Drosophila homologues of the transcriptional coactivation complex subunits
|
|
|
- Laura Dawson
- 5 years ago
- Views:
Transcription
1 Development 128, (2001) Printed in Great Britain The Company of Biologists Limited 2001 DEV Drosophila homologues of the transcriptional coactivation complex subunits TRAP240 and TRAP230 are required for identical processes in eye-antennal disc development Jessica E. Treisman Skirball Institute for Biomolecular Medicine and Department of Cell Biology, New York University School of Medicine, 540 First Avenue, New York, NY 10016, USA ( Accepted 5 December 2000; published on WWW 23 January 2001 SUMMARY We have identified mutations in two genes, blind spot and kohtalo, that encode Drosophila homologues of human TRAP240 and TRAP230, components of a large transcriptional coactivation complex homologous to the yeast Mediator complex. Loss of either blind spot or kohtalo has identical effects on the development of the eye-antennal disc. Eye disc cells mutant for either gene can express decapentaplegic and atonal in response to Hedgehog signaling, but they maintain inappropriate expression of these genes and fail to differentiate further. Mutant cells in the antennal disc lose expression of Distal-less and misexpress eyeless, suggesting a partial transformation towards the eye fate. blind spot and kohtalo are not required for cell proliferation or survival, and their absence cannot be rescued by activation of the Hedgehog or Notch signaling pathways. These novel and specific phenotypes suggest that TRAP240 and TRAP230 act in concert to mediate an unknown developmental signal or a combination of signals. Key words: TRAP240, TRAP230, blind spot, Transcription, Mediator, Drosophila melanogaster, Eye development INTRODUCTION Development requires the interpretation of specific temporal and spatial signals to yield tightly coordinated patterns of gene transcription. This process has been well studied in the Drosophila eye imaginal disc. The eye disc is determined during embryonic and larval stages by the action of the Pax6 transcription factors Eyeless (Ey; Quiring et al., 1994) and Twin of Eyeless (Toy; Czerny et al., 1999) and the downstream transcription factors encoded by eyes absent (eya; Bonini et al., 1993), sine oculis (so; Cheyette et al., 1994; Serikaku et al., 1994) and dachshund (dac; Mardon et al., 1994). Misexpression of these genes in other imaginal discs, particularly in combination, can lead to ectopic eye development (Bonini et al., 1997; Chen et al., 1997; Czerny et al., 1999; Halder et al., 1995; Pignoni et al., 1997; Shen and Mardon, 1997). The antennal disc primordium also expresses ey and toy at embryonic stages (Halder et al., 1998) and is more easily transformed toward the eye fate than other discs (Bonini et al., 1997; Chen et al., 1997; Pignoni et al., 1997). In the third larval instar a wave of photoreceptor differentiation initiates at the posterior margin of the eye disc and propagates towards the anterior; its front is marked by an indentation in the disc known as the morphogenetic furrow (Ready et al., 1976). Initiation of the morphogenetic furrow requires the secreted factors Hedgehog (Hh) and Decapentaplegic (Dpp), which are expressed at the posterior margin prior to initiation (Borod and Heberlein, 1998; Burke and Basler, 1996; Cho et al., 2000; Dominguez and Hafen, 1997; Royet and Finkelstein, 1997; Wiersdorff et al., 1996). Progression of differentiation also requires both these signals (Chanut and Heberlein, 1997; Heberlein et al., 1993; Ma et al., 1993), although at this stage they are partially redundant. Clones of cells mutant for smoothened (smo), which encodes the Hh receptor, show a partial inhibition of differentiation (Strutt and Mlodzik, 1997), while internal clones of cells mutant for components of the Dpp pathway differentiate almost normally (Burke and Basler, 1996; Wiersdorff et al., 1996). Only in cells unable to respond to either Hh or Dpp is differentiation completely blocked (Curtiss and Mlodzik, 2000; Greenwood and Struhl, 1999). A third signal, Wingless (Wg), is present at the dorsal and ventral margins of the eye disc and prevents ectopic initiation from occurring at those locations (Ma and Moses, 1995; Treisman and Rubin, 1995). Hh is required for the expression of atonal (ato; Borod and Heberlein, 1998; Dominguez, 1999; Greenwood and Struhl, 1999), which encodes a helix-loop-helix transcription factor (Jarman et al., 1993), in a stripe of cells anterior to the morphogenetic furrow. ato expression is subsequently refined to groups of cells and finally to single R8 photoreceptors (Jarman et al., 1994). Both the initial high-level expression of ato and its refinement require the Notch (N) receptor and its ligand Delta (Dl; Baker et al., 1996; Baker and Yu, 1997; Cagan and Ready, 1989). ato is required for the formation of
2 604 J. E. Treisman R8 photoreceptors and thus indirectly for all photoreceptors (Jarman et al., 1994; Jarman et al., 1995), as R8 recruits the remaining cells in the cluster by producing the ligand Spitz for the EGF receptor (Freeman, 1996; Tio and Moses, 1997). Ato function is antagonized anterior to the furrow by the helixloop-helix proteins Hairy (H) and Extramacrochaetae (Emc; Brown et al., 1995); interestingly, expression of h in a stripe anterior to the ato domain appears to be directly activated by Dpp and Hh signaling (Greenwood and Struhl, 1999; Heberlein et al., 1995). Regulation of gene transcription is a complex process requiring not only specific transcriptional activators and repressors, the activity of which may be modified by cell-cell signaling, but also more general factors. Chromatin remodeling complexes are thought to control the accessibility of regulatory sequences to other factors (Kingston and Narlikar, 1999). Another class of mammalian complexes related to the yeast Mediator complex (Kim et al., 1994) has recently been described; such complexes appear to be required as coactivators or in some cases corepressors that link specific factors to the basal transcriptional machinery (Malik and Roeder, 2000). The thyroid hormone receptor-associated protein (TRAP) 25-subunit complex was identified through its ligand-dependent association with the thyroid hormone receptor (Fondell et al., 1996). It appears to be very similar or identical to the SRB and MED-containing cofactor (SMCC) complex isolated by binding to the epitope-tagged SRB10 subunit (Gu et al., 1999; Ito et al., 1999), as well as to the vitamin D receptor interacting protein (DRIP) complex isolated by its association with liganded vitamin D receptor (Rachez et al., 1998; Rachez et al., 1999) and the activatorrecruited cofactor (ARC) complex isolated by binding to SREBP (Naar et al., 1999). Other complexes with similar composition include negative regulator of activated transcription (NAT; Sun et al., 1998), murine Mediator (Jiang et al., 1998), human Mediator (Boyer et al., 1999), cofactor required for Sp1 (CRSP; Ryu et al., 1999) and positive cofactor 2 (PC2; Malik et al., 2000). Several of these complexes have been shown to act as coactivators that stimulate transcription in vitro with a variety of activators, including nuclear receptors but also VP16, p53, Sp1, SREBP, E1A and GAL4. The NAT and SMCC complexes can also repress transcription (Gu et al., 1999; Sun et al., 1998). The roles of the individual subunits of these complexes have mostly not been defined; however, nuclear receptors have been shown to bind directly to TRAP220/DRIP205 (Yuan et al., 1998; Hittelman et al., 1999). Fibroblasts from mice defective for the TRAP220 gene show a reduction in thyroid hormonestimulated transcription, while the mice have abnormalities of heart and brain development (Ito et al., 2000). The glucocorticoid receptor also binds to DRIP150/TRAP170 through its constitutive AF-1 activation domain (Hittelman et al., 1999). The VP16 and p53 activators bind to TRAP80 (Ito et al., 1999), and E1A interacts with human Sur-2 (Boyer et al., 1999). Interestingly, TRAP240 and TRAP230 are absent from the PC2 and CRSP complexes; as PC2 still exhibits a broad coactivator activity (Malik et al., 2000), no biochemical function has yet been ascribed to these subunits. We have isolated mutations in the blind spot (bli) and kohtalo (kto) genes, which we have shown to encode homologues of TRAP240 and TRAP230 respectively, in a screen for mutations affecting early eye development in Drosophila. Mutations in the two genes have identical phenotypes; both block photoreceptor differentiation after the initial expression of ato and dpp has been established, and prolong the expression of these genes and others expressed anterior to the morphogenetic furrow. Neither gene is required for growth, and mutant cells do not influence the growth or differentiation of surrounding cells. In the antennal disc, bli and kto are required to prevent the expression of ey and promote the expression of Distal-less (Dll). Cells mutant for bli or kto cannot be induced to differentiate as photoreceptors by activation of the Hh or N signaling pathways. TRAP240 and TRAP230 thus seem to act together to promote specific cell fate decisions, transmitting either a novel signal or a combination of signals. MATERIALS AND METHODS Genetics The mosaic screen for mutations affecting early eye development will be described in detail elsewhere. Briefly, males carrying an FRT site at position 80B were mutagenized with 35 mm EMS and crossed to females carrying the same FRT site, a P(w + ) element at position 70C, and FLP recombinase driven by the eyeless (ey) promoter on the X chromosome (eyflp1; Newsome et al., 2000). Flies were screened in the next generation for scars or head cuticle in the eye in conjunction with the absence of photoreceptors in w clones. In a screen of 76,230 F 1 flies, we identified 19 alleles of kto and 15 alleles of bli. Although these kto alleles fail to complement the lethality of kto 1 and Df(3L)kto 2, the kto 1 chromosome also carries a second cell-lethal mutation that prevented the recovery of any kto 1 homozygous clones in the eye disc. The transgenic lines used were dpp-lacz (Blackman et al., 1991), wg P (Kassis et al., 1992), arm-lacz (Vincent et al., 1994), Ubi-GFP (Davis et al., 1995), UAS-hh (Azpiazu et al., 1996), UAS- HACi(m1-4) (Chen et al., 1999), UAS-N intra (Doherty et al., 1996), ey-gal4 (Hazelett et al., 1998) and UAS-FLP (Duffy et al., 1998). The clones shown were generated using eyflp, which is active in the antennal disc as well as the eye disc, except those in Figs 2C-D and 3D, which were generated using hsflp122 induced by a 1 hour heat shock at 38.5 C in both first and second instars, and those in Fig. 5A- F, which were generated using ey-gal4, UAS-FLP. The ey-gal4 simultaneously drove the expression of UAS-hh, UAS-HACi(m1-4) or UAS-N intra. The clone in Fig. 2F was made by crossing FRT80, bli T413 /TM6B males to y, w, eyflp1; FRT80, M(3)67C, P(w + )/TM6B females (Ito and Rubin, 1999). Immunohistochemistry and in situ hybridization Third instar larval eye discs were stained with antibodies and X-gal as described (Hazelett et al., 1998). Antibodies used were rat anti-elav (1:1; Robinow and White, 1991), rabbit anti-atonal (1:5000; Jarman et al., 1994), rabbit anti-β-galactosidase (1:5000; Cappel), mouse antiβ-galactosidase (1:200; Promega), rat anti-ci (1:1; Motzny and Holmgren, 1995), mouse anti-h (1:4; Brown et al., 1995), rabbit anti- Ey (1:5000; a gift from Uwe Walldorf), and mouse anti-dac (1:200; Mardon et al., 1994). Fluorescence images were obtained using a Leica TCS NT confocal microscope. Antisense RNA probes labeled with digoxigenin-utp homologous to subfragments of the coding regions of bli and kto were used for in situ hybridization; imaginal discs were hybridized as described previously (Maurel-Zaffran and Treisman, 2000). Molecular biology Plasmid rescue from the l(3)l7062 line was performed as described (Mlodzik et al., 1990) and DNA adjacent to l(3)rk760 was isolated
3 TRAP240 acts with TRAP230 in the eye disc 605 by inverse PCR. Sequence from these rescued fragments was used to align the P insertions on genomic sequence available from the Berkeley Drosophila genome project (GenBank accession number AC013164). Nearby genomic sequence showed homology to the ESTs GM05325 and LD10285; these were obtained from the genome project and used to screen the LD embryonic cdna library also obtained from the genome project. The sequence of the longest cdna obtained has been submitted to GenBank (accession number AF324425). A predicted gene encoding a TRAP230 homologue (CG8491) was identified by searching the Drosophila genome using tblastn. We amplified a 1.5 kb fragment of this gene from wild-type genomic DNA by PCR and used it to screen the LD library. The sequence of the longest cdna obtained has been submitted to GenBank (accession number AF324426). To identify sequence changes in bli mutant alleles, bli T13, bli T606 and bli T413 were balanced over TM6B, P(Ubi-GFPS65T) and homozygous late embryos or first instar larvae were selected by lack of GFP fluorescence and used to prepare genomic DNA. PCR primers were designed to amplify bli exons in fragments of 500 bp-2 kb, which were then sequenced. All observed sequence changes were confirmed on a second independent PCR reaction. The same procedure was followed to identify sequence changes in kto T555, kto T241 and kto T631. RESULTS Mutations in blind spot and kohtalo block photoreceptor differentiation We have undertaken a systematic screen for genes that are required for cells in the Drosophila eye disc to differentiate into retinal tissue, regardless of whether these genes also act at earlier developmental stages (Benlali et al., 2000). The basis of the screen was to use the FLP-FRT system (Xu and Rubin, 1993) to create clones of cells homozygous for a newly induced mutation in the eye of an otherwise heterozygous fly. We used an enhancer from the eyeless (ey) gene to express FLP recombinase specifically in the eye disc (Newsome et al., 2000). Mosaic flies were screened for scars or head cuticle in the eye in conjunction with the absence of photoreceptors derived from mutant cells marked with white (w). In this way we identified mutations that prevented cells from differentiating as photoreceptors but allowed them to survive long enough to prevent their replacement by surrounding cells. In our screen on the left arm of the third chromosome, we identified two complementation groups with identical mutant phenotypes (Fig. 1). One of these groups failed to complement the lethality of the trithorax group gene kohtalo (kto; Kennison and Tamkun, 1988); the other is a novel gene that we have named blind spot (bli). We isolated 19 alleles of kto and 15 alleles of bli; at least 10 alleles of each gene were examined by staining larval eye discs with an antibody to the neuronalspecific protein Elav (Robinow and White, 1991) and were found to have indistinguishable phenotypes, suggesting that they are all at least strong loss-of-function alleles. Clones of cells mutant for either kto or bli contained few or no photoreceptors expressing Elav; when photoreceptors were present they were found in posterior regions of the clone (Fig. 1). Most of these cells apparently die later in development, as scars, rather than w clones, were observed in adult eyes (data not shown). Interestingly, the clones appeared to grow to the same size as their wild-type twin spots, and had no effect on the growth of surrounding cells or the shape of the disc as a whole (Figs 1, 2D). Wild-type cells anterior to mutant clones Fig. 1. bli and kto mutations cause an identical block to photoreceptor differentiation. (A-D) Third instar larval eye discs stained with anti-elav antibody (brown) to mark developing photoreceptors, and with X-gal (blue) to reveal arm-lacz expression that marks the wild-type tissue. (A) Clones homozygous for bli T616 ; (B) clones homozygous for bli T773 ; (C) clones homozygous for kto T631 ; and (D) clones homozygous for kto T314. Anterior is to the left and arrows mark the morphogenetic furrow in this and subsequent figures. appeared to be at approximately the appropriate stage of development; thus mutant cells did not prevent the spread of differentiation-promoting signals even though they did not themselves respond to these signals. As the bli and kto phenotypes appeared indistinguishable, they will be shown interchangeably in future figures; however, almost all experiments have been carried out using alleles of both genes with identical results. bli and kto mutant cells initiate differentiation and then arrest To determine at what stage differentiation was blocked in bli and kto mutants, we examined the expression of atonal (ato), a proneural gene (Jarman et al., 1994) that is activated in the morphogenetic furrow by Hedgehog (Hh) signaling (Dominguez, 1999; Greenwood and Struhl, 1999; Heberlein et al., 1995) and subsequently restricted to the R8 photoreceptors (Jarman et al., 1994). In clones of cells mutant for either bli or kto, expression of ato was weaker than normal in the morphogenetic furrow, but persisted in many of the cells within the clone in regions posterior to the furrow (Fig. 2A,B).
4 606 J. E. Treisman Fig. 2. bli and kto mutant cells show prolonged expression of ato, dpp and Ci. All panels show third instar eye discs. A and B are stained with anti- Ato (brown) and X-gal to reveal armlacz expression in blue. (A) kto T314 clones; (B) bli T616 clones. Ato protein is present in many more cells posterior to the furrow in mutant clones than in wild-type tissue. C and D are stained with anti-β-galactosidase to reveal dpplacz expression in red; bli T413 mutant clones are marked by the absence of GFP expression (green in D). E is stained with X-gal to reveal dpp-lacz expression in blue; a bli T413 clone is marked by the lack of brown anti-elav staining. dpp expression is weaker than normal in the morphogenetic furrow but persists posterior to the furrow in bli clones. (F) An eye disc with bli T413 clones made in a Minute background, stained with anti-elav (brown) and X- gal (blue) to reveal dpp-lacz expression. Disorganized photoreceptor clusters can form in eye discs entirely mutant for bli. G and H are stained with an antibody to full-length Ci (red); kto T555 clones are marked by the absence of GFP (green in H). High levels of Ci persist posterior to the furrow in kto mutant cells. Expression could eventually resolve into single cells, but these were found much further posterior in the disc than in surrounding wild-type tissue. The expression of decapentaplegic (dpp) was affected in a similar manner. dpp is normally activated by Hh in a stripe in the morphogenetic furrow (Heberlein et al., 1993; Heberlein et al., 1995; Ma et al., 1993; Strutt and Mlodzik, 1997); clones mutant for bli or kto showed reduced dpp-lacz expression in this stripe, but ectopic expression of dpp-lacz in posterior regions of the eye disc (Fig. 2C-E). Thus cells mutant for bli or kto appear to respond to Hh by activating the expression of ato and dpp, though at lower levels than wild-type cells, and they fail to terminate the expression of these genes at the normal time. To test whether Hh signaling was affected in bli and kto mutant clones, we examined the distribution of full-length Cubitus interruptus (Ci). In the absence of Hh signaling the Ci transcription factor is cleaved to a repressor form that is not recognized by an antibody to the C-terminal domain (Aza- Blanc et al., 1997; Motzny and Holmgren, 1995). In bli or kto mutant clones, we found that the upregulation of full-length Ci was weaker than normal in the morphogenetic furrow, but high levels persisted posterior to the furrow (Fig. 2G,H and data not shown). Thus the response to Hh signaling is not terminated at the normal time; this is consistent with the effects on ato and dpp expression. It has been suggested that Hh is itself required to terminate this response (Dominguez, 1999), so the phenotype could also be due to a lack of late Hh signaling. To address whether the small number of photoreceptors that form in bli or kto mutant clones resulted from non-autonomous rescue by surrounding wild-type cells, we generated clones in a Minute background (Morata and Ripoll, 1975). This gives the bli or kto mutant cells a growth advantage over cells carrying a Minute mutation; in combination with eyflp it results in an eye disc entirely composed of mutant tissue (Benlali et al., 2000; Bohni et al., 1999; Newsome et al., 2000). In such eye discs, some photoreceptors were still able to differentiate, although the size and pattern of clusters were abnormal, and dpp was still expressed (Fig. 2F). The resulting pharate adults had very small eyes composed of bli or kto mutant tissue marked with w (data not shown). Thus a signal from nearby wild-type cells is not necessary for photoreceptors to form from bli or kto mutant cells. bli and kto affect genes expressed anterior to the morphogenetic furrow Because bli or kto mutant cells seemed to be arrested at an early step in their differentiation, at which they express ato and dpp but not Elav, we examined the effects of these mutations on other genes expressed at various stages of differentiation in the eye disc. The Pax-6 homologue Eyeless (Ey) is normally present only in undifferentiated cells anterior to the morphogenetic furrow at the third instar stage (Halder et al., 1998); however, in bli or kto mutant cells Ey continued to be expressed at high levels posterior to the furrow (Fig. 3A-C). Dachshund (Dac), which is also involved in early steps of eye specification (Chen et al., 1997), is expressed in a broad stripe both anterior and posterior to the morphogenetic furrow, but is down-regulated in more posterior regions of the eye disc (Mardon et al., 1994). Again, bli or kto mutant cells posterior to the normal Dac expression domain maintained high levels of Dac protein (Fig. 3D). However, not all genes expressed anterior to the furrow were maintained in bli or kto mutant cells. hairy (h) encodes a helixloop-helix protein that negatively regulates Ato activity and is expressed in a stripe just anterior to the furrow (Brown et al.,
5 TRAP240 acts with TRAP230 in the eye disc 607 Fig. 3. Effects of bli and kto on genes expressed anterior to the morphogenetic furrow. All panels show third instar eye discs. A and C are stained with anti-ey antibody (red); blit702 clones are marked in B and C by the absence of green antiβ-galactosidase staining reflecting armlacz expression. D is stained with antidac antibody (brown); blit751 clones are marked by the lack of blue X-gal staining reflecting arm-lacz expression. Ey and Dac expression persist posterior to the furrow in bli mutant clones. E and G are stained with anti-h antibody in green; blit342 clones are marked in F,G by the absence of red anti-βgalactosidase staining reflecting armlacz expression. H expression is autonomously reduced in bli mutant clones and low levels persist immediately posterior to the normal expression domain. (H) blit762 mutant clones marked by lack of brown antielav staining; wg-lacz expression is revealed by blue X-gal staining. bli clones do not ectopically express wg. 1991; Brown et al., 1995; Rushlow et al., 1989); its expression was autonomously reduced in bli or kto mutant cells, and low level expression was maintained only immediately posterior to its normal domain (Fig. 3E-G). This is similar to the effect on h of loss of the Dpp receptor encoded by thick veins (tkv; Greenwood and Struhl, 1999). Ectopic expression of wingless (wg) can block photoreceptor differentiation and lead to Ey maintenance (Treisman and Rubin, 1995; J. Lee and J. E. T., unpublished data). However, in discs containing clones of cells mutant for bli or kto, wg was still restricted to its normal expression domain at the anterior lateral margins (Fig. 3H). Antennal disc cell fates are altered in bli and kto mutants In addition to its maintenance in bli or kto mutant cells in the eye disc posterior to the furrow, we also observed ey misexpression in mutant cells in the antennal disc. However, Ey was not present in every mutant clone and did not always occupy the whole clone (Fig. 4A-D). It is not clear why some mutant cells failed to express ey, as this lack of expression did not appear to correlate with a particular position within the disc. The observed expression of ey suggested that mutant antennal disc cells could be partially transformed towards an eye fate. In agreement with this, expression of the homeobox gene Distal-less (Dll) was lost in bli or kto mutant clones in the antennal disc (Fig. 4E-F); Dll is normally expressed in a broad central domain of the antennal disc but is not found in the eye disc (Cavodeassi et al., 2000; Diaz-Benjumea et al., 1994). In contrast to the non-autonomy observed for ey, Dll expression was autonomously lost in all mutant cells. Although expression of ey in the antennal disc can induce ectopic eyes (Halder et al., 1995), we did not observe these because of the requirement for bli and kto in photoreceptor differentiation. However, mutant clones in the antennal disc did not misexpress ato, and in fact failed to express ato in its normal domain, suggesting that the transformation of antenna to eye was not complete (data not shown). Fig. 4. bli or kto mutant cells in the antennal disc misexpress ey and fail to express Dll. All panels show third instar antennal discs. A-D are stained with anti-ey antibody in red. (A,B) ktot631 clones are marked by the lack of GFP expression (green in B); (C,D) blit413 clones are marked by the lack of GFP expression (green in D). ey is misexpressed in some, but not all, mutant cells. E-F are stained with anti-β-galactosidase to reveal Dll-lacZ expression in red. blit413 clones are marked by the lack of GFP expression (green in F). Dll expression is autonomously lost in bli mutant clones.
6 608 J. E. Treisman Another interesting feature of clones in both the antennal disc and the eye disc was that their borders were smooth and rounded, indicating that cells mutant for bli or kto have an altered cell affinity preventing them from mixing normally with wild-type cells (Morata and Lawrence, 1975). This property was observed for clones occurring in several different proximal-distal domains of the antennal disc (Fig. 4 and data not shown). Hh or N signaling is not sufficient to rescue bli or kto mutants Because the effects on ato, dpp and Ci suggested that Hh signaling might be altered in bli and kto mutants, we attempted to rescue mutant clones by misexpressing hh throughout the eye disc. However, photoreceptors were still absent from bli or kto mutant clones induced in this background (Fig. 5A and data not shown). To test whether the mutations were blocking Hh signaling downstream of the expression of Hh, we also tried rescuing with an activated form of Ci in which four sites phosphorylated by protein kinase A have been mutated (Chen et al., 1999). However, clones of bli or kto mutant cells generated in eye discs expressing activated Ci still lacked Elavstained photoreceptors, failed to refine Ato, and maintained Ey expression (Fig. 5B-D and data not shown). Thus the action of bli and kto is epistatic to activation of the Ci protein. The Notch (N) pathway is also important in refining the expression of Ato (Baker et al., 1996) and could be affected by bli and kto. However, expression of the constitutively active intracellular domain of N (Doherty et al., 1996) throughout the eye disc did not alter the misexpression of Ato or the lack of Elav in bli or kto mutant clones (Fig. 5E-F and data not shown). bli and kto encode homologues of the human TRAP240 and TRAP230 proteins We identified two P element alleles of bli in the stock center collections, l(3)l7062 and l(3)rk760. Genomic DNA surrounding these P element insertions at 78A2 was isolated by plasmid rescue (for l(3)l7062) or inverse PCR (for l(3)rk760), and its sequence was used to align the insertions on genomic sequence available from the Berkeley Drosophila genome project. Nearby genomic sequence showed homology to several expressed sequence tags (ESTs) that we obtained from the genome project. We then used these ESTs as probes to isolate a full-length 9.8kb cdna from an embryonic cdna library. Both P elements are inserted in the third intron of the corresponding transcript (Fig. 6A). The sequence of the encoded 2618 amino acid protein is shown in Fig. 6C. To confirm that this transcript corresponded to the bli gene, we amplified genomic DNA from several of our EMS-induced alleles of bli and looked for sequence changes. We found changes in three alleles that would introduce stop codons into the predicted open reading frame and would thus be likely to abolish the function of the protein (Fig. 6B). The protein showed two regions of strong homology to the human protein TRAP240 (Fig. 6B,C), which was isolated as part of a large transcriptional coactivation complex that associates with ligand-bound thyroid hormone receptor and vitamin D receptor (Fondell et al., 1996; Ito et al., 1999). The function of TRAP240 within this complex has not been defined. We used deficiency and recombination mapping to map our kto alleles to 76B1-D5, in agreement with the previously Fig. 5. Activation of the Hh or N pathways does not rescue bli or kto mutant cells. All panels show third instar eye discs from flies with ey-gal4 and (A) UAS-hh; (B-D) UAS-ci(m1-4); (E,F) UAS-N intra. A and B show anti-elav staining in brown; kto T843 clones (A) and bli T606 clones (B) are marked by the lack of blue X-gal staining reflecting arm-lacz expression. Photoreceptors are absent from bli or kto clones even in the presence of ectopic Hh or activated Ci. (C-F) show anti-ato staining in green; (C,D) kto T631 clones are marked by the lack of anti-β-galactosidase staining reflecting armlacz expression (red in D); (E,F) bli T606 clones are marked by the lack of anti-β-galactosidase staining reflecting arm-lacz expression (red in F). Ato expression persists in posterior bli or kto mutant clones even in the presence of activated Ci or activated N. defined map position of the gene (Kennison and Tamkun, 1988). We were unable to find any P elements that failed to complement kto. However, we reasoned that kto was likely to encode another component of the TRAP complex, since its mutant phenotype was identical to that of bli. Searching the Drosophila genome project database for homologues of the human TRAP complex proteins revealed that a predicted TRAP230 homologue, CG8491, mapped to 76D1. We used PCR to amplify a fragment of this predicted gene, which we used as a probe to screen an embryonic cdna library and isolate full-length 7.7 kb cdnas. The genomic organization of this transcript is shown in Fig. 7A and the sequence of the
7 TRAP240 acts with TRAP230 in the eye disc 609 l(3)l7062 l(3)rk760 1 kb A Q7 stop Q369 stop Q1152 stop T606 T13 T413 B 32% identity 32% identity C Fly 1 MTHQNHQTNGASLEDCHTNFYALTDLCGIKWRKFV-NGERPNASSDPLA--DPILRSYSRCIQADMLCVWRRVQSTK 74 Human 1 MSASFVPNGASLEDCHCNLFCLADLTGIKWKKYVWQGPTSAPILFPVTEEDPILSSFSRCLKADVLGVWRRDQRP- TDHADPNALTFEMTTSTKVHPPLSLAAAKELWIFWYGEEPDLSELVDAELLRVAANQALWNGTWKGALTYECRSLLFKALHNLMERF GRRELWIFWWGEDPVLLTLFTMTYQKKKMECGRMDFPMNAVLCFS------KAVHNLLERC VLTKDIVRFGKWFVQPCTSSDRLFGRSSQHLSFSFTFFVHGDT-VCASIDLREHPAVRPLTKEHLTEAAAAFAAASSPPGSNGSAAS 247 LMNRNFVRIGKWFVKPYEKDEKPINK-SEHLSCSFTFFLHGDSNVCTSVEINQHQPVYLLSEEHITLAQQSNSPF AGGAVPNPGQDPNGASMDGLDGGEGAAKAAPPPHARKVMLAPFGIAGILTGNSYKASDPIAEKILEDWASFFPL -CNKD QVILCPFGLNGTLTGQAFKMSDSATKKLIGEWKQFYPISCCLKEMSEEKQE ----NTDVPPVVEVVSGGHKMYHPTNYVLVTDLDDMEHMEFVEMQKMQSSVAGAAAESAVLASLSCPPGAASAPSSMGAAPSATAVP 408 DMDWEDDSLAAVEVLVAGVRMIYPACFVLVPQSD 289 APSATAVPLGPASLNAPALAGAGVGATASSAAKEISRKAAPQAVSALERLAFQPYYDQRPTSGFTFNTNNTHIPASAAVEMPERTWQ 487 DCVMNTLHVDAAAAAAAVASSTPASGTGSLSADGDENEQNKPPQDSKQLVQQQIQQQQQRQKLWNFVDPMQKAPCICTKHLGNTPHG 574 TPHGGASTYSRNSLGGDSSMPVASVESPATPAPSPHPNSAHSQPTSVPPAEQLLNMSPHAPTSVSNLQQPPTPIDHLLDKNTPAPTP 661 TDQHDSKSITASPYVHQTPSVEPPSYTDHAAGGGPAGGQGLGTGPGSVPAQQPATPTAATSAGGAGSGGPSNAIGANAVGTISVKKL 748 EMQQQTPSAAMAIKQEPGAQGRGVGGVTSTTEALNNFKRLYNPPKLTLKDPDSFYDEEWLKEVIYDFQYQEYWDYSTVKRPKMEKQR 835 RPRYAKNLYEGQNHVKPVMPSPGSVYGSQLLSLDESASQAGGRGGGGQAAGSSGGGLVGIANSTASGSDVEADGSSFFQGLNIKTEP 922 GLHSPSCKETSKSSGGNSSGGGSGSGGNLFTAEGLNPSLNDLEQLFETSSNDECSSVQIHTPPDSNNPSNGGCSAVTNTIEDLKRST 1009 AVASAAVAAAAAAAASGAGNIQAEDLTKMFPTPPSHEQQHPNSSPCQTDVVMTDLSVDTTTTIITSSITTTCNTTITSSINTTTTAC 1096 SNPSNSIMLAAAQAPVTVVAIQTVSKMVKQEYNLELGSPMEEPINDWDYVYRPPQQEKFVGSTRYAPLTNLPSQTQPPLTLPTGCFY 1183 QPTWSSHKSRAATLAKAAAAQQQQHQKHQALQQRIQLHQQKLQQLQLQNQQQQQAAAAAAAAASGGVGHQKHQHQHLHDLLSAAPRT 1270 PLTPSTVPQPLSSGGSQYLLNQLNCPQAPPGASMQQLMHRAGMSPISPGPGMGPYAARSSPMSRATPTHPPPPYPYDLAVASPATST 1357 SSYLNRPLHSQEHPHMHGLGGGATGVVGHGTGGGGHMGMVAYTGDAGIVSGGTAMAAGSSSLLQELPEVNSVLVNILLYDTALNVFR IPEAHSLYVNLILSESVMNLFK DHNFDSSSVCVCNADTQKIGNIRGADSGVYVPLPYVPLPGVSFNPFPSGAGGALAGQRMLNGPSSAGFGGMRMISAFGGSPASASMP 1526 DCNSDSCCICVCNM------NIKGADVGVYIPDP 1114 Fig. 6. bli encodes a TRAP240 homologue. (A) Genomic organization of the bli locus. Exons are shown as boxes with the translated regions filled in. The two P element insertions causing bli mutations are shown as inverted triangles at their insertion positions in the third intron. (B) Diagram of the Bli protein. The blocks of homology to TRAP240 are shaded. The positions of the mutations in bli T606, bli T13 and bli T413 are shown. (C) Amino acid sequence of the Bli protein (fly) with homologous regions of TRAP240 (human) aligned below. Identical amino acids appear white on a black background, and similar amino acids appear white on a gray background. Only regions identified as homologous by the blastp program are shown. GAGSGHGHGPNGGSNSSSCTPPSSNPHITGYVDDDPVE----CTCGFSAVVNRRLSHRAGLFYEDEVEITGIADDPGRNKQPTLLSI TQEAQYRCTCGFSAVMNRKFGNNSGLFLEDELDIIGRNTDCGKEAEKRF-EA Figure 6C IQSLSRKNQNKQGPGETSSALDKIGAGGLPNGQLEQLGHAVFDLLLDQCSIIQTSSSSVHRALQSHRRRMSRQRRIFGNNGAPTASL 1696 LRATSAEHVN GGLKES--EKLSDDLILLLQDQCTNLFSPFGAADQ DPFPKSGV ASIANVLEFMDAHDVISLALEQSRLAFENQRMDNMMDFHGNGSSSSHQQQQLTAFHAPPPALRHKLAGIGAGRLTVHKWPYLPVGFT 1783 ISNWVRVEERDCCNDCYLALEHGR-----QFMDNM-----SGGKVDEALVKSSCLH -PWSKRNDVS M RSNKEIVRTMNAIQPMLQNAFHCKSRGGSGSKDASSYNTVSGPLTWRQFHRLAGRAS----GQCEPQPIPSVVVGYEKDWISVAPHS 1866 QCSQDILRMLLSLQPVLQDAIQKKRTVRPWG VQGPLTWQQFHKMAGRGSYGTDESPEPLPIPTFLLGYDYDYLVLSPFA IHYWDKFLLEPYSYARDVVYVVVCPDNEHVVNCTRSYFRELSSTYEMCKLGKHTPIRG--WDGFLQVGAARNNVPADRETTPLDDWL 1951 LPYWERLMLEPYGSQRDIAYVVLCPENEALLNGAKSFFRDLTAIYESCRLGQHRPVSRLLTDGIMRVGSTASKKLSEKLVAEWFSQA RTLEHAALAEQIRRYAVAFIHQLAPYLSRVPNDKTLLNPPDGSGNSHSKGGSSCSSNSSSVSGLPGGDLPTDNIKLEPGTEPQVQPM 2038 ADGNNEAFS-KLKLYAQVCRYDLGPYLASLPLDSSLLSQPN 1478 ETNEIKQEPGVGKGGTAAGETKPTLILGDPLGMGETLEDINPSAIVLYVVNPFTFAS--DSCELERLALIALLRCYAELLKAVPDSV PPAIVVYIIDPFTYENTDESTNSSSVWTLGLLRCFLEMVQTLPPHI RSQMNIQIISLESVMELGPCGNRKRFSDEIRCLALNIFSQCRRHLVHAQSVKSLTGFGTAANMEAFLKTKDEPNRRAYKMYTAPFVL 2210 KSTVSVQIIPCQYLLQPVKHEDREIYPQHLKSLAFSAFTQCRRPLPTSTNVKTLTGFGPGLAMETALRSPDRPE--CIRLYAPPFIL APMHERNDKTDFSRSAGSMHGQNEHRYSVMYCNYCLSEDQAWLLATATDERGEMLEKICINIDVPNRARRRKAPARYVALKKLMDFI 2297 APVKDKQTEL------GETFGEAGQKYNVLFVGYCLSHDQRWILASCTDLYGELLETCIINIDVPNRARRKKSSARKFGLQKLWEWC MGIISQTSQMWRLVIGRIGRIGHSELKSWSFLLSKQQLQKASKQFKDMCKQCTLMY--PPTILSACLVTLEPDAKLRVMPDQFTPDE 2382 LGLVQMSSLPWRVVIGRLGRIGHGELKDWSCLLSRRNLQSLSKRLKDMCRMCGISAADSPSILSACLVAMEPQGSFVIMPDSVSTGS RFSQ---ISMQ-NPLATPQDVTCTHILVFPTSA----VCAPFTRQ-----FQNEPQVDDDFLTFEEEGNEDFSDADIGDLFWDTHMD 2456 VFGRSTTLNMQTSQLNTPQDTSCTHILVFPTSASVQVASATYTTENLDLAFNPNNDGADGMGIFDLLDTGDDLDPDIINIL--PASP RVSNHGSPGRMDDNRSWQSAGGNNFKCTPPQEVEEVGSLNQQPISVGYMVSTAPTGRMPAWFWSACPHLEDVCPVFLKTALHLHVPS 2543 TGSPVHSPGSHYPHGGDAGKGQSTDRLLSTEPHEEVPNILQQPLALGYFVSTAKAGPLPDWFWSACPQAQYQCPLFLKASLHLHVPS IQSADDILNSTNAHQSGNDHPLDSNLTADVLRFVLEGYNALSWLALDSNTHDRLSCLPINVQTLMDLYYLTAAIA 2618 VQS-DELLHSKHS------HPLDSNQTSDVLRFVLEQYNALSWLTCDPATQDRRSCLPIHFVVLNQLYNFIMNML 2174
8 610 J. E. Treisman A 1 kb 8bpdeletion: T44 frameshift and stop Q270 stop Q1946 stop T555 T241 T631 B 38% identity Q-rich Fig. 7. kto encodes a TRAP230 homologue. (A) Genomic organization of the kto locus. Exons are shown as boxes with the translated regions filled in. (B) Diagram of the Kto protein. The region of homology to TRAP240 is shaded. The positions of the mutations in kto T555, kto T241 and kto T631 are shown. (C) Amino acid sequence of the Kto protein (fly) with homologous regions of TRAP230 (human) aligned below. Identical amino acids appear white on a black background, and similar amino acids appear white on a gray background. Only regions identified as homologous by the blastp program are shown. C Fly 1 MLSMLQEKRPLKRTR--LGPPDIYPQDAKQREDELTPTNVKHGFTTTPPLS-DEFGTAHNSNVNASKVSAFFSGV 72 Human 43 EHRPLKRPRPRLGPPDVYPQDPKQKEDELTALNVKQGFNNQPAVSGDEHGSAKNVSFNPAKISSNFSSI LAKKEELMTLPDTGRKKQQINCKDNFWPVSPRRKCTVDAWFKDLAGNKPLLSLAKRAPSFNKKEEIFITLCENQVNMQRATWFIK 157 IAEKLRCNTLPDTGRRKPQVNQKDNFWLVTARSQSAINTWFTDLAGTKPLTQLAKKVPIFSKKEEVFGYLAKYTVPVMRAAWLIK LSAAYTLSFTESKNKKRSIYDPAAEWTGNMIKFMKELLPKLQEYYQQNHDKSSSNGTTSGSLTAAGNGPASNGSTGTSSINSVTG 242 MTCAYYAAISETKVKKRHV-DPFMEWTQIITKYLWEQLQKMAEYYR 241 SSASTNVIPVPSMASPLPPIHSPANGQQAAPGGGVNAGSVMPTTGSLGGVVGGPGSSVVGGAAGAGAAVPPGSTISGIGSQFEDS STIGPLPHDV RNALKYWKYCHQLSKYMYEESLLDRQEFLNWILDLLDKMRTQASFDEPLKKLVLSFALQYMHDFVQSERLCRKMAYIVSKKLAQL 412 EVAIRQWDYTEKLAMFMFQDGMLDRHEFLTWVLECFEKIRPG---EDELLKLLLPLLLRYSGEFVQSAYLSRRLAYFCTRRLALQ LNTV VEQQTIKELDEPKLQQDPYELA----LQEQMSCPHHRDIVLYLSTILQIITIECPTALVWS-GIAAHRAPSSLL 486 LDGVSSHSSHVISAQSTSTLPTTPAPQPPTSSTPSTPFSDLLMCPQHRPLVFGLSCILQTILLCCPSALVWHYSLTDSRIKT--- GSPLDHLPLAPSVLPMPTRCPRTNHEIRRQLRAAESDIVLRTQHAEQRWFAAKWLSAGKN-QYTSVLATLDHLDTHCFDRMEHNN 570 GSPLDHLPIAPSNLPMPEGNSAFTQQVRAKLREIEQQIKERGQAVEVRWSFDKCQEATAGFTIGRVLHTLEVLDSHSFERSDFSN SIDTLYAQIF---PSPTVSRRREEDQVEPRPPYEPKQDKDTV-RILCEWAVSGQRWGEHRAMVVAILLDKRQIDVTSTPADQQSS 650 SLDSLCNRIFGLGPS KDGHEISSDDDAVVSLLCEWAVSCKRSGRHRAMVVAKLLEKRQAEIEAERCGESEA DKDDKDSLASGAGLIDGLPVFQHVLMHFLDHDAPVLDEHVSSPQQRTEFTNLVQLFSALIRHDVFSHNAYMHTLISRGDLLLESV 735 ADEKGS-IASGSLSAPSAPIFQDVLLQFLDTQAPMLTDP-RSESERVEFFNLVLLFCELIRHDVFSHNMYTCTLISRGDL 659 LVIKSGTTATKTSPPPPAPPPTTTHGFDDDGFGGGLDFKHNEFDDSNVDDDLDKLVQNIKEKGQQHEAPDSPKIGPPGDGETNPG 820 GSISRHYVYTKHFPIPQDDPSMSSYSSESNQRYILLFGVGKERDEKKHAVKKMSKEIGKLFTKKFSIDV RHVQYATHFPIPQEE----SCSHECNQRLVVLFGVGKQRDDARHAIKKITKDILKVLNRKGTAETDQLAPIVPLNPGDLTF --AAAGHVKKHSRNEF--NFEATTSKCQQMAYFDQHVVTAQCAANVLEQLNGFALGNNNYLPVQEHVAFLFDLMELALNIYSLLE 970 LGGEDGQKRRRNRPEAFPTAEDIFAKFQHLSHYDQHQVTAQVSRNVLEQITSFALGMSYHLPLVQHVQFIFDLMEYSLSISGLID LCDSLLKELPEVEHQLQLKKSNLVRSYTTSLALYIVSILRRYHSCLLLSPEQTLSVFEGVCRTIRHVSNPSECTSAERCIIAYLS 1055 FAIQLLNELSVVEAELLLKSSDLVGSYTTSLCLCIVAVLRHYHACLILNQDQMAQVFEGLCGVVKHGMNRSDGSSAERCILAYLY DLHESCVLLQGK--EQSTEYYQQLQCIKRFKDIFNTPEQLDLPPQGYNPLLLQELFMAPRRGGKLDPHWLGTLHESPANVYSFVS 1138 DLYTSCSHLKNKFGELFSDF-----CSKVKNTIYCNVEPSE-SNMRWAPEFMIDTLENPAAHTFTYTGLGKSLSENPANRYSFVC Figure 7C NALIAVC-RETDNERLNDVALACAELTASCNVLSEEWIYALQSLCSGSKSPR--YPHLGGQVDIGQLKTHNALAVFVCILVARHC 1220 NALMHVCVGHHDPDRVNDIAILCAELTGYCKSLSAEWLGVLKALCCSSNNGTCGFNDLLCNVDVSDLSFHDSLATFVAILIARQC FSLADFVSKFALPTLARSVSAGGAELSVDAEAGARLTCHLVLKLFKTLEIPQPGMYSVSTSPNPLHAVGN--DFSIRLSCDRHLL 1303 LLLEDLIRCAAIPSLLNAACS-----EQDSEPGARLTCRILLHLFKTPQL NPCQSDGNKPTVGIRSSCDRHLL VGAHKTIPIAAVLAVLKAILIVVDNAALKTPLASGSGTSSGGLGGAFGSGKRSGFNTPVHPGSTPKSNEQRPADLSQILGTSDLQ 1388 AAS VLGDAELK -LGSSLTSEPEALQQPSVGG MEQISLLEFAQAVLKQICAQEHVLERCLKNAEQLCDMIIDEMLTAKQAQRVL 1459 GSGFTVTGGTEELPEEEGGGGSGGRRQGGRNISVETASLDVYAKYVLRSICQQEWVGERCLKSLCEDSNDLQDPVLSSAQAQRLM HMICYPEPEF-NIISELDQRSMIVRILENLGQWTLRISWLDLQLMYRQSLSNNAELNVWLDTVARAAIDVFHME-EVVLPGAVKA 1543 QLICYPHRLLDNEDGENPQRQRIKRILQNLDQWTMRQSSLELQLMIKQTPNN--EMNSLLENIAKATIEVFQRSAETGSSSGSTA THKPKPS TWLVAPLIAKLTPAVQGRILRVAGQVLESMNYFSKVSKSDCNSSGSGDEREKSNSCHSSNSYGLG 1614 SNMPSSSKTKPVLSSLERSGVWLVAPLIAKLPTSVQGHVLKAAGEELEKGQHL GSSSRKERDRQKQKSMSL--- GVPARNKKMPLDYQPFLGLILTCLKGQDEYKENLLVSLYAQLSQCLQSFAELDTIGGIDEPQAREEILDALQLRFSLVGGMFEAI LSQQPFLSLVLTCLKGQDEQREGLLTSLYSQVHQIVNNWRDDQYL---DDCKPKQLMHEALKLRLNLVGGMFDTV QKNSTPTTDWAILLAQLVCQGVVDLSCNRELFTTVVDMLATLVHSTLVSD DERHYMNLMKKLKKEIGEKNNASI 1779 QRSTQQTTEWAMLLLEIIISGTVDMQSNNELFTTVLDMLSVLINGTLAADMSSISQGSMEENKRAYMNLAKKLQKELGERQSDSL RVIRQLLPLYKQPTEVIACEHSG--MDTKGNKICDID----KKQLRISDKQRISVWDILEGHKNPAPLSWVWFGAVKLERKPLTY 1852 EKVRQLLPLPKQTRDVITCEPQGSLIDTKGNKIAGFDSIFKKEGLQVSTKQKISPWDLFEGLKPSAPLSWGWFGTVRVDRRVARG EEAHRNLKYHTHSLVKPSSYYYEPLPLPPEDIEPVPEKICIKDEMKADTPSSVDQSPSAVVGGTGRGRGKGTTTRKRKPKNPKTP 1937 EEQQRLLLYHTHLRPRPRAYYLEPLPLPPEDEEP-PAPTLLEPEKKAPEPPKTDKPGAA 1810 PVVNTQQQQPQLAQQPQQPQNVQQQQLQQQQQQQQHMQQQHMQQQQMQPNQMGQMPMNMPMNMQQFAPNPNNMMQQNAMLQQQQQ 2022 QQMQQMGNNPMQQQLNVGGGNGQPNPQMNFMQQGPGGGGAGPQGMPGQQQQWHNAPQQQQPPQPYHNQYAPHQQNMQSNRIERPP 2107 LNANSKQALSQMLRQRQPFQQQAQQGPGGGFNPMQQQPQASQQQPGPQQQMNPNQMRQQQMNPQQNPQSVAAFNAMQQQPQQNAQ 2192 QQQMNPNQQQQQQFMRGGNMRPGMAPNQMNQMNMGGQGMSQNPMMQQQIPQNMVGMVNPNANQMMQSGGAQGGNGVGVGVGVGVG 2277 GAGNNPNMGMGGMPQQGMIQQQPQQQPQQQVQFQNFQNQYQQQQQQGMQQQGGGAGVGVGVGMAPNQQQQQQANMMGNFNPQMQQ 2362 GNRNNPDFMAAAAVAQQQQQQQQQQRVVPGGMMAGNRNQYMNQAPNVTMSTMMGPGPGGVVGQVPPYARQQSAGGGKPGVLNTQQ 2447 QFQQQQQQQQQLRHQMMQLQGMGGGAGGGMGAGPQQGGGAVGGGAGGGMVPQQQSMNQQQTPNLVAQLQRQNMMGQQQYQPPPY 2531
9 TRAP240 acts with TRAP230 in the eye disc 611 encoded 2531 amino acid protein in Fig. 7C. Homology to TRAP230 is predominantly in the N-terminal two-thirds of the protein, with the remainder of the protein being glutamine-rich in both species (Fig. 7B,C; Ito et al., 1999). We sequenced genomic DNA corresponding to the TRAP230 transcript amplified from several of our kto mutant alleles, and again found changes in three alleles that would cause premature termination of the TRAP230 protein (Fig. 7B). Thus the kto mutant phenotype is likely to result from disruption of TRAP230. We used in situ hybridization to show that both TRAP230 and TRAP240 transcripts were present throughout the eye disc and other imaginal discs (data not shown). This is in agreement with the mutant phenotypes seen when bli or kto function is removed from cells at any location in the eyeantennal disc. DISCUSSION Functions of TRAP240 and TRAP230 in the TRAP complex We have identified two Drosophila homologues of components of the TRAP/DRIP complex based on their requirement for photoreceptor differentiation. The very specific mutant phenotypes of these genes offer a new avenue for understanding the roles of the encoded proteins in transcriptional regulation. The TRAP complex and complexes related to it have been isolated by several different procedures. TRAP240 and TRAP230 are present in the complexes isolated by binding to liganded thyroid hormone receptor, liganded vitamin D receptor, sterol response element binding protein (SREBP), VP16, the p65 subunit of NF-κB, or tagged Srb10 (Fondell et al., 1996; Gu et al., 1999; Ito et al., 1999; Naar et al., 1999; Rachez et al., 1998). However, they are not present in the Sp1 cofactor CRSP (Ryu et al., 1999) or in the PC2 coactivation factor isolated from the USA fraction, which contains many other components of the TRAP complex (Malik et al., 2000). PC2 is sufficient for the general transcriptional coactivation functions previously ascribed to the TRAP complex (Malik et al., 2000); thus TRAP240 and TRAP230 cannot be essential for activation. Our finding that they are required for specific developmental processes in the Drosophila eye, but not for cell growth or viability, suggests that they are not core components required for all activities of the complex. Thus their presence or absence in individual mediator-related complexes may reflect real differences in activity rather than being an artefact of the purification method used. Mutations in kto were previously isolated as suppressors of the Polycomb phenotype, making it a member of the trithorax group of genes (Kennison and Tamkun, 1988). This is consistent with a general role in transcription, as other members of this group have been shown to encode components of a chromatin remodeling complex (Papoulas et al., 1998; Crosby et al., 1999; Collins et al., 1999). TRAP220, which is not consistently present in PC2 (Malik et al., 2000), has been shown to bind directly to the thyroid hormone receptor and other nuclear receptors (Hittelman et al., 1999; Yuan et al., 1998), and is thought to link these to the coactivation complex; the mouse homologue is required for cells to respond normally to thyroid hormone (Ito et al., 2000). It seems unlikely that the nuclear receptor for the steroid hormone ecdysone is the target of bli and kto; loss of ecdysone receptor function does not prevent photoreceptor differentiation (Brennan et al., 2001), and loss of the coreceptor encoded by ultraspiracle causes an acceleration of the morphogenetic furrow (Zelhof et al., 1997). Although there are many other potential nuclear receptors in the genome for which the phenotype in the eye is not known, the interaction of human nuclear receptors with the TRAP complex seems to be specifically through TRAP220 (Yuan et al., 1998). As the Drosophila homologue of TRAP220 also maps to the left arm of the third chromosome, we would have expected to identify mutations in it in our screen if they produced the same phenotype as bli and kto. However, we did not find any other complementation groups with this phenotype. Thus one possibility is that TRAP240 and TRAP230 serve a similar function for an unidentified transcription factor. Alternatively, they may regulate the activity of the TRAP complex. The identical mutant phenotypes we observe for bli and kto strongly suggest that TRAP240 and TRAP230 act as an obligate heterodimer. Interestingly, homologues of TRAP240 and TRAP230 are present in the C. elegans genomic sequence, while homologues of several other components of the TRAP complex, including TRAP220, TRAP100 and TRAP95, are apparently absent; the functions of TRAP240 and TRAP230 may thus be more ancient than that of TRAP220. Mutations in the C. elegans TRAP230 homologue, sop-1, have recently been shown to act as recessive suppressors of a mutation in an enhancer required for the transcription of the pal-1 gene (Zhang and Emmons, 2000). However, sop-1 mutations and RNA interference with sop-1 have no phenotype in animals wild type for pal-1 (Zhang and Emmons, 2000), suggesting either that TRAP230 activity is partially redundant in C. elegans, or that the function of TRAP230 was not completely abolished. C. elegans homologues of the MED6, MED7 and MED10 proteins also found in PC2 have been shown to be required for embryonic survival and the expression of two genes activated early in embryogenesis (Kwon et al., 1999). Functions of bli and kto in eye development The phenotypes caused by bli or kto mutations do not correspond exactly to those due to inhibition or activation of any signaling pathway known to act in eye development. Some aspects of the bli and kto phenotypes resemble defects in Hh or N signaling. Loss of the response to Hh signaling, for instance, in clones lacking the Hh receptor Smoothened (Smo), prevents ato and dpp expression (Dominguez, 1999; Greenwood and Struhl, 1999; Strutt and Mlodzik, 1997); although these responses are diminished in bli and kto mutants they also persist posterior to the morphogenetic furrow, where cells are normally unresponsive to Hh. The basis for this Hh insensitivity is unknown; it could directly require the functions of bli and kto, or it could be a downstream effect of differentiation, which does not occur in bli or kto mutants. Supporting the second model, cells mutant for N also fail to differentiate and continue to express ato in all cells posterior to its normal expression domain (Baker et al., 1996; Baker and Yu, 1997). Like bli or kto mutant cells, they also maintain high levels of full-length Ci (Baker and Yu, 1997). This may indicate that the Hh pathway remains active; however, it has also been suggested that Hh acts at short range to down-
10 612 J. E. Treisman regulate Ci and terminate ato expression (Dominguez, 1999). The ability of the Hh signal to pass through large bli or kto clones and trigger anterior cells to differentiate suggests that the mutant cells may be able to express hh and relay the signal. The Hh and N signaling pathways are both required for normal growth of cells in the eye disc (Cho and Choi, 1998; Dominguez and de Celis, 1998; Heberlein et al., 1995; Kurata et al., 2000; Ma et al., 1993; Pan and Rubin, 1995; Papayannopoulos et al., 1998; Strutt et al., 1995); mutations in bli or kto appear to have no effect on growth, arguing that they do not mediate all the functions of Hh or N. We were also unable to induce photoreceptor differentiation in bli or kto mutant cells by activating the Hh or N pathways. However, this cannot be taken as definitive evidence that TRAP240 and TRAP230 do not influence signaling through these pathways; if they act as transcriptional coactivators, they may be required for the activated forms of Ci and N that we used to function effectively. The bli and kto mutant phenotypes also share some features with misregulation of other signaling pathways. Loss of the response to Dpp signaling, like loss of bli or kto, prevents expression of h and leads to prolonged expression of ey (Curtiss and Mlodzik, 2000; Greenwood and Struhl, 1999). However, the phenotypes differ in that cells mutant for components of the Dpp pathway, but not for bli or kto, can differentiate normally if they are not at the posterior margin (Burke and Basler, 1996; Wiersdorff et al., 1996), and loss of Dpp signaling interferes with cell growth and survival (Burke and Basler, 1996). Ectopic activation of the Wg pathway blocks photoreceptor differentiation, but it also promotes proliferation and does not cause persistent dpp expression (Theisen et al., 1996; Treisman and Rubin, 1995). EGF receptor signaling is unlikely to be blocked by bli or kto mutations, as it is strongly required for cell proliferation and survival, but not for the refinement of Ato into single cells or the expression of Elav in those cells (Dominguez et al., 1998; Kumar et al., 1998; Lesokhin et al., 1999). It is possible that bli and kto could antagonize the function of the homeodomain protein Homothorax (Hth), which has been shown to be capable of blocking photoreceptor formation downstream of dpp expression (Pai et al., 1998; Pichaud and Casares, 2000). However, hth is normally restricted to the anterior margin and was not ectopically expressed in bli or kto clones (data not shown). The presence of small numbers of photoreceptors in clones mutant for null alleles of bli or kto shows that these genes are not absolutely required for photoreceptors to differentiate. As photoreceptors form even in clones that occupy the whole eye disc, their presence cannot be due to rescue by a signal from surrounding wild-type cells. Instead it seems that the process requiring bli and kto function can be autonomously bypassed in a small number of cells. Other developmental functions of bli and kto In the antennal disc, Dll expression is lost in all cells of bli or kto mutant clones, but ey misexpression is observed only in a subset of mutant cells; we do not know whether this is due to rescue by surrounding wild-type cells. Dll is thought to be activated in response to the combined signaling activity of Dpp and Wg (Lecuit and Cohen, 1997) and to be repressed by Iroquois complex genes in the dorsal eye disc (Cavodeassi et al., 2000). It is not clear which of these regulators might be affected by bli and kto. It is also not known what normally prevents ey expression in the antennal disc, except that this block can be overcome by the ectopic expression of toy, eya, dac, optix or tsh (Bonini et al., 1997; Chen et al., 1997; Czerny et al., 1999; Pan and Rubin, 1998; Pignoni et al., 1997; Seimiya and Gehring, 2000; Shen and Mardon, 1997). bli or kto mutant clones in the wing disc did not misexpress ey (data not shown), suggesting that its activation requires factors specific to the antennal disc as well as the absence of bli and kto. One possibility is that normal ey expression in the embryonic antennal disc (Halder et al., 1998) is inappropriately maintained in bli or kto mutant cells, which could then be considered arrested at an early stage of their development like cells in the eye disc. A further interesting feature of bli and kto mutant clones is their apparent alteration in cell affinity, shown by the smooth borders formed with surrounding cells. Such borders can form as a result of changes in proximal-distal (P-D) identity in the leg disc (Wu and Cohen, 1999), although the same genes do not appear to control this property in the antennal disc (Casares and Mann, 1998; Dong et al., 2000). It is not clear whether altered P-D identity is the basis for the failure of bli or kto mutant cells to mix normally, as we have not found any marker of P-D identity to be misexpressed in these cells (data not shown). While this paper was under review, Boube et al. (Boube et al., 2000) also described mutations in the Drosophila homologue of TRAP240, using the name poils aux pattes (pap). They found that a deletion of the first 23 amino acids of TRAP240 caused a distal to proximal transformation of mutant cells in the tarsus of the first leg, but it did not affect the development of the antenna or wing. In contrast, all the alleles we have analyzed cause specific cell fate changes in the antennal and wing discs leading to pronounced malformations in the adult (Fig. 4 and data not shown). The reason for this difference is unclear. The lack of previously described mutations with the same phenotypes as bli and kto suggests several possible models for their function. TRAP240 and TRAP230 could be required to link a novel transcription factor to the TRAP coactivation complex. They could be required for some, but not all, of the functions of one or several known transcription factors acting in the pathways discussed above. Or they could themselves be transcription factors that were copurified with the TRAP complex by virtue of their strong interactions with it. In this case, their ubiquitous expression would suggest that their activity is modified in some way to produce specific patterns of expression of their target genes. Further biochemical and genetic analysis of the functions of the TRAP proteins will be required to distinguish these alternatives. We thank Sean Carroll, Barry Dickson, Manfred Frasch, Ulrike Heberlein, Robert Holmgren, Yuh-Nung Jan, Armen Manoukian, Graeme Mardon, Sarah Smolik, Uwe Walldorf, the Bloomington Drosophila stock center and the Berkeley Drosophila genome project for fly stocks and reagents. We are grateful to Florence Janody for in situ hybridization with kto and to Aude Benlali, Ian Oliver and Zara Martirosyan for technical assistance. We thank Ruth Lehmann and Michelle Starz-Gaiano for help with confocal microscopy. The manuscript was improved by the critical comments of Russ Collins, Florence Janody, Jeff Lee and Corinne Zaffran. This work was
decapentaplegic and wingless are regulated by eyes absent and eyegone and
 Development 125, 3741-3751 (1998) Printed in Great Britain The Company of Biologists Limited 1998 DEV8522 3741 decapentaplegic and wingless are regulated by eyes absent and eyegone and interact to direct
Development 125, 3741-3751 (1998) Printed in Great Britain The Company of Biologists Limited 1998 DEV8522 3741 decapentaplegic and wingless are regulated by eyes absent and eyegone and interact to direct
Control of Drosophila eye specification by Wingless signalling
 Development 129, 5313-5322 2002 The Company of Biologists Ltd doi:10.1242/dev.00096 5313 Control of Drosophila eye specification by Wingless signalling Antonio Baonza and Matthew Freeman* MRC Laboratory
Development 129, 5313-5322 2002 The Company of Biologists Ltd doi:10.1242/dev.00096 5313 Control of Drosophila eye specification by Wingless signalling Antonio Baonza and Matthew Freeman* MRC Laboratory
Dorsal eye selector pannier (pnr) suppresses the eye fate to define dorsal margin of the Drosophila eye
 Accepted Manuscript Dorsal eye selector pannier (pnr) suppresses the eye fate to define dorsal margin of the Drosophila eye Sarah M. Oros, Meghana Tare, Madhuri Kango-Singh, Amit Singh PII: S0012-1606(10)00975-9
Accepted Manuscript Dorsal eye selector pannier (pnr) suppresses the eye fate to define dorsal margin of the Drosophila eye Sarah M. Oros, Meghana Tare, Madhuri Kango-Singh, Amit Singh PII: S0012-1606(10)00975-9
Developmental Biology
 Developmental Biology xxx (2010) xxx xxx YDBIO-04944; No. of pages: 14; 4C: 3, 4, 6, 7, 8, 9, 10, 11 Contents lists available at ScienceDirect Developmental Biology journal homepage: www.elsevier.com/developmentalbiology
Developmental Biology xxx (2010) xxx xxx YDBIO-04944; No. of pages: 14; 4C: 3, 4, 6, 7, 8, 9, 10, 11 Contents lists available at ScienceDirect Developmental Biology journal homepage: www.elsevier.com/developmentalbiology
Developmental Biology
 Developmental Biology 346 (2010) 258 271 Contents lists available at ScienceDirect Developmental Biology journal homepage: www.elsevier.com/developmentalbiology Dorsal eye selector pannier (pnr) suppresses
Developmental Biology 346 (2010) 258 271 Contents lists available at ScienceDirect Developmental Biology journal homepage: www.elsevier.com/developmentalbiology Dorsal eye selector pannier (pnr) suppresses
Midterm 1. Average score: 74.4 Median score: 77
 Midterm 1 Average score: 74.4 Median score: 77 NAME: TA (circle one) Jody Westbrook or Jessica Piel Section (circle one) Tue Wed Thur MCB 141 First Midterm Feb. 21, 2008 Only answer 4 of these 5 problems.
Midterm 1 Average score: 74.4 Median score: 77 NAME: TA (circle one) Jody Westbrook or Jessica Piel Section (circle one) Tue Wed Thur MCB 141 First Midterm Feb. 21, 2008 Only answer 4 of these 5 problems.
purpose of this Chapter is to highlight some problems that will likely provide new
 119 Chapter 6 Future Directions Besides our contributions discussed in previous chapters to the problem of developmental pattern formation, this work has also brought new questions that remain unanswered.
119 Chapter 6 Future Directions Besides our contributions discussed in previous chapters to the problem of developmental pattern formation, this work has also brought new questions that remain unanswered.
Two subunits of the Drosophila mediator complex act together to control cell affinity
 Development 130, 3691-3701 2003 The Company of Biologists Ltd doi:10.1242/dev.00607 3691 Two subunits of the Drosophila mediator complex act together to control cell affinity Florence Janody, Zara Martirosyan,
Development 130, 3691-3701 2003 The Company of Biologists Ltd doi:10.1242/dev.00607 3691 Two subunits of the Drosophila mediator complex act together to control cell affinity Florence Janody, Zara Martirosyan,
Chapter 4 Evaluating a potential interaction between deltex and git in Drosophila: genetic interaction, gene overexpression and cell biology assays.
 Evaluating a potential interaction between deltex and git in Drosophila: genetic interaction, gene overexpression and cell biology assays. The data described in chapter 3 presented evidence that endogenous
Evaluating a potential interaction between deltex and git in Drosophila: genetic interaction, gene overexpression and cell biology assays. The data described in chapter 3 presented evidence that endogenous
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/301/ra98/dc1 Supplementary Materials for Regulation of Epithelial Morphogenesis by the G Protein Coupled Receptor Mist and Its Ligand Fog Alyssa J. Manning,
www.sciencesignaling.org/cgi/content/full/6/301/ra98/dc1 Supplementary Materials for Regulation of Epithelial Morphogenesis by the G Protein Coupled Receptor Mist and Its Ligand Fog Alyssa J. Manning,
Why Flies? stages of embryogenesis. The Fly in History
 The Fly in History 1859 Darwin 1866 Mendel c. 1890 Driesch, Roux (experimental embryology) 1900 rediscovery of Mendel (birth of genetics) 1910 first mutant (white) (Morgan) 1913 first genetic map (Sturtevant
The Fly in History 1859 Darwin 1866 Mendel c. 1890 Driesch, Roux (experimental embryology) 1900 rediscovery of Mendel (birth of genetics) 1910 first mutant (white) (Morgan) 1913 first genetic map (Sturtevant
Segment boundary formation in Drosophila embryos
 Segment boundary formation in Drosophila embryos Development 130, August 2003 Camilla W. Larsen, Elizabeth Hirst, Cyrille Alexandre and Jean Paul Vincent 1. Introduction: - Segment boundary formation:
Segment boundary formation in Drosophila embryos Development 130, August 2003 Camilla W. Larsen, Elizabeth Hirst, Cyrille Alexandre and Jean Paul Vincent 1. Introduction: - Segment boundary formation:
Drosophila wing. Temporal regulation of Apterous activity during development of the. Marco Milán and Stephen M. Cohen* SUMMARY
 Development 127, 3069-3078 (2000) Printed in Great Britain The Company of Biologists Limited 2000 DEV2547 3069 Temporal regulation of Apterous activity during development of the Drosophila wing Marco Milán
Development 127, 3069-3078 (2000) Printed in Great Britain The Company of Biologists Limited 2000 DEV2547 3069 Temporal regulation of Apterous activity during development of the Drosophila wing Marco Milán
mirror controls planar polarity and equator formation through repression of fringe expression and through control of cell affinities
 Development 126, 5857-5866 (1999) Printed in Great Britain The Company of Biologists Limited 1999 DEV5359 5857 mirror controls planar polarity and equator formation through repression of fringe expression
Development 126, 5857-5866 (1999) Printed in Great Britain The Company of Biologists Limited 1999 DEV5359 5857 mirror controls planar polarity and equator formation through repression of fringe expression
IMAGINAL discs of Drosophila are sac-like epithelial
 Copyright Ó 2005 by the Genetics Society of America DOI: 10.1534/genetics.105.044180 Genetic Interaction of Lobe With Its Modifiers in Dorsoventral Patterning and Growth of the Drosophila Eye Amit Singh,*
Copyright Ó 2005 by the Genetics Society of America DOI: 10.1534/genetics.105.044180 Genetic Interaction of Lobe With Its Modifiers in Dorsoventral Patterning and Growth of the Drosophila Eye Amit Singh,*
RNA Synthesis and Processing
 RNA Synthesis and Processing Introduction Regulation of gene expression allows cells to adapt to environmental changes and is responsible for the distinct activities of the differentiated cell types that
RNA Synthesis and Processing Introduction Regulation of gene expression allows cells to adapt to environmental changes and is responsible for the distinct activities of the differentiated cell types that
Lecture 7. Development of the Fruit Fly Drosophila
 BIOLOGY 205/SECTION 7 DEVELOPMENT- LILJEGREN Lecture 7 Development of the Fruit Fly Drosophila 1. The fruit fly- a highly successful, specialized organism a. Quick life cycle includes three larval stages
BIOLOGY 205/SECTION 7 DEVELOPMENT- LILJEGREN Lecture 7 Development of the Fruit Fly Drosophila 1. The fruit fly- a highly successful, specialized organism a. Quick life cycle includes three larval stages
The Pax and large Maf families of genes in mammalian eye development
 The Pax and large Maf families of genes in mammalian eye development Vertebrate eye development is dependent on the coordinated action of thousands of genes. A specific group of over one hundred of regulatory
The Pax and large Maf families of genes in mammalian eye development Vertebrate eye development is dependent on the coordinated action of thousands of genes. A specific group of over one hundred of regulatory
Chapter 18 Lecture. Concepts of Genetics. Tenth Edition. Developmental Genetics
 Chapter 18 Lecture Concepts of Genetics Tenth Edition Developmental Genetics Chapter Contents 18.1 Differentiated States Develop from Coordinated Programs of Gene Expression 18.2 Evolutionary Conservation
Chapter 18 Lecture Concepts of Genetics Tenth Edition Developmental Genetics Chapter Contents 18.1 Differentiated States Develop from Coordinated Programs of Gene Expression 18.2 Evolutionary Conservation
TGF-β/BMP superfamily members, Gbb-60A and Dpp, cooperate to provide pattern information and establish cell identity in the Drosophila wing
 Development 125, 2723-2734 (1998) Printed in Great Britain The Company of Biologists Limited 1998 DEV5189 2723 TGF-β/BMP superfamily members, Gbb-60A and Dpp, cooperate to provide pattern information and
Development 125, 2723-2734 (1998) Printed in Great Britain The Company of Biologists Limited 1998 DEV5189 2723 TGF-β/BMP superfamily members, Gbb-60A and Dpp, cooperate to provide pattern information and
A complementation test would be done by crossing the haploid strains and scoring the phenotype in the diploids.
 Problem set H answers 1. To study DNA repair mechanisms, geneticists isolated yeast mutants that were sensitive to various types of radiation; for example, mutants that were more sensitive to UV light.
Problem set H answers 1. To study DNA repair mechanisms, geneticists isolated yeast mutants that were sensitive to various types of radiation; for example, mutants that were more sensitive to UV light.
BIS &003 Answers to Assigned Problems May 23, Week /18.6 How would you distinguish between an enhancer and a promoter?
 Week 9 Study Questions from the textbook: 6 th Edition: Chapter 19-19.6, 19.7, 19.15, 19.17 OR 7 th Edition: Chapter 18-18.6 18.7, 18.15, 18.17 19.6/18.6 How would you distinguish between an enhancer and
Week 9 Study Questions from the textbook: 6 th Edition: Chapter 19-19.6, 19.7, 19.15, 19.17 OR 7 th Edition: Chapter 18-18.6 18.7, 18.15, 18.17 19.6/18.6 How would you distinguish between an enhancer and
Evolution of Transcription factor function: Homeotic (Hox) proteins
 Evolution of Transcription factor function: Homeotic (Hox) proteins Hox proteins regulate morphology in cellular zones on the anterior-posterior axis of embryos via the activation/repression of unknown
Evolution of Transcription factor function: Homeotic (Hox) proteins Hox proteins regulate morphology in cellular zones on the anterior-posterior axis of embryos via the activation/repression of unknown
A Glimpse Into Dorso-Ventral Patterning of the Drosophila Eye
 a DEVELOPMENTAL DYNAMICS 241:69 84, 2012 SPECIAL ISSUE REVIEWS-A PEER REVIEWED FORUM A Glimpse Into Dorso-Ventral Patterning of the Drosophila Eye Amit Singh, 1,2,3 * Meghana Tare, 1 Oorvashi Roy Puli,
a DEVELOPMENTAL DYNAMICS 241:69 84, 2012 SPECIAL ISSUE REVIEWS-A PEER REVIEWED FORUM A Glimpse Into Dorso-Ventral Patterning of the Drosophila Eye Amit Singh, 1,2,3 * Meghana Tare, 1 Oorvashi Roy Puli,
Bypass and interaction suppressors; pathway analysis
 Bypass and interaction suppressors; pathway analysis The isolation of extragenic suppressors is a powerful tool for identifying genes that encode proteins that function in the same process as a gene of
Bypass and interaction suppressors; pathway analysis The isolation of extragenic suppressors is a powerful tool for identifying genes that encode proteins that function in the same process as a gene of
JAK/STAT signaling promotes regional specification by negatively regulating wingless expression in Drosophila
 Development Access Development Advance the most recent Online epress version Articles. at online First http://dev.biologists.org/lookup/doi/10.1242/dev.02675 posted publication online on date 1 November
Development Access Development Advance the most recent Online epress version Articles. at online First http://dev.biologists.org/lookup/doi/10.1242/dev.02675 posted publication online on date 1 November
Suppression of the rbf null mutants by a de2f1 allele that lacks transactivation domain
 Development 127, 367-379 (2000) Printed in Great Britain The Company of Biologists Limited 2000 DEV5342 367 Suppression of the rbf null mutants by a de2f1 allele that lacks transactivation domain Wei Du
Development 127, 367-379 (2000) Printed in Great Britain The Company of Biologists Limited 2000 DEV5342 367 Suppression of the rbf null mutants by a de2f1 allele that lacks transactivation domain Wei Du
Reading: Chapter 5, pp ; Reference chapter D, pp Problem set F
 Mosaic Analysis Reading: Chapter 5, pp140-141; Reference chapter D, pp820-823 Problem set F Twin spots in Drosophila Although segregation and recombination in mitosis do not occur at the same frequency
Mosaic Analysis Reading: Chapter 5, pp140-141; Reference chapter D, pp820-823 Problem set F Twin spots in Drosophila Although segregation and recombination in mitosis do not occur at the same frequency
Axis Specification in Drosophila
 Developmental Biology Biology 4361 Axis Specification in Drosophila November 2, 2006 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
Developmental Biology Biology 4361 Axis Specification in Drosophila November 2, 2006 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
Signals transmitted along retinal axons in Drosophila: Hedgehog signal reception and the cell circuitry of lamina cartridge assembly
 Development 125, 3753-3764 (1998) Printed in Great Britain The Company of Biologists Limited 1998 DEV9614 3753 Signals transmitted along retinal axons in Drosophila: Hedgehog signal reception and the cell
Development 125, 3753-3764 (1998) Printed in Great Britain The Company of Biologists Limited 1998 DEV9614 3753 Signals transmitted along retinal axons in Drosophila: Hedgehog signal reception and the cell
Welcome to Class 21!
 Welcome to Class 21! Introductory Biochemistry! Lecture 21: Outline and Objectives l Regulation of Gene Expression in Prokaryotes! l transcriptional regulation! l principles! l lac operon! l trp attenuation!
Welcome to Class 21! Introductory Biochemistry! Lecture 21: Outline and Objectives l Regulation of Gene Expression in Prokaryotes! l transcriptional regulation! l principles! l lac operon! l trp attenuation!
Lecture 18 June 2 nd, Gene Expression Regulation Mutations
 Lecture 18 June 2 nd, 2016 Gene Expression Regulation Mutations From Gene to Protein Central Dogma Replication DNA RNA PROTEIN Transcription Translation RNA Viruses: genome is RNA Reverse Transcriptase
Lecture 18 June 2 nd, 2016 Gene Expression Regulation Mutations From Gene to Protein Central Dogma Replication DNA RNA PROTEIN Transcription Translation RNA Viruses: genome is RNA Reverse Transcriptase
Introduction. Gene expression is the combined process of :
 1 To know and explain: Regulation of Bacterial Gene Expression Constitutive ( house keeping) vs. Controllable genes OPERON structure and its role in gene regulation Regulation of Eukaryotic Gene Expression
1 To know and explain: Regulation of Bacterial Gene Expression Constitutive ( house keeping) vs. Controllable genes OPERON structure and its role in gene regulation Regulation of Eukaryotic Gene Expression
Eukaryotic Gene Expression
 Eukaryotic Gene Expression Lectures 22-23 Several Features Distinguish Eukaryotic Processes From Mechanisms in Bacteria 123 Eukaryotic Gene Expression Several Features Distinguish Eukaryotic Processes
Eukaryotic Gene Expression Lectures 22-23 Several Features Distinguish Eukaryotic Processes From Mechanisms in Bacteria 123 Eukaryotic Gene Expression Several Features Distinguish Eukaryotic Processes
Dissecting the roles of the Drosophila EGF receptor in eye development and MAP kinase activation
 Development 125, 3875-3885 (1998) Printed in Great Britain The Company of Biologists Limited 1998 DEV5225 3875 Dissecting the roles of the Drosophila EGF receptor in eye development and MAP kinase activation
Development 125, 3875-3885 (1998) Printed in Great Britain The Company of Biologists Limited 1998 DEV5225 3875 Dissecting the roles of the Drosophila EGF receptor in eye development and MAP kinase activation
Signaling circuitries in development: insights from the retinal determination gene network
 3 Signaling circuitries in development: insights from the retinal determination gene network Serena J. Silver 1,2, * and Ilaria Rebay 1,2, 1 Whitehead Institute for Biomedical Research, Cambridge, MA 02142,
3 Signaling circuitries in development: insights from the retinal determination gene network Serena J. Silver 1,2, * and Ilaria Rebay 1,2, 1 Whitehead Institute for Biomedical Research, Cambridge, MA 02142,
Illegitimate translation causes unexpected gene expression from on-target out-of-frame alleles
 Illegitimate translation causes unexpected gene expression from on-target out-of-frame alleles created by CRISPR-Cas9 Shigeru Makino, Ryutaro Fukumura, Yoichi Gondo* Mutagenesis and Genomics Team, RIKEN
Illegitimate translation causes unexpected gene expression from on-target out-of-frame alleles created by CRISPR-Cas9 Shigeru Makino, Ryutaro Fukumura, Yoichi Gondo* Mutagenesis and Genomics Team, RIKEN
THE Hedgehog (Hh) signaling pathway regulates
 Copyright Ó 2006 by the Genetics Society of America DOI: 10.1534/genetics.106.061036 The Contributions of Protein Kinase A and Smoothened Phosphorylation to Hedgehog Signal Transduction in Drosophila melanogaster
Copyright Ó 2006 by the Genetics Society of America DOI: 10.1534/genetics.106.061036 The Contributions of Protein Kinase A and Smoothened Phosphorylation to Hedgehog Signal Transduction in Drosophila melanogaster
Axis determination in flies. Sem 9.3.B.5 Animal Science
 Axis determination in flies Sem 9.3.B.5 Animal Science All embryos are in lateral view (anterior to the left). Endoderm, midgut; mesoderm; central nervous system; foregut, hindgut and pole cells in yellow.
Axis determination in flies Sem 9.3.B.5 Animal Science All embryos are in lateral view (anterior to the left). Endoderm, midgut; mesoderm; central nervous system; foregut, hindgut and pole cells in yellow.
The JAK/STAT Pathway Regulates Proximo- Distal Patterning in Drosophila
 DEVELOPMENTAL DYNAMICS 236:2721 2730, 2007 RESEARCH ARTICLE The JAK/STAT Pathway Regulates Proximo- Distal Patterning in Drosophila Aidee Ayala-Camargo, 1 Laura A. Ekas, 1 Maria Sol Flaherty, 1 Gyeong-Hun
DEVELOPMENTAL DYNAMICS 236:2721 2730, 2007 RESEARCH ARTICLE The JAK/STAT Pathway Regulates Proximo- Distal Patterning in Drosophila Aidee Ayala-Camargo, 1 Laura A. Ekas, 1 Maria Sol Flaherty, 1 Gyeong-Hun
Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Tuesday, December 27, 16
 Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Enduring understanding 3.B: Expression of genetic information involves cellular and molecular
Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Enduring understanding 3.B: Expression of genetic information involves cellular and molecular
SIGNALLING PATHWAYS IN DROSOPHILA AND VERTEBRATE RETINAL DEVELOPMENT
 SIGNALLING PATHWAYS IN DROSOPHILA AND VERTEBRATE RETINAL DEVELOPMENT Justin P. Kumar The near-catholic conservation of paired box gene 6 (Pax6) and its supporting cast of retinal determination genes throughout
SIGNALLING PATHWAYS IN DROSOPHILA AND VERTEBRATE RETINAL DEVELOPMENT Justin P. Kumar The near-catholic conservation of paired box gene 6 (Pax6) and its supporting cast of retinal determination genes throughout
Homeotic genes in flies. Sem 9.3.B.6 Animal Science
 Homeotic genes in flies Sem 9.3.B.6 Animal Science So far We have seen that identities of each segment is determined by various regulators of segment polarity genes In arthopods, and in flies, each segment
Homeotic genes in flies Sem 9.3.B.6 Animal Science So far We have seen that identities of each segment is determined by various regulators of segment polarity genes In arthopods, and in flies, each segment
The knirps and knirps-related genes organize development of the second wing vein in Drosophila
 Development 125, 4145-4154 (1998) Printed in Great Britain The Company of Biologists Limited 1998 DEV3896 4145 The knirps and knirps-related genes organize development of the second wing vein in Drosophila
Development 125, 4145-4154 (1998) Printed in Great Britain The Company of Biologists Limited 1998 DEV3896 4145 The knirps and knirps-related genes organize development of the second wing vein in Drosophila
Development of Drosophila
 Development of Drosophila Hand-out CBT Chapter 2 Wolpert, 5 th edition March 2018 Introduction 6. Introduction Drosophila melanogaster, the fruit fly, is found in all warm countries. In cooler regions,
Development of Drosophila Hand-out CBT Chapter 2 Wolpert, 5 th edition March 2018 Introduction 6. Introduction Drosophila melanogaster, the fruit fly, is found in all warm countries. In cooler regions,
Developmental genetics: finding the genes that regulate development
 Developmental Biology BY1101 P. Murphy Lecture 9 Developmental genetics: finding the genes that regulate development Introduction The application of genetic analysis and DNA technology to the study of
Developmental Biology BY1101 P. Murphy Lecture 9 Developmental genetics: finding the genes that regulate development Introduction The application of genetic analysis and DNA technology to the study of
Evolution of the Complex Eye and Pax6 Gene
 Evolution of the Complex Eye and Pax6 Gene Rachel Thomsen Sarah Kim Jenia Ostrovskaya Key Points Definition of an Eye o Types of Eyes Origin of Species: Difficulties on Theory Pax family Pax6 gene why
Evolution of the Complex Eye and Pax6 Gene Rachel Thomsen Sarah Kim Jenia Ostrovskaya Key Points Definition of an Eye o Types of Eyes Origin of Species: Difficulties on Theory Pax family Pax6 gene why
MCB 141 Midterm I Feb. 19, 2009
 Write your name and student ID# on EVERY PAGE of your exam MCB 141 Midterm I Feb. 19, 2009 Circle the name of your TA Jessica Lyons Alberto Stolfi Question #1 Question #2 Question #3 Question #4 TOTAL
Write your name and student ID# on EVERY PAGE of your exam MCB 141 Midterm I Feb. 19, 2009 Circle the name of your TA Jessica Lyons Alberto Stolfi Question #1 Question #2 Question #3 Question #4 TOTAL
Axis Specification in Drosophila
 Developmental Biology Biology 4361 Axis Specification in Drosophila November 6, 2007 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
Developmental Biology Biology 4361 Axis Specification in Drosophila November 6, 2007 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
Drosophila. Cubitus interruptus-independent transduction of the Hedgehog signal in
 Development 127, 5509-5522 (2000) Printed in Great Britain The Company of Biologists Limited 2000 DEV7818 5509 Cubitus interruptus-independent transduction of the Hedgehog signal in Drosophila Armel Gallet
Development 127, 5509-5522 (2000) Printed in Great Britain The Company of Biologists Limited 2000 DEV7818 5509 Cubitus interruptus-independent transduction of the Hedgehog signal in Drosophila Armel Gallet
Supplementary Information
 Supplementary Information Supplementary Figure 1. JAK/STAT in early wing development (a-f) Wing primordia of second instar larvae of the indicated genotypes labeled to visualize expression of upd mrna
Supplementary Information Supplementary Figure 1. JAK/STAT in early wing development (a-f) Wing primordia of second instar larvae of the indicated genotypes labeled to visualize expression of upd mrna
orthodenticle homeobox gene
 Development 121, 3561-3572 (1995) Printed in Great Britain The Company of Biologists Limited 1995 DEV8275 3561 Pattern formation in Drosophila head development: the role of the orthodenticle homeobox gene
Development 121, 3561-3572 (1995) Printed in Great Britain The Company of Biologists Limited 1995 DEV8275 3561 Pattern formation in Drosophila head development: the role of the orthodenticle homeobox gene
THE near-perfect structure of the Drosophila melanogaster
 Copyright Ó 2005 by the Genetics Society of America DOI: 10.1534/genetics.105.045450 A Genetic Screen Identifies Putative Targets and Binding Partners of CREB-Binding Protein in the Developing Drosophila
Copyright Ó 2005 by the Genetics Society of America DOI: 10.1534/genetics.105.045450 A Genetic Screen Identifies Putative Targets and Binding Partners of CREB-Binding Protein in the Developing Drosophila
Three Distinct Roles for Notch in Drosophila R7 Photoreceptor Specification
 Three Distinct Roles for Notch in Drosophila R7 Photoreceptor Specification Andrew Tomlinson 1. *, Yannis Emmanuel Mavromatakis 1., Gary Struhl 1,2 1 Department of Genetics and Development, College of
Three Distinct Roles for Notch in Drosophila R7 Photoreceptor Specification Andrew Tomlinson 1. *, Yannis Emmanuel Mavromatakis 1., Gary Struhl 1,2 1 Department of Genetics and Development, College of
Drosophila limb development
 Drosophila limb development Principal Investigator Marco Milán (ICREA) Posdoctoral Fellows Isabel Becam Fernando Bejarano Héctor Herranz Carlos Luque PhD Students Duarte Mesquita Neus Rafel Georgina Sorrosal
Drosophila limb development Principal Investigator Marco Milán (ICREA) Posdoctoral Fellows Isabel Becam Fernando Bejarano Héctor Herranz Carlos Luque PhD Students Duarte Mesquita Neus Rafel Georgina Sorrosal
Functional and regulatory interactions between Hox and extradenticle genes
 Functional and regulatory interactions between Hox and extradenticle genes Natalia Azpiazu and Ginés Morata 1 Centro de Biologia Molecular Centro Superior de Investigaciones Cientificas-Universidad Autońoma
Functional and regulatory interactions between Hox and extradenticle genes Natalia Azpiazu and Ginés Morata 1 Centro de Biologia Molecular Centro Superior de Investigaciones Cientificas-Universidad Autońoma
Evolution of Insect Eye Development: First Insights from Fruit Fly, Grasshopper and Flour Beetle 1
 INTEGR. COMP. BIOL., 43:508 521 (2003) Evolution of Insect Eye Development: First Insights from Fruit Fly, Grasshopper and Flour Beetle 1 MARKUS FRIEDRICH 2 Department of Biological Sciences, Wayne State
INTEGR. COMP. BIOL., 43:508 521 (2003) Evolution of Insect Eye Development: First Insights from Fruit Fly, Grasshopper and Flour Beetle 1 MARKUS FRIEDRICH 2 Department of Biological Sciences, Wayne State
The function of engrailed and the specification of Drosophila wing pattern
 Development 121, 3447-3456 (1995) Printed in Great Britain The Company of Biologists Limited 1995 3447 The function of engrailed and the specification of Drosophila wing pattern Isabel Guillén 1, *, Jose
Development 121, 3447-3456 (1995) Printed in Great Britain The Company of Biologists Limited 1995 3447 The function of engrailed and the specification of Drosophila wing pattern Isabel Guillén 1, *, Jose
REVIEW SESSION. Wednesday, September 15 5:30 PM SHANTZ 242 E
 REVIEW SESSION Wednesday, September 15 5:30 PM SHANTZ 242 E Gene Regulation Gene Regulation Gene expression can be turned on, turned off, turned up or turned down! For example, as test time approaches,
REVIEW SESSION Wednesday, September 15 5:30 PM SHANTZ 242 E Gene Regulation Gene Regulation Gene expression can be turned on, turned off, turned up or turned down! For example, as test time approaches,
Peter Pristas. Gene regulation in eukaryotes
 Peter Pristas BNK1 Gene regulation in eukaryotes Gene Expression in Eukaryotes Only about 3-5% of all the genes in a human cell are expressed at any given time. The genes expressed can be specific for
Peter Pristas BNK1 Gene regulation in eukaryotes Gene Expression in Eukaryotes Only about 3-5% of all the genes in a human cell are expressed at any given time. The genes expressed can be specific for
CHAPTER 13 PROKARYOTE GENES: E. COLI LAC OPERON
 PROKARYOTE GENES: E. COLI LAC OPERON CHAPTER 13 CHAPTER 13 PROKARYOTE GENES: E. COLI LAC OPERON Figure 1. Electron micrograph of growing E. coli. Some show the constriction at the location where daughter
PROKARYOTE GENES: E. COLI LAC OPERON CHAPTER 13 CHAPTER 13 PROKARYOTE GENES: E. COLI LAC OPERON Figure 1. Electron micrograph of growing E. coli. Some show the constriction at the location where daughter
Compartments and the control of growth in the Drosophila wing imaginal disc
 Access Development the most First recent posted version epress online at online on http://dev.biologists.org/lookup/doi/10.1242/dev.02618 11 October publication 2006 as date 10.1242/dev.02618 11 October
Access Development the most First recent posted version epress online at online on http://dev.biologists.org/lookup/doi/10.1242/dev.02618 11 October publication 2006 as date 10.1242/dev.02618 11 October
A re-evaluation of the contributions of Apterous and Notch to the dorsoventral lineage restriction boundary in the Drosophila wing
 Development 130, 553-562 2003 The Company of Biologists Ltd doi:10.1242/dev.00276 553 A re-evaluation of the contributions of Apterous and Notch to the dorsoventral lineage restriction boundary in the
Development 130, 553-562 2003 The Company of Biologists Ltd doi:10.1242/dev.00276 553 A re-evaluation of the contributions of Apterous and Notch to the dorsoventral lineage restriction boundary in the
16 CONTROL OF GENE EXPRESSION
 16 CONTROL OF GENE EXPRESSION Chapter Outline 16.1 REGULATION OF GENE EXPRESSION IN PROKARYOTES The operon is the unit of transcription in prokaryotes The lac operon for lactose metabolism is transcribed
16 CONTROL OF GENE EXPRESSION Chapter Outline 16.1 REGULATION OF GENE EXPRESSION IN PROKARYOTES The operon is the unit of transcription in prokaryotes The lac operon for lactose metabolism is transcribed
Genetics 275 Notes Week 7
 Cytoplasmic Inheritance Genetics 275 Notes Week 7 Criteriafor recognition of cytoplasmic inheritance: 1. Reciprocal crosses give different results -mainly due to the fact that the female parent contributes
Cytoplasmic Inheritance Genetics 275 Notes Week 7 Criteriafor recognition of cytoplasmic inheritance: 1. Reciprocal crosses give different results -mainly due to the fact that the female parent contributes
Regulation and signaling. Overview. Control of gene expression. Cells need to regulate the amounts of different proteins they express, depending on
 Regulation and signaling Overview Cells need to regulate the amounts of different proteins they express, depending on cell development (skin vs liver cell) cell stage environmental conditions (food, temperature,
Regulation and signaling Overview Cells need to regulate the amounts of different proteins they express, depending on cell development (skin vs liver cell) cell stage environmental conditions (food, temperature,
MOLECULAR CONTROL OF EMBRYONIC PATTERN FORMATION
 MOLECULAR CONTROL OF EMBRYONIC PATTERN FORMATION Drosophila is the best understood of all developmental systems, especially at the genetic level, and although it is an invertebrate it has had an enormous
MOLECULAR CONTROL OF EMBRYONIC PATTERN FORMATION Drosophila is the best understood of all developmental systems, especially at the genetic level, and although it is an invertebrate it has had an enormous
BILD7: Problem Set. 2. What did Chargaff discover and why was this important?
 BILD7: Problem Set 1. What is the general structure of DNA? 2. What did Chargaff discover and why was this important? 3. What was the major contribution of Rosalind Franklin? 4. How did solving the structure
BILD7: Problem Set 1. What is the general structure of DNA? 2. What did Chargaff discover and why was this important? 3. What was the major contribution of Rosalind Franklin? 4. How did solving the structure
10/2/2015. Chapter 4. Determination and Differentiation. Neuroanatomical Diversity
 Chapter 4 Determination and Differentiation Neuroanatomical Diversity 1 Neurochemical diversity: another important aspect of neuronal fate Neurotransmitters and their receptors Excitatory Glutamate Acetylcholine
Chapter 4 Determination and Differentiation Neuroanatomical Diversity 1 Neurochemical diversity: another important aspect of neuronal fate Neurotransmitters and their receptors Excitatory Glutamate Acetylcholine
Honors Biology Reading Guide Chapter 11
 Honors Biology Reading Guide Chapter 11 v Promoter a specific nucleotide sequence in DNA located near the start of a gene that is the binding site for RNA polymerase and the place where transcription begins
Honors Biology Reading Guide Chapter 11 v Promoter a specific nucleotide sequence in DNA located near the start of a gene that is the binding site for RNA polymerase and the place where transcription begins
Indiana University PubMed for Justin Kumar January 6, 2012
 1. Kumar JP. Building an ommatidium one cell at a time. Dev Dyn. 2012 Jan;241(1):136-49. doi: 10.1002/dvdy.23707. Department of Biology, Indiana University, Bloomington, Indiana. jkumar@indiana.edu. PMID:
1. Kumar JP. Building an ommatidium one cell at a time. Dev Dyn. 2012 Jan;241(1):136-49. doi: 10.1002/dvdy.23707. Department of Biology, Indiana University, Bloomington, Indiana. jkumar@indiana.edu. PMID:
Caenorhabditis elegans
 Caenorhabditis elegans Why C. elegans? Sea urchins have told us much about embryogenesis. They are suited well for study in the lab; however, they do not tell us much about the genetics involved in embryogenesis.
Caenorhabditis elegans Why C. elegans? Sea urchins have told us much about embryogenesis. They are suited well for study in the lab; however, they do not tell us much about the genetics involved in embryogenesis.
Chapter 18 Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression Differential gene expression Every somatic cell in an individual organism contains the same genetic information and replicated from the same original fertilized
Chapter 18 Regulation of Gene Expression Differential gene expression Every somatic cell in an individual organism contains the same genetic information and replicated from the same original fertilized
AP Biology Gene Regulation and Development Review
 AP Biology Gene Regulation and Development Review 1. What does the regulatory gene code for? 2. Is the repressor by default active/inactive? 3. What changes the repressor activity? 4. What does repressor
AP Biology Gene Regulation and Development Review 1. What does the regulatory gene code for? 2. Is the repressor by default active/inactive? 3. What changes the repressor activity? 4. What does repressor
Biology. Biology. Slide 1 of 26. End Show. Copyright Pearson Prentice Hall
 Biology Biology 1 of 26 Fruit fly chromosome 12-5 Gene Regulation Mouse chromosomes Fruit fly embryo Mouse embryo Adult fruit fly Adult mouse 2 of 26 Gene Regulation: An Example Gene Regulation: An Example
Biology Biology 1 of 26 Fruit fly chromosome 12-5 Gene Regulation Mouse chromosomes Fruit fly embryo Mouse embryo Adult fruit fly Adult mouse 2 of 26 Gene Regulation: An Example Gene Regulation: An Example
Gonadal mesoderm and fat body initially follow a common developmental path in Drosophila
 Development 125, 837-844 (1998) Printed in Great Britain The Company of Biologists Limited 1998 DEV7627 837 Gonadal and fat body initially follow a common developmental path in Drosophila Lisa A. Moore
Development 125, 837-844 (1998) Printed in Great Britain The Company of Biologists Limited 1998 DEV7627 837 Gonadal and fat body initially follow a common developmental path in Drosophila Lisa A. Moore
A conserved blueprint for the eye?
 A conserved blueprint for the eye? Jessica E. Treisman Summary Although the eyes of all organisms have a common function, visual perception, their structures and developmental mechanisms are quite diverse.
A conserved blueprint for the eye? Jessica E. Treisman Summary Although the eyes of all organisms have a common function, visual perception, their structures and developmental mechanisms are quite diverse.
Lesson Overview. Gene Regulation and Expression. Lesson Overview Gene Regulation and Expression
 13.4 Gene Regulation and Expression THINK ABOUT IT Think of a library filled with how-to books. Would you ever need to use all of those books at the same time? Of course not. Now picture a tiny bacterium
13.4 Gene Regulation and Expression THINK ABOUT IT Think of a library filled with how-to books. Would you ever need to use all of those books at the same time? Of course not. Now picture a tiny bacterium
MBios 401/501: Lecture 14.2 Cell Differentiation I. Slide #1. Cell Differentiation
 MBios 401/501: Lecture 14.2 Cell Differentiation I Slide #1 Cell Differentiation Cell Differentiation I -Basic principles of differentiation (p1305-1320) -C-elegans (p1321-1327) Cell Differentiation II
MBios 401/501: Lecture 14.2 Cell Differentiation I Slide #1 Cell Differentiation Cell Differentiation I -Basic principles of differentiation (p1305-1320) -C-elegans (p1321-1327) Cell Differentiation II
Chapter 11. Development: Differentiation and Determination
 KAP Biology Dept Kenyon College Differential gene expression and development Mechanisms of cellular determination Induction Pattern formation Chapter 11. Development: Differentiation and Determination
KAP Biology Dept Kenyon College Differential gene expression and development Mechanisms of cellular determination Induction Pattern formation Chapter 11. Development: Differentiation and Determination
Wan-Ju Liu 1,2, John S Reece-Hoyes 3, Albertha JM Walhout 3 and David M Eisenmann 1*
 Liu et al. BMC Developmental Biology 2014, 14:17 RESEARCH ARTICLE Open Access Multiple transcription factors directly regulate Hox gene lin-39 expression in ventral hypodermal cells of the C. elegans embryo
Liu et al. BMC Developmental Biology 2014, 14:17 RESEARCH ARTICLE Open Access Multiple transcription factors directly regulate Hox gene lin-39 expression in ventral hypodermal cells of the C. elegans embryo
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11589 Supplementary Figure 1 Ciona intestinalis and Petromyzon marinus neural crest expression domain comparison. Cartoon shows dorsal views of Ciona mid gastrula (left) and Petromyzon
doi:10.1038/nature11589 Supplementary Figure 1 Ciona intestinalis and Petromyzon marinus neural crest expression domain comparison. Cartoon shows dorsal views of Ciona mid gastrula (left) and Petromyzon
1. Draw, label and describe the structure of DNA and RNA including bonding mechanisms.
 Practicing Biology BIG IDEA 3.A 1. Draw, label and describe the structure of DNA and RNA including bonding mechanisms. 2. Using at least 2 well-known experiments, describe which features of DNA and RNA
Practicing Biology BIG IDEA 3.A 1. Draw, label and describe the structure of DNA and RNA including bonding mechanisms. 2. Using at least 2 well-known experiments, describe which features of DNA and RNA
Cell Cell Communication in Development
 Biology 4361 Developmental Biology Cell Cell Communication in Development June 25, 2008 Cell Cell Communication Concepts Cells in developing organisms develop in the context of their environment, including
Biology 4361 Developmental Biology Cell Cell Communication in Development June 25, 2008 Cell Cell Communication Concepts Cells in developing organisms develop in the context of their environment, including
SIGNIFICANCE OF EMBRYOLOGY
 This lecture will discuss the following topics : Definition of Embryology Significance of Embryology Old and New Frontiers Introduction to Molecular Regulation and Signaling Descriptive terms in Embryology
This lecture will discuss the following topics : Definition of Embryology Significance of Embryology Old and New Frontiers Introduction to Molecular Regulation and Signaling Descriptive terms in Embryology
Drosophila Somatic Anterior-Posterior Axis (A-P Axis) Formation
 Home Biol 4241 Luria-Delbruck 1943 Hershey-Chase 1952 Meselson-Stahl 1958 Garapin et al. 1978 McClintock 1953 King-Wilson 1975 Sanger et al. 1977 Rothberg et al. 2011 Jeffreys et al. 1985 Bacterial Genetics
Home Biol 4241 Luria-Delbruck 1943 Hershey-Chase 1952 Meselson-Stahl 1958 Garapin et al. 1978 McClintock 1953 King-Wilson 1975 Sanger et al. 1977 Rothberg et al. 2011 Jeffreys et al. 1985 Bacterial Genetics
Cell Death & Trophic Factors II. Steven McLoon Department of Neuroscience University of Minnesota
 Cell Death & Trophic Factors II Steven McLoon Department of Neuroscience University of Minnesota 1 Remember? Neurotrophins are cell survival factors that neurons get from their target cells! There is a
Cell Death & Trophic Factors II Steven McLoon Department of Neuroscience University of Minnesota 1 Remember? Neurotrophins are cell survival factors that neurons get from their target cells! There is a
Supplementary Figure 1. Markedly decreased numbers of marginal zone B cells in DOCK8 mutant mice Supplementary Figure 2.
 Supplementary Figure 1. Markedly decreased numbers of marginal zone B cells in DOCK8 mutant mice. Percentage of marginal zone B cells in the spleen of wild-type mice (+/+), mice homozygous for cpm or pri
Supplementary Figure 1. Markedly decreased numbers of marginal zone B cells in DOCK8 mutant mice. Percentage of marginal zone B cells in the spleen of wild-type mice (+/+), mice homozygous for cpm or pri
Exam 1 ID#: October 4, 2007
 Biology 4361 Name: KEY Exam 1 ID#: October 4, 2007 Multiple choice (one point each) (1-25) 1. The process of cells forming tissues and organs is called a. morphogenesis. b. differentiation. c. allometry.
Biology 4361 Name: KEY Exam 1 ID#: October 4, 2007 Multiple choice (one point each) (1-25) 1. The process of cells forming tissues and organs is called a. morphogenesis. b. differentiation. c. allometry.
Principles of Genetics
 Principles of Genetics Snustad, D ISBN-13: 9780470903599 Table of Contents C H A P T E R 1 The Science of Genetics 1 An Invitation 2 Three Great Milestones in Genetics 2 DNA as the Genetic Material 6 Genetics
Principles of Genetics Snustad, D ISBN-13: 9780470903599 Table of Contents C H A P T E R 1 The Science of Genetics 1 An Invitation 2 Three Great Milestones in Genetics 2 DNA as the Genetic Material 6 Genetics
Shavenbaby Couples Patterning to Epidermal Cell Shape Control. Chanut-Delalande H, Fernandes I, Roch F, Payre F, Plaza S (2006) PLoS Biol 4(9): e290
 Shavenbaby Couples Patterning to Epidermal Cell Shape Control. Chanut-Delalande H, Fernandes I, Roch F, Payre F, Plaza S (2006) PLoS Biol 4(9): e290 Question (from Introduction): How does svb control the
Shavenbaby Couples Patterning to Epidermal Cell Shape Control. Chanut-Delalande H, Fernandes I, Roch F, Payre F, Plaza S (2006) PLoS Biol 4(9): e290 Question (from Introduction): How does svb control the
Biology 218, practise Exam 2, 2011
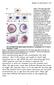 Figure 3 The long-range effect of Sqt does not depend on the induction of the endogenous cyc or sqt genes. a, Design and predictions for the experiments shown in b-e. b-e, Single-cell injection of 4 pg
Figure 3 The long-range effect of Sqt does not depend on the induction of the endogenous cyc or sqt genes. a, Design and predictions for the experiments shown in b-e. b-e, Single-cell injection of 4 pg
Supplementary Figure 1. Nature Genetics: doi: /ng.3848
 Supplementary Figure 1 Phenotypes and epigenetic properties of Fab2L flies. A- Phenotypic classification based on eye pigment levels in Fab2L male (orange bars) and female (yellow bars) flies (n>150).
Supplementary Figure 1 Phenotypes and epigenetic properties of Fab2L flies. A- Phenotypic classification based on eye pigment levels in Fab2L male (orange bars) and female (yellow bars) flies (n>150).
allosteric cis-acting DNA element coding strand dominant constitutive mutation coordinate regulation of genes denatured
 A B C D E F G H I J K L M N O P Q R S T U V W X Y Z AA BB CC DD EE FF GG HH II JJ KK LL MM NN OO PP QQ RR SS TT UU VV allosteric cis-acting DNA element coding strand codominant constitutive mutation coordinate
A B C D E F G H I J K L M N O P Q R S T U V W X Y Z AA BB CC DD EE FF GG HH II JJ KK LL MM NN OO PP QQ RR SS TT UU VV allosteric cis-acting DNA element coding strand codominant constitutive mutation coordinate
2. Der Dissertation zugrunde liegende Publikationen und Manuskripte. 2.1 Fine scale mapping in the sex locus region of the honey bee (Apis mellifera)
 2. Der Dissertation zugrunde liegende Publikationen und Manuskripte 2.1 Fine scale mapping in the sex locus region of the honey bee (Apis mellifera) M. Hasselmann 1, M. K. Fondrk², R. E. Page Jr.² und
2. Der Dissertation zugrunde liegende Publikationen und Manuskripte 2.1 Fine scale mapping in the sex locus region of the honey bee (Apis mellifera) M. Hasselmann 1, M. K. Fondrk², R. E. Page Jr.² und
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Overexpression of YFP::GPR-1 in the germline.
 Supplementary Figure 1 Overexpression of YFP::GPR-1 in the germline. The pie-1 promoter and 3 utr were used to express yfp::gpr-1 in the germline. Expression levels from the yfp::gpr-1(cai 1.0)-expressing
Supplementary Figure 1 Overexpression of YFP::GPR-1 in the germline. The pie-1 promoter and 3 utr were used to express yfp::gpr-1 in the germline. Expression levels from the yfp::gpr-1(cai 1.0)-expressing
Wingless and Hedgehog pattern Drosophila denticle belts by regulating the production of short-range signals
 Development 126, 5689-5698 (1999) Printed in Great Britain The Company of Biologists Limited 1999 DEV7768 5689 Wingless and Hedgehog pattern Drosophila denticle belts by regulating the production of short-range
Development 126, 5689-5698 (1999) Printed in Great Britain The Company of Biologists Limited 1999 DEV7768 5689 Wingless and Hedgehog pattern Drosophila denticle belts by regulating the production of short-range
Principles of Experimental Embryology
 Biology 4361 Developmental Biology Principles of Experimental Embryology September 19, 2006 Major Research Questions How do forces outside the embryo affect its development? (Environmental Developmental
Biology 4361 Developmental Biology Principles of Experimental Embryology September 19, 2006 Major Research Questions How do forces outside the embryo affect its development? (Environmental Developmental
Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression Edited by Shawn Lester PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley
Chapter 18 Regulation of Gene Expression Edited by Shawn Lester PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley
Solutions to Problem Set 4
 Question 1 Solutions to 7.014 Problem Set 4 Because you have not read much scientific literature, you decide to study the genetics of garden peas. You have two pure breeding pea strains. One that is tall
Question 1 Solutions to 7.014 Problem Set 4 Because you have not read much scientific literature, you decide to study the genetics of garden peas. You have two pure breeding pea strains. One that is tall
