Undirected graphical models
|
|
|
- Juliana Tyler
- 5 years ago
- Views:
Transcription
1 Undirected graphical models Kevin P. Murphy Last updated November 16, 2006 * Denotes advanced sections that may be omitted on a first reading. 1 Introduction We have seen that conditional independence assumptions are a useful way to decompose joint probability distributions into a simpler form. For example, the bag of words model assumes all variables are independent p(x 1:D ) = D p(x i ) (1) The naive Bayes model assumes all the x variables (features) are independent conditional on y (class): i=1 p(y, x 1:D ) = p(y)p(x 1:D y) = p(y) A Markov model assumes the variables only depend on their immediate predecessor p(x 1:D ) = p(x 1 ) D p(x i y) (2) i=1 D p(x i x i 1 ) (3) Graphical models provide a convenient way to represent conditional independence assumptions using graphs. Nodes represent random variables and (lack of) edges represent independence assumptions, in a way to be described below. Such independence assumptions reduce the number of parameters in the model, and therefore the amount of data that is needed to learn the model from data. Independence assumptions can also be exploited computationally to speed up the inference process (i.e., estimating one variable given evidence on some of the others). We will give details later. There are many kinds of graphical model, but the two most popular are based on directed acylic graphs (DAGs), and on undirected graphs. In the directed graph, we can (informally) think of an edge from node X i to node X j as meaning X i causes or directly influences X j, whereas in an undirected graph, edges represent correlation or constraints rather than causation. In this chapter, we focus on undirected graphs, also called Markov random fields (MRFs) or Markov networks. 2 Conditional independence properties In an MRF, we say that a set of nodes A is independent of another set B given a third set C (written more concisely as X A X B X C ) if C separates A from B in the graph. This means that, if we remove the nodes in C, there are no paths connecting any node in A to any node in B. For example, in Figure 1, we see that C = {2, 5} separates A = {1} from B = {4, 6}. We also see that C = {2} separates A = {3} from B = {4}. And so on. These are called the global Markov properties encoded by a graph. (There are various other kinds of Markov properties, discussed below, but these can be shown to be equivalent to the global Markov property under various assumptions.) Let us now consider some more examples. In Figure 2(left), we see that i=2 X i X j Y (4) 1
2 Figure 1: An example of an undirected graphical model. Figure 2: Left: a naive Bayes classifier represented as an MRF. Right: a Markov chain represented as an MRF. which is the naive Bayes assumption. Hence p(x 1:D, Y ) D ψ i (X i, Y ) (5) We can set ψ i (X i, Y ) = p(x i Y ). What about the p(y ) term? We can multiply that onto any of the clique potentials, say the first, so ψ 1 (X 1, Y ) = p(x 1 Y )p(y ) = p(x 1, Y ). Thus some of the potentials are conditional probabilities are some are joint probabilities. We will discuss this more below. In Figure 2(right), we see that X i+1 X i 1 X i (6) which is the Markov assumption. Hence p(x 1:D ) Again we can set ψ i (X i 1, X i ) = p(x i X i 1 ), and ψ 1 (X 1, X 2 ) = p(x 1 )p(x 2 X 1 ). 2.1 Varieties of conditional independencies * i=1 D ψ i (X i 1, X i ) (7) i=2 An undirected graph can represent various kinds of conditional independences. Define the pairwise Markov independencies of a graph G to be I p (G) = {X i X j X rest : (i j) G} (8) where X rest = V \ {i, j} are all the other variables apart from i and j, and V is all the nodes in the graph. In other words, for every missing edge i j, we create a conditional independence assertion. For example, in Figure 1, we 2
3 Figure 3: Nested sets of independence statements. have I p (G) = {(X 1 X 5 X 2, X 3, X 4, X 6 ), (X 1 X 4 X 2, X 3, X 5, X 6 ),..., } (9) (An alternative to listing all the independence properties is to think of the graph as an oracle, which can answer any conditional independence query.) A second kind of independence are the local Markov independencies, defined as I l (G) = {X i X rest X nbr(i) } (10) where nbr(i) are the neighbors of node i, and X rest = V \ {i} \ X nbr(i). (The neighbors of a node are also called its Markov blanket.) For example, in Figure 1, we have I l (G) = {(X 3 X 2, X 4, X 6 X 1, X 5 ),...} (11) since 1 and 5 shield 3 from the other nodes. The third kind of independence are the global Markov independencies, defined as I(G) = {X A X B X C : C separates A from B} (12) We can check if C separates A and B by removing all the nodes in C from the graph, and checking if there is a path from any node in A to any node in B. If so, they are not separated. For example, in Figure 1, C = {2, 5} separates A = {1} from B = {4, 6}, etc. Hence for this example I(G) = {(X 1 X 4, X 6 X 2, X 5 ),...} (13) Now we discuss the relationship between these three notions. Let I(p) be the set of conditional independencies that are true of distribution p. We say that G is an I-map (independency map) for p if I(G) I(p). In otherwords, the graph does not make any false assertions of independence. Note that the fully connected graph is an I-map of all distributions, since it makes no assertions at all. Theorem 1 If I(p) contains I(G)) (i.e., G is an I-map for p), then it will also contain I l (G) and I p (G): see Figure 3. In other words, global Markov properties imply local Markov properties imply pairwise Markov properties. If p is a positive distribution (i.e., p(x) > 0 for all x) then the converse is also true, hence I p (G) = I l (G) = I(G). 3 Factorization Above we saw that a graph encodes conditional independence assumptions. An alternative way to view graphical models is as a means to define probability distributions in terms of a product of local factors. To explain this, we need some definitions. 3
4 A clique is a set of variables in a graph that are all connected with each other. A maximal clique is a clique which cannot be made larger without ceasing to be a clique. For example, in Figure 1, the maximal cliques are {X 1, X 2 }, {X 1, X 3 }, {X 2, X 4 }, {X 3, X 5 }, {X 2, X 5, X 6 } (14) A potential function ψ( ) is any non negative function of its arguments. We say that a probability distribution p factorizes over G, or p is Markov wrt G (wrt = with respect to) if it can be written in this form: p(x) = 1 Z ψ c (x c ) (15) c C where C are the maximal cliques of the graph, ψ c (x c ) is a for the nodes in clique c and Z is the partition function Z = ψ c (x c ) (16) x 1:D c C The global normalization constant Z is needed because locally the ψ c terms need not sum to one. For example, in Figure 1, we have 3.1 Hammersley-Clifford theorem * p(x 1:6 ) ψ 12 (x 1, x 2 )ψ 13 (x 1, x 3 )ψ 24 (x 2, x 4 )ψ 35 (x 3, x 5 )ψ 256 (x 2, x 5, x 6 ) (17) The celebrated Hammersley-Clifford theorem states that, if the graph correctly captures the conditional independencies of the distribution, then there must exist potential functions ψ c such that the distribution can be represented as p(x) c ψ c(x c ). More precisely, Theorem 2 For any probability distribution p, if p factorizes over G, then G is an I-map of p (i.e., I(G) I(p)). Conversely, if p is everywhere non-negative (i.e, p(x) > 0 for all x), and G is an I-map of p, then p must factorize over G, i.e., p must have the form p(x) = 1 ψ c (x c ) (18) Z See [KF06] for a proof. 3.2 MRFs are exponential family distributions c C Let us assume a particular parametric form for each potential ψ c, and denote the corresponding parameters by θ c. Then we can write p(x θ) = 1 ψ c (x c θ c ) (19) Z(θ) c C We can rewrite this as an exponential family model as follows ( ) p(x θ) = exp log ψ c (x c θ c ) log Z(θ) c C If we restrict potentials to be strictly positive (which will imply that p(x) > 0 for all x, so we cannot encode hard constraints), then we can represent potentials using energy functions E c (20) ψ c (x c ) = exp[ E c (x c )] (21) This is called the Boltzmann distribution. Low energy states are more probable than high energy ones. We will examples of this below. 4
5 3.3 Maxent models * A maximum entropy (maxent) model is an MRF in which all the potentials have the form ψ c (x c ) = exp[ j=1 θ cj f cj (x c )] (22) where f cj (x c ) is the j th feature for clique c, applied to x c, and θ cj is the corresponding weight. In other words, the potentials are linear functions of a fixed set of features. Hence the joint is p(x θ) = = 1 Z(θ) 1 exp[ c j Z(θ) exp[ k θ cj f cj (x c )] (23) θ k f k (x)] (24) where we have combined all the features into one long feature vector f, and all the parameters into one long parameter vector θ. This is obviously a member of the exponential family. In machine, this model is sometimes called a loglinear model, although this means something slightly different in statistics, so we don t recommend this usage. We will explain the maxent term below. Note that feature based potentials include tabular potentials as a special case: we create one binary feature for every possible value of x c, and just set θ cj to be the log of the original potential entries. For example, consider a clique c = (i, j) on binary nodes. If the original potential ( ) ψ c = (25) 1 20 then we set f c1 (x i, x j ) = I(x i = 0, x j = 0), f c2 (x i, x j ) = I(x i = 0, x j = 1), f c3 (x i, x j ) = I(x i = 1, x j = 0), and f c4 (x i, x j ) = I(x i = 1, x j = 1), and then set θ c1 = log 0.1, θ c2 = log 0.5, θ c3 = log 1, and θ c4 = log 20. One example where a feature based representation of potentials is useful is when the number of states K is large and/or the cliques are large. In this case, the tabular representation of potentials may require too many parameters. For example, if we create a Markov model of language, we have one state per word, so K 50, 000. But K 2 parameters in each ψ(x i, X i+1 ) potential is too many to reasonably estimate. Instead of considering all possible word pairs, we can use log-linear potentials. Features might include does the first word begin with a capital letter, is the first word a noun and the second word a verb, etc. Typically these features are hand-constructed, but the weights (parameters) θ c are learned from data (see below). Such models have been used for unsupervised learning of the rules of English spelling [PPL97]. We now explain the origin of the term maximum entropy. Suppose we want to build a probability distribution that satisfies certain expectation constraints wrt some features f j : E[f j (X)] = α j (26) For example, we might want our model to generate English words with a certain proportion of each letter of the alphabet. Also, we may want to enforce hard constraints like q is always followed by u ; we just set f j (X) = 1 if X satisfies the constraint, and f j (X) = 0 otherwise, and set α j =. Apart from these constraints, we want pick as general a distribution as possible. In other words, we want to find a distribution p with maximum entropy, subject to the above constraints, plus the sum-to-one constraint. (We assume p x is a finite length vector of elements.) We can formalize this as a convex optimization problem subject to linear constraints: max p subject to p x log p x (27) x p x f j (x) = α j (28) x p x = 1 (29) x 5
6 Figure 4: Chain graphs in which the clique potentials are explicitly represented as random variables (square nodes). Left: one potential per maximal clique. Right: one potential per edge. We write the Lagrangian as J = x p x log p x + λ( x p x 1) + j θ j ( x p x f j (x) α j ) (30) Taking derivatives wrt a specific element p x J = p x [ log p x ] [ p x ] log p x + λ + p x p x p x j θ j f j (x) (31) = 1 log p x + j θ j f j (x) (32) = 0 (33) Hence log p x = 1 + λ + j θ j f j (x) (34) p x = e λ 1 exp[ j θ j f j (x)] (35) Since x p x = 1, we have Hence e λ 1 = 1 exp[ j θ jf j (x)] = Z (36) p x = 1 Z exp[ j θ j f j (x)] (37) So we have derived the exponential family using the maximum entropy principle, where the weights (parameters) θ j are the Lagrange multipliers. Typically the α j values are derived from data, as we discuss below. 4 Factor graphs * In a Bayesian model, the parameters are treated as random variables, and can therefore be viewed as nodes in the graph. Although the main graph of an MRF is undirected, the edges from the parameter nodes will be directed, since the priors on the parameters will usually be locally normalized. For example, referring to Figure 4(left), we 6
7 have p(x 1:4, ψ) = p(x 1:4 θ)p(ψ) (38) [ ] 1 = Z(ψ) ψ 124(x 124 ) ψ 234 (x 234 ) [p(ψ 124 )p(ψ 234 )] (39) Z(ψ) = ψ 124 (x 124 ) ψ 234 (x 234 ) (40) x 1,x 2,x 3,x 4 where ψ = (ψ 234, ψ 124 ) are the parameters. This is an example of a chain graph, which (loosely speaking) is a graph in which some nodes have directed arrows to sets of nodes connected together in undirected maximal cliques. Now imagine that instead of associating a potential with each maximal clique, we associate potentials with non maximal cliques. For example, referring to Figure 4(right), suppose ψ 124 = ψ 12 ψ 14 and ψ 234 = ψ 23 ψ 34 ψ 24, so we have one potential for each edge. (This is called a pairwise MRF.) Then p(x 1:4 ψ) = 1 Z <ij> ψ ij (x ij ) (41) = 1 Z ψ 12(x 12 )ψ 14 (x 14 )ψ 23 (x 23 )ψ 24 (x 24 )ψ 34 (x 34 ) (42) Note that the ψ 24 edge potential is shared (tied) across different max cliques, so the clique potentials are no longer independent. One way to make the parameterization of the MRF graphically explicit, without any committment to a Bayesian approach, is to use a factor graph. A factor graph is an undirected bipartite graph with two kinds of nodes. Round nodes represent variables, square nodes represent factors (potentials), and there is an edge from each variable to every factor that mentions it. In Figure 5, we show factor graphs for the two models mentioned above. It is possible to extend the conditional independence semantics of graphical model to include factor graphs [Fre03], but here we just use them as a convenient notation for making the parameterization more explicit. 5 Decomposable graphs * An undirected graph is said to be chordal or triangulated if every undirected loop/ cycle X 1 X 2 X k X 1 of length k 4 has a chord, i.e., an edge connects X i, X j for all non-adjacent nodes i,j in the cycle. The process of converting a non-chordal graph to chordal form is called triangulation. The importance of triangulated graphs will be explained later. Some simple examples of chordal and non chordal graphs are shown in Figure 6. A less obvious example is shown in Figure 7: this illustrates that triangulation is more than covering the graph with little triangles. In particular, we may have to add a lot of edges to ensure the graph is triangulated (such edges are called fill-in edges). Furthermore, there may be many ways to do this. Let us define the optimal triangulation as the one which adds the least number of fill-in edges. Let C(G) be the size of the largest maximal clique in the optimal triangulation of graph G. Then we define the tree width of the graph as T(G) = C(G) 1. (It is defined this way so that for trees, T(G) = 1.) For example, in Figure 7(right), we see that C(G) = 4 (look at the clique or the clique), which is larger than the maximal cliques in the non-triangulated graph in Figure 7(left). In this case, the max clique size is not very large, but in general, one can show that for planar 2D grids of size D = d d, the treewidth is O(d) [LT79]. The importance of this result will be discussed later. Triangulated graphs are also called decomposable graphs. A simple example of a decomposable graph is a chain, X 1 X 2 X 3 X 4. In this case the cliques are C 1 = {X 1, X 2 }, C 2 = {X 2, X 3 }, and C 3 = {X 3, X 4 }. Define the separators as the intersection of neighboring cliques: S ij = C i C j. For the chain, the separators are the internal nodes, S 1,2 = {X 2 }, S 2,3 = {X 3 }. See Figure 8. Let us denote the clique potentials by ψ c and the separator potentials by ψ s. If we define ψ c = p(x c ) and ψ s = p(x s ), then we can write the joint as p(x 1:D ) = c ψ c(x c ) s ψ (43) s(x s ) 7
8 (a) (b) (c) (d) Figure 5: Some factor graphs. (a) and (b) are topologically equivalent graphs, and represent the MRF in Figure 4(left), with one potential per maximal clique. (c) and (d) are topologically equivalent graphs, and represent the MRF in Figure 4(right), with one potential per edge. Figure 6: Left: a non triangulated graph. Right: one possible triangulated version. 8
9 Figure 7: Left: a non triangulated grid. Note the dotted loop is a chordless 4-cycle. Right: one possible triangulation. Figure 8: Cliques (ovals) and separators (squares) for a chain structured MRF. For example, in the case of the chain, we have p(x 1:4 ) = p(x 1, X 2 )p(x 2, X 3 )p(x 3, X 4 ) (44) p(x 2 )p(x 3 ) = p(x 1, X 2 )p(x 3 X 2 )p(x 4 X 3 ) (45) = p(x 1 X 2 )p(x 2 X 3 )p(x 3, X 4 ) (46) which corresponds to the joint distribution of a Markov chain going forwards or backwards in time. (Undirected graphs are acausal, and have no notion of directionality.) It turns out that Equation 43 applies to any decomposable graph, not just chains. The reason is that by dividing by the separator potentials, we convert marginals into conditionals, and then the chain rule applies. 6 Applications of MRFs 6.1 Image denoising One important application of MRFs is to low level vision. Consider the image in Figure 9 (top left). Let each true pixel be x i { 1, +1}. Suppose this image gets sent over some channel and gets corrupted by noise. What we 9
10 sigma=2.0 initial guess sample sample sample posterior mean after samples Figure 9: Example of image denoising using Gibbs sampling. We use an Ising prior with J = 1 and a Gaussian noise model with σ = 2. Figure 10: Grid-structured MRF. The shaded nodes y i are observed, the unshaded nodes x i are hidden and need to be estimated. 10
11 receive is the image in Figure 9 (top middle). Let each observed observed pixel be y i IR. Our goal is to estimate the underlying image from the noisy signal. There are two formulations of this. The first is to find the single best interpretation: x = argmax p(x 1:D y 1:D, θ) (47) x This is called the maximum a posteriori (MAP) estimate. An alternative is to find the single most probable guess for each pixel locally: x i = arg maxp(x i y 1:D, θ) (48) x i These are called the max marginals. Both of these are examples of state estimation (inference) which we will discuss later. The simplest model would be to assume that pixels are independent and have a uniform prior (i.e., +1 and -1 are equally likely). For the likelihood, let us assume a Gaussian noise model p(y i x i ) = N(y i x i, σ) (49) If σ is very small, then if x i = 1 then y i 1, and similarly if x i = +1 then y i +1, so we will can easily estimate x from y by thresholding each value separately. But if the noise level is high, we need to use some prior to help. A simple prior is to assume that nearby pixels like to be in the same state. In other words, if x i is on, its neighbors are more likely to be on; and vice versa if x i is off. This is called a smoothness prior. More precisely, we can write the probability model as p(x, y) = p(x)p(y x) (50) [ ] = 1 ψ ij (x i, x j ) p(y i x i ) (51) Z <ij> where φ i (x i ) = p(y i x i ) is called the local evidence potential for node i, and ψ ij is called the edge potential for edge i j. (We write φ i (x i ) as a function of x i only, since the y i s are observed constants.) The notation <ij> means product over all edges in the graph. We define the edge potential as ( ) e J ij e ψ ij (x i, x j ) = exp[j ij x i x j ] = Jij (52) where J ij = 0 if i j are not neighbors in the graph. If i j are neighbors, we set J ij = J > 0 which gives higher probability (lower energy) to configurations in which x i = x j (since x i x j = 1 if x i = x j, and x i x j = 1 if x i x j ). J is the strength of our smoothness prior. Using this model, we can then perform inference to estimate p(x y, θ), where θ = (J, σ) are the parameters of the prior and likelihood. In Figure 9, we show the result of using Gibbs sampling to draw samples from p(x y, θ). We also show an estimate of the posterior mean, E[X y, θ]. We will explain this and other algorithms for inference later. 6.2 Ising models The image denoising model above is closely related to the Ising model, which is used in statistical physics for modelling magnets. In particular, x i { 1, +} represents the atomic spin of a particle, either spin down or up. In an Ising model we assume that the coupling strength J ij = J is the same for all edges, and that the external field h i = H is the same for all nodes. In this case, the model becomes p(x) = E(x) e Jij i e Jij 1 exp[ βe(x)] (53) Z(θ) = [ <ij> Jx i x j + i Hx i ] (54) where E(x) is the energy of the system, and β = 1/(kT), where T is the temperature of the system, and k is Boltzmann s constant. Here θ = (J, H, β) are the parameters of the system. Often we take H = 0. 11
12 Figure 11: Markov random fields and some variants. BM = Boltzmann machine. RBM = Restricted Boltzmann machine (bipartite graph). PoE = product of experts. AMN = Associative Markov network. In a ferromagnet, neighboring spins want to be the same, so J > 0. In this case, the lowest energy (most probable) states are all -1s or all +1s. In an anti-ferromagnet, neighboring spins want to be different, so J < 0. In this case, the lowest energy (most probable) states are alternating checkerboards of +1/-1; these are called the ground states of the system. However, the system can only enter this state at temperature T = 0 (so β = ). At higher temperatures, differences in energy matter less, so the system will fluctuate around, and will spend time in other states (this is the motivation behind simulated annealing). If the J ij s and h i s are not constant, the result is called a spin glass in physics, or a Boltzmann machine or Hopfield network in machine learning. If the J ij s have mixed sign, then resulting system might exhibit frustration, since some states will want to be +1, and some -1. We discuss methods to learn the parameters J ij below. If we generalize the Ising model from binary to multiple states, we get a Potts model. For example, for 3 state variables, the edge potentials look like this e J e J e J ψ ij (x i, x j ) = exp[j I(x i = x j )] = e J e J e J (55) e J e J e J Obviously we can make J ij be non-constant in this case, too. If all J ij > 0, this is called an associative Markov network. See Figure 11 for some other model variants. 7 Parameter estimation* In this section, we will assume that all the variables are discrete, and that there is one parameter for each possible assignment to a clique potential, i.e., ψ c (x c ) can be represented as a table of K c numbers, where c is the size of clique c. (The results in this section also apply to Gaussian MRFs.) In Section 7.3, we consider a more general parameterization in which ψ c is defined in terms of a set of features. This can require fewer than K c parameters, which is useful if K or c is large. We will focus on maximum likelihood estimation, although Bayesian approaches can also be used (see e.g., [MG04]). 12
13 The log-likelihood of a set of N iid datasets x n is l(θ) = log 1 ψ c (x n,c θ c ) (56) Z n c = log ψ c (x nc θ c ) N log Z (57) n c = N(x c )log ψ c (x c ) N log Z (58) c x c where N(x c ) = n I(x c, x nc ) is the number of times clique c is in configuration x c in the data. For example, consider a chain X 1 X 2 X 3 X 4. Then l(θ) = N(x 1, x 2 )log ψ 12 (x 1, x 2 ) + N(x 2, x 3 )log ψ 23 (x 2, x 3 ) (59) x 1,x 2 x 2,x 3 + N(x 3, x 4 )log ψ 34 (x 3, x 4 ) N log Z (60) x 3,x 4 So the derivative wrt one of the clique potentials, say ψ 12, is l ψ 12 (x 1, x 2 ) = N(x 1, x 2 ) ψ 12 (x 1, x 2 ) N ψ 12 (x 1, log Z (61) x 2 ) The key question is: what are the derivatives wrt the log partition function? In this example we find And in general we have log Z ψ 12 (x 1, x 2 ) = 1 Z ψ 12 (x 1, x 2 ) ψ 12 (x 1, x 2 )ψ 23 (x 2, x 3 )ψ 34 (x 3, x 4 ) x 1,x 2,x 3,x 4 (62) = 1 Z ψ 12 (x 1, x 2 )ψ 12(x 1, x 2 )ψ 23(x 2, x 3)ψ 34 (x 3, x 4 ) (63) x 3,x 4 = 1 ψ 23 (x 2 Z 3)ψ 34 (x 3, x 4 ) x 3,x 4 (64) = 1 ψ 12 (x 1, x 2)ψ 23 (x 2, x 3 )ψ 34 (x 3, x 4 ) Z ψ x 12 (x 3,x 4 1, x 2 ) (65) = p(x 1, x 2, x 3, x 4 ) ψ x 12 (x 3,x 4 1, x 2 ) (66) = p(x 1, x 2) ψ 12 (x 1, x 2 ) (67) log Z ψ c (x c ) = p(x c) ψ c (x c ) (68) so the derivative of the log-likelihood is l ψ c (x c ) = N(x c) ψ c (x c ) N p(x c)) ψ c (x c ) (69) Hence l ψ c(x c) = 0 implies the following constraint must hold at the MLE: p ML (x c ) = N(x c) N 13 def = p(x c ) (70)
14 where p(x c ) is the empirical distribution on clique c. In other words, at the maximum likelihood setting of the parameters, for each clique, the model marginals must be equal to the observed marginals (empirical counts). This doesn t tell us how to get the ML parameters, it just gives us a condition that must be satisfied when we have them. The method we have to use depends on whether the graph is decomposable or not, and whether the potentials are defined on maximal cliques or on sub cliques, and on whether the potentials are fully parameterized (one parameter per clique configuration, as in a tabular representation), or whether they are parameterized more generally. We summarize the options below. Decomposable? Max. Cliques Potential Method Yes Yes Tabular Closed form - - Tabular IPF Gradient descent 7.1 MLE for decomposable graphs with tabular potentials on the max cliques If the graph is decomposable, and the clique potentials are defined on maximal cliques, and the potentials are tabular, then we can just write down the MLEs in terms of the empirical counts, since these simultaneously satisfy all the above constraints. Recall that for a decomposable MRF p(x 1:d ) = c ψ c(x c ) s φ (71) s(x s ) The MLE is gotten by setting ψ c = p(x c ) and ψ s = p(x s ). For example, consider a chain X 1 X 2 X 3. We set 7.2 Iterative proportional fitting (IPF) Consider the graph in Figure 5(d): ˆψ 12 ML (x 1, x 2 ) = p(x 1, x 2 ) (72) ˆψ 23 ML 2, x 3 ) = p(x 2, x 3 ), (73) ˆφ ML 12 (x 2 ) = p(x 2 ) (74) p(x 1:4 ) = 1 Z <ij> ψ ij (x ij ) (75) Although this is decomposable, the potentials are not defined on the maximal cliques. So the ψ 24 term contributes to both ψ 124 and ψ 234. Hence we cannot just set the potentials to the empirical marginals. Instead, we will use an iterative scheme. Recall that From this we infer l ψ c (x c ) = N(x c) ψ c (x c ) N p(x c)) ψ c (x c ) = 0 (76) N(x c ) N 1 ψc(x t c ) = pt (x c ) ψ t c(x c ) (77) where the t superscript denotes the values of the parameters at iteration t. We can solve this fixed point equation iteratively ψ t+1 c (x c ) = p(x c) p t (x c ) (78) This is called the iterative proportional fitting (IPF) algorithm. In pseudo code, we have for each clique c p c = normalize(empirical counts(x c )) 14
15 while not converged for each clique c ˆp t c = p(x c ψ t ) (*) ψ t+1 c = ψ t c pc ˆp t c The line marked (*) requires inference to compute the marginals. Note that if the graph is decomposable, IPF will terminate in one iteration. IPF is essentially a coordinate ascent method, where we update a whole clique potential at each step. Below we will consider gradient-based methods that update all the parameters at each step. We now give an example of fitting an MRF with two disconnected nodes. We use brute force enumeration for inference. Since there are only two variables, we can use a 2D matrix to represent the joint, and 1D vectors to represent the marginals. We observed some count data C(X 1, X 2 ) and want to choose potentials ψ 1 and ψ 2 such that they match the marginals C(X 1 ) = , , and C(X 2 ) = , , respectively. We find that the potentials stop changing after the first iteration, as expected. The final answers are ψ 1 = , , (79) ψ 2 = , , (80) If J(x 1, x 2 ) = ψ 1 (x 1 ) ψ 2 (x 2 ), You can check manually that x 1 J(x 1, x 2 ) = c 2 (x 2 ) and x 2 J(x 1, x 2 ) = c 1 (x 1 ). % IPFdemo % Approximate joint density as a product of two marginals % i.e., fit a 2 node disconnedted MRF X1 X2 C12 = [ ; ; ]; C12 = normalise(c12); C1 = sum(c12,2); C2 = sum(c12, 1); nstates = [3 3]; psi1 = ones(1,3); psi2 = ones(1,3); for iter=1:2 joint = psi1(:) * psi2(:) ; M1 = sum(joint,2); psi1 = psi1.* (C1./ M1) joint = psi1(:) * psi2(:) ; M2 = sum(joint,1); psi2 = psi2.* (C2./ M2) end joint = psi1(:) * psi2(:) ; assert(approxeq(c1, sum(joint,2))) assert(approxeq(c2, sum(joint,1))) We now give an example of fitting a loopy graph. Again we use brute force enumeration for inference. This takes about 10 iterations to converge. Because the joint is now on three variables, we use the tabular potential class to take care of all the bookkeeping involved in matching dimensions. % IPFdemo3 % Fit loopy MRF using iterative proportional fitting clqs = {[1 2], [2 3], [1 3]}; NC = length(clqs); N = 3; % Some count data C = reshape([ ], [2 2 2]); C = normalise(c); Cpot = tabularpot(1:n, 2*ones(1,N), C); for c=1:nc 15
16 counts{c} = marginalizepot(cpot, clqs{c}); end % Initial guess is all 1 s for c=1:nc pots{c} = tabularpot(clqs{c}, 2*ones(1,length(clqs{c}))); end converged = 0; iter = 0; thresh = 1e-3; % convergence threshold while converged converged = 1; for c=1:nc potsold{c} = pots{c}; end iter = iter + 1; fprintf( iter %d\n, iter); for c=1:nc J = multiplypots(pots{:}); Mc = marginalizepot(j, clqs{c}); pots{c}.t = pots{c}.t.* (counts{c}.t./ Mc.T); if approxeq(pots{c}.t, potsold{c}.t, thresh) converged = 0; end fprintf( c=%d\n, c) printtable(pots{c}) end end J = multiplypots(pots{:}); for c=1:nc Mc = marginalizepot(j, clqs{c}); assert(approxeq(counts{c}.t, Mc.T)) end 7.3 Gradient based methods for maxent models For a maxent model with general features, where ψ c (x c ) = θc T f c (x c ), the previous derivation simplifies somewhat. The log-likelihood becomes l(θ) = θ k f k (x n ) N log Z(θ) (81) n Hence the derivative is k l θ j = n f j (x n ) N θ j log Z (82) 16
17 Let us focus on the second term, the derivative wrt the log partition function. Hence the gradient is log Z(θ) θ j = = = = = = 1 Z(θ) (83) Z(θ) θ j 1 exp[ θ k f k (x)] (84) Z(θ) θ x j k 1 exp[θ k f k (x)] (85) Z(θ) θ x j k 1 [ exp[θ j f j (x)] exp[θ k f k (x)] (86) Z(θ) θ x j k j 1 [f j (x)exp[θ j f j (x)]] exp[θ k f k (x)] (87) Z(θ) k j x 1 f j (x) exp[θ k f k (x)] (88) Z(θ) x k = f j (x)p(x) x (89) = E[f j (X)] (90) l θ j = n f j (x n ) NE[f j (X)] (91) We can find the globally optimal MLE by passing this gradient to any gradient-based optimization algorithm, such as conjugate gradient, quasi-newton, etc. (In matlab, you can use fminunc (function minimization unconstrained) in the optimimization toolbox.) However, to compute the gradient, we have to evaluate E[f j (X)] = x p(x)f j (x) (92) for every feature f j, which might be expensive. Typically, each feature only depends on a subset of the variables. In particular, for a graphical model, each feature only depends on the variables in its clique, so what we need are ways to compute p(x c ) for each clique. We discuss this later. l At the optimum, = 0, so the expected features according to the model should match the expected features according to the data: θ k 1 N f j (x n ) = E[f j (X)] (93) n Hence maximum likelihood of an exponential family model gives the same results as maximum entropy, where the constraints are that the expected features match the empirical features. In the case that there is one feature for each possible clique value, then E[f cj (X c )] = p(x c = j), so this is saying the model s marginal probabilities should equal the empirical marginal probabilities, as we saw before. 7.4 Regularization Since maximum likelihood often overfits, it is very common to find MAP estimates ˆθ MAP = arg max θ l(θ) = arg max log p(θ) + log p(d θ) (94) θ where l(θ) is the penalized (regularized) log likelihood. It is common to use a Gaussian prior, p(θ) = N(θ µ, Σ) = 1 (2π) p/2 Σ 1/2 exp[ 1 2 (θ µ)t Σ 1 (θ µ)] (95) 17
18 . Figure 12: A Laplace prior If we set µ = 0 and Σ = λ 1 I, where λ is called the precision, then the log prior becomes where θ 2 2 = j θ2 j. The objective function becomes log p(θ) = λ θ 2 + const (96) l(θ) = log p(d θ) λ θ 2 2 (97) Hence λ reflects the degree of regularization. This is called an L2 penalty term, since it penalizes the L 2 norm of the parameters. It is also called weight decay, since it encourages parameters to become small. (A priori parameters are most likely to be zero.) The derivative of the penalized log likelihood becomes l θ j = [ n f j (x n )] NE[f j (X)] 2λθ j (98) and can be optimized using gradient methods. The term λ is often set by cross validation. An alternative to a Gaussian prior is to use a Laplace (double-sided exponential) prior: p(θ) = p j=1 Laplace(θ j λ) = j ( ) p λ λ 2 exp( λ θ j ) = exp( λ θ 1 ) (99) 2 See Figure 12. Hence the objective function becomes l(θ) = log p(d θ) λ θ 1 (100) This is called L1 regularization. Note that this function is non-differentiable if any θ j = 0, so care must be taken when optimizing this objective. The details are beyond the scope of this chapter. In the context of linear regression, L2 regularization is called ridge regression and L1 regularization is called lasso (least absolute shrinkage and selection operator). The key difference between them is that L2 encourages all 18
19 Figure 13: Illustration of L1 (left) vs L2 (right) regularization. Source: [HTF01] Figure parameters to become small, whereas L1 encourages some parameters to become exactly zero, and the rest to remain more or less unchanged. Thus L1 can be used to perform feature selection. To see why this happens, note that an L2 penalty on the parameter vector θ = (1, 0) is the same as on θ = (1/ (2), 1/ (2)), since (1, 0) 2 = (1/ (2), 1/ (2) 2 = 1 (101) However, for L1, setting θ = (1, 0) is cheaper than setting θ = (1/ (2), 1/ (2)), since (1, 0) 1 = 1 < (1/ (2), 1/ (2) 1 = (2) (102) Hence lasso prefers sparse solutions. Figure 13 shows this result visually. The ellipses represent the quadratic log likelihood function. The shaded blobs represent the log prior, whose size is controlled by λ. The MAP estimate is at the intersection point. It is apparent that with the L1 prior, the solution will always be at one of the corners (since they will touch the likelihood surface first). When performing regression with discrete features (factors), we need to include or exclude a block of columns for the result to be meaningful. This is called the group lasso. See [YL06] for details. 8 Structure learning * Given a dataset, we might want to find a graph structure that represents it. More formally, the dataset defines an empirical distribution ˆp emp (x) = 1 I(x = x n ) (103) N and we want to find a graph that models the independencies in ˆp e mp. Since the fully connected graph is an I-map of all distributions, this can represent ˆp emp. However, what we really want is a minimal I-map, i.e., the one that makes as few additional independence assumptions as necessary. One approach to this is to apply a series of conditional independence tests to the data (e.g., using χ 2 -tests), and then to fit a model consistent with those test results (see [Edw00] for details). Since there are O(2 D2 ) possible graphs on D nodes (since each edge in the adjacency matrix can be present or absent), this can be computationally expensive. In addition to computational cost, there are fundamental statistical problems: conditional independency tests return yes/no answers (based on a threshold). When combining such results, inconsistencies can arise. An n 19
20 alternative approach is to perform Bayesian model selection, which solves the statistical problem, at the cost of making the computational problem harder. We discuss this later. A final approach, in the case of log-linear models, is to put a sparsity promoting prior on the parameters θ c, which encourages them to go to zero, and then to perform MAP estimation. If all the weights between two nodes are zero, then the edge is effectively removed from the graph. References [Bis06] C. Bishop. Pattern recognition and machine learning. Springer, Draft version [Edw00] D. Edwards. Introduction to graphical modelling. Springer, nd edition. [Fre03] B. Frey. Extending factor graphs so as to unify directed and undirected graphical models. In UAI, [HTF01] T. Hastie, R. Tibshirani, and J. Friedman. The Elements of Statistical Learning. Springer, [KF06] D. Koller and N. Friedman. Bayesian networks and beyond To appear. [LT79] Richard J. Lipton and Robert E. Tarjan. A separator theorem for planar graphs. SIAM Journal of Applied Math, 36: , [MG04] I. Murray and Z. Ghahramani. Bayesian Learning in Undirected Graphical Models: Approximate MCMC algorithms. In UAI, [PPL97] S. Della Pietra, V. Della Pietra, and J. Lafferty. Inducing features of random fields. IEEE Trans. on Pattern Analysis and Machine Intelligence, 19(4), [YL06] M. Yuan and Y. Lin. Model selection and estimation in regression with grouped variables. J. Royal Statistical Society, Series B, 68(1):49 67,
CSC 412 (Lecture 4): Undirected Graphical Models
 CSC 412 (Lecture 4): Undirected Graphical Models Raquel Urtasun University of Toronto Feb 2, 2016 R Urtasun (UofT) CSC 412 Feb 2, 2016 1 / 37 Today Undirected Graphical Models: Semantics of the graph:
CSC 412 (Lecture 4): Undirected Graphical Models Raquel Urtasun University of Toronto Feb 2, 2016 R Urtasun (UofT) CSC 412 Feb 2, 2016 1 / 37 Today Undirected Graphical Models: Semantics of the graph:
Undirected Graphical Models: Markov Random Fields
 Undirected Graphical Models: Markov Random Fields 40-956 Advanced Topics in AI: Probabilistic Graphical Models Sharif University of Technology Soleymani Spring 2015 Markov Random Field Structure: undirected
Undirected Graphical Models: Markov Random Fields 40-956 Advanced Topics in AI: Probabilistic Graphical Models Sharif University of Technology Soleymani Spring 2015 Markov Random Field Structure: undirected
3 : Representation of Undirected GM
 10-708: Probabilistic Graphical Models 10-708, Spring 2016 3 : Representation of Undirected GM Lecturer: Eric P. Xing Scribes: Longqi Cai, Man-Chia Chang 1 MRF vs BN There are two types of graphical models:
10-708: Probabilistic Graphical Models 10-708, Spring 2016 3 : Representation of Undirected GM Lecturer: Eric P. Xing Scribes: Longqi Cai, Man-Chia Chang 1 MRF vs BN There are two types of graphical models:
Undirected Graphical Models
 Outline Hong Chang Institute of Computing Technology, Chinese Academy of Sciences Machine Learning Methods (Fall 2012) Outline Outline I 1 Introduction 2 Properties Properties 3 Generative vs. Conditional
Outline Hong Chang Institute of Computing Technology, Chinese Academy of Sciences Machine Learning Methods (Fall 2012) Outline Outline I 1 Introduction 2 Properties Properties 3 Generative vs. Conditional
Chris Bishop s PRML Ch. 8: Graphical Models
 Chris Bishop s PRML Ch. 8: Graphical Models January 24, 2008 Introduction Visualize the structure of a probabilistic model Design and motivate new models Insights into the model s properties, in particular
Chris Bishop s PRML Ch. 8: Graphical Models January 24, 2008 Introduction Visualize the structure of a probabilistic model Design and motivate new models Insights into the model s properties, in particular
Review: Directed Models (Bayes Nets)
 X Review: Directed Models (Bayes Nets) Lecture 3: Undirected Graphical Models Sam Roweis January 2, 24 Semantics: x y z if z d-separates x and y d-separation: z d-separates x from y if along every undirected
X Review: Directed Models (Bayes Nets) Lecture 3: Undirected Graphical Models Sam Roweis January 2, 24 Semantics: x y z if z d-separates x and y d-separation: z d-separates x from y if along every undirected
Probabilistic Graphical Models
 Probabilistic Graphical Models Brown University CSCI 295-P, Spring 213 Prof. Erik Sudderth Lecture 11: Inference & Learning Overview, Gaussian Graphical Models Some figures courtesy Michael Jordan s draft
Probabilistic Graphical Models Brown University CSCI 295-P, Spring 213 Prof. Erik Sudderth Lecture 11: Inference & Learning Overview, Gaussian Graphical Models Some figures courtesy Michael Jordan s draft
Probabilistic Graphical Models
 Probabilistic Graphical Models Lecture 11 CRFs, Exponential Family CS/CNS/EE 155 Andreas Krause Announcements Homework 2 due today Project milestones due next Monday (Nov 9) About half the work should
Probabilistic Graphical Models Lecture 11 CRFs, Exponential Family CS/CNS/EE 155 Andreas Krause Announcements Homework 2 due today Project milestones due next Monday (Nov 9) About half the work should
Graphical Models for Collaborative Filtering
 Graphical Models for Collaborative Filtering Le Song Machine Learning II: Advanced Topics CSE 8803ML, Spring 2012 Sequence modeling HMM, Kalman Filter, etc.: Similarity: the same graphical model topology,
Graphical Models for Collaborative Filtering Le Song Machine Learning II: Advanced Topics CSE 8803ML, Spring 2012 Sequence modeling HMM, Kalman Filter, etc.: Similarity: the same graphical model topology,
Lecture 6: Graphical Models: Learning
 Lecture 6: Graphical Models: Learning 4F13: Machine Learning Zoubin Ghahramani and Carl Edward Rasmussen Department of Engineering, University of Cambridge February 3rd, 2010 Ghahramani & Rasmussen (CUED)
Lecture 6: Graphical Models: Learning 4F13: Machine Learning Zoubin Ghahramani and Carl Edward Rasmussen Department of Engineering, University of Cambridge February 3rd, 2010 Ghahramani & Rasmussen (CUED)
Undirected graphical models
 Undirected graphical models Semantics of probabilistic models over undirected graphs Parameters of undirected models Example applications COMP-652 and ECSE-608, February 16, 2017 1 Undirected graphical
Undirected graphical models Semantics of probabilistic models over undirected graphs Parameters of undirected models Example applications COMP-652 and ECSE-608, February 16, 2017 1 Undirected graphical
Machine Learning Summer School
 Machine Learning Summer School Lecture 3: Learning parameters and structure Zoubin Ghahramani zoubin@eng.cam.ac.uk http://learning.eng.cam.ac.uk/zoubin/ Department of Engineering University of Cambridge,
Machine Learning Summer School Lecture 3: Learning parameters and structure Zoubin Ghahramani zoubin@eng.cam.ac.uk http://learning.eng.cam.ac.uk/zoubin/ Department of Engineering University of Cambridge,
Probabilistic Graphical Models
 2016 Robert Nowak Probabilistic Graphical Models 1 Introduction We have focused mainly on linear models for signals, in particular the subspace model x = Uθ, where U is a n k matrix and θ R k is a vector
2016 Robert Nowak Probabilistic Graphical Models 1 Introduction We have focused mainly on linear models for signals, in particular the subspace model x = Uθ, where U is a n k matrix and θ R k is a vector
Based on slides by Richard Zemel
 CSC 412/2506 Winter 2018 Probabilistic Learning and Reasoning Lecture 3: Directed Graphical Models and Latent Variables Based on slides by Richard Zemel Learning outcomes What aspects of a model can we
CSC 412/2506 Winter 2018 Probabilistic Learning and Reasoning Lecture 3: Directed Graphical Models and Latent Variables Based on slides by Richard Zemel Learning outcomes What aspects of a model can we
Directed and Undirected Graphical Models
 Directed and Undirected Davide Bacciu Dipartimento di Informatica Università di Pisa bacciu@di.unipi.it Machine Learning: Neural Networks and Advanced Models (AA2) Last Lecture Refresher Lecture Plan Directed
Directed and Undirected Davide Bacciu Dipartimento di Informatica Università di Pisa bacciu@di.unipi.it Machine Learning: Neural Networks and Advanced Models (AA2) Last Lecture Refresher Lecture Plan Directed
A graph contains a set of nodes (vertices) connected by links (edges or arcs)
 BOLTZMANN MACHINES Generative Models Graphical Models A graph contains a set of nodes (vertices) connected by links (edges or arcs) In a probabilistic graphical model, each node represents a random variable,
BOLTZMANN MACHINES Generative Models Graphical Models A graph contains a set of nodes (vertices) connected by links (edges or arcs) In a probabilistic graphical model, each node represents a random variable,
Part I. C. M. Bishop PATTERN RECOGNITION AND MACHINE LEARNING CHAPTER 8: GRAPHICAL MODELS
 Part I C. M. Bishop PATTERN RECOGNITION AND MACHINE LEARNING CHAPTER 8: GRAPHICAL MODELS Probabilistic Graphical Models Graphical representation of a probabilistic model Each variable corresponds to a
Part I C. M. Bishop PATTERN RECOGNITION AND MACHINE LEARNING CHAPTER 8: GRAPHICAL MODELS Probabilistic Graphical Models Graphical representation of a probabilistic model Each variable corresponds to a
Variational Inference (11/04/13)
 STA561: Probabilistic machine learning Variational Inference (11/04/13) Lecturer: Barbara Engelhardt Scribes: Matt Dickenson, Alireza Samany, Tracy Schifeling 1 Introduction In this lecture we will further
STA561: Probabilistic machine learning Variational Inference (11/04/13) Lecturer: Barbara Engelhardt Scribes: Matt Dickenson, Alireza Samany, Tracy Schifeling 1 Introduction In this lecture we will further
Representation of undirected GM. Kayhan Batmanghelich
 Representation of undirected GM Kayhan Batmanghelich Review Review: Directed Graphical Model Represent distribution of the form ny p(x 1,,X n = p(x i (X i i=1 Factorizes in terms of local conditional probabilities
Representation of undirected GM Kayhan Batmanghelich Review Review: Directed Graphical Model Represent distribution of the form ny p(x 1,,X n = p(x i (X i i=1 Factorizes in terms of local conditional probabilities
An Introduction to Bayesian Machine Learning
 1 An Introduction to Bayesian Machine Learning José Miguel Hernández-Lobato Department of Engineering, Cambridge University April 8, 2013 2 What is Machine Learning? The design of computational systems
1 An Introduction to Bayesian Machine Learning José Miguel Hernández-Lobato Department of Engineering, Cambridge University April 8, 2013 2 What is Machine Learning? The design of computational systems
Logistic Regression: Online, Lazy, Kernelized, Sequential, etc.
 Logistic Regression: Online, Lazy, Kernelized, Sequential, etc. Harsha Veeramachaneni Thomson Reuter Research and Development April 1, 2010 Harsha Veeramachaneni (TR R&D) Logistic Regression April 1, 2010
Logistic Regression: Online, Lazy, Kernelized, Sequential, etc. Harsha Veeramachaneni Thomson Reuter Research and Development April 1, 2010 Harsha Veeramachaneni (TR R&D) Logistic Regression April 1, 2010
1 Undirected Graphical Models. 2 Markov Random Fields (MRFs)
 Machine Learning (ML, F16) Lecture#07 (Thursday Nov. 3rd) Lecturer: Byron Boots Undirected Graphical Models 1 Undirected Graphical Models In the previous lecture, we discussed directed graphical models.
Machine Learning (ML, F16) Lecture#07 (Thursday Nov. 3rd) Lecturer: Byron Boots Undirected Graphical Models 1 Undirected Graphical Models In the previous lecture, we discussed directed graphical models.
Undirected Graphical Models
 Undirected Graphical Models 1 Conditional Independence Graphs Let G = (V, E) be an undirected graph with vertex set V and edge set E, and let A, B, and C be subsets of vertices. We say that C separates
Undirected Graphical Models 1 Conditional Independence Graphs Let G = (V, E) be an undirected graph with vertex set V and edge set E, and let A, B, and C be subsets of vertices. We say that C separates
High dimensional Ising model selection
 High dimensional Ising model selection Pradeep Ravikumar UT Austin (based on work with John Lafferty, Martin Wainwright) Sparse Ising model US Senate 109th Congress Banerjee et al, 2008 Estimate a sparse
High dimensional Ising model selection Pradeep Ravikumar UT Austin (based on work with John Lafferty, Martin Wainwright) Sparse Ising model US Senate 109th Congress Banerjee et al, 2008 Estimate a sparse
Learning discrete graphical models via generalized inverse covariance matrices
 Learning discrete graphical models via generalized inverse covariance matrices Duzhe Wang, Yiming Lv, Yongjoon Kim, Young Lee Department of Statistics University of Wisconsin-Madison {dwang282, lv23, ykim676,
Learning discrete graphical models via generalized inverse covariance matrices Duzhe Wang, Yiming Lv, Yongjoon Kim, Young Lee Department of Statistics University of Wisconsin-Madison {dwang282, lv23, ykim676,
Machine Learning 4771
 Machine Learning 4771 Instructor: Tony Jebara Topic 16 Undirected Graphs Undirected Separation Inferring Marginals & Conditionals Moralization Junction Trees Triangulation Undirected Graphs Separation
Machine Learning 4771 Instructor: Tony Jebara Topic 16 Undirected Graphs Undirected Separation Inferring Marginals & Conditionals Moralization Junction Trees Triangulation Undirected Graphs Separation
STA 4273H: Statistical Machine Learning
 STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 3 Linear
STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 3 Linear
Probabilistic Graphical Models
 Probabilistic Graphical Models Lecture 10 Undirected Models CS/CNS/EE 155 Andreas Krause Announcements Homework 2 due this Wednesday (Nov 4) in class Project milestones due next Monday (Nov 9) About half
Probabilistic Graphical Models Lecture 10 Undirected Models CS/CNS/EE 155 Andreas Krause Announcements Homework 2 due this Wednesday (Nov 4) in class Project milestones due next Monday (Nov 9) About half
Lecture 9: PGM Learning
 13 Oct 2014 Intro. to Stats. Machine Learning COMP SCI 4401/7401 Table of Contents I Learning parameters in MRFs 1 Learning parameters in MRFs Inference and Learning Given parameters (of potentials) and
13 Oct 2014 Intro. to Stats. Machine Learning COMP SCI 4401/7401 Table of Contents I Learning parameters in MRFs 1 Learning parameters in MRFs Inference and Learning Given parameters (of potentials) and
3 Undirected Graphical Models
 Massachusetts Institute of Technology Department of Electrical Engineering and Computer Science 6.438 Algorithms For Inference Fall 2014 3 Undirected Graphical Models In this lecture, we discuss undirected
Massachusetts Institute of Technology Department of Electrical Engineering and Computer Science 6.438 Algorithms For Inference Fall 2014 3 Undirected Graphical Models In this lecture, we discuss undirected
Introduction to graphical models: Lecture III
 Introduction to graphical models: Lecture III Martin Wainwright UC Berkeley Departments of Statistics, and EECS Martin Wainwright (UC Berkeley) Some introductory lectures January 2013 1 / 25 Introduction
Introduction to graphical models: Lecture III Martin Wainwright UC Berkeley Departments of Statistics, and EECS Martin Wainwright (UC Berkeley) Some introductory lectures January 2013 1 / 25 Introduction
Probabilistic Graphical Models
 Probabilistic Graphical Models David Sontag New York University Lecture 4, February 16, 2012 David Sontag (NYU) Graphical Models Lecture 4, February 16, 2012 1 / 27 Undirected graphical models Reminder
Probabilistic Graphical Models David Sontag New York University Lecture 4, February 16, 2012 David Sontag (NYU) Graphical Models Lecture 4, February 16, 2012 1 / 27 Undirected graphical models Reminder
CPSC 540: Machine Learning
 CPSC 540: Machine Learning Undirected Graphical Models Mark Schmidt University of British Columbia Winter 2016 Admin Assignment 3: 2 late days to hand it in today, Thursday is final day. Assignment 4:
CPSC 540: Machine Learning Undirected Graphical Models Mark Schmidt University of British Columbia Winter 2016 Admin Assignment 3: 2 late days to hand it in today, Thursday is final day. Assignment 4:
Conditional Independence and Factorization
 Conditional Independence and Factorization Seungjin Choi Department of Computer Science and Engineering Pohang University of Science and Technology 77 Cheongam-ro, Nam-gu, Pohang 37673, Korea seungjin@postech.ac.kr
Conditional Independence and Factorization Seungjin Choi Department of Computer Science and Engineering Pohang University of Science and Technology 77 Cheongam-ro, Nam-gu, Pohang 37673, Korea seungjin@postech.ac.kr
Cheng Soon Ong & Christian Walder. Canberra February June 2018
 Cheng Soon Ong & Christian Walder Research Group and College of Engineering and Computer Science Canberra February June 2018 Outlines Overview Introduction Linear Algebra Probability Linear Regression
Cheng Soon Ong & Christian Walder Research Group and College of Engineering and Computer Science Canberra February June 2018 Outlines Overview Introduction Linear Algebra Probability Linear Regression
Bayesian Learning in Undirected Graphical Models
 Bayesian Learning in Undirected Graphical Models Zoubin Ghahramani Gatsby Computational Neuroscience Unit University College London, UK http://www.gatsby.ucl.ac.uk/ Work with: Iain Murray and Hyun-Chul
Bayesian Learning in Undirected Graphical Models Zoubin Ghahramani Gatsby Computational Neuroscience Unit University College London, UK http://www.gatsby.ucl.ac.uk/ Work with: Iain Murray and Hyun-Chul
CS Lecture 4. Markov Random Fields
 CS 6347 Lecture 4 Markov Random Fields Recap Announcements First homework is available on elearning Reminder: Office hours Tuesday from 10am-11am Last Time Bayesian networks Today Markov random fields
CS 6347 Lecture 4 Markov Random Fields Recap Announcements First homework is available on elearning Reminder: Office hours Tuesday from 10am-11am Last Time Bayesian networks Today Markov random fields
Alternative Parameterizations of Markov Networks. Sargur Srihari
 Alternative Parameterizations of Markov Networks Sargur srihari@cedar.buffalo.edu 1 Topics Three types of parameterization 1. Gibbs Parameterization 2. Factor Graphs 3. Log-linear Models with Energy functions
Alternative Parameterizations of Markov Networks Sargur srihari@cedar.buffalo.edu 1 Topics Three types of parameterization 1. Gibbs Parameterization 2. Factor Graphs 3. Log-linear Models with Energy functions
Learning MN Parameters with Approximation. Sargur Srihari
 Learning MN Parameters with Approximation Sargur srihari@cedar.buffalo.edu 1 Topics Iterative exact learning of MN parameters Difficulty with exact methods Approximate methods Approximate Inference Belief
Learning MN Parameters with Approximation Sargur srihari@cedar.buffalo.edu 1 Topics Iterative exact learning of MN parameters Difficulty with exact methods Approximate methods Approximate Inference Belief
11. Learning graphical models
 Learning graphical models 11-1 11. Learning graphical models Maximum likelihood Parameter learning Structural learning Learning partially observed graphical models Learning graphical models 11-2 statistical
Learning graphical models 11-1 11. Learning graphical models Maximum likelihood Parameter learning Structural learning Learning partially observed graphical models Learning graphical models 11-2 statistical
CSC321 Lecture 18: Learning Probabilistic Models
 CSC321 Lecture 18: Learning Probabilistic Models Roger Grosse Roger Grosse CSC321 Lecture 18: Learning Probabilistic Models 1 / 25 Overview So far in this course: mainly supervised learning Language modeling
CSC321 Lecture 18: Learning Probabilistic Models Roger Grosse Roger Grosse CSC321 Lecture 18: Learning Probabilistic Models 1 / 25 Overview So far in this course: mainly supervised learning Language modeling
Intelligent Systems:
 Intelligent Systems: Undirected Graphical models (Factor Graphs) (2 lectures) Carsten Rother 15/01/2015 Intelligent Systems: Probabilistic Inference in DGM and UGM Roadmap for next two lectures Definition
Intelligent Systems: Undirected Graphical models (Factor Graphs) (2 lectures) Carsten Rother 15/01/2015 Intelligent Systems: Probabilistic Inference in DGM and UGM Roadmap for next two lectures Definition
13 : Variational Inference: Loopy Belief Propagation and Mean Field
 10-708: Probabilistic Graphical Models 10-708, Spring 2012 13 : Variational Inference: Loopy Belief Propagation and Mean Field Lecturer: Eric P. Xing Scribes: Peter Schulam and William Wang 1 Introduction
10-708: Probabilistic Graphical Models 10-708, Spring 2012 13 : Variational Inference: Loopy Belief Propagation and Mean Field Lecturer: Eric P. Xing Scribes: Peter Schulam and William Wang 1 Introduction
Lecture 3: More on regularization. Bayesian vs maximum likelihood learning
 Lecture 3: More on regularization. Bayesian vs maximum likelihood learning L2 and L1 regularization for linear estimators A Bayesian interpretation of regularization Bayesian vs maximum likelihood fitting
Lecture 3: More on regularization. Bayesian vs maximum likelihood learning L2 and L1 regularization for linear estimators A Bayesian interpretation of regularization Bayesian vs maximum likelihood fitting
Statistical Learning
 Statistical Learning Lecture 5: Bayesian Networks and Graphical Models Mário A. T. Figueiredo Instituto Superior Técnico & Instituto de Telecomunicações University of Lisbon, Portugal May 2018 Mário A.
Statistical Learning Lecture 5: Bayesian Networks and Graphical Models Mário A. T. Figueiredo Instituto Superior Técnico & Instituto de Telecomunicações University of Lisbon, Portugal May 2018 Mário A.
Probabilistic Graphical Models
 School of Computer Science Probabilistic Graphical Models Variational Inference II: Mean Field Method and Variational Principle Junming Yin Lecture 15, March 7, 2012 X 1 X 1 X 1 X 1 X 2 X 3 X 2 X 2 X 3
School of Computer Science Probabilistic Graphical Models Variational Inference II: Mean Field Method and Variational Principle Junming Yin Lecture 15, March 7, 2012 X 1 X 1 X 1 X 1 X 2 X 3 X 2 X 2 X 3
MAP Examples. Sargur Srihari
 MAP Examples Sargur srihari@cedar.buffalo.edu 1 Potts Model CRF for OCR Topics Image segmentation based on energy minimization 2 Examples of MAP Many interesting examples of MAP inference are instances
MAP Examples Sargur srihari@cedar.buffalo.edu 1 Potts Model CRF for OCR Topics Image segmentation based on energy minimization 2 Examples of MAP Many interesting examples of MAP inference are instances
Graphical Models and Kernel Methods
 Graphical Models and Kernel Methods Jerry Zhu Department of Computer Sciences University of Wisconsin Madison, USA MLSS June 17, 2014 1 / 123 Outline Graphical Models Probabilistic Inference Directed vs.
Graphical Models and Kernel Methods Jerry Zhu Department of Computer Sciences University of Wisconsin Madison, USA MLSS June 17, 2014 1 / 123 Outline Graphical Models Probabilistic Inference Directed vs.
Bayesian Networks: Construction, Inference, Learning and Causal Interpretation. Volker Tresp Summer 2016
 Bayesian Networks: Construction, Inference, Learning and Causal Interpretation Volker Tresp Summer 2016 1 Introduction So far we were mostly concerned with supervised learning: we predicted one or several
Bayesian Networks: Construction, Inference, Learning and Causal Interpretation Volker Tresp Summer 2016 1 Introduction So far we were mostly concerned with supervised learning: we predicted one or several
Generative and Discriminative Approaches to Graphical Models CMSC Topics in AI
 Generative and Discriminative Approaches to Graphical Models CMSC 35900 Topics in AI Lecture 2 Yasemin Altun January 26, 2007 Review of Inference on Graphical Models Elimination algorithm finds single
Generative and Discriminative Approaches to Graphical Models CMSC 35900 Topics in AI Lecture 2 Yasemin Altun January 26, 2007 Review of Inference on Graphical Models Elimination algorithm finds single
10708 Graphical Models: Homework 2
 10708 Graphical Models: Homework 2 Due Monday, March 18, beginning of class Feburary 27, 2013 Instructions: There are five questions (one for extra credit) on this assignment. There is a problem involves
10708 Graphical Models: Homework 2 Due Monday, March 18, beginning of class Feburary 27, 2013 Instructions: There are five questions (one for extra credit) on this assignment. There is a problem involves
Conditional Random Fields and beyond DANIEL KHASHABI CS 546 UIUC, 2013
 Conditional Random Fields and beyond DANIEL KHASHABI CS 546 UIUC, 2013 Outline Modeling Inference Training Applications Outline Modeling Problem definition Discriminative vs. Generative Chain CRF General
Conditional Random Fields and beyond DANIEL KHASHABI CS 546 UIUC, 2013 Outline Modeling Inference Training Applications Outline Modeling Problem definition Discriminative vs. Generative Chain CRF General
Basic math for biology
 Basic math for biology Lei Li Florida State University, Feb 6, 2002 The EM algorithm: setup Parametric models: {P θ }. Data: full data (Y, X); partial data Y. Missing data: X. Likelihood and maximum likelihood
Basic math for biology Lei Li Florida State University, Feb 6, 2002 The EM algorithm: setup Parametric models: {P θ }. Data: full data (Y, X); partial data Y. Missing data: X. Likelihood and maximum likelihood
Junction Tree, BP and Variational Methods
 Junction Tree, BP and Variational Methods Adrian Weller MLSALT4 Lecture Feb 21, 2018 With thanks to David Sontag (MIT) and Tony Jebara (Columbia) for use of many slides and illustrations For more information,
Junction Tree, BP and Variational Methods Adrian Weller MLSALT4 Lecture Feb 21, 2018 With thanks to David Sontag (MIT) and Tony Jebara (Columbia) for use of many slides and illustrations For more information,
Bayesian Networks: Construction, Inference, Learning and Causal Interpretation. Volker Tresp Summer 2014
 Bayesian Networks: Construction, Inference, Learning and Causal Interpretation Volker Tresp Summer 2014 1 Introduction So far we were mostly concerned with supervised learning: we predicted one or several
Bayesian Networks: Construction, Inference, Learning and Causal Interpretation Volker Tresp Summer 2014 1 Introduction So far we were mostly concerned with supervised learning: we predicted one or several
High-dimensional graphical model selection: Practical and information-theoretic limits
 1 High-dimensional graphical model selection: Practical and information-theoretic limits Martin Wainwright Departments of Statistics, and EECS UC Berkeley, California, USA Based on joint work with: John
1 High-dimensional graphical model selection: Practical and information-theoretic limits Martin Wainwright Departments of Statistics, and EECS UC Berkeley, California, USA Based on joint work with: John
Discrete Mathematics and Probability Theory Fall 2015 Lecture 21
 CS 70 Discrete Mathematics and Probability Theory Fall 205 Lecture 2 Inference In this note we revisit the problem of inference: Given some data or observations from the world, what can we infer about
CS 70 Discrete Mathematics and Probability Theory Fall 205 Lecture 2 Inference In this note we revisit the problem of inference: Given some data or observations from the world, what can we infer about
Notes on Machine Learning for and
 Notes on Machine Learning for 16.410 and 16.413 (Notes adapted from Tom Mitchell and Andrew Moore.) Choosing Hypotheses Generally want the most probable hypothesis given the training data Maximum a posteriori
Notes on Machine Learning for 16.410 and 16.413 (Notes adapted from Tom Mitchell and Andrew Moore.) Choosing Hypotheses Generally want the most probable hypothesis given the training data Maximum a posteriori
Lecture 2 Machine Learning Review
 Lecture 2 Machine Learning Review CMSC 35246: Deep Learning Shubhendu Trivedi & Risi Kondor University of Chicago March 29, 2017 Things we will look at today Formal Setup for Supervised Learning Things
Lecture 2 Machine Learning Review CMSC 35246: Deep Learning Shubhendu Trivedi & Risi Kondor University of Chicago March 29, 2017 Things we will look at today Formal Setup for Supervised Learning Things
Alternative Parameterizations of Markov Networks. Sargur Srihari
 Alternative Parameterizations of Markov Networks Sargur srihari@cedar.buffalo.edu 1 Topics Three types of parameterization 1. Gibbs Parameterization 2. Factor Graphs 3. Log-linear Models Features (Ising,
Alternative Parameterizations of Markov Networks Sargur srihari@cedar.buffalo.edu 1 Topics Three types of parameterization 1. Gibbs Parameterization 2. Factor Graphs 3. Log-linear Models Features (Ising,
Markov Random Fields
 Markov Random Fields Umamahesh Srinivas ipal Group Meeting February 25, 2011 Outline 1 Basic graph-theoretic concepts 2 Markov chain 3 Markov random field (MRF) 4 Gauss-Markov random field (GMRF), and
Markov Random Fields Umamahesh Srinivas ipal Group Meeting February 25, 2011 Outline 1 Basic graph-theoretic concepts 2 Markov chain 3 Markov random field (MRF) 4 Gauss-Markov random field (GMRF), and
Bayesian Machine Learning
 Bayesian Machine Learning Andrew Gordon Wilson ORIE 6741 Lecture 4 Occam s Razor, Model Construction, and Directed Graphical Models https://people.orie.cornell.edu/andrew/orie6741 Cornell University September
Bayesian Machine Learning Andrew Gordon Wilson ORIE 6741 Lecture 4 Occam s Razor, Model Construction, and Directed Graphical Models https://people.orie.cornell.edu/andrew/orie6741 Cornell University September
Does Better Inference mean Better Learning?
 Does Better Inference mean Better Learning? Andrew E. Gelfand, Rina Dechter & Alexander Ihler Department of Computer Science University of California, Irvine {agelfand,dechter,ihler}@ics.uci.edu Abstract
Does Better Inference mean Better Learning? Andrew E. Gelfand, Rina Dechter & Alexander Ihler Department of Computer Science University of California, Irvine {agelfand,dechter,ihler}@ics.uci.edu Abstract
CPSC 540: Machine Learning
 CPSC 540: Machine Learning More Approximate Inference Mark Schmidt University of British Columbia Winter 2018 Last Time: Approximate Inference We ve been discussing graphical models for density estimation,
CPSC 540: Machine Learning More Approximate Inference Mark Schmidt University of British Columbia Winter 2018 Last Time: Approximate Inference We ve been discussing graphical models for density estimation,
14 : Theory of Variational Inference: Inner and Outer Approximation
 10-708: Probabilistic Graphical Models 10-708, Spring 2017 14 : Theory of Variational Inference: Inner and Outer Approximation Lecturer: Eric P. Xing Scribes: Maria Ryskina, Yen-Chia Hsu 1 Introduction
10-708: Probabilistic Graphical Models 10-708, Spring 2017 14 : Theory of Variational Inference: Inner and Outer Approximation Lecturer: Eric P. Xing Scribes: Maria Ryskina, Yen-Chia Hsu 1 Introduction
Directed and Undirected Graphical Models
 Directed and Undirected Graphical Models Adrian Weller MLSALT4 Lecture Feb 26, 2016 With thanks to David Sontag (NYU) and Tony Jebara (Columbia) for use of many slides and illustrations For more information,
Directed and Undirected Graphical Models Adrian Weller MLSALT4 Lecture Feb 26, 2016 With thanks to David Sontag (NYU) and Tony Jebara (Columbia) for use of many slides and illustrations For more information,
Inference in Graphical Models Variable Elimination and Message Passing Algorithm
 Inference in Graphical Models Variable Elimination and Message Passing lgorithm Le Song Machine Learning II: dvanced Topics SE 8803ML, Spring 2012 onditional Independence ssumptions Local Markov ssumption
Inference in Graphical Models Variable Elimination and Message Passing lgorithm Le Song Machine Learning II: dvanced Topics SE 8803ML, Spring 2012 onditional Independence ssumptions Local Markov ssumption
Lecture 2: Simple Classifiers
 CSC 412/2506 Winter 2018 Probabilistic Learning and Reasoning Lecture 2: Simple Classifiers Slides based on Rich Zemel s All lecture slides will be available on the course website: www.cs.toronto.edu/~jessebett/csc412
CSC 412/2506 Winter 2018 Probabilistic Learning and Reasoning Lecture 2: Simple Classifiers Slides based on Rich Zemel s All lecture slides will be available on the course website: www.cs.toronto.edu/~jessebett/csc412
Chapter 16. Structured Probabilistic Models for Deep Learning
 Peng et al.: Deep Learning and Practice 1 Chapter 16 Structured Probabilistic Models for Deep Learning Peng et al.: Deep Learning and Practice 2 Structured Probabilistic Models way of using graphs to describe
Peng et al.: Deep Learning and Practice 1 Chapter 16 Structured Probabilistic Models for Deep Learning Peng et al.: Deep Learning and Practice 2 Structured Probabilistic Models way of using graphs to describe
Learning MN Parameters with Alternative Objective Functions. Sargur Srihari
 Learning MN Parameters with Alternative Objective Functions Sargur srihari@cedar.buffalo.edu 1 Topics Max Likelihood & Contrastive Objectives Contrastive Objective Learning Methods Pseudo-likelihood Gradient
Learning MN Parameters with Alternative Objective Functions Sargur srihari@cedar.buffalo.edu 1 Topics Max Likelihood & Contrastive Objectives Contrastive Objective Learning Methods Pseudo-likelihood Gradient
Linear Models for Regression
 Linear Models for Regression Machine Learning Torsten Möller Möller/Mori 1 Reading Chapter 3 of Pattern Recognition and Machine Learning by Bishop Chapter 3+5+6+7 of The Elements of Statistical Learning
Linear Models for Regression Machine Learning Torsten Möller Möller/Mori 1 Reading Chapter 3 of Pattern Recognition and Machine Learning by Bishop Chapter 3+5+6+7 of The Elements of Statistical Learning
A brief introduction to Conditional Random Fields
 A brief introduction to Conditional Random Fields Mark Johnson Macquarie University April, 2005, updated October 2010 1 Talk outline Graphical models Maximum likelihood and maximum conditional likelihood
A brief introduction to Conditional Random Fields Mark Johnson Macquarie University April, 2005, updated October 2010 1 Talk outline Graphical models Maximum likelihood and maximum conditional likelihood
Probabilistic Graphical Models & Applications
 Probabilistic Graphical Models & Applications Learning of Graphical Models Bjoern Andres and Bernt Schiele Max Planck Institute for Informatics The slides of today s lecture are authored by and shown with
Probabilistic Graphical Models & Applications Learning of Graphical Models Bjoern Andres and Bernt Schiele Max Planck Institute for Informatics The slides of today s lecture are authored by and shown with
High-dimensional graphical model selection: Practical and information-theoretic limits
 1 High-dimensional graphical model selection: Practical and information-theoretic limits Martin Wainwright Departments of Statistics, and EECS UC Berkeley, California, USA Based on joint work with: John
1 High-dimensional graphical model selection: Practical and information-theoretic limits Martin Wainwright Departments of Statistics, and EECS UC Berkeley, California, USA Based on joint work with: John
Sparse Graph Learning via Markov Random Fields
 Sparse Graph Learning via Markov Random Fields Xin Sui, Shao Tang Sep 23, 2016 Xin Sui, Shao Tang Sparse Graph Learning via Markov Random Fields Sep 23, 2016 1 / 36 Outline 1 Introduction to graph learning
Sparse Graph Learning via Markov Random Fields Xin Sui, Shao Tang Sep 23, 2016 Xin Sui, Shao Tang Sparse Graph Learning via Markov Random Fields Sep 23, 2016 1 / 36 Outline 1 Introduction to graph learning
6.867 Machine learning, lecture 23 (Jaakkola)
 Lecture topics: Markov Random Fields Probabilistic inference Markov Random Fields We will briefly go over undirected graphical models or Markov Random Fields (MRFs) as they will be needed in the context
Lecture topics: Markov Random Fields Probabilistic inference Markov Random Fields We will briefly go over undirected graphical models or Markov Random Fields (MRFs) as they will be needed in the context
Bayesian Learning in Undirected Graphical Models
 Bayesian Learning in Undirected Graphical Models Zoubin Ghahramani Gatsby Computational Neuroscience Unit University College London, UK http://www.gatsby.ucl.ac.uk/ and Center for Automated Learning and
Bayesian Learning in Undirected Graphical Models Zoubin Ghahramani Gatsby Computational Neuroscience Unit University College London, UK http://www.gatsby.ucl.ac.uk/ and Center for Automated Learning and
Lecture 15. Probabilistic Models on Graph
 Lecture 15. Probabilistic Models on Graph Prof. Alan Yuille Spring 2014 1 Introduction We discuss how to define probabilistic models that use richly structured probability distributions and describe how
Lecture 15. Probabilistic Models on Graph Prof. Alan Yuille Spring 2014 1 Introduction We discuss how to define probabilistic models that use richly structured probability distributions and describe how
Conditional Random Field
 Introduction Linear-Chain General Specific Implementations Conclusions Corso di Elaborazione del Linguaggio Naturale Pisa, May, 2011 Introduction Linear-Chain General Specific Implementations Conclusions
Introduction Linear-Chain General Specific Implementations Conclusions Corso di Elaborazione del Linguaggio Naturale Pisa, May, 2011 Introduction Linear-Chain General Specific Implementations Conclusions
Rapid Introduction to Machine Learning/ Deep Learning
 Rapid Introduction to Machine Learning/ Deep Learning Hyeong In Choi Seoul National University 1/24 Lecture 5b Markov random field (MRF) November 13, 2015 2/24 Table of contents 1 1. Objectives of Lecture
Rapid Introduction to Machine Learning/ Deep Learning Hyeong In Choi Seoul National University 1/24 Lecture 5b Markov random field (MRF) November 13, 2015 2/24 Table of contents 1 1. Objectives of Lecture
13: Variational inference II
 10-708: Probabilistic Graphical Models, Spring 2015 13: Variational inference II Lecturer: Eric P. Xing Scribes: Ronghuo Zheng, Zhiting Hu, Yuntian Deng 1 Introduction We started to talk about variational
10-708: Probabilistic Graphical Models, Spring 2015 13: Variational inference II Lecturer: Eric P. Xing Scribes: Ronghuo Zheng, Zhiting Hu, Yuntian Deng 1 Introduction We started to talk about variational
Markov Networks. l Like Bayes Nets. l Graphical model that describes joint probability distribution using tables (AKA potentials)
 Markov Networks l Like Bayes Nets l Graphical model that describes joint probability distribution using tables (AKA potentials) l Nodes are random variables l Labels are outcomes over the variables Markov
Markov Networks l Like Bayes Nets l Graphical model that describes joint probability distribution using tables (AKA potentials) l Nodes are random variables l Labels are outcomes over the variables Markov
Probabilistic Graphical Models. Guest Lecture by Narges Razavian Machine Learning Class April
 Probabilistic Graphical Models Guest Lecture by Narges Razavian Machine Learning Class April 14 2017 Today What is probabilistic graphical model and why it is useful? Bayesian Networks Basic Inference
Probabilistic Graphical Models Guest Lecture by Narges Razavian Machine Learning Class April 14 2017 Today What is probabilistic graphical model and why it is useful? Bayesian Networks Basic Inference
Statistical and Learning Techniques in Computer Vision Lecture 2: Maximum Likelihood and Bayesian Estimation Jens Rittscher and Chuck Stewart
 Statistical and Learning Techniques in Computer Vision Lecture 2: Maximum Likelihood and Bayesian Estimation Jens Rittscher and Chuck Stewart 1 Motivation and Problem In Lecture 1 we briefly saw how histograms
Statistical and Learning Techniques in Computer Vision Lecture 2: Maximum Likelihood and Bayesian Estimation Jens Rittscher and Chuck Stewart 1 Motivation and Problem In Lecture 1 we briefly saw how histograms
Graphical models: parameter learning
 Graphical models: parameter learning Zoubin Ghahramani Gatsby Computational Neuroscience Unit University College London London WC1N 3AR, England http://www.gatsby.ucl.ac.uk/ zoubin/ zoubin@gatsby.ucl.ac.uk
Graphical models: parameter learning Zoubin Ghahramani Gatsby Computational Neuroscience Unit University College London London WC1N 3AR, England http://www.gatsby.ucl.ac.uk/ zoubin/ zoubin@gatsby.ucl.ac.uk
Linear & nonlinear classifiers
 Linear & nonlinear classifiers Machine Learning Hamid Beigy Sharif University of Technology Fall 1394 Hamid Beigy (Sharif University of Technology) Linear & nonlinear classifiers Fall 1394 1 / 34 Table
Linear & nonlinear classifiers Machine Learning Hamid Beigy Sharif University of Technology Fall 1394 Hamid Beigy (Sharif University of Technology) Linear & nonlinear classifiers Fall 1394 1 / 34 Table
Lecture 16 Deep Neural Generative Models
 Lecture 16 Deep Neural Generative Models CMSC 35246: Deep Learning Shubhendu Trivedi & Risi Kondor University of Chicago May 22, 2017 Approach so far: We have considered simple models and then constructed
Lecture 16 Deep Neural Generative Models CMSC 35246: Deep Learning Shubhendu Trivedi & Risi Kondor University of Chicago May 22, 2017 Approach so far: We have considered simple models and then constructed
Introduction to Graphical Models. Srikumar Ramalingam School of Computing University of Utah
 Introduction to Graphical Models Srikumar Ramalingam School of Computing University of Utah Reference Christopher M. Bishop, Pattern Recognition and Machine Learning, Jonathan S. Yedidia, William T. Freeman,
Introduction to Graphical Models Srikumar Ramalingam School of Computing University of Utah Reference Christopher M. Bishop, Pattern Recognition and Machine Learning, Jonathan S. Yedidia, William T. Freeman,
Bayesian Learning. Two Roles for Bayesian Methods. Bayes Theorem. Choosing Hypotheses
 Bayesian Learning Two Roles for Bayesian Methods Probabilistic approach to inference. Quantities of interest are governed by prob. dist. and optimal decisions can be made by reasoning about these prob.
Bayesian Learning Two Roles for Bayesian Methods Probabilistic approach to inference. Quantities of interest are governed by prob. dist. and optimal decisions can be made by reasoning about these prob.
The Ising model and Markov chain Monte Carlo
 The Ising model and Markov chain Monte Carlo Ramesh Sridharan These notes give a short description of the Ising model for images and an introduction to Metropolis-Hastings and Gibbs Markov Chain Monte
The Ising model and Markov chain Monte Carlo Ramesh Sridharan These notes give a short description of the Ising model for images and an introduction to Metropolis-Hastings and Gibbs Markov Chain Monte
Learning Parameters of Undirected Models. Sargur Srihari
 Learning Parameters of Undirected Models Sargur srihari@cedar.buffalo.edu 1 Topics Log-linear Parameterization Likelihood Function Maximum Likelihood Parameter Estimation Simple and Conjugate Gradient
Learning Parameters of Undirected Models Sargur srihari@cedar.buffalo.edu 1 Topics Log-linear Parameterization Likelihood Function Maximum Likelihood Parameter Estimation Simple and Conjugate Gradient
Machine Learning and Computational Statistics, Spring 2017 Homework 2: Lasso Regression
 Machine Learning and Computational Statistics, Spring 2017 Homework 2: Lasso Regression Due: Monday, February 13, 2017, at 10pm (Submit via Gradescope) Instructions: Your answers to the questions below,
Machine Learning and Computational Statistics, Spring 2017 Homework 2: Lasso Regression Due: Monday, February 13, 2017, at 10pm (Submit via Gradescope) Instructions: Your answers to the questions below,
Probabilistic Graphical Models. Theory of Variational Inference: Inner and Outer Approximation. Lecture 15, March 4, 2013
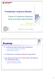 School of Computer Science Probabilistic Graphical Models Theory of Variational Inference: Inner and Outer Approximation Junming Yin Lecture 15, March 4, 2013 Reading: W & J Book Chapters 1 Roadmap Two
School of Computer Science Probabilistic Graphical Models Theory of Variational Inference: Inner and Outer Approximation Junming Yin Lecture 15, March 4, 2013 Reading: W & J Book Chapters 1 Roadmap Two
13 : Variational Inference: Loopy Belief Propagation
 10-708: Probabilistic Graphical Models 10-708, Spring 2014 13 : Variational Inference: Loopy Belief Propagation Lecturer: Eric P. Xing Scribes: Rajarshi Das, Zhengzhong Liu, Dishan Gupta 1 Introduction
10-708: Probabilistic Graphical Models 10-708, Spring 2014 13 : Variational Inference: Loopy Belief Propagation Lecturer: Eric P. Xing Scribes: Rajarshi Das, Zhengzhong Liu, Dishan Gupta 1 Introduction
Chapter 17: Undirected Graphical Models
 Chapter 17: Undirected Graphical Models The Elements of Statistical Learning Biaobin Jiang Department of Biological Sciences Purdue University bjiang@purdue.edu October 30, 2014 Biaobin Jiang (Purdue)
Chapter 17: Undirected Graphical Models The Elements of Statistical Learning Biaobin Jiang Department of Biological Sciences Purdue University bjiang@purdue.edu October 30, 2014 Biaobin Jiang (Purdue)
Computer Vision Group Prof. Daniel Cremers. 10a. Markov Chain Monte Carlo
 Group Prof. Daniel Cremers 10a. Markov Chain Monte Carlo Markov Chain Monte Carlo In high-dimensional spaces, rejection sampling and importance sampling are very inefficient An alternative is Markov Chain
Group Prof. Daniel Cremers 10a. Markov Chain Monte Carlo Markov Chain Monte Carlo In high-dimensional spaces, rejection sampling and importance sampling are very inefficient An alternative is Markov Chain
ECE521 week 3: 23/26 January 2017
 ECE521 week 3: 23/26 January 2017 Outline Probabilistic interpretation of linear regression - Maximum likelihood estimation (MLE) - Maximum a posteriori (MAP) estimation Bias-variance trade-off Linear
ECE521 week 3: 23/26 January 2017 Outline Probabilistic interpretation of linear regression - Maximum likelihood estimation (MLE) - Maximum a posteriori (MAP) estimation Bias-variance trade-off Linear
Learning Bayesian network : Given structure and completely observed data
 Learning Bayesian network : Given structure and completely observed data Probabilistic Graphical Models Sharif University of Technology Spring 2017 Soleymani Learning problem Target: true distribution
Learning Bayesian network : Given structure and completely observed data Probabilistic Graphical Models Sharif University of Technology Spring 2017 Soleymani Learning problem Target: true distribution
Statistical Data Mining and Machine Learning Hilary Term 2016
 Statistical Data Mining and Machine Learning Hilary Term 2016 Dino Sejdinovic Department of Statistics Oxford Slides and other materials available at: http://www.stats.ox.ac.uk/~sejdinov/sdmml Naïve Bayes
Statistical Data Mining and Machine Learning Hilary Term 2016 Dino Sejdinovic Department of Statistics Oxford Slides and other materials available at: http://www.stats.ox.ac.uk/~sejdinov/sdmml Naïve Bayes
Statistical Approaches to Learning and Discovery
 Statistical Approaches to Learning and Discovery Graphical Models Zoubin Ghahramani & Teddy Seidenfeld zoubin@cs.cmu.edu & teddy@stat.cmu.edu CALD / CS / Statistics / Philosophy Carnegie Mellon University
Statistical Approaches to Learning and Discovery Graphical Models Zoubin Ghahramani & Teddy Seidenfeld zoubin@cs.cmu.edu & teddy@stat.cmu.edu CALD / CS / Statistics / Philosophy Carnegie Mellon University
