Supporting information
|
|
|
- Barbara Owen
- 5 years ago
- Views:
Transcription
1 Supporting information Fluorescent derivatives of AC-42 to probe bitopic orthosteric/allosteric binding mechanisms on muscarinic M1 receptors Sandrine B. Daval, Céline Valant, Dominique Bonnet, Esther Kellenberger, Marcel Hibert, Jean-Luc Galzi and Brigitte Ilien * Table of contents : Supplemental Figure S1 : Normalized absorption and emission spectra of EGFP and para-lrb-ac42 in Hepes buffer. Supplemental Figure S2 : Structural variation of the M1 receptor. Supplemental Figure S3 : Sequence alignment for M1 homology modelling. Supplemental Procedures : Solubility measurements Estimation of donor-acceptor distances through FRET Biological data analyses S1
2 Supplemental Figure S1 Normalized absorption and emission spectra of EGFP and para-lrb-ac42 in Hepes buffer. Absorption (open symbols; dashed line) and fluorescence (closed symbols; solid line) spectra for EGFP (triangles) and para-lrb-ac42 (circles) are shown together with the overlap (grey area) for normalized EGFP emission and para-lrb-ac42 absorption. Arrows at 470 nm and 510 nm point the excitation and emission wavelengths, respectively, which were selected for FRET studies. S2
3 Supplemental Figure S2 Structural variation of the M1 receptor. Left and right panels show views from the extracellular side or in the plane of the plasma membrane, respectively. The 7TMs are represented by cylinders. The side chains of key residues discussed in the manuscript are displayed as capped sticks. The M1 receptor structure modeled by homology to the D3 receptor is colored in white, the structure obtained after a short molecular dynamic simulation is colored in green. The molecular dynamics was carried out using AMBER9 (University of California, San Francisco, CA, USA) as follows. The receptor was embedded in a pre-equilibrated lipid bilayer consisting of 82 molecules of 1-palmitoyl-2-oleoylphosphatidylcholine (POPC) and solvated with TIP3P water molecules (box dimensions: 88.2 Å x 93.1 Å x 83.8 Å). Energy minimization was first applied to solute molecules (by setting a positional harmonic constraint of 100 kcal/mol. Å on the receptor), then to the entire system. For the MD, the system was heated up with a linear increase of temperature from 100 to 300 K over 20 ps (periodic boundary with constant volume and no pressure control). A second equilibration stage of 100 ps at 300 K was performed using pressure and temperature control. The system was then subjected to a 1.8 ns simulation at 300 K. During all simulations, all bonds involving hydrogen atoms were frozen with the SHAKE algorithm. Interactions were calculated according to the AMBER 03 force field, using particle-mesh-ewald summation to include the long range electrostatic forces. The Van der Waals interactions were calculated using a cut-off of 8.0 Å. S3
4 Supplemental Figure S3 Sequence alignment for M1 homology modeling. Amino acids are coloured according to their physico-chemical properties. Sequence variations introduced in the recombinant proteins used for structure determination are written in lowercase (black letters). The cysteine residues involved in the conserved disulfide bridge (between the third TM and the second extracellular loop) are highlighted using a grey background. For the sake of clarity, long stretches of non conserved sequences were omitted. They are given below the alignment TM SWISSPROT ACM1_HUMAN...MNTSAPPAVSPNITVLAPGKGPWQVAFIGITTGLLSLATVTGN PDB 3PBL, chain A...dykddddgapASLSQLSSHLNYTCGAENSTGASQARPHAYYALSYCALILAIVFGN TM SWISSPROT ACM1_HUMAN LLVLISFKVNTELKTVNNYFLLSLACADLIIGTFSMNLYTTYLLMG.HWALGTLACDLWLA PDB 3PBL, chain A GLVCMAVLKERALQTTTNYLVVSLAVADLLVATLVMPWVVYLEVTGGVWNFSRICCDVFVT -TM TM SWISSPROT ACM1_HUMAN LDYVASNASVMNLLLISFDRYFSVTRPLSYR...AKRTPRRAALMIGLAWLVSFVLWAPAI PDB 3PBL, chain A LDVMMCTASIwNLCAISIDRYTAVVMPVHYQHGTGQSSCRRVALMITAVWVLAFAVSCPL. PDB 2RH1, chain A...QM TM SWISSPROT ACM1_HUMAN LFWQ...YLVGERTVLAGQCYIQFL...SQPIITFGTAMAAFYLPVTVMCTLYWRI PDB 3PBL, chain A LFGF...NTTGDPTVCSI...SNPDFVIYSSVVSFYLPFGVTVLVYARI PDB 2RH1, chain A HWYRATHQEAINCYAeETC... PDB 3EML...CLFEDV...VPMN SWISSPROT ACM1_HUMAN YRETENRARELAALQGSETPGKGGGSS...icl3 *...LVKEKKAAR PDB 3PBL, chain A YVVLKQRRRK...icl3 insert**...plrekkatq TM TM SWISSPROT ACM1_HUMAN TLSAILLAFILTWTPYNIMVLVSTFCKDC..VPETLWELGYWLCYVNSTINPMCYALCNK PDB 3PBL, chain A MVAIVLGAFIVCWLPFFLTHVLNTHCQTC.HVSPELYSATTWLGYVNSALNPVIYTTFNI SWISSPROT ACM1_HUMAN AFRDTFRLLLLCRWDKRRWRKIPKRPGSVHRTPSRQC... PDB 3PBL, chain A EFRKAFLKILSCgrplevlfq *SWISSPROT ACM1_HUMAN, icl3 SSSERSQPGAEGSPETPPGRCCRCCRAPRLLQAYSWKEEEEEDEGSMESLTSSEGEEPGSEVVIKMPMVDPEAQAPTKQPPRS SPNTVKRPTKKGRDRAGKGQKPRGKEQLAKRKTFS **PDB 3PBL, chain A, icl3 insert NIFEMLRIDEGLRLKIYKDTEGYYTIGIGHLLTKSPSLNAAKSELDKAIGRNTNGVITKDEAEKLFNQDVDAAVRGILRNAKL KPVYDSLDAVRRAALINMVFQMGETGVAGFTNSLRMLQQKRWDEAAVNLAKSRWYNQTPNRAKRVITTFRTGTWDAYGV S4
5 Supplemental Procedures Solubility measurements. Aliquots (1.5 µl) of DMSO stock solutions of AC-42, para-lrb-nbutyl, para-lrb-ac42 and ortho-lrb-ac42 were submitted to a 100 fold dilution either in Hepes buffer (10 mm Hepes, mm NaCl, 1.25 mm MgCl 2, 1.25 mm CaCl 2, 6 mm KCl and 10 mm glucose, 1 mg/ml bovine serum albumin; ph 7.4) or in CH 3 CN:H 2 O (1:1; v/v) to reach final concentrations ranging from 10 to 100µM. After mixing, samples were allowed to stand in the dark for 2 h at room temperature, in plastic test tubes used for binding assays. Following centrifugation at 15,000g for 10 min, supernatants were collected, diluted with an equivalent volume of CH 3 CN:H 2 O (1:1; v/v) in order to precipitate BSA contained in Hepes samples, and centrifuged again. Final clear supernatants were obtained and 20 µl of each of them were injected onto a Luna C18(2) column (4.6 x 50 mm; 5 µ; Phenomenex) for analytical RP-HPLC analysis. Elution was performed using a linear gradient (100% A for 0.2 min then 0 to 100% B in 2.5 min, then 100% B for 0.5 min, flow rate 2 ml/min) of solvent B (CH 3 CN 100%, 0.05% TFA, v/v) in solvent A (H 2 O 100%, 0.05% TFA, v/v). Detection was set at 280 and 565 nm. All compounds eluted as single peaks at retention times (given in min) that did not differ between assay (Hepes) or reference (CH 3 CN:H 2 O) series : AC-42 (2.03), para-lrb-nbutyl (2.00), para-lrb-ac42 (2.16) and ortho-lrb-ac42 (2.08). Peak surface areas of reference samples were calculated and plotted as a function of compound concentration to generate calibration curves. The strict linear dependency of peak surface areas with compound concentration (which was verified in all cases) allowed the absolute concentrations of assay samples to be quantified from their proper peak surface areas. Concentrations (mean values for two separate determinations) of compounds remaining in solution (following dilution in Hepes buffer), as opposed to theoretical concentrations (reference samples; 10, 30, 100 µm) were as follows : AC-42 (9.8, n.d., 98.1), para-lrb-nbutyl (2.5, 2.4, 2.2), para-lrb-ac42 (9.8, 29.5, 44.9) and ortho-lrb-ac42 (2.1, 2.7, 1.8). Estimation of donor-acceptor distances. The distance R 0 (Förster radius) for 50% transfer efficiency E, typical for each donor (EGFP)-acceptor (ligand fluorophore) pair, was calculated from: R 0 = 9790 (κ 2 n -4 Φ D J) 1/6. 63 Values for κ 2, a geometric factor accounting for the relative orientation in space of the donor emission and acceptor absorption transition dipoles, for N, the refractive index of the medium and for Φ D, the quantum yield of the EGFP donor together with calculation of J, the spectral overlap integral for the combined emission of EGFP and absorbance of fluorophore species, were as reported. 38 The distance R (Å) between EGFP and the fluorophore moiety of bound ligands was calculated using Förster s equation: R = R 0 (1/E 1) 1/6. 49 The efficiency of fluorescence energy transfer (E) was defined as the fractional decrease in EGFP fluorescence due to ligand binding and was expressed by: E = 1 F DA /F D, where F DA and F D are specific donor fluorescence emission in the presence or the absence of saturating concentrations of ligand, respectively. Specific EGFP fluorescence in the absence (F D ) or the presence (F DA ) of ligand was determined by subtracting autofluorescence of non transfected HEK cells (in the absence or presence of ligand) from total fluorescence of EGFP(Δ17)hM1 expressing cells. Biological data analyses. Nonlinear regression analyses of functional or binding data were performed using Kaleidagraph 4.0 (Synergy software, Reading, PA). Functional assays : Dose-effect curves were generated by plotting the normalized response amplitude, E, as a function of agonist or inhibitor concentration, X. Data were analysed using the equation : E = E max /(1 + (X 50 / X) nh ), to derive the maximal response E max, the drug concentration leading to halfmaximal response (EC 50 ) or inhibition (IC 50 ), and nh the slope factor. Competition-type experiments : Fractional receptor occupancy B/B 0 was plotted against the concentration X of unlabelled agent, on a logarithmic scale, and analysed using the general Hill equation : B/ B 0 = Bottom + [(1-Bottom) /(1 + (IC 50 / X) nh )], to estimate the bottom plateau, the IC 50 value and the slope factor nh. With bottom and slope values not significantly different from 0 and 1, respectively, interactions were assumed to be competitive and IC 50 values were converted into K i values by applying the Cheng and Prusoff correction. 64 S5
6 Competition curves with submaximal inhibition at high displacer concentration were analysed using the equation : B/B 0 = (L + K d ) / [L + K d.((1+(x / K X ) / (1+(X / α.k X )))], based on the allosteric ternary complex model of ligand interaction. 8 L and X are the concentrations of tracer and allosteric agent, respectively. K d and K X are equilibrium dissociation constants for the binding of the tracer and the allosteric drug at the free receptor, respectively. α denotes the cooperativity factor by which the two drugs mutually increase (α >1; negative cooperativity) or decrease (α <1; positive cooperativity) their respective equilibrium dissociation constants. Saturation experiments : Occupancy curves were generated by plotting specific tracer binding (B) at equilibrium, as a function of X, the radioactive or fluorescent tracer concentration. Data fitting to the empirical Hill equation derived for saturation : B = (B max ) /(1 + (K d / X) nh ), yields estimates of B max (density of receptor sites or maximal FRET amplitude), of K d (apparent equilibrium dissociation constant) and of nh (the slope factor). [ 3 H]-NMS dissociation kinetics : Off-rate constants (k off values) were determined from kinetic data fitting to a monoexponential time course reaction : B/B 0 = exp (-k off. t), where B 0 and B denote [ 3 H]- NMS binding at equilibrium (t = 0) and after t min of dissociation, respectively. Concentration-effect curves for the allosteric retardation of radioligand dissociation rate were generated by plotting k off obs / k off control ratios (i.e. dissociation rate constants measured in the presence and the absence of an allosteric agent) as a function of X, the alloster concentration. Fitting to the equation : k off obs / k off control = Bottom + [(1-Bottom) /(1 + (IC 50diss / X) nh )], allows the determination of the maximal amplitude E max diss (or (1-Bottom) value) of the retardation effect, of the alloster concentration IC 50diss leading to half-maximal reduction of [ 3 H]-NMS dissociation rate and of the slope factor nh. The IC 50diss parameter denotes the affinity (α.k X ) of the alloster for tracer-occupied receptors. 40 Additional References (63) Lakowicz, J. R. Principles of fluorescence spectroscopy; Kluwer Academic/Plenum Publishers: New York, (64) Cheng, Y. C.; Prusoff, W. H. Relationship between the inhibition constant (K i ) and the concentration of inhibitor which causes 50% inhibition (IC 50 ) of an enzymatic reaction. Biochem. Pharmacol. 1973, 22, S6
Supplementary figure 1 Application of tmfret in LeuT. (a) To assess the feasibility of using tmfret for distance-dependent measurements in LeuT, a
 Supplementary figure 1 Application of tmfret in LeuT. (a) To assess the feasibility of using tmfret for distance-dependent measurements in LeuT, a series of tmfret-pairs comprised of single cysteine mutants
Supplementary figure 1 Application of tmfret in LeuT. (a) To assess the feasibility of using tmfret for distance-dependent measurements in LeuT, a series of tmfret-pairs comprised of single cysteine mutants
A primer on pharmacology pharmacodynamics
 A primer on pharmacology pharmacodynamics Drug binding & effect Universidade do Algarve Faro 2017 by Ferdi Engels, Ph.D. 1 Pharmacodynamics Relation with pharmacokinetics? dosage plasma concentration site
A primer on pharmacology pharmacodynamics Drug binding & effect Universidade do Algarve Faro 2017 by Ferdi Engels, Ph.D. 1 Pharmacodynamics Relation with pharmacokinetics? dosage plasma concentration site
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Crystallization. a, Crystallization constructs of the ET B receptor are shown, with all of the modifications to the human wild-type the ET B receptor indicated. Residues interacting
Supplementary Figure 1 Crystallization. a, Crystallization constructs of the ET B receptor are shown, with all of the modifications to the human wild-type the ET B receptor indicated. Residues interacting
LanthaScreen Eu Kinase Binding Assay Validation Packet. Optimization of a LanthaScreen Eu Kinase Binding Assay for AURKB
 Page 1 of 18 LanthaScreen Eu Kinase Binding Assay for AURKB Overview This protocol describes how to perform a LanthaScreen Eu Kinase Binding Assay designed to detect and characterize kinase inhibitors.
Page 1 of 18 LanthaScreen Eu Kinase Binding Assay for AURKB Overview This protocol describes how to perform a LanthaScreen Eu Kinase Binding Assay designed to detect and characterize kinase inhibitors.
Supplemental Materials and Methods
 Supplemental Materials and Methods Time-resolved FRET (trfret) to probe for changes in the Box A/A stem upon complex assembly U3 MINI was folded and the decay of Fl fluorescence was measured at 20 ºC (see
Supplemental Materials and Methods Time-resolved FRET (trfret) to probe for changes in the Box A/A stem upon complex assembly U3 MINI was folded and the decay of Fl fluorescence was measured at 20 ºC (see
LanthaScreen Eu Kinase Binding Assay Validation Packet. Optimization of a LanthaScreen Eu Kinase Binding Assay for RPS6KA1
 Page 1 of 18 Optimization of a Assay for RPS6KA1 Assay for RPS6KA1 Overview This protocol describes how to perform a Assay designed to detect and characterize kinase inhibitors. Procedure 1 describes an
Page 1 of 18 Optimization of a Assay for RPS6KA1 Assay for RPS6KA1 Overview This protocol describes how to perform a Assay designed to detect and characterize kinase inhibitors. Procedure 1 describes an
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11085 Supplementary Tables: Supplementary Table 1. Summary of crystallographic and structure refinement data Structure BRIL-NOP receptor Data collection Number of crystals 23 Space group
doi:10.1038/nature11085 Supplementary Tables: Supplementary Table 1. Summary of crystallographic and structure refinement data Structure BRIL-NOP receptor Data collection Number of crystals 23 Space group
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature11524 Supplementary discussion Functional analysis of the sugar porter family (SP) signature motifs. As seen in Fig. 5c, single point mutation of the conserved
SUPPLEMENTARY INFORMATION doi:10.1038/nature11524 Supplementary discussion Functional analysis of the sugar porter family (SP) signature motifs. As seen in Fig. 5c, single point mutation of the conserved
Supplementary Methods
 Supplementary Methods MMPBSA Free energy calculation Molecular Mechanics/Poisson Boltzmann Surface Area (MM/PBSA) has been widely used to calculate binding free energy for protein-ligand systems (1-7).
Supplementary Methods MMPBSA Free energy calculation Molecular Mechanics/Poisson Boltzmann Surface Area (MM/PBSA) has been widely used to calculate binding free energy for protein-ligand systems (1-7).
Nuclear Medicine Department, Academic Medical Center, Amsterdam, The Netherlands;
 [ 3 H]-Spiperone Competition Binding to Dopamine D2, D3 and D4 Receptors Jan-Peter van Wieringen 1 and Martin C. Michel 2* 1 Nuclear Medicine Department, Academic Medical Center, Amsterdam, The Netherlands;
[ 3 H]-Spiperone Competition Binding to Dopamine D2, D3 and D4 Receptors Jan-Peter van Wieringen 1 and Martin C. Michel 2* 1 Nuclear Medicine Department, Academic Medical Center, Amsterdam, The Netherlands;
LanthaScreen Eu Kinase Binding Assay Validation Packet
 Page of 7 LanthaScreen Eu kinase binding assay for RF Overview This protocol describes how to perform a LanthaScreen Eu Kinase indng ssay designed to detect and characterize kinase inhibitors. Procedure
Page of 7 LanthaScreen Eu kinase binding assay for RF Overview This protocol describes how to perform a LanthaScreen Eu Kinase indng ssay designed to detect and characterize kinase inhibitors. Procedure
SIMPLE MODEL Direct Binding Analysis
 Neurochemistry, 56:120:575 Dr. Patrick J. McIlroy Supplementary Notes SIMPLE MODEL Direct Binding Analysis The interaction of a (radio)ligand, L, with its receptor, R, to form a non-covalent complex, RL,
Neurochemistry, 56:120:575 Dr. Patrick J. McIlroy Supplementary Notes SIMPLE MODEL Direct Binding Analysis The interaction of a (radio)ligand, L, with its receptor, R, to form a non-covalent complex, RL,
CHAPTER 8 Analysis of FP Binding Data
 CHAPTER 8 Analysis of FP Binding Data Determination of Binding Constants............................................................8-2 Definitions.........................................................................8-2
CHAPTER 8 Analysis of FP Binding Data Determination of Binding Constants............................................................8-2 Definitions.........................................................................8-2
Analyzing Radioligand Binding Data
 Analyzing Radioligand Binding Data APPENDIX 3H A radioligand is a radioactively labeled drug that can associate with a receptor, transporter, enzyme, or any protein of interest. The term ligand derives
Analyzing Radioligand Binding Data APPENDIX 3H A radioligand is a radioactively labeled drug that can associate with a receptor, transporter, enzyme, or any protein of interest. The term ligand derives
MOLECULAR DRUG TARGETS
 MOLECULAR DRUG TARGETS LEARNING OUTCOMES At the end of this session student shall be able to: List different types of druggable targets Describe forces involved in drug-receptor interactions Describe theories
MOLECULAR DRUG TARGETS LEARNING OUTCOMES At the end of this session student shall be able to: List different types of druggable targets Describe forces involved in drug-receptor interactions Describe theories
LanthaScreen Eu Kinase Binding Assay Validation Packet. Optimization of a LanthaScreen Eu Kinase Binding Assay for STK33
 Page of 7 Optimization of a ssay for STK33 ssay for STK33 Overview This protocol describes how to perform a ssay designed to detect and characterize kinase inhibitors. Procedure describes an experiment
Page of 7 Optimization of a ssay for STK33 ssay for STK33 Overview This protocol describes how to perform a ssay designed to detect and characterize kinase inhibitors. Procedure describes an experiment
COMPUTER AIDED DRUG DESIGN (CADD) AND DEVELOPMENT METHODS
 COMPUTER AIDED DRUG DESIGN (CADD) AND DEVELOPMENT METHODS DRUG DEVELOPMENT Drug development is a challenging path Today, the causes of many diseases (rheumatoid arthritis, cancer, mental diseases, etc.)
COMPUTER AIDED DRUG DESIGN (CADD) AND DEVELOPMENT METHODS DRUG DEVELOPMENT Drug development is a challenging path Today, the causes of many diseases (rheumatoid arthritis, cancer, mental diseases, etc.)
Measurement of Ligand G Protein-coupled Receptor Interactions. Katie Leach, Celine Valant, Patrick M. Sexton and Arthur Christopoulos
 1 Measurement of Ligand G Protein-coupled Receptor Interactions Katie Leach, Celine Valant, Patrick M. Sexton and Arthur Christopoulos Drug Discovery Biology Laboratory, Monash Institute of Pharmaceutical
1 Measurement of Ligand G Protein-coupled Receptor Interactions Katie Leach, Celine Valant, Patrick M. Sexton and Arthur Christopoulos Drug Discovery Biology Laboratory, Monash Institute of Pharmaceutical
A mitochondria-targeting fluorescent probe for detection of mitochondrial labile Fe(II) ion
 Electronic Supplementary Material (ESI) for Metallomics. This journal is The Royal Society of Chemistry 2018 A mitochondria-targeting fluorescent probe for detection of mitochondrial labile Fe(II) ion
Electronic Supplementary Material (ESI) for Metallomics. This journal is The Royal Society of Chemistry 2018 A mitochondria-targeting fluorescent probe for detection of mitochondrial labile Fe(II) ion
Lec.1 Chemistry Of Water
 Lec.1 Chemistry Of Water Biochemistry & Medicine Biochemistry can be defined as the science concerned with the chemical basis of life. Biochemistry can be described as the science concerned with the chemical
Lec.1 Chemistry Of Water Biochemistry & Medicine Biochemistry can be defined as the science concerned with the chemical basis of life. Biochemistry can be described as the science concerned with the chemical
single-molecule fluorescence resonance energy transfer
 single-molecule fluorescence resonance energy transfer (2) determing the Förster radius: quantum yield, donor lifetime, spectral overlap, anisotropy michael börsch 26/05/2004 1 fluorescence (1) absorbance
single-molecule fluorescence resonance energy transfer (2) determing the Förster radius: quantum yield, donor lifetime, spectral overlap, anisotropy michael börsch 26/05/2004 1 fluorescence (1) absorbance
CD Basis Set of Spectra that is used is that derived from comparing the spectra of globular proteins whose secondary structures are known from X-ray
 CD Basis Set of Spectra that is used is that derived from comparing the spectra of globular proteins whose secondary structures are known from X-ray crystallography An example of the use of CD Modeling
CD Basis Set of Spectra that is used is that derived from comparing the spectra of globular proteins whose secondary structures are known from X-ray crystallography An example of the use of CD Modeling
Lecture 5. Anisotropy decay/data analysis. Enrico Gratton
 Lecture 5. Anisotropy decay/data analysis Enrico Gratton Anisotropy decay Energy-transfer distance distributions Time resolved spectra Excited-state reactions Basic physics concept in polarization The
Lecture 5. Anisotropy decay/data analysis Enrico Gratton Anisotropy decay Energy-transfer distance distributions Time resolved spectra Excited-state reactions Basic physics concept in polarization The
Fluorescence Spectroscopy
 Fluorescence Spectroscopy Thomas Schmidt Department of Biophysics Leiden University, The Netherlands tschmidt@biophys.leidenuniv.nl www.biophys.leidenuniv.nl/research/fvl Biophysical Structural Biology
Fluorescence Spectroscopy Thomas Schmidt Department of Biophysics Leiden University, The Netherlands tschmidt@biophys.leidenuniv.nl www.biophys.leidenuniv.nl/research/fvl Biophysical Structural Biology
Caspase-1 Specific Light-up Probe with Aggregation-Induced Emission. Characteristics for Inhibitor Screening of Coumarin-Originated Natural.
 Supporting Information Caspase-1 Specific Light-up Probe with Aggregation-Induced Emission Characteristics for Inhibitor Screening of Coumarin-Originated Natural Products Hao Lin, ^ Haitao Yang, ^ Shuai
Supporting Information Caspase-1 Specific Light-up Probe with Aggregation-Induced Emission Characteristics for Inhibitor Screening of Coumarin-Originated Natural Products Hao Lin, ^ Haitao Yang, ^ Shuai
Supporting information for
 Supporting information for Rewiring multi-domain protein switches: transforming a fluorescent Zn 2+ -sensor into a light-responsive Zn 2+ binding protein Stijn J.A. Aper and Maarten Merkx Laboratory of
Supporting information for Rewiring multi-domain protein switches: transforming a fluorescent Zn 2+ -sensor into a light-responsive Zn 2+ binding protein Stijn J.A. Aper and Maarten Merkx Laboratory of
A Fluorescence Turn-On Sensor for the Detection of Palladium Ions that Operates Through In-Situ Generation of Palladium Nanoparticles
 This journal is The Royal Society of Chemistry 214 Supporting Information for A Fluorescence Turn-On Sensor for the Detection of Palladium Ions that Operates Through In-Situ Generation of Palladium Nanoparticles
This journal is The Royal Society of Chemistry 214 Supporting Information for A Fluorescence Turn-On Sensor for the Detection of Palladium Ions that Operates Through In-Situ Generation of Palladium Nanoparticles
schematic diagram; EGF binding, dimerization, phosphorylation, Grb2 binding, etc.
 Lecture 1: Noncovalent Biomolecular Interactions Bioengineering and Modeling of biological processes -e.g. tissue engineering, cancer, autoimmune disease Example: RTK signaling, e.g. EGFR Growth responses
Lecture 1: Noncovalent Biomolecular Interactions Bioengineering and Modeling of biological processes -e.g. tissue engineering, cancer, autoimmune disease Example: RTK signaling, e.g. EGFR Growth responses
Supporting Information
 Supporting Information Cyclodextrin Supramolecular Complex as Water Soluble Ratiometric Sensor for ferric Ion Sensing Meiyun Xu, Shuizhu Wu,* Fang Zeng, Changmin Yu College of Materials Science & Engineering,
Supporting Information Cyclodextrin Supramolecular Complex as Water Soluble Ratiometric Sensor for ferric Ion Sensing Meiyun Xu, Shuizhu Wu,* Fang Zeng, Changmin Yu College of Materials Science & Engineering,
Supplementary Figures
 1 Supplementary Figures Supplementary Figure 1 Type I FGFR1 inhibitors (a) Chemical structures of a pyrazolylaminopyrimidine inhibitor (henceforth referred to as PAPI; PDB-code of the FGFR1-PAPI complex:
1 Supplementary Figures Supplementary Figure 1 Type I FGFR1 inhibitors (a) Chemical structures of a pyrazolylaminopyrimidine inhibitor (henceforth referred to as PAPI; PDB-code of the FGFR1-PAPI complex:
Protein assay. Absorbance Fluorescence Emission Colorimetric detection BIO/MDT 325. Absorbance
 Protein assay Absorbance Fluorescence Emission Colorimetric detection BIO/MDT 325 Absorbance Using A280 to Determine Protein Concentration Determination of protein concentration by measuring absorbance
Protein assay Absorbance Fluorescence Emission Colorimetric detection BIO/MDT 325 Absorbance Using A280 to Determine Protein Concentration Determination of protein concentration by measuring absorbance
Interaction of Gold Nanoparticle with Proteins
 Chapter 7 Interaction of Gold Nanoparticle with Proteins 7.1. Introduction The interfacing of nanoparticle with biomolecules such as protein is useful for applications ranging from nano-biotechnology (molecular
Chapter 7 Interaction of Gold Nanoparticle with Proteins 7.1. Introduction The interfacing of nanoparticle with biomolecules such as protein is useful for applications ranging from nano-biotechnology (molecular
Analysis of Allosterism in Functional Assays a
 JPET This Fast article Forward. has not been Published copyedited and on formatted. July 26, The 2005 final as version DOI:10.1124/jpet.105.090886 may differ from this version. Analysis of Allosterism
JPET This Fast article Forward. has not been Published copyedited and on formatted. July 26, The 2005 final as version DOI:10.1124/jpet.105.090886 may differ from this version. Analysis of Allosterism
Optimization of a LanthaScreen Eu Kinase Binding Assay for PIK3CA/PIK3R1
 Page 1 of 18 LanthaScreen Eu Kinase Binding Assay for PIK3CA/PIK3R1 (p110α/p85α) Overview This protocol describes how to perform a LanthaScreen Eu Kinase Binding Assay designed to detect and characterize
Page 1 of 18 LanthaScreen Eu Kinase Binding Assay for PIK3CA/PIK3R1 (p110α/p85α) Overview This protocol describes how to perform a LanthaScreen Eu Kinase Binding Assay designed to detect and characterize
Appendix II- Bioanalytical Method Development and Validation
 A2. Bioanalytical method development 1. Optimization of chromatographic conditions Method development and optimization of chromatographic parameters is of utmost important for validating a method in biological
A2. Bioanalytical method development 1. Optimization of chromatographic conditions Method development and optimization of chromatographic parameters is of utmost important for validating a method in biological
Predictor Assay Setup Guide on the Tecan Safire 2 Microplate Reader
 Page 1 of 16 Predictor Assay Setup Guide on the Tecan Safire 2 Microplate Reader NOTE: The Tecan Safire 2 Microplate Reader was tested for compatibility with Invitrogen's Predictor herg FP Assay (PV5365)
Page 1 of 16 Predictor Assay Setup Guide on the Tecan Safire 2 Microplate Reader NOTE: The Tecan Safire 2 Microplate Reader was tested for compatibility with Invitrogen's Predictor herg FP Assay (PV5365)
Chapter 6. The interaction of Src SH2 with the focal adhesion kinase catalytic domain studied by NMR
 The interaction of Src SH2 with the focal adhesion kinase catalytic domain studied by NMR 103 Abstract The interaction of the Src SH2 domain with the catalytic domain of FAK, including the Y397 SH2 domain
The interaction of Src SH2 with the focal adhesion kinase catalytic domain studied by NMR 103 Abstract The interaction of the Src SH2 domain with the catalytic domain of FAK, including the Y397 SH2 domain
SUPPLEMENTARY INFORMATION
 UPPEER ORO doi:10.1038/nature10753 D D D D P E ntracellular C1 W P P C EC1 D Q R H C D W D R C C2 D E D E C R Q Q W P W W R P P EC2 EC3 P C C P W P W W P C W H R C R E C3 P R R P P P C Extracellular embrane
UPPEER ORO doi:10.1038/nature10753 D D D D P E ntracellular C1 W P P C EC1 D Q R H C D W D R C C2 D E D E C R Q Q W P W W R P P EC2 EC3 P C C P W P W W P C W H R C R E C3 P R R P P P C Extracellular embrane
type GroEL-GroES complex. Crystals were grown in buffer D (100 mm HEPES, ph 7.5,
 Supplementary Material Supplementary Materials and Methods Structure Determination of SR1-GroES-ADP AlF x SR1-GroES-ADP AlF x was purified as described in Materials and Methods for the wild type GroEL-GroES
Supplementary Material Supplementary Materials and Methods Structure Determination of SR1-GroES-ADP AlF x SR1-GroES-ADP AlF x was purified as described in Materials and Methods for the wild type GroEL-GroES
Potential Energy (hyper)surface
 The Molecular Dynamics Method Thermal motion of a lipid bilayer Water permeation through channels Selective sugar transport Potential Energy (hyper)surface What is Force? Energy U(x) F = " d dx U(x) Conformation
The Molecular Dynamics Method Thermal motion of a lipid bilayer Water permeation through channels Selective sugar transport Potential Energy (hyper)surface What is Force? Energy U(x) F = " d dx U(x) Conformation
Supporting Information
 Supporting Information Ottmann et al. 10.1073/pnas.0907587106 Fig. S1. Primary structure alignment of SBT3 with C5 peptidase from Streptococcus pyogenes. The Matchmaker tool in UCSF Chimera (http:// www.cgl.ucsf.edu/chimera)
Supporting Information Ottmann et al. 10.1073/pnas.0907587106 Fig. S1. Primary structure alignment of SBT3 with C5 peptidase from Streptococcus pyogenes. The Matchmaker tool in UCSF Chimera (http:// www.cgl.ucsf.edu/chimera)
Supporting Information
 Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2014 Supporting Information Performance comparison of two cascade reaction models in fluorescence
Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2014 Supporting Information Performance comparison of two cascade reaction models in fluorescence
SUPPLEMENTARY INFORMATION
 Ultrafast proton transport in sub-1-nm diameter carbon nanotube porins Ramya H. Tunuguntla, 1 Frances I. Allen, 2 Kyunghoon Kim, 1, Allison Belliveau, 1 Aleksandr Noy 1,3, * * Correspondence to AN: noy1@llnl.gov
Ultrafast proton transport in sub-1-nm diameter carbon nanotube porins Ramya H. Tunuguntla, 1 Frances I. Allen, 2 Kyunghoon Kim, 1, Allison Belliveau, 1 Aleksandr Noy 1,3, * * Correspondence to AN: noy1@llnl.gov
Solutions and Non-Covalent Binding Forces
 Chapter 3 Solutions and Non-Covalent Binding Forces 3.1 Solvent and solution properties Molecules stick together using the following forces: dipole-dipole, dipole-induced dipole, hydrogen bond, van der
Chapter 3 Solutions and Non-Covalent Binding Forces 3.1 Solvent and solution properties Molecules stick together using the following forces: dipole-dipole, dipole-induced dipole, hydrogen bond, van der
Advanced Fluorescence Microscopy I: Fluorescence (Foster) Resonance Energy Transfer
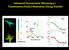 Advanced Fluorescence Microscopy I: Fluorescence (Foster) Resonance Energy Transfer 3.0 Pax 2.5 2.0 1200 800 400 GFP- Pax GFP-Pax + FATmCherry FAT FAT 1.5 Lifetime (ns) 0 1500 2000 2500 3000 Average lifetime
Advanced Fluorescence Microscopy I: Fluorescence (Foster) Resonance Energy Transfer 3.0 Pax 2.5 2.0 1200 800 400 GFP- Pax GFP-Pax + FATmCherry FAT FAT 1.5 Lifetime (ns) 0 1500 2000 2500 3000 Average lifetime
CAP 5510 Lecture 3 Protein Structures
 CAP 5510 Lecture 3 Protein Structures Su-Shing Chen Bioinformatics CISE 8/19/2005 Su-Shing Chen, CISE 1 Protein Conformation 8/19/2005 Su-Shing Chen, CISE 2 Protein Conformational Structures Hydrophobicity
CAP 5510 Lecture 3 Protein Structures Su-Shing Chen Bioinformatics CISE 8/19/2005 Su-Shing Chen, CISE 1 Protein Conformation 8/19/2005 Su-Shing Chen, CISE 2 Protein Conformational Structures Hydrophobicity
Application Note. Authors. Introduction. Lauren E. Frick and William A. LaMarr Agilent Technologies, Inc. Wakefield, MA, USA
 Fragment-Based Drug Discovery: Comparing Labeled and Label-Free Screening of β-amyloid Secretase (BACE-1) Using Fluorescence Spectroscopy and Ultrafast SPE/MS/MS Application Note Authors Lauren E. Frick
Fragment-Based Drug Discovery: Comparing Labeled and Label-Free Screening of β-amyloid Secretase (BACE-1) Using Fluorescence Spectroscopy and Ultrafast SPE/MS/MS Application Note Authors Lauren E. Frick
SUPPLEMENTARY INFORMATION
 Table of Contents Page Supplementary Table 1. Diffraction data collection statistics 2 Supplementary Table 2. Crystallographic refinement statistics 3 Supplementary Fig. 1. casic1mfc packing in the R3
Table of Contents Page Supplementary Table 1. Diffraction data collection statistics 2 Supplementary Table 2. Crystallographic refinement statistics 3 Supplementary Fig. 1. casic1mfc packing in the R3
Recommended Procedures for Labeling. Labeling Proteins with Amine-Reactive ATTO-Labels (NHS-Esters) Introduction
 Recommended Procedures for Labeling Introduction ATTO-TEC offers a large variety of high-quality dyes for labeling amino and thiol groups. ATTO reactive dyes cover the spectral region from 350 nm in the
Recommended Procedures for Labeling Introduction ATTO-TEC offers a large variety of high-quality dyes for labeling amino and thiol groups. ATTO reactive dyes cover the spectral region from 350 nm in the
Alkaline Phosphatase Labeling Kit-NH2
 Alkaline Phosphatase Labeling Kit-NH2 Catalog Number KA0001 1 Kit Version: 02 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 Principle of the Assay...
Alkaline Phosphatase Labeling Kit-NH2 Catalog Number KA0001 1 Kit Version: 02 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 Principle of the Assay...
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Identification of the ScDcp2 minimal region interacting with both ScDcp1 and the ScEdc3 LSm domain. Pull-down experiment of untagged ScEdc3 LSm with various ScDcp1-Dcp2-His 6 fragments.
Supplementary Figure 1 Identification of the ScDcp2 minimal region interacting with both ScDcp1 and the ScEdc3 LSm domain. Pull-down experiment of untagged ScEdc3 LSm with various ScDcp1-Dcp2-His 6 fragments.
Optimization of an Adapta Kinase Assay for CAMK1
 Overview Optimization of an Adapta Kinase Assay for CAMK1 This protocol describes how to perform an Adapta assay with the kinase CAMK1. To maximize the ability of the assay to detect ATP-competitive inhibitors,
Overview Optimization of an Adapta Kinase Assay for CAMK1 This protocol describes how to perform an Adapta assay with the kinase CAMK1. To maximize the ability of the assay to detect ATP-competitive inhibitors,
FLUORESCENCE STUDY ON TRYPTOPHAN POTASSIUM IODIDE INTERACTION
 PROCEEDINGS OF THE YEREVAN STATE UNIVERSITY C h e m i s t r y a n d B i o l o g y 08, 5(), p. 75 79 FLUORESCENCE STUDY ON TRYPTOPHAN POTASSIUM IODIDE INTERACTION C h emistr y H. A. SHILAJYAN, K. R. GRIGORYAN
PROCEEDINGS OF THE YEREVAN STATE UNIVERSITY C h e m i s t r y a n d B i o l o g y 08, 5(), p. 75 79 FLUORESCENCE STUDY ON TRYPTOPHAN POTASSIUM IODIDE INTERACTION C h emistr y H. A. SHILAJYAN, K. R. GRIGORYAN
Mathematical model of serodiagnostic immunochromatographic assay
 SUPPORTING INFORMATION Mathematical model of serodiagnostic immunochromatographic assay Dmitriy V. Sotnikov, Anatoly V. Zherdev, Boris B. Dzantiev* A.N. Bach Institute of Biochemistry, Federal Research
SUPPORTING INFORMATION Mathematical model of serodiagnostic immunochromatographic assay Dmitriy V. Sotnikov, Anatoly V. Zherdev, Boris B. Dzantiev* A.N. Bach Institute of Biochemistry, Federal Research
LanthaScreen Eu Kinase Binding Assay for GAK
 LanthaScreen Eu Kinase Binding Assay for GAK Catalog Number: A30973 Size: 10 µg Storage: 80 C, immediately upon receipt A32891 100 µg A33376 1 mg Pub. No. MAN0017224 Rev. A.0 Overview This user guide describes
LanthaScreen Eu Kinase Binding Assay for GAK Catalog Number: A30973 Size: 10 µg Storage: 80 C, immediately upon receipt A32891 100 µg A33376 1 mg Pub. No. MAN0017224 Rev. A.0 Overview This user guide describes
Supporting Information
 Supporting Information Monoamine oxidase isoform-dependent tautomeric influence in the recognition of 3,5 diaryl pyrazole inhibitors. Franco Chimenti, a Rossella Fioravanti,* a Adriana Bolasco, a Fedele
Supporting Information Monoamine oxidase isoform-dependent tautomeric influence in the recognition of 3,5 diaryl pyrazole inhibitors. Franco Chimenti, a Rossella Fioravanti,* a Adriana Bolasco, a Fedele
Problem solving steps
 Problem solving steps Determine the reaction Write the (balanced) equation ΔG K v Write the equilibrium constant v Find the equilibrium constant using v If necessary, solve for components K K = [ p ] ν
Problem solving steps Determine the reaction Write the (balanced) equation ΔG K v Write the equilibrium constant v Find the equilibrium constant using v If necessary, solve for components K K = [ p ] ν
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES Supplementary Figure 1 Protein sequence alignment of Vibrionaceae with either a 40-residue insertion or a 44-residue insertion. Identical residues are indicated by red background.
SUPPLEMENTARY FIGURES Supplementary Figure 1 Protein sequence alignment of Vibrionaceae with either a 40-residue insertion or a 44-residue insertion. Identical residues are indicated by red background.
Antibiotic Selectivity for Prokaryotic vs. Eukaryotic Decoding Sites Yun Xie, Andrew Dix, and Yitzhak Tor*
 Antibiotic Selectivity for Prokaryotic vs. Eukaryotic Decoding Sites Yun Xie, Andrew Dix, and Yitzhak Tor* Department of Chemistry and Biochemistry, University of California, San Diego, La Jolla, CA 92093-0358
Antibiotic Selectivity for Prokaryotic vs. Eukaryotic Decoding Sites Yun Xie, Andrew Dix, and Yitzhak Tor* Department of Chemistry and Biochemistry, University of California, San Diego, La Jolla, CA 92093-0358
Statistical concepts in QSAR.
 Statistical concepts in QSAR. Computational chemistry represents molecular structures as a numerical models and simulates their behavior with the equations of quantum and classical physics. Available programs
Statistical concepts in QSAR. Computational chemistry represents molecular structures as a numerical models and simulates their behavior with the equations of quantum and classical physics. Available programs
Chemistry in Biology. Section 1. Atoms, Elements, and Compounds
 Section 1 Atoms, Elements, and Compounds Atoms! Chemistry is the study of matter.! Atoms are the building blocks of matter.! Neutrons and protons are located at the center of the atom.! Protons are positively
Section 1 Atoms, Elements, and Compounds Atoms! Chemistry is the study of matter.! Atoms are the building blocks of matter.! Neutrons and protons are located at the center of the atom.! Protons are positively
Fluorescence (Notes 16)
 Fluorescence - 2014 (Notes 16) XV 74 Jablonski diagram Where does the energy go? Can be viewed like multistep kinetic pathway 1) Excite system through A Absorbance S 0 S n Excite from ground excited singlet
Fluorescence - 2014 (Notes 16) XV 74 Jablonski diagram Where does the energy go? Can be viewed like multistep kinetic pathway 1) Excite system through A Absorbance S 0 S n Excite from ground excited singlet
12 Nicarbazin Nicarbazin (4,4 -dinitro carbanilid (DNC) and 2-hydroxy-4,6-dimethyl pyrimidine (HDP))
 12 Nicarbazin Nicarbazin (4,4 -dinitro carbanilid (DNC) and 2-hydroxy-4,6-dimethyl pyrimidine (HDP)) O - O - O N + O N + O N NH N H N H O 1,3-bis(4-nitrophenyl)urea, 4,6-dimethyl-1H-pyrimidin-2-one C 13
12 Nicarbazin Nicarbazin (4,4 -dinitro carbanilid (DNC) and 2-hydroxy-4,6-dimethyl pyrimidine (HDP)) O - O - O N + O N + O N NH N H N H O 1,3-bis(4-nitrophenyl)urea, 4,6-dimethyl-1H-pyrimidin-2-one C 13
Supporting Information for. Jesinghaus, Rachael Barry, Zemer Gitai, Justin Kollman and Enoch P. Baldwin
 Supporting Information for Inhibition of E. coli CTP synthetase by NADH and other nicotinamides, and their mutual interactions with CTP and GTP Chris Habrian, Adithi Chandrasekhara, Bita Shahrvini, Brian
Supporting Information for Inhibition of E. coli CTP synthetase by NADH and other nicotinamides, and their mutual interactions with CTP and GTP Chris Habrian, Adithi Chandrasekhara, Bita Shahrvini, Brian
Enhancing Specificity in the Janus Kinases: A Study on the Thienopyridine. JAK2 Selective Mechanism Combined Molecular Dynamics Simulation
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2015 Supporting Information Enhancing Specificity in the Janus Kinases: A Study on the Thienopyridine
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2015 Supporting Information Enhancing Specificity in the Janus Kinases: A Study on the Thienopyridine
Supplementary Information. The protease GtgE from Salmonella exclusively targets. inactive Rab GTPases
 Supplementary Information The protease GtgE from Salmonella exclusively targets inactive Rab GTPases Table of Contents Supplementary Figures... 2 Supplementary Figure 1... 2 Supplementary Figure 2... 3
Supplementary Information The protease GtgE from Salmonella exclusively targets inactive Rab GTPases Table of Contents Supplementary Figures... 2 Supplementary Figure 1... 2 Supplementary Figure 2... 3
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11054 Supplementary Fig. 1 Sequence alignment of Na v Rh with NaChBac, Na v Ab, and eukaryotic Na v and Ca v homologs. Secondary structural elements of Na v Rh are indicated above the
doi:10.1038/nature11054 Supplementary Fig. 1 Sequence alignment of Na v Rh with NaChBac, Na v Ab, and eukaryotic Na v and Ca v homologs. Secondary structural elements of Na v Rh are indicated above the
SUPPLEMENTARY INFORMATION. doi: /nature07461
 Figure S1 Electrophysiology. a ph-activation of. Two-electrode voltage clamp recordings of Xenopus oocytes expressing in comparison to waterinjected oocytes. Currents were recorded at 40 mv. The ph of
Figure S1 Electrophysiology. a ph-activation of. Two-electrode voltage clamp recordings of Xenopus oocytes expressing in comparison to waterinjected oocytes. Currents were recorded at 40 mv. The ph of
Bradford Reagent, 5x
 INSTRUCTION MANUAL Bradford Reagent, 5x Reagent for protein quantification (Cat. No. 39222) SERVA Electrophoresis GmbH Carl-Benz-Str. 7 D-69115 HeidelbergPhone +49-6221-138400, Fax +49-6221-1384010 e-mail:
INSTRUCTION MANUAL Bradford Reagent, 5x Reagent for protein quantification (Cat. No. 39222) SERVA Electrophoresis GmbH Carl-Benz-Str. 7 D-69115 HeidelbergPhone +49-6221-138400, Fax +49-6221-1384010 e-mail:
S2004 Methods for characterization of biomolecular interactions - classical versus modern
 S2004 Methods for characterization of biomolecular interactions - classical versus modern Isothermal Titration Calorimetry (ITC) Eva Dubská email: eva.dubska@ceitec.cz Outline Calorimetry - history + a
S2004 Methods for characterization of biomolecular interactions - classical versus modern Isothermal Titration Calorimetry (ITC) Eva Dubská email: eva.dubska@ceitec.cz Outline Calorimetry - history + a
State Key Laboratory of Coordination Chemistry, School of Chemistry and Chemical
 Electronic Supplementary Information Evidence for inhibition of HIF-1α Prolyl hydroxylase 3 activity by four biological active tetraazamacrocycles Jing Cao, Zhirong Geng, Xiaoyan Ma, Jinghan Wen, Yuxin
Electronic Supplementary Information Evidence for inhibition of HIF-1α Prolyl hydroxylase 3 activity by four biological active tetraazamacrocycles Jing Cao, Zhirong Geng, Xiaoyan Ma, Jinghan Wen, Yuxin
Structural Bioinformatics (C3210) Molecular Mechanics
 Structural Bioinformatics (C3210) Molecular Mechanics How to Calculate Energies Calculation of molecular energies is of key importance in protein folding, molecular modelling etc. There are two main computational
Structural Bioinformatics (C3210) Molecular Mechanics How to Calculate Energies Calculation of molecular energies is of key importance in protein folding, molecular modelling etc. There are two main computational
Supporting Information. Labeled Ligand Displacement: Extending NMR-based Screening of Protein Targets
 Supporting Information Labeled Ligand Displacement: Extending NMR-based Screening of Protein Targets Steven L. Swann, Danying Song, Chaohong Sun, Philip J. Hajduk, and Andrew M. Petros Global Pharmaceutical
Supporting Information Labeled Ligand Displacement: Extending NMR-based Screening of Protein Targets Steven L. Swann, Danying Song, Chaohong Sun, Philip J. Hajduk, and Andrew M. Petros Global Pharmaceutical
Supporting Information
 Supporting Information Discovery of Hydrocarbon-Stapled Short α-helical Peptides as Promising Middle East Respiratory Syndrome Coronavirus (MERS-CoV) Fusion Inhibitors Chao Wang,, Shuai Xia,, Peiyu Zhang,
Supporting Information Discovery of Hydrocarbon-Stapled Short α-helical Peptides as Promising Middle East Respiratory Syndrome Coronavirus (MERS-CoV) Fusion Inhibitors Chao Wang,, Shuai Xia,, Peiyu Zhang,
Data sheet. UV-Tracer TM Biotin-Maleimide. For Labeling of Thiol-groups with UV-detectable Biotin CLK-B Description
 Cat. No. CLK-B105-10 CLK-B105-100 Amount 10 mg 100 mg For in vitro use only! Quality guaranteed for 12 months Store at -20 C 1. Description UV-Tracer TM Biotin Maleimide for biotinylation reactions of
Cat. No. CLK-B105-10 CLK-B105-100 Amount 10 mg 100 mg For in vitro use only! Quality guaranteed for 12 months Store at -20 C 1. Description UV-Tracer TM Biotin Maleimide for biotinylation reactions of
Supplemental Information. Molecular Basis of Spectral Diversity. in Near-Infrared Phytochrome-Based. Fluorescent Proteins
 Chemistry & Biology, Volume 22 Supplemental Information Molecular Basis of Spectral Diversity in Near-Infrared Phytochrome-Based Fluorescent Proteins Daria M. Shcherbakova, Mikhail Baloban, Sergei Pletnev,
Chemistry & Biology, Volume 22 Supplemental Information Molecular Basis of Spectral Diversity in Near-Infrared Phytochrome-Based Fluorescent Proteins Daria M. Shcherbakova, Mikhail Baloban, Sergei Pletnev,
Supporting Protocol This protocol describes the construction and the force-field parameters of the non-standard residue for the Ag + -site using CNS
 Supporting Protocol This protocol describes the construction and the force-field parameters of the non-standard residue for the Ag + -site using CNS CNS input file generatemetal.inp: remarks file generate/generatemetal.inp
Supporting Protocol This protocol describes the construction and the force-field parameters of the non-standard residue for the Ag + -site using CNS CNS input file generatemetal.inp: remarks file generate/generatemetal.inp
Amphiphilic diselenide-containing supramolecular polymers
 Electronic Supplementary Material (ESI) for Polymer Chemistry. This journal is The Royal Society of Chemistry 2014 Amphiphilic diselenide-containing supramolecular polymers Xinxin Tan, Liulin Yang, Zehuan
Electronic Supplementary Material (ESI) for Polymer Chemistry. This journal is The Royal Society of Chemistry 2014 Amphiphilic diselenide-containing supramolecular polymers Xinxin Tan, Liulin Yang, Zehuan
Water, water everywhere,; not a drop to drink. Consumption resulting from how environment inhabited Deforestation disrupts water cycle
 Chapter 3 Water: The Matrix of Life Overview n n n Water, water everywhere,; not a drop to drink Only 3% of world s water is fresh How has this happened Consumption resulting from how environment inhabited
Chapter 3 Water: The Matrix of Life Overview n n n Water, water everywhere,; not a drop to drink Only 3% of world s water is fresh How has this happened Consumption resulting from how environment inhabited
Lipid Regulated Intramolecular Conformational Dynamics of SNARE-Protein Ykt6
 Supplementary Information for: Lipid Regulated Intramolecular Conformational Dynamics of SNARE-Protein Ykt6 Yawei Dai 1, 2, Markus Seeger 3, Jingwei Weng 4, Song Song 1, 2, Wenning Wang 4, Yan-Wen 1, 2,
Supplementary Information for: Lipid Regulated Intramolecular Conformational Dynamics of SNARE-Protein Ykt6 Yawei Dai 1, 2, Markus Seeger 3, Jingwei Weng 4, Song Song 1, 2, Wenning Wang 4, Yan-Wen 1, 2,
Detection of Mercury(II) and Lead(II) with Graphene Oxide- Based Biosensors
 Detection of Mercury(II) and Lead(II) with Graphene Oxide- Based Biosensors Ming Li, Nianqiang (Nick) Wu* Mechanical & Aerospace Engineering West Virginia University Morgantown, WV 26506 Presentation Outline
Detection of Mercury(II) and Lead(II) with Graphene Oxide- Based Biosensors Ming Li, Nianqiang (Nick) Wu* Mechanical & Aerospace Engineering West Virginia University Morgantown, WV 26506 Presentation Outline
Polyethylene Glycol (PEG), High Sensitive ELISA
 K-ASSAY Polyethylene Glycol (PEG), High Sensitive ELISA For the high sensitive quantitative determination of PEG and PEGylated proteins in serum or plasma Cat. No. KT-657 For Research Use Only. 1 K-ASSAY
K-ASSAY Polyethylene Glycol (PEG), High Sensitive ELISA For the high sensitive quantitative determination of PEG and PEGylated proteins in serum or plasma Cat. No. KT-657 For Research Use Only. 1 K-ASSAY
Backbone modification of a parathyroid hormone receptor-1 antagonist/inverse agonist
 SUPPORTING INFORMATION Backbone modification of a parathyroid hormone receptor-1 antagonist/inverse agonist Ross W. Cheloha, Tomoyuki Watanabe, Thomas Dean, Samuel H. Gellman*, Thomas J. Gardella* *Corresponding
SUPPORTING INFORMATION Backbone modification of a parathyroid hormone receptor-1 antagonist/inverse agonist Ross W. Cheloha, Tomoyuki Watanabe, Thomas Dean, Samuel H. Gellman*, Thomas J. Gardella* *Corresponding
Malachite Green Phosphate Detection Kit Catalog Number: DY996
 Malachite Green Phosphate Detection Kit Catalog Number: DY996 This Malachite Green Phosphate Detection Kit employs a simple, sensitive, reproducible, and non-radioactive method for measuring inorganic
Malachite Green Phosphate Detection Kit Catalog Number: DY996 This Malachite Green Phosphate Detection Kit employs a simple, sensitive, reproducible, and non-radioactive method for measuring inorganic
Concept review: Binding equilibria
 Concept review: Binding equilibria 1 Binding equilibria and association/dissociation constants 2 The binding of a protein to a ligand at equilibrium can be written as: P + L PL And so the equilibrium constant
Concept review: Binding equilibria 1 Binding equilibria and association/dissociation constants 2 The binding of a protein to a ligand at equilibrium can be written as: P + L PL And so the equilibrium constant
Kinase-Glo Luminescent Kinase Assay Platform
 Technical Bulletin Kinase-Glo Luminescent Kinase Assay Platform INSTRUCTIONS FOR USE OF PRODUCTS V6711, V6712, V6713, V6714, V3771, V3772, V3773, V3774, V6071, V6072, V6073 AND V6074. PRINTED IN USA. Revised
Technical Bulletin Kinase-Glo Luminescent Kinase Assay Platform INSTRUCTIONS FOR USE OF PRODUCTS V6711, V6712, V6713, V6714, V3771, V3772, V3773, V3774, V6071, V6072, V6073 AND V6074. PRINTED IN USA. Revised
Supporting Information. Diffusion Limited Cargo Loading of an Engineered Protein Container Reinhard Zschoche and Donald Hilvert
 Supporting Information Diffusion Limited Cargo Loading of an Engineered Protein Container Reinhard Zschoche and Donald Hilvert Laboratory of Organic Chemistry, ETH Zürich, 8093 Zürich, Switzerland Supporting
Supporting Information Diffusion Limited Cargo Loading of an Engineered Protein Container Reinhard Zschoche and Donald Hilvert Laboratory of Organic Chemistry, ETH Zürich, 8093 Zürich, Switzerland Supporting
CHAPTER - 3 ANALYTICAL PROFILE. 3.1 Estimation of Drug in Pharmaceutical Formulation Estimation of Drugs
 CHAPTER - 3 ANALYTICAL PROFILE 3.1 Estimation of Drug in Pharmaceutical Formulation 3.1.1 Estimation of Drugs ANALYTICAL PROFILE 84 3.1 ESTIMATION OF DRUG IN PHARMACEUTICAL FORMULATION. Agrawal A et al
CHAPTER - 3 ANALYTICAL PROFILE 3.1 Estimation of Drug in Pharmaceutical Formulation 3.1.1 Estimation of Drugs ANALYTICAL PROFILE 84 3.1 ESTIMATION OF DRUG IN PHARMACEUTICAL FORMULATION. Agrawal A et al
BCA Protein Macro Assay Kit
 INSTRUCTION MANUAL BCA Protein Macro Assay Kit Kit for Protein Quantification (Cat. No. 39228) Contents 1. BCA PROTEIN MACRO ASSAY KIT 2 1.1. General information 2 1.2. Kit components 2 1.3. Additionally
INSTRUCTION MANUAL BCA Protein Macro Assay Kit Kit for Protein Quantification (Cat. No. 39228) Contents 1. BCA PROTEIN MACRO ASSAY KIT 2 1.1. General information 2 1.2. Kit components 2 1.3. Additionally
Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1.
 Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1. PDZK1 constru cts Amino acids MW [kda] KD [μm] PEPT2-CT- FITC KD [μm] NHE3-CT- FITC KD [μm] PDZK1-CT-
Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1. PDZK1 constru cts Amino acids MW [kda] KD [μm] PEPT2-CT- FITC KD [μm] NHE3-CT- FITC KD [μm] PDZK1-CT-
In vivo monitoring of hydrogen sulfide using a cresyl violet-based ratiometric fluorescence probe
 Electronic Supplementary Information for: In vivo monitoring of hydrogen sulfide using a cresyl violet-based ratiometric fluorescence probe Qiongqiong Wan, Yanchao Song, Zhao Li, Xinghui Gao and Huimin
Electronic Supplementary Information for: In vivo monitoring of hydrogen sulfide using a cresyl violet-based ratiometric fluorescence probe Qiongqiong Wan, Yanchao Song, Zhao Li, Xinghui Gao and Huimin
Enzymatic Assay of NITRIC OXIDE SYNTHETASE (EC ) ß-NADP = ß-Nicotinamide Adenine Dinucleotide Phosphate,
 PRINCIPLE: L-Arginine + ß-NADPH + O2 Nitric Oxide Synthetase > Citrulline + NO + ß-NADP Ca 2+, Tetrahydrobiopterin, Calmodulin Abbreviations used: ß-NADPH = ß-Nicotinamide Adenine Dinucleotide Phosphate,
PRINCIPLE: L-Arginine + ß-NADPH + O2 Nitric Oxide Synthetase > Citrulline + NO + ß-NADP Ca 2+, Tetrahydrobiopterin, Calmodulin Abbreviations used: ß-NADPH = ß-Nicotinamide Adenine Dinucleotide Phosphate,
The photoluminescent graphene oxide serves as an acceptor rather. than a donor in the fluorescence resonance energy transfer pair of
 Supplementary Material (ESI) for Chemical Communications This journal is (c) The Royal Society of Chemistry 20XX The photoluminescent graphene oxide serves as an acceptor rather than a donor in the fluorescence
Supplementary Material (ESI) for Chemical Communications This journal is (c) The Royal Society of Chemistry 20XX The photoluminescent graphene oxide serves as an acceptor rather than a donor in the fluorescence
BCA Protein Quantitation Kit
 BCA Protein Quantitation Kit Catalog Number KA3840 100 assays Version: 01 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 General Information... 4 Materials
BCA Protein Quantitation Kit Catalog Number KA3840 100 assays Version: 01 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 General Information... 4 Materials
Protocol for 2D-E. Protein Extraction
 Protocol for 2D-E Protein Extraction Reagent 1 inside the ReadyPrep TM Sequential Extraction kit (in powder form) 50ml of deionized water is used to dissolve all the Reagent 1. The solution is known as
Protocol for 2D-E Protein Extraction Reagent 1 inside the ReadyPrep TM Sequential Extraction kit (in powder form) 50ml of deionized water is used to dissolve all the Reagent 1. The solution is known as
A label-free DNA reduced graphene oxide-based fluorescent. sensor for highly sensitive and selective detection of hemin
 A label-free DNA reduced graphene oxide-based fluorescent sensor for highly sensitive and selective detection of hemin Yan Shi, Wei Tao Huang, Hong Qun Luo and Nian Bing Li* Key Laboratory on Luminescence
A label-free DNA reduced graphene oxide-based fluorescent sensor for highly sensitive and selective detection of hemin Yan Shi, Wei Tao Huang, Hong Qun Luo and Nian Bing Li* Key Laboratory on Luminescence
Supporting Information
 Supporting Information Development of a human vasopressin V 1a -receptor antagonist from an evolutionary-related insect neuropeptide Maria Giulia Di Giglio, Markus Muttenthaler, Kasper Harpsøe, Zita Liutkeviciute,
Supporting Information Development of a human vasopressin V 1a -receptor antagonist from an evolutionary-related insect neuropeptide Maria Giulia Di Giglio, Markus Muttenthaler, Kasper Harpsøe, Zita Liutkeviciute,
Biochemical / Biophysical Kinetics Made Easy
 Biochemical / Biophysical Kinetics Made Easy Software DYNAFIT in drug discovery research Petr Kuzmič, Ph.D. BioKin, Ltd. 1. Theory: differential equation models - DYNAFIT software 2. Example: lanthascreen
Biochemical / Biophysical Kinetics Made Easy Software DYNAFIT in drug discovery research Petr Kuzmič, Ph.D. BioKin, Ltd. 1. Theory: differential equation models - DYNAFIT software 2. Example: lanthascreen
Identifying Interaction Hot Spots with SuperStar
 Identifying Interaction Hot Spots with SuperStar Version 1.0 November 2017 Table of Contents Identifying Interaction Hot Spots with SuperStar... 2 Case Study... 3 Introduction... 3 Generate SuperStar Maps
Identifying Interaction Hot Spots with SuperStar Version 1.0 November 2017 Table of Contents Identifying Interaction Hot Spots with SuperStar... 2 Case Study... 3 Introduction... 3 Generate SuperStar Maps
QuickZyme Total Protein Assay (to be used with acid hydrolyzates)
 QuickZyme Total Protein Assay (to be used with acid hydrolyzates) August 2014 This package insert must be read in its entirety before using this product FOR RESEARCH USE ONLY. NOT FOR USE IN DIAGNOSTIC
QuickZyme Total Protein Assay (to be used with acid hydrolyzates) August 2014 This package insert must be read in its entirety before using this product FOR RESEARCH USE ONLY. NOT FOR USE IN DIAGNOSTIC
