Tensor Biclustering. Hamid Javadi Stanford University Abstract
|
|
|
- Alison Holt
- 5 years ago
- Views:
Transcription
1 Tensor Biclustering Soheil Feizi Stanford University Hamid Javadi Stanford University David Tse Stanford University Abstract Consider a dataset where data is collected on multiple features of multiple individuals over multiple times. This type of data can be represented as a three dimensional individual/feature/time tensor and has become increasingly prominent in various areas of science. The tensor biclustering problem computes a subset of individuals and a subset of features whose signal trajectories over time lie in a low-dimensional subspace, modeling similarity among the signal trajectories while allowing different scalings across different individuals or different features. We study the information-theoretic limit of this problem under a generative model. Moreover, we propose an efficient spectral algorithm to solve the tensor biclustering problem and analyze its achievability bound in an asymptotic regime. Finally, we show the efficiency of our proposed method in several synthetic and real datasets. 1 Introduction Let T R n1 n be a data matrix whose rows and columns represent individuals and features, respectively. Given T, the matrix biclustering problem aims to find a subset of individuals (i.e., J 1 {1,,..., n 1 }) which exhibit similar values across a subset of features (i.e., J {1,,..., n }) (Figure 1-a). The matrix biclustering problem has been studied extensively in machine learning and statistics and is closely related to problems of sub-matrix localization, planted clique and community detection [1,, 3]. In modern datasets, however, instead of collecting data on every individual-feature pair at a single time, we may collect data at multiple times. One can visualize a trajectory over time for each individual-feature pair. This type of datasets has become increasingly prominent in different areas of science. For example, the roadmap epigenomics dataset [4] provides multiple histon modification marks for genome-tissue pairs, the genotype-tissue expression dataset [5] provides expression data on multiple genes for individual-tissue pairs, while there have been recent efforts to collect various omics data in individuals at different times [6]. Suppose we have n 1 individuals, n features, and we collect data for every individual-feature pair at m different times. This data can be represented as a three dimensional tensor T R n1 n m (Figure 1-b). The tensor biclustering problem aims to compute a subset of individuals and a subset of features whose trajectories are highly similar. Similarity is modeled as the trajectories as lying in a low-dimensional (say one-dimensional) subspace (Figure 1-d). This definition allows different scalings across different individuals or different features, and is important in many applications such as in omics datasets [6] because individual-feature trajectories often have their own intrinsic scalings. In particular, at each time the individual-feature data matrix may not exhibit a matrix bicluster separately. This means that repeated applications of matrix biclustering cannot solve the tensor biclustering problem. Moreover, owing to the same reason, trajectories in a bicluster can have large distances among themselves (Figure 1-d). Thus, a distance-based clustering of signal trajectories is likely to fail as well. 31st Conference on Neural Information Processing Systems (NIPS 17), Long Beach, CA, USA.
2 (a) Matrix Biclustering (b) Our Model Tensor Biclustering (c) Tensor Triclustering individuals times times features features individuals features individuals (d) All Trajectories Trajectories in the Bicluster Tensor Biclustering Figure 1: (a) The matrix biclustering problem. (b) The tensor biclustering problem. (c) The tensor triclustering problem. (d) A visualization of a bicluster in a three dimensional tensor. Trajectories in the bicluster (red points) form a low dimensional subspace. This problem formulation has two main differences with tensor triclustering, which is a natural generalization of matrix biclustering to a three dimensional tensor (Figure 1-c). Firstly, unlike tensor triclustering, tensor biclustering has an asymmetric structure along tensor dimensions inspired by aforementioned applications. That is, since a tensor bicluster is defined as a subset of individuals and a subset of features with similar trajectories, the third dimension of the tensor (i.e., the time dimension) plays a different role compared to the other two dimensions. This is in contrast with tensor triclustering where there is not such a difference between roles of tensor dimensions in defining the cluster. Secondly, in tensor biclustering, the notion of a cluster is defined regarding to trajectories lying in a low-dimensional subspace while in tensor triclustering, a cluster is defined as a sub-cube with similar entries. Finding statistically significant patterns in multi-dimensional data tensors has been studied in dimensionality reduction [7, 8, 9, 1, 11, 1, 13, 14], topic modeling [15, 16, 17], among others. One related model is the spiked tensor model [7]. Unlike the tensor biclustering model that is asymmetric along tensor dimensions, the spiked tensor model has a symmetric structure. Computational and statistical limits for the spiked tensor model have been studied in [8, 9, 1, 14], among others. For more details, see Supplementary Materials (SM) Section 1.3. In this paper, we study information-theoretic and computational limits for the tensor biclustering problem under a statistical model described in Section. From a computational perspective, we present four polynomial time methods and analyze their asymptotic achievability bounds. In particular, one of our proposed methods, namely tensor folding+spectral, outperforms other methods both theoretically (under realistic model parameters) and numerically in several synthetic and real data experiments. Moreover, we characterize a fundamental limit under which no algorithm can solve the tensor biclustering problem reliably in a minimax sense. We show that above this limit, a maximum likelihood estimator (MLE) which has an exponential computational complexity can solve this problem with vanishing error probability. 1.1 Notation We use T, X, and Z to represent input, signal, and noise tensors, respectively. For any set J, J denotes its cardinality. [n] represents the set {1,,..., n}. J = [n] J. x = (x t x) 1/ is the second norm of the vector x. x y is the Kronecker product of two vectors x and y. The asymptotic notation a(n) = O(b(n)) means that, there exists a universal constant c such that for sufficiently
3 large n, we have a(n) < cb(n). If there exists c > such that a(n) = O(b(n) log(n) c ), we use the notation a(n) = Õ(b(n)). The asymptotic notation a(n) = Ω(b(n)) and a(n) = Ω(b(n)) is the same as b(n) = O(a(n)) and b(n) = Õ(a(n)), respectively. Moreover, we write a(n) = Θ(b(n)) iff a(n) = Ω(b(n)) and b(n) = Ω(a(n)). Similarly, we write a(n) = Θ(b(n)) iff a(n) = Ω(b(n)) and b(n) = Ω(a(n)). Problem Formulation Let T = X + Z where X is the signal tensor and Z is the noise tensor. Consider q T = X + Z = σ r u r (J1) w r (J) v r + Z, (1) r=1 where u (J1) r and w r (J) have zero entries outside of J 1 and J index sets, respectively. We assume σ 1 σ... σ q >. Under this model, trajectories X (J 1, J, :) form an at most q dimensional subspace. We assume q min(m, J 1 J ). Definition 1 (Tensor Biclustering). The problem of tensor biclustering aims to compute bicluster index sets (J 1, J ) given T according to (1). In this paper, we make the following simplifying assumptions: we assume q = 1, n = n 1 = n, and k = J 1 = J. To simplify notation, we drop superscripts (J 1 ) and (J ) from u (J1) 1 and w (J) 1, respectively. Without loss of generality, we normalize signal vectors such that u 1 = w 1 = v 1 = 1. Moreover, we assume that for every (j 1, j ) J 1 J, u 1 (j 1 ) c and w 1 (j ) c, where c is a constant. Under these assumptions, a signal trajectory can be written as X (j 1, j, :) = u 1 (j 1 )w 1 (j )v 1. The scaling of this trajectory depends on row and column specific parameters u 1 (j 1 ) and w 1 (j ). Note that our analysis can be extended naturally to a more general setup of having multiple embedded biclusters with q > 1. We discuss this in Section 7. Next we describe the noise model. If (j 1, j ) / J 1 J, we assume that entries of the noise trajectory Z(j 1, j, :) are i.i.d. and each entry has a standard normal distribution. If (j 1, j ) J 1 J, we assume that entries of Z(j 1, j, :) are i.i.d. and each entry has a Gaussian distribution with zero mean and σ z variance. We analyze the tensor biclustering problem under two noise models for σ z: - Noise Model I: In this model, we assume σ z = 1, i.e., the variance of the noise within and outside of the bicluster is assumed to be the same. This is the noise model often considered in analysis of sub-matrix localization [, 3] and tensor PCA [7, 8, 9, 1, 11, 1, 14]. Although this model simplifies the analysis, it has the following drawback: under this noise model, for every value of σ 1, the average trajectory lengths in the bicluster is larger than the average trajectory lengths outside of the bicluster. See SM Section 1. for more details. - Noise Model II: In this model, we assume σ z = max(, 1 σ 1 mk ), i.e., σ z is modeled to minimize the difference between the average trajectory lengths within and outside of the bicluster. If σ 1 < mk, noise is added to make the average trajectory lengths within and outside of the bicluster comparable. See SM Section 1. for more details. 3 Computational Limits of the Tensor Biclustering Problem 3.1 Tensor Folding+Spectral Recall the formulation of the tensor biclustering problem (1). Let T (j1,1) T (j 1, :, :) and T (j,) T (:, j, :), () be horizontal (the first mode) and lateral (the second mode) matrix slices of the tensor T, respectively. One way to learn the embedded bicluster in the tensor is to compute row and column indices whose trajectories are highly correlated with each other. To do that, we compute n n C 1 T t (j T,) (j,) and C T t (j T 1,1) (j 1,1). (3) j =1 j 1=1 3
4 T T(1, :, : ) T(k, :, : ) T(k+1, :, : ) T(n 1, :, : ) Input Tensor m Matrix Slices m n n 1 n t T(1, :, : ) T(1, :, : ) t T(n 1, :, : ) T(n 1, :, : ) Bicluster Index Set (J ) Spectral Decomposition n n Combined Covariance n n Figure : A visualization of the tensor folding+spectral algorithm 1 to compute the bicluster index set J. The bicluster index set J 1 can be computed similarly. Algorithm 1 Tensor Folding+Spectral Input: T, k Compute û 1, the top eigenvector of C 1 Compute ŵ 1, the top eigenvector of C Compute Ĵ1, indices of the k largest values of ŵ 1 Compute Ĵ, indices of the k largest values of û 1 Output: Ĵ1 and Ĵ C 1 represents a combined covariance matrix along the tensor columns (Figure ). We refer to it as the folded tensor over the columns. If there was no noise, this matrix would be equal to σ 1u 1 u t 1. Thus, its eigenvector corresponding to the largest eigenvalue would be equal to u 1. On the other hand, we have u 1 (j 1 ) = if j 1 / J 1 and u 1 (j 1 ) >, otherwise. Therefore, selecting k indices of the top eigenvector with largest magnitudes would recover the index set J 1. However, with added noise, the top eigenvector of the folded tensor would be a perturbed version of u 1. Nevertheless one can estimate J 1 similarly (Algorithm 1). A similar argument holds for C. Theorem 1. Let û 1 and ŵ 1 be top eigenvectors of C 1 and C, respectively. Under both noise models I and II, - for m < Õ( n), if σ 1 = Ω(n), - for m = Ω( n), if σ 1 = Ω( n max(n, m)), as n, with high probability, we have û 1 (j 1 ) > û 1 (j 1) and ŵ 1 (j ) > ŵ 1 (j ) for every j 1 J 1, j 1 J 1, j J and j J. In the proof of Theorem 1, following the result of [18] for a Wigner noise matrix, we have proved an l version of the Davis-Kahan Lemma for a Wishart noise matrix. This lemma can be of independent interest for the readers. 3. Tensor Unfolding+Spectral Let T unfolded R m n be the unfolded tensor T such that T unfolded (:, (j 1 1)n + j ) = T (j 1, j, :) for 1 j 1, j n. Without noise, the right singular vector of this matrix is u 1 w 1 which corresponds to the singular value σ 1. Therefore, selecting k indices of this singular vector with largest magnitudes would recover the index set J 1 J. With added noise, however, the top singular vector of the unfolded tensor will be perturbed. Nevertheless one can estimate J 1 J similarly (SM Section ). 4
5 Theorem. Let ˆx be the top right singular vector of T unfolded. Under both noise models I and II, if σ 1 = Ω(max(n, m)), as n, with high probability, we have ˆx(j ) < ˆx(j) for every j in the bicluster and j outside of the bicluster. 3.3 Thresholding Sum of Squared and Individual Trajectory Lengths If the average trajectory lengths in the bicluster is larger than the one outside of the bicluster, methods based on trajectory length statistics can be successful in solving the tensor biclustering problem. One such method is thresholding individual trajectory lengths. In this method, we select k indices (j 1, j ) with the largest trajectory length T (j 1, j, :) (SM Section ). Theorem 3. As n, with high probability, Ĵ1 = J 1 and Ĵ = J - if σ 1 = Ω( mk ), under noise model I. - if σ 1 = Ω(mk ), under noise model II. Another method to solve the tensor biclustering problem is thresholding sum of squared trajectory lengths. In this method, we select k row indices with the largest sum of squared trajectory lengths along the columns as an estimation of J 1. We estimate J similarly (SM Section ). Theorem 4. As n, with high probability, Ĵ1 = J 1 and Ĵ = J - if σ 1 = Ω(k nm), under noise model I. - if σ 1 = Ω(mk + k nm), under noise model II. 4 Statistical (Information-Theoretic) Limits of the Tensor Biclustering Problem 4.1 Coherent Case In this section, we study a statistical (information theoretic) boundary for the tensor biclustering problem under the following statistical model: We assume u 1 (j 1 ) = 1/ k for j 1 J 1. Similarly, we assume w 1 (j ) = 1/ k for j J. Moreover, we assume v 1 is a fixed given vector with v 1 = 1. In the next section, we consider a non-coherent model where v 1 is random and unknown. Let T be an observed tensor from the tensor biclustering model (J 1, J ). Let J all be the set of all possible (J 1, J ). Thus, J all = ( n. k) A maximum likelihood estimator (MLE) for the tensor biclustering problem can be written as: max Ĵ J all v t 1 (j 1,j ) Ĵ1 Ĵ T (j 1, j, :) (Ĵ1, Ĵ) J all. k(1 σ z) σ 1 (j 1,j ) Ĵ1 Ĵ T (j 1, j, :) (4) Note that under the noise model I, the second term is zero. To solve this optimization, one needs to compute the likelihood function for ( n k) possible bicluster indices. Thus, the computational complexity of the MLE is exponential in n. Theorem 5. Under noise model I, if σ 1 = Ω(k), as n, with high probability, (J 1, J ) is the optimal solution of optimization (4). A similar result holds under noise model II if mk = Ω(log(n/k)). Next, we establish an upper bound on σ 1 under which no computational method can solve the tensor biclustering problem with vanishing probability of error. This upper bound indeed matches with the MLE achievability bound of Theorem 5 indicating its tightness. Theorem 6. Let T be an observed tensor from the tensor biclustering model with bicluster indices (J 1, J ). Let A be an algorithm that uses T and computes (Ĵ1, Ĵ). Under noise model I, for any 5
6 fixed < α < 1, if σ1 < c α k log(n/k), as n, we have ] inf sup [Ĵ1 J 1 or Ĵ J > 1 α P A AllAlg (J 1,J ) J all A similar result holds under noise model II if mk = Ω(log(n/k)). 4. Non-coherent Case log() k log(ne/k). (5) In this section we consider a similar setup to the one of Section 4.1 with the difference that v 1 is assumed to be uniformly distributed over a unit sphere. For simplicity, in this section we only consider noise model I. The ML optimization in this setup can be written as follows: max Ĵ J all (j 1,j ) Ĵ1 Ĵ T (j 1, j, :) (6) (Ĵ1, Ĵ) J all. Theorem 7. Under noise model I, if σ 1 = Ω(max(k, km)), as n, with high probability, (J 1, J ) is the optimal solution of optimization (6). If k > Ω(m), the achievability bound of Theorem 7 simplifies to the one of Theorem 5. In this case, using the result of Theorem 6, this bound is tight. If k < O(m), the achievability bound of Theorem 7 simplifies to Ω( mk) which is larger than the one of Theorem 5 (this is the price we pay for not knowing v 1 ). In the following, we show that this bound is also tight. To show the converse of Theorem 7, we consider the detection task which is presumably easier than the estimation task. Consider two probability distributions: (1) P σ1 under which the observed tensor is T = σ 1 u 1 w 1 v 1 + Z where J 1 and J have uniform distributions over k subsets of [n] and v 1 is uniform over a unit sphere. () P under which the observed tensor is T = Z. Noise entries are i.i.d. normal. We need the following definition of contiguous distributions ([8]): Definition. For every n N, let P,n and P 1,n be two probability measures on the same measure space. We say that the sequence (P 1,n ) is contiguous with respect to (P,n ), if, for any sequence of events A n, we have lim P,n(A n ) = lim P 1,n(A n ) =. (7) n n Theorem 8. If σ 1 < Õ( mk), P σ1 is contiguous with respect to P. This theorem with Lemma of [8] establishes the converse of Theorem 7. The proof is based on bounding the second moment of the Radon-Nikodym derivative of P σ1 with respect to P (SM Section 4.9). 5 Summary of Asymptotic Results Table 1 summarizes asymptotic bounds for the case of = 1/ k and m = Θ(n). For the MLE we consider the coherent model of Section 4.1. Also in Table 1 we summarize computational complexity of different tensor biclustering methods. We discuss analytical and empirical running time of these methods in SM Section.. Table 1: Comparative analysis of tensor biclustering methods. Results have been simplified for the case of m = Θ(n) and = 1/ k. Methods σ 1, noise model I σ 1, noise model II Comp. Complexity Tensor Folding+Spectral Ω(n 3/ ) Ω(n 3/ ) O(n 4 ) Tensor Unfolding+Spectral Ω(n ) Ω(n ) O(n 3 ) Th. Sum of Squared Trajectory Lengths Ω(nk) Ω(nk ) O(n 3 ) Th. Individual Trajectory Lengths Ω(k n) Ω(nk ) O(n 3 ) Maximum Likelihood Ω(k) Ω(k) exp(n) Statistical Lower Bound Õ(k) Õ(k) - 6
7 (a) noise model I (b) noise model II Bicluster Recovery Rate Bicluster Recovery Rate Signal Strength Signal Strength Tensor folding+spectral Tensor unfolding+spectral Th. sum of squared trajectory lengths Th. individual trajectory lengths Figure 3: Performance of different tensor biclustering methods in various values of σ 1 (i.e., the signal strength), under both noise models I and II. We consider n =, m = 5, k = 4. Experiments have been repeated 1 times for each point. In both noise models, the maximum likelihood estimator which has an exponential computational complexity leads to the best achievability bound compared to other methods. Below this bound, the inference is statistically impossible. Tensor folding+spectral method outperforms other methods with polynomial computational complexity if k > n under noise model I, and k > n 1/4 under noise model II. For smaller values of k, thresholding individual trajectory lengths lead to a better achievability bound. This case is a part of the high-snr regime where the average trajectory lengths within the bicluster is significantly larger than the one outside of the bicluster. Unlike thresholding individual trajectory lengths, other methods use the entire tensor to solve the tensor biclustering problem. Thus, when k is very small, the accumulated noise can dominate the signal strength. Moreover, the performance of the tensor unfolding method is always worst than the one of the tensor folding method. The reason is that, the tensor unfolding method merely infers a low dimensional subspace of trajectories, ignoring the block structure that true low dimensional trajectories form. 6 Numerical Results 6.1 Synthetic Data In this section we evaluate the performance of different tensor biclustering methods in synthetic datasets. We use the statistical model described in Section 4.1 to generate the input tensor T. Let (Ĵ1, Ĵ) be estimated bicluster indices (J 1, J ) where Ĵ1 = Ĵ = k. To evaluate the inference quality we compute the fraction of correctly recovered bicluster indices (SM Section 3.1). In our simulations we consider n =, m = 5, k = 4. Figure 3 shows the performance of four tensor biclustering methods in different values of σ 1 (i.e., the signal strength), under both noise models I and II. Tensor folding+spectral algorithm outperforms other methods in both noise models. The gain is larger in the setup of noise model II compared to the one of noise model I. 6. Real Data In this section we apply tensor biclustering methods to the roadmap epigenomics dataset [4] which provides histon mark signal strengths in different segments of human genome in various tissues and cell types. In this dataset, finding a subset of genome segments and a subset of tissues (cell-types) with highly correlated histon mark values can provide insight on tissue-specific functional roles of genome segments [4]. After pre-processing the data (SM Section 3.), we obtain a data tensor T R n1 n m where n 1 = 49 is the number of tissues (cell-types), n = 1457 is the number of 7
8 (a) (b) (c) 5 i-th largest eigenvalue i-th largest eigenvalue i-th largest singular value (d) tissues i genome segments (chromosome ) (e) H3K7ac H3K7me3 H3K36me3 H3K4me1 H3K4me3 H3K9ac i H3K9me tissues x genome segments inferred bicluster quality (f) i Tensor folding Tensor Th. Th. unfolding sum of individual squared TL TL Figure 4: An application of tensor biclustering methods to the the roadmap epigenomics data. random genome segments, and m = 7 is the number of histon marks. Note that although in our analytical results for simplicity we assume n 1 = n, our proposed methods can be used in a more general case such as the one considered in this section. We form two combined covariance matrices C 1 R n1 n1 and C R n n according to (3). Figure 4-(a,b) shows largest eigenvalues of C 1 and C, respectively. As illustrated in these figures, spectral gaps (i.e., λ 1 λ ) of these matrices are large, indicating the existence of a low dimensional signal tensor in the input tensor. We also form an unfolded tensor T unfolded R m n1n. Similarly, there is a large gap between the first and second largest singular values of T unfolded (Figure 4-c). We use the tensor folding+spectral algorithm 1 with J 1 = 1 and J = 4 (we consider other values for the bicluster size in SM Section 3.). The output of the algorithm (Ĵ1, Ĵ) is illustrated in Figure 4-d (note that for visualization purposes, we re-order rows and columns to have the bicluster appear in the corner). Figure 4-e illustrates the unfolded subspace {T (j 1, j, :) : (j 1, j ) Ĵ1 Ĵ}. In this inferred bicluster, Histon marks H3K4me3, H3K9ac, and H3K7ac have relatively high values. Reference [4] shows that these histon marks indicate a promoter region with an increased activation in the genome. To evaluate the quality of the inferred bicluster, we compute total absolute pairwise correlations among vectors in the inferred bicluster. As illustrated in Figure 4-f, the quality of inferred bicluster by tensor folding+spectral algorithm is larger than the one of other methods. Next, we compute the bicluster quality by choosing bicluster indices uniformly at random with the same cardinality. We repeat this experiment 1 times. There is a significant gap between the quality of these random biclusters and the ones inferred by tensor biclustering methods indicating the significance of our inferred biclusters. For more details on these experiment, see SM Section Discussion In this paper, we introduced and analyzed the tensor biclustering problem. The goal is to compute a subset of tensor rows and columns whose corresponding trajectories form a low dimensional subspace. To solve this problem, we proposed a method called tensor folding+spectral which demonstrated improved analytical and empirical performance compared to other considered methods. Moreover, we characterized computational and statistical (information theoretic) limits for the tensor biclustering problem in an asymptotic regime, under both coherent and non-coherent statistical models. Our results consider the case when the rank of the subspace is equal to one (i.e., q = 1). When q > 1, in both tensor folding+spectral and tensor unfolding+spectral methods, the embedded subspace in the signal matrix will have a rank of q > 1, with singular values σ 1 σ... σ q >. In this 8
9 setup, we need the spectral radius of the noise matrix to be smaller than σ q in order to guarantee the recovery of the subspace. The procedure to characterize asymptotic achievability bounds would follow from similar steps of the rank one case with some technical differences. For example, we would need to extend Lemma 6 to the case where the signal matrix has rank q > 1. Moreover, in our problem setup, we assumed that the size of the bicluster k and the rank of its subspace q are know parameters. In practice, these parameters can be learned approximately from the data. For example, in the tensor folding+spectral method, a good choice for the q parameter would be the index where eigenvalues of the folded matrix decrease significantly. Knowing q, one can determine the size of the bicluster similarly as the number of indices in top eigenvectors with significantly larger absolute values. Another practical approach to estimate model parameters would be trial and error plus cross validations. Some of the developed proof techniques may be of independent interest as well. For example, we proved an l version of the Davis-Kahan lemma for a Wishart noise matrix. Solving the tensor biclustering problem for the case of having multiple overlapping biclusters, for the case of having incomplete tensor, and for the case of a priori unknown bicluster sizes are among future directions. 8 Code We provide code for tensor biclustering methods in the following link: SoheilFeizi/Tensor-Biclustering. 9 Acknowledgment We thank Prof. Ofer Zeitouni for the helpful discussion on detectably proof techniques of probability measures. References [1] Amos Tanay, Roded Sharan, and Ron Shamir. Biclustering algorithms: A survey. Handbook of computational molecular biology, 9(1-):1 14, 5. [] Yudong Chen and Jiaming Xu. Statistical-computational tradeoffs in planted problems and submatrix localization with a growing number of clusters and submatrices. arxiv preprint arxiv:14.167, 14. [3] T Tony Cai, Tengyuan Liang, and Alexander Rakhlin. Computational and statistical boundaries for submatrix localization in a large noisy matrix. arxiv preprint arxiv: , 15. [4] Anshul Kundaje, Wouter Meuleman, Jason Ernst, Misha Bilenky, Angela Yen, Alireza Heravi- Moussavi, Pouya Kheradpour, Zhizhuo Zhang, Jianrong Wang, Michael J Ziller, et al. Integrative analysis of 111 reference human epigenomes. Nature, 518(7539):317 33, 15. [5] GTEx Consortium et al. The genotype-tissue expression (gtex) pilot analysis: Multitissue gene regulation in humans. Science, 348(635):648 66, 15. [6] Rui Chen, George I Mias, Jennifer Li-Pook-Than, Lihua Jiang, Hugo YK Lam, Rong Chen, Elana Miriami, Konrad J Karczewski, Manoj Hariharan, Frederick E Dewey, et al. Personal omics profiling reveals dynamic molecular and medical phenotypes. Cell, 148(6): , 1. [7] Emile Richard and Andrea Montanari. A statistical model for tensor pca. In Advances in Neural Information Processing Systems, pages , 14. [8] Andrea Montanari, Daniel Reichman, and Ofer Zeitouni. On the limitation of spectral methods: From the gaussian hidden clique problem to rank-one perturbations of gaussian tensors. In Advances in Neural Information Processing Systems, pages 17 5, 15. [9] Samuel B Hopkins, Tselil Schramm, Jonathan Shi, and David Steurer. Fast spectral algorithms from sum-of-squares proofs: tensor decomposition and planted sparse vectors. arxiv preprint arxiv: , 15. 9
10 [1] Samuel B Hopkins, Jonathan Shi, and David Steurer. Tensor principal component analysis via sum-of-square proofs. In COLT, pages , 15. [11] Amelia Perry, Alexander S Wein, and Afonso S Bandeira. Statistical limits of spiked tensor models. arxiv preprint arxiv: , 16. [1] Thibault Lesieur, Léo Miolane, Marc Lelarge, Florent Krzakala, and Lenka Zdeborová. Statistical and computational phase transitions in spiked tensor estimation. arxiv preprint arxiv:171.81, 17. [13] Animashree Anandkumar, Rong Ge, and Majid Janzamin. Guaranteed non-orthogonal tensor decomposition via alternating rank-1 updates. arxiv preprint arxiv:14.518, 14. [14] Anru Zhang and Dong Xia. Guaranteed tensor pca with optimality in statistics and computation. arxiv preprint arxiv:173.74, 17. [15] Animashree Anandkumar, Rong Ge, Daniel J Hsu, and Sham M Kakade. A tensor approach to learning mixed membership community models. Journal of Machine Learning Research, 15(1):39 31, 14. [16] Animashree Anandkumar, Rong Ge, Daniel J Hsu, Sham M Kakade, and Matus Telgarsky. Tensor decompositions for learning latent variable models. Journal of Machine Learning Research, 15(1):773 83, 14. [17] Victoria Hore, Ana Viñuela, Alfonso Buil, Julian Knight, Mark I McCarthy, Kerrin Small, and Jonathan Marchini. Tensor decomposition for multiple-tissue gene expression experiments. Nature Genetics, 48(9):194 11, 16. [18] Yiqiao Zhong and Nicolas Boumal. Near-optimal bounds for phase synchronization. arxiv preprint arxiv: , 17. 1
11 Supplementary Materials Tensor Biclustering Soheil Feizi Stanford University Hamid Javadi Stanford University David Tse Stanford University Contents 1 Problem Formulation Notation Signal and Noise Models Related Work Details of Tensor Biclustering Methods 3.1 Algorithms Computational Complexity Details of Numerical Experiments Synthetic Data Real Data Proofs Preliminary Lemmas Proof of Theorem MT Proof of Theorem MT Proof of Theorem MT Proof of Theorem MT Proof of Theorem MT Proof of Theorem MT Proof of Theorem MT Proof of Theorem MT Problem Formulation 1.1 Notation In this document, we refer to pointers in the main text using the prefix MT. For example, equation MT-1 refers to equation 1 in the main text. 31st Conference on Neural Information Processing Systems (NIPS 17), Long Beach, CA, USA.
12 We use T, X, and Z to represent input, signal, and noise tensors. For matrices we use bold-faced upper case letters, for vectors we use bold-faced lower case letters, and for scalars we use regular lower case letters. For example, X represents a matrix, x represents a vector, and x represents a scalar number. For any set J, J denotes its cardinality. I n1 n and 1 n1 n are the identity and all one matrices of size n 1 n, respectively. When no confusion arises, we drop the subscripts. [n] represents the set {1,,..., n}. For J [n], u (J) means that u(j) = if j J. J = [n] J. Ix=y is the indicator function of the event x = y. e i is a vector whose i-th entry is one and its other entries are zero. X d = Y means random variables X and Y have the same distribution. T r(x) and X t represent the trace and the transpose of the matrix X, respectively. diag(x) is a diagonal matrix whose diagonal elements are equal to x, while diag(x) is a vector of the diagonal elements of the matrix X. x = (x t x) 1/ is the second norm of the vector x. When no confusion arises, we drop the subscript. x is the infinity norm of the vector x (i.e., x = max( x i )). X is the operator norm of the matrix X, while X F is its Frobenius norm. < x, y > is the inner product between vectors x and y. x y indicates that vectors x and y are orthogonal. The matrix inner product is defined as < X, Y >= T r(xy t ). X F =< X, X >. Inner product and Frobenius norm of a tensor are defined similarly. det(x) is the determinant of X. X Y indicates kronecker product of matrices X and Y. D KL (P 1 P ) represents the Kullback Leibler (KL) divergence between two distributions P 1 and P. The asymptotic notation a(n) = O(b(n)) means that, there exists a universal constant c such that for sufficiently large n, we have a(n) < cb(n). If there exists c > such that a(n) = O(b(n) log(n) c ), we use the notation a(n) = Õ(b(n)). The asymptotic notation a(n) = Ω(b(n)) and a(n) = Ω(b(n)) is the same as b(n) = O(a(n)) and b(n) = Õ(a(n)), respectively. Moreover, we write a(n) = Θ(b(n)) iff a(n) = Ω(b(n)) and b(n) = Ω(a(n)). Similarly, we write a(n) = Θ(b(n)) iff a(n) = Ω(b(n)) and b(n) = Ω(a(n)). 1. Signal and Noise Models Consider q = 1 in MT-(1), the tensor biclustering model simplifies to T = X + Z = σ 1 u 1 w 1 v r + Z, (1) In this section, we explain noise models I and II with more details: - Noise Model I: In this model, the variance of the noise within and outside of bicluster indices is assumed to be the same. Thus, under this model we have σ z = 1. () This is the noise model often considered in analysis of sub-matrix localization [1, ] and tensor PCA [3, 4, 5, 6, 7, 8, 9]. Although this model simplifies the analysis, it has the following drawback: under this noise model, for every value of σ 1, the average trajectory length within the bicluster is larger than the average trajectory length outside of the bicluster. To see this, let T 1 R m k be a matrix whose columns include trajectories T (j 1, j, :) for (j 1, j ) J 1 J (i.e., T 1 is the unfolded T (J 1, J, :)). We can write T 1 = X 1 + Z 1 where X 1 and Z 1 are unfolded X (J 1, J, :) and Z(J 1, J, :), respectively. The squared Frobenius norm of X 1 is equal to X 1 F = σ 1. Moreover, the squared Frobenius norm of Z 1 has a χ-squared distribution with mk degrees of freedom. Thus, the average squared Frobenius norm of T 1 is equal to σ 1 + σ zmk. Let T R m k be a matrix whose columns include only noise trajectories. Using a similar argument, we have E[ T F ] = mk, which is smaller than σ 1 + mk. - Noise Model II: In this model, σ z is modeled to minimize the difference between the average trajectory lengths within and outside of the bicluster. If σ 1 < mk, without noise, the average trajectory lengths in the bicluster is smaller than the one outside of the bicluster. In this regime, having σ z = 1 σ 1/mk makes the average trajectory lengths within and outside of the bicluster comparable. This regime is called the low-snr regime. If σ 1 > mk, the average trajectory lengths in the bicluster is larger than the one outside of the bicluster. This regime is called the high-snr regime. In this regime, adding noise to signal trajectories increases their lengths and makes solving the tensor biclustering problem easier. Therefore, in this regime we assume σ z =
13 to minimize the difference between average trajectory lengths within and outside of the bicluster. Therefore, under the noise model II, we have 1.3 Related Work σz = max(, 1 σ 1 ). (3) mk A related model to tensor biclustering (1) is the spiked tensor model [3]: T = σ 1 v v v + Z. (4) Unlike the tensor biclustering model which is asymmetric along tensor dimensions, the spiked tensor model has a symmetric structure. Assuming the noise tensor Z has i.i.d. standard normal entries, reference [4] has shown that, if σ 1 < (1 ɛ)n, no algorithm based on a spectral statistical test can detect the existence of such signal structure, with error probability vanishing as n. References [6, 5] have shown that the inference of the signal tensor is possible using a polynomial time algorithm if σ 1 = Ω(n 3/ ). A variation of this bound also appears in the analysis of our tensor folding method (Theorem MT-1). Moreover, statistical and computational trade-offs of a generalized tensor PCA model have been studied in [9]. In the model (1), if m = 1, the tensor can be viewed as a matrix. Let u 1 (j 1 ) = 1/ k and w 1 (j ) = 1/ k for (j 1, j ) J 1 J, and consider noise model I. In this case, elements of the sub-matrix T (J 1, J, 1) have i.i.d. Gaussian distributions with µ = σ 1 /k means and unit variances. Elements of the matrix outside of indices J 1 J have normal distributions. In this special case, the tensor biclustering problem simplifies to the sub-matrix localization problem [1, ]. Note that the bicluster structure in this special case comes from scaling coefficients u 1 and w 1 since v 1 = 1. This problem is closely related to the planted clique, bi-clustering (co-clustering), and community detection problems. In this case, our statistical lower bound (Theorem MT-6) and the achievability bound of the MLE (Theorem MT-5) match with the ones derived specifically for the sub-matrix localization problem [1, ]. Details of Tensor Biclustering Methods.1 Algorithms In this section, we provide more details on three tensor biclustering methods, namely tensor unfolding+spectral, thresholding sum of squared trajectory lengths, and thresholding individual trajectory lengths. Algorithm 1 Tensor Unfolding+Spectral Input: T, k Compute ˆx, the top right singular vector of T unfolded Let Ĵ1 be the set of tensor row indices of k largest entries of ˆx Let Ĵ be the set of tensor column indices of k largest entries of ˆx Output: Ĵ1 and Ĵ Algorithm Thresholding Individual Trajectory Lengths Input: T, k Compute Ĵ1, the set of first indices of k largest trajectories Compute Ĵ, the set of second indices of k largest trajectories Output: Ĵ1 and Ĵ 3
14 Algorithm 3 Thresholding Sum of Squared Trajectory Lengths Input: T, k Compute d 1 (j 1 ) = n j T (j =1 1, j, :) for 1 j 1 n Compute d (j ) = n j T (j 1=1 1, j, :) for 1 j n Compute Ĵ1, the index set of k largest components of d 1 Compute Ĵ, the index set of k largest components of d Output: Ĵ1 and Ĵ. Computational Complexity In Table MT-1 we summarize computational complexity of different tensor biclustering methods. Tensor unfolding+spectral, thresholding sum of squared trajectory lengths, and thresholding individual trajectory lengths have linear computational complexity with respect to the tensor size n m. Computational complexity of the tensor folding+spectral is O(n m ) which is higher than linear and lower than quadratic with respect to the tensor size. Computational complexity of the MLE method is exponential in n. Figure 1 shows the empirical running time of different tensor biclustering methods with respect to the tensor size N = n m. In Figure 1-a, we vary the tensor size by varying m, while in Figure 1-b we increase the tensor size by increasing the size of all dimensions. In both setups, tensor unfolding+spectral method has the worst empirical running time compared to other methods. In the setup of panel (a), tensor folding+spectral method has a larger running time compared to thresholding individual and sum of squared trajectory lengths since its computational complexity depends on m while the computational complexity of other methods depend on m. Our empirical running time analysis has been performed on an ordinary laptop using implementations of tensor biclustering methods in MATLAB. 3 Details of Numerical Experiments 3.1 Synthetic Data In Section MT-6.1, we evaluate the performance of different tensor biclustering methods in synthetic datasets. We use the statistical model described in Section MT-4.1 to generate the input tensor T. Let (Ĵ1, Ĵ) be estimated bicluster indices (J 1, J ) where Ĵ1 = Ĵ = k. To evaluate the inference quality we compute the following score: Ĵ1 J 1 k + Ĵ J. (5) k This score is always between zero and one. If (Ĵ1, Ĵ) = (J 1, J ), this score achieves its maximum value one. Tensor unfolding+spectral method (Algorithm 1) and thresholding individual trajectory lengths method (Algorithm ) may have an output (Ĵ1, Ĵ) where Ĵ1 > k or Ĵ > k. This is because these algorithms ignore the block structure formed by bicluster indices. To have a fair comparison with other methods, we select k most repeated indices in their outputs as an estimate of bicluster indices. 3. Real Data In Section MT-6., we apply different tensor biclustering methods to the roadmap epigenomics dataset [1] which provides histon mark signal strengths in different segments of human genome in various tissues and cell types. This dataset can be viewed as a three dimensional tensor whose dimensions represent segments of genome, tissues (cell types), and histon marks. Reference [1] has shown that in a tissue, segments of genome with similar histon mark values are often have similar functional roles (e.g., they are enhancers, promoters, etc.). Moreover, histon marks of a specific genome segment can vary across different tissues and cell-types. Here we consider a portion of the roadmap epigenomics dataset to demonstrate applicability of tensor biclustering methods to this data type. A full analysis of the roadmap epigenomics dataset along 4
15 (a) running time (seconds) Tensor size 1 7 (b) running time (seconds) Tensor size 1 7 Tensor folding+spectral Tensor unfolding+spectral Th. sum of squared trajectory lengths Th. individual trajectory lengths Figure 1: Empirical running time of different tensor biclustering methods with respect to the tensor size n m. In panel (a) we consider n = 4, m = α4 and vary α, while in panel (b) we consider n = m = α 1/3 4. Experiments have been repeated 1 times for each point. with biological validations of inferences are beyond the scope of the present paper. We consider genome segments of chromosome in human. Each segment has 1 base pairs. In each genome segment, we consider the average value of the signal strength for every histon mark. We only consider segments with at least one non-zero histon mark value. In the roadmap epigenomics dataset, some tissues (cell-types) do not have data for some histon marks. Thus, we only consider a subset of tissues (cell-types) and a subset of histon marks with complete data. After these filtering steps, we obtain a data tensor T R n1 n m where n 1 = 49 is the number of tissues (cell-types), n = 1457 is the number of genome segments, and m = 7 is the number of histon marks. Our seven histon marks include the core set of five histone modification marks reported in reference [1] (i.e., H3K4me3, H3K4me1, H3K36me3, H3K7me3, and H3K9me3), along with two additional marks (i.e., H3K7ac and H3K9ac). T (j 1, j, i) provides the signal strength of histon mark i in the genome segment j of tissue j 1. Our goal is to find J 1 [n 1 ] and J [n ] where {T (j 1, j, :) : (j 1, j ) J 1 J } form a low dimensional subspace. To evaluate the quality of the inferred bicluster, we compute the total absolute pairwise correlations among vectors in the inferred bicluster, i.e., (j ρ(t (j 1,j ) (j 1, j 1, :), T (j 1, j, :)),j ) Ĵ1 Ĵ (6) ( Ĵ1 Ĵ ) Ĵ1 Ĵ where ρ(.,.) indicates the Pearson s correlation between two vectors. If all vectors in the inferred bicluster are parallel to each other, this value will be one. If vectors in the inferred bicluster are orthogonal to each other, this value will be zero. We evaluate the quality of inferred biclusters for different cluster sizes in Figure. Similar to the setup considered in the main text, in these cases the tensor folding+spectral method continues to outperform other tensor biclustering methods. 4 Proofs 4.1 Preliminary Lemmas For a sub-gaussian variable X, X ψ denotes the sub-gaussian norm of X defined as X ψ sup p 1 (E [ X p ]) 1/p. (7) p 1 If X is a centered Gaussian variable with variance σ, then X ψ cσ where c is an absolute constant. 5
16 (a) inferred subspace quality Tensor folding Tensor unfolding Th. sum of Th. individual squared TL TL random (b) inferred subspace quality Tensor folding Tensor unfolding Th. sum of Th. individual random squared TL TL Figure : The quality of inferred biclusters by different tensor biclustering methods and uniformly randomly selected bicluster indices when (a) Ĵ1 = 5, Ĵ =, and (b) Ĵ1 = and Ĵ = 8. Lemma 1. Let X 1,...,X N be independent, centered sub-gaussian random variables. Then, for every a = (a 1,..., a N ) T R N and every t, we have [ ] N ) P a i X i > t < e exp ( ct σm a, (8) i=1 where σ m = max i X i ψ and c > is an absolute constant. Proof. See Proposition 5.1 in [11]. Let Y be a sub-exponential random variable. Y ψ1 denotes the sub-exponential norm of Y, defined as Y ψ1 sup p 1 (E[ Y p ]) 1/p. (9) p 1 If X is sub-gaussian, Y = X is sub-exponential, and vice versa. Moreover, we have X ψ Y ψ1 X ψ. (1) Lemma. Let Y 1,...,Y N be independent, centered sub-exponential random variables. Then, for every a = (a 1,..., a N ) T R N and every t, we have [ ] N [ ( )] t t P a i Y i > t < exp c min σm a,, (11) σ m a i=1 where σ m = max i Y i ψ1 and c > is an absolute constant. Proof. See Proposition 5.16 in [11]. To bound the operator norm of sum of random matrices we use the following lemma: Lemma 3. Let Y 1,...,Y N be n 1 n independent random matrices such that for all j [N] [ ] P Y j E[Y j ] β p 1. (1) Moreover suppose we have Let E[Yj ] E [ Y j I Yj <β] p. (13) N µ = max E[Y j Yj] t E[Y j ]E[Yj] t, N E[YjY t j ] E[Yj]E[Y t j ] (14) Then for Y = N Y j, we have ( P [ Y E[Y] t] Np 1 + (n 1 + n ) exp 6 (t Np ) (µ + β(t Np )/3) ). (15)
17 Proof. See Proposition A.7 in [5]. Lemma 4. Let Z 1,...,Z N be m n independent random matrices such that Z r (i, j) has a standard normal distribution for every i, j. Then, for some constant c >, with high probability, we have N Z t jz j NmI < c max(n, m) N log(n). (16) Proof. Let n 1 = max(n, m). Let Y j = Z t j Z j. We have Y j = Z j. Since Z j has a sub-gaussian tail distribution, for some constant c >, we have Therefore, we have P [ Z j > tn 1 ] exp( ct). (17) P [ Y j E[Y j ] > (t + 1)n 1 ] P [ Y j > tn 1 ] exp( ct). (18) Moreover, we have E [Yj ] E [ ( ) ( ) Y j I Yj <β] cβ cβ = P [ Yj > β] E[Y j ] = m exp < n 1 exp. Thus, for a large enough constant c > and having β = c n 1 log(n) in (18) and (19), we can satisfy conditions (1) and (13) of Lemma 3 for p 1 = p = n c, for a sufficiently large c >. Next we compute µ in (14). Since Y j is symmetric we have N µ = E[Y j Yj] t E[Y j ]E[Yj] t N E[Y j Yj] t + Nm N E[ Y j Yj t ] + Nm, where the last step follows form Jenson s inequality. Moreover, we have E[ Y j Y t j ] = = = n 1 n 1 (19) () P [ Yj Yj t ] > t dt (1) [ P Z j > t 1/4] dt ( ) c t exp n 1 = n 1 c Therefore, with high probability, µ = Õ(Nn 1). Substituting p 1, p and µ in (15) completes the proof of this lemma. Lemma 5. Let x = (x 1, x,..., x n ) R n be a random vector with independent sub-gaussian entries x i with Ex i = and x i ψ K. For A R n n, t, we have { ( )} t t P { x, Ax E x, Ax } exp c min K 4 A, K () A F for some constant c >. Proof. See reference [1]. Lemma 6. Let Y = xx T + σw, where x R n, x = 1 and W R n n = ( zj z T j Ez ) jz T j where zj are i.i.d. N (, I n n ) gaussian random vectors. Let x R n, N 7
18 x = 1 be the eigenvector corresponding to the largest eigenvalue of the matrix Y. Let the operator norm of the matrix W be such that with probability at least 1 O(n ). Further, let W λ n,n, (3) σ c λ n,n, (4) for some positive constant c < 1/6. Letting M = x, we have ( ( )) x x O σ N log N + Mλn,N, (5) with high probability as n. Proof. Our proof is based on the technique used in [13]. However, in [13], x C n, W C n n and x i = 1 for 1 i n. In addition, in [13], it is assumed that W is a complex Wigner random matrix which is not the case in our model. Hence, we will develop a bound that fits our model. For 1 l n, we denote the l-th row (or column) of W by w l. Further, we define W (l) R n n as W (l) i,j W i,j, for i l, j l, (6) W (l) i,j, if i = l or j = l. Further, we define W (l) W W (l). Note that (3) results in W (l) λ n,n, W (l) λ n,n, w l λ n,n, (7) with probability at least 1 O(n ). Let x (l) be the eigenvector corresponding to the top eigenvalue of the matrix Y (l) = xx T + W (l). Note that for any 1 l n, we can write x l x l = (Y x) l λ 1 (Y) x ( l = xxt x ) + σ (W x) l l x l (8) λ 1 (Y) x, x λ 1 (Y) 1 M + σ (W x) l. λ 1 (Y) Hence, x x x, x λ 1 (Y) 1 M + σ W x. (9) λ 1 (Y) Next we bound the term W x. Note that for 1 l n, we have (W x) l = w l, x = w l, x (l) + w l, x x (l) w l, x (l) + wl x x (l). Note that Y = Y (l) + σ W (l). Hence, using the Davis-Kahan sin Θ Theorem (see, e.g., Lemma 11 in [13]), we have (3) x x (l) σ W (l) x (l) δ ( Y (l)) σ W (l), (31) where δ ( Y (l)) = λ 1 (Y (l) ) λ (Y (l) ) is the spectral gap of the matrix Y (l). Note that using the Weyl s inequality we can write ( δ Y (l)) δ ( xx T ) σ W (l) 1 σλ n,n, (3) where we have used (7) in the last inequality. Therefore, using (31), (7), (4) we get x x (l) σ 1 3σλ n,n W (l) x (l) 3λ n,n W (l) x (l), (33) 8
19 with probability at least 1 O(n ). Putting this in (3) we get, (W x) l w l, x (l) wl + W (l) x (l), (34) 3λ n,n with probability at least 1 O(n ). Note that W (l) x (l) w l, x (l). Hence, by (7), we get (W x) l W (l) x (l), (35) with probability at least 1 O(n ). Thus, we need to bound the term W (l) x (l). Note that here we can leverage the independence between W (l) and x (l) to get a tight bound on x x. We can write W (l) x (l) ( = W (l) x (l)) l + k l w l, x (l) + x (l) l ( W (l) x (l)) k w l w l, x (l) (l) + x λ n,n, with probability at least 1 O(n ). In order to bound the term w l, x (l), it suffices to bound the term w l, u, where u is a fixed vector. Hence, using the independence between w l, x (l), the bound for w l, x (l) follows by first conditioning on x (l) and then using the bound for w l, u. Now for a fixed vector u R n, u = 1, we can write w l, u = ( W T u ) = W T N u, e l l = zj z T N j u, e l E zj z T j u, e l N N N N = u T z j z T j e l E u T z j z T j e l = z T j ue T l z j E z T j ue T l z j where U 1 U U l = ue T l R n n, U = l N N = z j, U l z j E z j, U l z j = z, Uz E z, Uz, U 3... U N RnN nn, w R nn = Now, using Lemma 5 for t > we have { { }} t t P { w l, u > t} = P { z, Uz E z, Uz > t} exp C min U, (39) U F { { }} t = exp C min N, t, (4) for some constant C >. Therefore, by taking t = C N log N, where C C 3, we have with probability at least 1 n 3, w 1 w w 3.. w N (36) (37) N (, I nn nn ). (38) w l, u C N log N. (41) Hence, using the union bound over 1 l n, with probability at least 1 O(n ), w l, u C N log N, for 1 l n. (4) 9
Optimality and Sub-optimality of Principal Component Analysis for Spiked Random Matrices
 Optimality and Sub-optimality of Principal Component Analysis for Spiked Random Matrices Alex Wein MIT Joint work with: Amelia Perry (MIT), Afonso Bandeira (Courant NYU), Ankur Moitra (MIT) 1 / 22 Random
Optimality and Sub-optimality of Principal Component Analysis for Spiked Random Matrices Alex Wein MIT Joint work with: Amelia Perry (MIT), Afonso Bandeira (Courant NYU), Ankur Moitra (MIT) 1 / 22 Random
Computational and Statistical Boundaries for Submatrix Localization in a Large Noisy Matrix
 University of Pennsylvania ScholarlyCommons Statistics Papers Wharton Faculty Research 2017 Computational and Statistical Boundaries for Submatrix Localization in a Large Noisy Matrix Tony Cai University
University of Pennsylvania ScholarlyCommons Statistics Papers Wharton Faculty Research 2017 Computational and Statistical Boundaries for Submatrix Localization in a Large Noisy Matrix Tony Cai University
PCA with random noise. Van Ha Vu. Department of Mathematics Yale University
 PCA with random noise Van Ha Vu Department of Mathematics Yale University An important problem that appears in various areas of applied mathematics (in particular statistics, computer science and numerical
PCA with random noise Van Ha Vu Department of Mathematics Yale University An important problem that appears in various areas of applied mathematics (in particular statistics, computer science and numerical
arxiv: v5 [math.na] 16 Nov 2017
![arxiv: v5 [math.na] 16 Nov 2017 arxiv: v5 [math.na] 16 Nov 2017](/thumbs/78/78774947.jpg) RANDOM PERTURBATION OF LOW RANK MATRICES: IMPROVING CLASSICAL BOUNDS arxiv:3.657v5 [math.na] 6 Nov 07 SEAN O ROURKE, VAN VU, AND KE WANG Abstract. Matrix perturbation inequalities, such as Weyl s theorem
RANDOM PERTURBATION OF LOW RANK MATRICES: IMPROVING CLASSICAL BOUNDS arxiv:3.657v5 [math.na] 6 Nov 07 SEAN O ROURKE, VAN VU, AND KE WANG Abstract. Matrix perturbation inequalities, such as Weyl s theorem
Appendix A. Proof to Theorem 1
 Appendix A Proof to Theorem In this section, we prove the sample complexity bound given in Theorem The proof consists of three main parts In Appendix A, we prove perturbation lemmas that bound the estimation
Appendix A Proof to Theorem In this section, we prove the sample complexity bound given in Theorem The proof consists of three main parts In Appendix A, we prove perturbation lemmas that bound the estimation
ECE 598: Representation Learning: Algorithms and Models Fall 2017
 ECE 598: Representation Learning: Algorithms and Models Fall 2017 Lecture 1: Tensor Methods in Machine Learning Lecturer: Pramod Viswanathan Scribe: Bharath V Raghavan, Oct 3, 2017 11 Introduction Tensors
ECE 598: Representation Learning: Algorithms and Models Fall 2017 Lecture 1: Tensor Methods in Machine Learning Lecturer: Pramod Viswanathan Scribe: Bharath V Raghavan, Oct 3, 2017 11 Introduction Tensors
8.1 Concentration inequality for Gaussian random matrix (cont d)
 MGMT 69: Topics in High-dimensional Data Analysis Falll 26 Lecture 8: Spectral clustering and Laplacian matrices Lecturer: Jiaming Xu Scribe: Hyun-Ju Oh and Taotao He, October 4, 26 Outline Concentration
MGMT 69: Topics in High-dimensional Data Analysis Falll 26 Lecture 8: Spectral clustering and Laplacian matrices Lecturer: Jiaming Xu Scribe: Hyun-Ju Oh and Taotao He, October 4, 26 Outline Concentration
Principal Component Analysis
 Machine Learning Michaelmas 2017 James Worrell Principal Component Analysis 1 Introduction 1.1 Goals of PCA Principal components analysis (PCA) is a dimensionality reduction technique that can be used
Machine Learning Michaelmas 2017 James Worrell Principal Component Analysis 1 Introduction 1.1 Goals of PCA Principal components analysis (PCA) is a dimensionality reduction technique that can be used
Porcupine Neural Networks: (Almost) All Local Optima are Global
 Porcupine Neural Networs: (Almost) All Local Optima are Global Soheil Feizi, Hamid Javadi, Jesse Zhang and David Tse arxiv:1710.0196v1 [stat.ml] 5 Oct 017 Stanford University Abstract Neural networs have
Porcupine Neural Networs: (Almost) All Local Optima are Global Soheil Feizi, Hamid Javadi, Jesse Zhang and David Tse arxiv:1710.0196v1 [stat.ml] 5 Oct 017 Stanford University Abstract Neural networs have
ELE 538B: Mathematics of High-Dimensional Data. Spectral methods. Yuxin Chen Princeton University, Fall 2018
 ELE 538B: Mathematics of High-Dimensional Data Spectral methods Yuxin Chen Princeton University, Fall 2018 Outline A motivating application: graph clustering Distance and angles between two subspaces Eigen-space
ELE 538B: Mathematics of High-Dimensional Data Spectral methods Yuxin Chen Princeton University, Fall 2018 Outline A motivating application: graph clustering Distance and angles between two subspaces Eigen-space
Rank Determination for Low-Rank Data Completion
 Journal of Machine Learning Research 18 017) 1-9 Submitted 7/17; Revised 8/17; Published 9/17 Rank Determination for Low-Rank Data Completion Morteza Ashraphijuo Columbia University New York, NY 1007,
Journal of Machine Learning Research 18 017) 1-9 Submitted 7/17; Revised 8/17; Published 9/17 Rank Determination for Low-Rank Data Completion Morteza Ashraphijuo Columbia University New York, NY 1007,
Statistical and Computational Phase Transitions in Planted Models
 Statistical and Computational Phase Transitions in Planted Models Jiaming Xu Joint work with Yudong Chen (UC Berkeley) Acknowledgement: Prof. Bruce Hajek November 4, 203 Cluster/Community structure in
Statistical and Computational Phase Transitions in Planted Models Jiaming Xu Joint work with Yudong Chen (UC Berkeley) Acknowledgement: Prof. Bruce Hajek November 4, 203 Cluster/Community structure in
Supremum of simple stochastic processes
 Subspace embeddings Daniel Hsu COMS 4772 1 Supremum of simple stochastic processes 2 Recap: JL lemma JL lemma. For any ε (0, 1/2), point set S R d of cardinality 16 ln n S = n, and k N such that k, there
Subspace embeddings Daniel Hsu COMS 4772 1 Supremum of simple stochastic processes 2 Recap: JL lemma JL lemma. For any ε (0, 1/2), point set S R d of cardinality 16 ln n S = n, and k N such that k, there
arxiv: v2 [math.st] 21 Oct 2015
![arxiv: v2 [math.st] 21 Oct 2015 arxiv: v2 [math.st] 21 Oct 2015](/thumbs/87/95700936.jpg) Computational and Statistical Boundaries for Submatrix Localization in a Large Noisy Matrix T. Tony Cai, Tengyuan Liang and Alexander Rakhlin arxiv:1502.01988v2 [math.st] 21 Oct 2015 Department of Statistics
Computational and Statistical Boundaries for Submatrix Localization in a Large Noisy Matrix T. Tony Cai, Tengyuan Liang and Alexander Rakhlin arxiv:1502.01988v2 [math.st] 21 Oct 2015 Department of Statistics
DISCUSSION OF INFLUENTIAL FEATURE PCA FOR HIGH DIMENSIONAL CLUSTERING. By T. Tony Cai and Linjun Zhang University of Pennsylvania
 Submitted to the Annals of Statistics DISCUSSION OF INFLUENTIAL FEATURE PCA FOR HIGH DIMENSIONAL CLUSTERING By T. Tony Cai and Linjun Zhang University of Pennsylvania We would like to congratulate the
Submitted to the Annals of Statistics DISCUSSION OF INFLUENTIAL FEATURE PCA FOR HIGH DIMENSIONAL CLUSTERING By T. Tony Cai and Linjun Zhang University of Pennsylvania We would like to congratulate the
Reconstruction in the Generalized Stochastic Block Model
 Reconstruction in the Generalized Stochastic Block Model Marc Lelarge 1 Laurent Massoulié 2 Jiaming Xu 3 1 INRIA-ENS 2 INRIA-Microsoft Research Joint Centre 3 University of Illinois, Urbana-Champaign GDR
Reconstruction in the Generalized Stochastic Block Model Marc Lelarge 1 Laurent Massoulié 2 Jiaming Xu 3 1 INRIA-ENS 2 INRIA-Microsoft Research Joint Centre 3 University of Illinois, Urbana-Champaign GDR
PCA and admixture models
 PCA and admixture models CM226: Machine Learning for Bioinformatics. Fall 2016 Sriram Sankararaman Acknowledgments: Fei Sha, Ameet Talwalkar, Alkes Price PCA and admixture models 1 / 57 Announcements HW1
PCA and admixture models CM226: Machine Learning for Bioinformatics. Fall 2016 Sriram Sankararaman Acknowledgments: Fei Sha, Ameet Talwalkar, Alkes Price PCA and admixture models 1 / 57 Announcements HW1
sparse and low-rank tensor recovery Cubic-Sketching
 Sparse and Low-Ran Tensor Recovery via Cubic-Setching Guang Cheng Department of Statistics Purdue University www.science.purdue.edu/bigdata CCAM@Purdue Math Oct. 27, 2017 Joint wor with Botao Hao and Anru
Sparse and Low-Ran Tensor Recovery via Cubic-Setching Guang Cheng Department of Statistics Purdue University www.science.purdue.edu/bigdata CCAM@Purdue Math Oct. 27, 2017 Joint wor with Botao Hao and Anru
Structure in Data. A major objective in data analysis is to identify interesting features or structure in the data.
 Structure in Data A major objective in data analysis is to identify interesting features or structure in the data. The graphical methods are very useful in discovering structure. There are basically two
Structure in Data A major objective in data analysis is to identify interesting features or structure in the data. The graphical methods are very useful in discovering structure. There are basically two
Introduction to Machine Learning. PCA and Spectral Clustering. Introduction to Machine Learning, Slides: Eran Halperin
 1 Introduction to Machine Learning PCA and Spectral Clustering Introduction to Machine Learning, 2013-14 Slides: Eran Halperin Singular Value Decomposition (SVD) The singular value decomposition (SVD)
1 Introduction to Machine Learning PCA and Spectral Clustering Introduction to Machine Learning, 2013-14 Slides: Eran Halperin Singular Value Decomposition (SVD) The singular value decomposition (SVD)
Orthogonal tensor decomposition
 Orthogonal tensor decomposition Daniel Hsu Columbia University Largely based on 2012 arxiv report Tensor decompositions for learning latent variable models, with Anandkumar, Ge, Kakade, and Telgarsky.
Orthogonal tensor decomposition Daniel Hsu Columbia University Largely based on 2012 arxiv report Tensor decompositions for learning latent variable models, with Anandkumar, Ge, Kakade, and Telgarsky.
Sum-of-Squares and Spectral Algorithms
 Sum-of-Squares and Spectral Algorithms Tselil Schramm June 23, 2017 Workshop on SoS @ STOC 2017 Spectral algorithms as a tool for analyzing SoS. SoS Semidefinite Programs Spectral Algorithms SoS suggests
Sum-of-Squares and Spectral Algorithms Tselil Schramm June 23, 2017 Workshop on SoS @ STOC 2017 Spectral algorithms as a tool for analyzing SoS. SoS Semidefinite Programs Spectral Algorithms SoS suggests
CS281 Section 4: Factor Analysis and PCA
 CS81 Section 4: Factor Analysis and PCA Scott Linderman At this point we have seen a variety of machine learning models, with a particular emphasis on models for supervised learning. In particular, we
CS81 Section 4: Factor Analysis and PCA Scott Linderman At this point we have seen a variety of machine learning models, with a particular emphasis on models for supervised learning. In particular, we
High-dimensional two-sample tests under strongly spiked eigenvalue models
 1 High-dimensional two-sample tests under strongly spiked eigenvalue models Makoto Aoshima and Kazuyoshi Yata University of Tsukuba Abstract: We consider a new two-sample test for high-dimensional data
1 High-dimensional two-sample tests under strongly spiked eigenvalue models Makoto Aoshima and Kazuyoshi Yata University of Tsukuba Abstract: We consider a new two-sample test for high-dimensional data
Supplementary Material for: Spectral Unsupervised Parsing with Additive Tree Metrics
 Supplementary Material for: Spectral Unsupervised Parsing with Additive Tree Metrics Ankur P. Parikh School of Computer Science Carnegie Mellon University apparikh@cs.cmu.edu Shay B. Cohen School of Informatics
Supplementary Material for: Spectral Unsupervised Parsing with Additive Tree Metrics Ankur P. Parikh School of Computer Science Carnegie Mellon University apparikh@cs.cmu.edu Shay B. Cohen School of Informatics
IV. Matrix Approximation using Least-Squares
 IV. Matrix Approximation using Least-Squares The SVD and Matrix Approximation We begin with the following fundamental question. Let A be an M N matrix with rank R. What is the closest matrix to A that
IV. Matrix Approximation using Least-Squares The SVD and Matrix Approximation We begin with the following fundamental question. Let A be an M N matrix with rank R. What is the closest matrix to A that
Donald Goldfarb IEOR Department Columbia University UCLA Mathematics Department Distinguished Lecture Series May 17 19, 2016
 Optimization for Tensor Models Donald Goldfarb IEOR Department Columbia University UCLA Mathematics Department Distinguished Lecture Series May 17 19, 2016 1 Tensors Matrix Tensor: higher-order matrix
Optimization for Tensor Models Donald Goldfarb IEOR Department Columbia University UCLA Mathematics Department Distinguished Lecture Series May 17 19, 2016 1 Tensors Matrix Tensor: higher-order matrix
Robust Principal Component Analysis
 ELE 538B: Mathematics of High-Dimensional Data Robust Principal Component Analysis Yuxin Chen Princeton University, Fall 2018 Disentangling sparse and low-rank matrices Suppose we are given a matrix M
ELE 538B: Mathematics of High-Dimensional Data Robust Principal Component Analysis Yuxin Chen Princeton University, Fall 2018 Disentangling sparse and low-rank matrices Suppose we are given a matrix M
Invertibility of random matrices
 University of Michigan February 2011, Princeton University Origins of Random Matrix Theory Statistics (Wishart matrices) PCA of a multivariate Gaussian distribution. [Gaël Varoquaux s blog gael-varoquaux.info]
University of Michigan February 2011, Princeton University Origins of Random Matrix Theory Statistics (Wishart matrices) PCA of a multivariate Gaussian distribution. [Gaël Varoquaux s blog gael-varoquaux.info]
Information-theoretic bounds and phase transitions in clustering, sparse PCA, and submatrix localization
 Information-theoretic bounds and phase transitions in clustering, sparse PCA, and submatrix localization Jess Banks Cristopher Moore Roman Vershynin Nicolas Verzelen Jiaming Xu Abstract We study the problem
Information-theoretic bounds and phase transitions in clustering, sparse PCA, and submatrix localization Jess Banks Cristopher Moore Roman Vershynin Nicolas Verzelen Jiaming Xu Abstract We study the problem
Lecture 7 MIMO Communica2ons
 Wireless Communications Lecture 7 MIMO Communica2ons Prof. Chun-Hung Liu Dept. of Electrical and Computer Engineering National Chiao Tung University Fall 2014 1 Outline MIMO Communications (Chapter 10
Wireless Communications Lecture 7 MIMO Communica2ons Prof. Chun-Hung Liu Dept. of Electrical and Computer Engineering National Chiao Tung University Fall 2014 1 Outline MIMO Communications (Chapter 10
Non-convex Robust PCA: Provable Bounds
 Non-convex Robust PCA: Provable Bounds Anima Anandkumar U.C. Irvine Joint work with Praneeth Netrapalli, U.N. Niranjan, Prateek Jain and Sujay Sanghavi. Learning with Big Data High Dimensional Regime Missing
Non-convex Robust PCA: Provable Bounds Anima Anandkumar U.C. Irvine Joint work with Praneeth Netrapalli, U.N. Niranjan, Prateek Jain and Sujay Sanghavi. Learning with Big Data High Dimensional Regime Missing
ECE G: Special Topics in Signal Processing: Sparsity, Structure, and Inference
 ECE 18-898G: Special Topics in Signal Processing: Sparsity, Structure, and Inference Low-rank matrix recovery via nonconvex optimization Yuejie Chi Department of Electrical and Computer Engineering Spring
ECE 18-898G: Special Topics in Signal Processing: Sparsity, Structure, and Inference Low-rank matrix recovery via nonconvex optimization Yuejie Chi Department of Electrical and Computer Engineering Spring
Analysis of Robust PCA via Local Incoherence
 Analysis of Robust PCA via Local Incoherence Huishuai Zhang Department of EECS Syracuse University Syracuse, NY 3244 hzhan23@syr.edu Yi Zhou Department of EECS Syracuse University Syracuse, NY 3244 yzhou35@syr.edu
Analysis of Robust PCA via Local Incoherence Huishuai Zhang Department of EECS Syracuse University Syracuse, NY 3244 hzhan23@syr.edu Yi Zhou Department of EECS Syracuse University Syracuse, NY 3244 yzhou35@syr.edu
Fast Angular Synchronization for Phase Retrieval via Incomplete Information
 Fast Angular Synchronization for Phase Retrieval via Incomplete Information Aditya Viswanathan a and Mark Iwen b a Department of Mathematics, Michigan State University; b Department of Mathematics & Department
Fast Angular Synchronization for Phase Retrieval via Incomplete Information Aditya Viswanathan a and Mark Iwen b a Department of Mathematics, Michigan State University; b Department of Mathematics & Department
Lecture 14: SVD, Power method, and Planted Graph problems (+ eigenvalues of random matrices) Lecturer: Sanjeev Arora
 princeton univ. F 13 cos 521: Advanced Algorithm Design Lecture 14: SVD, Power method, and Planted Graph problems (+ eigenvalues of random matrices) Lecturer: Sanjeev Arora Scribe: Today we continue the
princeton univ. F 13 cos 521: Advanced Algorithm Design Lecture 14: SVD, Power method, and Planted Graph problems (+ eigenvalues of random matrices) Lecturer: Sanjeev Arora Scribe: Today we continue the
Deep Linear Networks with Arbitrary Loss: All Local Minima Are Global
 homas Laurent * 1 James H. von Brecht * 2 Abstract We consider deep linear networks with arbitrary convex differentiable loss. We provide a short and elementary proof of the fact that all local minima
homas Laurent * 1 James H. von Brecht * 2 Abstract We consider deep linear networks with arbitrary convex differentiable loss. We provide a short and elementary proof of the fact that all local minima
CS264: Beyond Worst-Case Analysis Lecture #15: Topic Modeling and Nonnegative Matrix Factorization
 CS264: Beyond Worst-Case Analysis Lecture #15: Topic Modeling and Nonnegative Matrix Factorization Tim Roughgarden February 28, 2017 1 Preamble This lecture fulfills a promise made back in Lecture #1,
CS264: Beyond Worst-Case Analysis Lecture #15: Topic Modeling and Nonnegative Matrix Factorization Tim Roughgarden February 28, 2017 1 Preamble This lecture fulfills a promise made back in Lecture #1,
An Introduction to Spectral Learning
 An Introduction to Spectral Learning Hanxiao Liu November 8, 2013 Outline 1 Method of Moments 2 Learning topic models using spectral properties 3 Anchor words Preliminaries X 1,, X n p (x; θ), θ = (θ 1,
An Introduction to Spectral Learning Hanxiao Liu November 8, 2013 Outline 1 Method of Moments 2 Learning topic models using spectral properties 3 Anchor words Preliminaries X 1,, X n p (x; θ), θ = (θ 1,
Conditions for Robust Principal Component Analysis
 Rose-Hulman Undergraduate Mathematics Journal Volume 12 Issue 2 Article 9 Conditions for Robust Principal Component Analysis Michael Hornstein Stanford University, mdhornstein@gmail.com Follow this and
Rose-Hulman Undergraduate Mathematics Journal Volume 12 Issue 2 Article 9 Conditions for Robust Principal Component Analysis Michael Hornstein Stanford University, mdhornstein@gmail.com Follow this and
arxiv: v1 [cs.it] 21 Feb 2013
![arxiv: v1 [cs.it] 21 Feb 2013 arxiv: v1 [cs.it] 21 Feb 2013](/thumbs/88/115041122.jpg) q-ary Compressive Sensing arxiv:30.568v [cs.it] Feb 03 Youssef Mroueh,, Lorenzo Rosasco, CBCL, CSAIL, Massachusetts Institute of Technology LCSL, Istituto Italiano di Tecnologia and IIT@MIT lab, Istituto
q-ary Compressive Sensing arxiv:30.568v [cs.it] Feb 03 Youssef Mroueh,, Lorenzo Rosasco, CBCL, CSAIL, Massachusetts Institute of Technology LCSL, Istituto Italiano di Tecnologia and IIT@MIT lab, Istituto
A statistical model for tensor PCA
 A statistical model for tensor PCA Andrea Montanari Statistics & Electrical Engineering Stanford University Emile Richard Electrical Engineering Stanford University Abstract We consider the Principal Component
A statistical model for tensor PCA Andrea Montanari Statistics & Electrical Engineering Stanford University Emile Richard Electrical Engineering Stanford University Abstract We consider the Principal Component
Finding High-Order Correlations in High-Dimensional Biological Data
 Finding High-Order Correlations in High-Dimensional Biological Data Xiang Zhang, Feng Pan, and Wei Wang Department of Computer Science University of North Carolina at Chapel Hill 1 Introduction Many real
Finding High-Order Correlations in High-Dimensional Biological Data Xiang Zhang, Feng Pan, and Wei Wang Department of Computer Science University of North Carolina at Chapel Hill 1 Introduction Many real
First Efficient Convergence for Streaming k-pca: a Global, Gap-Free, and Near-Optimal Rate
 58th Annual IEEE Symposium on Foundations of Computer Science First Efficient Convergence for Streaming k-pca: a Global, Gap-Free, and Near-Optimal Rate Zeyuan Allen-Zhu Microsoft Research zeyuan@csail.mit.edu
58th Annual IEEE Symposium on Foundations of Computer Science First Efficient Convergence for Streaming k-pca: a Global, Gap-Free, and Near-Optimal Rate Zeyuan Allen-Zhu Microsoft Research zeyuan@csail.mit.edu
Approximate Message Passing
 Approximate Message Passing Mohammad Emtiyaz Khan CS, UBC February 8, 2012 Abstract In this note, I summarize Sections 5.1 and 5.2 of Arian Maleki s PhD thesis. 1 Notation We denote scalars by small letters
Approximate Message Passing Mohammad Emtiyaz Khan CS, UBC February 8, 2012 Abstract In this note, I summarize Sections 5.1 and 5.2 of Arian Maleki s PhD thesis. 1 Notation We denote scalars by small letters
1-Bit Matrix Completion
 1-Bit Matrix Completion Mark A. Davenport School of Electrical and Computer Engineering Georgia Institute of Technology Yaniv Plan Mary Wootters Ewout van den Berg Matrix Completion d When is it possible
1-Bit Matrix Completion Mark A. Davenport School of Electrical and Computer Engineering Georgia Institute of Technology Yaniv Plan Mary Wootters Ewout van den Berg Matrix Completion d When is it possible
Reconstruction from Anisotropic Random Measurements
 Reconstruction from Anisotropic Random Measurements Mark Rudelson and Shuheng Zhou The University of Michigan, Ann Arbor Coding, Complexity, and Sparsity Workshop, 013 Ann Arbor, Michigan August 7, 013
Reconstruction from Anisotropic Random Measurements Mark Rudelson and Shuheng Zhou The University of Michigan, Ann Arbor Coding, Complexity, and Sparsity Workshop, 013 Ann Arbor, Michigan August 7, 013
Gaussian Processes. Le Song. Machine Learning II: Advanced Topics CSE 8803ML, Spring 2012
 Gaussian Processes Le Song Machine Learning II: Advanced Topics CSE 8803ML, Spring 01 Pictorial view of embedding distribution Transform the entire distribution to expected features Feature space Feature
Gaussian Processes Le Song Machine Learning II: Advanced Topics CSE 8803ML, Spring 01 Pictorial view of embedding distribution Transform the entire distribution to expected features Feature space Feature
Dictionary Learning Using Tensor Methods
 Dictionary Learning Using Tensor Methods Anima Anandkumar U.C. Irvine Joint work with Rong Ge, Majid Janzamin and Furong Huang. Feature learning as cornerstone of ML ML Practice Feature learning as cornerstone
Dictionary Learning Using Tensor Methods Anima Anandkumar U.C. Irvine Joint work with Rong Ge, Majid Janzamin and Furong Huang. Feature learning as cornerstone of ML ML Practice Feature learning as cornerstone
Analysis of Spectral Kernel Design based Semi-supervised Learning
 Analysis of Spectral Kernel Design based Semi-supervised Learning Tong Zhang IBM T. J. Watson Research Center Yorktown Heights, NY 10598 Rie Kubota Ando IBM T. J. Watson Research Center Yorktown Heights,
Analysis of Spectral Kernel Design based Semi-supervised Learning Tong Zhang IBM T. J. Watson Research Center Yorktown Heights, NY 10598 Rie Kubota Ando IBM T. J. Watson Research Center Yorktown Heights,
Linear Algebra Massoud Malek
 CSUEB Linear Algebra Massoud Malek Inner Product and Normed Space In all that follows, the n n identity matrix is denoted by I n, the n n zero matrix by Z n, and the zero vector by θ n An inner product
CSUEB Linear Algebra Massoud Malek Inner Product and Normed Space In all that follows, the n n identity matrix is denoted by I n, the n n zero matrix by Z n, and the zero vector by θ n An inner product
Mathematical foundations - linear algebra
 Mathematical foundations - linear algebra Andrea Passerini passerini@disi.unitn.it Machine Learning Vector space Definition (over reals) A set X is called a vector space over IR if addition and scalar
Mathematical foundations - linear algebra Andrea Passerini passerini@disi.unitn.it Machine Learning Vector space Definition (over reals) A set X is called a vector space over IR if addition and scalar
U.C. Berkeley Better-than-Worst-Case Analysis Handout 3 Luca Trevisan May 24, 2018
 U.C. Berkeley Better-than-Worst-Case Analysis Handout 3 Luca Trevisan May 24, 2018 Lecture 3 In which we show how to find a planted clique in a random graph. 1 Finding a Planted Clique We will analyze
U.C. Berkeley Better-than-Worst-Case Analysis Handout 3 Luca Trevisan May 24, 2018 Lecture 3 In which we show how to find a planted clique in a random graph. 1 Finding a Planted Clique We will analyze
15 Singular Value Decomposition
 15 Singular Value Decomposition For any high-dimensional data analysis, one s first thought should often be: can I use an SVD? The singular value decomposition is an invaluable analysis tool for dealing
15 Singular Value Decomposition For any high-dimensional data analysis, one s first thought should often be: can I use an SVD? The singular value decomposition is an invaluable analysis tool for dealing
Performance Limits on the Classification of Kronecker-structured Models
 Performance Limits on the Classification of Kronecker-structured Models Ishan Jindal and Matthew Nokleby Electrical and Computer Engineering Wayne State University Motivation: Subspace models (Union of)
Performance Limits on the Classification of Kronecker-structured Models Ishan Jindal and Matthew Nokleby Electrical and Computer Engineering Wayne State University Motivation: Subspace models (Union of)
ECE 8201: Low-dimensional Signal Models for High-dimensional Data Analysis
 ECE 8201: Low-dimensional Signal Models for High-dimensional Data Analysis Lecture 7: Matrix completion Yuejie Chi The Ohio State University Page 1 Reference Guaranteed Minimum-Rank Solutions of Linear
ECE 8201: Low-dimensional Signal Models for High-dimensional Data Analysis Lecture 7: Matrix completion Yuejie Chi The Ohio State University Page 1 Reference Guaranteed Minimum-Rank Solutions of Linear
Lecture 3. Random Fourier measurements
 Lecture 3. Random Fourier measurements 1 Sampling from Fourier matrices 2 Law of Large Numbers and its operator-valued versions 3 Frames. Rudelson s Selection Theorem Sampling from Fourier matrices Our
Lecture 3. Random Fourier measurements 1 Sampling from Fourier matrices 2 Law of Large Numbers and its operator-valued versions 3 Frames. Rudelson s Selection Theorem Sampling from Fourier matrices Our
Community detection in stochastic block models via spectral methods
 Community detection in stochastic block models via spectral methods Laurent Massoulié (MSR-Inria Joint Centre, Inria) based on joint works with: Dan Tomozei (EPFL), Marc Lelarge (Inria), Jiaming Xu (UIUC),
Community detection in stochastic block models via spectral methods Laurent Massoulié (MSR-Inria Joint Centre, Inria) based on joint works with: Dan Tomozei (EPFL), Marc Lelarge (Inria), Jiaming Xu (UIUC),
STAT 200C: High-dimensional Statistics
 STAT 200C: High-dimensional Statistics Arash A. Amini May 30, 2018 1 / 59 Classical case: n d. Asymptotic assumption: d is fixed and n. Basic tools: LLN and CLT. High-dimensional setting: n d, e.g. n/d
STAT 200C: High-dimensional Statistics Arash A. Amini May 30, 2018 1 / 59 Classical case: n d. Asymptotic assumption: d is fixed and n. Basic tools: LLN and CLT. High-dimensional setting: n d, e.g. n/d
An algebraic perspective on integer sparse recovery
 An algebraic perspective on integer sparse recovery Lenny Fukshansky Claremont McKenna College (joint work with Deanna Needell and Benny Sudakov) Combinatorics Seminar USC October 31, 2018 From Wikipedia:
An algebraic perspective on integer sparse recovery Lenny Fukshansky Claremont McKenna College (joint work with Deanna Needell and Benny Sudakov) Combinatorics Seminar USC October 31, 2018 From Wikipedia:
Solving Corrupted Quadratic Equations, Provably
 Solving Corrupted Quadratic Equations, Provably Yuejie Chi London Workshop on Sparse Signal Processing September 206 Acknowledgement Joint work with Yuanxin Li (OSU), Huishuai Zhuang (Syracuse) and Yingbin
Solving Corrupted Quadratic Equations, Provably Yuejie Chi London Workshop on Sparse Signal Processing September 206 Acknowledgement Joint work with Yuanxin Li (OSU), Huishuai Zhuang (Syracuse) and Yingbin
Review (probability, linear algebra) CE-717 : Machine Learning Sharif University of Technology
 Review (probability, linear algebra) CE-717 : Machine Learning Sharif University of Technology M. Soleymani Fall 2012 Some slides have been adopted from Prof. H.R. Rabiee s and also Prof. R. Gutierrez-Osuna
Review (probability, linear algebra) CE-717 : Machine Learning Sharif University of Technology M. Soleymani Fall 2012 Some slides have been adopted from Prof. H.R. Rabiee s and also Prof. R. Gutierrez-Osuna
Asymptotic Distribution of the Largest Eigenvalue via Geometric Representations of High-Dimension, Low-Sample-Size Data
 Sri Lankan Journal of Applied Statistics (Special Issue) Modern Statistical Methodologies in the Cutting Edge of Science Asymptotic Distribution of the Largest Eigenvalue via Geometric Representations
Sri Lankan Journal of Applied Statistics (Special Issue) Modern Statistical Methodologies in the Cutting Edge of Science Asymptotic Distribution of the Largest Eigenvalue via Geometric Representations
Information-theoretically Optimal Sparse PCA
 Information-theoretically Optimal Sparse PCA Yash Deshpande Department of Electrical Engineering Stanford, CA. Andrea Montanari Departments of Electrical Engineering and Statistics Stanford, CA. Abstract
Information-theoretically Optimal Sparse PCA Yash Deshpande Department of Electrical Engineering Stanford, CA. Andrea Montanari Departments of Electrical Engineering and Statistics Stanford, CA. Abstract
Machine Learning. Principal Components Analysis. Le Song. CSE6740/CS7641/ISYE6740, Fall 2012
 Machine Learning CSE6740/CS7641/ISYE6740, Fall 2012 Principal Components Analysis Le Song Lecture 22, Nov 13, 2012 Based on slides from Eric Xing, CMU Reading: Chap 12.1, CB book 1 2 Factor or Component
Machine Learning CSE6740/CS7641/ISYE6740, Fall 2012 Principal Components Analysis Le Song Lecture 22, Nov 13, 2012 Based on slides from Eric Xing, CMU Reading: Chap 12.1, CB book 1 2 Factor or Component
Tensor Decompositions for Machine Learning. G. Roeder 1. UBC Machine Learning Reading Group, June University of British Columbia
 Network Feature s Decompositions for Machine Learning 1 1 Department of Computer Science University of British Columbia UBC Machine Learning Group, June 15 2016 1/30 Contact information Network Feature
Network Feature s Decompositions for Machine Learning 1 1 Department of Computer Science University of British Columbia UBC Machine Learning Group, June 15 2016 1/30 Contact information Network Feature
Analysis of a Privacy-preserving PCA Algorithm using Random Matrix Theory
 Analysis of a Privacy-preserving PCA Algorithm using Random Matrix Theory Lu Wei, Anand D. Sarwate, Jukka Corander, Alfred Hero, and Vahid Tarokh Department of Electrical and Computer Engineering, University
Analysis of a Privacy-preserving PCA Algorithm using Random Matrix Theory Lu Wei, Anand D. Sarwate, Jukka Corander, Alfred Hero, and Vahid Tarokh Department of Electrical and Computer Engineering, University
This job market paper consists of three papers.
 Computational Concerns in Statistical Inference and Learning for Networ Data Analysis Tengyuan Liang Wharton School at University of Pennsylvania Abstract Networ data analysis has wide applications in
Computational Concerns in Statistical Inference and Learning for Networ Data Analysis Tengyuan Liang Wharton School at University of Pennsylvania Abstract Networ data analysis has wide applications in
Lecture 7: Positive Semidefinite Matrices
 Lecture 7: Positive Semidefinite Matrices Rajat Mittal IIT Kanpur The main aim of this lecture note is to prepare your background for semidefinite programming. We have already seen some linear algebra.
Lecture 7: Positive Semidefinite Matrices Rajat Mittal IIT Kanpur The main aim of this lecture note is to prepare your background for semidefinite programming. We have already seen some linear algebra.
Spectral k-support Norm Regularization
 Spectral k-support Norm Regularization Andrew McDonald Department of Computer Science, UCL (Joint work with Massimiliano Pontil and Dimitris Stamos) 25 March, 2015 1 / 19 Problem: Matrix Completion Goal:
Spectral k-support Norm Regularization Andrew McDonald Department of Computer Science, UCL (Joint work with Massimiliano Pontil and Dimitris Stamos) 25 March, 2015 1 / 19 Problem: Matrix Completion Goal:
Singular Value Decomposition and Principal Component Analysis (PCA) I
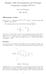 Singular Value Decomposition and Principal Component Analysis (PCA) I Prof Ned Wingreen MOL 40/50 Microarray review Data per array: 0000 genes, I (green) i,i (red) i 000 000+ data points! The expression
Singular Value Decomposition and Principal Component Analysis (PCA) I Prof Ned Wingreen MOL 40/50 Microarray review Data per array: 0000 genes, I (green) i,i (red) i 000 000+ data points! The expression
Spectral Learning of Mixture of Hidden Markov Models
 Spectral Learning of Mixture of Hidden Markov Models Y Cem Sübakan, Johannes raa, Paris Smaragdis,, Department of Computer Science, University of Illinois at Urbana-Champaign Department of Electrical and
Spectral Learning of Mixture of Hidden Markov Models Y Cem Sübakan, Johannes raa, Paris Smaragdis,, Department of Computer Science, University of Illinois at Urbana-Champaign Department of Electrical and
Lecture Notes 1: Vector spaces
 Optimization-based data analysis Fall 2017 Lecture Notes 1: Vector spaces In this chapter we review certain basic concepts of linear algebra, highlighting their application to signal processing. 1 Vector
Optimization-based data analysis Fall 2017 Lecture Notes 1: Vector spaces In this chapter we review certain basic concepts of linear algebra, highlighting their application to signal processing. 1 Vector
Linear Regression and Its Applications
 Linear Regression and Its Applications Predrag Radivojac October 13, 2014 Given a data set D = {(x i, y i )} n the objective is to learn the relationship between features and the target. We usually start
Linear Regression and Its Applications Predrag Radivojac October 13, 2014 Given a data set D = {(x i, y i )} n the objective is to learn the relationship between features and the target. We usually start
A Tutorial on Data Reduction. Principal Component Analysis Theoretical Discussion. By Shireen Elhabian and Aly Farag
 A Tutorial on Data Reduction Principal Component Analysis Theoretical Discussion By Shireen Elhabian and Aly Farag University of Louisville, CVIP Lab November 2008 PCA PCA is A backbone of modern data
A Tutorial on Data Reduction Principal Component Analysis Theoretical Discussion By Shireen Elhabian and Aly Farag University of Louisville, CVIP Lab November 2008 PCA PCA is A backbone of modern data
High-dimensional covariance estimation based on Gaussian graphical models
 High-dimensional covariance estimation based on Gaussian graphical models Shuheng Zhou Department of Statistics, The University of Michigan, Ann Arbor IMA workshop on High Dimensional Phenomena Sept. 26,
High-dimensional covariance estimation based on Gaussian graphical models Shuheng Zhou Department of Statistics, The University of Michigan, Ann Arbor IMA workshop on High Dimensional Phenomena Sept. 26,
Convergence of Eigenspaces in Kernel Principal Component Analysis
 Convergence of Eigenspaces in Kernel Principal Component Analysis Shixin Wang Advanced machine learning April 19, 2016 Shixin Wang Convergence of Eigenspaces April 19, 2016 1 / 18 Outline 1 Motivation
Convergence of Eigenspaces in Kernel Principal Component Analysis Shixin Wang Advanced machine learning April 19, 2016 Shixin Wang Convergence of Eigenspaces April 19, 2016 1 / 18 Outline 1 Motivation
Randomized Algorithms
 Randomized Algorithms Saniv Kumar, Google Research, NY EECS-6898, Columbia University - Fall, 010 Saniv Kumar 9/13/010 EECS6898 Large Scale Machine Learning 1 Curse of Dimensionality Gaussian Mixture Models
Randomized Algorithms Saniv Kumar, Google Research, NY EECS-6898, Columbia University - Fall, 010 Saniv Kumar 9/13/010 EECS6898 Large Scale Machine Learning 1 Curse of Dimensionality Gaussian Mixture Models
Deep Learning Basics Lecture 7: Factor Analysis. Princeton University COS 495 Instructor: Yingyu Liang
 Deep Learning Basics Lecture 7: Factor Analysis Princeton University COS 495 Instructor: Yingyu Liang Supervised v.s. Unsupervised Math formulation for supervised learning Given training data x i, y i
Deep Learning Basics Lecture 7: Factor Analysis Princeton University COS 495 Instructor: Yingyu Liang Supervised v.s. Unsupervised Math formulation for supervised learning Given training data x i, y i
SVD, Power method, and Planted Graph problems (+ eigenvalues of random matrices)
 Chapter 14 SVD, Power method, and Planted Graph problems (+ eigenvalues of random matrices) Today we continue the topic of low-dimensional approximation to datasets and matrices. Last time we saw the singular
Chapter 14 SVD, Power method, and Planted Graph problems (+ eigenvalues of random matrices) Today we continue the topic of low-dimensional approximation to datasets and matrices. Last time we saw the singular
Unsupervised Learning
 2018 EE448, Big Data Mining, Lecture 7 Unsupervised Learning Weinan Zhang Shanghai Jiao Tong University http://wnzhang.net http://wnzhang.net/teaching/ee448/index.html ML Problem Setting First build and
2018 EE448, Big Data Mining, Lecture 7 Unsupervised Learning Weinan Zhang Shanghai Jiao Tong University http://wnzhang.net http://wnzhang.net/teaching/ee448/index.html ML Problem Setting First build and
Concentration Inequalities for Random Matrices
 Concentration Inequalities for Random Matrices M. Ledoux Institut de Mathématiques de Toulouse, France exponential tail inequalities classical theme in probability and statistics quantify the asymptotic
Concentration Inequalities for Random Matrices M. Ledoux Institut de Mathématiques de Toulouse, France exponential tail inequalities classical theme in probability and statistics quantify the asymptotic
of Orthogonal Matching Pursuit
 A Sharp Restricted Isometry Constant Bound of Orthogonal Matching Pursuit Qun Mo arxiv:50.0708v [cs.it] 8 Jan 205 Abstract We shall show that if the restricted isometry constant (RIC) δ s+ (A) of the measurement
A Sharp Restricted Isometry Constant Bound of Orthogonal Matching Pursuit Qun Mo arxiv:50.0708v [cs.it] 8 Jan 205 Abstract We shall show that if the restricted isometry constant (RIC) δ s+ (A) of the measurement
Fundamental limits of symmetric low-rank matrix estimation
 Proceedings of Machine Learning Research vol 65:1 5, 2017 Fundamental limits of symmetric low-rank matrix estimation Marc Lelarge Safran Tech, Magny-Les-Hameaux, France Département d informatique, École
Proceedings of Machine Learning Research vol 65:1 5, 2017 Fundamental limits of symmetric low-rank matrix estimation Marc Lelarge Safran Tech, Magny-Les-Hameaux, France Département d informatique, École
Homework 1. Yuan Yao. September 18, 2011
 Homework 1 Yuan Yao September 18, 2011 1. Singular Value Decomposition: The goal of this exercise is to refresh your memory about the singular value decomposition and matrix norms. A good reference to
Homework 1 Yuan Yao September 18, 2011 1. Singular Value Decomposition: The goal of this exercise is to refresh your memory about the singular value decomposition and matrix norms. A good reference to
Stein s Method for Matrix Concentration
 Stein s Method for Matrix Concentration Lester Mackey Collaborators: Michael I. Jordan, Richard Y. Chen, Brendan Farrell, and Joel A. Tropp University of California, Berkeley California Institute of Technology
Stein s Method for Matrix Concentration Lester Mackey Collaborators: Michael I. Jordan, Richard Y. Chen, Brendan Farrell, and Joel A. Tropp University of California, Berkeley California Institute of Technology
Robustness of Principal Components
 PCA for Clustering An objective of principal components analysis is to identify linear combinations of the original variables that are useful in accounting for the variation in those original variables.
PCA for Clustering An objective of principal components analysis is to identify linear combinations of the original variables that are useful in accounting for the variation in those original variables.
Singular Value Decomposition and its. SVD and its Applications in Computer Vision
 Singular Value Decomposition and its Applications in Computer Vision Subhashis Banerjee Department of Computer Science and Engineering IIT Delhi October 24, 2013 Overview Linear algebra basics Singular
Singular Value Decomposition and its Applications in Computer Vision Subhashis Banerjee Department of Computer Science and Engineering IIT Delhi October 24, 2013 Overview Linear algebra basics Singular
1-Bit Matrix Completion
 1-Bit Matrix Completion Mark A. Davenport School of Electrical and Computer Engineering Georgia Institute of Technology Yaniv Plan Mary Wootters Ewout van den Berg Matrix Completion d When is it possible
1-Bit Matrix Completion Mark A. Davenport School of Electrical and Computer Engineering Georgia Institute of Technology Yaniv Plan Mary Wootters Ewout van den Berg Matrix Completion d When is it possible
Tensor Methods for Feature Learning
 Tensor Methods for Feature Learning Anima Anandkumar U.C. Irvine Feature Learning For Efficient Classification Find good transformations of input for improved classification Figures used attributed to
Tensor Methods for Feature Learning Anima Anandkumar U.C. Irvine Feature Learning For Efficient Classification Find good transformations of input for improved classification Figures used attributed to
Sum-of-Squares Method, Tensor Decomposition, Dictionary Learning
 Sum-of-Squares Method, Tensor Decomposition, Dictionary Learning David Steurer Cornell Approximation Algorithms and Hardness, Banff, August 2014 for many problems (e.g., all UG-hard ones): better guarantees
Sum-of-Squares Method, Tensor Decomposition, Dictionary Learning David Steurer Cornell Approximation Algorithms and Hardness, Banff, August 2014 for many problems (e.g., all UG-hard ones): better guarantees
Tutorial on Principal Component Analysis
 Tutorial on Principal Component Analysis Copyright c 1997, 2003 Javier R. Movellan. This is an open source document. Permission is granted to copy, distribute and/or modify this document under the terms
Tutorial on Principal Component Analysis Copyright c 1997, 2003 Javier R. Movellan. This is an open source document. Permission is granted to copy, distribute and/or modify this document under the terms
METHOD OF MOMENTS LEARNING FOR LEFT TO RIGHT HIDDEN MARKOV MODELS
 METHOD OF MOMENTS LEARNING FOR LEFT TO RIGHT HIDDEN MARKOV MODELS Y. Cem Subakan, Johannes Traa, Paris Smaragdis,,, Daniel Hsu UIUC Computer Science Department, Columbia University Computer Science Department
METHOD OF MOMENTS LEARNING FOR LEFT TO RIGHT HIDDEN MARKOV MODELS Y. Cem Subakan, Johannes Traa, Paris Smaragdis,,, Daniel Hsu UIUC Computer Science Department, Columbia University Computer Science Department
IEEE TRANSACTIONS ON INFORMATION THEORY, VOL. 57, NO. 11, NOVEMBER On the Performance of Sparse Recovery
 IEEE TRANSACTIONS ON INFORMATION THEORY, VOL. 57, NO. 11, NOVEMBER 2011 7255 On the Performance of Sparse Recovery Via `p-minimization (0 p 1) Meng Wang, Student Member, IEEE, Weiyu Xu, and Ao Tang, Senior
IEEE TRANSACTIONS ON INFORMATION THEORY, VOL. 57, NO. 11, NOVEMBER 2011 7255 On the Performance of Sparse Recovery Via `p-minimization (0 p 1) Meng Wang, Student Member, IEEE, Weiyu Xu, and Ao Tang, Senior
Fourier PCA. Navin Goyal (MSR India), Santosh Vempala (Georgia Tech) and Ying Xiao (Georgia Tech)
 Fourier PCA Navin Goyal (MSR India), Santosh Vempala (Georgia Tech) and Ying Xiao (Georgia Tech) Introduction 1. Describe a learning problem. 2. Develop an efficient tensor decomposition. Independent component
Fourier PCA Navin Goyal (MSR India), Santosh Vempala (Georgia Tech) and Ying Xiao (Georgia Tech) Introduction 1. Describe a learning problem. 2. Develop an efficient tensor decomposition. Independent component
COMMUNITY DETECTION IN SPARSE NETWORKS VIA GROTHENDIECK S INEQUALITY
 COMMUNITY DETECTION IN SPARSE NETWORKS VIA GROTHENDIECK S INEQUALITY OLIVIER GUÉDON AND ROMAN VERSHYNIN Abstract. We present a simple and flexible method to prove consistency of semidefinite optimization
COMMUNITY DETECTION IN SPARSE NETWORKS VIA GROTHENDIECK S INEQUALITY OLIVIER GUÉDON AND ROMAN VERSHYNIN Abstract. We present a simple and flexible method to prove consistency of semidefinite optimization
Unsupervised Kernel Dimension Reduction Supplemental Material
 Unsupervised Kernel Dimension Reduction Supplemental Material Meihong Wang Dept. of Computer Science U. of Southern California Los Angeles, CA meihongw@usc.edu Fei Sha Dept. of Computer Science U. of Southern
Unsupervised Kernel Dimension Reduction Supplemental Material Meihong Wang Dept. of Computer Science U. of Southern California Los Angeles, CA meihongw@usc.edu Fei Sha Dept. of Computer Science U. of Southern
Upper Bound for Intermediate Singular Values of Random Sub-Gaussian Matrices 1
 Upper Bound for Intermediate Singular Values of Random Sub-Gaussian Matrices 1 Feng Wei 2 University of Michigan July 29, 2016 1 This presentation is based a project under the supervision of M. Rudelson.
Upper Bound for Intermediate Singular Values of Random Sub-Gaussian Matrices 1 Feng Wei 2 University of Michigan July 29, 2016 1 This presentation is based a project under the supervision of M. Rudelson.
Provable Alternating Minimization Methods for Non-convex Optimization
 Provable Alternating Minimization Methods for Non-convex Optimization Prateek Jain Microsoft Research, India Joint work with Praneeth Netrapalli, Sujay Sanghavi, Alekh Agarwal, Animashree Anandkumar, Rashish
Provable Alternating Minimization Methods for Non-convex Optimization Prateek Jain Microsoft Research, India Joint work with Praneeth Netrapalli, Sujay Sanghavi, Alekh Agarwal, Animashree Anandkumar, Rashish
Estimation of large dimensional sparse covariance matrices
 Estimation of large dimensional sparse covariance matrices Department of Statistics UC, Berkeley May 5, 2009 Sample covariance matrix and its eigenvalues Data: n p matrix X n (independent identically distributed)
Estimation of large dimensional sparse covariance matrices Department of Statistics UC, Berkeley May 5, 2009 Sample covariance matrix and its eigenvalues Data: n p matrix X n (independent identically distributed)
