Online appendix to accompany:
|
|
|
- Ashley Payne
- 5 years ago
- Views:
Transcription
1 Online appendix to accompany: Preacher, K. J., & Hancock, G. R. (submitted). Meaningful aspects of change as novel random coefficients: A general method for reparameterizing longitudinal models. Contents 1. Mplus syntax for a reparameterized inverse quadratic function 2. Mplus syntax for an unconditional bilinear spline model 3. Mplus syntax for a conditional bilinear spline model 4. Mplus syntax for a negative exponential function with intercept, asymptote, and rate 5. Mplus syntax for a negative exponential function with intercept, rhalf-life, and ARC 6. Mathematica code for obtaining partial derivatives of an exponential function 7. Mplus syntax for a quadratic function fit to Kanfer & Ackerman's data 8. Derivation of Equation 11: Reparameterization of a bilinear spline target function 9. Annotated output: inverse quadratic function 10. Annotated output: unconditional bilinear spline model 11. Annotated output: conditional bilinear spline model 12. Annotated output: negative exponential function with intercept, asymptote, and rate 13. Annotated output: negative exponential function with intercept, half-life, and ARC 14. Annotated output: quadratic function fit to Kanfer & Ackerman s data (traditional) 15. Annotated output: quadratic function fit to Kanfer & Ackerman s data (modified) 1. Mplus syntax for a reparameterized inverse quadratic function The data may be obtained from Marie Davidian's website: The data file should be transformed to wide format prior to analysis.) TITLE: davidian theophylline data, inverse quadratic; DATA: FILE IS theophylline_wide.dat; data file in ascii format VARIABLE: NAMES ARE subject dose wt t1-t11 conc1-conc11; all 25 variables subject: subject ID dose: oral dose of theophylline in mg/kg wt: subject weight in kg t1-t11: time since administration of dose in hours conc1-conc11: theophylline concentration in mg/l USEVARIABLES ARE conc1-conc11; we plan to model 11 variables CONSTRAINT ARE t1-t11; individual-specific time scores for use in loadings ANALYSIS: ESTIMATOR=ML; ITERATIONS=2000; CONVERGENCE=.001; estimation options BOOTSTRAP=5000; for bootstrap confidence intervals if desired MODEL: [conc1-conc11*](z1-z11); mean trend (z's defined below) conc1-conc11*.7(ve); residual variances constrained equal conc1-conc10 PWITH conc2-conc11*.2(ve2); optional lag-1 residual covariances b*.002(vb); c@0; m*.5(vm); random coeff. variances (only for maximizer and theta1) b WITH c@0; b WITH m*; c WITH m@0; random coefficient covariances [b@0 c@0 m@0]; factor means constrained to zero b conc1*(b1); b conc2-conc11*(b2-b11); loadings for the theta1 parameter (b) c conc1*(c1); c conc2-conc11*(c2-c11); loadings for the theta2 parameter (c) m conc1*(m1); m conc2-conc11*(m2-m11); loadings for the maximizer parameter (m)
2 MODEL CONSTRAINT: NEW(mb*.07 mc*.02 mm*1.77); starts for theta1, theta2, and maximizer parameters intercepts constrained equal to target function z1=t1/(mb*t1+mc*(mm^2+t1^2)); z2=t2/(mb*t2+mc*(mm^2+t2^2)); z3=t3/(mb*t3+mc*(mm^2+t3^2)); z4=t4/(mb*t4+mc*(mm^2+t4^2)); z5=t5/(mb*t5+mc*(mm^2+t5^2)); z6=t6/(mb*t6+mc*(mm^2+t6^2)); z7=t7/(mb*t7+mc*(mm^2+t7^2)); z8=t8/(mb*t8+mc*(mm^2+t8^2)); z9=t9/(mb*t9+mc*(mm^2+t9^2)); z10=t10/(mb*t10+mc*(mm^2+t10^2)); z11=t11/(mb*t11+mc*(mm^2+t11^2)); loadings for "theta1" factor, constrained equal to d(y)/d(mb) b1=-1*t1^2/(mb*t1+mc*(mm^2+t1^2))^2; b2=-1*t2^2/(mb*t2+mc*(mm^2+t2^2))^2; b3=-1*t3^2/(mb*t3+mc*(mm^2+t3^2))^2; b4=-1*t4^2/(mb*t4+mc*(mm^2+t4^2))^2; b5=-1*t5^2/(mb*t5+mc*(mm^2+t5^2))^2; b6=-1*t6^2/(mb*t6+mc*(mm^2+t6^2))^2; b7=-1*t7^2/(mb*t7+mc*(mm^2+t7^2))^2; b8=-1*t8^2/(mb*t8+mc*(mm^2+t8^2))^2; b9=-1*t9^2/(mb*t9+mc*(mm^2+t9^2))^2; b10=-1*t10^2/(mb*t10+mc*(mm^2+t10^2))^2; b11=-1*t11^2/(mb*t11+mc*(mm^2+t11^2))^2; loadings for "theta2" factor, constrained equal to d(y)/d(mc) c1=-1*t1*(mm^2+t1^2)/(mb*t1+mc*(mm^2+t1^2))^2; not strictly necessary c2=-1*t2*(mm^2+t2^2)/(mb*t2+mc*(mm^2+t2^2))^2; here because this c3=-1*t3*(mm^2+t3^2)/(mb*t3+mc*(mm^2+t3^2))^2; coefficient is fixed, c4=-1*t4*(mm^2+t4^2)/(mb*t4+mc*(mm^2+t4^2))^2; but included for c5=-1*t5*(mm^2+t5^2)/(mb*t5+mc*(mm^2+t5^2))^2; completeness. c6=-1*t6*(mm^2+t6^2)/(mb*t6+mc*(mm^2+t6^2))^2; c7=-1*t7*(mm^2+t7^2)/(mb*t7+mc*(mm^2+t7^2))^2; c8=-1*t8*(mm^2+t8^2)/(mb*t8+mc*(mm^2+t8^2))^2; c9=-1*t9*(mm^2+t9^2)/(mb*t9+mc*(mm^2+t9^2))^2; c10=-1*t10*(mm^2+t10^2)/(mb*t10+mc*(mm^2+t10^2))^2; c11=-1*t11*(mm^2+t11^2)/(mb*t11+mc*(mm^2+t11^2))^2; loadings for "maximizer" factor, constrained equal to d(y)/d(mm) m1=-2*mm*mc*t1/(mb*t1+mc*(mm^2+t1^2))^2; m2=-2*mm*mc*t2/(mb*t2+mc*(mm^2+t2^2))^2; m3=-2*mm*mc*t3/(mb*t3+mc*(mm^2+t3^2))^2; m4=-2*mm*mc*t4/(mb*t4+mc*(mm^2+t4^2))^2; m5=-2*mm*mc*t5/(mb*t5+mc*(mm^2+t5^2))^2; m6=-2*mm*mc*t6/(mb*t6+mc*(mm^2+t6^2))^2; m7=-2*mm*mc*t7/(mb*t7+mc*(mm^2+t7^2))^2; m8=-2*mm*mc*t8/(mb*t8+mc*(mm^2+t8^2))^2; m9=-2*mm*mc*t9/(mb*t9+mc*(mm^2+t9^2))^2; m10=-2*mm*mc*t10/(mb*t10+mc*(mm^2+t10^2))^2; m11=-2*mm*mc*t11/(mb*t11+mc*(mm^2+t11^2))^2; OUTPUT: TECH1; CINTERVAL(BCBOOTSTRAP); 2. Mplus syntax for an unconditional bilinear spline model TITLE: zerbe glucose;
3 DATA: FILE IS glucose.dat; data file in ascii format VARIABLE: NAMES ARE obese ph1-ph8; all 9 variables obese: obesity status, 0=control, 1=obese ph1-ph8: plasma phosphate concentration repeated measures USEVARIABLES ARE ph1-ph8; we plan to model 8 variables ANALYSIS: BOOTSTRAP IS 1000; for bootstrap confidence intervals if desired MODEL: [ph1-ph8*](z1-z8); mean trend (z's defined below) ph1-ph8*(ve); residual variances constrained equal w1-w4*; random coefficient variances w1 WITH w2*; w1 WITH w3*; w1 WITH w4*; random coefficient covariances w2 WITH w3*; w2 WITH w4*; w3 WITH w4*; random coefficient covariances [w1@0 w2@0 w3@0 w4@0]; factor means constrained to zero w1 ph1-ph8@1; loadings for 1st factor, constrained to 1 w2 ph1*(w21); w2 ph2-ph8(w22-w28); loadings for 2nd factor, defined below w3 ph1*(w31); w3 ph2-ph8(w32-w38); loadings for 3rd factor, defined below w4 ph1*(w41); w4 ph2-ph8(w42-w48); loadings for knot factor, defined below MODEL CONSTRAINT: starts for parameters of the target function NEW(mw1*3.4 mw2*-.26 mw3*.54 mw4*1.6 b1-b4); intercepts constrained equal to target function z1=mw1+mw2*0.0+mw3*sqrt((0.0-mw4)^2); z2=mw1+mw2*0.5+mw3*sqrt((0.5-mw4)^2); z3=mw1+mw2*1.0+mw3*sqrt((1.0-mw4)^2); z4=mw1+mw2*1.5+mw3*sqrt((1.5-mw4)^2); z5=mw1+mw2*2.0+mw3*sqrt((2.0-mw4)^2); z6=mw1+mw2*3.0+mw3*sqrt((3.0-mw4)^2); z7=mw1+mw2*4.0+mw3*sqrt((4.0-mw4)^2); z8=mw1+mw2*5.0+mw3*sqrt((5.0-mw4)^2); loadings for 2nd and 3rd factors, constrained to d(y)/d(mw2) and d(y)/d(mw3) w21=0.0; w31=sqrt((0.0-mw4)^2); w22=0.5; w32=sqrt((0.5-mw4)^2); w23=1.0; w33=sqrt((1.0-mw4)^2); w24=1.5; w34=sqrt((1.5-mw4)^2); w25=2.0; w35=sqrt((2.0-mw4)^2); w26=3.0; w36=sqrt((3.0-mw4)^2); w27=4.0; w37=sqrt((4.0-mw4)^2); w28=5.0; w38=sqrt((5.0-mw4)^2); loadings for knot factor, constrained to d(y)/d(mw4) w41=(mw3*(mw4-0.0))/sqrt((0.0-mw4)^2); w42=(mw3*(mw4-0.5))/sqrt((0.5-mw4)^2); w43=(mw3*(mw4-1.0))/sqrt((1.0-mw4)^2); w44=(mw3*(mw4-1.5))/sqrt((1.5-mw4)^2); w45=(mw3*(mw4-2.0))/sqrt((2.0-mw4)^2); w46=(mw3*(mw4-3.0))/sqrt((3.0-mw4)^2); w47=(mw3*(mw4-4.0))/sqrt((4.0-mw4)^2); w48=(mw3*(mw4-5.0))/sqrt((5.0-mw4)^2); parameters of original parameterization b1=mw1+mw3*mw4; b2=mw2-mw3; b3=mw1-mw3*mw4; b4=mw2+mw3; OUTPUT: TECH1; CINTERVAL(BCBOOTSTRAP);
4 Zerbe's (1979) data (N = 33), obtained from Zerbe (1979, p. 219) Mplus syntax for a conditional bilinear spline model The data may be obtained from Zerbe (1979, p. 219). TITLE: zerbe glucose; DATA: FILE IS glucose.dat; data file in ascii format DEFINE: xobese=abs(obese-1); 1=control; 0=obese VARIABLE: NAMES ARE obese ph1-ph8; all 9 variables obese: obesity status, 0=control, 1=obese ph1-ph8: plasma phosphate concentration repeated measures USEVARIABLES ARE ph1-ph8 xobese; we plan to model 9 variables ANALYSIS: BOOTSTRAP IS 1000; for bootstrap confidence intervals if desired MODEL: [ph1-ph8*](z1-z8); mean trend (z's defined below) ph1-ph8*(ve); residual variances constrained equal w1-w4*; random coefficient residual variances w1 WITH w2*; w1 WITH w3*; w1 WITH w4*; random coefficient residual covariances w2 WITH w3*; w2 WITH w4*; w3 WITH w4*; random coefficient residual covariances [w1@0 w2@0 w3@0 w4@0]; factor intercepts constrained to zero w1 ph1-ph8@1; loadings for 1st factor, constrained to 1 w2 ph1*(w21); w2 ph2-ph8(w22-w28); loadings for 2nd factor, defined below w3 ph1*(w31); w3 ph2-ph8(w32-w38); loadings for 3rd factor, defined below w4 ph1*(w41); w4 ph2-ph8(w42-w48); loadings for knot factor, defined below
5 w1-w4 ON xobese; regress random coefficients on obese. MODEL CONSTRAINT: starts for parameters of the target function NEW(mw1*3.5 mw2*-.21 mw3*.49 mw4*1.95 b1-b4); intercepts constrained equal to target function z1=mw1+mw2*0.0+mw3*sqrt((0.0-mw4)^2); z2=mw1+mw2*0.5+mw3*sqrt((0.5-mw4)^2); z3=mw1+mw2*1.0+mw3*sqrt((1.0-mw4)^2); z4=mw1+mw2*1.5+mw3*sqrt((1.5-mw4)^2); z5=mw1+mw2*2.0+mw3*sqrt((2.0-mw4)^2); z6=mw1+mw2*3.0+mw3*sqrt((3.0-mw4)^2); z7=mw1+mw2*4.0+mw3*sqrt((4.0-mw4)^2); z8=mw1+mw2*5.0+mw3*sqrt((5.0-mw4)^2); loadings for 2nd and 3rd factors, constrained to d(y)/d(mw2) and d(y)/d(mw3) w21=0.0; w31=sqrt((0.0-mw4)^2); w22=0.5; w32=sqrt((0.5-mw4)^2); w23=1.0; w33=sqrt((1.0-mw4)^2); w24=1.5; w34=sqrt((1.5-mw4)^2); w25=2.0; w35=sqrt((2.0-mw4)^2); w26=3.0; w36=sqrt((3.0-mw4)^2); w27=4.0; w37=sqrt((4.0-mw4)^2); w28=5.0; w38=sqrt((5.0-mw4)^2); loadings for knot factor, constrained to d(y)/d(mw4) w41=(mw3*(mw4-0.0))/sqrt((0.0-mw4)^2); w42=(mw3*(mw4-0.5))/sqrt((0.5-mw4)^2); w43=(mw3*(mw4-1.0))/sqrt((1.0-mw4)^2); w44=(mw3*(mw4-1.5))/sqrt((1.5-mw4)^2); w45=(mw3*(mw4-2.0))/sqrt((2.0-mw4)^2); w46=(mw3*(mw4-3.0))/sqrt((3.0-mw4)^2); w47=(mw3*(mw4-4.0))/sqrt((4.0-mw4)^2); w48=(mw3*(mw4-5.0))/sqrt((5.0-mw4)^2); parameters of original parameterization b1=mw1+mw3*mw4; b2=mw2-mw3; b3=mw1-mw3*mw4; b4=mw2+mw3; OUTPUT: TECH1; CINTERVAL(BCBOOTSTRAP); 4. Mplus syntax for a negative exponential function with intercept, asymptote, and rate Chaiken's data are included in installations of LISREL: TITLE: chaiken, negative exponential, original parameterization; DATA: FILE IS schaiken2.dat; data file in ascii format VARIABLE: NAMES ARE v1-v12 q1-q12; all 24 variables v1-v12: verbal skill acquisition repeated measures q1-q12: quantitative skill acquisition repeated measures ANALYSIS: ESTIMATOR IS ML; use maximum likelihood estimation MODEL: verbal [v1-v12*](tv1-tv12); mean trend for verbal v1-v12*1.97(vv1); residual variances constrained equal
6 v1-v11 PWITH v2-v12*(dv1); lag-1 error covariance v1-v10 PWITH v3-v12*(dv2); lag-2, etc... v1-v9 PWITH v4-v12*(dv3); v1-v8 PWITH v5-v12*(dv4); v1-v7 PWITH v6-v12*(dv5); v1-v6 PWITH v7-v12*(dv6); v1-v5 PWITH v8-v12*(dv7); v1-v4 PWITH v9-v12*(dv8); v1-v3 PWITH v10-v12*(dv9); v1-v2 PWITH v11-v12*(dv10); v1 WITH v12*(dv11); factor means constrained to zero av*; bv*; cv*; random coefficient variances av WITH bv* cv*; bv WITH cv*; random coefficient covariances av v1*(av1); av v2-v12(av2-av12); loadings for 1st factor bv v1*(bv1); bv v2-v12(bv2-bv12); loadings for 2nd factor cv v1*(cv1); cv v2-v12(cv2-cv12); loadings for 3rd factor quantitative [q1-q12*](tq1-tq12); mean trend for quantitative q1-q12*1.21(vq1); residual variances constrained equal q1-q11 PWITH q2-q12*(dq1); lag-1 error covariance q1-q10 PWITH q3-q12*(dq2); lat-2, etc... q1-q9 PWITH q4-q12*(dq3); q1-q8 PWITH q5-q12*(dq4); q1-q7 PWITH q6-q12*(dq5); q1-q6 PWITH q7-q12*(dq6); q1-q5 PWITH q8-q12*(dq7); q1-q4 PWITH q9-q12*(dq8); q1-q3 PWITH q10-q12*(dq9); q1-q2 PWITH q11-q12*(dq10); q1 WITH q12*(dq11); factor means constrained to zero aq*; bq*; cq*; random coefficient variances aq WITH bq* cq*; bq WITH cq; random coefficient covariances aq q1*(aq1); aq q2-q12(aq2-aq12); loadings for 1st factor bq q1*(bq1); bq q2-q12(bq2-bq12); loadings for 2nd factor cq q1*(cq1); cq q2-q12(cq2-cq12); loadings for 3rd factor cross-domain covariances av WITH aq* bq*; bv WITH aq* bq*; MODEL CONSTRAINT: starts for parameters of the target function and for lagged error covariances NEW(mav*6.8 mbv*21 mcv*.7 maq*8.6 mbq*16.5 mcq*.7 rhov*.27 rhoq*.31); error covariances dv1= vv1*rhov; dq1= vq1*rhoq; dv2= vv1*rhov^2; dq2= vq1*rhoq^2; dv3= vv1*rhov^3; dq3= vq1*rhoq^3; dv4= vv1*rhov^4; dq4= vq1*rhoq^4; dv5= vv1*rhov^5; dq5= vq1*rhoq^5; dv6= vv1*rhov^6; dq6= vq1*rhoq^6; dv7= vv1*rhov^7; dq7= vq1*rhoq^7; dv8= vv1*rhov^8; dq8= vq1*rhoq^8; dv9= vv1*rhov^9; dq9= vq1*rhoq^9; dv10=vv1*rhov^10; dq10=vq1*rhoq^10; dv11=vv1*rhov^11; dq11=vq1*rhoq^11; verbal and quantitative intercepts constrained equal to target function tv1 =mav-(mav-mbv)*exp(-1*mcv*( 1-1)); tq1 =maq-(maq-mbq)*exp(-1*mcq*( 1-1)); tv2 =mav-(mav-mbv)*exp(-1*mcv*( 2-1)); tq2 =maq-(maq-mbq)*exp(-1*mcq*( 2-1));
7 tv3 =mav-(mav-mbv)*exp(-1*mcv*( 3-1)); tq3 =maq-(maq-mbq)*exp(-1*mcq*( 3-1)); tv4 =mav-(mav-mbv)*exp(-1*mcv*( 4-1)); tq4 =maq-(maq-mbq)*exp(-1*mcq*( 4-1)); tv5 =mav-(mav-mbv)*exp(-1*mcv*( 5-1)); tq5 =maq-(maq-mbq)*exp(-1*mcq*( 5-1)); tv6 =mav-(mav-mbv)*exp(-1*mcv*( 6-1)); tq6 =maq-(maq-mbq)*exp(-1*mcq*( 6-1)); tv7 =mav-(mav-mbv)*exp(-1*mcv*( 7-1)); tq7 =maq-(maq-mbq)*exp(-1*mcq*( 7-1)); tv8 =mav-(mav-mbv)*exp(-1*mcv*( 8-1)); tq8 =maq-(maq-mbq)*exp(-1*mcq*( 8-1)); tv9 =mav-(mav-mbv)*exp(-1*mcv*( 9-1)); tq9 =maq-(maq-mbq)*exp(-1*mcq*( 9-1)); tv10=mav-(mav-mbv)*exp(-1*mcv*(10-1)); tq10=maq-(maq-mbq)*exp(-1*mcq*(10-1)); tv11=mav-(mav-mbv)*exp(-1*mcv*(11-1)); tq11=maq-(maq-mbq)*exp(-1*mcq*(11-1)); tv12=mav-(mav-mbv)*exp(-1*mcv*(12-1)); tq12=maq-(maq-mbq)*exp(-1*mcq*(12-1)); loadings on 1st factor, constrained to d(y)/d(mav) and d(y)/d(maq) av1 =1-exp(-1*mcv*( 1-1)); aq1 =1-exp(-1*mcq*( 1-1)); av2 =1-exp(-1*mcv*( 2-1)); aq2 =1-exp(-1*mcq*( 2-1)); av3 =1-exp(-1*mcv*( 3-1)); aq3 =1-exp(-1*mcq*( 3-1)); av4 =1-exp(-1*mcv*( 4-1)); aq4 =1-exp(-1*mcq*( 4-1)); av5 =1-exp(-1*mcv*( 5-1)); aq5 =1-exp(-1*mcq*( 5-1)); av6 =1-exp(-1*mcv*( 6-1)); aq6 =1-exp(-1*mcq*( 6-1)); av7 =1-exp(-1*mcv*( 7-1)); aq7 =1-exp(-1*mcq*( 7-1)); av8 =1-exp(-1*mcv*( 8-1)); aq8 =1-exp(-1*mcq*( 8-1)); av9 =1-exp(-1*mcv*( 9-1)); aq9 =1-exp(-1*mcq*( 9-1)); av10=1-exp(-1*mcv*(10-1)); aq10=1-exp(-1*mcq*(10-1)); av11=1-exp(-1*mcv*(11-1)); aq11=1-exp(-1*mcq*(11-1)); av12=1-exp(-1*mcv*(12-1)); aq12=1-exp(-1*mcq*(12-1)); loadings on 2nd and 3rd factors for verbal, constrained to d(y)/d(mbv) and d(y)/d(mcv) bv1 =exp(-1*mcv*( 1-1)); cv1 =(mav-mbv)*( 1-1)*exp(-1*mcv*( 1-1)); bv2 =exp(-1*mcv*( 2-1)); cv2 =(mav-mbv)*( 2-1)*exp(-1*mcv*( 2-1)); bv3 =exp(-1*mcv*( 3-1)); cv3 =(mav-mbv)*( 3-1)*exp(-1*mcv*( 3-1)); bv4 =exp(-1*mcv*( 4-1)); cv4 =(mav-mbv)*( 4-1)*exp(-1*mcv*( 4-1)); bv5 =exp(-1*mcv*( 5-1)); cv5 =(mav-mbv)*( 5-1)*exp(-1*mcv*( 5-1)); bv6 =exp(-1*mcv*( 6-1)); cv6 =(mav-mbv)*( 6-1)*exp(-1*mcv*( 6-1)); bv7 =exp(-1*mcv*( 7-1)); cv7 =(mav-mbv)*( 7-1)*exp(-1*mcv*( 7-1)); bv8 =exp(-1*mcv*( 8-1)); cv8 =(mav-mbv)*( 8-1)*exp(-1*mcv*( 8-1)); bv9 =exp(-1*mcv*( 9-1)); cv9 =(mav-mbv)*( 9-1)*exp(-1*mcv*( 9-1)); bv10=exp(-1*mcv*(10-1)); cv10=(mav-mbv)*(10-1)*exp(-1*mcv*(10-1)); bv11=exp(-1*mcv*(11-1)); cv11=(mav-mbv)*(11-1)*exp(-1*mcv*(11-1)); bv12=exp(-1*mcv*(12-1)); cv12=(mav-mbv)*(12-1)*exp(-1*mcv*(12-1)); loadings on 2nd and 3rd factors for quant., constrained to d(y)/d(mbv) and d(y)/d(mcv) bq1 =exp(-1*mcq*( 1-1)); cq1 =(maq-mbq)*( 1-1)*exp(-1*mcq*( 1-1)); bq2 =exp(-1*mcq*( 2-1)); cq2 =(maq-mbq)*( 2-1)*exp(-1*mcq*( 2-1)); bq3 =exp(-1*mcq*( 3-1)); cq3 =(maq-mbq)*( 3-1)*exp(-1*mcq*( 3-1)); bq4 =exp(-1*mcq*( 4-1)); cq4 =(maq-mbq)*( 4-1)*exp(-1*mcq*( 4-1)); bq5 =exp(-1*mcq*( 5-1)); cq5 =(maq-mbq)*( 5-1)*exp(-1*mcq*( 5-1)); bq6 =exp(-1*mcq*( 6-1)); cq6 =(maq-mbq)*( 6-1)*exp(-1*mcq*( 6-1)); bq7 =exp(-1*mcq*( 7-1)); cq7 =(maq-mbq)*( 7-1)*exp(-1*mcq*( 7-1)); bq8 =exp(-1*mcq*( 8-1)); cq8 =(maq-mbq)*( 8-1)*exp(-1*mcq*( 8-1)); bq9 =exp(-1*mcq*( 9-1)); cq9 =(maq-mbq)*( 9-1)*exp(-1*mcq*( 9-1)); bq10=exp(-1*mcq*(10-1)); cq10=(maq-mbq)*(10-1)*exp(-1*mcq*(10-1)); bq11=exp(-1*mcq*(11-1)); cq11=(maq-mbq)*(11-1)*exp(-1*mcq*(11-1)); bq12=exp(-1*mcq*(12-1)); cq12=(maq-mbq)*(12-1)*exp(-1*mcq*(12-1)); OUTPUT: TECH1 TECH4; 5. Mplus syntax for a negative exponential function with intercept, half-life, and ARC Chaiken's data are included in installations of LISREL: TITLE: chaiken, negative exponential with halflife and arc;
8 DATA: FILE IS schaiken2.dat; data file in ascii format VARIABLE: NAMES ARE v1-v12 q1-q12; all 24 variables v1-v12: verbal skill acquisition repeated measures q1-q12: quantitative skill acquisition repeated measures ANALYSIS: ESTIMATOR IS ML; use maximum likelihood estimation MODEL: verbal [v1-v12*](tv1-tv12); mean trend for verbal v1-v12*1.97(vv1); residual variances constrained equal v1-v11 PWITH v2-v12*(dv1); lag-1 error covariance v1-v10 PWITH v3-v12*(dv2); lag-2, etc... v1-v9 PWITH v4-v12*(dv3); v1-v8 PWITH v5-v12*(dv4); v1-v7 PWITH v6-v12*(dv5); v1-v6 PWITH v7-v12*(dv6); v1-v5 PWITH v8-v12*(dv7); v1-v4 PWITH v9-v12*(dv8); v1-v3 PWITH v10-v12*(dv9); v1-v2 PWITH v11-v12*(dv10); v1 WITH v12*(dv11); factor means constrained to zero av*1.6; bv*92; cv*10; random coefficient variances av WITH bv*3.7 cv*; bv WITH cv*; random coefficient covariances av v1*(av1); av v2-v12(av2-av12); loadings for 1st factor bv v1*(bv1); bv v2-v12(bv2-bv12); loadings for 2nd factor cv v1*(cv1); cv v2-v12(cv2-cv12); loadings for 3rd factor quantitative [q1-q12*](tq1-tq12); mean trend for quantitative q1-q12*1.21(vq1); residual variances constrained equal q1-q11 PWITH q2-q12*(dq1); lag-1 error covariance q1-q10 PWITH q3-q12*(dq2); lag-2, etc... q1-q9 PWITH q4-q12*(dq3); q1-q8 PWITH q5-q12*(dq4); q1-q7 PWITH q6-q12*(dq5); q1-q6 PWITH q7-q12*(dq6); q1-q5 PWITH q8-q12*(dq7); q1-q4 PWITH q9-q12*(dq8); q1-q3 PWITH q10-q12*(dq9); q1-q2 PWITH q11-q12*(dq10); q1 WITH q12*(dq11); factor means constrained to zero aq*4; bq*35; cq*10; random coefficient variances aq WITH bq*7.4 cq*; bq WITH cq; random coefficient covariances aq q1*(aq1); aq q2-q12(aq2-aq12); loadings for 1st factor bq q1*(bq1); bq q2-q12(bq2-bq12); loadings for 2nd factor cq q1*(cq1); cq q2-q12(cq2-cq12); loadings for 3rd factor cross-domain covariances av WITH aq* bq* cq*; bv WITH aq* bq* cq*; cv WITH aq* bq* cq*; MODEL CONSTRAINT: starts for parameters of the target function and for lagged error covariances NEW(mav*1.7 mbv*-1.5 mcv*1.96 maq*4.2 mbq*-.9 mcq*1.99 rhov*.27 rhoq*.3 diff*0); error covariances dv1= vv1*rhov; dq1= vq1*rhoq; dv2= vv1*rhov^2; dq2= vq1*rhoq^2; dv3= vv1*rhov^3; dq3= vq1*rhoq^3; dv4= vv1*rhov^4; dq4= vq1*rhoq^4; dv5= vv1*rhov^5; dq5= vq1*rhoq^5;
9 dv6= vv1*rhov^6; dq6= vq1*rhoq^6; dv7= vv1*rhov^7; dq7= vq1*rhoq^7; dv8= vv1*rhov^8; dq8= vq1*rhoq^8; dv9= vv1*rhov^9; dq9= vq1*rhoq^9; dv10=vv1*rhov^10; dq10=vq1*rhoq^10; dv11=vv1*rhov^11; dq11=vq1*rhoq^11; verbal intercepts constrained equal to target function tv1 =mav+(mbv*11*(.5^(( 1-1)/(mcv-1))))/((.5^(11/(mcv-1)))-1); tv2 =mav+(mbv*11*(.5^(( 2-1)/(mcv-1))))/((.5^(11/(mcv-1)))-1); tv3 =mav+(mbv*11*(.5^(( 3-1)/(mcv-1))))/((.5^(11/(mcv-1)))-1); tv4 =mav+(mbv*11*(.5^(( 4-1)/(mcv-1))))/((.5^(11/(mcv-1)))-1); tv5 =mav+(mbv*11*(.5^(( 5-1)/(mcv-1))))/((.5^(11/(mcv-1)))-1); tv6 =mav+(mbv*11*(.5^(( 6-1)/(mcv-1))))/((.5^(11/(mcv-1)))-1); tv7 =mav+(mbv*11*(.5^(( 7-1)/(mcv-1))))/((.5^(11/(mcv-1)))-1); tv8 =mav+(mbv*11*(.5^(( 8-1)/(mcv-1))))/((.5^(11/(mcv-1)))-1); tv9 =mav+(mbv*11*(.5^(( 9-1)/(mcv-1))))/((.5^(11/(mcv-1)))-1); tv10=mav+(mbv*11*(.5^((10-1)/(mcv-1))))/((.5^(11/(mcv-1)))-1); tv11=mav+(mbv*11*(.5^((11-1)/(mcv-1))))/((.5^(11/(mcv-1)))-1); tv12=mav+(mbv*11*(.5^((12-1)/(mcv-1))))/((.5^(11/(mcv-1)))-1); loadings on 1st factor, constrained to d(y)/d(mav) and d(y)/d(maq) 0=1-av1 ; 0=1-aq1 ; 0=1-av2 ; 0=1-aq2 ; 0=1-av3 ; 0=1-aq3 ; 0=1-av4 ; 0=1-aq4 ; 0=1-av5 ; 0=1-aq5 ; 0=1-av6 ; 0=1-aq6 ; 0=1-av7 ; 0=1-aq7 ; 0=1-av8 ; 0=1-aq8 ; 0=1-av9 ; 0=1-aq9 ; 0=1-av10; 0=1-aq10; 0=1-av11; 0=1-aq11; 0=1-av12; 0=1-aq12; loadings on 2nd factor for verbal, constrained to d(y)/d(mbv) bv1 =(11*(.5^(( 1-1)/(mcv-1))))/((.5^(11/(mcv-1)))-1); bv2 =(11*(.5^(( 2-1)/(mcv-1))))/((.5^(11/(mcv-1)))-1); bv3 =(11*(.5^(( 3-1)/(mcv-1))))/((.5^(11/(mcv-1)))-1); bv4 =(11*(.5^(( 4-1)/(mcv-1))))/((.5^(11/(mcv-1)))-1); bv5 =(11*(.5^(( 5-1)/(mcv-1))))/((.5^(11/(mcv-1)))-1); bv6 =(11*(.5^(( 6-1)/(mcv-1))))/((.5^(11/(mcv-1)))-1); bv7 =(11*(.5^(( 7-1)/(mcv-1))))/((.5^(11/(mcv-1)))-1); bv8 =(11*(.5^(( 8-1)/(mcv-1))))/((.5^(11/(mcv-1)))-1); bv9 =(11*(.5^(( 9-1)/(mcv-1))))/((.5^(11/(mcv-1)))-1); bv10=(11*(.5^((10-1)/(mcv-1))))/((.5^(11/(mcv-1)))-1); bv11=(11*(.5^((11-1)/(mcv-1))))/((.5^(11/(mcv-1)))-1); bv12=(11*(.5^((12-1)/(mcv-1))))/((.5^(11/(mcv-1)))-1); loadings on 3rd factor for verbal, constrained to d(y)/d(mcv) cv1 =(.5^( 1/(mcv-1))*mcv*((.5^(1/(mcv-1)))*( 1-1)+(.5^(12/(mcv-1)))*(12-1))*11*ln(.5))/ (((mcv-1)^2)*(((.5^(11/(mcv-1)))-(.5^(1/(mcv-1))))^2)); cv2 =(.5^( 2/(mcv-1))*mcv*((.5^(1/(mcv-1)))*( 2-1)+(.5^(12/(mcv-1)))*(12-2))*11*ln(.5))/ (((mcv-1)^2)*(((.5^(11/(mcv-1)))-(.5^(1/(mcv-1))))^2)); cv3 =(.5^( 3/(mcv-1))*mcv*((.5^(1/(mcv-1)))*( 3-1)+(.5^(12/(mcv-1)))*(12-3))*11*ln(.5))/ (((mcv-1)^2)*(((.5^(11/(mcv-1)))-(.5^(1/(mcv-1))))^2)); cv4 =(.5^( 4/(mcv-1))*mcv*((.5^(1/(mcv-1)))*( 4-1)+(.5^(12/(mcv-1)))*(12-4))*11*ln(.5))/ (((mcv-1)^2)*(((.5^(11/(mcv-1)))-(.5^(1/(mcv-1))))^2)); cv5 =(.5^( 5/(mcv-1))*mcv*((.5^(1/(mcv-1)))*( 5-1)+(.5^(12/(mcv-1)))*(12-5))*11*ln(.5))/ (((mcv-1)^2)*(((.5^(11/(mcv-1)))-(.5^(1/(mcv-1))))^2)); cv6 =(.5^( 6/(mcv-1))*mcv*((.5^(1/(mcv-1)))*( 6-1)+(.5^(12/(mcv-1)))*(12-6))*11*ln(.5))/ (((mcv-1)^2)*(((.5^(11/(mcv-1)))-(.5^(1/(mcv-1))))^2)); cv7 =(.5^( 7/(mcv-1))*mcv*((.5^(1/(mcv-1)))*( 7-1)+(.5^(12/(mcv-1)))*(12-7))*11*ln(.5))/
10 (((mcv-1)^2)*(((.5^(11/(mcv-1)))-(.5^(1/(mcv-1))))^2)); cv8 =(.5^( 8/(mcv-1))*mcv*((.5^(1/(mcv-1)))*( 8-1)+(.5^(12/(mcv-1)))*(12-8))*11*ln(.5))/ (((mcv-1)^2)*(((.5^(11/(mcv-1)))-(.5^(1/(mcv-1))))^2)); cv9 =(.5^( 9/(mcv-1))*mcv*((.5^(1/(mcv-1)))*( 9-1)+(.5^(12/(mcv-1)))*(12-9))*11*ln(.5))/ (((mcv-1)^2)*(((.5^(11/(mcv-1)))-(.5^(1/(mcv-1))))^2)); cv10=(.5^(10/(mcv-1))*mcv*((.5^(1/(mcv-1)))*(10-1)+(.5^(12/(mcv-1)))*(12-10))*11*ln(.5))/ (((mcv-1)^2)*(((.5^(11/(mcv-1)))-(.5^(1/(mcv-1))))^2)); cv11=(.5^(11/(mcv-1))*mcv*((.5^(1/(mcv-1)))*(11-1)+(.5^(12/(mcv-1)))*(12-11))*11*ln(.5))/ (((mcv-1)^2)*(((.5^(11/(mcv-1)))-(.5^(1/(mcv-1))))^2)); cv12=(.5^(12/(mcv-1))*mcv*((.5^(1/(mcv-1)))*(12-1)+(.5^(12/(mcv-1)))*(12-12))*11*ln(.5))/ (((mcv-1)^2)*(((.5^(11/(mcv-1)))-(.5^(1/(mcv-1))))^2)); quantitative intercepts constrained equal to target function tq1 =maq+(mbq*11*(.5^(( 1-1)/(mcq-1))))/((.5^(11/(mcq-1)))-1); tq2 =maq+(mbq*11*(.5^(( 2-1)/(mcq-1))))/((.5^(11/(mcq-1)))-1); tq3 =maq+(mbq*11*(.5^(( 3-1)/(mcq-1))))/((.5^(11/(mcq-1)))-1); tq4 =maq+(mbq*11*(.5^(( 4-1)/(mcq-1))))/((.5^(11/(mcq-1)))-1); tq5 =maq+(mbq*11*(.5^(( 5-1)/(mcq-1))))/((.5^(11/(mcq-1)))-1); tq6 =maq+(mbq*11*(.5^(( 6-1)/(mcq-1))))/((.5^(11/(mcq-1)))-1); tq7 =maq+(mbq*11*(.5^(( 7-1)/(mcq-1))))/((.5^(11/(mcq-1)))-1); tq8 =maq+(mbq*11*(.5^(( 8-1)/(mcq-1))))/((.5^(11/(mcq-1)))-1); tq9 =maq+(mbq*11*(.5^(( 9-1)/(mcq-1))))/((.5^(11/(mcq-1)))-1); tq10=maq+(mbq*11*(.5^((10-1)/(mcq-1))))/((.5^(11/(mcq-1)))-1); tq11=maq+(mbq*11*(.5^((11-1)/(mcq-1))))/((.5^(11/(mcq-1)))-1); tq12=maq+(mbq*11*(.5^((12-1)/(mcq-1))))/((.5^(11/(mcq-1)))-1); loadings on 2nd factor for quantitative, constrained to d(y)/d(mbv) bq1 =(11*(.5^(( 1-1)/(mcq-1))))/((.5^(11/(mcq-1)))-1); bq2 =(11*(.5^(( 2-1)/(mcq-1))))/((.5^(11/(mcq-1)))-1); bq3 =(11*(.5^(( 3-1)/(mcq-1))))/((.5^(11/(mcq-1)))-1); bq4 =(11*(.5^(( 4-1)/(mcq-1))))/((.5^(11/(mcq-1)))-1); bq5 =(11*(.5^(( 5-1)/(mcq-1))))/((.5^(11/(mcq-1)))-1); bq6 =(11*(.5^(( 6-1)/(mcq-1))))/((.5^(11/(mcq-1)))-1); bq7 =(11*(.5^(( 7-1)/(mcq-1))))/((.5^(11/(mcq-1)))-1); bq8 =(11*(.5^(( 8-1)/(mcq-1))))/((.5^(11/(mcq-1)))-1); bq9 =(11*(.5^(( 9-1)/(mcq-1))))/((.5^(11/(mcq-1)))-1); bq10=(11*(.5^((10-1)/(mcq-1))))/((.5^(11/(mcq-1)))-1); bq11=(11*(.5^((11-1)/(mcq-1))))/((.5^(11/(mcq-1)))-1); bq12=(11*(.5^((12-1)/(mcq-1))))/((.5^(11/(mcq-1)))-1); loadings on 3rd factor for quantitative, constrained to d(y)/d(mcv) cq1 =(.5^( 1/(mcq-1))*mcq*((.5^(1/(mcq-1)))*( 1-1)+(.5^(12/(mcq-1)))*(12-1))*11*ln(.5))/ (((mcq-1)^2)*(((.5^(11/(mcq-1)))-(.5^(1/(mcq-1))))^2)); cq2 =(.5^( 2/(mcq-1))*mcq*((.5^(1/(mcq-1)))*( 2-1)+(.5^(12/(mcq-1)))*(12-2))*11*ln(.5))/ (((mcq-1)^2)*(((.5^(11/(mcq-1)))-(.5^(1/(mcq-1))))^2)); cq3 =(.5^( 3/(mcq-1))*mcq*((.5^(1/(mcq-1)))*( 3-1)+(.5^(12/(mcq-1)))*(12-3))*11*ln(.5))/ (((mcq-1)^2)*(((.5^(11/(mcq-1)))-(.5^(1/(mcq-1))))^2)); cq4 =(.5^( 4/(mcq-1))*mcq*((.5^(1/(mcq-1)))*( 4-1)+(.5^(12/(mcq-1)))*(12-4))*11*ln(.5))/ (((mcq-1)^2)*(((.5^(11/(mcq-1)))-(.5^(1/(mcq-1))))^2)); cq5 =(.5^( 5/(mcq-1))*mcq*((.5^(1/(mcq-1)))*( 5-1)+(.5^(12/(mcq-1)))*(12-5))*11*ln(.5))/ (((mcq-1)^2)*(((.5^(11/(mcq-1)))-(.5^(1/(mcq-1))))^2)); cq6 =(.5^( 6/(mcq-1))*mcq*((.5^(1/(mcq-1)))*( 6-1)+(.5^(12/(mcq-1)))*(12-6))*11*ln(.5))/ (((mcq-1)^2)*(((.5^(11/(mcq-1)))-(.5^(1/(mcq-1))))^2)); cq7 =(.5^( 7/(mcq-1))*mcq*((.5^(1/(mcq-1)))*( 7-1)+(.5^(12/(mcq-1)))*(12-7))*11*ln(.5))/ (((mcq-1)^2)*(((.5^(11/(mcq-1)))-(.5^(1/(mcq-1))))^2)); cq8 =(.5^( 8/(mcq-1))*mcq*((.5^(1/(mcq-1)))*( 8-1)+(.5^(12/(mcq-1)))*(12-8))*11*ln(.5))/ (((mcq-1)^2)*(((.5^(11/(mcq-1)))-(.5^(1/(mcq-1))))^2)); cq9 =(.5^( 9/(mcq-1))*mcq*((.5^(1/(mcq-1)))*( 9-1)+(.5^(12/(mcq-1)))*(12-9))*11*ln(.5))/ (((mcq-1)^2)*(((.5^(11/(mcq-1)))-(.5^(1/(mcq-1))))^2)); cq10=(.5^(10/(mcq-1))*mcq*((.5^(1/(mcq-1)))*(10-1)+(.5^(12/(mcq-1)))*(12-10))*11*ln(.5))/ (((mcq-1)^2)*(((.5^(11/(mcq-1)))-(.5^(1/(mcq-1))))^2)); cq11=(.5^(11/(mcq-1))*mcq*((.5^(1/(mcq-1)))*(11-1)+(.5^(12/(mcq-1)))*(12-11))*11*ln(.5))/ (((mcq-1)^2)*(((.5^(11/(mcq-1)))-(.5^(1/(mcq-1))))^2));
11 cq12=(.5^(12/(mcq-1))*mcq*((.5^(1/(mcq-1)))*(12-1)+(.5^(12/(mcq-1)))*(12-12))*11*ln(.5))/ (((mcq-1)^2)*(((.5^(11/(mcq-1)))-(.5^(1/(mcq-1))))^2)); diff=mcv-mcq; difference in the ARC parameters for verbal and quantitative OUTPUT: TECH1 TECH4; 6. Mathematica code for obtaining partial derivatives of an exponential function 7. Mplus syntax for a quadratic function fit to Kanfer & Ackerman's data Traditional structured latent curve: TITLE: Cudeck and du Toit (2002) quadratic curve; DATA: FILE IS kanfer.dat; data file in ascii format TYPE IS COVARIANCE MEANS; input data are covariances and means NOBSERVATIONS IS 140; sample size is 140 VARIABLE: NAMES ARE y1-y9; all 9 variables y1-y9: ANALYSIS: ESTIMATOR=ML; use maximum likelihood estimation ITERATIONS=5000; use 5000 iterations MODEL: y1-y9*9.9(v); [y1-y9@0]; residual covariances constrained equal, intercepts = 0 [a0*16 ay*39](ma0 may); [ax@0]; intercept and maximum means estimated, maximizer = 0 a0*123; ax*18; ay*53; random coefficient variances a0 WITH ax*-.8 ay*35; ax WITH ay*8.9; random coefficient covariances a0 y1-y9*(a01-a09); ax y1-y9*(ax1-ax9); ay y1-y9*(ay1-ay9); loadings MODEL CONSTRAINT: NEW(max*8.8); define maximizer mean loadings on maximum, intercept, and maximizer factors, constrained to d(y)/d(may), d(y)/d(ma0), and d(y)/d(max) ay1=1-((0/max)-1)^2; a01=((0/max)-1)^2; ax1=2*(may-ma0)*(0-max)*0/max^3; ay2=1-((1/max)-1)^2; a02=((1/max)-1)^2; ax2=2*(may-ma0)*(1-max)*1/max^3; ay3=1-((2/max)-1)^2; a03=((2/max)-1)^2; ax3=2*(may-ma0)*(2-max)*2/max^3; ay4=1-((3/max)-1)^2; a04=((3/max)-1)^2; ax4=2*(may-ma0)*(3-max)*3/max^3; ay5=1-((4/max)-1)^2; a05=((4/max)-1)^2; ax5=2*(may-ma0)*(4-max)*4/max^3; ay6=1-((5/max)-1)^2; a06=((5/max)-1)^2; ax6=2*(may-ma0)*(5-max)*5/max^3; ay7=1-((6/max)-1)^2; a07=((6/max)-1)^2; ax7=2*(may-ma0)*(6-max)*6/max^3; ay8=1-((7/max)-1)^2; a08=((7/max)-1)^2; ax8=2*(may-ma0)*(7-max)*7/max^3;
12 ay9=1-((8/max)-1)^2; a09=((8/max)-1)^2; ax9=2*(may-ma0)*(8-max)*8/max^3; OUTPUT: TECH1 TECH4; Modified structured latent curve: TITLE: Cudeck and du Toit (2002) quadratic curve; DATA: FILE IS kanfer.dat; data file in ascii format TYPE IS COVARIANCE MEANS; input data are covariances and means NOBSERVATIONS IS 140; sample size is 140 VARIABLE: NAMES ARE y1-y9; all 9 variables ANALYSIS: ESTIMATOR=ML; use maximum likelihood estimation ITERATIONS=5000; use 5000 iterations MODEL: [y1-y9*](my1-my9); intercepts constrained equal to target function y1-y9*9.9(v); residual covariances constrained equal [a0@0 ax@0 ay@0]; factor means constrained to zero a0*123; ax*18; ay*53; random coefficient variances a0 WITH ax*-.8 ay*35; ax WITH ay*8.9; random coefficient covariances a0 y1-y9*(a01-a09); ax y1-y9*(ax1-ax9); ay y1-y9*(ay1-ay9); loadings MODEL CONSTRAINT: NEW(ma0*16 max*8.8 may*39); define mean intercept, maximizer, and maximum intercepts constrained equal to target function my1=may-(may-ma0)*((0/max)-1)^2; my2=may-(may-ma0)*((1/max)-1)^2; my3=may-(may-ma0)*((2/max)-1)^2; my4=may-(may-ma0)*((3/max)-1)^2; my5=may-(may-ma0)*((4/max)-1)^2; my6=may-(may-ma0)*((5/max)-1)^2; my7=may-(may-ma0)*((6/max)-1)^2; my8=may-(may-ma0)*((7/max)-1)^2; my9=may-(may-ma0)*((8/max)-1)^2; loadings on maximum, intercept, and maximizer factors, constrained to d(y)/d(may), d(y)/d(ma0), and d(y)/d(max) ay1=1-((0/max)-1)^2; a01=((0/max)-1)^2; ax1=2*(may-ma0)*(0-max)*0/max^3; ay2=1-((1/max)-1)^2; a02=((1/max)-1)^2; ax2=2*(may-ma0)*(1-max)*1/max^3; ay3=1-((2/max)-1)^2; a03=((2/max)-1)^2; ax3=2*(may-ma0)*(2-max)*2/max^3; ay4=1-((3/max)-1)^2; a04=((3/max)-1)^2; ax4=2*(may-ma0)*(3-max)*3/max^3; ay5=1-((4/max)-1)^2; a05=((4/max)-1)^2; ax5=2*(may-ma0)*(4-max)*4/max^3; ay6=1-((5/max)-1)^2; a06=((5/max)-1)^2; ax6=2*(may-ma0)*(5-max)*5/max^3; ay7=1-((6/max)-1)^2; a07=((6/max)-1)^2; ax7=2*(may-ma0)*(6-max)*6/max^3; ay8=1-((7/max)-1)^2; a08=((7/max)-1)^2; ax8=2*(may-ma0)*(7-max)*7/max^3; ay9=1-((8/max)-1)^2; a09=((8/max)-1)^2; ax9=2*(may-ma0)*(8-max)*8/max^3; OUTPUT: TECH1 TECH4; Kanfer & Ackerman's (1989) data (N = 140), a mean vector and covariance matrix:
13 8. Derivation of Equation 11: Reparameterization of a bilinear spline target function A common expression for the bilinear spline model is: θ1 + θ2t t θκ y = θ3 + θ4t t > θκ where θ and 1 θ are the intercept and slope for the first segment, 2 θ and 3 θ are the intercept and 4 slope for the second segment, and θ is the knot. This expression is not convenient for κ linearization, so it needs to be changed. To perform the first step of the reparameterization, it is first necessary to decide whether the slope of the second segment is larger or smaller than the slope of the first segment. In most applications, this choice will be clear by inspection. In cases for which the slope of the second segment is greater than the slope of the first segment (as in our example), a bilinear spline model can be understood as the maximum of the two functions at any point t. Visually, at the knot point the linear equation for the second segment "takes over" where the equation for the first segment leaves off, so it is clear why the maximum would be chosen in this instance (if the slope of the second segment is expected to be less than that of the first, we would use the minimum rather than the minimum): Harring, Cudeck, and du Toit (2006, Appx. C) and Kohli and Harring (2013, Appx. A) provide a convenient expression for the minimum of two functions: min ( uv, ) = ½ u + v ( u v) 2, from which we may derive a convenient expression for the maximum:
14 max ( uv, ) = ½ u + v + ( u v) 2. In words, the maximum of two functions u and v is the sum of the two functions plus the absolute value of their difference, all divided by 2. The first step of the reparameterization consists of substituting the earlier expressions for each segment into this max(.) function: min ( uv, ) = ½ u + v + ( u v) 2 = ½ θ1 + θ2t + θ3 + θ4t + θ1 + θ2t θ3 + θ4t ( ( )) = ½ ( θ θ ) ( θ θ ) t ( θ θ ) t ( θ θ ) Typically, the two functions join at the knot, in which case there are effectively four parameters, not five. The second step in this reparameterization involves eliminating one of the parameters by replacing it with an equivalent expression involving some of the other parameters. Returning to the original expressions for each segment, we observe that where t = θ, κ θ + θ θ = θ + θ θ 1 2 κ 3 4 κ ( ) θ θ = θ θ θ κ Therefore, we can substitute ( θ θ ) θ for ( θ θ ) max, ½ 4 2 κ 1 3 ( uv) = ( θ + θ ) + ( θ + θ ) t + ( θ θ ) t + ( θ θ ) = ½ ( θ1 + θ3 ) + ( θ2 + θ4 ) t + ( θ2 θ4 ) t + ( θ4 θ2 ) θ κ = ½ ( θ θ ) ( θ θ ) t ( θ θ ) θ ( θ θ ) κ 4 2 = ½ κ ( θ θ ) ( θ θ ) t ( θ θ )( θ t ) = ½ κ ( θ θ ) ( θ θ ) t ( θ θ ) ( θ t ) = ½ ( θ + θ ) + ( θ + θ ) t + ( θ θ ) ( θ ) In the final step, bringing ( θ4 θ2 ) θ > θ, implying that ( θ θ ) assumption that in the max(.) function, yielding: 2 2 t κ 2 2 t 2 2 outside the radical is permissible because we started with the is a positive square root. Finally, given that we are 4 2
15 emphasizing the importance of θ and do not care much about the other parts of the equation, we κ can make the following substitutions with no loss: θ1 + θ3 θ2 + θ4 θ4 θ2 ω1 = ω2 = ω3 = so that ( ) 2 y = ω + ωt + ω t θκ. 9. Annotated output: inverse quadratic function Only the most relevant output has been included below. Mplus VERSION 7.2 MUTHEN & MUTHEN 10/02/2014 3:18 PM INPUT INSTRUCTIONS TITLE: davidian theophylline data, inverse quadratic; DATA: FILE IS theophylline_wide.dat; data file in ascii format VARIABLE: NAMES ARE subject dose wt t1-t11 conc1-conc11; all 25 variables subject: subject ID dose: oral dose of theophylline in mg/kg wt: subject weight in kg t1-t11: time since administration of dose in hours conc1-conc11: theophylline concentration in mg/l USEVARIABLES ARE conc1-conc11; we plan to model 11 variables CONSTRAINT ARE t1-t11; individual-specific time scores for use in loadings ANALYSIS: ESTIMATOR=ML; ITERATIONS=2000; CONVERGENCE=.001; estimation options BOOTSTRAP=5000; for bootstrap confidence intervals if desired MODEL: [conc1-conc11*](z1-z11); mean trend (z's defined below) conc1-conc11*.7(ve); residual variances constrained equal conc1-conc10 PWITH conc2-conc11*.2(ve2); optional lag-1 residual covariances b*.002(vb); c@0; m*.5(vm); random coeff. variances (only for maximizer and theta1) b WITH c@0; b WITH m*; c WITH m@0; random coefficient covariances [b@0 c@0 m@0]; factor means constrained to zero b conc1*(b1); b conc2-conc11*(b2-b11); loadings for the theta1 parameter (b) c conc1*(c1); c conc2-conc11*(c2-c11); loadings for the theta2 parameter (c) m conc1*(m1); m conc2-conc11*(m2-m11); loadings for the maximizer parameter (m) MODEL CONSTRAINT: NEW(mb*.07 mc*.02 mm*1.77); starts for theta1, theta2, and maximizer parameters intercepts constrained equal to target function z1=t1/(mb*t1+mc*(mm^2+t1^2)); z2=t2/(mb*t2+mc*(mm^2+t2^2)); z3=t3/(mb*t3+mc*(mm^2+t3^2)); z4=t4/(mb*t4+mc*(mm^2+t4^2)); z5=t5/(mb*t5+mc*(mm^2+t5^2)); z6=t6/(mb*t6+mc*(mm^2+t6^2)); z7=t7/(mb*t7+mc*(mm^2+t7^2));
16 z8=t8/(mb*t8+mc*(mm^2+t8^2)); z9=t9/(mb*t9+mc*(mm^2+t9^2)); z10=t10/(mb*t10+mc*(mm^2+t10^2)); z11=t11/(mb*t11+mc*(mm^2+t11^2)); loadings for "theta1" factor, constrained equal to d(y)/d(mb) b1=-1*t1^2/(mb*t1+mc*(mm^2+t1^2))^2; b2=-1*t2^2/(mb*t2+mc*(mm^2+t2^2))^2; b3=-1*t3^2/(mb*t3+mc*(mm^2+t3^2))^2; b4=-1*t4^2/(mb*t4+mc*(mm^2+t4^2))^2; b5=-1*t5^2/(mb*t5+mc*(mm^2+t5^2))^2; b6=-1*t6^2/(mb*t6+mc*(mm^2+t6^2))^2; b7=-1*t7^2/(mb*t7+mc*(mm^2+t7^2))^2; b8=-1*t8^2/(mb*t8+mc*(mm^2+t8^2))^2; b9=-1*t9^2/(mb*t9+mc*(mm^2+t9^2))^2; b10=-1*t10^2/(mb*t10+mc*(mm^2+t10^2))^2; b11=-1*t11^2/(mb*t11+mc*(mm^2+t11^2))^2; loadings for "theta2" factor, constrained equal to d(y)/d(mc) c1=-1*t1*(mm^2+t1^2)/(mb*t1+mc*(mm^2+t1^2))^2; not strictly necessary c2=-1*t2*(mm^2+t2^2)/(mb*t2+mc*(mm^2+t2^2))^2; here because this c3=-1*t3*(mm^2+t3^2)/(mb*t3+mc*(mm^2+t3^2))^2; coefficient is fixed, c4=-1*t4*(mm^2+t4^2)/(mb*t4+mc*(mm^2+t4^2))^2; but included for c5=-1*t5*(mm^2+t5^2)/(mb*t5+mc*(mm^2+t5^2))^2; completeness. c6=-1*t6*(mm^2+t6^2)/(mb*t6+mc*(mm^2+t6^2))^2; c7=-1*t7*(mm^2+t7^2)/(mb*t7+mc*(mm^2+t7^2))^2; c8=-1*t8*(mm^2+t8^2)/(mb*t8+mc*(mm^2+t8^2))^2; c9=-1*t9*(mm^2+t9^2)/(mb*t9+mc*(mm^2+t9^2))^2; c10=-1*t10*(mm^2+t10^2)/(mb*t10+mc*(mm^2+t10^2))^2; c11=-1*t11*(mm^2+t11^2)/(mb*t11+mc*(mm^2+t11^2))^2; loadings for "maximizer" factor, constrained equal to d(y)/d(mm) m1=-2*mm*mc*t1/(mb*t1+mc*(mm^2+t1^2))^2; m2=-2*mm*mc*t2/(mb*t2+mc*(mm^2+t2^2))^2; m3=-2*mm*mc*t3/(mb*t3+mc*(mm^2+t3^2))^2; m4=-2*mm*mc*t4/(mb*t4+mc*(mm^2+t4^2))^2; m5=-2*mm*mc*t5/(mb*t5+mc*(mm^2+t5^2))^2; m6=-2*mm*mc*t6/(mb*t6+mc*(mm^2+t6^2))^2; m7=-2*mm*mc*t7/(mb*t7+mc*(mm^2+t7^2))^2; m8=-2*mm*mc*t8/(mb*t8+mc*(mm^2+t8^2))^2; m9=-2*mm*mc*t9/(mb*t9+mc*(mm^2+t9^2))^2; m10=-2*mm*mc*t10/(mb*t10+mc*(mm^2+t10^2))^2; m11=-2*mm*mc*t11/(mb*t11+mc*(mm^2+t11^2))^2; OUTPUT: TECH1; CINTERVAL(BCBOOTSTRAP); INPUT READING TERMINATED NORMALLY davidian theophylline data, inverse quadratic; SUMMARY OF ANALYSIS Number of groups 1 Number of observations 12 Number of dependent variables 11 Number of independent variables 0 Number of continuous latent variables 3 Observed dependent variables
17 Continuous CONC1 CONC2 CONC3 CONC4 CONC5 CONC6 CONC7 CONC8 CONC9 CONC10 CONC11 Continuous latent variables B C M Estimator ML Information matrix OBSERVED Maximum number of iterations 2000 Convergence criterion 0.100D-02 Maximum number of steepest descent iterations 20 Input data file(s) theophylline_wide.dat Input data format FREE THE MODEL ESTIMATION TERMINATED NORMALLY MODEL FIT INFORMATION Number of Free Parameters 8 Loglikelihood Information Criteria MODEL RESULTS H0 Value Akaike (AIC) Bayesian (BIC) Sample-Size Adjusted BIC (n* = (n + 2) / 24) Two-Tailed Estimate S.E. Est./S.E. P-Value B C CONC CONC CONC CONC CONC CONC CONC CONC CONC CONC CONC CONC CONC CONC CONC CONC CONC CONC CONC CONC
18 CONC CONC M CONC CONC CONC CONC CONC CONC CONC CONC CONC CONC CONC B WITH C M COVARIANCE OF MAXIMIZER AND THETA1 RANDOM C WITH COEFFICIENTS M CONC1 WITH CONC LAG-1 ERROR COVARIANCE CONC2 WITH CONC CONC3 WITH CONC CONC4 WITH CONC CONC5 WITH CONC CONC6 WITH CONC CONC7 WITH CONC CONC8 WITH CONC CONC9 WITH CONC CONC10 WITH CONC Means B FACTOR MEANS FIXED = 0 C M Intercepts CONC CONC CONC CONC
19 CONC CONC CONC CONC CONC CONC CONC Variances B VARIANCES OF THETA1, C THETA2, AND MAXIMIZER M COEFFICIENTS Residual Variances CONC RESIDUAL VARIANCE, HELD CONC EQUAL OVER TIME CONC CONC CONC CONC CONC CONC CONC CONC CONC New/Additional Parameters MB MEAN THETA1 ESTIMATE MC MEAN THETA2 ESTIMATE MM MEAN MAXIMIZER ESTIMATE QUALITY OF NUMERICAL RESULTS Condition Number for the Information Matrix (ratio of smallest to largest eigenvalue) 0.496E Annotated output: unconditional bilinear spline model Mplus VERSION 7.2 MUTHEN & MUTHEN 06/10/2014 5:36 PM INPUT INSTRUCTIONS TITLE: zerbe glucose; DATA: FILE IS glucose.dat; data file in ascii format VARIABLE: NAMES ARE obese ph1-ph8; all 9 variables USEVARIABLES ARE ph1-ph8; we plan to model 8 variables obese: obesity status, 0=control, 1=obese ph1-ph8: plasma phosphate concentration repeated measures ANALYSIS: BOOTSTRAP IS 1000; for bootstrap confidence intervals if desired MODEL: [ph1-ph8*](z1-z8); mean trend (z's defined below) ph1-ph8*(ve); residual variances constrained equal w1-w4*; random coefficient variances w1 WITH w2*; w1 WITH w3*; w1 WITH w4*; random coefficient covariances w2 WITH w3*; w2 WITH w4*; w3 WITH w4*; random coefficient covariances [w1@0 w2@0 w3@0 w4@0]; factor means constrained to zero w1 ph1-ph8@1; loadings for 1st factor, constrained to 1 w2 ph1*(w21); w2 ph2-ph8(w22-w28); loadings for 2nd factor, defined below w3 ph1*(w31); w3 ph2-ph8(w32-w38); loadings for 3rd factor, defined below
20 w4 ph1*(w41); w4 ph2-ph8(w42-w48); loadings for knot factor, defined below MODEL CONSTRAINT: starts for parameters of the target function NEW(mw1*3.4 mw2*-.26 mw3*.54 mw4*1.6 b1-b4); intercepts constrained equal to target function z1=mw1+mw2*0.0+mw3*sqrt((0.0-mw4)^2); z2=mw1+mw2*0.5+mw3*sqrt((0.5-mw4)^2); z3=mw1+mw2*1.0+mw3*sqrt((1.0-mw4)^2); z4=mw1+mw2*1.5+mw3*sqrt((1.5-mw4)^2); z5=mw1+mw2*2.0+mw3*sqrt((2.0-mw4)^2); z6=mw1+mw2*3.0+mw3*sqrt((3.0-mw4)^2); z7=mw1+mw2*4.0+mw3*sqrt((4.0-mw4)^2); z8=mw1+mw2*5.0+mw3*sqrt((5.0-mw4)^2); loadings for 2nd and 3rd factors, constrained to d(y)/d(mw2) and d(y)/d(mw3) w21=0.0; w31=sqrt((0.0-mw4)^2); w22=0.5; w32=sqrt((0.5-mw4)^2); w23=1.0; w33=sqrt((1.0-mw4)^2); w24=1.5; w34=sqrt((1.5-mw4)^2); w25=2.0; w35=sqrt((2.0-mw4)^2); w26=3.0; w36=sqrt((3.0-mw4)^2); w27=4.0; w37=sqrt((4.0-mw4)^2); w28=5.0; w38=sqrt((5.0-mw4)^2); loadings for knot factor, constrained to d(y)/d(mw4) w41=(mw3*(mw4-0.0))/sqrt((0.0-mw4)^2); w42=(mw3*(mw4-0.5))/sqrt((0.5-mw4)^2); w43=(mw3*(mw4-1.0))/sqrt((1.0-mw4)^2); w44=(mw3*(mw4-1.5))/sqrt((1.5-mw4)^2); w45=(mw3*(mw4-2.0))/sqrt((2.0-mw4)^2); w46=(mw3*(mw4-3.0))/sqrt((3.0-mw4)^2); w47=(mw3*(mw4-4.0))/sqrt((4.0-mw4)^2); w48=(mw3*(mw4-5.0))/sqrt((5.0-mw4)^2); parameters of original parameterization b1=mw1+mw3*mw4; b2=mw2-mw3; b3=mw1-mw3*mw4; b4=mw2+mw3; OUTPUT: TECH1; CINTERVAL(BCBOOTSTRAP); INPUT READING TERMINATED NORMALLY zerbe glucose; SUMMARY OF ANALYSIS Number of groups 1 Number of observations 33 Number of dependent variables 8 Number of independent variables 0 Number of continuous latent variables 4 Observed dependent variables Continuous PH1 PH2 PH3 PH4 PH5 PH6 PH7 PH8
21 Continuous latent variables W1 W2 W3 W4 Estimator ML Information matrix OBSERVED Maximum number of iterations 1000 Convergence criterion 0.500D-04 Maximum number of steepest descent iterations 20 Input data file(s) glucose.dat Input data format FREE THE MODEL ESTIMATION TERMINATED NORMALLY MODEL FIT INFORMATION Number of Free Parameters 15 Loglikelihood H0 Value H1 Value Information Criteria Akaike (AIC) Bayesian (BIC) Sample-Size Adjusted BIC (n* = (n + 2) / 24) Chi-Square Test of Model Fit Value Degrees of Freedom 29 P-Value RMSEA (Root Mean Square Error Of Approximation) CFI/TLI Estimate Percent C.I Probability RMSEA <= CFI TLI Chi-Square Test of Model Fit for the Baseline Model Value Degrees of Freedom 28 P-Value SRMR (Standardized Root Mean Square Residual) MODEL RESULTS Value Two-Tailed
22 W1 W2 W3 W4 Estimate S.E. Est./S.E. P-Value PH PH PH PH PH PH PH PH PH PH PH PH PH PH PH PH PH PH PH PH PH PH PH PH PH PH PH PH PH PH PH PH W1 WITH COVARIANCES OF RANDOM W COEFFICIENTS W W W2 WITH W W W3 WITH W Means W FACTOR MEANS FIXED = 0 W W W Intercepts PH PH
23 PH PH PH PH PH PH Variances W VARIANCES OF RANDOM W COEFFICIENTS W W Residual Variances PH RESIDUAL VARIANCE, HELD EQUAL PH OVER TIME PH PH PH PH PH PH New/Additional Parameters MW MEAN OF 1ST GROWTH PARAMETER MW MEAN OF 2ND GROWTH PARAMETER MW MEAN OF 3RD GROWTH PARAMETER MW MEAN OF KNOT PARAMETER B INTERCEPT FOR FIRST SEGMENT B SLOPE FOR FIRST SEGMENT B INTERCEPT FOR SECOND SEGMENT B SLOPE FOR SECOND SEGMENT QUALITY OF NUMERICAL RESULTS Condition Number for the Information Matrix (ratio of smallest to largest eigenvalue) 0.260E Annotated output: conditional bilinear spline model Mplus VERSION 7.2 MUTHEN & MUTHEN 06/10/2014 5:37 PM INPUT INSTRUCTIONS TITLE: zerbe glucose; DATA: FILE IS glucose.dat; data file in ascii format DEFINE: xobese=abs(obese-1); 1=control; 0=obese VARIABLE: NAMES ARE obese ph1-ph8; all 9 variables obese: obesity status, 0=control, 1=obese ph1-ph8: plasma phosphate concentration repeated measures USEVARIABLES ARE ph1-ph8 xobese; we plan to model 9 variables ANALYSIS: BOOTSTRAP IS 1000; for bootstrap confidence intervals if desired MODEL: [ph1-ph8*](z1-z8); mean trend (z's defined below) ph1-ph8*(ve); residual variances constrained equal w1-w4*; random coefficient residual variances w1 WITH w2*; w1 WITH w3*; w1 WITH w4*; random coefficient residual covariances w2 WITH w3*; w2 WITH w4*; w3 WITH w4*; random coefficient residual covariances [w1@0 w2@0 w3@0 w4@0]; factor intercepts constrained to zero
24 w1 loadings for 1st factor, constrained to 1 w2 ph1*(w21); w2 ph2-ph8(w22-w28); loadings for 2nd factor, defined below w3 ph1*(w31); w3 ph2-ph8(w32-w38); loadings for 3rd factor, defined below w4 ph1*(w41); w4 ph2-ph8(w42-w48); loadings for knot factor, defined below w1-w4 ON xobese; regress random coefficients on obese MODEL CONSTRAINT: starts for parameters of the target function NEW(mw1*3.5 mw2*-.21 mw3*.49 mw4*1.95 b1-b4); intercepts constrained equal to target function z1=mw1+mw2*0.0+mw3*sqrt((0.0-mw4)^2); z2=mw1+mw2*0.5+mw3*sqrt((0.5-mw4)^2); z3=mw1+mw2*1.0+mw3*sqrt((1.0-mw4)^2); z4=mw1+mw2*1.5+mw3*sqrt((1.5-mw4)^2); z5=mw1+mw2*2.0+mw3*sqrt((2.0-mw4)^2); z6=mw1+mw2*3.0+mw3*sqrt((3.0-mw4)^2); z7=mw1+mw2*4.0+mw3*sqrt((4.0-mw4)^2); z8=mw1+mw2*5.0+mw3*sqrt((5.0-mw4)^2); loadings for 2nd and 3rd factors, constrained to d(y)/d(mw2) and d(y)/d(mw3) w21=0.0; w31=sqrt((0.0-mw4)^2); w22=0.5; w32=sqrt((0.5-mw4)^2); w23=1.0; w33=sqrt((1.0-mw4)^2); w24=1.5; w34=sqrt((1.5-mw4)^2); w25=2.0; w35=sqrt((2.0-mw4)^2); w26=3.0; w36=sqrt((3.0-mw4)^2); w27=4.0; w37=sqrt((4.0-mw4)^2); w28=5.0; w38=sqrt((5.0-mw4)^2); loadings for knot factor, constrained to d(y)/d(mw4) w41=(mw3*(mw4-0.0))/sqrt((0.0-mw4)^2); w42=(mw3*(mw4-0.5))/sqrt((0.5-mw4)^2); w43=(mw3*(mw4-1.0))/sqrt((1.0-mw4)^2); w44=(mw3*(mw4-1.5))/sqrt((1.5-mw4)^2); w45=(mw3*(mw4-2.0))/sqrt((2.0-mw4)^2); w46=(mw3*(mw4-3.0))/sqrt((3.0-mw4)^2); w47=(mw3*(mw4-4.0))/sqrt((4.0-mw4)^2); w48=(mw3*(mw4-5.0))/sqrt((5.0-mw4)^2); parameters of original parameterization b1=mw1+mw3*mw4; b2=mw2-mw3; b3=mw1-mw3*mw4; b4=mw2+mw3; OUTPUT: TECH1; CINTERVAL(BCBOOTSTRAP); INPUT READING TERMINATED NORMALLY zerbe glucose; SUMMARY OF ANALYSIS Number of groups 1 Number of observations 33 Number of dependent variables 8 Number of independent variables 1 Number of continuous latent variables 4 Observed dependent variables
Longitudinal Invariance CFA (using MLR) Example in Mplus v. 7.4 (N = 151; 6 items over 3 occasions)
 Longitudinal Invariance CFA (using MLR) Example in Mplus v. 7.4 (N = 151; 6 items over 3 occasions) CLP 948 Example 7b page 1 These data measuring a latent trait of social functioning were collected at
Longitudinal Invariance CFA (using MLR) Example in Mplus v. 7.4 (N = 151; 6 items over 3 occasions) CLP 948 Example 7b page 1 These data measuring a latent trait of social functioning were collected at
Multiple Group CFA Invariance Example (data from Brown Chapter 7) using MLR Mplus 7.4: Major Depression Criteria across Men and Women (n = 345 each)
 Multiple Group CFA Invariance Example (data from Brown Chapter 7) using MLR Mplus 7.4: Major Depression Criteria across Men and Women (n = 345 each) 9 items rated by clinicians on a scale of 0 to 8 (0
Multiple Group CFA Invariance Example (data from Brown Chapter 7) using MLR Mplus 7.4: Major Depression Criteria across Men and Women (n = 345 each) 9 items rated by clinicians on a scale of 0 to 8 (0
ADVANCED C. MEASUREMENT INVARIANCE SEM REX B KLINE CONCORDIA
 ADVANCED SEM C. MEASUREMENT INVARIANCE REX B KLINE CONCORDIA C C2 multiple model 2 data sets simultaneous C3 multiple 2 populations 2 occasions 2 methods C4 multiple unstandardized constrain to equal fit
ADVANCED SEM C. MEASUREMENT INVARIANCE REX B KLINE CONCORDIA C C2 multiple model 2 data sets simultaneous C3 multiple 2 populations 2 occasions 2 methods C4 multiple unstandardized constrain to equal fit
Specifying Latent Curve and Other Growth Models Using Mplus. (Revised )
 Ronald H. Heck 1 University of Hawai i at Mānoa Handout #20 Specifying Latent Curve and Other Growth Models Using Mplus (Revised 12-1-2014) The SEM approach offers a contrasting framework for use in analyzing
Ronald H. Heck 1 University of Hawai i at Mānoa Handout #20 Specifying Latent Curve and Other Growth Models Using Mplus (Revised 12-1-2014) The SEM approach offers a contrasting framework for use in analyzing
An Introduction to Path Analysis
 An Introduction to Path Analysis PRE 905: Multivariate Analysis Lecture 10: April 15, 2014 PRE 905: Lecture 10 Path Analysis Today s Lecture Path analysis starting with multivariate regression then arriving
An Introduction to Path Analysis PRE 905: Multivariate Analysis Lecture 10: April 15, 2014 PRE 905: Lecture 10 Path Analysis Today s Lecture Path analysis starting with multivariate regression then arriving
An Introduction to Mplus and Path Analysis
 An Introduction to Mplus and Path Analysis PSYC 943: Fundamentals of Multivariate Modeling Lecture 10: October 30, 2013 PSYC 943: Lecture 10 Today s Lecture Path analysis starting with multivariate regression
An Introduction to Mplus and Path Analysis PSYC 943: Fundamentals of Multivariate Modeling Lecture 10: October 30, 2013 PSYC 943: Lecture 10 Today s Lecture Path analysis starting with multivariate regression
Advanced Structural Equations Models I
 This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike License. Your use of this material constitutes acceptance of that license and the conditions of use of materials on this
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike License. Your use of this material constitutes acceptance of that license and the conditions of use of materials on this
Structural Equation Modeling and Confirmatory Factor Analysis. Types of Variables
 /4/04 Structural Equation Modeling and Confirmatory Factor Analysis Advanced Statistics for Researchers Session 3 Dr. Chris Rakes Website: http://csrakes.yolasite.com Email: Rakes@umbc.edu Twitter: @RakesChris
/4/04 Structural Equation Modeling and Confirmatory Factor Analysis Advanced Statistics for Researchers Session 3 Dr. Chris Rakes Website: http://csrakes.yolasite.com Email: Rakes@umbc.edu Twitter: @RakesChris
Subject-specific observed profiles of log(fev1) vs age First 50 subjects in Six Cities Study
 Subject-specific observed profiles of log(fev1) vs age First 50 subjects in Six Cities Study 1.4 0.0-6 7 8 9 10 11 12 13 14 15 16 17 18 19 age Model 1: A simple broken stick model with knot at 14 fit with
Subject-specific observed profiles of log(fev1) vs age First 50 subjects in Six Cities Study 1.4 0.0-6 7 8 9 10 11 12 13 14 15 16 17 18 19 age Model 1: A simple broken stick model with knot at 14 fit with
Categorical and Zero Inflated Growth Models
 Categorical and Zero Inflated Growth Models Alan C. Acock* Summer, 2009 *Alan C. Acock, Department of Human Development and Family Sciences, Oregon State University, Corvallis OR 97331 (alan.acock@oregonstate.edu).
Categorical and Zero Inflated Growth Models Alan C. Acock* Summer, 2009 *Alan C. Acock, Department of Human Development and Family Sciences, Oregon State University, Corvallis OR 97331 (alan.acock@oregonstate.edu).
Online Appendix for Sterba, S.K. (2013). Understanding linkages among mixture models. Multivariate Behavioral Research, 48,
 Online Appendix for, S.K. (2013). Understanding linkages among mixture models. Multivariate Behavioral Research, 48, 775-815. Table of Contents. I. Full presentation of parallel-process groups-based trajectory
Online Appendix for, S.K. (2013). Understanding linkages among mixture models. Multivariate Behavioral Research, 48, 775-815. Table of Contents. I. Full presentation of parallel-process groups-based trajectory
NELS 88. Latent Response Variable Formulation Versus Probability Curve Formulation
 NELS 88 Table 2.3 Adjusted odds ratios of eighth-grade students in 988 performing below basic levels of reading and mathematics in 988 and dropping out of school, 988 to 990, by basic demographics Variable
NELS 88 Table 2.3 Adjusted odds ratios of eighth-grade students in 988 performing below basic levels of reading and mathematics in 988 and dropping out of school, 988 to 990, by basic demographics Variable
The Role of Leader Motivating Language in Employee Absenteeism (Mayfield: 2009)
 DATE: 12/15/2009 TIME: 5:50 Page 1 LISREL 8.80 (STUDENT EDITION) BY Karl G. J reskog & Dag S rbom This program is published exclusively by Scientific Software International, Inc. 7383 N. Lincoln Avenue,
DATE: 12/15/2009 TIME: 5:50 Page 1 LISREL 8.80 (STUDENT EDITION) BY Karl G. J reskog & Dag S rbom This program is published exclusively by Scientific Software International, Inc. 7383 N. Lincoln Avenue,
Estimation of Curvilinear Effects in SEM. Rex B. Kline, September 2009
 Estimation of Curvilinear Effects in SEM Supplement to Principles and Practice of Structural Equation Modeling (3rd ed.) Rex B. Kline, September 009 Curvlinear Effects of Observed Variables Consider the
Estimation of Curvilinear Effects in SEM Supplement to Principles and Practice of Structural Equation Modeling (3rd ed.) Rex B. Kline, September 009 Curvlinear Effects of Observed Variables Consider the
The use of structural equation modeling to examine consumption of pornographic materials in Chinese adolescents in Hong Kong
 The use of structural equation modeling to examine consumption of pornographic materials in Chinese adolescents in Hong Kong Appendix 1 Creating matrices and checking for normality!prelis SYNTAX: Can be
The use of structural equation modeling to examine consumption of pornographic materials in Chinese adolescents in Hong Kong Appendix 1 Creating matrices and checking for normality!prelis SYNTAX: Can be
What is Latent Class Analysis. Tarani Chandola
 What is Latent Class Analysis Tarani Chandola methods@manchester Many names similar methods (Finite) Mixture Modeling Latent Class Analysis Latent Profile Analysis Latent class analysis (LCA) LCA is a
What is Latent Class Analysis Tarani Chandola methods@manchester Many names similar methods (Finite) Mixture Modeling Latent Class Analysis Latent Profile Analysis Latent class analysis (LCA) LCA is a
Title. Description. Remarks and examples. stata.com. stata.com. Variable notation. methods and formulas for sem Methods and formulas for sem
 Title stata.com methods and formulas for sem Methods and formulas for sem Description Remarks and examples References Also see Description The methods and formulas for the sem commands are presented below.
Title stata.com methods and formulas for sem Methods and formulas for sem Description Remarks and examples References Also see Description The methods and formulas for the sem commands are presented below.
Mplus Short Courses Topic 3. Growth Modeling With Latent Variables Using Mplus: Introductory And Intermediate Growth Models
 Mplus Short Courses Topic 3 Growth Modeling With Latent Variables Using Mplus: Introductory And Intermediate Growth Models Linda K. Muthén Bengt Muthén Copyright 2008 Muthén & Muthén www.statmodel.com
Mplus Short Courses Topic 3 Growth Modeling With Latent Variables Using Mplus: Introductory And Intermediate Growth Models Linda K. Muthén Bengt Muthén Copyright 2008 Muthén & Muthén www.statmodel.com
Using Mplus individual residual plots for. diagnostics and model evaluation in SEM
 Using Mplus individual residual plots for diagnostics and model evaluation in SEM Tihomir Asparouhov and Bengt Muthén Mplus Web Notes: No. 20 October 31, 2017 1 Introduction A variety of plots are available
Using Mplus individual residual plots for diagnostics and model evaluation in SEM Tihomir Asparouhov and Bengt Muthén Mplus Web Notes: No. 20 October 31, 2017 1 Introduction A variety of plots are available
Measurement Invariance Testing with Many Groups: A Comparison of Five Approaches (Online Supplements)
 University of South Florida Scholar Commons Educational and Psychological Studies Faculty Publications Educational and Psychological Studies 2017 Measurement Invariance Testing with Many Groups: A Comparison
University of South Florida Scholar Commons Educational and Psychological Studies Faculty Publications Educational and Psychological Studies 2017 Measurement Invariance Testing with Many Groups: A Comparison
Class Notes: Week 8. Probit versus Logit Link Functions and Count Data
 Ronald Heck Class Notes: Week 8 1 Class Notes: Week 8 Probit versus Logit Link Functions and Count Data This week we ll take up a couple of issues. The first is working with a probit link function. While
Ronald Heck Class Notes: Week 8 1 Class Notes: Week 8 Probit versus Logit Link Functions and Count Data This week we ll take up a couple of issues. The first is working with a probit link function. While
PRESENTATION TITLE. Is my survey biased? The importance of measurement invariance. Yusuke Kuroki Sunny Moon November 9 th, 2017
 PRESENTATION TITLE Is my survey biased? The importance of measurement invariance Yusuke Kuroki Sunny Moon November 9 th, 2017 Measurement Invariance Measurement invariance: the same construct is being
PRESENTATION TITLE Is my survey biased? The importance of measurement invariance Yusuke Kuroki Sunny Moon November 9 th, 2017 Measurement Invariance Measurement invariance: the same construct is being
Plausible Values for Latent Variables Using Mplus
 Plausible Values for Latent Variables Using Mplus Tihomir Asparouhov and Bengt Muthén August 21, 2010 1 1 Introduction Plausible values are imputed values for latent variables. All latent variables can
Plausible Values for Latent Variables Using Mplus Tihomir Asparouhov and Bengt Muthén August 21, 2010 1 1 Introduction Plausible values are imputed values for latent variables. All latent variables can
Univariate ARIMA Models
 Univariate ARIMA Models ARIMA Model Building Steps: Identification: Using graphs, statistics, ACFs and PACFs, transformations, etc. to achieve stationary and tentatively identify patterns and model components.
Univariate ARIMA Models ARIMA Model Building Steps: Identification: Using graphs, statistics, ACFs and PACFs, transformations, etc. to achieve stationary and tentatively identify patterns and model components.
Introduction to Structural Equation Modeling
 Introduction to Structural Equation Modeling Notes Prepared by: Lisa Lix, PhD Manitoba Centre for Health Policy Topics Section I: Introduction Section II: Review of Statistical Concepts and Regression
Introduction to Structural Equation Modeling Notes Prepared by: Lisa Lix, PhD Manitoba Centre for Health Policy Topics Section I: Introduction Section II: Review of Statistical Concepts and Regression
C:\Users\Rex Kline\AppData\Local\Temp\AmosTemp\AmosScratch.amw. Date: Friday, December 5, 2014 Time: 11:20:30 AM
 Page 1 of 7 C:\Users\Rex Kline\AppData\Local\Temp\AmosTemp\AmosScratch.amw Analysis Summary Date and Time Date: Friday, December 5, 2014 Time: 11:20:30 AM Title Groups Group number 1 (Group number 1) Notes
Page 1 of 7 C:\Users\Rex Kline\AppData\Local\Temp\AmosTemp\AmosScratch.amw Analysis Summary Date and Time Date: Friday, December 5, 2014 Time: 11:20:30 AM Title Groups Group number 1 (Group number 1) Notes
Mplus Short Courses Day 2. Growth Modeling With Latent Variables Using Mplus
 Mplus Short Courses Day 2 Growth Modeling With Latent Variables Using Mplus Linda K. Muthén Bengt Muthén Copyright 2007 Muthén & Muthén www.statmodel.com 1 Table Of Contents General Latent Variable Modeling
Mplus Short Courses Day 2 Growth Modeling With Latent Variables Using Mplus Linda K. Muthén Bengt Muthén Copyright 2007 Muthén & Muthén www.statmodel.com 1 Table Of Contents General Latent Variable Modeling
Maximum Likelihood Estimation; Robust Maximum Likelihood; Missing Data with Maximum Likelihood
 Maximum Likelihood Estimation; Robust Maximum Likelihood; Missing Data with Maximum Likelihood PRE 906: Structural Equation Modeling Lecture #3 February 4, 2015 PRE 906, SEM: Estimation Today s Class An
Maximum Likelihood Estimation; Robust Maximum Likelihood; Missing Data with Maximum Likelihood PRE 906: Structural Equation Modeling Lecture #3 February 4, 2015 PRE 906, SEM: Estimation Today s Class An
CHAPTER 9 EXAMPLES: MULTILEVEL MODELING WITH COMPLEX SURVEY DATA
 Examples: Multilevel Modeling With Complex Survey Data CHAPTER 9 EXAMPLES: MULTILEVEL MODELING WITH COMPLEX SURVEY DATA Complex survey data refers to data obtained by stratification, cluster sampling and/or
Examples: Multilevel Modeling With Complex Survey Data CHAPTER 9 EXAMPLES: MULTILEVEL MODELING WITH COMPLEX SURVEY DATA Complex survey data refers to data obtained by stratification, cluster sampling and/or
2/26/2017. PSY 512: Advanced Statistics for Psychological and Behavioral Research 2
 PSY 512: Advanced Statistics for Psychological and Behavioral Research 2 What is SEM? When should we use SEM? What can SEM tell us? SEM Terminology and Jargon Technical Issues Types of SEM Models Limitations
PSY 512: Advanced Statistics for Psychological and Behavioral Research 2 What is SEM? When should we use SEM? What can SEM tell us? SEM Terminology and Jargon Technical Issues Types of SEM Models Limitations
Example 7b: Generalized Models for Ordinal Longitudinal Data using SAS GLIMMIX, STATA MEOLOGIT, and MPLUS (last proportional odds model only)
 CLDP945 Example 7b page 1 Example 7b: Generalized Models for Ordinal Longitudinal Data using SAS GLIMMIX, STATA MEOLOGIT, and MPLUS (last proportional odds model only) This example comes from real data
CLDP945 Example 7b page 1 Example 7b: Generalized Models for Ordinal Longitudinal Data using SAS GLIMMIX, STATA MEOLOGIT, and MPLUS (last proportional odds model only) This example comes from real data
Walkthrough for Illustrations. Illustration 1
 Tay, L., Meade, A. W., & Cao, M. (in press). An overview and practical guide to IRT measurement equivalence analysis. Organizational Research Methods. doi: 10.1177/1094428114553062 Walkthrough for Illustrations
Tay, L., Meade, A. W., & Cao, M. (in press). An overview and practical guide to IRT measurement equivalence analysis. Organizational Research Methods. doi: 10.1177/1094428114553062 Walkthrough for Illustrations
Step 2: Select Analyze, Mixed Models, and Linear.
 Example 1a. 20 employees were given a mood questionnaire on Monday, Wednesday and again on Friday. The data will be first be analyzed using a Covariance Pattern model. Step 1: Copy Example1.sav data file
Example 1a. 20 employees were given a mood questionnaire on Monday, Wednesday and again on Friday. The data will be first be analyzed using a Covariance Pattern model. Step 1: Copy Example1.sav data file
CHAPTER 3. SPECIALIZED EXTENSIONS
 03-Preacher-45609:03-Preacher-45609.qxd 6/3/2008 3:36 PM Page 57 CHAPTER 3. SPECIALIZED EXTENSIONS We have by no means exhausted the possibilities of LGM with the examples presented thus far. As scientific
03-Preacher-45609:03-Preacher-45609.qxd 6/3/2008 3:36 PM Page 57 CHAPTER 3. SPECIALIZED EXTENSIONS We have by no means exhausted the possibilities of LGM with the examples presented thus far. As scientific
Introduction to Confirmatory Factor Analysis
 Introduction to Confirmatory Factor Analysis Multivariate Methods in Education ERSH 8350 Lecture #12 November 16, 2011 ERSH 8350: Lecture 12 Today s Class An Introduction to: Confirmatory Factor Analysis
Introduction to Confirmatory Factor Analysis Multivariate Methods in Education ERSH 8350 Lecture #12 November 16, 2011 ERSH 8350: Lecture 12 Today s Class An Introduction to: Confirmatory Factor Analysis
Supplemental material for Autoregressive Latent Trajectory 1
 Supplemental material for Autoregressive Latent Trajectory 1 Supplemental Materials for The Longitudinal Interplay of Adolescents Self-Esteem and Body Image: A Conditional Autoregressive Latent Trajectory
Supplemental material for Autoregressive Latent Trajectory 1 Supplemental Materials for The Longitudinal Interplay of Adolescents Self-Esteem and Body Image: A Conditional Autoregressive Latent Trajectory
Multilevel Structural Equation Modeling
 Multilevel Structural Equation Modeling Joop Hox Utrecht University j.hox@uu.nl http://www.joophox.net 14_15_mlevsem Multilevel Regression Three level data structure Groups at different levels may have
Multilevel Structural Equation Modeling Joop Hox Utrecht University j.hox@uu.nl http://www.joophox.net 14_15_mlevsem Multilevel Regression Three level data structure Groups at different levels may have
Latent Variable Analysis
 Latent Variable Analysis Path Analysis Recap I. Path Diagram a. Exogeneous vs. Endogeneous Variables b. Dependent vs, Independent Variables c. Recursive vs. on-recursive Models II. Structural (Regression)
Latent Variable Analysis Path Analysis Recap I. Path Diagram a. Exogeneous vs. Endogeneous Variables b. Dependent vs, Independent Variables c. Recursive vs. on-recursive Models II. Structural (Regression)
Confirmatory Factor Models (CFA: Confirmatory Factor Analysis)
 Confirmatory Factor Models (CFA: Confirmatory Factor Analysis) Today s topics: Comparison of EFA and CFA CFA model parameters and identification CFA model estimation CFA model fit evaluation CLP 948: Lecture
Confirmatory Factor Models (CFA: Confirmatory Factor Analysis) Today s topics: Comparison of EFA and CFA CFA model parameters and identification CFA model estimation CFA model fit evaluation CLP 948: Lecture
SEM Day 3 Lab Exercises SPIDA 2007 Dave Flora
 SEM Day 3 Lab Exercises SPIDA 2007 Dave Flora 1 Today we will see how to estimate SEM conditional latent trajectory models and interpret output using both SAS and LISREL. Exercise 1 Using SAS PROC CALIS,
SEM Day 3 Lab Exercises SPIDA 2007 Dave Flora 1 Today we will see how to estimate SEM conditional latent trajectory models and interpret output using both SAS and LISREL. Exercise 1 Using SAS PROC CALIS,
Estimation and Model Selection in Mixed Effects Models Part I. Adeline Samson 1
 Estimation and Model Selection in Mixed Effects Models Part I Adeline Samson 1 1 University Paris Descartes Summer school 2009 - Lipari, Italy These slides are based on Marc Lavielle s slides Outline 1
Estimation and Model Selection in Mixed Effects Models Part I Adeline Samson 1 1 University Paris Descartes Summer school 2009 - Lipari, Italy These slides are based on Marc Lavielle s slides Outline 1
CONFIRMATORY FACTOR ANALYSIS
 1 CONFIRMATORY FACTOR ANALYSIS The purpose of confirmatory factor analysis (CFA) is to explain the pattern of associations among a set of observed variables in terms of a smaller number of underlying latent
1 CONFIRMATORY FACTOR ANALYSIS The purpose of confirmatory factor analysis (CFA) is to explain the pattern of associations among a set of observed variables in terms of a smaller number of underlying latent
Thursday Morning. Growth Modelling in Mplus. Using a set of repeated continuous measures of bodyweight
 Thursday Morning Growth Modelling in Mplus Using a set of repeated continuous measures of bodyweight 1 Growth modelling Continuous Data Mplus model syntax refresher ALSPAC Confirmatory Factor Analysis
Thursday Morning Growth Modelling in Mplus Using a set of repeated continuous measures of bodyweight 1 Growth modelling Continuous Data Mplus model syntax refresher ALSPAC Confirmatory Factor Analysis
EPSY 905: Fundamentals of Multivariate Modeling Online Lecture #7
 Introduction to Generalized Univariate Models: Models for Binary Outcomes EPSY 905: Fundamentals of Multivariate Modeling Online Lecture #7 EPSY 905: Intro to Generalized In This Lecture A short review
Introduction to Generalized Univariate Models: Models for Binary Outcomes EPSY 905: Fundamentals of Multivariate Modeling Online Lecture #7 EPSY 905: Intro to Generalized In This Lecture A short review
Computationally Efficient Estimation of Multilevel High-Dimensional Latent Variable Models
 Computationally Efficient Estimation of Multilevel High-Dimensional Latent Variable Models Tihomir Asparouhov 1, Bengt Muthen 2 Muthen & Muthen 1 UCLA 2 Abstract Multilevel analysis often leads to modeling
Computationally Efficient Estimation of Multilevel High-Dimensional Latent Variable Models Tihomir Asparouhov 1, Bengt Muthen 2 Muthen & Muthen 1 UCLA 2 Abstract Multilevel analysis often leads to modeling
Fitting PK Models with SAS NLMIXED Procedure Halimu Haridona, PPD Inc., Beijing
 PharmaSUG China 1 st Conference, 2012 Fitting PK Models with SAS NLMIXED Procedure Halimu Haridona, PPD Inc., Beijing ABSTRACT Pharmacokinetic (PK) models are important for new drug development. Statistical
PharmaSUG China 1 st Conference, 2012 Fitting PK Models with SAS NLMIXED Procedure Halimu Haridona, PPD Inc., Beijing ABSTRACT Pharmacokinetic (PK) models are important for new drug development. Statistical
Confidence Intervals for the Odds Ratio in Logistic Regression with One Binary X
 Chapter 864 Confidence Intervals for the Odds Ratio in Logistic Regression with One Binary X Introduction Logistic regression expresses the relationship between a binary response variable and one or more
Chapter 864 Confidence Intervals for the Odds Ratio in Logistic Regression with One Binary X Introduction Logistic regression expresses the relationship between a binary response variable and one or more
Standard Errors & Confidence Intervals. N(0, I( β) 1 ), I( β) = [ 2 l(β, φ; y) β i β β= β j
 Standard Errors & Confidence Intervals β β asy N(0, I( β) 1 ), where I( β) = [ 2 l(β, φ; y) ] β i β β= β j We can obtain asymptotic 100(1 α)% confidence intervals for β j using: β j ± Z 1 α/2 se( β j )
Standard Errors & Confidence Intervals β β asy N(0, I( β) 1 ), where I( β) = [ 2 l(β, φ; y) ] β i β β= β j We can obtain asymptotic 100(1 α)% confidence intervals for β j using: β j ± Z 1 α/2 se( β j )
Trajectories. Kevin Grimm a, Zhiyong Zhang b, Fumiaki Hamagami c & Michèle Mazzocco d a Department of Psychology, University of California,
 This article was downloaded by: [University of California, Los Angeles (UCLA)] On: 01 April 2013, At: 06:52 Publisher: Routledge Informa Ltd Registered in England and Wales Registered Number: 1072954 Registered
This article was downloaded by: [University of California, Los Angeles (UCLA)] On: 01 April 2013, At: 06:52 Publisher: Routledge Informa Ltd Registered in England and Wales Registered Number: 1072954 Registered
Psychology 454: Latent Variable Modeling How do you know if a model works?
 Psychology 454: Latent Variable Modeling How do you know if a model works? William Revelle Department of Psychology Northwestern University Evanston, Illinois USA November, 2012 1 / 18 Outline 1 Goodness
Psychology 454: Latent Variable Modeling How do you know if a model works? William Revelle Department of Psychology Northwestern University Evanston, Illinois USA November, 2012 1 / 18 Outline 1 Goodness
EVALUATION OF STRUCTURAL EQUATION MODELS
 1 EVALUATION OF STRUCTURAL EQUATION MODELS I. Issues related to the initial specification of theoretical models of interest 1. Model specification: a. Measurement model: (i) EFA vs. CFA (ii) reflective
1 EVALUATION OF STRUCTURAL EQUATION MODELS I. Issues related to the initial specification of theoretical models of interest 1. Model specification: a. Measurement model: (i) EFA vs. CFA (ii) reflective
Basic IRT Concepts, Models, and Assumptions
 Basic IRT Concepts, Models, and Assumptions Lecture #2 ICPSR Item Response Theory Workshop Lecture #2: 1of 64 Lecture #2 Overview Background of IRT and how it differs from CFA Creating a scale An introduction
Basic IRT Concepts, Models, and Assumptions Lecture #2 ICPSR Item Response Theory Workshop Lecture #2: 1of 64 Lecture #2 Overview Background of IRT and how it differs from CFA Creating a scale An introduction
3 Results. Part I. 3.1 Base/primary model
 3 Results Part I 3.1 Base/primary model For the development of the base/primary population model the development dataset (for data details see Table 5 and sections 2.1 and 2.2), which included 1256 serum
3 Results Part I 3.1 Base/primary model For the development of the base/primary population model the development dataset (for data details see Table 5 and sections 2.1 and 2.2), which included 1256 serum
Path Analysis. PRE 906: Structural Equation Modeling Lecture #5 February 18, PRE 906, SEM: Lecture 5 - Path Analysis
 Path Analysis PRE 906: Structural Equation Modeling Lecture #5 February 18, 2015 PRE 906, SEM: Lecture 5 - Path Analysis Key Questions for Today s Lecture What distinguishes path models from multivariate
Path Analysis PRE 906: Structural Equation Modeling Lecture #5 February 18, 2015 PRE 906, SEM: Lecture 5 - Path Analysis Key Questions for Today s Lecture What distinguishes path models from multivariate
MIXED MODELS FOR REPEATED (LONGITUDINAL) DATA PART 2 DAVID C. HOWELL 4/1/2010
 MIXED MODELS FOR REPEATED (LONGITUDINAL) DATA PART 2 DAVID C. HOWELL 4/1/2010 Part 1 of this document can be found at http://www.uvm.edu/~dhowell/methods/supplements/mixed Models for Repeated Measures1.pdf
MIXED MODELS FOR REPEATED (LONGITUDINAL) DATA PART 2 DAVID C. HOWELL 4/1/2010 Part 1 of this document can be found at http://www.uvm.edu/~dhowell/methods/supplements/mixed Models for Repeated Measures1.pdf
Misspecification in Nonrecursive SEMs 1. Nonrecursive Latent Variable Models under Misspecification
 Misspecification in Nonrecursive SEMs 1 Nonrecursive Latent Variable Models under Misspecification Misspecification in Nonrecursive SEMs 2 Abstract A problem central to structural equation modeling is
Misspecification in Nonrecursive SEMs 1 Nonrecursive Latent Variable Models under Misspecification Misspecification in Nonrecursive SEMs 2 Abstract A problem central to structural equation modeling is
2.2 Classical Regression in the Time Series Context
 48 2 Time Series Regression and Exploratory Data Analysis context, and therefore we include some material on transformations and other techniques useful in exploratory data analysis. 2.2 Classical Regression
48 2 Time Series Regression and Exploratory Data Analysis context, and therefore we include some material on transformations and other techniques useful in exploratory data analysis. 2.2 Classical Regression
Preface. List of examples
 Contents Preface List of examples i xix 1 LISREL models and methods 1 1.1 The general LISREL model 1 Assumptions 2 The covariance matrix of the observations as implied by the LISREL model 3 Fixed, free,
Contents Preface List of examples i xix 1 LISREL models and methods 1 1.1 The general LISREL model 1 Assumptions 2 The covariance matrix of the observations as implied by the LISREL model 3 Fixed, free,
The Impact of Model Misspecification in Clustered and Continuous Growth Modeling
 The Impact of Model Misspecification in Clustered and Continuous Growth Modeling Daniel J. Bauer Odum Institute for Research in Social Science The University of North Carolina at Chapel Hill Patrick J.
The Impact of Model Misspecification in Clustered and Continuous Growth Modeling Daniel J. Bauer Odum Institute for Research in Social Science The University of North Carolina at Chapel Hill Patrick J.
Confirmatory Factor Analysis. Psych 818 DeShon
 Confirmatory Factor Analysis Psych 818 DeShon Purpose Takes factor analysis a few steps further. Impose theoretically interesting constraints on the model and examine the resulting fit of the model with
Confirmatory Factor Analysis Psych 818 DeShon Purpose Takes factor analysis a few steps further. Impose theoretically interesting constraints on the model and examine the resulting fit of the model with
An Introduction to SEM in Mplus
 An Introduction to SEM in Mplus Ben Kite Saturday Seminar Series Quantitative Training Program Center for Research Methods and Data Analysis Goals Provide an introduction to Mplus Demonstrate Mplus with
An Introduction to SEM in Mplus Ben Kite Saturday Seminar Series Quantitative Training Program Center for Research Methods and Data Analysis Goals Provide an introduction to Mplus Demonstrate Mplus with
Nesting and Equivalence Testing
 Nesting and Equivalence Testing Tihomir Asparouhov and Bengt Muthén August 13, 2018 Abstract In this note, we discuss the nesting and equivalence testing (NET) methodology developed in Bentler and Satorra
Nesting and Equivalence Testing Tihomir Asparouhov and Bengt Muthén August 13, 2018 Abstract In this note, we discuss the nesting and equivalence testing (NET) methodology developed in Bentler and Satorra
Inference using structural equations with latent variables
 This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike License. Your use of this material constitutes acceptance of that license and the conditions of use of materials on this
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike License. Your use of this material constitutes acceptance of that license and the conditions of use of materials on this
Week 5 Quantitative Analysis of Financial Markets Modeling and Forecasting Trend
 Week 5 Quantitative Analysis of Financial Markets Modeling and Forecasting Trend Christopher Ting http://www.mysmu.edu/faculty/christophert/ Christopher Ting : christopherting@smu.edu.sg : 6828 0364 :
Week 5 Quantitative Analysis of Financial Markets Modeling and Forecasting Trend Christopher Ting http://www.mysmu.edu/faculty/christophert/ Christopher Ting : christopherting@smu.edu.sg : 6828 0364 :
36-309/749 Experimental Design for Behavioral and Social Sciences. Dec 1, 2015 Lecture 11: Mixed Models (HLMs)
 36-309/749 Experimental Design for Behavioral and Social Sciences Dec 1, 2015 Lecture 11: Mixed Models (HLMs) Independent Errors Assumption An error is the deviation of an individual observed outcome (DV)
36-309/749 Experimental Design for Behavioral and Social Sciences Dec 1, 2015 Lecture 11: Mixed Models (HLMs) Independent Errors Assumption An error is the deviation of an individual observed outcome (DV)
Semiparametric Mixed Effects Models with Flexible Random Effects Distribution
 Semiparametric Mixed Effects Models with Flexible Random Effects Distribution Marie Davidian North Carolina State University davidian@stat.ncsu.edu www.stat.ncsu.edu/ davidian Joint work with A. Tsiatis,
Semiparametric Mixed Effects Models with Flexible Random Effects Distribution Marie Davidian North Carolina State University davidian@stat.ncsu.edu www.stat.ncsu.edu/ davidian Joint work with A. Tsiatis,
Model Estimation Example
 Ronald H. Heck 1 EDEP 606: Multivariate Methods (S2013) April 7, 2013 Model Estimation Example As we have moved through the course this semester, we have encountered the concept of model estimation. Discussions
Ronald H. Heck 1 EDEP 606: Multivariate Methods (S2013) April 7, 2013 Model Estimation Example As we have moved through the course this semester, we have encountered the concept of model estimation. Discussions
Brandon C. Kelly (Harvard Smithsonian Center for Astrophysics)
 Brandon C. Kelly (Harvard Smithsonian Center for Astrophysics) Probability quantifies randomness and uncertainty How do I estimate the normalization and logarithmic slope of a X ray continuum, assuming
Brandon C. Kelly (Harvard Smithsonian Center for Astrophysics) Probability quantifies randomness and uncertainty How do I estimate the normalization and logarithmic slope of a X ray continuum, assuming
Time-Invariant Predictors in Longitudinal Models
 Time-Invariant Predictors in Longitudinal Models Today s Class (or 3): Summary of steps in building unconditional models for time What happens to missing predictors Effects of time-invariant predictors
Time-Invariant Predictors in Longitudinal Models Today s Class (or 3): Summary of steps in building unconditional models for time What happens to missing predictors Effects of time-invariant predictors
TIME SERIES ANALYSIS AND FORECASTING USING THE STATISTICAL MODEL ARIMA
 CHAPTER 6 TIME SERIES ANALYSIS AND FORECASTING USING THE STATISTICAL MODEL ARIMA 6.1. Introduction A time series is a sequence of observations ordered in time. A basic assumption in the time series analysis
CHAPTER 6 TIME SERIES ANALYSIS AND FORECASTING USING THE STATISTICAL MODEL ARIMA 6.1. Introduction A time series is a sequence of observations ordered in time. A basic assumption in the time series analysis
Nonlinear mixed-effects models using Stata
 Nonlinear mixed-effects models using Stata Yulia Marchenko Executive Director of Statistics StataCorp LP 2017 German Stata Users Group meeting Yulia Marchenko (StataCorp) 1 / 48 Outline What is NLMEM?
Nonlinear mixed-effects models using Stata Yulia Marchenko Executive Director of Statistics StataCorp LP 2017 German Stata Users Group meeting Yulia Marchenko (StataCorp) 1 / 48 Outline What is NLMEM?
MLMED. User Guide. Nicholas J. Rockwood The Ohio State University Beta Version May, 2017
 MLMED User Guide Nicholas J. Rockwood The Ohio State University rockwood.19@osu.edu Beta Version May, 2017 MLmed is a computational macro for SPSS that simplifies the fitting of multilevel mediation and
MLMED User Guide Nicholas J. Rockwood The Ohio State University rockwood.19@osu.edu Beta Version May, 2017 MLmed is a computational macro for SPSS that simplifies the fitting of multilevel mediation and
Copyright 2013 The Guilford Press
 This is a chapter excerpt from Guilford Publications. Longitudinal Structural Equation Modeling, by Todd D. Little. Copyright 2013. Purchase this book now: www.guilford.com/p/little 7 Multiple-Group Models
This is a chapter excerpt from Guilford Publications. Longitudinal Structural Equation Modeling, by Todd D. Little. Copyright 2013. Purchase this book now: www.guilford.com/p/little 7 Multiple-Group Models
Preface Introduction to Statistics and Data Analysis Overview: Statistical Inference, Samples, Populations, and Experimental Design The Role of
 Preface Introduction to Statistics and Data Analysis Overview: Statistical Inference, Samples, Populations, and Experimental Design The Role of Probability Sampling Procedures Collection of Data Measures
Preface Introduction to Statistics and Data Analysis Overview: Statistical Inference, Samples, Populations, and Experimental Design The Role of Probability Sampling Procedures Collection of Data Measures
Final Review. Yang Feng. Yang Feng (Columbia University) Final Review 1 / 58
 Final Review Yang Feng http://www.stat.columbia.edu/~yangfeng Yang Feng (Columbia University) Final Review 1 / 58 Outline 1 Multiple Linear Regression (Estimation, Inference) 2 Special Topics for Multiple
Final Review Yang Feng http://www.stat.columbia.edu/~yangfeng Yang Feng (Columbia University) Final Review 1 / 58 Outline 1 Multiple Linear Regression (Estimation, Inference) 2 Special Topics for Multiple
SEM Day 1 Lab Exercises SPIDA 2007 Dave Flora
 SEM Day 1 Lab Exercises SPIDA 2007 Dave Flora 1 Today we will see how to estimate CFA models and interpret output using both SAS and LISREL. In SAS, commands for specifying SEMs are given using linear
SEM Day 1 Lab Exercises SPIDA 2007 Dave Flora 1 Today we will see how to estimate CFA models and interpret output using both SAS and LISREL. In SAS, commands for specifying SEMs are given using linear
bmuthen posted on Tuesday, August 23, :06 am The following question appeared on SEMNET Aug 19, 2005.
 Count modeling with different length of exposure uses an offset, that is, a term in the regression which has its coefficient fixed at 1. This can also be used for modeling proportions in that a count is
Count modeling with different length of exposure uses an offset, that is, a term in the regression which has its coefficient fixed at 1. This can also be used for modeling proportions in that a count is
Determining the number of components in mixture models for hierarchical data
 Determining the number of components in mixture models for hierarchical data Olga Lukočienė 1 and Jeroen K. Vermunt 2 1 Department of Methodology and Statistics, Tilburg University, P.O. Box 90153, 5000
Determining the number of components in mixture models for hierarchical data Olga Lukočienė 1 and Jeroen K. Vermunt 2 1 Department of Methodology and Statistics, Tilburg University, P.O. Box 90153, 5000
Mplus Code Corresponding to the Web Portal Customization Example
 Online supplement to Hayes, A. F., & Preacher, K. J. (2014). Statistical mediation analysis with a multicategorical independent variable. British Journal of Mathematical and Statistical Psychology, 67,
Online supplement to Hayes, A. F., & Preacher, K. J. (2014). Statistical mediation analysis with a multicategorical independent variable. British Journal of Mathematical and Statistical Psychology, 67,
Biostatistics-Lecture 16 Model Selection. Ruibin Xi Peking University School of Mathematical Sciences
 Biostatistics-Lecture 16 Model Selection Ruibin Xi Peking University School of Mathematical Sciences Motivating example1 Interested in factors related to the life expectancy (50 US states,1969-71 ) Per
Biostatistics-Lecture 16 Model Selection Ruibin Xi Peking University School of Mathematical Sciences Motivating example1 Interested in factors related to the life expectancy (50 US states,1969-71 ) Per
SIMS Variance Decomposition. SIMS Variance Decomposition (Continued)
 SIMS Variance Decomposition The Second International Mathematics Study (SIMS; Muthén, 1991, JEM). National probability sample of school districts selected proportional to size; a probability sample of
SIMS Variance Decomposition The Second International Mathematics Study (SIMS; Muthén, 1991, JEM). National probability sample of school districts selected proportional to size; a probability sample of
Mediation question: Does executive functioning mediate the relation between shyness and vocabulary? Plot data, descriptives, etc. Check for outliers
 Plot data, descriptives, etc. Check for outliers A. Nayena Blankson, Ph.D. Spelman College University of Southern California GC3 Lecture Series September 6, 2013 Treat missing i data Listwise Pairwise
Plot data, descriptives, etc. Check for outliers A. Nayena Blankson, Ph.D. Spelman College University of Southern California GC3 Lecture Series September 6, 2013 Treat missing i data Listwise Pairwise
Fall 2017 STAT 532 Homework Peter Hoff. 1. Let P be a probability measure on a collection of sets A.
 1. Let P be a probability measure on a collection of sets A. (a) For each n N, let H n be a set in A such that H n H n+1. Show that P (H n ) monotonically converges to P ( k=1 H k) as n. (b) For each n
1. Let P be a probability measure on a collection of sets A. (a) For each n N, let H n be a set in A such that H n H n+1. Show that P (H n ) monotonically converges to P ( k=1 H k) as n. (b) For each n
Logistic And Probit Regression
 Further Readings On Multilevel Regression Analysis Ludtke Marsh, Robitzsch, Trautwein, Asparouhov, Muthen (27). Analysis of group level effects using multilevel modeling: Probing a latent covariate approach.
Further Readings On Multilevel Regression Analysis Ludtke Marsh, Robitzsch, Trautwein, Asparouhov, Muthen (27). Analysis of group level effects using multilevel modeling: Probing a latent covariate approach.
Generalized Linear Models for Non-Normal Data
 Generalized Linear Models for Non-Normal Data Today s Class: 3 parts of a generalized model Models for binary outcomes Complications for generalized multivariate or multilevel models SPLH 861: Lecture
Generalized Linear Models for Non-Normal Data Today s Class: 3 parts of a generalized model Models for binary outcomes Complications for generalized multivariate or multilevel models SPLH 861: Lecture
PAPER 206 APPLIED STATISTICS
 MATHEMATICAL TRIPOS Part III Thursday, 1 June, 2017 9:00 am to 12:00 pm PAPER 206 APPLIED STATISTICS Attempt no more than FOUR questions. There are SIX questions in total. The questions carry equal weight.
MATHEMATICAL TRIPOS Part III Thursday, 1 June, 2017 9:00 am to 12:00 pm PAPER 206 APPLIED STATISTICS Attempt no more than FOUR questions. There are SIX questions in total. The questions carry equal weight.
Evaluation of structural equation models. Hans Baumgartner Penn State University
 Evaluation of structural equation models Hans Baumgartner Penn State University Issues related to the initial specification of theoretical models of interest Model specification: Measurement model: EFA
Evaluation of structural equation models Hans Baumgartner Penn State University Issues related to the initial specification of theoretical models of interest Model specification: Measurement model: EFA
Research Design: Topic 18 Hierarchical Linear Modeling (Measures within Persons) 2010 R.C. Gardner, Ph.d.
 Research Design: Topic 8 Hierarchical Linear Modeling (Measures within Persons) R.C. Gardner, Ph.d. General Rationale, Purpose, and Applications Linear Growth Models HLM can also be used with repeated
Research Design: Topic 8 Hierarchical Linear Modeling (Measures within Persons) R.C. Gardner, Ph.d. General Rationale, Purpose, and Applications Linear Growth Models HLM can also be used with repeated
One-stage dose-response meta-analysis
 One-stage dose-response meta-analysis Nicola Orsini, Alessio Crippa Biostatistics Team Department of Public Health Sciences Karolinska Institutet http://ki.se/en/phs/biostatistics-team 2017 Nordic and
One-stage dose-response meta-analysis Nicola Orsini, Alessio Crippa Biostatistics Team Department of Public Health Sciences Karolinska Institutet http://ki.se/en/phs/biostatistics-team 2017 Nordic and
Appendix 6. Selection procedure and results of best model fit for fatigue severity (CIS- FS)
 Aendix 6. Selection rocedure and results of best model fit for fatigue severity (CIS- FS) The figure below shows the various tested models for fatigue severity trajectory (CIS-FS). We tested if a linear
Aendix 6. Selection rocedure and results of best model fit for fatigue severity (CIS- FS) The figure below shows the various tested models for fatigue severity trajectory (CIS-FS). We tested if a linear
STAT 501 EXAM I NAME Spring 1999
 STAT 501 EXAM I NAME Spring 1999 Instructions: You may use only your calculator and the attached tables and formula sheet. You can detach the tables and formula sheet from the rest of this exam. Show your
STAT 501 EXAM I NAME Spring 1999 Instructions: You may use only your calculator and the attached tables and formula sheet. You can detach the tables and formula sheet from the rest of this exam. Show your
STAT Financial Time Series
 STAT 6104 - Financial Time Series Chapter 4 - Estimation in the time Domain Chun Yip Yau (CUHK) STAT 6104:Financial Time Series 1 / 46 Agenda 1 Introduction 2 Moment Estimates 3 Autoregressive Models (AR
STAT 6104 - Financial Time Series Chapter 4 - Estimation in the time Domain Chun Yip Yau (CUHK) STAT 6104:Financial Time Series 1 / 46 Agenda 1 Introduction 2 Moment Estimates 3 Autoregressive Models (AR
Contrasting Marginal and Mixed Effects Models Recall: two approaches to handling dependence in Generalized Linear Models:
 Contrasting Marginal and Mixed Effects Models Recall: two approaches to handling dependence in Generalized Linear Models: Marginal models: based on the consequences of dependence on estimating model parameters.
Contrasting Marginal and Mixed Effects Models Recall: two approaches to handling dependence in Generalized Linear Models: Marginal models: based on the consequences of dependence on estimating model parameters.
Outline. Mixed models in R using the lme4 package Part 3: Longitudinal data. Sleep deprivation data. Simple longitudinal data
 Outline Mixed models in R using the lme4 package Part 3: Longitudinal data Douglas Bates Longitudinal data: sleepstudy A model with random effects for intercept and slope University of Wisconsin - Madison
Outline Mixed models in R using the lme4 package Part 3: Longitudinal data Douglas Bates Longitudinal data: sleepstudy A model with random effects for intercept and slope University of Wisconsin - Madison
4. MA(2) +drift: y t = µ + ɛ t + θ 1 ɛ t 1 + θ 2 ɛ t 2. Mean: where θ(l) = 1 + θ 1 L + θ 2 L 2. Therefore,
 61 4. MA(2) +drift: y t = µ + ɛ t + θ 1 ɛ t 1 + θ 2 ɛ t 2 Mean: y t = µ + θ(l)ɛ t, where θ(l) = 1 + θ 1 L + θ 2 L 2. Therefore, E(y t ) = µ + θ(l)e(ɛ t ) = µ 62 Example: MA(q) Model: y t = ɛ t + θ 1 ɛ
61 4. MA(2) +drift: y t = µ + ɛ t + θ 1 ɛ t 1 + θ 2 ɛ t 2 Mean: y t = µ + θ(l)ɛ t, where θ(l) = 1 + θ 1 L + θ 2 L 2. Therefore, E(y t ) = µ + θ(l)e(ɛ t ) = µ 62 Example: MA(q) Model: y t = ɛ t + θ 1 ɛ
Dimensionality Assessment: Additional Methods
 Dimensionality Assessment: Additional Methods In Chapter 3 we use a nonlinear factor analytic model for assessing dimensionality. In this appendix two additional approaches are presented. The first strategy
Dimensionality Assessment: Additional Methods In Chapter 3 we use a nonlinear factor analytic model for assessing dimensionality. In this appendix two additional approaches are presented. The first strategy
How to run the RI CLPM with Mplus By Ellen Hamaker March 21, 2018
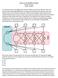 How to run the RI CLPM with Mplus By Ellen Hamaker March 21, 2018 The random intercept cross lagged panel model (RI CLPM) as proposed by Hamaker, Kuiper and Grasman (2015, Psychological Methods) is a model
How to run the RI CLPM with Mplus By Ellen Hamaker March 21, 2018 The random intercept cross lagged panel model (RI CLPM) as proposed by Hamaker, Kuiper and Grasman (2015, Psychological Methods) is a model
STAT 6350 Analysis of Lifetime Data. Probability Plotting
 STAT 6350 Analysis of Lifetime Data Probability Plotting Purpose of Probability Plots Probability plots are an important tool for analyzing data and have been particular popular in the analysis of life
STAT 6350 Analysis of Lifetime Data Probability Plotting Purpose of Probability Plots Probability plots are an important tool for analyzing data and have been particular popular in the analysis of life
FIT CRITERIA PERFORMANCE AND PARAMETER ESTIMATE BIAS IN LATENT GROWTH MODELS WITH SMALL SAMPLES
 FIT CRITERIA PERFORMANCE AND PARAMETER ESTIMATE BIAS IN LATENT GROWTH MODELS WITH SMALL SAMPLES Daniel M. McNeish Measurement, Statistics, and Evaluation University of Maryland, College Park Background
FIT CRITERIA PERFORMANCE AND PARAMETER ESTIMATE BIAS IN LATENT GROWTH MODELS WITH SMALL SAMPLES Daniel M. McNeish Measurement, Statistics, and Evaluation University of Maryland, College Park Background
Using Structural Equation Modeling to Conduct Confirmatory Factor Analysis
 Using Structural Equation Modeling to Conduct Confirmatory Factor Analysis Advanced Statistics for Researchers Session 3 Dr. Chris Rakes Website: http://csrakes.yolasite.com Email: Rakes@umbc.edu Twitter:
Using Structural Equation Modeling to Conduct Confirmatory Factor Analysis Advanced Statistics for Researchers Session 3 Dr. Chris Rakes Website: http://csrakes.yolasite.com Email: Rakes@umbc.edu Twitter:
