Using rjags for survival data with right censoring
|
|
|
- Tyrone Burns
- 6 years ago
- Views:
Transcription
1 Using rjags for survival data with right censoring Malcolm Farrow Newcastle University October 4, Method using data augmentation Here is a simple example of a model specification for a survival problem with right censoring. model{ for(i in 1:119) { is.censored[i]~dinterval(t[i],t.cen[i]) t[i]~dexp(lambda[i]) lambda[i]<-exp(beta0+beta.trt*trt[i]) beta0~dnorm(0.0,0.001) beta.trt~dnorm(0.0,0.001) Suppose that this model specification is in a file called MFbug.txt. Suppose that the data are initially in the form of a data frame called data. This contains elements data$t : the event or censoring times data$status : 1 if the event is observed or 0 if the time is censored data$trt : a covariate The data set used by rjags contains four vectors. t: Element i of t is the event time if the event was observed for case i. If case i was censored then element i if t is NA. We can construct this as follows. 1
2 t<-data$t is.na(t)<-status==0 is.censored: This is an indicator taking the value 1 if the time is censored and 0 if the event is observed. We can construct this as follows. is.censored<-1-data$status t.cen: Element i of t.cen is the censoring time for case i is this case was censored. If the event was observed for case i then the censoring time must be greater than the observed event time. We can construct a suitable vector as follows. t.cen<-data$t+data$status trt: A covariate. trt<-data$trt Notice that, in the model specification, is.censored is given a special distribution is.censored[i]~dinterval(t[i],t.cen[i]) Experience has shown that it is usually necessary to initialise the missing values of t. This is because the specification if is.censored declares these cases to be censored. Therefore the unobserved event times must be greater than the censoring times. Allowing JAGS to choose initial values at random is likely to lead to some values for unobserved event times which are less than the censoring times and therefore inconsistent with this. So we should initialise the missing values of t to values greater than the corresponding values of t.cen. For example, to construct two sets of initial values, one for each of two parallel chains: tinits1<-data$t+5 is.na(tinits1)<-data$status==1 tinits2<-tinits1+5 We can now form the data and initial values lists and build the model as follows. kidneydata<-list(t=t,t.cen=t.cen,trt=trt,is.censored=is.censored) kidneyinits<-list(list(t=tinits1),list(t=tinits2)) kidneyjags<-jags.model("mfbug.txt",data=kidneydata,inits=kidneyinits,n.chains=2) Now away we go! update(kidneyjags,5000) kidneysamples<-coda.samples(kidneyjags,c( beta0, beta.trt ),100000) 2
3 2 Examples 2.1 Renal transplant data The data in Table 1 are taken from Henderson and Milner (1991). They show the graft survival times (t) in months of 148 renal transplant patients. There is one covariate, x, the total number of HLA-B or DR antigen mismatches between donor and recipient. The data can be obtained in the file renal.txt from You can read the data and put them into a form ready for us with commands such as the following. renal<-read.table("renal.txt",header=true) n<-length(renal[,1]) t<-renal$t is.na(t)<-renal$status==0 is.censored<-1-renal$status t.cen<-renal$t+renal$status renaldata<-list(n=n,t=t,is.censored=is.censored,t.cen=t.cen,x=renal$x) A suitable model specification is shown in Table 2. called renalbug.txt. Create initial values as follows. Type this into a file tinits1<-renal$t+5 is.na(tinits1)<-renal$status==1 tinits2<-tinits1+5 renalinits<-list(list(t=tinits1),list(t=tinits2)) Build the rjags model as follows. renaljags<-jags.model("renalbug.txt",data=renaldata,inits=renalinits,n.chains=2) Use a burn-in. update(renaljags,5000) Collect samples. As the mixing is rather poor, we will use a large number of iterations. renalsamples<-coda.samples(renaljags,c( beta0, beta.x ),50000) Look at the results. par(ask=true) traceplot(renalsamples) summary(renalsamples) densplot(renalsamples) 3
4 t Status x t Status x t Status x t Status x Table 1: Graft survival times in months for 148 renal transplant patients, with the number of antigen mismatches, x. Status is 1 for an observed graft failure and 0 for a censored observation. 4
5 model{ for (i in 1:n) { is.censored[i]~dinterval(t[i],t.cen[i]) t[i]~dexp(lambda[i]) lambda[i]<-exp(beta0+beta.x*x[i]) beta0~dnorm(0.0,0.001) beta.x~dnorm(0.0,0.001) Table 2: Model specification for renal transplant data. You might like to try fitting a Weibull model instead of the exponential model. In the model specification, change the definition of t[i] to t[i]~dweib(alpha,lambda[i]) and add the prior for α at the end. alpha~dgamma(1.1,1.1) 2.2 Gastric cancer data Table 3 shows survival times for two groups of patients with gastric cancer. Group 1 received chemotherapy and radiation. Group 2 just received chemotherapy. The data are taken from Gamerman (1991). The data are available in the file gastric.txt from You can read the data and put them into a form ready for us with commands such as the following. gastric<-read.table("gastric.txt",header=true) n<-length(gastric[,1]) t<-gastric$t is.na(t)<-gastric$status==0 is.censored<-1-gastric$status t.cen<-gastric$t+gastric$status gastricdata<-list(n=n,t=t,is.censored=is.censored,t.cen=t.cen, group=gastric$group) 5
6 Group 1 Group 2 t Status t Status t Status t Status t Status t Status Table 3: Survival times of patients with gastric cancer. Status is 1 for an observed death and 0 for a censored observation. A suitable model specification, with an exponential survival distibution, is shown in Table 4. Type this into a file called gastricbug.txt. Create initial values as follows. tinits1<-gastric$t+5 is.na(tinits1)<-gastric$status==1 tinits2<-tinits1+5 gastricinits<-list(list(t=tinits1),list(t=tinits2)) Build the rjags model as follows. gastricjags<-jags.model("gastricbug.txt",data=gastricdata,inits=gastricinits, n.chains=2) Try computing and examining the posterior distribution as in the case of the renal transplant data. You might also try using a Weibull distribution. 3 Alternative method by splitting the data As an alternative to using data augmentation, we can divide the data into observed and censored subsets. This method may be more efficient. As an example, we will use the gastric cancer data as above. We read the data as before but then partition the data into two subsets. gastric<-read.table("gastric.txt",header=true) 6
7 model{ for (i in 1:n) { is.censored[i]~dinterval(t[i],t.cen[i]) t[i]~dexp(lambda[i]) lambda[i]<-exp(beta0+beta.g[group[i]]) beta0~dnorm(0.0,0.001) beta1~dnorm(0.0,0.001) beta.g[1]<-beta1 beta.g[2]<- -beta1 Table 4: Model specification for gastric cancer data. model{ for (i in 1:n.obs) {t.obs[i]~dexp(lambda.obs[i]) lambda.obs[i]<-exp(beta0+beta.g[group.obs[i]]) for (i in 1:n.cen) {status.cen[i]~dbern(s[i]) S[i]<-pexp(t.cen[i],lambda.cen[i]) lambda.cen[i]<-exp(beta0+beta.g[group.cen[i]]) beta0~dnorm(0.0,0.001) beta1~dnorm(0.0,0.001) beta.g[1]<-beta1 beta.g[2]<- -beta1 Table 5: Alternative model specification for the gastric cancer example, without data augmentation. 7
8 gastric.obs<-subset(gastric,status==1) gastric.cen<-subset(gastric,status==0) n.obs<-length(gastric.obs$t) n.cen<-length(gastric.cen$t) gastricdata<-list(n.obs=n.obs,n.cen=n.cen,t.obs=gastric.obs$t,t.cen=gastric.cen$t, status.cen=gastric.cen$status, group.obs=gastric.obs$group,group.cen=gastric.cen$group) It is advisable to use initial values for the parameters so that the survival probabilities do do become to close to 0 or 1 at the first iteration. gastricinits<-list(list(beta0=-100,beta1=0),list(beta0=-100,beta1=0)) The model specification is shown in Table 5. The two parts of the likelihood, corresponding to the observed events (at times t.obs) and the censored observations (at times t.cen), are defined separately. The likelihood for the censored observations uses the distribution function pexp( ). Try this. You might also like to try using a Weibull distribution, using the following. S[i]<-pexp(t.cen[i],alpha,lambda.cen[i]) References Gamerman, D. (1991). Dynamic Bayesian models for survival data. Applied Statistics, 40, Henderson, R. and Milner, A. (1991). Aalen plots under proportional hazards. Applied Statistics, 40,
You know I m not goin diss you on the internet Cause my mama taught me better than that I m a survivor (What?) I m not goin give up (What?
 You know I m not goin diss you on the internet Cause my mama taught me better than that I m a survivor (What?) I m not goin give up (What?) I m not goin stop (What?) I m goin work harder (What?) Sir David
You know I m not goin diss you on the internet Cause my mama taught me better than that I m a survivor (What?) I m not goin give up (What?) I m not goin stop (What?) I m goin work harder (What?) Sir David
Package crrsc. R topics documented: February 19, 2015
 Package crrsc February 19, 2015 Title Competing risks regression for Stratified and Clustered data Version 1.1 Author Bingqing Zhou and Aurelien Latouche Extension of cmprsk to Stratified and Clustered
Package crrsc February 19, 2015 Title Competing risks regression for Stratified and Clustered data Version 1.1 Author Bingqing Zhou and Aurelien Latouche Extension of cmprsk to Stratified and Clustered
Package SimSCRPiecewise
 Package SimSCRPiecewise July 27, 2016 Type Package Title 'Simulates Univariate and Semi-Competing Risks Data Given Covariates and Piecewise Exponential Baseline Hazards' Version 0.1.1 Author Andrew G Chapple
Package SimSCRPiecewise July 27, 2016 Type Package Title 'Simulates Univariate and Semi-Competing Risks Data Given Covariates and Piecewise Exponential Baseline Hazards' Version 0.1.1 Author Andrew G Chapple
Multivariate Survival Analysis
 Multivariate Survival Analysis Previously we have assumed that either (X i, δ i ) or (X i, δ i, Z i ), i = 1,..., n, are i.i.d.. This may not always be the case. Multivariate survival data can arise in
Multivariate Survival Analysis Previously we have assumed that either (X i, δ i ) or (X i, δ i, Z i ), i = 1,..., n, are i.i.d.. This may not always be the case. Multivariate survival data can arise in
Mixture modelling of recurrent event times with long-term survivors: Analysis of Hutterite birth intervals. John W. Mac McDonald & Alessandro Rosina
 Mixture modelling of recurrent event times with long-term survivors: Analysis of Hutterite birth intervals John W. Mac McDonald & Alessandro Rosina Quantitative Methods in the Social Sciences Seminar -
Mixture modelling of recurrent event times with long-term survivors: Analysis of Hutterite birth intervals John W. Mac McDonald & Alessandro Rosina Quantitative Methods in the Social Sciences Seminar -
Ronald Christensen. University of New Mexico. Albuquerque, New Mexico. Wesley Johnson. University of California, Irvine. Irvine, California
 Texts in Statistical Science Bayesian Ideas and Data Analysis An Introduction for Scientists and Statisticians Ronald Christensen University of New Mexico Albuquerque, New Mexico Wesley Johnson University
Texts in Statistical Science Bayesian Ideas and Data Analysis An Introduction for Scientists and Statisticians Ronald Christensen University of New Mexico Albuquerque, New Mexico Wesley Johnson University
Package ICBayes. September 24, 2017
 Package ICBayes September 24, 2017 Title Bayesian Semiparametric Models for Interval-Censored Data Version 1.1 Date 2017-9-24 Author Chun Pan, Bo Cai, Lianming Wang, and Xiaoyan Lin Maintainer Chun Pan
Package ICBayes September 24, 2017 Title Bayesian Semiparametric Models for Interval-Censored Data Version 1.1 Date 2017-9-24 Author Chun Pan, Bo Cai, Lianming Wang, and Xiaoyan Lin Maintainer Chun Pan
5. Parametric Regression Model
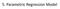 5. Parametric Regression Model The Accelerated Failure Time (AFT) Model Denote by S (t) and S 2 (t) the survival functions of two populations. The AFT model says that there is a constant c > 0 such that
5. Parametric Regression Model The Accelerated Failure Time (AFT) Model Denote by S (t) and S 2 (t) the survival functions of two populations. The AFT model says that there is a constant c > 0 such that
MAS3301 / MAS8311 Biostatistics Part II: Survival
 MAS330 / MAS83 Biostatistics Part II: Survival M. Farrow School of Mathematics and Statistics Newcastle University Semester 2, 2009-0 8 Parametric models 8. Introduction In the last few sections (the KM
MAS330 / MAS83 Biostatistics Part II: Survival M. Farrow School of Mathematics and Statistics Newcastle University Semester 2, 2009-0 8 Parametric models 8. Introduction In the last few sections (the KM
Survival Analysis. Stat 526. April 13, 2018
 Survival Analysis Stat 526 April 13, 2018 1 Functions of Survival Time Let T be the survival time for a subject Then P [T < 0] = 0 and T is a continuous random variable The Survival function is defined
Survival Analysis Stat 526 April 13, 2018 1 Functions of Survival Time Let T be the survival time for a subject Then P [T < 0] = 0 and T is a continuous random variable The Survival function is defined
Multistate models and recurrent event models
 Multistate models Multistate models and recurrent event models Patrick Breheny December 10 Patrick Breheny Survival Data Analysis (BIOS 7210) 1/22 Introduction Multistate models In this final lecture,
Multistate models Multistate models and recurrent event models Patrick Breheny December 10 Patrick Breheny Survival Data Analysis (BIOS 7210) 1/22 Introduction Multistate models In this final lecture,
Chapter 7 Fall Chapter 7 Hypothesis testing Hypotheses of interest: (A) 1-sample
 Bios 323: Applied Survival Analysis Qingxia (Cindy) Chen Chapter 7 Fall 2012 Chapter 7 Hypothesis testing Hypotheses of interest: (A) 1-sample H 0 : S(t) = S 0 (t), where S 0 ( ) is known survival function,
Bios 323: Applied Survival Analysis Qingxia (Cindy) Chen Chapter 7 Fall 2012 Chapter 7 Hypothesis testing Hypotheses of interest: (A) 1-sample H 0 : S(t) = S 0 (t), where S 0 ( ) is known survival function,
ADVANCED STATISTICAL ANALYSIS OF EPIDEMIOLOGICAL STUDIES. Cox s regression analysis Time dependent explanatory variables
 ADVANCED STATISTICAL ANALYSIS OF EPIDEMIOLOGICAL STUDIES Cox s regression analysis Time dependent explanatory variables Henrik Ravn Bandim Health Project, Statens Serum Institut 4 November 2011 1 / 53
ADVANCED STATISTICAL ANALYSIS OF EPIDEMIOLOGICAL STUDIES Cox s regression analysis Time dependent explanatory variables Henrik Ravn Bandim Health Project, Statens Serum Institut 4 November 2011 1 / 53
MAS3301 / MAS8311 Biostatistics Part II: Survival
 MAS3301 / MAS8311 Biostatistics Part II: Survival M. Farrow School of Mathematics and Statistics Newcastle University Semester 2, 2009-10 1 13 The Cox proportional hazards model 13.1 Introduction In the
MAS3301 / MAS8311 Biostatistics Part II: Survival M. Farrow School of Mathematics and Statistics Newcastle University Semester 2, 2009-10 1 13 The Cox proportional hazards model 13.1 Introduction In the
Extensions of Cox Model for Non-Proportional Hazards Purpose
 PhUSE Annual Conference 2013 Paper SP07 Extensions of Cox Model for Non-Proportional Hazards Purpose Author: Jadwiga Borucka PAREXEL, Warsaw, Poland Brussels 13 th - 16 th October 2013 Presentation Plan
PhUSE Annual Conference 2013 Paper SP07 Extensions of Cox Model for Non-Proportional Hazards Purpose Author: Jadwiga Borucka PAREXEL, Warsaw, Poland Brussels 13 th - 16 th October 2013 Presentation Plan
Multistate models and recurrent event models
 and recurrent event models Patrick Breheny December 6 Patrick Breheny University of Iowa Survival Data Analysis (BIOS:7210) 1 / 22 Introduction In this final lecture, we will briefly look at two other
and recurrent event models Patrick Breheny December 6 Patrick Breheny University of Iowa Survival Data Analysis (BIOS:7210) 1 / 22 Introduction In this final lecture, we will briefly look at two other
9 Estimating the Underlying Survival Distribution for a
 9 Estimating the Underlying Survival Distribution for a Proportional Hazards Model So far the focus has been on the regression parameters in the proportional hazards model. These parameters describe the
9 Estimating the Underlying Survival Distribution for a Proportional Hazards Model So far the focus has been on the regression parameters in the proportional hazards model. These parameters describe the
Simple techniques for comparing survival functions with interval-censored data
 Simple techniques for comparing survival functions with interval-censored data Jinheum Kim, joint with Chung Mo Nam jinhkim@suwon.ac.kr Department of Applied Statistics University of Suwon Comparing survival
Simple techniques for comparing survival functions with interval-censored data Jinheum Kim, joint with Chung Mo Nam jinhkim@suwon.ac.kr Department of Applied Statistics University of Suwon Comparing survival
Textbook: Survivial Analysis Techniques for Censored and Truncated Data 2nd edition, by Klein and Moeschberger
 Lecturer: James Degnan Office: SMLC 342 Office hours: MW 12:00 1:00 or by appointment E-mail: jamdeg@unm.edu Please include STAT474 or STAT574 in the subject line of the email to make sure I don t overlook
Lecturer: James Degnan Office: SMLC 342 Office hours: MW 12:00 1:00 or by appointment E-mail: jamdeg@unm.edu Please include STAT474 or STAT574 in the subject line of the email to make sure I don t overlook
Introduction to BGPhazard
 Introduction to BGPhazard José A. García-Bueno and Luis E. Nieto-Barajas February 11, 2016 Abstract We present BGPhazard, an R package which computes hazard rates from a Bayesian nonparametric view. This
Introduction to BGPhazard José A. García-Bueno and Luis E. Nieto-Barajas February 11, 2016 Abstract We present BGPhazard, an R package which computes hazard rates from a Bayesian nonparametric view. This
Multistate models in survival and event history analysis
 Multistate models in survival and event history analysis Dorota M. Dabrowska UCLA November 8, 2011 Research supported by the grant R01 AI067943 from NIAID. The content is solely the responsibility of the
Multistate models in survival and event history analysis Dorota M. Dabrowska UCLA November 8, 2011 Research supported by the grant R01 AI067943 from NIAID. The content is solely the responsibility of the
Lecture 6 PREDICTING SURVIVAL UNDER THE PH MODEL
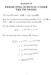 Lecture 6 PREDICTING SURVIVAL UNDER THE PH MODEL The Cox PH model: λ(t Z) = λ 0 (t) exp(β Z). How do we estimate the survival probability, S z (t) = S(t Z) = P (T > t Z), for an individual with covariates
Lecture 6 PREDICTING SURVIVAL UNDER THE PH MODEL The Cox PH model: λ(t Z) = λ 0 (t) exp(β Z). How do we estimate the survival probability, S z (t) = S(t Z) = P (T > t Z), for an individual with covariates
MAS8303 Modern Bayesian Inference Part 2
 MAS8303 Modern Bayesian Inference Part 2 M. Farrow School of Mathematics and Statistics Newcastle University Semester 1, 2012-13 48 Chapter 2 Generalised Linear Models 2.1 Generalised Linear Models 2.1.1
MAS8303 Modern Bayesian Inference Part 2 M. Farrow School of Mathematics and Statistics Newcastle University Semester 1, 2012-13 48 Chapter 2 Generalised Linear Models 2.1 Generalised Linear Models 2.1.1
Multilevel Statistical Models: 3 rd edition, 2003 Contents
 Multilevel Statistical Models: 3 rd edition, 2003 Contents Preface Acknowledgements Notation Two and three level models. A general classification notation and diagram Glossary Chapter 1 An introduction
Multilevel Statistical Models: 3 rd edition, 2003 Contents Preface Acknowledgements Notation Two and three level models. A general classification notation and diagram Glossary Chapter 1 An introduction
Available online Journal of Scientific and Engineering Research, 2018, 5(6): Research Article
 Available online www.jsaer.com, 2018, 5(6):32-51 Research Article ISSN: 2394-2630 CODEN(USA): JSERBR A Bayesian approach to relaxing the proportional hazard model and the form of baseline hazard in Survival
Available online www.jsaer.com, 2018, 5(6):32-51 Research Article ISSN: 2394-2630 CODEN(USA): JSERBR A Bayesian approach to relaxing the proportional hazard model and the form of baseline hazard in Survival
Lecture 7 Time-dependent Covariates in Cox Regression
 Lecture 7 Time-dependent Covariates in Cox Regression So far, we ve been considering the following Cox PH model: λ(t Z) = λ 0 (t) exp(β Z) = λ 0 (t) exp( β j Z j ) where β j is the parameter for the the
Lecture 7 Time-dependent Covariates in Cox Regression So far, we ve been considering the following Cox PH model: λ(t Z) = λ 0 (t) exp(β Z) = λ 0 (t) exp( β j Z j ) where β j is the parameter for the the
CTDL-Positive Stable Frailty Model
 CTDL-Positive Stable Frailty Model M. Blagojevic 1, G. MacKenzie 2 1 Department of Mathematics, Keele University, Staffordshire ST5 5BG,UK and 2 Centre of Biostatistics, University of Limerick, Ireland
CTDL-Positive Stable Frailty Model M. Blagojevic 1, G. MacKenzie 2 1 Department of Mathematics, Keele University, Staffordshire ST5 5BG,UK and 2 Centre of Biostatistics, University of Limerick, Ireland
Individualized Treatment Effects with Censored Data via Nonparametric Accelerated Failure Time Models
 Individualized Treatment Effects with Censored Data via Nonparametric Accelerated Failure Time Models Nicholas C. Henderson Thomas A. Louis Gary Rosner Ravi Varadhan Johns Hopkins University July 31, 2018
Individualized Treatment Effects with Censored Data via Nonparametric Accelerated Failure Time Models Nicholas C. Henderson Thomas A. Louis Gary Rosner Ravi Varadhan Johns Hopkins University July 31, 2018
Cox regression: Estimation
 Cox regression: Estimation Patrick Breheny October 27 Patrick Breheny Survival Data Analysis (BIOS 7210) 1/19 Introduction The Cox Partial Likelihood In our last lecture, we introduced the Cox partial
Cox regression: Estimation Patrick Breheny October 27 Patrick Breheny Survival Data Analysis (BIOS 7210) 1/19 Introduction The Cox Partial Likelihood In our last lecture, we introduced the Cox partial
Package LBLGXE. R topics documented: July 20, Type Package
 Type Package Package LBLGXE July 20, 2015 Title Bayesian Lasso for detecting Rare (or Common) Haplotype Association and their interactions with Environmental Covariates Version 1.2 Date 2015-07-09 Author
Type Package Package LBLGXE July 20, 2015 Title Bayesian Lasso for detecting Rare (or Common) Haplotype Association and their interactions with Environmental Covariates Version 1.2 Date 2015-07-09 Author
Two-stage Adaptive Randomization for Delayed Response in Clinical Trials
 Two-stage Adaptive Randomization for Delayed Response in Clinical Trials Guosheng Yin Department of Statistics and Actuarial Science The University of Hong Kong Joint work with J. Xu PSI and RSS Journal
Two-stage Adaptive Randomization for Delayed Response in Clinical Trials Guosheng Yin Department of Statistics and Actuarial Science The University of Hong Kong Joint work with J. Xu PSI and RSS Journal
Practical considerations for survival models
 Including historical data in the analysis of clinical trials using the modified power prior Practical considerations for survival models David Dejardin 1 2, Joost van Rosmalen 3 and Emmanuel Lesaffre 1
Including historical data in the analysis of clinical trials using the modified power prior Practical considerations for survival models David Dejardin 1 2, Joost van Rosmalen 3 and Emmanuel Lesaffre 1
Accelerated Failure Time Models
 Accelerated Failure Time Models Patrick Breheny October 12 Patrick Breheny University of Iowa Survival Data Analysis (BIOS 7210) 1 / 29 The AFT model framework Last time, we introduced the Weibull distribution
Accelerated Failure Time Models Patrick Breheny October 12 Patrick Breheny University of Iowa Survival Data Analysis (BIOS 7210) 1 / 29 The AFT model framework Last time, we introduced the Weibull distribution
Survival Regression Models
 Survival Regression Models David M. Rocke May 18, 2017 David M. Rocke Survival Regression Models May 18, 2017 1 / 32 Background on the Proportional Hazards Model The exponential distribution has constant
Survival Regression Models David M. Rocke May 18, 2017 David M. Rocke Survival Regression Models May 18, 2017 1 / 32 Background on the Proportional Hazards Model The exponential distribution has constant
Multistate Modeling and Applications
 Multistate Modeling and Applications Yang Yang Department of Statistics University of Michigan, Ann Arbor IBM Research Graduate Student Workshop: Statistics for a Smarter Planet Yang Yang (UM, Ann Arbor)
Multistate Modeling and Applications Yang Yang Department of Statistics University of Michigan, Ann Arbor IBM Research Graduate Student Workshop: Statistics for a Smarter Planet Yang Yang (UM, Ann Arbor)
Dynamic Prediction of Disease Progression Using Longitudinal Biomarker Data
 Dynamic Prediction of Disease Progression Using Longitudinal Biomarker Data Xuelin Huang Department of Biostatistics M. D. Anderson Cancer Center The University of Texas Joint Work with Jing Ning, Sangbum
Dynamic Prediction of Disease Progression Using Longitudinal Biomarker Data Xuelin Huang Department of Biostatistics M. D. Anderson Cancer Center The University of Texas Joint Work with Jing Ning, Sangbum
3003 Cure. F. P. Treasure
 3003 Cure F. P. reasure November 8, 2000 Peter reasure / November 8, 2000/ Cure / 3003 1 Cure A Simple Cure Model he Concept of Cure A cure model is a survival model where a fraction of the population
3003 Cure F. P. reasure November 8, 2000 Peter reasure / November 8, 2000/ Cure / 3003 1 Cure A Simple Cure Model he Concept of Cure A cure model is a survival model where a fraction of the population
Parametric Maximum Likelihood Estimation of Cure Fraction Using Interval-Censored Data
 Columbia International Publishing Journal of Advanced Computing (2013) 1: 43-58 doi:107726/jac20131004 Research Article Parametric Maximum Likelihood Estimation of Cure Fraction Using Interval-Censored
Columbia International Publishing Journal of Advanced Computing (2013) 1: 43-58 doi:107726/jac20131004 Research Article Parametric Maximum Likelihood Estimation of Cure Fraction Using Interval-Censored
Rank Regression with Normal Residuals using the Gibbs Sampler
 Rank Regression with Normal Residuals using the Gibbs Sampler Stephen P Smith email: hucklebird@aol.com, 2018 Abstract Yu (2000) described the use of the Gibbs sampler to estimate regression parameters
Rank Regression with Normal Residuals using the Gibbs Sampler Stephen P Smith email: hucklebird@aol.com, 2018 Abstract Yu (2000) described the use of the Gibbs sampler to estimate regression parameters
Semi-Competing Risks on A Trivariate Weibull Survival Model
 Semi-Competing Risks on A Trivariate Weibull Survival Model Cheng K. Lee Department of Targeting Modeling Insight & Innovation Marketing Division Wachovia Corporation Charlotte NC 28244 Jenq-Daw Lee Graduate
Semi-Competing Risks on A Trivariate Weibull Survival Model Cheng K. Lee Department of Targeting Modeling Insight & Innovation Marketing Division Wachovia Corporation Charlotte NC 28244 Jenq-Daw Lee Graduate
Multi-state Models: An Overview
 Multi-state Models: An Overview Andrew Titman Lancaster University 14 April 2016 Overview Introduction to multi-state modelling Examples of applications Continuously observed processes Intermittently observed
Multi-state Models: An Overview Andrew Titman Lancaster University 14 April 2016 Overview Introduction to multi-state modelling Examples of applications Continuously observed processes Intermittently observed
8. Parametric models in survival analysis General accelerated failure time models for parametric regression
 8. Parametric models in survival analysis 8.1. General accelerated failure time models for parametric regression The accelerated failure time model Let T be the time to event and x be a vector of covariates.
8. Parametric models in survival analysis 8.1. General accelerated failure time models for parametric regression The accelerated failure time model Let T be the time to event and x be a vector of covariates.
LSS: An S-Plus/R program for the accelerated failure time model to right censored data based on least-squares principle
 computer methods and programs in biomedicine 86 (2007) 45 50 journal homepage: www.intl.elsevierhealth.com/journals/cmpb LSS: An S-Plus/R program for the accelerated failure time model to right censored
computer methods and programs in biomedicine 86 (2007) 45 50 journal homepage: www.intl.elsevierhealth.com/journals/cmpb LSS: An S-Plus/R program for the accelerated failure time model to right censored
FAILURE-TIME WITH DELAYED ONSET
 REVSTAT Statistical Journal Volume 13 Number 3 November 2015 227 231 FAILURE-TIME WITH DELAYED ONSET Authors: Man Yu Wong Department of Mathematics Hong Kong University of Science and Technology Hong Kong
REVSTAT Statistical Journal Volume 13 Number 3 November 2015 227 231 FAILURE-TIME WITH DELAYED ONSET Authors: Man Yu Wong Department of Mathematics Hong Kong University of Science and Technology Hong Kong
REGRESSION ANALYSIS FOR TIME-TO-EVENT DATA THE PROPORTIONAL HAZARDS (COX) MODEL ST520
 REGRESSION ANALYSIS FOR TIME-TO-EVENT DATA THE PROPORTIONAL HAZARDS (COX) MODEL ST520 Department of Statistics North Carolina State University Presented by: Butch Tsiatis, Department of Statistics, NCSU
REGRESSION ANALYSIS FOR TIME-TO-EVENT DATA THE PROPORTIONAL HAZARDS (COX) MODEL ST520 Department of Statistics North Carolina State University Presented by: Butch Tsiatis, Department of Statistics, NCSU
Estimation for Modified Data
 Definition. Estimation for Modified Data 1. Empirical distribution for complete individual data (section 11.) An observation X is truncated from below ( left truncated) at d if when it is at or below d
Definition. Estimation for Modified Data 1. Empirical distribution for complete individual data (section 11.) An observation X is truncated from below ( left truncated) at d if when it is at or below d
Incorporating unobserved heterogeneity in Weibull survival models: A Bayesian approach
 Incorporating unobserved heterogeneity in Weibull survival models: A Bayesian approach Catalina A. Vallejos 1 Mark F.J. Steel 2 1 MRC Biostatistics Unit, EMBL-European Bioinformatics Institute. 2 Dept.
Incorporating unobserved heterogeneity in Weibull survival models: A Bayesian approach Catalina A. Vallejos 1 Mark F.J. Steel 2 1 MRC Biostatistics Unit, EMBL-European Bioinformatics Institute. 2 Dept.
Lecture 9. Statistics Survival Analysis. Presented February 23, Dan Gillen Department of Statistics University of California, Irvine
 Statistics 255 - Survival Analysis Presented February 23, 2016 Dan Gillen Department of Statistics University of California, Irvine 9.1 Survival analysis involves subjects moving through time Hazard may
Statistics 255 - Survival Analysis Presented February 23, 2016 Dan Gillen Department of Statistics University of California, Irvine 9.1 Survival analysis involves subjects moving through time Hazard may
Survival Analysis Math 434 Fall 2011
 Survival Analysis Math 434 Fall 2011 Part IV: Chap. 8,9.2,9.3,11: Semiparametric Proportional Hazards Regression Jimin Ding Math Dept. www.math.wustl.edu/ jmding/math434/fall09/index.html Basic Model Setup
Survival Analysis Math 434 Fall 2011 Part IV: Chap. 8,9.2,9.3,11: Semiparametric Proportional Hazards Regression Jimin Ding Math Dept. www.math.wustl.edu/ jmding/math434/fall09/index.html Basic Model Setup
Proportional hazards regression
 Proportional hazards regression Patrick Breheny October 8 Patrick Breheny Survival Data Analysis (BIOS 7210) 1/28 Introduction The model Solving for the MLE Inference Today we will begin discussing regression
Proportional hazards regression Patrick Breheny October 8 Patrick Breheny Survival Data Analysis (BIOS 7210) 1/28 Introduction The model Solving for the MLE Inference Today we will begin discussing regression
The coxvc_1-1-1 package
 Appendix A The coxvc_1-1-1 package A.1 Introduction The coxvc_1-1-1 package is a set of functions for survival analysis that run under R2.1.1 [81]. This package contains a set of routines to fit Cox models
Appendix A The coxvc_1-1-1 package A.1 Introduction The coxvc_1-1-1 package is a set of functions for survival analysis that run under R2.1.1 [81]. This package contains a set of routines to fit Cox models
Research & Reviews: Journal of Statistics and Mathematical Sciences
 Research & Reviews: Journal of Statistics and Mathematical Sciences Bayesian Estimation in Shared Compound Negative Binomial Frailty Models David D Hanagal * and Asmita T Kamble Department of Statistics,
Research & Reviews: Journal of Statistics and Mathematical Sciences Bayesian Estimation in Shared Compound Negative Binomial Frailty Models David D Hanagal * and Asmita T Kamble Department of Statistics,
Package GORCure. January 13, 2017
 Package GORCure January 13, 2017 Type Package Title Fit Generalized Odds Rate Mixture Cure Model with Interval Censored Data Version 2.0 Date 2017-01-12 Author Jie Zhou, Jiajia Zhang, Wenbin Lu Maintainer
Package GORCure January 13, 2017 Type Package Title Fit Generalized Odds Rate Mixture Cure Model with Interval Censored Data Version 2.0 Date 2017-01-12 Author Jie Zhou, Jiajia Zhang, Wenbin Lu Maintainer
Lecture 12. Multivariate Survival Data Statistics Survival Analysis. Presented March 8, 2016
 Statistics 255 - Survival Analysis Presented March 8, 2016 Dan Gillen Department of Statistics University of California, Irvine 12.1 Examples Clustered or correlated survival times Disease onset in family
Statistics 255 - Survival Analysis Presented March 8, 2016 Dan Gillen Department of Statistics University of California, Irvine 12.1 Examples Clustered or correlated survival times Disease onset in family
Probability Plots. Summary. Sample StatFolio: probplots.sgp
 STATGRAPHICS Rev. 9/6/3 Probability Plots Summary... Data Input... 2 Analysis Summary... 2 Analysis Options... 3 Uniform Plot... 3 Normal Plot... 4 Lognormal Plot... 4 Weibull Plot... Extreme Value Plot...
STATGRAPHICS Rev. 9/6/3 Probability Plots Summary... Data Input... 2 Analysis Summary... 2 Analysis Options... 3 Uniform Plot... 3 Normal Plot... 4 Lognormal Plot... 4 Weibull Plot... Extreme Value Plot...
Package ICGOR. January 13, 2017
 Package ICGOR January 13, 2017 Type Package Title Fit Generalized Odds Rate Hazards Model with Interval Censored Data Version 2.0 Date 2017-01-12 Author Jie Zhou, Jiajia Zhang, Wenbin Lu Maintainer Jie
Package ICGOR January 13, 2017 Type Package Title Fit Generalized Odds Rate Hazards Model with Interval Censored Data Version 2.0 Date 2017-01-12 Author Jie Zhou, Jiajia Zhang, Wenbin Lu Maintainer Jie
Optimal Treatment Regimes for Survival Endpoints from a Classification Perspective. Anastasios (Butch) Tsiatis and Xiaofei Bai
 Optimal Treatment Regimes for Survival Endpoints from a Classification Perspective Anastasios (Butch) Tsiatis and Xiaofei Bai Department of Statistics North Carolina State University 1/35 Optimal Treatment
Optimal Treatment Regimes for Survival Endpoints from a Classification Perspective Anastasios (Butch) Tsiatis and Xiaofei Bai Department of Statistics North Carolina State University 1/35 Optimal Treatment
Bayesian Graphical Models
 Graphical Models and Inference, Lecture 16, Michaelmas Term 2009 December 4, 2009 Parameter θ, data X = x, likelihood L(θ x) p(x θ). Express knowledge about θ through prior distribution π on θ. Inference
Graphical Models and Inference, Lecture 16, Michaelmas Term 2009 December 4, 2009 Parameter θ, data X = x, likelihood L(θ x) p(x θ). Express knowledge about θ through prior distribution π on θ. Inference
Joint Modeling of Longitudinal Item Response Data and Survival
 Joint Modeling of Longitudinal Item Response Data and Survival Jean-Paul Fox University of Twente Department of Research Methodology, Measurement and Data Analysis Faculty of Behavioural Sciences Enschede,
Joint Modeling of Longitudinal Item Response Data and Survival Jean-Paul Fox University of Twente Department of Research Methodology, Measurement and Data Analysis Faculty of Behavioural Sciences Enschede,
Bayesian semiparametric analysis of short- and long- term hazard ratios with covariates
 Bayesian semiparametric analysis of short- and long- term hazard ratios with covariates Department of Statistics, ITAM, Mexico 9th Conference on Bayesian nonparametrics Amsterdam, June 10-14, 2013 Contents
Bayesian semiparametric analysis of short- and long- term hazard ratios with covariates Department of Statistics, ITAM, Mexico 9th Conference on Bayesian nonparametrics Amsterdam, June 10-14, 2013 Contents
STAT 331. Accelerated Failure Time Models. Previously, we have focused on multiplicative intensity models, where
 STAT 331 Accelerated Failure Time Models Previously, we have focused on multiplicative intensity models, where h t z) = h 0 t) g z). These can also be expressed as H t z) = H 0 t) g z) or S t z) = e Ht
STAT 331 Accelerated Failure Time Models Previously, we have focused on multiplicative intensity models, where h t z) = h 0 t) g z). These can also be expressed as H t z) = H 0 t) g z) or S t z) = e Ht
Bayesian Nonparametric Accelerated Failure Time Models for Analyzing Heterogeneous Treatment Effects
 Bayesian Nonparametric Accelerated Failure Time Models for Analyzing Heterogeneous Treatment Effects Nicholas C. Henderson Thomas A. Louis Gary Rosner Ravi Varadhan Johns Hopkins University September 28,
Bayesian Nonparametric Accelerated Failure Time Models for Analyzing Heterogeneous Treatment Effects Nicholas C. Henderson Thomas A. Louis Gary Rosner Ravi Varadhan Johns Hopkins University September 28,
Kernel density estimation in R
 Kernel density estimation in R Kernel density estimation can be done in R using the density() function in R. The default is a Guassian kernel, but others are possible also. It uses it s own algorithm to
Kernel density estimation in R Kernel density estimation can be done in R using the density() function in R. The default is a Guassian kernel, but others are possible also. It uses it s own algorithm to
[Part 2] Model Development for the Prediction of Survival Times using Longitudinal Measurements
![[Part 2] Model Development for the Prediction of Survival Times using Longitudinal Measurements [Part 2] Model Development for the Prediction of Survival Times using Longitudinal Measurements](/thumbs/89/99255856.jpg) [Part 2] Model Development for the Prediction of Survival Times using Longitudinal Measurements Aasthaa Bansal PhD Pharmaceutical Outcomes Research & Policy Program University of Washington 69 Biomarkers
[Part 2] Model Development for the Prediction of Survival Times using Longitudinal Measurements Aasthaa Bansal PhD Pharmaceutical Outcomes Research & Policy Program University of Washington 69 Biomarkers
Statistical Inference and Methods
 Department of Mathematics Imperial College London d.stephens@imperial.ac.uk http://stats.ma.ic.ac.uk/ das01/ 31st January 2006 Part VI Session 6: Filtering and Time to Event Data Session 6: Filtering and
Department of Mathematics Imperial College London d.stephens@imperial.ac.uk http://stats.ma.ic.ac.uk/ das01/ 31st January 2006 Part VI Session 6: Filtering and Time to Event Data Session 6: Filtering and
A multistate additive relative survival semi-markov model
 46 e Journées de Statistique de la SFdS A multistate additive semi-markov Florence Gillaizeau,, & Etienne Dantan & Magali Giral, & Yohann Foucher, E-mail: florence.gillaizeau@univ-nantes.fr EA 4275 - SPHERE
46 e Journées de Statistique de la SFdS A multistate additive semi-markov Florence Gillaizeau,, & Etienne Dantan & Magali Giral, & Yohann Foucher, E-mail: florence.gillaizeau@univ-nantes.fr EA 4275 - SPHERE
The Weibull Distribution
 The Weibull Distribution Patrick Breheny October 10 Patrick Breheny University of Iowa Survival Data Analysis (BIOS 7210) 1 / 19 Introduction Today we will introduce an important generalization of the
The Weibull Distribution Patrick Breheny October 10 Patrick Breheny University of Iowa Survival Data Analysis (BIOS 7210) 1 / 19 Introduction Today we will introduce an important generalization of the
A Dirichlet Process Mixture Model for Survival Outcome Data: Assessing Nationwide Kidney Transplant Centers
 Research Article Received XXXX (www.interscience.wiley.com) DOI:.02/sim.0000 A Dirichlet Process Mixture Model for Survival Outcome Data: Assessing Nationwide Kidney Transplant s Lili Zhao a, Jingchunzi
Research Article Received XXXX (www.interscience.wiley.com) DOI:.02/sim.0000 A Dirichlet Process Mixture Model for Survival Outcome Data: Assessing Nationwide Kidney Transplant s Lili Zhao a, Jingchunzi
Chapter 6. Logistic Regression. 6.1 A linear model for the log odds
 Chapter 6 Logistic Regression In logistic regression, there is a categorical response variables, often coded 1=Yes and 0=No. Many important phenomena fit this framework. The patient survives the operation,
Chapter 6 Logistic Regression In logistic regression, there is a categorical response variables, often coded 1=Yes and 0=No. Many important phenomena fit this framework. The patient survives the operation,
Luke B Smith and Brian J Reich North Carolina State University May 21, 2013
 BSquare: An R package for Bayesian simultaneous quantile regression Luke B Smith and Brian J Reich North Carolina State University May 21, 2013 BSquare in an R package to conduct Bayesian quantile regression
BSquare: An R package for Bayesian simultaneous quantile regression Luke B Smith and Brian J Reich North Carolina State University May 21, 2013 BSquare in an R package to conduct Bayesian quantile regression
The Design of a Survival Study
 The Design of a Survival Study The design of survival studies are usually based on the logrank test, and sometimes assumes the exponential distribution. As in standard designs, the power depends on The
The Design of a Survival Study The design of survival studies are usually based on the logrank test, and sometimes assumes the exponential distribution. As in standard designs, the power depends on The
for Time-to-event Data Mei-Ling Ting Lee University of Maryland, College Park
 Threshold Regression for Time-to-event Data Mei-Ling Ting Lee University of Maryland, College Park MLTLEE@UMD.EDU Outline The proportional hazards (PH) model is widely used in analyzing time-to-event data.
Threshold Regression for Time-to-event Data Mei-Ling Ting Lee University of Maryland, College Park MLTLEE@UMD.EDU Outline The proportional hazards (PH) model is widely used in analyzing time-to-event data.
Multi-state models: prediction
 Department of Medical Statistics and Bioinformatics Leiden University Medical Center Course on advanced survival analysis, Copenhagen Outline Prediction Theory Aalen-Johansen Computational aspects Applications
Department of Medical Statistics and Bioinformatics Leiden University Medical Center Course on advanced survival analysis, Copenhagen Outline Prediction Theory Aalen-Johansen Computational aspects Applications
HOW TO USE SURVIVAL FORESTS (SFPDV1)
 HOW TO USE SURVIVAL FORESTS (SFPDV1) This program in f77 is an unorthodox approach to survival analysis. it allows the analyst to view the importance of the covariates as the experiment evolves over. It
HOW TO USE SURVIVAL FORESTS (SFPDV1) This program in f77 is an unorthodox approach to survival analysis. it allows the analyst to view the importance of the covariates as the experiment evolves over. It
Adaptive Prediction of Event Times in Clinical Trials
 Adaptive Prediction of Event Times in Clinical Trials Yu Lan Southern Methodist University Advisor: Daniel F. Heitjan May 8, 2017 Yu Lan (SMU) May 8, 2017 1 / 19 Clinical Trial Prediction Event-based trials:
Adaptive Prediction of Event Times in Clinical Trials Yu Lan Southern Methodist University Advisor: Daniel F. Heitjan May 8, 2017 Yu Lan (SMU) May 8, 2017 1 / 19 Clinical Trial Prediction Event-based trials:
Why Bayesian approaches? The average height of a rare plant
 Why Bayesian approaches? The average height of a rare plant Estimation and comparison of averages is an important step in many ecological analyses and demographic models. In this demonstration you will
Why Bayesian approaches? The average height of a rare plant Estimation and comparison of averages is an important step in many ecological analyses and demographic models. In this demonstration you will
Analysis of Time-to-Event Data: Chapter 6 - Regression diagnostics
 Analysis of Time-to-Event Data: Chapter 6 - Regression diagnostics Steffen Unkel Department of Medical Statistics University Medical Center Göttingen, Germany Winter term 2018/19 1/25 Residuals for the
Analysis of Time-to-Event Data: Chapter 6 - Regression diagnostics Steffen Unkel Department of Medical Statistics University Medical Center Göttingen, Germany Winter term 2018/19 1/25 Residuals for the
Estimating Causal Effects of Organ Transplantation Treatment Regimes
 Estimating Causal Effects of Organ Transplantation Treatment Regimes David M. Vock, Jeffrey A. Verdoliva Boatman Division of Biostatistics University of Minnesota July 31, 2018 1 / 27 Hot off the Press
Estimating Causal Effects of Organ Transplantation Treatment Regimes David M. Vock, Jeffrey A. Verdoliva Boatman Division of Biostatistics University of Minnesota July 31, 2018 1 / 27 Hot off the Press
ST495: Survival Analysis: Maximum likelihood
 ST495: Survival Analysis: Maximum likelihood Eric B. Laber Department of Statistics, North Carolina State University February 11, 2014 Everything is deception: seeking the minimum of illusion, keeping
ST495: Survival Analysis: Maximum likelihood Eric B. Laber Department of Statistics, North Carolina State University February 11, 2014 Everything is deception: seeking the minimum of illusion, keeping
Longitudinal + Reliability = Joint Modeling
 Longitudinal + Reliability = Joint Modeling Carles Serrat Institute of Statistics and Mathematics Applied to Building CYTED-HAROSA International Workshop November 21-22, 2013 Barcelona Mainly from Rizopoulos,
Longitudinal + Reliability = Joint Modeling Carles Serrat Institute of Statistics and Mathematics Applied to Building CYTED-HAROSA International Workshop November 21-22, 2013 Barcelona Mainly from Rizopoulos,
Lecture 7. Proportional Hazards Model - Handling Ties and Survival Estimation Statistics Survival Analysis. Presented February 4, 2016
 Proportional Hazards Model - Handling Ties and Survival Estimation Statistics 255 - Survival Analysis Presented February 4, 2016 likelihood - Discrete Dan Gillen Department of Statistics University of
Proportional Hazards Model - Handling Ties and Survival Estimation Statistics 255 - Survival Analysis Presented February 4, 2016 likelihood - Discrete Dan Gillen Department of Statistics University of
PENALIZED LIKELIHOOD PARAMETER ESTIMATION FOR ADDITIVE HAZARD MODELS WITH INTERVAL CENSORED DATA
 PENALIZED LIKELIHOOD PARAMETER ESTIMATION FOR ADDITIVE HAZARD MODELS WITH INTERVAL CENSORED DATA Kasun Rathnayake ; A/Prof Jun Ma Department of Statistics Faculty of Science and Engineering Macquarie University
PENALIZED LIKELIHOOD PARAMETER ESTIMATION FOR ADDITIVE HAZARD MODELS WITH INTERVAL CENSORED DATA Kasun Rathnayake ; A/Prof Jun Ma Department of Statistics Faculty of Science and Engineering Macquarie University
The influence of categorising survival time on parameter estimates in a Cox model
 The influence of categorising survival time on parameter estimates in a Cox model Anika Buchholz 1,2, Willi Sauerbrei 2, Patrick Royston 3 1 Freiburger Zentrum für Datenanalyse und Modellbildung, Albert-Ludwigs-Universität
The influence of categorising survival time on parameter estimates in a Cox model Anika Buchholz 1,2, Willi Sauerbrei 2, Patrick Royston 3 1 Freiburger Zentrum für Datenanalyse und Modellbildung, Albert-Ludwigs-Universität
CURE FRACTION MODELS USING MIXTURE AND NON-MIXTURE MODELS. 1. Introduction
 ØÑ Å Ø Ñ Ø Ð ÈÙ Ð Ø ÓÒ DOI: 10.2478/v10127-012-0001-4 Tatra Mt. Math. Publ. 51 (2012), 1 9 CURE FRACTION MODELS USING MIXTURE AND NON-MIXTURE MODELS Jorge A. Achcar Emílio A. Coelho-Barros Josmar Mazucheli
ØÑ Å Ø Ñ Ø Ð ÈÙ Ð Ø ÓÒ DOI: 10.2478/v10127-012-0001-4 Tatra Mt. Math. Publ. 51 (2012), 1 9 CURE FRACTION MODELS USING MIXTURE AND NON-MIXTURE MODELS Jorge A. Achcar Emílio A. Coelho-Barros Josmar Mazucheli
4. Comparison of Two (K) Samples
 4. Comparison of Two (K) Samples K=2 Problem: compare the survival distributions between two groups. E: comparing treatments on patients with a particular disease. Z: Treatment indicator, i.e. Z = 1 for
4. Comparison of Two (K) Samples K=2 Problem: compare the survival distributions between two groups. E: comparing treatments on patients with a particular disease. Z: Treatment indicator, i.e. Z = 1 for
UNIVERSITY OF CALIFORNIA, SAN DIEGO
 UNIVERSITY OF CALIFORNIA, SAN DIEGO Estimation of the primary hazard ratio in the presence of a secondary covariate with non-proportional hazards An undergraduate honors thesis submitted to the Department
UNIVERSITY OF CALIFORNIA, SAN DIEGO Estimation of the primary hazard ratio in the presence of a secondary covariate with non-proportional hazards An undergraduate honors thesis submitted to the Department
Step-Stress Models and Associated Inference
 Department of Mathematics & Statistics Indian Institute of Technology Kanpur August 19, 2014 Outline Accelerated Life Test 1 Accelerated Life Test 2 3 4 5 6 7 Outline Accelerated Life Test 1 Accelerated
Department of Mathematics & Statistics Indian Institute of Technology Kanpur August 19, 2014 Outline Accelerated Life Test 1 Accelerated Life Test 2 3 4 5 6 7 Outline Accelerated Life Test 1 Accelerated
Practice Exam 1. (A) (B) (C) (D) (E) You are given the following data on loss sizes:
 Practice Exam 1 1. Losses for an insurance coverage have the following cumulative distribution function: F(0) = 0 F(1,000) = 0.2 F(5,000) = 0.4 F(10,000) = 0.9 F(100,000) = 1 with linear interpolation
Practice Exam 1 1. Losses for an insurance coverage have the following cumulative distribution function: F(0) = 0 F(1,000) = 0.2 F(5,000) = 0.4 F(10,000) = 0.9 F(100,000) = 1 with linear interpolation
Frailty Modeling for clustered survival data: a simulation study
 Frailty Modeling for clustered survival data: a simulation study IAA Oslo 2015 Souad ROMDHANE LaREMFiQ - IHEC University of Sousse (Tunisia) souad_romdhane@yahoo.fr Lotfi BELKACEM LaREMFiQ - IHEC University
Frailty Modeling for clustered survival data: a simulation study IAA Oslo 2015 Souad ROMDHANE LaREMFiQ - IHEC University of Sousse (Tunisia) souad_romdhane@yahoo.fr Lotfi BELKACEM LaREMFiQ - IHEC University
Package CoxRidge. February 27, 2015
 Type Package Title Cox Models with Dynamic Ridge Penalties Version 0.9.2 Date 2015-02-12 Package CoxRidge February 27, 2015 Author Aris Perperoglou Maintainer Aris Perperoglou
Type Package Title Cox Models with Dynamic Ridge Penalties Version 0.9.2 Date 2015-02-12 Package CoxRidge February 27, 2015 Author Aris Perperoglou Maintainer Aris Perperoglou
36-463/663Multilevel and Hierarchical Models
 36-463/663Multilevel and Hierarchical Models From Bayes to MCMC to MLMs Brian Junker 132E Baker Hall brian@stat.cmu.edu 1 Outline Bayesian Statistics and MCMC Distribution of Skill Mastery in a Population
36-463/663Multilevel and Hierarchical Models From Bayes to MCMC to MLMs Brian Junker 132E Baker Hall brian@stat.cmu.edu 1 Outline Bayesian Statistics and MCMC Distribution of Skill Mastery in a Population
Residuals and model diagnostics
 Residuals and model diagnostics Patrick Breheny November 10 Patrick Breheny Survival Data Analysis (BIOS 7210) 1/42 Introduction Residuals Many assumptions go into regression models, and the Cox proportional
Residuals and model diagnostics Patrick Breheny November 10 Patrick Breheny Survival Data Analysis (BIOS 7210) 1/42 Introduction Residuals Many assumptions go into regression models, and the Cox proportional
Survival Analysis for Interval Censored Data - Nonparametric Estimation
 Survival Analysis for Interval Censored Data - Nonparametric Estimation Seminar of Statistics, ETHZ Group 8, 02. May 2011 Martina Albers, Nanina Anderegg, Urs Müller Overview Examples Nonparametric MLE
Survival Analysis for Interval Censored Data - Nonparametric Estimation Seminar of Statistics, ETHZ Group 8, 02. May 2011 Martina Albers, Nanina Anderegg, Urs Müller Overview Examples Nonparametric MLE
Bayesian Inference on Joint Mixture Models for Survival-Longitudinal Data with Multiple Features. Yangxin Huang
 Bayesian Inference on Joint Mixture Models for Survival-Longitudinal Data with Multiple Features Yangxin Huang Department of Epidemiology and Biostatistics, COPH, USF, Tampa, FL yhuang@health.usf.edu January
Bayesian Inference on Joint Mixture Models for Survival-Longitudinal Data with Multiple Features Yangxin Huang Department of Epidemiology and Biostatistics, COPH, USF, Tampa, FL yhuang@health.usf.edu January
β j = coefficient of x j in the model; β = ( β1, β2,
 Regression Modeling of Survival Time Data Why regression models? Groups similar except for the treatment under study use the nonparametric methods discussed earlier. Groups differ in variables (covariates)
Regression Modeling of Survival Time Data Why regression models? Groups similar except for the treatment under study use the nonparametric methods discussed earlier. Groups differ in variables (covariates)
How To Do Piecewise Exponential Survival Analysis in Stata 7 (Allison 1995:Output 4.20) revised
 WM Mason, Soc 213B, S 02, UCLA Page 1 of 15 How To Do Piecewise Exponential Survival Analysis in Stata 7 (Allison 1995:Output 420) revised 4-25-02 This document can function as a "how to" for setting up
WM Mason, Soc 213B, S 02, UCLA Page 1 of 15 How To Do Piecewise Exponential Survival Analysis in Stata 7 (Allison 1995:Output 420) revised 4-25-02 This document can function as a "how to" for setting up
Some Statistical Properties of Exponentiated Weighted Weibull Distribution
 Global Journal of Science Frontier Research: F Mathematics and Decision Sciences Volume 4 Issue 2 Version. Year 24 Type : Double Blind Peer Reviewed International Research Journal Publisher: Global Journals
Global Journal of Science Frontier Research: F Mathematics and Decision Sciences Volume 4 Issue 2 Version. Year 24 Type : Double Blind Peer Reviewed International Research Journal Publisher: Global Journals
Likelihood Construction, Inference for Parametric Survival Distributions
 Week 1 Likelihood Construction, Inference for Parametric Survival Distributions In this section we obtain the likelihood function for noninformatively rightcensored survival data and indicate how to make
Week 1 Likelihood Construction, Inference for Parametric Survival Distributions In this section we obtain the likelihood function for noninformatively rightcensored survival data and indicate how to make
ST5212: Survival Analysis
 ST51: Survival Analysis 8/9: Semester II Tutorial 1. A model for lifetimes, with a bathtub-shaped hazard rate, is the exponential power distribution with survival fumction S(x) =exp{1 exp[(λx) α ]}. (a)
ST51: Survival Analysis 8/9: Semester II Tutorial 1. A model for lifetimes, with a bathtub-shaped hazard rate, is the exponential power distribution with survival fumction S(x) =exp{1 exp[(λx) α ]}. (a)
Hybrid Censoring Scheme: An Introduction
 Department of Mathematics & Statistics Indian Institute of Technology Kanpur August 19, 2014 Outline 1 2 3 4 5 Outline 1 2 3 4 5 What is? Lifetime data analysis is used to analyze data in which the time
Department of Mathematics & Statistics Indian Institute of Technology Kanpur August 19, 2014 Outline 1 2 3 4 5 Outline 1 2 3 4 5 What is? Lifetime data analysis is used to analyze data in which the time
