Higher order patterns via factor models
|
|
|
- Darren Green
- 5 years ago
- Views:
Transcription
1 1/39 Higher order patterns via factor models 567 Statistical analysis of social networks Peter Hoff Statistics, University of Washington
2 2/39 Conflict data Y<-conflict90s$conflicts Xn<-conflict90s$nodevars colnames(xn) ## [1] "pop" "gdp" "polity" Xn[,1:2]<-log(Xn[,1:2]) table(y) ## Y ## ##
3 Dichotomized conflict data AFG ALB ANG ARG AUL BAH BEL BEN BNG BOT BUI CAM CAN CAO CDI CHA CHL CHN COL CON COS CUB CYP DOM DRC EGY FRN GHA GNB GRC GUI GUY HAI HON IND INS IRN IRQ ISR ITA JOR JPN KEN LBR LES LIB MAA MAL MLI MON MOR MYA MZM NAM NIC NIG NIR NTH OMA PAK PHI PNG PRK QAT ROK RWA SAF SAL SAU SEN SIE SIN SPN SRI SUD SWA SYR TAW TAZ THI TOG TRI TUR UAE UGA UKG USA VEN YEM ZAM ZIM 3/39
4 4/39 SRM fit to ordinal data #### nodal covariates only fit_n<-ame(y,xrow=xn,xcol=xn,model="ord",nscan=10000) summary(fit_n) ## ## beta: ## pmean psd z-stat p-val ## pop.row ## gdp.row ## polity.row ## pop.col ## gdp.col ## polity.col ## ## Sigma_ab pmean: ## a b ## a ## b ## ## rho pmean: ## 0.804
5 5/39 SRM fit to ordinal data plot(fit_n) SABR BETA sd.rowmean sd.colmean dyad.dep triad.dep
6 6/39 Triad dependence measure gofstats ## function (Y) ## { ## sd.rowmean <- sd(rowmeans(y, na.rm = TRUE), na.rm = TRUE) ## sd.colmean <- sd(colmeans(y, na.rm = TRUE), na.rm = TRUE) ## dyad.dep <- cor(c(y), c(t(y)), use = "complete.obs") ## E <- Y - mean(y, na.rm = TRUE) ## D <- 1 * (!is.na(e)) ## E[is.na(E)] <- 0 ## triad.dep <- sum(diag(e %*% E %*% E))/(sum(diag(D %*% D %*% ## D)) * sd(c(y), na.rm = TRUE)^3) ## gof <- c(sd.rowmean, sd.colmean, dyad.dep, triad.dep) ## names(gof) <- c("sd.rowmean", "sd.colmean", "dyad.dep", "triad.dep") ## gof ## } ## <environment: namespace:amen>
7 7/39 Triad dependence measure Let E = (Y 11 T ȳ )/s y, that is, e i,j = (y i,j ȳ )/s y. ȳ is the grand mean; s y is the sample standard deviation of the y i,j s e i,j is like a scaled residual from the simple mean model y i,j = µ + ɛ i,j. The sum of the diagonal of E 3 gives the scaled third-order moment: trace(e 3 ) = e i,j e j,k e k,i i j The GOF statistic used in amen is the average of the e i,j e j,k e k,i s: i j k t(y) = e i,je j,k e i,j # of ordered triads k
8 7/39 Triad dependence measure Let E = (Y 11 T ȳ )/s y, that is, e i,j = (y i,j ȳ )/s y. ȳ is the grand mean; s y is the sample standard deviation of the y i,j s e i,j is like a scaled residual from the simple mean model y i,j = µ + ɛ i,j. The sum of the diagonal of E 3 gives the scaled third-order moment: trace(e 3 ) = e i,j e j,k e k,i i j The GOF statistic used in amen is the average of the e i,j e j,k e k,i s: i j k t(y) = e i,je j,k e i,j # of ordered triads k
9 7/39 Triad dependence measure Let E = (Y 11 T ȳ )/s y, that is, e i,j = (y i,j ȳ )/s y. ȳ is the grand mean; s y is the sample standard deviation of the y i,j s e i,j is like a scaled residual from the simple mean model y i,j = µ + ɛ i,j. The sum of the diagonal of E 3 gives the scaled third-order moment: trace(e 3 ) = e i,j e j,k e k,i i j The GOF statistic used in amen is the average of the e i,j e j,k e k,i s: i j k t(y) = e i,je j,k e i,j # of ordered triads k
10 7/39 Triad dependence measure Let E = (Y 11 T ȳ )/s y, that is, e i,j = (y i,j ȳ )/s y. ȳ is the grand mean; s y is the sample standard deviation of the y i,j s e i,j is like a scaled residual from the simple mean model y i,j = µ + ɛ i,j. The sum of the diagonal of E 3 gives the scaled third-order moment: trace(e 3 ) = e i,j e j,k e k,i i j The GOF statistic used in amen is the average of the e i,j e j,k e k,i s: i j k t(y) = e i,je j,k e i,j # of ordered triads k
11 8/39 Triad dependence and transitivity This triadic dependence measure is related to transitivity for binary data: Transitive triple: An ordered triple (i, j, k) is transitive if i j k i (y i,j = y j,k = y k,i ). trans(y) = i y i,j y j,k y k,i j k
12 8/39 Triad dependence and transitivity This triadic dependence measure is related to transitivity for binary data: Transitive triple: An ordered triple (i, j, k) is transitive if i j k i (y i,j = y j,k = y k,i ). trans(y) = i y i,j y j,k y k,i j k
13 8/39 Triad dependence and transitivity This triadic dependence measure is related to transitivity for binary data: Transitive triple: An ordered triple (i, j, k) is transitive if i j k i (y i,j = y j,k = y k,i ). trans(y) = i y i,j y j,k y k,i j k
14 9/39 SRM fit to ordinal data with dyadic covariates #### dyadic and nodal covariates fit_dn<-ame(y,xdyad=xd,xrow=xn,xcol=xn,model="ord",nscan=10000) summary(fit_dn) ## ## beta: ## pmean psd z-stat p-val ## pop.row ## gdp.row ## polity.row ## pop.col ## gdp.col ## polity.col ## polity_int.dyad ## imports.dyad ## shared_igos.dyad ## distance.dyad ## ## Sigma_ab pmean: ## a b ## a ## b ## ## rho pmean: ## 0.566
15 10/39 SRM fit to ordinal data with dyadic covariates plot(fit_dn) SABR BETA sd.rowmean sd.colmean dyad.dep triad.dep
16 11/39 GOF comparison Density th Density th
17 12/39 Dyadic functions of nodal characteristics Adding these dyadic covariates improved the fit with regard to t(y). Note that each of these dyadic covariates was a function of nodal covariates: x i,j,1 = f 1(polity i, polity j ) x i,j,2 = f 2(igo i, igo j ) x i,j,3 = f 3(location i, location j ) This suggests models of the form y i,j β 0 + β 1x i,j x i,j = s(x i, x j ) where s(, ) is some (non-additive) function of the nodal characteristics x i, x j.
18 12/39 Dyadic functions of nodal characteristics Adding these dyadic covariates improved the fit with regard to t(y). Note that each of these dyadic covariates was a function of nodal covariates: x i,j,1 = f 1(polity i, polity j ) x i,j,2 = f 2(igo i, igo j ) x i,j,3 = f 3(location i, location j ) This suggests models of the form y i,j β 0 + β 1x i,j x i,j = s(x i, x j ) where s(, ) is some (non-additive) function of the nodal characteristics x i, x j.
19 12/39 Dyadic functions of nodal characteristics Adding these dyadic covariates improved the fit with regard to t(y). Note that each of these dyadic covariates was a function of nodal covariates: x i,j,1 = f 1(polity i, polity j ) x i,j,2 = f 2(igo i, igo j ) x i,j,3 = f 3(location i, location j ) This suggests models of the form y i,j β 0 + β 1x i,j x i,j = s(x i, x j ) where s(, ) is some (non-additive) function of the nodal characteristics x i, x j.
20 12/39 Dyadic functions of nodal characteristics Adding these dyadic covariates improved the fit with regard to t(y). Note that each of these dyadic covariates was a function of nodal covariates: x i,j,1 = f 1(polity i, polity j ) x i,j,2 = f 2(igo i, igo j ) x i,j,3 = f 3(location i, location j ) This suggests models of the form y i,j β 0 + β 1x i,j x i,j = s(x i, x j ) where s(, ) is some (non-additive) function of the nodal characteristics x i, x j.
21 12/39 Dyadic functions of nodal characteristics Adding these dyadic covariates improved the fit with regard to t(y). Note that each of these dyadic covariates was a function of nodal covariates: x i,j,1 = f 1(polity i, polity j ) x i,j,2 = f 2(igo i, igo j ) x i,j,3 = f 3(location i, location j ) This suggests models of the form y i,j β 0 + β 1x i,j x i,j = s(x i, x j ) where s(, ) is some (non-additive) function of the nodal characteristics x i, x j.
22 12/39 Dyadic functions of nodal characteristics Adding these dyadic covariates improved the fit with regard to t(y). Note that each of these dyadic covariates was a function of nodal covariates: x i,j,1 = f 1(polity i, polity j ) x i,j,2 = f 2(igo i, igo j ) x i,j,3 = f 3(location i, location j ) This suggests models of the form y i,j β 0 + β 1x i,j x i,j = s(x i, x j ) where s(, ) is some (non-additive) function of the nodal characteristics x i, x j.
23 13/39 Homophily and stochastic equivalence More generally, let u i v j be a covariate of i as a sender of ties; be covariate of j as a receiver of ties. y i,j β 0 + β 1 s(u i, v j ) Such a model can describe various types of higher-order dependence, including transitivity stochastic equivalence
24 13/39 Homophily and stochastic equivalence More generally, let u i v j be a covariate of i as a sender of ties; be covariate of j as a receiver of ties. y i,j β 0 + β 1 s(u i, v j ) Such a model can describe various types of higher-order dependence, including transitivity stochastic equivalence
25 13/39 Homophily and stochastic equivalence More generally, let u i v j be a covariate of i as a sender of ties; be covariate of j as a receiver of ties. y i,j β 0 + β 1 s(u i, v j ) Such a model can describe various types of higher-order dependence, including transitivity stochastic equivalence
26 14/39 Transitivity via homophily Homophily on covariates can explain transitivity: y i,j β 0 + β 1 s(x i, x j ) If β 1 > 0 and the y i,j s are binary, then y i,j = 1 x i x j ; y i,k = 1 x i x k ; x i x j, x i x k x j x k. x j x k y j,k = 1. Similarly, homophily can explain triadic dependence for ordinal y i,j s.
27 14/39 Transitivity via homophily Homophily on covariates can explain transitivity: y i,j β 0 + β 1 s(x i, x j ) If β 1 > 0 and the y i,j s are binary, then y i,j = 1 x i x j ; y i,k = 1 x i x k ; x i x j, x i x k x j x k. x j x k y j,k = 1. Similarly, homophily can explain triadic dependence for ordinal y i,j s.
28 14/39 Transitivity via homophily Homophily on covariates can explain transitivity: y i,j β 0 + β 1 s(x i, x j ) If β 1 > 0 and the y i,j s are binary, then y i,j = 1 x i x j ; y i,k = 1 x i x k ; x i x j, x i x k x j x k. x j x k y j,k = 1. Similarly, homophily can explain triadic dependence for ordinal y i,j s.
29 4/39 Transitivity via homophily Homophily on covariates can explain transitivity: y i,j β 0 + β 1 s(x i, x j ) If β 1 > 0 and the y i,j s are binary, then y i,j = 1 x i x j ; y i,k = 1 x i x k ; x i x j, x i x k x j x k. x j x k y j,k = 1. Similarly, homophily can explain triadic dependence for ordinal y i,j s.
30 4/39 Transitivity via homophily Homophily on covariates can explain transitivity: y i,j β 0 + β 1 s(x i, x j ) If β 1 > 0 and the y i,j s are binary, then y i,j = 1 x i x j ; y i,k = 1 x i x k ; x i x j, x i x k x j x k. x j x k y j,k = 1. Similarly, homophily can explain triadic dependence for ordinal y i,j s.
31 4/39 Transitivity via homophily Homophily on covariates can explain transitivity: y i,j β 0 + β 1 s(x i, x j ) If β 1 > 0 and the y i,j s are binary, then y i,j = 1 x i x j ; y i,k = 1 x i x k ; x i x j, x i x k x j x k. x j x k y j,k = 1. Similarly, homophily can explain triadic dependence for ordinal y i,j s.
32 15/39 Stochastic equivalence Returning to the more general model: y i,j β 0 + β 1 s(u i, v j ) If u i = u k then i and j are equivalent as senders in terms of the model. Example: Logistic regression If u i = u k = u, then Pr(Y i,j = 1) = eβ 0+β 1 s(u i,v j ) 1 + e β 0+β 1 s(u i,v j ) Pr(Y i,j = 1) = eβ 0+β 1 s(u i,v j ) 1 + e β 0+β 1 s(u i,v j ) = Pr(Y k,j = 1), and nodes i and k are stochastically equivalent (as senders).
33 15/39 Stochastic equivalence Returning to the more general model: y i,j β 0 + β 1 s(u i, v j ) If u i = u k then i and j are equivalent as senders in terms of the model. Example: Logistic regression If u i = u k = u, then Pr(Y i,j = 1) = eβ 0+β 1 s(u i,v j ) 1 + e β 0+β 1 s(u i,v j ) Pr(Y i,j = 1) = eβ 0+β 1 s(u i,v j ) 1 + e β 0+β 1 s(u i,v j ) = Pr(Y k,j = 1), and nodes i and k are stochastically equivalent (as senders).
34 15/39 Stochastic equivalence Returning to the more general model: y i,j β 0 + β 1 s(u i, v j ) If u i = u k then i and j are equivalent as senders in terms of the model. Example: Logistic regression If u i = u k = u, then Pr(Y i,j = 1) = eβ 0+β 1 s(u i,v j ) 1 + e β 0+β 1 s(u i,v j ) Pr(Y i,j = 1) = eβ 0+β 1 s(u i,v j ) 1 + e β 0+β 1 s(u i,v j ) = Pr(Y k,j = 1), and nodes i and k are stochastically equivalent (as senders).
35 16/39 Homophily and stochastic equivalence In our hypothetical model of social relations, we ve seen how nodal characteristics relate to homophily and stochastic equivalence. Homophily: Similar nodes link to each other similar in terms of characteristics (potentially unobserved) homophily leads to transitive or clustered social networks observed transitivity may be due to exogenous or endogenous factors ( See Shalizi and Thomas 2010 for a more careful discussion ) Stochastic equivalence: Similar nodes have similar relational patterns similar nodes may or may not link to each other equivalent nodes can be thought of as having the same role
36 16/39 Homophily and stochastic equivalence In our hypothetical model of social relations, we ve seen how nodal characteristics relate to homophily and stochastic equivalence. Homophily: Similar nodes link to each other similar in terms of characteristics (potentially unobserved) homophily leads to transitive or clustered social networks observed transitivity may be due to exogenous or endogenous factors ( See Shalizi and Thomas 2010 for a more careful discussion ) Stochastic equivalence: Similar nodes have similar relational patterns similar nodes may or may not link to each other equivalent nodes can be thought of as having the same role
37 16/39 Homophily and stochastic equivalence In our hypothetical model of social relations, we ve seen how nodal characteristics relate to homophily and stochastic equivalence. Homophily: Similar nodes link to each other similar in terms of characteristics (potentially unobserved) homophily leads to transitive or clustered social networks observed transitivity may be due to exogenous or endogenous factors ( See Shalizi and Thomas 2010 for a more careful discussion ) Stochastic equivalence: Similar nodes have similar relational patterns similar nodes may or may not link to each other equivalent nodes can be thought of as having the same role
38 16/39 Homophily and stochastic equivalence In our hypothetical model of social relations, we ve seen how nodal characteristics relate to homophily and stochastic equivalence. Homophily: Similar nodes link to each other similar in terms of characteristics (potentially unobserved) homophily leads to transitive or clustered social networks observed transitivity may be due to exogenous or endogenous factors ( See Shalizi and Thomas 2010 for a more careful discussion ) Stochastic equivalence: Similar nodes have similar relational patterns similar nodes may or may not link to each other equivalent nodes can be thought of as having the same role
39 16/39 Homophily and stochastic equivalence In our hypothetical model of social relations, we ve seen how nodal characteristics relate to homophily and stochastic equivalence. Homophily: Similar nodes link to each other similar in terms of characteristics (potentially unobserved) homophily leads to transitive or clustered social networks observed transitivity may be due to exogenous or endogenous factors ( See Shalizi and Thomas 2010 for a more careful discussion ) Stochastic equivalence: Similar nodes have similar relational patterns similar nodes may or may not link to each other equivalent nodes can be thought of as having the same role
40 16/39 Homophily and stochastic equivalence In our hypothetical model of social relations, we ve seen how nodal characteristics relate to homophily and stochastic equivalence. Homophily: Similar nodes link to each other similar in terms of characteristics (potentially unobserved) homophily leads to transitive or clustered social networks observed transitivity may be due to exogenous or endogenous factors ( See Shalizi and Thomas 2010 for a more careful discussion ) Stochastic equivalence: Similar nodes have similar relational patterns similar nodes may or may not link to each other equivalent nodes can be thought of as having the same role
41 16/39 Homophily and stochastic equivalence In our hypothetical model of social relations, we ve seen how nodal characteristics relate to homophily and stochastic equivalence. Homophily: Similar nodes link to each other similar in terms of characteristics (potentially unobserved) homophily leads to transitive or clustered social networks observed transitivity may be due to exogenous or endogenous factors ( See Shalizi and Thomas 2010 for a more careful discussion ) Stochastic equivalence: Similar nodes have similar relational patterns similar nodes may or may not link to each other equivalent nodes can be thought of as having the same role
42 16/39 Homophily and stochastic equivalence In our hypothetical model of social relations, we ve seen how nodal characteristics relate to homophily and stochastic equivalence. Homophily: Similar nodes link to each other similar in terms of characteristics (potentially unobserved) homophily leads to transitive or clustered social networks observed transitivity may be due to exogenous or endogenous factors ( See Shalizi and Thomas 2010 for a more careful discussion ) Stochastic equivalence: Similar nodes have similar relational patterns similar nodes may or may not link to each other equivalent nodes can be thought of as having the same role
43 16/39 Homophily and stochastic equivalence In our hypothetical model of social relations, we ve seen how nodal characteristics relate to homophily and stochastic equivalence. Homophily: Similar nodes link to each other similar in terms of characteristics (potentially unobserved) homophily leads to transitive or clustered social networks observed transitivity may be due to exogenous or endogenous factors ( See Shalizi and Thomas 2010 for a more careful discussion ) Stochastic equivalence: Similar nodes have similar relational patterns similar nodes may or may not link to each other equivalent nodes can be thought of as having the same role
44 16/39 Homophily and stochastic equivalence In our hypothetical model of social relations, we ve seen how nodal characteristics relate to homophily and stochastic equivalence. Homophily: Similar nodes link to each other similar in terms of characteristics (potentially unobserved) homophily leads to transitive or clustered social networks observed transitivity may be due to exogenous or endogenous factors ( See Shalizi and Thomas 2010 for a more careful discussion ) Stochastic equivalence: Similar nodes have similar relational patterns similar nodes may or may not link to each other equivalent nodes can be thought of as having the same role
45 16/39 Homophily and stochastic equivalence In our hypothetical model of social relations, we ve seen how nodal characteristics relate to homophily and stochastic equivalence. Homophily: Similar nodes link to each other similar in terms of characteristics (potentially unobserved) homophily leads to transitive or clustered social networks observed transitivity may be due to exogenous or endogenous factors ( See Shalizi and Thomas 2010 for a more careful discussion ) Stochastic equivalence: Similar nodes have similar relational patterns similar nodes may or may not link to each other equivalent nodes can be thought of as having the same role
46 Visualizing stochastic equivalence For which network is homophily a plausible explanation? Which network exhibits a large degree of stochastic equivalence? 17/39
47 Visualizing stochastic equivalence For which network is homophily a plausible explanation? Which network exhibits a large degree of stochastic equivalence? 17/39
48 Homophily and stochastic equivalence in real networks, ; :. a above abundantly after air all also and appear be bearing beast beginning behold blessed bring brought called cattle created creature creepeth creeping darkness day days deep divide divided dominion dry earth evening every face female fifth fill finished firmament first fish fly for form forth fourth fowl from fruit fruitful gathered gathering give given god good grass great greater green had hath have he heaven heavens herb him his host i image in is it itself kind land lesser let life light lights likeness living made make male man may meat midst morning moved moveth moving multiply night of one open our over own place replenish rule said saw saying sea seas seasons second seed set shall signs sixth so spirit stars subdue that the their them there thing third thus to together tree two under unto upon us very void was waters were whales wherein which whose winged without years yielding you b0185 b2316 b1094 b4187 b2697 b1731 b2097 b2496 b4131 b1000 b0882 b0437 b0438b1120 b3556 b1557 b1823 b0880 b0623 b3162 b3702 b0184 b0015 b0014 b0215 b2779 b1288 b0180 b2925 b3895 b0684 b3463 b0095 b3517 b1493 b3181 b3406 b2614 b2231 b3699 b0059 b4172 b1099 b4259 b4372 b1842 b2526 b3931 b3932 b4000 b0440 b2728 b2729 b3687 b1718 b2530 b2529 b1215 b0186 b0179 b2890 b4129 b3418 b2942 b4143 b4142 b3251 b0924 b0923 b3189 b1224 b1225 b1226 b1467 b3169 b3982 b0133 b3019 b1127 b0903 b3164 b3863 b4226 b1207 b2585 b3725 b2476 b1854 b2892 b3822 b3619 b2038 b3780 b1084 b0214 b3704 b3319 b3295b3987 b3988 b3067 b3461 b3202 b2741 b3649 b0098 b3609 b3590 b2620 b3650 b2576 b4059 b3115 b0406 b1274 b1763 b3919 b0170 b3339 b3980 b2319 b1913 b4179 b0101 b0119 b0528 b0607 b0660 b0877 b0948 b1055 b1086 b1103 b1269 b2330 b2517 b2579 b2581 b2830 b2960 b3180 b3470 b3746 b4191 b0661 b2836 b2323 b4041 b2563 b1095 b1093 b3730 b0736 b3888 b0492 b3701 b3783 b3317 b4203 b3315 b3314 b0169 b3341 b1468 b2592 b0990 b3298 b3320 b3308 b3231 b0911 b3296 b3984b3303 b3309 b3321 b0023 b1552 b3054 b2606 b3186 b4200 b3306 b3230 b3307 b3165 b2609 b4202 b0470 b0640 b2699 b0595 b1294 b2610 b3305 b2515 b2311 b3340 b3247 b1413 b0439 b0436 b0178 b1808 b3935 b3417 b2487 b2721 b3686 b3986 b2096 b0922 b3168 b0480 b2011 b3301 b1716 b2372 b4051 b1817 b0925 b3637 b3652 b3294 b3499 b3304 b3602 b3318 b2804 b3342 AddHealth friendships: friendships among th-graders Word neighbors in Genesis: neighboring occurrences among 158 words Protein binding interactions: binding patterns among 230 proteins 18/39
49 Homophily and stochastic equivalence in real networks, ; :. a above abundantly after air all also and appear be bearing beast beginning behold blessed bring brought called cattle created creature creepeth creeping darkness day days deep divide divided dominion dry earth evening every face female fifth fill finished firmament first fish fly for form forth fourth fowl from fruit fruitful gathered gathering give given god good grass great greater green had hath have he heaven heavens herb him his host i image in is it itself kind land lesser let life light lights likeness living made make male man may meat midst morning moved moveth moving multiply night of one open our over own place replenish rule said saw saying sea seas seasons second seed set shall signs sixth so spirit stars subdue that the their them there thing third thus to together tree two under unto upon us very void was waters were whales wherein which whose winged without years yielding you b0185 b2316 b1094 b4187 b2697 b1731 b2097 b2496 b4131 b1000 b0882 b0437 b0438b1120 b3556 b1557 b1823 b0880 b0623 b3162 b3702 b0184 b0015 b0014 b0215 b2779 b1288 b0180 b2925 b3895 b0684 b3463 b0095 b3517 b1493 b3181 b3406 b2614 b2231 b3699 b0059 b4172 b1099 b4259 b4372 b1842 b2526 b3931 b3932 b4000 b0440 b2728 b2729 b3687 b1718 b2530 b2529 b1215 b0186 b0179 b2890 b4129 b3418 b2942 b4143 b4142 b3251 b0924 b0923 b3189 b1224 b1225 b1226 b1467 b3169 b3982 b0133 b3019 b1127 b0903 b3164 b3863 b4226 b1207 b2585 b3725 b2476 b1854 b2892 b3822 b3619 b2038 b3780 b1084 b0214 b3704 b3319 b3295b3987 b3988 b3067 b3461 b3202 b2741 b3649 b0098 b3609 b3590 b2620 b3650 b2576 b4059 b3115 b0406 b1274 b1763 b3919 b0170 b3339 b3980 b2319 b1913 b4179 b0101 b0119 b0528 b0607 b0660 b0877 b0948 b1055 b1086 b1103 b1269 b2330 b2517 b2579 b2581 b2830 b2960 b3180 b3470 b3746 b4191 b0661 b2836 b2323 b4041 b2563 b1095 b1093 b3730 b0736 b3888 b0492 b3701 b3783 b3317 b4203 b3315 b3314 b0169 b3341 b1468 b2592 b0990 b3298 b3320 b3308 b3231 b0911 b3296 b3984b3303 b3309 b3321 b0023 b1552 b3054 b2606 b3186 b4200 b3306 b3230 b3307 b3165 b2609 b4202 b0470 b0640 b2699 b0595 b1294 b2610 b3305 b2515 b2311 b3340 b3247 b1413 b0439 b0436 b0178 b1808 b3935 b3417 b2487 b2721 b3686 b3986 b2096 b0922 b3168 b0480 b2011 b3301 b1716 b2372 b4051 b1817 b0925 b3637 b3652 b3294 b3499 b3304 b3602 b3318 b2804 b3342 AddHealth friendships: friendships among th-graders Word neighbors in Genesis: neighboring occurrences among 158 words Protein binding interactions: binding patterns among 230 proteins 18/39
50 Homophily and stochastic equivalence in real networks, ; :. a above abundantly after air all also and appear be bearing beast beginning behold blessed bring brought called cattle created creature creepeth creeping darkness day days deep divide divided dominion dry earth evening every face female fifth fill finished firmament first fish fly for form forth fourth fowl from fruit fruitful gathered gathering give given god good grass great greater green had hath have he heaven heavens herb him his host i image in is it itself kind land lesser let life light lights likeness living made make male man may meat midst morning moved moveth moving multiply night of one open our over own place replenish rule said saw saying sea seas seasons second seed set shall signs sixth so spirit stars subdue that the their them there thing third thus to together tree two under unto upon us very void was waters were whales wherein which whose winged without years yielding you b0185 b2316 b1094 b4187 b2697 b1731 b2097 b2496 b4131 b1000 b0882 b0437 b0438b1120 b3556 b1557 b1823 b0880 b0623 b3162 b3702 b0184 b0015 b0014 b0215 b2779 b1288 b0180 b2925 b3895 b0684 b3463 b0095 b3517 b1493 b3181 b3406 b2614 b2231 b3699 b0059 b4172 b1099 b4259 b4372 b1842 b2526 b3931 b3932 b4000 b0440 b2728 b2729 b3687 b1718 b2530 b2529 b1215 b0186 b0179 b2890 b4129 b3418 b2942 b4143 b4142 b3251 b0924 b0923 b3189 b1224 b1225 b1226 b1467 b3169 b3982 b0133 b3019 b1127 b0903 b3164 b3863 b4226 b1207 b2585 b3725 b2476 b1854 b2892 b3822 b3619 b2038 b3780 b1084 b0214 b3704 b3319 b3295b3987 b3988 b3067 b3461 b3202 b2741 b3649 b0098 b3609 b3590 b2620 b3650 b2576 b4059 b3115 b0406 b1274 b1763 b3919 b0170 b3339 b3980 b2319 b1913 b4179 b0101 b0119 b0528 b0607 b0660 b0877 b0948 b1055 b1086 b1103 b1269 b2330 b2517 b2579 b2581 b2830 b2960 b3180 b3470 b3746 b4191 b0661 b2836 b2323 b4041 b2563 b1095 b1093 b3730 b0736 b3888 b0492 b3701 b3783 b3317 b4203 b3315 b3314 b0169 b3341 b1468 b2592 b0990 b3298 b3320 b3308 b3231 b0911 b3296 b3984b3303 b3309 b3321 b0023 b1552 b3054 b2606 b3186 b4200 b3306 b3230 b3307 b3165 b2609 b4202 b0470 b0640 b2699 b0595 b1294 b2610 b3305 b2515 b2311 b3340 b3247 b1413 b0439 b0436 b0178 b1808 b3935 b3417 b2487 b2721 b3686 b3986 b2096 b0922 b3168 b0480 b2011 b3301 b1716 b2372 b4051 b1817 b0925 b3637 b3652 b3294 b3499 b3304 b3602 b3318 b2804 b3342 AddHealth friendships: friendships among th-graders Word neighbors in Genesis: neighboring occurrences among 158 words Protein binding interactions: binding patterns among 230 proteins 8/39
51 Homophily and stochastic equivalence in real networks, ; :. a above abundantly after air all also and appear be bearing beast beginning behold blessed bring brought called cattle created creature creepeth creeping darkness day days deep divide divided dominion dry earth evening every face female fifth fill finished firmament first fish fly for form forth fourth fowl from fruit fruitful gathered gathering give given god good grass great greater green had hath have he heaven heavens herb him his host i image in is it itself kind land lesser let life light lights likeness living made make male man may meat midst morning moved moveth moving multiply night of one open our over own place replenish rule said saw saying sea seas seasons second seed set shall signs sixth so spirit stars subdue that the their them there thing third thus to together tree two under unto upon us very void was waters were whales wherein which whose winged without years yielding you b0185 b2316 b1094 b4187 b2697 b1731 b2097 b2496 b4131 b1000 b0882 b0437 b0438b1120 b3556 b1557 b1823 b0880 b0623 b3162 b3702 b0184 b0015 b0014 b0215 b2779 b1288 b0180 b2925 b3895 b0684 b3463 b0095 b3517 b1493 b3181 b3406 b2614 b2231 b3699 b0059 b4172 b1099 b4259 b4372 b1842 b2526 b3931 b3932 b4000 b0440 b2728 b2729 b3687 b1718 b2530 b2529 b1215 b0186 b0179 b2890 b4129 b3418 b2942 b4143 b4142 b3251 b0924 b0923 b3189 b1224 b1225 b1226 b1467 b3169 b3982 b0133 b3019 b1127 b0903 b3164 b3863 b4226 b1207 b2585 b3725 b2476 b1854 b2892 b3822 b3619 b2038 b3780 b1084 b0214 b3704 b3319 b3295b3987 b3988 b3067 b3461 b3202 b2741 b3649 b0098 b3609 b3590 b2620 b3650 b2576 b4059 b3115 b0406 b1274 b1763 b3919 b0170 b3339 b3980 b2319 b1913 b4179 b0101 b0119 b0528 b0607 b0660 b0877 b0948 b1055 b1086 b1103 b1269 b2330 b2517 b2579 b2581 b2830 b2960 b3180 b3470 b3746 b4191 b0661 b2836 b2323 b4041 b2563 b1095 b1093 b3730 b0736 b3888 b0492 b3701 b3783 b3317 b4203 b3315 b3314 b0169 b3341 b1468 b2592 b0990 b3298 b3320 b3308 b3231 b0911 b3296 b3984b3303 b3309 b3321 b0023 b1552 b3054 b2606 b3186 b4200 b3306 b3230 b3307 b3165 b2609 b4202 b0470 b0640 b2699 b0595 b1294 b2610 b3305 b2515 b2311 b3340 b3247 b1413 b0439 b0436 b0178 b1808 b3935 b3417 b2487 b2721 b3686 b3986 b2096 b0922 b3168 b0480 b2011 b3301 b1716 b2372 b4051 b1817 b0925 b3637 b3652 b3294 b3499 b3304 b3602 b3318 b2804 b3342 AddHealth friendships: friendships among th-graders Word neighbors in Genesis: neighboring occurrences among 158 words Protein binding interactions: binding patterns among 230 proteins 8/39
52 Homophily and stochastic equivalence in real networks, ; :. a above abundantly after air all also and appear be bearing beast beginning behold blessed bring brought called cattle created creature creepeth creeping darkness day days deep divide divided dominion dry earth evening every face female fifth fill finished firmament first fish fly for form forth fourth fowl from fruit fruitful gathered gathering give given god good grass great greater green had hath have he heaven heavens herb him his host i image in is it itself kind land lesser let life light lights likeness living made make male man may meat midst morning moved moveth moving multiply night of one open our over own place replenish rule said saw saying sea seas seasons second seed set shall signs sixth so spirit stars subdue that the their them there thing third thus to together tree two under unto upon us very void was waters were whales wherein which whose winged without years yielding you b0185 b2316 b1094 b4187 b2697 b1731 b2097 b2496 b4131 b1000 b0882 b0437 b0438b1120 b3556 b1557 b1823 b0880 b0623 b3162 b3702 b0184 b0015 b0014 b0215 b2779 b1288 b0180 b2925 b3895 b0684 b3463 b0095 b3517 b1493 b3181 b3406 b2614 b2231 b3699 b0059 b4172 b1099 b4259 b4372 b1842 b2526 b3931 b3932 b4000 b0440 b2728 b2729 b3687 b1718 b2530 b2529 b1215 b0186 b0179 b2890 b4129 b3418 b2942 b4143 b4142 b3251 b0924 b0923 b3189 b1224 b1225 b1226 b1467 b3169 b3982 b0133 b3019 b1127 b0903 b3164 b3863 b4226 b1207 b2585 b3725 b2476 b1854 b2892 b3822 b3619 b2038 b3780 b1084 b0214 b3704 b3319 b3295b3987 b3988 b3067 b3461 b3202 b2741 b3649 b0098 b3609 b3590 b2620 b3650 b2576 b4059 b3115 b0406 b1274 b1763 b3919 b0170 b3339 b3980 b2319 b1913 b4179 b0101 b0119 b0528 b0607 b0660 b0877 b0948 b1055 b1086 b1103 b1269 b2330 b2517 b2579 b2581 b2830 b2960 b3180 b3470 b3746 b4191 b0661 b2836 b2323 b4041 b2563 b1095 b1093 b3730 b0736 b3888 b0492 b3701 b3783 b3317 b4203 b3315 b3314 b0169 b3341 b1468 b2592 b0990 b3298 b3320 b3308 b3231 b0911 b3296 b3984b3303 b3309 b3321 b0023 b1552 b3054 b2606 b3186 b4200 b3306 b3230 b3307 b3165 b2609 b4202 b0470 b0640 b2699 b0595 b1294 b2610 b3305 b2515 b2311 b3340 b3247 b1413 b0439 b0436 b0178 b1808 b3935 b3417 b2487 b2721 b3686 b3986 b2096 b0922 b3168 b0480 b2011 b3301 b1716 b2372 b4051 b1817 b0925 b3637 b3652 b3294 b3499 b3304 b3602 b3318 b2804 b3342 AddHealth friendships: friendships among th-graders Word neighbors in Genesis: neighboring occurrences among 158 words Protein binding interactions: binding patterns among 230 proteins 8/39
53 19/39 Latent factor models Homophily and stochastic equivalence from unobserved variables can be represented with a latent factor model: y i,j u T i Dv j, u i is a vector of latent factors describing i as a sender of ties; v j is a vector of latent factors describing j as a receiver of ties; D is a diagonal matrix of factor weights. Normal, binomial, ordinal data can be represented with this structure as follows: z i,j = u T i Dv j + ɛ i,j y i,j = g(z i,j ) where g is some increasing function: g(z) = z for normal data; g(z) = 1(z > 0) for binomial data; g(z) = some increasing step function for ordinal data.
54 19/39 Latent factor models Homophily and stochastic equivalence from unobserved variables can be represented with a latent factor model: y i,j u T i Dv j, u i is a vector of latent factors describing i as a sender of ties; v j is a vector of latent factors describing j as a receiver of ties; D is a diagonal matrix of factor weights. Normal, binomial, ordinal data can be represented with this structure as follows: z i,j = u T i Dv j + ɛ i,j y i,j = g(z i,j ) where g is some increasing function: g(z) = z for normal data; g(z) = 1(z > 0) for binomial data; g(z) = some increasing step function for ordinal data.
55 9/39 Latent factor models Homophily and stochastic equivalence from unobserved variables can be represented with a latent factor model: y i,j u T i Dv j, u i is a vector of latent factors describing i as a sender of ties; v j is a vector of latent factors describing j as a receiver of ties; D is a diagonal matrix of factor weights. Normal, binomial, ordinal data can be represented with this structure as follows: z i,j = u T i Dv j + ɛ i,j y i,j = g(z i,j ) where g is some increasing function: g(z) = z for normal data; g(z) = 1(z > 0) for binomial data; g(z) = some increasing step function for ordinal data.
56 9/39 Latent factor models Homophily and stochastic equivalence from unobserved variables can be represented with a latent factor model: y i,j u T i Dv j, u i is a vector of latent factors describing i as a sender of ties; v j is a vector of latent factors describing j as a receiver of ties; D is a diagonal matrix of factor weights. Normal, binomial, ordinal data can be represented with this structure as follows: z i,j = u T i Dv j + ɛ i,j y i,j = g(z i,j ) where g is some increasing function: g(z) = z for normal data; g(z) = 1(z > 0) for binomial data; g(z) = some increasing step function for ordinal data.
57 9/39 Latent factor models Homophily and stochastic equivalence from unobserved variables can be represented with a latent factor model: y i,j u T i Dv j, u i is a vector of latent factors describing i as a sender of ties; v j is a vector of latent factors describing j as a receiver of ties; D is a diagonal matrix of factor weights. Normal, binomial, ordinal data can be represented with this structure as follows: z i,j = u T i Dv j + ɛ i,j y i,j = g(z i,j ) where g is some increasing function: g(z) = z for normal data; g(z) = 1(z > 0) for binomial data; g(z) = some increasing step function for ordinal data.
58 9/39 Latent factor models Homophily and stochastic equivalence from unobserved variables can be represented with a latent factor model: y i,j u T i Dv j, u i is a vector of latent factors describing i as a sender of ties; v j is a vector of latent factors describing j as a receiver of ties; D is a diagonal matrix of factor weights. Normal, binomial, ordinal data can be represented with this structure as follows: z i,j = u T i Dv j + ɛ i,j y i,j = g(z i,j ) where g is some increasing function: g(z) = z for normal data; g(z) = 1(z > 0) for binomial data; g(z) = some increasing step function for ordinal data.
59 19/39 Latent factor models Homophily and stochastic equivalence from unobserved variables can be represented with a latent factor model: y i,j u T i Dv j, u i is a vector of latent factors describing i as a sender of ties; v j is a vector of latent factors describing j as a receiver of ties; D is a diagonal matrix of factor weights. Normal, binomial, ordinal data can be represented with this structure as follows: z i,j = u T i Dv j + ɛ i,j y i,j = g(z i,j ) where g is some increasing function: g(z) = z for normal data; g(z) = 1(z > 0) for binomial data; g(z) = some increasing step function for ordinal data.
60 19/39 Latent factor models Homophily and stochastic equivalence from unobserved variables can be represented with a latent factor model: y i,j u T i Dv j, u i is a vector of latent factors describing i as a sender of ties; v j is a vector of latent factors describing j as a receiver of ties; D is a diagonal matrix of factor weights. Normal, binomial, ordinal data can be represented with this structure as follows: z i,j = u T i Dv j + ɛ i,j y i,j = g(z i,j ) where g is some increasing function: g(z) = z for normal data; g(z) = 1(z > 0) for binomial data; g(z) = some increasing step function for ordinal data.
61 19/39 Latent factor models Homophily and stochastic equivalence from unobserved variables can be represented with a latent factor model: y i,j u T i Dv j, u i is a vector of latent factors describing i as a sender of ties; v j is a vector of latent factors describing j as a receiver of ties; D is a diagonal matrix of factor weights. Normal, binomial, ordinal data can be represented with this structure as follows: z i,j = u T i Dv j + ɛ i,j y i,j = g(z i,j ) where g is some increasing function: g(z) = z for normal data; g(z) = 1(z > 0) for binomial data; g(z) = some increasing step function for ordinal data.
62 20/39 Understanding latent factors Z = U T DV + E z i,j = u T i Dv j + ɛ i,j R = d r u i,r v j,r + ɛ i,j For example, in a 2 factor model, we have z i,j = d 1(u i,1 v j,1 ) + d 2(u i,2 v j,2 ) + ɛ i,j r=1 Interpretation u i u j : similarity of latent factors implies approximate stoch equivalence; u i v j : similarity of latent factors implies high probability of a tie.
63 20/39 Understanding latent factors Z = U T DV + E z i,j = u T i Dv j + ɛ i,j R = d r u i,r v j,r + ɛ i,j For example, in a 2 factor model, we have z i,j = d 1(u i,1 v j,1 ) + d 2(u i,2 v j,2 ) + ɛ i,j r=1 Interpretation u i u j : similarity of latent factors implies approximate stoch equivalence; u i v j : similarity of latent factors implies high probability of a tie.
64 20/39 Understanding latent factors Z = U T DV + E z i,j = u T i Dv j + ɛ i,j R = d r u i,r v j,r + ɛ i,j For example, in a 2 factor model, we have z i,j = d 1(u i,1 v j,1 ) + d 2(u i,2 v j,2 ) + ɛ i,j r=1 Interpretation u i u j : similarity of latent factors implies approximate stoch equivalence; u i v j : similarity of latent factors implies high probability of a tie.
65 20/39 Understanding latent factors Z = U T DV + E z i,j = u T i Dv j + ɛ i,j R = d r u i,r v j,r + ɛ i,j For example, in a 2 factor model, we have z i,j = d 1(u i,1 v j,1 ) + d 2(u i,2 v j,2 ) + ɛ i,j r=1 Interpretation u i u j : similarity of latent factors implies approximate stoch equivalence; u i v j : similarity of latent factors implies high probability of a tie.
66 20/39 Understanding latent factors Z = U T DV + E z i,j = u T i Dv j + ɛ i,j R = d r u i,r v j,r + ɛ i,j For example, in a 2 factor model, we have z i,j = d 1(u i,1 v j,1 ) + d 2(u i,2 v j,2 ) + ɛ i,j r=1 Interpretation u i u j : similarity of latent factors implies approximate stoch equivalence; u i v j : similarity of latent factors implies high probability of a tie.
67 20/39 Understanding latent factors Z = U T DV + E z i,j = u T i Dv j + ɛ i,j R = d r u i,r v j,r + ɛ i,j For example, in a 2 factor model, we have z i,j = d 1(u i,1 v j,1 ) + d 2(u i,2 v j,2 ) + ɛ i,j r=1 Interpretation u i u j : similarity of latent factors implies approximate stoch equivalence; u i v j : similarity of latent factors implies high probability of a tie.
68 20/39 Understanding latent factors Z = U T DV + E z i,j = u T i Dv j + ɛ i,j R = d r u i,r v j,r + ɛ i,j For example, in a 2 factor model, we have z i,j = d 1(u i,1 v j,1 ) + d 2(u i,2 v j,2 ) + ɛ i,j r=1 Interpretation u i u j : similarity of latent factors implies approximate stoch equivalence; u i v j : similarity of latent factors implies high probability of a tie.
69 1/39 Matrix decomposition interpretation Recall from linear algebra: Every m n matrix Z can be written Z = UDV T where D = diag(d 1,..., d n), U and V are orthonormal. If UDV T is the svd of Z, then Ẑ k U [,1:k] D [1:k,1:k] V T [,1:k] is the least-squares rank-k approximation to Z.
70 1/39 Matrix decomposition interpretation Recall from linear algebra: Every m n matrix Z can be written Z = UDV T where D = diag(d 1,..., d n), U and V are orthonormal. If UDV T is the svd of Z, then Ẑ k U [,1:k] D [1:k,1:k] V T [,1:k] is the least-squares rank-k approximation to Z.
71 21/39 Matrix decomposition interpretation Recall from linear algebra: Every m n matrix Z can be written Z = UDV T where D = diag(d 1,..., d n), U and V are orthonormal. If UDV T is the svd of Z, then Ẑ k U [,1:k] D [1:k,1:k] V T [,1:k] is the least-squares rank-k approximation to Z.
72 22/39 Least squares matrix approximations singular value Z^ r=1 Z^ r=3 Z^ r=6 Z^ r= Z Z Z Z
73 23/39 LFM for symmetric data Probit version of the symmetric latent factor model: y i,j = g(z i,j ), where g is a nondecreasing function z i,j = u T i Λu j + ɛ i,j, where u i R K, Λ=diag(λ 1,..., λ K ) {ɛ i,j } iid normal(0, 1) Writing {z i,j } as a matrix, Recall from linear algebra: Z = UΛU T + E Every n n symmetric matrix Z can be written Z = UΛU T where Λ = diag(λ 1,..., λ n) and U is orthonormal. If UΛU T is the eigendecomposition of Z, then Ẑ k U [,1:k] Λ [1:k,1:k] U T [,1:k] is the least-squares rank-k approximation to Z.
74 23/39 LFM for symmetric data Probit version of the symmetric latent factor model: y i,j = g(z i,j ), where g is a nondecreasing function z i,j = u T i Λu j + ɛ i,j, where u i R K, Λ=diag(λ 1,..., λ K ) {ɛ i,j } iid normal(0, 1) Writing {z i,j } as a matrix, Recall from linear algebra: Z = UΛU T + E Every n n symmetric matrix Z can be written Z = UΛU T where Λ = diag(λ 1,..., λ n) and U is orthonormal. If UΛU T is the eigendecomposition of Z, then Ẑ k U [,1:k] Λ [1:k,1:k] U T [,1:k] is the least-squares rank-k approximation to Z.
75 23/39 LFM for symmetric data Probit version of the symmetric latent factor model: y i,j = g(z i,j ), where g is a nondecreasing function z i,j = u T i Λu j + ɛ i,j, where u i R K, Λ=diag(λ 1,..., λ K ) {ɛ i,j } iid normal(0, 1) Writing {z i,j } as a matrix, Recall from linear algebra: Z = UΛU T + E Every n n symmetric matrix Z can be written Z = UΛU T where Λ = diag(λ 1,..., λ n) and U is orthonormal. If UΛU T is the eigendecomposition of Z, then Ẑ k U [,1:k] Λ [1:k,1:k] U T [,1:k] is the least-squares rank-k approximation to Z.
76 23/39 LFM for symmetric data Probit version of the symmetric latent factor model: y i,j = g(z i,j ), where g is a nondecreasing function z i,j = u T i Λu j + ɛ i,j, where u i R K, Λ=diag(λ 1,..., λ K ) {ɛ i,j } iid normal(0, 1) Writing {z i,j } as a matrix, Recall from linear algebra: Z = UΛU T + E Every n n symmetric matrix Z can be written Z = UΛU T where Λ = diag(λ 1,..., λ n) and U is orthonormal. If UΛU T is the eigendecomposition of Z, then Ẑ k U [,1:k] Λ [1:k,1:k] U T [,1:k] is the least-squares rank-k approximation to Z.
77 23/39 LFM for symmetric data Probit version of the symmetric latent factor model: y i,j = g(z i,j ), where g is a nondecreasing function z i,j = u T i Λu j + ɛ i,j, where u i R K, Λ=diag(λ 1,..., λ K ) {ɛ i,j } iid normal(0, 1) Writing {z i,j } as a matrix, Recall from linear algebra: Z = UΛU T + E Every n n symmetric matrix Z can be written Z = UΛU T where Λ = diag(λ 1,..., λ n) and U is orthonormal. If UΛU T is the eigendecomposition of Z, then Ẑ k U [,1:k] Λ [1:k,1:k] U T [,1:k] is the least-squares rank-k approximation to Z.
78 23/39 LFM for symmetric data Probit version of the symmetric latent factor model: y i,j = g(z i,j ), where g is a nondecreasing function z i,j = u T i Λu j + ɛ i,j, where u i R K, Λ=diag(λ 1,..., λ K ) {ɛ i,j } iid normal(0, 1) Writing {z i,j } as a matrix, Recall from linear algebra: Z = UΛU T + E Every n n symmetric matrix Z can be written Z = UΛU T where Λ = diag(λ 1,..., λ n) and U is orthonormal. If UΛU T is the eigendecomposition of Z, then Ẑ k U [,1:k] Λ [1:k,1:k] U T [,1:k] is the least-squares rank-k approximation to Z.
79 23/39 LFM for symmetric data Probit version of the symmetric latent factor model: y i,j = g(z i,j ), where g is a nondecreasing function z i,j = u T i Λu j + ɛ i,j, where u i R K, Λ=diag(λ 1,..., λ K ) {ɛ i,j } iid normal(0, 1) Writing {z i,j } as a matrix, Recall from linear algebra: Z = UΛU T + E Every n n symmetric matrix Z can be written Z = UΛU T where Λ = diag(λ 1,..., λ n) and U is orthonormal. If UΛU T is the eigendecomposition of Z, then Ẑ k U [,1:k] Λ [1:k,1:k] U T [,1:k] is the least-squares rank-k approximation to Z.
80 24/39 Least squares approximations of increasing rank eigenvalue Index ^ Z Z ^ ^ Z ^ Z range(c(x, y)) range(c(x, y)) range(c(x, y)) range(c(x, y))
81 25/39 Understanding eigenvectors and eigenvalues Z = U T ΛU + E z i,j = u T i Λu j + ɛ i,j R = λ r u i,r u j,r + ɛ i,j For example, in a rank-2 model, we have z i,j = λ 1(u i,1 u j,1 ) + λ 2(u i,2 u j,2 ) + ɛ i,j r=1 Interpretation u i,r u j,r : equality of latent factors; represents stochastic equivalence; λ r > 0: positive eigenvalues represent homophily; λ r < 0: negative eigenvalues represent antihomophily.
82 25/39 Understanding eigenvectors and eigenvalues Z = U T ΛU + E z i,j = u T i Λu j + ɛ i,j R = λ r u i,r u j,r + ɛ i,j For example, in a rank-2 model, we have z i,j = λ 1(u i,1 u j,1 ) + λ 2(u i,2 u j,2 ) + ɛ i,j r=1 Interpretation u i,r u j,r : equality of latent factors; represents stochastic equivalence; λ r > 0: positive eigenvalues represent homophily; λ r < 0: negative eigenvalues represent antihomophily.
83 25/39 Understanding eigenvectors and eigenvalues Z = U T ΛU + E z i,j = u T i Λu j + ɛ i,j R = λ r u i,r u j,r + ɛ i,j For example, in a rank-2 model, we have z i,j = λ 1(u i,1 u j,1 ) + λ 2(u i,2 u j,2 ) + ɛ i,j r=1 Interpretation u i,r u j,r : equality of latent factors; represents stochastic equivalence; λ r > 0: positive eigenvalues represent homophily; λ r < 0: negative eigenvalues represent antihomophily.
84 25/39 Understanding eigenvectors and eigenvalues Z = U T ΛU + E z i,j = u T i Λu j + ɛ i,j R = λ r u i,r u j,r + ɛ i,j For example, in a rank-2 model, we have z i,j = λ 1(u i,1 u j,1 ) + λ 2(u i,2 u j,2 ) + ɛ i,j r=1 Interpretation u i,r u j,r : equality of latent factors; represents stochastic equivalence; λ r > 0: positive eigenvalues represent homophily; λ r < 0: negative eigenvalues represent antihomophily.
85 25/39 Understanding eigenvectors and eigenvalues Z = U T ΛU + E z i,j = u T i Λu j + ɛ i,j R = λ r u i,r u j,r + ɛ i,j For example, in a rank-2 model, we have z i,j = λ 1(u i,1 u j,1 ) + λ 2(u i,2 u j,2 ) + ɛ i,j r=1 Interpretation u i,r u j,r : equality of latent factors; represents stochastic equivalence; λ r > 0: positive eigenvalues represent homophily; λ r < 0: negative eigenvalues represent antihomophily.
86 25/39 Understanding eigenvectors and eigenvalues Z = U T ΛU + E z i,j = u T i Λu j + ɛ i,j R = λ r u i,r u j,r + ɛ i,j For example, in a rank-2 model, we have z i,j = λ 1(u i,1 u j,1 ) + λ 2(u i,2 u j,2 ) + ɛ i,j r=1 Interpretation u i,r u j,r : equality of latent factors; represents stochastic equivalence; λ r > 0: positive eigenvalues represent homophily; λ r < 0: negative eigenvalues represent antihomophily.
87 25/39 Understanding eigenvectors and eigenvalues Z = U T ΛU + E z i,j = u T i Λu j + ɛ i,j R = λ r u i,r u j,r + ɛ i,j For example, in a rank-2 model, we have z i,j = λ 1(u i,1 u j,1 ) + λ 2(u i,2 u j,2 ) + ɛ i,j r=1 Interpretation u i,r u j,r : equality of latent factors; represents stochastic equivalence; λ r > 0: positive eigenvalues represent homophily; λ r < 0: negative eigenvalues represent antihomophily.
88 25/39 Understanding eigenvectors and eigenvalues Z = U T ΛU + E z i,j = u T i Λu j + ɛ i,j R = λ r u i,r u j,r + ɛ i,j For example, in a rank-2 model, we have z i,j = λ 1(u i,1 u j,1 ) + λ 2(u i,2 u j,2 ) + ɛ i,j r=1 Interpretation u i,r u j,r : equality of latent factors; represents stochastic equivalence; λ r > 0: positive eigenvalues represent homophily; λ r < 0: negative eigenvalues represent antihomophily.
89 25/39 Understanding eigenvectors and eigenvalues Z = U T ΛU + E z i,j = u T i Λu j + ɛ i,j R = λ r u i,r u j,r + ɛ i,j For example, in a rank-2 model, we have z i,j = λ 1(u i,1 u j,1 ) + λ 2(u i,2 u j,2 ) + ɛ i,j r=1 Interpretation u i,r u j,r : equality of latent factors; represents stochastic equivalence; λ r > 0: positive eigenvalues represent homophily; λ r < 0: negative eigenvalues represent antihomophily.
90 25/39 Understanding eigenvectors and eigenvalues Z = U T ΛU + E z i,j = u T i Λu j + ɛ i,j R = λ r u i,r u j,r + ɛ i,j For example, in a rank-2 model, we have z i,j = λ 1(u i,1 u j,1 ) + λ 2(u i,2 u j,2 ) + ɛ i,j r=1 Interpretation u i,r u j,r : equality of latent factors; represents stochastic equivalence; λ r > 0: positive eigenvalues represent homophily; λ r < 0: negative eigenvalues represent antihomophily.
91 26/39 Returning to directed relations: AME models SRM: We have motivated the SRM in order to represent 2nd order dependence: within row dependence, within column dependence; within dyad dependence. z i,j = β T x i,j + a i + b j + ɛ i,j y i,j = g(z i,j ) This model is made up of additive random effects. LFM: We have motivated the LFM to model more complex structures: third order dependence and transitivity; stochastic equivalence. z i,j = u T i Dv j + ɛ i,j y i,j = g(z i,j ) This model is made up of multiplicative random effects.
Latent Factor Models for Relational Data
 Latent Factor Models for Relational Data Peter Hoff Statistics, Biostatistics and Center for Statistics and the Social Sciences University of Washington Outline Part 1: Multiplicative factor models for
Latent Factor Models for Relational Data Peter Hoff Statistics, Biostatistics and Center for Statistics and the Social Sciences University of Washington Outline Part 1: Multiplicative factor models for
Latent SVD Models for Relational Data
 Latent SVD Models for Relational Data Peter Hoff Statistics, iostatistics and enter for Statistics and the Social Sciences University of Washington 1 Outline 1 Relational data 2 Exchangeability for arrays
Latent SVD Models for Relational Data Peter Hoff Statistics, iostatistics and enter for Statistics and the Social Sciences University of Washington 1 Outline 1 Relational data 2 Exchangeability for arrays
4. "God called the light (BRIGHT, DAY), and the darkness He called (NIGHT, SPACE). So the evening and the morning were the first day.
 MEMORY VERSE: "In the beginning was the Word, and the Word was with God, and the Word was God. He was in the beginning with God. All things were made through Him, and without Him nothing was made that
MEMORY VERSE: "In the beginning was the Word, and the Word was with God, and the Word was God. He was in the beginning with God. All things were made through Him, and without Him nothing was made that
Hierarchical models for multiway data
 Hierarchical models for multiway data Peter Hoff Statistics, Biostatistics and the CSSS University of Washington Array-valued data y i,j,k = jth variable of ith subject under condition k (psychometrics).
Hierarchical models for multiway data Peter Hoff Statistics, Biostatistics and the CSSS University of Washington Array-valued data y i,j,k = jth variable of ith subject under condition k (psychometrics).
Mean and covariance models for relational arrays
 Mean and covariance models for relational arrays Peter Hoff Statistics, Biostatistics and the CSSS University of Washington Outline Introduction and examples Models for multiway mean structure Models for
Mean and covariance models for relational arrays Peter Hoff Statistics, Biostatistics and the CSSS University of Washington Outline Introduction and examples Models for multiway mean structure Models for
Modeling homophily and stochastic equivalence in symmetric relational data
 Modeling homophily and stochastic equivalence in symmetric relational data Peter D. Hoff Departments of Statistics and Biostatistics University of Washington Seattle, WA 98195-4322. hoff@stat.washington.edu
Modeling homophily and stochastic equivalence in symmetric relational data Peter D. Hoff Departments of Statistics and Biostatistics University of Washington Seattle, WA 98195-4322. hoff@stat.washington.edu
Probability models for multiway data
 Probability models for multiway data Peter Hoff Statistics, Biostatistics and the CSSS University of Washington Outline Introduction and examples Hierarchical models for multiway factors Deep interactions
Probability models for multiway data Peter Hoff Statistics, Biostatistics and the CSSS University of Washington Outline Introduction and examples Hierarchical models for multiway factors Deep interactions
Dyadic data analysis with amen
 Dyadic data analysis with amen Peter D. Hoff May 23, 2017 Abstract Dyadic data on pairs of objects, such as relational or social network data, often exhibit strong statistical dependencies. Certain types
Dyadic data analysis with amen Peter D. Hoff May 23, 2017 Abstract Dyadic data on pairs of objects, such as relational or social network data, often exhibit strong statistical dependencies. Certain types
OT004. Genesis 1:26-2:3
 OT004 Genesis 1:26-2:3 Calvary Curriculum s CHILDREN S CURRICULUM - OT004 Copyright 2016 Calvary Curriculum. All rights reserved. You may not, under any circumstances, sell, distribute, or commercially
OT004 Genesis 1:26-2:3 Calvary Curriculum s CHILDREN S CURRICULUM - OT004 Copyright 2016 Calvary Curriculum. All rights reserved. You may not, under any circumstances, sell, distribute, or commercially
OT002. Genesis 1:14-25
 OT002 Genesis 1:14-25 Calvary Curriculum s CHILDREN S CURRICULUM - OT002 Copyright 2016 Calvary Curriculum. All rights reserved. You may not, under any circumstances, sell, distribute, or commercially
OT002 Genesis 1:14-25 Calvary Curriculum s CHILDREN S CURRICULUM - OT002 Copyright 2016 Calvary Curriculum. All rights reserved. You may not, under any circumstances, sell, distribute, or commercially
fish birds stars cattle mountains creeping things
 MEMORY VERSE: "So God created man in His own image; in the image of God He created him; male and female He created them." GENESIS 1:27 CIRCLE THE CORRECT WORDS: 1. "Then God said, 'Let Us make man in Our
MEMORY VERSE: "So God created man in His own image; in the image of God He created him; male and female He created them." GENESIS 1:27 CIRCLE THE CORRECT WORDS: 1. "Then God said, 'Let Us make man in Our
Bible Story 2 GOD MAKES ADAM & EVE GENESIS 1:26-4:1
 Bible Story 2 GOD MAKES ADAM & EVE GENESIS 1:26-4:1 "So God created man in His own image; in the image of God He created him; male and female He created them." GENESIS 1:27 MEMORY VERSE: "So God created
Bible Story 2 GOD MAKES ADAM & EVE GENESIS 1:26-4:1 "So God created man in His own image; in the image of God He created him; male and female He created them." GENESIS 1:27 MEMORY VERSE: "So God created
The International-Trade Network: Gravity Equations and Topological Properties
 The International-Trade Network: Gravity Equations and Topological Properties Giorgio Fagiolo Sant Anna School of Advanced Studies Laboratory of Economics and Management Piazza Martiri della Libertà 33
The International-Trade Network: Gravity Equations and Topological Properties Giorgio Fagiolo Sant Anna School of Advanced Studies Laboratory of Economics and Management Piazza Martiri della Libertà 33
Inferring Latent Preferences from Network Data
 Inferring Latent Preferences from Network John S. Ahlquist 1 Arturas 2 1 UC San Diego GPS 2 NYU 14 November 2015 very early stages Methodological extend latent space models (Hoff et al 2002) to partial
Inferring Latent Preferences from Network John S. Ahlquist 1 Arturas 2 1 UC San Diego GPS 2 NYU 14 November 2015 very early stages Methodological extend latent space models (Hoff et al 2002) to partial
Modeling networks: regression with additive and multiplicative effects
 Modeling networks: regression with additive and multiplicative effects Alexander Volfovsky Department of Statistical Science, Duke May 25 2017 May 25, 2017 Health Networks 1 Why model networks? Interested
Modeling networks: regression with additive and multiplicative effects Alexander Volfovsky Department of Statistical Science, Duke May 25 2017 May 25, 2017 Health Networks 1 Why model networks? Interested
Sampling and incomplete network data
 1/58 Sampling and incomplete network data 567 Statistical analysis of social networks Peter Hoff Statistics, University of Washington 2/58 Network sampling methods It is sometimes difficult to obtain a
1/58 Sampling and incomplete network data 567 Statistical analysis of social networks Peter Hoff Statistics, University of Washington 2/58 Network sampling methods It is sometimes difficult to obtain a
Creation Genesis 1:1-2:25
 Lesson 001 Creation Genesis 1:1-2:25 MEMORY VERSE John 1:1-3 In the beginning was the Word, and the Word was with God, and the Word w as God. He w as in the beginning w ith God. All things w ere m ade
Lesson 001 Creation Genesis 1:1-2:25 MEMORY VERSE John 1:1-3 In the beginning was the Word, and the Word was with God, and the Word w as God. He w as in the beginning w ith God. All things w ere m ade
Quick Tour of Linear Algebra and Graph Theory
 Quick Tour of Linear Algebra and Graph Theory CS224w: Social and Information Network Analysis Fall 2012 Yu Wayne Wu Based on Borja Pelato s version in Fall 2011 Matrices and Vectors Matrix: A rectangular
Quick Tour of Linear Algebra and Graph Theory CS224w: Social and Information Network Analysis Fall 2012 Yu Wayne Wu Based on Borja Pelato s version in Fall 2011 Matrices and Vectors Matrix: A rectangular
And I looked, and I heard an angel flying through the midst of heaven, saying with a loud voice, Woe, woe, woe to the inhabitants of the earth,
 And I looked, and I heard an angel flying through the midst of heaven, saying with a loud voice, Woe, woe, woe to the inhabitants of the earth, because of the remaining blasts of the trumpet of the three
And I looked, and I heard an angel flying through the midst of heaven, saying with a loud voice, Woe, woe, woe to the inhabitants of the earth, because of the remaining blasts of the trumpet of the three
Linear Algebra (Review) Volker Tresp 2018
 Linear Algebra (Review) Volker Tresp 2018 1 Vectors k, M, N are scalars A one-dimensional array c is a column vector. Thus in two dimensions, ( ) c1 c = c 2 c i is the i-th component of c c T = (c 1, c
Linear Algebra (Review) Volker Tresp 2018 1 Vectors k, M, N are scalars A one-dimensional array c is a column vector. Thus in two dimensions, ( ) c1 c = c 2 c i is the i-th component of c c T = (c 1, c
Review of Linear Algebra
 Review of Linear Algebra Dr Gerhard Roth COMP 40A Winter 05 Version Linear algebra Is an important area of mathematics It is the basis of computer vision Is very widely taught, and there are many resources
Review of Linear Algebra Dr Gerhard Roth COMP 40A Winter 05 Version Linear algebra Is an important area of mathematics It is the basis of computer vision Is very widely taught, and there are many resources
Lecture Note 13 The Gains from International Trade: Empirical Evidence Using the Method of Instrumental Variables
 Lecture Note 13 The Gains from International Trade: Empirical Evidence Using the Method of Instrumental Variables David Autor, MIT and NBER 14.03/14.003 Microeconomic Theory and Public Policy, Fall 2016
Lecture Note 13 The Gains from International Trade: Empirical Evidence Using the Method of Instrumental Variables David Autor, MIT and NBER 14.03/14.003 Microeconomic Theory and Public Policy, Fall 2016
14 Singular Value Decomposition
 14 Singular Value Decomposition For any high-dimensional data analysis, one s first thought should often be: can I use an SVD? The singular value decomposition is an invaluable analysis tool for dealing
14 Singular Value Decomposition For any high-dimensional data analysis, one s first thought should often be: can I use an SVD? The singular value decomposition is an invaluable analysis tool for dealing
Mathematical Optimisation, Chpt 2: Linear Equations and inequalities
 Mathematical Optimisation, Chpt 2: Linear Equations and inequalities Peter J.C. Dickinson p.j.c.dickinson@utwente.nl http://dickinson.website version: 12/02/18 Monday 5th February 2018 Peter J.C. Dickinson
Mathematical Optimisation, Chpt 2: Linear Equations and inequalities Peter J.C. Dickinson p.j.c.dickinson@utwente.nl http://dickinson.website version: 12/02/18 Monday 5th February 2018 Peter J.C. Dickinson
Statistics 202: Data Mining. c Jonathan Taylor. Week 2 Based in part on slides from textbook, slides of Susan Holmes. October 3, / 1
 Week 2 Based in part on slides from textbook, slides of Susan Holmes October 3, 2012 1 / 1 Part I Other datatypes, preprocessing 2 / 1 Other datatypes Document data You might start with a collection of
Week 2 Based in part on slides from textbook, slides of Susan Holmes October 3, 2012 1 / 1 Part I Other datatypes, preprocessing 2 / 1 Other datatypes Document data You might start with a collection of
Part I. Other datatypes, preprocessing. Other datatypes. Other datatypes. Week 2 Based in part on slides from textbook, slides of Susan Holmes
 Week 2 Based in part on slides from textbook, slides of Susan Holmes Part I Other datatypes, preprocessing October 3, 2012 1 / 1 2 / 1 Other datatypes Other datatypes Document data You might start with
Week 2 Based in part on slides from textbook, slides of Susan Holmes Part I Other datatypes, preprocessing October 3, 2012 1 / 1 2 / 1 Other datatypes Other datatypes Document data You might start with
Quick Tour of Linear Algebra and Graph Theory
 Quick Tour of Linear Algebra and Graph Theory CS224W: Social and Information Network Analysis Fall 2014 David Hallac Based on Peter Lofgren, Yu Wayne Wu, and Borja Pelato s previous versions Matrices and
Quick Tour of Linear Algebra and Graph Theory CS224W: Social and Information Network Analysis Fall 2014 David Hallac Based on Peter Lofgren, Yu Wayne Wu, and Borja Pelato s previous versions Matrices and
Lecture 10 Optimal Growth Endogenous Growth. Noah Williams
 Lecture 10 Optimal Growth Endogenous Growth Noah Williams University of Wisconsin - Madison Economics 702 Spring 2018 Optimal Growth Path Recall we assume exogenous growth in population and productivity:
Lecture 10 Optimal Growth Endogenous Growth Noah Williams University of Wisconsin - Madison Economics 702 Spring 2018 Optimal Growth Path Recall we assume exogenous growth in population and productivity:
Christ and the Geological Periods
 Christ and the Geological Periods There are uncertainties associated in any discussion of geological periods. The most obvious uncertainty is that there were no observers during these geological periods
Christ and the Geological Periods There are uncertainties associated in any discussion of geological periods. The most obvious uncertainty is that there were no observers during these geological periods
Properties of Matrices and Operations on Matrices
 Properties of Matrices and Operations on Matrices A common data structure for statistical analysis is a rectangular array or matris. Rows represent individual observational units, or just observations,
Properties of Matrices and Operations on Matrices A common data structure for statistical analysis is a rectangular array or matris. Rows represent individual observational units, or just observations,
CS 143 Linear Algebra Review
 CS 143 Linear Algebra Review Stefan Roth September 29, 2003 Introductory Remarks This review does not aim at mathematical rigor very much, but instead at ease of understanding and conciseness. Please see
CS 143 Linear Algebra Review Stefan Roth September 29, 2003 Introductory Remarks This review does not aim at mathematical rigor very much, but instead at ease of understanding and conciseness. Please see
PCA and admixture models
 PCA and admixture models CM226: Machine Learning for Bioinformatics. Fall 2016 Sriram Sankararaman Acknowledgments: Fei Sha, Ameet Talwalkar, Alkes Price PCA and admixture models 1 / 57 Announcements HW1
PCA and admixture models CM226: Machine Learning for Bioinformatics. Fall 2016 Sriram Sankararaman Acknowledgments: Fei Sha, Ameet Talwalkar, Alkes Price PCA and admixture models 1 / 57 Announcements HW1
Random Effects Models for Network Data
 Random Effects Models for Network Data Peter D. Hoff 1 Working Paper no. 28 Center for Statistics and the Social Sciences University of Washington Seattle, WA 98195-4320 January 14, 2003 1 Department of
Random Effects Models for Network Data Peter D. Hoff 1 Working Paper no. 28 Center for Statistics and the Social Sciences University of Washington Seattle, WA 98195-4320 January 14, 2003 1 Department of
forms Christopher Engström November 14, 2014 MAA704: Matrix factorization and canonical forms Matrix properties Matrix factorization Canonical forms
 Christopher Engström November 14, 2014 Hermitian LU QR echelon Contents of todays lecture Some interesting / useful / important of matrices Hermitian LU QR echelon Rewriting a as a product of several matrices.
Christopher Engström November 14, 2014 Hermitian LU QR echelon Contents of todays lecture Some interesting / useful / important of matrices Hermitian LU QR echelon Rewriting a as a product of several matrices.
Clustering and blockmodeling
 ISEG Technical University of Lisbon Introductory Workshop to Network Analysis of Texts Clustering and blockmodeling Vladimir Batagelj University of Ljubljana Lisbon, Portugal: 2nd to 5th February 2004
ISEG Technical University of Lisbon Introductory Workshop to Network Analysis of Texts Clustering and blockmodeling Vladimir Batagelj University of Ljubljana Lisbon, Portugal: 2nd to 5th February 2004
Agriculture, Transportation and the Timing of Urbanization
 Agriculture, Transportation and the Timing of Urbanization Global Analysis at the Grid Cell Level Mesbah Motamed Raymond Florax William Masters Department of Agricultural Economics Purdue University SHaPE
Agriculture, Transportation and the Timing of Urbanization Global Analysis at the Grid Cell Level Mesbah Motamed Raymond Florax William Masters Department of Agricultural Economics Purdue University SHaPE
Stat 206: Linear algebra
 Stat 206: Linear algebra James Johndrow (adapted from Iain Johnstone s notes) 2016-11-02 Vectors We have already been working with vectors, but let s review a few more concepts. The inner product of two
Stat 206: Linear algebra James Johndrow (adapted from Iain Johnstone s notes) 2016-11-02 Vectors We have already been working with vectors, but let s review a few more concepts. The inner product of two
Foundations of Computer Vision
 Foundations of Computer Vision Wesley. E. Snyder North Carolina State University Hairong Qi University of Tennessee, Knoxville Last Edited February 8, 2017 1 3.2. A BRIEF REVIEW OF LINEAR ALGEBRA Apply
Foundations of Computer Vision Wesley. E. Snyder North Carolina State University Hairong Qi University of Tennessee, Knoxville Last Edited February 8, 2017 1 3.2. A BRIEF REVIEW OF LINEAR ALGEBRA Apply
CS 340 Lec. 6: Linear Dimensionality Reduction
 CS 340 Lec. 6: Linear Dimensionality Reduction AD January 2011 AD () January 2011 1 / 46 Linear Dimensionality Reduction Introduction & Motivation Brief Review of Linear Algebra Principal Component Analysis
CS 340 Lec. 6: Linear Dimensionality Reduction AD January 2011 AD () January 2011 1 / 46 Linear Dimensionality Reduction Introduction & Motivation Brief Review of Linear Algebra Principal Component Analysis
ECON 581. The Solow Growth Model, Continued. Instructor: Dmytro Hryshko
 ECON 581. The Solow Growth Model, Continued Instructor: Dmytro Hryshko 1 / 38 The Solow model in continuous time Consider the following (difference) equation x(t + 1) x(t) = g(x(t)), where g( ) is some
ECON 581. The Solow Growth Model, Continued Instructor: Dmytro Hryshko 1 / 38 The Solow model in continuous time Consider the following (difference) equation x(t + 1) x(t) = g(x(t)), where g( ) is some
Dimension reduction, PCA & eigenanalysis Based in part on slides from textbook, slides of Susan Holmes. October 3, Statistics 202: Data Mining
 Dimension reduction, PCA & eigenanalysis Based in part on slides from textbook, slides of Susan Holmes October 3, 2012 1 / 1 Combinations of features Given a data matrix X n p with p fairly large, it can
Dimension reduction, PCA & eigenanalysis Based in part on slides from textbook, slides of Susan Holmes October 3, 2012 1 / 1 Combinations of features Given a data matrix X n p with p fairly large, it can
Linear Algebra Review. Vectors
 Linear Algebra Review 9/4/7 Linear Algebra Review By Tim K. Marks UCSD Borrows heavily from: Jana Kosecka http://cs.gmu.edu/~kosecka/cs682.html Virginia de Sa (UCSD) Cogsci 8F Linear Algebra review Vectors
Linear Algebra Review 9/4/7 Linear Algebra Review By Tim K. Marks UCSD Borrows heavily from: Jana Kosecka http://cs.gmu.edu/~kosecka/cs682.html Virginia de Sa (UCSD) Cogsci 8F Linear Algebra review Vectors
Singular Value Decomposition and Principal Component Analysis (PCA) I
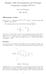 Singular Value Decomposition and Principal Component Analysis (PCA) I Prof Ned Wingreen MOL 40/50 Microarray review Data per array: 0000 genes, I (green) i,i (red) i 000 000+ data points! The expression
Singular Value Decomposition and Principal Component Analysis (PCA) I Prof Ned Wingreen MOL 40/50 Microarray review Data per array: 0000 genes, I (green) i,i (red) i 000 000+ data points! The expression
Landlocked or Policy Locked?
 Landlocked or Policy Locked? How Services Trade Protection Deepens Economic Isolation Ingo Borchert University of Sussex Based on research with Batshur Gootiiz, Arti Grover and Aaditya Mattoo FERDI ITC
Landlocked or Policy Locked? How Services Trade Protection Deepens Economic Isolation Ingo Borchert University of Sussex Based on research with Batshur Gootiiz, Arti Grover and Aaditya Mattoo FERDI ITC
Deep Learning Book Notes Chapter 2: Linear Algebra
 Deep Learning Book Notes Chapter 2: Linear Algebra Compiled By: Abhinaba Bala, Dakshit Agrawal, Mohit Jain Section 2.1: Scalars, Vectors, Matrices and Tensors Scalar Single Number Lowercase names in italic
Deep Learning Book Notes Chapter 2: Linear Algebra Compiled By: Abhinaba Bala, Dakshit Agrawal, Mohit Jain Section 2.1: Scalars, Vectors, Matrices and Tensors Scalar Single Number Lowercase names in italic
Structure in Data. A major objective in data analysis is to identify interesting features or structure in the data.
 Structure in Data A major objective in data analysis is to identify interesting features or structure in the data. The graphical methods are very useful in discovering structure. There are basically two
Structure in Data A major objective in data analysis is to identify interesting features or structure in the data. The graphical methods are very useful in discovering structure. There are basically two
Multivariate Statistical Analysis
 Multivariate Statistical Analysis Fall 2011 C. L. Williams, Ph.D. Lecture 4 for Applied Multivariate Analysis Outline 1 Eigen values and eigen vectors Characteristic equation Some properties of eigendecompositions
Multivariate Statistical Analysis Fall 2011 C. L. Williams, Ph.D. Lecture 4 for Applied Multivariate Analysis Outline 1 Eigen values and eigen vectors Characteristic equation Some properties of eigendecompositions
More Linear Algebra. Edps/Soc 584, Psych 594. Carolyn J. Anderson
 More Linear Algebra Edps/Soc 584, Psych 594 Carolyn J. Anderson Department of Educational Psychology I L L I N O I S university of illinois at urbana-champaign c Board of Trustees, University of Illinois
More Linear Algebra Edps/Soc 584, Psych 594 Carolyn J. Anderson Department of Educational Psychology I L L I N O I S university of illinois at urbana-champaign c Board of Trustees, University of Illinois
Introduction to Machine Learning
 10-701 Introduction to Machine Learning PCA Slides based on 18-661 Fall 2018 PCA Raw data can be Complex, High-dimensional To understand a phenomenon we measure various related quantities If we knew what
10-701 Introduction to Machine Learning PCA Slides based on 18-661 Fall 2018 PCA Raw data can be Complex, High-dimensional To understand a phenomenon we measure various related quantities If we knew what
Economic Growth: Lecture 1, Questions and Evidence
 14.452 Economic Growth: Lecture 1, Questions and Evidence Daron Acemoglu MIT October 23, 2018 Daron Acemoglu (MIT) Economic Growth Lecture 1 October 23, 2018 1 / 38 Cross-Country Income Differences Cross-Country
14.452 Economic Growth: Lecture 1, Questions and Evidence Daron Acemoglu MIT October 23, 2018 Daron Acemoglu (MIT) Economic Growth Lecture 1 October 23, 2018 1 / 38 Cross-Country Income Differences Cross-Country
Principal Component Analysis
 Principal Component Analysis CS5240 Theoretical Foundations in Multimedia Leow Wee Kheng Department of Computer Science School of Computing National University of Singapore Leow Wee Kheng (NUS) Principal
Principal Component Analysis CS5240 Theoretical Foundations in Multimedia Leow Wee Kheng Department of Computer Science School of Computing National University of Singapore Leow Wee Kheng (NUS) Principal
Lecture 7 Spectral methods
 CSE 291: Unsupervised learning Spring 2008 Lecture 7 Spectral methods 7.1 Linear algebra review 7.1.1 Eigenvalues and eigenvectors Definition 1. A d d matrix M has eigenvalue λ if there is a d-dimensional
CSE 291: Unsupervised learning Spring 2008 Lecture 7 Spectral methods 7.1 Linear algebra review 7.1.1 Eigenvalues and eigenvectors Definition 1. A d d matrix M has eigenvalue λ if there is a d-dimensional
A re examination of the Columbian exchange: Agriculture and Economic Development in the Long Run
 Are examinationofthecolumbianexchange: AgricultureandEconomicDevelopmentintheLongRun AlfonsoDíezMinguela MªDoloresAñónHigón UniversitatdeValéncia UniversitatdeValéncia May2012 [PRELIMINARYRESEARCH,PLEASEDONOTCITE]
Are examinationofthecolumbianexchange: AgricultureandEconomicDevelopmentintheLongRun AlfonsoDíezMinguela MªDoloresAñónHigón UniversitatdeValéncia UniversitatdeValéncia May2012 [PRELIMINARYRESEARCH,PLEASEDONOTCITE]
Dimensionality Reduction
 394 Chapter 11 Dimensionality Reduction There are many sources of data that can be viewed as a large matrix. We saw in Chapter 5 how the Web can be represented as a transition matrix. In Chapter 9, the
394 Chapter 11 Dimensionality Reduction There are many sources of data that can be viewed as a large matrix. We saw in Chapter 5 how the Web can be represented as a transition matrix. In Chapter 9, the
CS 246 Review of Linear Algebra 01/17/19
 1 Linear algebra In this section we will discuss vectors and matrices. We denote the (i, j)th entry of a matrix A as A ij, and the ith entry of a vector as v i. 1.1 Vectors and vector operations A vector
1 Linear algebra In this section we will discuss vectors and matrices. We denote the (i, j)th entry of a matrix A as A ij, and the ith entry of a vector as v i. 1.1 Vectors and vector operations A vector
Probabilistic Latent Semantic Analysis
 Probabilistic Latent Semantic Analysis Seungjin Choi Department of Computer Science and Engineering Pohang University of Science and Technology 77 Cheongam-ro, Nam-gu, Pohang 37673, Korea seungjin@postech.ac.kr
Probabilistic Latent Semantic Analysis Seungjin Choi Department of Computer Science and Engineering Pohang University of Science and Technology 77 Cheongam-ro, Nam-gu, Pohang 37673, Korea seungjin@postech.ac.kr
TRUE OR FALSE: 3. "And Noah did according to all that the LORD commanded him." GENESIS 7:5 TRUE OR FALSE
 MEMORY VERSE: "The Lord is not slack concerning His promise, as some count slackness, but is longsuffering toward us, not willing that any should perish but that all should come to repentance." 2 PETER
MEMORY VERSE: "The Lord is not slack concerning His promise, as some count slackness, but is longsuffering toward us, not willing that any should perish but that all should come to repentance." 2 PETER
7 Principal Component Analysis
 7 Principal Component Analysis This topic will build a series of techniques to deal with high-dimensional data. Unlike regression problems, our goal is not to predict a value (the y-coordinate), it is
7 Principal Component Analysis This topic will build a series of techniques to deal with high-dimensional data. Unlike regression problems, our goal is not to predict a value (the y-coordinate), it is
REVIEW 8/2/2017 陈芳华东师大英语系
 REVIEW Hypothesis testing starts with a null hypothesis and a null distribution. We compare what we have to the null distribution, if the result is too extreme to belong to the null distribution (p
REVIEW Hypothesis testing starts with a null hypothesis and a null distribution. We compare what we have to the null distribution, if the result is too extreme to belong to the null distribution (p
Review of Linear Algebra
 Review of Linear Algebra Definitions An m n (read "m by n") matrix, is a rectangular array of entries, where m is the number of rows and n the number of columns. 2 Definitions (Con t) A is square if m=
Review of Linear Algebra Definitions An m n (read "m by n") matrix, is a rectangular array of entries, where m is the number of rows and n the number of columns. 2 Definitions (Con t) A is square if m=
Matrices and Vectors. Definition of Matrix. An MxN matrix A is a two-dimensional array of numbers A =
 30 MATHEMATICS REVIEW G A.1.1 Matrices and Vectors Definition of Matrix. An MxN matrix A is a two-dimensional array of numbers A = a 11 a 12... a 1N a 21 a 22... a 2N...... a M1 a M2... a MN A matrix can
30 MATHEMATICS REVIEW G A.1.1 Matrices and Vectors Definition of Matrix. An MxN matrix A is a two-dimensional array of numbers A = a 11 a 12... a 1N a 21 a 22... a 2N...... a M1 a M2... a MN A matrix can
Lecture 8. Principal Component Analysis. Luigi Freda. ALCOR Lab DIAG University of Rome La Sapienza. December 13, 2016
 Lecture 8 Principal Component Analysis Luigi Freda ALCOR Lab DIAG University of Rome La Sapienza December 13, 2016 Luigi Freda ( La Sapienza University) Lecture 8 December 13, 2016 1 / 31 Outline 1 Eigen
Lecture 8 Principal Component Analysis Luigi Freda ALCOR Lab DIAG University of Rome La Sapienza December 13, 2016 Luigi Freda ( La Sapienza University) Lecture 8 December 13, 2016 1 / 31 Outline 1 Eigen
Maximum Likelihood Estimation
 Maximum Likelihood Estimation Merlise Clyde STA721 Linear Models Duke University August 31, 2017 Outline Topics Likelihood Function Projections Maximum Likelihood Estimates Readings: Christensen Chapter
Maximum Likelihood Estimation Merlise Clyde STA721 Linear Models Duke University August 31, 2017 Outline Topics Likelihood Function Projections Maximum Likelihood Estimates Readings: Christensen Chapter
. = V c = V [x]v (5.1) c 1. c k
![. = V c = V [x]v (5.1) c 1. c k . = V c = V [x]v (5.1) c 1. c k](/thumbs/90/101201642.jpg) Chapter 5 Linear Algebra It can be argued that all of linear algebra can be understood using the four fundamental subspaces associated with a matrix Because they form the foundation on which we later work,
Chapter 5 Linear Algebra It can be argued that all of linear algebra can be understood using the four fundamental subspaces associated with a matrix Because they form the foundation on which we later work,
VECTORS, TENSORS AND INDEX NOTATION
 VECTORS, TENSORS AND INDEX NOTATION Enrico Nobile Dipartimento di Ingegneria e Architettura Università degli Studi di Trieste, 34127 TRIESTE March 5, 2018 Vectors & Tensors, E. Nobile March 5, 2018 1 /
VECTORS, TENSORS AND INDEX NOTATION Enrico Nobile Dipartimento di Ingegneria e Architettura Università degli Studi di Trieste, 34127 TRIESTE March 5, 2018 Vectors & Tensors, E. Nobile March 5, 2018 1 /
GI07/COMPM012: Mathematical Programming and Research Methods (Part 2) 2. Least Squares and Principal Components Analysis. Massimiliano Pontil
 GI07/COMPM012: Mathematical Programming and Research Methods (Part 2) 2. Least Squares and Principal Components Analysis Massimiliano Pontil 1 Today s plan SVD and principal component analysis (PCA) Connection
GI07/COMPM012: Mathematical Programming and Research Methods (Part 2) 2. Least Squares and Principal Components Analysis Massimiliano Pontil 1 Today s plan SVD and principal component analysis (PCA) Connection
Specification and estimation of exponential random graph models for social (and other) networks
 Specification and estimation of exponential random graph models for social (and other) networks Tom A.B. Snijders University of Oxford March 23, 2009 c Tom A.B. Snijders (University of Oxford) Models for
Specification and estimation of exponential random graph models for social (and other) networks Tom A.B. Snijders University of Oxford March 23, 2009 c Tom A.B. Snijders (University of Oxford) Models for
Fundamentals of Matrices
 Maschinelles Lernen II Fundamentals of Matrices Christoph Sawade/Niels Landwehr/Blaine Nelson Tobias Scheffer Matrix Examples Recap: Data Linear Model: f i x = w i T x Let X = x x n be the data matrix
Maschinelles Lernen II Fundamentals of Matrices Christoph Sawade/Niels Landwehr/Blaine Nelson Tobias Scheffer Matrix Examples Recap: Data Linear Model: f i x = w i T x Let X = x x n be the data matrix
DS-GA 1002 Lecture notes 10 November 23, Linear models
 DS-GA 2 Lecture notes November 23, 2 Linear functions Linear models A linear model encodes the assumption that two quantities are linearly related. Mathematically, this is characterized using linear functions.
DS-GA 2 Lecture notes November 23, 2 Linear functions Linear models A linear model encodes the assumption that two quantities are linearly related. Mathematically, this is characterized using linear functions.
The Blood Moon Tetrad
 The 2014-2015 Blood Moon Tetrad What is it? Does it mean anything??? PART 1 of 6 Introduction What is a blood moon? A blood moon is a name for a complete lunar eclipse when the sun s shadow completely
The 2014-2015 Blood Moon Tetrad What is it? Does it mean anything??? PART 1 of 6 Introduction What is a blood moon? A blood moon is a name for a complete lunar eclipse when the sun s shadow completely
Large Scale Data Analysis Using Deep Learning
 Large Scale Data Analysis Using Deep Learning Linear Algebra U Kang Seoul National University U Kang 1 In This Lecture Overview of linear algebra (but, not a comprehensive survey) Focused on the subset
Large Scale Data Analysis Using Deep Learning Linear Algebra U Kang Seoul National University U Kang 1 In This Lecture Overview of linear algebra (but, not a comprehensive survey) Focused on the subset
Linear Algebra (Review) Volker Tresp 2017
 Linear Algebra (Review) Volker Tresp 2017 1 Vectors k is a scalar (a number) c is a column vector. Thus in two dimensions, c = ( c1 c 2 ) (Advanced: More precisely, a vector is defined in a vector space.
Linear Algebra (Review) Volker Tresp 2017 1 Vectors k is a scalar (a number) c is a column vector. Thus in two dimensions, c = ( c1 c 2 ) (Advanced: More precisely, a vector is defined in a vector space.
Manning & Schuetze, FSNLP (c) 1999,2000
 558 15 Topics in Information Retrieval (15.10) y 4 3 2 1 0 0 1 2 3 4 5 6 7 8 Figure 15.7 An example of linear regression. The line y = 0.25x + 1 is the best least-squares fit for the four points (1,1),
558 15 Topics in Information Retrieval (15.10) y 4 3 2 1 0 0 1 2 3 4 5 6 7 8 Figure 15.7 An example of linear regression. The line y = 0.25x + 1 is the best least-squares fit for the four points (1,1),
Vector Space Models. wine_spectral.r
 Vector Space Models 137 wine_spectral.r Latent Semantic Analysis Problem with words Even a small vocabulary as in wine example is challenging LSA Reduce number of columns of DTM by principal components
Vector Space Models 137 wine_spectral.r Latent Semantic Analysis Problem with words Even a small vocabulary as in wine example is challenging LSA Reduce number of columns of DTM by principal components
PETER'S VISION (ACTS 10:9-16) MEMORY VERSE: "...What God has cleansed you must not call common." ACTS 10:15
 MEMORY VERSE: "...What God has cleansed you must not call common." ACTS 10:15 FILL IN THE BLANK: 1. "The next day, as they went on their journey and drew near to the city, Peter went up on the housetop
MEMORY VERSE: "...What God has cleansed you must not call common." ACTS 10:15 FILL IN THE BLANK: 1. "The next day, as they went on their journey and drew near to the city, Peter went up on the housetop
God s Amazingly Astonishing World
 a script from God s Amazingly Astonishing World by Ginny Neil What In this interactive Readers Theater, children and adults present the story of creation in a fun way. Works with as few as two children,
a script from God s Amazingly Astonishing World by Ginny Neil What In this interactive Readers Theater, children and adults present the story of creation in a fun way. Works with as few as two children,
IDENTIFYING MULTILATERAL DEPENDENCIES IN THE WORLD TRADE NETWORK
 IDENTIFYING MULTILATERAL DEPENDENCIES IN THE WORLD TRADE NETWORK PETER R. HERMAN Abstract. When studying the formation of trade between two countries, traditional modeling has described this decision as
IDENTIFYING MULTILATERAL DEPENDENCIES IN THE WORLD TRADE NETWORK PETER R. HERMAN Abstract. When studying the formation of trade between two countries, traditional modeling has described this decision as
Principal Component Analysis-I Geog 210C Introduction to Spatial Data Analysis. Chris Funk. Lecture 17
 Principal Component Analysis-I Geog 210C Introduction to Spatial Data Analysis Chris Funk Lecture 17 Outline Filters and Rotations Generating co-varying random fields Translating co-varying fields into
Principal Component Analysis-I Geog 210C Introduction to Spatial Data Analysis Chris Funk Lecture 17 Outline Filters and Rotations Generating co-varying random fields Translating co-varying fields into
Agent-Based Methods for Dynamic Social Networks. Duke University
 Agent-Based Methods for Dynamic Social Networks Eric Vance Institute of Statistics & Decision Sciences Duke University STA 395 Talk October 24, 2005 Outline Introduction Social Network Models Agent-based
Agent-Based Methods for Dynamic Social Networks Eric Vance Institute of Statistics & Decision Sciences Duke University STA 395 Talk October 24, 2005 Outline Introduction Social Network Models Agent-based
Days of Creation Mini Puzzle Unit
 Mini Puzzle Unit By: Annette @ In All You Do 2015 Thank you for visiting In All You Do and finding a resource you d like to use! Please feel free to use these files for your own personal use or for your
Mini Puzzle Unit By: Annette @ In All You Do 2015 Thank you for visiting In All You Do and finding a resource you d like to use! Please feel free to use these files for your own personal use or for your
STATISTICAL LEARNING SYSTEMS
 STATISTICAL LEARNING SYSTEMS LECTURE 8: UNSUPERVISED LEARNING: FINDING STRUCTURE IN DATA Institute of Computer Science, Polish Academy of Sciences Ph. D. Program 2013/2014 Principal Component Analysis
STATISTICAL LEARNING SYSTEMS LECTURE 8: UNSUPERVISED LEARNING: FINDING STRUCTURE IN DATA Institute of Computer Science, Polish Academy of Sciences Ph. D. Program 2013/2014 Principal Component Analysis
Lecture Notes 2: Matrices
 Optimization-based data analysis Fall 2017 Lecture Notes 2: Matrices Matrices are rectangular arrays of numbers, which are extremely useful for data analysis. They can be interpreted as vectors in a vector
Optimization-based data analysis Fall 2017 Lecture Notes 2: Matrices Matrices are rectangular arrays of numbers, which are extremely useful for data analysis. They can be interpreted as vectors in a vector
Introduction to statistical analysis of Social Networks
 The Social Statistics Discipline Area, School of Social Sciences Introduction to statistical analysis of Social Networks Mitchell Centre for Network Analysis Johan Koskinen http://www.ccsr.ac.uk/staff/jk.htm!
The Social Statistics Discipline Area, School of Social Sciences Introduction to statistical analysis of Social Networks Mitchell Centre for Network Analysis Johan Koskinen http://www.ccsr.ac.uk/staff/jk.htm!
Latent Semantic Analysis. Hongning Wang
 Latent Semantic Analysis Hongning Wang CS@UVa VS model in practice Document and query are represented by term vectors Terms are not necessarily orthogonal to each other Synonymy: car v.s. automobile Polysemy:
Latent Semantic Analysis Hongning Wang CS@UVa VS model in practice Document and query are represented by term vectors Terms are not necessarily orthogonal to each other Synonymy: car v.s. automobile Polysemy:
MA 575 Linear Models: Cedric E. Ginestet, Boston University Regularization: Ridge Regression and Lasso Week 14, Lecture 2
 MA 575 Linear Models: Cedric E. Ginestet, Boston University Regularization: Ridge Regression and Lasso Week 14, Lecture 2 1 Ridge Regression Ridge regression and the Lasso are two forms of regularized
MA 575 Linear Models: Cedric E. Ginestet, Boston University Regularization: Ridge Regression and Lasso Week 14, Lecture 2 1 Ridge Regression Ridge regression and the Lasso are two forms of regularized
Matrix Factorizations
 1 Stat 540, Matrix Factorizations Matrix Factorizations LU Factorization Definition... Given a square k k matrix S, the LU factorization (or decomposition) represents S as the product of two triangular
1 Stat 540, Matrix Factorizations Matrix Factorizations LU Factorization Definition... Given a square k k matrix S, the LU factorization (or decomposition) represents S as the product of two triangular
Lecture 13. Principal Component Analysis. Brett Bernstein. April 25, CDS at NYU. Brett Bernstein (CDS at NYU) Lecture 13 April 25, / 26
 Principal Component Analysis Brett Bernstein CDS at NYU April 25, 2017 Brett Bernstein (CDS at NYU) Lecture 13 April 25, 2017 1 / 26 Initial Question Intro Question Question Let S R n n be symmetric. 1
Principal Component Analysis Brett Bernstein CDS at NYU April 25, 2017 Brett Bernstein (CDS at NYU) Lecture 13 April 25, 2017 1 / 26 Initial Question Intro Question Question Let S R n n be symmetric. 1
Matrices A brief introduction
 Matrices A brief introduction Basilio Bona DAUIN Politecnico di Torino Semester 1, 2014-15 B. Bona (DAUIN) Matrices Semester 1, 2014-15 1 / 41 Definitions Definition A matrix is a set of N real or complex
Matrices A brief introduction Basilio Bona DAUIN Politecnico di Torino Semester 1, 2014-15 B. Bona (DAUIN) Matrices Semester 1, 2014-15 1 / 41 Definitions Definition A matrix is a set of N real or complex
Econometrics I KS. Module 1: Bivariate Linear Regression. Alexander Ahammer. This version: March 12, 2018
 Econometrics I KS Module 1: Bivariate Linear Regression Alexander Ahammer Department of Economics Johannes Kepler University of Linz This version: March 12, 2018 Alexander Ahammer (JKU) Module 1: Bivariate
Econometrics I KS Module 1: Bivariate Linear Regression Alexander Ahammer Department of Economics Johannes Kepler University of Linz This version: March 12, 2018 Alexander Ahammer (JKU) Module 1: Bivariate
j=1 u 1jv 1j. 1/ 2 Lemma 1. An orthogonal set of vectors must be linearly independent.
 Lecture Notes: Orthogonal and Symmetric Matrices Yufei Tao Department of Computer Science and Engineering Chinese University of Hong Kong taoyf@cse.cuhk.edu.hk Orthogonal Matrix Definition. Let u = [u
Lecture Notes: Orthogonal and Symmetric Matrices Yufei Tao Department of Computer Science and Engineering Chinese University of Hong Kong taoyf@cse.cuhk.edu.hk Orthogonal Matrix Definition. Let u = [u
Department of Statistics. Bayesian Modeling for a Generalized Social Relations Model. Tyler McCormick. Introduction.
 A University of Connecticut and Columbia University A models for dyadic data are extensions of the (). y i,j = a i + b j + γ i,j (1) Here, y i,j is a measure of the tie from actor i to actor j. The random
A University of Connecticut and Columbia University A models for dyadic data are extensions of the (). y i,j = a i + b j + γ i,j (1) Here, y i,j is a measure of the tie from actor i to actor j. The random
Linear Algebra & Geometry why is linear algebra useful in computer vision?
 Linear Algebra & Geometry why is linear algebra useful in computer vision? References: -Any book on linear algebra! -[HZ] chapters 2, 4 Some of the slides in this lecture are courtesy to Prof. Octavia
Linear Algebra & Geometry why is linear algebra useful in computer vision? References: -Any book on linear algebra! -[HZ] chapters 2, 4 Some of the slides in this lecture are courtesy to Prof. Octavia
Lecture 2: Linear Algebra Review
 EE 227A: Convex Optimization and Applications January 19 Lecture 2: Linear Algebra Review Lecturer: Mert Pilanci Reading assignment: Appendix C of BV. Sections 2-6 of the web textbook 1 2.1 Vectors 2.1.1
EE 227A: Convex Optimization and Applications January 19 Lecture 2: Linear Algebra Review Lecturer: Mert Pilanci Reading assignment: Appendix C of BV. Sections 2-6 of the web textbook 1 2.1 Vectors 2.1.1
Multilinear tensor regression for longitudinal relational data
 Multilinear tensor regression for longitudinal relational data Peter D. Hoff 1 Working Paper no. 149 Center for Statistics and the Social Sciences University of Washington Seattle, WA 98195-4320 November
Multilinear tensor regression for longitudinal relational data Peter D. Hoff 1 Working Paper no. 149 Center for Statistics and the Social Sciences University of Washington Seattle, WA 98195-4320 November
Eigenvalues and diagonalization
 Eigenvalues and diagonalization Patrick Breheny November 15 Patrick Breheny BST 764: Applied Statistical Modeling 1/20 Introduction The next topic in our course, principal components analysis, revolves
Eigenvalues and diagonalization Patrick Breheny November 15 Patrick Breheny BST 764: Applied Statistical Modeling 1/20 Introduction The next topic in our course, principal components analysis, revolves
Definition 2.3. We define addition and multiplication of matrices as follows.
 14 Chapter 2 Matrices In this chapter, we review matrix algebra from Linear Algebra I, consider row and column operations on matrices, and define the rank of a matrix. Along the way prove that the row
14 Chapter 2 Matrices In this chapter, we review matrix algebra from Linear Algebra I, consider row and column operations on matrices, and define the rank of a matrix. Along the way prove that the row
Fiedler s Theorems on Nodal Domains
 Spectral Graph Theory Lecture 7 Fiedler s Theorems on Nodal Domains Daniel A Spielman September 9, 202 7 About these notes These notes are not necessarily an accurate representation of what happened in
Spectral Graph Theory Lecture 7 Fiedler s Theorems on Nodal Domains Daniel A Spielman September 9, 202 7 About these notes These notes are not necessarily an accurate representation of what happened in
TRUE OR FALSE: 3. "So God blessed Noah and his sons, and said to them: 'Be fruitful and multiply, and fill the earth.' " GENESIS 9:1 TRUE OR FALSE
 MEMORY VERSE: "The rainbow shall be in the cloud, and I will look on it to remember the everlasting covenant between God and every living creature of all flesh that is on the earth." GENESIS 9:16 CIRCLE
MEMORY VERSE: "The rainbow shall be in the cloud, and I will look on it to remember the everlasting covenant between God and every living creature of all flesh that is on the earth." GENESIS 9:16 CIRCLE
1 Vocabulary. Chapter 5 Ecology. Lesson
 1 Vocabulary Symbiosis a close, long-term relationship between organisms that benefits at least one of the organisms Decomposer living thing that breaks down waste and things that have died Energy pyramid
1 Vocabulary Symbiosis a close, long-term relationship between organisms that benefits at least one of the organisms Decomposer living thing that breaks down waste and things that have died Energy pyramid
Bargaining, Information Networks and Interstate
 Bargaining, Information Networks and Interstate Conflict Erik Gartzke Oliver Westerwinter UC, San Diego Department of Political Sciene egartzke@ucsd.edu European University Institute Department of Political
Bargaining, Information Networks and Interstate Conflict Erik Gartzke Oliver Westerwinter UC, San Diego Department of Political Sciene egartzke@ucsd.edu European University Institute Department of Political
