Univariate shrinkage in the Cox model for high dimensional data
|
|
|
- Horatio Hutchinson
- 6 years ago
- Views:
Transcription
1 Univariate shrinkage in the Cox model for high dimensional data Robert Tibshirani January 6, 2009 Abstract We propose a method for prediction in Cox s proportional model, when the number of features (regressors) p exceeds the number of observations n. The method assumes that features are independent in each risk set, so that the partial likelihood factors into a product. As such, it is analogous to univariate thresholding in linear regression and nearest shrunken centroids in classification. We call the procedure Cox univariate shrinkage and demonstrate its usefulness on real and simulated data. The method has the attractive property of being essentially univariate in its operation: the features are entered into the model based on the size of their Cox score statistics. We illustrate the new method on real and simulated data, and compare it to other proposed methods for survival prediction with a large number of predictors. Keywords: survival data, Proportional hazards model, Cox model, shrinkage, subset selection 1 Introduction High-dimensional regression problems are challenging, and a current focus of statistical research. One simplifying assumption that can be useful is that of independence of features. While this assumption will rarely be even approximately true, in practice, it leads to simple estimates with predictive performance sometimes as good or better than more ambitious multivariate methods. Depts. of Health, Research & Policy, and Statistics, Stanford Univ, Stanford CA, 94305; tibs@stat.stanford.edu 1
2 In linear regression with squared error loss and a lasso penalty, independence of the features leads to the univariate soft-thresholding estimates, that often behave well. In classification problems, the idea of class-wise independence forms the basis for diagonal discriminant analysis. In that setting, if we assume that the features are independent within each class, and incorporate a lasso penalty, the resulting estimator is the nearest shrunken centroid procedure proposed by Tibshirani et al. (2001): we review this fact in the next section. The nearest shrunken centroid classifier is very simple but useful in practice, and has many attractive properties. It has a single tuning parameter that automatically selects the features to retain in the model, and it provides a single ranking of all features, that is independent of the value of the tuning parameter. We review the details of the regression and classification problems in the next section. Then we extend the idea of class-wise feature independence to the Cox model for survival data: this is the main contribution of the paper. We study this new procedure on both real and simulated datasets, and compare it to competing methods for this problem. 2 Regression and classification using feature independence 2.1 Regression Consider the usual regression situation: we have data (x i, y i ), i = 1, 2,... n where x i = (x i1,... x ip ) T and y i are the regressors and response for the ith observation. Assume the x ij are standardized so that i x ij/n = 0, i x2 ij /n = 1. The lasso finds β = (β 1,... β p ) T to minimize N i=1 (y i j p β j x ij ) 2 + λ β j. (1) j=1 Now if we assume that the features are independent (uncorrelated), it is easy to show that ˆβ j = sign( ˆβ o j )( ˆβ o j λ) + (2) where ˆβ o j are the ordinary (univariate) least squares estimates. These are univariate soft-thresholded estimates, and perform particularly well when p n. Zou & Hastie (2005) study these estimates as a special case of their elastic net procedure. See Fan & Fan (2008) for some theoretical support of univariate soft-thresholding. 2
3 2.2 Classification Here we review the nearest shrunken centroid method for high-dimensional classification and its connection to the lasso. Consider the diagonal-covariance linear discriminant rule for classification. The discriminant score for class k is p (x δ k (x j x kj ) 2 ) = + 2 log π k. (3) j=1 Here x = (x 1, x 2,..., x p )T is a vector of expression values for a test observation, s j is the pooled within-class standard deviation of the jth gene, and x kj = i C k x ij /n k is the mean of the n k values for gene j in class k, with C k being the index set for class k. We call x k = ( x k1, x k2,... x kp ) T the centroid of class k. The first part of (3) is simply the (negative) standardized squared distance of x to the kth centroid. The second part is a correction based on the class prior probability π k, where K k=1 π k = 1. The classification rule is then s 2 j C(x ) = l if δ l (x ) = max k δ k (x ). (4) We see that the diagonal LDA classifier is equivalent to a nearest centroid classifier after appropriate standardization. The nearest shrunken centroid classifier regularizes this further, and is defined as follows. Let d kj = x kj x j m k (s j + s 0 ), (5) where x j is the overall mean for gene j, m 2 k = 1/n k 1/n and s 0 is a small positive constant, typically chosen to be the median of the s j values. This constant guards against large d kj values that arise from expression values near zero. Finally, we can derive the nearest shrunken centroid classifier as the solution to a lasso-regularized problem that assumes independence of the features. Consider a (naive Bayes) Gaussian model for classification in which the features j = 1, 2,..., p are assumed to be independent within each class k = 1, 2,..., K. With observations i = 1, 2,..., n and C k equal to the set of indices of the n k observations in class k, we observe x ij N(µ j + µ jk, σj 2) for i C k with K k=1 µ jk = 0. We set ˆσ j 2 = s2 j, the pooled within-class variance for feature j, and consider the lasso-style minimization problem { 1 p K min {µ j,µ jk } 2 j=1 k=1 (x ij µ j µ jk)2 s 2 i C j k + λ n k p K j=1 k=1 } µ jk. (6) s j By direct calculation, it is easy to show that the solution is equivalent to the nearest shrunken centroid estimator (5), with s 0 set to zero, and M k equal to 1/n k instead of 1/n k 1/n as before. Some interesting theoretical results for diagonal linear discriminant analysis are given in Bickel & Levina (2004). Efron (2008) proposes an Empirical Bayes variation on nearest shrunken centroids. 3
4 3 Survival data and Cox s proportional hazards model For the rest of this paper we consider the right-censored survival data setting. The data available are of the form (y 1, x 1, δ 1, ),..., (y n, x n, δ n ), the survival time y i being complete if δ i = 1 and right censored if δ i = 0, with x i denoting the usual vector of predictors (x 1, x 2,... x p ) for the ith individual. Denote the distinct failure times by t 1 < < t K, there being d k failures at time t k. The proportional-hazards model for survival data, also known as the Cox model, assumes that ( ) λ(t x) = λ 0 (t) exp x j β j (7) where λ(t x) is the hazard at time t given predictor values x = (x 1,..., x p ), and λ 0 (t) is an arbitrary baseline hazard function. One usually estimates the parameter β = (β 1, β 2,... β p ) T in the proportionalhazards model (7) without specification of λ 0 (t) through maximization of the partial likelihood PL(β) = exp(x k β) { k D m R k exp(x m β)}. Here D is the set of indices of the failure times, R k is the set of indices of the individuals at risk at time t k 0, and we have assumed there are no ties in the survival times. In the case of ties, we use the Breslow approximation to the partial likelihood. We also assume that the censoring is noninformative, so that the construction of the partial likelihood is justified. If p n then maximizer of the partial likelihood is not unique, and some regularization must be used. One possibility is to include a lasso penalty and jointly optimize over the parameters, as suggested by Tibshirani (1997). Here we extend the idea of feature independence to the Cox model for survival data. Now the partial likelihood term exp(x k β)/ m R k exp(x m β) equals Pr(k x i, i R k ) the probability that individual k is the one who fails at time t k, given the risk set and their feature vectors. Now we assume that both conditionally on each risk set, and marginally, the features are independent of one another. Then using Bayes theorem we obtain Pr(k x i, i R k ) j Pr(k x ij, i R k ). As a result, up to a proportionality constant, the partial likelihood becomes a simple product p exp(x kj β j ) PL(β) = c { m R k exp(x mj β j )} j=1 k D j 4
5 The log partial likelihood is l(β) = p K xkj β j log k=1( m R k p g j (β j ). (9) j=1 j=1 exp(x mj β j ) ) (8) We propose the Cox univariate shrinkage () estimator as the maximizer of the penalized partial log-likelihood J(β) = p g j (β j ) λ β j. (10) j=1 Here λ 0 is a tuning parameter, and we solve this problem for a range of λ values. Note that is just a set of one-dimensional maximizations, since we can maximize each function g j (β j ) λ β j separately. The minimizers of (10) are not simply soft-thresholded versions of the unpenalized estimates ˆβ j, as they are in the least squares regression case. We discuss and illustrate this fact in the next section. From this, we obtain solutions paths ( ˆβ 1 (λ), ˆβ 2 (λ),... ˆβ p (λ)), and we can use these to predict the survival in a separate test sample. Note that the estimates ˆβ will tend to be biased towards zero, especially for larger values of λ. In the test set and cross-validation computations in this paper, we fit the predictor z = x ˆβ in the new (test or validation) data, and so obtain a scale factor to debias the estimate. If we are interested in the values of ˆβ themselves, a better approach would be fit the single predictor γx i ˆβ in a Cox model on the training data, and hence obtain the adjusted estimates ˆγ ˆβ. The lasso penalty above might not make sense if the predictors are in different units. Therefore we first standardize each predictor x j by s j, the square root of the observed (negative) Fisher information at β j = Example Zhao et al. (2005) collected gene expression data on 14, 814 genes from 177 kidney patients. Survival times (possibly censored) were also measured for each patient. The data were split into 88 samples to form the training set and the remaining 89 formed the test set. Figure 1 shows the solution paths for Cox univariate shrinkage for 500 randomly chosen genes. Notice that the solutions paths have a particularly nice form: they are monotonically decreasing in absolute value and once they hit zero, they stay at zero. It is easy to prove that this holds in general. The proof is given in the Appendix. Note also that the profiles decrease at different rates. This is somewhat surprising. In the least squares regression case under orthogonality of the predictors, 5
6 Coefficients PSfrag replacements λ Figure 1: Kidney cancer example: solution paths for univariate shrinkage, for 500 randomly chosen genes. the lasso estimates are simply the soft-thresholded versions of the least squares estimates ˆβ j :, that is, sign( ˆβ j )( ˆβ j t) +. Thus every profile decreases at the same rate. Here, each profile decreases at rate 1/ g ( ˆβ j ), which is the reciprocal of the observed Fisher information. Let U j, V j be the gradient of the (unpenalized) log-partial likelihood and the (negative) observed Fisher information. Then we can also show that ˆβ λ 0 U j / V j > λ. (11) The proof is given in the Appendix. Therefore we have the following useful fact: For any value of λ, the set of predictors that appear in the model estimated by Cox univariate shrinkage are just those whose score statistic exceeds λ in absolute value. That is, the Cox univariate shrinkage method used a fixed ranking of all features based on the Cox score statistic. This is very convenient for interpretation, as the Cox score is often used for determining the univariate significance of features (e.g. in the SAM procedure of Tusher et al. (2001)). However the non-zero coefficients given to these predictors are not simply the score statistics or soft thresholded versions of them. Figure 3 shows the drop in test sample deviance for (Cox univariate shrinkage), lasso, SPC (Supervised principal components) and UST (univariate 6
7 Drop in deviance Number of non zero genes Figure 2: Kidney cancer example: cross-validation curve. Drop in deviance on validation set, with plus and minus one standard error bars. soft thresholding), plotted against the number of genes in the model. The and SPC methods work best. Figure 4 shows the univariate Cox coefficients plotted against the Cox score statistics, for the kidney cancer data. Although the two measures are correlated, this correlation is far from perfect, and again this underlines the difference between our proposed univariate shrinkage method, which enters features based on the size of the Cox score, and simple soft-thresholding of the univariate coefficients. The latter ranks features based on the size of their unregularized coefficient, while the former looks at the standardized effect size at the null end of the scale. To estimate the tuning parameter λ we can use cross-validation. However it is not clear how to apply cross-validation in the setting of partial likelihood, since this function is not a simple sum of independent terms. Verweij & van Houwelingen (1993) propose an interesting method for leave-one-out cross-validation of the partial likelihood: for each left-out observation they compute the probability of its sample path over the risk sets. For the fitting methods presented in this paper, this leave-one-out cross-validation was too slow, requiring too many model fits. When we instead tried 5 or 10- fold partial likelihood cross-validation, the resulting curve was quite unstable. So we settled on a simpler approach, 5-fold cross-validation where we fit a Cox model to the left out one-fifth and record the drop in deviance. The results for the kidney cancer data is shown in Figure 2. The curve is a reasonable estimate of the test deviance curve of Figure 3. In some microarray datasets, there are duplicate genes, or multiple clones representing the same gene. In this case, the data for the different features will 7
8 Drop in test set deviance SPC UST Figure 3: Kidney cancer example: drop in test sample deviance for (Cox univariate shrinkage), lasso, SPC (Supervised principal components) and UST (univariate soft thresholding). Univariate Cox coefficients Cox score Figure 4: Kidney cancer data: the univariate Cox coefficients plotted against the Cox score statistics. 8
9 be highly correlated, and it makes sense to average the data for each gene, before applying UC. One can also average genes whose pairwise correlation is very high, whether or not they re supposed to represent the same gene. For example, when we averaged genes with correlation greater than 0.9 in the kidney cancer dataset, it reduced the number of genes by a few hundred and caused very little change in the test deviance curve. 3.2 Selection of a more parsimonius model The procedure enters predictors into the model based on this univariate Cox scores. Thus if two predictors are strongly predictive and correlated with each other, both will appear in the model. In some situations it is desirable pare down the predictors to a smaller, mode independent set. The lasso does this selection automatically, but sometimes doesn t predict as well as other methods (as in Figure 3.) An alternative approach is pre-conditioning (Paul et al. 2008), in which we apply the standard (regression) lasso to the fitted values from a model. In Paul et al. (2008) the initial fitting method used was supervised principal components. Here we start with the fit using 500 genes, which approximately the best model from Figure 2. We then apply the (regression version) of the lasso to the fitted values, and we report the drop in test deviance in Figure 5, along with that for itself (green curve copied from Figure 3.) We see that pre-conditioning gives about the same drop in test deviance as the 500 gene model, but using fewer than 100 genes. And it performs better here than the Cox model lasso (blue curve in Figure 3, a finding supported theoretically in Paul et al. (2008). Figure 6 shows the univariate Cox scores for the predictors entered by each method. We see that the pre-conditioning approach enters only predictors that are individually strong, while the usual lasso enter many predictors that are individually weak. 4 Simulation study We generated Gaussian samples of size n = 50 with p = 1000 predictors, with correlation ρ(x i, x j ) between each pair of predictors. The outcome y was generated as an exponential random variable with mean exp( x j β j ), with a randomly chosen s out of the p coefficients equal to value ±β and the rest equal to zero. We considered two scenarios s = 20 and s = 200, and tried both ρ(x i, x j ) = 0 and ρ(x i, x j ) = 0.5 i j. Three fitting methods for the Cox model were compared: lasso, supervised principal components, and, over a range of the tuning parameter for each method. A test set of size 400 was generated and the test set deviance of each fitting model was computed. Figure 7 shows the median test deviance plus and minus one standard error over 10 simulations, plotted against the number of genes in each model. 9
10 Drop in test set deviance Pre cond Figure 5: Kidney cancer example: drop in test sample deviance for (Cox univariate shrinkage), and pre-conditioning applied to. In the latter we apply the regression version of the lasso to the fitted values from. Cox model lasso applied to fit Univariate Cox score Univariate Cox score Number of features entered Number of features entered Figure 6: Kidney cancer example: univariate Cox scores for all predictors (black) and those entered into the model (red). Panel on the left corresponds to L 1 - penalized partial likelihood, fit by coxpath. On the right is the standard regression version of the lasso applied to the fitted values from. 10
11 Corr= 0, #nonzero= 20 Corr= 0.5, #nonzero= 20 Drop in test set deviance SPC Drop in test set deviance SPC Corr= 0, #nonzero= 200 Corr= 0.5, #nonzero= 200 Drop in test set deviance SPC Drop in test set deviance SPC Figure 7: Simulation results: average drop in test set deviance (± one standard error), for three different estimators, over four different simulation settings. 11
12 Figure 8 shows the number of genes correctly included in the model versus the number of genes in the model, for each of and lasso. Neither method is very good at isolating the true non-zero predictors, but surprisingly the lasso does no better than the simple procedure. Corr= 0, #nonzero= 20 Corr= 0.5, #nonzero= 20 Number correct Number correct chosen chosen Corr= 0, #nonzero= 200 Corr= 0.5, #nonzero= 200 Number correct Number correct chosen chosen Figure 8: Simulation results: number of correctly included predictors in the model, for and lasso, over the four different simulation settings. 12
13 5 Some real data examples We applied three methods- lasso, supervised principal components, and, to three real datasets. The first is the kidney data discussed earlier; the other datasets are the diffuse large cell lymphoma data of Rosenwald et al. (2002) (7399 genes and 240 patients), and the breast cancer data of van t Veer et al. (2002) (24881 genes, 295 patients). We randomly divided each dataset into training and test sets ten times, and averaged the results. The first two datasets included test sets of size 88 and 80 respectively, so we retained those test set sizes. For the third dataset, we took a test set of about one-third (98 patients), and to ease the computation load, restricted attention to the 10, 000 genes with largest variance. Figure 9 shows the median test deviance plus and minus one standard error over 10 simulations, plotted against the number of genes in each model. The method yields a larger drop in test deviance than the lasso, in at least two of the datasets and performs a little better than SPC overall. Acknowledgments. I thank Daniela Witten for helpful comments, and acknowledge support from National Science Foundation Grant DMS and National Institutes of Health Contract N01-HV Appendix Monotonicity of profiles. The subgradient equation for each β j is: K ( m R s kj d k x mj exp(x mj β j )) k λ t j = 0 (12) m R k exp(x mj β j ) k=1 where t j sign(β j ), that is t j = sign(β j ) if β j 0 and t j [ 1, 1] if β j = 0 Claim: ˆβ j (λ) is strictly decreasing in λ when ˆβ j (λ) is non-zero, and if ˆβ j (λ) = 0 then ˆβ j (λ ) = 0 for all λ > λ. Proof: for ease of notation, suppose we have a single β having the sub-gradient equation g(β) λt = 0 (13) with t sign(β). Then g (β) is the negative of the observed information and it is easy to show that g (β) < 0. Denote the solution to (13) for parameter λ by ˆβ(λ). Suppose that we have a solution ˆβ(λ) 0 for some λ. WLOG assume ˆβ(λ) > 0. Then by (13) we have g( ˆβ(λ)) = λ 13
14 Drop in test set deviance SPC Drop in test set deviance SPC Drop in test set deviance SPC Figure 9: Top to bottom: results for Kidney, Lymphoma and Breast cancer datasets; shown is the average drop in test set deviance (± one standard error) for (Cox univariate shrinkage), lasso and SPC (supervised principal components). 14
15 Then if λ > λ, we can t have ˆβ(λ ) ˆβ(λ) since this would imply g( ˆβ(λ )) < g( ˆβ(λ)) = λ < λ. Hence ˆβ(λ ) < ˆβ(λ). On the other hand, if ˆβ(λ) = 0 then g(0) λt = 0 for some t with t 1. Then if λ > λ the equation g(0) λ t = 0 can be solved by taking t = t(λ/λ ). Hence ˆβ(λ ) = 0. Proof of (11) Let f j (β j ) = K ( m R s kj d k x mj exp(x mj β j )) k m R k exp(x mj β j ) k=1 Suppose ˆβ j (0) > 0. Then ˆβ j (λ) 0 the equation f j (β j ) λ = 0 has a solution in (0, ˆβ j (0)). Since the x j have been standardized by V j, this equivalent to f j (0)/ V j > λ. This latter quantity is the (square root) of the usual score statistic for β j = 0. References Bickel, P. & Levina, E. (2004), Some theory for Fisher s linear discriminant function, naive Bayes, and some alternatives when there are many more variables than observations, Bernoulli 10, Efron, B. (2008), Empirical bayes estimates for large scale prediction problems. Unpublished. Fan, J. & Fan, Y. (2008), Sure independence screening for ultra-high dimensional feature space, Journal of the Royal Statistical Society Series B, to appear. Paul, D., Bair, E., Hastie, T. & Tibshirani, R. (2008), pre-conditioning for feature selection and regression in high-dimensional problems, Annals of Statistics 36(4), Rosenwald, A., Wright, G., Chan, W. C., Connors, J. M., Campo, E., Fisher, R. I., Gascoyne, R. D., Muller-Hermelink, H. K., Smeland, E. B. & Staudt, L. M. (2002), The use of molecular profiling to predict survival after chemotherapy for diffuse large b-cell lymphoma, The New England Journal of Medicine 346, Tibshirani, R. (1997), The lasso method for variable selection in the cox model, Statistics in Medicine 16,
16 Tibshirani, R., Hastie, T., Narasimhan, B. & Chu, G. (2001), Diagnosis of multiple cancer types by shrunken centroids of gene expression, Proceedings of the National Academy of Sciences 99, Tusher, V., Tibshirani, R. & Chu, G. (2001), Significance analysis of microarrays applied to transcriptional responses to ionizing radiation, Proc. Natl. Acad. Sci. USA. 98, van t Veer, L. J., van de Vijver, H. D. M. J., He, Y. D., Hart, A. A., Mao, M., Peterse, H. L., van der Kooy, K., Marton, M. J., Anke T. Witteveen and, G. J. S., Kerkhoven, R. M., Roberts, C., Linsley, P. S., Bernards, R., & Friend, S. H. (2002), Gene expression profiling predicts clinical outcome of breast cancer, Nature 415, Verweij, P. & van Houwelingen, H. (1993), Cross-validation in survival analysis, Statistics in Medicine 12, Zhao, H., Tibshirani, R. & Brooks, J. (2005), Gene expression profiling predicts survival in conventional renal cell carcinoma, PLOS Medicine pp Zou, H. & Hastie, T. (2005), Regularization and variable selection via the elastic net, Journal of the Royal Statistical Society Series B. 67(2),
A Bias Correction for the Minimum Error Rate in Cross-validation
 A Bias Correction for the Minimum Error Rate in Cross-validation Ryan J. Tibshirani Robert Tibshirani Abstract Tuning parameters in supervised learning problems are often estimated by cross-validation.
A Bias Correction for the Minimum Error Rate in Cross-validation Ryan J. Tibshirani Robert Tibshirani Abstract Tuning parameters in supervised learning problems are often estimated by cross-validation.
Simultaneous variable selection and class fusion for high-dimensional linear discriminant analysis
 Biostatistics (2010), 11, 4, pp. 599 608 doi:10.1093/biostatistics/kxq023 Advance Access publication on May 26, 2010 Simultaneous variable selection and class fusion for high-dimensional linear discriminant
Biostatistics (2010), 11, 4, pp. 599 608 doi:10.1093/biostatistics/kxq023 Advance Access publication on May 26, 2010 Simultaneous variable selection and class fusion for high-dimensional linear discriminant
Covariance-regularized regression and classification for high-dimensional problems
 Covariance-regularized regression and classification for high-dimensional problems Daniela M. Witten Department of Statistics, Stanford University, 390 Serra Mall, Stanford CA 94305, USA. E-mail: dwitten@stanford.edu
Covariance-regularized regression and classification for high-dimensional problems Daniela M. Witten Department of Statistics, Stanford University, 390 Serra Mall, Stanford CA 94305, USA. E-mail: dwitten@stanford.edu
A STUDY OF PRE-VALIDATION
 A STUDY OF PRE-VALIDATION Holger Höfling Robert Tibshirani July 3, 2007 Abstract Pre-validation is a useful technique for the analysis of microarray and other high dimensional data. It allows one to derive
A STUDY OF PRE-VALIDATION Holger Höfling Robert Tibshirani July 3, 2007 Abstract Pre-validation is a useful technique for the analysis of microarray and other high dimensional data. It allows one to derive
CMSC858P Supervised Learning Methods
 CMSC858P Supervised Learning Methods Hector Corrada Bravo March, 2010 Introduction Today we discuss the classification setting in detail. Our setting is that we observe for each subject i a set of p predictors
CMSC858P Supervised Learning Methods Hector Corrada Bravo March, 2010 Introduction Today we discuss the classification setting in detail. Our setting is that we observe for each subject i a set of p predictors
Lecture 6: Methods for high-dimensional problems
 Lecture 6: Methods for high-dimensional problems Hector Corrada Bravo and Rafael A. Irizarry March, 2010 In this Section we will discuss methods where data lies on high-dimensional spaces. In particular,
Lecture 6: Methods for high-dimensional problems Hector Corrada Bravo and Rafael A. Irizarry March, 2010 In this Section we will discuss methods where data lies on high-dimensional spaces. In particular,
Building a Prognostic Biomarker
 Building a Prognostic Biomarker Noah Simon and Richard Simon July 2016 1 / 44 Prognostic Biomarker for a Continuous Measure On each of n patients measure y i - single continuous outcome (eg. blood pressure,
Building a Prognostic Biomarker Noah Simon and Richard Simon July 2016 1 / 44 Prognostic Biomarker for a Continuous Measure On each of n patients measure y i - single continuous outcome (eg. blood pressure,
Fast Regularization Paths via Coordinate Descent
 August 2008 Trevor Hastie, Stanford Statistics 1 Fast Regularization Paths via Coordinate Descent Trevor Hastie Stanford University joint work with Jerry Friedman and Rob Tibshirani. August 2008 Trevor
August 2008 Trevor Hastie, Stanford Statistics 1 Fast Regularization Paths via Coordinate Descent Trevor Hastie Stanford University joint work with Jerry Friedman and Rob Tibshirani. August 2008 Trevor
Statistical Applications in Genetics and Molecular Biology
 Statistical Applications in Genetics and Molecular Biology Volume 1, Issue 1 2002 Article 1 Pre-validation and inference in microarrays Robert J. Tibshirani Brad Efron Stanford University, tibs@stat.stanford.edu
Statistical Applications in Genetics and Molecular Biology Volume 1, Issue 1 2002 Article 1 Pre-validation and inference in microarrays Robert J. Tibshirani Brad Efron Stanford University, tibs@stat.stanford.edu
TESTING SIGNIFICANCE OF FEATURES BY LASSOED PRINCIPAL COMPONENTS. BY DANIELA M. WITTEN 1 AND ROBERT TIBSHIRANI 2 Stanford University
 The Annals of Applied Statistics 2008, Vol. 2, No. 3, 986 1012 DOI: 10.1214/08-AOAS182 Institute of Mathematical Statistics, 2008 TESTING SIGNIFICANCE OF FEATURES BY LASSOED PRINCIPAL COMPONENTS BY DANIELA
The Annals of Applied Statistics 2008, Vol. 2, No. 3, 986 1012 DOI: 10.1214/08-AOAS182 Institute of Mathematical Statistics, 2008 TESTING SIGNIFICANCE OF FEATURES BY LASSOED PRINCIPAL COMPONENTS BY DANIELA
Journal of Statistical Software
 JSS Journal of Statistical Software March 2011, Volume 39, Issue 5. http://www.jstatsoft.org/ Regularization Paths for Cox s Proportional Hazards Model via Coordinate Descent Noah Simon Stanford University
JSS Journal of Statistical Software March 2011, Volume 39, Issue 5. http://www.jstatsoft.org/ Regularization Paths for Cox s Proportional Hazards Model via Coordinate Descent Noah Simon Stanford University
Regularized Discriminant Analysis and Its Application in Microarrays
 Biostatistics (2005), 1, 1, pp. 1 18 Printed in Great Britain Regularized Discriminant Analysis and Its Application in Microarrays By YAQIAN GUO Department of Statistics, Stanford University Stanford,
Biostatistics (2005), 1, 1, pp. 1 18 Printed in Great Britain Regularized Discriminant Analysis and Its Application in Microarrays By YAQIAN GUO Department of Statistics, Stanford University Stanford,
Regularization and Variable Selection via the Elastic Net
 p. 1/1 Regularization and Variable Selection via the Elastic Net Hui Zou and Trevor Hastie Journal of Royal Statistical Society, B, 2005 Presenter: Minhua Chen, Nov. 07, 2008 p. 2/1 Agenda Introduction
p. 1/1 Regularization and Variable Selection via the Elastic Net Hui Zou and Trevor Hastie Journal of Royal Statistical Society, B, 2005 Presenter: Minhua Chen, Nov. 07, 2008 p. 2/1 Agenda Introduction
The lasso: some novel algorithms and applications
 1 The lasso: some novel algorithms and applications Newton Institute, June 25, 2008 Robert Tibshirani Stanford University Collaborations with Trevor Hastie, Jerome Friedman, Holger Hoefling, Gen Nowak,
1 The lasso: some novel algorithms and applications Newton Institute, June 25, 2008 Robert Tibshirani Stanford University Collaborations with Trevor Hastie, Jerome Friedman, Holger Hoefling, Gen Nowak,
A Blockwise Descent Algorithm for Group-penalized Multiresponse and Multinomial Regression
 A Blockwise Descent Algorithm for Group-penalized Multiresponse and Multinomial Regression Noah Simon Jerome Friedman Trevor Hastie November 5, 013 Abstract In this paper we purpose a blockwise descent
A Blockwise Descent Algorithm for Group-penalized Multiresponse and Multinomial Regression Noah Simon Jerome Friedman Trevor Hastie November 5, 013 Abstract In this paper we purpose a blockwise descent
EVALUATING THE REPEATABILITY OF TWO STUDIES OF A LARGE NUMBER OF OBJECTS: MODIFIED KENDALL RANK-ORDER ASSOCIATION TEST
 EVALUATING THE REPEATABILITY OF TWO STUDIES OF A LARGE NUMBER OF OBJECTS: MODIFIED KENDALL RANK-ORDER ASSOCIATION TEST TIAN ZHENG, SHAW-HWA LO DEPARTMENT OF STATISTICS, COLUMBIA UNIVERSITY Abstract. In
EVALUATING THE REPEATABILITY OF TWO STUDIES OF A LARGE NUMBER OF OBJECTS: MODIFIED KENDALL RANK-ORDER ASSOCIATION TEST TIAN ZHENG, SHAW-HWA LO DEPARTMENT OF STATISTICS, COLUMBIA UNIVERSITY Abstract. In
Linear Model Selection and Regularization
 Linear Model Selection and Regularization Recall the linear model Y = β 0 + β 1 X 1 + + β p X p + ɛ. In the lectures that follow, we consider some approaches for extending the linear model framework. In
Linear Model Selection and Regularization Recall the linear model Y = β 0 + β 1 X 1 + + β p X p + ɛ. In the lectures that follow, we consider some approaches for extending the linear model framework. In
Sparse inverse covariance estimation with the lasso
 Sparse inverse covariance estimation with the lasso Jerome Friedman Trevor Hastie and Robert Tibshirani November 8, 2007 Abstract We consider the problem of estimating sparse graphs by a lasso penalty
Sparse inverse covariance estimation with the lasso Jerome Friedman Trevor Hastie and Robert Tibshirani November 8, 2007 Abstract We consider the problem of estimating sparse graphs by a lasso penalty
application in microarrays
 Biostatistics Advance Access published April 7, 2006 Regularized linear discriminant analysis and its application in microarrays Yaqian Guo, Trevor Hastie and Robert Tibshirani Abstract In this paper,
Biostatistics Advance Access published April 7, 2006 Regularized linear discriminant analysis and its application in microarrays Yaqian Guo, Trevor Hastie and Robert Tibshirani Abstract In this paper,
Classification 2: Linear discriminant analysis (continued); logistic regression
 Classification 2: Linear discriminant analysis (continued); logistic regression Ryan Tibshirani Data Mining: 36-462/36-662 April 4 2013 Optional reading: ISL 4.4, ESL 4.3; ISL 4.3, ESL 4.4 1 Reminder:
Classification 2: Linear discriminant analysis (continued); logistic regression Ryan Tibshirani Data Mining: 36-462/36-662 April 4 2013 Optional reading: ISL 4.4, ESL 4.3; ISL 4.3, ESL 4.4 1 Reminder:
Regularized Discriminant Analysis. Part I. Linear and Quadratic Discriminant Analysis. Discriminant Analysis. Example. Example. Class distribution
 Part I 09.06.2006 Discriminant Analysis The purpose of discriminant analysis is to assign objects to one of several (K) groups based on a set of measurements X = (X 1, X 2,..., X p ) which are obtained
Part I 09.06.2006 Discriminant Analysis The purpose of discriminant analysis is to assign objects to one of several (K) groups based on a set of measurements X = (X 1, X 2,..., X p ) which are obtained
FEATURE SCREENING IN ULTRAHIGH DIMENSIONAL
 Statistica Sinica 26 (2016), 881-901 doi:http://dx.doi.org/10.5705/ss.2014.171 FEATURE SCREENING IN ULTRAHIGH DIMENSIONAL COX S MODEL Guangren Yang 1, Ye Yu 2, Runze Li 2 and Anne Buu 3 1 Jinan University,
Statistica Sinica 26 (2016), 881-901 doi:http://dx.doi.org/10.5705/ss.2014.171 FEATURE SCREENING IN ULTRAHIGH DIMENSIONAL COX S MODEL Guangren Yang 1, Ye Yu 2, Runze Li 2 and Anne Buu 3 1 Jinan University,
Classification. Classification is similar to regression in that the goal is to use covariates to predict on outcome.
 Classification Classification is similar to regression in that the goal is to use covariates to predict on outcome. We still have a vector of covariates X. However, the response is binary (or a few classes),
Classification Classification is similar to regression in that the goal is to use covariates to predict on outcome. We still have a vector of covariates X. However, the response is binary (or a few classes),
Prediction by supervised principal components
 Prediction by supervised principal components Eric Bair Trevor Hastie Debashis Paul and Robert Tibshirani September 15, 2004 Summary In regression problems where the number of predictors greatly exceeds
Prediction by supervised principal components Eric Bair Trevor Hastie Debashis Paul and Robert Tibshirani September 15, 2004 Summary In regression problems where the number of predictors greatly exceeds
On testing the significance of sets of genes
 On testing the significance of sets of genes Bradley Efron and Robert Tibshirani August 17, 2006 Abstract This paper discusses the problem of identifying differentially expressed groups of genes from a
On testing the significance of sets of genes Bradley Efron and Robert Tibshirani August 17, 2006 Abstract This paper discusses the problem of identifying differentially expressed groups of genes from a
Regularized Discriminant Analysis and Its Application in Microarray
 Regularized Discriminant Analysis and Its Application in Microarray Yaqian Guo, Trevor Hastie and Robert Tibshirani May 5, 2004 Abstract In this paper, we introduce a family of some modified versions of
Regularized Discriminant Analysis and Its Application in Microarray Yaqian Guo, Trevor Hastie and Robert Tibshirani May 5, 2004 Abstract In this paper, we introduce a family of some modified versions of
Pre-conditioning for feature selection and. regression in high-dimensional problems
 Pre-conditioning for feature selection and regression in high-dimensional problems Debashis Paul Eric Bair Trevor Hastie Robert Tibshirani December 1, 2006 Abstract We consider regression problems where
Pre-conditioning for feature selection and regression in high-dimensional problems Debashis Paul Eric Bair Trevor Hastie Robert Tibshirani December 1, 2006 Abstract We consider regression problems where
Chris Fraley and Daniel Percival. August 22, 2008, revised May 14, 2010
 Model-Averaged l 1 Regularization using Markov Chain Monte Carlo Model Composition Technical Report No. 541 Department of Statistics, University of Washington Chris Fraley and Daniel Percival August 22,
Model-Averaged l 1 Regularization using Markov Chain Monte Carlo Model Composition Technical Report No. 541 Department of Statistics, University of Washington Chris Fraley and Daniel Percival August 22,
Regularization Path Algorithms for Detecting Gene Interactions
 Regularization Path Algorithms for Detecting Gene Interactions Mee Young Park Trevor Hastie July 16, 2006 Abstract In this study, we consider several regularization path algorithms with grouped variable
Regularization Path Algorithms for Detecting Gene Interactions Mee Young Park Trevor Hastie July 16, 2006 Abstract In this study, we consider several regularization path algorithms with grouped variable
Penalized classification using Fisher s linear discriminant
 J. R. Statist. Soc. B (2011) 73, Part 5, pp. 753 772 Penalized classification using Fisher s linear discriminant Daniela M. Witten University of Washington, Seattle, USA and Robert Tibshirani Stanford
J. R. Statist. Soc. B (2011) 73, Part 5, pp. 753 772 Penalized classification using Fisher s linear discriminant Daniela M. Witten University of Washington, Seattle, USA and Robert Tibshirani Stanford
Regularization in Cox Frailty Models
 Regularization in Cox Frailty Models Andreas Groll 1, Trevor Hastie 2, Gerhard Tutz 3 1 Ludwig-Maximilians-Universität Munich, Department of Mathematics, Theresienstraße 39, 80333 Munich, Germany 2 University
Regularization in Cox Frailty Models Andreas Groll 1, Trevor Hastie 2, Gerhard Tutz 3 1 Ludwig-Maximilians-Universität Munich, Department of Mathematics, Theresienstraße 39, 80333 Munich, Germany 2 University
Sparse Discriminant Analysis
 Sparse Discriminant Analysis Line Clemmensen Trevor Hastie Daniela Witten + Bjarne Ersbøll Department of Informatics and Mathematical Modelling, Technical University of Denmark, Kgs. Lyngby, Denmark Department
Sparse Discriminant Analysis Line Clemmensen Trevor Hastie Daniela Witten + Bjarne Ersbøll Department of Informatics and Mathematical Modelling, Technical University of Denmark, Kgs. Lyngby, Denmark Department
Feature Screening in Ultrahigh Dimensional Cox s Model
 Feature Screening in Ultrahigh Dimensional Cox s Model Guangren Yang School of Economics, Jinan University, Guangzhou, P.R. China Ye Yu Runze Li Department of Statistics, Penn State Anne Buu Indiana University
Feature Screening in Ultrahigh Dimensional Cox s Model Guangren Yang School of Economics, Jinan University, Guangzhou, P.R. China Ye Yu Runze Li Department of Statistics, Penn State Anne Buu Indiana University
Other likelihoods. Patrick Breheny. April 25. Multinomial regression Robust regression Cox regression
 Other likelihoods Patrick Breheny April 25 Patrick Breheny High-Dimensional Data Analysis (BIOS 7600) 1/29 Introduction In principle, the idea of penalized regression can be extended to any sort of regression
Other likelihoods Patrick Breheny April 25 Patrick Breheny High-Dimensional Data Analysis (BIOS 7600) 1/29 Introduction In principle, the idea of penalized regression can be extended to any sort of regression
Variable Selection for Highly Correlated Predictors
 Variable Selection for Highly Correlated Predictors Fei Xue and Annie Qu Department of Statistics, University of Illinois at Urbana-Champaign WHOA-PSI, Aug, 2017 St. Louis, Missouri 1 / 30 Background Variable
Variable Selection for Highly Correlated Predictors Fei Xue and Annie Qu Department of Statistics, University of Illinois at Urbana-Champaign WHOA-PSI, Aug, 2017 St. Louis, Missouri 1 / 30 Background Variable
Classification 1: Linear regression of indicators, linear discriminant analysis
 Classification 1: Linear regression of indicators, linear discriminant analysis Ryan Tibshirani Data Mining: 36-462/36-662 April 2 2013 Optional reading: ISL 4.1, 4.2, 4.4, ESL 4.1 4.3 1 Classification
Classification 1: Linear regression of indicators, linear discriminant analysis Ryan Tibshirani Data Mining: 36-462/36-662 April 2 2013 Optional reading: ISL 4.1, 4.2, 4.4, ESL 4.1 4.3 1 Classification
Survival Prediction Under Dependent Censoring: A Copula-based Approach
 Survival Prediction Under Dependent Censoring: A Copula-based Approach Yi-Hau Chen Institute of Statistical Science, Academia Sinica 2013 AMMS, National Sun Yat-Sen University December 7 2013 Joint work
Survival Prediction Under Dependent Censoring: A Copula-based Approach Yi-Hau Chen Institute of Statistical Science, Academia Sinica 2013 AMMS, National Sun Yat-Sen University December 7 2013 Joint work
Pre-Selection in Cluster Lasso Methods for Correlated Variable Selection in High-Dimensional Linear Models
 Pre-Selection in Cluster Lasso Methods for Correlated Variable Selection in High-Dimensional Linear Models Niharika Gauraha and Swapan Parui Indian Statistical Institute Abstract. We consider variable
Pre-Selection in Cluster Lasso Methods for Correlated Variable Selection in High-Dimensional Linear Models Niharika Gauraha and Swapan Parui Indian Statistical Institute Abstract. We consider variable
STAT331. Cox s Proportional Hazards Model
 STAT331 Cox s Proportional Hazards Model In this unit we introduce Cox s proportional hazards (Cox s PH) model, give a heuristic development of the partial likelihood function, and discuss adaptations
STAT331 Cox s Proportional Hazards Model In this unit we introduce Cox s proportional hazards (Cox s PH) model, give a heuristic development of the partial likelihood function, and discuss adaptations
The miss rate for the analysis of gene expression data
 Biostatistics (2005), 6, 1,pp. 111 117 doi: 10.1093/biostatistics/kxh021 The miss rate for the analysis of gene expression data JONATHAN TAYLOR Department of Statistics, Stanford University, Stanford,
Biostatistics (2005), 6, 1,pp. 111 117 doi: 10.1093/biostatistics/kxh021 The miss rate for the analysis of gene expression data JONATHAN TAYLOR Department of Statistics, Stanford University, Stanford,
Chapter 3. Linear Models for Regression
 Chapter 3. Linear Models for Regression Wei Pan Division of Biostatistics, School of Public Health, University of Minnesota, Minneapolis, MN 55455 Email: weip@biostat.umn.edu PubH 7475/8475 c Wei Pan Linear
Chapter 3. Linear Models for Regression Wei Pan Division of Biostatistics, School of Public Health, University of Minnesota, Minneapolis, MN 55455 Email: weip@biostat.umn.edu PubH 7475/8475 c Wei Pan Linear
Fast Regularization Paths via Coordinate Descent
 KDD August 2008 Trevor Hastie, Stanford Statistics 1 Fast Regularization Paths via Coordinate Descent Trevor Hastie Stanford University joint work with Jerry Friedman and Rob Tibshirani. KDD August 2008
KDD August 2008 Trevor Hastie, Stanford Statistics 1 Fast Regularization Paths via Coordinate Descent Trevor Hastie Stanford University joint work with Jerry Friedman and Rob Tibshirani. KDD August 2008
Effective Linear Discriminant Analysis for High Dimensional, Low Sample Size Data
 Effective Linear Discriant Analysis for High Dimensional, Low Sample Size Data Zhihua Qiao, Lan Zhou and Jianhua Z. Huang Abstract In the so-called high dimensional, low sample size (HDLSS) settings, LDA
Effective Linear Discriant Analysis for High Dimensional, Low Sample Size Data Zhihua Qiao, Lan Zhou and Jianhua Z. Huang Abstract In the so-called high dimensional, low sample size (HDLSS) settings, LDA
An Introduction to Graphical Lasso
 An Introduction to Graphical Lasso Bo Chang Graphical Models Reading Group May 15, 2015 Bo Chang (UBC) Graphical Lasso May 15, 2015 1 / 16 Undirected Graphical Models An undirected graph, each vertex represents
An Introduction to Graphical Lasso Bo Chang Graphical Models Reading Group May 15, 2015 Bo Chang (UBC) Graphical Lasso May 15, 2015 1 / 16 Undirected Graphical Models An undirected graph, each vertex represents
Bayesian variable selection via. Penalized credible regions. Brian Reich, NCSU. Joint work with. Howard Bondell and Ander Wilson
 Bayesian variable selection via penalized credible regions Brian Reich, NC State Joint work with Howard Bondell and Ander Wilson Brian Reich, NCSU Penalized credible regions 1 Motivation big p, small n
Bayesian variable selection via penalized credible regions Brian Reich, NC State Joint work with Howard Bondell and Ander Wilson Brian Reich, NCSU Penalized credible regions 1 Motivation big p, small n
Pathwise coordinate optimization
 Stanford University 1 Pathwise coordinate optimization Jerome Friedman, Trevor Hastie, Holger Hoefling, Robert Tibshirani Stanford University Acknowledgements: Thanks to Stephen Boyd, Michael Saunders,
Stanford University 1 Pathwise coordinate optimization Jerome Friedman, Trevor Hastie, Holger Hoefling, Robert Tibshirani Stanford University Acknowledgements: Thanks to Stephen Boyd, Michael Saunders,
Modelling geoadditive survival data
 Modelling geoadditive survival data Thomas Kneib & Ludwig Fahrmeir Department of Statistics, Ludwig-Maximilians-University Munich 1. Leukemia survival data 2. Structured hazard regression 3. Mixed model
Modelling geoadditive survival data Thomas Kneib & Ludwig Fahrmeir Department of Statistics, Ludwig-Maximilians-University Munich 1. Leukemia survival data 2. Structured hazard regression 3. Mixed model
Stability and the elastic net
 Stability and the elastic net Patrick Breheny March 28 Patrick Breheny High-Dimensional Data Analysis (BIOS 7600) 1/32 Introduction Elastic Net Our last several lectures have concentrated on methods for
Stability and the elastic net Patrick Breheny March 28 Patrick Breheny High-Dimensional Data Analysis (BIOS 7600) 1/32 Introduction Elastic Net Our last several lectures have concentrated on methods for
Generalized Boosted Models: A guide to the gbm package
 Generalized Boosted Models: A guide to the gbm package Greg Ridgeway April 15, 2006 Boosting takes on various forms with different programs using different loss functions, different base models, and different
Generalized Boosted Models: A guide to the gbm package Greg Ridgeway April 15, 2006 Boosting takes on various forms with different programs using different loss functions, different base models, and different
Graphical Model Selection
 May 6, 2013 Trevor Hastie, Stanford Statistics 1 Graphical Model Selection Trevor Hastie Stanford University joint work with Jerome Friedman, Rob Tibshirani, Rahul Mazumder and Jason Lee May 6, 2013 Trevor
May 6, 2013 Trevor Hastie, Stanford Statistics 1 Graphical Model Selection Trevor Hastie Stanford University joint work with Jerome Friedman, Rob Tibshirani, Rahul Mazumder and Jason Lee May 6, 2013 Trevor
EXAM IN STATISTICAL MACHINE LEARNING STATISTISK MASKININLÄRNING
 EXAM IN STATISTICAL MACHINE LEARNING STATISTISK MASKININLÄRNING DATE AND TIME: August 30, 2018, 14.00 19.00 RESPONSIBLE TEACHER: Niklas Wahlström NUMBER OF PROBLEMS: 5 AIDING MATERIAL: Calculator, mathematical
EXAM IN STATISTICAL MACHINE LEARNING STATISTISK MASKININLÄRNING DATE AND TIME: August 30, 2018, 14.00 19.00 RESPONSIBLE TEACHER: Niklas Wahlström NUMBER OF PROBLEMS: 5 AIDING MATERIAL: Calculator, mathematical
Least Absolute Shrinkage is Equivalent to Quadratic Penalization
 Least Absolute Shrinkage is Equivalent to Quadratic Penalization Yves Grandvalet Heudiasyc, UMR CNRS 6599, Université de Technologie de Compiègne, BP 20.529, 60205 Compiègne Cedex, France Yves.Grandvalet@hds.utc.fr
Least Absolute Shrinkage is Equivalent to Quadratic Penalization Yves Grandvalet Heudiasyc, UMR CNRS 6599, Université de Technologie de Compiègne, BP 20.529, 60205 Compiègne Cedex, France Yves.Grandvalet@hds.utc.fr
TECHNICAL REPORT NO. 1091r. A Note on the Lasso and Related Procedures in Model Selection
 DEPARTMENT OF STATISTICS University of Wisconsin 1210 West Dayton St. Madison, WI 53706 TECHNICAL REPORT NO. 1091r April 2004, Revised December 2004 A Note on the Lasso and Related Procedures in Model
DEPARTMENT OF STATISTICS University of Wisconsin 1210 West Dayton St. Madison, WI 53706 TECHNICAL REPORT NO. 1091r April 2004, Revised December 2004 A Note on the Lasso and Related Procedures in Model
MSA220/MVE440 Statistical Learning for Big Data
 MSA220/MVE440 Statistical Learning for Big Data Lecture 7/8 - High-dimensional modeling part 1 Rebecka Jörnsten Mathematical Sciences University of Gothenburg and Chalmers University of Technology Classification
MSA220/MVE440 Statistical Learning for Big Data Lecture 7/8 - High-dimensional modeling part 1 Rebecka Jörnsten Mathematical Sciences University of Gothenburg and Chalmers University of Technology Classification
Reduced-rank hazard regression
 Chapter 2 Reduced-rank hazard regression Abstract The Cox proportional hazards model is the most common method to analyze survival data. However, the proportional hazards assumption might not hold. The
Chapter 2 Reduced-rank hazard regression Abstract The Cox proportional hazards model is the most common method to analyze survival data. However, the proportional hazards assumption might not hold. The
A significance test for the lasso
 1 First part: Joint work with Richard Lockhart (SFU), Jonathan Taylor (Stanford), and Ryan Tibshirani (Carnegie-Mellon Univ.) Second part: Joint work with Max Grazier G Sell, Stefan Wager and Alexandra
1 First part: Joint work with Richard Lockhart (SFU), Jonathan Taylor (Stanford), and Ryan Tibshirani (Carnegie-Mellon Univ.) Second part: Joint work with Max Grazier G Sell, Stefan Wager and Alexandra
Classification. Chapter Introduction. 6.2 The Bayes classifier
 Chapter 6 Classification 6.1 Introduction Often encountered in applications is the situation where the response variable Y takes values in a finite set of labels. For example, the response Y could encode
Chapter 6 Classification 6.1 Introduction Often encountered in applications is the situation where the response variable Y takes values in a finite set of labels. For example, the response Y could encode
Accepted author version posted online: 28 Nov 2012.
 This article was downloaded by: [University of Minnesota Libraries, Twin Cities] On: 15 May 2013, At: 12:23 Publisher: Taylor & Francis Informa Ltd Registered in England and Wales Registered Number: 1072954
This article was downloaded by: [University of Minnesota Libraries, Twin Cities] On: 15 May 2013, At: 12:23 Publisher: Taylor & Francis Informa Ltd Registered in England and Wales Registered Number: 1072954
Proteomics and Variable Selection
 Proteomics and Variable Selection p. 1/55 Proteomics and Variable Selection Alex Lewin With thanks to Paul Kirk for some graphs Department of Epidemiology and Biostatistics, School of Public Health, Imperial
Proteomics and Variable Selection p. 1/55 Proteomics and Variable Selection Alex Lewin With thanks to Paul Kirk for some graphs Department of Epidemiology and Biostatistics, School of Public Health, Imperial
Regression Model In The Analysis Of Micro Array Data-Gene Expression Detection
 Jamal Fathima.J.I 1 and P.Venkatesan 1. Research Scholar -Department of statistics National Institute For Research In Tuberculosis, Indian Council For Medical Research,Chennai,India,.Department of statistics
Jamal Fathima.J.I 1 and P.Venkatesan 1. Research Scholar -Department of statistics National Institute For Research In Tuberculosis, Indian Council For Medical Research,Chennai,India,.Department of statistics
Selection of Smoothing Parameter for One-Step Sparse Estimates with L q Penalty
 Journal of Data Science 9(2011), 549-564 Selection of Smoothing Parameter for One-Step Sparse Estimates with L q Penalty Masaru Kanba and Kanta Naito Shimane University Abstract: This paper discusses the
Journal of Data Science 9(2011), 549-564 Selection of Smoothing Parameter for One-Step Sparse Estimates with L q Penalty Masaru Kanba and Kanta Naito Shimane University Abstract: This paper discusses the
Properties of optimizations used in penalized Gaussian likelihood inverse covariance matrix estimation
 Properties of optimizations used in penalized Gaussian likelihood inverse covariance matrix estimation Adam J. Rothman School of Statistics University of Minnesota October 8, 2014, joint work with Liliana
Properties of optimizations used in penalized Gaussian likelihood inverse covariance matrix estimation Adam J. Rothman School of Statistics University of Minnesota October 8, 2014, joint work with Liliana
Sparse regression. Optimization-Based Data Analysis. Carlos Fernandez-Granda
 Sparse regression Optimization-Based Data Analysis http://www.cims.nyu.edu/~cfgranda/pages/obda_spring16 Carlos Fernandez-Granda 3/28/2016 Regression Least-squares regression Example: Global warming Logistic
Sparse regression Optimization-Based Data Analysis http://www.cims.nyu.edu/~cfgranda/pages/obda_spring16 Carlos Fernandez-Granda 3/28/2016 Regression Least-squares regression Example: Global warming Logistic
Consistent high-dimensional Bayesian variable selection via penalized credible regions
 Consistent high-dimensional Bayesian variable selection via penalized credible regions Howard Bondell bondell@stat.ncsu.edu Joint work with Brian Reich Howard Bondell p. 1 Outline High-Dimensional Variable
Consistent high-dimensional Bayesian variable selection via penalized credible regions Howard Bondell bondell@stat.ncsu.edu Joint work with Brian Reich Howard Bondell p. 1 Outline High-Dimensional Variable
High-dimensional regression
 High-dimensional regression Advanced Methods for Data Analysis 36-402/36-608) Spring 2014 1 Back to linear regression 1.1 Shortcomings Suppose that we are given outcome measurements y 1,... y n R, and
High-dimensional regression Advanced Methods for Data Analysis 36-402/36-608) Spring 2014 1 Back to linear regression 1.1 Shortcomings Suppose that we are given outcome measurements y 1,... y n R, and
Post-selection inference with an application to internal inference
 Post-selection inference with an application to internal inference Robert Tibshirani, Stanford University November 23, 2015 Seattle Symposium in Biostatistics, 2015 Joint work with Sam Gross, Will Fithian,
Post-selection inference with an application to internal inference Robert Tibshirani, Stanford University November 23, 2015 Seattle Symposium in Biostatistics, 2015 Joint work with Sam Gross, Will Fithian,
A significance test for the lasso
 1 Gold medal address, SSC 2013 Joint work with Richard Lockhart (SFU), Jonathan Taylor (Stanford), and Ryan Tibshirani (Carnegie-Mellon Univ.) Reaping the benefits of LARS: A special thanks to Brad Efron,
1 Gold medal address, SSC 2013 Joint work with Richard Lockhart (SFU), Jonathan Taylor (Stanford), and Ryan Tibshirani (Carnegie-Mellon Univ.) Reaping the benefits of LARS: A special thanks to Brad Efron,
Regression Shrinkage and Selection via the Elastic Net, with Applications to Microarrays
 Regression Shrinkage and Selection via the Elastic Net, with Applications to Microarrays Hui Zou and Trevor Hastie Department of Statistics, Stanford University December 5, 2003 Abstract We propose the
Regression Shrinkage and Selection via the Elastic Net, with Applications to Microarrays Hui Zou and Trevor Hastie Department of Statistics, Stanford University December 5, 2003 Abstract We propose the
Tuning Parameter Selection in Regularized Estimations of Large Covariance Matrices
 Tuning Parameter Selection in Regularized Estimations of Large Covariance Matrices arxiv:1308.3416v1 [stat.me] 15 Aug 2013 Yixin Fang 1, Binhuan Wang 1, and Yang Feng 2 1 New York University and 2 Columbia
Tuning Parameter Selection in Regularized Estimations of Large Covariance Matrices arxiv:1308.3416v1 [stat.me] 15 Aug 2013 Yixin Fang 1, Binhuan Wang 1, and Yang Feng 2 1 New York University and 2 Columbia
ISyE 6416: Computational Statistics Spring Lecture 5: Discriminant analysis and classification
 ISyE 6416: Computational Statistics Spring 2017 Lecture 5: Discriminant analysis and classification Prof. Yao Xie H. Milton Stewart School of Industrial and Systems Engineering Georgia Institute of Technology
ISyE 6416: Computational Statistics Spring 2017 Lecture 5: Discriminant analysis and classification Prof. Yao Xie H. Milton Stewart School of Industrial and Systems Engineering Georgia Institute of Technology
Classification Methods II: Linear and Quadratic Discrimminant Analysis
 Classification Methods II: Linear and Quadratic Discrimminant Analysis Rebecca C. Steorts, Duke University STA 325, Chapter 4 ISL Agenda Linear Discrimminant Analysis (LDA) Classification Recall that linear
Classification Methods II: Linear and Quadratic Discrimminant Analysis Rebecca C. Steorts, Duke University STA 325, Chapter 4 ISL Agenda Linear Discrimminant Analysis (LDA) Classification Recall that linear
An Algorithm for Bayesian Variable Selection in High-dimensional Generalized Linear Models
 Proceedings 59th ISI World Statistics Congress, 25-30 August 2013, Hong Kong (Session CPS023) p.3938 An Algorithm for Bayesian Variable Selection in High-dimensional Generalized Linear Models Vitara Pungpapong
Proceedings 59th ISI World Statistics Congress, 25-30 August 2013, Hong Kong (Session CPS023) p.3938 An Algorithm for Bayesian Variable Selection in High-dimensional Generalized Linear Models Vitara Pungpapong
Estimating subgroup specific treatment effects via concave fusion
 Estimating subgroup specific treatment effects via concave fusion Jian Huang University of Iowa April 6, 2016 Outline 1 Motivation and the problem 2 The proposed model and approach Concave pairwise fusion
Estimating subgroup specific treatment effects via concave fusion Jian Huang University of Iowa April 6, 2016 Outline 1 Motivation and the problem 2 The proposed model and approach Concave pairwise fusion
MAS3301 / MAS8311 Biostatistics Part II: Survival
 MAS3301 / MAS8311 Biostatistics Part II: Survival M. Farrow School of Mathematics and Statistics Newcastle University Semester 2, 2009-10 1 13 The Cox proportional hazards model 13.1 Introduction In the
MAS3301 / MAS8311 Biostatistics Part II: Survival M. Farrow School of Mathematics and Statistics Newcastle University Semester 2, 2009-10 1 13 The Cox proportional hazards model 13.1 Introduction In the
Final Overview. Introduction to ML. Marek Petrik 4/25/2017
 Final Overview Introduction to ML Marek Petrik 4/25/2017 This Course: Introduction to Machine Learning Build a foundation for practice and research in ML Basic machine learning concepts: max likelihood,
Final Overview Introduction to ML Marek Petrik 4/25/2017 This Course: Introduction to Machine Learning Build a foundation for practice and research in ML Basic machine learning concepts: max likelihood,
High-dimensional regression modeling
 High-dimensional regression modeling David Causeur Department of Statistics and Computer Science Agrocampus Ouest IRMAR CNRS UMR 6625 http://www.agrocampus-ouest.fr/math/causeur/ Course objectives Making
High-dimensional regression modeling David Causeur Department of Statistics and Computer Science Agrocampus Ouest IRMAR CNRS UMR 6625 http://www.agrocampus-ouest.fr/math/causeur/ Course objectives Making
SUPPLEMENTARY APPENDICES FOR WAVELET-DOMAIN REGRESSION AND PREDICTIVE INFERENCE IN PSYCHIATRIC NEUROIMAGING
 Submitted to the Annals of Applied Statistics SUPPLEMENTARY APPENDICES FOR WAVELET-DOMAIN REGRESSION AND PREDICTIVE INFERENCE IN PSYCHIATRIC NEUROIMAGING By Philip T. Reiss, Lan Huo, Yihong Zhao, Clare
Submitted to the Annals of Applied Statistics SUPPLEMENTARY APPENDICES FOR WAVELET-DOMAIN REGRESSION AND PREDICTIVE INFERENCE IN PSYCHIATRIC NEUROIMAGING By Philip T. Reiss, Lan Huo, Yihong Zhao, Clare
Machine Learning for OR & FE
 Machine Learning for OR & FE Regression II: Regularization and Shrinkage Methods Martin Haugh Department of Industrial Engineering and Operations Research Columbia University Email: martin.b.haugh@gmail.com
Machine Learning for OR & FE Regression II: Regularization and Shrinkage Methods Martin Haugh Department of Industrial Engineering and Operations Research Columbia University Email: martin.b.haugh@gmail.com
Lecture 6 PREDICTING SURVIVAL UNDER THE PH MODEL
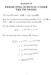 Lecture 6 PREDICTING SURVIVAL UNDER THE PH MODEL The Cox PH model: λ(t Z) = λ 0 (t) exp(β Z). How do we estimate the survival probability, S z (t) = S(t Z) = P (T > t Z), for an individual with covariates
Lecture 6 PREDICTING SURVIVAL UNDER THE PH MODEL The Cox PH model: λ(t Z) = λ 0 (t) exp(β Z). How do we estimate the survival probability, S z (t) = S(t Z) = P (T > t Z), for an individual with covariates
Permutation-invariant regularization of large covariance matrices. Liza Levina
 Liza Levina Permutation-invariant covariance regularization 1/42 Permutation-invariant regularization of large covariance matrices Liza Levina Department of Statistics University of Michigan Joint work
Liza Levina Permutation-invariant covariance regularization 1/42 Permutation-invariant regularization of large covariance matrices Liza Levina Department of Statistics University of Michigan Joint work
A Significance Test for the Lasso
 A Significance Test for the Lasso Lockhart R, Taylor J, Tibshirani R, and Tibshirani R Ashley Petersen May 14, 2013 1 Last time Problem: Many clinical covariates which are important to a certain medical
A Significance Test for the Lasso Lockhart R, Taylor J, Tibshirani R, and Tibshirani R Ashley Petersen May 14, 2013 1 Last time Problem: Many clinical covariates which are important to a certain medical
TGDR: An Introduction
 TGDR: An Introduction Julian Wolfson Student Seminar March 28, 2007 1 Variable Selection 2 Penalization, Solution Paths and TGDR 3 Applying TGDR 4 Extensions 5 Final Thoughts Some motivating examples We
TGDR: An Introduction Julian Wolfson Student Seminar March 28, 2007 1 Variable Selection 2 Penalization, Solution Paths and TGDR 3 Applying TGDR 4 Extensions 5 Final Thoughts Some motivating examples We
Robust Bayesian Variable Selection for Modeling Mean Medical Costs
 Robust Bayesian Variable Selection for Modeling Mean Medical Costs Grace Yoon 1,, Wenxin Jiang 2, Lei Liu 3 and Ya-Chen T. Shih 4 1 Department of Statistics, Texas A&M University 2 Department of Statistics,
Robust Bayesian Variable Selection for Modeling Mean Medical Costs Grace Yoon 1,, Wenxin Jiang 2, Lei Liu 3 and Ya-Chen T. Shih 4 1 Department of Statistics, Texas A&M University 2 Department of Statistics,
ESL Chap3. Some extensions of lasso
 ESL Chap3 Some extensions of lasso 1 Outline Consistency of lasso for model selection Adaptive lasso Elastic net Group lasso 2 Consistency of lasso for model selection A number of authors have studied
ESL Chap3 Some extensions of lasso 1 Outline Consistency of lasso for model selection Adaptive lasso Elastic net Group lasso 2 Consistency of lasso for model selection A number of authors have studied
Smoothly Clipped Absolute Deviation (SCAD) for Correlated Variables
 Smoothly Clipped Absolute Deviation (SCAD) for Correlated Variables LIB-MA, FSSM Cadi Ayyad University (Morocco) COMPSTAT 2010 Paris, August 22-27, 2010 Motivations Fan and Li (2001), Zou and Li (2008)
Smoothly Clipped Absolute Deviation (SCAD) for Correlated Variables LIB-MA, FSSM Cadi Ayyad University (Morocco) COMPSTAT 2010 Paris, August 22-27, 2010 Motivations Fan and Li (2001), Zou and Li (2008)
MSA200/TMS041 Multivariate Analysis
 MSA200/TMS041 Multivariate Analysis Lecture 8 Rebecka Jörnsten Mathematical Sciences University of Gothenburg and Chalmers University of Technology Back to Discriminant analysis As mentioned in the previous
MSA200/TMS041 Multivariate Analysis Lecture 8 Rebecka Jörnsten Mathematical Sciences University of Gothenburg and Chalmers University of Technology Back to Discriminant analysis As mentioned in the previous
Ensemble estimation and variable selection with semiparametric regression models
 Ensemble estimation and variable selection with semiparametric regression models Sunyoung Shin Department of Mathematical Sciences University of Texas at Dallas Joint work with Jason Fine, Yufeng Liu,
Ensemble estimation and variable selection with semiparametric regression models Sunyoung Shin Department of Mathematical Sciences University of Texas at Dallas Joint work with Jason Fine, Yufeng Liu,
The exam is closed book, closed notes except your one-page (two sides) or two-page (one side) crib sheet.
 CS 189 Spring 013 Introduction to Machine Learning Final You have 3 hours for the exam. The exam is closed book, closed notes except your one-page (two sides) or two-page (one side) crib sheet. Please
CS 189 Spring 013 Introduction to Machine Learning Final You have 3 hours for the exam. The exam is closed book, closed notes except your one-page (two sides) or two-page (one side) crib sheet. Please
UNIVERSITY of PENNSYLVANIA CIS 520: Machine Learning Final, Fall 2013
 UNIVERSITY of PENNSYLVANIA CIS 520: Machine Learning Final, Fall 2013 Exam policy: This exam allows two one-page, two-sided cheat sheets; No other materials. Time: 2 hours. Be sure to write your name and
UNIVERSITY of PENNSYLVANIA CIS 520: Machine Learning Final, Fall 2013 Exam policy: This exam allows two one-page, two-sided cheat sheets; No other materials. Time: 2 hours. Be sure to write your name and
Part III. Hypothesis Testing. III.1. Log-rank Test for Right-censored Failure Time Data
 1 Part III. Hypothesis Testing III.1. Log-rank Test for Right-censored Failure Time Data Consider a survival study consisting of n independent subjects from p different populations with survival functions
1 Part III. Hypothesis Testing III.1. Log-rank Test for Right-censored Failure Time Data Consider a survival study consisting of n independent subjects from p different populations with survival functions
Compressed Sensing in Cancer Biology? (A Work in Progress)
 Compressed Sensing in Cancer Biology? (A Work in Progress) M. Vidyasagar FRS Cecil & Ida Green Chair The University of Texas at Dallas M.Vidyasagar@utdallas.edu www.utdallas.edu/ m.vidyasagar University
Compressed Sensing in Cancer Biology? (A Work in Progress) M. Vidyasagar FRS Cecil & Ida Green Chair The University of Texas at Dallas M.Vidyasagar@utdallas.edu www.utdallas.edu/ m.vidyasagar University
Generalized Elastic Net Regression
 Abstract Generalized Elastic Net Regression Geoffroy MOURET Jean-Jules BRAULT Vahid PARTOVINIA This work presents a variation of the elastic net penalization method. We propose applying a combined l 1
Abstract Generalized Elastic Net Regression Geoffroy MOURET Jean-Jules BRAULT Vahid PARTOVINIA This work presents a variation of the elastic net penalization method. We propose applying a combined l 1
Machine Learning Linear Classification. Prof. Matteo Matteucci
 Machine Learning Linear Classification Prof. Matteo Matteucci Recall from the first lecture 2 X R p Regression Y R Continuous Output X R p Y {Ω 0, Ω 1,, Ω K } Classification Discrete Output X R p Y (X)
Machine Learning Linear Classification Prof. Matteo Matteucci Recall from the first lecture 2 X R p Regression Y R Continuous Output X R p Y {Ω 0, Ω 1,, Ω K } Classification Discrete Output X R p Y (X)
Fused elastic net EPoC
 Fused elastic net EPoC Tobias Abenius November 28, 2013 Data Set and Method Use mrna and copy number aberration (CNA) to build a network using an extension of our EPoC method (Jörnsten et al., 2011; Abenius
Fused elastic net EPoC Tobias Abenius November 28, 2013 Data Set and Method Use mrna and copy number aberration (CNA) to build a network using an extension of our EPoC method (Jörnsten et al., 2011; Abenius
Pre-validation and inference in microarrays
 Pre-validation and inference in microarrays Robert Tibshirani and Bradley Efron June 26, 2002 Abstract In microarray studies, an important problem is to compare a predictor of disease outcome derived from
Pre-validation and inference in microarrays Robert Tibshirani and Bradley Efron June 26, 2002 Abstract In microarray studies, an important problem is to compare a predictor of disease outcome derived from
Sparse statistical modelling
 Sparse statistical modelling Tom Bartlett Sparse statistical modelling Tom Bartlett 1 / 28 Introduction A sparse statistical model is one having only a small number of nonzero parameters or weights. [1]
Sparse statistical modelling Tom Bartlett Sparse statistical modelling Tom Bartlett 1 / 28 Introduction A sparse statistical model is one having only a small number of nonzero parameters or weights. [1]
On Measurement Error Problems with Predictors Derived from Stationary Stochastic Processes and Application to Cocaine Dependence Treatment Data
 On Measurement Error Problems with Predictors Derived from Stationary Stochastic Processes and Application to Cocaine Dependence Treatment Data Yehua Li Department of Statistics University of Georgia Yongtao
On Measurement Error Problems with Predictors Derived from Stationary Stochastic Processes and Application to Cocaine Dependence Treatment Data Yehua Li Department of Statistics University of Georgia Yongtao
Penalized versus constrained generalized eigenvalue problems
 Penalized versus constrained generalized eigenvalue problems Irina Gaynanova, James G. Booth and Martin T. Wells. arxiv:141.6131v3 [stat.co] 4 May 215 Abstract We investigate the difference between using
Penalized versus constrained generalized eigenvalue problems Irina Gaynanova, James G. Booth and Martin T. Wells. arxiv:141.6131v3 [stat.co] 4 May 215 Abstract We investigate the difference between using
Simple techniques for comparing survival functions with interval-censored data
 Simple techniques for comparing survival functions with interval-censored data Jinheum Kim, joint with Chung Mo Nam jinhkim@suwon.ac.kr Department of Applied Statistics University of Suwon Comparing survival
Simple techniques for comparing survival functions with interval-censored data Jinheum Kim, joint with Chung Mo Nam jinhkim@suwon.ac.kr Department of Applied Statistics University of Suwon Comparing survival
Simultaneous regression shrinkage, variable selection, and supervised clustering of predictors with OSCAR
 Simultaneous regression shrinkage, variable selection, and supervised clustering of predictors with OSCAR Howard D. Bondell and Brian J. Reich Department of Statistics, North Carolina State University,
Simultaneous regression shrinkage, variable selection, and supervised clustering of predictors with OSCAR Howard D. Bondell and Brian J. Reich Department of Statistics, North Carolina State University,
