Force fields in computer simulation of soft nanomaterials
|
|
|
- Pierce Hardy
- 6 years ago
- Views:
Transcription
1 Lecture 2 Part B Force fields in computer simulation of soft nanomaterials Recommended reading: Leach Chapter 4 1
2 Force Field Methods Force field methods are simulation methods that use classical force fields to describe the interactions between atoms or molecules or particles. (E.g. molecular mechanics and molecular dynamics) Force field methods rely on Born-Oppenheimer, ignore electronic motions and calculate the energy of a system as a function of nuclear positions only. Thus simulations of much larger systems can be performed. Up to billions of atoms possible on the largest supercomputers (nearly a cubic micrometer - beyond that should not need to resolve everything at the atomic or molecular level)! 2
3 Enter Force Field Methods In many cases, just as accurate as QM calculations, but much faster. Also allows statistical phenomena to be explored, and classical and statistical mechanical theories to be explicitly tested. Thus force field methods are not just to save time or if system too large for QM; also to probe qualitatively different processes from what is probed with QM methods. If some phenomena not understood on a fundamental level can be mimicked accurately or even just qualitatively by more coarse-grained methods, no reason to bring out the big guns. 3
4 Force Fields What is a force field? An expression for the interactions between atoms (or molecules, or particles). Force fields rely on: Validity of Born-Oppenheimer approximation. Relatively simple expressions that capture the stretching of bonds, the opening and closing of angles, rotations about bonds Transferability: the ability to apply a given form for a force field to many materials by tweaking parameters (e.g. polystyrene vs. polyethylene) 4
5 Force Field Methods Time Scale (log10 seconds) Simulation methods that use force fields/ interaction potentials Ab Initio/ Electronic Structure MD and MC Explicit atomistic models Particle-based mesoscale simulation Coarse-grained models BD DPD Time and length scales overlap with neutron scattering, NMR, etc. Field-based mesoscale and continuum mechanics Length Scale (log 10 meters) -3 5
6 Force Fields The force can be written in terms of the gradient of the interaction energy (interaction potential): F = "#U({r i }) The term force field refers to the interaction potential U (i.e. the potential field from which forces are derived)! Recall when a force is derivable from the gradient of a scalar potential, the force is conservative. 6
7 Force Fields Many molecular modeling force fields in use today (especially for soft materials) can be written in terms of five simple types of interactions: Bond stretching Non-bonded (electrostatic) Angle bending Bond rotation (torsion) Non-bonded (van der Waals) δ+ δ+ δ- 7
8 U({r i }) = + + $ torsions N $ i, j=1 V n 2 Electrostatic (Coulomb) A typical force field Bond stretch k j l k j # $ 2 (l " l 0 j j ) ( # j "# 0 j ) 2 j (1+ cos(n% " &)) N / q i q ) j (, + $ 4' ij ij 1 r + ij * r. i, j=1 1 0 ij - 12 $ j ) " (, ij + * r. ij - van der Waals (Lennard-Jones) Valence angle bend torsional Intraand Intermolecular Intra- Molecular 8
9 Example: Propane C 3 H 8 has ten bonds: 2 C-C bonds and 8 C-H bonds. The C-H bonds fall into two classes The H s bonded to the central (methylene) carbon The H s bonded to the end (methyl) carbons Some fancier force fields treat these two differently. H H H H-C-C-C-H H H H 18 valence angles & 18 torsional terms 27 non-bonded interactions (if 1,4 s neglected) 73 total terms to calculate for a single molecule: still many fewer than the number of integrals that would be involved in an equivalent ab initio calculation. 9
10 General Features of Force Fields Defining a force field means defining both the functional form and the parameters. Two or more FF s with the same form but different parameters, or with different forms, may give results of comparable accuracy for the same material and property. Typically, FF s are developed to reproduce structures or spectra or thermodynamic quantities. Just because it does one or two things well, doesn t mean it gets everything right! This is not a failure of the FF. A single FF can t do everything, even just for one material. 10
11 General Features of Force Fields Force fields are empirical; there is no one correct form. Although most of the FF s in use today have similar overall forms, that doesn t mean better forms can t be found. FF s are a compromise between accuracy and computational efficiency. We ll see this when we look at the van der Waals term. Forms should be nice since first and second derivatives of the energy wrt atomic coordinates are required in molecular mechanics/molecular dynamics. 11
12 Force Fields Let s delve more deeply into the individual terms that make up a force field (read Ch. 4 in Leach). In the many force fields in use today (MM2, MM3, Amber, Compass, Dreiding, UFF, ), some of these terms are written slightly differently, and treated with different emphasis, and some force fields contain additional terms. When using a force field, it is important to know what is being included and how, and what isn t. 12
13 Bond Stretching U(l) The potential energy curve measured for a typical bond looks like the red curve at the right. l 13
14 Bond Stretching This can be modeled by a Morse potential: { [ ( )]} 2 u(l) = D e 1" exp "a l " l 0 D e = depth of potential energy minimum a = ω (µ/2d e ) µ = reduced mass ω = frequency of bond vibration, related to stretching constant by ω (k/µ) l 0 = reference bond length Rarely used except for when bond-breaking needed*, since exponential is computationally slow. 14
15 Bond Stretching: Approximating Morse Harmonic: u(l) = k ( 2 l " l ) 2 0 Harmonic + higher orders: u(l) = k ( 2 l " l ) 2 0 # [ 1" k' ( l " l ) ] cubic - used in MM2 MM3 uses quartic to fix large l catastrophe 15
16 Bond Stretching Bond l 0 (Å) k (kcal mol -1 Å -2 ) Csp 3 - Csp 3 Csp 3 - Csp 2 Csp 2 - Csp 2 Csp 3 = 0 Csp 3 - Nsp 3 C - N Allinger 1977 (from Leach) 16
17 Bond Angle Bending Interactions The deviation of the angle between two bonds is also often modeled with a harmonic (Hooke s law) potential: u(") = k ( 2 " #" ) 2 0 Angles between different types of bonds have different force constants k and different reference angles θ 0. As with bond stretching terms, higher accuracy is achieved by including higher order terms. 17
18 Bond Angle Bending Interactions Angle θ 0 (deg) k (kcal mol -1 deg -1 ) Csp 3 - Csp 3 - Csp 3 Csp 3 - Csp 3 - H H - Csp 3 - H Csp 3 - Csp 2 - Csp 3 Csp 3 - Csp 2 = Csp 2 Csp 3 - Csp 2 = Allinger 1977 (from Leach) Smaller force constants compared to bonds = much less energy req d to bend than to stretch or compress 18
19 Bond Rotation (Torsional) Interactions Barriers to rotation about chemical bonds arise from antibonding interactions between the H atoms on the (1,4) atoms. Antibonding interactions are minimized when conformation is staggered and maximized when it is eclipsed. In organic macromolecules, torsional potentials are nearly always used for each A-B-C-D since major conformational changes are due to rotations about bonds. In inorganic materials, torsional potentials are often neglected and non-bonded interactions between every pair of (1,4) atoms are tweaked to achieve the desired energy profile. 19
20 Bond Rotation (Torsional) Interactions Torsional energy profile along dihedral angle Si-C-C-C in propyl-poss 6 5 Energy (kcal/mol) From Cummings, et al Rotation Angle 20
21 Bond Rotation (Torsional) Interactions Torsional potentials modeled by a cosine series expansion: u(") = % n= 0 V n 2 (1+ cos(n" # $)) or u(") = # C n cos(") n n= 0 n is multiplicity = number of minima as bond rotated through 360 γ is phase factor: determines where ϕ passes through minimum V n is called barrier height, but not exactly, since non-bonded contributions from (1,4) atoms affect rotation. V n for double bond > V n for single bond 21
22 Bond Rotation (Torsional) Interactions Again, different force fields may use different numbers of terms in cosine series (e.g. Amber contains just one, MM2 uses more). More accurate potentials will use more terms, since the more parameters you have in your FF, the better you can generally fit experimental data. Drawback of using more terms: Many more parameters to fit is more complicated and time consuming. Necessary for some systems. 22
23 Cross Terms b/t Stretch, Bend, Torsion Many force fields also include cross terms, such as: Stretch - stretch Stretch - torsion Stretch - bend Bend - torsion θ θ Bend - bend Again, in general, the more terms you include, the more accurate the FF, but the harder it is to parameterize and use. θ θ 23
24 Coulomb Interactions The distribution of charge in a molecule may be represented by an arrangement of fractional point charges (partial charges). The electrostatic interaction between two molecules or between different parts of the same molecule is calculated as a sum of interactions between pairs of point charges using Coulomb s law: V = N N $ A $ B i=1 j=1 q i q j 4"# 0 r ij 24
25 Coulomb Interactions: multipole expansion Electrostatic interactions can be calculated via a multipole expansion, in which one sums over the individual electric moments or multipoles in an efficient way. Charged molecules have the charge (a monopole) as their lowest non-zero term; uncharged molecules may have the dipole or quadrupole or octupole or as their lowest nonzero term, depending on the asymmetry of the charge distribution within the molecule. V = N N $ A $ B i=1 j=1 q i q j 4"# 0 r ij 25
26 Coulomb Interactions: multipole expansion But, the multipole expansion approach to summing the electrostatic interactions is typically used only for small molecules whose conformations are kept fixed during the calculation, and where the interactions between molecules act at their centers of mass. Leach describes several other approaches to dealing with charges; it is a complicated business, and can be important when studying, e.g., charged macromolecules in aqueous solutions. For many studies, especially of uncharged species, the electrostatic interaction is ignored. 26
27 van der Waals Interactions Electrostatic forces don t account for all the non-bonded interactions in a system. Van der Waals forces describe deviations from ideal gas behavior not accounted for by electrostatics. Johannes Diderik Van der Waals
28 van der Waals Interactions The interaction energy between two isolated Argon atoms has the following dependence on the distance between the atoms: U = 0 at infinite distance, negligible at long distances. As separation is reduced, U decreases, passing thru a minimum at r =.38 nm for Argon. U then rapidly increases as separation decreases further. F ij =-du ij /dr ij 28
29 van der Waals Interactions The VdW curve arises from a balance between long-range attractive forces and short-range repulsive forces. The attractive contribution is due to dispersive forces (London forces), which arise from instantaneous dipoles that arise from fluctuations in the electron density distribution (electron cloud), and which induce dipoles in neighboring atoms. The Drude model predicts that the dispersion interaction varies as 1/r 6, and is negative (attractive). Additional negative, higher order even powers result if higher order induced multipoles are considered. 29
30 van der Waals Interactions The repulsive contribution has a quantum mechanical origin and arises due to the Pauli exchange principle, which prohibits any two electrons in a system from having the same set of quantum numbers. The exchange force prohibits the electrons in the two atoms from occupying the same region of space. This results in a reduced electron density between the two nuclei, which leads to repulsion between the incompletely shielded nuclei. At very small separations, the interaction energy due to nuclear repulsion varies as 1/r. At larger (but still small) separations, the energy decays exponentially as exp(-2r/a 0 ), where a 0 is the Bohr radius. 30
31 van der Waals Interactions The dispersive (attractive) and exchange (repulsive) interactions can be calculated using QM (not trivial, however). There are several empirical models for the van der Waals potential. Required for molecular simulations: a simple expression, because there are MANY non-bonded interactions to include (at least N*(N-1)/2, not counting Coulomb). 31
32 The Lennard-Jones Interaction Potential The most popular model of the van der Waals non-bonded interactions between atoms is the Lennard-Jones model. U(r) $ $ # ' U(r) = 4" && ) %% r ( 12 $ *& # % r ' ) ( 6 ' ) ( Collision diameter σ ε typical cutoff at 2.5σ minimum at r=1.122σ Sir John Edward Lennard-Jones r = r ij 32
33 Lennard-Jones Model The 1/r 6 term in the LJ potential has the same form as that found from theoretical calculations of the Drude model for the dispersive interactions. There are no strong theoretical arguments in support of the 1/r 12 term, which is not a particularly good approximation to 1/r times an exponential. While the 1/r 12 term works well for rare gases, it is usually too steep for hydrocarbons. 33
34 Lennard-Jones Model Nevertheless, the 6-12 LJ potential is widely used, especially for large simulations, since the 1/r 12 is rapidly calculated by squaring the 1/r 6 term, which in turn is found from r 2 without any costly square roots. These are the types of considerations that go into developing force fields! 34
35 Lennard-Jones Model LJ original potential was written in the general form: $ $ U(r) = k" # ' && ) %% r ( Several force fields use n 12. n $ * # ' & ) % r ( m ' ) k = ( n n * m Several formulations in which 1/r 12 term is replaced by a more realistic potential have been proposed, such as the Buckingham potential: % 6 U(r) = "' '# $ 6 e$# & % r ( ' $1 & r m ) * # % $ ' # $ 6& r $ & % r m n m ( * ) 6 ' ) ( ( * * ) m ( n*m) Careful, when separation very small U is steeply attractive. 35
36 Lennard-Jones Model Between different types of atoms, the LJ interaction takes the form: &&% U "# (r) = 4$ "# ) "# (( + '' r * A system with N different types of atoms has N(N-1)/2 different sets of parameters for interactions between unlike atoms. Since vdw parameters are difficult to obtain, we assume parameters for cross interactions between unlike atoms can be obtained from those for like atoms through mixing rules. 12 &, % ) "# ( + ' r * 6 ) + * 36
37 Lorenz-Berthelot Mixing Rules A common mixing rule used to calculate LJ parameters between unlike atoms is the Lorenz-Berthelot mixing rule. Collision diameter σ AB = (σ AA + σ BB )/2 - (arithmetic mean) Well depth ε AB = (ε AA ε BB ) 1/2 - (geometric mean) These mixing rules are most successful between similar types of atoms. Major drawback: well depth can be overestimated by geometric mean. Some force fields also use geometric mean for collision diameter. VdW parameters are typically optimized to reproduce thermodynamic properties such as densities and enthalpies of vaporization, and structure. 37
38 Many Body Effects b/t Non-bonded Atoms In real materials, there are many body effects between non-bonded atoms that cannot be simply described by pairwise forces. The interaction between two atoms can be affected by the presence of other atoms. 38
39 Many Body Effects b/t Non-bonded Atoms Some force fields include three-body terms at the expense of greatly increased computational effort. # of intramolecular terms increases linearly with N # of non-bonded forces increases as N 2 N(N-1)/2 interactions to evaluate in a pair-wise potential. N(N-1)(N-2)/6 unique three-body interactions. For 1000 atom system: 499,500 pairwise terms 166,167,000 three-body terms In gen l, N/3 times more three-body than two-body terms. 39
40 Comparison of Force Fields and QM UFF (Universal Force Field) A.K. Rappe, et al, UFF, a Full Periodic-Table Force-Field for Molecular Mechanics and Molecular Dynamics Simulations, J. Am. Chem. Society, 114, Used for many nanoscale carbon structures (Buckyballs, nanotubes) COMPASS (Condensed-phase Optimized Molecular Potentials for Atomistic Simulation Studies) H. Sun and D. Rigby, Polysilsesquioxanes: Ab Initio Force Field and Structural Conformational and Thermophysical Properties, Spectrochimica Acta (A) 1779, 53, 130, and refs therein. 40
41 Comparison of Force Fields and QM Structure of POSS(H) 8 cube determined from various QM and MM energy minimizations. Exp. Plane Wave (VASP) Atomic Orbital (DMOL) RHF (cc-pvdz) (GAUSSIAN 98) Molecular Mechanics Cerius 2 UFF Cerius 2 Compass Kieffer RFF Si-O Si-H Si-O-Si O-Si-O
42 Biological and Biochemical Force Fields CHARMM, GROMOS, AMBER, OPLS, Optimized to work in the temperature and density range relevant to living systems Little concern for accuracy beyond this regime 42
43 United Atom Force Fields There are essentially three classes of force field models, each of which corresponds to a different level of detail: Explicit atom (all atoms represented explicitly) Used to model a specific system United atom (coarse-grained) Used to model a specific system Mesoscopic (coarse-grained) Used to model phenomena characteristic of a class of systems 43
44 United Atom Force Fields Since the number of non-bonded interactions scales as N 2, for large systems explicitly including every atom is prohibitive. In united atom force fields, some atoms (usually just the hydrogens) are subsumed into the atoms to which they are bonded. E.g. CH 3 is treated as a single united atom. The interaction parameters for each term in the force field are modified to account for the subsumed atoms. Many hydrocarbons are modeled with united atom force fields. In some UA models of proteins, only some of the hydrogens are eliminated. Good alkane UA models: Toxvaerd, Siepmann, Smith & Paul 44
45 Explicit Atom & United Atom Force Fields Force field development is something of an art. Experimental data and QM calculations are used to help parameterize force fields. Incredibly important to have good force fields, since your simulation is only as good as the model you use. Few simulators develop force fields; many use them. Force fields developed for metals, ceramics, polymers, liquids, solids, gases have different requirements and there is a different culture. Challenge for nanoscience: Force fields for dissimilar materials are not easy to obtain. Future studies rely on new force fields. Major area of research for nanoscience researchers. 45
46 Explicit Atom & United Atom Force Fields How well can these force fields do? Structure: within a few percent considered good. Thermodynamics: within a few percent considered good. Dynamics: within 20% considered good. Limitations With MD, atom-steps on large supercomputer possible, (10 12 atom-steps usual) depending on terms in force fields, approximations, tricks used, etc. For many problems, less computationally intensive models are needed in order to study larger systems. 46
47 Mesoscopic Molecular Models Explicit atom and united atom force fields are used to model particular systems, and to predict properties with quantitative accuracy. Sometimes we do not seek to model specific systems, but rather we seek to model phenomena characteristic of a class of systems. For such studies, further coarse-graining is done and mesoscopic molecular models are used. These models contain some of the same terms used in force fields and thus look like force fields; they re used like force field models, but not generally called force fields. 47
48 A Mesoscopic Polymer Model Bead-spring polymer V(r) = 4ε[(σ/r) 12 (σ/r) 6 ] - (k/2)r o2 ln[1-(r/r o ) 2 ] all monomer pairs bonded pairs only Kremer- Grest/ Rudisill- Cummings V(r) Coarse-grained mesoscopic molecular model of a polymer melt. Eg. k=30, R o =1.5, ε=1, σ=1 48
49 A mesoscopic model for colloids The Yukawa potential is commonly used to model the interaction between charged colloids in solvent where the interactions are screened due to the solvent: Here σ = particle diameter, κ = inverse screening length, and ε = surface potential, r = distance between two colloid centers. 49
50 Mapping Mesoscopic Models to EA or UA Models Important area of current research: Coarse-graining detailed EA or UA models to mesoscopic models Mapping mesoscopic models to EA or UA models We ll learn more about this later in the class. 50
CE 530 Molecular Simulation
 1 CE 530 Molecular Simulation Lecture 14 Molecular Models David A. Kofke Department of Chemical Engineering SUNY Buffalo kofke@eng.buffalo.edu 2 Review Monte Carlo ensemble averaging, no dynamics easy
1 CE 530 Molecular Simulation Lecture 14 Molecular Models David A. Kofke Department of Chemical Engineering SUNY Buffalo kofke@eng.buffalo.edu 2 Review Monte Carlo ensemble averaging, no dynamics easy
Lecture 11: Potential Energy Functions
 Lecture 11: Potential Energy Functions Dr. Ronald M. Levy ronlevy@temple.edu Originally contributed by Lauren Wickstrom (2011) Microscopic/Macroscopic Connection The connection between microscopic interactions
Lecture 11: Potential Energy Functions Dr. Ronald M. Levy ronlevy@temple.edu Originally contributed by Lauren Wickstrom (2011) Microscopic/Macroscopic Connection The connection between microscopic interactions
Structural Bioinformatics (C3210) Molecular Mechanics
 Structural Bioinformatics (C3210) Molecular Mechanics How to Calculate Energies Calculation of molecular energies is of key importance in protein folding, molecular modelling etc. There are two main computational
Structural Bioinformatics (C3210) Molecular Mechanics How to Calculate Energies Calculation of molecular energies is of key importance in protein folding, molecular modelling etc. There are two main computational
Example questions for Molecular modelling (Level 4) Dr. Adrian Mulholland
 Example questions for Molecular modelling (Level 4) Dr. Adrian Mulholland 1) Question. Two methods which are widely used for the optimization of molecular geometies are the Steepest descents and Newton-Raphson
Example questions for Molecular modelling (Level 4) Dr. Adrian Mulholland 1) Question. Two methods which are widely used for the optimization of molecular geometies are the Steepest descents and Newton-Raphson
Session 1. Introduction to Computational Chemistry. Computational (chemistry education) and/or (Computational chemistry) education
 Session 1 Introduction to Computational Chemistry 1 Introduction to Computational Chemistry Computational (chemistry education) and/or (Computational chemistry) education First one: Use computational tools
Session 1 Introduction to Computational Chemistry 1 Introduction to Computational Chemistry Computational (chemistry education) and/or (Computational chemistry) education First one: Use computational tools
Bioengineering 215. An Introduction to Molecular Dynamics for Biomolecules
 Bioengineering 215 An Introduction to Molecular Dynamics for Biomolecules David Parker May 18, 2007 ntroduction A principal tool to study biological molecules is molecular dynamics simulations (MD). MD
Bioengineering 215 An Introduction to Molecular Dynamics for Biomolecules David Parker May 18, 2007 ntroduction A principal tool to study biological molecules is molecular dynamics simulations (MD). MD
All-atom Molecular Mechanics. Trent E. Balius AMS 535 / CHE /27/2010
 All-atom Molecular Mechanics Trent E. Balius AMS 535 / CHE 535 09/27/2010 Outline Molecular models Molecular mechanics Force Fields Potential energy function functional form parameters and parameterization
All-atom Molecular Mechanics Trent E. Balius AMS 535 / CHE 535 09/27/2010 Outline Molecular models Molecular mechanics Force Fields Potential energy function functional form parameters and parameterization
Molecular Mechanics. Yohann Moreau. November 26, 2015
 Molecular Mechanics Yohann Moreau yohann.moreau@ujf-grenoble.fr November 26, 2015 Yohann Moreau (UJF) Molecular Mechanics, Label RFCT 2015 November 26, 2015 1 / 29 Introduction A so-called Force-Field
Molecular Mechanics Yohann Moreau yohann.moreau@ujf-grenoble.fr November 26, 2015 Yohann Moreau (UJF) Molecular Mechanics, Label RFCT 2015 November 26, 2015 1 / 29 Introduction A so-called Force-Field
Scuola di Chimica Computazionale
 Societa Chimica Italiana Gruppo Interdivisionale di Chimica Computazionale Scuola di Chimica Computazionale Introduzione, per Esercizi, all Uso del Calcolatore in Chimica Organica e Biologica Modellistica
Societa Chimica Italiana Gruppo Interdivisionale di Chimica Computazionale Scuola di Chimica Computazionale Introduzione, per Esercizi, all Uso del Calcolatore in Chimica Organica e Biologica Modellistica
Introduction to molecular dynamics
 1 Introduction to molecular dynamics Yves Lansac Université François Rabelais, Tours, France Visiting MSE, GIST for the summer Molecular Simulation 2 Molecular simulation is a computational experiment.
1 Introduction to molecular dynamics Yves Lansac Université François Rabelais, Tours, France Visiting MSE, GIST for the summer Molecular Simulation 2 Molecular simulation is a computational experiment.
Molecular mechanics. classical description of molecules. Marcus Elstner and Tomáš Kubař. April 29, 2016
 classical description of molecules April 29, 2016 Chemical bond Conceptual and chemical basis quantum effect solution of the SR numerically expensive (only small molecules can be treated) approximations
classical description of molecules April 29, 2016 Chemical bond Conceptual and chemical basis quantum effect solution of the SR numerically expensive (only small molecules can be treated) approximations
k θ (θ θ 0 ) 2 angles r i j r i j
 1 Force fields 1.1 Introduction The term force field is slightly misleading, since it refers to the parameters of the potential used to calculate the forces (via gradient) in molecular dynamics simulations.
1 Force fields 1.1 Introduction The term force field is slightly misleading, since it refers to the parameters of the potential used to calculate the forces (via gradient) in molecular dynamics simulations.
Molecular Mechanics. C. David Sherrill School of Chemistry and Biochemistry Georgia Institute of Technology. January 2001
 Molecular Mechanics C. David Sherrill School of Chemistry and Biochemistry Georgia Institute of Technology January 2001 Introduction Molecular Mechanics uses classical type models to predict the energy
Molecular Mechanics C. David Sherrill School of Chemistry and Biochemistry Georgia Institute of Technology January 2001 Introduction Molecular Mechanics uses classical type models to predict the energy
Intro to ab initio methods
 Lecture 2 Part A Intro to ab initio methods Recommended reading: Leach, Chapters 2 & 3 for QM methods For more QM methods: Essentials of Computational Chemistry by C.J. Cramer, Wiley (2002) 1 ab initio
Lecture 2 Part A Intro to ab initio methods Recommended reading: Leach, Chapters 2 & 3 for QM methods For more QM methods: Essentials of Computational Chemistry by C.J. Cramer, Wiley (2002) 1 ab initio
This semester. Books
 Models mostly proteins from detailed to more abstract models Some simulation methods This semester Books None necessary for my group and Prof Rarey Molecular Modelling: Principles and Applications Leach,
Models mostly proteins from detailed to more abstract models Some simulation methods This semester Books None necessary for my group and Prof Rarey Molecular Modelling: Principles and Applications Leach,
T6.2 Molecular Mechanics
 T6.2 Molecular Mechanics We have seen that Benson group additivities are capable of giving heats of formation of molecules with accuracies comparable to those of the best ab initio procedures. However,
T6.2 Molecular Mechanics We have seen that Benson group additivities are capable of giving heats of formation of molecules with accuracies comparable to those of the best ab initio procedures. However,
CE 530 Molecular Simulation
 1 CE 530 Molecular Simulation Lecture 1 David A. Kofke Department of Chemical Engineering SUNY Buffalo kofke@eng.buffalo.edu 2 Time/s Multi-Scale Modeling Based on SDSC Blue Horizon (SP3) 1.728 Tflops
1 CE 530 Molecular Simulation Lecture 1 David A. Kofke Department of Chemical Engineering SUNY Buffalo kofke@eng.buffalo.edu 2 Time/s Multi-Scale Modeling Based on SDSC Blue Horizon (SP3) 1.728 Tflops
Computational Methods. Chem 561
 Computational Methods Chem 561 Lecture Outline 1. Ab initio methods a) HF SCF b) Post-HF methods 2. Density Functional Theory 3. Semiempirical methods 4. Molecular Mechanics Computational Chemistry " Computational
Computational Methods Chem 561 Lecture Outline 1. Ab initio methods a) HF SCF b) Post-HF methods 2. Density Functional Theory 3. Semiempirical methods 4. Molecular Mechanics Computational Chemistry " Computational
An introduction to Molecular Dynamics. EMBO, June 2016
 An introduction to Molecular Dynamics EMBO, June 2016 What is MD? everything that living things do can be understood in terms of the jiggling and wiggling of atoms. The Feynman Lectures in Physics vol.
An introduction to Molecular Dynamics EMBO, June 2016 What is MD? everything that living things do can be understood in terms of the jiggling and wiggling of atoms. The Feynman Lectures in Physics vol.
The Potential Energy Surface (PES) And the Basic Force Field Chem 4021/8021 Video II.iii
 The Potential Energy Surface (PES) And the Basic Force Field Chem 4021/8021 Video II.iii Fundamental Points About Which to Be Thinking It s clear the PES is useful, so how can I construct it for an arbitrary
The Potential Energy Surface (PES) And the Basic Force Field Chem 4021/8021 Video II.iii Fundamental Points About Which to Be Thinking It s clear the PES is useful, so how can I construct it for an arbitrary
Hyeyoung Shin a, Tod A. Pascal ab, William A. Goddard III abc*, and Hyungjun Kim a* Korea
 The Scaled Effective Solvent Method for Predicting the Equilibrium Ensemble of Structures with Analysis of Thermodynamic Properties of Amorphous Polyethylene Glycol-Water Mixtures Hyeyoung Shin a, Tod
The Scaled Effective Solvent Method for Predicting the Equilibrium Ensemble of Structures with Analysis of Thermodynamic Properties of Amorphous Polyethylene Glycol-Water Mixtures Hyeyoung Shin a, Tod
The Effect of Model Internal Flexibility Upon NEMD Simulations of Viscosity
 Draft: September 29, 1999 The Effect of Model Internal Flexibility Upon NEMD Simulations of Viscosity N. G. Fuller 1 and R. L. Rowley 1,2 Abstract The influence of model flexibility upon simulated viscosity
Draft: September 29, 1999 The Effect of Model Internal Flexibility Upon NEMD Simulations of Viscosity N. G. Fuller 1 and R. L. Rowley 1,2 Abstract The influence of model flexibility upon simulated viscosity
3rd Advanced in silico Drug Design KFC/ADD Molecular mechanics intro Karel Berka, Ph.D. Martin Lepšík, Ph.D. Pavel Polishchuk, Ph.D.
 3rd Advanced in silico Drug Design KFC/ADD Molecular mechanics intro Karel Berka, Ph.D. Martin Lepšík, Ph.D. Pavel Polishchuk, Ph.D. Thierry Langer, Ph.D. Jana Vrbková, Ph.D. UP Olomouc, 23.1.-26.1. 2018
3rd Advanced in silico Drug Design KFC/ADD Molecular mechanics intro Karel Berka, Ph.D. Martin Lepšík, Ph.D. Pavel Polishchuk, Ph.D. Thierry Langer, Ph.D. Jana Vrbková, Ph.D. UP Olomouc, 23.1.-26.1. 2018
Multiscale Materials Modeling
 Multiscale Materials Modeling Lecture 09 Quantum Mechanics/Molecular Mechanics (QM/MM) Techniques Fundamentals of Sustainable Technology These notes created by David Keffer, University of Tennessee, Knoxville,
Multiscale Materials Modeling Lecture 09 Quantum Mechanics/Molecular Mechanics (QM/MM) Techniques Fundamentals of Sustainable Technology These notes created by David Keffer, University of Tennessee, Knoxville,
Reactive potentials and applications
 1.021, 3.021, 10.333, 22.00 Introduction to Modeling and Simulation Spring 2011 Part I Continuum and particle methods Reactive potentials and applications Lecture 8 Markus J. Buehler Laboratory for Atomistic
1.021, 3.021, 10.333, 22.00 Introduction to Modeling and Simulation Spring 2011 Part I Continuum and particle methods Reactive potentials and applications Lecture 8 Markus J. Buehler Laboratory for Atomistic
Solutions to Assignment #4 Getting Started with HyperChem
 Solutions to Assignment #4 Getting Started with HyperChem 1. This first exercise is meant to familiarize you with the different methods for visualizing molecules available in HyperChem. (a) Create a molecule
Solutions to Assignment #4 Getting Started with HyperChem 1. This first exercise is meant to familiarize you with the different methods for visualizing molecules available in HyperChem. (a) Create a molecule
Coarse-Grained Models!
 Coarse-Grained Models! Large and complex molecules (e.g. long polymers) can not be simulated on the all-atom level! Requires coarse-graining of the model! Coarse-grained models are usually also particles
Coarse-Grained Models! Large and complex molecules (e.g. long polymers) can not be simulated on the all-atom level! Requires coarse-graining of the model! Coarse-grained models are usually also particles
Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015,
 Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015, Course,Informa5on, BIOC%530% GraduateAlevel,discussion,of,the,structure,,func5on,,and,chemistry,of,proteins,and, nucleic,acids,,control,of,enzyma5c,reac5ons.,please,see,the,course,syllabus,and,
Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015, Course,Informa5on, BIOC%530% GraduateAlevel,discussion,of,the,structure,,func5on,,and,chemistry,of,proteins,and, nucleic,acids,,control,of,enzyma5c,reac5ons.,please,see,the,course,syllabus,and,
A Molecular Dynamics Simulation of a Homogeneous Organic-Inorganic Hybrid Silica Membrane
 A Molecular Dynamics Simulation of a Homogeneous Organic-Inorganic Hybrid Silica Membrane Supplementary Information: Simulation Procedure and Physical Property Analysis Simulation Procedure The molecular
A Molecular Dynamics Simulation of a Homogeneous Organic-Inorganic Hybrid Silica Membrane Supplementary Information: Simulation Procedure and Physical Property Analysis Simulation Procedure The molecular
Molecular Dynamics. A very brief introduction
 Molecular Dynamics A very brief introduction Sander Pronk Dept. of Theoretical Physics KTH Royal Institute of Technology & Science For Life Laboratory Stockholm, Sweden Why computer simulations? Two primary
Molecular Dynamics A very brief introduction Sander Pronk Dept. of Theoretical Physics KTH Royal Institute of Technology & Science For Life Laboratory Stockholm, Sweden Why computer simulations? Two primary
Scientific Computing II
 Scientific Computing II Molecular Dynamics Simulation Michael Bader SCCS Summer Term 2015 Molecular Dynamics Simulation, Summer Term 2015 1 Continuum Mechanics for Fluid Mechanics? Molecular Dynamics the
Scientific Computing II Molecular Dynamics Simulation Michael Bader SCCS Summer Term 2015 Molecular Dynamics Simulation, Summer Term 2015 1 Continuum Mechanics for Fluid Mechanics? Molecular Dynamics the
Lecture C2 Microscopic to Macroscopic, Part 2: Intermolecular Interactions. Let's get together.
 Lecture C2 Microscopic to Macroscopic, Part 2: Intermolecular Interactions Let's get together. Most gases are NOT ideal except at very low pressures: Z=1 for ideal gases Intermolecular interactions come
Lecture C2 Microscopic to Macroscopic, Part 2: Intermolecular Interactions Let's get together. Most gases are NOT ideal except at very low pressures: Z=1 for ideal gases Intermolecular interactions come
Peptide folding in non-aqueous environments investigated with molecular dynamics simulations Soto Becerra, Patricia
 University of Groningen Peptide folding in non-aqueous environments investigated with molecular dynamics simulations Soto Becerra, Patricia IMPORTANT NOTE: You are advised to consult the publisher's version
University of Groningen Peptide folding in non-aqueous environments investigated with molecular dynamics simulations Soto Becerra, Patricia IMPORTANT NOTE: You are advised to consult the publisher's version
Force Fields in Molecular Mechanics
 Force Fields in Molecular Mechanics Rajarshi Guha (9915607) and Rajesh Sardar (9915610) March 21, 2001 1 Introduction With the advent of computers chemists have realized the utility of carrying out simulations
Force Fields in Molecular Mechanics Rajarshi Guha (9915607) and Rajesh Sardar (9915610) March 21, 2001 1 Introduction With the advent of computers chemists have realized the utility of carrying out simulations
Energy functions and their relationship to molecular conformation. CS/CME/BioE/Biophys/BMI 279 Oct. 3 and 5, 2017 Ron Dror
 Energy functions and their relationship to molecular conformation CS/CME/BioE/Biophys/BMI 279 Oct. 3 and 5, 2017 Ron Dror Yesterday s Nobel Prize: single-particle cryoelectron microscopy 2 Outline Energy
Energy functions and their relationship to molecular conformation CS/CME/BioE/Biophys/BMI 279 Oct. 3 and 5, 2017 Ron Dror Yesterday s Nobel Prize: single-particle cryoelectron microscopy 2 Outline Energy
Water models in classical simulations
 Water models in classical simulations Maria Fyta Institut für Computerphysik, Universität Stuttgart Stuttgart, Germany Water transparent, odorless, tasteless and ubiquitous really simple: two H atoms attached
Water models in classical simulations Maria Fyta Institut für Computerphysik, Universität Stuttgart Stuttgart, Germany Water transparent, odorless, tasteless and ubiquitous really simple: two H atoms attached
Computational Modeling of Protein-Ligand Interactions
 Computational Modeling of Protein-Ligand Interactions Steven R. Gwaltney Department of Chemistry Mississippi State University Mississippi State, MS 39762 Auguste Comte, 1830 Every attempt to refer chemical
Computational Modeling of Protein-Ligand Interactions Steven R. Gwaltney Department of Chemistry Mississippi State University Mississippi State, MS 39762 Auguste Comte, 1830 Every attempt to refer chemical
Imperfect Gases. NC State University
 Chemistry 431 Lecture 3 Imperfect Gases NC State University The Compression Factor One way to represent the relationship between ideal and real gases is to plot the deviation from ideality as the gas is
Chemistry 431 Lecture 3 Imperfect Gases NC State University The Compression Factor One way to represent the relationship between ideal and real gases is to plot the deviation from ideality as the gas is
4 th Advanced in silico Drug Design KFC/ADD Molecular Modelling Intro. Karel Berka, Ph.D.
 4 th Advanced in silico Drug Design KFC/ADD Molecular Modelling Intro Karel Berka, Ph.D. UP Olomouc, 21.1.-25.1. 2019 Motto A theory is something nobody believes, except the person who made it An experiment
4 th Advanced in silico Drug Design KFC/ADD Molecular Modelling Intro Karel Berka, Ph.D. UP Olomouc, 21.1.-25.1. 2019 Motto A theory is something nobody believes, except the person who made it An experiment
Ideal Gas Behavior. NC State University
 Chemistry 331 Lecture 6 Ideal Gas Behavior NC State University Macroscopic variables P, T Pressure is a force per unit area (P= F/A) The force arises from the change in momentum as particles hit an object
Chemistry 331 Lecture 6 Ideal Gas Behavior NC State University Macroscopic variables P, T Pressure is a force per unit area (P= F/A) The force arises from the change in momentum as particles hit an object
ICCP Project 2 - Advanced Monte Carlo Methods Choose one of the three options below
 ICCP Project 2 - Advanced Monte Carlo Methods Choose one of the three options below Introduction In statistical physics Monte Carlo methods are considered to have started in the Manhattan project (1940
ICCP Project 2 - Advanced Monte Carlo Methods Choose one of the three options below Introduction In statistical physics Monte Carlo methods are considered to have started in the Manhattan project (1940
Computational Chemistry. An Introduction to Molecular Dynamic Simulations
 Computational Chemistry An Introduction to Molecular Dynamic Simulations Computational chemistry simulates chemical structures and reactions numerically, based in full or in part on the fundamental laws
Computational Chemistry An Introduction to Molecular Dynamic Simulations Computational chemistry simulates chemical structures and reactions numerically, based in full or in part on the fundamental laws
General Physical Chemistry II
 General Physical Chemistry II Lecture 13 Aleksey Kocherzhenko October 16, 2014" Last time " The Hückel method" Ø Used to study π systems of conjugated molecules" Ø π orbitals are treated separately from
General Physical Chemistry II Lecture 13 Aleksey Kocherzhenko October 16, 2014" Last time " The Hückel method" Ø Used to study π systems of conjugated molecules" Ø π orbitals are treated separately from
Computational Molecular Biophysics. Computational Biophysics, GRS Jülich SS 2013
 Computational Molecular Biophysics Computational Biophysics, GRS Jülich SS 2013 Computational Considerations Number of terms for different energy contributions / E el ~N ~N ~N ~N 2 Computational cost for
Computational Molecular Biophysics Computational Biophysics, GRS Jülich SS 2013 Computational Considerations Number of terms for different energy contributions / E el ~N ~N ~N ~N 2 Computational cost for
Molecular Mechanics / ReaxFF
 Molecular Dynamics simulations Lecture 09: Molecular Mechanics / ReaxFF Dr. Olli Pakarinen University of Helsinki Fall 2012 Lecture notes based on notes by Dr. Jani Kotakoski, 2010 CONTENTS Molecular mechanics
Molecular Dynamics simulations Lecture 09: Molecular Mechanics / ReaxFF Dr. Olli Pakarinen University of Helsinki Fall 2012 Lecture notes based on notes by Dr. Jani Kotakoski, 2010 CONTENTS Molecular mechanics
Colloid Chemistry. La chimica moderna e la sua comunicazione Silvia Gross.
 Colloid Chemistry La chimica moderna e la sua comunicazione Silvia Gross Istituto Dipartimento di Scienze di e Scienze Tecnologie Chimiche Molecolari ISTM-CNR, Università Università degli Studi degli Studi
Colloid Chemistry La chimica moderna e la sua comunicazione Silvia Gross Istituto Dipartimento di Scienze di e Scienze Tecnologie Chimiche Molecolari ISTM-CNR, Università Università degli Studi degli Studi
Subject of the Lecture:
 Subject of the Lecture: Conceptual basis for the development of force fields. Implementation/validation Water - a worked example Extensions - combining molecular mechanics and quantum mechanics (QM/MM)
Subject of the Lecture: Conceptual basis for the development of force fields. Implementation/validation Water - a worked example Extensions - combining molecular mechanics and quantum mechanics (QM/MM)
What is Classical Molecular Dynamics?
 What is Classical Molecular Dynamics? Simulation of explicit particles (atoms, ions,... ) Particles interact via relatively simple analytical potential functions Newton s equations of motion are integrated
What is Classical Molecular Dynamics? Simulation of explicit particles (atoms, ions,... ) Particles interact via relatively simple analytical potential functions Newton s equations of motion are integrated
Dihedral Angles. Homayoun Valafar. Department of Computer Science and Engineering, USC 02/03/10 CSCE 769
 Dihedral Angles Homayoun Valafar Department of Computer Science and Engineering, USC The precise definition of a dihedral or torsion angle can be found in spatial geometry Angle between to planes Dihedral
Dihedral Angles Homayoun Valafar Department of Computer Science and Engineering, USC The precise definition of a dihedral or torsion angle can be found in spatial geometry Angle between to planes Dihedral
Close agreement between the orientation dependence of hydrogen bonds observed in protein structures and quantum mechanical calculations
 Close agreement between the orientation dependence of hydrogen bonds observed in protein structures and quantum mechanical calculations Alexandre V. Morozov, Tanja Kortemme, Kiril Tsemekhman, David Baker
Close agreement between the orientation dependence of hydrogen bonds observed in protein structures and quantum mechanical calculations Alexandre V. Morozov, Tanja Kortemme, Kiril Tsemekhman, David Baker
Lecture 1. Conformational Analysis in Acyclic Systems
 Lecture 1 Conformational Analysis in Acyclic Systems Learning Outcomes: by the end of this lecture and after answering the associated problems, you will be able to: 1. use Newman and saw-horse projections
Lecture 1 Conformational Analysis in Acyclic Systems Learning Outcomes: by the end of this lecture and after answering the associated problems, you will be able to: 1. use Newman and saw-horse projections
Chapter 3. Crystal Binding
 Chapter 3. Crystal Binding Energy of a crystal and crystal binding Cohesive energy of Molecular crystals Ionic crystals Metallic crystals Elasticity What causes matter to exist in three different forms?
Chapter 3. Crystal Binding Energy of a crystal and crystal binding Cohesive energy of Molecular crystals Ionic crystals Metallic crystals Elasticity What causes matter to exist in three different forms?
74 these states cannot be reliably obtained from experiments. In addition, the barriers between the local minima can also not be obtained reliably fro
 73 Chapter 5 Development of Adiabatic Force Field for Polyvinyl Chloride (PVC) and Chlorinated PVC (CPVC) 5.1 Introduction Chlorinated polyvinyl chloride has become an important specialty polymer due to
73 Chapter 5 Development of Adiabatic Force Field for Polyvinyl Chloride (PVC) and Chlorinated PVC (CPVC) 5.1 Introduction Chlorinated polyvinyl chloride has become an important specialty polymer due to
Materials 218/UCSB: Class III Cohesion in solids van der Waals, ionic, covalent, metallic
 Materials 218/UCSB: Class III Cohesion in solids van der Waals, ionic, covalent, metallic Ram Seshadri (seshadri@mrl.ucsb.edu) Introduction There are four forces in nature. The strong and the weak interactions
Materials 218/UCSB: Class III Cohesion in solids van der Waals, ionic, covalent, metallic Ram Seshadri (seshadri@mrl.ucsb.edu) Introduction There are four forces in nature. The strong and the weak interactions
Molecular Dynamics Simulations. Dr. Noelia Faginas Lago Dipartimento di Chimica,Biologia e Biotecnologie Università di Perugia
 Molecular Dynamics Simulations Dr. Noelia Faginas Lago Dipartimento di Chimica,Biologia e Biotecnologie Università di Perugia 1 An Introduction to Molecular Dynamics Simulations Macroscopic properties
Molecular Dynamics Simulations Dr. Noelia Faginas Lago Dipartimento di Chimica,Biologia e Biotecnologie Università di Perugia 1 An Introduction to Molecular Dynamics Simulations Macroscopic properties
Energy functions and their relationship to molecular conformation. CS/CME/BioE/Biophys/BMI 279 Oct. 3 and 5, 2017 Ron Dror
 Energy functions and their relationship to molecular conformation CS/CME/BioE/Biophys/BMI 279 Oct. 3 and 5, 2017 Ron Dror Outline Energy functions for proteins (or biomolecular systems more generally)
Energy functions and their relationship to molecular conformation CS/CME/BioE/Biophys/BMI 279 Oct. 3 and 5, 2017 Ron Dror Outline Energy functions for proteins (or biomolecular systems more generally)
3. An Introduction to Molecular Mechanics
 3. An Introduction to Molecular Mechanics Introduction When you use Chem3D to draw molecules, the program assigns bond lengths and bond angles based on experimental data. The program does not contain real
3. An Introduction to Molecular Mechanics Introduction When you use Chem3D to draw molecules, the program assigns bond lengths and bond angles based on experimental data. The program does not contain real
Potentials, periodicity
 Potentials, periodicity Lecture 2 1/23/18 1 Survey responses 2 Topic requests DFT (10), Molecular dynamics (7), Monte Carlo (5) Machine Learning (4), High-throughput, Databases (4) NEB, phonons, Non-equilibrium
Potentials, periodicity Lecture 2 1/23/18 1 Survey responses 2 Topic requests DFT (10), Molecular dynamics (7), Monte Carlo (5) Machine Learning (4), High-throughput, Databases (4) NEB, phonons, Non-equilibrium
Equations of State. Equations of State (EoS)
 Equations of State (EoS) Equations of State From molecular considerations, identify which intermolecular interactions are significant (including estimating relative strengths of dipole moments, polarizability,
Equations of State (EoS) Equations of State From molecular considerations, identify which intermolecular interactions are significant (including estimating relative strengths of dipole moments, polarizability,
510 Subject Index. Hamiltonian 33, 86, 88, 89 Hamilton operator 34, 164, 166
 Subject Index Ab-initio calculation 24, 122, 161. 165 Acentric factor 279, 338 Activity absolute 258, 295 coefficient 7 definition 7 Atom 23 Atomic units 93 Avogadro number 5, 92 Axilrod-Teller-forces
Subject Index Ab-initio calculation 24, 122, 161. 165 Acentric factor 279, 338 Activity absolute 258, 295 coefficient 7 definition 7 Atom 23 Atomic units 93 Avogadro number 5, 92 Axilrod-Teller-forces
Atomic and molecular interaction forces in biology
 Atomic and molecular interaction forces in biology 1 Outline Types of interactions relevant to biology Van der Waals interactions H-bond interactions Some properties of water Hydrophobic effect 2 Types
Atomic and molecular interaction forces in biology 1 Outline Types of interactions relevant to biology Van der Waals interactions H-bond interactions Some properties of water Hydrophobic effect 2 Types
Reactive Empirical Force Fields
 Reactive Empirical Force Fields Jason Quenneville jasonq@lanl.gov X-1: Solid Mechanics, EOS and Materials Properties Applied Physics Division Los Alamos National Laboratory Timothy C. Germann, Los Alamos
Reactive Empirical Force Fields Jason Quenneville jasonq@lanl.gov X-1: Solid Mechanics, EOS and Materials Properties Applied Physics Division Los Alamos National Laboratory Timothy C. Germann, Los Alamos
UNIT 5 : STATES OF MATTER Concept 1. INTERMOLECULAR FORCES
 www.tiwariacademy.in UNIT 5 : STATES OF MATTER CONCEPT WISE HANDOUTS KEY CONCEPTS : 1. Intermolecular Forces 2. Gas Laws 3. Behaviour of gases Concept 1. INTERMOLECULAR FORCES Intermolecular forces- forces
www.tiwariacademy.in UNIT 5 : STATES OF MATTER CONCEPT WISE HANDOUTS KEY CONCEPTS : 1. Intermolecular Forces 2. Gas Laws 3. Behaviour of gases Concept 1. INTERMOLECULAR FORCES Intermolecular forces- forces
Fondamenti di Chimica Farmaceutica. Computer Chemistry in Drug Research: Introduction
 Fondamenti di Chimica Farmaceutica Computer Chemistry in Drug Research: Introduction Introduction Introduction Introduction Computer Chemistry in Drug Design Drug Discovery: Target identification Lead
Fondamenti di Chimica Farmaceutica Computer Chemistry in Drug Research: Introduction Introduction Introduction Introduction Computer Chemistry in Drug Design Drug Discovery: Target identification Lead
Biophysics II. Hydrophobic Bio-molecules. Key points to be covered. Molecular Interactions in Bio-molecular Structures - van der Waals Interaction
 Biophysics II Key points to be covered By A/Prof. Xiang Yang Liu Biophysics & Micro/nanostructures Lab Department of Physics, NUS 1. van der Waals Interaction 2. Hydrogen bond 3. Hydrophilic vs hydrophobic
Biophysics II Key points to be covered By A/Prof. Xiang Yang Liu Biophysics & Micro/nanostructures Lab Department of Physics, NUS 1. van der Waals Interaction 2. Hydrogen bond 3. Hydrophilic vs hydrophobic
CHEM-UA 127: Advanced General Chemistry I
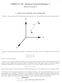 1 CHEM-UA 127: Advanced General Chemistry I Notes for Lecture 6 I MOLECULAR GEOMETRY AND COORDINATES Consider a diatomic molecule AB Imagine fixing this molecule at a very specific spatial location, as
1 CHEM-UA 127: Advanced General Chemistry I Notes for Lecture 6 I MOLECULAR GEOMETRY AND COORDINATES Consider a diatomic molecule AB Imagine fixing this molecule at a very specific spatial location, as
Why study protein dynamics?
 Why study protein dynamics? Protein flexibility is crucial for function. One average structure is not enough. Proteins constantly sample configurational space. Transport - binding and moving molecules
Why study protein dynamics? Protein flexibility is crucial for function. One average structure is not enough. Proteins constantly sample configurational space. Transport - binding and moving molecules
DENSITY FUNCTIONAL THEORY FOR NON-THEORISTS JOHN P. PERDEW DEPARTMENTS OF PHYSICS AND CHEMISTRY TEMPLE UNIVERSITY
 DENSITY FUNCTIONAL THEORY FOR NON-THEORISTS JOHN P. PERDEW DEPARTMENTS OF PHYSICS AND CHEMISTRY TEMPLE UNIVERSITY A TUTORIAL FOR PHYSICAL SCIENTISTS WHO MAY OR MAY NOT HATE EQUATIONS AND PROOFS REFERENCES
DENSITY FUNCTIONAL THEORY FOR NON-THEORISTS JOHN P. PERDEW DEPARTMENTS OF PHYSICS AND CHEMISTRY TEMPLE UNIVERSITY A TUTORIAL FOR PHYSICAL SCIENTISTS WHO MAY OR MAY NOT HATE EQUATIONS AND PROOFS REFERENCES
The Molecular Dynamics Method
 The Molecular Dynamics Method Thermal motion of a lipid bilayer Water permeation through channels Selective sugar transport Potential Energy (hyper)surface What is Force? Energy U(x) F = d dx U(x) Conformation
The Molecular Dynamics Method Thermal motion of a lipid bilayer Water permeation through channels Selective sugar transport Potential Energy (hyper)surface What is Force? Energy U(x) F = d dx U(x) Conformation
Interatomic Potentials. The electronic-structure problem
 Interatomic Potentials Before we can start a simulation, we need the model! Interaction between atoms and molecules is determined by quantum mechanics: Schrödinger Equation + Born-Oppenheimer approximation
Interatomic Potentials Before we can start a simulation, we need the model! Interaction between atoms and molecules is determined by quantum mechanics: Schrödinger Equation + Born-Oppenheimer approximation
Why Is Molecular Interaction Important in Our Life
 Why Is Molecular Interaction Important in ur Life QuLiS and Graduate School of Science iroshima University http://www.nabit.hiroshima-u.ac.jp/iwatasue/indexe.htm Suehiro Iwata Sept. 29, 2007 Department
Why Is Molecular Interaction Important in ur Life QuLiS and Graduate School of Science iroshima University http://www.nabit.hiroshima-u.ac.jp/iwatasue/indexe.htm Suehiro Iwata Sept. 29, 2007 Department
Adsorption of gases on solids (focus on physisorption)
 Adsorption of gases on solids (focus on physisorption) Adsorption Solid surfaces show strong affinity towards gas molecules that it comes in contact with and some amt of them are trapped on the surface
Adsorption of gases on solids (focus on physisorption) Adsorption Solid surfaces show strong affinity towards gas molecules that it comes in contact with and some amt of them are trapped on the surface
Modeling & Simulation of Glass Structure
 Modeling & Simulation of Glass Structure VCG Lecture 21 John Kieffer Department of Materials Science and Engineering University of Michigan 1 Overview Historical perspective Simulation methodologies Theoretical
Modeling & Simulation of Glass Structure VCG Lecture 21 John Kieffer Department of Materials Science and Engineering University of Michigan 1 Overview Historical perspective Simulation methodologies Theoretical
CHEMISTRY 4021/8021 MIDTERM EXAM 1 SPRING 2014
 CHEMISTRY 4021/8021 Q1) Propose a simple, united-atom molecular mechanics force-field needed to generate a potential energy surface for an isolated molecule of acetone (Me 2 CO). I.e., provide an energy
CHEMISTRY 4021/8021 Q1) Propose a simple, united-atom molecular mechanics force-field needed to generate a potential energy surface for an isolated molecule of acetone (Me 2 CO). I.e., provide an energy
The broad topic of physical metallurgy provides a basis that links the structure of materials with their properties, focusing primarily on metals.
 Physical Metallurgy The broad topic of physical metallurgy provides a basis that links the structure of materials with their properties, focusing primarily on metals. Crystal Binding In our discussions
Physical Metallurgy The broad topic of physical metallurgy provides a basis that links the structure of materials with their properties, focusing primarily on metals. Crystal Binding In our discussions
Free energy, electrostatics, and the hydrophobic effect
 Protein Physics 2016 Lecture 3, January 26 Free energy, electrostatics, and the hydrophobic effect Magnus Andersson magnus.andersson@scilifelab.se Theoretical & Computational Biophysics Recap Protein structure
Protein Physics 2016 Lecture 3, January 26 Free energy, electrostatics, and the hydrophobic effect Magnus Andersson magnus.andersson@scilifelab.se Theoretical & Computational Biophysics Recap Protein structure
Ionic Bonds. H He: ... Li Be B C :N :O :F: :Ne:
 Ionic Bonds Valence electrons - the electrons in the highest occupied energy level - always electrons in the s and p orbitals - maximum of 8 valence electrons - elements in the same group have the same
Ionic Bonds Valence electrons - the electrons in the highest occupied energy level - always electrons in the s and p orbitals - maximum of 8 valence electrons - elements in the same group have the same
Molecular Mechanics, Dynamics & Docking
 Molecular Mechanics, Dynamics & Docking Lawrence Hunter, Ph.D. Director, Computational Bioscience Program University of Colorado School of Medicine Larry.Hunter@uchsc.edu http://compbio.uchsc.edu/hunter
Molecular Mechanics, Dynamics & Docking Lawrence Hunter, Ph.D. Director, Computational Bioscience Program University of Colorado School of Medicine Larry.Hunter@uchsc.edu http://compbio.uchsc.edu/hunter
The liquid-vapour interface of QDO water. Flaviu Cipcigan Andrew Jones Jason Crain Vlad Sokhan Glenn Martyna
 The liquid-vapour interface of QDO water Flaviu Cipcigan Andrew Jones Jason Crain Vlad Sokhan Glenn Martyna The liquid-vapour interface of QDO water 1. Molecular models 2. The Quantum Drude Oscillator
The liquid-vapour interface of QDO water Flaviu Cipcigan Andrew Jones Jason Crain Vlad Sokhan Glenn Martyna The liquid-vapour interface of QDO water 1. Molecular models 2. The Quantum Drude Oscillator
Supporting Information
 Projection of atomistic simulation data for the dynamics of entangled polymers onto the tube theory: Calculation of the segment survival probability function and comparison with modern tube models Pavlos
Projection of atomistic simulation data for the dynamics of entangled polymers onto the tube theory: Calculation of the segment survival probability function and comparison with modern tube models Pavlos
Electrons and Molecular Forces
 Electrons and Molecular Forces Chemistry 30 Ms. Hayduk Electron Configuration Atomic Structure Atomic Number Number of protons in the nucleus Defines the element Used to organize the periodic table 1 Bohr
Electrons and Molecular Forces Chemistry 30 Ms. Hayduk Electron Configuration Atomic Structure Atomic Number Number of protons in the nucleus Defines the element Used to organize the periodic table 1 Bohr
3. An Introduction to Molecular Mechanics
 3. An Introduction to Molecular Mechanics Introduction When you use Chem3D to draw molecules, the program assigns bond lengths and bond angles based on experimental data. The program does not contain real
3. An Introduction to Molecular Mechanics Introduction When you use Chem3D to draw molecules, the program assigns bond lengths and bond angles based on experimental data. The program does not contain real
Biomolecules are dynamic no single structure is a perfect model
 Molecular Dynamics Simulations of Biomolecules References: A. R. Leach Molecular Modeling Principles and Applications Prentice Hall, 2001. M. P. Allen and D. J. Tildesley "Computer Simulation of Liquids",
Molecular Dynamics Simulations of Biomolecules References: A. R. Leach Molecular Modeling Principles and Applications Prentice Hall, 2001. M. P. Allen and D. J. Tildesley "Computer Simulation of Liquids",
Can a continuum solvent model reproduce the free energy landscape of a β-hairpin folding in water?
 Can a continuum solvent model reproduce the free energy landscape of a β-hairpin folding in water? Ruhong Zhou 1 and Bruce J. Berne 2 1 IBM Thomas J. Watson Research Center; and 2 Department of Chemistry,
Can a continuum solvent model reproduce the free energy landscape of a β-hairpin folding in water? Ruhong Zhou 1 and Bruce J. Berne 2 1 IBM Thomas J. Watson Research Center; and 2 Department of Chemistry,
Introduction to Computational Chemistry
 Introduction to Computational Chemistry Vesa Hänninen Laboratory of Physical Chemistry room B430, Chemicum 4th floor vesa.hanninen@helsinki.fi September 3, 2013 Introduction and theoretical backround September
Introduction to Computational Chemistry Vesa Hänninen Laboratory of Physical Chemistry room B430, Chemicum 4th floor vesa.hanninen@helsinki.fi September 3, 2013 Introduction and theoretical backround September
CHEM-UA 652: Thermodynamics and Kinetics
 1 CHEM-UA 652: Thermodynamics and Kinetics Notes for Lecture 4 I. THE ISOTHERMAL-ISOBARIC ENSEMBLE The isothermal-isobaric ensemble is the closest mimic to the conditions under which most experiments are
1 CHEM-UA 652: Thermodynamics and Kinetics Notes for Lecture 4 I. THE ISOTHERMAL-ISOBARIC ENSEMBLE The isothermal-isobaric ensemble is the closest mimic to the conditions under which most experiments are
Biomolecular modeling I
 2016, December 6 Biomolecular structure Structural elements of life Biomolecules proteins, nucleic acids, lipids, carbohydrates... Biomolecular structure Biomolecules biomolecular complexes aggregates...
2016, December 6 Biomolecular structure Structural elements of life Biomolecules proteins, nucleic acids, lipids, carbohydrates... Biomolecular structure Biomolecules biomolecular complexes aggregates...
Chapter II: Interactions of ions with matter
 Chapter II: Interactions of ions with matter 1 Trajectories of α particles of 5.5 MeV Source: SRIM www.srim.org 2 Incident proton on Al: Bohr model v=v 0 E p =0.025 MeV relativistic effect E p =938 MeV
Chapter II: Interactions of ions with matter 1 Trajectories of α particles of 5.5 MeV Source: SRIM www.srim.org 2 Incident proton on Al: Bohr model v=v 0 E p =0.025 MeV relativistic effect E p =938 MeV
Atoms, Molecules and Solids (selected topics)
 Atoms, Molecules and Solids (selected topics) Part I: Electronic configurations and transitions Transitions between atomic states (Hydrogen atom) Transition probabilities are different depending on the
Atoms, Molecules and Solids (selected topics) Part I: Electronic configurations and transitions Transitions between atomic states (Hydrogen atom) Transition probabilities are different depending on the
Chapter 2 Experimental sources of intermolecular potentials
 Chapter 2 Experimental sources of intermolecular potentials 2.1 Overview thermodynamical properties: heat of vaporization (Trouton s rule) crystal structures ionic crystals rare gas solids physico-chemical
Chapter 2 Experimental sources of intermolecular potentials 2.1 Overview thermodynamical properties: heat of vaporization (Trouton s rule) crystal structures ionic crystals rare gas solids physico-chemical
Big Idea #5: The laws of thermodynamics describe the essential role of energy and explain and predict the direction of changes in matter.
 KUDs for Unit 6: Chemical Bonding Textbook Reading: Chapters 8 & 9 Big Idea #2: Chemical and physical properties of materials can be explained by the structure and the arrangement of atoms, ion, or molecules
KUDs for Unit 6: Chemical Bonding Textbook Reading: Chapters 8 & 9 Big Idea #2: Chemical and physical properties of materials can be explained by the structure and the arrangement of atoms, ion, or molecules
PH 548 Atomistic Simulation Techniques
 PH 548 Atomistic Simulation Techniques Lectures: Lab: 2-0-2-6 Tuesday 12-1 PM Thursday 12-1 PM Monday 2-5 PM P. K. Padmanabhan PH548: Two Excellent Books How to do it? 1 2 F 12 F 23 3 F i = F ij j i F
PH 548 Atomistic Simulation Techniques Lectures: Lab: 2-0-2-6 Tuesday 12-1 PM Thursday 12-1 PM Monday 2-5 PM P. K. Padmanabhan PH548: Two Excellent Books How to do it? 1 2 F 12 F 23 3 F i = F ij j i F
Molecular Simulation II. Classical Mechanical Treatment
 Molecular Simulation II Quantum Chemistry Classical Mechanics E = Ψ H Ψ ΨΨ U = E bond +E angle +E torsion +E non-bond Jeffry D. Madura Department of Chemistry & Biochemistry Center for Computational Sciences
Molecular Simulation II Quantum Chemistry Classical Mechanics E = Ψ H Ψ ΨΨ U = E bond +E angle +E torsion +E non-bond Jeffry D. Madura Department of Chemistry & Biochemistry Center for Computational Sciences
From Dynamics to Thermodynamics using Molecular Simulation
 From Dynamics to Thermodynamics using Molecular Simulation David van der Spoel Computational Chemistry Physical models to describe molecules Software to evaluate models and do predictions - GROMACS Model
From Dynamics to Thermodynamics using Molecular Simulation David van der Spoel Computational Chemistry Physical models to describe molecules Software to evaluate models and do predictions - GROMACS Model
Complete and precise descriptions based on quantum mechanics exist for the Coulombic/Electrostatic force. These are used to describe materials.
 The forces of nature: 1. Strong forces hold protons and neutrons together (exchange of mesons) 2. Weak interactions are involved in some kinds of radioactive decay (β-decay) 3. Coulombic or electrostatic
The forces of nature: 1. Strong forces hold protons and neutrons together (exchange of mesons) 2. Weak interactions are involved in some kinds of radioactive decay (β-decay) 3. Coulombic or electrostatic
Force Fields for Classical Molecular Dynamics simulations of Biomolecules. Emad Tajkhorshid
 Force Fields for Classical Molecular Dynamics simulations of Biomolecules Emad Tajkhorshid Theoretical and Computational Biophysics Group, Beckman Institute Departments of Biochemistry and Pharmacology,
Force Fields for Classical Molecular Dynamics simulations of Biomolecules Emad Tajkhorshid Theoretical and Computational Biophysics Group, Beckman Institute Departments of Biochemistry and Pharmacology,
NMR BMB 173 Lecture 16, February
 NMR The Structural Biology Continuum Today s lecture: NMR Lots of slides adapted from Levitt, Spin Dynamics; Creighton, Proteins; And Andy Rawlinson There are three types of particles in the universe Quarks
NMR The Structural Biology Continuum Today s lecture: NMR Lots of slides adapted from Levitt, Spin Dynamics; Creighton, Proteins; And Andy Rawlinson There are three types of particles in the universe Quarks
Why Proteins Fold? (Parts of this presentation are based on work of Ashok Kolaskar) CS490B: Introduction to Bioinformatics Mar.
 Why Proteins Fold? (Parts of this presentation are based on work of Ashok Kolaskar) CS490B: Introduction to Bioinformatics Mar. 25, 2002 Molecular Dynamics: Introduction At physiological conditions, the
Why Proteins Fold? (Parts of this presentation are based on work of Ashok Kolaskar) CS490B: Introduction to Bioinformatics Mar. 25, 2002 Molecular Dynamics: Introduction At physiological conditions, the
Swelling and Collapse of Single Polymer Molecules and Gels.
 Swelling and Collapse of Single Polymer Molecules and Gels. Coil-Globule Transition in Single Polymer Molecules. the coil-globule transition If polymer chains are not ideal, interactions of non-neighboring
Swelling and Collapse of Single Polymer Molecules and Gels. Coil-Globule Transition in Single Polymer Molecules. the coil-globule transition If polymer chains are not ideal, interactions of non-neighboring
Nanotube AFM Probe Resolution
 Influence of Elastic Deformation on Single-Wall Carbon Nanotube AFM Probe Resolution Ian R. Shapiro, Santiago D. Solares, Maria J. Esplandiu, Lawrence A. Wade, William A. Goddard,* and C. Patrick Collier*
Influence of Elastic Deformation on Single-Wall Carbon Nanotube AFM Probe Resolution Ian R. Shapiro, Santiago D. Solares, Maria J. Esplandiu, Lawrence A. Wade, William A. Goddard,* and C. Patrick Collier*
