MODEL ASSESSMENT FOR MODELS WITH MISSING DATA. Xiaolei Zhou. Chapel Hill 2015
|
|
|
- Archibald Potter
- 5 years ago
- Views:
Transcription
1 MODEL ASSESSMENT FOR MODELS WITH MISSING DATA Xiaolei Zhou A dissertation submitted to the faculty of the University of North Carolina at Chapel Hill in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the Department of Biostatistics in the Gillings School of Global Public Health. Chapel Hill 2015 Approved by: Hongtu Zhu Elizabeth Andrews Shrikant Bangdiwala Yun Li Wen Sun
2 2015 Xiaolei Zhou ALL RIGHTS RESERVED ii
3 ABSTRACT Xiaolei Zhou: Model Assessment for Models with Missing Data (Under the direction of Hongtu Zhu) Missing data commonly occur in various study setting. In this dissertation, we first investigate three likelihood-based models for missing data in longitudinal studies: mixed effects models, pattern mixture models (PMM), and selection models. Extensive simulations from ten missing mechanisms are performed with the focus on treatment effect. Results suggest that no model consistently performs better than others under various missing data mechanism. However, PMM using the treatment-specific proportion and selection model provide some correction of the estimate compared with mixed-effects model in several missing not at random situations, even when the mechanism of missing data is not exactly the same as the model assumption. Secondly, we focus on the case deletion diagnostic measures for general linear models (GLMs) with missing covariate data. Cook's distance is one of the most important diagnostic tools to identify influential observations on the parametric models. However, Cook's distance may not be directly comparable because its scale stochastically depends on the degree of the perturbation. We define the degree of perturbation for GLM with missing covariates. Then, we derive the Cook's distance based on likelihood function and compare it to the Cook's distance based on the Q-function used in the EM algorithm for models with missing data. We further develop the scaled Cook's distance in the GLM with missing covariate data, which resolves the size issue of Cook's distance. Simulation data are used to illustrate the size iii
4 matters issue in GLM with missing covariates. The applications of scaled Cook's distances in a formal influence analysis are examined in simulations and real data examples. At last, we examine the connection between case deletion measures and cross validation method for GLM with missing covariates models. Based on such connection, we develop case-deletion model complexity (CMC) measures for quantifying the model complexity and case-deletion information criteria (CIC) for model selection. We develop these new measures and criteria based on the likelihood function and the Q-function, respectively. Some properties of CMC and CIC are investigated. Simulations and real data analysis show that CIC is a valuable tool for analysis of models with missing data. iv
5 ACKNOWLEDGMENTS Foremost, I would like to thank my advisor, Dr. Hongtu Zhu, for his guidance, wisdom, and patience throughout the development of this dissertation. In addition, this dissertation would not have been possible without the encouragement from my colleagues at RTI Health Solutions and the support from my husband, Yong Yang. v
6 TABLE OF CONTENTS LIST OF TABLES ix LIST OF FIGURES x CHAPTER 1: INTRODUCTION Missing Data and Treatment Effect Model Assessment Case Influence Measures Criterion-based Model Assessment CHAPTER 2: COMPARISON OF STATISTICAL MODELS IN ESTIMATING TREATMENT EFFECT FOR MISSING DATA IN LONGITUDINAL STUDIES Introduction Treatment Effect Existing Methods Mixed-effects Models Pattern-mixture Models Selection Models Simulation Data Generation Analysis of the Simulated Data Results vi
7 2.6 Conclusions CHAPTER 3: DIAGNOSTIC MEASURES FOR GENERALIZED LINEAR MODELS WITH MISSING COVARIATES Introduction Generalized Linear Model with Missing Covariate Data Degree of Perturbation Cook's Distance Scaled Cook's Distance Simulation Studies Using One Dataset Additional Simulation Studies Real Data Examples National Survey Cholesterol Data Liver Cancer Data Conclusions CHAPTER 4: INFORMATION CRITERIA FOR GENERALIZED LINEAR MODELS WITH MISSING COVARIATES Introduction Method Case Deletion Measures Cross Validation and Model Complexity Case-deletion Information Criterion Simulation Real Data Analysis Conclusions vii
8 APPENDIX: ASSUMPTIONS AND PROOFS BIBLIOGRAPHY viii
9 LIST OF TABLES 2.1 Treatment Effect for Four Commonly Used Models Examples of Mixed-effects Models, PMMs, and Selection Models Missing Data Mechanism and Missing Rate in Simulation Data Average Treatment Effect and Its Standard Error Estimated from the Mixed-Effects Model, the Pattern-Mixture Model, and the Selection Model for Scenarios 1 to 6 (true treatment effect = 1) Average Treatment Effect and Its Standard Error Estimated from the Mixed-Effects Model, the Pattern-Mixture Model, and the Selection model for Scenarios 7 to Summary of Simulation Scenarios Comparison of Ranks of the True Model M1 from Various Model Selection Criteria in GLM with Missing Covariates Comparison of Ranks for M1 to M5 from Various Model Selection Criteria in GLM with Missing Covariates Model Selection Results of Liver Cancer Data Model Selection Results of Liver Cancer Data, Excluding One Outlier ix
10 LIST OF FIGURES 3.1 Index Plots and Scatter Plots of CD and QCD from the Simulation Data with No Outlier. The three plots in the top are from the scenario of MAR (Scenario 1). The three plots in the middle are from the scenario of MNAR (Scenario 5). The three plots in the bottom are from the scenario of MNAR with a greater missing rate (Scenario 9) Index Plots and Scatter Plots of CD and QCD from the Simulation Data with One Outlier in z Domain. The three plots in the top are from the scenario of MAR (Scenario 2a). The three plots in the middle are from the scenario of MNAR (Scenario 6a). The three plots in the bottom are from the scenario of MNAR with a greater missing rate (Scenario 10a) Index Plots and Scatter Plots of CD and QCD from the Simulation Data with Two Outliers in z Domain in Same Direction. The three plots in the top are from the scenario of MAR (Scenario 2b). The three plots in the middle are from the scenario of MNAR (Scenario 6b). The three plots in the bottom are from the scenario of MNAR with a greater missing rate (Scenario 10b) Index Plots and Scatter Plots of CD and QCD from the Simulation Data with Two Outliers in z Domain in Opposite Direction. The three plots in the top are from the scenario of MAR (Scenario 2c). The three plots in the middle are from the scenario of MNAR (Scenario 6c). The three plots in the bottom are from the scenario of MNAR with a greater missing rate (Scenario 10c) Index Plots and Scatter Plots of CD and QCD from the Simulation Data with One Outlier in y Domain. The three plots in the top are from the scenario of MAR (Scenario 3a). The three plots in the middle are from the scenario of MNAR (Scenario 7a). The three plots in the bottom are from the scenario of MNAR with a greater missing rate (Scenario 11a) Index Plots and Scatter Plots of CD and QCD from the Simulation Data with Two Outliers in y Domain in Same Direction. The three plots in the top are from the scenario of MAR (Scenario 3b). The three plots in the middle are from the scenario of MNAR (Scenario 7b). The three plots in the bottom are from the scenario of MNAR with a greater missing rate (Scenario 11b) x
11 3.7 Index Plots and Scatter Plots of CD and QCD from the Simulation Data with Two Outliers in y Domain in Opposite Direction. The three plots in the top are from the scenario of MAR (Scenario 3c). The three plots in the middle are from the scenario of MNAR (Scenario 7c). The three plots in the bottom are from the scenario of MNAR with a greater missing rate (Scenario 11c) Index Plots and Scatter Plots of CD and QCD from the Simulation Data with One Outlier in x Domain. The three plots in the top are from the scenario of MAR (Scenario 4a). The three plots in the middle are from the scenario of MNAR (Scenario 8a). The three plots in the bottom are from the scenario of MNAR with a greater missing rate (Scenario 12a) Index Plots and Scatter Plots of CD and QCD from the Simulation Data with Two Outliers in x Domain in Same Direction. The three plots in the top are from the scenario of MAR (Scenario 4b). The three plots in the middle are from the scenario of MNAR (Scenario 8b). The three plots in the bottom are from the scenario of MNAR with a greater missing rate (Scenario 12b) Index Plots and Scatter Plots of CD and QCD from the Simulation Data with Two Outliers in x Domain in Opposite Direction. The three plots in the top are from the scenario of MAR (Scenario 4c). The three plots in the middle are from the scenario of MNAR (Scenario 8c). The three plots in the bottom are from the scenario of MNAR with a greater missing rate (Scenario 12c) Box Plots for CD, Scaled CD, Pr and Scatter Plots with Q-based Approximation. Results from 100 Simulation Samples for Scenario 1. Green dots indicates means. Red bars (dots) are observed, and blue bars (dots) are subjects with missing z Scatter Plots of Mean Degree of Perturbation with x, Mean CD, and Mean Scaled CD. Results from 100 Simulation Samples for Scenario 1. Red dots are observed subjects, and blue dots are subjects with missing z Box Plots for CD, Scaled CD, and Pr. Results from 100 Simulation Samples for Scenarios 2a (MAR, left panels) and 6a (MNAR, right panels) - 5 Outliers in z Domain in the Same Direction. Green dots indicates means. Red bars (dots) are observed, and blue bars (dots) are subjects with missing z xii
12 3.14 Box Plots for CD, Scaled CD, and Pr. Results from 100 Simulation Samples for Scenarios 2b (MAR, left panels) and 6b (MNAR, right panels) - 5 Outliers in z Domain in the Opposite Direction. Green dots indicates means. Red bars (dots) are observed, and blue bars (dots) are subjects with missing z Box Plots for CD, Scaled CD, and Pr. Results from 100 Simulation Samples for Scenarios 3a (MAR, left panels) and 7a (MNAR, right panels) - 5 Outliers in y Domain in the Same Direction. Green dots indicates means. Red bars (dots) are observed, and blue bars (dots) are subjects with missing z Box Plots for CD, Scaled CD, and Pr. Results from 100 Simulation Samples for Scenarios 3b (MAR, left panels) and 7b (MNAR, right panels) - 5 Outliers in y Domain in the Opposite Direction. Green dots indicates means. Red bars (dots) are observed, and blue bars (dots) are subjects with missing z Box Plots for CD, Scaled CD, and Pr. Results from 100 Simulation Samples for Scenarios 4a (MAR, left panels) and 8a (MNAR, right panels) - 5 Outliers in x Domain in the Same Direction. Green dots indicates means. Red bars (dots) are observed, and blue bars (dots) are subjects with missing z Box Plots for CD, Scaled CD, and Pr. Results from 100 Simulation Samples for Scenarios 4b (MAR, left panels) and 8b (MNAR, right panels) - 5 Outliers in x Domain in the Opposite Direction. Green dots indicates means. Red bars (dots) are observed, and blue bars (dots) are subjects with missing z Results from Cholesterol Data Results from Liver Cancer Data xiii
13 CHAPTER 1 INTRODUCTION In the big data era, a large amount of data is available and waiting for people to find out what is hidden inside. These data may come from a true complicated process. Statisticians use statistical models to interpret data and to approximate the true complicate process. However, the fitted model is nearly always not the true process. How to use statistical tools (diagnostic measures) to detect the discrepancies between fitted model and true process is a very important question for statisticians. We can distinguish two types of discrepancies: i) discrepancy existing between isolated observations (influential points and outliers) and the rest of the observations, and ii) systematic discrepancies between the data and the fitted value obtained from statistical models. The existence of missing data further increases the complexity of model fitting and diagnosis. Although several methods have been developed to handle missing data, it remains a challenging and active field for statisticians. In this dissertation, we first present literature reviews. Then in Chapter 2, we compare mixed-effects model, pattern-mixture model, and selection model in estimating treatment effect for missing data in longitudinal studies. In Chapter 3, we develop the scaled Cook s distance for generalized linear models with missing covariates. In Chapter 4, we develop the case-deletion information criterion for model selection on generalized linear models with missing covariates. 1
14 1.1 Missing Data and Treatment Effect Missing data commonly occur in longitudinal studies. In clinical trial studies, patientreported outcomes (PROs), such as health-related quality of life (HRQOL) and symptoms collected via validated questionnaires, often have a higher missing rate compared with clinical outcomes evaluated by physicians. In some severe diseases, such as metastatic breast cancer, it is not uncommon for 15% of patients to be missing HRQOL data even at baseline (Zhou et al., 2009). Additional missingness occurring after baseline further reduces the proportion of patients with data available for analysis. The reasons for and amount of missing HRQOL data in clinical trials depend on the disease and when and how the study is conducted and may not be similar among different treatment groups. Fairclough (2010) listed various reasons why subjects fail to complete HRQOL assessments. Some missing data may be caused by administrative reasons, such as staff forgetting to administer the questionnaire or translation not available in the patient s language. The missing value may also be related to the patient s condition; for example, the patient stated that he or she was too ill to complete the questionnaire. The high missing rate in HRQOL data can also be caused by a self-assessment questionnaire that contains a long series of questions. Once a patient has missed an HRQOL assessment, the retrospective collection of these data is usually impossible. It is well known that missing data may not only reduce the power to detect change from baseline but, more important, will lead to biased estimates of response when missingness depends on the response. At the end of 2009, the Food and Drug Administration (FDA, 2009) published a guidance for industry in using PROs in clinical trials for label claims. The guidance encourages the study to minimize patients dropouts and collect PRO data even after patients have discontinued treatment. The study s protocol and statistical analysis plan should describe how missing data will be handled in the analysis. However, the FDA does not consider any single method as preferred regarding statisti- 2
15 cal strategies to deal with missing data due to early termination of patients before the planned completion of a trial. European Medicines Agency (2010) guideline on missing data in confirmatory clinical trials concur with this. In 2010, the Panel on the Handling of Missing Data in Clinical Trials under the National Research Council (2010) published a report with recommendations that will be used not only to the FDA but also to the entire clinical trial community (Little et al., 2012, O Neill and Temple, 2012). The panel classified four types of approaches to adjust for missing data: complete-case analysis (excluding subjects with missing data from analysis), single imputation methods (such as last observation carried forward or baseline value carried forward, was used in some clinical trials), estimating-equation methods, and methods based on a statistical model. In estimating-equation methods, complete cases are weighted by the inverse of an estimate of the probability of being observed, which may be modeled with the use of observed variables, for example, baseline data. The statistical model based methods includes likelihood function-based models, Bayesian methods, and multiple imputation. For the missing data problem, the applicability of the different methods is based on a classification of the following missingness mechanisms: missing completely at random (MCAR), missing at random (MAR), and missing not at random (MNAR). If the probability of an observation being missing does not depend on observed or unobserved measurements, then the observation is MCAR. If the probability of an observation being missing depends only on observed measurements, then the observation is MAR. If the probability of an observation being missing depends on unobserved measurements (e.g., patients with a poor outcome score are more likely to miss the assessment), then the observation is MNAR. This type of missing data is also called nonignorable missing data. Complete-case analysis is based on MCAR. Single imputation methods is arbitrary. Weighted estimating equations and multiple imputation are computationally available, 3
16 but they are built on ignorable missingness or MAR (Little et al., 2012, Ali and Siddiqui, 2000). The mixed-effects model, as a likelihood function-based model, is a frequent choice for analyzing continuous outcomes in clinical trials because it uses all available observed data and are valid when missing data are MCAR or MAR. However, when missing data depend on an unobserved outcome (MNAR), the parameter estimates from the mixed-effects model can be biased. In clinical trial studies, we cannot rule out the MNAR scenario and sometimes it may be more realistic than MAR; thus, it is recommended to assess the robustness of the results by performing sensitivity analysis under the assumption of MNAR. Development of statistical methods to handle MNAR data is a very important and promising area. Ibrahim and colleagues (2005) and Ibrahim and Molenberghs (2009) provided several overview articles of various models for missing data problems. Pattern-mixture model (PMM) and selection model are two major methods to handle missing data under MNAR based on likelihood functions. To account for nonignorable missing data, in addition to random variables in the mixed-effects model, PMM and selection model include an additional random variable for missingness in the likelihood functions (missing pattern in PMM and missingness indicator in selection model). In both models, the random variable for missing mechanism is not independent with the response variable. Pattern-mixture models (Little, 1995) have been used widely to analyze continuous PRO data. A popular PPM for PRO data analysis assumes that the missing mechanism depends only on treatment (Hedeker and Gibbons, 1997). Pauler and colleagues (2003) further provided details on how to estimate overall treatment effect from pattern-specific estimates based on the treatment-specific proportion of the pattern when HRQOL changes linearly over time. The missing mechanism may also depend on other covariates or response. The selection model is based on the assumption of MNAR that whether or not the dependent variable is missing depends directly on the value of the dependent variable at the time of missing. The parameters in selection models can be estimated using the Monte Carlo expectation- 4
17 maximization (MCEM) algorithm (Ibrahim et al., 2001, 2005, Ibrahim and Molenberghs, 2009). Both PMM and selection model have been applied using the Bayesian approach (Daniels and Hogan, 2008, Little et al., 2011). Note that the pattern-mixture model and selection model factorizations of the likelihood functions can be used to develop more complex methods of joint modeling of responses and missing data process such as the shared-parameter models (Ibrahim and Molenberghs, 2009, Daniels and Hogan, 2008), where the missingness may depend on the random effect. All models for handling missing data under MNAR make specific assumptions, which are often untestable, and the statistical results obtained from different MNAR models can be different. Therefore, the models under MNAR are often considered as part of a sensitivity analysis (Molenberghs et al., 2001, 2004, Verbeke et al., 2001b). Although it is well known that the estimates of response obtained from mixed-effects models fitted to MNAR missing data can be biased, the impact of missing data on the estimate of treatment effect (the difference between treatments) is more complex, depending on the proportion of and reason for missing data in all treatment groups. Several studies (Pauler et al., 2003, Michiels et al., 2002, Post et al., 2010) have included treatment effects estimated from mixed-effects model and PMM or selection model using collected data. However, because the true treatment effect is unknown in collected data, it is impossible to evaluate the bias of the estimate using collected data. A simulation study is needed to evaluate the impact of missing data on the estimate of treatment effect. 1.2 Model Assessment The goal of the diagnostic measures is to assess how well the model fits the data and how robust it is. Residuals (the difference between the observed response and the modelpredicted response) provide very important information about the model fitting, not only for the individual observations, but also for the global model fitting. Instead of comparing observed value to model-predicted value, another approach to assess model fitting is the 5
18 influence measure. If a minor modification of the model seriously influences key results of an analysis, it will be a cause for concern. On the other hand, if such modifications do not have large impact the results, the model is robust with respect to the induced perturbations (Cook, 1986) Case Influence Measures Case deletion measures assess the influence of deleting one or a set of observations from the data on certain statistics. They are often used to identify influential points or the impact of one or a few observations on overall model fitting. Two widely used case deletion measures are Cook s distance (Cook, 1977) and likelihood displacement (Cook and Weisberg, 1982, Cook, 1986). The likelihood displacement measures the difference in log-likelihood when one or a set of observations are removed. The likelihood displacement is defined by LD(i) = 2[l(ˆθ) l(ˆθ [i] )], where θ is a p vector of the parameter of interest, ˆθ is the parameter estimated with full data, ˆ θ[i] is the parameter estimates using data with the ith case deleted, and l(θ) is the log-likelihood function for θ. The Cook s distance measures the impact of deleting one or a set of observations on parameter estimates. The generalized Cook s distance is defined by CD(i) = (ˆθ [i] ˆθ) G(ˆθ [i] ˆθ), where G is a positive definite matrix, e.g., θ 2l(ˆθ). It has been shown that Cook s distance combines information from the studentized residuals and the variance of predicted values for general linear model (Cook, 1977). Most of the diagnostic measures were originally developed under linear regression models (Cook, 1977, Cook and Weisberg, 1982, Chatterjee and Hadi, 1986), and then for more complicated models, such as generalized linear models (Davison and Tsai, 6
19 1992), generalized estimating equations (Preisser and Qaqish, 1996), models for clustered data(christensen et al., 1992, Banerjee and Frees, 1997, Haslett and Dillane, 2004), and survival data (Weissfeld, 1990, Lin et al., 1993). In addition, considerable research has been conducted to develop case influence measures in Bayesian analysis (Johnson and Geisser, 1983, 1985, Pettit, 1986, Carlin and Carlin, 1991, Gelfand et al., 1992, Weiss and Cook, 1992, Blyth, 1994, Peng and Dey, 1995, Weiss, 1996, Bradlow and Zaslavsky, 1997). Zhu et al. (2010) provided a comprehensive review of various Bayesian case influence measures and their properties. In Bayesian analysis, the influence of individual observations (or a set of observations) is often assessed by comparing the posterior (or predictive) distribution of the full data to the distribution after deleting these observations (case deletion). For example, the Cook s posterior mode distance, denoted by CP (i), quantifies the discrepancy between the posterior mode of θ with and without the ith case (Cook and Weisberg, 1982). The posterior modes of θ for the full sample Y and a subsample Y [i] are defined as ˆθ = argmax θ log p(θ Y ) and ˆθ [i] = argmax θ log p(θ Y [i] ), respectively. CP (i) is given by CP (i) = (ˆθ [i] ˆθ) G θ (ˆθ [i] ˆθ), where G θ is chosen to be a positive definite matrix. For instance, G θ can be θ 2 log p(θ Y ) = θ 2 log p(y θ) 2 θ log p(θ) evaluated at ˆθ. Similarly, the Cook s posterior mean distance, denoted by CM(i), quantifies the discrepancy between the posterior mean of θ with and without the ith case. The posterior means of θ for the full sample Y and a subsample Y [i] are defined as θ = θp(θ Y )dθ and θ [i] = θṗ(θ Y [i] )dθ, respectively. CM(i) is given by CM(i) = ( θ [i] θ) W θ ( θ [i] θ), where W θ is chosen to be a positive definite matrix. Diagnostic measures have also been developed for models with missing data (Zhu 7
20 et al., 2001, 2009, 2012b). The models for missing data usually use the EM algorithm to obtain the maximum likelihood estimates (Lipsitz and Ibrahim, 1996, Ibrahim et al., 1999). In EM algorithm, the MLE of the parameters θ in the complete likelihood function (based on complete data) is obtained through iterations that maximizes the Q-function Q(θ ˆθ) = E[l c (θ D c ) D o, ˆθ], where D c being the complete data, D o being the observed data, and l c (θ D c ) is the complete-data log-likelihood function. Zhu et al. (2001) used Q-function to replace the log likelihood in the likelihood displacement and showed that the analytic results were very similar to those obtained from a classical local influence approach based on the observed data likelihood function. Q-function is also used to obtain Cook s distance for generalized linear model with missing covariates (Zhu et al., 2009). However, to our knowledge, there is no literature to assess how close the Cook s distance obtained from the Q-function compares to that obtained from the classical likelihood function. Cook s distance is one of the most important diagnostic tools. A large value of Cook s distance indicates that the observation is influential. However, Zhu et al. (2012a) deliberated size matters issue of Cook s distance. Cook s distance may not be directly comparable because the scale of Cook s distance stochastically depends on the degree of the perturbation. Dr. Zhu et al. introduced the scaled Cook s distance to detect the relatively influential subjects in the sense that the Cook s distance is large relative to the degree of perturbation. How the missing data impacts the degree of perturbation has not been evaluated. In addition to case deletion measures, a broader range of sensitive analyses have been developed to assess the robustness of a model when perturbing the model assumptions and/or individual observations. In frequentist analysis, extensive literature exists on sensitivity analysis for missing data problems (Troxel, 1998, Zhu and Lee, 2001, Little 8
21 and Rubin, 2002, Verbeke et al., 2001a, van Steen et al., 2001, Jansen et al., 2003, 2006, Copas and Eguchi, 2005, Daniels and Hogan, 2008, Shi et al., 2009). The general questions are how to introduce an appropriate perturbation and how to assess its influence on a model. In the Bayesian analysis of statistical models with missing data, Zhu et al. (2014b) in their recent paper introduced various perturbations to modeling assumptions and individual observations, and then developed a formal sensitivity analysis to assess theses perturbations Criterion-based Model Assessment To select an optimal model from a pool of statistical models for a given dataset, we often need to consider both goodness of fit and model complexity. A model which balances model fitting and complexity is preferred. To achieve this, various information criteria have been proposed for model comparisons, which incorporate measures of fit and complexity for model choice. Such information criteria include Akaiki Information Criterion (AIC) (Akaike, 1974), Takeuchi Information Criterion (TIC) (Takeuchi, 1976), Generalized Information Criterion (GIC), (Konishi and Kitagawa, 1996), Network Information Criterion (NIC) (Murata et al., 1994) the Bayesian Information Criterion (BIC) (Schwarz et al., 1978, Lv and Liu, 2014, Konishi et al., 2004), the Deviance Information Criterion (DIC) (Spiegelhalter et al., 2002), Bayesian Predictive Information Criterion (BPIC) (Ando, 2007), and many others. In these criteria, the goodness of fit are based on the deviance component 2 log p(y ˆθ) or the posterior mean of the deviance component 2E θ y log p(y θ), while the measures of model complexity vary from simply the number of parameters to more complicated form. (Ibrahim et al., 2008) Recently, Zhu et al. (2014a) proposed a set of Bayesian case-deletion model complexity measure to quantify the effective number of parameters in a given statistical model, which leads to a Bayesian case-deletion information criterion (BCIC) for model comparison. The above mentioned model selection criteria can be difficult to obtain for models 9
22 including missing data, because these model selection criteria depend on the likelihood function based on the observed data. Some research (Garcia et al., 2010, Ibrahim et al., 2008) has been conducted to use the key components of the EM algorithm, such as the Q-function, to develop an easily computable model selection criterion. 10
23 CHAPTER 2 COMPARISON OF STATISTICAL MODELS IN ESTIMATING TREATMENT EFFECT FOR MISSING DATA IN LONGITUDINAL STUDIES 2.1 Introduction The aim of this chapter is to examine the magnitude of the bias and the robustness of mixed-effects models, PMMs, and selection models under different MNAR mechanisms that are at different degrees of perturbation to the model assumptions. Our primary interest is on the estimate of treatment effect, which is the interest of most clinical trials and many observational studies. We also perform sensitivity analyses and simulations to evaluate the robustness of the PMM, selection model, and mixed-effects model on the estimates of treatment effects. The methods discussed in this manuscript and the simulations performed were motivated by the analysis of PROs. However, they are appropriate for the broader problem of missing data. These analyses are the first we know to systematically evaluate and compare the mixed-effects models, PMMs, and selection models using simulation data. Section 2.2 defines treatment effect, and Section 2.3 reviews the three existing models: mixed-effects models, PMMs, and selection models. In Section 2.4, we describe an extensive simulation study on comparing the three statistical models under several common scenarios of missing-data mechanisms, focusing on the treatment effect. Results are presented in Section 2.5. Section 2.6 provides concluding remarks. 11
24 2.2 Treatment Effect Treatment effect is the primary interest in clinical trials. A large amount of literature has been developed regard the effect of treatments on HRQOL in various disease areas (Zhou et al., 2009, Sherrill et al., 2010). Different analysis methods have been used to evaluate the treatment effect on HRQOL. However, the statistical definition of treatment effect is not always clear or correct, even in some published methods articles. Depending on the structure of models, treatment effect can be obtained from different regression coefficients. For continuous outcomes, there are two popular types of models based on the form of outcome. One type directly uses the response score as the dependent variable, whereas the other uses change from baseline score as the dependent variable. These models can also be further classified based on whether baseline measure is included. The baseline measure is the one assessed before treatment initiation. We denote this time point as t = 0. Table 2.1 summarizes how to obtain the treatment effects for four commonly used models. We define treatment effect as the difference in the expected value of the response scores between treatments after accounting for baseline difference. In models 1 and 2, baseline value is not included in the models as a covariate. Therefore, the treatment effect is obtained by subtracting the baseline difference from the response difference. In models 3 and 4, baseline difference is accounted for by including the baseline value in the model as a covariate. 12
25 Table 2.1: Treatment Effect for Four Commonly Used Models Model Response Variable Treatment Effect Model 1 E(y t) = β 0+β 1 trt+β 2 t+β 3 trt t, t 0 Response score is the response variable, including y 0; y 0 is not a covariate Difference in E(y t) at t > 0 minus difference in E(y 0) = E(y t trt = 1) E(y t trt = 0) - [E(y 0 trt = 1) E(y 0 trt = 0)] = β 3 t, where E(y t trt = 1) = β 0+β 1+(β 2+β 3) t, E(y t trt = 0) = β 0 + β 2 t Model 2 E(y t - y 0) = β 0+β 1 trt+β 2 t+β 3 trt t, t > 0 Model 3 E(y t)=β 0+β 1 trt+β 2 t+β 3 trt t+β 4y 0, t > 0 Change from baseline score is the response variable; y 0 is not a covariate Response score after baseline is the response variable; y 0 is a covariate Difference in E(y t) at t > 0 minus difference in E(y 0) = E[y t - y 0 trt = 1]-E[y t - y 0 trt =0] = β 1+β 3 t, where E(y t - y 0 trt = 1) = β 0+β 1+(β 2+β 3)t, E(y t - y 0 trt = 0) = β 0 + β 2 t Difference in E(y t) at t > 0 given a fixed y 0 = E[y t trt = 1, y 0] - E[y t trt = 0, y 0] = β 1+β 3 t, where E(y t trt=1,y 0)= β 0+β 1+(β 2+β 3)t+β 4y 0, E(y t trt = 0, y 0)= β 0+β 2 t +β 4y 0 Model 4 E(y t - y 0)=β 0+β 1trt+β 2t+β 3 trt t+β 4y 0, t > 0 Change from baseline score is the response variable; y 0 is a covariate Difference in E(y t) at t > 0 given a fixed y 0 = E[y t trt = 1, y 0] - E[y t trt = 0, y 0] = E[y t - y 0 trt = 1, y 0] - E[y t - y 0 trt = 0, y 0] = β 1+β 3 t, where E(y t -y 0 trt=1, y 0)=β 0+β 1+(β 2+β 3)t+β 4y 0, E(y t -y 0 trt=0, y 0)=β 0+β 2t+β 4y 0 Note: trt is a 0/1 variable indicating treatment. t is used to indicate the time variable and the time index for the dependent variable ; t = 0 indicates baseline. 13
26 Note that by including the treatment-by-time interaction term, these models allow treatment effect varies by time. In model 1, treatment effect depends only on the coefficient for the trt t interaction (i.e., slope difference, β 3 ). The slope difference obtained from model 1 is the treatment effect at t = 1. If this coefficient is 0 or the interaction is not included in the model, there is no treatment effect. However, in the other three models, it is possible to estimate the treatment effect when the trt t interaction is not included. In other words, when treatment effect does not vary by time, the trt t interaction can be removed from models 2, 3, and 4. Model 3 and model 4 in Table 2.1 are equivalent. The only difference is that β 4 in model 4 equals β 4 in model 3 minus 1. When using model 4, it is often observed that the change from baseline score (y t y 0, t > 0) is negatively related to baseline score (i.e., β 4 < 0). This seems counterintuitive. However, as long as β 4 in model 4 is greater than -1, the response score (y t ) is still positively correlated to y 0. On the other hand, if β 4 in model 4 is less than -1, caution should be used because it indicates a negative correlation between y t and y 0. The treatment effects can be obtained from a linear combination of the fixed-effects coefficients. When assuming a linear relationship between the dependent variable and time (e.g., models shown in Table 2.1), researchers sometimes naively compare the difference between treatments in slope and intercept or simply compare the difference in the estimated value of the dependent variable. However, these comparisons are not always appropriate depending on the setting of the model and whether treatment groups are different at baseline. Although in clinical trials patients are randomized into two treatment groups, in some clinical trials, less than 70% of patients complete baseline PRO assessments, and the baseline scores in the two treatment groups are not always comparable. Therefore, to obtain treatment effect, the treatment difference in response scores must be adjusted for baseline difference. Otherwise, the difference between post-treatment scores may be due to the difference at baseline, not to treatment. 14
27 In Table 2.1, time t is a continuous variable. However, it can also be generalized to the case that time is a categorical variable, which allows a non-linear relationship between treatment effect and time. In general, the methods described in the rest of this paper to estimate treament effect when missing data exist are applicable to all above model structures. 2.3 Existing Methods In this section, we describe mixed-effects models, PMMs, and selection models, which are used to estimate treatment effect when missing data exist in the response variable. Let y i = (y i1,..., y ini ), where y ij denotes the response score (or the change from baseline score) of the ith subject on the jth visit for i = 1,..., N and j = 1,..., n i. In some clinical trials, n i may be the same for all subjects. For example, in a study to treat irritable bowel syndrome, all patients received 12 weeks of treatment, and adequate relief by patients as an endpoint was reported weekly (Mangel et al., 2008). However, n i may vary across subjects for example, in cancer trials where HRQOL is periodically reported by patients until disease progression, death, or withdrawal from the study for toxicity or other reasons (Zhou et al., 2009). In clinical trial data, the response y i may contain missing values with nonmonotone patterns that is, some response values are observed again after a missing value occurs. For example, y ik may be missing and y ik may be observed for some k > k. In this case, we call y ik intermittent missing. A subject may also have dropout missing, which is the missing value after the last nonmissing value that is, no response values are observed again after the missing value occurs. One subject may have an intermittent missing or a dropout missing value or both. When data are missing, we write y i = (y mis,i, y obs,i ) for convenience, where y mis,i denotes the missing components of y i, and y obs,i denotes the observed components of y i. 15
28 2.3.1 Mixed-effects Models Consider the normal mixed-effects model without missing data given by y i = X i β + Z i b i + ɛ i, i = 1,..., N, where X i is an n i p matrix of the fixed-effect covariates, and Z i is an n i q matrix of the random-effect covariates. Both X i and Z i are fixed and known, and Z i is usually a subset of X i with fewer covariates. In addition, β is a p 1 vector of unknown regression parameters, b i is a q 1 vector of random effects, and ɛ i is an n i 1 vector of errors. It is commonly assumed that all ɛ i s and b i s are independent, b i N q (0, G) and ɛ i N ni (θ, φi ni ), where G is a q q matrix and I ni is an n i n i identity matrix. Table 2.2 presents the likelihood function of the joint distribution of y i and b i for subject i. Upon integration over the random effects, the marginal distribution of y i is N ni (X i β, Z i GZi +φin i ). When missing data exist in y i, the standard mixed-effects model assumes that missing data are ignorable and the regression coefficients are estimated using the observed data, y obs,i. Specifically, we can write y obs,i = X obs,i β + Z obs,i b i + ɛ obs,i, i = 1,..., N, where X obs,i, Z obs,i, and ɛ obs,i are the subsets of X i, Z i, and ɛ i, respectively, corresponding to y obs,i. 16
29 Table 2.2: Examples of Mixed-effects Models, PMMs, and Selection Models Model Likelihood Function Distributions Mixed f(y i, b i) = y i = X iβ + Z ib i + ε i, f(y i b i ; β, φ) f(b i ; i = 1,, N G) where b i ~ N q(0, G) ε i ~ N ni(0, φ I ni ) b i ε i Note PMM f(y i, b i, s i) = f(y i b i, s i ; β k, φ) f(b i; G) f(s i), given s i b i Selection f(y i, b i, r i) = f(r i y i) f(y i b i ; β, φ) f(b i ; G), given r i y i b i s i ~ multinomial (1, π i1,, π ik) For missing pattern k, y i (k) = X i β (k) + Z ib i + ε i y i = X iβ + Z ib i + ε i, i = 1,, N r i = (r i1,, r ini) T, r ij y ij ~ Bernoulli (π ij) Logit(π ij ) = ξ 0 + ξ 1 y ij s i is the random variable that subject i belongs to a missing pattern k. π ik, k = 1,.., K, is the proportion of subjects in missing pattern k in the true population, which may depend on covariates, such as treatment, or on response. r ij is the random variable that the observation j for subject i is missing. π ij is the proportion of y ij being missing given the value of y ij in the true population. For MNAR, π ij is dependent on y ij, but it can also depend on response variables at other time points or on covariates. Note: X i is an n i p matrix of the fixed-effect covariates. Z i is an n i q matrix of the random-effect covariates. Both X i and Z i are fixed and known, and Z i is usually a subset of X i with fewer covariates. β is a p 1 vector of unknown regression parameters, b i is a q 1 vector of random effects, and ε i is an n i 1 vector of errors. G is a q q matrix and I ni is an n i n i identity matrix. 17
30 2.3.2 Pattern-mixture Models In addition to the response variables and the random effects, the likelihood function of a PMM includes the missing pattern stratum variable s i, which is usually assumed independent of b i. For subject i, which is in pattern k, the likelihood function is given by L i,p MM = f(y i, b i, s i ) = f(s i )f(y i b i, s i ; β k, φ)f(b i ; G), where s i multinomial(1, π i1,..., π ik ), and π i1,..., π ik are the proportion of subjects in all K missing patterns in the true population, which may depend on covariates, such as treatment, or on response. Moreover, y i, b i, and s i are assumed independent for different subjects. Conditional on the kth missing pattern, for subject i, the complete data y i = (y mis,i, y obs,i ) are given by where the superscript (k) on y (k) i y (k) i = X i β (k) + Z i b i + ɛ i, indicates that subject i is in missing pattern k, and β (k) is a p 1 vector of unknown regression parameters specifically for pattern k. Other model assumptions are the same as those in the mixed-effects model. The PMM is conducted by simply adding the covariate missing pattern and its interaction terms with other fixed-effect covariates into a mixed-effect model. Parameter estimates can be obtained by standard statistical software such as PROC MIXED in SAS. The estimated response value given each dropout pattern can be obtained from pattern-specific regression parameters. However, when estimating the response variable for time points after dropout, additional assumptions (restriction) must be made, for example, based on complete case, available case, or neighboring case (Fairclough, 2010). These assumptions are not testable due to lack of data. The estimation of the response value across all missing patterns can be obtained by 18
31 a weighted sum of pattern-specific estimates given by E(y X) = K E(y X, pattern = k)π k, k=1 where π k is the proportion of patients with the missing pattern, and X is fixed such as treatment and baseline score. A PMM assumes that a subject belongs to a specific missing pattern. When performing a PMM analysis, initially, K strata are created based on missing patterns. There are several different ways of defining strata. In practice, the missing pattern strata are often defined based on last visit before or after a certain time point (Hedeker and Gibbons, 1997). However, creating strata based only on last visit makes a strong assumption that missing data within each stratum (i.e., intermittent missing) are ignorable (MCAR, or MAR). Strata have also been defined based on reasons for withdrawal (e.g., death, disease progression, toxicity, or other reason) (Pauler et al., 2003, Post et al., 2010). In summary, the choice of strata (or pattern) is a clinical judgment or is made for computational convenience. The parameters π ik, k = 1,..., K, may or may not vary by subpopulation. In HRQOL analysis, two popular assumptions are made on the mechanism, by which data are missing: (i) π ik does not depend on any variable, or (ii) π ik depends on treatment. Under assumption (i), π ik is estimated by the proportion of the kth missing pattern in the whole sample. Under assumption (ii), π ik is estimated by the proportion of the kth missing pattern in each treatment group, separately Selection Models Selection models get the name that observations were selected to be missing. Selection models assume that the complete data, y i, follow a distribution and the probability of missingness depends on the current value of y i, which may be missing. The conditional 19
32 probability of missingness, π ij, may be assumed to follow a logistic regression in which response may be included as a covariate. In selection models, the complete data y i = (y mis,i, y obs,i ) are typically assumed to follow the same model given by y i = X i β + Z i b i + ɛ i, i = 1,..., N. The selection model introduces an n i vector of the missing data mechanism variable r i = (r i1,..., r i,ni ), where r ij = 1 indicating that y ij is missing. Conditional on y i, r i is commonly assumed to be independent of b i. Thus, for subject i, the joint distribution of y i, b i, and r i is given by L i,sm = f(y i, b i, r i ) = f(r i y i )f(y i b i ; β, φ)f(b i ; G), where r ij y ij Bernoulli(π ij ), and π ij is the proportion of y ij being missing given the value of y ij in the true population. f(r i y i ) is often modeled by logistic regression or probit models. For MNAR, π ij is dependent on y ij, but it can also depend on response variables at other time points or on covariates. For different subjects, y i, b i, and r i are assumed independent. Ibrahim and Molenberghs (2009) and Ibrahim and colleagues (2001) described the MCEM algorithm used for parametric estimation in selection models (Ibrahim et al., 2001, Ibrahim and Molenberghs, 2009). 2.4 Simulation We conducted an extensive simulation study to compare the three existing models handling of missing data under various missing patterns and mechanisms Data Generation We simulated missing data on the response variable according to 10 different missing mechanisms (Table 2.3). These missing mechanisms included MCAR (scenario 1); MAR 20
33 (missing depending on treatment [scenario 2], the baseline score [scenario 3], or the response at the previous assessment [scenario 4]); and MNAR (missing depending on response at the current assessment [scenarios 5a and 5b]). We also considered intermittent missing data in scenario 5b. Scenario 6 assumed that the missing mechanism was different between two treatment groups: MCAR in one treatment group and MNAR in the other treatment group. In scenarios 7 to 10, data were generated based on the assumption of PMM. Specifically, subjects first were assigned randomly to one of the four dropout groups: pattern 1, only one visit at time 1; pattern 2, last visit at time 2; pattern 3, last visit at time 3; and pattern 4, last visit at time 4 (completer). In these scenarios, the response variable depended on the dropout group to which a particular subject belonged. In all scenarios, subjects were assigned randomly at a 1:1 ratio to one of the two treatment groups. For each subject, a continuous baseline covariate x i was generated from N(0, 1), and the random-effect variable b i was generated from N(0, 1). The measurement errors ɛ ij, i = 1,..., N, j = 1,..., n i, were generated independently from N(0, 1). 21
BAYESIAN INFLUENCE DIAGNOSTIC METHODS FOR PARAMETRIC REGRESSION MODELS
 BAYESIAN INFLUENCE DIAGNOSTIC METHODS FOR PARAMETRIC REGRESSION MODELS Hyunsoon Cho A dissertation submitted to the faculty of the University of North Carolina at Chapel Hill in partial fulfillment of
BAYESIAN INFLUENCE DIAGNOSTIC METHODS FOR PARAMETRIC REGRESSION MODELS Hyunsoon Cho A dissertation submitted to the faculty of the University of North Carolina at Chapel Hill in partial fulfillment of
A Bayesian Nonparametric Approach to Monotone Missing Data in Longitudinal Studies with Informative Missingness
 A Bayesian Nonparametric Approach to Monotone Missing Data in Longitudinal Studies with Informative Missingness A. Linero and M. Daniels UF, UT-Austin SRC 2014, Galveston, TX 1 Background 2 Working model
A Bayesian Nonparametric Approach to Monotone Missing Data in Longitudinal Studies with Informative Missingness A. Linero and M. Daniels UF, UT-Austin SRC 2014, Galveston, TX 1 Background 2 Working model
Fractional Imputation in Survey Sampling: A Comparative Review
 Fractional Imputation in Survey Sampling: A Comparative Review Shu Yang Jae-Kwang Kim Iowa State University Joint Statistical Meetings, August 2015 Outline Introduction Fractional imputation Features Numerical
Fractional Imputation in Survey Sampling: A Comparative Review Shu Yang Jae-Kwang Kim Iowa State University Joint Statistical Meetings, August 2015 Outline Introduction Fractional imputation Features Numerical
Comparing Group Means When Nonresponse Rates Differ
 UNF Digital Commons UNF Theses and Dissertations Student Scholarship 2015 Comparing Group Means When Nonresponse Rates Differ Gabriela M. Stegmann University of North Florida Suggested Citation Stegmann,
UNF Digital Commons UNF Theses and Dissertations Student Scholarship 2015 Comparing Group Means When Nonresponse Rates Differ Gabriela M. Stegmann University of North Florida Suggested Citation Stegmann,
Discussion of Missing Data Methods in Longitudinal Studies: A Review by Ibrahim and Molenberghs
 Discussion of Missing Data Methods in Longitudinal Studies: A Review by Ibrahim and Molenberghs Michael J. Daniels and Chenguang Wang Jan. 18, 2009 First, we would like to thank Joe and Geert for a carefully
Discussion of Missing Data Methods in Longitudinal Studies: A Review by Ibrahim and Molenberghs Michael J. Daniels and Chenguang Wang Jan. 18, 2009 First, we would like to thank Joe and Geert for a carefully
Statistical Methods. Missing Data snijders/sm.htm. Tom A.B. Snijders. November, University of Oxford 1 / 23
 1 / 23 Statistical Methods Missing Data http://www.stats.ox.ac.uk/ snijders/sm.htm Tom A.B. Snijders University of Oxford November, 2011 2 / 23 Literature: Joseph L. Schafer and John W. Graham, Missing
1 / 23 Statistical Methods Missing Data http://www.stats.ox.ac.uk/ snijders/sm.htm Tom A.B. Snijders University of Oxford November, 2011 2 / 23 Literature: Joseph L. Schafer and John W. Graham, Missing
Statistical Practice
 Statistical Practice A Note on Bayesian Inference After Multiple Imputation Xiang ZHOU and Jerome P. REITER This article is aimed at practitioners who plan to use Bayesian inference on multiply-imputed
Statistical Practice A Note on Bayesian Inference After Multiple Imputation Xiang ZHOU and Jerome P. REITER This article is aimed at practitioners who plan to use Bayesian inference on multiply-imputed
Whether to use MMRM as primary estimand.
 Whether to use MMRM as primary estimand. James Roger London School of Hygiene & Tropical Medicine, London. PSI/EFSPI European Statistical Meeting on Estimands. Stevenage, UK: 28 September 2015. 1 / 38
Whether to use MMRM as primary estimand. James Roger London School of Hygiene & Tropical Medicine, London. PSI/EFSPI European Statistical Meeting on Estimands. Stevenage, UK: 28 September 2015. 1 / 38
A Note on Bayesian Inference After Multiple Imputation
 A Note on Bayesian Inference After Multiple Imputation Xiang Zhou and Jerome P. Reiter Abstract This article is aimed at practitioners who plan to use Bayesian inference on multiplyimputed datasets in
A Note on Bayesian Inference After Multiple Imputation Xiang Zhou and Jerome P. Reiter Abstract This article is aimed at practitioners who plan to use Bayesian inference on multiplyimputed datasets in
Factor Analytic Models of Clustered Multivariate Data with Informative Censoring (refer to Dunson and Perreault, 2001, Biometrics 57, )
 Factor Analytic Models of Clustered Multivariate Data with Informative Censoring (refer to Dunson and Perreault, 2001, Biometrics 57, 302-308) Consider data in which multiple outcomes are collected for
Factor Analytic Models of Clustered Multivariate Data with Informative Censoring (refer to Dunson and Perreault, 2001, Biometrics 57, 302-308) Consider data in which multiple outcomes are collected for
Multi-state Models: An Overview
 Multi-state Models: An Overview Andrew Titman Lancaster University 14 April 2016 Overview Introduction to multi-state modelling Examples of applications Continuously observed processes Intermittently observed
Multi-state Models: An Overview Andrew Titman Lancaster University 14 April 2016 Overview Introduction to multi-state modelling Examples of applications Continuously observed processes Intermittently observed
Bayesian Inference on Joint Mixture Models for Survival-Longitudinal Data with Multiple Features. Yangxin Huang
 Bayesian Inference on Joint Mixture Models for Survival-Longitudinal Data with Multiple Features Yangxin Huang Department of Epidemiology and Biostatistics, COPH, USF, Tampa, FL yhuang@health.usf.edu January
Bayesian Inference on Joint Mixture Models for Survival-Longitudinal Data with Multiple Features Yangxin Huang Department of Epidemiology and Biostatistics, COPH, USF, Tampa, FL yhuang@health.usf.edu January
A weighted simulation-based estimator for incomplete longitudinal data models
 To appear in Statistics and Probability Letters, 113 (2016), 16-22. doi 10.1016/j.spl.2016.02.004 A weighted simulation-based estimator for incomplete longitudinal data models Daniel H. Li 1 and Liqun
To appear in Statistics and Probability Letters, 113 (2016), 16-22. doi 10.1016/j.spl.2016.02.004 A weighted simulation-based estimator for incomplete longitudinal data models Daniel H. Li 1 and Liqun
MISSING or INCOMPLETE DATA
 MISSING or INCOMPLETE DATA A (fairly) complete review of basic practice Don McLeish and Cyntha Struthers University of Waterloo Dec 5, 2015 Structure of the Workshop Session 1 Common methods for dealing
MISSING or INCOMPLETE DATA A (fairly) complete review of basic practice Don McLeish and Cyntha Struthers University of Waterloo Dec 5, 2015 Structure of the Workshop Session 1 Common methods for dealing
Sparse Linear Models (10/7/13)
 STA56: Probabilistic machine learning Sparse Linear Models (0/7/) Lecturer: Barbara Engelhardt Scribes: Jiaji Huang, Xin Jiang, Albert Oh Sparsity Sparsity has been a hot topic in statistics and machine
STA56: Probabilistic machine learning Sparse Linear Models (0/7/) Lecturer: Barbara Engelhardt Scribes: Jiaji Huang, Xin Jiang, Albert Oh Sparsity Sparsity has been a hot topic in statistics and machine
ECLT 5810 Linear Regression and Logistic Regression for Classification. Prof. Wai Lam
 ECLT 5810 Linear Regression and Logistic Regression for Classification Prof. Wai Lam Linear Regression Models Least Squares Input vectors is an attribute / feature / predictor (independent variable) The
ECLT 5810 Linear Regression and Logistic Regression for Classification Prof. Wai Lam Linear Regression Models Least Squares Input vectors is an attribute / feature / predictor (independent variable) The
Bayesian methods for missing data: part 1. Key Concepts. Nicky Best and Alexina Mason. Imperial College London
 Bayesian methods for missing data: part 1 Key Concepts Nicky Best and Alexina Mason Imperial College London BAYES 2013, May 21-23, Erasmus University Rotterdam Missing Data: Part 1 BAYES2013 1 / 68 Outline
Bayesian methods for missing data: part 1 Key Concepts Nicky Best and Alexina Mason Imperial College London BAYES 2013, May 21-23, Erasmus University Rotterdam Missing Data: Part 1 BAYES2013 1 / 68 Outline
Niche Modeling. STAMPS - MBL Course Woods Hole, MA - August 9, 2016
 Niche Modeling Katie Pollard & Josh Ladau Gladstone Institutes UCSF Division of Biostatistics, Institute for Human Genetics and Institute for Computational Health Science STAMPS - MBL Course Woods Hole,
Niche Modeling Katie Pollard & Josh Ladau Gladstone Institutes UCSF Division of Biostatistics, Institute for Human Genetics and Institute for Computational Health Science STAMPS - MBL Course Woods Hole,
Empirical Likelihood Methods for Two-sample Problems with Data Missing-by-Design
 1 / 32 Empirical Likelihood Methods for Two-sample Problems with Data Missing-by-Design Changbao Wu Department of Statistics and Actuarial Science University of Waterloo (Joint work with Min Chen and Mary
1 / 32 Empirical Likelihood Methods for Two-sample Problems with Data Missing-by-Design Changbao Wu Department of Statistics and Actuarial Science University of Waterloo (Joint work with Min Chen and Mary
Statistical Methods for Handling Incomplete Data Chapter 2: Likelihood-based approach
 Statistical Methods for Handling Incomplete Data Chapter 2: Likelihood-based approach Jae-Kwang Kim Department of Statistics, Iowa State University Outline 1 Introduction 2 Observed likelihood 3 Mean Score
Statistical Methods for Handling Incomplete Data Chapter 2: Likelihood-based approach Jae-Kwang Kim Department of Statistics, Iowa State University Outline 1 Introduction 2 Observed likelihood 3 Mean Score
Generalized Linear Models
 Generalized Linear Models Lecture 3. Hypothesis testing. Goodness of Fit. Model diagnostics GLM (Spring, 2018) Lecture 3 1 / 34 Models Let M(X r ) be a model with design matrix X r (with r columns) r n
Generalized Linear Models Lecture 3. Hypothesis testing. Goodness of Fit. Model diagnostics GLM (Spring, 2018) Lecture 3 1 / 34 Models Let M(X r ) be a model with design matrix X r (with r columns) r n
Latent Variable Model for Weight Gain Prevention Data with Informative Intermittent Missingness
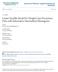 Journal of Modern Applied Statistical Methods Volume 15 Issue 2 Article 36 11-1-2016 Latent Variable Model for Weight Gain Prevention Data with Informative Intermittent Missingness Li Qin Yale University,
Journal of Modern Applied Statistical Methods Volume 15 Issue 2 Article 36 11-1-2016 Latent Variable Model for Weight Gain Prevention Data with Informative Intermittent Missingness Li Qin Yale University,
Analyzing Group by Time Effects in Longitudinal Two-Group Randomized Trial Designs With Missing Data
 Journal of Modern Applied Statistical Methods Volume Issue 1 Article 6 5-1-003 Analyzing Group by Time Effects in Longitudinal Two-Group Randomized Trial Designs With Missing Data James Algina University
Journal of Modern Applied Statistical Methods Volume Issue 1 Article 6 5-1-003 Analyzing Group by Time Effects in Longitudinal Two-Group Randomized Trial Designs With Missing Data James Algina University
Semiparametric Mixed Effects Models with Flexible Random Effects Distribution
 Semiparametric Mixed Effects Models with Flexible Random Effects Distribution Marie Davidian North Carolina State University davidian@stat.ncsu.edu www.stat.ncsu.edu/ davidian Joint work with A. Tsiatis,
Semiparametric Mixed Effects Models with Flexible Random Effects Distribution Marie Davidian North Carolina State University davidian@stat.ncsu.edu www.stat.ncsu.edu/ davidian Joint work with A. Tsiatis,
Missing Data in Longitudinal Studies: Mixed-effects Pattern-Mixture and Selection Models
 Missing Data in Longitudinal Studies: Mixed-effects Pattern-Mixture and Selection Models Hedeker D & Gibbons RD (1997). Application of random-effects pattern-mixture models for missing data in longitudinal
Missing Data in Longitudinal Studies: Mixed-effects Pattern-Mixture and Selection Models Hedeker D & Gibbons RD (1997). Application of random-effects pattern-mixture models for missing data in longitudinal
A STRATEGY FOR STEPWISE REGRESSION PROCEDURES IN SURVIVAL ANALYSIS WITH MISSING COVARIATES. by Jia Li B.S., Beijing Normal University, 1998
 A STRATEGY FOR STEPWISE REGRESSION PROCEDURES IN SURVIVAL ANALYSIS WITH MISSING COVARIATES by Jia Li B.S., Beijing Normal University, 1998 Submitted to the Graduate Faculty of the Graduate School of Public
A STRATEGY FOR STEPWISE REGRESSION PROCEDURES IN SURVIVAL ANALYSIS WITH MISSING COVARIATES by Jia Li B.S., Beijing Normal University, 1998 Submitted to the Graduate Faculty of the Graduate School of Public
Analyzing Pilot Studies with Missing Observations
 Analyzing Pilot Studies with Missing Observations Monnie McGee mmcgee@smu.edu. Department of Statistical Science Southern Methodist University, Dallas, Texas Co-authored with N. Bergasa (SUNY Downstate
Analyzing Pilot Studies with Missing Observations Monnie McGee mmcgee@smu.edu. Department of Statistical Science Southern Methodist University, Dallas, Texas Co-authored with N. Bergasa (SUNY Downstate
DIAGNOSTICS FOR STRATIFIED CLINICAL TRIALS IN PROPORTIONAL ODDS MODELS
 DIAGNOSTICS FOR STRATIFIED CLINICAL TRIALS IN PROPORTIONAL ODDS MODELS Ivy Liu and Dong Q. Wang School of Mathematics, Statistics and Computer Science Victoria University of Wellington New Zealand Corresponding
DIAGNOSTICS FOR STRATIFIED CLINICAL TRIALS IN PROPORTIONAL ODDS MODELS Ivy Liu and Dong Q. Wang School of Mathematics, Statistics and Computer Science Victoria University of Wellington New Zealand Corresponding
2 Naïve Methods. 2.1 Complete or available case analysis
 2 Naïve Methods Before discussing methods for taking account of missingness when the missingness pattern can be assumed to be MAR in the next three chapters, we review some simple methods for handling
2 Naïve Methods Before discussing methods for taking account of missingness when the missingness pattern can be assumed to be MAR in the next three chapters, we review some simple methods for handling
Shu Yang and Jae Kwang Kim. Harvard University and Iowa State University
 Statistica Sinica 27 (2017), 000-000 doi:https://doi.org/10.5705/ss.202016.0155 DISCUSSION: DISSECTING MULTIPLE IMPUTATION FROM A MULTI-PHASE INFERENCE PERSPECTIVE: WHAT HAPPENS WHEN GOD S, IMPUTER S AND
Statistica Sinica 27 (2017), 000-000 doi:https://doi.org/10.5705/ss.202016.0155 DISCUSSION: DISSECTING MULTIPLE IMPUTATION FROM A MULTI-PHASE INFERENCE PERSPECTIVE: WHAT HAPPENS WHEN GOD S, IMPUTER S AND
Analysis of Incomplete Non-Normal Longitudinal Lipid Data
 Analysis of Incomplete Non-Normal Longitudinal Lipid Data Jiajun Liu*, Devan V. Mehrotra, Xiaoming Li, and Kaifeng Lu 2 Merck Research Laboratories, PA/NJ 2 Forrest Laboratories, NY *jiajun_liu@merck.com
Analysis of Incomplete Non-Normal Longitudinal Lipid Data Jiajun Liu*, Devan V. Mehrotra, Xiaoming Li, and Kaifeng Lu 2 Merck Research Laboratories, PA/NJ 2 Forrest Laboratories, NY *jiajun_liu@merck.com
Bayesian Model Diagnostics and Checking
 Earvin Balderama Quantitative Ecology Lab Department of Forestry and Environmental Resources North Carolina State University April 12, 2013 1 / 34 Introduction MCMCMC 2 / 34 Introduction MCMCMC Steps in
Earvin Balderama Quantitative Ecology Lab Department of Forestry and Environmental Resources North Carolina State University April 12, 2013 1 / 34 Introduction MCMCMC 2 / 34 Introduction MCMCMC Steps in
Review. Timothy Hanson. Department of Statistics, University of South Carolina. Stat 770: Categorical Data Analysis
 Review Timothy Hanson Department of Statistics, University of South Carolina Stat 770: Categorical Data Analysis 1 / 22 Chapter 1: background Nominal, ordinal, interval data. Distributions: Poisson, binomial,
Review Timothy Hanson Department of Statistics, University of South Carolina Stat 770: Categorical Data Analysis 1 / 22 Chapter 1: background Nominal, ordinal, interval data. Distributions: Poisson, binomial,
Impact of serial correlation structures on random effect misspecification with the linear mixed model.
 Impact of serial correlation structures on random effect misspecification with the linear mixed model. Brandon LeBeau University of Iowa file:///c:/users/bleb/onedrive%20 %20University%20of%20Iowa%201/JournalArticlesInProgress/Diss/Study2/Pres/pres.html#(2)
Impact of serial correlation structures on random effect misspecification with the linear mixed model. Brandon LeBeau University of Iowa file:///c:/users/bleb/onedrive%20 %20University%20of%20Iowa%201/JournalArticlesInProgress/Diss/Study2/Pres/pres.html#(2)
Basics of Modern Missing Data Analysis
 Basics of Modern Missing Data Analysis Kyle M. Lang Center for Research Methods and Data Analysis University of Kansas March 8, 2013 Topics to be Covered An introduction to the missing data problem Missing
Basics of Modern Missing Data Analysis Kyle M. Lang Center for Research Methods and Data Analysis University of Kansas March 8, 2013 Topics to be Covered An introduction to the missing data problem Missing
For more information about how to cite these materials visit
 Author(s): Kerby Shedden, Ph.D., 2010 License: Unless otherwise noted, this material is made available under the terms of the Creative Commons Attribution Share Alike 3.0 License: http://creativecommons.org/licenses/by-sa/3.0/
Author(s): Kerby Shedden, Ph.D., 2010 License: Unless otherwise noted, this material is made available under the terms of the Creative Commons Attribution Share Alike 3.0 License: http://creativecommons.org/licenses/by-sa/3.0/
Statistical Practice. Selecting the Best Linear Mixed Model Under REML. Matthew J. GURKA
 Matthew J. GURKA Statistical Practice Selecting the Best Linear Mixed Model Under REML Restricted maximum likelihood (REML) estimation of the parameters of the mixed model has become commonplace, even
Matthew J. GURKA Statistical Practice Selecting the Best Linear Mixed Model Under REML Restricted maximum likelihood (REML) estimation of the parameters of the mixed model has become commonplace, even
Variable selection for model-based clustering
 Variable selection for model-based clustering Matthieu Marbac (Ensai - Crest) Joint works with: M. Sedki (Univ. Paris-sud) and V. Vandewalle (Univ. Lille 2) The problem Objective: Estimation of a partition
Variable selection for model-based clustering Matthieu Marbac (Ensai - Crest) Joint works with: M. Sedki (Univ. Paris-sud) and V. Vandewalle (Univ. Lille 2) The problem Objective: Estimation of a partition
Longitudinal + Reliability = Joint Modeling
 Longitudinal + Reliability = Joint Modeling Carles Serrat Institute of Statistics and Mathematics Applied to Building CYTED-HAROSA International Workshop November 21-22, 2013 Barcelona Mainly from Rizopoulos,
Longitudinal + Reliability = Joint Modeling Carles Serrat Institute of Statistics and Mathematics Applied to Building CYTED-HAROSA International Workshop November 21-22, 2013 Barcelona Mainly from Rizopoulos,
Introduction to Bayesian Statistics and Markov Chain Monte Carlo Estimation. EPSY 905: Multivariate Analysis Spring 2016 Lecture #10: April 6, 2016
 Introduction to Bayesian Statistics and Markov Chain Monte Carlo Estimation EPSY 905: Multivariate Analysis Spring 2016 Lecture #10: April 6, 2016 EPSY 905: Intro to Bayesian and MCMC Today s Class An
Introduction to Bayesian Statistics and Markov Chain Monte Carlo Estimation EPSY 905: Multivariate Analysis Spring 2016 Lecture #10: April 6, 2016 EPSY 905: Intro to Bayesian and MCMC Today s Class An
LOGISTIC REGRESSION Joseph M. Hilbe
 LOGISTIC REGRESSION Joseph M. Hilbe Arizona State University Logistic regression is the most common method used to model binary response data. When the response is binary, it typically takes the form of
LOGISTIC REGRESSION Joseph M. Hilbe Arizona State University Logistic regression is the most common method used to model binary response data. When the response is binary, it typically takes the form of
Lifetime prediction and confidence bounds in accelerated degradation testing for lognormal response distributions with an Arrhenius rate relationship
 Scholars' Mine Doctoral Dissertations Student Research & Creative Works Spring 01 Lifetime prediction and confidence bounds in accelerated degradation testing for lognormal response distributions with
Scholars' Mine Doctoral Dissertations Student Research & Creative Works Spring 01 Lifetime prediction and confidence bounds in accelerated degradation testing for lognormal response distributions with
A Flexible Bayesian Approach to Monotone Missing. Data in Longitudinal Studies with Nonignorable. Missingness with Application to an Acute
 A Flexible Bayesian Approach to Monotone Missing Data in Longitudinal Studies with Nonignorable Missingness with Application to an Acute Schizophrenia Clinical Trial Antonio R. Linero, Michael J. Daniels
A Flexible Bayesian Approach to Monotone Missing Data in Longitudinal Studies with Nonignorable Missingness with Application to an Acute Schizophrenia Clinical Trial Antonio R. Linero, Michael J. Daniels
On Estimating the Relationship between Longitudinal Measurements and Time-to-Event Data Using a Simple Two-Stage Procedure
 Biometrics DOI: 10.1111/j.1541-0420.2009.01324.x On Estimating the Relationship between Longitudinal Measurements and Time-to-Event Data Using a Simple Two-Stage Procedure Paul S. Albert 1, and Joanna
Biometrics DOI: 10.1111/j.1541-0420.2009.01324.x On Estimating the Relationship between Longitudinal Measurements and Time-to-Event Data Using a Simple Two-Stage Procedure Paul S. Albert 1, and Joanna
Joint Modeling of Longitudinal Item Response Data and Survival
 Joint Modeling of Longitudinal Item Response Data and Survival Jean-Paul Fox University of Twente Department of Research Methodology, Measurement and Data Analysis Faculty of Behavioural Sciences Enschede,
Joint Modeling of Longitudinal Item Response Data and Survival Jean-Paul Fox University of Twente Department of Research Methodology, Measurement and Data Analysis Faculty of Behavioural Sciences Enschede,
T E C H N I C A L R E P O R T KERNEL WEIGHTED INFLUENCE MEASURES. HENS, N., AERTS, M., MOLENBERGHS, G., THIJS, H. and G. VERBEKE
 T E C H N I C A L R E P O R T 0465 KERNEL WEIGHTED INFLUENCE MEASURES HENS, N., AERTS, M., MOLENBERGHS, G., THIJS, H. and G. VERBEKE * I A P S T A T I S T I C S N E T W O R K INTERUNIVERSITY ATTRACTION
T E C H N I C A L R E P O R T 0465 KERNEL WEIGHTED INFLUENCE MEASURES HENS, N., AERTS, M., MOLENBERGHS, G., THIJS, H. and G. VERBEKE * I A P S T A T I S T I C S N E T W O R K INTERUNIVERSITY ATTRACTION
Fractional Hot Deck Imputation for Robust Inference Under Item Nonresponse in Survey Sampling
 Fractional Hot Deck Imputation for Robust Inference Under Item Nonresponse in Survey Sampling Jae-Kwang Kim 1 Iowa State University June 26, 2013 1 Joint work with Shu Yang Introduction 1 Introduction
Fractional Hot Deck Imputation for Robust Inference Under Item Nonresponse in Survey Sampling Jae-Kwang Kim 1 Iowa State University June 26, 2013 1 Joint work with Shu Yang Introduction 1 Introduction
COS513 LECTURE 8 STATISTICAL CONCEPTS
 COS513 LECTURE 8 STATISTICAL CONCEPTS NIKOLAI SLAVOV AND ANKUR PARIKH 1. MAKING MEANINGFUL STATEMENTS FROM JOINT PROBABILITY DISTRIBUTIONS. A graphical model (GM) represents a family of probability distributions
COS513 LECTURE 8 STATISTICAL CONCEPTS NIKOLAI SLAVOV AND ANKUR PARIKH 1. MAKING MEANINGFUL STATEMENTS FROM JOINT PROBABILITY DISTRIBUTIONS. A graphical model (GM) represents a family of probability distributions
Analysing longitudinal data when the visit times are informative
 Analysing longitudinal data when the visit times are informative Eleanor Pullenayegum, PhD Scientist, Hospital for Sick Children Associate Professor, University of Toronto eleanor.pullenayegum@sickkids.ca
Analysing longitudinal data when the visit times are informative Eleanor Pullenayegum, PhD Scientist, Hospital for Sick Children Associate Professor, University of Toronto eleanor.pullenayegum@sickkids.ca
A note on multiple imputation for general purpose estimation
 A note on multiple imputation for general purpose estimation Shu Yang Jae Kwang Kim SSC meeting June 16, 2015 Shu Yang, Jae Kwang Kim Multiple Imputation June 16, 2015 1 / 32 Introduction Basic Setup Assume
A note on multiple imputation for general purpose estimation Shu Yang Jae Kwang Kim SSC meeting June 16, 2015 Shu Yang, Jae Kwang Kim Multiple Imputation June 16, 2015 1 / 32 Introduction Basic Setup Assume
9/12/17. Types of learning. Modeling data. Supervised learning: Classification. Supervised learning: Regression. Unsupervised learning: Clustering
 Types of learning Modeling data Supervised: we know input and targets Goal is to learn a model that, given input data, accurately predicts target data Unsupervised: we know the input only and want to make
Types of learning Modeling data Supervised: we know input and targets Goal is to learn a model that, given input data, accurately predicts target data Unsupervised: we know the input only and want to make
Comparison of methods for repeated measures binary data with missing values. Farhood Mohammadi. A thesis submitted in partial fulfillment of the
 Comparison of methods for repeated measures binary data with missing values by Farhood Mohammadi A thesis submitted in partial fulfillment of the requirements for the degree of Master of Science in Biostatistics
Comparison of methods for repeated measures binary data with missing values by Farhood Mohammadi A thesis submitted in partial fulfillment of the requirements for the degree of Master of Science in Biostatistics
Likelihood-based inference with missing data under missing-at-random
 Likelihood-based inference with missing data under missing-at-random Jae-kwang Kim Joint work with Shu Yang Department of Statistics, Iowa State University May 4, 014 Outline 1. Introduction. Parametric
Likelihood-based inference with missing data under missing-at-random Jae-kwang Kim Joint work with Shu Yang Department of Statistics, Iowa State University May 4, 014 Outline 1. Introduction. Parametric
MARGINALIZED REGRESSION MODELS FOR LONGITUDINAL CATEGORICAL DATA
 MARGINALIZED REGRESSION MODELS FOR LONGITUDINAL CATEGORICAL DATA By KEUNBAIK LEE A DISSERTATION PRESENTED TO THE GRADUATE SCHOOL OF THE UNIVERSITY OF FLORIDA IN PARTIAL FULFILLMENT OF THE REQUIREMENTS
MARGINALIZED REGRESSION MODELS FOR LONGITUDINAL CATEGORICAL DATA By KEUNBAIK LEE A DISSERTATION PRESENTED TO THE GRADUATE SCHOOL OF THE UNIVERSITY OF FLORIDA IN PARTIAL FULFILLMENT OF THE REQUIREMENTS
Missing Data: Theory & Methods. Introduction
 STAT 992/BMI 826 University of Wisconsin-Madison Missing Data: Theory & Methods Chapter 1 Introduction Lu Mao lmao@biostat.wisc.edu 1-1 Introduction Statistical analysis with missing data is a rich and
STAT 992/BMI 826 University of Wisconsin-Madison Missing Data: Theory & Methods Chapter 1 Introduction Lu Mao lmao@biostat.wisc.edu 1-1 Introduction Statistical analysis with missing data is a rich and
Case Study in the Use of Bayesian Hierarchical Modeling and Simulation for Design and Analysis of a Clinical Trial
 Case Study in the Use of Bayesian Hierarchical Modeling and Simulation for Design and Analysis of a Clinical Trial William R. Gillespie Pharsight Corporation Cary, North Carolina, USA PAGE 2003 Verona,
Case Study in the Use of Bayesian Hierarchical Modeling and Simulation for Design and Analysis of a Clinical Trial William R. Gillespie Pharsight Corporation Cary, North Carolina, USA PAGE 2003 Verona,
Penalized Loss functions for Bayesian Model Choice
 Penalized Loss functions for Bayesian Model Choice Martyn International Agency for Research on Cancer Lyon, France 13 November 2009 The pure approach For a Bayesian purist, all uncertainty is represented
Penalized Loss functions for Bayesian Model Choice Martyn International Agency for Research on Cancer Lyon, France 13 November 2009 The pure approach For a Bayesian purist, all uncertainty is represented
Extending causal inferences from a randomized trial to a target population
 Extending causal inferences from a randomized trial to a target population Issa Dahabreh Center for Evidence Synthesis in Health, Brown University issa dahabreh@brown.edu January 16, 2019 Issa Dahabreh
Extending causal inferences from a randomized trial to a target population Issa Dahabreh Center for Evidence Synthesis in Health, Brown University issa dahabreh@brown.edu January 16, 2019 Issa Dahabreh
7. Estimation and hypothesis testing. Objective. Recommended reading
 7. Estimation and hypothesis testing Objective In this chapter, we show how the election of estimators can be represented as a decision problem. Secondly, we consider the problem of hypothesis testing
7. Estimation and hypothesis testing Objective In this chapter, we show how the election of estimators can be represented as a decision problem. Secondly, we consider the problem of hypothesis testing
Nonrespondent subsample multiple imputation in two-phase random sampling for nonresponse
 Nonrespondent subsample multiple imputation in two-phase random sampling for nonresponse Nanhua Zhang Division of Biostatistics & Epidemiology Cincinnati Children s Hospital Medical Center (Joint work
Nonrespondent subsample multiple imputation in two-phase random sampling for nonresponse Nanhua Zhang Division of Biostatistics & Epidemiology Cincinnati Children s Hospital Medical Center (Joint work
Variational Bayesian Inference for Parametric and Non-Parametric Regression with Missing Predictor Data
 for Parametric and Non-Parametric Regression with Missing Predictor Data August 23, 2010 Introduction Bayesian inference For parametric regression: long history (e.g. Box and Tiao, 1973; Gelman, Carlin,
for Parametric and Non-Parametric Regression with Missing Predictor Data August 23, 2010 Introduction Bayesian inference For parametric regression: long history (e.g. Box and Tiao, 1973; Gelman, Carlin,
Performance Comparison of K-Means and Expectation Maximization with Gaussian Mixture Models for Clustering EE6540 Final Project
 Performance Comparison of K-Means and Expectation Maximization with Gaussian Mixture Models for Clustering EE6540 Final Project Devin Cornell & Sushruth Sastry May 2015 1 Abstract In this article, we explore
Performance Comparison of K-Means and Expectation Maximization with Gaussian Mixture Models for Clustering EE6540 Final Project Devin Cornell & Sushruth Sastry May 2015 1 Abstract In this article, we explore
Statistics 203: Introduction to Regression and Analysis of Variance Course review
 Statistics 203: Introduction to Regression and Analysis of Variance Course review Jonathan Taylor - p. 1/?? Today Review / overview of what we learned. - p. 2/?? General themes in regression models Specifying
Statistics 203: Introduction to Regression and Analysis of Variance Course review Jonathan Taylor - p. 1/?? Today Review / overview of what we learned. - p. 2/?? General themes in regression models Specifying
Some methods for handling missing values in outcome variables. Roderick J. Little
 Some methods for handling missing values in outcome variables Roderick J. Little Missing data principles Likelihood methods Outline ML, Bayes, Multiple Imputation (MI) Robust MAR methods Predictive mean
Some methods for handling missing values in outcome variables Roderick J. Little Missing data principles Likelihood methods Outline ML, Bayes, Multiple Imputation (MI) Robust MAR methods Predictive mean
1 Theoretical Examples
 Supplementary Document of Perturbation and Scaled Cook s Distances Hongtu Zhu, Joseph G. Ibrahim and Hyunsoon Cho Abstract We investigate two theoretical examples on generalized linear models and linear
Supplementary Document of Perturbation and Scaled Cook s Distances Hongtu Zhu, Joseph G. Ibrahim and Hyunsoon Cho Abstract We investigate two theoretical examples on generalized linear models and linear
Pattern Mixture Models for the Analysis of Repeated Attempt Designs
 Biometrics 71, 1160 1167 December 2015 DOI: 10.1111/biom.12353 Mixture Models for the Analysis of Repeated Attempt Designs Michael J. Daniels, 1, * Dan Jackson, 2, ** Wei Feng, 3, *** and Ian R. White
Biometrics 71, 1160 1167 December 2015 DOI: 10.1111/biom.12353 Mixture Models for the Analysis of Repeated Attempt Designs Michael J. Daniels, 1, * Dan Jackson, 2, ** Wei Feng, 3, *** and Ian R. White
Shared Random Parameter Models for Informative Missing Data
 Shared Random Parameter Models for Informative Missing Data Dean Follmann NIAID NIAID/NIH p. A General Set-up Longitudinal data (Y ij,x ij,r ij ) Y ij = outcome for person i on visit j R ij = 1 if observed
Shared Random Parameter Models for Informative Missing Data Dean Follmann NIAID NIAID/NIH p. A General Set-up Longitudinal data (Y ij,x ij,r ij ) Y ij = outcome for person i on visit j R ij = 1 if observed
Discussion of Identifiability and Estimation of Causal Effects in Randomized. Trials with Noncompliance and Completely Non-ignorable Missing Data
 Biometrics 000, 000 000 DOI: 000 000 0000 Discussion of Identifiability and Estimation of Causal Effects in Randomized Trials with Noncompliance and Completely Non-ignorable Missing Data Dylan S. Small
Biometrics 000, 000 000 DOI: 000 000 0000 Discussion of Identifiability and Estimation of Causal Effects in Randomized Trials with Noncompliance and Completely Non-ignorable Missing Data Dylan S. Small
ST 790, Homework 1 Spring 2017
 ST 790, Homework 1 Spring 2017 1. In EXAMPLE 1 of Chapter 1 of the notes, it is shown at the bottom of page 22 that the complete case estimator for the mean µ of an outcome Y given in (1.18) under MNAR
ST 790, Homework 1 Spring 2017 1. In EXAMPLE 1 of Chapter 1 of the notes, it is shown at the bottom of page 22 that the complete case estimator for the mean µ of an outcome Y given in (1.18) under MNAR
Ronald Christensen. University of New Mexico. Albuquerque, New Mexico. Wesley Johnson. University of California, Irvine. Irvine, California
 Texts in Statistical Science Bayesian Ideas and Data Analysis An Introduction for Scientists and Statisticians Ronald Christensen University of New Mexico Albuquerque, New Mexico Wesley Johnson University
Texts in Statistical Science Bayesian Ideas and Data Analysis An Introduction for Scientists and Statisticians Ronald Christensen University of New Mexico Albuquerque, New Mexico Wesley Johnson University
Linear Mixed Models for Longitudinal Data with Nonrandom Dropouts
 Journal of Data Science 4(2006), 447-460 Linear Mixed Models for Longitudinal Data with Nonrandom Dropouts Ahmed M. Gad and Noha A. Youssif Cairo University Abstract: Longitudinal studies represent one
Journal of Data Science 4(2006), 447-460 Linear Mixed Models for Longitudinal Data with Nonrandom Dropouts Ahmed M. Gad and Noha A. Youssif Cairo University Abstract: Longitudinal studies represent one
MISSING or INCOMPLETE DATA
 MISSING or INCOMPLETE DATA A (fairly) complete review of basic practice Don McLeish and Cyntha Struthers University of Waterloo Dec 5, 2015 Structure of the Workshop Session 1 Common methods for dealing
MISSING or INCOMPLETE DATA A (fairly) complete review of basic practice Don McLeish and Cyntha Struthers University of Waterloo Dec 5, 2015 Structure of the Workshop Session 1 Common methods for dealing
Alexina Mason. Department of Epidemiology and Biostatistics Imperial College, London. 16 February 2010
 Strategy for modelling non-random missing data mechanisms in longitudinal studies using Bayesian methods: application to income data from the Millennium Cohort Study Alexina Mason Department of Epidemiology
Strategy for modelling non-random missing data mechanisms in longitudinal studies using Bayesian methods: application to income data from the Millennium Cohort Study Alexina Mason Department of Epidemiology
Mixed-Effects Pattern-Mixture Models for Incomplete Longitudinal Data. Don Hedeker University of Illinois at Chicago
 Mixed-Effects Pattern-Mixture Models for Incomplete Longitudinal Data Don Hedeker University of Illinois at Chicago This work was supported by National Institute of Mental Health Contract N44MH32056. 1
Mixed-Effects Pattern-Mixture Models for Incomplete Longitudinal Data Don Hedeker University of Illinois at Chicago This work was supported by National Institute of Mental Health Contract N44MH32056. 1
Hierarchical Models & Bayesian Model Selection
 Hierarchical Models & Bayesian Model Selection Geoffrey Roeder Departments of Computer Science and Statistics University of British Columbia Jan. 20, 2016 Contact information Please report any typos or
Hierarchical Models & Bayesian Model Selection Geoffrey Roeder Departments of Computer Science and Statistics University of British Columbia Jan. 20, 2016 Contact information Please report any typos or
Regression Clustering
 Regression Clustering In regression clustering, we assume a model of the form y = f g (x, θ g ) + ɛ g for observations y and x in the g th group. Usually, of course, we assume linear models of the form
Regression Clustering In regression clustering, we assume a model of the form y = f g (x, θ g ) + ɛ g for observations y and x in the g th group. Usually, of course, we assume linear models of the form
Generating Half-normal Plot for Zero-inflated Binomial Regression
 Paper SP05 Generating Half-normal Plot for Zero-inflated Binomial Regression Zhao Yang, Xuezheng Sun Department of Epidemiology & Biostatistics University of South Carolina, Columbia, SC 29208 SUMMARY
Paper SP05 Generating Half-normal Plot for Zero-inflated Binomial Regression Zhao Yang, Xuezheng Sun Department of Epidemiology & Biostatistics University of South Carolina, Columbia, SC 29208 SUMMARY
Introduction An approximated EM algorithm Simulation studies Discussion
 1 / 33 An Approximated Expectation-Maximization Algorithm for Analysis of Data with Missing Values Gong Tang Department of Biostatistics, GSPH University of Pittsburgh NISS Workshop on Nonignorable Nonresponse
1 / 33 An Approximated Expectation-Maximization Algorithm for Analysis of Data with Missing Values Gong Tang Department of Biostatistics, GSPH University of Pittsburgh NISS Workshop on Nonignorable Nonresponse
A CONNECTION BETWEEN LOCAL AND DELETION INFLUENCE
 Sankhyā : The Indian Journal of Statistics 2000, Volume 62, Series A, Pt. 1, pp. 144 149 A CONNECTION BETWEEN LOCAL AND DELETION INFLUENCE By M. MERCEDES SUÁREZ RANCEL and MIGUEL A. GONZÁLEZ SIERRA University
Sankhyā : The Indian Journal of Statistics 2000, Volume 62, Series A, Pt. 1, pp. 144 149 A CONNECTION BETWEEN LOCAL AND DELETION INFLUENCE By M. MERCEDES SUÁREZ RANCEL and MIGUEL A. GONZÁLEZ SIERRA University
Multivariate Survival Analysis
 Multivariate Survival Analysis Previously we have assumed that either (X i, δ i ) or (X i, δ i, Z i ), i = 1,..., n, are i.i.d.. This may not always be the case. Multivariate survival data can arise in
Multivariate Survival Analysis Previously we have assumed that either (X i, δ i ) or (X i, δ i, Z i ), i = 1,..., n, are i.i.d.. This may not always be the case. Multivariate survival data can arise in
REGRESSION ANALYSIS IN LONGITUDINAL STUDIES WITH NON-IGNORABLE MISSING OUTCOMES
 REGRESSION ANALYSIS IN LONGITUDINAL STUDIES WITH NON-IGNORABLE MISSING OUTCOMES by Changyu Shen B.S. in Biology University of Science and Technology of China, 1998 Submitted to the Graduate Faculty of
REGRESSION ANALYSIS IN LONGITUDINAL STUDIES WITH NON-IGNORABLE MISSING OUTCOMES by Changyu Shen B.S. in Biology University of Science and Technology of China, 1998 Submitted to the Graduate Faculty of
Stat 5101 Lecture Notes
 Stat 5101 Lecture Notes Charles J. Geyer Copyright 1998, 1999, 2000, 2001 by Charles J. Geyer May 7, 2001 ii Stat 5101 (Geyer) Course Notes Contents 1 Random Variables and Change of Variables 1 1.1 Random
Stat 5101 Lecture Notes Charles J. Geyer Copyright 1998, 1999, 2000, 2001 by Charles J. Geyer May 7, 2001 ii Stat 5101 (Geyer) Course Notes Contents 1 Random Variables and Change of Variables 1 1.1 Random
Data Integration for Big Data Analysis for finite population inference
 for Big Data Analysis for finite population inference Jae-kwang Kim ISU January 23, 2018 1 / 36 What is big data? 2 / 36 Data do not speak for themselves Knowledge Reproducibility Information Intepretation
for Big Data Analysis for finite population inference Jae-kwang Kim ISU January 23, 2018 1 / 36 What is big data? 2 / 36 Data do not speak for themselves Knowledge Reproducibility Information Intepretation
Guideline on adjustment for baseline covariates in clinical trials
 26 February 2015 EMA/CHMP/295050/2013 Committee for Medicinal Products for Human Use (CHMP) Guideline on adjustment for baseline covariates in clinical trials Draft Agreed by Biostatistics Working Party
26 February 2015 EMA/CHMP/295050/2013 Committee for Medicinal Products for Human Use (CHMP) Guideline on adjustment for baseline covariates in clinical trials Draft Agreed by Biostatistics Working Party
Model Selection in GLMs. (should be able to implement frequentist GLM analyses!) Today: standard frequentist methods for model selection
 Model Selection in GLMs Last class: estimability/identifiability, analysis of deviance, standard errors & confidence intervals (should be able to implement frequentist GLM analyses!) Today: standard frequentist
Model Selection in GLMs Last class: estimability/identifiability, analysis of deviance, standard errors & confidence intervals (should be able to implement frequentist GLM analyses!) Today: standard frequentist
Model-based cluster analysis: a Defence. Gilles Celeux Inria Futurs
 Model-based cluster analysis: a Defence Gilles Celeux Inria Futurs Model-based cluster analysis Model-based clustering (MBC) consists of assuming that the data come from a source with several subpopulations.
Model-based cluster analysis: a Defence Gilles Celeux Inria Futurs Model-based cluster analysis Model-based clustering (MBC) consists of assuming that the data come from a source with several subpopulations.
STAT 5500/6500 Conditional Logistic Regression for Matched Pairs
 STAT 5500/6500 Conditional Logistic Regression for Matched Pairs Motivating Example: The data we will be using comes from a subset of data taken from the Los Angeles Study of the Endometrial Cancer Data
STAT 5500/6500 Conditional Logistic Regression for Matched Pairs Motivating Example: The data we will be using comes from a subset of data taken from the Los Angeles Study of the Endometrial Cancer Data
TECHNICAL REPORT # 59 MAY Interim sample size recalculation for linear and logistic regression models: a comprehensive Monte-Carlo study
 TECHNICAL REPORT # 59 MAY 2013 Interim sample size recalculation for linear and logistic regression models: a comprehensive Monte-Carlo study Sergey Tarima, Peng He, Tao Wang, Aniko Szabo Division of Biostatistics,
TECHNICAL REPORT # 59 MAY 2013 Interim sample size recalculation for linear and logistic regression models: a comprehensive Monte-Carlo study Sergey Tarima, Peng He, Tao Wang, Aniko Szabo Division of Biostatistics,
BAYESIAN ESTIMATION OF LINEAR STATISTICAL MODEL BIAS
 BAYESIAN ESTIMATION OF LINEAR STATISTICAL MODEL BIAS Andrew A. Neath 1 and Joseph E. Cavanaugh 1 Department of Mathematics and Statistics, Southern Illinois University, Edwardsville, Illinois 606, USA
BAYESIAN ESTIMATION OF LINEAR STATISTICAL MODEL BIAS Andrew A. Neath 1 and Joseph E. Cavanaugh 1 Department of Mathematics and Statistics, Southern Illinois University, Edwardsville, Illinois 606, USA
Longitudinal analysis of ordinal data
 Longitudinal analysis of ordinal data A report on the external research project with ULg Anne-Françoise Donneau, Murielle Mauer June 30 th 2009 Generalized Estimating Equations (Liang and Zeger, 1986)
Longitudinal analysis of ordinal data A report on the external research project with ULg Anne-Françoise Donneau, Murielle Mauer June 30 th 2009 Generalized Estimating Equations (Liang and Zeger, 1986)
Analysis of Matched Case Control Data in Presence of Nonignorable Missing Exposure
 Biometrics DOI: 101111/j1541-0420200700828x Analysis of Matched Case Control Data in Presence of Nonignorable Missing Exposure Samiran Sinha 1, and Tapabrata Maiti 2, 1 Department of Statistics, Texas
Biometrics DOI: 101111/j1541-0420200700828x Analysis of Matched Case Control Data in Presence of Nonignorable Missing Exposure Samiran Sinha 1, and Tapabrata Maiti 2, 1 Department of Statistics, Texas
ANALYSIS OF ORDINAL SURVEY RESPONSES WITH DON T KNOW
 SSC Annual Meeting, June 2015 Proceedings of the Survey Methods Section ANALYSIS OF ORDINAL SURVEY RESPONSES WITH DON T KNOW Xichen She and Changbao Wu 1 ABSTRACT Ordinal responses are frequently involved
SSC Annual Meeting, June 2015 Proceedings of the Survey Methods Section ANALYSIS OF ORDINAL SURVEY RESPONSES WITH DON T KNOW Xichen She and Changbao Wu 1 ABSTRACT Ordinal responses are frequently involved
Model Assessment and Comparisons
 Model Assessment and Comparisons Sudipto Banerjee 1 and Andrew O. Finley 2 1 Biostatistics, School of Public Health, University of Minnesota, Minneapolis, Minnesota, U.S.A. 2 Department of Forestry & Department
Model Assessment and Comparisons Sudipto Banerjee 1 and Andrew O. Finley 2 1 Biostatistics, School of Public Health, University of Minnesota, Minneapolis, Minnesota, U.S.A. 2 Department of Forestry & Department
an introduction to bayesian inference
 with an application to network analysis http://jakehofman.com january 13, 2010 motivation would like models that: provide predictive and explanatory power are complex enough to describe observed phenomena
with an application to network analysis http://jakehofman.com january 13, 2010 motivation would like models that: provide predictive and explanatory power are complex enough to describe observed phenomena
Harvard University. Harvard University Biostatistics Working Paper Series
 Harvard University Harvard University Biostatistics Working Paper Series Year 2010 Paper 117 Estimating Causal Effects in Trials Involving Multi-treatment Arms Subject to Non-compliance: A Bayesian Frame-work
Harvard University Harvard University Biostatistics Working Paper Series Year 2010 Paper 117 Estimating Causal Effects in Trials Involving Multi-treatment Arms Subject to Non-compliance: A Bayesian Frame-work
Chapter 4. Parametric Approach. 4.1 Introduction
 Chapter 4 Parametric Approach 4.1 Introduction The missing data problem is already a classical problem that has not been yet solved satisfactorily. This problem includes those situations where the dependent
Chapter 4 Parametric Approach 4.1 Introduction The missing data problem is already a classical problem that has not been yet solved satisfactorily. This problem includes those situations where the dependent
Maximum Likelihood, Logistic Regression, and Stochastic Gradient Training
 Maximum Likelihood, Logistic Regression, and Stochastic Gradient Training Charles Elkan elkan@cs.ucsd.edu January 17, 2013 1 Principle of maximum likelihood Consider a family of probability distributions
Maximum Likelihood, Logistic Regression, and Stochastic Gradient Training Charles Elkan elkan@cs.ucsd.edu January 17, 2013 1 Principle of maximum likelihood Consider a family of probability distributions
Prerequisite: STATS 7 or STATS 8 or AP90 or (STATS 120A and STATS 120B and STATS 120C). AP90 with a minimum score of 3
 University of California, Irvine 2017-2018 1 Statistics (STATS) Courses STATS 5. Seminar in Data Science. 1 Unit. An introduction to the field of Data Science; intended for entering freshman and transfers.
University of California, Irvine 2017-2018 1 Statistics (STATS) Courses STATS 5. Seminar in Data Science. 1 Unit. An introduction to the field of Data Science; intended for entering freshman and transfers.
6. Fractional Imputation in Survey Sampling
 6. Fractional Imputation in Survey Sampling 1 Introduction Consider a finite population of N units identified by a set of indices U = {1, 2,, N} with N known. Associated with each unit i in the population
6. Fractional Imputation in Survey Sampling 1 Introduction Consider a finite population of N units identified by a set of indices U = {1, 2,, N} with N known. Associated with each unit i in the population
Causal Inference with General Treatment Regimes: Generalizing the Propensity Score
 Causal Inference with General Treatment Regimes: Generalizing the Propensity Score David van Dyk Department of Statistics, University of California, Irvine vandyk@stat.harvard.edu Joint work with Kosuke
Causal Inference with General Treatment Regimes: Generalizing the Propensity Score David van Dyk Department of Statistics, University of California, Irvine vandyk@stat.harvard.edu Joint work with Kosuke
