CLASSIFIER FUSION FOR POORLY-DIFFERENTIATED TUMOR CLASSIFICATION USING BOTH MESSENGER RNA AND MICRORNA EXPRESSION PROFILES
|
|
|
- Teresa Houston
- 6 years ago
- Views:
Transcription
1 1 CLASSIFIER FUSION FOR POORLY-DIFFERENTIATED TUMOR CLASSIFICATION USING BOTH MESSENGER RNA AND MICRORNA EXPRESSION PROFILES Yuhang Wang and Margaret H. Dunham Department of Computer Science and Engineering, Southern Methodist University, James A. Waddle Department of Biology, Southern Methodist University, Monnie McGee Department of Statistical Science, Southern Methodist University, MicroRNAs (mirnas) are an important class of small non-coding RNAs that regulate diverse biological processes. MiRNAs are thought to regulate gene expression by degrading or repressing target messenger RNAs (mrnas) at the post-transcriptional level. Recent studies suggest that mirnas are implicated in human cancers. In a recent paper, Lu et al. showed that the expression profile of 217 mammalian mirnas could be used to successfully classify poorly differentiated tumor samples at the accuracy of 70.6%, whereas the same classifier using mrna profiles resulted in a low accuracy of 5.9%. Because mirnas regulate gene expression at the post-transcriptional level, we hypothesize that mirna expression profiles can provide information that is complementary to mrna expression profiles. Therefore, a data fusion approach could lead to improved classification accuracy. As a proof of concept, we re-analyzed the data in the paper by Lu et al. using a classifier fusion approach that utilizes both mrna and mirna expression data. We built a meta-classifier from two bagged k-nearest-neighbor classifiers. Experimental results showed that our meta-classifier was able to classify the same set of poorly differentiated tumor samples at an improved accuracy of 76.5%, when trained only with the expression profiles of more-differentiated tumor samples. 1. INTRODUCTION MicroRNAs (mirnas) are an important class of small non-coding RNAs that regulate diverse biological processes 3. MiRNAs are thought to regulate gene expression by degrading or repressing target messenger RNAs (mrnas) at the post-transcriptional level 1. They have been shown to control cell growth, differentiation and apoptosis 3, 1. Consequently, recent studies suggest that mirnas are implicated in human tumorigenesis 5, 8 and differentially expressed in cancers 17, 20. Most interestingly, Lu et al. 17 recently showed that the expression profile of 217 mammalian mirnas could be used to successfully classify poorly differentiated tumor samples at the accuracy of 70.6%, whereas the same classifier using messenger RNA profiles (microarray gene expression data) resulted in a low accuracy of 5.9%. On the other hand, significant work has been done in cancer classification based on microarray gene expression data 9, 2, 22. Because mirnas regulate gene expression at the post-transcriptional level, we hypothesize that mirna expression profiles can provide information that is complementary to mrna expression profiles. Therefore, a data fusion approach could lead to improved classification accuracy. To investigate this issue, we propose a metaclassifier that uses both mirna and mrna expression profiles to classify poorly differentiated tumor samples. As a proof of concept, we then re-analyze the data set used in the paper by Lu et al. 17 using our approach. The remainder of the paper is organized as follows. We give a brief review of the data fusion methods in the literature in the next section. In Section 3, we describe our proposed classifier fusion approach. Corresponding author.
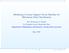
2 2 Section 4 presents the experiment results on the data set used in the paper by Lu et al. 17. Section 5 concludes the paper. 2. RELATED WORK ON DATA FUSION Data fusion is a well-studied subject in machine learning. By means of data fusion, different sources of information are combined to improve the performances of a system. Using an appropriate fusion scheme, one may expect improved classification accuracy due to the use of complementary information. Data fusion processes are often categorized in three levels, depending on the processing stage at which fusion takes place: (1) Low-level fusion 11 combines several sources of raw data to produce new raw data that is expected to be more informative and synthetic than the inputs. This kind of fusion is not feasible for integrating mrna and microrna expression data. (2) Intermediate-level fusion or feature-level fusion 24, 10. Here various features extracted from several sources of raw data are combined into a composite feature that may then be used by further processing stages. In the context of cancer classification using both mrna and microrna expression data, one possible featurelevel fusion approach can be a linear concatenation of the mrna feature vectors (mrna expression profiles for tissue samples) and the mirna feature vectors (mirna expression profiles for tissue samples). However, we tested this approach on data in Lu et al. 17 and found that it actually lead to poorer classification performance (data not shown) than what was reported by Lu et al. 17 using mirna expression profiles only. (3) High-level fusion 7, 12 (or decision fusion, classifier fusion) in which each source of input yields a decision and the decisions are fused. The classifiers may return a soft class label and not a hard decision. To distinguish both cases, one speaks of hard and soft fusion. Methods of decision fusion 16 include voting methods, statistical methods, fuzzy logic based methods, etc. The classifier fusion approach we propose in this paper falls under this category. In our data fusion problem, we only have two data sources (mirna and mrna expression profiles). Therefore, simple classifier fusion schemes such as majority voting cannot be used here. Our proposed meta-classifier utilizes two bagged fuzzy k-nearestneighbor classifiers and picks the output of the more confident one as the final classification output. Thus our approach is well-suited for integrating two data sources. We described the proposed approach in detail in the next section. 3. CLASSIFIER FUSION FOR mrna AND mirna EXPRESSION DATA INTEGRATION An overview of our proposed classifier fusion system is illustrated in Figure 1. In this approach, we first apply the Relief-F feature selection algorithm 15 to the training data to select top-ranked informative genes from mrna expression data and mirna expression data separately. Then, using only the selected genes as features, for each of the two types of expression data, a fuzzy k-nearest-neighbor (k-nn) classifier augmented with bagging is trained on the training data and applied to the test data. For each tissue sample in the test data set, the classification outputs (soft class labels) from the two classifiers are compared, and the output from the classifier with better confidence is then chosen as the final class assignment Gene Selection using Relief-F Relief-F 15 is one of the most widely used feature selection algorithm. The basic idea of Relief-F is to draw instances at random, compute their nearest neighbors, and adjust a feature weighting vector to give more weight to features that discriminate the instance from neighbors of different classes. Specifically, it tries to find a good estimate of the following probability to assign as the weight for each feature f. w f = P (different value of f different class) P (different value of f same class) This approach has shown good performance
3 3 mirna Expression Data for Traning Set mrna Expression Data for Traning Set Relief-F Feature Selection Relief-F Feature Selection mirna Expression Data for Bagged Fuzzy k-nn Classifier Bagged Fuzzy k-nn Classifier mrna Expression Data for Soft Class Label for Soft Class Label for Decision Fusion Final Class Label for Fig. 1. Classifier fusion system for cancer classification. in various domains 19, including informative gene selection Fuzzy k-nn Classifiers Here we chose to use the fuzzy k-nn classifier because it supports non-linear decision boundaries and is naturally applicable to multi-class classification problems. The k-nn classifier 6 is a well-known nonparametric classifier. Fuzzy k-nn extends k-nn by replacing the crisp class labels with soft labels, l(v i ) [0, 1] c. Different ways of combining the soft labels of the k nearest neighbors to compute the soft output label l(x) for x X have been proposed 13, 14, 6. In this work, we use the scheme by Keller et al. 14. To find the ith component of the soft class label l i (x) for x, the method by Keller et al. 14 takes distances into consideration: k j=1 l i (x) = l i(z j )(d j ) 2 m 1 k j=1 (d (1) j) 2 m 1 d j is the distance between x and its jth nearest neighbor z j. In our case, however, the class labels of the training data are actually crisp labels. If a training sample z belongs to class i, the ith component of the soft class label l i (z) for z is 1, and all other components are Bagging Bagging classifiers 4 is a method for generating multiple versions of a classifier and using them to obtain an aggregated classifier. It is also used to address the inherent instability of results when applying classifiers to relatively small data sets. Here, multiple versions of the fuzzy k-nna classifier are obtained by repeatedly sub-sample (with replacement) from the training data. The outputs (soft class labels) from those classifiers are then simply averaged. where m is a fuzzification parameter;
4 4 Table 1. Comparison of classification accuracies and P values. Classification accuracy P value Classifier fusion 76.5% Bagged fuzzy k-nn classifier using mirna expression data 70.6% Bagged fuzzy k-nn classifier using mrna expression data 47.1% Original approach using mirna expression data 70.6% Original approach using mrnaexpression data 5.9% Decision fusion Two bagged fuzzy k-nn classifiers are trained with mrna and mirna expression data separately. For each tissue sample in the test data set, the decision of the two classifiers are aggregated by choosing the decision of the classifier with higher confidence. Note that we do not average the outputs from the two classifiers. One of the two classifiers is actually chosen based on its confidence. Here, we use the highest value in the soft class label as the degree of confidence of the classifier about the class assignment of a test sample. 4. RESULTS 4.1. Data We downloaded the processed mirna expression data and the corresponding mrna expression data (published in Ref. 18) from edu/cancer/pub/migcm. The data has two parts: a training set of 68 more differentiated tumors, representing 11 tumour types; and a test set of 17 poorly differentiated test samples. Each sample was profiled in the space of 217 mammalian mirnas and 16, 000 mrnas Experiment Setup For the mrna expression data, top 40 genes were selected using Relief-F algorithm. For the mirna expression data, top 40 mirna genes were selected using Relief-F. In both cases, 5 nearest neighbors were used in Relief-F for estimating feature weights. For the bagging algorithm, the number of bagging iterations was 10 and the size of each bag was the same as the size of the training data. For the fuzzy k-nn classifiers, we used the Euclidean distance as the distance metric in the experiments, and the best k between 1 and 5 was found by performing leaveone-out cross-validation on the training data. The system was implemented using Perl and Weka Empirical Results Table 1 shows the classification accuracies and P values of our approach and the original approaches used by Luet al. 17. We can observe that our classifier fusion approach was able to classify the same set of poorly differentiated tumor samples at an improved accuracy of 76.5%, as compared to the accuracy of 70.6% obtained using the original approach (a probabilistic neural network algorithm) described in the paper by Lu et al. 17. The bagged fuzzy k-nn classifier using only mirna expression data achieved the same classification accuracy of 70.6% as the original approach. Interestingly, the bagged fuzzy k-nn classifier using only mrna expression data achieved a classification accuracy of 47.1% (P = ), far exceeding the classification accuracy of 5.9% (P = 0.47) obtained by the original approach on the same data. 5. CONCLUSIONS In this paper, a classifier fusion approach is proposed for poorly-differentiated tumor classification using both mrna and mirna expression profiles. Preliminary results on a public data set showed improved classification accuracy compared to that obtained using only mrna or mirna expression profile. We will test the proposed classifier fusion approach on more data sets when they become available. References 1. Ambros, V. The functions of animal micrornas. Nature 431, 7006 (2004), Armstrong, S. A., Staunton, J. E., Silverman, L. B., Pieters, R., den Boer, M. L., Minden,
5 5 M. D., Sallan, S. E., Lander, E. S., Golub, T. R., and Korsmeyer, S. J. MLL translocations specify a distinct gene expression profile that distinguishes a unique leukemia. Nat Genet 30, 1 (2002), Bartel, D. P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 2 (2004), Breiman, L. Bagging predictors. Machine Learning 24, 2 (1996), Croce, C. M., and Calin, G. A. mirnas, cancer, and stem cell division. Cell 122, 1 (2005), Dasarathy, B. Nearest Neighbor Norms: NN Pattern Classification Techniques. IEEE Computer Society Press, Dasarathy, B. Decision Fusion. Computer Society Press, Los Alamitos, California, Esquela-Kerscher, A., and Slack, F. J. Oncomirs micrornas with a role in cancer. Nat Rev Cancer 6, 4 (2006), Golub, T. R., Slonim, D. K., Tamayo, P., Huard, C., Gaasenbeek, M., Mesirov, J. P., Coller, H., Loh, M. L., Downing, J. R., Caligiuri, M. A., Bloomfield, C. D., and Lander, E. S. Molecular classification of cancer: Class discovery and class prediction by gene expression monitoring. Science 286, 5439 (1999), Gunatilaka, A., and Baertlein, B. Feature-level and decision-level fusion of noncoincidently sampled sensors for land mine detection. PAMI 23, 6 (June 2001), Hall, D., and Linas, J. Handbook of Multisensor Data Fusion. CRC Press, Ho, T., Hull, J., and Srihari, S. Decision combination in multiple classifier systems. PAMI 16, 1 (January 1994), Jóźwik, A. A learning scheme for a fuzzy k-nn rule. Pattern Recognition Letters 1 (1983), Keller, J. M., Gray, M. R., and Givens, J. A. A fuzzy k-nearest neighbor algorithm. IEEE Transactions on Systems, Man and Cybernetics 15, 4 (1985), Kononenko, I. Estimating attributes: analysis and extensions of relief. In Proceedings of the European conference on machine learning on Machine Learning (1994), Springer-Verlag New York, Inc., pp Kuncheva, L. I. Combining Pattern Classifiers: Methods and Algorithms. Wiley-Interscience, Lu, J., Getz, G., Miska, E. A., Alvarez- Saavedra, E., Lamb, J., Peck, D., Sweet- Cordero, A., Ebert, B. L., Mak, R. H., Ferrando, A. A., Downing, J. R., Jacks, T., Horvitz, H. R., and Golub, T. R. MicroRNA expression profiles classify human cancers. Nature 435, 7043 (2005), Ramaswamy, S., Tamayo, P., Rifkin, R., Mukherjee, S., Yeang, C. H., Angelo, M., Ladd, C., Reich, M., Latulippe, E., Mesirov, J. P., Poggio, T., Gerald, W., Loda, M., Lander, E. S., and Golub, T. R. Multiclass cancer diagnosis using tumor gene expression signatures. Proc Natl Acad Sci U S A 98, 26 (2001), Robnik-Sikonja, M., and Kononenko, I. Theoretical and empirical analysis of relieff and rrelieff. Mach. Learn. 53, 1-2 (2003), Volinia, S., Calin, G. A., Liu, C. G., Ambs, S., Cimmino, A., Petrocca, F., Visone, R., Iorio, M., Roldo, C., Ferracin, M., Prueitt, R. L., Yanaihara, N., Lanza, G., Scarpa, A., Vecchione, A., Negrini, M., Harris, C. C., and Croce, C. M. A microrna expression signature of human solid tumors defines cancer gene targets. Proc Natl Acad Sci U S A 103, 7 (2006), Wang, Y., and Makedon, F. Application of relieff feature filtering algorithm to selecting informative genes for cancer classification using microarray data. In Proceedings of the 2004 IEEE Computational Systems Bioinformatics Conference (Stanford, California, 2004), pp Wang, Y., Makedon, F. S., Ford, J. C., and Pearlman, J. HyKGene: a hybrid approach for selecting marker genes for phenotype classification using microarray gene expression data. Bioinformatics 21, 8 (2005), Witten, I. H., and Frank, E. Data mining : practical machine learning tools and techniques with Java implementations. Morgan Kaufmann, San Francisco, Calif., Yang, J., Yang, J., Zhang, D., and Lu, J. Feature fusion: parallel strategy vs. serial strategy. Pattern Recognition 36, 6 (June 2003),
Regression Model In The Analysis Of Micro Array Data-Gene Expression Detection
 Jamal Fathima.J.I 1 and P.Venkatesan 1. Research Scholar -Department of statistics National Institute For Research In Tuberculosis, Indian Council For Medical Research,Chennai,India,.Department of statistics
Jamal Fathima.J.I 1 and P.Venkatesan 1. Research Scholar -Department of statistics National Institute For Research In Tuberculosis, Indian Council For Medical Research,Chennai,India,.Department of statistics
Regularized Discriminant Analysis and Its Application in Microarrays
 Biostatistics (2005), 1, 1, pp. 1 18 Printed in Great Britain Regularized Discriminant Analysis and Its Application in Microarrays By YAQIAN GUO Department of Statistics, Stanford University Stanford,
Biostatistics (2005), 1, 1, pp. 1 18 Printed in Great Britain Regularized Discriminant Analysis and Its Application in Microarrays By YAQIAN GUO Department of Statistics, Stanford University Stanford,
application in microarrays
 Biostatistics Advance Access published April 7, 2006 Regularized linear discriminant analysis and its application in microarrays Yaqian Guo, Trevor Hastie and Robert Tibshirani Abstract In this paper,
Biostatistics Advance Access published April 7, 2006 Regularized linear discriminant analysis and its application in microarrays Yaqian Guo, Trevor Hastie and Robert Tibshirani Abstract In this paper,
Feature Selection for SVMs
 Feature Selection for SVMs J. Weston, S. Mukherjee, O. Chapelle, M. Pontil T. Poggio, V. Vapnik, Barnhill BioInformatics.com, Savannah, Georgia, USA. CBCL MIT, Cambridge, Massachusetts, USA. AT&T Research
Feature Selection for SVMs J. Weston, S. Mukherjee, O. Chapelle, M. Pontil T. Poggio, V. Vapnik, Barnhill BioInformatics.com, Savannah, Georgia, USA. CBCL MIT, Cambridge, Massachusetts, USA. AT&T Research
Dimensional reduction of clustered data sets
 Dimensional reduction of clustered data sets Guido Sanguinetti 5th February 2007 Abstract We present a novel probabilistic latent variable model to perform linear dimensional reduction on data sets which
Dimensional reduction of clustered data sets Guido Sanguinetti 5th February 2007 Abstract We present a novel probabilistic latent variable model to perform linear dimensional reduction on data sets which
Data Dependence in Combining Classifiers
 in Combining Classifiers Mohamed Kamel, Nayer Wanas Pattern Analysis and Machine Intelligence Lab University of Waterloo CANADA ! Dependence! Dependence Architecture! Algorithm Outline Pattern Recognition
in Combining Classifiers Mohamed Kamel, Nayer Wanas Pattern Analysis and Machine Intelligence Lab University of Waterloo CANADA ! Dependence! Dependence Architecture! Algorithm Outline Pattern Recognition
Internet Electronic Journal of Molecular Design
 ISSN 1538 6414 Internet Electronic Journal of Molecular Design May 2003, Volume 2, Number 5, Pages 348 357 Editor: Ovidiu Ivanciuc Special issue dedicated to Professor Haruo Hosoya on the occasion of the
ISSN 1538 6414 Internet Electronic Journal of Molecular Design May 2003, Volume 2, Number 5, Pages 348 357 Editor: Ovidiu Ivanciuc Special issue dedicated to Professor Haruo Hosoya on the occasion of the
Gene Expression Data Classification with Revised Kernel Partial Least Squares Algorithm
 Gene Expression Data Classification with Revised Kernel Partial Least Squares Algorithm Zhenqiu Liu, Dechang Chen 2 Department of Computer Science Wayne State University, Market Street, Frederick, MD 273,
Gene Expression Data Classification with Revised Kernel Partial Least Squares Algorithm Zhenqiu Liu, Dechang Chen 2 Department of Computer Science Wayne State University, Market Street, Frederick, MD 273,
hsnim: Hyper Scalable Network Inference Machine for Scale-Free Protein-Protein Interaction Networks Inference
 CS 229 Project Report (TR# MSB2010) Submitted 12/10/2010 hsnim: Hyper Scalable Network Inference Machine for Scale-Free Protein-Protein Interaction Networks Inference Muhammad Shoaib Sehgal Computer Science
CS 229 Project Report (TR# MSB2010) Submitted 12/10/2010 hsnim: Hyper Scalable Network Inference Machine for Scale-Free Protein-Protein Interaction Networks Inference Muhammad Shoaib Sehgal Computer Science
Localized Sliced Inverse Regression
 Localized Sliced Inverse Regression Qiang Wu, Sayan Mukherjee Department of Statistical Science Institute for Genome Sciences & Policy Department of Computer Science Duke University, Durham NC 2778-251,
Localized Sliced Inverse Regression Qiang Wu, Sayan Mukherjee Department of Statistical Science Institute for Genome Sciences & Policy Department of Computer Science Duke University, Durham NC 2778-251,
Module Based Neural Networks for Modeling Gene Regulatory Networks
 Module Based Neural Networks for Modeling Gene Regulatory Networks Paresh Chandra Barman, Std 1 ID: 20044523 Term Project: BiS732 Bio-Network Department of BioSystems, Korea Advanced Institute of Science
Module Based Neural Networks for Modeling Gene Regulatory Networks Paresh Chandra Barman, Std 1 ID: 20044523 Term Project: BiS732 Bio-Network Department of BioSystems, Korea Advanced Institute of Science
A short introduction to supervised learning, with applications to cancer pathway analysis Dr. Christina Leslie
 A short introduction to supervised learning, with applications to cancer pathway analysis Dr. Christina Leslie Computational Biology Program Memorial Sloan-Kettering Cancer Center http://cbio.mskcc.org/leslielab
A short introduction to supervised learning, with applications to cancer pathway analysis Dr. Christina Leslie Computational Biology Program Memorial Sloan-Kettering Cancer Center http://cbio.mskcc.org/leslielab
Improving Fusion of Dimensionality Reduction Methods for Nearest Neighbor Classification
 Improving Fusion of Dimensionality Reduction Methods for Nearest Neighbor Classification Sampath Deegalla and Henrik Boström Department of Computer and Systems Sciences Stockholm University Forum 100,
Improving Fusion of Dimensionality Reduction Methods for Nearest Neighbor Classification Sampath Deegalla and Henrik Boström Department of Computer and Systems Sciences Stockholm University Forum 100,
Comparison of Shannon, Renyi and Tsallis Entropy used in Decision Trees
 Comparison of Shannon, Renyi and Tsallis Entropy used in Decision Trees Tomasz Maszczyk and W lodzis law Duch Department of Informatics, Nicolaus Copernicus University Grudzi adzka 5, 87-100 Toruń, Poland
Comparison of Shannon, Renyi and Tsallis Entropy used in Decision Trees Tomasz Maszczyk and W lodzis law Duch Department of Informatics, Nicolaus Copernicus University Grudzi adzka 5, 87-100 Toruń, Poland
FEATURE SELECTION COMBINED WITH RANDOM SUBSPACE ENSEMBLE FOR GENE EXPRESSION BASED DIAGNOSIS OF MALIGNANCIES
 FEATURE SELECTION COMBINED WITH RANDOM SUBSPACE ENSEMBLE FOR GENE EXPRESSION BASED DIAGNOSIS OF MALIGNANCIES Alberto Bertoni, 1 Raffaella Folgieri, 1 Giorgio Valentini, 1 1 DSI, Dipartimento di Scienze
FEATURE SELECTION COMBINED WITH RANDOM SUBSPACE ENSEMBLE FOR GENE EXPRESSION BASED DIAGNOSIS OF MALIGNANCIES Alberto Bertoni, 1 Raffaella Folgieri, 1 Giorgio Valentini, 1 1 DSI, Dipartimento di Scienze
Identification of Statistically Significant Features from Random Forests
 Identification of Statistically Significant Features from Random Forests Jérôme Paul, Michel Verleysen, and Pierre Dupont Université catholique de Louvain ICTEAM/Machine Learning Group Place Sainte Barbe
Identification of Statistically Significant Features from Random Forests Jérôme Paul, Michel Verleysen, and Pierre Dupont Université catholique de Louvain ICTEAM/Machine Learning Group Place Sainte Barbe
Feature Selection and Classification on Matrix Data: From Large Margins To Small Covering Numbers
 Feature Selection and Classification on Matrix Data: From Large Margins To Small Covering Numbers Sepp Hochreiter and Klaus Obermayer Department of Electrical Engineering and Computer Science Technische
Feature Selection and Classification on Matrix Data: From Large Margins To Small Covering Numbers Sepp Hochreiter and Klaus Obermayer Department of Electrical Engineering and Computer Science Technische
arxiv: v1 [stat.ml] 18 Jan 2010
![arxiv: v1 [stat.ml] 18 Jan 2010 arxiv: v1 [stat.ml] 18 Jan 2010](/thumbs/85/91711997.jpg) Improving stability and interpretability of gene expression signatures arxiv:1001.3109v1 [stat.ml] 18 Jan 2010 Anne-Claire Haury Mines ParisTech, CBIO Institut Curie, Paris, F-75248 INSERM, U900, Paris,
Improving stability and interpretability of gene expression signatures arxiv:1001.3109v1 [stat.ml] 18 Jan 2010 Anne-Claire Haury Mines ParisTech, CBIO Institut Curie, Paris, F-75248 INSERM, U900, Paris,
An Improved 1-norm SVM for Simultaneous Classification and Variable Selection
 An Improved 1-norm SVM for Simultaneous Classification and Variable Selection Hui Zou School of Statistics University of Minnesota Minneapolis, MN 55455 hzou@stat.umn.edu Abstract We propose a novel extension
An Improved 1-norm SVM for Simultaneous Classification and Variable Selection Hui Zou School of Statistics University of Minnesota Minneapolis, MN 55455 hzou@stat.umn.edu Abstract We propose a novel extension
Microarray Data Analysis: Discovery
 Microarray Data Analysis: Discovery Lecture 5 Classification Classification vs. Clustering Classification: Goal: Placing objects (e.g. genes) into meaningful classes Supervised Clustering: Goal: Discover
Microarray Data Analysis: Discovery Lecture 5 Classification Classification vs. Clustering Classification: Goal: Placing objects (e.g. genes) into meaningful classes Supervised Clustering: Goal: Discover
Multiclass Classifiers Based on Dimension Reduction. with Generalized LDA
 Multiclass Classifiers Based on Dimension Reduction with Generalized LDA Hyunsoo Kim a Barry L Drake a Haesun Park a a College of Computing, Georgia Institute of Technology, Atlanta, GA 30332, USA Abstract
Multiclass Classifiers Based on Dimension Reduction with Generalized LDA Hyunsoo Kim a Barry L Drake a Haesun Park a a College of Computing, Georgia Institute of Technology, Atlanta, GA 30332, USA Abstract
Iterative Laplacian Score for Feature Selection
 Iterative Laplacian Score for Feature Selection Linling Zhu, Linsong Miao, and Daoqiang Zhang College of Computer Science and echnology, Nanjing University of Aeronautics and Astronautics, Nanjing 2006,
Iterative Laplacian Score for Feature Selection Linling Zhu, Linsong Miao, and Daoqiang Zhang College of Computer Science and echnology, Nanjing University of Aeronautics and Astronautics, Nanjing 2006,
Data Mining. 3.6 Regression Analysis. Fall Instructor: Dr. Masoud Yaghini. Numeric Prediction
 Data Mining 3.6 Regression Analysis Fall 2008 Instructor: Dr. Masoud Yaghini Outline Introduction Straight-Line Linear Regression Multiple Linear Regression Other Regression Models References Introduction
Data Mining 3.6 Regression Analysis Fall 2008 Instructor: Dr. Masoud Yaghini Outline Introduction Straight-Line Linear Regression Multiple Linear Regression Other Regression Models References Introduction
Proceedings of the Twenty-Seventh AAAI Conference on Artificial Intelligence. Uncorrelated Lasso
 Proceedings of the Twenty-Seventh AAAI Conference on Artificial Intelligence Uncorrelated Lasso Si-Bao Chen School of Computer Science and Technology, Anhui University, Hefei, 23060, China Chris Ding Department
Proceedings of the Twenty-Seventh AAAI Conference on Artificial Intelligence Uncorrelated Lasso Si-Bao Chen School of Computer Science and Technology, Anhui University, Hefei, 23060, China Chris Ding Department
An Empirical Study of Building Compact Ensembles
 An Empirical Study of Building Compact Ensembles Huan Liu, Amit Mandvikar, and Jigar Mody Computer Science & Engineering Arizona State University Tempe, AZ 85281 {huan.liu,amitm,jigar.mody}@asu.edu Abstract.
An Empirical Study of Building Compact Ensembles Huan Liu, Amit Mandvikar, and Jigar Mody Computer Science & Engineering Arizona State University Tempe, AZ 85281 {huan.liu,amitm,jigar.mody}@asu.edu Abstract.
Classification Ensemble That Maximizes the Area Under Receiver Operating Characteristic Curve (AUC)
 Classification Ensemble That Maximizes the Area Under Receiver Operating Characteristic Curve (AUC) Eunsik Park 1 and Y-c Ivan Chang 2 1 Chonnam National University, Gwangju, Korea 2 Academia Sinica, Taipei,
Classification Ensemble That Maximizes the Area Under Receiver Operating Characteristic Curve (AUC) Eunsik Park 1 and Y-c Ivan Chang 2 1 Chonnam National University, Gwangju, Korea 2 Academia Sinica, Taipei,
COMPARING PERFORMANCE OF NEURAL NETWORKS RECOGNIZING MACHINE GENERATED CHARACTERS
 Proceedings of the First Southern Symposium on Computing The University of Southern Mississippi, December 4-5, 1998 COMPARING PERFORMANCE OF NEURAL NETWORKS RECOGNIZING MACHINE GENERATED CHARACTERS SEAN
Proceedings of the First Southern Symposium on Computing The University of Southern Mississippi, December 4-5, 1998 COMPARING PERFORMANCE OF NEURAL NETWORKS RECOGNIZING MACHINE GENERATED CHARACTERS SEAN
Fusion of Dimensionality Reduction Methods: a Case Study in Microarray Classification
 Fusion of Dimensionality Reduction Methods: a Case Study in Microarray Classification Sampath Deegalla Dept. of Computer and Systems Sciences Stockholm University Sweden si-sap@dsv.su.se Henrik Boström
Fusion of Dimensionality Reduction Methods: a Case Study in Microarray Classification Sampath Deegalla Dept. of Computer and Systems Sciences Stockholm University Sweden si-sap@dsv.su.se Henrik Boström
ASYMPTOTIC THEORY FOR DISCRIMINANT ANALYSIS IN HIGH DIMENSION LOW SAMPLE SIZE
 Mem. Gra. Sci. Eng. Shimane Univ. Series B: Mathematics 48 (2015), pp. 15 26 ASYMPTOTIC THEORY FOR DISCRIMINANT ANALYSIS IN HIGH DIMENSION LOW SAMPLE SIZE MITSURU TAMATANI Communicated by Kanta Naito (Received:
Mem. Gra. Sci. Eng. Shimane Univ. Series B: Mathematics 48 (2015), pp. 15 26 ASYMPTOTIC THEORY FOR DISCRIMINANT ANALYSIS IN HIGH DIMENSION LOW SAMPLE SIZE MITSURU TAMATANI Communicated by Kanta Naito (Received:
P leiades: Subspace Clustering and Evaluation
 P leiades: Subspace Clustering and Evaluation Ira Assent, Emmanuel Müller, Ralph Krieger, Timm Jansen, and Thomas Seidl Data management and exploration group, RWTH Aachen University, Germany {assent,mueller,krieger,jansen,seidl}@cs.rwth-aachen.de
P leiades: Subspace Clustering and Evaluation Ira Assent, Emmanuel Müller, Ralph Krieger, Timm Jansen, and Thomas Seidl Data management and exploration group, RWTH Aachen University, Germany {assent,mueller,krieger,jansen,seidl}@cs.rwth-aachen.de
Gene Expression Data Classification With Kernel Principal Component Analysis
 Journal of Biomedicine and Biotechnology 25:2 25 55 59 DOI:.55/JBB.25.55 RESEARCH ARTICLE Gene Expression Data Classification With Kernel Principal Component Analysis Zhenqiu Liu, Dechang Chen, 2 and Halima
Journal of Biomedicine and Biotechnology 25:2 25 55 59 DOI:.55/JBB.25.55 RESEARCH ARTICLE Gene Expression Data Classification With Kernel Principal Component Analysis Zhenqiu Liu, Dechang Chen, 2 and Halima
Feature Selection with Fuzzy Decision Reducts
 Feature Selection with Fuzzy Decision Reducts Chris Cornelis 1, Germán Hurtado Martín 1,2, Richard Jensen 3, and Dominik Ślȩzak4 1 Dept. of Mathematics and Computer Science, Ghent University, Gent, Belgium
Feature Selection with Fuzzy Decision Reducts Chris Cornelis 1, Germán Hurtado Martín 1,2, Richard Jensen 3, and Dominik Ślȩzak4 1 Dept. of Mathematics and Computer Science, Ghent University, Gent, Belgium
MINIMUM EXPECTED RISK PROBABILITY ESTIMATES FOR NONPARAMETRIC NEIGHBORHOOD CLASSIFIERS. Maya Gupta, Luca Cazzanti, and Santosh Srivastava
 MINIMUM EXPECTED RISK PROBABILITY ESTIMATES FOR NONPARAMETRIC NEIGHBORHOOD CLASSIFIERS Maya Gupta, Luca Cazzanti, and Santosh Srivastava University of Washington Dept. of Electrical Engineering Seattle,
MINIMUM EXPECTED RISK PROBABILITY ESTIMATES FOR NONPARAMETRIC NEIGHBORHOOD CLASSIFIERS Maya Gupta, Luca Cazzanti, and Santosh Srivastava University of Washington Dept. of Electrical Engineering Seattle,
Investigating the Performance of a Linear Regression Combiner on Multi-class Data Sets
 Investigating the Performance of a Linear Regression Combiner on Multi-class Data Sets Chun-Xia Zhang 1,2 Robert P.W. Duin 2 1 School of Science and State Key Laboratory for Manufacturing Systems Engineering,
Investigating the Performance of a Linear Regression Combiner on Multi-class Data Sets Chun-Xia Zhang 1,2 Robert P.W. Duin 2 1 School of Science and State Key Laboratory for Manufacturing Systems Engineering,
Constrained Optimization and Support Vector Machines
 Constrained Optimization and Support Vector Machines Man-Wai MAK Dept. of Electronic and Information Engineering, The Hong Kong Polytechnic University enmwmak@polyu.edu.hk http://www.eie.polyu.edu.hk/
Constrained Optimization and Support Vector Machines Man-Wai MAK Dept. of Electronic and Information Engineering, The Hong Kong Polytechnic University enmwmak@polyu.edu.hk http://www.eie.polyu.edu.hk/
CS7267 MACHINE LEARNING
 CS7267 MACHINE LEARNING ENSEMBLE LEARNING Ref: Dr. Ricardo Gutierrez-Osuna at TAMU, and Aarti Singh at CMU Mingon Kang, Ph.D. Computer Science, Kennesaw State University Definition of Ensemble Learning
CS7267 MACHINE LEARNING ENSEMBLE LEARNING Ref: Dr. Ricardo Gutierrez-Osuna at TAMU, and Aarti Singh at CMU Mingon Kang, Ph.D. Computer Science, Kennesaw State University Definition of Ensemble Learning
Linear Classification and SVM. Dr. Xin Zhang
 Linear Classification and SVM Dr. Xin Zhang Email: eexinzhang@scut.edu.cn What is linear classification? Classification is intrinsically non-linear It puts non-identical things in the same class, so a
Linear Classification and SVM Dr. Xin Zhang Email: eexinzhang@scut.edu.cn What is linear classification? Classification is intrinsically non-linear It puts non-identical things in the same class, so a
(134/ !"#!$%&%' ()*)$+,-.)//01*$2*345/0/ M<39$N1$;<)5$")3/B.)O ;).C0*141=5 " /7$83.9$:
 (134/!"#!$%&%' ()*)$+,-)//01*$2*345/0/ "061335/7$839$: ;$9
(134/!"#!$%&%' ()*)$+,-)//01*$2*345/0/ "061335/7$839$: ;$9
Dyadic Classification Trees via Structural Risk Minimization
 Dyadic Classification Trees via Structural Risk Minimization Clayton Scott and Robert Nowak Department of Electrical and Computer Engineering Rice University Houston, TX 77005 cscott,nowak @rice.edu Abstract
Dyadic Classification Trees via Structural Risk Minimization Clayton Scott and Robert Nowak Department of Electrical and Computer Engineering Rice University Houston, TX 77005 cscott,nowak @rice.edu Abstract
10-810: Advanced Algorithms and Models for Computational Biology. Optimal leaf ordering and classification
 10-810: Advanced Algorithms and Models for Computational Biology Optimal leaf ordering and classification Hierarchical clustering As we mentioned, its one of the most popular methods for clustering gene
10-810: Advanced Algorithms and Models for Computational Biology Optimal leaf ordering and classification Hierarchical clustering As we mentioned, its one of the most popular methods for clustering gene
HYPERGRAPH BASED SEMI-SUPERVISED LEARNING ALGORITHMS APPLIED TO SPEECH RECOGNITION PROBLEM: A NOVEL APPROACH
 HYPERGRAPH BASED SEMI-SUPERVISED LEARNING ALGORITHMS APPLIED TO SPEECH RECOGNITION PROBLEM: A NOVEL APPROACH Hoang Trang 1, Tran Hoang Loc 1 1 Ho Chi Minh City University of Technology-VNU HCM, Ho Chi
HYPERGRAPH BASED SEMI-SUPERVISED LEARNING ALGORITHMS APPLIED TO SPEECH RECOGNITION PROBLEM: A NOVEL APPROACH Hoang Trang 1, Tran Hoang Loc 1 1 Ho Chi Minh City University of Technology-VNU HCM, Ho Chi
Introduction to Bioinformatics
 CSCI8980: Applied Machine Learning in Computational Biology Introduction to Bioinformatics Rui Kuang Department of Computer Science and Engineering University of Minnesota kuang@cs.umn.edu History of Bioinformatics
CSCI8980: Applied Machine Learning in Computational Biology Introduction to Bioinformatics Rui Kuang Department of Computer Science and Engineering University of Minnesota kuang@cs.umn.edu History of Bioinformatics
The miss rate for the analysis of gene expression data
 Biostatistics (2005), 6, 1,pp. 111 117 doi: 10.1093/biostatistics/kxh021 The miss rate for the analysis of gene expression data JONATHAN TAYLOR Department of Statistics, Stanford University, Stanford,
Biostatistics (2005), 6, 1,pp. 111 117 doi: 10.1093/biostatistics/kxh021 The miss rate for the analysis of gene expression data JONATHAN TAYLOR Department of Statistics, Stanford University, Stanford,
Model Accuracy Measures
 Model Accuracy Measures Master in Bioinformatics UPF 2017-2018 Eduardo Eyras Computational Genomics Pompeu Fabra University - ICREA Barcelona, Spain Variables What we can measure (attributes) Hypotheses
Model Accuracy Measures Master in Bioinformatics UPF 2017-2018 Eduardo Eyras Computational Genomics Pompeu Fabra University - ICREA Barcelona, Spain Variables What we can measure (attributes) Hypotheses
CLASSIFICATION METHODS FOR MAGIC TELESCOPE IMAGES ON A PIXEL-BY-PIXEL BASE. 1 Introduction
 CLASSIFICATION METHODS FOR MAGIC TELESCOPE IMAGES ON A PIXEL-BY-PIXEL BASE CONSTANTINO MALAGÓN Departamento de Ingeniería Informática, Univ. Antonio de Nebrija, 28040 Madrid, Spain. JUAN ABEL BARRIO 2,
CLASSIFICATION METHODS FOR MAGIC TELESCOPE IMAGES ON A PIXEL-BY-PIXEL BASE CONSTANTINO MALAGÓN Departamento de Ingeniería Informática, Univ. Antonio de Nebrija, 28040 Madrid, Spain. JUAN ABEL BARRIO 2,
Wavelet-Based Feature Extraction for Microarray Data Classification
 26 International Joint Conference on Neural Networks Sheraton Vancouver Wall Centre Hotel, Vancouver, BC, Canada July 16-21, 26 Wavelet-Based Feature Extraction for Microarray Data Classification Shutao
26 International Joint Conference on Neural Networks Sheraton Vancouver Wall Centre Hotel, Vancouver, BC, Canada July 16-21, 26 Wavelet-Based Feature Extraction for Microarray Data Classification Shutao
Sensitivity of the Elucidative Fusion System to the Choice of the Underlying Similarity Metric
 Sensitivity of the Elucidative Fusion System to the Choice of the Underlying Similarity Metric Belur V. Dasarathy Dynetics, Inc., P. O. Box 5500 Huntsville, AL. 35814-5500, USA Belur.d@dynetics.com Abstract
Sensitivity of the Elucidative Fusion System to the Choice of the Underlying Similarity Metric Belur V. Dasarathy Dynetics, Inc., P. O. Box 5500 Huntsville, AL. 35814-5500, USA Belur.d@dynetics.com Abstract
Support Vector Machine & Its Applications
 Support Vector Machine & Its Applications A portion (1/3) of the slides are taken from Prof. Andrew Moore s SVM tutorial at http://www.cs.cmu.edu/~awm/tutorials Mingyue Tan The University of British Columbia
Support Vector Machine & Its Applications A portion (1/3) of the slides are taken from Prof. Andrew Moore s SVM tutorial at http://www.cs.cmu.edu/~awm/tutorials Mingyue Tan The University of British Columbia
Brief Introduction to Machine Learning
 Brief Introduction to Machine Learning Yuh-Jye Lee Lab of Data Science and Machine Intelligence Dept. of Applied Math. at NCTU August 29, 2016 1 / 49 1 Introduction 2 Binary Classification 3 Support Vector
Brief Introduction to Machine Learning Yuh-Jye Lee Lab of Data Science and Machine Intelligence Dept. of Applied Math. at NCTU August 29, 2016 1 / 49 1 Introduction 2 Binary Classification 3 Support Vector
Discriminative Learning can Succeed where Generative Learning Fails
 Discriminative Learning can Succeed where Generative Learning Fails Philip M. Long, a Rocco A. Servedio, b,,1 Hans Ulrich Simon c a Google, Mountain View, CA, USA b Columbia University, New York, New York,
Discriminative Learning can Succeed where Generative Learning Fails Philip M. Long, a Rocco A. Servedio, b,,1 Hans Ulrich Simon c a Google, Mountain View, CA, USA b Columbia University, New York, New York,
Will it rain tomorrow?
 Will it rain tomorrow? Bilal Ahmed - 561539 Department of Computing and Information Systems, The University of Melbourne, Victoria, Australia bahmad@student.unimelb.edu.au Abstract With the availability
Will it rain tomorrow? Bilal Ahmed - 561539 Department of Computing and Information Systems, The University of Melbourne, Victoria, Australia bahmad@student.unimelb.edu.au Abstract With the availability
A Simple Algorithm for Learning Stable Machines
 A Simple Algorithm for Learning Stable Machines Savina Andonova and Andre Elisseeff and Theodoros Evgeniou and Massimiliano ontil Abstract. We present an algorithm for learning stable machines which is
A Simple Algorithm for Learning Stable Machines Savina Andonova and Andre Elisseeff and Theodoros Evgeniou and Massimiliano ontil Abstract. We present an algorithm for learning stable machines which is
A genome sequence based discriminator for vancomycin intermediate Staphyolococcus aureus Supplementary Methods
 1 Journal of Bacteriology Computational Biology Section 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 Supplementary Information for: A genome sequence based discriminator
1 Journal of Bacteriology Computational Biology Section 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 Supplementary Information for: A genome sequence based discriminator
Optimizing Abstaining Classifiers using ROC Analysis. Tadek Pietraszek / 'tʌ dek pɪe 'trʌ ʃek / ICML 2005 August 9, 2005
 IBM Zurich Research Laboratory, GSAL Optimizing Abstaining Classifiers using ROC Analysis Tadek Pietraszek / 'tʌ dek pɪe 'trʌ ʃek / pie@zurich.ibm.com ICML 2005 August 9, 2005 To classify, or not to classify:
IBM Zurich Research Laboratory, GSAL Optimizing Abstaining Classifiers using ROC Analysis Tadek Pietraszek / 'tʌ dek pɪe 'trʌ ʃek / pie@zurich.ibm.com ICML 2005 August 9, 2005 To classify, or not to classify:
Data Mining und Maschinelles Lernen
 Data Mining und Maschinelles Lernen Ensemble Methods Bias-Variance Trade-off Basic Idea of Ensembles Bagging Basic Algorithm Bagging with Costs Randomization Random Forests Boosting Stacking Error-Correcting
Data Mining und Maschinelles Lernen Ensemble Methods Bias-Variance Trade-off Basic Idea of Ensembles Bagging Basic Algorithm Bagging with Costs Randomization Random Forests Boosting Stacking Error-Correcting
A Modified Incremental Principal Component Analysis for On-line Learning of Feature Space and Classifier
 A Modified Incremental Principal Component Analysis for On-line Learning of Feature Space and Classifier Seiichi Ozawa, Shaoning Pang, and Nikola Kasabov Graduate School of Science and Technology, Kobe
A Modified Incremental Principal Component Analysis for On-line Learning of Feature Space and Classifier Seiichi Ozawa, Shaoning Pang, and Nikola Kasabov Graduate School of Science and Technology, Kobe
#33 - Genomics 11/09/07
 BCB 444/544 Required Reading (before lecture) Lecture 33 Mon Nov 5 - Lecture 31 Phylogenetics Parsimony and ML Chp 11 - pp 142 169 Genomics Wed Nov 7 - Lecture 32 Machine Learning Fri Nov 9 - Lecture 33
BCB 444/544 Required Reading (before lecture) Lecture 33 Mon Nov 5 - Lecture 31 Phylogenetics Parsimony and ML Chp 11 - pp 142 169 Genomics Wed Nov 7 - Lecture 32 Machine Learning Fri Nov 9 - Lecture 33
Classification of High Spatial Resolution Remote Sensing Images Based on Decision Fusion
 Journal of Advances in Information Technology Vol. 8, No. 1, February 2017 Classification of High Spatial Resolution Remote Sensing Images Based on Decision Fusion Guizhou Wang Institute of Remote Sensing
Journal of Advances in Information Technology Vol. 8, No. 1, February 2017 Classification of High Spatial Resolution Remote Sensing Images Based on Decision Fusion Guizhou Wang Institute of Remote Sensing
Non-Negative Factorization for Clustering of Microarray Data
 INT J COMPUT COMMUN, ISSN 1841-9836 9(1):16-23, February, 2014. Non-Negative Factorization for Clustering of Microarray Data L. Morgos Lucian Morgos Dept. of Electronics and Telecommunications Faculty
INT J COMPUT COMMUN, ISSN 1841-9836 9(1):16-23, February, 2014. Non-Negative Factorization for Clustering of Microarray Data L. Morgos Lucian Morgos Dept. of Electronics and Telecommunications Faculty
Effect of Rule Weights in Fuzzy Rule-Based Classification Systems
 506 IEEE TRANSACTIONS ON FUZZY SYSTEMS, VOL. 9, NO. 4, AUGUST 2001 Effect of Rule Weights in Fuzzy Rule-Based Classification Systems Hisao Ishibuchi, Member, IEEE, and Tomoharu Nakashima, Member, IEEE
506 IEEE TRANSACTIONS ON FUZZY SYSTEMS, VOL. 9, NO. 4, AUGUST 2001 Effect of Rule Weights in Fuzzy Rule-Based Classification Systems Hisao Ishibuchi, Member, IEEE, and Tomoharu Nakashima, Member, IEEE
Principles of Pattern Recognition. C. A. Murthy Machine Intelligence Unit Indian Statistical Institute Kolkata
 Principles of Pattern Recognition C. A. Murthy Machine Intelligence Unit Indian Statistical Institute Kolkata e-mail: murthy@isical.ac.in Pattern Recognition Measurement Space > Feature Space >Decision
Principles of Pattern Recognition C. A. Murthy Machine Intelligence Unit Indian Statistical Institute Kolkata e-mail: murthy@isical.ac.in Pattern Recognition Measurement Space > Feature Space >Decision
Visualize Biological Database for Protein in Homosapiens Using Classification Searching Models
 International Journal of Computational Intelligence Research ISSN 0973-1873 Volume 13, Number 2 (2017), pp. 213-224 Research India Publications http://www.ripublication.com Visualize Biological Database
International Journal of Computational Intelligence Research ISSN 0973-1873 Volume 13, Number 2 (2017), pp. 213-224 Research India Publications http://www.ripublication.com Visualize Biological Database
Clustering and Network
 Clustering and Network Jing-Dong Jackie Han jdhan@picb.ac.cn http://www.picb.ac.cn/~jdhan Copy Right: Jing-Dong Jackie Han What is clustering? A way of grouping together data samples that are similar in
Clustering and Network Jing-Dong Jackie Han jdhan@picb.ac.cn http://www.picb.ac.cn/~jdhan Copy Right: Jing-Dong Jackie Han What is clustering? A way of grouping together data samples that are similar in
Regularized Linear Models in Stacked Generalization
 Regularized Linear Models in Stacked Generalization Sam Reid and Greg Grudic University of Colorado at Boulder, Boulder CO 80309-0430, USA Abstract Stacked generalization is a flexible method for multiple
Regularized Linear Models in Stacked Generalization Sam Reid and Greg Grudic University of Colorado at Boulder, Boulder CO 80309-0430, USA Abstract Stacked generalization is a flexible method for multiple
Multi-Metric and Multi-Substructure Biclustering Analysis for Gene Expression Data
 Multi-Metric and Multi-Substructure Biclustering Analysis for Gene Expression Data S.Y. Kung Princeton University kung@princeton.edu Man-Wai Mak The Hong Kong Polytechnic University enmwmak@polyu.edu.hk
Multi-Metric and Multi-Substructure Biclustering Analysis for Gene Expression Data S.Y. Kung Princeton University kung@princeton.edu Man-Wai Mak The Hong Kong Polytechnic University enmwmak@polyu.edu.hk
Single gene analysis of differential expression
 Single gene analysis of differential expression Giorgio Valentini DSI Dipartimento di Scienze dell Informazione Università degli Studi di Milano valentini@dsi.unimi.it Comparing two conditions Each condition
Single gene analysis of differential expression Giorgio Valentini DSI Dipartimento di Scienze dell Informazione Università degli Studi di Milano valentini@dsi.unimi.it Comparing two conditions Each condition
AN ENHANCED INITIALIZATION METHOD FOR NON-NEGATIVE MATRIX FACTORIZATION. Liyun Gong 1, Asoke K. Nandi 2,3 L69 3BX, UK; 3PH, UK;
 213 IEEE INERNAIONAL WORKSHOP ON MACHINE LEARNING FOR SIGNAL PROCESSING, SEP. 22 25, 213, SOUHAMPON, UK AN ENHANCED INIIALIZAION MEHOD FOR NON-NEGAIVE MARIX FACORIZAION Liyun Gong 1, Asoke K. Nandi 2,3
213 IEEE INERNAIONAL WORKSHOP ON MACHINE LEARNING FOR SIGNAL PROCESSING, SEP. 22 25, 213, SOUHAMPON, UK AN ENHANCED INIIALIZAION MEHOD FOR NON-NEGAIVE MARIX FACORIZAION Liyun Gong 1, Asoke K. Nandi 2,3
A Bias Correction for the Minimum Error Rate in Cross-validation
 A Bias Correction for the Minimum Error Rate in Cross-validation Ryan J. Tibshirani Robert Tibshirani Abstract Tuning parameters in supervised learning problems are often estimated by cross-validation.
A Bias Correction for the Minimum Error Rate in Cross-validation Ryan J. Tibshirani Robert Tibshirani Abstract Tuning parameters in supervised learning problems are often estimated by cross-validation.
Penalized versus constrained generalized eigenvalue problems
 Penalized versus constrained generalized eigenvalue problems Irina Gaynanova, James G. Booth and Martin T. Wells. arxiv:141.6131v3 [stat.co] 4 May 215 Abstract We investigate the difference between using
Penalized versus constrained generalized eigenvalue problems Irina Gaynanova, James G. Booth and Martin T. Wells. arxiv:141.6131v3 [stat.co] 4 May 215 Abstract We investigate the difference between using
MATHEMATICS OF DATA FUSION
 MATHEMATICS OF DATA FUSION by I. R. GOODMAN NCCOSC RDTE DTV, San Diego, California, U.S.A. RONALD P. S. MAHLER Lockheed Martin Tactical Defences Systems, Saint Paul, Minnesota, U.S.A. and HUNG T. NGUYEN
MATHEMATICS OF DATA FUSION by I. R. GOODMAN NCCOSC RDTE DTV, San Diego, California, U.S.A. RONALD P. S. MAHLER Lockheed Martin Tactical Defences Systems, Saint Paul, Minnesota, U.S.A. and HUNG T. NGUYEN
A Modified Incremental Principal Component Analysis for On-Line Learning of Feature Space and Classifier
 A Modified Incremental Principal Component Analysis for On-Line Learning of Feature Space and Classifier Seiichi Ozawa 1, Shaoning Pang 2, and Nikola Kasabov 2 1 Graduate School of Science and Technology,
A Modified Incremental Principal Component Analysis for On-Line Learning of Feature Space and Classifier Seiichi Ozawa 1, Shaoning Pang 2, and Nikola Kasabov 2 1 Graduate School of Science and Technology,
Department of Computer Science, University of Waikato, Hamilton, New Zealand. 2
 Predicting Apple Bruising using Machine Learning G.Holmes 1, S.J.Cunningham 1, B.T. Dela Rue 2 and A.F. Bollen 2 1 Department of Computer Science, University of Waikato, Hamilton, New Zealand. 2 Lincoln
Predicting Apple Bruising using Machine Learning G.Holmes 1, S.J.Cunningham 1, B.T. Dela Rue 2 and A.F. Bollen 2 1 Department of Computer Science, University of Waikato, Hamilton, New Zealand. 2 Lincoln
Enhancing Generalization Capability of SVM Classifiers with Feature Weight Adjustment
 Enhancing Generalization Capability of SVM Classifiers ith Feature Weight Adjustment Xizhao Wang and Qiang He College of Mathematics and Computer Science, Hebei University, Baoding 07002, Hebei, China
Enhancing Generalization Capability of SVM Classifiers ith Feature Weight Adjustment Xizhao Wang and Qiang He College of Mathematics and Computer Science, Hebei University, Baoding 07002, Hebei, China
Intro. ANN & Fuzzy Systems. Lecture 15. Pattern Classification (I): Statistical Formulation
 Lecture 15. Pattern Classification (I): Statistical Formulation Outline Statistical Pattern Recognition Maximum Posterior Probability (MAP) Classifier Maximum Likelihood (ML) Classifier K-Nearest Neighbor
Lecture 15. Pattern Classification (I): Statistical Formulation Outline Statistical Pattern Recognition Maximum Posterior Probability (MAP) Classifier Maximum Likelihood (ML) Classifier K-Nearest Neighbor
A Clustering Coefficient for Weighted Networks, with Application to Gene Expression Data
 A Clustering Coefficient for Weighted Networks, with Application to Gene Expression Data Gabriela Kalna a, and Desmond J. Higham a a Department of Mathematics, University of Strathclyde, Glasgow G XH,
A Clustering Coefficient for Weighted Networks, with Application to Gene Expression Data Gabriela Kalna a, and Desmond J. Higham a a Department of Mathematics, University of Strathclyde, Glasgow G XH,
Variable Selection and Weighting by Nearest Neighbor Ensembles
 Variable Selection and Weighting by Nearest Neighbor Ensembles Jan Gertheiss (joint work with Gerhard Tutz) Department of Statistics University of Munich WNI 2008 Nearest Neighbor Methods Introduction
Variable Selection and Weighting by Nearest Neighbor Ensembles Jan Gertheiss (joint work with Gerhard Tutz) Department of Statistics University of Munich WNI 2008 Nearest Neighbor Methods Introduction
Unsupervised Classification via Convex Absolute Value Inequalities
 Unsupervised Classification via Convex Absolute Value Inequalities Olvi L. Mangasarian Abstract We consider the problem of classifying completely unlabeled data by using convex inequalities that contain
Unsupervised Classification via Convex Absolute Value Inequalities Olvi L. Mangasarian Abstract We consider the problem of classifying completely unlabeled data by using convex inequalities that contain
KERNEL LOGISTIC REGRESSION-LINEAR FOR LEUKEMIA CLASSIFICATION USING HIGH DIMENSIONAL DATA
 Rahayu, Kernel Logistic Regression-Linear for Leukemia Classification using High Dimensional Data KERNEL LOGISTIC REGRESSION-LINEAR FOR LEUKEMIA CLASSIFICATION USING HIGH DIMENSIONAL DATA S.P. Rahayu 1,2
Rahayu, Kernel Logistic Regression-Linear for Leukemia Classification using High Dimensional Data KERNEL LOGISTIC REGRESSION-LINEAR FOR LEUKEMIA CLASSIFICATION USING HIGH DIMENSIONAL DATA S.P. Rahayu 1,2
Classification using stochastic ensembles
 July 31, 2014 Topics Introduction Topics Classification Application and classfication Classification and Regression Trees Stochastic ensemble methods Our application: USAID Poverty Assessment Tools Topics
July 31, 2014 Topics Introduction Topics Classification Application and classfication Classification and Regression Trees Stochastic ensemble methods Our application: USAID Poverty Assessment Tools Topics
Selection of Classifiers based on Multiple Classifier Behaviour
 Selection of Classifiers based on Multiple Classifier Behaviour Giorgio Giacinto, Fabio Roli, and Giorgio Fumera Dept. of Electrical and Electronic Eng. - University of Cagliari Piazza d Armi, 09123 Cagliari,
Selection of Classifiers based on Multiple Classifier Behaviour Giorgio Giacinto, Fabio Roli, and Giorgio Fumera Dept. of Electrical and Electronic Eng. - University of Cagliari Piazza d Armi, 09123 Cagliari,
Introduction to clustering methods for gene expression data analysis
 Introduction to clustering methods for gene expression data analysis Giorgio Valentini e-mail: valentini@dsi.unimi.it Outline Levels of analysis of DNA microarray data Clustering methods for functional
Introduction to clustering methods for gene expression data analysis Giorgio Valentini e-mail: valentini@dsi.unimi.it Outline Levels of analysis of DNA microarray data Clustering methods for functional
Online Estimation of Discrete Densities using Classifier Chains
 Online Estimation of Discrete Densities using Classifier Chains Michael Geilke 1 and Eibe Frank 2 and Stefan Kramer 1 1 Johannes Gutenberg-Universtität Mainz, Germany {geilke,kramer}@informatik.uni-mainz.de
Online Estimation of Discrete Densities using Classifier Chains Michael Geilke 1 and Eibe Frank 2 and Stefan Kramer 1 1 Johannes Gutenberg-Universtität Mainz, Germany {geilke,kramer}@informatik.uni-mainz.de
Neural Networks and Ensemble Methods for Classification
 Neural Networks and Ensemble Methods for Classification NEURAL NETWORKS 2 Neural Networks A neural network is a set of connected input/output units (neurons) where each connection has a weight associated
Neural Networks and Ensemble Methods for Classification NEURAL NETWORKS 2 Neural Networks A neural network is a set of connected input/output units (neurons) where each connection has a weight associated
Click Prediction and Preference Ranking of RSS Feeds
 Click Prediction and Preference Ranking of RSS Feeds 1 Introduction December 11, 2009 Steven Wu RSS (Really Simple Syndication) is a family of data formats used to publish frequently updated works. RSS
Click Prediction and Preference Ranking of RSS Feeds 1 Introduction December 11, 2009 Steven Wu RSS (Really Simple Syndication) is a family of data formats used to publish frequently updated works. RSS
Classifier Selection. Nicholas Ver Hoeve Craig Martek Ben Gardner
 Classifier Selection Nicholas Ver Hoeve Craig Martek Ben Gardner Classifier Ensembles Assume we have an ensemble of classifiers with a well-chosen feature set. We want to optimize the competence of this
Classifier Selection Nicholas Ver Hoeve Craig Martek Ben Gardner Classifier Ensembles Assume we have an ensemble of classifiers with a well-chosen feature set. We want to optimize the competence of this
Computational Systems Biology
 Computational Systems Biology Vasant Honavar Artificial Intelligence Research Laboratory Bioinformatics and Computational Biology Graduate Program Center for Computational Intelligence, Learning, & Discovery
Computational Systems Biology Vasant Honavar Artificial Intelligence Research Laboratory Bioinformatics and Computational Biology Graduate Program Center for Computational Intelligence, Learning, & Discovery
International Journal "Information Theories & Applications" Vol.14 /
 International Journal "Information Theories & Applications" Vol.4 / 2007 87 or 2) Nˆ t N. That criterion and parameters F, M, N assign method of constructing sample decision function. In order to estimate
International Journal "Information Theories & Applications" Vol.4 / 2007 87 or 2) Nˆ t N. That criterion and parameters F, M, N assign method of constructing sample decision function. In order to estimate
Support Vector Machine. Industrial AI Lab.
 Support Vector Machine Industrial AI Lab. Classification (Linear) Autonomously figure out which category (or class) an unknown item should be categorized into Number of categories / classes Binary: 2 different
Support Vector Machine Industrial AI Lab. Classification (Linear) Autonomously figure out which category (or class) an unknown item should be categorized into Number of categories / classes Binary: 2 different
Machine Learning Ensemble Learning I Hamid R. Rabiee Jafar Muhammadi, Alireza Ghasemi Spring /
 Machine Learning Ensemble Learning I Hamid R. Rabiee Jafar Muhammadi, Alireza Ghasemi Spring 2015 http://ce.sharif.edu/courses/93-94/2/ce717-1 / Agenda Combining Classifiers Empirical view Theoretical
Machine Learning Ensemble Learning I Hamid R. Rabiee Jafar Muhammadi, Alireza Ghasemi Spring 2015 http://ce.sharif.edu/courses/93-94/2/ce717-1 / Agenda Combining Classifiers Empirical view Theoretical
Machine Learning Basics
 Security and Fairness of Deep Learning Machine Learning Basics Anupam Datta CMU Spring 2019 Image Classification Image Classification Image classification pipeline Input: A training set of N images, each
Security and Fairness of Deep Learning Machine Learning Basics Anupam Datta CMU Spring 2019 Image Classification Image Classification Image classification pipeline Input: A training set of N images, each
Introduction to Support Vector Machines
 Introduction to Support Vector Machines Hsuan-Tien Lin Learning Systems Group, California Institute of Technology Talk in NTU EE/CS Speech Lab, November 16, 2005 H.-T. Lin (Learning Systems Group) Introduction
Introduction to Support Vector Machines Hsuan-Tien Lin Learning Systems Group, California Institute of Technology Talk in NTU EE/CS Speech Lab, November 16, 2005 H.-T. Lin (Learning Systems Group) Introduction
Mathematics, Genomics, and Cancer
 School of Informatics IUB April 6, 2009 Outline Introduction Class Comparison Class Discovery Class Prediction Example Biological states and state modulation Software Tools Research directions Math & Biology
School of Informatics IUB April 6, 2009 Outline Introduction Class Comparison Class Discovery Class Prediction Example Biological states and state modulation Software Tools Research directions Math & Biology
CS 229 Final report A Study Of Ensemble Methods In Machine Learning
 A Study Of Ensemble Methods In Machine Learning Abstract The idea of ensemble methodology is to build a predictive model by integrating multiple models. It is well-known that ensemble methods can be used
A Study Of Ensemble Methods In Machine Learning Abstract The idea of ensemble methodology is to build a predictive model by integrating multiple models. It is well-known that ensemble methods can be used
Modifying A Linear Support Vector Machine for Microarray Data Classification
 Modifying A Linear Support Vector Machine for Microarray Data Classification Prof. Rosemary A. Renaut Dr. Hongbin Guo & Wang Juh Chen Department of Mathematics and Statistics, Arizona State University
Modifying A Linear Support Vector Machine for Microarray Data Classification Prof. Rosemary A. Renaut Dr. Hongbin Guo & Wang Juh Chen Department of Mathematics and Statistics, Arizona State University
MACHINE LEARNING. Support Vector Machines. Alessandro Moschitti
 MACHINE LEARNING Support Vector Machines Alessandro Moschitti Department of information and communication technology University of Trento Email: moschitti@dit.unitn.it Summary Support Vector Machines
MACHINE LEARNING Support Vector Machines Alessandro Moschitti Department of information and communication technology University of Trento Email: moschitti@dit.unitn.it Summary Support Vector Machines
Introduction to clustering methods for gene expression data analysis
 Introduction to clustering methods for gene expression data analysis Giorgio Valentini e-mail: valentini@dsi.unimi.it Outline Levels of analysis of DNA microarray data Clustering methods for functional
Introduction to clustering methods for gene expression data analysis Giorgio Valentini e-mail: valentini@dsi.unimi.it Outline Levels of analysis of DNA microarray data Clustering methods for functional
DATA MINING WITH DIFFERENT TYPES OF X-RAY DATA
 DATA MINING WITH DIFFERENT TYPES OF X-RAY DATA 315 C. K. Lowe-Ma, A. E. Chen, D. Scholl Physical & Environmental Sciences, Research and Advanced Engineering Ford Motor Company, Dearborn, Michigan, USA
DATA MINING WITH DIFFERENT TYPES OF X-RAY DATA 315 C. K. Lowe-Ma, A. E. Chen, D. Scholl Physical & Environmental Sciences, Research and Advanced Engineering Ford Motor Company, Dearborn, Michigan, USA
THE DOUBLY REGULARIZED SUPPORT VECTOR MACHINE
 Statistica Sinica 16(006), 589-615 THE DOUBLY REGULARIZED SUPPORT VECTOR MACHINE Li Wang, Ji Zhu and Hui Zou University of Michigan and University of Minnesota Abstract: The standard -norm support vector
Statistica Sinica 16(006), 589-615 THE DOUBLY REGULARIZED SUPPORT VECTOR MACHINE Li Wang, Ji Zhu and Hui Zou University of Michigan and University of Minnesota Abstract: The standard -norm support vector
An experimental study of the intrinsic stability of random forest variable importance measures
 Wang et al. BMC Bioinformatics (2016) 17:60 DOI 10.1186/s12859-016-0900-5 RESEARCH ARTICLE Open Access An experimental study of the intrinsic stability of random forest variable importance measures Huazhen
Wang et al. BMC Bioinformatics (2016) 17:60 DOI 10.1186/s12859-016-0900-5 RESEARCH ARTICLE Open Access An experimental study of the intrinsic stability of random forest variable importance measures Huazhen
False discovery control for multiple tests of association under general dependence
 False discovery control for multiple tests of association under general dependence Nicolai Meinshausen Seminar für Statistik ETH Zürich December 2, 2004 Abstract We propose a confidence envelope for false
False discovery control for multiple tests of association under general dependence Nicolai Meinshausen Seminar für Statistik ETH Zürich December 2, 2004 Abstract We propose a confidence envelope for false
