CS1820 Notes. hgupta1, kjline, smechery. April 3-April 5. output: plausible Ancestral Recombination Graph (ARG)
|
|
|
- Meredith Austin
- 5 years ago
- Views:
Transcription
1 CS1820 Notes hgupta1, kjline, smechery April 3-April 5 April 3 Notes 1 Minichiello-Durbin Algorithm input: set of sequences output: plausible Ancestral Recombination Graph (ARG) note: the optimal ARG is the one with the minimum of recombinations. 2 Definitions 0,1 are alleles, is an undefined allele sequences involved: haplotype sequences of length m over alphabet 0, 1, Let C be a sequence, C[i] is the ith symbol, 1 i m C 1 [i] C 2 [i] iff C 1 [i] = C 2 [i] or C 1 [i] = or C 2 [i] = T is the argument for the time step S T is a sample of sequences operations 1. Coalesce Rule: If there are two sequences C 1 and C 2 in S T such that for all instances of i the condition C 1 [i] C 2 [i] is true, then C 1 and C 2 coalesce into an ancestor sequence Transition: S T +1 = (S T \ {C 1, C 2 }) {C} such that C[i] = C 1 [i] when C 1 [i] and C 2 [i] and C[i] = C 2 [i] otherwise. 2. Mutation 1
2 Rule: If there exists a sequence C 1 in S T and a marker i where, for all of C 2 in (S T \ {C 1 }), we have C 2 [i] = C 1 [i] or, then we can remove the derived allele (C 1 [i]) from the population Transition: S T +1 = (S T \ C 1 ) {C }, where C [i] = C 1 [i] and C [j] = C 1 [j] for all j i 3. Recombination Rule: When the rules of mutation and coalesce do not apply, must apply recombination or pair of recombinations. Denote the recombination break point as (α, β) as meaning that it occurs between markers α and β. Picking a shared tract C 1, C 2 [α, β] from those available in S T, we aim to put recombination parent of C 1 and one recombination parent of C 2 satisfy the rule of coalescence. To do this, we must put a break point at (α 1, α), if α 1 and put a break point at (β, β + 1) if β m. Transition: From the tract {C 1, C 2 }[α, β], pick (1) a valid breakpoint (α, β), where either (α, β) = (α 1, α) or (α, β) = (β, β+1) and (2) a recombinant sequence C R, where either the C R = 1 or C R = 2. Then, S T +1 = (S T \{C R }){C 1, C 2}, where C 1[i] = C R [i] for all i α and C 1[i] = otherwise, C 2[i] = C R [i] for all i β and C 2[i] = otherwise. If both (α 1, α) and (β, β +1) are valid breakpoints, we must put the second recombination (taking us to S T +2 ), on an appropriate ancestor of C 1 or C 2. 3 The Algorithm The goal of the algorithm is to find a single ancestral sequence of a sample of sequences. 1. The algorithm starts at T=1. 2. For each iteration S i (where i = T ) from 1 i m, apply the Coalesce, Mutation, and Recombination rules to S i 3. Stop when S i contains one sequence. 2
3 April 5 Notes Chapter 4: Hidden Markov Models - The Learning Problem and Algorithm Three Fundamental Problems A Hidden Markov Model has the following inputs The observation sequence: Θ = θ 1 θ 2...θ T The Model: λ = (A, B, π) Problem 1: the Equation problem or Model Scoring problem. Given: Θ, λ Compute: P (Θ λ), the probability of observing the observation given the word. Problem 2: The Decoding Problem or Uncovering the Hidden Part Given: Θ, λ Compute: the sequence of sites q 1, q 2,..., q T that optimally explains the observed sequences The Viturbi Algorithm (Maximum Likelihood) is used for this problem. Problem 3: The Learning problem or Training problem Given: Θ Compute: the parameters of a model λ = (A, B, π) that maximize P (Θ λ) Definition of a Hidden Markov Model An HMM has 5 elements: N, M, A, B, π N 1. N = of states s = s 1, s 2,..., s N 2. M = of distinct observation symbols per state. 3. A = transition probability distribution, A = a ij, a ij = P [q t+1 = s j q i = s i ], 1 i, j N 4. B = observation symbols probability distribution, B = b j (k) b j (k) = P [v k at time t q t = s j ], 1 j N, 1 k M 5. π N = The initial state distribution. π = π i, π i = P [q i = s i ], 1 i N We need a number of variables: α: The forward variable α t (i) = P (θ 1 θ 2...θ T, q T = s i ) Note: θ 1 θ 2 is the prefix of the observation sequences. The backward variable: B t (i) = P (θ t+1 θ t+2...θ T, q t = s i ) θ t+1 θ t+2 is the suffix of the observation sequence. 3
4 Delta δ = δ t (i) = MAX P [q 1 q 2...q t = s i, θ 1 θ 2...θ T Solution to Problem 3: We want to construct parameters of the model λ = (A, Bπ) to maximize the probability of observing the sequence Θ in λ. There is no analytical exact solution (like for problems 1 and 2) We are going to construct a λ = (A, B π that is a local max pf P(Θ λ ) An Iterative algorithm: ζ t (i, j) = probability of being in state i at time t and transitioning to state s j at time t + 1. ζ t (i, j) = P (q t = s i, q t+1 = s j Θ, λ) ζ t (i, j) = αt(i)aijbj(θt+1)βt+1(j) P (θ λ) ζ t (i, j) = N α t(i)a ijbj(θ t+1)β t+1(j) N i =1 j =1 αt(i )a i j b j (θ t+1)β t(j ) β t (j ) is the normalizing factor Gamma: γ t = the probability of being in state s i at the time t given the observation sequence. γ t (i) = N j=1 ζ t(i, j) The expected number of times that state s i is visited: T 1 t=1 γ t(i) The expected number of transitions from s i : T 1 t=1 ζ t(i, j)=the expected number of transitions from s i to s j Suppose that we have a model λ = (A, B, π) Construct a new model λ = (Ā, B, π) π i =expected frequency (n number of times) in state s i at t=1 = γ t (i) A. a ij a ij = expected number of transitions from s i to s j = expected number of transitions from s j B. b j (k) b j (k) = expected of times in state observing symbol v k expected of times in state s j T γ t=1,θ = k =v t(j) k T γt(j) t=1 T 1 t=1 ζ T (i,j) T 1 t=1 γt(i) 4
5 λ = (Ā, B, π) 5
6 Figure 1: An example of a full ARG 6
7 Figure 2: First part of a partial recombination Figure 3: Second part of a partial recombination 7
Hidden Markov Models. Three classic HMM problems
 An Introduction to Bioinformatics Algorithms www.bioalgorithms.info Hidden Markov Models Slides revised and adapted to Computational Biology IST 2015/2016 Ana Teresa Freitas Three classic HMM problems
An Introduction to Bioinformatics Algorithms www.bioalgorithms.info Hidden Markov Models Slides revised and adapted to Computational Biology IST 2015/2016 Ana Teresa Freitas Three classic HMM problems
Implementation of EM algorithm in HMM training. Jens Lagergren
 Implementation of EM algorithm in HMM training Jens Lagergren April 22, 2009 EM Algorithms: An expectation-maximization (EM) algorithm is used in statistics for finding maximum likelihood estimates of
Implementation of EM algorithm in HMM training Jens Lagergren April 22, 2009 EM Algorithms: An expectation-maximization (EM) algorithm is used in statistics for finding maximum likelihood estimates of
Hidden Markov Model. Ying Wu. Electrical Engineering and Computer Science Northwestern University Evanston, IL 60208
 Hidden Markov Model Ying Wu Electrical Engineering and Computer Science Northwestern University Evanston, IL 60208 http://www.eecs.northwestern.edu/~yingwu 1/19 Outline Example: Hidden Coin Tossing Hidden
Hidden Markov Model Ying Wu Electrical Engineering and Computer Science Northwestern University Evanston, IL 60208 http://www.eecs.northwestern.edu/~yingwu 1/19 Outline Example: Hidden Coin Tossing Hidden
Basic math for biology
 Basic math for biology Lei Li Florida State University, Feb 6, 2002 The EM algorithm: setup Parametric models: {P θ }. Data: full data (Y, X); partial data Y. Missing data: X. Likelihood and maximum likelihood
Basic math for biology Lei Li Florida State University, Feb 6, 2002 The EM algorithm: setup Parametric models: {P θ }. Data: full data (Y, X); partial data Y. Missing data: X. Likelihood and maximum likelihood
CISC 889 Bioinformatics (Spring 2004) Hidden Markov Models (II)
 CISC 889 Bioinformatics (Spring 24) Hidden Markov Models (II) a. Likelihood: forward algorithm b. Decoding: Viterbi algorithm c. Model building: Baum-Welch algorithm Viterbi training Hidden Markov models
CISC 889 Bioinformatics (Spring 24) Hidden Markov Models (II) a. Likelihood: forward algorithm b. Decoding: Viterbi algorithm c. Model building: Baum-Welch algorithm Viterbi training Hidden Markov models
Hidden Markov Models
 Hidden Markov Models Slides revised and adapted to Bioinformática 55 Engª Biomédica/IST 2005 Ana Teresa Freitas Forward Algorithm For Markov chains we calculate the probability of a sequence, P(x) How
Hidden Markov Models Slides revised and adapted to Bioinformática 55 Engª Biomédica/IST 2005 Ana Teresa Freitas Forward Algorithm For Markov chains we calculate the probability of a sequence, P(x) How
Parametric Models Part III: Hidden Markov Models
 Parametric Models Part III: Hidden Markov Models Selim Aksoy Department of Computer Engineering Bilkent University saksoy@cs.bilkent.edu.tr CS 551, Spring 2014 CS 551, Spring 2014 c 2014, Selim Aksoy (Bilkent
Parametric Models Part III: Hidden Markov Models Selim Aksoy Department of Computer Engineering Bilkent University saksoy@cs.bilkent.edu.tr CS 551, Spring 2014 CS 551, Spring 2014 c 2014, Selim Aksoy (Bilkent
Machine Learning for natural language processing
 Machine Learning for natural language processing Hidden Markov Models Laura Kallmeyer Heinrich-Heine-Universität Düsseldorf Summer 2016 1 / 33 Introduction So far, we have classified texts/observations
Machine Learning for natural language processing Hidden Markov Models Laura Kallmeyer Heinrich-Heine-Universität Düsseldorf Summer 2016 1 / 33 Introduction So far, we have classified texts/observations
Computational Genomics and Molecular Biology, Fall
 Computational Genomics and Molecular Biology, Fall 2011 1 HMM Lecture Notes Dannie Durand and Rose Hoberman October 11th 1 Hidden Markov Models In the last few lectures, we have focussed on three problems
Computational Genomics and Molecular Biology, Fall 2011 1 HMM Lecture Notes Dannie Durand and Rose Hoberman October 11th 1 Hidden Markov Models In the last few lectures, we have focussed on three problems
Computational Biology Lecture #3: Probability and Statistics. Bud Mishra Professor of Computer Science, Mathematics, & Cell Biology Sept
 Computational Biology Lecture #3: Probability and Statistics Bud Mishra Professor of Computer Science, Mathematics, & Cell Biology Sept 26 2005 L2-1 Basic Probabilities L2-2 1 Random Variables L2-3 Examples
Computational Biology Lecture #3: Probability and Statistics Bud Mishra Professor of Computer Science, Mathematics, & Cell Biology Sept 26 2005 L2-1 Basic Probabilities L2-2 1 Random Variables L2-3 Examples
CS 7180: Behavioral Modeling and Decision- making in AI
 CS 7180: Behavioral Modeling and Decision- making in AI Learning Probabilistic Graphical Models Prof. Amy Sliva October 31, 2012 Hidden Markov model Stochastic system represented by three matrices N =
CS 7180: Behavioral Modeling and Decision- making in AI Learning Probabilistic Graphical Models Prof. Amy Sliva October 31, 2012 Hidden Markov model Stochastic system represented by three matrices N =
Lecture 9. Intro to Hidden Markov Models (finish up)
 Lecture 9 Intro to Hidden Markov Models (finish up) Review Structure Number of states Q 1.. Q N M output symbols Parameters: Transition probability matrix a ij Emission probabilities b i (a), which is
Lecture 9 Intro to Hidden Markov Models (finish up) Review Structure Number of states Q 1.. Q N M output symbols Parameters: Transition probability matrix a ij Emission probabilities b i (a), which is
Lab 3: Practical Hidden Markov Models (HMM)
 Advanced Topics in Bioinformatics Lab 3: Practical Hidden Markov Models () Maoying, Wu Department of Bioinformatics & Biostatistics Shanghai Jiao Tong University November 27, 2014 Hidden Markov Models
Advanced Topics in Bioinformatics Lab 3: Practical Hidden Markov Models () Maoying, Wu Department of Bioinformatics & Biostatistics Shanghai Jiao Tong University November 27, 2014 Hidden Markov Models
Hidden Markov Models The three basic HMM problems (note: change in notation) Mitch Marcus CSE 391
 Hidden Markov Models The three basic HMM problems (note: change in notation) Mitch Marcus CSE 391 Parameters of an HMM States: A set of states S=s 1, s n Transition probabilities: A= a 1,1, a 1,2,, a n,n
Hidden Markov Models The three basic HMM problems (note: change in notation) Mitch Marcus CSE 391 Parameters of an HMM States: A set of states S=s 1, s n Transition probabilities: A= a 1,1, a 1,2,, a n,n
An Introduction to Bioinformatics Algorithms Hidden Markov Models
 Hidden Markov Models Outline 1. CG-Islands 2. The Fair Bet Casino 3. Hidden Markov Model 4. Decoding Algorithm 5. Forward-Backward Algorithm 6. Profile HMMs 7. HMM Parameter Estimation 8. Viterbi Training
Hidden Markov Models Outline 1. CG-Islands 2. The Fair Bet Casino 3. Hidden Markov Model 4. Decoding Algorithm 5. Forward-Backward Algorithm 6. Profile HMMs 7. HMM Parameter Estimation 8. Viterbi Training
Hidden Markov Models Hamid R. Rabiee
 Hidden Markov Models Hamid R. Rabiee 1 Hidden Markov Models (HMMs) In the previous slides, we have seen that in many cases the underlying behavior of nature could be modeled as a Markov process. However
Hidden Markov Models Hamid R. Rabiee 1 Hidden Markov Models (HMMs) In the previous slides, we have seen that in many cases the underlying behavior of nature could be modeled as a Markov process. However
Hidden Markov Models
 Hidden Markov Models Outline 1. CG-Islands 2. The Fair Bet Casino 3. Hidden Markov Model 4. Decoding Algorithm 5. Forward-Backward Algorithm 6. Profile HMMs 7. HMM Parameter Estimation 8. Viterbi Training
Hidden Markov Models Outline 1. CG-Islands 2. The Fair Bet Casino 3. Hidden Markov Model 4. Decoding Algorithm 5. Forward-Backward Algorithm 6. Profile HMMs 7. HMM Parameter Estimation 8. Viterbi Training
11.3 Decoding Algorithm
 11.3 Decoding Algorithm 393 For convenience, we have introduced π 0 and π n+1 as the fictitious initial and terminal states begin and end. This model defines the probability P(x π) for a given sequence
11.3 Decoding Algorithm 393 For convenience, we have introduced π 0 and π n+1 as the fictitious initial and terminal states begin and end. This model defines the probability P(x π) for a given sequence
Data-Intensive Computing with MapReduce
 Data-Intensive Computing with MapReduce Session 8: Sequence Labeling Jimmy Lin University of Maryland Thursday, March 14, 2013 This work is licensed under a Creative Commons Attribution-Noncommercial-Share
Data-Intensive Computing with MapReduce Session 8: Sequence Labeling Jimmy Lin University of Maryland Thursday, March 14, 2013 This work is licensed under a Creative Commons Attribution-Noncommercial-Share
Lecture 4: Hidden Markov Models: An Introduction to Dynamic Decision Making. November 11, 2010
 Hidden Lecture 4: Hidden : An Introduction to Dynamic Decision Making November 11, 2010 Special Meeting 1/26 Markov Model Hidden When a dynamical system is probabilistic it may be determined by the transition
Hidden Lecture 4: Hidden : An Introduction to Dynamic Decision Making November 11, 2010 Special Meeting 1/26 Markov Model Hidden When a dynamical system is probabilistic it may be determined by the transition
Dynamic Approaches: The Hidden Markov Model
 Dynamic Approaches: The Hidden Markov Model Davide Bacciu Dipartimento di Informatica Università di Pisa bacciu@di.unipi.it Machine Learning: Neural Networks and Advanced Models (AA2) Inference as Message
Dynamic Approaches: The Hidden Markov Model Davide Bacciu Dipartimento di Informatica Università di Pisa bacciu@di.unipi.it Machine Learning: Neural Networks and Advanced Models (AA2) Inference as Message
Hidden Markov Models. x 1 x 2 x 3 x K
 Hidden Markov Models 1 1 1 1 2 2 2 2 K K K K x 1 x 2 x 3 x K Viterbi, Forward, Backward VITERBI FORWARD BACKWARD Initialization: V 0 (0) = 1 V k (0) = 0, for all k > 0 Initialization: f 0 (0) = 1 f k (0)
Hidden Markov Models 1 1 1 1 2 2 2 2 K K K K x 1 x 2 x 3 x K Viterbi, Forward, Backward VITERBI FORWARD BACKWARD Initialization: V 0 (0) = 1 V k (0) = 0, for all k > 0 Initialization: f 0 (0) = 1 f k (0)
p(d θ ) l(θ ) 1.2 x x x
 p(d θ ).2 x 0-7 0.8 x 0-7 0.4 x 0-7 l(θ ) -20-40 -60-80 -00 2 3 4 5 6 7 θ ˆ 2 3 4 5 6 7 θ ˆ 2 3 4 5 6 7 θ θ x FIGURE 3.. The top graph shows several training points in one dimension, known or assumed to
p(d θ ).2 x 0-7 0.8 x 0-7 0.4 x 0-7 l(θ ) -20-40 -60-80 -00 2 3 4 5 6 7 θ ˆ 2 3 4 5 6 7 θ ˆ 2 3 4 5 6 7 θ θ x FIGURE 3.. The top graph shows several training points in one dimension, known or assumed to
Hidden Markov Models. Ivan Gesteira Costa Filho IZKF Research Group Bioinformatics RWTH Aachen Adapted from:
 Hidden Markov Models Ivan Gesteira Costa Filho IZKF Research Group Bioinformatics RWTH Aachen Adapted from: www.ioalgorithms.info Outline CG-islands The Fair Bet Casino Hidden Markov Model Decoding Algorithm
Hidden Markov Models Ivan Gesteira Costa Filho IZKF Research Group Bioinformatics RWTH Aachen Adapted from: www.ioalgorithms.info Outline CG-islands The Fair Bet Casino Hidden Markov Model Decoding Algorithm
Hidden Markov Models. Terminology, Representation and Basic Problems
 Hidden Markov Models Terminology, Representation and Basic Problems Data analysis? Machine learning? In bioinformatics, we analyze a lot of (sequential) data (biological sequences) to learn unknown parameters
Hidden Markov Models Terminology, Representation and Basic Problems Data analysis? Machine learning? In bioinformatics, we analyze a lot of (sequential) data (biological sequences) to learn unknown parameters
Hidden Markov Models. By Parisa Abedi. Slides courtesy: Eric Xing
 Hidden Markov Models By Parisa Abedi Slides courtesy: Eric Xing i.i.d to sequential data So far we assumed independent, identically distributed data Sequential (non i.i.d.) data Time-series data E.g. Speech
Hidden Markov Models By Parisa Abedi Slides courtesy: Eric Xing i.i.d to sequential data So far we assumed independent, identically distributed data Sequential (non i.i.d.) data Time-series data E.g. Speech
Hidden Markov Modelling
 Hidden Markov Modelling Introduction Problem formulation Forward-Backward algorithm Viterbi search Baum-Welch parameter estimation Other considerations Multiple observation sequences Phone-based models
Hidden Markov Modelling Introduction Problem formulation Forward-Backward algorithm Viterbi search Baum-Welch parameter estimation Other considerations Multiple observation sequences Phone-based models
Note Set 5: Hidden Markov Models
 Note Set 5: Hidden Markov Models Probabilistic Learning: Theory and Algorithms, CS 274A, Winter 2016 1 Hidden Markov Models (HMMs) 1.1 Introduction Consider observed data vectors x t that are d-dimensional
Note Set 5: Hidden Markov Models Probabilistic Learning: Theory and Algorithms, CS 274A, Winter 2016 1 Hidden Markov Models (HMMs) 1.1 Introduction Consider observed data vectors x t that are d-dimensional
Conditional Random Fields
 Conditional Random Fields Micha Elsner February 14, 2013 2 Sums of logs Issue: computing α forward probabilities can undeflow Normally we d fix this using logs But α requires a sum of probabilities Not
Conditional Random Fields Micha Elsner February 14, 2013 2 Sums of logs Issue: computing α forward probabilities can undeflow Normally we d fix this using logs But α requires a sum of probabilities Not
An Introduction to Bioinformatics Algorithms Hidden Markov Models
 Hidden Markov Models Hidden Markov Models Outline CG-islands The Fair Bet Casino Hidden Markov Model Decoding Algorithm Forward-Backward Algorithm Profile HMMs HMM Parameter Estimation Viterbi training
Hidden Markov Models Hidden Markov Models Outline CG-islands The Fair Bet Casino Hidden Markov Model Decoding Algorithm Forward-Backward Algorithm Profile HMMs HMM Parameter Estimation Viterbi training
Hidden Markov Models. Terminology and Basic Algorithms
 Hidden Markov Models Terminology and Basic Algorithms The next two weeks Hidden Markov models (HMMs): Wed 9/11: Terminology and basic algorithms Mon 14/11: Implementing the basic algorithms Wed 16/11:
Hidden Markov Models Terminology and Basic Algorithms The next two weeks Hidden Markov models (HMMs): Wed 9/11: Terminology and basic algorithms Mon 14/11: Implementing the basic algorithms Wed 16/11:
Hidden Markov Models. Aarti Singh Slides courtesy: Eric Xing. Machine Learning / Nov 8, 2010
 Hidden Markov Models Aarti Singh Slides courtesy: Eric Xing Machine Learning 10-701/15-781 Nov 8, 2010 i.i.d to sequential data So far we assumed independent, identically distributed data Sequential data
Hidden Markov Models Aarti Singh Slides courtesy: Eric Xing Machine Learning 10-701/15-781 Nov 8, 2010 i.i.d to sequential data So far we assumed independent, identically distributed data Sequential data
Hidden Markov Models
 CS769 Spring 2010 Advanced Natural Language Processing Hidden Markov Models Lecturer: Xiaojin Zhu jerryzhu@cs.wisc.edu 1 Part-of-Speech Tagging The goal of Part-of-Speech (POS) tagging is to label each
CS769 Spring 2010 Advanced Natural Language Processing Hidden Markov Models Lecturer: Xiaojin Zhu jerryzhu@cs.wisc.edu 1 Part-of-Speech Tagging The goal of Part-of-Speech (POS) tagging is to label each
A brief introduction to Conditional Random Fields
 A brief introduction to Conditional Random Fields Mark Johnson Macquarie University April, 2005, updated October 2010 1 Talk outline Graphical models Maximum likelihood and maximum conditional likelihood
A brief introduction to Conditional Random Fields Mark Johnson Macquarie University April, 2005, updated October 2010 1 Talk outline Graphical models Maximum likelihood and maximum conditional likelihood
Introduction to Machine Learning CMU-10701
 Introduction to Machine Learning CMU-10701 Hidden Markov Models Barnabás Póczos & Aarti Singh Slides courtesy: Eric Xing i.i.d to sequential data So far we assumed independent, identically distributed
Introduction to Machine Learning CMU-10701 Hidden Markov Models Barnabás Póczos & Aarti Singh Slides courtesy: Eric Xing i.i.d to sequential data So far we assumed independent, identically distributed
Log-Linear Models, MEMMs, and CRFs
 Log-Linear Models, MEMMs, and CRFs Michael Collins 1 Notation Throughout this note I ll use underline to denote vectors. For example, w R d will be a vector with components w 1, w 2,... w d. We use expx
Log-Linear Models, MEMMs, and CRFs Michael Collins 1 Notation Throughout this note I ll use underline to denote vectors. For example, w R d will be a vector with components w 1, w 2,... w d. We use expx
Hidden Markov Models. Terminology and Basic Algorithms
 Hidden Markov Models Terminology and Basic Algorithms What is machine learning? From http://en.wikipedia.org/wiki/machine_learning Machine learning, a branch of artificial intelligence, is about the construction
Hidden Markov Models Terminology and Basic Algorithms What is machine learning? From http://en.wikipedia.org/wiki/machine_learning Machine learning, a branch of artificial intelligence, is about the construction
Sequence Modelling with Features: Linear-Chain Conditional Random Fields. COMP-599 Oct 6, 2015
 Sequence Modelling with Features: Linear-Chain Conditional Random Fields COMP-599 Oct 6, 2015 Announcement A2 is out. Due Oct 20 at 1pm. 2 Outline Hidden Markov models: shortcomings Generative vs. discriminative
Sequence Modelling with Features: Linear-Chain Conditional Random Fields COMP-599 Oct 6, 2015 Announcement A2 is out. Due Oct 20 at 1pm. 2 Outline Hidden Markov models: shortcomings Generative vs. discriminative
CS 136a Lecture 7 Speech Recognition Architecture: Training models with the Forward backward algorithm
 + September13, 2016 Professor Meteer CS 136a Lecture 7 Speech Recognition Architecture: Training models with the Forward backward algorithm Thanks to Dan Jurafsky for these slides + ASR components n Feature
+ September13, 2016 Professor Meteer CS 136a Lecture 7 Speech Recognition Architecture: Training models with the Forward backward algorithm Thanks to Dan Jurafsky for these slides + ASR components n Feature
Steven L. Scott. Presented by Ahmet Engin Ural
 Steven L. Scott Presented by Ahmet Engin Ural Overview of HMM Evaluating likelihoods The Likelihood Recursion The Forward-Backward Recursion Sampling HMM DG and FB samplers Autocovariance of samplers Some
Steven L. Scott Presented by Ahmet Engin Ural Overview of HMM Evaluating likelihoods The Likelihood Recursion The Forward-Backward Recursion Sampling HMM DG and FB samplers Autocovariance of samplers Some
Alignment Algorithms. Alignment Algorithms
 Midterm Results Big improvement over scores from the previous two years. Since this class grade is based on the previous years curve, that means this class will get higher grades than the previous years.
Midterm Results Big improvement over scores from the previous two years. Since this class grade is based on the previous years curve, that means this class will get higher grades than the previous years.
Counting All Possible Ancestral Configurations of Sample Sequences in Population Genetics
 1 Counting All Possible Ancestral Configurations of Sample Sequences in Population Genetics Yun S. Song, Rune Lyngsø, and Jotun Hein Abstract Given a set D of input sequences, a genealogy for D can be
1 Counting All Possible Ancestral Configurations of Sample Sequences in Population Genetics Yun S. Song, Rune Lyngsø, and Jotun Hein Abstract Given a set D of input sequences, a genealogy for D can be
CSCE 471/871 Lecture 3: Markov Chains and
 and and 1 / 26 sscott@cse.unl.edu 2 / 26 Outline and chains models (s) Formal definition Finding most probable state path (Viterbi algorithm) Forward and backward algorithms State sequence known State
and and 1 / 26 sscott@cse.unl.edu 2 / 26 Outline and chains models (s) Formal definition Finding most probable state path (Viterbi algorithm) Forward and backward algorithms State sequence known State
O 3 O 4 O 5. q 3. q 4. Transition
 Hidden Markov Models Hidden Markov models (HMM) were developed in the early part of the 1970 s and at that time mostly applied in the area of computerized speech recognition. They are first described in
Hidden Markov Models Hidden Markov models (HMM) were developed in the early part of the 1970 s and at that time mostly applied in the area of computerized speech recognition. They are first described in
Conditional Random Field
 Introduction Linear-Chain General Specific Implementations Conclusions Corso di Elaborazione del Linguaggio Naturale Pisa, May, 2011 Introduction Linear-Chain General Specific Implementations Conclusions
Introduction Linear-Chain General Specific Implementations Conclusions Corso di Elaborazione del Linguaggio Naturale Pisa, May, 2011 Introduction Linear-Chain General Specific Implementations Conclusions
HMM: Parameter Estimation
 I529: Machine Learning in Bioinformatics (Spring 2017) HMM: Parameter Estimation Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Content Review HMM: three problems
I529: Machine Learning in Bioinformatics (Spring 2017) HMM: Parameter Estimation Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Content Review HMM: three problems
Hidden Markov Models
 Hidden Markov Models Outline CG-islands The Fair Bet Casino Hidden Markov Model Decoding Algorithm Forward-Backward Algorithm Profile HMMs HMM Parameter Estimation Viterbi training Baum-Welch algorithm
Hidden Markov Models Outline CG-islands The Fair Bet Casino Hidden Markov Model Decoding Algorithm Forward-Backward Algorithm Profile HMMs HMM Parameter Estimation Viterbi training Baum-Welch algorithm
10. Hidden Markov Models (HMM) for Speech Processing. (some slides taken from Glass and Zue course)
 10. Hidden Markov Models (HMM) for Speech Processing (some slides taken from Glass and Zue course) Definition of an HMM The HMM are powerful statistical methods to characterize the observed samples of
10. Hidden Markov Models (HMM) for Speech Processing (some slides taken from Glass and Zue course) Definition of an HMM The HMM are powerful statistical methods to characterize the observed samples of
Stephen Scott.
 1 / 27 sscott@cse.unl.edu 2 / 27 Useful for modeling/making predictions on sequential data E.g., biological sequences, text, series of sounds/spoken words Will return to graphical models that are generative
1 / 27 sscott@cse.unl.edu 2 / 27 Useful for modeling/making predictions on sequential data E.g., biological sequences, text, series of sounds/spoken words Will return to graphical models that are generative
Conditional Random Fields and beyond DANIEL KHASHABI CS 546 UIUC, 2013
 Conditional Random Fields and beyond DANIEL KHASHABI CS 546 UIUC, 2013 Outline Modeling Inference Training Applications Outline Modeling Problem definition Discriminative vs. Generative Chain CRF General
Conditional Random Fields and beyond DANIEL KHASHABI CS 546 UIUC, 2013 Outline Modeling Inference Training Applications Outline Modeling Problem definition Discriminative vs. Generative Chain CRF General
1 What is a hidden Markov model?
 1 What is a hidden Markov model? Consider a Markov chain {X k }, where k is a non-negative integer. Suppose {X k } embedded in signals corrupted by some noise. Indeed, {X k } is hidden due to noise and
1 What is a hidden Markov model? Consider a Markov chain {X k }, where k is a non-negative integer. Suppose {X k } embedded in signals corrupted by some noise. Indeed, {X k } is hidden due to noise and
Hidden Markov Models (I)
 GLOBEX Bioinformatics (Summer 2015) Hidden Markov Models (I) a. The model b. The decoding: Viterbi algorithm Hidden Markov models A Markov chain of states At each state, there are a set of possible observables
GLOBEX Bioinformatics (Summer 2015) Hidden Markov Models (I) a. The model b. The decoding: Viterbi algorithm Hidden Markov models A Markov chain of states At each state, there are a set of possible observables
Hidden Markov Model and Speech Recognition
 1 Dec,2006 Outline Introduction 1 Introduction 2 3 4 5 Introduction What is Speech Recognition? Understanding what is being said Mapping speech data to textual information Speech Recognition is indeed
1 Dec,2006 Outline Introduction 1 Introduction 2 3 4 5 Introduction What is Speech Recognition? Understanding what is being said Mapping speech data to textual information Speech Recognition is indeed
Hidden Markov models in population genetics and evolutionary biology
 Hidden Markov models in population genetics and evolutionary biology Gerton Lunter Wellcome Trust Centre for Human Genetics Oxford, UK April 29, 2013 Topics for today Markov chains Hidden Markov models
Hidden Markov models in population genetics and evolutionary biology Gerton Lunter Wellcome Trust Centre for Human Genetics Oxford, UK April 29, 2013 Topics for today Markov chains Hidden Markov models
Automatic Speech Recognition (CS753)
 Automatic Speech Recognition (CS753) Lecture 6: Hidden Markov Models (Part II) Instructor: Preethi Jyothi Aug 10, 2017 Recall: Computing Likelihood Problem 1 (Likelihood): Given an HMM l =(A, B) and an
Automatic Speech Recognition (CS753) Lecture 6: Hidden Markov Models (Part II) Instructor: Preethi Jyothi Aug 10, 2017 Recall: Computing Likelihood Problem 1 (Likelihood): Given an HMM l =(A, B) and an
2 : Directed GMs: Bayesian Networks
 10-708: Probabilistic Graphical Models 10-708, Spring 2017 2 : Directed GMs: Bayesian Networks Lecturer: Eric P. Xing Scribes: Jayanth Koushik, Hiroaki Hayashi, Christian Perez Topic: Directed GMs 1 Types
10-708: Probabilistic Graphical Models 10-708, Spring 2017 2 : Directed GMs: Bayesian Networks Lecturer: Eric P. Xing Scribes: Jayanth Koushik, Hiroaki Hayashi, Christian Perez Topic: Directed GMs 1 Types
6.864: Lecture 5 (September 22nd, 2005) The EM Algorithm
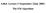 6.864: Lecture 5 (September 22nd, 2005) The EM Algorithm Overview The EM algorithm in general form The EM algorithm for hidden markov models (brute force) The EM algorithm for hidden markov models (dynamic
6.864: Lecture 5 (September 22nd, 2005) The EM Algorithm Overview The EM algorithm in general form The EM algorithm for hidden markov models (brute force) The EM algorithm for hidden markov models (dynamic
Giri Narasimhan. CAP 5510: Introduction to Bioinformatics. ECS 254; Phone: x3748
 CAP 5510: Introduction to Bioinformatics Giri Narasimhan ECS 254; Phone: x3748 giri@cis.fiu.edu www.cis.fiu.edu/~giri/teach/bioinfs07.html 2/14/07 CAP5510 1 CpG Islands Regions in DNA sequences with increased
CAP 5510: Introduction to Bioinformatics Giri Narasimhan ECS 254; Phone: x3748 giri@cis.fiu.edu www.cis.fiu.edu/~giri/teach/bioinfs07.html 2/14/07 CAP5510 1 CpG Islands Regions in DNA sequences with increased
Example: The Dishonest Casino. Hidden Markov Models. Question # 1 Evaluation. The dishonest casino model. Question # 3 Learning. Question # 2 Decoding
 Example: The Dishonest Casino Hidden Markov Models Durbin and Eddy, chapter 3 Game:. You bet $. You roll 3. Casino player rolls 4. Highest number wins $ The casino has two dice: Fair die P() = P() = P(3)
Example: The Dishonest Casino Hidden Markov Models Durbin and Eddy, chapter 3 Game:. You bet $. You roll 3. Casino player rolls 4. Highest number wins $ The casino has two dice: Fair die P() = P() = P(3)
Machine Learning & Data Mining Caltech CS/CNS/EE 155 Hidden Markov Models Last Updated: Feb 7th, 2017
 1 Introduction Let x = (x 1,..., x M ) denote a sequence (e.g. a sequence of words), and let y = (y 1,..., y M ) denote a corresponding hidden sequence that we believe explains or influences x somehow
1 Introduction Let x = (x 1,..., x M ) denote a sequence (e.g. a sequence of words), and let y = (y 1,..., y M ) denote a corresponding hidden sequence that we believe explains or influences x somehow
4 : Exact Inference: Variable Elimination
 10-708: Probabilistic Graphical Models 10-708, Spring 2014 4 : Exact Inference: Variable Elimination Lecturer: Eric P. ing Scribes: Soumya Batra, Pradeep Dasigi, Manzil Zaheer 1 Probabilistic Inference
10-708: Probabilistic Graphical Models 10-708, Spring 2014 4 : Exact Inference: Variable Elimination Lecturer: Eric P. ing Scribes: Soumya Batra, Pradeep Dasigi, Manzil Zaheer 1 Probabilistic Inference
Hidden Markov Models: Maxing and Summing
 Hidden Markov Models: Maxing and Summing Introduction to Natural Language Processing Computer Science 585 Fall 2009 University of Massachusetts Amherst David Smith 1 Markov vs. Hidden Markov Models Fed
Hidden Markov Models: Maxing and Summing Introduction to Natural Language Processing Computer Science 585 Fall 2009 University of Massachusetts Amherst David Smith 1 Markov vs. Hidden Markov Models Fed
CRF for human beings
 CRF for human beings Arne Skjærholt LNS seminar CRF for human beings LNS seminar 1 / 29 Let G = (V, E) be a graph such that Y = (Y v ) v V, so that Y is indexed by the vertices of G. Then (X, Y) is a conditional
CRF for human beings Arne Skjærholt LNS seminar CRF for human beings LNS seminar 1 / 29 Let G = (V, E) be a graph such that Y = (Y v ) v V, so that Y is indexed by the vertices of G. Then (X, Y) is a conditional
Hidden Markov Models for biological sequence analysis
 Hidden Markov Models for biological sequence analysis Master in Bioinformatics UPF 2017-2018 http://comprna.upf.edu/courses/master_agb/ Eduardo Eyras Computational Genomics Pompeu Fabra University - ICREA
Hidden Markov Models for biological sequence analysis Master in Bioinformatics UPF 2017-2018 http://comprna.upf.edu/courses/master_agb/ Eduardo Eyras Computational Genomics Pompeu Fabra University - ICREA
COMS 4771 Probabilistic Reasoning via Graphical Models. Nakul Verma
 COMS 4771 Probabilistic Reasoning via Graphical Models Nakul Verma Last time Dimensionality Reduction Linear vs non-linear Dimensionality Reduction Principal Component Analysis (PCA) Non-linear methods
COMS 4771 Probabilistic Reasoning via Graphical Models Nakul Verma Last time Dimensionality Reduction Linear vs non-linear Dimensionality Reduction Principal Component Analysis (PCA) Non-linear methods
Grundlagen der Bioinformatik, SS 09, D. Huson, June 16, S. Durbin, S. Eddy, A. Krogh and G. Mitchison, Biological Sequence
 rundlagen der Bioinformatik, SS 09,. Huson, June 16, 2009 81 7 Markov chains and Hidden Markov Models We will discuss: Markov chains Hidden Markov Models (HMMs) Profile HMMs his chapter is based on: nalysis,
rundlagen der Bioinformatik, SS 09,. Huson, June 16, 2009 81 7 Markov chains and Hidden Markov Models We will discuss: Markov chains Hidden Markov Models (HMMs) Profile HMMs his chapter is based on: nalysis,
( ).666 Information Extraction from Speech and Text
 (520 600).666 Information Extraction from Speech and Text HMM Parameters Estimation for Gaussian Output Densities April 27, 205. Generalization of the Results of Section 9.4. It is suggested in Section
(520 600).666 Information Extraction from Speech and Text HMM Parameters Estimation for Gaussian Output Densities April 27, 205. Generalization of the Results of Section 9.4. It is suggested in Section
Basic Text Analysis. Hidden Markov Models. Joakim Nivre. Uppsala University Department of Linguistics and Philology
 Basic Text Analysis Hidden Markov Models Joakim Nivre Uppsala University Department of Linguistics and Philology joakimnivre@lingfiluuse Basic Text Analysis 1(33) Hidden Markov Models Markov models are
Basic Text Analysis Hidden Markov Models Joakim Nivre Uppsala University Department of Linguistics and Philology joakimnivre@lingfiluuse Basic Text Analysis 1(33) Hidden Markov Models Markov models are
Data Mining in Bioinformatics HMM
 Data Mining in Bioinformatics HMM Microarray Problem: Major Objective n Major Objective: Discover a comprehensive theory of life s organization at the molecular level 2 1 Data Mining in Bioinformatics
Data Mining in Bioinformatics HMM Microarray Problem: Major Objective n Major Objective: Discover a comprehensive theory of life s organization at the molecular level 2 1 Data Mining in Bioinformatics
Grundlagen der Bioinformatik, SS 08, D. Huson, June 16, S. Durbin, S. Eddy, A. Krogh and G. Mitchison, Biological Sequence
 rundlagen der Bioinformatik, SS 08,. Huson, June 16, 2008 89 8 Markov chains and Hidden Markov Models We will discuss: Markov chains Hidden Markov Models (HMMs) Profile HMMs his chapter is based on: nalysis,
rundlagen der Bioinformatik, SS 08,. Huson, June 16, 2008 89 8 Markov chains and Hidden Markov Models We will discuss: Markov chains Hidden Markov Models (HMMs) Profile HMMs his chapter is based on: nalysis,
CSci 8980: Advanced Topics in Graphical Models Analysis of Genetic Variation
 CSci 8980: Advanced Topics in Graphical Models Analysis of Genetic Variation Instructor: Arindam Banerjee November 26, 2007 Genetic Polymorphism Single nucleotide polymorphism (SNP) Genetic Polymorphism
CSci 8980: Advanced Topics in Graphical Models Analysis of Genetic Variation Instructor: Arindam Banerjee November 26, 2007 Genetic Polymorphism Single nucleotide polymorphism (SNP) Genetic Polymorphism
CSCE 478/878 Lecture 9: Hidden. Markov. Models. Stephen Scott. Introduction. Outline. Markov. Chains. Hidden Markov Models. CSCE 478/878 Lecture 9:
 Useful for modeling/making predictions on sequential data E.g., biological sequences, text, series of sounds/spoken words Will return to graphical models that are generative sscott@cse.unl.edu 1 / 27 2
Useful for modeling/making predictions on sequential data E.g., biological sequences, text, series of sounds/spoken words Will return to graphical models that are generative sscott@cse.unl.edu 1 / 27 2
Hidden Markov Models. based on chapters from the book Durbin, Eddy, Krogh and Mitchison Biological Sequence Analysis via Shamir s lecture notes
 Hidden Markov Models based on chapters from the book Durbin, Eddy, Krogh and Mitchison Biological Sequence Analysis via Shamir s lecture notes music recognition deal with variations in - actual sound -
Hidden Markov Models based on chapters from the book Durbin, Eddy, Krogh and Mitchison Biological Sequence Analysis via Shamir s lecture notes music recognition deal with variations in - actual sound -
Hidden Markov Models
 Andrea Passerini passerini@disi.unitn.it Statistical relational learning The aim Modeling temporal sequences Model signals which vary over time (e.g. speech) Two alternatives: deterministic models directly
Andrea Passerini passerini@disi.unitn.it Statistical relational learning The aim Modeling temporal sequences Model signals which vary over time (e.g. speech) Two alternatives: deterministic models directly
Multiscale Systems Engineering Research Group
 Hidden Markov Model Prof. Yan Wang Woodruff School of Mechanical Engineering Georgia Institute of echnology Atlanta, GA 30332, U.S.A. yan.wang@me.gatech.edu Learning Objectives o familiarize the hidden
Hidden Markov Model Prof. Yan Wang Woodruff School of Mechanical Engineering Georgia Institute of echnology Atlanta, GA 30332, U.S.A. yan.wang@me.gatech.edu Learning Objectives o familiarize the hidden
Hidden Markov Models (HMMs)
 Hidden Markov Models (HMMs) Reading Assignments R. Duda, P. Hart, and D. Stork, Pattern Classification, John-Wiley, 2nd edition, 2001 (section 3.10, hard-copy). L. Rabiner, "A tutorial on HMMs and selected
Hidden Markov Models (HMMs) Reading Assignments R. Duda, P. Hart, and D. Stork, Pattern Classification, John-Wiley, 2nd edition, 2001 (section 3.10, hard-copy). L. Rabiner, "A tutorial on HMMs and selected
Hidden Markov Models. Hosein Mohimani GHC7717
 Hidden Markov Models Hosein Mohimani GHC7717 hoseinm@andrew.cmu.edu Fair et Casino Problem Dealer flips a coin and player bets on outcome Dealer use either a fair coin (head and tail equally likely) or
Hidden Markov Models Hosein Mohimani GHC7717 hoseinm@andrew.cmu.edu Fair et Casino Problem Dealer flips a coin and player bets on outcome Dealer use either a fair coin (head and tail equally likely) or
Markov Chains and Hidden Markov Models. COMP 571 Luay Nakhleh, Rice University
 Markov Chains and Hidden Markov Models COMP 571 Luay Nakhleh, Rice University Markov Chains and Hidden Markov Models Modeling the statistical properties of biological sequences and distinguishing regions
Markov Chains and Hidden Markov Models COMP 571 Luay Nakhleh, Rice University Markov Chains and Hidden Markov Models Modeling the statistical properties of biological sequences and distinguishing regions
Hidden Markov Models (HMMs) November 14, 2017
 Hidden Markov Models (HMMs) November 14, 2017 inferring a hidden truth 1) You hear a static-filled radio transmission. how can you determine what did the sender intended to say? 2) You know that genes
Hidden Markov Models (HMMs) November 14, 2017 inferring a hidden truth 1) You hear a static-filled radio transmission. how can you determine what did the sender intended to say? 2) You know that genes
Hidden Markov Models. Representing sequence data. Markov Models. A dice-y example 4/5/2017. CISC 5800 Professor Daniel Leeds Π A = 0.3, Π B = 0.
 Representing sequence data Hidden Markov Models CISC 5800 Professor Daniel Leeds Spoken language DNA sequences Daily stock values Example: spoken language F?r plu? fi?e is nine Between F and r expect a
Representing sequence data Hidden Markov Models CISC 5800 Professor Daniel Leeds Spoken language DNA sequences Daily stock values Example: spoken language F?r plu? fi?e is nine Between F and r expect a
Modeling conditional distributions with mixture models: Applications in finance and financial decision-making
 Modeling conditional distributions with mixture models: Applications in finance and financial decision-making John Geweke University of Iowa, USA Journal of Applied Econometrics Invited Lecture Università
Modeling conditional distributions with mixture models: Applications in finance and financial decision-making John Geweke University of Iowa, USA Journal of Applied Econometrics Invited Lecture Università
Lecture 8 Learning Sequence Motif Models Using Expectation Maximization (EM) Colin Dewey February 14, 2008
 Lecture 8 Learning Sequence Motif Models Using Expectation Maximization (EM) Colin Dewey February 14, 2008 1 Sequence Motifs what is a sequence motif? a sequence pattern of biological significance typically
Lecture 8 Learning Sequence Motif Models Using Expectation Maximization (EM) Colin Dewey February 14, 2008 1 Sequence Motifs what is a sequence motif? a sequence pattern of biological significance typically
Graphical Models for Collaborative Filtering
 Graphical Models for Collaborative Filtering Le Song Machine Learning II: Advanced Topics CSE 8803ML, Spring 2012 Sequence modeling HMM, Kalman Filter, etc.: Similarity: the same graphical model topology,
Graphical Models for Collaborative Filtering Le Song Machine Learning II: Advanced Topics CSE 8803ML, Spring 2012 Sequence modeling HMM, Kalman Filter, etc.: Similarity: the same graphical model topology,
Statistical Machine Learning from Data
 Samy Bengio Statistical Machine Learning from Data Statistical Machine Learning from Data Samy Bengio IDIAP Research Institute, Martigny, Switzerland, and Ecole Polytechnique Fédérale de Lausanne (EPFL),
Samy Bengio Statistical Machine Learning from Data Statistical Machine Learning from Data Samy Bengio IDIAP Research Institute, Martigny, Switzerland, and Ecole Polytechnique Fédérale de Lausanne (EPFL),
HIDDEN MARKOV MODELS
 HIDDEN MARKOV MODELS Outline CG-islands The Fair Bet Casino Hidden Markov Model Decoding Algorithm Forward-Backward Algorithm Profile HMMs HMM Parameter Estimation Viterbi training Baum-Welch algorithm
HIDDEN MARKOV MODELS Outline CG-islands The Fair Bet Casino Hidden Markov Model Decoding Algorithm Forward-Backward Algorithm Profile HMMs HMM Parameter Estimation Viterbi training Baum-Welch algorithm
CS838-1 Advanced NLP: Hidden Markov Models
 CS838-1 Advanced NLP: Hidden Markov Models Xiaojin Zhu 2007 Send comments to jerryzhu@cs.wisc.edu 1 Part of Speech Tagging Tag each word in a sentence with its part-of-speech, e.g., The/AT representative/nn
CS838-1 Advanced NLP: Hidden Markov Models Xiaojin Zhu 2007 Send comments to jerryzhu@cs.wisc.edu 1 Part of Speech Tagging Tag each word in a sentence with its part-of-speech, e.g., The/AT representative/nn
Sequence Labeling: HMMs & Structured Perceptron
 Sequence Labeling: HMMs & Structured Perceptron CMSC 723 / LING 723 / INST 725 MARINE CARPUAT marine@cs.umd.edu HMM: Formal Specification Q: a finite set of N states Q = {q 0, q 1, q 2, q 3, } N N Transition
Sequence Labeling: HMMs & Structured Perceptron CMSC 723 / LING 723 / INST 725 MARINE CARPUAT marine@cs.umd.edu HMM: Formal Specification Q: a finite set of N states Q = {q 0, q 1, q 2, q 3, } N N Transition
Lecture 12: Algorithms for HMMs
 Lecture 12: Algorithms for HMMs Nathan Schneider (some slides from Sharon Goldwater; thanks to Jonathan May for bug fixes) ENLP 17 October 2016 updated 9 September 2017 Recap: tagging POS tagging is a
Lecture 12: Algorithms for HMMs Nathan Schneider (some slides from Sharon Goldwater; thanks to Jonathan May for bug fixes) ENLP 17 October 2016 updated 9 September 2017 Recap: tagging POS tagging is a
Effect of Genetic Divergence in Identifying Ancestral Origin using HAPAA
 Effect of Genetic Divergence in Identifying Ancestral Origin using HAPAA Andreas Sundquist*, Eugene Fratkin*, Chuong B. Do, Serafim Batzoglou Department of Computer Science, Stanford University, Stanford,
Effect of Genetic Divergence in Identifying Ancestral Origin using HAPAA Andreas Sundquist*, Eugene Fratkin*, Chuong B. Do, Serafim Batzoglou Department of Computer Science, Stanford University, Stanford,
Hidden Markov Models
 Hidden Markov Models Slides mostly from Mitch Marcus and Eric Fosler (with lots of modifications). Have you seen HMMs? Have you seen Kalman filters? Have you seen dynamic programming? HMMs are dynamic
Hidden Markov Models Slides mostly from Mitch Marcus and Eric Fosler (with lots of modifications). Have you seen HMMs? Have you seen Kalman filters? Have you seen dynamic programming? HMMs are dynamic
Hidden Markov Models
 Hidden Markov Models CI/CI(CS) UE, SS 2015 Christian Knoll Signal Processing and Speech Communication Laboratory Graz University of Technology June 23, 2015 CI/CI(CS) SS 2015 June 23, 2015 Slide 1/26 Content
Hidden Markov Models CI/CI(CS) UE, SS 2015 Christian Knoll Signal Processing and Speech Communication Laboratory Graz University of Technology June 23, 2015 CI/CI(CS) SS 2015 June 23, 2015 Slide 1/26 Content
Phylogenetic Networks with Recombination
 Phylogenetic Networks with Recombination October 17 2012 Recombination All DNA is recombinant DNA... [The] natural process of recombination and mutation have acted throughout evolution... Genetic exchange
Phylogenetic Networks with Recombination October 17 2012 Recombination All DNA is recombinant DNA... [The] natural process of recombination and mutation have acted throughout evolution... Genetic exchange
A Higher-Order Interactive Hidden Markov Model and Its Applications Wai-Ki Ching Department of Mathematics The University of Hong Kong
 A Higher-Order Interactive Hidden Markov Model and Its Applications Wai-Ki Ching Department of Mathematics The University of Hong Kong Abstract: In this talk, a higher-order Interactive Hidden Markov Model
A Higher-Order Interactive Hidden Markov Model and Its Applications Wai-Ki Ching Department of Mathematics The University of Hong Kong Abstract: In this talk, a higher-order Interactive Hidden Markov Model
Hidden Markov Models for biological sequence analysis I
 Hidden Markov Models for biological sequence analysis I Master in Bioinformatics UPF 2014-2015 Eduardo Eyras Computational Genomics Pompeu Fabra University - ICREA Barcelona, Spain Example: CpG Islands
Hidden Markov Models for biological sequence analysis I Master in Bioinformatics UPF 2014-2015 Eduardo Eyras Computational Genomics Pompeu Fabra University - ICREA Barcelona, Spain Example: CpG Islands
Hidden Markov Models. Representing sequence data. Markov Models. A dice-y example 4/26/2018. CISC 5800 Professor Daniel Leeds Π A = 0.3, Π B = 0.
 Representing sequence data Hidden Markov Models CISC 5800 Professor Daniel Leeds Spoken language DNA sequences Daily stock values Example: spoken language F?r plu? fi?e is nine Between F and r expect a
Representing sequence data Hidden Markov Models CISC 5800 Professor Daniel Leeds Spoken language DNA sequences Daily stock values Example: spoken language F?r plu? fi?e is nine Between F and r expect a
Today s Lecture: HMMs
 Today s Lecture: HMMs Definitions Examples Probability calculations WDAG Dynamic programming algorithms: Forward Viterbi Parameter estimation Viterbi training 1 Hidden Markov Models Probability models
Today s Lecture: HMMs Definitions Examples Probability calculations WDAG Dynamic programming algorithms: Forward Viterbi Parameter estimation Viterbi training 1 Hidden Markov Models Probability models
CS839: Probabilistic Graphical Models. Lecture 7: Learning Fully Observed BNs. Theo Rekatsinas
 CS839: Probabilistic Graphical Models Lecture 7: Learning Fully Observed BNs Theo Rekatsinas 1 Exponential family: a basic building block For a numeric random variable X p(x ) =h(x)exp T T (x) A( ) = 1
CS839: Probabilistic Graphical Models Lecture 7: Learning Fully Observed BNs Theo Rekatsinas 1 Exponential family: a basic building block For a numeric random variable X p(x ) =h(x)exp T T (x) A( ) = 1
An Evolutionary Programming Based Algorithm for HMM training
 An Evolutionary Programming Based Algorithm for HMM training Ewa Figielska,Wlodzimierz Kasprzak Institute of Control and Computation Engineering, Warsaw University of Technology ul. Nowowiejska 15/19,
An Evolutionary Programming Based Algorithm for HMM training Ewa Figielska,Wlodzimierz Kasprzak Institute of Control and Computation Engineering, Warsaw University of Technology ul. Nowowiejska 15/19,
Logistic Regression: Online, Lazy, Kernelized, Sequential, etc.
 Logistic Regression: Online, Lazy, Kernelized, Sequential, etc. Harsha Veeramachaneni Thomson Reuter Research and Development April 1, 2010 Harsha Veeramachaneni (TR R&D) Logistic Regression April 1, 2010
Logistic Regression: Online, Lazy, Kernelized, Sequential, etc. Harsha Veeramachaneni Thomson Reuter Research and Development April 1, 2010 Harsha Veeramachaneni (TR R&D) Logistic Regression April 1, 2010
COGS Q250 Fall Homework 7: Learning in Neural Networks Due: 9:00am, Friday 2nd November.
 COGS Q250 Fall 2012 Homework 7: Learning in Neural Networks Due: 9:00am, Friday 2nd November. For the first two questions of the homework you will need to understand the learning algorithm using the delta
COGS Q250 Fall 2012 Homework 7: Learning in Neural Networks Due: 9:00am, Friday 2nd November. For the first two questions of the homework you will need to understand the learning algorithm using the delta
