Hierachical Name Entity Recognition
|
|
|
- Logan Adams
- 5 years ago
- Views:
Transcription
1 Hierachical Name Entity Recognition Dakan Wang, Yu Wu Mentor: David Mcclosky, Mihai Surdeanu March 10, Introduction In this project, we investigte the hierarchical name entity recognition problem implement three modesl to empirically verify that it is probable to utilize the hierarchical relationship between entity types to improve the tranditional NER task. Specifically, our three models are all non-trivial extensions of the classical MEMM classifier. We believe some of the ideas can be conveniently adapted to sequence classifiers such as conditional random fields. 2 MEMM and Hierachical NER 2.1 Definition Assuming that we have a bunch of entities X = {x (i) } Nx i=1, and a collection of entity types C = {c i } Nc i=1. An name-entity recognition task is to identify entities in the text and map them to their belonged classes. One commonly used model to do this is the Maximum Entropy Markov Model, which can be specified as follows, exp( j P λ (c x, c ) = λ jf j (x, c, c)) c C exp( j λ jf j (x, c, c )) In the above formulation, c is the class for the previous word of x. In training, this information is available for training the MEMM. However, in testing, such information is hidden and a verterbi decoding algorithm is usually used to get the most probable sequence of entity classes. MEMM is a very useful technique used in various applications concerning sequence-based classification. Though there are some more principled sequence model such as conditional random fields, in practice, MEMM is comparable with those advanced sequenced models. In traditional NER, each word is assumed to be belonged to one entity class and entity classes are always by default assumed to be independent. However, in real world applications, even entity classes can be dependent. One such dependence is called nested entity relationship[2], in which one entity class is allowed to include several other classes. For instance, in the word sequence Bank Of America, bank and America are both 1 (1)
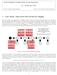
2 Figure 1: Ontology Tree Figure 2: Tree Normalization 2
3 assumed to belong to some entity classes, and the whole world Bank of America also belongs to some entity class. Such implicit relationships about entity classes are very interesting and may help improve the performance of the NER tasks. In this report, we consider another entity class relaitonship and propose large scale hierachical NER. Unlike traditional NER which only involve 3 or 4 entity classes, we considerthe domain which includes many entity classes. Biomedicine is just one such domain which has more than ten entity types. These entities form the leaves of a hierarchical tree structure and the dependency distance of two entity types can be measured by their distance in the tree. One typical example of this is shown in Figure 1. It will be interesting to see if the given tree structure can help the traditional NER taks which ignore the tree structure and take the leaves in the tree to be independent of each other. Formally, suppose that we have a hierarchical tree structure T, and each node in the tree has a label indicating the node s corresponding class. The labels of the leaves of T forms a set C = {c i } Nc i=1, which is just consisted of the entity classes in our NER task. For a node u in the tree. we use A(u) to denote the set of u s ancestors in the tree. Also, let A(c) denote the ancestor nodes of node u whose label is c. Furthermore, we use D(T, d) to denote all the nodes at depth d in the tree, and the root of the tree is considered to be at depth 0. The concept of tree normalization refers to converting the tree such that all leaves in the tree are at the same depth. For instance, if in the original tree, one leaf u 1 is at depth 2 and the other one u 2 is at depth 5, then tree normalization attaches a path of length 2 to leaf u 1. We show an example of tree normalization in Figure 2. With these concepts and notations, we now describe our three proposed models. 2.2 model 0 Model 0 is our baseline model, which does not exploit any tree stucture and only takes the labels of the leaves in the tree to be entity types and does a traditional NER task. 2.3 model 1 The first model is the simplest model but also the most intuitive one. The basic idea is just to use the information in the hierarchical tree to enrich features and expect to improve the model performance. We modify the MEMM formulation a bit as follows. exp( j P λ (c x, c ) = λ jf j (x, A(c ), c)) c C exp( j λ jf j (x, A(c ), c )) Notice that here instead of only being only a function of the label of the previous label c, the feature function is now a function of a path from the leaf with label c to the root. We take the tree in to be an example of this idea. Suppose that we have now an word whose previous word belongs to entity class DNA. In traditional NER, when we extract features, we will on do the following, features.add( PREVLABEL-DNA ) However, in our model1, we will instead do the following (2) 3
4 Figure 3: Weight Construction for model 2 features.add( PREVLABEL-DNA ) features.add( PREVLABEL-Nulceid Acid ) features.add( PREVLABEL- Is Entity ) In prediction, the viterbi decoding algorithm is a dynamic programming on the paths from root to leaves since we need all ancesters of a leaf to make the prediction. 2.4 Model 2 In this model, the basic idea is to train a classifier at each depth and combine all classifier s prediction result to make the final prediction. Suppose the tree T is already normalized, and at the d-th level, the set of entity classes is C d and the model parameters are λ d, the prediction of the d-th level can be written to be P λ d(c d x, c d exp( j ) = λ jf j (x, c d, c d )) c d C exp( d j λ jf j (x, c d (3), c d )) An illustration for the weight vector for each node is shown in Figure 3. Since we want to combine the prediction from classifier s result, we proposed two approaches. The first approach is a heuristic which is shown not to very useful in the experiments. But as an approach we have tried in our experiments, we also report it here. 2.5 Model 2.1 The idea here is to ignore the sequence and let the classifier vote for the entity type for a word. Since the label of the previous word is one of the features for the classifier, the probability of a word s label is conditional on the previous label. Therefore when calculating the type of a word, we need to eliminate the conditional variable and do something like belief prorogation, that is P (c x t,.., x 0 ) = P (c x t, c )P (c x t 1,.., x 0 ) (4) c C 4
5 The above equation can be easily calculate by going through all words in the sequence. Recall that we are in the hierarchical setting, and for each depth d, we can calculate P (c d x t,.., x 0 ). Then for each label path from the root to a leaf p = (c 1,.., c d,..), we calculate i c P (c i x t,.., x 0 ). Then we take path with the largest probability and the leaf of the path to be the entity of x t. 2.6 Model 2.2 In model 2.2, we extend the viterbi algorithm as follows. Suppose that V t 1,p is the maximum probability of sequence in which the (t-1)-th word in the sentence corresponds to path p. Here p is a path from the root to a leaf, and we say a word corresponds to a path p when the word s label was the leaf corresponding to the path. V t,p = max{p (p x t, p )V t 1,p } (5) In the above function, x t is the word at position t, p is the possible label path for the previous word and P (p x t, p ) = u p P (u x t, u ) (6) Here u is a node on the path, and P (u x t, u ) can be calculated from each independently trained classifier. We give an example on the extended Viterbi algorithm, suppose that in the tree in Figure 2, we want to calculate the maximum probability of the sequence in which x t belongs to label RNA. The label path corresponding to the label rna is is p = (is entity,nucleic acid compound, rna). To calculate the maximum probability, we need to enumerate all the label paths of the previous word, P = {(O, O, O), (is entity, protein, protein) (is entity, nucleic acid compound, rna), (is entity, nucleic acid compound, dna)} (7) For p = (is entity, nucleic acid compound, rna), P (p x t, p ) = P (is entity x t, is entity) P (nucleic acid compound x t, nucleic acid compound) P (rna x t, dna) (8) And here P (is entity x t, is entity), P (nucleic acid compound x t, nucleic acid compound), and P (rna x t, dna) are calculated from classifiers for nodes in depth 2, depth 3, and depth 4 respectively. 2.7 Model 3 All the above mentioned models rely on some sort of heuristics. We finally describe a model which can principally exploits the tree structure. The trick here is that we 5
6 Figure 4: Weight Construction for Model 3 construct the weight vector using the tree structure. For convenience of defining the model, we use a different formalization of MEMM: the logistic regression model. In LR model, we denote f(x) to be the feature vector for instance x, and W c to be the weight vector for class c. And the probability of x belonging to class c can be written to be P (c x) = exp(w c x) c exp(w c x) It is easy to see the equivalence of the above formulation and the MEMM formulation. Recall that in the Hierarchical NER setting, each entity class should belong to a leaf in the entity type tree. We now construct the weight vectors {W c } from the taxonomy. Assume that every node u in the tree is associated with a local weight vector w u, and the global weight vector W u is defined to be W root = w root W u = w u + W pa(u) Here pa(u) means the parent of node u. Writing the above definition in another way, we have W u = w u + w v (10) v A(u) Recall that A(u) is the ancesters of node u. An illustration of this weight vector construction is shown in Figure 4. Intuitively, the global weight of u is the sum of the local weights on the path from the root to the node u. Thus if we come back to the hierarchical NER problem in which each entity type corresponds to a leaf, we can rewrite the logistic regression formulation to be (9) P (c x) = exp( u A(c) wu x) c exp( u A(c ) wu x) (11) Our task here is to estimate all the local weights w c in the entity type tree. 6
7 We give some analysis of how the above formulation utilizes the tree structure. The basic intuition comes from the observation that if two leaves u 1 and u 2 (correponding to two entity types) has a shares common ancester v, then their global weight vectors W u 1 and W u 2 share a local weight vector w v. The closer u 1 and u 2 are in the tree, the more local weight vectors they share. In our example tree, the global weight vectors of rna and dna have a common part which is W u where u is the node corresponding to nucleic acid compound in our example tree. In the learning process, whenever the model receives instances belonging to either dna or ran, it will update all the local weight vectors from the root to the node corresponding to nucleic acid compound. This can be seen more clearly if we calculate the derivative with respect to w u when the model receives a set of instances {(x (i), c (i) )} N i=1. Notice that the error function for logistic regression can be written to be E = = N log(p (c (i) x i )) i=1 N log exp( u A(c (i) ) wu x (i) ) i=1 c exp( u A(c ) wu x (i) ) Let C v be the set of leaves of the subtree rooted at node v, the derivative of E with respect to a node v in the entity type tree can be written as follows E c w v = x (i) C v x(i) exp u A(c ) wu x (i) c (i) C v c exp( u A(c ) wu x (i) ) c = x (i) x (i) C v exp u A(c ) wu c (i) C v c exp( u A(c ) wu x (i) ) = x (i) (1 p(c x i )) c (i) Cv c C v The above formula can be interpreted as follows: The derivative of w v will be large only if the classifier cannot classify the instance to the leaves of the tree rooted in v. On the other hand, if the current classifier can classifier the instance into any of the leaves of the subtree rooted at v, then that instance will not contribute to the gradient of w v. This observation is also consistent with our intention of constructing the weight vectors: w v can be seen to be a sub-classifier which helps determine whether an instance should be classified into the subtree rooted at v. Since all instances with labels corresponding to leaves of that subtree can be seen to belong to entity type v, we should not differenetiate between these instances when we are leaning w v. To elabroate the above model, we further propose two extensions, 7
8 2.8 Model 3.1 We formulate the error function as follows N E = log(p (c (i) x i )) + λ d w u 2 (12) i=1 0<d depth(t ) u D(T,d) Recall that we have stated in the definition part that D(T,d) is set of all nodes at depth d of the tree T. Here λ d1 λ d2 for d 1 d 2. We briefly explain the intuition behind this. Basically, a node at a larger depth in the tree will have less training instances and will be more likely have the problem of overfitting. Therefore it is necessary to add a larger regularization term to weight vectors correponding to nodes at a larger depth to avoid overfitting. Another interesting way of thinking about the learned weight vectors can be expected to be consistent over the tree. Here consistency means that for two nodes u 1 and u 2 at depth d in the tree, if W u 1 f(x) > W u 2 f(x), then for any two nodes v 1 and v 2, where v 1 and v 2 are respectively the nodes in the subtrees rooted at u 1 and u 2, it satisfies W v 1 f(x) > W v 2 f(x). This can be roughly obtained (but no exact math proof!) as follows. Let p(u 1, v 1 ) be the path from one of u 1 s child to v 1, then we can write W v 1 f(x) = W u 1 f(x)+ u p(u 1,v 1 ) wu f(x). Since u is at a deeper position in the tree than u, we can think that (again, it s just high level, not maths proof!) w u W u 1. Thus it can be expected that W u 1 f(x) W u 2 f(x) > u p(u 1,v 1 ) wu f(x)+ u p(u 2,v 2 ) wu f(x), indicating the consistency. 2.9 Model 3.2 We do not implement this model, but it should be an interesting extension. We write error function to be N E = log(p (c (i) x i )) + λ d w u 1 (13) i=1 0<d depth(t ) u D(T,d) Here λ d1 λ d2 for d 1 d 2.. Notice that we only change the l 2 regularization to l 1 regularization. Our intention of proposing the above function is to enable to model to do feature selection. We would like to select the entity classes shared features and also their own specific features. Since the regularization term at a shallower node has a larger regularization term, we can expect the learned feature vector will be sparser, and will select those common features in the subtree rooted at the corresponding node. And the leaf nodes will be the least sparse, and it will enable the model to discover entity class specific features Features Used By the Model We extracted several categories of features, which we summarize below current word 8
9 previous word next word previous word+current word current word+next word previous word shape current word shape next word shape if the word contains a non-letter character(such as hyphen) if the word contains a capital letter substring of a word. The range of the substring length is [2,6] For wordshape feature, we just replace all letter characters with x, all digit characters with d, and all other weird characters with?. All the other feature templates are quite clear. Also notice that the features for model 1 should further include all the ancesters of the previous word s label. 3 Experimental Results The dataset we use is the genia corpus, which is an xml file. NER on this dataset has also been studied by [1]. In this dataset, each word can be associated with an entity, which can be mapped to a leaf in the ontology tree as in Figure 1. To simplify the task and have a quick result of whether hierarchical NER can improve the performance, we simpliy the originla ontology tree and use a very simple tree structure as defined in Figure 2. There are three entity types, DNA, protein, and RNA. The models used are model0, model1, model2.2, and model3.1. We have also implemented and evaluated model 2.1, but it does not work very well on the dataset, and it is even worse than the baseline. Thus we do not report its results here. From the table, we can see that the hierarchical structure always helps improve the performance. Model 1 and Model 2.2 is comparable with each other. Our last model 3.1 achieves the best performance and is consistently better than all the other model in the F1-performance. We have also compared the overall accuracy of the four models in Figure 5. From the figure we can also see that model3.1 achieves the best accuracy. 4 Conclusion and future work Ww in this report have built four models which can exploit the hierarchical entity type structure. Out of the four models, three of them have already been empirically proven 9
10 precision recall FB1 overall 67.73% 69.54% 68.62% DNA 51.26% 49.93% 50.58% protein 73.18% 76.62% 74.86% RNA 43.98% 39.85% 41.81% Table 1: baseline precision recall FB1 overall 72.40% 68.89% 70.60% DNA 59.74% 48.89% 53.77% protein 76.18% 76.24% 76.21% RNA 48.68% 34.59% 40.44% Table 2: model1 precision recall FB1 overall 72.12% 69.21% 70.64% DNA 58.47% 48.96% 53.30% protein 76.28% 76.60% 76.44% RNA 47.52% 36.09% 41.03% Table 3: model2.2 precision recall FB1 overall 72.25% 70.07% 71.14% DNA 59.38% 50.11% 54.35% protein 76.07% 77.29% 76.68% RNA 52.24% 39.47% 44.97% Table 4: model3.1 to be useful in improving the performance. Further work includes implmenting model 3.2 to see whehter the model can do automatic feature selection. It is also interesting to investigate whether the proposed ideas in this work can be adapted to more advanced sequence classifiers such as CRF. 5 Acknowledgement The authors would like to thank David and Mihai for insightful discussions. 10
11 Figure 5: Accuracy Comparison References [1] Jenny Finkel, Shipra Dingare, Christopher D. Manning, Malvina Nissim, Beatrice Alex, and Claire Grover. Exploring the boundaries: Gene and protein identification in biomedical text. In In Proceedings of the BioCreative Workshop, [2] Jenny Rose Finkel and Christopher D. Manning. Nested named entity recognition. In Proceedings of the 2009 Conference on Empirical Methods in Natural Language Processing: Volume 1 - Volume 1, EMNLP 09, pages , Stroudsburg, PA, USA, Association for Computational Linguistics. 11
Information Extraction from Text
 Information Extraction from Text Jing Jiang Chapter 2 from Mining Text Data (2012) Presented by Andrew Landgraf, September 13, 2013 1 What is Information Extraction? Goal is to discover structured information
Information Extraction from Text Jing Jiang Chapter 2 from Mining Text Data (2012) Presented by Andrew Landgraf, September 13, 2013 1 What is Information Extraction? Goal is to discover structured information
CLRG Biocreative V
 CLRG ChemTMiner @ Biocreative V Sobha Lalitha Devi., Sindhuja Gopalan., Vijay Sundar Ram R., Malarkodi C.S., Lakshmi S., Pattabhi RK Rao Computational Linguistics Research Group, AU-KBC Research Centre
CLRG ChemTMiner @ Biocreative V Sobha Lalitha Devi., Sindhuja Gopalan., Vijay Sundar Ram R., Malarkodi C.S., Lakshmi S., Pattabhi RK Rao Computational Linguistics Research Group, AU-KBC Research Centre
Lecture 13: Discriminative Sequence Models (MEMM and Struct. Perceptron)
 Lecture 13: Discriminative Sequence Models (MEMM and Struct. Perceptron) Intro to NLP, CS585, Fall 2014 http://people.cs.umass.edu/~brenocon/inlp2014/ Brendan O Connor (http://brenocon.com) 1 Models for
Lecture 13: Discriminative Sequence Models (MEMM and Struct. Perceptron) Intro to NLP, CS585, Fall 2014 http://people.cs.umass.edu/~brenocon/inlp2014/ Brendan O Connor (http://brenocon.com) 1 Models for
Statistical NLP for the Web Log Linear Models, MEMM, Conditional Random Fields
 Statistical NLP for the Web Log Linear Models, MEMM, Conditional Random Fields Sameer Maskey Week 13, Nov 28, 2012 1 Announcements Next lecture is the last lecture Wrap up of the semester 2 Final Project
Statistical NLP for the Web Log Linear Models, MEMM, Conditional Random Fields Sameer Maskey Week 13, Nov 28, 2012 1 Announcements Next lecture is the last lecture Wrap up of the semester 2 Final Project
Lecture 13: Structured Prediction
 Lecture 13: Structured Prediction Kai-Wei Chang CS @ University of Virginia kw@kwchang.net Couse webpage: http://kwchang.net/teaching/nlp16 CS6501: NLP 1 Quiz 2 v Lectures 9-13 v Lecture 12: before page
Lecture 13: Structured Prediction Kai-Wei Chang CS @ University of Virginia kw@kwchang.net Couse webpage: http://kwchang.net/teaching/nlp16 CS6501: NLP 1 Quiz 2 v Lectures 9-13 v Lecture 12: before page
Probabilistic Graphical Models
 Probabilistic Graphical Models David Sontag New York University Lecture 4, February 16, 2012 David Sontag (NYU) Graphical Models Lecture 4, February 16, 2012 1 / 27 Undirected graphical models Reminder
Probabilistic Graphical Models David Sontag New York University Lecture 4, February 16, 2012 David Sontag (NYU) Graphical Models Lecture 4, February 16, 2012 1 / 27 Undirected graphical models Reminder
Conditional Random Fields and beyond DANIEL KHASHABI CS 546 UIUC, 2013
 Conditional Random Fields and beyond DANIEL KHASHABI CS 546 UIUC, 2013 Outline Modeling Inference Training Applications Outline Modeling Problem definition Discriminative vs. Generative Chain CRF General
Conditional Random Fields and beyond DANIEL KHASHABI CS 546 UIUC, 2013 Outline Modeling Inference Training Applications Outline Modeling Problem definition Discriminative vs. Generative Chain CRF General
ToxiCat: Hybrid Named Entity Recognition services to support curation of the Comparative Toxicogenomic Database
 ToxiCat: Hybrid Named Entity Recognition services to support curation of the Comparative Toxicogenomic Database Dina Vishnyakova 1,2, 4, *, Julien Gobeill 1,3,4, Emilie Pasche 1,2,3,4 and Patrick Ruch
ToxiCat: Hybrid Named Entity Recognition services to support curation of the Comparative Toxicogenomic Database Dina Vishnyakova 1,2, 4, *, Julien Gobeill 1,3,4, Emilie Pasche 1,2,3,4 and Patrick Ruch
Generalization Error on Pruning Decision Trees
 Generalization Error on Pruning Decision Trees Ryan R. Rosario Computer Science 269 Fall 2010 A decision tree is a predictive model that can be used for either classification or regression [3]. Decision
Generalization Error on Pruning Decision Trees Ryan R. Rosario Computer Science 269 Fall 2010 A decision tree is a predictive model that can be used for either classification or regression [3]. Decision
CSCI-567: Machine Learning (Spring 2019)
 CSCI-567: Machine Learning (Spring 2019) Prof. Victor Adamchik U of Southern California Mar. 19, 2019 March 19, 2019 1 / 43 Administration March 19, 2019 2 / 43 Administration TA3 is due this week March
CSCI-567: Machine Learning (Spring 2019) Prof. Victor Adamchik U of Southern California Mar. 19, 2019 March 19, 2019 1 / 43 Administration March 19, 2019 2 / 43 Administration TA3 is due this week March
Statistical Machine Learning Theory. From Multi-class Classification to Structured Output Prediction. Hisashi Kashima.
 http://goo.gl/jv7vj9 Course website KYOTO UNIVERSITY Statistical Machine Learning Theory From Multi-class Classification to Structured Output Prediction Hisashi Kashima kashima@i.kyoto-u.ac.jp DEPARTMENT
http://goo.gl/jv7vj9 Course website KYOTO UNIVERSITY Statistical Machine Learning Theory From Multi-class Classification to Structured Output Prediction Hisashi Kashima kashima@i.kyoto-u.ac.jp DEPARTMENT
Graphical models for part of speech tagging
 Indian Institute of Technology, Bombay and Research Division, India Research Lab Graphical models for part of speech tagging Different Models for POS tagging HMM Maximum Entropy Markov Models Conditional
Indian Institute of Technology, Bombay and Research Division, India Research Lab Graphical models for part of speech tagging Different Models for POS tagging HMM Maximum Entropy Markov Models Conditional
Statistical Methods for NLP
 Statistical Methods for NLP Sequence Models Joakim Nivre Uppsala University Department of Linguistics and Philology joakim.nivre@lingfil.uu.se Statistical Methods for NLP 1(21) Introduction Structured
Statistical Methods for NLP Sequence Models Joakim Nivre Uppsala University Department of Linguistics and Philology joakim.nivre@lingfil.uu.se Statistical Methods for NLP 1(21) Introduction Structured
Intelligent Systems (AI-2)
 Intelligent Systems (AI-2) Computer Science cpsc422, Lecture 19 Oct, 23, 2015 Slide Sources Raymond J. Mooney University of Texas at Austin D. Koller, Stanford CS - Probabilistic Graphical Models D. Page,
Intelligent Systems (AI-2) Computer Science cpsc422, Lecture 19 Oct, 23, 2015 Slide Sources Raymond J. Mooney University of Texas at Austin D. Koller, Stanford CS - Probabilistic Graphical Models D. Page,
Statistical Machine Learning Theory. From Multi-class Classification to Structured Output Prediction. Hisashi Kashima.
 http://goo.gl/xilnmn Course website KYOTO UNIVERSITY Statistical Machine Learning Theory From Multi-class Classification to Structured Output Prediction Hisashi Kashima kashima@i.kyoto-u.ac.jp DEPARTMENT
http://goo.gl/xilnmn Course website KYOTO UNIVERSITY Statistical Machine Learning Theory From Multi-class Classification to Structured Output Prediction Hisashi Kashima kashima@i.kyoto-u.ac.jp DEPARTMENT
Sequence Modelling with Features: Linear-Chain Conditional Random Fields. COMP-599 Oct 6, 2015
 Sequence Modelling with Features: Linear-Chain Conditional Random Fields COMP-599 Oct 6, 2015 Announcement A2 is out. Due Oct 20 at 1pm. 2 Outline Hidden Markov models: shortcomings Generative vs. discriminative
Sequence Modelling with Features: Linear-Chain Conditional Random Fields COMP-599 Oct 6, 2015 Announcement A2 is out. Due Oct 20 at 1pm. 2 Outline Hidden Markov models: shortcomings Generative vs. discriminative
Part of Speech Tagging: Viterbi, Forward, Backward, Forward- Backward, Baum-Welch. COMP-599 Oct 1, 2015
 Part of Speech Tagging: Viterbi, Forward, Backward, Forward- Backward, Baum-Welch COMP-599 Oct 1, 2015 Announcements Research skills workshop today 3pm-4:30pm Schulich Library room 313 Start thinking about
Part of Speech Tagging: Viterbi, Forward, Backward, Forward- Backward, Baum-Welch COMP-599 Oct 1, 2015 Announcements Research skills workshop today 3pm-4:30pm Schulich Library room 313 Start thinking about
Learning theory. Ensemble methods. Boosting. Boosting: history
 Learning theory Probability distribution P over X {0, 1}; let (X, Y ) P. We get S := {(x i, y i )} n i=1, an iid sample from P. Ensemble methods Goal: Fix ɛ, δ (0, 1). With probability at least 1 δ (over
Learning theory Probability distribution P over X {0, 1}; let (X, Y ) P. We get S := {(x i, y i )} n i=1, an iid sample from P. Ensemble methods Goal: Fix ɛ, δ (0, 1). With probability at least 1 δ (over
Supervised Learning. George Konidaris
 Supervised Learning George Konidaris gdk@cs.brown.edu Fall 2017 Machine Learning Subfield of AI concerned with learning from data. Broadly, using: Experience To Improve Performance On Some Task (Tom Mitchell,
Supervised Learning George Konidaris gdk@cs.brown.edu Fall 2017 Machine Learning Subfield of AI concerned with learning from data. Broadly, using: Experience To Improve Performance On Some Task (Tom Mitchell,
Machine Learning for natural language processing
 Machine Learning for natural language processing Classification: Maximum Entropy Models Laura Kallmeyer Heinrich-Heine-Universität Düsseldorf Summer 2016 1 / 24 Introduction Classification = supervised
Machine Learning for natural language processing Classification: Maximum Entropy Models Laura Kallmeyer Heinrich-Heine-Universität Düsseldorf Summer 2016 1 / 24 Introduction Classification = supervised
UNIVERSITY of PENNSYLVANIA CIS 520: Machine Learning Final, Fall 2013
 UNIVERSITY of PENNSYLVANIA CIS 520: Machine Learning Final, Fall 2013 Exam policy: This exam allows two one-page, two-sided cheat sheets; No other materials. Time: 2 hours. Be sure to write your name and
UNIVERSITY of PENNSYLVANIA CIS 520: Machine Learning Final, Fall 2013 Exam policy: This exam allows two one-page, two-sided cheat sheets; No other materials. Time: 2 hours. Be sure to write your name and
A Decision Stump. Decision Trees, cont. Boosting. Machine Learning 10701/15781 Carlos Guestrin Carnegie Mellon University. October 1 st, 2007
 Decision Trees, cont. Boosting Machine Learning 10701/15781 Carlos Guestrin Carnegie Mellon University October 1 st, 2007 1 A Decision Stump 2 1 The final tree 3 Basic Decision Tree Building Summarized
Decision Trees, cont. Boosting Machine Learning 10701/15781 Carlos Guestrin Carnegie Mellon University October 1 st, 2007 1 A Decision Stump 2 1 The final tree 3 Basic Decision Tree Building Summarized
Intelligent Systems (AI-2)
 Intelligent Systems (AI-2) Computer Science cpsc422, Lecture 19 Oct, 24, 2016 Slide Sources Raymond J. Mooney University of Texas at Austin D. Koller, Stanford CS - Probabilistic Graphical Models D. Page,
Intelligent Systems (AI-2) Computer Science cpsc422, Lecture 19 Oct, 24, 2016 Slide Sources Raymond J. Mooney University of Texas at Austin D. Koller, Stanford CS - Probabilistic Graphical Models D. Page,
Natural Language Processing Prof. Pawan Goyal Department of Computer Science and Engineering Indian Institute of Technology, Kharagpur
 Natural Language Processing Prof. Pawan Goyal Department of Computer Science and Engineering Indian Institute of Technology, Kharagpur Lecture - 18 Maximum Entropy Models I Welcome back for the 3rd module
Natural Language Processing Prof. Pawan Goyal Department of Computer Science and Engineering Indian Institute of Technology, Kharagpur Lecture - 18 Maximum Entropy Models I Welcome back for the 3rd module
A brief introduction to Conditional Random Fields
 A brief introduction to Conditional Random Fields Mark Johnson Macquarie University April, 2005, updated October 2010 1 Talk outline Graphical models Maximum likelihood and maximum conditional likelihood
A brief introduction to Conditional Random Fields Mark Johnson Macquarie University April, 2005, updated October 2010 1 Talk outline Graphical models Maximum likelihood and maximum conditional likelihood
Class 4: Classification. Quaid Morris February 11 th, 2011 ML4Bio
 Class 4: Classification Quaid Morris February 11 th, 211 ML4Bio Overview Basic concepts in classification: overfitting, cross-validation, evaluation. Linear Discriminant Analysis and Quadratic Discriminant
Class 4: Classification Quaid Morris February 11 th, 211 ML4Bio Overview Basic concepts in classification: overfitting, cross-validation, evaluation. Linear Discriminant Analysis and Quadratic Discriminant
Week 5: Logistic Regression & Neural Networks
 Week 5: Logistic Regression & Neural Networks Instructor: Sergey Levine 1 Summary: Logistic Regression In the previous lecture, we covered logistic regression. To recap, logistic regression models and
Week 5: Logistic Regression & Neural Networks Instructor: Sergey Levine 1 Summary: Logistic Regression In the previous lecture, we covered logistic regression. To recap, logistic regression models and
Symbolic methods in TC: Decision Trees
 Symbolic methods in TC: Decision Trees ML for NLP Lecturer: Kevin Koidl Assist. Lecturer Alfredo Maldonado https://www.cs.tcd.ie/kevin.koidl/cs0/ kevin.koidl@scss.tcd.ie, maldonaa@tcd.ie 01-017 A symbolic
Symbolic methods in TC: Decision Trees ML for NLP Lecturer: Kevin Koidl Assist. Lecturer Alfredo Maldonado https://www.cs.tcd.ie/kevin.koidl/cs0/ kevin.koidl@scss.tcd.ie, maldonaa@tcd.ie 01-017 A symbolic
Undirected Graphical Models
 Outline Hong Chang Institute of Computing Technology, Chinese Academy of Sciences Machine Learning Methods (Fall 2012) Outline Outline I 1 Introduction 2 Properties Properties 3 Generative vs. Conditional
Outline Hong Chang Institute of Computing Technology, Chinese Academy of Sciences Machine Learning Methods (Fall 2012) Outline Outline I 1 Introduction 2 Properties Properties 3 Generative vs. Conditional
From Binary to Multiclass Classification. CS 6961: Structured Prediction Spring 2018
 From Binary to Multiclass Classification CS 6961: Structured Prediction Spring 2018 1 So far: Binary Classification We have seen linear models Learning algorithms Perceptron SVM Logistic Regression Prediction
From Binary to Multiclass Classification CS 6961: Structured Prediction Spring 2018 1 So far: Binary Classification We have seen linear models Learning algorithms Perceptron SVM Logistic Regression Prediction
Support Vector Machine & Its Applications
 Support Vector Machine & Its Applications A portion (1/3) of the slides are taken from Prof. Andrew Moore s SVM tutorial at http://www.cs.cmu.edu/~awm/tutorials Mingyue Tan The University of British Columbia
Support Vector Machine & Its Applications A portion (1/3) of the slides are taken from Prof. Andrew Moore s SVM tutorial at http://www.cs.cmu.edu/~awm/tutorials Mingyue Tan The University of British Columbia
Randomized Decision Trees
 Randomized Decision Trees compiled by Alvin Wan from Professor Jitendra Malik s lecture Discrete Variables First, let us consider some terminology. We have primarily been dealing with real-valued data,
Randomized Decision Trees compiled by Alvin Wan from Professor Jitendra Malik s lecture Discrete Variables First, let us consider some terminology. We have primarily been dealing with real-valued data,
CS 446 Machine Learning Fall 2016 Nov 01, Bayesian Learning
 CS 446 Machine Learning Fall 206 Nov 0, 206 Bayesian Learning Professor: Dan Roth Scribe: Ben Zhou, C. Cervantes Overview Bayesian Learning Naive Bayes Logistic Regression Bayesian Learning So far, we
CS 446 Machine Learning Fall 206 Nov 0, 206 Bayesian Learning Professor: Dan Roth Scribe: Ben Zhou, C. Cervantes Overview Bayesian Learning Naive Bayes Logistic Regression Bayesian Learning So far, we
Click Prediction and Preference Ranking of RSS Feeds
 Click Prediction and Preference Ranking of RSS Feeds 1 Introduction December 11, 2009 Steven Wu RSS (Really Simple Syndication) is a family of data formats used to publish frequently updated works. RSS
Click Prediction and Preference Ranking of RSS Feeds 1 Introduction December 11, 2009 Steven Wu RSS (Really Simple Syndication) is a family of data formats used to publish frequently updated works. RSS
Maxent Models and Discriminative Estimation
 Maxent Models and Discriminative Estimation Generative vs. Discriminative models (Reading: J+M Ch6) Introduction So far we ve looked at generative models Language models, Naive Bayes But there is now much
Maxent Models and Discriminative Estimation Generative vs. Discriminative models (Reading: J+M Ch6) Introduction So far we ve looked at generative models Language models, Naive Bayes But there is now much
Qualifying Exam in Machine Learning
 Qualifying Exam in Machine Learning October 20, 2009 Instructions: Answer two out of the three questions in Part 1. In addition, answer two out of three questions in two additional parts (choose two parts
Qualifying Exam in Machine Learning October 20, 2009 Instructions: Answer two out of the three questions in Part 1. In addition, answer two out of three questions in two additional parts (choose two parts
Introduction to Machine Learning Midterm, Tues April 8
 Introduction to Machine Learning 10-701 Midterm, Tues April 8 [1 point] Name: Andrew ID: Instructions: You are allowed a (two-sided) sheet of notes. Exam ends at 2:45pm Take a deep breath and don t spend
Introduction to Machine Learning 10-701 Midterm, Tues April 8 [1 point] Name: Andrew ID: Instructions: You are allowed a (two-sided) sheet of notes. Exam ends at 2:45pm Take a deep breath and don t spend
CS145: INTRODUCTION TO DATA MINING
 CS145: INTRODUCTION TO DATA MINING 4: Vector Data: Decision Tree Instructor: Yizhou Sun yzsun@cs.ucla.edu October 10, 2017 Methods to Learn Vector Data Set Data Sequence Data Text Data Classification Clustering
CS145: INTRODUCTION TO DATA MINING 4: Vector Data: Decision Tree Instructor: Yizhou Sun yzsun@cs.ucla.edu October 10, 2017 Methods to Learn Vector Data Set Data Sequence Data Text Data Classification Clustering
Sequences and Information
 Sequences and Information Rahul Siddharthan The Institute of Mathematical Sciences, Chennai, India http://www.imsc.res.in/ rsidd/ Facets 16, 04/07/2016 This box says something By looking at the symbols
Sequences and Information Rahul Siddharthan The Institute of Mathematical Sciences, Chennai, India http://www.imsc.res.in/ rsidd/ Facets 16, 04/07/2016 This box says something By looking at the symbols
Learning Decision Trees
 Learning Decision Trees Machine Learning Spring 2018 1 This lecture: Learning Decision Trees 1. Representation: What are decision trees? 2. Algorithm: Learning decision trees The ID3 algorithm: A greedy
Learning Decision Trees Machine Learning Spring 2018 1 This lecture: Learning Decision Trees 1. Representation: What are decision trees? 2. Algorithm: Learning decision trees The ID3 algorithm: A greedy
Learning Decision Trees
 Learning Decision Trees Machine Learning Fall 2018 Some slides from Tom Mitchell, Dan Roth and others 1 Key issues in machine learning Modeling How to formulate your problem as a machine learning problem?
Learning Decision Trees Machine Learning Fall 2018 Some slides from Tom Mitchell, Dan Roth and others 1 Key issues in machine learning Modeling How to formulate your problem as a machine learning problem?
Naïve Bayes, Maxent and Neural Models
 Naïve Bayes, Maxent and Neural Models CMSC 473/673 UMBC Some slides adapted from 3SLP Outline Recap: classification (MAP vs. noisy channel) & evaluation Naïve Bayes (NB) classification Terminology: bag-of-words
Naïve Bayes, Maxent and Neural Models CMSC 473/673 UMBC Some slides adapted from 3SLP Outline Recap: classification (MAP vs. noisy channel) & evaluation Naïve Bayes (NB) classification Terminology: bag-of-words
Log-Linear Models, MEMMs, and CRFs
 Log-Linear Models, MEMMs, and CRFs Michael Collins 1 Notation Throughout this note I ll use underline to denote vectors. For example, w R d will be a vector with components w 1, w 2,... w d. We use expx
Log-Linear Models, MEMMs, and CRFs Michael Collins 1 Notation Throughout this note I ll use underline to denote vectors. For example, w R d will be a vector with components w 1, w 2,... w d. We use expx
Discovering molecular pathways from protein interaction and ge
 Discovering molecular pathways from protein interaction and gene expression data 9-4-2008 Aim To have a mechanism for inferring pathways from gene expression and protein interaction data. Motivation Why
Discovering molecular pathways from protein interaction and gene expression data 9-4-2008 Aim To have a mechanism for inferring pathways from gene expression and protein interaction data. Motivation Why
Decision trees. Special Course in Computer and Information Science II. Adam Gyenge Helsinki University of Technology
 Decision trees Special Course in Computer and Information Science II Adam Gyenge Helsinki University of Technology 6.2.2008 Introduction Outline: Definition of decision trees ID3 Pruning methods Bibliography:
Decision trees Special Course in Computer and Information Science II Adam Gyenge Helsinki University of Technology 6.2.2008 Introduction Outline: Definition of decision trees ID3 Pruning methods Bibliography:
Probabilistic Graphical Models: MRFs and CRFs. CSE628: Natural Language Processing Guest Lecturer: Veselin Stoyanov
 Probabilistic Graphical Models: MRFs and CRFs CSE628: Natural Language Processing Guest Lecturer: Veselin Stoyanov Why PGMs? PGMs can model joint probabilities of many events. many techniques commonly
Probabilistic Graphical Models: MRFs and CRFs CSE628: Natural Language Processing Guest Lecturer: Veselin Stoyanov Why PGMs? PGMs can model joint probabilities of many events. many techniques commonly
Text Mining. March 3, March 3, / 49
 Text Mining March 3, 2017 March 3, 2017 1 / 49 Outline Language Identification Tokenisation Part-Of-Speech (POS) tagging Hidden Markov Models - Sequential Taggers Viterbi Algorithm March 3, 2017 2 / 49
Text Mining March 3, 2017 March 3, 2017 1 / 49 Outline Language Identification Tokenisation Part-Of-Speech (POS) tagging Hidden Markov Models - Sequential Taggers Viterbi Algorithm March 3, 2017 2 / 49
Learning Decision Trees
 Learning Decision Trees CS194-10 Fall 2011 Lecture 8 CS194-10 Fall 2011 Lecture 8 1 Outline Decision tree models Tree construction Tree pruning Continuous input features CS194-10 Fall 2011 Lecture 8 2
Learning Decision Trees CS194-10 Fall 2011 Lecture 8 CS194-10 Fall 2011 Lecture 8 1 Outline Decision tree models Tree construction Tree pruning Continuous input features CS194-10 Fall 2011 Lecture 8 2
Boosting: Foundations and Algorithms. Rob Schapire
 Boosting: Foundations and Algorithms Rob Schapire Example: Spam Filtering problem: filter out spam (junk email) gather large collection of examples of spam and non-spam: From: yoav@ucsd.edu Rob, can you
Boosting: Foundations and Algorithms Rob Schapire Example: Spam Filtering problem: filter out spam (junk email) gather large collection of examples of spam and non-spam: From: yoav@ucsd.edu Rob, can you
The Perceptron. Volker Tresp Summer 2014
 The Perceptron Volker Tresp Summer 2014 1 Introduction One of the first serious learning machines Most important elements in learning tasks Collection and preprocessing of training data Definition of a
The Perceptron Volker Tresp Summer 2014 1 Introduction One of the first serious learning machines Most important elements in learning tasks Collection and preprocessing of training data Definition of a
Bayesian Hierarchical Classification. Seminar on Predicting Structured Data Jukka Kohonen
 Bayesian Hierarchical Classification Seminar on Predicting Structured Data Jukka Kohonen 17.4.2008 Overview Intro: The task of hierarchical gene annotation Approach I: SVM/Bayes hybrid Barutcuoglu et al:
Bayesian Hierarchical Classification Seminar on Predicting Structured Data Jukka Kohonen 17.4.2008 Overview Intro: The task of hierarchical gene annotation Approach I: SVM/Bayes hybrid Barutcuoglu et al:
Structure Learning in Sequential Data
 Structure Learning in Sequential Data Liam Stewart liam@cs.toronto.edu Richard Zemel zemel@cs.toronto.edu 2005.09.19 Motivation. Cau, R. Kuiper, and W.-P. de Roever. Formalising Dijkstra's development
Structure Learning in Sequential Data Liam Stewart liam@cs.toronto.edu Richard Zemel zemel@cs.toronto.edu 2005.09.19 Motivation. Cau, R. Kuiper, and W.-P. de Roever. Formalising Dijkstra's development
Lecture 6: Backpropagation
 Lecture 6: Backpropagation Roger Grosse 1 Introduction So far, we ve seen how to train shallow models, where the predictions are computed as a linear function of the inputs. We ve also observed that deeper
Lecture 6: Backpropagation Roger Grosse 1 Introduction So far, we ve seen how to train shallow models, where the predictions are computed as a linear function of the inputs. We ve also observed that deeper
10-701/ Machine Learning, Fall
 0-70/5-78 Machine Learning, Fall 2003 Homework 2 Solution If you have questions, please contact Jiayong Zhang .. (Error Function) The sum-of-squares error is the most common training
0-70/5-78 Machine Learning, Fall 2003 Homework 2 Solution If you have questions, please contact Jiayong Zhang .. (Error Function) The sum-of-squares error is the most common training
Machine Learning for NLP
 Machine Learning for NLP Linear Models Joakim Nivre Uppsala University Department of Linguistics and Philology Slides adapted from Ryan McDonald, Google Research Machine Learning for NLP 1(26) Outline
Machine Learning for NLP Linear Models Joakim Nivre Uppsala University Department of Linguistics and Philology Slides adapted from Ryan McDonald, Google Research Machine Learning for NLP 1(26) Outline
Notes on Noise Contrastive Estimation (NCE)
 Notes on Noise Contrastive Estimation NCE) David Meyer dmm@{-4-5.net,uoregon.edu,...} March 0, 207 Introduction In this note we follow the notation used in [2]. Suppose X x, x 2,, x Td ) is a sample of
Notes on Noise Contrastive Estimation NCE) David Meyer dmm@{-4-5.net,uoregon.edu,...} March 0, 207 Introduction In this note we follow the notation used in [2]. Suppose X x, x 2,, x Td ) is a sample of
Deep Feedforward Networks. Han Shao, Hou Pong Chan, and Hongyi Zhang
 Deep Feedforward Networks Han Shao, Hou Pong Chan, and Hongyi Zhang Deep Feedforward Networks Goal: approximate some function f e.g., a classifier, maps input to a class y = f (x) x y Defines a mapping
Deep Feedforward Networks Han Shao, Hou Pong Chan, and Hongyi Zhang Deep Feedforward Networks Goal: approximate some function f e.g., a classifier, maps input to a class y = f (x) x y Defines a mapping
Input-Output Hidden Markov Models for Trees
 Input-Output Hidden Markov Models for Trees Davide Bacciu 1 and Alessio Micheli 1 and Alessandro Sperduti 2 1- Dipartimento di Informatica - Università di Pisa- Italy 2- Dipartimento di Matematica Pura
Input-Output Hidden Markov Models for Trees Davide Bacciu 1 and Alessio Micheli 1 and Alessandro Sperduti 2 1- Dipartimento di Informatica - Università di Pisa- Italy 2- Dipartimento di Matematica Pura
word2vec Parameter Learning Explained
 word2vec Parameter Learning Explained Xin Rong ronxin@umich.edu Abstract The word2vec model and application by Mikolov et al. have attracted a great amount of attention in recent two years. The vector
word2vec Parameter Learning Explained Xin Rong ronxin@umich.edu Abstract The word2vec model and application by Mikolov et al. have attracted a great amount of attention in recent two years. The vector
The Perceptron. Volker Tresp Summer 2016
 The Perceptron Volker Tresp Summer 2016 1 Elements in Learning Tasks Collection, cleaning and preprocessing of training data Definition of a class of learning models. Often defined by the free model parameters
The Perceptron Volker Tresp Summer 2016 1 Elements in Learning Tasks Collection, cleaning and preprocessing of training data Definition of a class of learning models. Often defined by the free model parameters
lecture 6: modeling sequences (final part)
 Natural Language Processing 1 lecture 6: modeling sequences (final part) Ivan Titov Institute for Logic, Language and Computation Outline After a recap: } Few more words about unsupervised estimation of
Natural Language Processing 1 lecture 6: modeling sequences (final part) Ivan Titov Institute for Logic, Language and Computation Outline After a recap: } Few more words about unsupervised estimation of
Nonlinear Classification
 Nonlinear Classification INFO-4604, Applied Machine Learning University of Colorado Boulder October 5-10, 2017 Prof. Michael Paul Linear Classification Most classifiers we ve seen use linear functions
Nonlinear Classification INFO-4604, Applied Machine Learning University of Colorado Boulder October 5-10, 2017 Prof. Michael Paul Linear Classification Most classifiers we ve seen use linear functions
STA 414/2104: Machine Learning
 STA 414/2104: Machine Learning Russ Salakhutdinov Department of Computer Science! Department of Statistics! rsalakhu@cs.toronto.edu! http://www.cs.toronto.edu/~rsalakhu/ Lecture 9 Sequential Data So far
STA 414/2104: Machine Learning Russ Salakhutdinov Department of Computer Science! Department of Statistics! rsalakhu@cs.toronto.edu! http://www.cs.toronto.edu/~rsalakhu/ Lecture 9 Sequential Data So far
6.047 / Computational Biology: Genomes, Networks, Evolution Fall 2008
 MIT OpenCourseWare http://ocw.mit.edu 6.047 / 6.878 Computational Biology: Genomes, etworks, Evolution Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu 6.047 / 6.878 Computational Biology: Genomes, etworks, Evolution Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
Probabilistic Graphical Models
 School of Computer Science Probabilistic Graphical Models Max-margin learning of GM Eric Xing Lecture 28, Apr 28, 2014 b r a c e Reading: 1 Classical Predictive Models Input and output space: Predictive
School of Computer Science Probabilistic Graphical Models Max-margin learning of GM Eric Xing Lecture 28, Apr 28, 2014 b r a c e Reading: 1 Classical Predictive Models Input and output space: Predictive
CS229 Supplemental Lecture notes
 CS229 Supplemental Lecture notes John Duchi 1 Boosting We have seen so far how to solve classification (and other) problems when we have a data representation already chosen. We now talk about a procedure,
CS229 Supplemental Lecture notes John Duchi 1 Boosting We have seen so far how to solve classification (and other) problems when we have a data representation already chosen. We now talk about a procedure,
CSE 250a. Assignment Noisy-OR model. Out: Tue Oct 26 Due: Tue Nov 2
 CSE 250a. Assignment 4 Out: Tue Oct 26 Due: Tue Nov 2 4.1 Noisy-OR model X 1 X 2 X 3... X d Y For the belief network of binary random variables shown above, consider the noisy-or conditional probability
CSE 250a. Assignment 4 Out: Tue Oct 26 Due: Tue Nov 2 4.1 Noisy-OR model X 1 X 2 X 3... X d Y For the belief network of binary random variables shown above, consider the noisy-or conditional probability
Stochastic Gradient Descent
 Stochastic Gradient Descent Machine Learning CSE546 Carlos Guestrin University of Washington October 9, 2013 1 Logistic Regression Logistic function (or Sigmoid): Learn P(Y X) directly Assume a particular
Stochastic Gradient Descent Machine Learning CSE546 Carlos Guestrin University of Washington October 9, 2013 1 Logistic Regression Logistic function (or Sigmoid): Learn P(Y X) directly Assume a particular
Active learning in sequence labeling
 Active learning in sequence labeling Tomáš Šabata 11. 5. 2017 Czech Technical University in Prague Faculty of Information technology Department of Theoretical Computer Science Table of contents 1. Introduction
Active learning in sequence labeling Tomáš Šabata 11. 5. 2017 Czech Technical University in Prague Faculty of Information technology Department of Theoretical Computer Science Table of contents 1. Introduction
Biological Networks: Comparison, Conservation, and Evolution via Relative Description Length By: Tamir Tuller & Benny Chor
 Biological Networks:,, and via Relative Description Length By: Tamir Tuller & Benny Chor Presented by: Noga Grebla Content of the presentation Presenting the goals of the research Reviewing basic terms
Biological Networks:,, and via Relative Description Length By: Tamir Tuller & Benny Chor Presented by: Noga Grebla Content of the presentation Presenting the goals of the research Reviewing basic terms
UNIVERSITY of PENNSYLVANIA CIS 520: Machine Learning Final, Fall 2014
 UNIVERSITY of PENNSYLVANIA CIS 520: Machine Learning Final, Fall 2014 Exam policy: This exam allows two one-page, two-sided cheat sheets (i.e. 4 sides); No other materials. Time: 2 hours. Be sure to write
UNIVERSITY of PENNSYLVANIA CIS 520: Machine Learning Final, Fall 2014 Exam policy: This exam allows two one-page, two-sided cheat sheets (i.e. 4 sides); No other materials. Time: 2 hours. Be sure to write
CHAPTER-17. Decision Tree Induction
 CHAPTER-17 Decision Tree Induction 17.1 Introduction 17.2 Attribute selection measure 17.3 Tree Pruning 17.4 Extracting Classification Rules from Decision Trees 17.5 Bayesian Classification 17.6 Bayes
CHAPTER-17 Decision Tree Induction 17.1 Introduction 17.2 Attribute selection measure 17.3 Tree Pruning 17.4 Extracting Classification Rules from Decision Trees 17.5 Bayesian Classification 17.6 Bayes
Data Compression Techniques
 Data Compression Techniques Part 2: Text Compression Lecture 5: Context-Based Compression Juha Kärkkäinen 14.11.2017 1 / 19 Text Compression We will now look at techniques for text compression. These techniques
Data Compression Techniques Part 2: Text Compression Lecture 5: Context-Based Compression Juha Kärkkäinen 14.11.2017 1 / 19 Text Compression We will now look at techniques for text compression. These techniques
MACHINE LEARNING FOR NATURAL LANGUAGE PROCESSING
 MACHINE LEARNING FOR NATURAL LANGUAGE PROCESSING Outline Some Sample NLP Task [Noah Smith] Structured Prediction For NLP Structured Prediction Methods Conditional Random Fields Structured Perceptron Discussion
MACHINE LEARNING FOR NATURAL LANGUAGE PROCESSING Outline Some Sample NLP Task [Noah Smith] Structured Prediction For NLP Structured Prediction Methods Conditional Random Fields Structured Perceptron Discussion
Lecture 4: Training a Classifier
 Lecture 4: Training a Classifier Roger Grosse 1 Introduction Now that we ve defined what binary classification is, let s actually train a classifier. We ll approach this problem in much the same way as
Lecture 4: Training a Classifier Roger Grosse 1 Introduction Now that we ve defined what binary classification is, let s actually train a classifier. We ll approach this problem in much the same way as
Lecture 5 Neural models for NLP
 CS546: Machine Learning in NLP (Spring 2018) http://courses.engr.illinois.edu/cs546/ Lecture 5 Neural models for NLP Julia Hockenmaier juliahmr@illinois.edu 3324 Siebel Center Office hours: Tue/Thu 2pm-3pm
CS546: Machine Learning in NLP (Spring 2018) http://courses.engr.illinois.edu/cs546/ Lecture 5 Neural models for NLP Julia Hockenmaier juliahmr@illinois.edu 3324 Siebel Center Office hours: Tue/Thu 2pm-3pm
Decision Trees. Tirgul 5
 Decision Trees Tirgul 5 Using Decision Trees It could be difficult to decide which pet is right for you. We ll find a nice algorithm to help us decide what to choose without having to think about it. 2
Decision Trees Tirgul 5 Using Decision Trees It could be difficult to decide which pet is right for you. We ll find a nice algorithm to help us decide what to choose without having to think about it. 2
Real Estate Price Prediction with Regression and Classification CS 229 Autumn 2016 Project Final Report
 Real Estate Price Prediction with Regression and Classification CS 229 Autumn 2016 Project Final Report Hujia Yu, Jiafu Wu [hujiay, jiafuwu]@stanford.edu 1. Introduction Housing prices are an important
Real Estate Price Prediction with Regression and Classification CS 229 Autumn 2016 Project Final Report Hujia Yu, Jiafu Wu [hujiay, jiafuwu]@stanford.edu 1. Introduction Housing prices are an important
Symbolic methods in TC: Decision Trees
 Symbolic methods in TC: Decision Trees ML for NLP Lecturer: Kevin Koidl Assist. Lecturer Alfredo Maldonado https://www.cs.tcd.ie/kevin.koidl/cs4062/ kevin.koidl@scss.tcd.ie, maldonaa@tcd.ie 2016-2017 2
Symbolic methods in TC: Decision Trees ML for NLP Lecturer: Kevin Koidl Assist. Lecturer Alfredo Maldonado https://www.cs.tcd.ie/kevin.koidl/cs4062/ kevin.koidl@scss.tcd.ie, maldonaa@tcd.ie 2016-2017 2
SUPERVISED LEARNING: INTRODUCTION TO CLASSIFICATION
 SUPERVISED LEARNING: INTRODUCTION TO CLASSIFICATION 1 Outline Basic terminology Features Training and validation Model selection Error and loss measures Statistical comparison Evaluation measures 2 Terminology
SUPERVISED LEARNING: INTRODUCTION TO CLASSIFICATION 1 Outline Basic terminology Features Training and validation Model selection Error and loss measures Statistical comparison Evaluation measures 2 Terminology
Natural Language Processing with Deep Learning CS224N/Ling284
 Natural Language Processing with Deep Learning CS224N/Ling284 Lecture 4: Word Window Classification and Neural Networks Richard Socher Organization Main midterm: Feb 13 Alternative midterm: Friday Feb
Natural Language Processing with Deep Learning CS224N/Ling284 Lecture 4: Word Window Classification and Neural Networks Richard Socher Organization Main midterm: Feb 13 Alternative midterm: Friday Feb
More on HMMs and other sequence models. Intro to NLP - ETHZ - 18/03/2013
 More on HMMs and other sequence models Intro to NLP - ETHZ - 18/03/2013 Summary Parts of speech tagging HMMs: Unsupervised parameter estimation Forward Backward algorithm Bayesian variants Discriminative
More on HMMs and other sequence models Intro to NLP - ETHZ - 18/03/2013 Summary Parts of speech tagging HMMs: Unsupervised parameter estimation Forward Backward algorithm Bayesian variants Discriminative
Decision Trees. None Some Full > No Yes. No Yes. No Yes. No Yes. No Yes. No Yes. No Yes. Patrons? WaitEstimate? Hungry? Alternate?
 Decision rees Decision trees is one of the simplest methods for supervised learning. It can be applied to both regression & classification. Example: A decision tree for deciding whether to wait for a place
Decision rees Decision trees is one of the simplest methods for supervised learning. It can be applied to both regression & classification. Example: A decision tree for deciding whether to wait for a place
Introduction: MLE, MAP, Bayesian reasoning (28/8/13)
 STA561: Probabilistic machine learning Introduction: MLE, MAP, Bayesian reasoning (28/8/13) Lecturer: Barbara Engelhardt Scribes: K. Ulrich, J. Subramanian, N. Raval, J. O Hollaren 1 Classifiers In this
STA561: Probabilistic machine learning Introduction: MLE, MAP, Bayesian reasoning (28/8/13) Lecturer: Barbara Engelhardt Scribes: K. Ulrich, J. Subramanian, N. Raval, J. O Hollaren 1 Classifiers In this
Lecture 6: Graphical Models
 Lecture 6: Graphical Models Kai-Wei Chang CS @ Uniersity of Virginia kw@kwchang.net Some slides are adapted from Viek Skirmar s course on Structured Prediction 1 So far We discussed sequence labeling tasks:
Lecture 6: Graphical Models Kai-Wei Chang CS @ Uniersity of Virginia kw@kwchang.net Some slides are adapted from Viek Skirmar s course on Structured Prediction 1 So far We discussed sequence labeling tasks:
Linear Regression. Robot Image Credit: Viktoriya Sukhanova 123RF.com
 Linear Regression These slides were assembled by Eric Eaton, with grateful acknowledgement of the many others who made their course materials freely available online. Feel free to reuse or adapt these
Linear Regression These slides were assembled by Eric Eaton, with grateful acknowledgement of the many others who made their course materials freely available online. Feel free to reuse or adapt these
BACKPROPAGATION. Neural network training optimization problem. Deriving backpropagation
 BACKPROPAGATION Neural network training optimization problem min J(w) w The application of gradient descent to this problem is called backpropagation. Backpropagation is gradient descent applied to J(w)
BACKPROPAGATION Neural network training optimization problem min J(w) w The application of gradient descent to this problem is called backpropagation. Backpropagation is gradient descent applied to J(w)
CLUe Training An Introduction to Machine Learning in R with an example from handwritten digit recognition
 CLUe Training An Introduction to Machine Learning in R with an example from handwritten digit recognition Ad Feelders Universiteit Utrecht Department of Information and Computing Sciences Algorithmic Data
CLUe Training An Introduction to Machine Learning in R with an example from handwritten digit recognition Ad Feelders Universiteit Utrecht Department of Information and Computing Sciences Algorithmic Data
Conditional probabilities and graphical models
 Conditional probabilities and graphical models Thomas Mailund Bioinformatics Research Centre (BiRC), Aarhus University Probability theory allows us to describe uncertainty in the processes we model within
Conditional probabilities and graphical models Thomas Mailund Bioinformatics Research Centre (BiRC), Aarhus University Probability theory allows us to describe uncertainty in the processes we model within
Kernel Methods & Support Vector Machines
 Kernel Methods & Support Vector Machines Mahdi pakdaman Naeini PhD Candidate, University of Tehran Senior Researcher, TOSAN Intelligent Data Miners Outline Motivation Introduction to pattern recognition
Kernel Methods & Support Vector Machines Mahdi pakdaman Naeini PhD Candidate, University of Tehran Senior Researcher, TOSAN Intelligent Data Miners Outline Motivation Introduction to pattern recognition
Learning Multiple Tasks with a Sparse Matrix-Normal Penalty
 Learning Multiple Tasks with a Sparse Matrix-Normal Penalty Yi Zhang and Jeff Schneider NIPS 2010 Presented by Esther Salazar Duke University March 25, 2011 E. Salazar (Reading group) March 25, 2011 1
Learning Multiple Tasks with a Sparse Matrix-Normal Penalty Yi Zhang and Jeff Schneider NIPS 2010 Presented by Esther Salazar Duke University March 25, 2011 E. Salazar (Reading group) March 25, 2011 1
Linear Classification and SVM. Dr. Xin Zhang
 Linear Classification and SVM Dr. Xin Zhang Email: eexinzhang@scut.edu.cn What is linear classification? Classification is intrinsically non-linear It puts non-identical things in the same class, so a
Linear Classification and SVM Dr. Xin Zhang Email: eexinzhang@scut.edu.cn What is linear classification? Classification is intrinsically non-linear It puts non-identical things in the same class, so a
From perceptrons to word embeddings. Simon Šuster University of Groningen
 From perceptrons to word embeddings Simon Šuster University of Groningen Outline A basic computational unit Weighting some input to produce an output: classification Perceptron Classify tweets Written
From perceptrons to word embeddings Simon Šuster University of Groningen Outline A basic computational unit Weighting some input to produce an output: classification Perceptron Classify tweets Written
10 : HMM and CRF. 1 Case Study: Supervised Part-of-Speech Tagging
 10-708: Probabilistic Graphical Models 10-708, Spring 2018 10 : HMM and CRF Lecturer: Kayhan Batmanghelich Scribes: Ben Lengerich, Michael Kleyman 1 Case Study: Supervised Part-of-Speech Tagging We will
10-708: Probabilistic Graphical Models 10-708, Spring 2018 10 : HMM and CRF Lecturer: Kayhan Batmanghelich Scribes: Ben Lengerich, Michael Kleyman 1 Case Study: Supervised Part-of-Speech Tagging We will
Generative Clustering, Topic Modeling, & Bayesian Inference
 Generative Clustering, Topic Modeling, & Bayesian Inference INFO-4604, Applied Machine Learning University of Colorado Boulder December 12-14, 2017 Prof. Michael Paul Unsupervised Naïve Bayes Last week
Generative Clustering, Topic Modeling, & Bayesian Inference INFO-4604, Applied Machine Learning University of Colorado Boulder December 12-14, 2017 Prof. Michael Paul Unsupervised Naïve Bayes Last week
Logistic Regression. COMP 527 Danushka Bollegala
 Logistic Regression COMP 527 Danushka Bollegala Binary Classification Given an instance x we must classify it to either positive (1) or negative (0) class We can use {1,-1} instead of {1,0} but we will
Logistic Regression COMP 527 Danushka Bollegala Binary Classification Given an instance x we must classify it to either positive (1) or negative (0) class We can use {1,-1} instead of {1,0} but we will
Hidden Markov Models
 Hidden Markov Models A selection of slides taken from the following: Chris Bystroff Protein Folding Initiation Site Motifs Iosif Vaisman Bioinformatics and Gene Discovery Colin Cherry Hidden Markov Models
Hidden Markov Models A selection of slides taken from the following: Chris Bystroff Protein Folding Initiation Site Motifs Iosif Vaisman Bioinformatics and Gene Discovery Colin Cherry Hidden Markov Models
Classification. The goal: map from input X to a label Y. Y has a discrete set of possible values. We focused on binary Y (values 0 or 1).
 Regression and PCA Classification The goal: map from input X to a label Y. Y has a discrete set of possible values We focused on binary Y (values 0 or 1). But we also discussed larger number of classes
Regression and PCA Classification The goal: map from input X to a label Y. Y has a discrete set of possible values We focused on binary Y (values 0 or 1). But we also discussed larger number of classes
Data Mining Classification: Basic Concepts and Techniques. Lecture Notes for Chapter 3. Introduction to Data Mining, 2nd Edition
 Data Mining Classification: Basic Concepts and Techniques Lecture Notes for Chapter 3 by Tan, Steinbach, Karpatne, Kumar 1 Classification: Definition Given a collection of records (training set ) Each
Data Mining Classification: Basic Concepts and Techniques Lecture Notes for Chapter 3 by Tan, Steinbach, Karpatne, Kumar 1 Classification: Definition Given a collection of records (training set ) Each
Predictive Modeling: Classification. KSE 521 Topic 6 Mun Yi
 Predictive Modeling: Classification Topic 6 Mun Yi Agenda Models and Induction Entropy and Information Gain Tree-Based Classifier Probability Estimation 2 Introduction Key concept of BI: Predictive modeling
Predictive Modeling: Classification Topic 6 Mun Yi Agenda Models and Induction Entropy and Information Gain Tree-Based Classifier Probability Estimation 2 Introduction Key concept of BI: Predictive modeling
