The Probability of Pathogenicity in Clinical Genetic Testing: A Solution for the Variant of Uncertain Significance
|
|
|
- Norman Henry
- 5 years ago
- Views:
Transcription
1 The Probability of Pathogenicity in Clinical Genetic Testing: A Solution for the Variant of Uncertain Significance 1 Introduction by (in alphabetical order) Michael Gollob, Jeffrey S. Rosenthal, and Kevin Thorpe Peter Munk Cardiac Centre and the University of Toronto (November, 2015; last revised April 18, 2016) It frequently arises in medical genetics that a patient has a particular disease, and also has a genetic variant which may or may not be the cause of that disease. The variant is said to be of unknown or uncertain significance if its disease-causing probability cannot be determined, and this is a common challenge (see e.g. Richards et al. (2008), Cheon, Mozersky, and Cook- Deegan (2014), Domcheck and Weber (2008), and the references therein). The problem is put succinctly in the patient guide by Ambry Genetics (2015), which states, Variants of unknown significance are DNA changes with too little information known to classify as either pathogenic or benign, and it is unknown whether they contribute to a medical condition. Assessing whether or not the genetic variant did indeed cause the disease is important not only for future medical research and prevention, but also as a practical guide to whether or not the patient s family members are also at risk. However, in the case of a rare variant with little or no previous information available, it is unclear how this assessment should be made. In this paper, we present a direct calculation for determining the probability that a rare genetic variant is indeed the cause of an observed disease. Our calculation requires one assumption, namely the natural-seeming Variant Fraction Assumption that in the absence of any other evidence, the probability that a newly observed rare genetic variant of a gene causes a specified disease is equal to the fraction of all previously-observed rare variants of that gene which did cause that disease (see Section 4.3 for details). With this one assumption, we are able to compute the desired probability purely in terms of the disease s prevalence in the population, and the fraction of rare variants in the general population and in the diseased population. Our results are described below. 2 Formal Set-Up To set up our probability model, we make the following assumptions and notations: 1
2 A certain disease D has a known prevalence p in the general population. Among the healthy population, a certain known fraction q have some distinct rare variant of a certain gene G. Among patients with the disease D, a certain fraction r q have some distinct rare variant of the gene G. Some subset S of the rare variants of G always cause the disease D, i.e. the probability of disease given a rare variant in S is 1. Rare variants of G which are not in S have no effect on D, i.e. the probability of disease given a rare variant not in S is the same as for patients without a rare variant. A new patient X is found to have a never-before-seen variant V of the gene G. The question is, given all of the above, if X gets the disease D, then what is the probability that the genetic variant V was actually a cause of the disease D in X? That is, we wish to compute the conditional probability P(V S X has the disease D), i.e. the probability that X s variant V is in fact a cause of the disease D in X, given that X has the variant V and also has the disease D. We describe our calculation of this and related probabilities below. We first note that since the genetic variant V has never been seen before, it is impossible compute its probabilities without making some additional assumption. Thus, we also make the Variant Fraction Assumption that in the absence of any other evidence, the probability that a newly observed genetic variant of a gene G is a cause of D, is equal to the fraction of all of the previously-observed rare variants of G which were indeed found to cause D. For a more precise statement of this assumption, see Section 4.3 below. Remark. In fact, the assumption that r q follows directly from the other assumptions; see Appendix A2. 3 Main Result Our main results are as follows. (Below, unconditional probability means the probability without conditioning on whether or not X has the disease D. Also, we write V is the sole cause of D to mean that V caused D, and furthermore patient X would not have gotten D in the hypothetical case that they instead did not have the rare variant V.) 2
3 Theorem 1 For the above set-up, for a patient X having a rare variant V of gene G, under the Variant Fraction Assumption, we have the following probabilities, where y = (r q)/(1 q), z = p(1 y)/(1 py), w = py / [pr + (1 p)q], and u = (1 z)w. (a) The unconditional probability that the variant V is a cause of the disease D is given by: P(V S) = w. (b) The unconditional probability that patient X will get disease D is given by: P(X gets D) = z + u. (c) Conditional on patient X getting the disease D, the conditional probability that the variant V was the sole cause of D is given by: P(X s disease D was caused solely by V X has D) = u z + u. (d) Conditional on patient X getting the disease D, the conditional probability that the variant V is a cause of D is given by: P(V S X gets D) = w z + u = r q r(1 q). Theorem 1 thus gives precise probabilities for the relevant possibilities related to patient X and disease D and rare genetic variant V. In particular, Theorem 1(d) gives a precise estimate of the probability, given that patient X has disease D and has the rare genetic variant V, that V is in fact a cause of the disease D. Theorem 1 is proved in the next section. We first consider some numerical examples. For example, if p = 1/4, 000 is the prevalence of the disease in the general population, and q = 2% = 0.02 is prevalence of some rare variant of G in the healthy population, and r = 40% = 0.4 is the prevalence of some rare variant of G in patients with the disease, then conditional on X getting the disease, the probability that the variant V is a cause of the disease works out to: P(V S X gets D) = = 97.0%, or about 97 percent (i.e., nearly certain). Or, if p = 1/400 and q = 0.01 and r = 0.4, then P(V S X gets D). = 98.5%. By contrast, if p = 1/50 and q = 0.1 and r = 0.2, then P(V S X gets D). = 58.1%, which is much smaller. Or, if p = 1/400 and q = 0.1 and r = 0.15, then P(V S X gets D). = 39.5%. 3
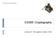
4 Figure 1: Disease cause probabilities as a function of r, with q = 0.1 fixed. A plot of other values of P(V S X gets D) is presented in Figure 1 as a function of r, with q = 0.1 fixed. Probabilities for other parameter values can be computed using the above formulae, or using our simple javascript online calculator available at: As a further check, we note that the formula in Theorem 1(d) gives answers which make sense even for certain extreme parameter values. For example, if r = 1 (i.e., 100%), then it is computed that the formula gives a value of 1. This makes sense, since if r = 1, then by the definition of r, every diseased patient has a rare variant of G, which means that the disease can only be caused by a rare variant of G, so that V must indeed cause D. Or, if q = 0, then again it is computed that the formula gives a value of 1. This also makes sense, since if q = 0, then by the definition of q, no healthy patients have rare variants of G, which means that rare variants of G always cause D, so again V must indeed cause D. By contrast, if r = q, then it is computed that the formula gives a value of 0. This again makes sense, since if the rate of rare variants of G is the same for diseased and healthy 4
5 patients, then the rare variants of G do not cause any additional disease at all, so none of them cause D. As a final comment, we note that since u = (1 z)w, the formula in Theorem 1(c) is smaller than that in Theorem 1(d) by a factor of (1 z). This is due to the possibility that V S but X still would have gotten D by chance alone, i.e. that V does indeed cause D, but X would have gotten D even in the absence of V (e.g. from a variant of some other gene besides G). Now, usually z will be very small, so the answers in (c) and (d) will be very similar, though not identical. 4 Proof of Theorem 1 In this section, we prove Theorem 1 using a sequence of probability calculations. We break up our argument into several steps. 4.1 Preliminary Population Prevalence Calculations We first note that our assumption above that rare variants of G which are not in S have no effect on D, can be written more formally as P(X has D X has a rare variant V S) = P(X has D X has no rare variant). From this it follows (see Appendix A1) that P(X has rare variant V / S X has D, and no variant in S) = P(X has some rare variant X does not have D). We next calculate two population fractions that will be important in our solution. First, we write y for the fraction of diseased patients who have a rare variant of G which is in S, i.e. which does cause D. (Thus, y is close to r, but slightly less since even among diseased patients without a variant in S, a fraction q of them will still happen to have a rare variant not in S by chance alone, just like for the healthy population.) In terms of y, the set of all diseased patients can be divided into three groups: a fraction y with a rare variant of G which is in S, a fraction (1 y)q with a rare variant of G which is not in S, and a fraction (1 y)(1 q) with no rare variant of G at all. Hence, the fraction of diseased patients with some rare variant of G is equal to y + (1 y)q. On the other hand, we know that the overall fraction of diseased patients with some rare variant of G is equal to r. For this to hold, we must have y + (1 y)q = r. Solving for y, we obtain that y = (r q)/(1 q). Then, since a fraction p of the population is diseased, and a fraction y of diseased patients have a variant in S, it follows that the fraction of the total population who have a rare variant 5
6 in S is equal to the product py. And, the fraction who do not have a rare variant in S is equal to 1 py. Next, we write z for the prevalence of the disease D among all people who specifically do not have a variant in S. Then the fraction of people who have D but do not have a variant in S is equal to (1 py)z. And, the fraction of people with a variant in S (who therefore have D) is equal to py. So, the total fraction of the population who have the disease D is equal to (1 py)z + py. On the other hand, we know that the overall prevalence of the disease is equal to p. For this to hold, we must have p = py + (1 py)z. Solving for z, we compute that z = p(1 y)/(1 py). 4.2 The Prior Probability of Disease Prior to diagnosis, what was the prior probability that a given patient X, who has some rare variant V, would get the disease D? That is, what is the conditional probability that X gets D, conditioning (throughout) on the fact that X has a rare variant V? To answer this question, let I be the indicator function of the subset S, so I(V)=1 if V S, otherwise I(V)=0. We know that the prior probability of X getting the disease D is equal to 1 (i.e., 100%) if I(V)=1, or is equal to z if I(V)=0. So, if we knew I(V), i.e. if we knew whether or not V causes D, then could write this as: P(X gets D I(V)) = z + (1 z) I(V ). In fact we do not know I(V), i.e. it could equal either 1 or 0. So, instead, we use the Law of Total Expectation (that the expected value of a conditional probability is the unconditional probability). This shows that: P(X gets D) = E [ P(X gets D I(V)) ] = z + (1 z) E[I(V )] = z + (1 z) P[I(V ) = 1] = z + (1 z) P(V S). This gives a formula for the prior probability that X would get D, in terms of the probability P(V S) that V is in S. However, we do not know P(V S), so it must be estimated. 4.3 The Variant Fraction Assumption To continue, we need to obtain an estimate for P(V S), the unconditional probability (in the absence of any other evidence) that the newly observed variant V of G is in fact a cause of the disease D. Now, since V was never before seen, there is no way to directly calculate this probability. Instead, as mentioned above, we use the Variant Fraction Assumption that 6
7 in the absence of any other evidence, the probability that a newly observed genetic variant of a gene G is a cause of D, is equal to the fraction of all of the previously-observed rare variants of G which were indeed found to cause D. That is, we assume that P(V S) = fraction of the population with a disease-causing rare variant of G fraction of the population with any rare variant of G. This assumption appears to be quite reasonable, in the absence of any other prior information about the new variant V. In any case, some such assumption must be made, otherwise no probabilities associated with V can possibly be computed. But under this one assumption, all of the remaining probability calculations can be completed. 4.4 Estimating the Variant Probability Using the above Variant Fraction Assumption, we are able to compute the desired probability P(V S). To do this, we need to compute the fraction of all of G s rare variants which are in S. We proceed as follows. Since a fraction p of the population is diseased, and since a fraction r of them have some rare variant of G, it follows that the fraction of the population who are diseased and have some rare variant of G is pr. Similarly, since y is the fraction of diseased patients who have a variant in S, it follows that the fraction of the population who are diseased and have a rare variant in S is py. Also, the fraction of the population who are healthy and have some rare variant of G is (1 p)q. That is, a fraction pr + (1 p)q of the population has a rare variant, and a fraction py of the population has a rare variant in S. So, assuming these rare variants are all distinct, the fraction of all the rare variants of G in the entire population which are in S is equal to py/[pr + (1 p)q]. We then estimate the probability P(V S) by the above fraction of all the rare variants of G which are in S, i.e. by P(V S) py / [pr + (1 p)q] =: w where w = py / [pr + (1 p)q], as claimed in Theorem 1(a). Now, since we earlier derived the prior-to-diagnosis probability P(X gets D) = z + (1 z) P(V S), it then follows from the above estimate that P(X gets D) = z + (1 z) w =: z + u where u = (1 z)w, as claimed in Theorem 1(b). 7
8 4.5 The Cause Probability The above formula for P(X gets D) can be interpreted as follows: Patient X can get the disease D either without any influence at all from the gene G (with probability z), or caused by the variant V of G (with probability f). Under this interpretation, given that X does in fact have the disease D, the conditional probability that the disease D in X was caused by the genetic variant V, and would not otherwise have arisen, is given by the second probability divided by the sum of the two probabilities, i.e. by: P(X s disease D was caused solely by V X has D) = u z + u, as claimed in Theorem 1(c). This formula thus gives an estimate of the probability, given that patient X has disease D, that the disease was caused solely by their rare genetic variant V of the gene G. This is similar to, but not quite the same as, our desired conditional probability, as we now explain. 4.6 Computing the Conditional Probability Putting the previous equations together, we can compute the required conditional probability, as follows: P(V S, and X gets D) P(V S X gets D) = P(X gets D) = P(V S) P(X gets D) = w z + u. This gives our first formula claimed in Theorem 1(d). w Finally, through careful algebraic simplification, it is verified directly that in fact = z+u r q (which, surprisingly, does not depend on p), thus giving the second formula claimed r(1 q) in Theorem 1(d). Remark. whence The above equations can be combined as follows. We have that u z + u = P(D was caused solely by V X gets D) = P(D was caused solely by V) / P(X gets D) = P(D was caused solely by V V in S) P(V in S X gets D) w = P(D was caused solely by V V in S) z + u, P(D was caused solely by V V in S) = u w = (1 z)w w = 1 z. 8
9 Taking the complementary event, P(X would have gotten D even without having V V in S) = z, which makes sense since z is the prevalence of the disease in people who do not have a rare variant, corresponding to the appropriate hypothetical. Final Remark. After originally completing this research, we became aware of the related recent paper by Ruklisa, Ware, Walsh, Balding, and Cook (2015). That paper mostly focuses on specific probability estimates for specific diseases and specific genetic variants. However, it does begin with what they call the prior odds of pathogenicity, which appears to correspond to odds for the event P(V S X gets D) considered in Theorem 1(d) above. For that case, they assert that the odds might be assumed to be given by Burden of rare variants in cases Burden of rare variants in controls Burden of rare variants in controls In our notation, this appears to correspond to asserting that or equivalently that P(V S X gets D) 1 P(V S X gets D) = r q, q. P(V S X gets D) = r q q 1 + r q q = r q r. This last expression is fairly similar to the final formula in our Theorem 1(d), but it differs by a factor of 1 q. It appears that the reason for this discrepency is their assertion that it is reasonable to assume that the burden of benign rare variants in cases is equal to the burden of rare variants in controls, which apparently corresponds to assuming that P(Rare variant not in S Diseased) = P(Rare variant not in S Not diseased), which is different from what we believe (for more see Appendix A3). Appendix: Additional Probability Calculations We here provide a few additional probability calculations, to further clarify some of the material in the main text. For these calculations, define a through f to be the proportions of the total population in each of the six categories implied by the following Table: 9
10 A1. We first show that the assumption Diseased Not diseased Rare variant in S a d Rare variant not in S b e No rare variant c f P(X has D X has a rare variant V S) = P(X has D X has no rare variant) ( ). implies the condition that P(X has rare variant V / S X has D, and no variant in S) = P(X has some rare variant X does not have D). ( ) In terms of the above Table, condition ( ) is equivalent to saying that b/(b + e) = c/(c + f), i.e. that b/e = c/f. Meanwhile, condition ( ) is equivalent to saying that b/(b + c) = (d + e)/(d+e+f). On the other hand, our assumption that variants in S always cause the disease D implies that d = 0. Hence, if b/e = c/f, then (d + e)/(d + e + f) = e/(e + f) = b/(b + c), thus establishing ( ). A2. We here show that the assumption r q actually follows from our other assumptions. Indeed, in terms of the above Table, using as above that d = 0 and e/(e + f) = b/(b + c), we have that Also q = P(Rare variant Not Diseased) = The result then follows since r q = a + b a + b + c r = P(Rare variant Diseased) = d + e d + e + f = e e + f = b b + c. a + b a + b + c. b b + c = (ab + ac + b2 + bc) (ab + b 2 + bc) (a + b + c)(b + c) = ac (a + b + c)(b + c) 0. A3. Finally, we consider the quantities which arise in the reference Ruklisa et al. (2015), as discussed in our Final Remark above, namely and P(Rare variant not in S Diseased) P(Rare variant not in S Not diseased). (&) (&&) In our notation, (&&) = P(Rare variant not in S Not diseased) = q which, in terms of the above Table, is equal to e/(e+f). By contrast, (&) = P(Rare variant not in S Diseased) = b/(a+b+c). Hence, assuming ( ), we have (&) = b/(a+b+c) b/(b+c) = e/(e+f) = (&&). Furthermore, we have non-zero equality only when a = 0, which corresponds to no variants in S, i.e. to the gene G having no effect whatsoever on the disease D. Otherwise, a > 0, and hence (&) < (&&) (assuming b > 0), leading to a different result from that of Ruklisa et al. 10
11 References [1] Ambry Genetics (2015), Your genetic test result. Available at: General%20VUS%20patient%20brochure%20Updates 0.pdf [2] Cheon, J.Y., Mozersky, J., and Cook-Deegan, R. (2014), Variants of uncertain significance in BRCA: a harbinger of ethical and policy issues to come? Genome Medicine 6:121, [3] Domchek, S. and Weber, B.L. (2008), Genetic variants of uncertain significance: flies in the ointment. Journal of Clinical Oncology 26(1), [4] Richards, C.S., Bale, S., Bellissimo, D.B., Das, S., Grody, W.W., Hegde, M.R., Lyon, E., and Ward, B.E. (2008), ACMG recommendations for standards for interpretation and reporting of sequence variations: Revisions Genetics in Medicine bf 10(4), [5] Ruklisa, D., Ware, J.S., Walsh, R., Balding, D.J., and Cook, S.A. (2015), Bayesian models for syndrome- and gene-specific probabilities of novel variant pathogenicity. Genome Medicine 7:5,
The Probability of Pathogenicity in Clinical Genetic Testing: A Solution for the Variant of Uncertain Significance
 International Journal of Statistics and Probability; Vol. 5, No. 4; July 2016 ISSN 1927-7032 E-ISSN 1927-7040 Published by Canadian Center of Science and Education The Probability of Pathogenicity in Clinical
International Journal of Statistics and Probability; Vol. 5, No. 4; July 2016 ISSN 1927-7032 E-ISSN 1927-7040 Published by Canadian Center of Science and Education The Probability of Pathogenicity in Clinical
Review. More Review. Things to know about Probability: Let Ω be the sample space for a probability measure P.
 1 2 Review Data for assessing the sensitivity and specificity of a test are usually of the form disease category test result diseased (+) nondiseased ( ) + A B C D Sensitivity: is the proportion of diseased
1 2 Review Data for assessing the sensitivity and specificity of a test are usually of the form disease category test result diseased (+) nondiseased ( ) + A B C D Sensitivity: is the proportion of diseased
Fundamentals of Machine Learning for Predictive Data Analytics
 Fundamentals of Machine Learning for Predictive Data Analytics Chapter 6: Probability-based Learning Sections 6.1, 6.2, 6.3 John Kelleher and Brian Mac Namee and Aoife D Arcy john.d.kelleher@dit.ie brian.macnamee@ucd.ie
Fundamentals of Machine Learning for Predictive Data Analytics Chapter 6: Probability-based Learning Sections 6.1, 6.2, 6.3 John Kelleher and Brian Mac Namee and Aoife D Arcy john.d.kelleher@dit.ie brian.macnamee@ucd.ie
Probability and Probability Distributions. Dr. Mohammed Alahmed
 Probability and Probability Distributions 1 Probability and Probability Distributions Usually we want to do more with data than just describing them! We might want to test certain specific inferences about
Probability and Probability Distributions 1 Probability and Probability Distributions Usually we want to do more with data than just describing them! We might want to test certain specific inferences about
Exact McNemar s Test and Matching Confidence Intervals Michael P. Fay April 25,
 Exact McNemar s Test and Matching Confidence Intervals Michael P. Fay April 25, 2016 1 McNemar s Original Test Consider paired binary response data. For example, suppose you have twins randomized to two
Exact McNemar s Test and Matching Confidence Intervals Michael P. Fay April 25, 2016 1 McNemar s Original Test Consider paired binary response data. For example, suppose you have twins randomized to two
Ignoring the matching variables in cohort studies - when is it valid, and why?
 Ignoring the matching variables in cohort studies - when is it valid, and why? Arvid Sjölander Abstract In observational studies of the effect of an exposure on an outcome, the exposure-outcome association
Ignoring the matching variables in cohort studies - when is it valid, and why? Arvid Sjölander Abstract In observational studies of the effect of an exposure on an outcome, the exposure-outcome association
Probability. Introduction to Biostatistics
 Introduction to Biostatistics Probability Second Semester 2014/2015 Text Book: Basic Concepts and Methodology for the Health Sciences By Wayne W. Daniel, 10 th edition Dr. Sireen Alkhaldi, BDS, MPH, DrPH
Introduction to Biostatistics Probability Second Semester 2014/2015 Text Book: Basic Concepts and Methodology for the Health Sciences By Wayne W. Daniel, 10 th edition Dr. Sireen Alkhaldi, BDS, MPH, DrPH
Footnotes to Linear Algebra (MA 540 fall 2013), T. Goodwillie, Bases
 Footnotes to Linear Algebra (MA 540 fall 2013), T. Goodwillie, Bases November 18, 2013 1 Spanning and linear independence I will outline a slightly different approach to the material in Chapter 2 of Axler
Footnotes to Linear Algebra (MA 540 fall 2013), T. Goodwillie, Bases November 18, 2013 1 Spanning and linear independence I will outline a slightly different approach to the material in Chapter 2 of Axler
The Bayes Theorema. Converting pre-diagnostic odds into post-diagnostic odds. Prof. Dr. F. Vanstapel, MD PhD Laboratoriumgeneeskunde UZ KULeuven
 slide 1 The Bayes Theorema Converting pre-diagnostic odds into post-diagnostic odds Prof. Dr. F. Vanstapel, MD PhD Laboratoriumgeneeskunde UZ KULeuven slide 2 Problem * The yearly incidence of TBC infections
slide 1 The Bayes Theorema Converting pre-diagnostic odds into post-diagnostic odds Prof. Dr. F. Vanstapel, MD PhD Laboratoriumgeneeskunde UZ KULeuven slide 2 Problem * The yearly incidence of TBC infections
Solutions to November 2006 Problems
 Solutions to November 2006 Problems Problem 1. A six-sided convex polygon is inscribed in a circle. All its angles are equal. Show that its sides need not be equal. What can be said about seven-sided equal-angled
Solutions to November 2006 Problems Problem 1. A six-sided convex polygon is inscribed in a circle. All its angles are equal. Show that its sides need not be equal. What can be said about seven-sided equal-angled
Basic Probabilistic Reasoning SEG
 Basic Probabilistic Reasoning SEG 7450 1 Introduction Reasoning under uncertainty using probability theory Dealing with uncertainty is one of the main advantages of an expert system over a simple decision
Basic Probabilistic Reasoning SEG 7450 1 Introduction Reasoning under uncertainty using probability theory Dealing with uncertainty is one of the main advantages of an expert system over a simple decision
Causality II: How does causal inference fit into public health and what it is the role of statistics?
 Causality II: How does causal inference fit into public health and what it is the role of statistics? Statistics for Psychosocial Research II November 13, 2006 1 Outline Potential Outcomes / Counterfactual
Causality II: How does causal inference fit into public health and what it is the role of statistics? Statistics for Psychosocial Research II November 13, 2006 1 Outline Potential Outcomes / Counterfactual
Discrete Mathematics and Probability Theory Spring 2016 Rao and Walrand Note 14
 CS 70 Discrete Mathematics and Probability Theory Spring 2016 Rao and Walrand Note 14 Introduction One of the key properties of coin flips is independence: if you flip a fair coin ten times and get ten
CS 70 Discrete Mathematics and Probability Theory Spring 2016 Rao and Walrand Note 14 Introduction One of the key properties of coin flips is independence: if you flip a fair coin ten times and get ten
1. When applied to an affected person, the test comes up positive in 90% of cases, and negative in 10% (these are called false negatives ).
 CS 70 Discrete Mathematics for CS Spring 2006 Vazirani Lecture 8 Conditional Probability A pharmaceutical company is marketing a new test for a certain medical condition. According to clinical trials,
CS 70 Discrete Mathematics for CS Spring 2006 Vazirani Lecture 8 Conditional Probability A pharmaceutical company is marketing a new test for a certain medical condition. According to clinical trials,
Document category: There is no restriction on the circulation of this document
 GA2-06 Agenda Item 2 Issued: 16 January 2018 CIMMYT Position on gene editing: An example to support the development of a common position on gene editing Purpose This document provides CIMMYT s Position
GA2-06 Agenda Item 2 Issued: 16 January 2018 CIMMYT Position on gene editing: An example to support the development of a common position on gene editing Purpose This document provides CIMMYT s Position
2. Introduction to commutative rings (continued)
 2. Introduction to commutative rings (continued) 2.1. New examples of commutative rings. Recall that in the first lecture we defined the notions of commutative rings and field and gave some examples of
2. Introduction to commutative rings (continued) 2.1. New examples of commutative rings. Recall that in the first lecture we defined the notions of commutative rings and field and gave some examples of
Brief Review of Probability
 Brief Review of Probability Nuno Vasconcelos (Ken Kreutz-Delgado) ECE Department, UCSD Probability Probability theory is a mathematical language to deal with processes or experiments that are non-deterministic
Brief Review of Probability Nuno Vasconcelos (Ken Kreutz-Delgado) ECE Department, UCSD Probability Probability theory is a mathematical language to deal with processes or experiments that are non-deterministic
Bayesian Inference. Introduction
 Bayesian Inference Introduction The frequentist approach to inference holds that probabilities are intrinsicially tied (unsurprisingly) to frequencies. This interpretation is actually quite natural. What,
Bayesian Inference Introduction The frequentist approach to inference holds that probabilities are intrinsicially tied (unsurprisingly) to frequencies. This interpretation is actually quite natural. What,
Genetic Changes Lesson 2 HW
 Guiding Question What theory serves as the basis of what we believe about how evolutionary changes occur? 7 th GRADE SCIENCE Genetic Changes Lesson 2 HW # Name: Date: Homeroom: Jean-Baptiste Lamarck (1744-1829)
Guiding Question What theory serves as the basis of what we believe about how evolutionary changes occur? 7 th GRADE SCIENCE Genetic Changes Lesson 2 HW # Name: Date: Homeroom: Jean-Baptiste Lamarck (1744-1829)
1 INFO 2950, 2 4 Feb 10
 First a few paragraphs of review from previous lectures: A finite probability space is a set S and a function p : S [0, 1] such that p(s) > 0 ( s S) and s S p(s) 1. We refer to S as the sample space, subsets
First a few paragraphs of review from previous lectures: A finite probability space is a set S and a function p : S [0, 1] such that p(s) > 0 ( s S) and s S p(s) 1. We refer to S as the sample space, subsets
More on conditioning and Mr. Bayes
 More on conditioning and Mr. Bayes Saad Mneimneh 1 Multiplication rule for conditioning We can generalize the formula P(A,B) P(A B)P(B) to more than two events. For instance, P(A,B,C) P(A)P(B A)P(C A,B).
More on conditioning and Mr. Bayes Saad Mneimneh 1 Multiplication rule for conditioning We can generalize the formula P(A,B) P(A B)P(B) to more than two events. For instance, P(A,B,C) P(A)P(B A)P(C A,B).
Citation for published version (APA): Ruíz Duarte, E. An invitation to algebraic number theory and class field theory
 University of Groningen An invitation to algebraic number theory and class field theory Ruíz Duarte, Eduardo IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's PDF) if you
University of Groningen An invitation to algebraic number theory and class field theory Ruíz Duarte, Eduardo IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's PDF) if you
MATH 201 Solutions: TEST 3-A (in class)
 MATH 201 Solutions: TEST 3-A (in class) (revised) God created infinity, and man, unable to understand infinity, had to invent finite sets. - Gian Carlo Rota Part I [5 pts each] 1. Let X be a set. Define
MATH 201 Solutions: TEST 3-A (in class) (revised) God created infinity, and man, unable to understand infinity, had to invent finite sets. - Gian Carlo Rota Part I [5 pts each] 1. Let X be a set. Define
Lectures 5 & 6: Hypothesis Testing
 Lectures 5 & 6: Hypothesis Testing in which you learn to apply the concept of statistical significance to OLS estimates, learn the concept of t values, how to use them in regression work and come across
Lectures 5 & 6: Hypothesis Testing in which you learn to apply the concept of statistical significance to OLS estimates, learn the concept of t values, how to use them in regression work and come across
PubH 7470: STATISTICS FOR TRANSLATIONAL & CLINICAL RESEARCH
 PubH 7470: STATISTICS FOR TRANSLATIONAL & CLINICAL RESEARCH The First Step: SAMPLE SIZE DETERMINATION THE ULTIMATE GOAL The most important, ultimate step of any of clinical research is to do draw inferences;
PubH 7470: STATISTICS FOR TRANSLATIONAL & CLINICAL RESEARCH The First Step: SAMPLE SIZE DETERMINATION THE ULTIMATE GOAL The most important, ultimate step of any of clinical research is to do draw inferences;
Environmental Economics Lectures 11, 12 Valuation and Cost-Benefit Analysis
 Environmental Economics Lectures 11, 12 Valuation and Cost-Benefit Analysis Florian K. Diekert April 9 and 30, 2014 Perman et al (2011) ch. 11-13 CON 4910, L11&12. Slide 1/ 28 Preview lecture 11 and 12
Environmental Economics Lectures 11, 12 Valuation and Cost-Benefit Analysis Florian K. Diekert April 9 and 30, 2014 Perman et al (2011) ch. 11-13 CON 4910, L11&12. Slide 1/ 28 Preview lecture 11 and 12
Formal Modeling in Cognitive Science
 Formal Modeling in Cognitive Science Lecture 9: Application of Bayes Theorem; Discrete Random Variables; Steve Renals (notes by Frank Keller) School of Informatics University of Edinburgh s.renals@ed.ac.uk
Formal Modeling in Cognitive Science Lecture 9: Application of Bayes Theorem; Discrete Random Variables; Steve Renals (notes by Frank Keller) School of Informatics University of Edinburgh s.renals@ed.ac.uk
An introduction to biostatistics: part 1
 An introduction to biostatistics: part 1 Cavan Reilly September 6, 2017 Table of contents Introduction to data analysis Uncertainty Probability Conditional probability Random variables Discrete random
An introduction to biostatistics: part 1 Cavan Reilly September 6, 2017 Table of contents Introduction to data analysis Uncertainty Probability Conditional probability Random variables Discrete random
Formal Modeling in Cognitive Science Lecture 19: Application of Bayes Theorem; Discrete Random Variables; Distributions. Background.
 Formal Modeling in Cognitive Science Lecture 9: ; Discrete Random Variables; Steve Renals (notes by Frank Keller) School of Informatics University of Edinburgh s.renals@ed.ac.uk February 7 Probability
Formal Modeling in Cognitive Science Lecture 9: ; Discrete Random Variables; Steve Renals (notes by Frank Keller) School of Informatics University of Edinburgh s.renals@ed.ac.uk February 7 Probability
Artificial Intelligence CS 6364
 Artificial Intelligence CS 6364 rofessor Dan Moldovan Section 12 robabilistic Reasoning Acting under uncertainty Logical agents assume propositions are - True - False - Unknown acting under uncertainty
Artificial Intelligence CS 6364 rofessor Dan Moldovan Section 12 robabilistic Reasoning Acting under uncertainty Logical agents assume propositions are - True - False - Unknown acting under uncertainty
Lecture 2: Probability and Distributions
 Lecture 2: Probability and Distributions Ani Manichaikul amanicha@jhsph.edu 17 April 2007 1 / 65 Probability: Why do we care? Probability helps us by: Allowing us to translate scientific questions info
Lecture 2: Probability and Distributions Ani Manichaikul amanicha@jhsph.edu 17 April 2007 1 / 65 Probability: Why do we care? Probability helps us by: Allowing us to translate scientific questions info
Naive Bayes classification
 Naive Bayes classification Christos Dimitrakakis December 4, 2015 1 Introduction One of the most important methods in machine learning and statistics is that of Bayesian inference. This is the most fundamental
Naive Bayes classification Christos Dimitrakakis December 4, 2015 1 Introduction One of the most important methods in machine learning and statistics is that of Bayesian inference. This is the most fundamental
Adaptive clinical trials with subgroup selection
 Adaptive clinical trials with subgroup selection Nigel Stallard 1, Tim Friede 2, Nick Parsons 1 1 Warwick Medical School, University of Warwick, Coventry, UK 2 University Medical Center Göttingen, Göttingen,
Adaptive clinical trials with subgroup selection Nigel Stallard 1, Tim Friede 2, Nick Parsons 1 1 Warwick Medical School, University of Warwick, Coventry, UK 2 University Medical Center Göttingen, Göttingen,
Philosophy 148 Announcements & Such
 Branden Fitelson Philosophy 148 Lecture 1 Philosophy 148 Announcements & Such Overall, people did very well on the mid-term (µ = 90, σ = 16). HW #2 graded will be posted very soon. Raul won t be able to
Branden Fitelson Philosophy 148 Lecture 1 Philosophy 148 Announcements & Such Overall, people did very well on the mid-term (µ = 90, σ = 16). HW #2 graded will be posted very soon. Raul won t be able to
Lecture 3: Probability
 Lecture 3: Probability 28th of October 2015 Lecture 3: Probability 28th of October 2015 1 / 36 Summary of previous lecture Define chance experiment, sample space and event Introduce the concept of the
Lecture 3: Probability 28th of October 2015 Lecture 3: Probability 28th of October 2015 1 / 36 Summary of previous lecture Define chance experiment, sample space and event Introduce the concept of the
Machine Learning (CS 567) Lecture 2
 Machine Learning (CS 567) Lecture 2 Time: T-Th 5:00pm - 6:20pm Location: GFS118 Instructor: Sofus A. Macskassy (macskass@usc.edu) Office: SAL 216 Office hours: by appointment Teaching assistant: Cheol
Machine Learning (CS 567) Lecture 2 Time: T-Th 5:00pm - 6:20pm Location: GFS118 Instructor: Sofus A. Macskassy (macskass@usc.edu) Office: SAL 216 Office hours: by appointment Teaching assistant: Cheol
The expansion of random regular graphs
 The expansion of random regular graphs David Ellis Introduction Our aim is now to show that for any d 3, almost all d-regular graphs on {1, 2,..., n} have edge-expansion ratio at least c d d (if nd is
The expansion of random regular graphs David Ellis Introduction Our aim is now to show that for any d 3, almost all d-regular graphs on {1, 2,..., n} have edge-expansion ratio at least c d d (if nd is
Confidence Intervals of the Simple Difference between the Proportions of a Primary Infection and a Secondary Infection, Given the Primary Infection
 Biometrical Journal 42 (2000) 1, 59±69 Confidence Intervals of the Simple Difference between the Proportions of a Primary Infection and a Secondary Infection, Given the Primary Infection Kung-Jong Lui
Biometrical Journal 42 (2000) 1, 59±69 Confidence Intervals of the Simple Difference between the Proportions of a Primary Infection and a Secondary Infection, Given the Primary Infection Kung-Jong Lui
Probability: Why do we care? Lecture 2: Probability and Distributions. Classical Definition. What is Probability?
 Probability: Why do we care? Lecture 2: Probability and Distributions Sandy Eckel seckel@jhsph.edu 22 April 2008 Probability helps us by: Allowing us to translate scientific questions into mathematical
Probability: Why do we care? Lecture 2: Probability and Distributions Sandy Eckel seckel@jhsph.edu 22 April 2008 Probability helps us by: Allowing us to translate scientific questions into mathematical
Predicate Calculus. Lila Kari. University of Waterloo. Predicate Calculus CS245, Logic and Computation 1 / 59
 Predicate Calculus Lila Kari University of Waterloo Predicate Calculus CS245, Logic and Computation 1 / 59 Predicate Calculus Alternative names: predicate logic, first order logic, elementary logic, restricted
Predicate Calculus Lila Kari University of Waterloo Predicate Calculus CS245, Logic and Computation 1 / 59 Predicate Calculus Alternative names: predicate logic, first order logic, elementary logic, restricted
Paucity, abundance, and the theory of number: Online Appendices
 Paucity, abundance, and the theory of number: Online Appendices Daniel Harbour Language, Volume 90, Number 1, March 2014, pp. s1-s4 (Article) Published by Linguistic Society of America DOI: https://doi.org/10.1353/lan.2014.0019
Paucity, abundance, and the theory of number: Online Appendices Daniel Harbour Language, Volume 90, Number 1, March 2014, pp. s1-s4 (Article) Published by Linguistic Society of America DOI: https://doi.org/10.1353/lan.2014.0019
Person-Time Data. Incidence. Cumulative Incidence: Example. Cumulative Incidence. Person-Time Data. Person-Time Data
 Person-Time Data CF Jeff Lin, MD., PhD. Incidence 1. Cumulative incidence (incidence proportion) 2. Incidence density (incidence rate) December 14, 2005 c Jeff Lin, MD., PhD. c Jeff Lin, MD., PhD. Person-Time
Person-Time Data CF Jeff Lin, MD., PhD. Incidence 1. Cumulative incidence (incidence proportion) 2. Incidence density (incidence rate) December 14, 2005 c Jeff Lin, MD., PhD. c Jeff Lin, MD., PhD. Person-Time
Stat 642, Lecture notes for 04/12/05 96
 Stat 642, Lecture notes for 04/12/05 96 Hosmer-Lemeshow Statistic The Hosmer-Lemeshow Statistic is another measure of lack of fit. Hosmer and Lemeshow recommend partitioning the observations into 10 equal
Stat 642, Lecture notes for 04/12/05 96 Hosmer-Lemeshow Statistic The Hosmer-Lemeshow Statistic is another measure of lack of fit. Hosmer and Lemeshow recommend partitioning the observations into 10 equal
CS Homework 2: Combinatorics & Discrete Events Due Date: September 25, 2018 at 2:20 PM
 CS1450 - Homework 2: Combinatorics & Discrete Events Due Date: September 25, 2018 at 2:20 PM Question 1 A website allows the user to create an 8-character password that consists of lower case letters (a-z)
CS1450 - Homework 2: Combinatorics & Discrete Events Due Date: September 25, 2018 at 2:20 PM Question 1 A website allows the user to create an 8-character password that consists of lower case letters (a-z)
Boardworks Ltd Evolution
 1 of 34 Boardworks Ltd 2011 Evolution 2 of 34 Boardworks Ltd 2011 Life on earth 3 of 34 Boardworks Ltd 2011 Life on earth began approximately 3,500 million years ago. What do you think the earliest life
1 of 34 Boardworks Ltd 2011 Evolution 2 of 34 Boardworks Ltd 2011 Life on earth 3 of 34 Boardworks Ltd 2011 Life on earth began approximately 3,500 million years ago. What do you think the earliest life
Local-Effect Games. Kevin Leyton-Brown. Moshe Tennenholtz. Stanford UBC. Technion
 Local-Effect Games Kevin Leyton-Brown Stanford UBC Moshe Tennenholtz Technion Computation-Friendly Game Representations In practice, interesting games are large; computing equilibrium is hard CS agenda
Local-Effect Games Kevin Leyton-Brown Stanford UBC Moshe Tennenholtz Technion Computation-Friendly Game Representations In practice, interesting games are large; computing equilibrium is hard CS agenda
Bayesian Learning in Undirected Graphical Models
 Bayesian Learning in Undirected Graphical Models Zoubin Ghahramani Gatsby Computational Neuroscience Unit University College London, UK http://www.gatsby.ucl.ac.uk/ Work with: Iain Murray and Hyun-Chul
Bayesian Learning in Undirected Graphical Models Zoubin Ghahramani Gatsby Computational Neuroscience Unit University College London, UK http://www.gatsby.ucl.ac.uk/ Work with: Iain Murray and Hyun-Chul
Proofs. Chapter 2 P P Q Q
 Chapter Proofs In this chapter we develop three methods for proving a statement. To start let s suppose the statement is of the form P Q or if P, then Q. Direct: This method typically starts with P. Then,
Chapter Proofs In this chapter we develop three methods for proving a statement. To start let s suppose the statement is of the form P Q or if P, then Q. Direct: This method typically starts with P. Then,
The Cross Product. MATH 311, Calculus III. J. Robert Buchanan. Fall Department of Mathematics. J. Robert Buchanan The Cross Product
 The Cross Product MATH 311, Calculus III J. Robert Buchanan Department of Mathematics Fall 2011 Introduction Recall: the dot product of two vectors is a scalar. There is another binary operation on vectors
The Cross Product MATH 311, Calculus III J. Robert Buchanan Department of Mathematics Fall 2011 Introduction Recall: the dot product of two vectors is a scalar. There is another binary operation on vectors
Feature selection and classifier performance in computer-aided diagnosis: The effect of finite sample size
 Feature selection and classifier performance in computer-aided diagnosis: The effect of finite sample size Berkman Sahiner, a) Heang-Ping Chan, Nicholas Petrick, Robert F. Wagner, b) and Lubomir Hadjiiski
Feature selection and classifier performance in computer-aided diagnosis: The effect of finite sample size Berkman Sahiner, a) Heang-Ping Chan, Nicholas Petrick, Robert F. Wagner, b) and Lubomir Hadjiiski
Proofs. Chapter 2 P P Q Q
 Chapter Proofs In this chapter we develop three methods for proving a statement. To start let s suppose the statement is of the form P Q or if P, then Q. Direct: This method typically starts with P. Then,
Chapter Proofs In this chapter we develop three methods for proving a statement. To start let s suppose the statement is of the form P Q or if P, then Q. Direct: This method typically starts with P. Then,
Lecture 01: Introduction
 Lecture 01: Introduction Dipankar Bandyopadhyay, Ph.D. BMTRY 711: Analysis of Categorical Data Spring 2011 Division of Biostatistics and Epidemiology Medical University of South Carolina Lecture 01: Introduction
Lecture 01: Introduction Dipankar Bandyopadhyay, Ph.D. BMTRY 711: Analysis of Categorical Data Spring 2011 Division of Biostatistics and Epidemiology Medical University of South Carolina Lecture 01: Introduction
PROBABILITY. Inference: constraint propagation
 PROBABILITY Inference: constraint propagation! Use the constraints to reduce the number of legal values for a variable! Possible to find a solution without searching! Node consistency " A node is node-consistent
PROBABILITY Inference: constraint propagation! Use the constraints to reduce the number of legal values for a variable! Possible to find a solution without searching! Node consistency " A node is node-consistent
Cristina Nita-Rotaru. CS355: Cryptography. Lecture 9: Encryption modes. AES
 CS355: Cryptography Lecture 9: Encryption modes. AES Encryption modes: ECB } Message is broken into independent blocks of block_size bits; } Electronic Code Book (ECB): each block encrypted separately.
CS355: Cryptography Lecture 9: Encryption modes. AES Encryption modes: ECB } Message is broken into independent blocks of block_size bits; } Electronic Code Book (ECB): each block encrypted separately.
Agreement Coefficients and Statistical Inference
 CHAPTER Agreement Coefficients and Statistical Inference OBJECTIVE This chapter describes several approaches for evaluating the precision associated with the inter-rater reliability coefficients of the
CHAPTER Agreement Coefficients and Statistical Inference OBJECTIVE This chapter describes several approaches for evaluating the precision associated with the inter-rater reliability coefficients of the
THE TWIN PRIMES CONJECTURE - SOLUTIONS Bertrand Wong, Eurotech, S pore,
 THE TWIN PRIMES CONJECTURE - SOLUTIONS Bertrand Wong, Eurotech, S pore, Email: bwong8@singnetcomsg ABSTRACT The author had published a paper on the solutions for the twin primes conjecture in an international
THE TWIN PRIMES CONJECTURE - SOLUTIONS Bertrand Wong, Eurotech, S pore, Email: bwong8@singnetcomsg ABSTRACT The author had published a paper on the solutions for the twin primes conjecture in an international
Pig organ transplants within 5 years
 www.breaking News English.com Ready-to-use ESL / EFL Lessons Pig organ transplants within 5 years URL: http://www.breakingnewsenglish.com/0509/050911-xenotransplant.html Today s contents The Article 2
www.breaking News English.com Ready-to-use ESL / EFL Lessons Pig organ transplants within 5 years URL: http://www.breakingnewsenglish.com/0509/050911-xenotransplant.html Today s contents The Article 2
The prime number is a natural number greater than 1, is divisible by itself and the only one.
 The prime number is a natural number greater than 1, is divisible by itself and the only one. And a set of prime numbers is finished. Euclid has proved that in 300 BC. Most Greeks did not consider the
The prime number is a natural number greater than 1, is divisible by itself and the only one. And a set of prime numbers is finished. Euclid has proved that in 300 BC. Most Greeks did not consider the
C Examples of Bayes Calculations
 There are two buckets containing poker chips. Bucket A has Bucket B has 700 red chips 300 blue chips 300 red chips 700 blue chips Flip a coin to choose a bucket at random. From the chosen bucket, select
There are two buckets containing poker chips. Bucket A has Bucket B has 700 red chips 300 blue chips 300 red chips 700 blue chips Flip a coin to choose a bucket at random. From the chosen bucket, select
Probability Notes (A) , Fall 2010
 Probability Notes (A) 18.310, Fall 2010 We are going to be spending around four lectures on probability theory this year. These notes cover approximately the first three lectures on it. Probability theory
Probability Notes (A) 18.310, Fall 2010 We are going to be spending around four lectures on probability theory this year. These notes cover approximately the first three lectures on it. Probability theory
CHL 5225H Advanced Statistical Methods for Clinical Trials: Multiplicity
 CHL 5225H Advanced Statistical Methods for Clinical Trials: Multiplicity Prof. Kevin E. Thorpe Dept. of Public Health Sciences University of Toronto Objectives 1. Be able to distinguish among the various
CHL 5225H Advanced Statistical Methods for Clinical Trials: Multiplicity Prof. Kevin E. Thorpe Dept. of Public Health Sciences University of Toronto Objectives 1. Be able to distinguish among the various
Supplementary Logic Notes CSE 321 Winter 2009
 1 Propositional Logic Supplementary Logic Notes CSE 321 Winter 2009 1.1 More efficient truth table methods The method of using truth tables to prove facts about propositional formulas can be a very tedious
1 Propositional Logic Supplementary Logic Notes CSE 321 Winter 2009 1.1 More efficient truth table methods The method of using truth tables to prove facts about propositional formulas can be a very tedious
Notes for Math 324, Part 20
 7 Notes for Math 34, Part Chapter Conditional epectations, variances, etc.. Conditional probability Given two events, the conditional probability of A given B is defined by P[A B] = P[A B]. P[B] P[A B]
7 Notes for Math 34, Part Chapter Conditional epectations, variances, etc.. Conditional probability Given two events, the conditional probability of A given B is defined by P[A B] = P[A B]. P[B] P[A B]
SAMPLE SIZE RE-ESTIMATION FOR ADAPTIVE SEQUENTIAL DESIGN IN CLINICAL TRIALS
 Journal of Biopharmaceutical Statistics, 18: 1184 1196, 2008 Copyright Taylor & Francis Group, LLC ISSN: 1054-3406 print/1520-5711 online DOI: 10.1080/10543400802369053 SAMPLE SIZE RE-ESTIMATION FOR ADAPTIVE
Journal of Biopharmaceutical Statistics, 18: 1184 1196, 2008 Copyright Taylor & Francis Group, LLC ISSN: 1054-3406 print/1520-5711 online DOI: 10.1080/10543400802369053 SAMPLE SIZE RE-ESTIMATION FOR ADAPTIVE
Languages, regular languages, finite automata
 Notes on Computer Theory Last updated: January, 2018 Languages, regular languages, finite automata Content largely taken from Richards [1] and Sipser [2] 1 Languages An alphabet is a finite set of characters,
Notes on Computer Theory Last updated: January, 2018 Languages, regular languages, finite automata Content largely taken from Richards [1] and Sipser [2] 1 Languages An alphabet is a finite set of characters,
Welcome! Webinar Biostatistics: sample size & power. Thursday, April 26, 12:30 1:30 pm (NDT)
 . Welcome! Webinar Biostatistics: sample size & power Thursday, April 26, 12:30 1:30 pm (NDT) Get started now: Please check if your speakers are working and mute your audio. Please use the chat box to
. Welcome! Webinar Biostatistics: sample size & power Thursday, April 26, 12:30 1:30 pm (NDT) Get started now: Please check if your speakers are working and mute your audio. Please use the chat box to
Analysis I. Classroom Notes. H.-D. Alber
 Analysis I Classroom Notes H-D Alber Contents 1 Fundamental notions 1 11 Sets 1 12 Product sets, relations 5 13 Composition of statements 7 14 Quantifiers, negation of statements 9 2 Real numbers 11 21
Analysis I Classroom Notes H-D Alber Contents 1 Fundamental notions 1 11 Sets 1 12 Product sets, relations 5 13 Composition of statements 7 14 Quantifiers, negation of statements 9 2 Real numbers 11 21
Section 1 (closed-book) Total points 30
 CS 454 Theory of Computation Fall 2011 Section 1 (closed-book) Total points 30 1. Which of the following are true? (a) a PDA can always be converted to an equivalent PDA that at each step pops or pushes
CS 454 Theory of Computation Fall 2011 Section 1 (closed-book) Total points 30 1. Which of the following are true? (a) a PDA can always be converted to an equivalent PDA that at each step pops or pushes
HMMT February 2018 February 10, 2018
 HMMT February 018 February 10, 018 Algebra and Number Theory 1. For some real number c, the graphs of the equation y = x 0 + x + 18 and the line y = x + c intersect at exactly one point. What is c? 18
HMMT February 018 February 10, 018 Algebra and Number Theory 1. For some real number c, the graphs of the equation y = x 0 + x + 18 and the line y = x + c intersect at exactly one point. What is c? 18
Explainable AI that Can be Used for Judgment with Responsibility
 Explain AI@Imperial Workshop Explainable AI that Can be Used for Judgment with Responsibility FUJITSU LABORATORIES LTD. 5th April 08 About me Name: Hajime Morita Research interests: Natural Language Processing
Explain AI@Imperial Workshop Explainable AI that Can be Used for Judgment with Responsibility FUJITSU LABORATORIES LTD. 5th April 08 About me Name: Hajime Morita Research interests: Natural Language Processing
1 Gambler s Ruin Problem
 1 Gambler s Ruin Problem Consider a gambler who starts with an initial fortune of $1 and then on each successive gamble either wins $1 or loses $1 independent of the past with probabilities p and q = 1
1 Gambler s Ruin Problem Consider a gambler who starts with an initial fortune of $1 and then on each successive gamble either wins $1 or loses $1 independent of the past with probabilities p and q = 1
Bayes Formula. MATH 107: Finite Mathematics University of Louisville. March 26, 2014
 Bayes Formula MATH 07: Finite Mathematics University of Louisville March 26, 204 Test Accuracy Conditional reversal 2 / 5 A motivating question A rare disease occurs in out of every 0,000 people. A test
Bayes Formula MATH 07: Finite Mathematics University of Louisville March 26, 204 Test Accuracy Conditional reversal 2 / 5 A motivating question A rare disease occurs in out of every 0,000 people. A test
Penalized Loss functions for Bayesian Model Choice
 Penalized Loss functions for Bayesian Model Choice Martyn International Agency for Research on Cancer Lyon, France 13 November 2009 The pure approach For a Bayesian purist, all uncertainty is represented
Penalized Loss functions for Bayesian Model Choice Martyn International Agency for Research on Cancer Lyon, France 13 November 2009 The pure approach For a Bayesian purist, all uncertainty is represented
OPTION-CLOSED GAMES RICHARD J. NOWAKOWSKI AND PAUL OTTAWAY
 Volume 6, Number 1, Pages 142 153 ISSN 1715-0868 OPTION-CLOSED GAMES RICHARD J. NOWAKOWSKI AND PAUL OTTAWAY Abstract. We consider the class of combinatorial games with the property that each player s move
Volume 6, Number 1, Pages 142 153 ISSN 1715-0868 OPTION-CLOSED GAMES RICHARD J. NOWAKOWSKI AND PAUL OTTAWAY Abstract. We consider the class of combinatorial games with the property that each player s move
32 Divisibility Theory in Integral Domains
 3 Divisibility Theory in Integral Domains As we have already mentioned, the ring of integers is the prototype of integral domains. There is a divisibility relation on * : an integer b is said to be divisible
3 Divisibility Theory in Integral Domains As we have already mentioned, the ring of integers is the prototype of integral domains. There is a divisibility relation on * : an integer b is said to be divisible
Applied Machine Learning Annalisa Marsico
 Applied Machine Learning Annalisa Marsico OWL RNA Bionformatics group Max Planck Institute for Molecular Genetics Free University of Berlin 22 April, SoSe 2015 Goals Feature Selection rather than Feature
Applied Machine Learning Annalisa Marsico OWL RNA Bionformatics group Max Planck Institute for Molecular Genetics Free University of Berlin 22 April, SoSe 2015 Goals Feature Selection rather than Feature
Practical Considerations Surrounding Normality
 Practical Considerations Surrounding Normality Prof. Kevin E. Thorpe Dalla Lana School of Public Health University of Toronto KE Thorpe (U of T) Normality 1 / 16 Objectives Objectives 1. Understand the
Practical Considerations Surrounding Normality Prof. Kevin E. Thorpe Dalla Lana School of Public Health University of Toronto KE Thorpe (U of T) Normality 1 / 16 Objectives Objectives 1. Understand the
Probability and Conditional Probability
 Probability and Conditional Probability Bret Hanlon and Bret Larget Department of Statistics University of Wisconsin Madison September 27 29, 2011 Probability 1 / 33 Parasitic Fish Case Study Example 9.3
Probability and Conditional Probability Bret Hanlon and Bret Larget Department of Statistics University of Wisconsin Madison September 27 29, 2011 Probability 1 / 33 Parasitic Fish Case Study Example 9.3
Check Your Understanding of the Lecture Material Finger Exercises with Solutions
 Check Your Understanding of the Lecture Material Finger Exercises with Solutions Instructor: David Dobor March 6, 2017 While the solutions to these questions are available, you are strongly encouraged
Check Your Understanding of the Lecture Material Finger Exercises with Solutions Instructor: David Dobor March 6, 2017 While the solutions to these questions are available, you are strongly encouraged
Leber s Hereditary Optic Neuropathy
 Leber s Hereditary Optic Neuropathy Dear Editor: It is well known that the majority of Leber s hereditary optic neuropathy (LHON) cases was caused by 3 mtdna primary mutations (m.3460g A, m.11778g A, and
Leber s Hereditary Optic Neuropathy Dear Editor: It is well known that the majority of Leber s hereditary optic neuropathy (LHON) cases was caused by 3 mtdna primary mutations (m.3460g A, m.11778g A, and
It applies to discrete and continuous random variables, and a mix of the two.
 3. Bayes Theorem 3.1 Bayes Theorem A straightforward application of conditioning: using p(x, y) = p(x y) p(y) = p(y x) p(x), we obtain Bayes theorem (also called Bayes rule) p(x y) = p(y x) p(x). p(y)
3. Bayes Theorem 3.1 Bayes Theorem A straightforward application of conditioning: using p(x, y) = p(x y) p(y) = p(y x) p(x), we obtain Bayes theorem (also called Bayes rule) p(x y) = p(y x) p(x). p(y)
The Lady Tasting Tea. How to deal with multiple testing. Need to explore many models. More Predictive Modeling
 The Lady Tasting Tea More Predictive Modeling R. A. Fisher & the Lady B. Muriel Bristol claimed she prefers tea added to milk rather than milk added to tea Fisher was skeptical that she could distinguish
The Lady Tasting Tea More Predictive Modeling R. A. Fisher & the Lady B. Muriel Bristol claimed she prefers tea added to milk rather than milk added to tea Fisher was skeptical that she could distinguish
Conditional probability
 CHAPTER 4 Conditional probability 4.1. Introduction Suppose there are 200 men, of which 100 are smokers, and 100 women, of which 20 are smokers. What is the probability that a person chosen at random will
CHAPTER 4 Conditional probability 4.1. Introduction Suppose there are 200 men, of which 100 are smokers, and 100 women, of which 20 are smokers. What is the probability that a person chosen at random will
EE512 Graphical Models Fall 2009
 EE512 Graphical Models Fall 2009 Prof. Jeff Bilmes University of Washington, Seattle Department of Electrical Engineering Fall Quarter, 2009 http://ssli.ee.washington.edu/~bilmes/ee512fa09 Lecture 3 -
EE512 Graphical Models Fall 2009 Prof. Jeff Bilmes University of Washington, Seattle Department of Electrical Engineering Fall Quarter, 2009 http://ssli.ee.washington.edu/~bilmes/ee512fa09 Lecture 3 -
Lecture 3: Error Correcting Codes
 CS 880: Pseudorandomness and Derandomization 1/30/2013 Lecture 3: Error Correcting Codes Instructors: Holger Dell and Dieter van Melkebeek Scribe: Xi Wu In this lecture we review some background on error
CS 880: Pseudorandomness and Derandomization 1/30/2013 Lecture 3: Error Correcting Codes Instructors: Holger Dell and Dieter van Melkebeek Scribe: Xi Wu In this lecture we review some background on error
Chapter 13 Meiosis and Sexual Reproduction
 Biology 110 Sec. 11 J. Greg Doheny Chapter 13 Meiosis and Sexual Reproduction Quiz Questions: 1. What word do you use to describe a chromosome or gene allele that we inherit from our Mother? From our Father?
Biology 110 Sec. 11 J. Greg Doheny Chapter 13 Meiosis and Sexual Reproduction Quiz Questions: 1. What word do you use to describe a chromosome or gene allele that we inherit from our Mother? From our Father?
Fooling Sets and. Lecture 5
 Fooling Sets and Introduction to Nondeterministic Finite Automata Lecture 5 Proving that a language is not regular Given a language, we saw how to prove it is regular (union, intersection, concatenation,
Fooling Sets and Introduction to Nondeterministic Finite Automata Lecture 5 Proving that a language is not regular Given a language, we saw how to prove it is regular (union, intersection, concatenation,
Hidden Markov Models (HMMs) November 14, 2017
 Hidden Markov Models (HMMs) November 14, 2017 inferring a hidden truth 1) You hear a static-filled radio transmission. how can you determine what did the sender intended to say? 2) You know that genes
Hidden Markov Models (HMMs) November 14, 2017 inferring a hidden truth 1) You hear a static-filled radio transmission. how can you determine what did the sender intended to say? 2) You know that genes
Chance Models & Probability
 1 Definition of Probability Chance Models & Probability Stacey Hancock A random process is one in which the outcome is unpredictable. We encounter random processes every day: will it rain today? how many
1 Definition of Probability Chance Models & Probability Stacey Hancock A random process is one in which the outcome is unpredictable. We encounter random processes every day: will it rain today? how many
Theory of Computation
 Theory of Computation Prof. Michael Mascagni Florida State University Department of Computer Science 1 / 33 This course aims to cover... the development of computability theory using an extremely simple
Theory of Computation Prof. Michael Mascagni Florida State University Department of Computer Science 1 / 33 This course aims to cover... the development of computability theory using an extremely simple
Probability, Entropy, and Inference / More About Inference
 Probability, Entropy, and Inference / More About Inference Mário S. Alvim (msalvim@dcc.ufmg.br) Information Theory DCC-UFMG (2018/02) Mário S. Alvim (msalvim@dcc.ufmg.br) Probability, Entropy, and Inference
Probability, Entropy, and Inference / More About Inference Mário S. Alvim (msalvim@dcc.ufmg.br) Information Theory DCC-UFMG (2018/02) Mário S. Alvim (msalvim@dcc.ufmg.br) Probability, Entropy, and Inference
Probability and Statistics. Terms and concepts
 Probability and Statistics Joyeeta Dutta Moscato June 30, 2014 Terms and concepts Sample vs population Central tendency: Mean, median, mode Variance, standard deviation Normal distribution Cumulative distribution
Probability and Statistics Joyeeta Dutta Moscato June 30, 2014 Terms and concepts Sample vs population Central tendency: Mean, median, mode Variance, standard deviation Normal distribution Cumulative distribution
Time: 1 hour 30 minutes
 Paper Reference(s) 6684/01 Edexcel GCE Statistics S2 Bronze Level B4 Time: 1 hour 30 minutes Materials required for examination papers Mathematical Formulae (Green) Items included with question Nil Candidates
Paper Reference(s) 6684/01 Edexcel GCE Statistics S2 Bronze Level B4 Time: 1 hour 30 minutes Materials required for examination papers Mathematical Formulae (Green) Items included with question Nil Candidates
Expression QTLs and Mapping of Complex Trait Loci. Paul Schliekelman Statistics Department University of Georgia
 Expression QTLs and Mapping of Complex Trait Loci Paul Schliekelman Statistics Department University of Georgia Definitions: Genes, Loci and Alleles A gene codes for a protein. Proteins due everything.
Expression QTLs and Mapping of Complex Trait Loci Paul Schliekelman Statistics Department University of Georgia Definitions: Genes, Loci and Alleles A gene codes for a protein. Proteins due everything.
Time: 1 hour 30 minutes
 Paper Reference(s) 668/0 Edexcel GCE Statistics S Silver Level S2 Time: hour 0 minutes Materials required for examination papers Mathematical Formulae (Green) Items included with question Nil Candidates
Paper Reference(s) 668/0 Edexcel GCE Statistics S Silver Level S2 Time: hour 0 minutes Materials required for examination papers Mathematical Formulae (Green) Items included with question Nil Candidates
Bayesian decision making
 Bayesian decision making Václav Hlaváč Czech Technical University in Prague Czech Institute of Informatics, Robotics and Cybernetics 166 36 Prague 6, Jugoslávských partyzánů 1580/3, Czech Republic http://people.ciirc.cvut.cz/hlavac,
Bayesian decision making Václav Hlaváč Czech Technical University in Prague Czech Institute of Informatics, Robotics and Cybernetics 166 36 Prague 6, Jugoslávských partyzánů 1580/3, Czech Republic http://people.ciirc.cvut.cz/hlavac,
Chapter 2: Describing Contingency Tables - II
 : Describing Contingency Tables - II Dipankar Bandyopadhyay Department of Biostatistics, Virginia Commonwealth University BIOS 625: Categorical Data & GLM [Acknowledgements to Tim Hanson and Haitao Chu]
: Describing Contingency Tables - II Dipankar Bandyopadhyay Department of Biostatistics, Virginia Commonwealth University BIOS 625: Categorical Data & GLM [Acknowledgements to Tim Hanson and Haitao Chu]
MATH 19B FINAL EXAM PROBABILITY REVIEW PROBLEMS SPRING, 2010
 MATH 9B FINAL EXAM PROBABILITY REVIEW PROBLEMS SPRING, 00 This handout is meant to provide a collection of exercises that use the material from the probability and statistics portion of the course The
MATH 9B FINAL EXAM PROBABILITY REVIEW PROBLEMS SPRING, 00 This handout is meant to provide a collection of exercises that use the material from the probability and statistics portion of the course The
Biology New Jersey 1. NATURE OF LIFE 2. THE CHEMISTRY OF LIFE. Tutorial Outline
 Tutorial Outline New Jersey Tutorials are designed specifically for the New Jersey Core Curriculum Content Standards to prepare students for the PARCC assessments, the New Jersey Biology Competency Test
Tutorial Outline New Jersey Tutorials are designed specifically for the New Jersey Core Curriculum Content Standards to prepare students for the PARCC assessments, the New Jersey Biology Competency Test
PHASES OF STATISTICAL ANALYSIS 1. Initial Data Manipulation Assembling data Checks of data quality - graphical and numeric
 PHASES OF STATISTICAL ANALYSIS 1. Initial Data Manipulation Assembling data Checks of data quality - graphical and numeric 2. Preliminary Analysis: Clarify Directions for Analysis Identifying Data Structure:
PHASES OF STATISTICAL ANALYSIS 1. Initial Data Manipulation Assembling data Checks of data quality - graphical and numeric 2. Preliminary Analysis: Clarify Directions for Analysis Identifying Data Structure:
