Demographic Inference with Coalescent Hidden Markov Model
|
|
|
- Sharleen Gibson
- 5 years ago
- Views:
Transcription
1 Demographic Inference with Coalescent Hidden Markov Model Jade Y. Cheng Thomas Mailund Bioinformatics Research Centre Aarhus University Denmark The Thirteenth Asia Pacific Bioinformatics Conference HsinChu, Taiwan, January 2015
2 Presentation Outline CoalHMM framework Overview Model construction Continuous time Markov chain (CTMC) Hidden Markov model (HMM) Numerical optimization Genetic algorithm Particle swarm optimzation Model construction and implementation Simulation case studies Isolation model IIM with nine epochs Simulation case studies
3 Framework Overview CoalHMM is a demorgraphic inference framework based on combining the sequential Markov coalescence with hidden Markov models. E.g. Ancestral Period Migration Period Isolation Period Migration Demorgraphic parameters: 1. Isolation duration 2. Migration duration 3. Coalescent rate 4. Recombination rate 5. Migration rate Our framework is available under open source licence GPLv2 at h ps://github.com/mailund/imcoalhmm
4 Presentation Outline CoalHMM framework Overview Model construction Continuous time Markov chain (CTMC) Hidden Markov model (HMM) Numerical optimization Genetic algorithm Particle swarm optimzation Model construction and implementation Simulation case studies Isolation model IIM with nine epochs Simulation case studies
5 Hidden Markov Model In the context of CoalHMM, hidden states are different coalescence trees. E.g. for two samples, the hidden states are the coalescence times. ^ t3 x /-\ t2 / \ x /-----\ /-\ t1 / \ / \ x / \ /-----\ /-\ t0 time o o o o o o HMM state #3 HMM state #2 HMM state #1
6 HMM Transition Probabilities Transition probability T ij is the normalized joint probability J ij, which is the probability of observing coalescence of the le nucleotide in time period i and coalescence of the right nucleotide in time period j. E.g. ^ t4 x x t3 x-x t2 o-o o o t1 o-o o-o t0 time o-o o-o E.g. ^ t4 x-x t3 o x-o t2 o o o-o t1 o-o o-o t0 time o-o o-o J 33 J 34
7 Continuous Time Markov Chain CTMC state space for two samples in two isolated populations C2 C1 0 1 R - 0 C1 2 R 0 - C2 3 0 R R - 0 [(1, ([1], [])), (1, ([], [1])), (2, ([2], [])), (2, ([], [2]))] 1 [(1, ([1], [])), (1, ([], [1])), (2, ([2], [2]))] 2 [(1, ([1], [1])), (2, ([2], [])), (2, ([], [2]))] 3 [(1, ([1], [1])), (2, ([2], [2]))]
8 Continuous Time Markov Chain CTMC state space for two samples in a single population C1 C1 C1 C1 0 0 C1 0 0 C R C1 0 0 C1 0 0 C R C1 0 0 C1 0 C R C1 0 C C R C C1 0 0 C R 0 0 R C R R C C1 C C R C R C C1 C1 C R C R 0-0 C C R -
9 Continuous Time Markov Chain e size of CTMC state space grows exponentially with the number of populations, samples, or loci. E.g. CTMC for a 3 samples 3 populations 2 loci 2 donor populations 1 receiver population demographic senario has 578 states.
10 CTMC Projections We need projection matrices to move samples between time slices that have different CTMC state spaces. No Match Match Single CTMC Isolation CTMC Isolation CTMC Single CTMC
11 HMM Transition Probabilities Transition probability T ij is the normalized joint probability J ij, which is the probability of observing coalescence of the le nucleotide in time period i and coalescence of the right nucleotide in time period j. E.g. ^ t4 x x t3 x-x t2 o-o o o t1 o-o o-o t0 time o-o o-o E.g. ^ t4 x-x t3 o x-o t2 o o o-o t1 o-o o-o t0 time o-o o-o J 33 J 34
12 HMM Transition Probabilities For a two-sample CTMC, we can split its rate and probability matrices into 16 sections using the 4 state types: begin (B), le (L), right (R), and end (E) E.g. for the two-sample single-population CTMC
13 HMM Transition Probabilities ere are three possible ways to reach an E state from a B state, which is the initial condition of two samples. Goal: Possible Paths: B to E Legal Moves: B to B B to L B to R B to E L to L L to E R to R R to E #1 B to B B to E #2 B to B B to L L to L L to E #3 B to B B to R R to R R to E
14 HMM Transition Probabilities e probability of taking path 3 is the same as taking path 2. Joint probability matrix, J, is symmetric. B L R E B B-B B-L B-R B-E L L-B L-L L-R L-E P = R R-B R-L R-R R-E E E-B E-L E-R E-E P LE
15 HMM Transition Probabilities E.g. a simple joint probability calculation ^ t4 x x t3 o x o t2 o x-o t1 o o o-o t0 time o-o o-o
16 HMM Transition Probabilities E.g. another joint probability calculation involving CTMC projection. ^ t4 x-x t3 o x-o t2 o o o-o t1 o o o-o t0 time o-o o-o
17 Presentation Outline CoalHMM framework Overview Model construction Continuous time Markov chain (CTMC) Hidden Markov model (HMM) Numerical optimization Genetic algorithm Particle swarm optimzation Model construction and implementation Simulation case studies Isolation model IIM with nine epochs Simulation case studies
18 optimization optimization minimises an objective function in a manydimensional space by continuously refining a simplex. reflect p e contract p r expand p c p min shrink p max
19 A is a type of evolutionary algorithm. evaluation selection mutation crossover
20 Fitness Proportion Selection Selection: a GA chooses a relatively fit subset of individuals for breeding. E.g. fitness 40; fitness 25; fitness 18; fitness 12; fitness 5 Roule e Wheel Selection Universal Sampling Breeding Pool Breeding Pool
21 Rank Based Selection Tournament selection selects individuals with the highest fitness values from random subsets of the population. E.g. fitness 40; fitness 25; fitness 18; fitness 12; fitness 5 Tournament #1 Tournament #2 Original Population Tournament #3 Breeding Pool
22 Crossover & Mutation Crossover: it is a genetic operation used to combine pairs of individuals previously selected for breeding the following generation. One-point crossover Two-point crossover Uniform crossover Mutation: each position has a certain probability to mutate, mutation
23 Genetic Optimization PSO is another heuristic based search algorithm. velocity cognitive influence global influence
24 Presentation Outline CoalHMM framework Overview Model construction Continuous time Markov chain (CTMC) Hidden Markov model (HMM) Numerical optimization Genetic algorithm Particle swarm optimzation Model construction and implementation Simulation case studies Isolation model IIM with nine epochs Simulation case studies
25 Pipeline & Isolation Model Simulation study pipeline. O P T I M ms seq-gen CoalHMM ziphmm I S A T I O N e simplest demographic model we consider is the clean isolation model. It has three parameters. Ancestral Period Isolation Period True Value: Isolation Time True Value: Coalescent Rate True Value: Recombination Rate
26 IIM-Nine Epoch Model 10-2 Isolation Time Migration Time Recombination Rate 10 4 Coalescent Rate #1 True Value: (31.3% out of range) True Value: (3.2% out of range) True Value: True Value: (20.4% out of range) Ancestral Epoch 9 Ancestral Epoch 8 Ancestral Epoch 7 Migration Epoch 6 True Value: (23.5% out of range) Coalescent Rate #2 True Value: (21.9% out of range) Coalescent Rate #3 True Value: (15.7% out of range) Coalescent Rate #4 True Value: (39.1% out of range) Coalescent Rate #5 Migration Epoch 5 Migration Epoch 4 Isolation Epoch 3 Isolation Epoch 2 Isolation Epoch 1 Migration True Value: Coalescent Rate # (51.6% out of range) Migration Rate #1 (1.6% out of range) True Value: (28.2% out of range) Coalescent Rate #7 Migration Rate #2 True Value: (42.2% out of range) Coalescent Rate #8 Migration Rate #3 True Value: (34.4% out of range) Coalescent Rate #9 True Value: True Value: True Value: (9.4% out of range) (43.8% out of range) (42.2% out of range)
27 Discussion We discussed the construction of our statistic inference framework, CoalHMM and showed how to infer demographics with it. CTMC HMM Numerical Optimization We compared the estimation accuracy between the previously-used with newly developed optimization methods GA PSO We presented the inference results on two demographic models, a Ancestral Epoch 9 simple one and a more complex one. Ancestral Epoch 8 Ancestral Period Isolation Period Ancestral Epoch 7 Migration Epoch 6 Migration Epoch 5 Migration Epoch 4 Isolation Epoch 3 Migration Isolation Epoch 2 Isolation Epoch 1
28 Acknowledgement Thanks!
Genetic Drift in Human Evolution
 Genetic Drift in Human Evolution (Part 2 of 2) 1 Ecology and Evolutionary Biology Center for Computational Molecular Biology Brown University Outline Introduction to genetic drift Modeling genetic drift
Genetic Drift in Human Evolution (Part 2 of 2) 1 Ecology and Evolutionary Biology Center for Computational Molecular Biology Brown University Outline Introduction to genetic drift Modeling genetic drift
Hidden Markov models in population genetics and evolutionary biology
 Hidden Markov models in population genetics and evolutionary biology Gerton Lunter Wellcome Trust Centre for Human Genetics Oxford, UK April 29, 2013 Topics for today Markov chains Hidden Markov models
Hidden Markov models in population genetics and evolutionary biology Gerton Lunter Wellcome Trust Centre for Human Genetics Oxford, UK April 29, 2013 Topics for today Markov chains Hidden Markov models
Challenges when applying stochastic models to reconstruct the demographic history of populations.
 Challenges when applying stochastic models to reconstruct the demographic history of populations. Willy Rodríguez Institut de Mathématiques de Toulouse October 11, 2017 Outline 1 Introduction 2 Inverse
Challenges when applying stochastic models to reconstruct the demographic history of populations. Willy Rodríguez Institut de Mathématiques de Toulouse October 11, 2017 Outline 1 Introduction 2 Inverse
Learning ancestral genetic processes using nonparametric Bayesian models
 Learning ancestral genetic processes using nonparametric Bayesian models Kyung-Ah Sohn October 31, 2011 Committee Members: Eric P. Xing, Chair Zoubin Ghahramani Russell Schwartz Kathryn Roeder Matthew
Learning ancestral genetic processes using nonparametric Bayesian models Kyung-Ah Sohn October 31, 2011 Committee Members: Eric P. Xing, Chair Zoubin Ghahramani Russell Schwartz Kathryn Roeder Matthew
Mathematical models in population genetics II
 Mathematical models in population genetics II Anand Bhaskar Evolutionary Biology and Theory of Computing Bootcamp January 1, 014 Quick recap Large discrete-time randomly mating Wright-Fisher population
Mathematical models in population genetics II Anand Bhaskar Evolutionary Biology and Theory of Computing Bootcamp January 1, 014 Quick recap Large discrete-time randomly mating Wright-Fisher population
CSci 8980: Advanced Topics in Graphical Models Analysis of Genetic Variation
 CSci 8980: Advanced Topics in Graphical Models Analysis of Genetic Variation Instructor: Arindam Banerjee November 26, 2007 Genetic Polymorphism Single nucleotide polymorphism (SNP) Genetic Polymorphism
CSci 8980: Advanced Topics in Graphical Models Analysis of Genetic Variation Instructor: Arindam Banerjee November 26, 2007 Genetic Polymorphism Single nucleotide polymorphism (SNP) Genetic Polymorphism
Estimating Evolutionary Trees. Phylogenetic Methods
 Estimating Evolutionary Trees v if the data are consistent with infinite sites then all methods should yield the same tree v it gets more complicated when there is homoplasy, i.e., parallel or convergent
Estimating Evolutionary Trees v if the data are consistent with infinite sites then all methods should yield the same tree v it gets more complicated when there is homoplasy, i.e., parallel or convergent
Gene Genealogies Coalescence Theory. Annabelle Haudry Glasgow, July 2009
 Gene Genealogies Coalescence Theory Annabelle Haudry Glasgow, July 2009 What could tell a gene genealogy? How much diversity in the population? Has the demographic size of the population changed? How?
Gene Genealogies Coalescence Theory Annabelle Haudry Glasgow, July 2009 What could tell a gene genealogy? How much diversity in the population? Has the demographic size of the population changed? How?
ECE521 Lecture 19 HMM cont. Inference in HMM
 ECE521 Lecture 19 HMM cont. Inference in HMM Outline Hidden Markov models Model definitions and notations Inference in HMMs Learning in HMMs 2 Formally, a hidden Markov model defines a generative process
ECE521 Lecture 19 HMM cont. Inference in HMM Outline Hidden Markov models Model definitions and notations Inference in HMMs Learning in HMMs 2 Formally, a hidden Markov model defines a generative process
Frequency Spectra and Inference in Population Genetics
 Frequency Spectra and Inference in Population Genetics Although coalescent models have come to play a central role in population genetics, there are some situations where genealogies may not lead to efficient
Frequency Spectra and Inference in Population Genetics Although coalescent models have come to play a central role in population genetics, there are some situations where genealogies may not lead to efficient
Hidden Markov Models. Terminology, Representation and Basic Problems
 Hidden Markov Models Terminology, Representation and Basic Problems Data analysis? Machine learning? In bioinformatics, we analyze a lot of (sequential) data (biological sequences) to learn unknown parameters
Hidden Markov Models Terminology, Representation and Basic Problems Data analysis? Machine learning? In bioinformatics, we analyze a lot of (sequential) data (biological sequences) to learn unknown parameters
STA 414/2104: Machine Learning
 STA 414/2104: Machine Learning Russ Salakhutdinov Department of Computer Science! Department of Statistics! rsalakhu@cs.toronto.edu! http://www.cs.toronto.edu/~rsalakhu/ Lecture 9 Sequential Data So far
STA 414/2104: Machine Learning Russ Salakhutdinov Department of Computer Science! Department of Statistics! rsalakhu@cs.toronto.edu! http://www.cs.toronto.edu/~rsalakhu/ Lecture 9 Sequential Data So far
EVOLUTIONARY DISTANCES
 EVOLUTIONARY DISTANCES FROM STRINGS TO TREES Luca Bortolussi 1 1 Dipartimento di Matematica ed Informatica Università degli studi di Trieste luca@dmi.units.it Trieste, 14 th November 2007 OUTLINE 1 STRINGS:
EVOLUTIONARY DISTANCES FROM STRINGS TO TREES Luca Bortolussi 1 1 Dipartimento di Matematica ed Informatica Università degli studi di Trieste luca@dmi.units.it Trieste, 14 th November 2007 OUTLINE 1 STRINGS:
Computational statistics
 Computational statistics Combinatorial optimization Thierry Denœux February 2017 Thierry Denœux Computational statistics February 2017 1 / 37 Combinatorial optimization Assume we seek the maximum of f
Computational statistics Combinatorial optimization Thierry Denœux February 2017 Thierry Denœux Computational statistics February 2017 1 / 37 Combinatorial optimization Assume we seek the maximum of f
Reading for Lecture 13 Release v10
 Reading for Lecture 13 Release v10 Christopher Lee November 15, 2011 Contents 1 Evolutionary Trees i 1.1 Evolution as a Markov Process...................................... ii 1.2 Rooted vs. Unrooted Trees........................................
Reading for Lecture 13 Release v10 Christopher Lee November 15, 2011 Contents 1 Evolutionary Trees i 1.1 Evolution as a Markov Process...................................... ii 1.2 Rooted vs. Unrooted Trees........................................
Bustamante et al., Supplementary Nature Manuscript # 1 out of 9 Information #
 Bustamante et al., Supplementary Nature Manuscript # 1 out of 9 Details of PRF Methodology In the Poisson Random Field PRF) model, it is assumed that non-synonymous mutations at a given gene are either
Bustamante et al., Supplementary Nature Manuscript # 1 out of 9 Details of PRF Methodology In the Poisson Random Field PRF) model, it is assumed that non-synonymous mutations at a given gene are either
How robust are the predictions of the W-F Model?
 How robust are the predictions of the W-F Model? As simplistic as the Wright-Fisher model may be, it accurately describes the behavior of many other models incorporating additional complexity. Many population
How robust are the predictions of the W-F Model? As simplistic as the Wright-Fisher model may be, it accurately describes the behavior of many other models incorporating additional complexity. Many population
Introduction to Advanced Population Genetics
 Introduction to Advanced Population Genetics Learning Objectives Describe the basic model of human evolutionary history Describe the key evolutionary forces How demography can influence the site frequency
Introduction to Advanced Population Genetics Learning Objectives Describe the basic model of human evolutionary history Describe the key evolutionary forces How demography can influence the site frequency
HMM part 1. Dr Philip Jackson
 Centre for Vision Speech & Signal Processing University of Surrey, Guildford GU2 7XH. HMM part 1 Dr Philip Jackson Probability fundamentals Markov models State topology diagrams Hidden Markov models -
Centre for Vision Speech & Signal Processing University of Surrey, Guildford GU2 7XH. HMM part 1 Dr Philip Jackson Probability fundamentals Markov models State topology diagrams Hidden Markov models -
Phylogenomics of closely related species and individuals
 Phylogenomics of closely related species and individuals Matthew Rasmussen Siepel lab, Cornell University In collaboration with Manolis Kellis, MIT CSAIL February, 2013 Short time scales 1kyr-1myrs Long
Phylogenomics of closely related species and individuals Matthew Rasmussen Siepel lab, Cornell University In collaboration with Manolis Kellis, MIT CSAIL February, 2013 Short time scales 1kyr-1myrs Long
CISC 889 Bioinformatics (Spring 2004) Hidden Markov Models (II)
 CISC 889 Bioinformatics (Spring 24) Hidden Markov Models (II) a. Likelihood: forward algorithm b. Decoding: Viterbi algorithm c. Model building: Baum-Welch algorithm Viterbi training Hidden Markov models
CISC 889 Bioinformatics (Spring 24) Hidden Markov Models (II) a. Likelihood: forward algorithm b. Decoding: Viterbi algorithm c. Model building: Baum-Welch algorithm Viterbi training Hidden Markov models
Bioinformatics 2 - Lecture 4
 Bioinformatics 2 - Lecture 4 Guido Sanguinetti School of Informatics University of Edinburgh February 14, 2011 Sequences Many data types are ordered, i.e. you can naturally say what is before and what
Bioinformatics 2 - Lecture 4 Guido Sanguinetti School of Informatics University of Edinburgh February 14, 2011 Sequences Many data types are ordered, i.e. you can naturally say what is before and what
Evolutionary Models. Evolutionary Models
 Edit Operators In standard pairwise alignment, what are the allowed edit operators that transform one sequence into the other? Describe how each of these edit operations are represented on a sequence alignment
Edit Operators In standard pairwise alignment, what are the allowed edit operators that transform one sequence into the other? Describe how each of these edit operations are represented on a sequence alignment
Conditional probabilities and graphical models
 Conditional probabilities and graphical models Thomas Mailund Bioinformatics Research Centre (BiRC), Aarhus University Probability theory allows us to describe uncertainty in the processes we model within
Conditional probabilities and graphical models Thomas Mailund Bioinformatics Research Centre (BiRC), Aarhus University Probability theory allows us to describe uncertainty in the processes we model within
O 3 O 4 O 5. q 3. q 4. Transition
 Hidden Markov Models Hidden Markov models (HMM) were developed in the early part of the 1970 s and at that time mostly applied in the area of computerized speech recognition. They are first described in
Hidden Markov Models Hidden Markov models (HMM) were developed in the early part of the 1970 s and at that time mostly applied in the area of computerized speech recognition. They are first described in
6 Introduction to Population Genetics
 70 Grundlagen der Bioinformatik, SoSe 11, D. Huson, May 19, 2011 6 Introduction to Population Genetics This chapter is based on: J. Hein, M.H. Schierup and C. Wuif, Gene genealogies, variation and evolution,
70 Grundlagen der Bioinformatik, SoSe 11, D. Huson, May 19, 2011 6 Introduction to Population Genetics This chapter is based on: J. Hein, M.H. Schierup and C. Wuif, Gene genealogies, variation and evolution,
Genetic Algorithms and Genetic Programming Lecture 17
 Genetic Algorithms and Genetic Programming Lecture 17 Gillian Hayes 28th November 2006 Selection Revisited 1 Selection and Selection Pressure The Killer Instinct Memetic Algorithms Selection and Schemas
Genetic Algorithms and Genetic Programming Lecture 17 Gillian Hayes 28th November 2006 Selection Revisited 1 Selection and Selection Pressure The Killer Instinct Memetic Algorithms Selection and Schemas
STA 4273H: Statistical Machine Learning
 STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 11 Project
STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 11 Project
CS1820 Notes. hgupta1, kjline, smechery. April 3-April 5. output: plausible Ancestral Recombination Graph (ARG)
 CS1820 Notes hgupta1, kjline, smechery April 3-April 5 April 3 Notes 1 Minichiello-Durbin Algorithm input: set of sequences output: plausible Ancestral Recombination Graph (ARG) note: the optimal ARG is
CS1820 Notes hgupta1, kjline, smechery April 3-April 5 April 3 Notes 1 Minichiello-Durbin Algorithm input: set of sequences output: plausible Ancestral Recombination Graph (ARG) note: the optimal ARG is
Matt Heavner CSE710 Fall 2009
 Matt Heavner mheavner@buffalo.edu CSE710 Fall 2009 Problem Statement: Given a set of cities and corresponding locations, what is the shortest closed circuit that visits all cities without loops?? Fitness
Matt Heavner mheavner@buffalo.edu CSE710 Fall 2009 Problem Statement: Given a set of cities and corresponding locations, what is the shortest closed circuit that visits all cities without loops?? Fitness
ABCME: Summary statistics selection for ABC inference in R
 ABCME: Summary statistics selection for ABC inference in R Matt Nunes and David Balding Lancaster University UCL Genetics Institute Outline Motivation: why the ABCME package? Description of the package
ABCME: Summary statistics selection for ABC inference in R Matt Nunes and David Balding Lancaster University UCL Genetics Institute Outline Motivation: why the ABCME package? Description of the package
6 Introduction to Population Genetics
 Grundlagen der Bioinformatik, SoSe 14, D. Huson, May 18, 2014 67 6 Introduction to Population Genetics This chapter is based on: J. Hein, M.H. Schierup and C. Wuif, Gene genealogies, variation and evolution,
Grundlagen der Bioinformatik, SoSe 14, D. Huson, May 18, 2014 67 6 Introduction to Population Genetics This chapter is based on: J. Hein, M.H. Schierup and C. Wuif, Gene genealogies, variation and evolution,
Anatomy of a species tree
 Anatomy of a species tree T 1 Size of current and ancestral Populations (N) N Confidence in branches of species tree t/2n = 1 coalescent unit T 2 Branch lengths and divergence times of species & populations
Anatomy of a species tree T 1 Size of current and ancestral Populations (N) N Confidence in branches of species tree t/2n = 1 coalescent unit T 2 Branch lengths and divergence times of species & populations
Full Likelihood Inference from the Site Frequency Spectrum based on the Optimal Tree Resolution
 Full Likelihood Inference from the Site Frequency Spectrum based on the Optimal Tree Resolution Raazesh Sainudiin 1, Amandine Véber 2 March 3, 2018 1 Department of Mathematics, Uppsala University, Uppsala,
Full Likelihood Inference from the Site Frequency Spectrum based on the Optimal Tree Resolution Raazesh Sainudiin 1, Amandine Véber 2 March 3, 2018 1 Department of Mathematics, Uppsala University, Uppsala,
Some of these slides have been borrowed from Dr. Paul Lewis, Dr. Joe Felsenstein. Thanks!
 Some of these slides have been borrowed from Dr. Paul Lewis, Dr. Joe Felsenstein. Thanks! Paul has many great tools for teaching phylogenetics at his web site: http://hydrodictyon.eeb.uconn.edu/people/plewis
Some of these slides have been borrowed from Dr. Paul Lewis, Dr. Joe Felsenstein. Thanks! Paul has many great tools for teaching phylogenetics at his web site: http://hydrodictyon.eeb.uconn.edu/people/plewis
Lecture 3: Markov chains.
 1 BIOINFORMATIK II PROBABILITY & STATISTICS Summer semester 2008 The University of Zürich and ETH Zürich Lecture 3: Markov chains. Prof. Andrew Barbour Dr. Nicolas Pétrélis Adapted from a course by Dr.
1 BIOINFORMATIK II PROBABILITY & STATISTICS Summer semester 2008 The University of Zürich and ETH Zürich Lecture 3: Markov chains. Prof. Andrew Barbour Dr. Nicolas Pétrélis Adapted from a course by Dr.
Detecting selection from differentiation between populations: the FLK and hapflk approach.
 Detecting selection from differentiation between populations: the FLK and hapflk approach. Bertrand Servin bservin@toulouse.inra.fr Maria-Ines Fariello, Simon Boitard, Claude Chevalet, Magali SanCristobal,
Detecting selection from differentiation between populations: the FLK and hapflk approach. Bertrand Servin bservin@toulouse.inra.fr Maria-Ines Fariello, Simon Boitard, Claude Chevalet, Magali SanCristobal,
TUTORIAL: HYPER-HEURISTICS AND COMPUTATIONAL INTELLIGENCE
 TUTORIAL: HYPER-HEURISTICS AND COMPUTATIONAL INTELLIGENCE Nelishia Pillay School of Mathematics, Statistics and Computer Science University of KwaZulu-Natal South Africa TUTORIAL WEBSITE URL: http://titancs.ukzn.ac.za/ssci2015tutorial.aspx
TUTORIAL: HYPER-HEURISTICS AND COMPUTATIONAL INTELLIGENCE Nelishia Pillay School of Mathematics, Statistics and Computer Science University of KwaZulu-Natal South Africa TUTORIAL WEBSITE URL: http://titancs.ukzn.ac.za/ssci2015tutorial.aspx
Modelling Genetic Variations with Fragmentation-Coagulation Processes
 Modelling Genetic Variations with Fragmentation-Coagulation Processes Yee Whye Teh, Charles Blundell, Lloyd Elliott Gatsby Computational Neuroscience Unit, UCL Genetic Variations in Populations Inferring
Modelling Genetic Variations with Fragmentation-Coagulation Processes Yee Whye Teh, Charles Blundell, Lloyd Elliott Gatsby Computational Neuroscience Unit, UCL Genetic Variations in Populations Inferring
Population Genetics I. Bio
 Population Genetics I. Bio5488-2018 Don Conrad dconrad@genetics.wustl.edu Why study population genetics? Functional Inference Demographic inference: History of mankind is written in our DNA. We can learn
Population Genetics I. Bio5488-2018 Don Conrad dconrad@genetics.wustl.edu Why study population genetics? Functional Inference Demographic inference: History of mankind is written in our DNA. We can learn
ms a program for generating samples under neutral models
 ms a program for generating samples under neutral models Richard R. Hudson December 11, 2004 This document describes how to use ms, a program to generate samples under a variety of neutral models. The
ms a program for generating samples under neutral models Richard R. Hudson December 11, 2004 This document describes how to use ms, a program to generate samples under a variety of neutral models. The
CS 68: BIOINFORMATICS. Prof. Sara Mathieson Swarthmore College Spring 2018
 CS 68: BIOINFORMTICS Prof. Sara Mathieson Swarthmore College Spring 2018 Outline: pr 4 Lab 5 Examples HMM example in population genetics Recap Viterbi lgorithm Forward-Backward lgorithm Posterior Decoding
CS 68: BIOINFORMTICS Prof. Sara Mathieson Swarthmore College Spring 2018 Outline: pr 4 Lab 5 Examples HMM example in population genetics Recap Viterbi lgorithm Forward-Backward lgorithm Posterior Decoding
Models of Molecular Evolution
 Models of Molecular Evolution Bret Larget Departments of Botany and of Statistics University of Wisconsin Madison September 15, 2007 Genetics 875 (Fall 2009) Molecular Evolution September 15, 2009 1 /
Models of Molecular Evolution Bret Larget Departments of Botany and of Statistics University of Wisconsin Madison September 15, 2007 Genetics 875 (Fall 2009) Molecular Evolution September 15, 2009 1 /
Contrasts for a within-species comparative method
 Contrasts for a within-species comparative method Joseph Felsenstein, Department of Genetics, University of Washington, Box 357360, Seattle, Washington 98195-7360, USA email address: joe@genetics.washington.edu
Contrasts for a within-species comparative method Joseph Felsenstein, Department of Genetics, University of Washington, Box 357360, Seattle, Washington 98195-7360, USA email address: joe@genetics.washington.edu
Supplementary Materials for
 www.sciencemag.org/content/343/6172/747/suppl/dc1 Supplementary Materials for A Genetic Atlas of Human Admixture History Garrett Hellenthal, George B. J. Busby, Gavin Band, James F. Wilson, Cristian Capelli,
www.sciencemag.org/content/343/6172/747/suppl/dc1 Supplementary Materials for A Genetic Atlas of Human Admixture History Garrett Hellenthal, George B. J. Busby, Gavin Band, James F. Wilson, Cristian Capelli,
Hidden Markov Models. By Parisa Abedi. Slides courtesy: Eric Xing
 Hidden Markov Models By Parisa Abedi Slides courtesy: Eric Xing i.i.d to sequential data So far we assumed independent, identically distributed data Sequential (non i.i.d.) data Time-series data E.g. Speech
Hidden Markov Models By Parisa Abedi Slides courtesy: Eric Xing i.i.d to sequential data So far we assumed independent, identically distributed data Sequential (non i.i.d.) data Time-series data E.g. Speech
BMI/CS 776 Lecture 4. Colin Dewey
 BMI/CS 776 Lecture 4 Colin Dewey 2007.02.01 Outline Common nucleotide substitution models Directed graphical models Ancestral sequence inference Poisson process continuous Markov process X t0 X t1 X t2
BMI/CS 776 Lecture 4 Colin Dewey 2007.02.01 Outline Common nucleotide substitution models Directed graphical models Ancestral sequence inference Poisson process continuous Markov process X t0 X t1 X t2
Note Set 5: Hidden Markov Models
 Note Set 5: Hidden Markov Models Probabilistic Learning: Theory and Algorithms, CS 274A, Winter 2016 1 Hidden Markov Models (HMMs) 1.1 Introduction Consider observed data vectors x t that are d-dimensional
Note Set 5: Hidden Markov Models Probabilistic Learning: Theory and Algorithms, CS 274A, Winter 2016 1 Hidden Markov Models (HMMs) 1.1 Introduction Consider observed data vectors x t that are d-dimensional
Noisy Pattern Search using Hidden Markov Models
 Noisy Pattern Search using Hidden Markov Models David Barber University College London March 3, 22 String search We have a pattern string david that we wish to search for in an observation string, say
Noisy Pattern Search using Hidden Markov Models David Barber University College London March 3, 22 String search We have a pattern string david that we wish to search for in an observation string, say
DNA Phylogeny. Signals and Systems in Biology Kushal EE, IIT Delhi
 DNA Phylogeny Signals and Systems in Biology Kushal Shah @ EE, IIT Delhi Phylogenetics Grouping and Division of organisms Keeps changing with time Splitting, hybridization and termination Cladistics :
DNA Phylogeny Signals and Systems in Biology Kushal Shah @ EE, IIT Delhi Phylogenetics Grouping and Division of organisms Keeps changing with time Splitting, hybridization and termination Cladistics :
Statistical Model Checking as Feedback Control
 Statistical Model Checking as Feedback Control, MSc Vienna University of Technology Supervisor: Radu Grosu Co-supervisor: Ezio Bartocci Analysis of CPS: Challenges State-space explosion: Open, physical
Statistical Model Checking as Feedback Control, MSc Vienna University of Technology Supervisor: Radu Grosu Co-supervisor: Ezio Bartocci Analysis of CPS: Challenges State-space explosion: Open, physical
Intraspecific gene genealogies: trees grafting into networks
 Intraspecific gene genealogies: trees grafting into networks by David Posada & Keith A. Crandall Kessy Abarenkov Tartu, 2004 Article describes: Population genetics principles Intraspecific genetic variation
Intraspecific gene genealogies: trees grafting into networks by David Posada & Keith A. Crandall Kessy Abarenkov Tartu, 2004 Article describes: Population genetics principles Intraspecific genetic variation
Sequence Analysis 17: lecture 5. Substitution matrices Multiple sequence alignment
 Sequence Analysis 17: lecture 5 Substitution matrices Multiple sequence alignment Substitution matrices Used to score aligned positions, usually of amino acids. Expressed as the log-likelihood ratio of
Sequence Analysis 17: lecture 5 Substitution matrices Multiple sequence alignment Substitution matrices Used to score aligned positions, usually of amino acids. Expressed as the log-likelihood ratio of
Cladistics and Bioinformatics Questions 2013
 AP Biology Name Cladistics and Bioinformatics Questions 2013 1. The following table shows the percentage similarity in sequences of nucleotides from a homologous gene derived from five different species
AP Biology Name Cladistics and Bioinformatics Questions 2013 1. The following table shows the percentage similarity in sequences of nucleotides from a homologous gene derived from five different species
Linear Dynamical Systems (Kalman filter)
 Linear Dynamical Systems (Kalman filter) (a) Overview of HMMs (b) From HMMs to Linear Dynamical Systems (LDS) 1 Markov Chains with Discrete Random Variables x 1 x 2 x 3 x T Let s assume we have discrete
Linear Dynamical Systems (Kalman filter) (a) Overview of HMMs (b) From HMMs to Linear Dynamical Systems (LDS) 1 Markov Chains with Discrete Random Variables x 1 x 2 x 3 x T Let s assume we have discrete
Today s Lecture: HMMs
 Today s Lecture: HMMs Definitions Examples Probability calculations WDAG Dynamic programming algorithms: Forward Viterbi Parameter estimation Viterbi training 1 Hidden Markov Models Probability models
Today s Lecture: HMMs Definitions Examples Probability calculations WDAG Dynamic programming algorithms: Forward Viterbi Parameter estimation Viterbi training 1 Hidden Markov Models Probability models
Populations in statistical genetics
 Populations in statistical genetics What are they, and how can we infer them from whole genome data? Daniel Lawson Heilbronn Institute, University of Bristol www.paintmychromosomes.com Work with: January
Populations in statistical genetics What are they, and how can we infer them from whole genome data? Daniel Lawson Heilbronn Institute, University of Bristol www.paintmychromosomes.com Work with: January
Hidden Markov Models. Aarti Singh Slides courtesy: Eric Xing. Machine Learning / Nov 8, 2010
 Hidden Markov Models Aarti Singh Slides courtesy: Eric Xing Machine Learning 10-701/15-781 Nov 8, 2010 i.i.d to sequential data So far we assumed independent, identically distributed data Sequential data
Hidden Markov Models Aarti Singh Slides courtesy: Eric Xing Machine Learning 10-701/15-781 Nov 8, 2010 i.i.d to sequential data So far we assumed independent, identically distributed data Sequential data
Supplementary Material to Full Likelihood Inference from the Site Frequency Spectrum based on the Optimal Tree Resolution
 Supplementary Material to Full Likelihood Inference from the Site Frequency Spectrum based on the Optimal Tree Resolution Raazesh Sainudiin, Amandine Véber 1 Pseudo-code First, the global process MakeHistory
Supplementary Material to Full Likelihood Inference from the Site Frequency Spectrum based on the Optimal Tree Resolution Raazesh Sainudiin, Amandine Véber 1 Pseudo-code First, the global process MakeHistory
HMM for modeling aligned multiple sequences: phylo-hmm & multivariate HMM
 I529: Machine Learning in Bioinformatics (Spring 2017) HMM for modeling aligned multiple sequences: phylo-hmm & multivariate HMM Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington
I529: Machine Learning in Bioinformatics (Spring 2017) HMM for modeling aligned multiple sequences: phylo-hmm & multivariate HMM Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington
Jed Chou. April 13, 2015
 of of CS598 AGB April 13, 2015 Overview of 1 2 3 4 5 Competing Approaches of Two competing approaches to species tree inference: Summary methods: estimate a tree on each gene alignment then combine gene
of of CS598 AGB April 13, 2015 Overview of 1 2 3 4 5 Competing Approaches of Two competing approaches to species tree inference: Summary methods: estimate a tree on each gene alignment then combine gene
Closed-form sampling formulas for the coalescent with recombination
 0 / 21 Closed-form sampling formulas for the coalescent with recombination Yun S. Song CS Division and Department of Statistics University of California, Berkeley September 7, 2009 Joint work with Paul
0 / 21 Closed-form sampling formulas for the coalescent with recombination Yun S. Song CS Division and Department of Statistics University of California, Berkeley September 7, 2009 Joint work with Paul
The Wright-Fisher Model and Genetic Drift
 The Wright-Fisher Model and Genetic Drift January 22, 2015 1 1 Hardy-Weinberg Equilibrium Our goal is to understand the dynamics of allele and genotype frequencies in an infinite, randomlymating population
The Wright-Fisher Model and Genetic Drift January 22, 2015 1 1 Hardy-Weinberg Equilibrium Our goal is to understand the dynamics of allele and genotype frequencies in an infinite, randomlymating population
A (short) introduction to phylogenetics
 A (short) introduction to phylogenetics Thibaut Jombart, Marie-Pauline Beugin MRC Centre for Outbreak Analysis and Modelling Imperial College London Genetic data analysis with PR Statistics, Millport Field
A (short) introduction to phylogenetics Thibaut Jombart, Marie-Pauline Beugin MRC Centre for Outbreak Analysis and Modelling Imperial College London Genetic data analysis with PR Statistics, Millport Field
Copyright 2000 N. AYDIN. All rights reserved. 1
 Introduction to Bioinformatics Prof. Dr. Nizamettin AYDIN naydin@yildiz.edu.tr Multiple Sequence Alignment Outline Multiple sequence alignment introduction to msa methods of msa progressive global alignment
Introduction to Bioinformatics Prof. Dr. Nizamettin AYDIN naydin@yildiz.edu.tr Multiple Sequence Alignment Outline Multiple sequence alignment introduction to msa methods of msa progressive global alignment
Hidden Markov Models. Ivan Gesteira Costa Filho IZKF Research Group Bioinformatics RWTH Aachen Adapted from:
 Hidden Markov Models Ivan Gesteira Costa Filho IZKF Research Group Bioinformatics RWTH Aachen Adapted from: www.ioalgorithms.info Outline CG-islands The Fair Bet Casino Hidden Markov Model Decoding Algorithm
Hidden Markov Models Ivan Gesteira Costa Filho IZKF Research Group Bioinformatics RWTH Aachen Adapted from: www.ioalgorithms.info Outline CG-islands The Fair Bet Casino Hidden Markov Model Decoding Algorithm
Evolutionary Genetics: Part 0.2 Introduction to Population genetics
 Evolutionary Genetics: Part 0.2 Introduction to Population genetics S. chilense S. peruvianum Winter Semester 2012-2013 Prof Aurélien Tellier FG Populationsgenetik Population genetics Evolution = changes
Evolutionary Genetics: Part 0.2 Introduction to Population genetics S. chilense S. peruvianum Winter Semester 2012-2013 Prof Aurélien Tellier FG Populationsgenetik Population genetics Evolution = changes
An Evolutionary Programming Based Algorithm for HMM training
 An Evolutionary Programming Based Algorithm for HMM training Ewa Figielska,Wlodzimierz Kasprzak Institute of Control and Computation Engineering, Warsaw University of Technology ul. Nowowiejska 15/19,
An Evolutionary Programming Based Algorithm for HMM training Ewa Figielska,Wlodzimierz Kasprzak Institute of Control and Computation Engineering, Warsaw University of Technology ul. Nowowiejska 15/19,
Introduction to Machine Learning CMU-10701
 Introduction to Machine Learning CMU-10701 Hidden Markov Models Barnabás Póczos & Aarti Singh Slides courtesy: Eric Xing i.i.d to sequential data So far we assumed independent, identically distributed
Introduction to Machine Learning CMU-10701 Hidden Markov Models Barnabás Póczos & Aarti Singh Slides courtesy: Eric Xing i.i.d to sequential data So far we assumed independent, identically distributed
Lecture 4: Hidden Markov Models: An Introduction to Dynamic Decision Making. November 11, 2010
 Hidden Lecture 4: Hidden : An Introduction to Dynamic Decision Making November 11, 2010 Special Meeting 1/26 Markov Model Hidden When a dynamical system is probabilistic it may be determined by the transition
Hidden Lecture 4: Hidden : An Introduction to Dynamic Decision Making November 11, 2010 Special Meeting 1/26 Markov Model Hidden When a dynamical system is probabilistic it may be determined by the transition
Sequence Modelling with Features: Linear-Chain Conditional Random Fields. COMP-599 Oct 6, 2015
 Sequence Modelling with Features: Linear-Chain Conditional Random Fields COMP-599 Oct 6, 2015 Announcement A2 is out. Due Oct 20 at 1pm. 2 Outline Hidden Markov models: shortcomings Generative vs. discriminative
Sequence Modelling with Features: Linear-Chain Conditional Random Fields COMP-599 Oct 6, 2015 Announcement A2 is out. Due Oct 20 at 1pm. 2 Outline Hidden Markov models: shortcomings Generative vs. discriminative
Genetic Algorithm. Outline
 Genetic Algorithm 056: 166 Production Systems Shital Shah SPRING 2004 Outline Genetic Algorithm (GA) Applications Search space Step-by-step GA Mechanism Examples GA performance Other GA examples 1 Genetic
Genetic Algorithm 056: 166 Production Systems Shital Shah SPRING 2004 Outline Genetic Algorithm (GA) Applications Search space Step-by-step GA Mechanism Examples GA performance Other GA examples 1 Genetic
Cheng Soon Ong & Christian Walder. Canberra February June 2018
 Cheng Soon Ong & Christian Walder Research Group and College of Engineering and Computer Science Canberra February June 218 Outlines Overview Introduction Linear Algebra Probability Linear Regression 1
Cheng Soon Ong & Christian Walder Research Group and College of Engineering and Computer Science Canberra February June 218 Outlines Overview Introduction Linear Algebra Probability Linear Regression 1
3/1/17. Content. TWINSCAN model. Example. TWINSCAN algorithm. HMM for modeling aligned multiple sequences: phylo-hmm & multivariate HMM
 I529: Machine Learning in Bioinformatics (Spring 2017) Content HMM for modeling aligned multiple sequences: phylo-hmm & multivariate HMM Yuzhen Ye School of Informatics and Computing Indiana University,
I529: Machine Learning in Bioinformatics (Spring 2017) Content HMM for modeling aligned multiple sequences: phylo-hmm & multivariate HMM Yuzhen Ye School of Informatics and Computing Indiana University,
Bioinformatics (GLOBEX, Summer 2015) Pairwise sequence alignment
 Bioinformatics (GLOBEX, Summer 2015) Pairwise sequence alignment Substitution score matrices, PAM, BLOSUM Needleman-Wunsch algorithm (Global) Smith-Waterman algorithm (Local) BLAST (local, heuristic) E-value
Bioinformatics (GLOBEX, Summer 2015) Pairwise sequence alignment Substitution score matrices, PAM, BLOSUM Needleman-Wunsch algorithm (Global) Smith-Waterman algorithm (Local) BLAST (local, heuristic) E-value
Supplemental Information Likelihood-based inference in isolation-by-distance models using the spatial distribution of low-frequency alleles
 Supplemental Information Likelihood-based inference in isolation-by-distance models using the spatial distribution of low-frequency alleles John Novembre and Montgomery Slatkin Supplementary Methods To
Supplemental Information Likelihood-based inference in isolation-by-distance models using the spatial distribution of low-frequency alleles John Novembre and Montgomery Slatkin Supplementary Methods To
Titles and Abstracts
 Titles and Abstracts Stability of the Nonlinear Filter for Random Expanding Maps Jochen Broecker A ubiquitous problem in science and engineering is to reconstruct the state of a hidden Markov process (the
Titles and Abstracts Stability of the Nonlinear Filter for Random Expanding Maps Jochen Broecker A ubiquitous problem in science and engineering is to reconstruct the state of a hidden Markov process (the
Hidden Markov Models Part 1: Introduction
 Hidden Markov Models Part 1: Introduction CSE 6363 Machine Learning Vassilis Athitsos Computer Science and Engineering Department University of Texas at Arlington 1 Modeling Sequential Data Suppose that
Hidden Markov Models Part 1: Introduction CSE 6363 Machine Learning Vassilis Athitsos Computer Science and Engineering Department University of Texas at Arlington 1 Modeling Sequential Data Suppose that
Sequence Alignment Techniques and Their Uses
 Sequence Alignment Techniques and Their Uses Sarah Fiorentino Since rapid sequencing technology and whole genomes sequencing, the amount of sequence information has grown exponentially. With all of this
Sequence Alignment Techniques and Their Uses Sarah Fiorentino Since rapid sequencing technology and whole genomes sequencing, the amount of sequence information has grown exponentially. With all of this
Week 7.2 Ch 4 Microevolutionary Proceses
 Week 7.2 Ch 4 Microevolutionary Proceses 1 Mendelian Traits vs Polygenic Traits Mendelian -discrete -single gene determines effect -rarely influenced by environment Polygenic: -continuous -multiple genes
Week 7.2 Ch 4 Microevolutionary Proceses 1 Mendelian Traits vs Polygenic Traits Mendelian -discrete -single gene determines effect -rarely influenced by environment Polygenic: -continuous -multiple genes
The Phylogenetic Handbook
 The Phylogenetic Handbook A Practical Approach to DNA and Protein Phylogeny Edited by Marco Salemi University of California, Irvine and Katholieke Universiteit Leuven, Belgium and Anne-Mieke Vandamme Rega
The Phylogenetic Handbook A Practical Approach to DNA and Protein Phylogeny Edited by Marco Salemi University of California, Irvine and Katholieke Universiteit Leuven, Belgium and Anne-Mieke Vandamme Rega
DNA Feature Sensors. B. Majoros
 DNA Feature Sensors B. Majoros What is Feature Sensing? A feature is any DNA subsequence of biological significance. For practical reasons, we recognize two broad classes of features: signals short, fixed-length
DNA Feature Sensors B. Majoros What is Feature Sensing? A feature is any DNA subsequence of biological significance. For practical reasons, we recognize two broad classes of features: signals short, fixed-length
Lecture 21: Spectral Learning for Graphical Models
 10-708: Probabilistic Graphical Models 10-708, Spring 2016 Lecture 21: Spectral Learning for Graphical Models Lecturer: Eric P. Xing Scribes: Maruan Al-Shedivat, Wei-Cheng Chang, Frederick Liu 1 Motivation
10-708: Probabilistic Graphical Models 10-708, Spring 2016 Lecture 21: Spectral Learning for Graphical Models Lecturer: Eric P. Xing Scribes: Maruan Al-Shedivat, Wei-Cheng Chang, Frederick Liu 1 Motivation
A concentration fluctuation model for virtual testing of detection systems
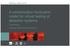 A concentration fluctuation model for virtual testing of detection systems Presented by: Dr Martyn Bull and Dr Robert Gordon Contents Rob Gordon - Overview Definition of concentration fluctuations Why
A concentration fluctuation model for virtual testing of detection systems Presented by: Dr Martyn Bull and Dr Robert Gordon Contents Rob Gordon - Overview Definition of concentration fluctuations Why
Chapter 8: Introduction to Evolutionary Computation
 Computational Intelligence: Second Edition Contents Some Theories about Evolution Evolution is an optimization process: the aim is to improve the ability of an organism to survive in dynamically changing
Computational Intelligence: Second Edition Contents Some Theories about Evolution Evolution is an optimization process: the aim is to improve the ability of an organism to survive in dynamically changing
Introduction to population genetics & evolution
 Introduction to population genetics & evolution Course Organization Exam dates: Feb 19 March 1st Has everybody registered? Did you get the email with the exam schedule Summer seminar: Hot topics in Bioinformatics
Introduction to population genetics & evolution Course Organization Exam dates: Feb 19 March 1st Has everybody registered? Did you get the email with the exam schedule Summer seminar: Hot topics in Bioinformatics
Reduced-Rank Hidden Markov Models
 Reduced-Rank Hidden Markov Models Sajid M. Siddiqi Byron Boots Geoffrey J. Gordon Carnegie Mellon University ... x 1 x 2 x 3 x τ y 1 y 2 y 3 y τ Sequence of observations: Y =[y 1 y 2 y 3... y τ ] Assume
Reduced-Rank Hidden Markov Models Sajid M. Siddiqi Byron Boots Geoffrey J. Gordon Carnegie Mellon University ... x 1 x 2 x 3 x τ y 1 y 2 y 3 y τ Sequence of observations: Y =[y 1 y 2 y 3... y τ ] Assume
26 : Spectral GMs. Lecturer: Eric P. Xing Scribes: Guillermo A Cidre, Abelino Jimenez G.
 10-708: Probabilistic Graphical Models, Spring 2015 26 : Spectral GMs Lecturer: Eric P. Xing Scribes: Guillermo A Cidre, Abelino Jimenez G. 1 Introduction A common task in machine learning is to work with
10-708: Probabilistic Graphical Models, Spring 2015 26 : Spectral GMs Lecturer: Eric P. Xing Scribes: Guillermo A Cidre, Abelino Jimenez G. 1 Introduction A common task in machine learning is to work with
Evolutionary Computation Theory. Jun He School of Computer Science University of Birmingham Web: jxh
 Evolutionary Computation Theory Jun He School of Computer Science University of Birmingham Web: www.cs.bham.ac.uk/ jxh Outline Motivation History Schema Theorem Convergence and Convergence Rate Computational
Evolutionary Computation Theory Jun He School of Computer Science University of Birmingham Web: www.cs.bham.ac.uk/ jxh Outline Motivation History Schema Theorem Convergence and Convergence Rate Computational
Bayesian Networks BY: MOHAMAD ALSABBAGH
 Bayesian Networks BY: MOHAMAD ALSABBAGH Outlines Introduction Bayes Rule Bayesian Networks (BN) Representation Size of a Bayesian Network Inference via BN BN Learning Dynamic BN Introduction Conditional
Bayesian Networks BY: MOHAMAD ALSABBAGH Outlines Introduction Bayes Rule Bayesian Networks (BN) Representation Size of a Bayesian Network Inference via BN BN Learning Dynamic BN Introduction Conditional
NEUROEVOLUTION. Contents. Evolutionary Computation. Neuroevolution. Types of neuro-evolution algorithms
 Contents Evolutionary Computation overview NEUROEVOLUTION Presenter: Vlad Chiriacescu Neuroevolution overview Issues in standard Evolutionary Computation NEAT method Complexification in competitive coevolution
Contents Evolutionary Computation overview NEUROEVOLUTION Presenter: Vlad Chiriacescu Neuroevolution overview Issues in standard Evolutionary Computation NEAT method Complexification in competitive coevolution
Bioinformatics: Network Analysis
 Bioinformatics: Network Analysis Model Fitting COMP 572 (BIOS 572 / BIOE 564) - Fall 2013 Luay Nakhleh, Rice University 1 Outline Parameter estimation Model selection 2 Parameter Estimation 3 Generally
Bioinformatics: Network Analysis Model Fitting COMP 572 (BIOS 572 / BIOE 564) - Fall 2013 Luay Nakhleh, Rice University 1 Outline Parameter estimation Model selection 2 Parameter Estimation 3 Generally
Algorithms in Bioinformatics FOUR Pairwise Sequence Alignment. Pairwise Sequence Alignment. Convention: DNA Sequences 5. Sequence Alignment
 Algorithms in Bioinformatics FOUR Sami Khuri Department of Computer Science San José State University Pairwise Sequence Alignment Homology Similarity Global string alignment Local string alignment Dot
Algorithms in Bioinformatics FOUR Sami Khuri Department of Computer Science San José State University Pairwise Sequence Alignment Homology Similarity Global string alignment Local string alignment Dot
Three Steps toward Tuning the Coordinate Systems in Nature-Inspired Optimization Algorithms
 Three Steps toward Tuning the Coordinate Systems in Nature-Inspired Optimization Algorithms Yong Wang and Zhi-Zhong Liu School of Information Science and Engineering Central South University ywang@csu.edu.cn
Three Steps toward Tuning the Coordinate Systems in Nature-Inspired Optimization Algorithms Yong Wang and Zhi-Zhong Liu School of Information Science and Engineering Central South University ywang@csu.edu.cn
Linear Dynamical Systems
 Linear Dynamical Systems Sargur N. srihari@cedar.buffalo.edu Machine Learning Course: http://www.cedar.buffalo.edu/~srihari/cse574/index.html Two Models Described by Same Graph Latent variables Observations
Linear Dynamical Systems Sargur N. srihari@cedar.buffalo.edu Machine Learning Course: http://www.cedar.buffalo.edu/~srihari/cse574/index.html Two Models Described by Same Graph Latent variables Observations
Stochastic Demography, Coalescents, and Effective Population Size
 Demography Stochastic Demography, Coalescents, and Effective Population Size Steve Krone University of Idaho Department of Mathematics & IBEST Demographic effects bottlenecks, expansion, fluctuating population
Demography Stochastic Demography, Coalescents, and Effective Population Size Steve Krone University of Idaho Department of Mathematics & IBEST Demographic effects bottlenecks, expansion, fluctuating population
A representation for the semigroup of a two-level Fleming Viot process in terms of the Kingman nested coalescent
 A representation for the semigroup of a two-level Fleming Viot process in terms of the Kingman nested coalescent Airam Blancas June 11, 017 Abstract Simple nested coalescent were introduced in [1] to model
A representation for the semigroup of a two-level Fleming Viot process in terms of the Kingman nested coalescent Airam Blancas June 11, 017 Abstract Simple nested coalescent were introduced in [1] to model
Geometry of Phylogenetic Inference
 Geometry of Phylogenetic Inference Matilde Marcolli CS101: Mathematical and Computational Linguistics Winter 2015 References N. Eriksson, K. Ranestad, B. Sturmfels, S. Sullivant, Phylogenetic algebraic
Geometry of Phylogenetic Inference Matilde Marcolli CS101: Mathematical and Computational Linguistics Winter 2015 References N. Eriksson, K. Ranestad, B. Sturmfels, S. Sullivant, Phylogenetic algebraic
Search. Search is a key component of intelligent problem solving. Get closer to the goal if time is not enough
 Search Search is a key component of intelligent problem solving Search can be used to Find a desired goal if time allows Get closer to the goal if time is not enough section 11 page 1 The size of the search
Search Search is a key component of intelligent problem solving Search can be used to Find a desired goal if time allows Get closer to the goal if time is not enough section 11 page 1 The size of the search
The Combinatorial Interpretation of Formulas in Coalescent Theory
 The Combinatorial Interpretation of Formulas in Coalescent Theory John L. Spouge National Center for Biotechnology Information NLM, NIH, DHHS spouge@ncbi.nlm.nih.gov Bldg. A, Rm. N 0 NCBI, NLM, NIH Bethesda
The Combinatorial Interpretation of Formulas in Coalescent Theory John L. Spouge National Center for Biotechnology Information NLM, NIH, DHHS spouge@ncbi.nlm.nih.gov Bldg. A, Rm. N 0 NCBI, NLM, NIH Bethesda
