Hidden Markov Models. Main source: Durbin et al., Biological Sequence Alignment (Cambridge, 98)
|
|
|
- Clarence Greer
- 5 years ago
- Views:
Transcription
1 Hidden Markov Models Main source: Durbin et al., Biological Sequence Alignment (Cambridge, 98) 1
2 The occasionally dishonest casino A P A (1) = P A (2) = = 1/6 P A->B = P B->A = 1/10 B P B (1)= P B (5)=0.1 P B (6) = Can we tell when the loaded die is used? 2
3 CpG islands: Example - CpG islands DNA stretches (100~1000bp) with frequent CG pairs (contiguous on same strand). Rare, appear in significant genomic parts. Problem (1): Given a short genome sequence, decide if it comes from a CpG island. 3
4 Preliminaries: Markov Chains (S, A, p) S: State set p: Initial state prob. vector {p(x 1 =s)} [alternatively, use a begin state] A: Transition prob. matrix a st = P(x i =t x i-1 =s) Assumption: X=x 1 x n is a random process with memory length 1, i.e.: s i S P(x i =s i x 1 =s 1,,x i-1 =s i-1 ) = P(x i =s i x i-1 =s i-1 ) = a s i-1,si Sequence probability: P(X) = p(x 1 ) Π i=2 L a x i-1,xi
5 Markov Models - Transition probs for non-cpg islands + Transition probs for CpG islands - A C G T A C G T A C G T A C G T
6 CpG islands: Fixed Window Problem (1): Given a short genome sequence X, decide if it comes from a CpG island. Solution: Model by a Markov chain. Let a + st : transition prob. in CpG islands, a - st : transition prob. outside CpG islands. Decide by log-likelihood ratio score: score( X ) P( X CpG island) = log P( X non CpG island) 1 bits _ score( X ) = n n + xi 1,x log2 i = 1 ax,x a i 1 i i = n log a a + x i 1 i = x,x 1 i 1,x i i 6
7 Discrimination of sequences via Markov Chains 48 CpG islands, tot length ~60K nt. Similar non-cpg. Durbin et. al, Fig
8 CpG islands the general case Problem(2): Detect CpG islands in a long DNA sequence. Naive Solution - Sliding windows: 1 k L-l, window: X k = (x k+1,,x k+l ) score: score(x k ) positive score potential CpG island Disadvantage: what is the length of the islands? How do we identify transitions? Idea: Use Markov chains as before, with additional (hidden) states 8
9 Hidden Markov Model (HMM) Alphabet of symbols Example: {A, C, G, T} path Π=π 1,,π n (sequence of states - simple Markov chain; convention: π 0 - begin, π L+1 - end) Given sequence X = (x 1,,x L ): a kl = P(π i =l π i-1 =k), e k (b) = P(x i =b π i =k) Finite set of states, capable of emitting symbols. Example: Q = {A +,C +,G +,T +,A -,C -,G -,T - } M=(Σ, Q, Θ) Θ=(A,E) A: Transition prob. a kl k,l Q E: Emission prob. e k (b) k Q, b Σ P(X,Π) = a π0,π 1 Π i=1 L e π i(x i ) a π i,πi+1 Ex.: express P(X,Π) in terms of counts 9
10 Viterbi s Decoding Algorithm (finding most probable state path) Want: path Π maximizing P(X, Π) v k (i) = prob. of most probable path ending in state k at step i. Init: v 0 (0) = 1; v k (0)=0 k>0 Step: v k (i+1)=e k (x i+1 ) max l {v l (i) a lk } End: P(X, Π * ) = max l {v l (L) a l0 } Time complexity: O(Ln 2 ) for n states, L steps Can find Π* using back pointers. 10
11 The occasionally dishonest casino (2) A B
12 The occasionally dishonest casino (2) 12
13 HMM for CpG Islands States: A + C + G + T + A - C - G - T - Symbols: A C G T A C G T Path Π=π 1,,π n : sequence of states + A C G T A C G T A C G T A C G transition prob. T
14 Posterior State Probabilities Goal: calculate P(π i =k X) Our strategy: P(X, π i =k) = = P(x 1,,x i, π i =k) P(x i+1,,x L x 1,,x i, π i =k) = P(x 1,,x i, π i =k) P(x i+1,,x L π i =k) P(π i =k X) = P(π i =k, X) / P(X) Need to compute these two terms - and P(X) 14
15 Forward Algorithm Goal: calculate P(X) = Σ Π P(X, Π) Approximation: take max path Π* from Viterbi alg. Not justified when several near maximal paths Exact alg : Forward Algorithm f k (i) = P(x 1,,x i, π i =k) Init: f 0 (0) = 1; f k (0)=0 k>0 Step: f k (i+1) = e k (x i+1 ) l f l (i) a lk End: P(X) = Σ l f l (L) a l0 15
16 Backward Algorithm b k (i) = P(x i+1, x L π i =k) init: k, b k (L) = a k0 step: b k (i) = l a kl e l (x i+1 ) b l (i+1) End: P(X) = Σ k a 0k e k (x 1 ) b k (1) 16
17 Posterior State Probabilities (2) Goal: calculate P(π i =k X) Recall: f k (i) = P(x 1,,x i, π i =k) b k (i) = P(x i+1, x L π i =k) Each can be used to compute P(X) P(X, π i =k) = = P(x 1,,x i, π i =k) P(x i+1,,x L x 1,,x i, π i =k) = P(x 1,,x i, π i =k) P(x i+1,,x L π i =k) = f k (i) b k (i) P(π i =k X) = P(π i =k, X) / P(X) 17
18 Dishonest Casino (3) Durbin et al. pp
19 Posterior Decoding Now we have P(π i =k X). How do we decode? 1. π i* =argmax k P(π i =k X) Good when interested in state at particular point path of states π 1*,.., π * L may not be legal 2. Define a function of interest g(i) on the states. Compute G(i X) = Σ k P(π i =k X) g(k) E.g.: g(i) =1 for states in S, 0 on the rest: G(i X) is posterior prob of symbol i coming from S e.g., CpG island 19 S={A +,C +,G +,T + }
20 Parameter Estimation for HMMs Log likelihood of model Input: X 1,,X n independent training sequences Goal: estimation of Θ = (A,E) (model parameters) Note: P(X 1,,X n Θ) = Π i=1 n P(X i Θ) (indep.) l(x 1,,x n Θ) = log P(X 1,,X n Θ) = i=1 n log P(X i Θ) Case 1 - Estimation When State Sequence is Known: A kl = #(occurred k l transitions) E k (b) = #(emissions of symbol b that occurred in state k) Max. Likelihood Estimators: a kl e k (b) = A kl / l A kl = E k (b) / b E k (b ) small sample, or prior knowledge correction: A kl = A kl + r kl E k(b) = E k (b) + r k (b) Dirichlet priors 20
21 Parameter Estimation in HMM Case 2: -Estimation When States are Unknown Input: X 1,,X n indep training sequences Baum-Welch alg. (1972): Expectation: guarantees convergence monotone many local optima special case of EM compute expected no. of k l state transitions: (ex.) P(π i =k, π i+1 =l X, Θ) = [1/P(x)] f k (i) a kl e l (x i+1 ) b l (i+1) A kl = j [1/P(X j )] i f kj (i) a kl e l (x j i+1) b lj (i+1) compute expected no. of symbol b appearances in state k E k (b) = j [1/P(X j )] {i x j i =b} f kj (i) b kj (i) (ex.) Maximization: re-compute new parameters from A, E using max. likelihood. repeat (1)+(2) until improvement ε 21
22 Profile HMMs (Haussler et al, 1993) Ungapped alignment AG---C of X against a profile M: e i (a) = prob. of A-AT-C observing a at position i. P(X M) = Π i=1 L AG-AA- e i (x i ), or Score(X M) --AAAC = i=1 L log [e i (x i ) / q x i] indels: AG---C background prob. insert I match states begin j M j end delete D j 22
23 Profile HMMs Gapped alignment of X against a profile M: assume e I j(a) = q a (backgroud prob.) gap of length k contributes to log-odds: log(a Mj,Ij ) + log(a Ij,Mj+1 ) + (k-1) log(a Ij,Ij ) { gap open } (gap extension) No log odds contribution from the emission insert I match states begin j M j end delete D j 23
24 Profile HMM Transition Probabilities Mi Mi+ 1 Mi Di+ 1 Mi Ii Emission probabilities M i a I i a Ii Mi+ 1 Ii Ii Ii Di+ 1 Di Di+ 1 Di Mi+ 1 Di Ii 24
25 Example Suppose we are given the aligned sequences **---* AG---C A-AT-C AG-AA- --AAAC AG---C Suppose also that the match positions are marked
26 Calculating A, E count transitions and emissions: transitions M-M M-D M-I I-M I-D I-I D-M D-D D-I **---* AG---C A-AT-C AG-AA- --AAAC AG---C A C G T A C T G emissions
27 Calculating A, E count transitions and emissions: transitions M-M M-D M-I I-M I-D I-I D-M D-D D-I **---* AG---C A-AT-C AG-AA- --AAAC AG---C emissions A C G T A C T G
28 Estimating Maximum Likelihood probabilities using fractions emissions A C G T A C T G A C G T A C T G
29 Estimating ML probabilities (contd) transitions M-M M-D M-I I-M I-D I-I D-M D-D D-I M-M M-D M-I I-M I-D I-I D-M D-D D-I
30 HMM from multiple alignment Shade: insert 30
31 .. and multiple alignment from a given HMM Align each sequence to the profile separately Right: a new sequence realigend with the model Inserts are unaligned. 31
32 Searching with Profile HMMs Compute: log P( X Model) P( X random) V jm (i): log odds of best path matching x 1,,x i to submodel up to level j, ending with x i emitted by state M j V ji (i) : same, ending with x i emitted by state I j V jd (i) : score for best path ending in D j after x i has been emitted (and x i+1 has not been emitted yet) V jm (i) = log [e Mj (x i ) / q x i] + max { V j-1m (i-1)+log(a Mj-1,Mj ), V j-1i (i-1)+log(a Ij-1,Mj ), V j-1d (i-1)+log(a Dj-1,Mj ) } V ji (i) = log [e Ij (x i ) / q x i] + max { V jm (i-1)+log(a Mj-1,Ij ), V ji (i-1)+log(a Ij,Ij ), V jd (i-1)+log(a Dj,Ij ) } V jd (i) = max {V j-1m (i)+log(a Mj-1,Dj ), V j-1i (i)+log(a Ij-1,Dj ), V j-1d (i)+log(a Dj-1,Dj ) } Π i q x i 32
33 Training a profile HMM from unaligned sequences choose length of the profile HMM, initialize parameters train the model using Baum-Welch obtain MA with the resulting profile as before. (Formulas inlc. forward/backward in handout) 33
34 An Illustrative Study HMMs in Computational Biology, Krogh, Brown, Mian, Sjoalnder, Haussler 93 Globin experiment: Heme-containing proteins, involved in the storage and transport of oxygen 625 globins from Swissprot, ave. legnth 145AA Training set: 400 sequences Built model and ran it against test globins and all Swissprot proteins (25K). 34
35 Representative globins Ron Shamir, CG 08 Alignment from Bashford et al 87 A-H: alpha helices 35
36 Realignment by trained HMM Ron Shamir, CG 08 36
37 Part of the final globin model 37
38 Score vs sequence length Negative LL. Length dependent using log odds is preferable For all Swissprot seqs of < 300AA NLL=-log(P(seq model)) 38
39 Z-score distribution Z-score: (S-E(S)) / sd(s) On ~25K proteins, cutoff 5 misses 2/628 globins with no fp 39
40 Discovering subfamilies 10 component HMM. Training set: 628 globins. Classified each sequence using the model. Generated 7 nonempty clusters, with ~560 falling into 4 biologically meaningful clusters. 40
41 Local alignment in HMM Add flanking states for regions of unaligned seq. Durbin et al. Ch
42 42
Hidden Markov Models. Ron Shamir, CG 08
 Hidden Markov Models 1 Dr Richard Durbin is a graduate in mathematics from Cambridge University and one of the founder members of the Sanger Institute. He has also held carried out research at the Laboratory
Hidden Markov Models 1 Dr Richard Durbin is a graduate in mathematics from Cambridge University and one of the founder members of the Sanger Institute. He has also held carried out research at the Laboratory
Hidden Markov Models
 Hidden Markov Models Slides revised and adapted to Bioinformática 55 Engª Biomédica/IST 2005 Ana Teresa Freitas Forward Algorithm For Markov chains we calculate the probability of a sequence, P(x) How
Hidden Markov Models Slides revised and adapted to Bioinformática 55 Engª Biomédica/IST 2005 Ana Teresa Freitas Forward Algorithm For Markov chains we calculate the probability of a sequence, P(x) How
CSCE 471/871 Lecture 3: Markov Chains and
 and and 1 / 26 sscott@cse.unl.edu 2 / 26 Outline and chains models (s) Formal definition Finding most probable state path (Viterbi algorithm) Forward and backward algorithms State sequence known State
and and 1 / 26 sscott@cse.unl.edu 2 / 26 Outline and chains models (s) Formal definition Finding most probable state path (Viterbi algorithm) Forward and backward algorithms State sequence known State
Stephen Scott.
 1 / 27 sscott@cse.unl.edu 2 / 27 Useful for modeling/making predictions on sequential data E.g., biological sequences, text, series of sounds/spoken words Will return to graphical models that are generative
1 / 27 sscott@cse.unl.edu 2 / 27 Useful for modeling/making predictions on sequential data E.g., biological sequences, text, series of sounds/spoken words Will return to graphical models that are generative
Hidden Markov Models
 Hidden Markov Models Outline 1. CG-Islands 2. The Fair Bet Casino 3. Hidden Markov Model 4. Decoding Algorithm 5. Forward-Backward Algorithm 6. Profile HMMs 7. HMM Parameter Estimation 8. Viterbi Training
Hidden Markov Models Outline 1. CG-Islands 2. The Fair Bet Casino 3. Hidden Markov Model 4. Decoding Algorithm 5. Forward-Backward Algorithm 6. Profile HMMs 7. HMM Parameter Estimation 8. Viterbi Training
An Introduction to Bioinformatics Algorithms Hidden Markov Models
 Hidden Markov Models Outline 1. CG-Islands 2. The Fair Bet Casino 3. Hidden Markov Model 4. Decoding Algorithm 5. Forward-Backward Algorithm 6. Profile HMMs 7. HMM Parameter Estimation 8. Viterbi Training
Hidden Markov Models Outline 1. CG-Islands 2. The Fair Bet Casino 3. Hidden Markov Model 4. Decoding Algorithm 5. Forward-Backward Algorithm 6. Profile HMMs 7. HMM Parameter Estimation 8. Viterbi Training
Hidden Markov Models
 Hidden Markov Models Outline CG-islands The Fair Bet Casino Hidden Markov Model Decoding Algorithm Forward-Backward Algorithm Profile HMMs HMM Parameter Estimation Viterbi training Baum-Welch algorithm
Hidden Markov Models Outline CG-islands The Fair Bet Casino Hidden Markov Model Decoding Algorithm Forward-Backward Algorithm Profile HMMs HMM Parameter Estimation Viterbi training Baum-Welch algorithm
Example: The Dishonest Casino. Hidden Markov Models. Question # 1 Evaluation. The dishonest casino model. Question # 3 Learning. Question # 2 Decoding
 Example: The Dishonest Casino Hidden Markov Models Durbin and Eddy, chapter 3 Game:. You bet $. You roll 3. Casino player rolls 4. Highest number wins $ The casino has two dice: Fair die P() = P() = P(3)
Example: The Dishonest Casino Hidden Markov Models Durbin and Eddy, chapter 3 Game:. You bet $. You roll 3. Casino player rolls 4. Highest number wins $ The casino has two dice: Fair die P() = P() = P(3)
Stephen Scott.
 1 / 21 sscott@cse.unl.edu 2 / 21 Introduction Designed to model (profile) a multiple alignment of a protein family (e.g., Fig. 5.1) Gives a probabilistic model of the proteins in the family Useful for
1 / 21 sscott@cse.unl.edu 2 / 21 Introduction Designed to model (profile) a multiple alignment of a protein family (e.g., Fig. 5.1) Gives a probabilistic model of the proteins in the family Useful for
HIDDEN MARKOV MODELS
 HIDDEN MARKOV MODELS Outline CG-islands The Fair Bet Casino Hidden Markov Model Decoding Algorithm Forward-Backward Algorithm Profile HMMs HMM Parameter Estimation Viterbi training Baum-Welch algorithm
HIDDEN MARKOV MODELS Outline CG-islands The Fair Bet Casino Hidden Markov Model Decoding Algorithm Forward-Backward Algorithm Profile HMMs HMM Parameter Estimation Viterbi training Baum-Welch algorithm
Hidden Markov Models. x 1 x 2 x 3 x K
 Hidden Markov Models 1 1 1 1 2 2 2 2 K K K K x 1 x 2 x 3 x K Viterbi, Forward, Backward VITERBI FORWARD BACKWARD Initialization: V 0 (0) = 1 V k (0) = 0, for all k > 0 Initialization: f 0 (0) = 1 f k (0)
Hidden Markov Models 1 1 1 1 2 2 2 2 K K K K x 1 x 2 x 3 x K Viterbi, Forward, Backward VITERBI FORWARD BACKWARD Initialization: V 0 (0) = 1 V k (0) = 0, for all k > 0 Initialization: f 0 (0) = 1 f k (0)
CISC 889 Bioinformatics (Spring 2004) Hidden Markov Models (II)
 CISC 889 Bioinformatics (Spring 24) Hidden Markov Models (II) a. Likelihood: forward algorithm b. Decoding: Viterbi algorithm c. Model building: Baum-Welch algorithm Viterbi training Hidden Markov models
CISC 889 Bioinformatics (Spring 24) Hidden Markov Models (II) a. Likelihood: forward algorithm b. Decoding: Viterbi algorithm c. Model building: Baum-Welch algorithm Viterbi training Hidden Markov models
Hidden Markov Models. Ivan Gesteira Costa Filho IZKF Research Group Bioinformatics RWTH Aachen Adapted from:
 Hidden Markov Models Ivan Gesteira Costa Filho IZKF Research Group Bioinformatics RWTH Aachen Adapted from: www.ioalgorithms.info Outline CG-islands The Fair Bet Casino Hidden Markov Model Decoding Algorithm
Hidden Markov Models Ivan Gesteira Costa Filho IZKF Research Group Bioinformatics RWTH Aachen Adapted from: www.ioalgorithms.info Outline CG-islands The Fair Bet Casino Hidden Markov Model Decoding Algorithm
CSCE 478/878 Lecture 9: Hidden. Markov. Models. Stephen Scott. Introduction. Outline. Markov. Chains. Hidden Markov Models. CSCE 478/878 Lecture 9:
 Useful for modeling/making predictions on sequential data E.g., biological sequences, text, series of sounds/spoken words Will return to graphical models that are generative sscott@cse.unl.edu 1 / 27 2
Useful for modeling/making predictions on sequential data E.g., biological sequences, text, series of sounds/spoken words Will return to graphical models that are generative sscott@cse.unl.edu 1 / 27 2
Lecture 5: December 13, 2001
 Algorithms for Molecular Biology Fall Semester, 2001 Lecture 5: December 13, 2001 Lecturer: Ron Shamir Scribe: Roi Yehoshua and Oren Danewitz 1 5.1 Hidden Markov Models 5.1.1 Preface: CpG islands CpG is
Algorithms for Molecular Biology Fall Semester, 2001 Lecture 5: December 13, 2001 Lecturer: Ron Shamir Scribe: Roi Yehoshua and Oren Danewitz 1 5.1 Hidden Markov Models 5.1.1 Preface: CpG islands CpG is
Hidden Markov Models. x 1 x 2 x 3 x K
 Hidden Markov Models 1 1 1 1 2 2 2 2 K K K K x 1 x 2 x 3 x K HiSeq X & NextSeq Viterbi, Forward, Backward VITERBI FORWARD BACKWARD Initialization: V 0 (0) = 1 V k (0) = 0, for all k > 0 Initialization:
Hidden Markov Models 1 1 1 1 2 2 2 2 K K K K x 1 x 2 x 3 x K HiSeq X & NextSeq Viterbi, Forward, Backward VITERBI FORWARD BACKWARD Initialization: V 0 (0) = 1 V k (0) = 0, for all k > 0 Initialization:
Hidden Markov Models. based on chapters from the book Durbin, Eddy, Krogh and Mitchison Biological Sequence Analysis via Shamir s lecture notes
 Hidden Markov Models based on chapters from the book Durbin, Eddy, Krogh and Mitchison Biological Sequence Analysis via Shamir s lecture notes music recognition deal with variations in - actual sound -
Hidden Markov Models based on chapters from the book Durbin, Eddy, Krogh and Mitchison Biological Sequence Analysis via Shamir s lecture notes music recognition deal with variations in - actual sound -
Hidden Markov Models. Three classic HMM problems
 An Introduction to Bioinformatics Algorithms www.bioalgorithms.info Hidden Markov Models Slides revised and adapted to Computational Biology IST 2015/2016 Ana Teresa Freitas Three classic HMM problems
An Introduction to Bioinformatics Algorithms www.bioalgorithms.info Hidden Markov Models Slides revised and adapted to Computational Biology IST 2015/2016 Ana Teresa Freitas Three classic HMM problems
Multiple Sequence Alignment using Profile HMM
 Multiple Sequence Alignment using Profile HMM. based on Chapter 5 and Section 6.5 from Biological Sequence Analysis by R. Durbin et al., 1998 Acknowledgements: M.Sc. students Beatrice Miron, Oana Răţoi,
Multiple Sequence Alignment using Profile HMM. based on Chapter 5 and Section 6.5 from Biological Sequence Analysis by R. Durbin et al., 1998 Acknowledgements: M.Sc. students Beatrice Miron, Oana Răţoi,
Hidden Markov Models for biological sequence analysis
 Hidden Markov Models for biological sequence analysis Master in Bioinformatics UPF 2017-2018 http://comprna.upf.edu/courses/master_agb/ Eduardo Eyras Computational Genomics Pompeu Fabra University - ICREA
Hidden Markov Models for biological sequence analysis Master in Bioinformatics UPF 2017-2018 http://comprna.upf.edu/courses/master_agb/ Eduardo Eyras Computational Genomics Pompeu Fabra University - ICREA
Page 1. References. Hidden Markov models and multiple sequence alignment. Markov chains. Probability review. Example. Markovian sequence
 Page Hidden Markov models and multiple sequence alignment Russ B Altman BMI 4 CS 74 Some slides borrowed from Scott C Schmidler (BMI graduate student) References Bioinformatics Classic: Krogh et al (994)
Page Hidden Markov models and multiple sequence alignment Russ B Altman BMI 4 CS 74 Some slides borrowed from Scott C Schmidler (BMI graduate student) References Bioinformatics Classic: Krogh et al (994)
An Introduction to Bioinformatics Algorithms Hidden Markov Models
 Hidden Markov Models Hidden Markov Models Outline CG-islands The Fair Bet Casino Hidden Markov Model Decoding Algorithm Forward-Backward Algorithm Profile HMMs HMM Parameter Estimation Viterbi training
Hidden Markov Models Hidden Markov Models Outline CG-islands The Fair Bet Casino Hidden Markov Model Decoding Algorithm Forward-Backward Algorithm Profile HMMs HMM Parameter Estimation Viterbi training
Hidden Markov Models. music recognition. deal with variations in - pitch - timing - timbre 2
 Hidden Markov Models based on chapters from the book Durbin, Eddy, Krogh and Mitchison Biological Sequence Analysis Shamir s lecture notes and Rabiner s tutorial on HMM 1 music recognition deal with variations
Hidden Markov Models based on chapters from the book Durbin, Eddy, Krogh and Mitchison Biological Sequence Analysis Shamir s lecture notes and Rabiner s tutorial on HMM 1 music recognition deal with variations
Hidden Markov Models (I)
 GLOBEX Bioinformatics (Summer 2015) Hidden Markov Models (I) a. The model b. The decoding: Viterbi algorithm Hidden Markov models A Markov chain of states At each state, there are a set of possible observables
GLOBEX Bioinformatics (Summer 2015) Hidden Markov Models (I) a. The model b. The decoding: Viterbi algorithm Hidden Markov models A Markov chain of states At each state, there are a set of possible observables
Markov Chains and Hidden Markov Models. COMP 571 Luay Nakhleh, Rice University
 Markov Chains and Hidden Markov Models COMP 571 Luay Nakhleh, Rice University Markov Chains and Hidden Markov Models Modeling the statistical properties of biological sequences and distinguishing regions
Markov Chains and Hidden Markov Models COMP 571 Luay Nakhleh, Rice University Markov Chains and Hidden Markov Models Modeling the statistical properties of biological sequences and distinguishing regions
Hidden Markov Models for biological sequence analysis I
 Hidden Markov Models for biological sequence analysis I Master in Bioinformatics UPF 2014-2015 Eduardo Eyras Computational Genomics Pompeu Fabra University - ICREA Barcelona, Spain Example: CpG Islands
Hidden Markov Models for biological sequence analysis I Master in Bioinformatics UPF 2014-2015 Eduardo Eyras Computational Genomics Pompeu Fabra University - ICREA Barcelona, Spain Example: CpG Islands
Pairwise alignment using HMMs
 Pairwise alignment using HMMs The states of an HMM fulfill the Markov property: probability of transition depends only on the last state. CpG islands and casino example: HMMs emit sequence of symbols (nucleotides
Pairwise alignment using HMMs The states of an HMM fulfill the Markov property: probability of transition depends only on the last state. CpG islands and casino example: HMMs emit sequence of symbols (nucleotides
Lecture 9. Intro to Hidden Markov Models (finish up)
 Lecture 9 Intro to Hidden Markov Models (finish up) Review Structure Number of states Q 1.. Q N M output symbols Parameters: Transition probability matrix a ij Emission probabilities b i (a), which is
Lecture 9 Intro to Hidden Markov Models (finish up) Review Structure Number of states Q 1.. Q N M output symbols Parameters: Transition probability matrix a ij Emission probabilities b i (a), which is
Hidden Markov Models
 Hidden Markov Models Slides revised and adapted to Bioinformática 55 Engª Biomédica/IST 2005 Ana Teresa Freitas CG-Islands Given 4 nucleotides: probability of occurrence is ~ 1/4. Thus, probability of
Hidden Markov Models Slides revised and adapted to Bioinformática 55 Engª Biomédica/IST 2005 Ana Teresa Freitas CG-Islands Given 4 nucleotides: probability of occurrence is ~ 1/4. Thus, probability of
6.047/6.878/HST.507 Computational Biology: Genomes, Networks, Evolution. Lecture 05. Hidden Markov Models Part II
 6.047/6.878/HST.507 Computational Biology: Genomes, Networks, Evolution Lecture 05 Hidden Markov Models Part II 1 2 Module 1: Aligning and modeling genomes Module 1: Computational foundations Dynamic programming:
6.047/6.878/HST.507 Computational Biology: Genomes, Networks, Evolution Lecture 05 Hidden Markov Models Part II 1 2 Module 1: Aligning and modeling genomes Module 1: Computational foundations Dynamic programming:
11.3 Decoding Algorithm
 11.3 Decoding Algorithm 393 For convenience, we have introduced π 0 and π n+1 as the fictitious initial and terminal states begin and end. This model defines the probability P(x π) for a given sequence
11.3 Decoding Algorithm 393 For convenience, we have introduced π 0 and π n+1 as the fictitious initial and terminal states begin and end. This model defines the probability P(x π) for a given sequence
Plan for today. ! Part 1: (Hidden) Markov models. ! Part 2: String matching and read mapping
 Plan for today! Part 1: (Hidden) Markov models! Part 2: String matching and read mapping! 2.1 Exact algorithms! 2.2 Heuristic methods for approximate search (Hidden) Markov models Why consider probabilistics
Plan for today! Part 1: (Hidden) Markov models! Part 2: String matching and read mapping! 2.1 Exact algorithms! 2.2 Heuristic methods for approximate search (Hidden) Markov models Why consider probabilistics
Hidden Markov Models. Hosein Mohimani GHC7717
 Hidden Markov Models Hosein Mohimani GHC7717 hoseinm@andrew.cmu.edu Fair et Casino Problem Dealer flips a coin and player bets on outcome Dealer use either a fair coin (head and tail equally likely) or
Hidden Markov Models Hosein Mohimani GHC7717 hoseinm@andrew.cmu.edu Fair et Casino Problem Dealer flips a coin and player bets on outcome Dealer use either a fair coin (head and tail equally likely) or
CS711008Z Algorithm Design and Analysis
 .. Lecture 6. Hidden Markov model and Viterbi s decoding algorithm Institute of Computing Technology Chinese Academy of Sciences, Beijing, China . Outline The occasionally dishonest casino: an example
.. Lecture 6. Hidden Markov model and Viterbi s decoding algorithm Institute of Computing Technology Chinese Academy of Sciences, Beijing, China . Outline The occasionally dishonest casino: an example
CAP 5510: Introduction to Bioinformatics CGS 5166: Bioinformatics Tools. Giri Narasimhan
 CAP 5510: Introduction to Bioinformatics CGS 5166: Bioinformatics Tools Giri Narasimhan ECS 254; Phone: x3748 giri@cis.fiu.edu www.cis.fiu.edu/~giri/teach/bioinfs15.html Describing & Modeling Patterns
CAP 5510: Introduction to Bioinformatics CGS 5166: Bioinformatics Tools Giri Narasimhan ECS 254; Phone: x3748 giri@cis.fiu.edu www.cis.fiu.edu/~giri/teach/bioinfs15.html Describing & Modeling Patterns
6 Markov Chains and Hidden Markov Models
 6 Markov Chains and Hidden Markov Models (This chapter 1 is primarily based on Durbin et al., chapter 3, [DEKM98] and the overview article by Rabiner [Rab89] on HMMs.) Why probabilistic models? In problems
6 Markov Chains and Hidden Markov Models (This chapter 1 is primarily based on Durbin et al., chapter 3, [DEKM98] and the overview article by Rabiner [Rab89] on HMMs.) Why probabilistic models? In problems
Markov Chains and Hidden Markov Models. = stochastic, generative models
 Markov Chains and Hidden Markov Models = stochastic, generative models (Drawing heavily from Durbin et al., Biological Sequence Analysis) BCH339N Systems Biology / Bioinformatics Spring 2016 Edward Marcotte,
Markov Chains and Hidden Markov Models = stochastic, generative models (Drawing heavily from Durbin et al., Biological Sequence Analysis) BCH339N Systems Biology / Bioinformatics Spring 2016 Edward Marcotte,
Lecture 4: Hidden Markov Models: An Introduction to Dynamic Decision Making. November 11, 2010
 Hidden Lecture 4: Hidden : An Introduction to Dynamic Decision Making November 11, 2010 Special Meeting 1/26 Markov Model Hidden When a dynamical system is probabilistic it may be determined by the transition
Hidden Lecture 4: Hidden : An Introduction to Dynamic Decision Making November 11, 2010 Special Meeting 1/26 Markov Model Hidden When a dynamical system is probabilistic it may be determined by the transition
HMMs and biological sequence analysis
 HMMs and biological sequence analysis Hidden Markov Model A Markov chain is a sequence of random variables X 1, X 2, X 3,... That has the property that the value of the current state depends only on the
HMMs and biological sequence analysis Hidden Markov Model A Markov chain is a sequence of random variables X 1, X 2, X 3,... That has the property that the value of the current state depends only on the
EECS730: Introduction to Bioinformatics
 EECS730: Introduction to Bioinformatics Lecture 07: profile Hidden Markov Model http://bibiserv.techfak.uni-bielefeld.de/sadr2/databasesearch/hmmer/profilehmm.gif Slides adapted from Dr. Shaojie Zhang
EECS730: Introduction to Bioinformatics Lecture 07: profile Hidden Markov Model http://bibiserv.techfak.uni-bielefeld.de/sadr2/databasesearch/hmmer/profilehmm.gif Slides adapted from Dr. Shaojie Zhang
Computational Genomics and Molecular Biology, Fall
 Computational Genomics and Molecular Biology, Fall 2014 1 HMM Lecture Notes Dannie Durand and Rose Hoberman November 6th Introduction In the last few lectures, we have focused on three problems related
Computational Genomics and Molecular Biology, Fall 2014 1 HMM Lecture Notes Dannie Durand and Rose Hoberman November 6th Introduction In the last few lectures, we have focused on three problems related
ROBI POLIKAR. ECE 402/504 Lecture Hidden Markov Models IGNAL PROCESSING & PATTERN RECOGNITION ROWAN UNIVERSITY
 BIOINFORMATICS Lecture 11-12 Hidden Markov Models ROBI POLIKAR 2011, All Rights Reserved, Robi Polikar. IGNAL PROCESSING & PATTERN RECOGNITION LABORATORY @ ROWAN UNIVERSITY These lecture notes are prepared
BIOINFORMATICS Lecture 11-12 Hidden Markov Models ROBI POLIKAR 2011, All Rights Reserved, Robi Polikar. IGNAL PROCESSING & PATTERN RECOGNITION LABORATORY @ ROWAN UNIVERSITY These lecture notes are prepared
Introduction to Hidden Markov Models for Gene Prediction ECE-S690
 Introduction to Hidden Markov Models for Gene Prediction ECE-S690 Outline Markov Models The Hidden Part How can we use this for gene prediction? Learning Models Want to recognize patterns (e.g. sequence
Introduction to Hidden Markov Models for Gene Prediction ECE-S690 Outline Markov Models The Hidden Part How can we use this for gene prediction? Learning Models Want to recognize patterns (e.g. sequence
Hidden Markov Models. By Parisa Abedi. Slides courtesy: Eric Xing
 Hidden Markov Models By Parisa Abedi Slides courtesy: Eric Xing i.i.d to sequential data So far we assumed independent, identically distributed data Sequential (non i.i.d.) data Time-series data E.g. Speech
Hidden Markov Models By Parisa Abedi Slides courtesy: Eric Xing i.i.d to sequential data So far we assumed independent, identically distributed data Sequential (non i.i.d.) data Time-series data E.g. Speech
Hidden Markov Models 1
 Hidden Markov Models Dinucleotide Frequency Consider all 2-mers in a sequence {AA,AC,AG,AT,CA,CC,CG,CT,GA,GC,GG,GT,TA,TC,TG,TT} Given 4 nucleotides: each with a probability of occurrence of. 4 Thus, one
Hidden Markov Models Dinucleotide Frequency Consider all 2-mers in a sequence {AA,AC,AG,AT,CA,CC,CG,CT,GA,GC,GG,GT,TA,TC,TG,TT} Given 4 nucleotides: each with a probability of occurrence of. 4 Thus, one
Hidden Markov Models
 Andrea Passerini passerini@disi.unitn.it Statistical relational learning The aim Modeling temporal sequences Model signals which vary over time (e.g. speech) Two alternatives: deterministic models directly
Andrea Passerini passerini@disi.unitn.it Statistical relational learning The aim Modeling temporal sequences Model signals which vary over time (e.g. speech) Two alternatives: deterministic models directly
O 3 O 4 O 5. q 3. q 4. Transition
 Hidden Markov Models Hidden Markov models (HMM) were developed in the early part of the 1970 s and at that time mostly applied in the area of computerized speech recognition. They are first described in
Hidden Markov Models Hidden Markov models (HMM) were developed in the early part of the 1970 s and at that time mostly applied in the area of computerized speech recognition. They are first described in
HMM: Parameter Estimation
 I529: Machine Learning in Bioinformatics (Spring 2017) HMM: Parameter Estimation Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Content Review HMM: three problems
I529: Machine Learning in Bioinformatics (Spring 2017) HMM: Parameter Estimation Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Content Review HMM: three problems
Data Mining in Bioinformatics HMM
 Data Mining in Bioinformatics HMM Microarray Problem: Major Objective n Major Objective: Discover a comprehensive theory of life s organization at the molecular level 2 1 Data Mining in Bioinformatics
Data Mining in Bioinformatics HMM Microarray Problem: Major Objective n Major Objective: Discover a comprehensive theory of life s organization at the molecular level 2 1 Data Mining in Bioinformatics
Hidden Markov Models. Aarti Singh Slides courtesy: Eric Xing. Machine Learning / Nov 8, 2010
 Hidden Markov Models Aarti Singh Slides courtesy: Eric Xing Machine Learning 10-701/15-781 Nov 8, 2010 i.i.d to sequential data So far we assumed independent, identically distributed data Sequential data
Hidden Markov Models Aarti Singh Slides courtesy: Eric Xing Machine Learning 10-701/15-781 Nov 8, 2010 i.i.d to sequential data So far we assumed independent, identically distributed data Sequential data
HMM applications. Applications of HMMs. Gene finding with HMMs. Using the gene finder
 HMM applications Applications of HMMs Gene finding Pairwise alignment (pair HMMs) Characterizing protein families (profile HMMs) Predicting membrane proteins, and membrane protein topology Gene finding
HMM applications Applications of HMMs Gene finding Pairwise alignment (pair HMMs) Characterizing protein families (profile HMMs) Predicting membrane proteins, and membrane protein topology Gene finding
Introduction to Machine Learning CMU-10701
 Introduction to Machine Learning CMU-10701 Hidden Markov Models Barnabás Póczos & Aarti Singh Slides courtesy: Eric Xing i.i.d to sequential data So far we assumed independent, identically distributed
Introduction to Machine Learning CMU-10701 Hidden Markov Models Barnabás Póczos & Aarti Singh Slides courtesy: Eric Xing i.i.d to sequential data So far we assumed independent, identically distributed
Today s Lecture: HMMs
 Today s Lecture: HMMs Definitions Examples Probability calculations WDAG Dynamic programming algorithms: Forward Viterbi Parameter estimation Viterbi training 1 Hidden Markov Models Probability models
Today s Lecture: HMMs Definitions Examples Probability calculations WDAG Dynamic programming algorithms: Forward Viterbi Parameter estimation Viterbi training 1 Hidden Markov Models Probability models
order is number of previous outputs
 Markov Models Lecture : Markov and Hidden Markov Models PSfrag Use past replacements as state. Next output depends on previous output(s): y t = f[y t, y t,...] order is number of previous outputs y t y
Markov Models Lecture : Markov and Hidden Markov Models PSfrag Use past replacements as state. Next output depends on previous output(s): y t = f[y t, y t,...] order is number of previous outputs y t y
Markov chains and Hidden Markov Models
 Discrete Math for Bioinformatics WS 10/11:, b A. Bockmar/K. Reinert, 7. November 2011, 10:24 2001 Markov chains and Hidden Markov Models We will discuss: Hidden Markov Models (HMMs) Algorithms: Viterbi,
Discrete Math for Bioinformatics WS 10/11:, b A. Bockmar/K. Reinert, 7. November 2011, 10:24 2001 Markov chains and Hidden Markov Models We will discuss: Hidden Markov Models (HMMs) Algorithms: Viterbi,
STA 414/2104: Machine Learning
 STA 414/2104: Machine Learning Russ Salakhutdinov Department of Computer Science! Department of Statistics! rsalakhu@cs.toronto.edu! http://www.cs.toronto.edu/~rsalakhu/ Lecture 9 Sequential Data So far
STA 414/2104: Machine Learning Russ Salakhutdinov Department of Computer Science! Department of Statistics! rsalakhu@cs.toronto.edu! http://www.cs.toronto.edu/~rsalakhu/ Lecture 9 Sequential Data So far
Giri Narasimhan. CAP 5510: Introduction to Bioinformatics. ECS 254; Phone: x3748
 CAP 5510: Introduction to Bioinformatics Giri Narasimhan ECS 254; Phone: x3748 giri@cis.fiu.edu www.cis.fiu.edu/~giri/teach/bioinfs07.html 2/14/07 CAP5510 1 CpG Islands Regions in DNA sequences with increased
CAP 5510: Introduction to Bioinformatics Giri Narasimhan ECS 254; Phone: x3748 giri@cis.fiu.edu www.cis.fiu.edu/~giri/teach/bioinfs07.html 2/14/07 CAP5510 1 CpG Islands Regions in DNA sequences with increased
Computational Genomics and Molecular Biology, Fall
 Computational Genomics and Molecular Biology, Fall 2011 1 HMM Lecture Notes Dannie Durand and Rose Hoberman October 11th 1 Hidden Markov Models In the last few lectures, we have focussed on three problems
Computational Genomics and Molecular Biology, Fall 2011 1 HMM Lecture Notes Dannie Durand and Rose Hoberman October 11th 1 Hidden Markov Models In the last few lectures, we have focussed on three problems
Lecture 7 Sequence analysis. Hidden Markov Models
 Lecture 7 Sequence analysis. Hidden Markov Models Nicolas Lartillot may 2012 Nicolas Lartillot (Universite de Montréal) BIN6009 may 2012 1 / 60 1 Motivation 2 Examples of Hidden Markov models 3 Hidden
Lecture 7 Sequence analysis. Hidden Markov Models Nicolas Lartillot may 2012 Nicolas Lartillot (Universite de Montréal) BIN6009 may 2012 1 / 60 1 Motivation 2 Examples of Hidden Markov models 3 Hidden
Pair Hidden Markov Models
 Pair Hidden Markov Models Scribe: Rishi Bedi Lecturer: Serafim Batzoglou January 29, 2015 1 Recap of HMMs alphabet: Σ = {b 1,...b M } set of states: Q = {1,..., K} transition probabilities: A = [a ij ]
Pair Hidden Markov Models Scribe: Rishi Bedi Lecturer: Serafim Batzoglou January 29, 2015 1 Recap of HMMs alphabet: Σ = {b 1,...b M } set of states: Q = {1,..., K} transition probabilities: A = [a ij ]
R. Durbin, S. Eddy, A. Krogh, G. Mitchison: Biological sequence analysis. Cambridge University Press, ISBN (Chapter 3)
 9 Markov chains and Hidden Markov Models We will discuss: Markov chains Hidden Markov Models (HMMs) lgorithms: Viterbi, forward, backward, posterior decoding Profile HMMs Baum-Welch algorithm This chapter
9 Markov chains and Hidden Markov Models We will discuss: Markov chains Hidden Markov Models (HMMs) lgorithms: Viterbi, forward, backward, posterior decoding Profile HMMs Baum-Welch algorithm This chapter
Advanced Data Science
 Advanced Data Science Dr. Kira Radinsky Slides Adapted from Tom M. Mitchell Agenda Topics Covered: Time series data Markov Models Hidden Markov Models Dynamic Bayes Nets Additional Reading: Bishop: Chapter
Advanced Data Science Dr. Kira Radinsky Slides Adapted from Tom M. Mitchell Agenda Topics Covered: Time series data Markov Models Hidden Markov Models Dynamic Bayes Nets Additional Reading: Bishop: Chapter
Comparative Gene Finding. BMI/CS 776 Spring 2015 Colin Dewey
 Comparative Gene Finding BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2015 Colin Dewey cdewey@biostat.wisc.edu Goals for Lecture the key concepts to understand are the following: using related genomes
Comparative Gene Finding BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2015 Colin Dewey cdewey@biostat.wisc.edu Goals for Lecture the key concepts to understand are the following: using related genomes
Grundlagen der Bioinformatik, SS 09, D. Huson, June 16, S. Durbin, S. Eddy, A. Krogh and G. Mitchison, Biological Sequence
 rundlagen der Bioinformatik, SS 09,. Huson, June 16, 2009 81 7 Markov chains and Hidden Markov Models We will discuss: Markov chains Hidden Markov Models (HMMs) Profile HMMs his chapter is based on: nalysis,
rundlagen der Bioinformatik, SS 09,. Huson, June 16, 2009 81 7 Markov chains and Hidden Markov Models We will discuss: Markov chains Hidden Markov Models (HMMs) Profile HMMs his chapter is based on: nalysis,
RNA Search and! Motif Discovery" Genome 541! Intro to Computational! Molecular Biology"
 RNA Search and! Motif Discovery" Genome 541! Intro to Computational! Molecular Biology" Day 1" Many biologically interesting roles for RNA" RNA secondary structure prediction" 3 4 Approaches to Structure
RNA Search and! Motif Discovery" Genome 541! Intro to Computational! Molecular Biology" Day 1" Many biologically interesting roles for RNA" RNA secondary structure prediction" 3 4 Approaches to Structure
Hidden Markov Models (HMMs) and Profiles
 Hidden Markov Models (HMMs) and Profiles Swiss Institute of Bioinformatics (SIB) 26-30 November 2001 Markov Chain Models A Markov Chain Model is a succession of states S i (i = 0, 1,...) connected by transitions.
Hidden Markov Models (HMMs) and Profiles Swiss Institute of Bioinformatics (SIB) 26-30 November 2001 Markov Chain Models A Markov Chain Model is a succession of states S i (i = 0, 1,...) connected by transitions.
Statistical Machine Learning Methods for Bioinformatics II. Hidden Markov Model for Biological Sequences
 Statistical Machine Learning Methods for Bioinformatics II. Hidden Markov Model for Biological Sequences Jianlin Cheng, PhD Department of Computer Science University of Missouri 2008 Free for Academic
Statistical Machine Learning Methods for Bioinformatics II. Hidden Markov Model for Biological Sequences Jianlin Cheng, PhD Department of Computer Science University of Missouri 2008 Free for Academic
Hidden Markov Models Part 2: Algorithms
 Hidden Markov Models Part 2: Algorithms CSE 6363 Machine Learning Vassilis Athitsos Computer Science and Engineering Department University of Texas at Arlington 1 Hidden Markov Model An HMM consists of:
Hidden Markov Models Part 2: Algorithms CSE 6363 Machine Learning Vassilis Athitsos Computer Science and Engineering Department University of Texas at Arlington 1 Hidden Markov Model An HMM consists of:
Hidden Markov Models and Their Applications in Biological Sequence Analysis
 Hidden Markov Models and Their Applications in Biological Sequence Analysis Byung-Jun Yoon Dept. of Electrical & Computer Engineering Texas A&M University, College Station, TX 77843-3128, USA Abstract
Hidden Markov Models and Their Applications in Biological Sequence Analysis Byung-Jun Yoon Dept. of Electrical & Computer Engineering Texas A&M University, College Station, TX 77843-3128, USA Abstract
What s an HMM? Extraction with Finite State Machines e.g. Hidden Markov Models (HMMs) Hidden Markov Models (HMMs) for Information Extraction
 Hidden Markov Models (HMMs) for Information Extraction Daniel S. Weld CSE 454 Extraction with Finite State Machines e.g. Hidden Markov Models (HMMs) standard sequence model in genomics, speech, NLP, What
Hidden Markov Models (HMMs) for Information Extraction Daniel S. Weld CSE 454 Extraction with Finite State Machines e.g. Hidden Markov Models (HMMs) standard sequence model in genomics, speech, NLP, What
Bioinformatics 1--lectures 15, 16. Markov chains Hidden Markov models Profile HMMs
 Bioinformatics 1--lectures 15, 16 Markov chains Hidden Markov models Profile HMMs target sequence database input to database search results are sequence family pseudocounts or background-weighted pseudocounts
Bioinformatics 1--lectures 15, 16 Markov chains Hidden Markov models Profile HMMs target sequence database input to database search results are sequence family pseudocounts or background-weighted pseudocounts
Statistical Machine Learning Methods for Biomedical Informatics II. Hidden Markov Model for Biological Sequences
 Statistical Machine Learning Methods for Biomedical Informatics II. Hidden Markov Model for Biological Sequences Jianlin Cheng, PhD William and Nancy Thompson Missouri Distinguished Professor Department
Statistical Machine Learning Methods for Biomedical Informatics II. Hidden Markov Model for Biological Sequences Jianlin Cheng, PhD William and Nancy Thompson Missouri Distinguished Professor Department
Hidden Markov Model. Ying Wu. Electrical Engineering and Computer Science Northwestern University Evanston, IL 60208
 Hidden Markov Model Ying Wu Electrical Engineering and Computer Science Northwestern University Evanston, IL 60208 http://www.eecs.northwestern.edu/~yingwu 1/19 Outline Example: Hidden Coin Tossing Hidden
Hidden Markov Model Ying Wu Electrical Engineering and Computer Science Northwestern University Evanston, IL 60208 http://www.eecs.northwestern.edu/~yingwu 1/19 Outline Example: Hidden Coin Tossing Hidden
STA 4273H: Statistical Machine Learning
 STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 11 Project
STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 11 Project
Lecture #5. Dependencies along the genome
 Markov Chains Lecture #5 Background Readings: Durbin et. al. Section 3., Polanski&Kimmel Section 2.8. Prepared by Shlomo Moran, based on Danny Geiger s and Nir Friedman s. Dependencies along the genome
Markov Chains Lecture #5 Background Readings: Durbin et. al. Section 3., Polanski&Kimmel Section 2.8. Prepared by Shlomo Moran, based on Danny Geiger s and Nir Friedman s. Dependencies along the genome
Lecture 8 Learning Sequence Motif Models Using Expectation Maximization (EM) Colin Dewey February 14, 2008
 Lecture 8 Learning Sequence Motif Models Using Expectation Maximization (EM) Colin Dewey February 14, 2008 1 Sequence Motifs what is a sequence motif? a sequence pattern of biological significance typically
Lecture 8 Learning Sequence Motif Models Using Expectation Maximization (EM) Colin Dewey February 14, 2008 1 Sequence Motifs what is a sequence motif? a sequence pattern of biological significance typically
Hidden Markov Models. Terminology, Representation and Basic Problems
 Hidden Markov Models Terminology, Representation and Basic Problems Data analysis? Machine learning? In bioinformatics, we analyze a lot of (sequential) data (biological sequences) to learn unknown parameters
Hidden Markov Models Terminology, Representation and Basic Problems Data analysis? Machine learning? In bioinformatics, we analyze a lot of (sequential) data (biological sequences) to learn unknown parameters
Grundlagen der Bioinformatik, SS 08, D. Huson, June 16, S. Durbin, S. Eddy, A. Krogh and G. Mitchison, Biological Sequence
 rundlagen der Bioinformatik, SS 08,. Huson, June 16, 2008 89 8 Markov chains and Hidden Markov Models We will discuss: Markov chains Hidden Markov Models (HMMs) Profile HMMs his chapter is based on: nalysis,
rundlagen der Bioinformatik, SS 08,. Huson, June 16, 2008 89 8 Markov chains and Hidden Markov Models We will discuss: Markov chains Hidden Markov Models (HMMs) Profile HMMs his chapter is based on: nalysis,
Sequence labeling. Taking collective a set of interrelated instances x 1,, x T and jointly labeling them
 HMM, MEMM and CRF 40-957 Special opics in Artificial Intelligence: Probabilistic Graphical Models Sharif University of echnology Soleymani Spring 2014 Sequence labeling aking collective a set of interrelated
HMM, MEMM and CRF 40-957 Special opics in Artificial Intelligence: Probabilistic Graphical Models Sharif University of echnology Soleymani Spring 2014 Sequence labeling aking collective a set of interrelated
Chapter 4: Hidden Markov Models
 Chapter 4: Hidden Markov Models 4.1 Introduction to HMM Prof. Yechiam Yemini (YY) Computer Science Department Columbia University Overview Markov models of sequence structures Introduction to Hidden Markov
Chapter 4: Hidden Markov Models 4.1 Introduction to HMM Prof. Yechiam Yemini (YY) Computer Science Department Columbia University Overview Markov models of sequence structures Introduction to Hidden Markov
Hidden Markov Models. x 1 x 2 x 3 x N
 Hidden Markov Models 1 1 1 1 K K K K x 1 x x 3 x N Example: The dishonest casino A casino has two dice: Fair die P(1) = P() = P(3) = P(4) = P(5) = P(6) = 1/6 Loaded die P(1) = P() = P(3) = P(4) = P(5)
Hidden Markov Models 1 1 1 1 K K K K x 1 x x 3 x N Example: The dishonest casino A casino has two dice: Fair die P(1) = P() = P(3) = P(4) = P(5) = P(6) = 1/6 Loaded die P(1) = P() = P(3) = P(4) = P(5)
Computational Biology Lecture #3: Probability and Statistics. Bud Mishra Professor of Computer Science, Mathematics, & Cell Biology Sept
 Computational Biology Lecture #3: Probability and Statistics Bud Mishra Professor of Computer Science, Mathematics, & Cell Biology Sept 26 2005 L2-1 Basic Probabilities L2-2 1 Random Variables L2-3 Examples
Computational Biology Lecture #3: Probability and Statistics Bud Mishra Professor of Computer Science, Mathematics, & Cell Biology Sept 26 2005 L2-1 Basic Probabilities L2-2 1 Random Variables L2-3 Examples
CS 136a Lecture 7 Speech Recognition Architecture: Training models with the Forward backward algorithm
 + September13, 2016 Professor Meteer CS 136a Lecture 7 Speech Recognition Architecture: Training models with the Forward backward algorithm Thanks to Dan Jurafsky for these slides + ASR components n Feature
+ September13, 2016 Professor Meteer CS 136a Lecture 7 Speech Recognition Architecture: Training models with the Forward backward algorithm Thanks to Dan Jurafsky for these slides + ASR components n Feature
Hidden Markov Models (HMMs) November 14, 2017
 Hidden Markov Models (HMMs) November 14, 2017 inferring a hidden truth 1) You hear a static-filled radio transmission. how can you determine what did the sender intended to say? 2) You know that genes
Hidden Markov Models (HMMs) November 14, 2017 inferring a hidden truth 1) You hear a static-filled radio transmission. how can you determine what did the sender intended to say? 2) You know that genes
Lecture 12: Algorithms for HMMs
 Lecture 12: Algorithms for HMMs Nathan Schneider (some slides from Sharon Goldwater; thanks to Jonathan May for bug fixes) ENLP 17 October 2016 updated 9 September 2017 Recap: tagging POS tagging is a
Lecture 12: Algorithms for HMMs Nathan Schneider (some slides from Sharon Goldwater; thanks to Jonathan May for bug fixes) ENLP 17 October 2016 updated 9 September 2017 Recap: tagging POS tagging is a
Markov Models & DNA Sequence Evolution
 7.91 / 7.36 / BE.490 Lecture #5 Mar. 9, 2004 Markov Models & DNA Sequence Evolution Chris Burge Review of Markov & HMM Models for DNA Markov Models for splice sites Hidden Markov Models - looking under
7.91 / 7.36 / BE.490 Lecture #5 Mar. 9, 2004 Markov Models & DNA Sequence Evolution Chris Burge Review of Markov & HMM Models for DNA Markov Models for splice sites Hidden Markov Models - looking under
Hidden Markov Models The three basic HMM problems (note: change in notation) Mitch Marcus CSE 391
 Hidden Markov Models The three basic HMM problems (note: change in notation) Mitch Marcus CSE 391 Parameters of an HMM States: A set of states S=s 1, s n Transition probabilities: A= a 1,1, a 1,2,, a n,n
Hidden Markov Models The three basic HMM problems (note: change in notation) Mitch Marcus CSE 391 Parameters of an HMM States: A set of states S=s 1, s n Transition probabilities: A= a 1,1, a 1,2,, a n,n
Biology 644: Bioinformatics
 A stochastic (probabilistic) model that assumes the Markov property Markov property is satisfied when the conditional probability distribution of future states of the process (conditional on both past
A stochastic (probabilistic) model that assumes the Markov property Markov property is satisfied when the conditional probability distribution of future states of the process (conditional on both past
Bioinformatics Introduction to Hidden Markov Models Hidden Markov Models and Multiple Sequence Alignment
 Bioinformatics Introduction to Hidden Markov Models Hidden Markov Models and Multiple Sequence Alignment Slides borrowed from Scott C. Schmidler (MIS graduated student) Outline! Probability Review! Markov
Bioinformatics Introduction to Hidden Markov Models Hidden Markov Models and Multiple Sequence Alignment Slides borrowed from Scott C. Schmidler (MIS graduated student) Outline! Probability Review! Markov
Hidden Markov Models and Applica2ons. Spring 2017 February 21,23, 2017
 Hidden Markov Models and Applica2ons Spring 2017 February 21,23, 2017 Gene finding in prokaryotes Reading frames A protein is coded by groups of three nucleo2des (codons): ACGTACGTACGTACGT ACG-TAC-GTA-CGT-ACG-T
Hidden Markov Models and Applica2ons Spring 2017 February 21,23, 2017 Gene finding in prokaryotes Reading frames A protein is coded by groups of three nucleo2des (codons): ACGTACGTACGTACGT ACG-TAC-GTA-CGT-ACG-T
Hidden Markov Models and Gaussian Mixture Models
 Hidden Markov Models and Gaussian Mixture Models Hiroshi Shimodaira and Steve Renals Automatic Speech Recognition ASR Lectures 4&5 23&27 January 2014 ASR Lectures 4&5 Hidden Markov Models and Gaussian
Hidden Markov Models and Gaussian Mixture Models Hiroshi Shimodaira and Steve Renals Automatic Speech Recognition ASR Lectures 4&5 23&27 January 2014 ASR Lectures 4&5 Hidden Markov Models and Gaussian
Biological Sequences and Hidden Markov Models CPBS7711
 Biological Sequences and Hidden Markov Models CPBS7711 Sept 27, 2011 Sonia Leach, PhD Assistant Professor Center for Genes, Environment, and Health National Jewish Health sonia.leach@gmail.com Slides created
Biological Sequences and Hidden Markov Models CPBS7711 Sept 27, 2011 Sonia Leach, PhD Assistant Professor Center for Genes, Environment, and Health National Jewish Health sonia.leach@gmail.com Slides created
1/22/13. Example: CpG Island. Question 2: Finding CpG Islands
 I529: Machine Learning in Bioinformatics (Spring 203 Hidden Markov Models Yuzhen Ye School of Informatics and Computing Indiana Univerty, Bloomington Spring 203 Outline Review of Markov chain & CpG island
I529: Machine Learning in Bioinformatics (Spring 203 Hidden Markov Models Yuzhen Ye School of Informatics and Computing Indiana Univerty, Bloomington Spring 203 Outline Review of Markov chain & CpG island
Lecture 12: Algorithms for HMMs
 Lecture 12: Algorithms for HMMs Nathan Schneider (some slides from Sharon Goldwater; thanks to Jonathan May for bug fixes) ENLP 26 February 2018 Recap: tagging POS tagging is a sequence labelling task.
Lecture 12: Algorithms for HMMs Nathan Schneider (some slides from Sharon Goldwater; thanks to Jonathan May for bug fixes) ENLP 26 February 2018 Recap: tagging POS tagging is a sequence labelling task.
Basic math for biology
 Basic math for biology Lei Li Florida State University, Feb 6, 2002 The EM algorithm: setup Parametric models: {P θ }. Data: full data (Y, X); partial data Y. Missing data: X. Likelihood and maximum likelihood
Basic math for biology Lei Li Florida State University, Feb 6, 2002 The EM algorithm: setup Parametric models: {P θ }. Data: full data (Y, X); partial data Y. Missing data: X. Likelihood and maximum likelihood
6.864: Lecture 5 (September 22nd, 2005) The EM Algorithm
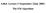 6.864: Lecture 5 (September 22nd, 2005) The EM Algorithm Overview The EM algorithm in general form The EM algorithm for hidden markov models (brute force) The EM algorithm for hidden markov models (dynamic
6.864: Lecture 5 (September 22nd, 2005) The EM Algorithm Overview The EM algorithm in general form The EM algorithm for hidden markov models (brute force) The EM algorithm for hidden markov models (dynamic
Hidden Markov models in population genetics and evolutionary biology
 Hidden Markov models in population genetics and evolutionary biology Gerton Lunter Wellcome Trust Centre for Human Genetics Oxford, UK April 29, 2013 Topics for today Markov chains Hidden Markov models
Hidden Markov models in population genetics and evolutionary biology Gerton Lunter Wellcome Trust Centre for Human Genetics Oxford, UK April 29, 2013 Topics for today Markov chains Hidden Markov models
COMS 4771 Probabilistic Reasoning via Graphical Models. Nakul Verma
 COMS 4771 Probabilistic Reasoning via Graphical Models Nakul Verma Last time Dimensionality Reduction Linear vs non-linear Dimensionality Reduction Principal Component Analysis (PCA) Non-linear methods
COMS 4771 Probabilistic Reasoning via Graphical Models Nakul Verma Last time Dimensionality Reduction Linear vs non-linear Dimensionality Reduction Principal Component Analysis (PCA) Non-linear methods
Recap: HMM. ANLP Lecture 9: Algorithms for HMMs. More general notation. Recap: HMM. Elements of HMM: Sharon Goldwater 4 Oct 2018.
 Recap: HMM ANLP Lecture 9: Algorithms for HMMs Sharon Goldwater 4 Oct 2018 Elements of HMM: Set of states (tags) Output alphabet (word types) Start state (beginning of sentence) State transition probabilities
Recap: HMM ANLP Lecture 9: Algorithms for HMMs Sharon Goldwater 4 Oct 2018 Elements of HMM: Set of states (tags) Output alphabet (word types) Start state (beginning of sentence) State transition probabilities
Gibbs Sampling Methods for Multiple Sequence Alignment
 Gibbs Sampling Methods for Multiple Sequence Alignment Scott C. Schmidler 1 Jun S. Liu 2 1 Section on Medical Informatics and 2 Department of Statistics Stanford University 11/17/99 1 Outline Statistical
Gibbs Sampling Methods for Multiple Sequence Alignment Scott C. Schmidler 1 Jun S. Liu 2 1 Section on Medical Informatics and 2 Department of Statistics Stanford University 11/17/99 1 Outline Statistical
