 1, ZHU Xiaopeng ( 2, ZHONG Lan ( 1,3 & CHEN Runsheng ( 1,2 1. Beijing Genomics Institute/Center of Genomics and
Vol. 45 No. 5 SCIENCE IN CHINA (Series C) October 2002 Small RNA in rice genome WANG Kai ( 1, ZHU Xiaopeng ( 2, ZHONG Lan ( 1,3 & CHEN Runsheng ( 1,2 1. Beijing Genomics Institute/Center of Genomics and
BLAST. Varieties of BLAST
 BLAST Basic Local Alignment Search Tool (1990) Altschul, Gish, Miller, Myers, & Lipman Uses short-cuts or heuristics to improve search speed Like speed-reading, does not examine every nucleotide of database
BLAST Basic Local Alignment Search Tool (1990) Altschul, Gish, Miller, Myers, & Lipman Uses short-cuts or heuristics to improve search speed Like speed-reading, does not examine every nucleotide of database
Pyrobayes: an improved base caller for SNP discovery in pyrosequences
 Pyrobayes: an improved base caller for SNP discovery in pyrosequences Aaron R Quinlan, Donald A Stewart, Michael P Strömberg & Gábor T Marth Supplementary figures and text: Supplementary Figure 1. The
Pyrobayes: an improved base caller for SNP discovery in pyrosequences Aaron R Quinlan, Donald A Stewart, Michael P Strömberg & Gábor T Marth Supplementary figures and text: Supplementary Figure 1. The
Sequence Database Search Techniques I: Blast and PatternHunter tools
 Sequence Database Search Techniques I: Blast and PatternHunter tools Zhang Louxin National University of Singapore Outline. Database search 2. BLAST (and filtration technique) 3. PatternHunter (empowered
Sequence Database Search Techniques I: Blast and PatternHunter tools Zhang Louxin National University of Singapore Outline. Database search 2. BLAST (and filtration technique) 3. PatternHunter (empowered
You are required to know all terms defined in lecture. EXPLORE THE COURSE WEB SITE 1/6/2010 MENDEL AND MODELS
 1/6/2010 MENDEL AND MODELS!!! GENETIC TERMINOLOGY!!! Essential to the mastery of genetics is a thorough knowledge and understanding of the vocabulary of this science. New terms will be introduced and defined
1/6/2010 MENDEL AND MODELS!!! GENETIC TERMINOLOGY!!! Essential to the mastery of genetics is a thorough knowledge and understanding of the vocabulary of this science. New terms will be introduced and defined
Heuristic Methods. Heuristic methods for alignment Sequence databases Multiple alignment Gene and protein prediction
 Heuristic methods for alignment Sequence databases Multiple alignment Gene and protein prediction Armstrong, 2010 Heuristic Methods! FASTA! BLAST! Gapped BLAST! PSI-BLAST Armstrong, 2010 1 Assumptions
Heuristic methods for alignment Sequence databases Multiple alignment Gene and protein prediction Armstrong, 2010 Heuristic Methods! FASTA! BLAST! Gapped BLAST! PSI-BLAST Armstrong, 2010 1 Assumptions
2 The Proteome. The Proteome 15
 The Proteome 15 2 The Proteome 2.1. The Proteome and the Genome Each of our cells contains all the information necessary to make a complete human being. However, not all the genes are expressed in all
The Proteome 15 2 The Proteome 2.1. The Proteome and the Genome Each of our cells contains all the information necessary to make a complete human being. However, not all the genes are expressed in all
CISC 636 Computational Biology & Bioinformatics (Fall 2016)
 CISC 636 Computational Biology & Bioinformatics (Fall 2016) Predicting Protein-Protein Interactions CISC636, F16, Lec22, Liao 1 Background Proteins do not function as isolated entities. Protein-Protein
CISC 636 Computational Biology & Bioinformatics (Fall 2016) Predicting Protein-Protein Interactions CISC636, F16, Lec22, Liao 1 Background Proteins do not function as isolated entities. Protein-Protein
Multiple Sequence Alignments
 Multiple Sequence Alignments...... Elements of Bioinformatics Spring, 2003 Tom Carter http://astarte.csustan.edu/ tom/ March, 2003 1 Sequence Alignments Often, we would like to make direct comparisons
Multiple Sequence Alignments...... Elements of Bioinformatics Spring, 2003 Tom Carter http://astarte.csustan.edu/ tom/ March, 2003 1 Sequence Alignments Often, we would like to make direct comparisons
Hands-On Nine The PAX6 Gene and Protein
 Hands-On Nine The PAX6 Gene and Protein Main Purpose of Hands-On Activity: Using bioinformatics tools to examine the sequences, homology, and disease relevance of the Pax6: a master gene of eye formation.
Hands-On Nine The PAX6 Gene and Protein Main Purpose of Hands-On Activity: Using bioinformatics tools to examine the sequences, homology, and disease relevance of the Pax6: a master gene of eye formation.
The Eukaryotic Genome and Its Expression. The Eukaryotic Genome and Its Expression. A. The Eukaryotic Genome. Lecture Series 11
 The Eukaryotic Genome and Its Expression Lecture Series 11 The Eukaryotic Genome and Its Expression A. The Eukaryotic Genome B. Repetitive Sequences (rem: teleomeres) C. The Structures of Protein-Coding
The Eukaryotic Genome and Its Expression Lecture Series 11 The Eukaryotic Genome and Its Expression A. The Eukaryotic Genome B. Repetitive Sequences (rem: teleomeres) C. The Structures of Protein-Coding
Unit 5: Cell Division and Development Guided Reading Questions (45 pts total)
 Name: AP Biology Biology, Campbell and Reece, 7th Edition Adapted from chapter reading guides originally created by Lynn Miriello Chapter 12 The Cell Cycle Unit 5: Cell Division and Development Guided
Name: AP Biology Biology, Campbell and Reece, 7th Edition Adapted from chapter reading guides originally created by Lynn Miriello Chapter 12 The Cell Cycle Unit 5: Cell Division and Development Guided
Comparing Genomes! Homologies and Families! Sequence Alignments!
 Comparing Genomes! Homologies and Families! Sequence Alignments! Allows us to achieve a greater understanding of vertebrate evolution! Tells us what is common and what is unique between different species
Comparing Genomes! Homologies and Families! Sequence Alignments! Allows us to achieve a greater understanding of vertebrate evolution! Tells us what is common and what is unique between different species
Introduction to Bioinformatics
 CSCI8980: Applied Machine Learning in Computational Biology Introduction to Bioinformatics Rui Kuang Department of Computer Science and Engineering University of Minnesota kuang@cs.umn.edu History of Bioinformatics
CSCI8980: Applied Machine Learning in Computational Biology Introduction to Bioinformatics Rui Kuang Department of Computer Science and Engineering University of Minnesota kuang@cs.umn.edu History of Bioinformatics
Proteomics. Yeast two hybrid. Proteomics - PAGE techniques. Data obtained. What is it?
 Proteomics What is it? Reveal protein interactions Protein profiling in a sample Yeast two hybrid screening High throughput 2D PAGE Automatic analysis of 2D Page Yeast two hybrid Use two mating strains
Proteomics What is it? Reveal protein interactions Protein profiling in a sample Yeast two hybrid screening High throughput 2D PAGE Automatic analysis of 2D Page Yeast two hybrid Use two mating strains
- conserved in Eukaryotes. - proteins in the cluster have identifiable conserved domains. - human gene should be included in the cluster.
 NCBI BLAST Services DELTA-BLAST BLAST (http://blast.ncbi.nlm.nih.gov/), Basic Local Alignment Search tool, is a suite of programs for finding similarities between biological sequences. DELTA-BLAST is a
NCBI BLAST Services DELTA-BLAST BLAST (http://blast.ncbi.nlm.nih.gov/), Basic Local Alignment Search tool, is a suite of programs for finding similarities between biological sequences. DELTA-BLAST is a
COMPARING DNA SEQUENCES TO UNDERSTAND EVOLUTIONARY RELATIONSHIPS WITH BLAST
 Big Idea 1 Evolution INVESTIGATION 3 COMPARING DNA SEQUENCES TO UNDERSTAND EVOLUTIONARY RELATIONSHIPS WITH BLAST How can bioinformatics be used as a tool to determine evolutionary relationships and to
Big Idea 1 Evolution INVESTIGATION 3 COMPARING DNA SEQUENCES TO UNDERSTAND EVOLUTIONARY RELATIONSHIPS WITH BLAST How can bioinformatics be used as a tool to determine evolutionary relationships and to
6.096 Algorithms for Computational Biology. Prof. Manolis Kellis
 6.096 Algorithms for Computational Biology Prof. Manolis Kellis Today s Goals Introduction Class introduction Challenges in Computational Biology Gene Regulation: Regulatory Motif Discovery Exhaustive
6.096 Algorithms for Computational Biology Prof. Manolis Kellis Today s Goals Introduction Class introduction Challenges in Computational Biology Gene Regulation: Regulatory Motif Discovery Exhaustive
Investigation 3: Comparing DNA Sequences to Understand Evolutionary Relationships with BLAST
 Investigation 3: Comparing DNA Sequences to Understand Evolutionary Relationships with BLAST Introduction Bioinformatics is a powerful tool which can be used to determine evolutionary relationships and
Investigation 3: Comparing DNA Sequences to Understand Evolutionary Relationships with BLAST Introduction Bioinformatics is a powerful tool which can be used to determine evolutionary relationships and
Introduction to Bioinformatics Integrated Science, 11/9/05
 1 Introduction to Bioinformatics Integrated Science, 11/9/05 Morris Levy Biological Sciences Research: Evolutionary Ecology, Plant- Fungal Pathogen Interactions Coordinator: BIOL 495S/CS490B/STAT490B Introduction
1 Introduction to Bioinformatics Integrated Science, 11/9/05 Morris Levy Biological Sciences Research: Evolutionary Ecology, Plant- Fungal Pathogen Interactions Coordinator: BIOL 495S/CS490B/STAT490B Introduction
CRISPRseek Workshop Design of target-specific guide RNAs in CRISPR-Cas9 genome-editing systems
 April 2008 CRISPRseek Workshop Design of target-specific guide RNAs in CRISPR-Cas9 genome-editing systems Lihua Julie Zhu Sept 10th 2014 Michael Brodsky Jianhong Ou INSTALLATION First install R 3.1.0 Windows:
April 2008 CRISPRseek Workshop Design of target-specific guide RNAs in CRISPR-Cas9 genome-editing systems Lihua Julie Zhu Sept 10th 2014 Michael Brodsky Jianhong Ou INSTALLATION First install R 3.1.0 Windows:
Lecture 10: Cyclins, cyclin kinases and cell division
 Chem*3560 Lecture 10: Cyclins, cyclin kinases and cell division The eukaryotic cell cycle Actively growing mammalian cells divide roughly every 24 hours, and follow a precise sequence of events know as
Chem*3560 Lecture 10: Cyclins, cyclin kinases and cell division The eukaryotic cell cycle Actively growing mammalian cells divide roughly every 24 hours, and follow a precise sequence of events know as
Comparative Features of Multicellular Eukaryotic Genomes
 Comparative Features of Multicellular Eukaryotic Genomes C elegans A thaliana O. Sativa D. melanogaster M. musculus H. sapiens Size (Mb) 97 115 389 120 2500 2900 # Genes 18,425 25,498 37,544 13,601 30,000
Comparative Features of Multicellular Eukaryotic Genomes C elegans A thaliana O. Sativa D. melanogaster M. musculus H. sapiens Size (Mb) 97 115 389 120 2500 2900 # Genes 18,425 25,498 37,544 13,601 30,000
Introduction of Biotechnology
 Handai Cyber University Introduction of Biotechnology No.5:Genome biotechnology and yeast Graduate School of Engineering, Osaka University Graduate School of Information Science, Osaka University 1 International
Handai Cyber University Introduction of Biotechnology No.5:Genome biotechnology and yeast Graduate School of Engineering, Osaka University Graduate School of Information Science, Osaka University 1 International
Ensembl focuses on metazoan (animal) genomes. The genomes currently available at the Ensembl site are:
 Comparative genomics and proteomics Species available Ensembl focuses on metazoan (animal) genomes. The genomes currently available at the Ensembl site are: Vertebrates: human, chimpanzee, mouse, rat,
Comparative genomics and proteomics Species available Ensembl focuses on metazoan (animal) genomes. The genomes currently available at the Ensembl site are: Vertebrates: human, chimpanzee, mouse, rat,
Genome Sequences and Evolution
 Jones & Bartlett Learning LLC, an Ascend Learning Company.. Chapter 5 Genome Sequences and Evolution Chapter Outline 5.1 Introduction 5.2 5.3 Prokaryotic Gene Numbers Range Over an Order of Magnitude Total
Jones & Bartlett Learning LLC, an Ascend Learning Company.. Chapter 5 Genome Sequences and Evolution Chapter Outline 5.1 Introduction 5.2 5.3 Prokaryotic Gene Numbers Range Over an Order of Magnitude Total
Bioinformatics for Biologists
 Bioinformatics for Biologists Sequence Analysis: Part I. Pairwise alignment and database searching Fran Lewitter, Ph.D. Head, Biocomputing Whitehead Institute Bioinformatics Definitions The use of computational
Bioinformatics for Biologists Sequence Analysis: Part I. Pairwise alignment and database searching Fran Lewitter, Ph.D. Head, Biocomputing Whitehead Institute Bioinformatics Definitions The use of computational
Inparanoid: a comprehensive database of eukaryotic orthologs
 D476 D480 Nucleic Acids Research, 2005, Vol. 33, Database issue doi:10.1093/nar/gki107 Inparanoid: a comprehensive database of eukaryotic orthologs Kevin P. O Brien, Maido Remm 1 and Erik L. L. Sonnhammer*
D476 D480 Nucleic Acids Research, 2005, Vol. 33, Database issue doi:10.1093/nar/gki107 Inparanoid: a comprehensive database of eukaryotic orthologs Kevin P. O Brien, Maido Remm 1 and Erik L. L. Sonnhammer*
Bio2. Heuristics, Databases ; Multiple Sequence Alignment ; Gene Finding. Biological Databases (sequences) Armstrong, 2007 Bioinformatics 2
 Bio2 Heuristics, Databases ; Multiple Sequence Alignment ; Gene Finding Biological Databases (sequences) 1 Biological Databases Introduction to Sequence Databases Overview of primary query tools and the
Bio2 Heuristics, Databases ; Multiple Sequence Alignment ; Gene Finding Biological Databases (sequences) 1 Biological Databases Introduction to Sequence Databases Overview of primary query tools and the
Protein function prediction based on sequence analysis
 Performing sequence searches Post-Blast analysis, Using profiles and pattern-matching Protein function prediction based on sequence analysis Slides from a lecture on MOL204 - Applied Bioinformatics 18-Oct-2005
Performing sequence searches Post-Blast analysis, Using profiles and pattern-matching Protein function prediction based on sequence analysis Slides from a lecture on MOL204 - Applied Bioinformatics 18-Oct-2005
Basic Local Alignment Search Tool
 Basic Local Alignment Search Tool Alignments used to uncover homologies between sequences combined with phylogenetic studies o can determine orthologous and paralogous relationships Local Alignment uses
Basic Local Alignment Search Tool Alignments used to uncover homologies between sequences combined with phylogenetic studies o can determine orthologous and paralogous relationships Local Alignment uses
Prediction of Protein-protein Interactions on the Basis of Evolutionary Conservation of Protein Functions
 Prediction of Protein-protein Interactions on the Basis of Evolutionary Conservation of Protein Functions Ekaterina Kotelnikova 1, Andrey Kalinin 1, Anton Yuryev 1, and Sergei Maslov 2 1 Ariadne Genomics
Prediction of Protein-protein Interactions on the Basis of Evolutionary Conservation of Protein Functions Ekaterina Kotelnikova 1, Andrey Kalinin 1, Anton Yuryev 1, and Sergei Maslov 2 1 Ariadne Genomics
BIOINFORMATICS. Improved Network-based Identification of Protein Orthologs. Nir Yosef a,, Roded Sharan a and William Stafford Noble b
 BIOINFORMATICS Vol. no. 28 Pages 7 Improved Network-based Identification of Protein Orthologs Nir Yosef a,, Roded Sharan a and William Stafford Noble b a School of Computer Science, Tel-Aviv University,
BIOINFORMATICS Vol. no. 28 Pages 7 Improved Network-based Identification of Protein Orthologs Nir Yosef a,, Roded Sharan a and William Stafford Noble b a School of Computer Science, Tel-Aviv University,
Bioinformatics 2. Yeast two hybrid. Proteomics. Proteomics
 GENOME Bioinformatics 2 Proteomics protein-gene PROTEOME protein-protein METABOLISM Slide from http://www.nd.edu/~networks/ Citrate Cycle Bio-chemical reactions What is it? Proteomics Reveal protein Protein
GENOME Bioinformatics 2 Proteomics protein-gene PROTEOME protein-protein METABOLISM Slide from http://www.nd.edu/~networks/ Citrate Cycle Bio-chemical reactions What is it? Proteomics Reveal protein Protein
NUCLEOTIDE SUBSTITUTIONS AND THE EVOLUTION OF DUPLICATE GENES
 Conery, J.S. and Lynch, M. Nucleotide substitutions and evolution of duplicate genes. Pacific Symposium on Biocomputing 6:167-178 (2001). NUCLEOTIDE SUBSTITUTIONS AND THE EVOLUTION OF DUPLICATE GENES JOHN
Conery, J.S. and Lynch, M. Nucleotide substitutions and evolution of duplicate genes. Pacific Symposium on Biocomputing 6:167-178 (2001). NUCLEOTIDE SUBSTITUTIONS AND THE EVOLUTION OF DUPLICATE GENES JOHN
Annotation of Drosophila grimashawi Contig12
 Annotation of Drosophila grimashawi Contig12 Marshall Strother April 27, 2009 Contents 1 Overview 3 2 Genes 3 2.1 Genscan Feature 12.4............................................. 3 2.1.1 Genome Browser:
Annotation of Drosophila grimashawi Contig12 Marshall Strother April 27, 2009 Contents 1 Overview 3 2 Genes 3 2.1 Genscan Feature 12.4............................................. 3 2.1.1 Genome Browser:
Systematic prediction of gene function in Arabidopsis thaliana using a probabilistic functional gene network
 Systematic prediction of gene function in Arabidopsis thaliana using a probabilistic functional gene network Sohyun Hwang 1, Seung Y Rhee 2, Edward M Marcotte 3,4 & Insuk Lee 1 protocol 1 Department of
Systematic prediction of gene function in Arabidopsis thaliana using a probabilistic functional gene network Sohyun Hwang 1, Seung Y Rhee 2, Edward M Marcotte 3,4 & Insuk Lee 1 protocol 1 Department of
Peeling the yeast protein network
 444 DOI 10.1002/pmic.200400962 Proteomics 2005, 5, 444 449 REGULAR ARTICLE Peeling the yeast protein network Stefan Wuchty and Eivind Almaas Department of Physics, University of Notre Dame, Notre Dame,
444 DOI 10.1002/pmic.200400962 Proteomics 2005, 5, 444 449 REGULAR ARTICLE Peeling the yeast protein network Stefan Wuchty and Eivind Almaas Department of Physics, University of Notre Dame, Notre Dame,
In Search of the Biological Significance of Modular Structures in Protein Networks
 In Search of the Biological Significance of Modular Structures in Protein Networks Zhi Wang, Jianzhi Zhang * Department of Ecology and Evolutionary Biology, University of Michigan, Ann Arbor, Michigan,
In Search of the Biological Significance of Modular Structures in Protein Networks Zhi Wang, Jianzhi Zhang * Department of Ecology and Evolutionary Biology, University of Michigan, Ann Arbor, Michigan,
Universal Rules Governing Genome Evolution Expressed by Linear Formulas
 The Open Genomics Journal, 8, 1, 33-3 33 Open Access Universal Rules Governing Genome Evolution Expressed by Linear Formulas Kenji Sorimachi *,1 and Teiji Okayasu 1 Educational Support Center and Center
The Open Genomics Journal, 8, 1, 33-3 33 Open Access Universal Rules Governing Genome Evolution Expressed by Linear Formulas Kenji Sorimachi *,1 and Teiji Okayasu 1 Educational Support Center and Center
GEP Annotation Report
 GEP Annotation Report Note: For each gene described in this annotation report, you should also prepare the corresponding GFF, transcript and peptide sequence files as part of your submission. Student name:
GEP Annotation Report Note: For each gene described in this annotation report, you should also prepare the corresponding GFF, transcript and peptide sequence files as part of your submission. Student name:
Fundamentally different strategies for transcriptional regulation are revealed by information-theoretical analysis of binding motifs
 Fundamentally different strategies for transcriptional regulation are revealed by information-theoretical analysis of binding motifs Zeba Wunderlich 1* and Leonid A. Mirny 1,2 1 Biophysics Program, Harvard
Fundamentally different strategies for transcriptional regulation are revealed by information-theoretical analysis of binding motifs Zeba Wunderlich 1* and Leonid A. Mirny 1,2 1 Biophysics Program, Harvard
Comparison of Protein-Protein Interaction Confidence Assignment Schemes
 Comparison of Protein-Protein Interaction Confidence Assignment Schemes Silpa Suthram 1, Tomer Shlomi 2, Eytan Ruppin 2, Roded Sharan 2, and Trey Ideker 1 1 Department of Bioengineering, University of
Comparison of Protein-Protein Interaction Confidence Assignment Schemes Silpa Suthram 1, Tomer Shlomi 2, Eytan Ruppin 2, Roded Sharan 2, and Trey Ideker 1 1 Department of Bioengineering, University of
Reminders about Eukaryotes
 BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 16: Eukaryotes at last! http://compbio.uchsc.edu/hunter/bio5099 Larry.Hunter@uchsc.edu Reminders about Eukaryotes Eukaryotes arose around
BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 16: Eukaryotes at last! http://compbio.uchsc.edu/hunter/bio5099 Larry.Hunter@uchsc.edu Reminders about Eukaryotes Eukaryotes arose around
Chapter 18 Lecture. Concepts of Genetics. Tenth Edition. Developmental Genetics
 Chapter 18 Lecture Concepts of Genetics Tenth Edition Developmental Genetics Chapter Contents 18.1 Differentiated States Develop from Coordinated Programs of Gene Expression 18.2 Evolutionary Conservation
Chapter 18 Lecture Concepts of Genetics Tenth Edition Developmental Genetics Chapter Contents 18.1 Differentiated States Develop from Coordinated Programs of Gene Expression 18.2 Evolutionary Conservation
7.06 Problem Set #4, Spring 2005
 7.06 Problem Set #4, Spring 2005 1. You re doing a mutant hunt in S. cerevisiae (budding yeast), looking for temperaturesensitive mutants that are defective in the cell cycle. You discover a mutant strain
7.06 Problem Set #4, Spring 2005 1. You re doing a mutant hunt in S. cerevisiae (budding yeast), looking for temperaturesensitive mutants that are defective in the cell cycle. You discover a mutant strain
Genome Sequencing & DNA Sequence Analysis
 7.91 / 7.36 / BE.490 Lecture #1 Feb. 24, 2004 Genome Sequencing & DNA Sequence Analysis Chris Burge What is a Genome? A genome is NOT a bag of proteins What s in the Human Genome? Outline of Unit II: DNA/RNA
7.91 / 7.36 / BE.490 Lecture #1 Feb. 24, 2004 Genome Sequencing & DNA Sequence Analysis Chris Burge What is a Genome? A genome is NOT a bag of proteins What s in the Human Genome? Outline of Unit II: DNA/RNA
The Gene The gene; Genes Genes Allele;
 Gene, genetic code and regulation of the gene expression, Regulating the Metabolism, The Lac- Operon system,catabolic repression, The Trp Operon system: regulating the biosynthesis of the tryptophan. Mitesh
Gene, genetic code and regulation of the gene expression, Regulating the Metabolism, The Lac- Operon system,catabolic repression, The Trp Operon system: regulating the biosynthesis of the tryptophan. Mitesh
Network alignment and querying
 Network biology minicourse (part 4) Algorithmic challenges in genomics Network alignment and querying Roded Sharan School of Computer Science, Tel Aviv University Multiple Species PPI Data Rapid growth
Network biology minicourse (part 4) Algorithmic challenges in genomics Network alignment and querying Roded Sharan School of Computer Science, Tel Aviv University Multiple Species PPI Data Rapid growth
Evolution and Development Evo-Devo
 Evolution and Development Evo-Devo Darwin wrote a book on barnacles. Plate 1 (Lepas), from A monograph on the sub-class Cirripedia, by Charles Darwin. Comparative embryology There is an obvious similarity
Evolution and Development Evo-Devo Darwin wrote a book on barnacles. Plate 1 (Lepas), from A monograph on the sub-class Cirripedia, by Charles Darwin. Comparative embryology There is an obvious similarity
G C T G U A. template strand
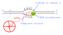 A 5 w 3 3 c 5 mrna RNA polymerase T G C A G C C T G U A coding or sense st AUG?? template strand The triplet code 3 bases = 1 amino acid Punctuation: sta rt: sto p: AUGA CUU CAGUAAC AU U AA C C AUG = methionine,
A 5 w 3 3 c 5 mrna RNA polymerase T G C A G C C T G U A coding or sense st AUG?? template strand The triplet code 3 bases = 1 amino acid Punctuation: sta rt: sto p: AUGA CUU CAGUAAC AU U AA C C AUG = methionine,
Example of Function Prediction
 Find similar genes Example of Function Prediction Suggesting functions of newly identified genes It was known that mutations of NF1 are associated with inherited disease neurofibromatosis 1; but little
Find similar genes Example of Function Prediction Suggesting functions of newly identified genes It was known that mutations of NF1 are associated with inherited disease neurofibromatosis 1; but little
Tools and Algorithms in Bioinformatics
 Tools and Algorithms in Bioinformatics GCBA815, Fall 2013 Week3: Blast Algorithm, theory and practice Babu Guda, Ph.D. Department of Genetics, Cell Biology & Anatomy Bioinformatics and Systems Biology
Tools and Algorithms in Bioinformatics GCBA815, Fall 2013 Week3: Blast Algorithm, theory and practice Babu Guda, Ph.D. Department of Genetics, Cell Biology & Anatomy Bioinformatics and Systems Biology
GATA family of transcription factors of vertebrates: phylogenetics and chromosomal synteny
 Phylogenetics and chromosomal synteny of the GATAs 1273 GATA family of transcription factors of vertebrates: phylogenetics and chromosomal synteny CHUNJIANG HE, HANHUA CHENG* and RONGJIA ZHOU* Department
Phylogenetics and chromosomal synteny of the GATAs 1273 GATA family of transcription factors of vertebrates: phylogenetics and chromosomal synteny CHUNJIANG HE, HANHUA CHENG* and RONGJIA ZHOU* Department
Proteins: Structure & Function. Ulf Leser
 Proteins: Structure & Function Ulf Leser This Lecture Proteins Structure Function Databases Predicting Protein Secondary Structure Many figures from Zvelebil, M. and Baum, J. O. (2008). "Understanding
Proteins: Structure & Function Ulf Leser This Lecture Proteins Structure Function Databases Predicting Protein Secondary Structure Many figures from Zvelebil, M. and Baum, J. O. (2008). "Understanding
08/21/2017 BLAST. Multiple Sequence Alignments: Clustal Omega
 BLAST Multiple Sequence Alignments: Clustal Omega What does basic BLAST do (e.g. what is input sequence and how does BLAST look for matches?) Susan Parrish McDaniel College Multiple Sequence Alignments
BLAST Multiple Sequence Alignments: Clustal Omega What does basic BLAST do (e.g. what is input sequence and how does BLAST look for matches?) Susan Parrish McDaniel College Multiple Sequence Alignments
RELATIONSHIPS BETWEEN GENES/PROTEINS HOMOLOGUES
 Molecular Biology-2018 1 Definitions: RELATIONSHIPS BETWEEN GENES/PROTEINS HOMOLOGUES Heterologues: Genes or proteins that possess different sequences and activities. Homologues: Genes or proteins that
Molecular Biology-2018 1 Definitions: RELATIONSHIPS BETWEEN GENES/PROTEINS HOMOLOGUES Heterologues: Genes or proteins that possess different sequences and activities. Homologues: Genes or proteins that
Understanding Science Through the Lens of Computation. Richard M. Karp Nov. 3, 2007
 Understanding Science Through the Lens of Computation Richard M. Karp Nov. 3, 2007 The Computational Lens Exposes the computational nature of natural processes and provides a language for their description.
Understanding Science Through the Lens of Computation Richard M. Karp Nov. 3, 2007 The Computational Lens Exposes the computational nature of natural processes and provides a language for their description.
Multiple Sequence Alignment. Sequences
 Multiple Sequence Alignment Sequences > YOR020c mstllksaksivplmdrvlvqrikaqaktasglylpe knveklnqaevvavgpgftdangnkvvpqvkvgdqvl ipqfggstiklgnddevilfrdaeilakiakd > crassa mattvrsvksliplldrvlvqrvkaeaktasgiflpe
Multiple Sequence Alignment Sequences > YOR020c mstllksaksivplmdrvlvqrikaqaktasglylpe knveklnqaevvavgpgftdangnkvvpqvkvgdqvl ipqfggstiklgnddevilfrdaeilakiakd > crassa mattvrsvksliplldrvlvqrvkaeaktasgiflpe
Improved network-based identification of protein orthologs
 BIOINFORMATICS Vol. 24 ECCB 28, pages i2 i26 doi:.93/bioinformatics/btn277 Improved network-based identification of protein orthologs Nir Yosef,, Roded Sharan and William Stafford Noble 2,3 School of Computer
BIOINFORMATICS Vol. 24 ECCB 28, pages i2 i26 doi:.93/bioinformatics/btn277 Improved network-based identification of protein orthologs Nir Yosef,, Roded Sharan and William Stafford Noble 2,3 School of Computer
Protein function studies: history, current status and future trends
 19 3 2007 6 Chinese Bulletin of Life Sciences Vol. 19, No. 3 Jun., 2007 1004-0374(2007)03-0294-07 ( 100871) Q51A Protein function studies: history, current status and future trends MA Jing, GE Xi, CHANG
19 3 2007 6 Chinese Bulletin of Life Sciences Vol. 19, No. 3 Jun., 2007 1004-0374(2007)03-0294-07 ( 100871) Q51A Protein function studies: history, current status and future trends MA Jing, GE Xi, CHANG
Computational identification and analysis of MADS box genes in Camellia sinensis
 www.bioinformation.net Hypothesis Volume 11(3) Computational identification and analysis of MADS box genes in Camellia sinensis Madhurjya Gogoi*, Sangeeta Borchetia & Tanoy Bandyopadhyay Department of
www.bioinformation.net Hypothesis Volume 11(3) Computational identification and analysis of MADS box genes in Camellia sinensis Madhurjya Gogoi*, Sangeeta Borchetia & Tanoy Bandyopadhyay Department of
Bioinformatics: Network Analysis
 Bioinformatics: Network Analysis Comparative Network Analysis COMP 572 (BIOS 572 / BIOE 564) - Fall 2013 Luay Nakhleh, Rice University 1 Biomolecular Network Components 2 Accumulation of Network Components
Bioinformatics: Network Analysis Comparative Network Analysis COMP 572 (BIOS 572 / BIOE 564) - Fall 2013 Luay Nakhleh, Rice University 1 Biomolecular Network Components 2 Accumulation of Network Components
Arabidopsis genomic information for interpreting wheat EST sequences
 Funct Integr Genomics (2003) 3:33 38 DOI 10.1007/s10142-002-0075-1 REVIEW Bryan Clarke Mark Lambrecht Seung Y. Rhee Arabidopsis genomic information for interpreting wheat EST sequences Received: 12 April
Funct Integr Genomics (2003) 3:33 38 DOI 10.1007/s10142-002-0075-1 REVIEW Bryan Clarke Mark Lambrecht Seung Y. Rhee Arabidopsis genomic information for interpreting wheat EST sequences Received: 12 April
How much non-coding DNA do eukaryotes require?
 How much non-coding DNA do eukaryotes require? Andrei Zinovyev UMR U900 Computational Systems Biology of Cancer Institute Curie/INSERM/Ecole de Mine Paritech Dr. Sebastian Ahnert Dr. Thomas Fink Bioinformatics
How much non-coding DNA do eukaryotes require? Andrei Zinovyev UMR U900 Computational Systems Biology of Cancer Institute Curie/INSERM/Ecole de Mine Paritech Dr. Sebastian Ahnert Dr. Thomas Fink Bioinformatics
The MANTiS Manual. Contents. MANTiS Version 1.1
 The MANTiS Manual MANTiS Version 1.1 Contents Connection to the MANTiS database... 2 Memory settings... 2 Main functionalities... 2 Character Mapping View... 4 Genome content View... 5 Biological processes
The MANTiS Manual MANTiS Version 1.1 Contents Connection to the MANTiS database... 2 Memory settings... 2 Main functionalities... 2 Character Mapping View... 4 Genome content View... 5 Biological processes
Genome-Wide Computational Prediction and Analysis of Core Promoter Elements across Plant Monocots and Dicots
 Genome-Wide Computational Prediction and Analysis of Core Promoter Elements across Plant Monocots and Dicots Sunita Kumari 1, Doreen Ware 1,2 * 1 Cold Spring Harbor Laboratory, Cold Spring Harbor, New
Genome-Wide Computational Prediction and Analysis of Core Promoter Elements across Plant Monocots and Dicots Sunita Kumari 1, Doreen Ware 1,2 * 1 Cold Spring Harbor Laboratory, Cold Spring Harbor, New
Comparative Analysis of Molecular Interaction Networks
 Comparative Analysis of Molecular Interaction Networks Jayesh Pandey 1, Mehmet Koyuturk 2, Shankar Subramaniam 3, Ananth Grama 1 1 Purdue University, 2 Case Western Reserve University 3 University of California,
Comparative Analysis of Molecular Interaction Networks Jayesh Pandey 1, Mehmet Koyuturk 2, Shankar Subramaniam 3, Ananth Grama 1 1 Purdue University, 2 Case Western Reserve University 3 University of California,
MCB142/ICB163 Overview. Instructors:
 MCB142/ICB163 Overview Instructors: Professor Sharon Amacher Professor Abby Dernburg Professor Monty Slatkin GSIs: Kristin Camfield Nat Hallihan Hana Lee Christine Preston Richard Price Kate Smallenburg
MCB142/ICB163 Overview Instructors: Professor Sharon Amacher Professor Abby Dernburg Professor Monty Slatkin GSIs: Kristin Camfield Nat Hallihan Hana Lee Christine Preston Richard Price Kate Smallenburg
Characterization of New Proteins Found by Analysis of Short Open Reading Frames from the Full Yeast Genome
 YEAST VOL. 13: 1363 1374 (1997) Characterization of New Proteins Found by Analysis of Short Open Reading Frames from the Full Yeast Genome MIGUEL A. ANDRADE 1 *, ANTOINE DARUVAR 1, GEORG CASARI 2, REINHARD
YEAST VOL. 13: 1363 1374 (1997) Characterization of New Proteins Found by Analysis of Short Open Reading Frames from the Full Yeast Genome MIGUEL A. ANDRADE 1 *, ANTOINE DARUVAR 1, GEORG CASARI 2, REINHARD
Emergence of gene regulatory networks under functional constraints
 under functional constraints Institute of Physics, Jagiellonian University, Kraków, Poland in collaboration with Zdzislaw Burda, Andre Krzywicki and Olivier C. Martin 30 November 2012, Leipzig, CompPhys12
under functional constraints Institute of Physics, Jagiellonian University, Kraków, Poland in collaboration with Zdzislaw Burda, Andre Krzywicki and Olivier C. Martin 30 November 2012, Leipzig, CompPhys12
Bioinformatics and BLAST
 Bioinformatics and BLAST Overview Recap of last time Similarity discussion Algorithms: Needleman-Wunsch Smith-Waterman BLAST Implementation issues and current research Recap from Last Time Genome consists
Bioinformatics and BLAST Overview Recap of last time Similarity discussion Algorithms: Needleman-Wunsch Smith-Waterman BLAST Implementation issues and current research Recap from Last Time Genome consists
Supplemental Materials
 JOURNAL OF MICROBIOLOGY & BIOLOGY EDUCATION, May 2013, p. 107-109 DOI: http://dx.doi.org/10.1128/jmbe.v14i1.496 Supplemental Materials for Engaging Students in a Bioinformatics Activity to Introduce Gene
JOURNAL OF MICROBIOLOGY & BIOLOGY EDUCATION, May 2013, p. 107-109 DOI: http://dx.doi.org/10.1128/jmbe.v14i1.496 Supplemental Materials for Engaging Students in a Bioinformatics Activity to Introduce Gene
Comparative Bioinformatics Midterm II Fall 2004
 Comparative Bioinformatics Midterm II Fall 2004 Objective Answer, part I: For each of the following, select the single best answer or completion of the phrase. (3 points each) 1. Deinococcus radiodurans
Comparative Bioinformatics Midterm II Fall 2004 Objective Answer, part I: For each of the following, select the single best answer or completion of the phrase. (3 points each) 1. Deinococcus radiodurans
SI Materials and Methods
 SI Materials and Methods Gibbs Sampling with Informative Priors. Full description of the PhyloGibbs algorithm, including comprehensive tests on synthetic and yeast data sets, can be found in Siddharthan
SI Materials and Methods Gibbs Sampling with Informative Priors. Full description of the PhyloGibbs algorithm, including comprehensive tests on synthetic and yeast data sets, can be found in Siddharthan
Bioinformatics Chapter 1. Introduction
 Bioinformatics Chapter 1. Introduction Outline! Biological Data in Digital Symbol Sequences! Genomes Diversity, Size, and Structure! Proteins and Proteomes! On the Information Content of Biological Sequences!
Bioinformatics Chapter 1. Introduction Outline! Biological Data in Digital Symbol Sequences! Genomes Diversity, Size, and Structure! Proteins and Proteomes! On the Information Content of Biological Sequences!
Comparative Gene Expression Analysis by a Differential Clustering Approach: Application to the Candida albicans Transcription Program
 Comparative Gene Expression Analysis by a Differential Clustering Approach: Application to the Candida albicans Transcription Program Jan Ihmels 1[, Sven Bergmann 1,2[, Judith Berman 3, Naama Barkai 1*
Comparative Gene Expression Analysis by a Differential Clustering Approach: Application to the Candida albicans Transcription Program Jan Ihmels 1[, Sven Bergmann 1,2[, Judith Berman 3, Naama Barkai 1*
CSE 427 Comp Bio. Sequence Alignment
 CSE 427 Comp Bio Sequence Alignment 1 Sequence Alignment What Why A Dynamic Programming Algorithm 2 Sequence Alignment Goal: position characters in two strings to best line up identical/similar ones with
CSE 427 Comp Bio Sequence Alignment 1 Sequence Alignment What Why A Dynamic Programming Algorithm 2 Sequence Alignment Goal: position characters in two strings to best line up identical/similar ones with
