Quantitative trait evolution with mutations of large effect
|
|
|
- Gwenda Mathews
- 5 years ago
- Views:
Transcription
1 Quantitative trait evolution with mutations of large effect May 1, 2014
2 Quantitative traits Traits that vary continuously in populations - Mass - Height - Bristle number (approx) Adaption - Low oxygen tolerance Disease - Obesity Agriculture - Fruit yield
3 Fisher s quantitative trait model 1 locus n biallelic loci impacting trait Frequency Allele j at locus i has effect a ij Phenotype is X k = i,j a ijz ijk - z ijk is number of copies of allele j at locus i in individual k E( X ) = 2nE(p)E(a) E(Var(X )) = 2nE(p(1 p))e(a 2 ) Environmental variation induces extra variability Frequency Frequency Trait 5 loci Trait 100 loci Trait
4 Neutral genetic variance Equilibrium genetic variance - E(V A ( )) = 2NV m V m is the additive variance of new mutations - V m 2µσ 2 - µ: is trait-wide mutation rate - σ 2 : variance of mutant effect size distribution Var(V A ( )) 4Nµσ 4 (Lynch and Hill 1986)
5 Lande s Brownian motion model Approximation to full model - Don t keep track of individual loci - Assume genetic variance constant in time - Mutations have small effect sizes Can incorporate selection via Ornstein-Uhlenbeck model Extremely influential in comparative biology - c.f. Felsenstein: independent contrasts X time
6 Evidence for non-brownian evolution
7 A coalescent model Haploid population of size N Trait governed by n loci Each locus has an independent coalescent tree Each locus has mutation rate θ 2 (coalescent time units) When a mutation happens, effect Y is drawn from distribution with density p(y).
8 Example
9 Mutational effects Common assumption: many loci of very small effect - Infinitesimal model
10 Large effect mutations In some cases, large effect mutations may occur - Transcription factor binding sites - Null mutants upstream in pathways
11 Characteristic functions Characteristic function For a random variable X drawn from the probability measure µ( ), the function φ X (k) = E(e ikx ) = e ikx dµ(x) is called the characteristic function Sums of random variables If X 1,..., X n are independent random variables, and X = i X i then φ X (k) = φ Xi (k) i
12 Compound Poisson process Definition Let N(t) be a rate λ Poisson process, and (Y i, i 1) a sequence of independent and identically distributed random variables. Then N(t) Y i X (t) = i=1 is called a compound Poisson process. X(t) t
13 Characteristic function of a compound Poisson process CF of a CP process If X (t) is a rate λ Compound Poisson process, and ψ(k) is the characteristic function of the jump distribution, then the characteristic function of X is φ t (k) = e λt(ψ(k) 1)
14 One locus, sample of size 2 Condition on T 2, the coalescence time - X 1 and X 2 are independent CP(θ/2) processes run for time T 2 - Joint distribution depends on root value Consider Z = X 2 X 1 - Doesn t depend on root
15 One locus, sample of size 2 (CF) φ Z (k) = = = = = E(e ikz )e t dt E(e ik(x 2 X 1 ) )e t dt φ t (k)φ t ( k)e t dt e θ 2 t(ψ(k)+ψ( k) 2) e t dt 1 1 θ 2 (ψ(k) + ψ( k) 2)
16 Phenotype at the root The root of the tree must be specified A new individual could coalesce more anciently than the current root - For each n, there is a fixed root - Ensuring consistency could be hard
17 The root problem
18 Getting rid of the root Normalization If X = (X 0,..., X n 1 ) are the trait values in the sample, then the random vector Z = (Z 1,..., Z n 1 ) does not depend on the root value = (X 1 X 0,..., X n 1 X 0 ) Normalizing by an arbitrary individual makes n unimportant - Intuition: now every sample has to follow the path through the tree to individual 0, so the root doesn t matter
19 One locus, sample of size 3 Characteristic function for n = 3, and symmetric mutational effects φ Z1,Z 2 (k 1, k 2 ) = 1 1+θ(1 ψ(k 1 )) θ(1 ψ(k 2 )) θ(1 ψ(k 1 +k 2 )) 3 θ 2 (ψ(k 1) + ψ(k 2 ) + ψ(k 1 + k 2 ) 3) Sketch of derivation: - Condition on topology (each of 3 topologies equally likely) - Compute characteristic function for each topology while conditioning on coalescence times - Integrate over coalescence times - Take weighted sum of characteristic functions for each topology
20 Sending the number of loci off to infinity We would like some nice limit as the number of loci increases - Trivial result: uniform on R or δ(x) Need to decrease the effect size or mutation rate of each locus Three nontrivial limits - Mutation rate per locus decreases but effect sizes remain constaint - Mutation rate per locus remains constant but effect sizes decrease and effect size distribution does not have fat tails - Mutation rate per locus remains constant but effect size decreases and effect size distribution has fat tail
21 Decreasing mutation rate Correlated CPP limit As n and θ 0 such that nθ Θ, φ Z1,Z 2 (k 1, k 2 ) e Θ 2 (ψ(k 1)+ψ(k 2 )+ψ(k 1 +k 2 ) 3) which is the characteristic function of two correlated compound Poisson processes. Perhaps not very biologically relevant Will ignore for rest of talk
22 Small effect mutations Bivariate Gaussian limit As n and the second moment of the effect distrbution, τ 2, 0 such that nτ 2 σ 2, φ Z1,Z 2 (k 1, k 2 ) e θ 2 σ2 (k 2 1 +k 1k 2 +k 2 2 ) which is the characteristic function of a Bivariate Gaussian distribution with mean vector (0, 0) and variance-covariance matrix [ ] Σ = θσ 2 1 1/2 1/2 1
23 Large effect mutations Bivariate stable distribution Assume that p(y) κ y (α+1) and set t = κπ (sin(απ/2)γ(α)α) 1. As n and t 0 such that nt c, where c = c φ Z1,Z 2 (k 1, k 2 ) e 1 2 θc( k 1 α + k 2 α + k 1 +k 2 α ) π sin( απ 2 ) bivariate α-stable distribution, which is the characteristic function of a
24 de Finetti s theorem Theorem If (X 1, X 2,...) is an infinitely exchangeable random vector, then the probability density of (X 1 = x 1,... X n = x n ) is a mixture over i.i.d. probability densities. That is, p(x 1,..., x n ) = p θ (x i )ν(dθ) Suggests that we can find a de Finetti measure such that all our normalized samples are i.i.d. i
25 A guess in the normal case Given the mean, the trait is distributed N (0, θ 2 σ2 ) The mean is itself distributed N (0, θ 2 σ2 ) Law of total variance gives Var(X ) = 2Nµσ 2 - Same as classical derivation ( φ(k 1, k 2 ) = 2 ) e ik l x l 1 (m x πθσ 2 e l ) 2 1 θσ 2 m2 dx l e θσ 2 dm πθσ 2 l=1 = e θ 4 σ2 (k1 2+k2 2) = e θ 4 σ2 (k 2 1 +k2 2) e θ 4 σ2 (k 1 +k 2 ) 2 = e θ 2 σ2 (k 2 1 +k 1k 2 +k 2 2) e im(k 1+k 2 ) 1 m2 e θσ 2 dm πθσ 2
26 Tempting interpretation and conjecture Conditioning on the amount of evolution exclusive to sample 0 - Integrate over whether sample 1 or sample 2 coalesces with sample 0 first Suggests that this can be extended to arbitrary sample sizes Conjecture about larger samples (Gaussian limit) When the mutation kernel has only small effect mutations, the limit distribution is normal with variance θ 2 σ2 and random mean.
27 Testing the Gaussian conjecture Simulate quantitative traits according to the coalescent model Use KS test to assess convergence in the limit Normal, KS D # samples value diff same # loci
28 Conjecture for the stable limit Bivariate case - Independently α-stable with random median - Analogous to Gaussian limit Conjecture about larger samples (Stable limit) When the mutation kernel has large mutational effects, the limit distribution is α-stable with scale ( θ 2 c) 1/α and random median.
29 Testing the stable conjecture Simulate quantitative traits according to the coalescent model Use KS test to assess convergence in the limit Stable, KS D # samples value diff same # loci
30 More randomness than expected Density Trait value
31 Stable limit for sample of size 4 Tedious computation - Need to average over 18 trees. Conjecture would require φ Z1,Z 2,Z 3 (k 1, k 2, k 3 ) e θ 2 c( k 1 α + k 2 α + k 3 α + k 1 +k 2 +k 3 α ) Trivariate characteristic function For a sample of size three, under the same conditions as the bivariate limit, φ Z1,Z 2,Z 3 (k 1, k 2, k 3 ) exp{ θ 3 c( k 1 α + k 2 α + k 3 α k 1 + k 2 α k 1 + k 3 α k 2 + k 3 α + k 1 + k 2 + k 3 α )}
32 Extremely weak conjecture for large effect mutations Conjecture (Stable limit) When the mutation kernel has large mutational effects, the limit distribution is α-stable with random parameters Not even sure that this is true - A mixture of stable distributions?
33 Empirical data Neurospora crassa mrna-seq - Ellison et al (2011), PNAS - Restrict to genes with FPKM > 1 For each gene fit normal distribution, α-stable distribution Bootstrap likelihood ratio test (H 0 : α = 2 vs. H 1 : α < 2) Preliminary! - Only analyzed 346 genes
34 Distribution of p-values Frequency p value 54 genes significant at FDR of 32%
35 Distribution of α in significant genes Frequency α^
36 Cherry-picked examples NCU09477 NCU00484 Density Density log(fpkm) log(fpkm) NCU02435 NCU07659 Density Density log(fpkm) log(fpkm)
37 Conclusions Introduced a coalescent model of quantitative trait evolution - Provided exact formulas for the characteristic functions for samples of size 2, 3, 4 - Found limiting distributions as the number of loci became large in each case - Conjectured about extending these limits to larger samples Extensions - Samples not contemporaneous - Population structure - Diploids Why a neutral model? - Analytically tractable calculations - Intuition for what to expect with weak selection - Null model
Optimum selection and OU processes
 Optimum selection and OU processes 29 November 2016. Joe Felsenstein Biology 550D Optimum selection and OU processes p.1/15 With selection... life is harder There is the Breeder s Equation of Wright and
Optimum selection and OU processes 29 November 2016. Joe Felsenstein Biology 550D Optimum selection and OU processes p.1/15 With selection... life is harder There is the Breeder s Equation of Wright and
Mathematical models in population genetics II
 Mathematical models in population genetics II Anand Bhaskar Evolutionary Biology and Theory of Computing Bootcamp January 1, 014 Quick recap Large discrete-time randomly mating Wright-Fisher population
Mathematical models in population genetics II Anand Bhaskar Evolutionary Biology and Theory of Computing Bootcamp January 1, 014 Quick recap Large discrete-time randomly mating Wright-Fisher population
Frequency Spectra and Inference in Population Genetics
 Frequency Spectra and Inference in Population Genetics Although coalescent models have come to play a central role in population genetics, there are some situations where genealogies may not lead to efficient
Frequency Spectra and Inference in Population Genetics Although coalescent models have come to play a central role in population genetics, there are some situations where genealogies may not lead to efficient
Association studies and regression
 Association studies and regression CM226: Machine Learning for Bioinformatics. Fall 2016 Sriram Sankararaman Acknowledgments: Fei Sha, Ameet Talwalkar Association studies and regression 1 / 104 Administration
Association studies and regression CM226: Machine Learning for Bioinformatics. Fall 2016 Sriram Sankararaman Acknowledgments: Fei Sha, Ameet Talwalkar Association studies and regression 1 / 104 Administration
Lecture 18 : Ewens sampling formula
 Lecture 8 : Ewens sampling formula MATH85K - Spring 00 Lecturer: Sebastien Roch References: [Dur08, Chapter.3]. Previous class In the previous lecture, we introduced Kingman s coalescent as a limit of
Lecture 8 : Ewens sampling formula MATH85K - Spring 00 Lecturer: Sebastien Roch References: [Dur08, Chapter.3]. Previous class In the previous lecture, we introduced Kingman s coalescent as a limit of
Expression QTLs and Mapping of Complex Trait Loci. Paul Schliekelman Statistics Department University of Georgia
 Expression QTLs and Mapping of Complex Trait Loci Paul Schliekelman Statistics Department University of Georgia Definitions: Genes, Loci and Alleles A gene codes for a protein. Proteins due everything.
Expression QTLs and Mapping of Complex Trait Loci Paul Schliekelman Statistics Department University of Georgia Definitions: Genes, Loci and Alleles A gene codes for a protein. Proteins due everything.
The Wright-Fisher Model and Genetic Drift
 The Wright-Fisher Model and Genetic Drift January 22, 2015 1 1 Hardy-Weinberg Equilibrium Our goal is to understand the dynamics of allele and genotype frequencies in an infinite, randomlymating population
The Wright-Fisher Model and Genetic Drift January 22, 2015 1 1 Hardy-Weinberg Equilibrium Our goal is to understand the dynamics of allele and genotype frequencies in an infinite, randomlymating population
Population Genetics: a tutorial
 : a tutorial Institute for Science and Technology Austria ThRaSh 2014 provides the basic mathematical foundation of evolutionary theory allows a better understanding of experiments allows the development
: a tutorial Institute for Science and Technology Austria ThRaSh 2014 provides the basic mathematical foundation of evolutionary theory allows a better understanding of experiments allows the development
MODEL-FREE LINKAGE AND ASSOCIATION MAPPING OF COMPLEX TRAITS USING QUANTITATIVE ENDOPHENOTYPES
 MODEL-FREE LINKAGE AND ASSOCIATION MAPPING OF COMPLEX TRAITS USING QUANTITATIVE ENDOPHENOTYPES Saurabh Ghosh Human Genetics Unit Indian Statistical Institute, Kolkata Most common diseases are caused by
MODEL-FREE LINKAGE AND ASSOCIATION MAPPING OF COMPLEX TRAITS USING QUANTITATIVE ENDOPHENOTYPES Saurabh Ghosh Human Genetics Unit Indian Statistical Institute, Kolkata Most common diseases are caused by
Solutions to Even-Numbered Exercises to accompany An Introduction to Population Genetics: Theory and Applications Rasmus Nielsen Montgomery Slatkin
 Solutions to Even-Numbered Exercises to accompany An Introduction to Population Genetics: Theory and Applications Rasmus Nielsen Montgomery Slatkin CHAPTER 1 1.2 The expected homozygosity, given allele
Solutions to Even-Numbered Exercises to accompany An Introduction to Population Genetics: Theory and Applications Rasmus Nielsen Montgomery Slatkin CHAPTER 1 1.2 The expected homozygosity, given allele
Spring 2012 Math 541B Exam 1
 Spring 2012 Math 541B Exam 1 1. A sample of size n is drawn without replacement from an urn containing N balls, m of which are red and N m are black; the balls are otherwise indistinguishable. Let X denote
Spring 2012 Math 541B Exam 1 1. A sample of size n is drawn without replacement from an urn containing N balls, m of which are red and N m are black; the balls are otherwise indistinguishable. Let X denote
URN MODELS: the Ewens Sampling Lemma
 Department of Computer Science Brown University, Providence sorin@cs.brown.edu October 3, 2014 1 2 3 4 Mutation Mutation: typical values for parameters Equilibrium Probability of fixation 5 6 Ewens Sampling
Department of Computer Science Brown University, Providence sorin@cs.brown.edu October 3, 2014 1 2 3 4 Mutation Mutation: typical values for parameters Equilibrium Probability of fixation 5 6 Ewens Sampling
The mathematical challenge. Evolution in a spatial continuum. The mathematical challenge. Other recruits... The mathematical challenge
 The mathematical challenge What is the relative importance of mutation, selection, random drift and population subdivision for standing genetic variation? Evolution in a spatial continuum Al lison Etheridge
The mathematical challenge What is the relative importance of mutation, selection, random drift and population subdivision for standing genetic variation? Evolution in a spatial continuum Al lison Etheridge
Computational Systems Biology: Biology X
 Bud Mishra Room 1002, 715 Broadway, Courant Institute, NYU, New York, USA Human Population Genomics Outline 1 2 Damn the Human Genomes. Small initial populations; genes too distant; pestered with transposons;
Bud Mishra Room 1002, 715 Broadway, Courant Institute, NYU, New York, USA Human Population Genomics Outline 1 2 Damn the Human Genomes. Small initial populations; genes too distant; pestered with transposons;
Population Genetics I. Bio
 Population Genetics I. Bio5488-2018 Don Conrad dconrad@genetics.wustl.edu Why study population genetics? Functional Inference Demographic inference: History of mankind is written in our DNA. We can learn
Population Genetics I. Bio5488-2018 Don Conrad dconrad@genetics.wustl.edu Why study population genetics? Functional Inference Demographic inference: History of mankind is written in our DNA. We can learn
Dynamics of the evolving Bolthausen-Sznitman coalescent. by Jason Schweinsberg University of California at San Diego.
 Dynamics of the evolving Bolthausen-Sznitman coalescent by Jason Schweinsberg University of California at San Diego Outline of Talk 1. The Moran model and Kingman s coalescent 2. The evolving Kingman s
Dynamics of the evolving Bolthausen-Sznitman coalescent by Jason Schweinsberg University of California at San Diego Outline of Talk 1. The Moran model and Kingman s coalescent 2. The evolving Kingman s
Notes, March 4, 2013, R. Dudley Maximum likelihood estimation: actual or supposed
 18.466 Notes, March 4, 2013, R. Dudley Maximum likelihood estimation: actual or supposed 1. MLEs in exponential families Let f(x,θ) for x X and θ Θ be a likelihood function, that is, for present purposes,
18.466 Notes, March 4, 2013, R. Dudley Maximum likelihood estimation: actual or supposed 1. MLEs in exponential families Let f(x,θ) for x X and θ Θ be a likelihood function, that is, for present purposes,
GARCH processes continuous counterparts (Part 2)
 GARCH processes continuous counterparts (Part 2) Alexander Lindner Centre of Mathematical Sciences Technical University of Munich D 85747 Garching Germany lindner@ma.tum.de http://www-m1.ma.tum.de/m4/pers/lindner/
GARCH processes continuous counterparts (Part 2) Alexander Lindner Centre of Mathematical Sciences Technical University of Munich D 85747 Garching Germany lindner@ma.tum.de http://www-m1.ma.tum.de/m4/pers/lindner/
First Year Examination Department of Statistics, University of Florida
 First Year Examination Department of Statistics, University of Florida August 19, 010, 8:00 am - 1:00 noon Instructions: 1. You have four hours to answer questions in this examination.. You must show your
First Year Examination Department of Statistics, University of Florida August 19, 010, 8:00 am - 1:00 noon Instructions: 1. You have four hours to answer questions in this examination.. You must show your
This article appeared in a journal published by Elsevier. The attached copy is furnished to the author for internal non-commercial research and
 This article appeared in a journal published by Elsevier. The attached copy is furnished to the author for internal non-commercial research education use, including for instruction at the authors institution
This article appeared in a journal published by Elsevier. The attached copy is furnished to the author for internal non-commercial research education use, including for instruction at the authors institution
Evolutionary Theory. Sinauer Associates, Inc. Publishers Sunderland, Massachusetts U.S.A.
 Evolutionary Theory Mathematical and Conceptual Foundations Sean H. Rice Sinauer Associates, Inc. Publishers Sunderland, Massachusetts U.S.A. Contents Preface ix Introduction 1 CHAPTER 1 Selection on One
Evolutionary Theory Mathematical and Conceptual Foundations Sean H. Rice Sinauer Associates, Inc. Publishers Sunderland, Massachusetts U.S.A. Contents Preface ix Introduction 1 CHAPTER 1 Selection on One
Physics Sep Example A Spin System
 Physics 30 7-Sep-004 4- Example A Spin System In the last lecture, we discussed the binomial distribution. Now, I would like to add a little physical content by considering a spin system. Actually this
Physics 30 7-Sep-004 4- Example A Spin System In the last lecture, we discussed the binomial distribution. Now, I would like to add a little physical content by considering a spin system. Actually this
Statistical nonmolecular phylogenetics: can molecular phylogenies illuminate morphological evolution?
 Statistical nonmolecular phylogenetics: can molecular phylogenies illuminate morphological evolution? 30 July 2011. Joe Felsenstein Workshop on Molecular Evolution, MBL, Woods Hole Statistical nonmolecular
Statistical nonmolecular phylogenetics: can molecular phylogenies illuminate morphological evolution? 30 July 2011. Joe Felsenstein Workshop on Molecular Evolution, MBL, Woods Hole Statistical nonmolecular
Introduction to population genetics & evolution
 Introduction to population genetics & evolution Course Organization Exam dates: Feb 19 March 1st Has everybody registered? Did you get the email with the exam schedule Summer seminar: Hot topics in Bioinformatics
Introduction to population genetics & evolution Course Organization Exam dates: Feb 19 March 1st Has everybody registered? Did you get the email with the exam schedule Summer seminar: Hot topics in Bioinformatics
UNIT 8 BIOLOGY: Meiosis and Heredity Page 148
 UNIT 8 BIOLOGY: Meiosis and Heredity Page 148 CP: CHAPTER 6, Sections 1-6; CHAPTER 7, Sections 1-4; HN: CHAPTER 11, Section 1-5 Standard B-4: The student will demonstrate an understanding of the molecular
UNIT 8 BIOLOGY: Meiosis and Heredity Page 148 CP: CHAPTER 6, Sections 1-6; CHAPTER 7, Sections 1-4; HN: CHAPTER 11, Section 1-5 Standard B-4: The student will demonstrate an understanding of the molecular
Evolution in a spatial continuum
 Evolution in a spatial continuum Drift, draft and structure Alison Etheridge University of Oxford Joint work with Nick Barton (Edinburgh) and Tom Kurtz (Wisconsin) New York, Sept. 2007 p.1 Kingman s Coalescent
Evolution in a spatial continuum Drift, draft and structure Alison Etheridge University of Oxford Joint work with Nick Barton (Edinburgh) and Tom Kurtz (Wisconsin) New York, Sept. 2007 p.1 Kingman s Coalescent
Some of these slides have been borrowed from Dr. Paul Lewis, Dr. Joe Felsenstein. Thanks!
 Some of these slides have been borrowed from Dr. Paul Lewis, Dr. Joe Felsenstein. Thanks! Paul has many great tools for teaching phylogenetics at his web site: http://hydrodictyon.eeb.uconn.edu/people/plewis
Some of these slides have been borrowed from Dr. Paul Lewis, Dr. Joe Felsenstein. Thanks! Paul has many great tools for teaching phylogenetics at his web site: http://hydrodictyon.eeb.uconn.edu/people/plewis
Contrasts for a within-species comparative method
 Contrasts for a within-species comparative method Joseph Felsenstein, Department of Genetics, University of Washington, Box 357360, Seattle, Washington 98195-7360, USA email address: joe@genetics.washington.edu
Contrasts for a within-species comparative method Joseph Felsenstein, Department of Genetics, University of Washington, Box 357360, Seattle, Washington 98195-7360, USA email address: joe@genetics.washington.edu
Empirical Likelihood Tests for High-dimensional Data
 Empirical Likelihood Tests for High-dimensional Data Department of Statistics and Actuarial Science University of Waterloo, Canada ICSA - Canada Chapter 2013 Symposium Toronto, August 2-3, 2013 Based on
Empirical Likelihood Tests for High-dimensional Data Department of Statistics and Actuarial Science University of Waterloo, Canada ICSA - Canada Chapter 2013 Symposium Toronto, August 2-3, 2013 Based on
Random Vectors and Multivariate Normal Distributions
 Chapter 3 Random Vectors and Multivariate Normal Distributions 3.1 Random vectors Definition 3.1.1. Random vector. Random vectors are vectors of random 75 variables. For instance, X = X 1 X 2., where each
Chapter 3 Random Vectors and Multivariate Normal Distributions 3.1 Random vectors Definition 3.1.1. Random vector. Random vectors are vectors of random 75 variables. For instance, X = X 1 X 2., where each
Derivation of Itô SDE and Relationship to ODE and CTMC Models
 Derivation of Itô SDE and Relationship to ODE and CTMC Models Biomathematics II April 23, 2015 Linda J. S. Allen Texas Tech University TTU 1 Euler-Maruyama Method for Numerical Solution of an Itô SDE dx(t)
Derivation of Itô SDE and Relationship to ODE and CTMC Models Biomathematics II April 23, 2015 Linda J. S. Allen Texas Tech University TTU 1 Euler-Maruyama Method for Numerical Solution of an Itô SDE dx(t)
Fall 2017 STAT 532 Homework Peter Hoff. 1. Let P be a probability measure on a collection of sets A.
 1. Let P be a probability measure on a collection of sets A. (a) For each n N, let H n be a set in A such that H n H n+1. Show that P (H n ) monotonically converges to P ( k=1 H k) as n. (b) For each n
1. Let P be a probability measure on a collection of sets A. (a) For each n N, let H n be a set in A such that H n H n+1. Show that P (H n ) monotonically converges to P ( k=1 H k) as n. (b) For each n
Supplemental Information Likelihood-based inference in isolation-by-distance models using the spatial distribution of low-frequency alleles
 Supplemental Information Likelihood-based inference in isolation-by-distance models using the spatial distribution of low-frequency alleles John Novembre and Montgomery Slatkin Supplementary Methods To
Supplemental Information Likelihood-based inference in isolation-by-distance models using the spatial distribution of low-frequency alleles John Novembre and Montgomery Slatkin Supplementary Methods To
Gaussian processes. Chuong B. Do (updated by Honglak Lee) November 22, 2008
 Gaussian processes Chuong B Do (updated by Honglak Lee) November 22, 2008 Many of the classical machine learning algorithms that we talked about during the first half of this course fit the following pattern:
Gaussian processes Chuong B Do (updated by Honglak Lee) November 22, 2008 Many of the classical machine learning algorithms that we talked about during the first half of this course fit the following pattern:
BioControl - Week 6, Lecture 1
 BioControl - Week 6, Lecture 1 Goals of this lecture Large metabolic networks organization Design principles for small genetic modules - Rules based on gene demand - Rules based on error minimization Suggested
BioControl - Week 6, Lecture 1 Goals of this lecture Large metabolic networks organization Design principles for small genetic modules - Rules based on gene demand - Rules based on error minimization Suggested
Spring 2012 Math 541A Exam 1. X i, S 2 = 1 n. n 1. X i I(X i < c), T n =
 Spring 2012 Math 541A Exam 1 1. (a) Let Z i be independent N(0, 1), i = 1, 2,, n. Are Z = 1 n n Z i and S 2 Z = 1 n 1 n (Z i Z) 2 independent? Prove your claim. (b) Let X 1, X 2,, X n be independent identically
Spring 2012 Math 541A Exam 1 1. (a) Let Z i be independent N(0, 1), i = 1, 2,, n. Are Z = 1 n n Z i and S 2 Z = 1 n 1 n (Z i Z) 2 independent? Prove your claim. (b) Let X 1, X 2,, X n be independent identically
Problems for 3505 (2011)
 Problems for 505 (2011) 1. In the simplex of genotype distributions x + y + z = 1, for two alleles, the Hardy- Weinberg distributions x = p 2, y = 2pq, z = q 2 (p + q = 1) are characterized by y 2 = 4xz.
Problems for 505 (2011) 1. In the simplex of genotype distributions x + y + z = 1, for two alleles, the Hardy- Weinberg distributions x = p 2, y = 2pq, z = q 2 (p + q = 1) are characterized by y 2 = 4xz.
How robust are the predictions of the W-F Model?
 How robust are the predictions of the W-F Model? As simplistic as the Wright-Fisher model may be, it accurately describes the behavior of many other models incorporating additional complexity. Many population
How robust are the predictions of the W-F Model? As simplistic as the Wright-Fisher model may be, it accurately describes the behavior of many other models incorporating additional complexity. Many population
EXAMINATIONS OF THE ROYAL STATISTICAL SOCIETY
 EXAMINATIONS OF THE ROYAL STATISTICAL SOCIETY GRADUATE DIPLOMA, 2016 MODULE 1 : Probability distributions Time allowed: Three hours Candidates should answer FIVE questions. All questions carry equal marks.
EXAMINATIONS OF THE ROYAL STATISTICAL SOCIETY GRADUATE DIPLOMA, 2016 MODULE 1 : Probability distributions Time allowed: Three hours Candidates should answer FIVE questions. All questions carry equal marks.
Part IA Probability. Definitions. Based on lectures by R. Weber Notes taken by Dexter Chua. Lent 2015
 Part IA Probability Definitions Based on lectures by R. Weber Notes taken by Dexter Chua Lent 2015 These notes are not endorsed by the lecturers, and I have modified them (often significantly) after lectures.
Part IA Probability Definitions Based on lectures by R. Weber Notes taken by Dexter Chua Lent 2015 These notes are not endorsed by the lecturers, and I have modified them (often significantly) after lectures.
COMS 4721: Machine Learning for Data Science Lecture 10, 2/21/2017
 COMS 4721: Machine Learning for Data Science Lecture 10, 2/21/2017 Prof. John Paisley Department of Electrical Engineering & Data Science Institute Columbia University FEATURE EXPANSIONS FEATURE EXPANSIONS
COMS 4721: Machine Learning for Data Science Lecture 10, 2/21/2017 Prof. John Paisley Department of Electrical Engineering & Data Science Institute Columbia University FEATURE EXPANSIONS FEATURE EXPANSIONS
Lecture 9: Kernel (Variance Component) Tests and Omnibus Tests for Rare Variants. Summer Institute in Statistical Genetics 2017
 Lecture 9: Kernel (Variance Component) Tests and Omnibus Tests for Rare Variants Timothy Thornton and Michael Wu Summer Institute in Statistical Genetics 2017 1 / 46 Lecture Overview 1. Variance Component
Lecture 9: Kernel (Variance Component) Tests and Omnibus Tests for Rare Variants Timothy Thornton and Michael Wu Summer Institute in Statistical Genetics 2017 1 / 46 Lecture Overview 1. Variance Component
Gaussian Processes. 1. Basic Notions
 Gaussian Processes 1. Basic Notions Let T be a set, and X : {X } T a stochastic process, defined on a suitable probability space (Ω P), that is indexed by T. Definition 1.1. We say that X is a Gaussian
Gaussian Processes 1. Basic Notions Let T be a set, and X : {X } T a stochastic process, defined on a suitable probability space (Ω P), that is indexed by T. Definition 1.1. We say that X is a Gaussian
Estimation Theory. as Θ = (Θ 1,Θ 2,...,Θ m ) T. An estimator
 Estimation Theory Estimation theory deals with finding numerical values of interesting parameters from given set of data. We start with formulating a family of models that could describe how the data were
Estimation Theory Estimation theory deals with finding numerical values of interesting parameters from given set of data. We start with formulating a family of models that could describe how the data were
WXML Final Report: Chinese Restaurant Process
 WXML Final Report: Chinese Restaurant Process Dr. Noah Forman, Gerandy Brito, Alex Forney, Yiruey Chou, Chengning Li Spring 2017 1 Introduction The Chinese Restaurant Process (CRP) generates random partitions
WXML Final Report: Chinese Restaurant Process Dr. Noah Forman, Gerandy Brito, Alex Forney, Yiruey Chou, Chengning Li Spring 2017 1 Introduction The Chinese Restaurant Process (CRP) generates random partitions
Sympatric Speciation
 Sympatric Speciation Summary Speciation mechanisms: allopatry x sympatry The model of Dieckmann & Doebeli Sex Charles Darwin 1809-1882 Alfred R. Wallace 1823-1913 Seleção Natural Evolution by natural selection
Sympatric Speciation Summary Speciation mechanisms: allopatry x sympatry The model of Dieckmann & Doebeli Sex Charles Darwin 1809-1882 Alfred R. Wallace 1823-1913 Seleção Natural Evolution by natural selection
Eco517 Fall 2004 C. Sims MIDTERM EXAM
 Eco517 Fall 2004 C. Sims MIDTERM EXAM Answer all four questions. Each is worth 23 points. Do not devote disproportionate time to any one question unless you have answered all the others. (1) We are considering
Eco517 Fall 2004 C. Sims MIDTERM EXAM Answer all four questions. Each is worth 23 points. Do not devote disproportionate time to any one question unless you have answered all the others. (1) We are considering
Evolution of quantitative traits
 Evolution of quantitative traits Introduction Let s stop and review quickly where we ve come and where we re going We started our survey of quantitative genetics by pointing out that our objective was
Evolution of quantitative traits Introduction Let s stop and review quickly where we ve come and where we re going We started our survey of quantitative genetics by pointing out that our objective was
CPSC 540: Machine Learning
 CPSC 540: Machine Learning Undirected Graphical Models Mark Schmidt University of British Columbia Winter 2016 Admin Assignment 3: 2 late days to hand it in today, Thursday is final day. Assignment 4:
CPSC 540: Machine Learning Undirected Graphical Models Mark Schmidt University of British Columbia Winter 2016 Admin Assignment 3: 2 late days to hand it in today, Thursday is final day. Assignment 4:
On the posterior structure of NRMI
 On the posterior structure of NRMI Igor Prünster University of Turin, Collegio Carlo Alberto and ICER Joint work with L.F. James and A. Lijoi Isaac Newton Institute, BNR Programme, 8th August 2007 Outline
On the posterior structure of NRMI Igor Prünster University of Turin, Collegio Carlo Alberto and ICER Joint work with L.F. James and A. Lijoi Isaac Newton Institute, BNR Programme, 8th August 2007 Outline
This is a Gaussian probability centered around m = 0 (the most probable and mean position is the origin) and the mean square displacement m 2 = n,or
 Physics 7b: Statistical Mechanics Brownian Motion Brownian motion is the motion of a particle due to the buffeting by the molecules in a gas or liquid. The particle must be small enough that the effects
Physics 7b: Statistical Mechanics Brownian Motion Brownian motion is the motion of a particle due to the buffeting by the molecules in a gas or liquid. The particle must be small enough that the effects
Inférence en génétique des populations IV.
 Inférence en génétique des populations IV. François Rousset & Raphaël Leblois M2 Biostatistiques 2015 2016 FR & RL Inférence en génétique des populations IV. M2 Biostatistiques 2015 2016 1 / 33 Modeling
Inférence en génétique des populations IV. François Rousset & Raphaël Leblois M2 Biostatistiques 2015 2016 FR & RL Inférence en génétique des populations IV. M2 Biostatistiques 2015 2016 1 / 33 Modeling
Heterozygosity is variance. How Drift Affects Heterozygosity. Decay of heterozygosity in Buri s two experiments
 eterozygosity is variance ow Drift Affects eterozygosity Alan R Rogers September 17, 2014 Assumptions Random mating Allele A has frequency p N diploid individuals Let X 0,1, or 2) be the number of copies
eterozygosity is variance ow Drift Affects eterozygosity Alan R Rogers September 17, 2014 Assumptions Random mating Allele A has frequency p N diploid individuals Let X 0,1, or 2) be the number of copies
Multiple Random Variables
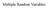 Multiple Random Variables Joint Probability Density Let X and Y be two random variables. Their joint distribution function is F ( XY x, y) P X x Y y. F XY ( ) 1, < x
Multiple Random Variables Joint Probability Density Let X and Y be two random variables. Their joint distribution function is F ( XY x, y) P X x Y y. F XY ( ) 1, < x
Tail probability of linear combinations of chi-square variables and its application to influence analysis in QTL detection
 Tail probability of linear combinations of chi-square variables and its application to influence analysis in QTL detection Satoshi Kuriki and Xiaoling Dou (Inst. Statist. Math., Tokyo) ISM Cooperative
Tail probability of linear combinations of chi-square variables and its application to influence analysis in QTL detection Satoshi Kuriki and Xiaoling Dou (Inst. Statist. Math., Tokyo) ISM Cooperative
Diffusion Models in Population Genetics
 Diffusion Models in Population Genetics Laura Kubatko kubatko.2@osu.edu MBI Workshop on Spatially-varying stochastic differential equations, with application to the biological sciences July 10, 2015 Laura
Diffusion Models in Population Genetics Laura Kubatko kubatko.2@osu.edu MBI Workshop on Spatially-varying stochastic differential equations, with application to the biological sciences July 10, 2015 Laura
Lecture 2: Basic Concepts of Statistical Decision Theory
 EE378A Statistical Signal Processing Lecture 2-03/31/2016 Lecture 2: Basic Concepts of Statistical Decision Theory Lecturer: Jiantao Jiao, Tsachy Weissman Scribe: John Miller and Aran Nayebi In this lecture
EE378A Statistical Signal Processing Lecture 2-03/31/2016 Lecture 2: Basic Concepts of Statistical Decision Theory Lecturer: Jiantao Jiao, Tsachy Weissman Scribe: John Miller and Aran Nayebi In this lecture
Linear Regression (1/1/17)
 STA613/CBB540: Statistical methods in computational biology Linear Regression (1/1/17) Lecturer: Barbara Engelhardt Scribe: Ethan Hada 1. Linear regression 1.1. Linear regression basics. Linear regression
STA613/CBB540: Statistical methods in computational biology Linear Regression (1/1/17) Lecturer: Barbara Engelhardt Scribe: Ethan Hada 1. Linear regression 1.1. Linear regression basics. Linear regression
Stationary Distribution of the Linkage Disequilibrium Coefficient r 2
 Stationary Distribution of the Linkage Disequilibrium Coefficient r 2 Wei Zhang, Jing Liu, Rachel Fewster and Jesse Goodman Department of Statistics, The University of Auckland December 1, 2015 Overview
Stationary Distribution of the Linkage Disequilibrium Coefficient r 2 Wei Zhang, Jing Liu, Rachel Fewster and Jesse Goodman Department of Statistics, The University of Auckland December 1, 2015 Overview
{σ x >t}p x. (σ x >t)=e at.
 3.11. EXERCISES 121 3.11 Exercises Exercise 3.1 Consider the Ornstein Uhlenbeck process in example 3.1.7(B). Show that the defined process is a Markov process which converges in distribution to an N(0,σ
3.11. EXERCISES 121 3.11 Exercises Exercise 3.1 Consider the Ornstein Uhlenbeck process in example 3.1.7(B). Show that the defined process is a Markov process which converges in distribution to an N(0,σ
Variance Component Models for Quantitative Traits. Biostatistics 666
 Variance Component Models for Quantitative Traits Biostatistics 666 Today Analysis of quantitative traits Modeling covariance for pairs of individuals estimating heritability Extending the model beyond
Variance Component Models for Quantitative Traits Biostatistics 666 Today Analysis of quantitative traits Modeling covariance for pairs of individuals estimating heritability Extending the model beyond
The Contour Process of Crump-Mode-Jagers Branching Processes
 The Contour Process of Crump-Mode-Jagers Branching Processes Emmanuel Schertzer (LPMA Paris 6), with Florian Simatos (ISAE Toulouse) June 24, 2015 Crump-Mode-Jagers trees Crump Mode Jagers (CMJ) branching
The Contour Process of Crump-Mode-Jagers Branching Processes Emmanuel Schertzer (LPMA Paris 6), with Florian Simatos (ISAE Toulouse) June 24, 2015 Crump-Mode-Jagers trees Crump Mode Jagers (CMJ) branching
Exercises. T 2T. e ita φ(t)dt.
 Exercises. Set #. Construct an example of a sequence of probability measures P n on R which converge weakly to a probability measure P but so that the first moments m,n = xdp n do not converge to m = xdp.
Exercises. Set #. Construct an example of a sequence of probability measures P n on R which converge weakly to a probability measure P but so that the first moments m,n = xdp n do not converge to m = xdp.
General Bayesian Inference I
 General Bayesian Inference I Outline: Basic concepts, One-parameter models, Noninformative priors. Reading: Chapters 10 and 11 in Kay-I. (Occasional) Simplified Notation. When there is no potential for
General Bayesian Inference I Outline: Basic concepts, One-parameter models, Noninformative priors. Reading: Chapters 10 and 11 in Kay-I. (Occasional) Simplified Notation. When there is no potential for
Looking Under the EA Hood with Price s Equation
 Looking Under the EA Hood with Price s Equation Jeffrey K. Bassett 1, Mitchell A. Potter 2, and Kenneth A. De Jong 1 1 George Mason University, Fairfax, VA 22030 {jbassett, kdejong}@cs.gmu.edu 2 Naval
Looking Under the EA Hood with Price s Equation Jeffrey K. Bassett 1, Mitchell A. Potter 2, and Kenneth A. De Jong 1 1 George Mason University, Fairfax, VA 22030 {jbassett, kdejong}@cs.gmu.edu 2 Naval
Mod-φ convergence I: examples and probabilistic estimates
 Mod-φ convergence I: examples and probabilistic estimates Valentin Féray (joint work with Pierre-Loïc Méliot and Ashkan Nikeghbali) Institut für Mathematik, Universität Zürich Summer school in Villa Volpi,
Mod-φ convergence I: examples and probabilistic estimates Valentin Féray (joint work with Pierre-Loïc Méliot and Ashkan Nikeghbali) Institut für Mathematik, Universität Zürich Summer school in Villa Volpi,
Machine Learning. Gaussian Mixture Models. Zhiyao Duan & Bryan Pardo, Machine Learning: EECS 349 Fall
 Machine Learning Gaussian Mixture Models Zhiyao Duan & Bryan Pardo, Machine Learning: EECS 349 Fall 2012 1 The Generative Model POV We think of the data as being generated from some process. We assume
Machine Learning Gaussian Mixture Models Zhiyao Duan & Bryan Pardo, Machine Learning: EECS 349 Fall 2012 1 The Generative Model POV We think of the data as being generated from some process. We assume
Density Modeling and Clustering Using Dirichlet Diffusion Trees
 p. 1/3 Density Modeling and Clustering Using Dirichlet Diffusion Trees Radford M. Neal Bayesian Statistics 7, 2003, pp. 619-629. Presenter: Ivo D. Shterev p. 2/3 Outline Motivation. Data points generation.
p. 1/3 Density Modeling and Clustering Using Dirichlet Diffusion Trees Radford M. Neal Bayesian Statistics 7, 2003, pp. 619-629. Presenter: Ivo D. Shterev p. 2/3 Outline Motivation. Data points generation.
Discrete & continuous characters: The threshold model
 Discrete & continuous characters: The threshold model Discrete & continuous characters: the threshold model So far we have discussed continuous & discrete character models separately for estimating ancestral
Discrete & continuous characters: The threshold model Discrete & continuous characters: the threshold model So far we have discussed continuous & discrete character models separately for estimating ancestral
ON COMPOUND POISSON POPULATION MODELS
 ON COMPOUND POISSON POPULATION MODELS Martin Möhle, University of Tübingen (joint work with Thierry Huillet, Université de Cergy-Pontoise) Workshop on Probability, Population Genetics and Evolution Centre
ON COMPOUND POISSON POPULATION MODELS Martin Möhle, University of Tübingen (joint work with Thierry Huillet, Université de Cergy-Pontoise) Workshop on Probability, Population Genetics and Evolution Centre
University of Regina. Lecture Notes. Michael Kozdron
 University of Regina Statistics 252 Mathematical Statistics Lecture Notes Winter 2005 Michael Kozdron kozdron@math.uregina.ca www.math.uregina.ca/ kozdron Contents 1 The Basic Idea of Statistics: Estimating
University of Regina Statistics 252 Mathematical Statistics Lecture Notes Winter 2005 Michael Kozdron kozdron@math.uregina.ca www.math.uregina.ca/ kozdron Contents 1 The Basic Idea of Statistics: Estimating
Some mathematical models from population genetics
 Some mathematical models from population genetics 5: Muller s ratchet and the rate of adaptation Alison Etheridge University of Oxford joint work with Peter Pfaffelhuber (Vienna), Anton Wakolbinger (Frankfurt)
Some mathematical models from population genetics 5: Muller s ratchet and the rate of adaptation Alison Etheridge University of Oxford joint work with Peter Pfaffelhuber (Vienna), Anton Wakolbinger (Frankfurt)
STAT 418: Probability and Stochastic Processes
 STAT 418: Probability and Stochastic Processes Spring 2016; Homework Assignments Latest updated on April 29, 2016 HW1 (Due on Jan. 21) Chapter 1 Problems 1, 8, 9, 10, 11, 18, 19, 26, 28, 30 Theoretical
STAT 418: Probability and Stochastic Processes Spring 2016; Homework Assignments Latest updated on April 29, 2016 HW1 (Due on Jan. 21) Chapter 1 Problems 1, 8, 9, 10, 11, 18, 19, 26, 28, 30 Theoretical
Problems on Evolutionary dynamics
 Problems on Evolutionary dynamics Doctoral Programme in Physics José A. Cuesta Lausanne, June 10 13, 2014 Replication 1. Consider the Galton-Watson process defined by the offspring distribution p 0 =
Problems on Evolutionary dynamics Doctoral Programme in Physics José A. Cuesta Lausanne, June 10 13, 2014 Replication 1. Consider the Galton-Watson process defined by the offspring distribution p 0 =
Phasing via the Expectation Maximization (EM) Algorithm
 Computing Haplotype Frequencies and Haplotype Phasing via the Expectation Maximization (EM) Algorithm Department of Computer Science Brown University, Providence sorin@cs.brown.edu September 14, 2010 Outline
Computing Haplotype Frequencies and Haplotype Phasing via the Expectation Maximization (EM) Algorithm Department of Computer Science Brown University, Providence sorin@cs.brown.edu September 14, 2010 Outline
Expectation Propagation Algorithm
 Expectation Propagation Algorithm 1 Shuang Wang School of Electrical and Computer Engineering University of Oklahoma, Tulsa, OK, 74135 Email: {shuangwang}@ou.edu This note contains three parts. First,
Expectation Propagation Algorithm 1 Shuang Wang School of Electrical and Computer Engineering University of Oklahoma, Tulsa, OK, 74135 Email: {shuangwang}@ou.edu This note contains three parts. First,
Wald s theorem and the Asimov data set
 Wald s theorem and the Asimov data set Eilam Gross & Ofer Vitells ATLAS statistics forum, Dec. 009 1 Outline We have previously guessed that the median significance of many toy MC experiments could be
Wald s theorem and the Asimov data set Eilam Gross & Ofer Vitells ATLAS statistics forum, Dec. 009 1 Outline We have previously guessed that the median significance of many toy MC experiments could be
Homework # , Spring Due 14 May Convergence of the empirical CDF, uniform samples
 Homework #3 36-754, Spring 27 Due 14 May 27 1 Convergence of the empirical CDF, uniform samples In this problem and the next, X i are IID samples on the real line, with cumulative distribution function
Homework #3 36-754, Spring 27 Due 14 May 27 1 Convergence of the empirical CDF, uniform samples In this problem and the next, X i are IID samples on the real line, with cumulative distribution function
Nonparametric hypothesis tests and permutation tests
 Nonparametric hypothesis tests and permutation tests 1.7 & 2.3. Probability Generating Functions 3.8.3. Wilcoxon Signed Rank Test 3.8.2. Mann-Whitney Test Prof. Tesler Math 283 Fall 2018 Prof. Tesler Wilcoxon
Nonparametric hypothesis tests and permutation tests 1.7 & 2.3. Probability Generating Functions 3.8.3. Wilcoxon Signed Rank Test 3.8.2. Mann-Whitney Test Prof. Tesler Math 283 Fall 2018 Prof. Tesler Wilcoxon
6 Introduction to Population Genetics
 Grundlagen der Bioinformatik, SoSe 14, D. Huson, May 18, 2014 67 6 Introduction to Population Genetics This chapter is based on: J. Hein, M.H. Schierup and C. Wuif, Gene genealogies, variation and evolution,
Grundlagen der Bioinformatik, SoSe 14, D. Huson, May 18, 2014 67 6 Introduction to Population Genetics This chapter is based on: J. Hein, M.H. Schierup and C. Wuif, Gene genealogies, variation and evolution,
Poisson Approximation for Two Scan Statistics with Rates of Convergence
 Poisson Approximation for Two Scan Statistics with Rates of Convergence Xiao Fang (Joint work with David Siegmund) National University of Singapore May 28, 2015 Outline The first scan statistic The second
Poisson Approximation for Two Scan Statistics with Rates of Convergence Xiao Fang (Joint work with David Siegmund) National University of Singapore May 28, 2015 Outline The first scan statistic The second
Random Averaging. Eli Ben-Naim Los Alamos National Laboratory. Paul Krapivsky (Boston University) John Machta (University of Massachusetts)
 Random Averaging Eli Ben-Naim Los Alamos National Laboratory Paul Krapivsky (Boston University) John Machta (University of Massachusetts) Talk, papers available from: http://cnls.lanl.gov/~ebn Plan I.
Random Averaging Eli Ben-Naim Los Alamos National Laboratory Paul Krapivsky (Boston University) John Machta (University of Massachusetts) Talk, papers available from: http://cnls.lanl.gov/~ebn Plan I.
Sample Spaces, Random Variables
 Sample Spaces, Random Variables Moulinath Banerjee University of Michigan August 3, 22 Probabilities In talking about probabilities, the fundamental object is Ω, the sample space. (elements) in Ω are denoted
Sample Spaces, Random Variables Moulinath Banerjee University of Michigan August 3, 22 Probabilities In talking about probabilities, the fundamental object is Ω, the sample space. (elements) in Ω are denoted
Gaussian processes for inference in stochastic differential equations
 Gaussian processes for inference in stochastic differential equations Manfred Opper, AI group, TU Berlin November 6, 2017 Manfred Opper, AI group, TU Berlin (TU Berlin) inference in SDE November 6, 2017
Gaussian processes for inference in stochastic differential equations Manfred Opper, AI group, TU Berlin November 6, 2017 Manfred Opper, AI group, TU Berlin (TU Berlin) inference in SDE November 6, 2017
Genetic Variation in Finite Populations
 Genetic Variation in Finite Populations The amount of genetic variation found in a population is influenced by two opposing forces: mutation and genetic drift. 1 Mutation tends to increase variation. 2
Genetic Variation in Finite Populations The amount of genetic variation found in a population is influenced by two opposing forces: mutation and genetic drift. 1 Mutation tends to increase variation. 2
Modelling Dependent Credit Risks
 Modelling Dependent Credit Risks Filip Lindskog, RiskLab, ETH Zürich 30 November 2000 Home page:http://www.math.ethz.ch/ lindskog E-mail:lindskog@math.ethz.ch RiskLab:http://www.risklab.ch Modelling Dependent
Modelling Dependent Credit Risks Filip Lindskog, RiskLab, ETH Zürich 30 November 2000 Home page:http://www.math.ethz.ch/ lindskog E-mail:lindskog@math.ethz.ch RiskLab:http://www.risklab.ch Modelling Dependent
Negative Association, Ordering and Convergence of Resampling Methods
 Negative Association, Ordering and Convergence of Resampling Methods Nicolas Chopin ENSAE, Paristech (Joint work with Mathieu Gerber and Nick Whiteley, University of Bristol) Resampling schemes: Informal
Negative Association, Ordering and Convergence of Resampling Methods Nicolas Chopin ENSAE, Paristech (Joint work with Mathieu Gerber and Nick Whiteley, University of Bristol) Resampling schemes: Informal
Remarks on Improper Ignorance Priors
 As a limit of proper priors Remarks on Improper Ignorance Priors Two caveats relating to computations with improper priors, based on their relationship with finitely-additive, but not countably-additive
As a limit of proper priors Remarks on Improper Ignorance Priors Two caveats relating to computations with improper priors, based on their relationship with finitely-additive, but not countably-additive
The Admixture Model in Linkage Analysis
 The Admixture Model in Linkage Analysis Jie Peng D. Siegmund Department of Statistics, Stanford University, Stanford, CA 94305 SUMMARY We study an appropriate version of the score statistic to test the
The Admixture Model in Linkage Analysis Jie Peng D. Siegmund Department of Statistics, Stanford University, Stanford, CA 94305 SUMMARY We study an appropriate version of the score statistic to test the
. Find E(V ) and var(v ).
 Math 6382/6383: Probability Models and Mathematical Statistics Sample Preliminary Exam Questions 1. A person tosses a fair coin until she obtains 2 heads in a row. She then tosses a fair die the same number
Math 6382/6383: Probability Models and Mathematical Statistics Sample Preliminary Exam Questions 1. A person tosses a fair coin until she obtains 2 heads in a row. She then tosses a fair die the same number
6.207/14.15: Networks Lecture 12: Generalized Random Graphs
 6.207/14.15: Networks Lecture 12: Generalized Random Graphs 1 Outline Small-world model Growing random networks Power-law degree distributions: Rich-Get-Richer effects Models: Uniform attachment model
6.207/14.15: Networks Lecture 12: Generalized Random Graphs 1 Outline Small-world model Growing random networks Power-law degree distributions: Rich-Get-Richer effects Models: Uniform attachment model
Normalized kernel-weighted random measures
 Normalized kernel-weighted random measures Jim Griffin University of Kent 1 August 27 Outline 1 Introduction 2 Ornstein-Uhlenbeck DP 3 Generalisations Bayesian Density Regression We observe data (x 1,
Normalized kernel-weighted random measures Jim Griffin University of Kent 1 August 27 Outline 1 Introduction 2 Ornstein-Uhlenbeck DP 3 Generalisations Bayesian Density Regression We observe data (x 1,
Lecture 24: Multivariate Response: Changes in G. Bruce Walsh lecture notes Synbreed course version 10 July 2013
 Lecture 24: Multivariate Response: Changes in G Bruce Walsh lecture notes Synbreed course version 10 July 2013 1 Overview Changes in G from disequilibrium (generalized Bulmer Equation) Fragility of covariances
Lecture 24: Multivariate Response: Changes in G Bruce Walsh lecture notes Synbreed course version 10 July 2013 1 Overview Changes in G from disequilibrium (generalized Bulmer Equation) Fragility of covariances
Coding sequence array Office hours Wednesday 3-4pm 304A Stanley Hall
 Coding sequence array Office hours Wednesday 3-4pm 304A Stanley Hall Review session 5pm Thursday, Dec. 11 GPB100 Fig. 1.13 RNA-seq Expression effects of cancer AAAAA AAAAA AAAAA Solexa sequencing counts
Coding sequence array Office hours Wednesday 3-4pm 304A Stanley Hall Review session 5pm Thursday, Dec. 11 GPB100 Fig. 1.13 RNA-seq Expression effects of cancer AAAAA AAAAA AAAAA Solexa sequencing counts
f (1 0.5)/n Z =
 Math 466/566 - Homework 4. We want to test a hypothesis involving a population proportion. The unknown population proportion is p. The null hypothesis is p = / and the alternative hypothesis is p > /.
Math 466/566 - Homework 4. We want to test a hypothesis involving a population proportion. The unknown population proportion is p. The null hypothesis is p = / and the alternative hypothesis is p > /.
FE 5204 Stochastic Differential Equations
 Instructor: Jim Zhu e-mail:zhu@wmich.edu http://homepages.wmich.edu/ zhu/ January 20, 2009 Preliminaries for dealing with continuous random processes. Brownian motions. Our main reference for this lecture
Instructor: Jim Zhu e-mail:zhu@wmich.edu http://homepages.wmich.edu/ zhu/ January 20, 2009 Preliminaries for dealing with continuous random processes. Brownian motions. Our main reference for this lecture
Stat 451 Lecture Notes Numerical Integration
 Stat 451 Lecture Notes 03 12 Numerical Integration Ryan Martin UIC www.math.uic.edu/~rgmartin 1 Based on Chapter 5 in Givens & Hoeting, and Chapters 4 & 18 of Lange 2 Updated: February 11, 2016 1 / 29
Stat 451 Lecture Notes 03 12 Numerical Integration Ryan Martin UIC www.math.uic.edu/~rgmartin 1 Based on Chapter 5 in Givens & Hoeting, and Chapters 4 & 18 of Lange 2 Updated: February 11, 2016 1 / 29
Model Specification Testing in Nonparametric and Semiparametric Time Series Econometrics. Jiti Gao
 Model Specification Testing in Nonparametric and Semiparametric Time Series Econometrics Jiti Gao Department of Statistics School of Mathematics and Statistics The University of Western Australia Crawley
Model Specification Testing in Nonparametric and Semiparametric Time Series Econometrics Jiti Gao Department of Statistics School of Mathematics and Statistics The University of Western Australia Crawley
Bayesian Methods for Machine Learning
 Bayesian Methods for Machine Learning CS 584: Big Data Analytics Material adapted from Radford Neal s tutorial (http://ftp.cs.utoronto.ca/pub/radford/bayes-tut.pdf), Zoubin Ghahramni (http://hunch.net/~coms-4771/zoubin_ghahramani_bayesian_learning.pdf),
Bayesian Methods for Machine Learning CS 584: Big Data Analytics Material adapted from Radford Neal s tutorial (http://ftp.cs.utoronto.ca/pub/radford/bayes-tut.pdf), Zoubin Ghahramni (http://hunch.net/~coms-4771/zoubin_ghahramani_bayesian_learning.pdf),
6. Brownian Motion. Q(A) = P [ ω : x(, ω) A )
 6. Brownian Motion. stochastic process can be thought of in one of many equivalent ways. We can begin with an underlying probability space (Ω, Σ, P) and a real valued stochastic process can be defined
6. Brownian Motion. stochastic process can be thought of in one of many equivalent ways. We can begin with an underlying probability space (Ω, Σ, P) and a real valued stochastic process can be defined
