Unsupervised clustering of COMBO-17 galaxy photometry
|
|
|
- Wilfred Lang
- 5 years ago
- Views:
Transcription
1 STScI Astrostatistics R tutorials Eric Feigelson (Penn State) November 2011 SESSION 2 Multivariate clustering and classification ***************** ***************** Unsupervised clustering of COMBO-17 galaxy photometry We illustrate unsupervised clustering algorithms using a twodimensional color-magnitude diagram constructed from the COMBO-17 (`Classifying Objects by Medium-Band Observations in 17 Filters') photometric survey of normal galaxies (Wolf et al. 2003). The R script below starts with the which function to filter the dataset, keeping only low-redshift galaxies with z < 0.3 and remove a few points with bad data values. Most of the original 65 variables are ignored, and we keep only the galaxy absolute magnitude in the blue band, M_B, and the ultraviolet-to-blue color index, M_{280 - M_B. The resulting color-magnitude diagram ( left panel) shows the wellknown concentrations of luminous red galaxies around (M_B,M_{280- M_B) \simeq (-16,-0.2) and fainter blue galaxies around (-13,-0.9). The clusters are more clearly seen after smoothing the point process with a two-dimensional kernel density estimator using kde2d in R's MASS library. The blue galaxies are spirals and irregular galaxies that have experienced recent active star formation, while the red galaxies are mostly ellipticals that have only older stars formed early in the Universe's history. Note that many galaxies have properties distributed around the major concentrations. For example, a few extremely luminous and red galaxies are seen around (-20,1.5); these are nearby examples of the `luminous red galaxies' that are very important in cosmological studies (Eisenstein et al. 2005). While the kernel density estimator provides a valuable visualization of the clustering pattern, it does not assign individual galaxies to specific clusters. We illustrate unsupervised clustering of the
2 dataset using three methods in the following R script. For the nonparametric procedures where we assume a Euclidean distances between points in the 2-space, we first standardize the variables by removing the means and dividing by the standard deviations. # Unsupervised clustering of red and blue galaxies # Color-magnitude diagram for low-redshift COMBO-17 galaxies COMBO=read.table(' COMBO17_lowz.dat',header=T,fill=T) dim(combo) ; names(combo) par(mfrow=c(1,2)) plot(combo,pch=20,cex=0.5,xlim=c(-22,-7), ylim=c(-2,2.5),xlab='m_b (mag)',ylab='m_280 - M_B (mag)',main='combo-17 galaxies (z<0.3)') # Two-dimensional kernel-density estimator library(mass) COMBO_sm=kde2d(COMBO[,1],COMBO[,2], h=c(1.6,0.4),lims = c (-22,-7,-2,2.5), n=500) image(combo_sm,col=grey(13:0/15),xlab='m_b (mag)',ylab='m_280 - M_B (mag)',,xlim=c(-22,-7), ylim=c(-2,2.5),xaxp=c(-20,-10,2)) text(-18,-1.2,'blue cloud', col='blue') text(-18,-0.8,'green valley', col='darkgreen') text(-10,-0.2,'red sequence', col='red') text(-13,1.5,'bright cluster galaxies', col='darkred') par(mfrow=c(1,1)) R and CRAN have a variety of agglomerative hierarchical clustering algorithms. We start with the most commonly used procedure, function hclust in base-r. The procedure runs on the matrix of pairwise distances between points constructed using the function dist. As the structures in the smoothed distribution seem roughly spherical, we choose the `complete linkage' definition of group locations. The product of hclust is a dendrogram which can be displayed using plclust. # Standardize variables
3 Mag_std=(COMBO[,1]-mean(COMBO[,1]))/sd(COMBO[,1]) Color_std=(COMBO[,2]-mean(COMBO[,2]))/sd(COMBO[,2]) COMBO_std=cbind(Mag_std,Color_std) plot(combo_std) # Hierarchical clustering COMBO_dist=dist(COMBO_std) COMBO_hc=hclust(COMBO_dist,method='complete') # Cutting the tree at k=5 clusters plclust(combo_hc,label=f) COMBO_hc5a=rect.hclust(COMBO_hc,k=5,border='black') ; str (COMBO_hc5a) COMBO_hc5b=cutree(COMBO_hc,k=5) ; str(combo_hc5b) plot(combo,pch=(combo_hc5b+19),cex=1.0,xlab='m_b (mag)',ylab='m_280 - M_B (mag)',main='combo-17 hier clustering (k=5)',cex.lab=1.3,cex.axis=1.3) There is no formal procedure to select branches of the dendrogram as physically valid clusters. The cophenetic correlation coefficient, a measure of the similarity of the hierarchical structure and the data, is 0.52 using functions cophenetic and cor, but this can not readily be converted to a probability. We investigated the tree by trial-and-error, and found that `cutting the tree' at $k=5$ clusters provides a useful result. Two procedures are shown here: {rect.hclust that shows rectangles in the dendrogram (top panel), and cutree which gives an output with individual galaxy memberships of the five clusters. These are shown as different symbols in the color-magnitude diagram of the bottom panel; the open triangles show the red galaxies and the open circles show the blue galaxies. These clusters include many outlying galaxies, and examination of smaller clusters in the hierarchy does not cleanly discriminate the cluster extents seen in the smoothed distribution seen in the earlier figure (left panel). Our second clustering method attempts to alleviate this problem with the hierarchical clustering results by using the density as a starting point for the clustering algorithm. We use the DBSCAN (density-based cluster analysis) in CRAN package fpc (`fixed point clusters') which implements the procedure of Ester et al. (1996). DBSCAN is widely used, particularly for problems where compact clusters of interest are embedded in multiscale structure. The
4 {dbscan function requires user input of two parameters: the minimum number of points within a radius (or `reach') associated with the clusters of interest. By trial-and-error, we found that a minimum of 10 points within 0.3 standardized magnitude units provided a useful result as shown in the figure. Here only the fraction of galaxies lying within the regions satisfying this local density criterion are classified; red and blue galaxy groups are clearly discriminated, and intermediate galaxies are not classified. # Density-based clustering algorithm install.packages('fpc') ; library(fpc) COMBO_dbs = dbscan(combo_std,eps=0.1,minpts=5,method='raw') print.dbscan(combo_dbs) ; COMBO_dbs$cluster plot(combo[combo_dbs$cluster==0,], pch=20,cex=0.7,xlab='m_b (mag)',ylab='m_280 - M_B (mag)',main='combo-17 density wt clustering',cex.lab=1.3,cex.axis=1.3) points(combo[combo_dbs$cluster==2,],pch=2,cex=1.0) points(combo[combo_dbs$cluster==1 COMBO_dbs $cluster==3,],pch=1,cex=1.0) Our third clustering method is the well-respected parametric mclust (`model-based clustering') package in CRAN that fits a multivariate normal (MVN) mixture model by maximum-likelihood estimation using the EM Algorithm with Bayesian regularization (Fraley & Raftery 2002, 2007). We run an unsupervised procedure, but the calculation can be initialized with the output of MVN hierarchical clustering and the user can specify conjugate priors for the means and variances. In function mclustbic, the `VVV' model name specifies multivariate ellipsoidal Gaussians with arbitrary orientations. Model selection is performed by maximizing the Bayesian Information Criterion (BIC) for different number of clusters. The model-based clustering algorithm is shown in the figure. The likelihood for the COMBO-17 color-magnitude diagram is maximized for three clusters, two of which distinguish the red and blue galaxy sequences. Detailed results are provided by the summary.mclustbic function including the probabilities of cluster membership for each galaxy and the uncertainties to these probabilities. Points lying between two clusters can be investigated using a
5 visualization tool known as `shadow' and `silhouette' plots coupled to centroid-based partitioning cluster analysis (Leisch 2009). Each data point has a shadow value equal to two-times the distance to the closest cluster centroid divided by the sum of distances to closest and second-closest centroids. Points with shadow values near unity lie equidistant from the two clusters. Silhouette values measure the difference between the average dissimilarity of a point to all points in its own cluster to the smallest average dissimilarity to the points of a different cluster. Small values again indicate points with ill-defined cluster memberships. These plots can be constructed using CRAN's flexclust package. # Model-based clustering library(mclust) COMBO_mclus=mclustBIC(COMBO,modelNames='VVV') plot(combo_mclus,col='black') COMBO_sum_mclus=summary.mclustBIC(COMBO_mclus,COMBO,3) COMBO_sum_mclus$parameters ; COMBO_sum_mclus$classification COMBO_sum_mclus$z ; COMBO_sum_mclus$uncertainty plot(combo,pch=(19+combo_sum_mclus$classification),cex=1.0,xlab='m_b (mag)',ylab='m_280 - M_B (mag)',main='combo-17 MVN model clustering (k=3)',cex.lab=1.3,cex.axis=1.3) Altogether, none of the unsupervised clustering techniques showed the `blue cloud', `green valley', `red sequence' and `BCGs' as well as a simple kernel smoother. The results of unsupervised clustering, including the astronomers' favorite `friends-of-friends' algorithm, are often unreliable in the sense that reasonable alternative algorithms give very different scientific results Supervised classification of SDSS point sources The Sloan Digital Sky Survey (SDSS) has produced some of the most impressive photometric catalogs in modern astronomy. A selection of 17,000 SDSS point sources, along with training sets for three spectroscopically confirmed classes (main sequence plus red giant stars, quasars, and white dwarfs). These are 4-dimensional datasets
6 with variables representing the ratios of brightness in the five SDSS photometric bands (u-g, g-r, r-i, and i-z). The resulting color-color scatterplots show distributions that cannot be wellmodeled by multinormal distributions, and distributions that are distinct in some variables but overlapping in others. The analysis here starts with the SDSS_train and SDSS_test obtained using the following R script. # SDSS point sources dataset, N=17,000 (mag<21, point sources, hiqual) SDSS=read.csv(' SDSS_test.csv',header=T) dim(sdss) ; summary(sdss) SDSS_test=data.frame(cbind((SDSS[,1]-SDSS[,2]),(SDSS[,2]-SDSS[,3]), (SDSS[,3]-SDSS[,4]),(SDSS[,4]-SDSS[,5]))) names(sdss_test)=c('u_g','g_r','r_i','i_z') str(sdss_test) par(mfrow=c(1,3)) plot(sdss_test[,1],sdss_test[,2],xlim=c(-0.7,3),ylim=c (-0.7,1.8),pch=20, cex=0.6,cex.lab=1.5,cex.axis=1.5,main='',xlab='ug (mag)',ylab='g-r (mag)') plot(sdss_test[,2],sdss_test[,3],xlim=c(-0.7,1.8),ylim=c (-0.7,1.8),pch=20, cex=0.6,cex.lab=1.5,cex.axis=1.5,main='',xlab='gr (mag)',ylab='r-i (mag)') plot(sdss_test[,3],sdss_test[,4],xlim=c(-0.7,1.8),ylim=c (-1.1,1.3),pch=20, cex=0.6,cex.lab=1.5,cex.axis=1.5,main='',xlab='ri (mag)',ylab='i-z (mag)') par(mfrow=c(1,1)) # Quasar training set, N=2000 (Class 1) temp1 = read.table(' SDSS_QSO.dat', header=t) dim(temp1) ; summary(temp1) qso = cbind(temp1[,c(5,7,9,11,13,6,8,10,12,14,2,3)]) # set same variables in both datasets
7 bad_phot_qso = which(qso[,1:6] > 21.0 qso[,9]==0) qso1 = qso[-bad_phot_qso,] qso2 = qso1[1:2000,] qso3=cbind((qso2[,1]-qso2[,2]),(qso2[,2]-qso2[,3]),(qso2[,3]-qso2[, 4]),(qso2[,4]-qso2[,5])) qso_train=data.frame(cbind(qso3,rep(1,length(qso2[,1])))) names(qso_train)=c('u_g','g_r','r_i','i_z','class') dim(qso_train) ; summary(qso_train) # Star training set, N=5000 (Class 2) temp2 = read.csv(' SDSS_stars.csv', header=t) dim(temp2) ; summary(temp2) star=cbind((temp2[,1]-temp2[,2]),(temp2[,2]-temp2[,3]),(temp2[,3]- temp2[,4]),(temp2[,4]-temp2[,5])) star_train=data.frame(cbind(star,rep(2,length(star[,1])))) names(star_train)=c('u_g','g_r','r_i','i_z','class') dim(star_train) ; summary(star_train) # White dwarf training set, N=2000 (Class 3) temp3 = read.csv(' SDSS_wd.csv', header=t) dim(temp3) ; summary(temp3) temp3=na.omit(temp3) wd =cbind((temp3[1:2000,2]-temp3[1:2000,3]),(temp3[1:2000,3]-temp3 [1:2000,4]),(temp3[1:2000,4]-temp3[1:2000,5]),(temp3[1:2000,5]-temp3 [1:2000,6])) wd_train=data.frame(cbind(wd,rep(3,length(wd[,1])))) names(wd_train)=c('u_g','g_r','r_i','i_z','class') dim(wd_train) ; summary(wd_train) # Combined training set (9000 objects) SDSS_train=data.frame(rbind(qso_train,star_train,wd_train)) names(sdss_train)=c('u_g','g_r','r_i','i_z','class') str(sdss_train) par(mfrow=c(1,3)) plot(sdss_train[,1],sdss_train[,2],xlim=c(-0.7,3),ylim=c (-0.7,1.8),pch=20, col=sdss_train[,
8 5],cex=0.6,cex.lab=1.5,cex.axis=1.5,main='',xlab='u-g (mag)',ylab='g-r (mag)') legend(-0.5,1.7,c('qso','ms + RG','WD'), pch=20,col=c ('black','red','green'),cex=1.8) plot(sdss_train[,2],sdss_train[,3],xlim=c(-0.7,1.8),ylim=c (-0.7,1.8),pch=20, col=sdss_train[, 5],cex=0.6,cex.lab=1.5,cex.axis=1.5,main='',xlab='g-r (mag)',ylab='r-i (mag)') plot(sdss_train[,3],sdss_train[,4],xlim=c(-0.7,1.8),ylim=c (-1.1,1.3),pch=20, col=sdss_train[, 5],cex=0.6,cex.lab=1.5,cex.axis=1.5,main='',xlab='r-i (mag)',ylab='i-z (mag)') par(mfrow=c(1,1)) Unsupervised clustering fails to recover the known distributions in the SDSS photometric distribution. We show, for example, the result of a k-means partitioning in the figure. The k-means partitioning with divides the main sequence into segments, even though there are no gaps in the distribution. Even a supervised k-means partitioning with three initial cluster roughly centered on the three training classes does not lead to a correct result. Similar problems arise when unsupervised hierarchical clustering (R function hclust) or model-based clustering (function Mclust in package mclust) are applied to the SDSS distributions. # Unsupervised k-means partitioning SDSS_kmean=kmeans(SDSS_test,6) plot(sdss_test[,1],sdss_test[,2],xlim=c(-0.5,3),ylim=c (-0.5,2),pch=20,col=SDSS_kmean$cluster, cex=0.6, cex.lab=1.3,cex.axis=1.3,cex.main=1.3,main='sdss unsupervised k- means',xlab='u-g (mag)',ylab='g-r (mag)') Discrimination analysis and k-nn classification The classification of SDSS objects is much improved when the training set used. We show here the result of linear discriminant analysis (LDA) using function lda in base-r's MASS package. The LDA
9 classification from the training set is applied to the test set using R's predict function. The figure (top left panel) shows the result for the test sample; a similar plot can be inspected for the training sample. Discriminant analysis gives a reasonable classification of stars, quasars and white dwarfs, with no difficulty following the elongated and curved distributions in 4-space. However, some classification errors are evident: a few main sequence stars are mislabeled as quasars (black dots), and the white dwarf class (green dots) is truncated by the quasar distribution. The closely related quadratic discriminate analysis using function qda has additional problems, classifying some main sequence and red giant stars as white dwarfs. CRAN package class implements a k-nearest neighbors classifier where a grid of classifications is constructed from the training set. The application to the SDSS test set is shown in the figure (top right panel) and shows good performance. Here we use k=4 neighbors, but the result is not sensitive to a range of k values. We consider two ways to examine the reliability of the these classifiers by applying it to the training set. First, a crossvalidation experiment can be made (e.g., using function knn.cv) where leave-one-out resamples of the test dataset give posterior probabilities for the classification of each object. Second, the class obtained by the classified can be plotted against the true class known for the training set objects. We show this in the bottom panels of the figure; the LDA clearly makes more misclassifications than the k-nn algorithm. For k-nn, misclassification of stars (Class 2) is rare (0.1%) but confusion between quasars (Class 1) and white dwarfs (Class 3) occurs in about 2% of cases. This is understandable given the overlap in their distributions in color-color plots. Note the use of R's jitter function to facilitate visualization of categorical data for scatterplots. # Linear discriminant analysis library(mass) SDSS_lda=lda(SDSS_train[,1:4],as.factor(SDSS_train[,5])) SDSS_train_lda=predict(SDSS_lda,SDSS_train[,1:4]) SDSS_test_lda=predict(SDSS_lda,SDSS_test[,1:4])
10 par(mfrow=c(1,2)) plot(sdss_test[,1],sdss_test[,2],xlim=c(-0.7,3),ylim=c (-0.7,1.8),pch=20, col=sdss_test_lda$class, cex=0.6,cex.lab=1.5,cex.axis=1.5,main='',xlab='u-g (mag)',ylab='g-r (mag)') # k-nn classification install.packages('class') ; library(class) SDSS_knn4 = knn(sdss_train[,1:4],sdss_test,sdss_train[, 5],k=4,prob=T) plot(sdss_test[,1],sdss_test[,2],xlim=c(-0.7,3),ylim=c (-0.7,1.8),pch=20, col=sdss_knn4, cex=0.6,cex.lab=1.5,cex.axis=1.5,main='',xlab='u-g (mag)',ylab='g-r (mag)') par(mfrow=c(1,1)) # Validation of k-nn classification SDSS_train_lda_cv=lda(SDSS_train[,1:4],as.factor(SDSS_train[, 5]),CV=T) SDSS_train_lda=lda(SDSS_train[,1:4],as.factor(SDSS_train[,5])) SDSS_train_knn4 = knn(sdss_train[,1:4],sdss_train[,1:4],sdss_train[, 5],k=4) par(mfrow=c(1,2)) plot(jitter(as.numeric(sdss_train_lda$class),factor=0.5),jitter (as.numeric(sdss_train[,5]),factor=0.5),pch=20,cex=1.0,xlab='lda class',ylab='true class',cex.lab=1.3,cex.axis=1.3, xaxp=c (1,3,2),yaxp=c(1,3,2)) plot(jitter(as.numeric(sdss_train_knn4),factor=0.5),jitter (as.numeric(sdss_train[,5]),factor=0.5),pch=20,cex=1.0,xlab='k-nn class',ylab='true class',cex.lab=1.3,cex.axis=1.3, xaxp=c (1,3,2),yaxp=c(1,3,2)) par(mfrow=c(1,1)) # Single layer neutral network install.packages('nnet') ; library(nnet) options(size=100, maxit=1000) SDSS_nnet <- multinom(as.factor(sdss_train[,5]) ~ SDSS_train[,1] +
11 SDSS_train[,2] + SDSS_train[,3] + SDSS_train[,4], data=sdss_train) SDSS_train_nnet <- predict(sdss_nnet,sdss_train[,1:4]) plot(jitter(as.numeric(sdss_train_nnet), factor=0.5), jitter (as.numeric(sdss_train[,5]), factor=0.5), pch=20, cex=0.5, xlab='nnet class', ylab='true class', xaxp=c(1,3,2),yaxp=c(1,3,2)) Machine learning classifiers Machine learning classifiers perform well for this problem. In the following R script, we apply CART using rpart (acronym for `recursive partitioning and regression trees') in base-r's rpart library, and the Support Vector Machine {svm implemented in CRAN's e1071 package. The procedure for running these and similar classifiers is straightforward. The `model' is produced by rpart or svm with a formula like `Known_classes ~.' to the training set. Examining the model using summary and str shows that the classifier output can be quite complicated; e.g., CART will give details on the decision tree nodes while SVM will give details on the support vectors. But the model predictions can be automatically applied to the training and test datasets using R's predict function without understanding these details. We plot the predicted classes against the known classes for the training set in the figure. CART does not perform as well as the k- nn shown above, but the SVM classifier does a better job. The figure shows the CART tree with the splits labeled, and the next figure shows how much of the variance is reduced by each split of the data. Considering the SVM classification as the best available, we show the final classifications of the test SDSS sample in the figure, and write them to an ASCII output file SDSS_test_svm.out. Note that R's write function produces tables that are difficult to read; we use the format function and other options in write to improve the appearance of the ASCII output. # Classification And Regression Tree model, prediction and validation library('rpart')
12 SDSS_rpart_mod = rpart(sdss_train[,5] ~.,data=sdss_train[,1:4]) SDSS_rpart_test_pred = predict(sdss_rpart_mod, SDSS_test) SDSS_rpart_train_pred = predict(sdss_rpart_mod, SDSS_train) summary(sdss_rpart_mod) ; str(sdss_rpart_mod) par(mfrow=c(1,2)) plot(jitter(sdss_rpart_train_pred,factor=5),jitter(sdss_train[, 5]),pch=20,cex=0.3,cex.axis=1.5, cex.lab=1.5,xlab='cart class',ylab='true class',yaxp=c(1,3,2)) plot(sdss_test[,1],sdss_test[,2],xlim=c(-0.7,3),ylim=c (-0.7,1.8),pch=20, col=round(sdss_rpart_test_pred), cex=0.6,cex.lab=1.5,cex.axis=1.5,main='',xlab='u-g (mag)',ylab='g-r (mag)') plot(sdss_rpart_mod) ; text(sdss_rpart_mod) plotcp(sdss_rpart_mod,lwd=2,cex.axis=1.3,cex.lab=1.3) # Support Vector Machine model, prediction and validation install.packages('e1071') ; library(e1071) SDSS_svm_mod = svm(sdss_train[,5] ~.,data=sdss_train[,1:4],cost = 100, gamma = 1) summary(sdss_svm_mod) ; str(sdss_svm_mod) SDSS_svm_test_pred = predict(sdss_svm_mod, SDSS_test) SDSS_svm_train_pred = predict(sdss_svm_mod, SDSS_train) par(mfrow=c(1,2)) plot(sdss_svm_train_pred,jitter(sdss_train[, 5]),pch=20,cex=0.3,cex.axis=1.5, cex.lab=1.5,xlab='svm class',ylab='true class',yaxp=c(1,3,2)) plot(sdss_test[,1],sdss_test[,2],xlim=c(-0.7,3),ylim=c (-0.7,1.8),pch=20, col=round(sdss_svm_test_pred), cex=0.6,cex.lab=1.5,cex.axis=1.5,main='',xlab='u-g (mag)',ylab='g-r (mag)') # 23 new R functions are used in this tutorial: # text, cbind, dist, hclust, plclust, rect.hclust, dbscan, print.dbscan, # points, mclustbic, read.csv, data.frame, which, kmeans, lda, predict, # knn, as.numeric, jitter, multinom, as.factor, rpart, svm
Lectures in AstroStatistics: Topics in Machine Learning for Astronomers
 Lectures in AstroStatistics: Topics in Machine Learning for Astronomers Jessi Cisewski Yale University American Astronomical Society Meeting Wednesday, January 6, 2016 1 Statistical Learning - learning
Lectures in AstroStatistics: Topics in Machine Learning for Astronomers Jessi Cisewski Yale University American Astronomical Society Meeting Wednesday, January 6, 2016 1 Statistical Learning - learning
Clustering. CSL465/603 - Fall 2016 Narayanan C Krishnan
 Clustering CSL465/603 - Fall 2016 Narayanan C Krishnan ckn@iitrpr.ac.in Supervised vs Unsupervised Learning Supervised learning Given x ", y " "%& ', learn a function f: X Y Categorical output classification
Clustering CSL465/603 - Fall 2016 Narayanan C Krishnan ckn@iitrpr.ac.in Supervised vs Unsupervised Learning Supervised learning Given x ", y " "%& ', learn a function f: X Y Categorical output classification
Data Preprocessing. Cluster Similarity
 1 Cluster Similarity Similarity is most often measured with the help of a distance function. The smaller the distance, the more similar the data objects (points). A function d: M M R is a distance on M
1 Cluster Similarity Similarity is most often measured with the help of a distance function. The smaller the distance, the more similar the data objects (points). A function d: M M R is a distance on M
Data Exploration and Unsupervised Learning with Clustering
 Data Exploration and Unsupervised Learning with Clustering Paul F Rodriguez,PhD San Diego Supercomputer Center Predictive Analytic Center of Excellence Clustering Idea Given a set of data can we find a
Data Exploration and Unsupervised Learning with Clustering Paul F Rodriguez,PhD San Diego Supercomputer Center Predictive Analytic Center of Excellence Clustering Idea Given a set of data can we find a
More on Unsupervised Learning
 More on Unsupervised Learning Two types of problems are to find association rules for occurrences in common in observations (market basket analysis), and finding the groups of values of observational data
More on Unsupervised Learning Two types of problems are to find association rules for occurrences in common in observations (market basket analysis), and finding the groups of values of observational data
Machine learning comes from Bayesian decision theory in statistics. There we want to minimize the expected value of the loss function.
 Bayesian learning: Machine learning comes from Bayesian decision theory in statistics. There we want to minimize the expected value of the loss function. Let y be the true label and y be the predicted
Bayesian learning: Machine learning comes from Bayesian decision theory in statistics. There we want to minimize the expected value of the loss function. Let y be the true label and y be the predicted
Mul$variate clustering & classifica$on with R
 Mul$variate clustering & classifica$on with R Eric Feigelson Penn State Cosmology on the Beach, 2014 Astronomical context Astronomers have constructed classifica@ons of celes@al objects for centuries:
Mul$variate clustering & classifica$on with R Eric Feigelson Penn State Cosmology on the Beach, 2014 Astronomical context Astronomers have constructed classifica@ons of celes@al objects for centuries:
Gaussian Models
 Gaussian Models ddebarr@uw.edu 2016-04-28 Agenda Introduction Gaussian Discriminant Analysis Inference Linear Gaussian Systems The Wishart Distribution Inferring Parameters Introduction Gaussian Density
Gaussian Models ddebarr@uw.edu 2016-04-28 Agenda Introduction Gaussian Discriminant Analysis Inference Linear Gaussian Systems The Wishart Distribution Inferring Parameters Introduction Gaussian Density
Part I. Linear regression & LASSO. Linear Regression. Linear Regression. Week 10 Based in part on slides from textbook, slides of Susan Holmes
 Week 10 Based in part on slides from textbook, slides of Susan Holmes Part I Linear regression & December 5, 2012 1 / 1 2 / 1 We ve talked mostly about classification, where the outcome categorical. If
Week 10 Based in part on slides from textbook, slides of Susan Holmes Part I Linear regression & December 5, 2012 1 / 1 2 / 1 We ve talked mostly about classification, where the outcome categorical. If
Semiparametric Discriminant Analysis of Mixture Populations Using Mahalanobis Distance. Probal Chaudhuri and Subhajit Dutta
 Semiparametric Discriminant Analysis of Mixture Populations Using Mahalanobis Distance Probal Chaudhuri and Subhajit Dutta Indian Statistical Institute, Kolkata. Workshop on Classification and Regression
Semiparametric Discriminant Analysis of Mixture Populations Using Mahalanobis Distance Probal Chaudhuri and Subhajit Dutta Indian Statistical Institute, Kolkata. Workshop on Classification and Regression
The Challenges of Heteroscedastic Measurement Error in Astronomical Data
 The Challenges of Heteroscedastic Measurement Error in Astronomical Data Eric Feigelson Penn State Measurement Error Workshop TAMU April 2016 Outline 1. Measurement errors are ubiquitous in astronomy,
The Challenges of Heteroscedastic Measurement Error in Astronomical Data Eric Feigelson Penn State Measurement Error Workshop TAMU April 2016 Outline 1. Measurement errors are ubiquitous in astronomy,
The Bayes classifier
 The Bayes classifier Consider where is a random vector in is a random variable (depending on ) Let be a classifier with probability of error/risk given by The Bayes classifier (denoted ) is the optimal
The Bayes classifier Consider where is a random vector in is a random variable (depending on ) Let be a classifier with probability of error/risk given by The Bayes classifier (denoted ) is the optimal
9. High-level processing (astronomical data analysis)
 Master ISTI / PARI / IV Introduction to Astronomical Image Processing 9. High-level processing (astronomical data analysis) André Jalobeanu LSIIT / MIV / PASEO group Jan. 2006 lsiit-miv.u-strasbg.fr/paseo
Master ISTI / PARI / IV Introduction to Astronomical Image Processing 9. High-level processing (astronomical data analysis) André Jalobeanu LSIIT / MIV / PASEO group Jan. 2006 lsiit-miv.u-strasbg.fr/paseo
CLUe Training An Introduction to Machine Learning in R with an example from handwritten digit recognition
 CLUe Training An Introduction to Machine Learning in R with an example from handwritten digit recognition Ad Feelders Universiteit Utrecht Department of Information and Computing Sciences Algorithmic Data
CLUe Training An Introduction to Machine Learning in R with an example from handwritten digit recognition Ad Feelders Universiteit Utrecht Department of Information and Computing Sciences Algorithmic Data
Introduction to Machine Learning Midterm Exam
 10-701 Introduction to Machine Learning Midterm Exam Instructors: Eric Xing, Ziv Bar-Joseph 17 November, 2015 There are 11 questions, for a total of 100 points. This exam is open book, open notes, but
10-701 Introduction to Machine Learning Midterm Exam Instructors: Eric Xing, Ziv Bar-Joseph 17 November, 2015 There are 11 questions, for a total of 100 points. This exam is open book, open notes, but
Advanced Statistical Methods: Beyond Linear Regression
 Advanced Statistical Methods: Beyond Linear Regression John R. Stevens Utah State University Notes 3. Statistical Methods II Mathematics Educators Worshop 28 March 2009 1 http://www.stat.usu.edu/~jrstevens/pcmi
Advanced Statistical Methods: Beyond Linear Regression John R. Stevens Utah State University Notes 3. Statistical Methods II Mathematics Educators Worshop 28 March 2009 1 http://www.stat.usu.edu/~jrstevens/pcmi
Unsupervised machine learning
 Chapter 9 Unsupervised machine learning Unsupervised machine learning (a.k.a. cluster analysis) is a set of methods to assign objects into clusters under a predefined distance measure when class labels
Chapter 9 Unsupervised machine learning Unsupervised machine learning (a.k.a. cluster analysis) is a set of methods to assign objects into clusters under a predefined distance measure when class labels
Pattern Recognition and Machine Learning
 Christopher M. Bishop Pattern Recognition and Machine Learning ÖSpri inger Contents Preface Mathematical notation Contents vii xi xiii 1 Introduction 1 1.1 Example: Polynomial Curve Fitting 4 1.2 Probability
Christopher M. Bishop Pattern Recognition and Machine Learning ÖSpri inger Contents Preface Mathematical notation Contents vii xi xiii 1 Introduction 1 1.1 Example: Polynomial Curve Fitting 4 1.2 Probability
Table of Contents. Multivariate methods. Introduction II. Introduction I
 Table of Contents Introduction Antti Penttilä Department of Physics University of Helsinki Exactum summer school, 04 Construction of multinormal distribution Test of multinormality with 3 Interpretation
Table of Contents Introduction Antti Penttilä Department of Physics University of Helsinki Exactum summer school, 04 Construction of multinormal distribution Test of multinormality with 3 Interpretation
Mul$variate clustering. Astro 585 Spring 2013
 Mul$variate clustering Astro 585 Spring 2013 Astronomical context Astronomers have constructed classifica
Mul$variate clustering Astro 585 Spring 2013 Astronomical context Astronomers have constructed classifica
Big Data Inference. Combining Hierarchical Bayes and Machine Learning to Improve Photometric Redshifts. Josh Speagle 1,
 ICHASC, Apr. 25 2017 Big Data Inference Combining Hierarchical Bayes and Machine Learning to Improve Photometric Redshifts Josh Speagle 1, jspeagle@cfa.harvard.edu In collaboration with: Boris Leistedt
ICHASC, Apr. 25 2017 Big Data Inference Combining Hierarchical Bayes and Machine Learning to Improve Photometric Redshifts Josh Speagle 1, jspeagle@cfa.harvard.edu In collaboration with: Boris Leistedt
Clustering using Mixture Models
 Clustering using Mixture Models The full posterior of the Gaussian Mixture Model is p(x, Z, µ,, ) =p(x Z, µ, )p(z )p( )p(µ, ) data likelihood (Gaussian) correspondence prob. (Multinomial) mixture prior
Clustering using Mixture Models The full posterior of the Gaussian Mixture Model is p(x, Z, µ,, ) =p(x Z, µ, )p(z )p( )p(µ, ) data likelihood (Gaussian) correspondence prob. (Multinomial) mixture prior
L11: Pattern recognition principles
 L11: Pattern recognition principles Bayesian decision theory Statistical classifiers Dimensionality reduction Clustering This lecture is partly based on [Huang, Acero and Hon, 2001, ch. 4] Introduction
L11: Pattern recognition principles Bayesian decision theory Statistical classifiers Dimensionality reduction Clustering This lecture is partly based on [Huang, Acero and Hon, 2001, ch. 4] Introduction
Regression Clustering
 Regression Clustering In regression clustering, we assume a model of the form y = f g (x, θ g ) + ɛ g for observations y and x in the g th group. Usually, of course, we assume linear models of the form
Regression Clustering In regression clustering, we assume a model of the form y = f g (x, θ g ) + ɛ g for observations y and x in the g th group. Usually, of course, we assume linear models of the form
Final Overview. Introduction to ML. Marek Petrik 4/25/2017
 Final Overview Introduction to ML Marek Petrik 4/25/2017 This Course: Introduction to Machine Learning Build a foundation for practice and research in ML Basic machine learning concepts: max likelihood,
Final Overview Introduction to ML Marek Petrik 4/25/2017 This Course: Introduction to Machine Learning Build a foundation for practice and research in ML Basic machine learning concepts: max likelihood,
Classifying Galaxy Morphology using Machine Learning
 Julian Kates-Harbeck, Introduction: Classifying Galaxy Morphology using Machine Learning The goal of this project is to classify galaxy morphologies. Generally, galaxy morphologies fall into one of two
Julian Kates-Harbeck, Introduction: Classifying Galaxy Morphology using Machine Learning The goal of this project is to classify galaxy morphologies. Generally, galaxy morphologies fall into one of two
Learning algorithms at the service of WISE survey
 Katarzyna Ma lek 1,2,3, T. Krakowski 1, M. Bilicki 4,3, A. Pollo 1,5,3, A. Solarz 2,3, M. Krupa 5,3, A. Kurcz 5,3, W. Hellwing 6,3, J. Peacock 7, T. Jarrett 4 1 National Centre for Nuclear Research, ul.
Katarzyna Ma lek 1,2,3, T. Krakowski 1, M. Bilicki 4,3, A. Pollo 1,5,3, A. Solarz 2,3, M. Krupa 5,3, A. Kurcz 5,3, W. Hellwing 6,3, J. Peacock 7, T. Jarrett 4 1 National Centre for Nuclear Research, ul.
Contents Lecture 4. Lecture 4 Linear Discriminant Analysis. Summary of Lecture 3 (II/II) Summary of Lecture 3 (I/II)
 Contents Lecture Lecture Linear Discriminant Analysis Fredrik Lindsten Division of Systems and Control Department of Information Technology Uppsala University Email: fredriklindsten@ituuse Summary of lecture
Contents Lecture Lecture Linear Discriminant Analysis Fredrik Lindsten Division of Systems and Control Department of Information Technology Uppsala University Email: fredriklindsten@ituuse Summary of lecture
Brief Introduction of Machine Learning Techniques for Content Analysis
 1 Brief Introduction of Machine Learning Techniques for Content Analysis Wei-Ta Chu 2008/11/20 Outline 2 Overview Gaussian Mixture Model (GMM) Hidden Markov Model (HMM) Support Vector Machine (SVM) Overview
1 Brief Introduction of Machine Learning Techniques for Content Analysis Wei-Ta Chu 2008/11/20 Outline 2 Overview Gaussian Mixture Model (GMM) Hidden Markov Model (HMM) Support Vector Machine (SVM) Overview
Introduction. Chapter 1
 Chapter 1 Introduction In this book we will be concerned with supervised learning, which is the problem of learning input-output mappings from empirical data (the training dataset). Depending on the characteristics
Chapter 1 Introduction In this book we will be concerned with supervised learning, which is the problem of learning input-output mappings from empirical data (the training dataset). Depending on the characteristics
Machine Learning Linear Classification. Prof. Matteo Matteucci
 Machine Learning Linear Classification Prof. Matteo Matteucci Recall from the first lecture 2 X R p Regression Y R Continuous Output X R p Y {Ω 0, Ω 1,, Ω K } Classification Discrete Output X R p Y (X)
Machine Learning Linear Classification Prof. Matteo Matteucci Recall from the first lecture 2 X R p Regression Y R Continuous Output X R p Y {Ω 0, Ω 1,, Ω K } Classification Discrete Output X R p Y (X)
Final Exam, Machine Learning, Spring 2009
 Name: Andrew ID: Final Exam, 10701 Machine Learning, Spring 2009 - The exam is open-book, open-notes, no electronics other than calculators. - The maximum possible score on this exam is 100. You have 3
Name: Andrew ID: Final Exam, 10701 Machine Learning, Spring 2009 - The exam is open-book, open-notes, no electronics other than calculators. - The maximum possible score on this exam is 100. You have 3
Learning from Examples
 Learning from Examples Data fitting Decision trees Cross validation Computational learning theory Linear classifiers Neural networks Nonparametric methods: nearest neighbor Support vector machines Ensemble
Learning from Examples Data fitting Decision trees Cross validation Computational learning theory Linear classifiers Neural networks Nonparametric methods: nearest neighbor Support vector machines Ensemble
Machine Learning. Nonparametric Methods. Space of ML Problems. Todo. Histograms. Instance-Based Learning (aka non-parametric methods)
 Machine Learning InstanceBased Learning (aka nonparametric methods) Supervised Learning Unsupervised Learning Reinforcement Learning Parametric Non parametric CSE 446 Machine Learning Daniel Weld March
Machine Learning InstanceBased Learning (aka nonparametric methods) Supervised Learning Unsupervised Learning Reinforcement Learning Parametric Non parametric CSE 446 Machine Learning Daniel Weld March
Logic and machine learning review. CS 540 Yingyu Liang
 Logic and machine learning review CS 540 Yingyu Liang Propositional logic Logic If the rules of the world are presented formally, then a decision maker can use logical reasoning to make rational decisions.
Logic and machine learning review CS 540 Yingyu Liang Propositional logic Logic If the rules of the world are presented formally, then a decision maker can use logical reasoning to make rational decisions.
Holdout and Cross-Validation Methods Overfitting Avoidance
 Holdout and Cross-Validation Methods Overfitting Avoidance Decision Trees Reduce error pruning Cost-complexity pruning Neural Networks Early stopping Adjusting Regularizers via Cross-Validation Nearest
Holdout and Cross-Validation Methods Overfitting Avoidance Decision Trees Reduce error pruning Cost-complexity pruning Neural Networks Early stopping Adjusting Regularizers via Cross-Validation Nearest
Clustering VS Classification
 MCQ Clustering VS Classification 1. What is the relation between the distance between clusters and the corresponding class discriminability? a. proportional b. inversely-proportional c. no-relation Ans:
MCQ Clustering VS Classification 1. What is the relation between the distance between clusters and the corresponding class discriminability? a. proportional b. inversely-proportional c. no-relation Ans:
Lecture 4 Discriminant Analysis, k-nearest Neighbors
 Lecture 4 Discriminant Analysis, k-nearest Neighbors Fredrik Lindsten Division of Systems and Control Department of Information Technology Uppsala University. Email: fredrik.lindsten@it.uu.se fredrik.lindsten@it.uu.se
Lecture 4 Discriminant Analysis, k-nearest Neighbors Fredrik Lindsten Division of Systems and Control Department of Information Technology Uppsala University. Email: fredrik.lindsten@it.uu.se fredrik.lindsten@it.uu.se
View of the Galaxy from within. Lecture 12: Galaxies. Comparison to an external disk galaxy. Where do we lie in our Galaxy?
 Lecture 12: Galaxies View of the Galaxy from within The Milky Way galaxy Rotation curves and dark matter External galaxies and the Hubble classification scheme Plotting the sky brightness in galactic coordinates,
Lecture 12: Galaxies View of the Galaxy from within The Milky Way galaxy Rotation curves and dark matter External galaxies and the Hubble classification scheme Plotting the sky brightness in galactic coordinates,
Multivariate Statistics
 Multivariate Statistics Chapter 6: Cluster Analysis Pedro Galeano Departamento de Estadística Universidad Carlos III de Madrid pedro.galeano@uc3m.es Course 2017/2018 Master in Mathematical Engineering
Multivariate Statistics Chapter 6: Cluster Analysis Pedro Galeano Departamento de Estadística Universidad Carlos III de Madrid pedro.galeano@uc3m.es Course 2017/2018 Master in Mathematical Engineering
Curve Fitting Re-visited, Bishop1.2.5
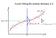 Curve Fitting Re-visited, Bishop1.2.5 Maximum Likelihood Bishop 1.2.5 Model Likelihood differentiation p(t x, w, β) = Maximum Likelihood N N ( t n y(x n, w), β 1). (1.61) n=1 As we did in the case of the
Curve Fitting Re-visited, Bishop1.2.5 Maximum Likelihood Bishop 1.2.5 Model Likelihood differentiation p(t x, w, β) = Maximum Likelihood N N ( t n y(x n, w), β 1). (1.61) n=1 As we did in the case of the
Modern Information Retrieval
 Modern Information Retrieval Chapter 8 Text Classification Introduction A Characterization of Text Classification Unsupervised Algorithms Supervised Algorithms Feature Selection or Dimensionality Reduction
Modern Information Retrieval Chapter 8 Text Classification Introduction A Characterization of Text Classification Unsupervised Algorithms Supervised Algorithms Feature Selection or Dimensionality Reduction
Computer Vision Group Prof. Daniel Cremers. 2. Regression (cont.)
 Prof. Daniel Cremers 2. Regression (cont.) Regression with MLE (Rep.) Assume that y is affected by Gaussian noise : t = f(x, w)+ where Thus, we have p(t x, w, )=N (t; f(x, w), 2 ) 2 Maximum A-Posteriori
Prof. Daniel Cremers 2. Regression (cont.) Regression with MLE (Rep.) Assume that y is affected by Gaussian noise : t = f(x, w)+ where Thus, we have p(t x, w, )=N (t; f(x, w), 2 ) 2 Maximum A-Posteriori
Classification: The rest of the story
 U NIVERSITY OF ILLINOIS AT URBANA-CHAMPAIGN CS598 Machine Learning for Signal Processing Classification: The rest of the story 3 October 2017 Today s lecture Important things we haven t covered yet Fisher
U NIVERSITY OF ILLINOIS AT URBANA-CHAMPAIGN CS598 Machine Learning for Signal Processing Classification: The rest of the story 3 October 2017 Today s lecture Important things we haven t covered yet Fisher
PATTERN RECOGNITION AND MACHINE LEARNING
 PATTERN RECOGNITION AND MACHINE LEARNING Chapter 1. Introduction Shuai Huang April 21, 2014 Outline 1 What is Machine Learning? 2 Curve Fitting 3 Probability Theory 4 Model Selection 5 The curse of dimensionality
PATTERN RECOGNITION AND MACHINE LEARNING Chapter 1. Introduction Shuai Huang April 21, 2014 Outline 1 What is Machine Learning? 2 Curve Fitting 3 Probability Theory 4 Model Selection 5 The curse of dimensionality
Overview of Statistical Tools. Statistical Inference. Bayesian Framework. Modeling. Very simple case. Things are usually more complicated
 Fall 3 Computer Vision Overview of Statistical Tools Statistical Inference Haibin Ling Observation inference Decision Prior knowledge http://www.dabi.temple.edu/~hbling/teaching/3f_5543/index.html Bayesian
Fall 3 Computer Vision Overview of Statistical Tools Statistical Inference Haibin Ling Observation inference Decision Prior knowledge http://www.dabi.temple.edu/~hbling/teaching/3f_5543/index.html Bayesian
ISyE 6416: Computational Statistics Spring Lecture 5: Discriminant analysis and classification
 ISyE 6416: Computational Statistics Spring 2017 Lecture 5: Discriminant analysis and classification Prof. Yao Xie H. Milton Stewart School of Industrial and Systems Engineering Georgia Institute of Technology
ISyE 6416: Computational Statistics Spring 2017 Lecture 5: Discriminant analysis and classification Prof. Yao Xie H. Milton Stewart School of Industrial and Systems Engineering Georgia Institute of Technology
Instance-based Learning CE-717: Machine Learning Sharif University of Technology. M. Soleymani Fall 2016
 Instance-based Learning CE-717: Machine Learning Sharif University of Technology M. Soleymani Fall 2016 Outline Non-parametric approach Unsupervised: Non-parametric density estimation Parzen Windows Kn-Nearest
Instance-based Learning CE-717: Machine Learning Sharif University of Technology M. Soleymani Fall 2016 Outline Non-parametric approach Unsupervised: Non-parametric density estimation Parzen Windows Kn-Nearest
There are three main ways to derive q 0 :
 Measuring q 0 Measuring the deceleration parameter, q 0, is much more difficult than measuring H 0. In order to measure the Hubble Constant, one needs to derive distances to objects at 100 Mpc; this corresponds
Measuring q 0 Measuring the deceleration parameter, q 0, is much more difficult than measuring H 0. In order to measure the Hubble Constant, one needs to derive distances to objects at 100 Mpc; this corresponds
Switch Mechanism Diagnosis using a Pattern Recognition Approach
 The 4th IET International Conference on Railway Condition Monitoring RCM 2008 Switch Mechanism Diagnosis using a Pattern Recognition Approach F. Chamroukhi, A. Samé, P. Aknin The French National Institute
The 4th IET International Conference on Railway Condition Monitoring RCM 2008 Switch Mechanism Diagnosis using a Pattern Recognition Approach F. Chamroukhi, A. Samé, P. Aknin The French National Institute
CS145: INTRODUCTION TO DATA MINING
 CS145: INTRODUCTION TO DATA MINING 4: Vector Data: Decision Tree Instructor: Yizhou Sun yzsun@cs.ucla.edu October 10, 2017 Methods to Learn Vector Data Set Data Sequence Data Text Data Classification Clustering
CS145: INTRODUCTION TO DATA MINING 4: Vector Data: Decision Tree Instructor: Yizhou Sun yzsun@cs.ucla.edu October 10, 2017 Methods to Learn Vector Data Set Data Sequence Data Text Data Classification Clustering
An introduction to clustering techniques
 - ABSTRACT Cluster analysis has been used in a wide variety of fields, such as marketing, social science, biology, pattern recognition etc. It is used to identify homogenous groups of cases to better understand
- ABSTRACT Cluster analysis has been used in a wide variety of fields, such as marketing, social science, biology, pattern recognition etc. It is used to identify homogenous groups of cases to better understand
Introduction to machine learning and pattern recognition Lecture 2 Coryn Bailer-Jones
 Introduction to machine learning and pattern recognition Lecture 2 Coryn Bailer-Jones http://www.mpia.de/homes/calj/mlpr_mpia2008.html 1 1 Last week... supervised and unsupervised methods need adaptive
Introduction to machine learning and pattern recognition Lecture 2 Coryn Bailer-Jones http://www.mpia.de/homes/calj/mlpr_mpia2008.html 1 1 Last week... supervised and unsupervised methods need adaptive
Unsupervised Anomaly Detection for High Dimensional Data
 Unsupervised Anomaly Detection for High Dimensional Data Department of Mathematics, Rowan University. July 19th, 2013 International Workshop in Sequential Methodologies (IWSM-2013) Outline of Talk Motivation
Unsupervised Anomaly Detection for High Dimensional Data Department of Mathematics, Rowan University. July 19th, 2013 International Workshop in Sequential Methodologies (IWSM-2013) Outline of Talk Motivation
Non-Bayesian Classifiers Part II: Linear Discriminants and Support Vector Machines
 Non-Bayesian Classifiers Part II: Linear Discriminants and Support Vector Machines Selim Aksoy Department of Computer Engineering Bilkent University saksoy@cs.bilkent.edu.tr CS 551, Fall 2018 CS 551, Fall
Non-Bayesian Classifiers Part II: Linear Discriminants and Support Vector Machines Selim Aksoy Department of Computer Engineering Bilkent University saksoy@cs.bilkent.edu.tr CS 551, Fall 2018 CS 551, Fall
Data Release 5. Sky coverage of imaging data in the DR5
 Data Release 5 The Sloan Digital Sky Survey has released its fifth Data Release (DR5). The spatial coverage of DR5 is about 20% larger than that of DR4. The photometric data in DR5 are based on five band
Data Release 5 The Sloan Digital Sky Survey has released its fifth Data Release (DR5). The spatial coverage of DR5 is about 20% larger than that of DR4. The photometric data in DR5 are based on five band
Clustering and classification with applications to microarrays and cellular phenotypes
 Clustering and classification with applications to microarrays and cellular phenotypes Gregoire Pau, EMBL Heidelberg gregoire.pau@embl.de European Molecular Biology Laboratory 1 Microarray data patients
Clustering and classification with applications to microarrays and cellular phenotypes Gregoire Pau, EMBL Heidelberg gregoire.pau@embl.de European Molecular Biology Laboratory 1 Microarray data patients
Chemometrics: Classification of spectra
 Chemometrics: Classification of spectra Vladimir Bochko Jarmo Alander University of Vaasa November 1, 2010 Vladimir Bochko Chemometrics: Classification 1/36 Contents Terminology Introduction Big picture
Chemometrics: Classification of spectra Vladimir Bochko Jarmo Alander University of Vaasa November 1, 2010 Vladimir Bochko Chemometrics: Classification 1/36 Contents Terminology Introduction Big picture
UNIVERSITY of PENNSYLVANIA CIS 520: Machine Learning Final, Fall 2013
 UNIVERSITY of PENNSYLVANIA CIS 520: Machine Learning Final, Fall 2013 Exam policy: This exam allows two one-page, two-sided cheat sheets; No other materials. Time: 2 hours. Be sure to write your name and
UNIVERSITY of PENNSYLVANIA CIS 520: Machine Learning Final, Fall 2013 Exam policy: This exam allows two one-page, two-sided cheat sheets; No other materials. Time: 2 hours. Be sure to write your name and
ECE 521. Lecture 11 (not on midterm material) 13 February K-means clustering, Dimensionality reduction
 ECE 521 Lecture 11 (not on midterm material) 13 February 2017 K-means clustering, Dimensionality reduction With thanks to Ruslan Salakhutdinov for an earlier version of the slides Overview K-means clustering
ECE 521 Lecture 11 (not on midterm material) 13 February 2017 K-means clustering, Dimensionality reduction With thanks to Ruslan Salakhutdinov for an earlier version of the slides Overview K-means clustering
Classification: Linear Discriminant Analysis
 Classification: Linear Discriminant Analysis Discriminant analysis uses sample information about individuals that are known to belong to one of several populations for the purposes of classification. Based
Classification: Linear Discriminant Analysis Discriminant analysis uses sample information about individuals that are known to belong to one of several populations for the purposes of classification. Based
Classification using stochastic ensembles
 July 31, 2014 Topics Introduction Topics Classification Application and classfication Classification and Regression Trees Stochastic ensemble methods Our application: USAID Poverty Assessment Tools Topics
July 31, 2014 Topics Introduction Topics Classification Application and classfication Classification and Regression Trees Stochastic ensemble methods Our application: USAID Poverty Assessment Tools Topics
CS145: INTRODUCTION TO DATA MINING
 CS145: INTRODUCTION TO DATA MINING Text Data: Topic Model Instructor: Yizhou Sun yzsun@cs.ucla.edu December 4, 2017 Methods to be Learnt Vector Data Set Data Sequence Data Text Data Classification Clustering
CS145: INTRODUCTION TO DATA MINING Text Data: Topic Model Instructor: Yizhou Sun yzsun@cs.ucla.edu December 4, 2017 Methods to be Learnt Vector Data Set Data Sequence Data Text Data Classification Clustering
Statistical Machine Learning
 Statistical Machine Learning Christoph Lampert Spring Semester 2015/2016 // Lecture 12 1 / 36 Unsupervised Learning Dimensionality Reduction 2 / 36 Dimensionality Reduction Given: data X = {x 1,..., x
Statistical Machine Learning Christoph Lampert Spring Semester 2015/2016 // Lecture 12 1 / 36 Unsupervised Learning Dimensionality Reduction 2 / 36 Dimensionality Reduction Given: data X = {x 1,..., x
Support Vector Machine (SVM) and Kernel Methods
 Support Vector Machine (SVM) and Kernel Methods CE-717: Machine Learning Sharif University of Technology Fall 2014 Soleymani Outline Margin concept Hard-Margin SVM Soft-Margin SVM Dual Problems of Hard-Margin
Support Vector Machine (SVM) and Kernel Methods CE-717: Machine Learning Sharif University of Technology Fall 2014 Soleymani Outline Margin concept Hard-Margin SVM Soft-Margin SVM Dual Problems of Hard-Margin
Fast Hierarchical Clustering from the Baire Distance
 Fast Hierarchical Clustering from the Baire Distance Pedro Contreras 1 and Fionn Murtagh 1,2 1 Department of Computer Science. Royal Holloway, University of London. 57 Egham Hill. Egham TW20 OEX, England.
Fast Hierarchical Clustering from the Baire Distance Pedro Contreras 1 and Fionn Murtagh 1,2 1 Department of Computer Science. Royal Holloway, University of London. 57 Egham Hill. Egham TW20 OEX, England.
Machine Learning for Signal Processing Bayes Classification and Regression
 Machine Learning for Signal Processing Bayes Classification and Regression Instructor: Bhiksha Raj 11755/18797 1 Recap: KNN A very effective and simple way of performing classification Simple model: For
Machine Learning for Signal Processing Bayes Classification and Regression Instructor: Bhiksha Raj 11755/18797 1 Recap: KNN A very effective and simple way of performing classification Simple model: For
Diffusion/Inference geometries of data features, situational awareness and visualization. Ronald R Coifman Mathematics Yale University
 Diffusion/Inference geometries of data features, situational awareness and visualization Ronald R Coifman Mathematics Yale University Digital data is generally converted to point clouds in high dimensional
Diffusion/Inference geometries of data features, situational awareness and visualization Ronald R Coifman Mathematics Yale University Digital data is generally converted to point clouds in high dimensional
Inferring Galaxy Morphology Through Texture Analysis
 Inferring Galaxy Morphology Through Texture Analysis 1 Galaxy Morphology Kinman Au, Christopher Genovese, Andrew Connolly Galaxies are not static objects; they evolve by interacting with the gas, dust
Inferring Galaxy Morphology Through Texture Analysis 1 Galaxy Morphology Kinman Au, Christopher Genovese, Andrew Connolly Galaxies are not static objects; they evolve by interacting with the gas, dust
Day 3: Classification, logistic regression
 Day 3: Classification, logistic regression Introduction to Machine Learning Summer School June 18, 2018 - June 29, 2018, Chicago Instructor: Suriya Gunasekar, TTI Chicago 20 June 2018 Topics so far Supervised
Day 3: Classification, logistic regression Introduction to Machine Learning Summer School June 18, 2018 - June 29, 2018, Chicago Instructor: Suriya Gunasekar, TTI Chicago 20 June 2018 Topics so far Supervised
Data Analysis Methods
 Data Analysis Methods Successes, Opportunities, and Challenges Chad M. Schafer Department of Statistics & Data Science Carnegie Mellon University cschafer@cmu.edu The Astro/Data Science Community LSST
Data Analysis Methods Successes, Opportunities, and Challenges Chad M. Schafer Department of Statistics & Data Science Carnegie Mellon University cschafer@cmu.edu The Astro/Data Science Community LSST
Text Mining. Dr. Yanjun Li. Associate Professor. Department of Computer and Information Sciences Fordham University
 Text Mining Dr. Yanjun Li Associate Professor Department of Computer and Information Sciences Fordham University Outline Introduction: Data Mining Part One: Text Mining Part Two: Preprocessing Text Data
Text Mining Dr. Yanjun Li Associate Professor Department of Computer and Information Sciences Fordham University Outline Introduction: Data Mining Part One: Text Mining Part Two: Preprocessing Text Data
ROBERTO BATTITI, MAURO BRUNATO. The LION Way: Machine Learning plus Intelligent Optimization. LIONlab, University of Trento, Italy, Apr 2015
 ROBERTO BATTITI, MAURO BRUNATO. The LION Way: Machine Learning plus Intelligent Optimization. LIONlab, University of Trento, Italy, Apr 2015 http://intelligentoptimization.org/lionbook Roberto Battiti
ROBERTO BATTITI, MAURO BRUNATO. The LION Way: Machine Learning plus Intelligent Optimization. LIONlab, University of Trento, Italy, Apr 2015 http://intelligentoptimization.org/lionbook Roberto Battiti
Probabilistic clustering
 Aprendizagem Automática Probabilistic clustering Ludwig Krippahl Probabilistic clustering Summary Fuzzy sets and clustering Fuzzy c-means Probabilistic Clustering: mixture models Expectation-Maximization,
Aprendizagem Automática Probabilistic clustering Ludwig Krippahl Probabilistic clustering Summary Fuzzy sets and clustering Fuzzy c-means Probabilistic Clustering: mixture models Expectation-Maximization,
Unsupervised Learning with Permuted Data
 Unsupervised Learning with Permuted Data Sergey Kirshner skirshne@ics.uci.edu Sridevi Parise sparise@ics.uci.edu Padhraic Smyth smyth@ics.uci.edu School of Information and Computer Science, University
Unsupervised Learning with Permuted Data Sergey Kirshner skirshne@ics.uci.edu Sridevi Parise sparise@ics.uci.edu Padhraic Smyth smyth@ics.uci.edu School of Information and Computer Science, University
day month year documentname/initials 1
 ECE471-571 Pattern Recognition Lecture 13 Decision Tree Hairong Qi, Gonzalez Family Professor Electrical Engineering and Computer Science University of Tennessee, Knoxville http://www.eecs.utk.edu/faculty/qi
ECE471-571 Pattern Recognition Lecture 13 Decision Tree Hairong Qi, Gonzalez Family Professor Electrical Engineering and Computer Science University of Tennessee, Knoxville http://www.eecs.utk.edu/faculty/qi
Photometric identification of blue horizontal branch stars
 Astronomy & Astrophysics manuscript no. 14381ms c ESO 2010 August 3, 2010 Photometric identification of blue horizontal branch stars K.W. Smith 1, C.A.L. Bailer-Jones 1, R.J. Klement 1, and X.X. Xue 2
Astronomy & Astrophysics manuscript no. 14381ms c ESO 2010 August 3, 2010 Photometric identification of blue horizontal branch stars K.W. Smith 1, C.A.L. Bailer-Jones 1, R.J. Klement 1, and X.X. Xue 2
Performance Comparison of K-Means and Expectation Maximization with Gaussian Mixture Models for Clustering EE6540 Final Project
 Performance Comparison of K-Means and Expectation Maximization with Gaussian Mixture Models for Clustering EE6540 Final Project Devin Cornell & Sushruth Sastry May 2015 1 Abstract In this article, we explore
Performance Comparison of K-Means and Expectation Maximization with Gaussian Mixture Models for Clustering EE6540 Final Project Devin Cornell & Sushruth Sastry May 2015 1 Abstract In this article, we explore
9/12/17. Types of learning. Modeling data. Supervised learning: Classification. Supervised learning: Regression. Unsupervised learning: Clustering
 Types of learning Modeling data Supervised: we know input and targets Goal is to learn a model that, given input data, accurately predicts target data Unsupervised: we know the input only and want to make
Types of learning Modeling data Supervised: we know input and targets Goal is to learn a model that, given input data, accurately predicts target data Unsupervised: we know the input only and want to make
Gaussian Mixtures. ## Type 'citation("mclust")' for citing this R package in publications.
 Gaussian Mixtures The galaxies data in the MASS package (Venables and Ripley, 2002) is a frequently used example for Gaussian mixture models. It contains the velocities of 82 galaxies from a redshift survey
Gaussian Mixtures The galaxies data in the MASS package (Venables and Ripley, 2002) is a frequently used example for Gaussian mixture models. It contains the velocities of 82 galaxies from a redshift survey
CS 6375 Machine Learning
 CS 6375 Machine Learning Decision Trees Instructor: Yang Liu 1 Supervised Classifier X 1 X 2. X M Ref class label 2 1 Three variables: Attribute 1: Hair = {blond, dark} Attribute 2: Height = {tall, short}
CS 6375 Machine Learning Decision Trees Instructor: Yang Liu 1 Supervised Classifier X 1 X 2. X M Ref class label 2 1 Three variables: Attribute 1: Hair = {blond, dark} Attribute 2: Height = {tall, short}
EEL 851: Biometrics. An Overview of Statistical Pattern Recognition EEL 851 1
 EEL 851: Biometrics An Overview of Statistical Pattern Recognition EEL 851 1 Outline Introduction Pattern Feature Noise Example Problem Analysis Segmentation Feature Extraction Classification Design Cycle
EEL 851: Biometrics An Overview of Statistical Pattern Recognition EEL 851 1 Outline Introduction Pattern Feature Noise Example Problem Analysis Segmentation Feature Extraction Classification Design Cycle
PATTERN CLASSIFICATION
 PATTERN CLASSIFICATION Second Edition Richard O. Duda Peter E. Hart David G. Stork A Wiley-lnterscience Publication JOHN WILEY & SONS, INC. New York Chichester Weinheim Brisbane Singapore Toronto CONTENTS
PATTERN CLASSIFICATION Second Edition Richard O. Duda Peter E. Hart David G. Stork A Wiley-lnterscience Publication JOHN WILEY & SONS, INC. New York Chichester Weinheim Brisbane Singapore Toronto CONTENTS
ECE521 week 3: 23/26 January 2017
 ECE521 week 3: 23/26 January 2017 Outline Probabilistic interpretation of linear regression - Maximum likelihood estimation (MLE) - Maximum a posteriori (MAP) estimation Bias-variance trade-off Linear
ECE521 week 3: 23/26 January 2017 Outline Probabilistic interpretation of linear regression - Maximum likelihood estimation (MLE) - Maximum a posteriori (MAP) estimation Bias-variance trade-off Linear
Final Exam, Fall 2002
 15-781 Final Exam, Fall 22 1. Write your name and your andrew email address below. Name: Andrew ID: 2. There should be 17 pages in this exam (excluding this cover sheet). 3. If you need more room to work
15-781 Final Exam, Fall 22 1. Write your name and your andrew email address below. Name: Andrew ID: 2. There should be 17 pages in this exam (excluding this cover sheet). 3. If you need more room to work
Parametric Models. Dr. Shuang LIANG. School of Software Engineering TongJi University Fall, 2012
 Parametric Models Dr. Shuang LIANG School of Software Engineering TongJi University Fall, 2012 Today s Topics Maximum Likelihood Estimation Bayesian Density Estimation Today s Topics Maximum Likelihood
Parametric Models Dr. Shuang LIANG School of Software Engineering TongJi University Fall, 2012 Today s Topics Maximum Likelihood Estimation Bayesian Density Estimation Today s Topics Maximum Likelihood
Applying cluster analysis to 2011 Census local authority data
 Applying cluster analysis to 2011 Census local authority data Kitty.Lymperopoulou@manchester.ac.uk SPSS User Group Conference November, 10 2017 Outline Basic ideas of cluster analysis How to choose variables
Applying cluster analysis to 2011 Census local authority data Kitty.Lymperopoulou@manchester.ac.uk SPSS User Group Conference November, 10 2017 Outline Basic ideas of cluster analysis How to choose variables
Introduction to Machine Learning Midterm Exam Solutions
 10-701 Introduction to Machine Learning Midterm Exam Solutions Instructors: Eric Xing, Ziv Bar-Joseph 17 November, 2015 There are 11 questions, for a total of 100 points. This exam is open book, open notes,
10-701 Introduction to Machine Learning Midterm Exam Solutions Instructors: Eric Xing, Ziv Bar-Joseph 17 November, 2015 There are 11 questions, for a total of 100 points. This exam is open book, open notes,
Detection of outliers in multivariate data:
 1 Detection of outliers in multivariate data: a method based on clustering and robust estimators Carla M. Santos-Pereira 1 and Ana M. Pires 2 1 Universidade Portucalense Infante D. Henrique, Oporto, Portugal
1 Detection of outliers in multivariate data: a method based on clustering and robust estimators Carla M. Santos-Pereira 1 and Ana M. Pires 2 1 Universidade Portucalense Infante D. Henrique, Oporto, Portugal
Support Vector Machine (SVM) & Kernel CE-717: Machine Learning Sharif University of Technology. M. Soleymani Fall 2012
 Support Vector Machine (SVM) & Kernel CE-717: Machine Learning Sharif University of Technology M. Soleymani Fall 2012 Linear classifier Which classifier? x 2 x 1 2 Linear classifier Margin concept x 2
Support Vector Machine (SVM) & Kernel CE-717: Machine Learning Sharif University of Technology M. Soleymani Fall 2012 Linear classifier Which classifier? x 2 x 1 2 Linear classifier Margin concept x 2
Lecture 15: Galaxy morphology and environment
 GALAXIES 626 Lecture 15: Galaxy morphology and environment Why classify galaxies? The Hubble system gives us our basic description of galaxies. The sequence of galaxy types may reflect an underlying physical
GALAXIES 626 Lecture 15: Galaxy morphology and environment Why classify galaxies? The Hubble system gives us our basic description of galaxies. The sequence of galaxy types may reflect an underlying physical
Contents. Preface to the second edition. Preface to the fírst edition. Acknowledgments PART I PRELIMINARIES
 Contents Foreword Preface to the second edition Preface to the fírst edition Acknowledgments xvll xix xxi xxiii PART I PRELIMINARIES CHAPTER 1 Introduction 3 1.1 What Is Data Mining? 3 1.2 Where Is Data
Contents Foreword Preface to the second edition Preface to the fírst edition Acknowledgments xvll xix xxi xxiii PART I PRELIMINARIES CHAPTER 1 Introduction 3 1.1 What Is Data Mining? 3 1.2 Where Is Data
D4.2. First release of on-line science-oriented tutorials
 EuroVO-AIDA Euro-VO Astronomical Infrastructure for Data Access D4.2 First release of on-line science-oriented tutorials Final version Grant agreement no: 212104 Combination of Collaborative Projects &
EuroVO-AIDA Euro-VO Astronomical Infrastructure for Data Access D4.2 First release of on-line science-oriented tutorials Final version Grant agreement no: 212104 Combination of Collaborative Projects &
Machine Learning Practice Page 2 of 2 10/28/13
 Machine Learning 10-701 Practice Page 2 of 2 10/28/13 1. True or False Please give an explanation for your answer, this is worth 1 pt/question. (a) (2 points) No classifier can do better than a naive Bayes
Machine Learning 10-701 Practice Page 2 of 2 10/28/13 1. True or False Please give an explanation for your answer, this is worth 1 pt/question. (a) (2 points) No classifier can do better than a naive Bayes
An improved technique for increasing the accuracy of photometrically determined redshifts for blended galaxies
 An improved technique for increasing the accuracy of photometrically determined redshifts for blended galaxies Ashley Marie Parker Marietta College, Marietta, Ohio Office of Science, Science Undergraduate
An improved technique for increasing the accuracy of photometrically determined redshifts for blended galaxies Ashley Marie Parker Marietta College, Marietta, Ohio Office of Science, Science Undergraduate
Classification for High Dimensional Problems Using Bayesian Neural Networks and Dirichlet Diffusion Trees
 Classification for High Dimensional Problems Using Bayesian Neural Networks and Dirichlet Diffusion Trees Rafdord M. Neal and Jianguo Zhang Presented by Jiwen Li Feb 2, 2006 Outline Bayesian view of feature
Classification for High Dimensional Problems Using Bayesian Neural Networks and Dirichlet Diffusion Trees Rafdord M. Neal and Jianguo Zhang Presented by Jiwen Li Feb 2, 2006 Outline Bayesian view of feature
Introduction to astrostatistics
 Introduction to astrostatistics Eric Feigelson (Astro & Astrophys) & G. Jogesh Babu (Stat) Center for Astrostatistics Penn State University Astronomy & statistics: A glorious history Hipparchus (4th c.
Introduction to astrostatistics Eric Feigelson (Astro & Astrophys) & G. Jogesh Babu (Stat) Center for Astrostatistics Penn State University Astronomy & statistics: A glorious history Hipparchus (4th c.
Computer Vision Group Prof. Daniel Cremers. 14. Clustering
 Group Prof. Daniel Cremers 14. Clustering Motivation Supervised learning is good for interaction with humans, but labels from a supervisor are hard to obtain Clustering is unsupervised learning, i.e. it
Group Prof. Daniel Cremers 14. Clustering Motivation Supervised learning is good for interaction with humans, but labels from a supervisor are hard to obtain Clustering is unsupervised learning, i.e. it
Machine Learning, Fall 2009: Midterm
 10-601 Machine Learning, Fall 009: Midterm Monday, November nd hours 1. Personal info: Name: Andrew account: E-mail address:. You are permitted two pages of notes and a calculator. Please turn off all
10-601 Machine Learning, Fall 009: Midterm Monday, November nd hours 1. Personal info: Name: Andrew account: E-mail address:. You are permitted two pages of notes and a calculator. Please turn off all
Machine Learning. Gaussian Mixture Models. Zhiyao Duan & Bryan Pardo, Machine Learning: EECS 349 Fall
 Machine Learning Gaussian Mixture Models Zhiyao Duan & Bryan Pardo, Machine Learning: EECS 349 Fall 2012 1 The Generative Model POV We think of the data as being generated from some process. We assume
Machine Learning Gaussian Mixture Models Zhiyao Duan & Bryan Pardo, Machine Learning: EECS 349 Fall 2012 1 The Generative Model POV We think of the data as being generated from some process. We assume
