MCMC: Markov Chain Monte Carlo
|
|
|
- Judith Reeves
- 5 years ago
- Views:
Transcription
1 I529: Machine Learning in Bioinformatics (Spring 2013) MCMC: Markov Chain Monte Carlo Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013
2 Contents Review of Markov Chains Monte Carlo simulation Introduction of MCMC Motivating problems MCMC updating schemes Practical implementation issues The choice of transition mechanism for the chain The number of chains to be run and their length MCMC algorithms & applications Gibbs sampler (motif finding problem)
3 Review: 1 st order Markov chain An integer time stochastic process, consisting of a set of m>1 states {s 1,,s m } and 1. An m dimensional initial distribution vector ( p(s 1 ),.., p(s m )) 2. An m m transition probabilities matrix M= (a si s j ) For example, for DNA sequence: the states are {A, C, T, G} (m=4) p(a) the probability of A to be the 1 st letter a AG the probability that G follows A in a sequence.
4 Motivating problems for MCMC The integration operation that plays a fundamental role in Bayesian statistics For calculating the normalizing constant Marginal distribution Expectation MCMC, first introduced by Metropolis (1953), provides an alternative whereby we sample from the posterior directly, and obtain sample estimates of the quantities of interest
5 Sampling and optimization To maximize a function, f(x): Brute force method: try all possible x Sample method: sample x from probability distribution: p(x) ~ f(x) Idea: suppose x max is a maximum of f(x), then it is also maximum of p(x), thus we have a high probability of sampling x max
6 Monte Carlo simulation The idea of Monte Carlo simulation is to draw an i.i.d set of N samples from a target density p(x) defined on a high-dimensional space X. I N (f) = 1 N NX Z f(x (i)! ) N!1I(f) = i=1 X The N samples can also be used to obtain a maximum of the objective function p(x) X f(x)p(x)dx
7 Rejection sampling Sample from a distribution p(x), which is known up to a proportionality constant, by sampling from another easy-to-sample proposal distribution q(x) that satisfies p(x) <= Mq(x), using an accept/reject procedure. Ref: An introduction to MCMC for Machine Learning, 2003
8 MCMC algorithms When samples cannot be drawn from p(x) directly but p(x) can be evaluated up to a normalizing constant, MCMC can be used, which is a strategy for generating samples x while exploring the state space X using a Markov chain mechanism.
9 MCMC: basics Any Markov chain which is irreducible and aperiodic will have a unique stationary distribution. Irreducibility: from any state of the Markov chain, there is a positive probability of visiting all other states (i.e., the transition matrix cannot be reduced to separate smaller matrices). Aperiodicity: the chain should not get trapped in cycles From any starting point, the chain will converge to the invariant distribution p(x), as long as T is a stochastic transition matrix that have the two properties: irreducibility & aperiodicity. MCMC samplers are irreducible and aperiodic Markov chains that have the target distribution a the invariant distribution" "
10 MCMC approaches The Metropolis-Hastings (MH) algorithm The MH algorithm is the most popular MCMC method Most practical MCMC algorithms can be interpreted as special cases or extensions of this algorithm Simulated annealing for global optimization Mixtures and cycles of MCMC kernels It is possible to combine several samplers into mixtures and cycles of the individual samplers The Gibbs sampler
11 The motif finding problem Given a set of DNA sequences: cctgatagacgctatctggctatccacgtacgtaggtcctctgtgcgaatctatgcgtttccaaccat agtactggtgtacatttgatacgtacgtacaccggcaacctgaaacaaacgctcagaaccagaagtgc aaacgtacgtgcaccctctttcttcgtggctctggccaacgagggctgatgtataagacgaaaatttt agcctccgatgtaagtcatagctgtaactattacctgccacccctattacatcttacgtacgtataca ctgttatacaacgcgtcatggcggggtatgcgttttggtcgtcgtacgctcgatcgttaacgtacgtc Find the motif in each of the individual sequences
12 The motif finding problem If starting positions s=(s 1, s 2, s t ) are given, finding consensus is easy because we can simply construct (and evaluate) the profile to find the motif. But the starting positions s are usually not given. How can we find the best profile matrix? Gibbs sampling Expectation-Maximization algorithm
13 Notations Σ Set of symbols: Sequences: S = {S 1, S 2,, S N } Starting positions of motifs: A = {a 1, a 2,, a N } Motif model (θ) : q ij = P(symbol at the i-th position = j) Background model (θ 0 ): p j = P(symbol = j) Count of symbols in each column: c ij = count of symbol j in the i-th column in the aligned motif instances
14 Motif finding problem Problem: find starting positions and model parameters simultaneously to maximize the posterior probability: max θ, A P( θ, A S) This is equivalent to maximizing the likelihood by Bayes Theorem, assuming a uniform prior distribution over different models: max θ, A P( S A, θ )
15 Equivalent scoring function Maximize the log-odds ratio: P W ( S A, θ ) = q ij ij P ( S A, θ ) 0 = i= 1 Σ j= 1 c W i= 1 Σ j= 1 cij p j Motif model (θ) : q ij = P(symbol at the i-th position = j) Background model (θ 0 ): p j = P(symbol = j) c ij : #(symbol j at position i) F = log P( S A, P( S A, Log of the ratio θ θ 0 ) ) = W Σ ij cij log i= 1 j= 1 p j q
16 Gibbs sampling Idea: a joint distribution in a high dimension may be hard to sample from, but it may be easy to sample from the conditional distributions where all variables are fixed except one To sample from p(x 1, x 2, x n ), let each state of the Markov chain represent (x 1, x 2, x n ), the probability of moving to a state (x 1, x 2, x n ) is: p(x i x 1, x i-1,x i+1, x n ). It is a algorithm in a class of sampling techniques called Markov Chain Monte Carlo (MCMC) method.
17 Gibbs sampling Best Location New Location Start with random motif locations and calculate a motif model Randomly select a sequence, remove its motif and recalculate temporary model A C G T With temporary model, calculate probability of motif at each position on sequence Select new position based on this distribution Update model and Iterate A C G T ETC
18 Gibbs sampling Initialization, t=0, sample ( x (0) 1,x (0) 2,x (0) (0) 3,...,x ) n (t x +1) 1 ~ P( (t x 1 x ) (t 2, x ) (t ) 3,..., x ) n (t x +1) 2 ~ P( (t x 2 x +1) (t 1,x ) (t ) 3,..., x ) n (t x +1) 3 ~ P( (t x 2 x +1) (t 1,x +1) (t 2,x ) (t ) 4,...,x ) n...,... (t x +1) n ~ P( (t x n x +1) (t 1, x +1) (t +1) 3,..., x ) n 1
19 Estimator of θ Given an alignment A, i.e. the starting positions of motifs, θ can be estimated by its MLE with prior probabilities (e.g. Dirichlet prior with parameter b j ): q ij = c ij + b N 1+ B j where B=Σ j b j
20 Finding TF binding sites TF CACGTGT Gene 1 TF CACGTGA Gene 2 TF CAAGTGA Gene 3 TF CAGGTGA Gene 4 Transcription factor binding site, or motif instances
21 Sampling motifs on trees Using both the overpresentation property and the evolutionary conservation property of motifs CACGTGACC CACGTGAAC CACGTGAAC Ref: Sampling motifs on phylogenetic trees, Li & Wong, PNAS, 2005
22 Implementation: Initialization Initialization Parameters are sampled using prior distributions; Motif instances in current species are sampled from sequences directly for each current species; Motif instances in ancestral species are randomly assigned with one of its immediate child motif instances.
23 Motif instance updating Updating motif instances in ancestral species (0) A 1 (0) (0) (1) (2) Pr( A1 A[ 1], A1, A1, Θ0, p1, p2, w, M11, M12) M 11 M 12 Updating motif instances in current species (1) (0) Pr( A1 A1, S, Θ0, p1, w, M11) (1) A 1 (2) A 1
24 Motif instance updating Updating motif instance in ancestral species Ancestral Motif Weight Matrix A C G T th position A: C: G: 8.4e-6 T: 2.5e-4 M 11 M 12 M 11 M 12 CCCGTGACC CACGTGAAC C A
25 Motif instance updating Updating motif instances for current species Updated ancestral motif instance CACTTGAAC M 11 M 12 CACACCACGTGAGCTT... CACATCACGTGAACTT
26 Parameter sampling step Metropolis-Hasting algorithm is used to increase or decrease the width w of the motif by 1 from the left or right side
27 Other applications of Gibbs sampling Biclustering microarray data by Gibbs sampling Microarray data is discretized Bioinformatics, 2003 Assignment of ambiguously mapped reads Bioinformatics, 2010
28 Practical implementation issues How many iterations? One run or many?
Introduction to Machine Learning CMU-10701
 Introduction to Machine Learning CMU-10701 Markov Chain Monte Carlo Methods Barnabás Póczos & Aarti Singh Contents Markov Chain Monte Carlo Methods Goal & Motivation Sampling Rejection Importance Markov
Introduction to Machine Learning CMU-10701 Markov Chain Monte Carlo Methods Barnabás Póczos & Aarti Singh Contents Markov Chain Monte Carlo Methods Goal & Motivation Sampling Rejection Importance Markov
Expectation-Maximization (EM) algorithm
 I529: Machine Learning in Bioinformatics (Spring 2017) Expectation-Maximization (EM) algorithm Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Contents Introduce
I529: Machine Learning in Bioinformatics (Spring 2017) Expectation-Maximization (EM) algorithm Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Contents Introduce
CPSC 540: Machine Learning
 CPSC 540: Machine Learning MCMC and Non-Parametric Bayes Mark Schmidt University of British Columbia Winter 2016 Admin I went through project proposals: Some of you got a message on Piazza. No news is
CPSC 540: Machine Learning MCMC and Non-Parametric Bayes Mark Schmidt University of British Columbia Winter 2016 Admin I went through project proposals: Some of you got a message on Piazza. No news is
Bayesian Inference and MCMC
 Bayesian Inference and MCMC Aryan Arbabi Partly based on MCMC slides from CSC412 Fall 2018 1 / 18 Bayesian Inference - Motivation Consider we have a data set D = {x 1,..., x n }. E.g each x i can be the
Bayesian Inference and MCMC Aryan Arbabi Partly based on MCMC slides from CSC412 Fall 2018 1 / 18 Bayesian Inference - Motivation Consider we have a data set D = {x 1,..., x n }. E.g each x i can be the
Convex Optimization CMU-10725
 Convex Optimization CMU-10725 Simulated Annealing Barnabás Póczos & Ryan Tibshirani Andrey Markov Markov Chains 2 Markov Chains Markov chain: Homogen Markov chain: 3 Markov Chains Assume that the state
Convex Optimization CMU-10725 Simulated Annealing Barnabás Póczos & Ryan Tibshirani Andrey Markov Markov Chains 2 Markov Chains Markov chain: Homogen Markov chain: 3 Markov Chains Assume that the state
Quantifying Uncertainty
 Sai Ravela M. I. T Last Updated: Spring 2013 1 Markov Chain Monte Carlo Monte Carlo sampling made for large scale problems via Markov Chains Monte Carlo Sampling Rejection Sampling Importance Sampling
Sai Ravela M. I. T Last Updated: Spring 2013 1 Markov Chain Monte Carlo Monte Carlo sampling made for large scale problems via Markov Chains Monte Carlo Sampling Rejection Sampling Importance Sampling
MCMC and Gibbs Sampling. Kayhan Batmanghelich
 MCMC and Gibbs Sampling Kayhan Batmanghelich 1 Approaches to inference l Exact inference algorithms l l l The elimination algorithm Message-passing algorithm (sum-product, belief propagation) The junction
MCMC and Gibbs Sampling Kayhan Batmanghelich 1 Approaches to inference l Exact inference algorithms l l l The elimination algorithm Message-passing algorithm (sum-product, belief propagation) The junction
Computer Vision Group Prof. Daniel Cremers. 10a. Markov Chain Monte Carlo
 Group Prof. Daniel Cremers 10a. Markov Chain Monte Carlo Markov Chain Monte Carlo In high-dimensional spaces, rejection sampling and importance sampling are very inefficient An alternative is Markov Chain
Group Prof. Daniel Cremers 10a. Markov Chain Monte Carlo Markov Chain Monte Carlo In high-dimensional spaces, rejection sampling and importance sampling are very inefficient An alternative is Markov Chain
Introduction to Computational Biology Lecture # 14: MCMC - Markov Chain Monte Carlo
 Introduction to Computational Biology Lecture # 14: MCMC - Markov Chain Monte Carlo Assaf Weiner Tuesday, March 13, 2007 1 Introduction Today we will return to the motif finding problem, in lecture 10
Introduction to Computational Biology Lecture # 14: MCMC - Markov Chain Monte Carlo Assaf Weiner Tuesday, March 13, 2007 1 Introduction Today we will return to the motif finding problem, in lecture 10
Computer Vision Group Prof. Daniel Cremers. 11. Sampling Methods: Markov Chain Monte Carlo
 Group Prof. Daniel Cremers 11. Sampling Methods: Markov Chain Monte Carlo Markov Chain Monte Carlo In high-dimensional spaces, rejection sampling and importance sampling are very inefficient An alternative
Group Prof. Daniel Cremers 11. Sampling Methods: Markov Chain Monte Carlo Markov Chain Monte Carlo In high-dimensional spaces, rejection sampling and importance sampling are very inefficient An alternative
STA 4273H: Statistical Machine Learning
 STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Computer Science! Department of Statistical Sciences! rsalakhu@cs.toronto.edu! h0p://www.cs.utoronto.ca/~rsalakhu/ Lecture 7 Approximate
STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Computer Science! Department of Statistical Sciences! rsalakhu@cs.toronto.edu! h0p://www.cs.utoronto.ca/~rsalakhu/ Lecture 7 Approximate
Bayesian Methods for Machine Learning
 Bayesian Methods for Machine Learning CS 584: Big Data Analytics Material adapted from Radford Neal s tutorial (http://ftp.cs.utoronto.ca/pub/radford/bayes-tut.pdf), Zoubin Ghahramni (http://hunch.net/~coms-4771/zoubin_ghahramani_bayesian_learning.pdf),
Bayesian Methods for Machine Learning CS 584: Big Data Analytics Material adapted from Radford Neal s tutorial (http://ftp.cs.utoronto.ca/pub/radford/bayes-tut.pdf), Zoubin Ghahramni (http://hunch.net/~coms-4771/zoubin_ghahramani_bayesian_learning.pdf),
Learning Sequence Motif Models Using Expectation Maximization (EM) and Gibbs Sampling
 Learning Sequence Motif Models Using Expectation Maximization (EM) and Gibbs Sampling BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 009 Mark Craven craven@biostat.wisc.edu Sequence Motifs what is a sequence
Learning Sequence Motif Models Using Expectation Maximization (EM) and Gibbs Sampling BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 009 Mark Craven craven@biostat.wisc.edu Sequence Motifs what is a sequence
Markov Chain Monte Carlo Lecture 6
 Sequential parallel tempering With the development of science and technology, we more and more need to deal with high dimensional systems. For example, we need to align a group of protein or DNA sequences
Sequential parallel tempering With the development of science and technology, we more and more need to deal with high dimensional systems. For example, we need to align a group of protein or DNA sequences
Markov chain Monte Carlo Lecture 9
 Markov chain Monte Carlo Lecture 9 David Sontag New York University Slides adapted from Eric Xing and Qirong Ho (CMU) Limitations of Monte Carlo Direct (unconditional) sampling Hard to get rare events
Markov chain Monte Carlo Lecture 9 David Sontag New York University Slides adapted from Eric Xing and Qirong Ho (CMU) Limitations of Monte Carlo Direct (unconditional) sampling Hard to get rare events
Bayesian Phylogenetics:
 Bayesian Phylogenetics: an introduction Marc A. Suchard msuchard@ucla.edu UCLA Who is this man? How sure are you? The one true tree? Methods we ve learned so far try to find a single tree that best describes
Bayesian Phylogenetics: an introduction Marc A. Suchard msuchard@ucla.edu UCLA Who is this man? How sure are you? The one true tree? Methods we ve learned so far try to find a single tree that best describes
LECTURE 15 Markov chain Monte Carlo
 LECTURE 15 Markov chain Monte Carlo There are many settings when posterior computation is a challenge in that one does not have a closed form expression for the posterior distribution. Markov chain Monte
LECTURE 15 Markov chain Monte Carlo There are many settings when posterior computation is a challenge in that one does not have a closed form expression for the posterior distribution. Markov chain Monte
17 : Markov Chain Monte Carlo
 10-708: Probabilistic Graphical Models, Spring 2015 17 : Markov Chain Monte Carlo Lecturer: Eric P. Xing Scribes: Heran Lin, Bin Deng, Yun Huang 1 Review of Monte Carlo Methods 1.1 Overview Monte Carlo
10-708: Probabilistic Graphical Models, Spring 2015 17 : Markov Chain Monte Carlo Lecturer: Eric P. Xing Scribes: Heran Lin, Bin Deng, Yun Huang 1 Review of Monte Carlo Methods 1.1 Overview Monte Carlo
Computational statistics
 Computational statistics Markov Chain Monte Carlo methods Thierry Denœux March 2017 Thierry Denœux Computational statistics March 2017 1 / 71 Contents of this chapter When a target density f can be evaluated
Computational statistics Markov Chain Monte Carlo methods Thierry Denœux March 2017 Thierry Denœux Computational statistics March 2017 1 / 71 Contents of this chapter When a target density f can be evaluated
Monte Carlo Methods. Leon Gu CSD, CMU
 Monte Carlo Methods Leon Gu CSD, CMU Approximate Inference EM: y-observed variables; x-hidden variables; θ-parameters; E-step: q(x) = p(x y, θ t 1 ) M-step: θ t = arg max E q(x) [log p(y, x θ)] θ Monte
Monte Carlo Methods Leon Gu CSD, CMU Approximate Inference EM: y-observed variables; x-hidden variables; θ-parameters; E-step: q(x) = p(x y, θ t 1 ) M-step: θ t = arg max E q(x) [log p(y, x θ)] θ Monte
Machine Learning for Data Science (CS4786) Lecture 24
 Machine Learning for Data Science (CS4786) Lecture 24 Graphical Models: Approximate Inference Course Webpage : http://www.cs.cornell.edu/courses/cs4786/2016sp/ BELIEF PROPAGATION OR MESSAGE PASSING Each
Machine Learning for Data Science (CS4786) Lecture 24 Graphical Models: Approximate Inference Course Webpage : http://www.cs.cornell.edu/courses/cs4786/2016sp/ BELIEF PROPAGATION OR MESSAGE PASSING Each
Markov Chain Monte Carlo
 1 Motivation 1.1 Bayesian Learning Markov Chain Monte Carlo Yale Chang In Bayesian learning, given data X, we make assumptions on the generative process of X by introducing hidden variables Z: p(z): prior
1 Motivation 1.1 Bayesian Learning Markov Chain Monte Carlo Yale Chang In Bayesian learning, given data X, we make assumptions on the generative process of X by introducing hidden variables Z: p(z): prior
MARKOV CHAIN MONTE CARLO
 MARKOV CHAIN MONTE CARLO RYAN WANG Abstract. This paper gives a brief introduction to Markov Chain Monte Carlo methods, which offer a general framework for calculating difficult integrals. We start with
MARKOV CHAIN MONTE CARLO RYAN WANG Abstract. This paper gives a brief introduction to Markov Chain Monte Carlo methods, which offer a general framework for calculating difficult integrals. We start with
Markov Chain Monte Carlo Inference. Siamak Ravanbakhsh Winter 2018
 Graphical Models Markov Chain Monte Carlo Inference Siamak Ravanbakhsh Winter 2018 Learning objectives Markov chains the idea behind Markov Chain Monte Carlo (MCMC) two important examples: Gibbs sampling
Graphical Models Markov Chain Monte Carlo Inference Siamak Ravanbakhsh Winter 2018 Learning objectives Markov chains the idea behind Markov Chain Monte Carlo (MCMC) two important examples: Gibbs sampling
6 Markov Chain Monte Carlo (MCMC)
 6 Markov Chain Monte Carlo (MCMC) The underlying idea in MCMC is to replace the iid samples of basic MC methods, with dependent samples from an ergodic Markov chain, whose limiting (stationary) distribution
6 Markov Chain Monte Carlo (MCMC) The underlying idea in MCMC is to replace the iid samples of basic MC methods, with dependent samples from an ergodic Markov chain, whose limiting (stationary) distribution
Markov Chains and MCMC
 Markov Chains and MCMC CompSci 590.02 Instructor: AshwinMachanavajjhala Lecture 4 : 590.02 Spring 13 1 Recap: Monte Carlo Method If U is a universe of items, and G is a subset satisfying some property,
Markov Chains and MCMC CompSci 590.02 Instructor: AshwinMachanavajjhala Lecture 4 : 590.02 Spring 13 1 Recap: Monte Carlo Method If U is a universe of items, and G is a subset satisfying some property,
April 20th, Advanced Topics in Machine Learning California Institute of Technology. Markov Chain Monte Carlo for Machine Learning
 for for Advanced Topics in California Institute of Technology April 20th, 2017 1 / 50 Table of Contents for 1 2 3 4 2 / 50 History of methods for Enrico Fermi used to calculate incredibly accurate predictions
for for Advanced Topics in California Institute of Technology April 20th, 2017 1 / 50 Table of Contents for 1 2 3 4 2 / 50 History of methods for Enrico Fermi used to calculate incredibly accurate predictions
The Particle Filter. PD Dr. Rudolph Triebel Computer Vision Group. Machine Learning for Computer Vision
 The Particle Filter Non-parametric implementation of Bayes filter Represents the belief (posterior) random state samples. by a set of This representation is approximate. Can represent distributions that
The Particle Filter Non-parametric implementation of Bayes filter Represents the belief (posterior) random state samples. by a set of This representation is approximate. Can represent distributions that
Computer Vision Group Prof. Daniel Cremers. 14. Sampling Methods
 Prof. Daniel Cremers 14. Sampling Methods Sampling Methods Sampling Methods are widely used in Computer Science as an approximation of a deterministic algorithm to represent uncertainty without a parametric
Prof. Daniel Cremers 14. Sampling Methods Sampling Methods Sampling Methods are widely used in Computer Science as an approximation of a deterministic algorithm to represent uncertainty without a parametric
Introduction to Markov Chain Monte Carlo & Gibbs Sampling
 Introduction to Markov Chain Monte Carlo & Gibbs Sampling Prof. Nicholas Zabaras Sibley School of Mechanical and Aerospace Engineering 101 Frank H. T. Rhodes Hall Ithaca, NY 14853-3801 Email: zabaras@cornell.edu
Introduction to Markov Chain Monte Carlo & Gibbs Sampling Prof. Nicholas Zabaras Sibley School of Mechanical and Aerospace Engineering 101 Frank H. T. Rhodes Hall Ithaca, NY 14853-3801 Email: zabaras@cornell.edu
27 : Distributed Monte Carlo Markov Chain. 1 Recap of MCMC and Naive Parallel Gibbs Sampling
 10-708: Probabilistic Graphical Models 10-708, Spring 2014 27 : Distributed Monte Carlo Markov Chain Lecturer: Eric P. Xing Scribes: Pengtao Xie, Khoa Luu In this scribe, we are going to review the Parallel
10-708: Probabilistic Graphical Models 10-708, Spring 2014 27 : Distributed Monte Carlo Markov Chain Lecturer: Eric P. Xing Scribes: Pengtao Xie, Khoa Luu In this scribe, we are going to review the Parallel
Stat 535 C - Statistical Computing & Monte Carlo Methods. Lecture February Arnaud Doucet
 Stat 535 C - Statistical Computing & Monte Carlo Methods Lecture 13-28 February 2006 Arnaud Doucet Email: arnaud@cs.ubc.ca 1 1.1 Outline Limitations of Gibbs sampling. Metropolis-Hastings algorithm. Proof
Stat 535 C - Statistical Computing & Monte Carlo Methods Lecture 13-28 February 2006 Arnaud Doucet Email: arnaud@cs.ubc.ca 1 1.1 Outline Limitations of Gibbs sampling. Metropolis-Hastings algorithm. Proof
16 : Approximate Inference: Markov Chain Monte Carlo
 10-708: Probabilistic Graphical Models 10-708, Spring 2017 16 : Approximate Inference: Markov Chain Monte Carlo Lecturer: Eric P. Xing Scribes: Yuan Yang, Chao-Ming Yen 1 Introduction As the target distribution
10-708: Probabilistic Graphical Models 10-708, Spring 2017 16 : Approximate Inference: Markov Chain Monte Carlo Lecturer: Eric P. Xing Scribes: Yuan Yang, Chao-Ming Yen 1 Introduction As the target distribution
Computer Vision Group Prof. Daniel Cremers. 11. Sampling Methods
 Prof. Daniel Cremers 11. Sampling Methods Sampling Methods Sampling Methods are widely used in Computer Science as an approximation of a deterministic algorithm to represent uncertainty without a parametric
Prof. Daniel Cremers 11. Sampling Methods Sampling Methods Sampling Methods are widely used in Computer Science as an approximation of a deterministic algorithm to represent uncertainty without a parametric
Introduction to Machine Learning CMU-10701
 Introduction to Machine Learning CMU-10701 Markov Chain Monte Carlo Methods Barnabás Póczos Contents Markov Chain Monte Carlo Methods Sampling Rejection Importance Hastings-Metropolis Gibbs Markov Chains
Introduction to Machine Learning CMU-10701 Markov Chain Monte Carlo Methods Barnabás Póczos Contents Markov Chain Monte Carlo Methods Sampling Rejection Importance Hastings-Metropolis Gibbs Markov Chains
STA 294: Stochastic Processes & Bayesian Nonparametrics
 MARKOV CHAINS AND CONVERGENCE CONCEPTS Markov chains are among the simplest stochastic processes, just one step beyond iid sequences of random variables. Traditionally they ve been used in modelling a
MARKOV CHAINS AND CONVERGENCE CONCEPTS Markov chains are among the simplest stochastic processes, just one step beyond iid sequences of random variables. Traditionally they ve been used in modelling a
 SC7/SM6 Bayes Methods HT18 Lecturer: Geoff Nicholls Lecture 2: Monte Carlo Methods Notes and Problem sheets are available at http://www.stats.ox.ac.uk/~nicholls/bayesmethods/ and via the MSc weblearn pages.
SC7/SM6 Bayes Methods HT18 Lecturer: Geoff Nicholls Lecture 2: Monte Carlo Methods Notes and Problem sheets are available at http://www.stats.ox.ac.uk/~nicholls/bayesmethods/ and via the MSc weblearn pages.
Markov Chain Monte Carlo
 Markov Chain Monte Carlo Recall: To compute the expectation E ( h(y ) ) we use the approximation E(h(Y )) 1 n n h(y ) t=1 with Y (1),..., Y (n) h(y). Thus our aim is to sample Y (1),..., Y (n) from f(y).
Markov Chain Monte Carlo Recall: To compute the expectation E ( h(y ) ) we use the approximation E(h(Y )) 1 n n h(y ) t=1 with Y (1),..., Y (n) h(y). Thus our aim is to sample Y (1),..., Y (n) from f(y).
Markov chain Monte Carlo
 1 / 26 Markov chain Monte Carlo Timothy Hanson 1 and Alejandro Jara 2 1 Division of Biostatistics, University of Minnesota, USA 2 Department of Statistics, Universidad de Concepción, Chile IAP-Workshop
1 / 26 Markov chain Monte Carlo Timothy Hanson 1 and Alejandro Jara 2 1 Division of Biostatistics, University of Minnesota, USA 2 Department of Statistics, Universidad de Concepción, Chile IAP-Workshop
Markov Chain Monte Carlo Methods
 Markov Chain Monte Carlo Methods p. /36 Markov Chain Monte Carlo Methods Michel Bierlaire michel.bierlaire@epfl.ch Transport and Mobility Laboratory Markov Chain Monte Carlo Methods p. 2/36 Markov Chains
Markov Chain Monte Carlo Methods p. /36 Markov Chain Monte Carlo Methods Michel Bierlaire michel.bierlaire@epfl.ch Transport and Mobility Laboratory Markov Chain Monte Carlo Methods p. 2/36 Markov Chains
Lecture 6: Markov Chain Monte Carlo
 Lecture 6: Markov Chain Monte Carlo D. Jason Koskinen koskinen@nbi.ku.dk Photo by Howard Jackman University of Copenhagen Advanced Methods in Applied Statistics Feb - Apr 2016 Niels Bohr Institute 2 Outline
Lecture 6: Markov Chain Monte Carlo D. Jason Koskinen koskinen@nbi.ku.dk Photo by Howard Jackman University of Copenhagen Advanced Methods in Applied Statistics Feb - Apr 2016 Niels Bohr Institute 2 Outline
SAMPLING ALGORITHMS. In general. Inference in Bayesian models
 SAMPLING ALGORITHMS SAMPLING ALGORITHMS In general A sampling algorithm is an algorithm that outputs samples x 1, x 2,... from a given distribution P or density p. Sampling algorithms can for example be
SAMPLING ALGORITHMS SAMPLING ALGORITHMS In general A sampling algorithm is an algorithm that outputs samples x 1, x 2,... from a given distribution P or density p. Sampling algorithms can for example be
Neyman-Pearson. More Motifs. Weight Matrix Models. What s best WMM?
 Neyman-Pearson More Motifs WMM, log odds scores, Neyman-Pearson, background; Greedy & EM for motif discovery Given a sample x 1, x 2,..., x n, from a distribution f(... #) with parameter #, want to test
Neyman-Pearson More Motifs WMM, log odds scores, Neyman-Pearson, background; Greedy & EM for motif discovery Given a sample x 1, x 2,..., x n, from a distribution f(... #) with parameter #, want to test
Who was Bayes? Bayesian Phylogenetics. What is Bayes Theorem?
 Who was Bayes? Bayesian Phylogenetics Bret Larget Departments of Botany and of Statistics University of Wisconsin Madison October 6, 2011 The Reverand Thomas Bayes was born in London in 1702. He was the
Who was Bayes? Bayesian Phylogenetics Bret Larget Departments of Botany and of Statistics University of Wisconsin Madison October 6, 2011 The Reverand Thomas Bayes was born in London in 1702. He was the
Answers and expectations
 Answers and expectations For a function f(x) and distribution P(x), the expectation of f with respect to P is The expectation is the average of f, when x is drawn from the probability distribution P E
Answers and expectations For a function f(x) and distribution P(x), the expectation of f with respect to P is The expectation is the average of f, when x is drawn from the probability distribution P E
Bayesian Phylogenetics
 Bayesian Phylogenetics Bret Larget Departments of Botany and of Statistics University of Wisconsin Madison October 6, 2011 Bayesian Phylogenetics 1 / 27 Who was Bayes? The Reverand Thomas Bayes was born
Bayesian Phylogenetics Bret Larget Departments of Botany and of Statistics University of Wisconsin Madison October 6, 2011 Bayesian Phylogenetics 1 / 27 Who was Bayes? The Reverand Thomas Bayes was born
Gibbs Sampling Methods for Multiple Sequence Alignment
 Gibbs Sampling Methods for Multiple Sequence Alignment Scott C. Schmidler 1 Jun S. Liu 2 1 Section on Medical Informatics and 2 Department of Statistics Stanford University 11/17/99 1 Outline Statistical
Gibbs Sampling Methods for Multiple Sequence Alignment Scott C. Schmidler 1 Jun S. Liu 2 1 Section on Medical Informatics and 2 Department of Statistics Stanford University 11/17/99 1 Outline Statistical
Stat 451 Lecture Notes Markov Chain Monte Carlo. Ryan Martin UIC
 Stat 451 Lecture Notes 07 12 Markov Chain Monte Carlo Ryan Martin UIC www.math.uic.edu/~rgmartin 1 Based on Chapters 8 9 in Givens & Hoeting, Chapters 25 27 in Lange 2 Updated: April 4, 2016 1 / 42 Outline
Stat 451 Lecture Notes 07 12 Markov Chain Monte Carlo Ryan Martin UIC www.math.uic.edu/~rgmartin 1 Based on Chapters 8 9 in Givens & Hoeting, Chapters 25 27 in Lange 2 Updated: April 4, 2016 1 / 42 Outline
3/1/17. Content. TWINSCAN model. Example. TWINSCAN algorithm. HMM for modeling aligned multiple sequences: phylo-hmm & multivariate HMM
 I529: Machine Learning in Bioinformatics (Spring 2017) Content HMM for modeling aligned multiple sequences: phylo-hmm & multivariate HMM Yuzhen Ye School of Informatics and Computing Indiana University,
I529: Machine Learning in Bioinformatics (Spring 2017) Content HMM for modeling aligned multiple sequences: phylo-hmm & multivariate HMM Yuzhen Ye School of Informatics and Computing Indiana University,
Markov Chain Monte Carlo (MCMC)
 Markov Chain Monte Carlo (MCMC Dependent Sampling Suppose we wish to sample from a density π, and we can evaluate π as a function but have no means to directly generate a sample. Rejection sampling can
Markov Chain Monte Carlo (MCMC Dependent Sampling Suppose we wish to sample from a density π, and we can evaluate π as a function but have no means to directly generate a sample. Rejection sampling can
References. Markov-Chain Monte Carlo. Recall: Sampling Motivation. Problem. Recall: Sampling Methods. CSE586 Computer Vision II
 References Markov-Chain Monte Carlo CSE586 Computer Vision II Spring 2010, Penn State Univ. Recall: Sampling Motivation If we can generate random samples x i from a given distribution P(x), then we can
References Markov-Chain Monte Carlo CSE586 Computer Vision II Spring 2010, Penn State Univ. Recall: Sampling Motivation If we can generate random samples x i from a given distribution P(x), then we can
Brief introduction to Markov Chain Monte Carlo
 Brief introduction to Department of Probability and Mathematical Statistics seminar Stochastic modeling in economics and finance November 7, 2011 Brief introduction to Content 1 and motivation Classical
Brief introduction to Department of Probability and Mathematical Statistics seminar Stochastic modeling in economics and finance November 7, 2011 Brief introduction to Content 1 and motivation Classical
HMM: Parameter Estimation
 I529: Machine Learning in Bioinformatics (Spring 2017) HMM: Parameter Estimation Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Content Review HMM: three problems
I529: Machine Learning in Bioinformatics (Spring 2017) HMM: Parameter Estimation Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Content Review HMM: three problems
Markov Chain Monte Carlo, Numerical Integration
 Markov Chain Monte Carlo, Numerical Integration (See Statistics) Trevor Gallen Fall 2015 1 / 1 Agenda Numerical Integration: MCMC methods Estimating Markov Chains Estimating latent variables 2 / 1 Numerical
Markov Chain Monte Carlo, Numerical Integration (See Statistics) Trevor Gallen Fall 2015 1 / 1 Agenda Numerical Integration: MCMC methods Estimating Markov Chains Estimating latent variables 2 / 1 Numerical
HMM for modeling aligned multiple sequences: phylo-hmm & multivariate HMM
 I529: Machine Learning in Bioinformatics (Spring 2017) HMM for modeling aligned multiple sequences: phylo-hmm & multivariate HMM Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington
I529: Machine Learning in Bioinformatics (Spring 2017) HMM for modeling aligned multiple sequences: phylo-hmm & multivariate HMM Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington
Markov chain Monte Carlo
 Markov chain Monte Carlo Markov chain Monte Carlo (MCMC) Gibbs and Metropolis Hastings Slice sampling Practical details Iain Murray http://iainmurray.net/ Reminder Need to sample large, non-standard distributions:
Markov chain Monte Carlo Markov chain Monte Carlo (MCMC) Gibbs and Metropolis Hastings Slice sampling Practical details Iain Murray http://iainmurray.net/ Reminder Need to sample large, non-standard distributions:
MCMC algorithms for fitting Bayesian models
 MCMC algorithms for fitting Bayesian models p. 1/1 MCMC algorithms for fitting Bayesian models Sudipto Banerjee sudiptob@biostat.umn.edu University of Minnesota MCMC algorithms for fitting Bayesian models
MCMC algorithms for fitting Bayesian models p. 1/1 MCMC algorithms for fitting Bayesian models Sudipto Banerjee sudiptob@biostat.umn.edu University of Minnesota MCMC algorithms for fitting Bayesian models
Stochastic optimization Markov Chain Monte Carlo
 Stochastic optimization Markov Chain Monte Carlo Ethan Fetaya Weizmann Institute of Science 1 Motivation Markov chains Stationary distribution Mixing time 2 Algorithms Metropolis-Hastings Simulated Annealing
Stochastic optimization Markov Chain Monte Carlo Ethan Fetaya Weizmann Institute of Science 1 Motivation Markov chains Stationary distribution Mixing time 2 Algorithms Metropolis-Hastings Simulated Annealing
Markov Chain Monte Carlo
 Chapter 5 Markov Chain Monte Carlo MCMC is a kind of improvement of the Monte Carlo method By sampling from a Markov chain whose stationary distribution is the desired sampling distributuion, it is possible
Chapter 5 Markov Chain Monte Carlo MCMC is a kind of improvement of the Monte Carlo method By sampling from a Markov chain whose stationary distribution is the desired sampling distributuion, it is possible
Bayesian GLMs and Metropolis-Hastings Algorithm
 Bayesian GLMs and Metropolis-Hastings Algorithm We have seen that with conjugate or semi-conjugate prior distributions the Gibbs sampler can be used to sample from the posterior distribution. In situations,
Bayesian GLMs and Metropolis-Hastings Algorithm We have seen that with conjugate or semi-conjugate prior distributions the Gibbs sampler can be used to sample from the posterior distribution. In situations,
Stat 516, Homework 1
 Stat 516, Homework 1 Due date: October 7 1. Consider an urn with n distinct balls numbered 1,..., n. We sample balls from the urn with replacement. Let N be the number of draws until we encounter a ball
Stat 516, Homework 1 Due date: October 7 1. Consider an urn with n distinct balls numbered 1,..., n. We sample balls from the urn with replacement. Let N be the number of draws until we encounter a ball
Phylogenetics: Bayesian Phylogenetic Analysis. COMP Spring 2015 Luay Nakhleh, Rice University
 Phylogenetics: Bayesian Phylogenetic Analysis COMP 571 - Spring 2015 Luay Nakhleh, Rice University Bayes Rule P(X = x Y = y) = P(X = x, Y = y) P(Y = y) = P(X = x)p(y = y X = x) P x P(X = x 0 )P(Y = y X
Phylogenetics: Bayesian Phylogenetic Analysis COMP 571 - Spring 2015 Luay Nakhleh, Rice University Bayes Rule P(X = x Y = y) = P(X = x, Y = y) P(Y = y) = P(X = x)p(y = y X = x) P x P(X = x 0 )P(Y = y X
Markov-Chain Monte Carlo
 Markov-Chain Monte Carlo CSE586 Computer Vision II Spring 2010, Penn State Univ. References Recall: Sampling Motivation If we can generate random samples x i from a given distribution P(x), then we can
Markov-Chain Monte Carlo CSE586 Computer Vision II Spring 2010, Penn State Univ. References Recall: Sampling Motivation If we can generate random samples x i from a given distribution P(x), then we can
Robert Collins CSE586, PSU Intro to Sampling Methods
 Robert Collins Intro to Sampling Methods CSE586 Computer Vision II Penn State Univ Robert Collins A Brief Overview of Sampling Monte Carlo Integration Sampling and Expected Values Inverse Transform Sampling
Robert Collins Intro to Sampling Methods CSE586 Computer Vision II Penn State Univ Robert Collins A Brief Overview of Sampling Monte Carlo Integration Sampling and Expected Values Inverse Transform Sampling
Bayesian Inference in GLMs. Frequentists typically base inferences on MLEs, asymptotic confidence
 Bayesian Inference in GLMs Frequentists typically base inferences on MLEs, asymptotic confidence limits, and log-likelihood ratio tests Bayesians base inferences on the posterior distribution of the unknowns
Bayesian Inference in GLMs Frequentists typically base inferences on MLEs, asymptotic confidence limits, and log-likelihood ratio tests Bayesians base inferences on the posterior distribution of the unknowns
Bayes Nets: Sampling
 Bayes Nets: Sampling [These slides were created by Dan Klein and Pieter Abbeel for CS188 Intro to AI at UC Berkeley. All CS188 materials are available at http://ai.berkeley.edu.] Approximate Inference:
Bayes Nets: Sampling [These slides were created by Dan Klein and Pieter Abbeel for CS188 Intro to AI at UC Berkeley. All CS188 materials are available at http://ai.berkeley.edu.] Approximate Inference:
MCMC notes by Mark Holder
 MCMC notes by Mark Holder Bayesian inference Ultimately, we want to make probability statements about true values of parameters, given our data. For example P(α 0 < α 1 X). According to Bayes theorem:
MCMC notes by Mark Holder Bayesian inference Ultimately, we want to make probability statements about true values of parameters, given our data. For example P(α 0 < α 1 X). According to Bayes theorem:
1 Probabilities. 1.1 Basics 1 PROBABILITIES
 1 PROBABILITIES 1 Probabilities Probability is a tricky word usually meaning the likelyhood of something occuring or how frequent something is. Obviously, if something happens frequently, then its probability
1 PROBABILITIES 1 Probabilities Probability is a tricky word usually meaning the likelyhood of something occuring or how frequent something is. Obviously, if something happens frequently, then its probability
Graphical Models and Kernel Methods
 Graphical Models and Kernel Methods Jerry Zhu Department of Computer Sciences University of Wisconsin Madison, USA MLSS June 17, 2014 1 / 123 Outline Graphical Models Probabilistic Inference Directed vs.
Graphical Models and Kernel Methods Jerry Zhu Department of Computer Sciences University of Wisconsin Madison, USA MLSS June 17, 2014 1 / 123 Outline Graphical Models Probabilistic Inference Directed vs.
CSE 527 Autumn Lectures 8-9 (& part of 10) Motifs: Representation & Discovery
 CSE 527 Autumn 2006 Lectures 8-9 (& part of 10) Motifs: Representation & Discovery 1 DNA Binding Proteins A variety of DNA binding proteins ( transcription factors ; a significant fraction, perhaps 5-10%,
CSE 527 Autumn 2006 Lectures 8-9 (& part of 10) Motifs: Representation & Discovery 1 DNA Binding Proteins A variety of DNA binding proteins ( transcription factors ; a significant fraction, perhaps 5-10%,
BAYESIAN METHODS FOR VARIABLE SELECTION WITH APPLICATIONS TO HIGH-DIMENSIONAL DATA
 BAYESIAN METHODS FOR VARIABLE SELECTION WITH APPLICATIONS TO HIGH-DIMENSIONAL DATA Intro: Course Outline and Brief Intro to Marina Vannucci Rice University, USA PASI-CIMAT 04/28-30/2010 Marina Vannucci
BAYESIAN METHODS FOR VARIABLE SELECTION WITH APPLICATIONS TO HIGH-DIMENSIONAL DATA Intro: Course Outline and Brief Intro to Marina Vannucci Rice University, USA PASI-CIMAT 04/28-30/2010 Marina Vannucci
Probabilistic Graphical Models Lecture 17: Markov chain Monte Carlo
 Probabilistic Graphical Models Lecture 17: Markov chain Monte Carlo Andrew Gordon Wilson www.cs.cmu.edu/~andrewgw Carnegie Mellon University March 18, 2015 1 / 45 Resources and Attribution Image credits,
Probabilistic Graphical Models Lecture 17: Markov chain Monte Carlo Andrew Gordon Wilson www.cs.cmu.edu/~andrewgw Carnegie Mellon University March 18, 2015 1 / 45 Resources and Attribution Image credits,
MCMC Methods: Gibbs and Metropolis
 MCMC Methods: Gibbs and Metropolis Patrick Breheny February 28 Patrick Breheny BST 701: Bayesian Modeling in Biostatistics 1/30 Introduction As we have seen, the ability to sample from the posterior distribution
MCMC Methods: Gibbs and Metropolis Patrick Breheny February 28 Patrick Breheny BST 701: Bayesian Modeling in Biostatistics 1/30 Introduction As we have seen, the ability to sample from the posterior distribution
Reminder of some Markov Chain properties:
 Reminder of some Markov Chain properties: 1. a transition from one state to another occurs probabilistically 2. only state that matters is where you currently are (i.e. given present, future is independent
Reminder of some Markov Chain properties: 1. a transition from one state to another occurs probabilistically 2. only state that matters is where you currently are (i.e. given present, future is independent
Probabilistic Graphical Networks: Definitions and Basic Results
 This document gives a cursory overview of Probabilistic Graphical Networks. The material has been gleaned from different sources. I make no claim to original authorship of this material. Bayesian Graphical
This document gives a cursory overview of Probabilistic Graphical Networks. The material has been gleaned from different sources. I make no claim to original authorship of this material. Bayesian Graphical
Markov Chain Monte Carlo methods
 Markov Chain Monte Carlo methods By Oleg Makhnin 1 Introduction a b c M = d e f g h i 0 f(x)dx 1.1 Motivation 1.1.1 Just here Supresses numbering 1.1.2 After this 1.2 Literature 2 Method 2.1 New math As
Markov Chain Monte Carlo methods By Oleg Makhnin 1 Introduction a b c M = d e f g h i 0 f(x)dx 1.1 Motivation 1.1.1 Just here Supresses numbering 1.1.2 After this 1.2 Literature 2 Method 2.1 New math As
Probabilistic Machine Learning
 Probabilistic Machine Learning Bayesian Nets, MCMC, and more Marek Petrik 4/18/2017 Based on: P. Murphy, K. (2012). Machine Learning: A Probabilistic Perspective. Chapter 10. Conditional Independence Independent
Probabilistic Machine Learning Bayesian Nets, MCMC, and more Marek Petrik 4/18/2017 Based on: P. Murphy, K. (2012). Machine Learning: A Probabilistic Perspective. Chapter 10. Conditional Independence Independent
Markov chain Monte Carlo methods in atmospheric remote sensing
 1 / 45 Markov chain Monte Carlo methods in atmospheric remote sensing Johanna Tamminen johanna.tamminen@fmi.fi ESA Summer School on Earth System Monitoring and Modeling July 3 Aug 11, 212, Frascati July,
1 / 45 Markov chain Monte Carlo methods in atmospheric remote sensing Johanna Tamminen johanna.tamminen@fmi.fi ESA Summer School on Earth System Monitoring and Modeling July 3 Aug 11, 212, Frascati July,
Lecture 8: The Metropolis-Hastings Algorithm
 30.10.2008 What we have seen last time: Gibbs sampler Key idea: Generate a Markov chain by updating the component of (X 1,..., X p ) in turn by drawing from the full conditionals: X (t) j Two drawbacks:
30.10.2008 What we have seen last time: Gibbs sampler Key idea: Generate a Markov chain by updating the component of (X 1,..., X p ) in turn by drawing from the full conditionals: X (t) j Two drawbacks:
Principles of Bayesian Inference
 Principles of Bayesian Inference Sudipto Banerjee University of Minnesota July 20th, 2008 1 Bayesian Principles Classical statistics: model parameters are fixed and unknown. A Bayesian thinks of parameters
Principles of Bayesian Inference Sudipto Banerjee University of Minnesota July 20th, 2008 1 Bayesian Principles Classical statistics: model parameters are fixed and unknown. A Bayesian thinks of parameters
Convergence Rate of Markov Chains
 Convergence Rate of Markov Chains Will Perkins April 16, 2013 Convergence Last class we saw that if X n is an irreducible, aperiodic, positive recurrent Markov chain, then there exists a stationary distribution
Convergence Rate of Markov Chains Will Perkins April 16, 2013 Convergence Last class we saw that if X n is an irreducible, aperiodic, positive recurrent Markov chain, then there exists a stationary distribution
1 Probabilities. 1.1 Basics 1 PROBABILITIES
 1 PROBABILITIES 1 Probabilities Probability is a tricky word usually meaning the likelyhood of something occuring or how frequent something is. Obviously, if something happens frequently, then its probability
1 PROBABILITIES 1 Probabilities Probability is a tricky word usually meaning the likelyhood of something occuring or how frequent something is. Obviously, if something happens frequently, then its probability
CS 343: Artificial Intelligence
 CS 343: Artificial Intelligence Bayes Nets: Sampling Prof. Scott Niekum The University of Texas at Austin [These slides based on those of Dan Klein and Pieter Abbeel for CS188 Intro to AI at UC Berkeley.
CS 343: Artificial Intelligence Bayes Nets: Sampling Prof. Scott Niekum The University of Texas at Austin [These slides based on those of Dan Klein and Pieter Abbeel for CS188 Intro to AI at UC Berkeley.
Theory of Stochastic Processes 8. Markov chain Monte Carlo
 Theory of Stochastic Processes 8. Markov chain Monte Carlo Tomonari Sei sei@mist.i.u-tokyo.ac.jp Department of Mathematical Informatics, University of Tokyo June 8, 2017 http://www.stat.t.u-tokyo.ac.jp/~sei/lec.html
Theory of Stochastic Processes 8. Markov chain Monte Carlo Tomonari Sei sei@mist.i.u-tokyo.ac.jp Department of Mathematical Informatics, University of Tokyo June 8, 2017 http://www.stat.t.u-tokyo.ac.jp/~sei/lec.html
Sampling Methods (11/30/04)
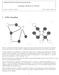 CS281A/Stat241A: Statistical Learning Theory Sampling Methods (11/30/04) Lecturer: Michael I. Jordan Scribe: Jaspal S. Sandhu 1 Gibbs Sampling Figure 1: Undirected and directed graphs, respectively, with
CS281A/Stat241A: Statistical Learning Theory Sampling Methods (11/30/04) Lecturer: Michael I. Jordan Scribe: Jaspal S. Sandhu 1 Gibbs Sampling Figure 1: Undirected and directed graphs, respectively, with
Markov Chain Monte Carlo
 Markov Chain Monte Carlo (and Bayesian Mixture Models) David M. Blei Columbia University October 14, 2014 We have discussed probabilistic modeling, and have seen how the posterior distribution is the critical
Markov Chain Monte Carlo (and Bayesian Mixture Models) David M. Blei Columbia University October 14, 2014 We have discussed probabilistic modeling, and have seen how the posterior distribution is the critical
(5) Multi-parameter models - Gibbs sampling. ST440/540: Applied Bayesian Analysis
 Summarizing a posterior Given the data and prior the posterior is determined Summarizing the posterior gives parameter estimates, intervals, and hypothesis tests Most of these computations are integrals
Summarizing a posterior Given the data and prior the posterior is determined Summarizing the posterior gives parameter estimates, intervals, and hypothesis tests Most of these computations are integrals
Introduction to Restricted Boltzmann Machines
 Introduction to Restricted Boltzmann Machines Ilija Bogunovic and Edo Collins EPFL {ilija.bogunovic,edo.collins}@epfl.ch October 13, 2014 Introduction Ingredients: 1. Probabilistic graphical models (undirected,
Introduction to Restricted Boltzmann Machines Ilija Bogunovic and Edo Collins EPFL {ilija.bogunovic,edo.collins}@epfl.ch October 13, 2014 Introduction Ingredients: 1. Probabilistic graphical models (undirected,
SAMSI Astrostatistics Tutorial. More Markov chain Monte Carlo & Demo of Mathematica software
 SAMSI Astrostatistics Tutorial More Markov chain Monte Carlo & Demo of Mathematica software Phil Gregory University of British Columbia 26 Bayesian Logical Data Analysis for the Physical Sciences Contents:
SAMSI Astrostatistics Tutorial More Markov chain Monte Carlo & Demo of Mathematica software Phil Gregory University of British Columbia 26 Bayesian Logical Data Analysis for the Physical Sciences Contents:
Monte Carlo in Bayesian Statistics
 Monte Carlo in Bayesian Statistics Matthew Thomas SAMBa - University of Bath m.l.thomas@bath.ac.uk December 4, 2014 Matthew Thomas (SAMBa) Monte Carlo in Bayesian Statistics December 4, 2014 1 / 16 Overview
Monte Carlo in Bayesian Statistics Matthew Thomas SAMBa - University of Bath m.l.thomas@bath.ac.uk December 4, 2014 Matthew Thomas (SAMBa) Monte Carlo in Bayesian Statistics December 4, 2014 1 / 16 Overview
Introduction to Machine Learning
 Introduction to Machine Learning Brown University CSCI 1950-F, Spring 2012 Prof. Erik Sudderth Lecture 25: Markov Chain Monte Carlo (MCMC) Course Review and Advanced Topics Many figures courtesy Kevin
Introduction to Machine Learning Brown University CSCI 1950-F, Spring 2012 Prof. Erik Sudderth Lecture 25: Markov Chain Monte Carlo (MCMC) Course Review and Advanced Topics Many figures courtesy Kevin
Particle-Based Approximate Inference on Graphical Model
 article-based Approimate Inference on Graphical Model Reference: robabilistic Graphical Model Ch. 2 Koller & Friedman CMU, 0-708, Fall 2009 robabilistic Graphical Models Lectures 8,9 Eric ing attern Recognition
article-based Approimate Inference on Graphical Model Reference: robabilistic Graphical Model Ch. 2 Koller & Friedman CMU, 0-708, Fall 2009 robabilistic Graphical Models Lectures 8,9 Eric ing attern Recognition
Introduction to MCMC. DB Breakfast 09/30/2011 Guozhang Wang
 Introduction to MCMC DB Breakfast 09/30/2011 Guozhang Wang Motivation: Statistical Inference Joint Distribution Sleeps Well Playground Sunny Bike Ride Pleasant dinner Productive day Posterior Estimation
Introduction to MCMC DB Breakfast 09/30/2011 Guozhang Wang Motivation: Statistical Inference Joint Distribution Sleeps Well Playground Sunny Bike Ride Pleasant dinner Productive day Posterior Estimation
Advanced Introduction to Machine Learning
 10-715 Advanced Introduction to Machine Learning Homework 3 Due Nov 12, 10.30 am Rules 1. Homework is due on the due date at 10.30 am. Please hand over your homework at the beginning of class. Please see
10-715 Advanced Introduction to Machine Learning Homework 3 Due Nov 12, 10.30 am Rules 1. Homework is due on the due date at 10.30 am. Please hand over your homework at the beginning of class. Please see
Lecture 7 and 8: Markov Chain Monte Carlo
 Lecture 7 and 8: Markov Chain Monte Carlo 4F13: Machine Learning Zoubin Ghahramani and Carl Edward Rasmussen Department of Engineering University of Cambridge http://mlg.eng.cam.ac.uk/teaching/4f13/ Ghahramani
Lecture 7 and 8: Markov Chain Monte Carlo 4F13: Machine Learning Zoubin Ghahramani and Carl Edward Rasmussen Department of Engineering University of Cambridge http://mlg.eng.cam.ac.uk/teaching/4f13/ Ghahramani
DNA Binding Proteins CSE 527 Autumn 2007
 DNA Binding Proteins CSE 527 Autumn 2007 A variety of DNA binding proteins ( transcription factors ; a significant fraction, perhaps 5-10%, of all human proteins) modulate transcription of protein coding
DNA Binding Proteins CSE 527 Autumn 2007 A variety of DNA binding proteins ( transcription factors ; a significant fraction, perhaps 5-10%, of all human proteins) modulate transcription of protein coding
Learning Bayesian Networks for Biomedical Data
 Learning Bayesian Networks for Biomedical Data Faming Liang (Texas A&M University ) Liang, F. and Zhang, J. (2009) Learning Bayesian Networks for Discrete Data. Computational Statistics and Data Analysis,
Learning Bayesian Networks for Biomedical Data Faming Liang (Texas A&M University ) Liang, F. and Zhang, J. (2009) Learning Bayesian Networks for Discrete Data. Computational Statistics and Data Analysis,
ELEC633: Graphical Models
 ELEC633: Graphical Models Tahira isa Saleem Scribe from 7 October 2008 References: Casella and George Exploring the Gibbs sampler (1992) Chib and Greenberg Understanding the Metropolis-Hastings algorithm
ELEC633: Graphical Models Tahira isa Saleem Scribe from 7 October 2008 References: Casella and George Exploring the Gibbs sampler (1992) Chib and Greenberg Understanding the Metropolis-Hastings algorithm
Reinforcement Learning: Part 3 Evolution
 1 Reinforcement Learning: Part 3 Evolution Chris Watkins Department of Computer Science Royal Holloway, University of London July 27, 2015 2 Cross-entropy method for TSP Simple genetic style methods can
1 Reinforcement Learning: Part 3 Evolution Chris Watkins Department of Computer Science Royal Holloway, University of London July 27, 2015 2 Cross-entropy method for TSP Simple genetic style methods can
I. Bayesian econometrics
 I. Bayesian econometrics A. Introduction B. Bayesian inference in the univariate regression model C. Statistical decision theory D. Large sample results E. Diffuse priors F. Numerical Bayesian methods
I. Bayesian econometrics A. Introduction B. Bayesian inference in the univariate regression model C. Statistical decision theory D. Large sample results E. Diffuse priors F. Numerical Bayesian methods
