Mixed model analysis of censored longitudinal data with flexible random-effects density
|
|
|
- Bethany Stewart
- 6 years ago
- Views:
Transcription
1 Biostatistics (2012), 13, 1, pp doi: /biostatistics/kxr026 Advance Access publication on September 13, 2011 Mixed model analysis of censored longitudinal data with flexible random-effects density DAVID M. VOCK, MARIE DAVIDIAN, ANASTASIOS A. TSIATIS Department of Statistics, North Carolina State University, Raleigh, NC 27695, USA ANDREW J. MUIR Duke Clinical Research Institute, Duke University, Durham, NC 27705, USA SUMMARY Mixed models are commonly used to represent longitudinal or repeated measures data. An additional complication arises when the response is censored, for example, due to limits of quantification of the assay used. While Gaussian random effects are routinely assumed, little work has characterized the consequences of misspecifying the random-effects distribution nor has a more flexible distribution been studied for censored longitudinal data. We show that, in general, maximum likelihood estimators will not be consistent when the random-effects density is misspecified, and the effect of misspecification is likely to be greatest when the true random-effects density deviates substantially from normality and the number of noncensored observations on each subject is small. We develop a mixed model framework for censored longitudinal data in which the random effects are represented by the flexible seminonparametric density and show how to obtain estimates in SAS procedure NLMIXED. Simulations show that this approach can lead to reduction in bias and increase in efficiency relative to assuming Gaussian random effects. The methods are demonstrated on data from a study of hepatitis C virus. Keywords: Censoring; HCV; HIV; Limit of quantification; Longitudinal data; Random effects. 1. INTRODUCTION Longitudinal or repeated measures data are commonly represented by mixed effects models. A complication occurs when the response is censored for some of the observations, which often arises when assay measures are collected over time and the assay procedure is subject to limits of quantification (Hughes, 1999; Jacqmin-Gadda and others, 2000; Wu, 2002, and references therein). As an example, we consider data from the Individual Dosing Efficacy versus Flat Dosing to Assess Optimal Pegylated Interferon Therapy (IDEAL) study, which tracked the viral load progression of treatment-naive patients with hepatitis C (Thompson and others, 2010). One objective was to characterize the viral load decline in patients receiving standard treatment, pegylated interferon-alpha and ribavirin, over the first 12 weeks of treatment. Within-subjects, the trajectory of the log 10 viral load over the first 12 weeks is approximately linear, but responses were censored from below at log 10 IU/mL, To whom correspondence should be addressed. c The Author Published by Oxford University Press. All rights reserved. For permissions, please journals.permissions@oup.com.
2 62 D. M. VOCK AND OTHERS Fig. 1. Trajectory of log 10 viral load of 25 randomly selected subjects with genotype CT from the IDEAL study. For graphical purposes, the lower limit of quantification, log 10 IU/mL, was imputed for the censored responses. the lower limit of quantification of the assay used. Over the first 12 weeks, 10.5% of all observations were censored, and 35.2% were censored at week 12. The trajectories of 25 randomly selected subjects are shown in Figure 1. A standard analysis would impute the censored responses as the limit or half limit of quantification and then fit a mixed model for uncensored longitudinal data. However, Jacqmin-Gadda and others (2000) showed through simulation studies that such crude parameter estimators are badly biased even for modest levels of censoring. A refined analysis would postulate a likelihood that accounts for the censoring and obtain the maximum likelihood estimates. However, methods using this full-likelihood approach have all assumed Gaussian random effects and intra-subject error (Hughes, 1999; Jacqmin-Gadda and others, 2000; Wu, 2002; Vaida and Liu, 2009). While intra-subject error may be reasonably assumed to be normally distributed, the assumption of Gaussian random effects may be too restrictive in many applications. In the IDEAL study, some patients do not respond to treatment, while others show marked declines in the viral load, so the distribution of subject-specific slopes is unlikely to be approximately Gaussian. For noncensored linear mixed-effects models, the maximum likelihood estimators for the fixed-effects and covariance components are consistent under broad regularity conditions even when the random-effects distribution is misspecified (Verbeke and Lesaffre, 1997). For nonlinear mixed-effects models, which include mixed models with censored responses and generalized linear mixed models, the maximum likelihood estimators will not, in general, be consistent if the random-effects distribution is misspecified. The effect of misspecification has been well studied in generalized linear mixed models (Neuhaus and others, 1992; Heagerty and Kurland, 2001; Agresti and others, 2004; Litière and others, 2008), but the potential effect of misspecification of the random-effects distribution when the response is censored has not been examined. A variety of methods have been proposed to relax the Gaussian assumption for the random effects (Magder and Zeger, 1996; Verbeke and Lesaffre, 1997; Kleinman and Ibrahim, 1998; Aitkin, 1999; Zhang
3 Mixed model analysis of censored longitudinal data 63 and Davidian, 2001; Chen and others, 2002; Lee and Thompson, 2008); however, none of these methods has been implemented for censored longitudinal data. For linear mixed models, Zhang and Davidian (2001) assumed that the random effects follow a smooth density that can be represented by the seminonparametric (SNP) formulation proposed by Gallant and Nychka (1987). They showed the log-likelihood can be written in a closed form leading to straightforward estimation. We show how the SNP representation can be implemented for the random-effects density in linear mixed models when the response is potentially censored. In Section 2, we formulate the censored-response mixed model with Gaussian random effects and show that incorrectly assuming the random effects are Gaussian leads to inconsistent estimators of model parameters. In addition, we suggest general scenarios where estimation is likely to be particularly poor. In Section 3, we discuss the censored-response linear mixed model, where the random effects are assumed to follow a smooth continuous density and introduce the SNP representation. Section 4 describes how to fit the SNP model using SAS procedure NLMIXED, how to select the degree of flexility in the model, and how to choose starting values. Section 5 presents the results of several simulations. In Section 6, we illustrate the method by application to data from the IDEAL study. 2. GAUSSIAN LINEAR MIXED MODEL WITH CENSORED RESPONSE For ease of presentation, we assume that responses are potentially left censored, but developments are easily generalized to right- or interval-censored data. Let Y i j, i = 1,..., m and j = 1,..., n i be the jth response for subject i that would have been observed had there been no censoring. Consider the usual linear mixed model for longitudinal data Y i j = v T i j δ + st i j u i + ɛ i j = x T i j β + st i j b i + ɛ i j, (2.1) where v i j = (xi T j, st i j )T, δ = (β T, δ T ), and b i = γ + u i ; β and γ are the p- and q-dimensional fixedeffects parameters associated with covariates x i j and s i j, respectively; u i are the q-dimensional, mean zero iid subject-specific random-effects vectors associated with covariates s i j, independent across i; and ɛ i j N(0, σ 2 ) are the intra-subject error, which are independent of u i. We adopt the rightmost representation in (2.1) and note that the centered random effects b i can be written as b i = μ + RZ i, where μ is a (q 1) vector, R is a (q q) lower triangular matrix, and Z i is a (q 1) vector of random effects. Typically, Z i are assumed to follow a standard q-dimensional normal distribution, which implies that b i are normally distributed with mean μ = γ and covariance matrix var(u i ) = R R T. As an example, a special case of (2.1) is the linear random coefficient model with baseline covariate x i given by Y i j = x i β + b 0i + b 1i t i j + ɛ i j, (2.2) where b i = (b 0i, b 1i ) T and s i j = (1, t i j ) T. Due to left censoring, we observe Q i j which takes the value Y i j for Y i j > l i j and takes the value l i j, the known lower limit of quantification for the jth response on subject i, otherwise. Letting r = vech(r) be the nonzero elements of R, the parameters of interest are θ = (β T, μ T, r T, σ 2 ) T, and the likelihood assuming Gaussian random effects is L(θ; Q) = m i=1 [ n i { 1 (m i j )} I (Q i j =l i j ) j=1 { } ] 1 I (Qi j >l i j ) σ φ 1(m i j ) φ q (z i )dz i, (2.3) where m i j = {Q i j x T i j β st i j (μ + RZ i)}/σ, Q is the vector of all responses observed, and φ n ( ) and n ( ) are the standard n-dimensional normal density and distribution.
4 64 D. M. VOCK AND OTHERS Alternatively, we can express (2.1) as Y i = V i δ + S i u i + ɛ i, where V i {n i (p + q)} and S i (n i q) are the matrices with rows v T i j and s T i j, respectively, and Y i = ( Y i1,..., Y ini ) T and similarly for ɛi, l i, and Q i. To simplify the following argument, we assume that the design matrix is fixed, V i = V and n i = n for i = 1,..., m and also suppress the index i throughout. To express (2.3) using vector notation, let f Y (y; θ) and F Y (y; θ) be the multivariate normal density and distribution, respectively, with mean V δ and covariance matrix = SRR T S T +σ 2 I n. Define f Y1 Y 2 (y 1 y 2 ; θ) and F Y1 Y 2 (y 1 y 2 ; θ) to be the conditional multivariate normal density and distribution, respectively, of the vector Y 1 given the vector Y 2. There are 2 n 2 patterns of censoring/noncensoring that could be observed for an individual with at least one censored and one noncensored observation. Index each of these 2 n 2 distinct censoring patterns by k. Let Y k,o (Y k,c, respectively) be the random vector containing elements of Y that would be observed (censored) under censoring pattern k. If the kth pattern were observed for a particular subject, then the likelihood contribution from that subject assuming normal random effects is given by F Yk,c Y k,o (l k,c Q k,o ; θ) f Yk,o (Q k,o ; θ), where Q k,o is the vector of noncensored responses and l k,c (l k,o, respectively) is the limit of quantification for the censored observations (noncensored observations) under censoring pattern k. The likelihood contribution from one subject can now be written as 2n 2 L(θ, Q) = f Y (Q; θ) I (Q>l) F Y (l; θ) I (Q=l) {F Yk,c Y k,o (l k,c Q k,o ; θ) f Yk,o (Q k,o ; θ)} I (Q k,c=l k,c,q k,o >l k,o ). k=1 If the random-effects distribution is incorrectly specified, the maximum likelihood estimator ˆθ will converge to value of θ that solves E { / θ log L(θ; Q)} = 0, where the expectation is taken with respect to the true random-effects distribution. Assuming that the covariance components are known and using a first-order Taylor series expansion, we show in the supplementary material (available at Biostatistics online) that ˆδ will converge to the value of δ that solves l { } 0 = D(δ, V )(δ δ) + V T 1 (q V δ GY (l) ) F Y (l) f Y (q) g Y (q) dq + 2 n 2 k=1 [ l k,o lk,c V T k,c 1 k,c {q k,c V k,c δ } { } ] GYk,c Y k,o (l k,c q k,o ) F Yk,c Y k,o (l k,c q k,o ) f Y k,c Y k,o (q k,c q k,o ) g Yk,c Y k,o (q k,c q k,o ) g Yk,o (q k,o )dq k,c dq k,o, where g Y (q; δ ) and G Y (q; δ ) are the true marginal density and distribution, respectively, of Y ; all densities in (2.4) are evaluated at δ, the true parameter value; and D(δ, V ), Q k,c, V k,c, and k,c are given in the supplementary material available at Biostatistics online. While complex, the approximation reveals important insights on the asymptotic bias when the random effects are incorrectly specified. In the likely case that the random-effects distribution is misspecified, the bias is driven by the difference in the true conditional distribution of the censored response given the noncensored response (i.e. g Yk,c Y k,o (q k,c q k,o )) and the same conditional distribution induced by the Gaussian random effects (i.e. f Yk,c Y k,o (q k,c q k,o )), weighted by the likelihood of the noncensored response (i.e. g Yk,o (q k,o )). Practically, this suggests that small deviations from normality will lead to only slight asymptotic bias. If we have correctly specified the intra-subject error distribution, then g Yk,c Y k,o (y k,c y k,o ) will be approximately normal if the dimension of Y k,o is large regardless of the true random-effects density. Thus, if there is a large probability of observing many noncensored observations on each subject, then the (2.4)
5 Mixed model analysis of censored longitudinal data 65 asymptotic bias is likely to be slight. This suggests that the overall percentage of censored responses is immaterial in determining the asymptotic bias. Instead, the absolute number of noncensored responses for each subject is critical. 3. SEMIPARAMETRIC LINEAR MIXED MODEL WITH CENSORED RESPONSE We offer a summary of semiparametric linear mixed models and refer the reader to Zhang and Davidian (2001) for a complete description. We now assume that the distribution of Z i belongs to the smooth class of continuous densities described by Gallant and Nychka (1987). This class is sufficiently flexible to include skewed, thick- and thin-tailed, and multimodal distributions but does not include densities with jumps, kinks, or oscillations. These densities can be represented as an infinite series but can be approximated by a truncated series. The densities that are part of the truncated series are referred to as seminonparametric (SNP). We thus assume that the density of Z i can be approximated by the SNP representation with degree of truncation K given by h K (z) = PK 2 (z)φ q(z) = ( j j q ) K a j1,..., j q (z j 1 1 z j q q ) 2 φ q (z), (3.1) where j l 0 for l = 1,..., q and K is the order of the polynomial P K (z). For example, with K = 2 and q = 2, P K (z) = a 00 + a 10 z 1 + a 01 z 2 + a 20 z1 2 + a 11z 1 z 2 + a 02 z2 2. When K = 0, we show below that a 0,...,0 must equal 1, and the density of Z i is a standard q-dimensional normal. K controls the degree of flexibility of the density h K (z); we discuss in Section 4 how to select K. The coefficients a j1,..., j q must be chosen so that h K (z) integrates to one. The constraint h k (z)dz = 1 is equivalent to imposing E{PK 2 (U)} = 1, where U follows a standard q-dimensional normal distribution (Zhang and Davidian, 2001). Let a be the d-dimensional vector of coefficients for the polynomial P K (z) and let j 1,..., j q be the subscripts corresponding to the jth element. Then the above constraint can be rewritten as E{PK 2 (U)} = at Aa = 1, where A is the matrix with ( j, k) element equal to {E(U j 1+k 1 1 ) E(U j q +k q 1 )} and U 1 is distributed as a standard normal. Rather than impose a constraint on a, we may rewrite the (d 1) vector a as a function of d 1 parameters. Because A must be positive definite, there exists a matrix B such that A = B 2. If we let c = Ba, then a T Aa = 1 implies c T c = 1. As c and c will result in the same density h K (z), c = (c 1,..., c d ) T must lie in the half-unit sphere of R d. Therefore, we can express c in terms of a polar coordinate transformation, where c 1 = sin(ξ 1 ), c 2 = cos(ξ 1 ) sin(ξ 2 ),..., c d 1 = cos(ξ 1 )... cos(ξ d 2 ) sin(ξ d 1 ), c d = cos(ξ 1 )... cos(ξ d 1 ), π/2 ξ j π/2 for j = 1,..., d 1, and ξ = (ξ 1,..., ξ d 1 ) T. The vector a can be written as B 1 c, which only involves d 1 parameters. The parameters of interest are now θsnp K = (βt, μ T, r T, σ 2, ξ T ) T, which includes an additional d 1 parameters compared to when normality is assumed. Note that the SNP density does not impose E(Z i ) = 0, so that E(b i ) = γ = μ + R{E(Z i )} and Var(b i ) = R{Var(Z i )}R T. The moments of Z i are linear combinations of the moments of a standard normal density, which can be found using recursion formulas. 4. ESTIMATION USING SAS NLMIXED In the case where there is no censoring of the response variable, the log-likelihood assuming the SNP representation of the random effects can be expressed in a closed form (Zhang and Davidian, 2001).
6 66 D. M. VOCK AND OTHERS Censoring necessitates numerical integration. For a fixed K, the likelihood of θsnp K is given by m L(θSNP K ; Q) = i=1 [ n i { 1 (m i j )} I (Q i j =l i j ) j=1 { } ] 1 I (Qi j >l i j ) σ φ 1(m i j ) PK 2 (z i)φ q (z i )dz i. (4.1) For each observation i, this requires evaluation of a q-dimensional integral, which, in practice, would likely only have dimension 1 or 2. In contrast, we could integrate over the marginal density of Y i for each censored observation. However, the dimension of that integral would equal the number of censored observations for subject i, which could be quite large and computationally intractable. The SAS procedure NLMIXED has been developed to obtain maximum likelihood estimates for mixed models with Gaussian random effects. Except when K = 0, the SNP random-effects density will not be normally distributed. However, if we consider [ n i { 1 (m i j )} I (Q i j =l i j ) j=1 { } ] 1 I (Qi j >l i j ) σ φ 1(m i j ) PK 2 (z i) (4.2) to be the likelihood for Q i j conditioned on the random effects, then the random effects Z i can be thought to follow a standard q-dimensional normal distribution (Liu and Yu, 2008). Example code to implement SNP for left-censored mixed models is given in the supplementary material (available at Biostatistics online). In practice, given that (4.2) is highly nonlinear, we have found optimization routines within NLMIXED to obtain the empirical Bayes estimates of Z i, required for the default adaptive Gaussian quadrature used to evaluate the integrals to be unstable and computationally intractable for q 2. To approximate the integral in (4.1), we recommend using nonadaptive Gaussian quadrature with quadrature points centered at the empirical Bayes estimates of b i from assuming Gaussian random effects. Among the optimization routines available in NLMIXED, dual-quasi Newton optimization works well. The inverse Hessian matrix may be used to obtain standard errors for the parameter estimates and is computed as part of the standard output in NLMIXED. The preceding discussion assumes a fixed K. We treat K as a tuning parameter and select K by visually comparing the estimated densities and computing information criteria evaluated at the estimates, ˆθ SNP K, of θ SNP K from model fits for different values of K (Zhang and Davidian, 2001). The information criteria are of the form 2L( ˆθ SNP K, Q) + C(m)p, where p is the number of parameters in the model. For the Akaike information criterion (AIC), C(m) = 2; Hannan Quinn information criterion (HQIC), C(m) = 2 log log m; and Bayesian information criterion (BIC), C(m) = log m. Because the HQIC will select a model that is of intermediate complexity compared to those chosen by AIC and BIC, this criterion is often preferred (Davidian and Gallant, 1993). Prior research has shown that K need not be greater than 2 to capture many complex densities (Davidian and Gallant, 1993; Zhang and Davidian, 2001; Chen and others, 2002). Optimization for SNP can be highly dependent on the starting values used for θsnp K and especially for ξ. We suggest the following approach advocated by Doehler and Davidian (2008). Initial estimates of β, E(b i ), var(b i ), and σ 2 should be obtained, perhaps by fitting a mixed model assuming Gaussian random effects. The log-likelihood (4.1) can then be evaluated over a grid of ξ with starting values for β and σ 2 set to the initial estimates and starting values for μ and r selected to give the same value for E(b i ) and var(b i ) as the initial estimates. The sets of starting values that yield local maxima among all the grid points can then be used for optimization.
7 Mixed model analysis of censored longitudinal data SIMULATION We conducted a variety of simulation studies both to assess the impact of erroneously assuming Gaussian random effects and to gauge the ability of SNP to represent a broad range of random-effects distributions. We report on a subset of these with q = 2 here; results of additional simulations are included in the supplementary material (available at Biostatistics online). The first part of the simulation study examined the effect of misspecification of the random-effects distribution on estimation and inference of model parameters. We considered (2.2) where x i is equal to iid 0 or 1 with equal probability, ɛ i j N(0, σ 2 ), and t i = (t i1,..., t i5 ) T = (0, 1, 2, 3, 4) T t A for all i = 1,..., m. For all simulations, β = 0.5, σ 2 = 0.25, and (b 0i, b 1i ) T were generated from distributions that were shifted and scaled so that E(b 0i ) = 5.75, E(b 1i ) = 0.60, var(b 0i ) = 0.36, var(b 1i ) = , and cov(b 0i, b 1i ) = The random effects, b i were drawn from 1 of 4 shifted and scaled distributions: (1) bivariate normal; (2) a bivariate t 5 distribution; (3) a mixture of normal densities with mean components (5.6, ) T (70% component) and (6.1, 1.75) T, which gives a skewed marginal density for b 1i ; and (4) a mixture of normal densities with mean components (5.6, 0.03) T (70% component) and (6.1, 1.93) T, which produces a bimodal marginal density for b 1i. For (3) and (4), the covariance matrices of the components were equal to each other. For each of the 4 distributions, 500 Monte Carlo simulations were generated with 500 subjects each. Here, we report the results for censoring level l i j 4 for all subjects and time points. The models were fit using SAS procedure NLMIXED assuming Gaussian random effects. The likelihood (2.3) was approximated using adaptive Gaussian quadrature with the number of quadrature points selected adaptively to achieve a tolerance of Dual-quasi Newton was used for optimization with the true values used as starting values. Within each of the simulation scenarios, nearly all subjects had at least one noncensored measurement, and approximately 90% had at least 2 noncensored observations. With the exception of the data generated from the bimodal random-effects density (4) where 42.4% of subjects had at least one censored observation, a slight majority in each of the other scenarios had at least one censored observation. A complete description of the censoring pattern is given in the supplementary material (available at Biostatistics online). The results are given in Table 1. When the random effects are correctly specified, parameter estimators are unbiased, and the coverage probabilities attain their stated level of confidence. When the random effects have heavy tails (t 5 distribution), parameter estimators are still unbiased although coverage probabilities for the covariance components do degrade slightly. Equation (2.4) suggests that the asymptotic bias in this scenario would be slight. When the random slope is slightly skewed, parameter estimators are biased, especially for E(b 1i ) (6.0%) and var(b 1i ) ( 10.0%). Coverage probabilities for these parameters in particular are far from nominal. This illustrates that even slight departures from normality can lead to erroneous inference when the number of noncensored observations for each subject is small. In the case of severe misspecification with bimodal random slopes, the bias of all the parameter estimators increases in comparison to that under skewed random effects; the bias in the estimators for E(b 1i ) and var(b 1i ) is substantial (13.0% and 18.9%, respectively), and coverage probabilities for these parameters are poor. We also examined the effect of adding design points where there was unlikely to be any censoring and where censoring was likely. We generated 500 data sets with bimodal random effects, where t i = (0, 1, 2, 3, 3.5, 4, 4.5) T t B and t i = ( 2, 1, 0, 1, 2, 3, 4) T t C for all subjects. With measurements at t i j = 2 and t i j = 1, 99.9% and 90.5% of subjects had at least 3 and 4 noncensored measurements, respectively. Adding measurements at t i j = 3.5 and t i j = 4.5 increased the overall percentage of censored observations to 25.5% from 21.8%. Simulation results with bimodal random effects and taking t i to be each of t A, t B, and t C are shown in Table 2. When there are more design points where censored observations are unlikely, bias is mitigated
8 68 D. M. VOCK AND OTHERS Table 1. Simulation results when Gaussian random effects were assumed for all models regardless of the true distribution of the random effects. The simulation included 500 data sets with 500 subjects each Distribution E(b 0i ) E(b 1i ) β 0 var(b 0i ) cov(b 0i, b 1i ) var(b 1i ) Truth MC Avg Normal t Skewed Bimodal MC SD Normal t Skewed Bimodal Avg SE Normal t Skewed Bimodal CP Normal t Skewed Bimodal MC Avg, Monte Carlo average of the parameter estimates; MC SD, Monte Carlo standard deviation of the parameter estimates; Avg SE, average of the standard error estimates; CP, Monte Carlo coverage probability of the 95% Wald-type confidence intervals. Table 2. Simulation results when Gaussian random effects were assumed for the bimodal random effects but t i in (2.2) was varied. The simulation included 500 data sets with 500 subjects each Time points E(b 0i ) E(b 1i ) β 0 var(b 0i ) cov(b 0i, b 1i ) var(b 1i ) Truth MC Avg t A t B t C MC SD t A t B t C Avg SE t A t B t C CP t A t B t C t A = (0, 1, 2, 3, 4) T ; t B = (0, 1, 2, 3, 3.5, 4, 4.5) T ; t C = ( 2, 1, 0, 1, 2, 3, 4) T ; MC Avg, Monte Carlo average of the parameter estimates; MC SD, Monte Carlo standard deviation of the parameter estimates; Avg SE, average of the standard error estimates; CP, Monte Carlo coverage probability of the 95% Wald-type confidence intervals. substantially and coverage probabilities improve, which is consistent with the asymptotic results in Section 2. Conversely, increasing the number of design points where censoring is likely has little effect on parameter estimation, illustrating that the overall censoring level has little to do with the bias. We also conducted simulation studies of the performance of SNP density estimation for censored longitudinal data analysis. We considered the same simulation scenarios as above except for the t 5
9 Mixed model analysis of censored longitudinal data 69 random-effects distribution, as assuming Gaussian random effects did not adversely affect inference. To obtain estimates of parameters with SNP random effects, we again used SAS procedure NLMIXED. Models were fit for K = 0, 1, and 2, and K was chosen using HQIC. The likelihood (4.1) was approximated using Gaussian quadrature with quadrature points centered at the empirical Bayes estimates of b i derived from assuming Gaussian random effects and with the number of quadrature points selected with a stated tolerance of Dual-quasi Newton was used for optimization. 500 Monte Carlo data sets were generated for each scenario. To obtain starting values, the log-likelihood was evaluated over a grid of ξ of 50 points for K = 1 and 150 points for K = 2 with starting values for β and σ 2 set to the true values and for μ and r set to the values that would give the true values for E(b i ) and var(b i ). The results when K was selected by HQIC are shown in Table 3. When the random effects are normally distributed, HQIC selects K = 0 for 94.8% of the data sets. However, when the random-effects density was skewed and bimodal, K = 0 was selected only 0.2% and 0.0% of the time, respectively, and K = 2 was selected for 95.2% and 36.4% of the data sets, respectively. This illustrates that even with a modest sample size and loss of information due to censoring, the method is able to detect slight departures from normality while not over-fitting models where the true random-effects density is Gaussian. A complete table on the proportion of data sets that selected K = 0, 1, and 2 by information criterion is given in the supplementary material (available at Biostatistics online). The SNP estimators when the random effects are not normally distributed are less biased than the estimators when Gaussian random effects were assumed. When the random effects were skewed, the bias for E(b 1i ) and var(b 1i ) is reduced to 0.5% and 0.3%, respectively, which leads to large efficiency gains (Table 3). The coverage probabilities for these parameters improve substantially although for bimodal random effects are still below the stated level. Because K = 0 is selected so frequently when the random effects are Gaussian, there is little loss in efficiency from considering a more flexible class of random Table 3. Simulation results when SNP was used to estimate the random effects. Models with K = 0, K = 1, and K = 2 were fit and K was selected using the HQIC. The simulation included 500 data sets with 500 subjects each Distribution E(b 0i ) E(b 1i ) β 0 var(b 0i ) cov(b 0i, b 1i ) var(b 1i ) Truth MC Avg Normal Skewed Bimodal MC SD Normal Skewed Bimodal Avg SE Normal Skewed Bimodal CP Normal Skewed Bimodal Ratio MSE Normal Skewed Bimodal MC Avg, Monte Carlo average of the parameter estimates; MC SD, Monte Carlo standard deviation of the parameter estimates; Avg SE, average of the standard error estimates; Ratio MSE, ratio of the Monte Carlo mean square error between K = 0 and K selected by the HQIC.
10 70 D. M. VOCK AND OTHERS effects. Estimated contour and marginal density plots of the random effects from assuming that the random effects follow the SNP density is provided in the supplementary material (available at Biostatistics online). 6. APPLICATION We now illustrate the proposed methods using data from 811 subjects in the IDEAL study with the CT genotype at polymorphic site upstream of interleukin (IL) 28B which is associated with virologic response (Thompson and others, 2010). Subjects had viral load measurements taken at baseline and at 2, 4, and 12 weeks after treatment began, some of which were censored at the lower limit of quantification of log 10 IU/mL. As shown in Figure 1, the viral load change over the first 12 weeks within subject can be well approximated by a linear trajectory, and the measurement error and biological fluctuations at each time point can be assumed reasonably to be independent and Gaussian. However, standard therapy is not effective for all subjects, so the assumption that the subject-specific slopes are normally distributed is questionable. Based on these observations, we consider the semiparametric model Y i j = b 0i + b 1i t i j + ɛ i j, (6.1) where Y i j is the log 10 viral load for patient i at the jth time, t i j is the time in weeks since starting treatment, x i j is null, s i j = (1, t i j ) T iid, ɛ i j N(0, σ 2 ), and b i = (b 0i, b 1i ) T is the vector of subject-specific intercept and slope, which we assume can be written as b i = μ + RZ i with Z i = (Z 0i, Z 1i ) T, μ = (μ 1, μ 2 ) T, and R a (2 2) lower triangular matrix. We assume that Z i follows the density (3.1) for the K described below. We do not observe Y i j but instead observe Q i j with l i j All patients included in this analysis have noncensored baseline measurements, and greater than 94% of measurements taken at 2 and 4 weeks after starting treatment are uncensored. Still, 12.8% of patients only have 1 or 2 noncensored measurements. At week 12, 35.2% of subjects responses are censored. The fit statistics and relevant parameter estimates from fitting model (6.1) with K = 0, K = 1, and K = 2 appear in Table 4; additional parameter estimates are given in the supplementary material (available at Biostatistics online). While most of the parameter estimates are only altered marginally as K increases, the estimate for E(b 1i ), the average weekly log viral load decline, changes substantially. When K = 1 and K = 2, the estimate is more than one standard error away from the estimate when K = 0. Each of the information criteria preferred K = 2, so we present the density estimates from that model in Figure 2. The contour plot for the bivariate random effects shows the presence of 2 or possibly 3 modes. SNP density estimation results in a spurious mode to capture mass in what is actually a long Table 4. Information criteria and parameter estimates from fitting model (6.1) concerning the IDEAL study with K = 0, K = 1, and K = 2 K = 0 K = 1 K = 2 AIC HQIC BIC Estimate Standard error Estimate Sandard error Estimate Standard error E(b 0i ) E(b 1i ) var(b 0i ) cov(b 0i, b 1i ) var(b 1i )
11 Mixed model analysis of censored longitudinal data 71 Fig. 2. Contour plot of the bivariate density estimate and marginal density estimates for the subject-specific intercepts and slopes with K = 2 from model (6.1) concerning the IDEAL study. tail, so one should be cautious about overinterpreting this third mode. The marginal density estimates show a large departure from normality for the subject-specific slope, confirming our prior hypothesis, but little departure from normality in the baseline log 10 viral load. The majority of patients with the CT genotype experience very modest weekly viral load changes around 0.25 log 10 IU/mL/week. However, the remaining patients experience greater viral load decline with the mode at approximately 0.85 log 10 IU/mL/week. One possible clinical explanation for the nonnormal random slopes is that some patients respond to standard therapy while the majority with the CT genotype do not show substantial improvement. The IL28B genotype has been shown to be a strong predictor of virological response for patients with hepatitis C undergoing standard therapy. However, the analysis here suggests that, even within this genotype, there are responders and nonresponders, and more research is required to determine why patients respond to therapy. This example illustrates that fitting a flexible model for the random effects when the response is censored cannot only substantially alter the estimates of clinically relevant parameters like
12 72 D. M. VOCK AND OTHERS E(b 1i ) but can also lead to a fuller understanding of the underlying process. Given the multimodality of the subject-specific slopes, some would even question if E(b 1i ) is a useful parameter with which to characterize the population. 7. DISCUSSION We have shown how the SNP random-effects density can be extended to linear mixed models with a censored response. The implementation within SAS and the example code given in the supplementary material (available at Biostatistics online) allow the method to be applied easily in practice. If one is interested purely in inference for the fixed-effects parameters, we have shown that the asymptotic bias from erroneously assuming Gaussian random effects is likely to be greatest when the true random-effects distribution deviates from normality and the probability of observing a small number of noncensored observations is not trivially small. That is, the overall level of censoring is unimportant, but rather the absolute number of noncensored responses for each subject is relevant. Simulations show that the deviation from normality need not be substantial to affect inference. Since there is little efficiency loss from using the SNP density when the random effects are Gaussian, we recommend using the SNP density for censored longitudinal data when the number of noncensored observations is small to avoid erroneous inference. More specifically, we suggest fitting SNP models for several K to determine if the random-effects density deviates from normality. Visual inspection of the estimated densities for each K > 0 as well as information criteria can be used to assess if the random-effects density is nonnormal. When the randomeffects distribution deviates from normality, the information criteria rarely select K = 0 even with modest sample sizes, indicating the method s ability to detect nonnormal distributions. In addition to improved inference, one gains insight into the data generating process if a flexible random-effects model is used. Within the economics literature, there has been substantial work on developing tests to determine if the error distribution is Gaussian in censored regression models with independent responses. Future work could extend those tests to random-effects densities. We have focused on linear mixed models with a censored response. However, nonlinear trajectories can be easily incorporated in procedure NLMIXED, so the methods could easily be transferred to a nonlinear mixed model with censored response. Future work could examine the benefit of assuming a flexible random-effects distribution in this setting. SUPPLEMENTARY MATERIAL Supplementary material is available at ACKNOWLEDGMENTS The authors thank Merck for approving the use of the IDEAL study and Oliver Schabenberger for helpful discussions on procedure NLMIXED. Conflict of Interest: None declared. FUNDING National Institutes of Health (R01CA085848, R01CA051962, R37AI031789, and P01CA142538). REFERENCES AGRESTI, A., CAFFO, B. AND OHMAN-STRICKLAND, P. (2004). Examples in which misspecification of a random effects distribution reduces efficiency, and possible remedies. Computational Statistics & Data Analysis 47,
13 Mixed model analysis of censored longitudinal data 73 AITKIN, M. (1999). A general maximum likelihood analysis of variance components in generalized linear models. Biometrics 55, CHEN, J., ZHANG, D. AND DAVIDIAN, M. (2002). A Monte Carlo EM algorithm for generalized linear mixed models with flexible random effects distribution. Biostatistics 3, DAVIDIAN, M. AND GALLANT, A. R. (1993). The nonlinear mixed effects model with a smooth random effects density. Biometrika 80, DOEHLER, K. AND DAVIDIAN, M. (2008). Smooth inference for survival functions with arbitrarily censored data. Statistics in Medicine 27, GALLANT, A. R. AND NYCHKA, D. W. (1987). Semi-nonparametric maximum likelihood estimation. Econometrica 55, HEAGERTY, P. J. AND KURLAND, B. F. (2001). Misspecified maximum likelihood estimates and generalised linear mixed models. Biometrika 88, HUGHES, J. P. (1999). Mixed effects models with censored data with application to HIV RNA levels. Biometrics 55, JACQMIN-GADDA, H., THIÉBAUT, R., CHÊNE, G. AND COMMENGES, D. (2000). Analysis of left-censored longitudinal data with application to viral load in HIV infection. Biostatistics 1, KLEINMAN, K. P. AND IBRAHIM, J. G. (1998). A semiparametric Bayesian approach to the random effects model. Biometrics 54, LEE, K. J. AND THOMPSON, S. G. (2008). Flexible parametric models for random-effects distributions. Statistics in Medicine 27, LITIÈRE, S., ALONSO, A. AND MOLENBERGHS, G. (2008). The impact of a misspecified random-effects distribution on the estimation and the performance of inferential procedures in generalized linear mixed models. Statistics in Medicine 27, LIU, L. AND YU, Z. (2008). A likelihood reformulation method in non-normal random effects models. Statistics in Medicine 27, MAGDER, L. S. AND ZEGER, S. L. (1996). A smooth nonparametric estimate of a mixing distribution using mixtures of Gaussians. Journal of the American Statistical Association 91, NEUHAUS, J. M., HAUCK, W. W. AND KALBFLEISCH, J. D. (1992). The effects of mixture distribution misspecification when fitting mixed-effects logistic models. Biometrika 79, THOMPSON, A. J., MUIR, A. J., SULKOWSKI, M. S., GE, D., FELLAY, J., SHIANNA, K. V., URBAN, T., AFDHAL, N. H., JACOBSON, I. M., ESTEBAN, R. and others. (2010). Interleukin-28b polymorphism improves viral kinetics and is the strongest pretreatment predictor of sustained virologic response in genotype 1 hepatitis C virus. Gastroenterology 139, VAIDA, F. AND LIU, L. (2009). Fast implementation for normal mixed effects models with censored response. Journal of Computational and Graphical Statistics 18, VERBEKE, G. AND LESAFFRE, E. (1997). The effect of misspecifying the random-effects distribution in linear mixed models for longitudinal data. Computational Statistics & Data Analysis 23, WU, L. (2002). A joint model for nonlinear mixed-effects models with censoring and covariates measured with error, with application to AIDS studies. Journal of the American Statistical Association 97, ZHANG, D. AND DAVIDIAN, M. (2001). Linear mixed models with flexible distributions of random effects for longitudinal data. Biometrics 57, [Received April 8, 2011; revised July 21, 2011; accepted for publication July 24, 2011]
Semiparametric Mixed Effects Models with Flexible Random Effects Distribution
 Semiparametric Mixed Effects Models with Flexible Random Effects Distribution Marie Davidian North Carolina State University davidian@stat.ncsu.edu www.stat.ncsu.edu/ davidian Joint work with A. Tsiatis,
Semiparametric Mixed Effects Models with Flexible Random Effects Distribution Marie Davidian North Carolina State University davidian@stat.ncsu.edu www.stat.ncsu.edu/ davidian Joint work with A. Tsiatis,
are censored. Because artificial censoring reduces the information available and leads to nonsmooth estimating functions, prior research has noted
 ABSTRACT VOCK, DAVID MICHAEL. Advanced Statistical Methods for Complex Longitudinal Data. (Under the direction of Marie Davidian and Anastasios A. Tsiatis.) Longitudinally collected data play an important
ABSTRACT VOCK, DAVID MICHAEL. Advanced Statistical Methods for Complex Longitudinal Data. (Under the direction of Marie Davidian and Anastasios A. Tsiatis.) Longitudinally collected data play an important
Some New Methods for Latent Variable Models and Survival Analysis. Latent-Model Robustness in Structural Measurement Error Models.
 Some New Methods for Latent Variable Models and Survival Analysis Marie Davidian Department of Statistics North Carolina State University 1. Introduction Outline 3. Empirically checking latent-model robustness
Some New Methods for Latent Variable Models and Survival Analysis Marie Davidian Department of Statistics North Carolina State University 1. Introduction Outline 3. Empirically checking latent-model robustness
T E C H N I C A L R E P O R T A SANDWICH-ESTIMATOR TEST FOR MISSPECIFICATION IN MIXED-EFFECTS MODELS. LITIERE S., ALONSO A., and G.
 T E C H N I C A L R E P O R T 0658 A SANDWICH-ESTIMATOR TEST FOR MISSPECIFICATION IN MIXED-EFFECTS MODELS LITIERE S., ALONSO A., and G. MOLENBERGHS * I A P S T A T I S T I C S N E T W O R K INTERUNIVERSITY
T E C H N I C A L R E P O R T 0658 A SANDWICH-ESTIMATOR TEST FOR MISSPECIFICATION IN MIXED-EFFECTS MODELS LITIERE S., ALONSO A., and G. MOLENBERGHS * I A P S T A T I S T I C S N E T W O R K INTERUNIVERSITY
Type I and type II error under random-effects misspecification in generalized linear mixed models Link Peer-reviewed author version
 Type I and type II error under random-effects misspecification in generalized linear mixed models Link Peer-reviewed author version Made available by Hasselt University Library in Document Server@UHasselt
Type I and type II error under random-effects misspecification in generalized linear mixed models Link Peer-reviewed author version Made available by Hasselt University Library in Document Server@UHasselt
The consequences of misspecifying the random effects distribution when fitting generalized linear mixed models
 The consequences of misspecifying the random effects distribution when fitting generalized linear mixed models John M. Neuhaus Charles E. McCulloch Division of Biostatistics University of California, San
The consequences of misspecifying the random effects distribution when fitting generalized linear mixed models John M. Neuhaus Charles E. McCulloch Division of Biostatistics University of California, San
Using Estimating Equations for Spatially Correlated A
 Using Estimating Equations for Spatially Correlated Areal Data December 8, 2009 Introduction GEEs Spatial Estimating Equations Implementation Simulation Conclusion Typical Problem Assess the relationship
Using Estimating Equations for Spatially Correlated Areal Data December 8, 2009 Introduction GEEs Spatial Estimating Equations Implementation Simulation Conclusion Typical Problem Assess the relationship
SEMIPARAMETRIC APPROACHES TO INFERENCE IN JOINT MODELS FOR LONGITUDINAL AND TIME-TO-EVENT DATA
 SEMIPARAMETRIC APPROACHES TO INFERENCE IN JOINT MODELS FOR LONGITUDINAL AND TIME-TO-EVENT DATA Marie Davidian and Anastasios A. Tsiatis http://www.stat.ncsu.edu/ davidian/ http://www.stat.ncsu.edu/ tsiatis/
SEMIPARAMETRIC APPROACHES TO INFERENCE IN JOINT MODELS FOR LONGITUDINAL AND TIME-TO-EVENT DATA Marie Davidian and Anastasios A. Tsiatis http://www.stat.ncsu.edu/ davidian/ http://www.stat.ncsu.edu/ tsiatis/
Score test for random changepoint in a mixed model
 Score test for random changepoint in a mixed model Corentin Segalas and Hélène Jacqmin-Gadda INSERM U1219, Biostatistics team, Bordeaux GDR Statistiques et Santé October 6, 2017 Biostatistics 1 / 27 Introduction
Score test for random changepoint in a mixed model Corentin Segalas and Hélène Jacqmin-Gadda INSERM U1219, Biostatistics team, Bordeaux GDR Statistiques et Santé October 6, 2017 Biostatistics 1 / 27 Introduction
Review Article Analysis of Longitudinal and Survival Data: Joint Modeling, Inference Methods, and Issues
 Journal of Probability and Statistics Volume 2012, Article ID 640153, 17 pages doi:10.1155/2012/640153 Review Article Analysis of Longitudinal and Survival Data: Joint Modeling, Inference Methods, and
Journal of Probability and Statistics Volume 2012, Article ID 640153, 17 pages doi:10.1155/2012/640153 Review Article Analysis of Longitudinal and Survival Data: Joint Modeling, Inference Methods, and
WEIGHTED QUANTILE REGRESSION THEORY AND ITS APPLICATION. Abstract
 Journal of Data Science,17(1). P. 145-160,2019 DOI:10.6339/JDS.201901_17(1).0007 WEIGHTED QUANTILE REGRESSION THEORY AND ITS APPLICATION Wei Xiong *, Maozai Tian 2 1 School of Statistics, University of
Journal of Data Science,17(1). P. 145-160,2019 DOI:10.6339/JDS.201901_17(1).0007 WEIGHTED QUANTILE REGRESSION THEORY AND ITS APPLICATION Wei Xiong *, Maozai Tian 2 1 School of Statistics, University of
Group Sequential Designs: Theory, Computation and Optimisation
 Group Sequential Designs: Theory, Computation and Optimisation Christopher Jennison Department of Mathematical Sciences, University of Bath, UK http://people.bath.ac.uk/mascj 8th International Conference
Group Sequential Designs: Theory, Computation and Optimisation Christopher Jennison Department of Mathematical Sciences, University of Bath, UK http://people.bath.ac.uk/mascj 8th International Conference
PQL Estimation Biases in Generalized Linear Mixed Models
 PQL Estimation Biases in Generalized Linear Mixed Models Woncheol Jang Johan Lim March 18, 2006 Abstract The penalized quasi-likelihood (PQL) approach is the most common estimation procedure for the generalized
PQL Estimation Biases in Generalized Linear Mixed Models Woncheol Jang Johan Lim March 18, 2006 Abstract The penalized quasi-likelihood (PQL) approach is the most common estimation procedure for the generalized
Estimation of Conditional Kendall s Tau for Bivariate Interval Censored Data
 Communications for Statistical Applications and Methods 2015, Vol. 22, No. 6, 599 604 DOI: http://dx.doi.org/10.5351/csam.2015.22.6.599 Print ISSN 2287-7843 / Online ISSN 2383-4757 Estimation of Conditional
Communications for Statistical Applications and Methods 2015, Vol. 22, No. 6, 599 604 DOI: http://dx.doi.org/10.5351/csam.2015.22.6.599 Print ISSN 2287-7843 / Online ISSN 2383-4757 Estimation of Conditional
On Estimating the Relationship between Longitudinal Measurements and Time-to-Event Data Using a Simple Two-Stage Procedure
 Biometrics DOI: 10.1111/j.1541-0420.2009.01324.x On Estimating the Relationship between Longitudinal Measurements and Time-to-Event Data Using a Simple Two-Stage Procedure Paul S. Albert 1, and Joanna
Biometrics DOI: 10.1111/j.1541-0420.2009.01324.x On Estimating the Relationship between Longitudinal Measurements and Time-to-Event Data Using a Simple Two-Stage Procedure Paul S. Albert 1, and Joanna
Approximate Median Regression via the Box-Cox Transformation
 Approximate Median Regression via the Box-Cox Transformation Garrett M. Fitzmaurice,StuartR.Lipsitz, and Michael Parzen Median regression is used increasingly in many different areas of applications. The
Approximate Median Regression via the Box-Cox Transformation Garrett M. Fitzmaurice,StuartR.Lipsitz, and Michael Parzen Median regression is used increasingly in many different areas of applications. The
Improving Efficiency of Inferences in Randomized Clinical Trials Using Auxiliary Covariates
 Improving Efficiency of Inferences in Randomized Clinical Trials Using Auxiliary Covariates Anastasios (Butch) Tsiatis Department of Statistics North Carolina State University http://www.stat.ncsu.edu/
Improving Efficiency of Inferences in Randomized Clinical Trials Using Auxiliary Covariates Anastasios (Butch) Tsiatis Department of Statistics North Carolina State University http://www.stat.ncsu.edu/
Bayesian Inference on Joint Mixture Models for Survival-Longitudinal Data with Multiple Features. Yangxin Huang
 Bayesian Inference on Joint Mixture Models for Survival-Longitudinal Data with Multiple Features Yangxin Huang Department of Epidemiology and Biostatistics, COPH, USF, Tampa, FL yhuang@health.usf.edu January
Bayesian Inference on Joint Mixture Models for Survival-Longitudinal Data with Multiple Features Yangxin Huang Department of Epidemiology and Biostatistics, COPH, USF, Tampa, FL yhuang@health.usf.edu January
Computationally Efficient Estimation of Multilevel High-Dimensional Latent Variable Models
 Computationally Efficient Estimation of Multilevel High-Dimensional Latent Variable Models Tihomir Asparouhov 1, Bengt Muthen 2 Muthen & Muthen 1 UCLA 2 Abstract Multilevel analysis often leads to modeling
Computationally Efficient Estimation of Multilevel High-Dimensional Latent Variable Models Tihomir Asparouhov 1, Bengt Muthen 2 Muthen & Muthen 1 UCLA 2 Abstract Multilevel analysis often leads to modeling
BIOS 2083: Linear Models
 BIOS 2083: Linear Models Abdus S Wahed September 2, 2009 Chapter 0 2 Chapter 1 Introduction to linear models 1.1 Linear Models: Definition and Examples Example 1.1.1. Estimating the mean of a N(μ, σ 2
BIOS 2083: Linear Models Abdus S Wahed September 2, 2009 Chapter 0 2 Chapter 1 Introduction to linear models 1.1 Linear Models: Definition and Examples Example 1.1.1. Estimating the mean of a N(μ, σ 2
A TWO-STAGE LINEAR MIXED-EFFECTS/COX MODEL FOR LONGITUDINAL DATA WITH MEASUREMENT ERROR AND SURVIVAL
 A TWO-STAGE LINEAR MIXED-EFFECTS/COX MODEL FOR LONGITUDINAL DATA WITH MEASUREMENT ERROR AND SURVIVAL Christopher H. Morrell, Loyola College in Maryland, and Larry J. Brant, NIA Christopher H. Morrell,
A TWO-STAGE LINEAR MIXED-EFFECTS/COX MODEL FOR LONGITUDINAL DATA WITH MEASUREMENT ERROR AND SURVIVAL Christopher H. Morrell, Loyola College in Maryland, and Larry J. Brant, NIA Christopher H. Morrell,
Logistic regression analysis with multidimensional random effects: A comparison of three approaches
 Quality & Quantity manuscript No. (will be inserted by the editor) Logistic regression analysis with multidimensional random effects: A comparison of three approaches Olga Lukočienė Jeroen K. Vermunt Received:
Quality & Quantity manuscript No. (will be inserted by the editor) Logistic regression analysis with multidimensional random effects: A comparison of three approaches Olga Lukočienė Jeroen K. Vermunt Received:
Multivariate Survival Analysis
 Multivariate Survival Analysis Previously we have assumed that either (X i, δ i ) or (X i, δ i, Z i ), i = 1,..., n, are i.i.d.. This may not always be the case. Multivariate survival data can arise in
Multivariate Survival Analysis Previously we have assumed that either (X i, δ i ) or (X i, δ i, Z i ), i = 1,..., n, are i.i.d.. This may not always be the case. Multivariate survival data can arise in
Web-based Supplementary Materials for A Robust Method for Estimating. Optimal Treatment Regimes
 Biometrics 000, 000 000 DOI: 000 000 0000 Web-based Supplementary Materials for A Robust Method for Estimating Optimal Treatment Regimes Baqun Zhang, Anastasios A. Tsiatis, Eric B. Laber, and Marie Davidian
Biometrics 000, 000 000 DOI: 000 000 0000 Web-based Supplementary Materials for A Robust Method for Estimating Optimal Treatment Regimes Baqun Zhang, Anastasios A. Tsiatis, Eric B. Laber, and Marie Davidian
International Journal of PharmTech Research CODEN (USA): IJPRIF, ISSN: , ISSN(Online): Vol.9, No.9, pp , 2016
 International Journal of PharmTech Research CODEN (USA): IJPRIF, ISSN: 0974-4304, ISSN(Online): 2455-9563 Vol.9, No.9, pp 477-487, 2016 Modeling of Multi-Response Longitudinal DataUsing Linear Mixed Modelin
International Journal of PharmTech Research CODEN (USA): IJPRIF, ISSN: 0974-4304, ISSN(Online): 2455-9563 Vol.9, No.9, pp 477-487, 2016 Modeling of Multi-Response Longitudinal DataUsing Linear Mixed Modelin
Approximation of Survival Function by Taylor Series for General Partly Interval Censored Data
 Malaysian Journal of Mathematical Sciences 11(3): 33 315 (217) MALAYSIAN JOURNAL OF MATHEMATICAL SCIENCES Journal homepage: http://einspem.upm.edu.my/journal Approximation of Survival Function by Taylor
Malaysian Journal of Mathematical Sciences 11(3): 33 315 (217) MALAYSIAN JOURNAL OF MATHEMATICAL SCIENCES Journal homepage: http://einspem.upm.edu.my/journal Approximation of Survival Function by Taylor
Conditional Inference Functions for Mixed-Effects Models with Unspecified Random-Effects Distribution
 Conditional Inference Functions for Mixed-Effects Models with Unspecified Random-Effects Distribution Peng WANG, Guei-feng TSAI and Annie QU 1 Abstract In longitudinal studies, mixed-effects models are
Conditional Inference Functions for Mixed-Effects Models with Unspecified Random-Effects Distribution Peng WANG, Guei-feng TSAI and Annie QU 1 Abstract In longitudinal studies, mixed-effects models are
6 Pattern Mixture Models
 6 Pattern Mixture Models A common theme underlying the methods we have discussed so far is that interest focuses on making inference on parameters in a parametric or semiparametric model for the full data
6 Pattern Mixture Models A common theme underlying the methods we have discussed so far is that interest focuses on making inference on parameters in a parametric or semiparametric model for the full data
Calibration Estimation of Semiparametric Copula Models with Data Missing at Random
 Calibration Estimation of Semiparametric Copula Models with Data Missing at Random Shigeyuki Hamori 1 Kaiji Motegi 1 Zheng Zhang 2 1 Kobe University 2 Renmin University of China Institute of Statistics
Calibration Estimation of Semiparametric Copula Models with Data Missing at Random Shigeyuki Hamori 1 Kaiji Motegi 1 Zheng Zhang 2 1 Kobe University 2 Renmin University of China Institute of Statistics
Generalized linear mixed models for biologists
 Generalized linear mixed models for biologists McMaster University 7 May 2009 Outline 1 2 Outline 1 2 Coral protection by symbionts 10 Number of predation events Number of blocks 8 6 4 2 2 2 1 0 2 0 2
Generalized linear mixed models for biologists McMaster University 7 May 2009 Outline 1 2 Outline 1 2 Coral protection by symbionts 10 Number of predation events Number of blocks 8 6 4 2 2 2 1 0 2 0 2
Fall 2017 STAT 532 Homework Peter Hoff. 1. Let P be a probability measure on a collection of sets A.
 1. Let P be a probability measure on a collection of sets A. (a) For each n N, let H n be a set in A such that H n H n+1. Show that P (H n ) monotonically converges to P ( k=1 H k) as n. (b) For each n
1. Let P be a probability measure on a collection of sets A. (a) For each n N, let H n be a set in A such that H n H n+1. Show that P (H n ) monotonically converges to P ( k=1 H k) as n. (b) For each n
Parametric Modelling of Over-dispersed Count Data. Part III / MMath (Applied Statistics) 1
 Parametric Modelling of Over-dispersed Count Data Part III / MMath (Applied Statistics) 1 Introduction Poisson regression is the de facto approach for handling count data What happens then when Poisson
Parametric Modelling of Over-dispersed Count Data Part III / MMath (Applied Statistics) 1 Introduction Poisson regression is the de facto approach for handling count data What happens then when Poisson
Subject CS1 Actuarial Statistics 1 Core Principles
 Institute of Actuaries of India Subject CS1 Actuarial Statistics 1 Core Principles For 2019 Examinations Aim The aim of the Actuarial Statistics 1 subject is to provide a grounding in mathematical and
Institute of Actuaries of India Subject CS1 Actuarial Statistics 1 Core Principles For 2019 Examinations Aim The aim of the Actuarial Statistics 1 subject is to provide a grounding in mathematical and
Biostatistics Workshop Longitudinal Data Analysis. Session 4 GARRETT FITZMAURICE
 Biostatistics Workshop 2008 Longitudinal Data Analysis Session 4 GARRETT FITZMAURICE Harvard University 1 LINEAR MIXED EFFECTS MODELS Motivating Example: Influence of Menarche on Changes in Body Fat Prospective
Biostatistics Workshop 2008 Longitudinal Data Analysis Session 4 GARRETT FITZMAURICE Harvard University 1 LINEAR MIXED EFFECTS MODELS Motivating Example: Influence of Menarche on Changes in Body Fat Prospective
Default Priors and Effcient Posterior Computation in Bayesian
 Default Priors and Effcient Posterior Computation in Bayesian Factor Analysis January 16, 2010 Presented by Eric Wang, Duke University Background and Motivation A Brief Review of Parameter Expansion Literature
Default Priors and Effcient Posterior Computation in Bayesian Factor Analysis January 16, 2010 Presented by Eric Wang, Duke University Background and Motivation A Brief Review of Parameter Expansion Literature
Longitudinal + Reliability = Joint Modeling
 Longitudinal + Reliability = Joint Modeling Carles Serrat Institute of Statistics and Mathematics Applied to Building CYTED-HAROSA International Workshop November 21-22, 2013 Barcelona Mainly from Rizopoulos,
Longitudinal + Reliability = Joint Modeling Carles Serrat Institute of Statistics and Mathematics Applied to Building CYTED-HAROSA International Workshop November 21-22, 2013 Barcelona Mainly from Rizopoulos,
Estimation in Generalized Linear Models with Heterogeneous Random Effects. Woncheol Jang Johan Lim. May 19, 2004
 Estimation in Generalized Linear Models with Heterogeneous Random Effects Woncheol Jang Johan Lim May 19, 2004 Abstract The penalized quasi-likelihood (PQL) approach is the most common estimation procedure
Estimation in Generalized Linear Models with Heterogeneous Random Effects Woncheol Jang Johan Lim May 19, 2004 Abstract The penalized quasi-likelihood (PQL) approach is the most common estimation procedure
On the Behavior of Marginal and Conditional Akaike Information Criteria in Linear Mixed Models
 On the Behavior of Marginal and Conditional Akaike Information Criteria in Linear Mixed Models Thomas Kneib Department of Mathematics Carl von Ossietzky University Oldenburg Sonja Greven Department of
On the Behavior of Marginal and Conditional Akaike Information Criteria in Linear Mixed Models Thomas Kneib Department of Mathematics Carl von Ossietzky University Oldenburg Sonja Greven Department of
GOODNESS-OF-FIT TEST ISSUES IN GENERALIZED LINEAR MIXED MODELS. A Dissertation NAI-WEI CHEN
 GOODNESS-OF-FIT TEST ISSUES IN GENERALIZED LINEAR MIXED MODELS A Dissertation by NAI-WEI CHEN Submitted to the Office of Graduate Studies of Texas A&M University in partial fulfillment of the requirements
GOODNESS-OF-FIT TEST ISSUES IN GENERALIZED LINEAR MIXED MODELS A Dissertation by NAI-WEI CHEN Submitted to the Office of Graduate Studies of Texas A&M University in partial fulfillment of the requirements
Conditional Estimation for Generalized Linear Models When Covariates Are Subject-specific Parameters in a Mixed Model for Longitudinal Measurements
 Conditional Estimation for Generalized Linear Models When Covariates Are Subject-specific Parameters in a Mixed Model for Longitudinal Measurements Erning Li, Daowen Zhang, and Marie Davidian Department
Conditional Estimation for Generalized Linear Models When Covariates Are Subject-specific Parameters in a Mixed Model for Longitudinal Measurements Erning Li, Daowen Zhang, and Marie Davidian Department
Web-based Supplementary Material for A Two-Part Joint. Model for the Analysis of Survival and Longitudinal Binary. Data with excess Zeros
 Web-based Supplementary Material for A Two-Part Joint Model for the Analysis of Survival and Longitudinal Binary Data with excess Zeros Dimitris Rizopoulos, 1 Geert Verbeke, 1 Emmanuel Lesaffre 1 and Yves
Web-based Supplementary Material for A Two-Part Joint Model for the Analysis of Survival and Longitudinal Binary Data with excess Zeros Dimitris Rizopoulos, 1 Geert Verbeke, 1 Emmanuel Lesaffre 1 and Yves
Modification and Improvement of Empirical Likelihood for Missing Response Problem
 UW Biostatistics Working Paper Series 12-30-2010 Modification and Improvement of Empirical Likelihood for Missing Response Problem Kwun Chuen Gary Chan University of Washington - Seattle Campus, kcgchan@u.washington.edu
UW Biostatistics Working Paper Series 12-30-2010 Modification and Improvement of Empirical Likelihood for Missing Response Problem Kwun Chuen Gary Chan University of Washington - Seattle Campus, kcgchan@u.washington.edu
Bayesian Estimation of Prediction Error and Variable Selection in Linear Regression
 Bayesian Estimation of Prediction Error and Variable Selection in Linear Regression Andrew A. Neath Department of Mathematics and Statistics; Southern Illinois University Edwardsville; Edwardsville, IL,
Bayesian Estimation of Prediction Error and Variable Selection in Linear Regression Andrew A. Neath Department of Mathematics and Statistics; Southern Illinois University Edwardsville; Edwardsville, IL,
σ(a) = a N (x; 0, 1 2 ) dx. σ(a) = Φ(a) =
 Until now we have always worked with likelihoods and prior distributions that were conjugate to each other, allowing the computation of the posterior distribution to be done in closed form. Unfortunately,
Until now we have always worked with likelihoods and prior distributions that were conjugate to each other, allowing the computation of the posterior distribution to be done in closed form. Unfortunately,
Joint Modeling of Survival and Longitudinal Data: Likelihood Approach Revisited
 Biometrics 62, 1037 1043 December 2006 DOI: 10.1111/j.1541-0420.2006.00570.x Joint Modeling of Survival and Longitudinal Data: Likelihood Approach Revisited Fushing Hsieh, 1 Yi-Kuan Tseng, 2 and Jane-Ling
Biometrics 62, 1037 1043 December 2006 DOI: 10.1111/j.1541-0420.2006.00570.x Joint Modeling of Survival and Longitudinal Data: Likelihood Approach Revisited Fushing Hsieh, 1 Yi-Kuan Tseng, 2 and Jane-Ling
University of California, Berkeley
 University of California, Berkeley U.C. Berkeley Division of Biostatistics Working Paper Series Year 2008 Paper 241 A Note on Risk Prediction for Case-Control Studies Sherri Rose Mark J. van der Laan Division
University of California, Berkeley U.C. Berkeley Division of Biostatistics Working Paper Series Year 2008 Paper 241 A Note on Risk Prediction for Case-Control Studies Sherri Rose Mark J. van der Laan Division
STA 4273H: Statistical Machine Learning
 STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 3 Linear
STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 3 Linear
Calibration Estimation of Semiparametric Copula Models with Data Missing at Random
 Calibration Estimation of Semiparametric Copula Models with Data Missing at Random Shigeyuki Hamori 1 Kaiji Motegi 1 Zheng Zhang 2 1 Kobe University 2 Renmin University of China Econometrics Workshop UNC
Calibration Estimation of Semiparametric Copula Models with Data Missing at Random Shigeyuki Hamori 1 Kaiji Motegi 1 Zheng Zhang 2 1 Kobe University 2 Renmin University of China Econometrics Workshop UNC
A Fully Nonparametric Modeling Approach to. BNP Binary Regression
 A Fully Nonparametric Modeling Approach to Binary Regression Maria Department of Applied Mathematics and Statistics University of California, Santa Cruz SBIES, April 27-28, 2012 Outline 1 2 3 Simulation
A Fully Nonparametric Modeling Approach to Binary Regression Maria Department of Applied Mathematics and Statistics University of California, Santa Cruz SBIES, April 27-28, 2012 Outline 1 2 3 Simulation
FULL LIKELIHOOD INFERENCES IN THE COX MODEL
 October 20, 2007 FULL LIKELIHOOD INFERENCES IN THE COX MODEL BY JIAN-JIAN REN 1 AND MAI ZHOU 2 University of Central Florida and University of Kentucky Abstract We use the empirical likelihood approach
October 20, 2007 FULL LIKELIHOOD INFERENCES IN THE COX MODEL BY JIAN-JIAN REN 1 AND MAI ZHOU 2 University of Central Florida and University of Kentucky Abstract We use the empirical likelihood approach
Pairwise rank based likelihood for estimating the relationship between two homogeneous populations and their mixture proportion
 Pairwise rank based likelihood for estimating the relationship between two homogeneous populations and their mixture proportion Glenn Heller and Jing Qin Department of Epidemiology and Biostatistics Memorial
Pairwise rank based likelihood for estimating the relationship between two homogeneous populations and their mixture proportion Glenn Heller and Jing Qin Department of Epidemiology and Biostatistics Memorial
Discussion of Missing Data Methods in Longitudinal Studies: A Review by Ibrahim and Molenberghs
 Discussion of Missing Data Methods in Longitudinal Studies: A Review by Ibrahim and Molenberghs Michael J. Daniels and Chenguang Wang Jan. 18, 2009 First, we would like to thank Joe and Geert for a carefully
Discussion of Missing Data Methods in Longitudinal Studies: A Review by Ibrahim and Molenberghs Michael J. Daniels and Chenguang Wang Jan. 18, 2009 First, we would like to thank Joe and Geert for a carefully
Bayesian estimation of the discrepancy with misspecified parametric models
 Bayesian estimation of the discrepancy with misspecified parametric models Pierpaolo De Blasi University of Torino & Collegio Carlo Alberto Bayesian Nonparametrics workshop ICERM, 17-21 September 2012
Bayesian estimation of the discrepancy with misspecified parametric models Pierpaolo De Blasi University of Torino & Collegio Carlo Alberto Bayesian Nonparametrics workshop ICERM, 17-21 September 2012
Biost 518 Applied Biostatistics II. Purpose of Statistics. First Stage of Scientific Investigation. Further Stages of Scientific Investigation
 Biost 58 Applied Biostatistics II Scott S. Emerson, M.D., Ph.D. Professor of Biostatistics University of Washington Lecture 5: Review Purpose of Statistics Statistics is about science (Science in the broadest
Biost 58 Applied Biostatistics II Scott S. Emerson, M.D., Ph.D. Professor of Biostatistics University of Washington Lecture 5: Review Purpose of Statistics Statistics is about science (Science in the broadest
Likelihood ratio testing for zero variance components in linear mixed models
 Likelihood ratio testing for zero variance components in linear mixed models Sonja Greven 1,3, Ciprian Crainiceanu 2, Annette Peters 3 and Helmut Küchenhoff 1 1 Department of Statistics, LMU Munich University,
Likelihood ratio testing for zero variance components in linear mixed models Sonja Greven 1,3, Ciprian Crainiceanu 2, Annette Peters 3 and Helmut Küchenhoff 1 1 Department of Statistics, LMU Munich University,
Likelihood-based inference with missing data under missing-at-random
 Likelihood-based inference with missing data under missing-at-random Jae-kwang Kim Joint work with Shu Yang Department of Statistics, Iowa State University May 4, 014 Outline 1. Introduction. Parametric
Likelihood-based inference with missing data under missing-at-random Jae-kwang Kim Joint work with Shu Yang Department of Statistics, Iowa State University May 4, 014 Outline 1. Introduction. Parametric
Minimum Hellinger Distance Estimation in a. Semiparametric Mixture Model
 Minimum Hellinger Distance Estimation in a Semiparametric Mixture Model Sijia Xiang 1, Weixin Yao 1, and Jingjing Wu 2 1 Department of Statistics, Kansas State University, Manhattan, Kansas, USA 66506-0802.
Minimum Hellinger Distance Estimation in a Semiparametric Mixture Model Sijia Xiang 1, Weixin Yao 1, and Jingjing Wu 2 1 Department of Statistics, Kansas State University, Manhattan, Kansas, USA 66506-0802.
Statistical Practice. Selecting the Best Linear Mixed Model Under REML. Matthew J. GURKA
 Matthew J. GURKA Statistical Practice Selecting the Best Linear Mixed Model Under REML Restricted maximum likelihood (REML) estimation of the parameters of the mixed model has become commonplace, even
Matthew J. GURKA Statistical Practice Selecting the Best Linear Mixed Model Under REML Restricted maximum likelihood (REML) estimation of the parameters of the mixed model has become commonplace, even
Semiparametric Regression
 Semiparametric Regression Patrick Breheny October 22 Patrick Breheny Survival Data Analysis (BIOS 7210) 1/23 Introduction Over the past few weeks, we ve introduced a variety of regression models under
Semiparametric Regression Patrick Breheny October 22 Patrick Breheny Survival Data Analysis (BIOS 7210) 1/23 Introduction Over the past few weeks, we ve introduced a variety of regression models under
Bios 6649: Clinical Trials - Statistical Design and Monitoring
 Bios 6649: Clinical Trials - Statistical Design and Monitoring Spring Semester 2015 John M. Kittelson Department of Biostatistics & Informatics Colorado School of Public Health University of Colorado Denver
Bios 6649: Clinical Trials - Statistical Design and Monitoring Spring Semester 2015 John M. Kittelson Department of Biostatistics & Informatics Colorado School of Public Health University of Colorado Denver
Lasso Maximum Likelihood Estimation of Parametric Models with Singular Information Matrices
 Article Lasso Maximum Likelihood Estimation of Parametric Models with Singular Information Matrices Fei Jin 1,2 and Lung-fei Lee 3, * 1 School of Economics, Shanghai University of Finance and Economics,
Article Lasso Maximum Likelihood Estimation of Parametric Models with Singular Information Matrices Fei Jin 1,2 and Lung-fei Lee 3, * 1 School of Economics, Shanghai University of Finance and Economics,
Estimation and Model Selection in Mixed Effects Models Part I. Adeline Samson 1
 Estimation and Model Selection in Mixed Effects Models Part I Adeline Samson 1 1 University Paris Descartes Summer school 2009 - Lipari, Italy These slides are based on Marc Lavielle s slides Outline 1
Estimation and Model Selection in Mixed Effects Models Part I Adeline Samson 1 1 University Paris Descartes Summer school 2009 - Lipari, Italy These slides are based on Marc Lavielle s slides Outline 1
Smooth Semiparametric Regression Analysis for Arbitrarily Censored Time-to-Event Data
 Smooth Semiparametric Regression Analysis for Arbitrarily Censored Time-to-Event Data Min Zhang and Marie Davidian Department of Statistics, North Carolina State University, Raleigh, North Carolina 27695-8203,
Smooth Semiparametric Regression Analysis for Arbitrarily Censored Time-to-Event Data Min Zhang and Marie Davidian Department of Statistics, North Carolina State University, Raleigh, North Carolina 27695-8203,
Likelihood-Based Methods
 Likelihood-Based Methods Handbook of Spatial Statistics, Chapter 4 Susheela Singh September 22, 2016 OVERVIEW INTRODUCTION MAXIMUM LIKELIHOOD ESTIMATION (ML) RESTRICTED MAXIMUM LIKELIHOOD ESTIMATION (REML)
Likelihood-Based Methods Handbook of Spatial Statistics, Chapter 4 Susheela Singh September 22, 2016 OVERVIEW INTRODUCTION MAXIMUM LIKELIHOOD ESTIMATION (ML) RESTRICTED MAXIMUM LIKELIHOOD ESTIMATION (REML)
Generalized Linear. Mixed Models. Methods and Applications. Modern Concepts, Walter W. Stroup. Texts in Statistical Science.
 Texts in Statistical Science Generalized Linear Mixed Models Modern Concepts, Methods and Applications Walter W. Stroup CRC Press Taylor & Francis Croup Boca Raton London New York CRC Press is an imprint
Texts in Statistical Science Generalized Linear Mixed Models Modern Concepts, Methods and Applications Walter W. Stroup CRC Press Taylor & Francis Croup Boca Raton London New York CRC Press is an imprint
On the Behavior of Marginal and Conditional Akaike Information Criteria in Linear Mixed Models
 On the Behavior of Marginal and Conditional Akaike Information Criteria in Linear Mixed Models Thomas Kneib Institute of Statistics and Econometrics Georg-August-University Göttingen Department of Statistics
On the Behavior of Marginal and Conditional Akaike Information Criteria in Linear Mixed Models Thomas Kneib Institute of Statistics and Econometrics Georg-August-University Göttingen Department of Statistics
Supporting Information for Estimating restricted mean. treatment effects with stacked survival models
 Supporting Information for Estimating restricted mean treatment effects with stacked survival models Andrew Wey, David Vock, John Connett, and Kyle Rudser Section 1 presents several extensions to the simulation
Supporting Information for Estimating restricted mean treatment effects with stacked survival models Andrew Wey, David Vock, John Connett, and Kyle Rudser Section 1 presents several extensions to the simulation
Robust covariance estimator for small-sample adjustment in the generalized estimating equations: A simulation study
 Science Journal of Applied Mathematics and Statistics 2014; 2(1): 20-25 Published online February 20, 2014 (http://www.sciencepublishinggroup.com/j/sjams) doi: 10.11648/j.sjams.20140201.13 Robust covariance
Science Journal of Applied Mathematics and Statistics 2014; 2(1): 20-25 Published online February 20, 2014 (http://www.sciencepublishinggroup.com/j/sjams) doi: 10.11648/j.sjams.20140201.13 Robust covariance
Restricted Maximum Likelihood in Linear Regression and Linear Mixed-Effects Model
 Restricted Maximum Likelihood in Linear Regression and Linear Mixed-Effects Model Xiuming Zhang zhangxiuming@u.nus.edu A*STAR-NUS Clinical Imaging Research Center October, 015 Summary This report derives
Restricted Maximum Likelihood in Linear Regression and Linear Mixed-Effects Model Xiuming Zhang zhangxiuming@u.nus.edu A*STAR-NUS Clinical Imaging Research Center October, 015 Summary This report derives
Marginal Specifications and a Gaussian Copula Estimation
 Marginal Specifications and a Gaussian Copula Estimation Kazim Azam Abstract Multivariate analysis involving random variables of different type like count, continuous or mixture of both is frequently required
Marginal Specifications and a Gaussian Copula Estimation Kazim Azam Abstract Multivariate analysis involving random variables of different type like count, continuous or mixture of both is frequently required
ON THE CONSEQUENCES OF MISSPECIFING ASSUMPTIONS CONCERNING RESIDUALS DISTRIBUTION IN A REPEATED MEASURES AND NONLINEAR MIXED MODELLING CONTEXT
 ON THE CONSEQUENCES OF MISSPECIFING ASSUMPTIONS CONCERNING RESIDUALS DISTRIBUTION IN A REPEATED MEASURES AND NONLINEAR MIXED MODELLING CONTEXT Rachid el Halimi and Jordi Ocaña Departament d Estadística
ON THE CONSEQUENCES OF MISSPECIFING ASSUMPTIONS CONCERNING RESIDUALS DISTRIBUTION IN A REPEATED MEASURES AND NONLINEAR MIXED MODELLING CONTEXT Rachid el Halimi and Jordi Ocaña Departament d Estadística
A Sampling of IMPACT Research:
 A Sampling of IMPACT Research: Methods for Analysis with Dropout and Identifying Optimal Treatment Regimes Marie Davidian Department of Statistics North Carolina State University http://www.stat.ncsu.edu/
A Sampling of IMPACT Research: Methods for Analysis with Dropout and Identifying Optimal Treatment Regimes Marie Davidian Department of Statistics North Carolina State University http://www.stat.ncsu.edu/
Indirect estimation of a simultaneous limited dependent variable model for patient costs and outcome
 Indirect estimation of a simultaneous limited dependent variable model for patient costs and outcome Per Hjertstrand Research Institute of Industrial Economics (IFN), Stockholm, Sweden Per.Hjertstrand@ifn.se
Indirect estimation of a simultaneous limited dependent variable model for patient costs and outcome Per Hjertstrand Research Institute of Industrial Economics (IFN), Stockholm, Sweden Per.Hjertstrand@ifn.se
401 Review. 6. Power analysis for one/two-sample hypothesis tests and for correlation analysis.
 401 Review Major topics of the course 1. Univariate analysis 2. Bivariate analysis 3. Simple linear regression 4. Linear algebra 5. Multiple regression analysis Major analysis methods 1. Graphical analysis
401 Review Major topics of the course 1. Univariate analysis 2. Bivariate analysis 3. Simple linear regression 4. Linear algebra 5. Multiple regression analysis Major analysis methods 1. Graphical analysis
Supplemental Material for KERNEL-BASED INFERENCE IN TIME-VARYING COEFFICIENT COINTEGRATING REGRESSION. September 2017
 Supplemental Material for KERNEL-BASED INFERENCE IN TIME-VARYING COEFFICIENT COINTEGRATING REGRESSION By Degui Li, Peter C. B. Phillips, and Jiti Gao September 017 COWLES FOUNDATION DISCUSSION PAPER NO.
Supplemental Material for KERNEL-BASED INFERENCE IN TIME-VARYING COEFFICIENT COINTEGRATING REGRESSION By Degui Li, Peter C. B. Phillips, and Jiti Gao September 017 COWLES FOUNDATION DISCUSSION PAPER NO.
Introduction to mtm: An R Package for Marginalized Transition Models
 Introduction to mtm: An R Package for Marginalized Transition Models Bryan A. Comstock and Patrick J. Heagerty Department of Biostatistics University of Washington 1 Introduction Marginalized transition
Introduction to mtm: An R Package for Marginalized Transition Models Bryan A. Comstock and Patrick J. Heagerty Department of Biostatistics University of Washington 1 Introduction Marginalized transition
Akaike Information Criterion
 Akaike Information Criterion Shuhua Hu Center for Research in Scientific Computation North Carolina State University Raleigh, NC February 7, 2012-1- background Background Model statistical model: Y j =
Akaike Information Criterion Shuhua Hu Center for Research in Scientific Computation North Carolina State University Raleigh, NC February 7, 2012-1- background Background Model statistical model: Y j =
Stat 5101 Lecture Notes
 Stat 5101 Lecture Notes Charles J. Geyer Copyright 1998, 1999, 2000, 2001 by Charles J. Geyer May 7, 2001 ii Stat 5101 (Geyer) Course Notes Contents 1 Random Variables and Change of Variables 1 1.1 Random
Stat 5101 Lecture Notes Charles J. Geyer Copyright 1998, 1999, 2000, 2001 by Charles J. Geyer May 7, 2001 ii Stat 5101 (Geyer) Course Notes Contents 1 Random Variables and Change of Variables 1 1.1 Random
Sparse Linear Models (10/7/13)
 STA56: Probabilistic machine learning Sparse Linear Models (0/7/) Lecturer: Barbara Engelhardt Scribes: Jiaji Huang, Xin Jiang, Albert Oh Sparsity Sparsity has been a hot topic in statistics and machine
STA56: Probabilistic machine learning Sparse Linear Models (0/7/) Lecturer: Barbara Engelhardt Scribes: Jiaji Huang, Xin Jiang, Albert Oh Sparsity Sparsity has been a hot topic in statistics and machine
Lecture 3. G. Cowan. Lecture 3 page 1. Lectures on Statistical Data Analysis
 Lecture 3 1 Probability (90 min.) Definition, Bayes theorem, probability densities and their properties, catalogue of pdfs, Monte Carlo 2 Statistical tests (90 min.) general concepts, test statistics,
Lecture 3 1 Probability (90 min.) Definition, Bayes theorem, probability densities and their properties, catalogue of pdfs, Monte Carlo 2 Statistical tests (90 min.) general concepts, test statistics,
Plausible Values for Latent Variables Using Mplus
 Plausible Values for Latent Variables Using Mplus Tihomir Asparouhov and Bengt Muthén August 21, 2010 1 1 Introduction Plausible values are imputed values for latent variables. All latent variables can
Plausible Values for Latent Variables Using Mplus Tihomir Asparouhov and Bengt Muthén August 21, 2010 1 1 Introduction Plausible values are imputed values for latent variables. All latent variables can
Power and Sample Size Calculations with the Additive Hazards Model
 Journal of Data Science 10(2012), 143-155 Power and Sample Size Calculations with the Additive Hazards Model Ling Chen, Chengjie Xiong, J. Philip Miller and Feng Gao Washington University School of Medicine
Journal of Data Science 10(2012), 143-155 Power and Sample Size Calculations with the Additive Hazards Model Ling Chen, Chengjie Xiong, J. Philip Miller and Feng Gao Washington University School of Medicine
Latent Variable Model for Weight Gain Prevention Data with Informative Intermittent Missingness
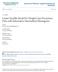 Journal of Modern Applied Statistical Methods Volume 15 Issue 2 Article 36 11-1-2016 Latent Variable Model for Weight Gain Prevention Data with Informative Intermittent Missingness Li Qin Yale University,
Journal of Modern Applied Statistical Methods Volume 15 Issue 2 Article 36 11-1-2016 Latent Variable Model for Weight Gain Prevention Data with Informative Intermittent Missingness Li Qin Yale University,
Contents. Part I: Fundamentals of Bayesian Inference 1
 Contents Preface xiii Part I: Fundamentals of Bayesian Inference 1 1 Probability and inference 3 1.1 The three steps of Bayesian data analysis 3 1.2 General notation for statistical inference 4 1.3 Bayesian
Contents Preface xiii Part I: Fundamentals of Bayesian Inference 1 1 Probability and inference 3 1.1 The three steps of Bayesian data analysis 3 1.2 General notation for statistical inference 4 1.3 Bayesian
Analysing geoadditive regression data: a mixed model approach
 Analysing geoadditive regression data: a mixed model approach Institut für Statistik, Ludwig-Maximilians-Universität München Joint work with Ludwig Fahrmeir & Stefan Lang 25.11.2005 Spatio-temporal regression
Analysing geoadditive regression data: a mixed model approach Institut für Statistik, Ludwig-Maximilians-Universität München Joint work with Ludwig Fahrmeir & Stefan Lang 25.11.2005 Spatio-temporal regression
Ronald Christensen. University of New Mexico. Albuquerque, New Mexico. Wesley Johnson. University of California, Irvine. Irvine, California
 Texts in Statistical Science Bayesian Ideas and Data Analysis An Introduction for Scientists and Statisticians Ronald Christensen University of New Mexico Albuquerque, New Mexico Wesley Johnson University
Texts in Statistical Science Bayesian Ideas and Data Analysis An Introduction for Scientists and Statisticians Ronald Christensen University of New Mexico Albuquerque, New Mexico Wesley Johnson University
Discrepancy-Based Model Selection Criteria Using Cross Validation
 33 Discrepancy-Based Model Selection Criteria Using Cross Validation Joseph E. Cavanaugh, Simon L. Davies, and Andrew A. Neath Department of Biostatistics, The University of Iowa Pfizer Global Research
33 Discrepancy-Based Model Selection Criteria Using Cross Validation Joseph E. Cavanaugh, Simon L. Davies, and Andrew A. Neath Department of Biostatistics, The University of Iowa Pfizer Global Research
Econometrics I, Estimation
 Econometrics I, Estimation Department of Economics Stanford University September, 2008 Part I Parameter, Estimator, Estimate A parametric is a feature of the population. An estimator is a function of the
Econometrics I, Estimation Department of Economics Stanford University September, 2008 Part I Parameter, Estimator, Estimate A parametric is a feature of the population. An estimator is a function of the
Modelling Survival Events with Longitudinal Data Measured with Error
 Modelling Survival Events with Longitudinal Data Measured with Error Hongsheng Dai, Jianxin Pan & Yanchun Bao First version: 14 December 29 Research Report No. 16, 29, Probability and Statistics Group
Modelling Survival Events with Longitudinal Data Measured with Error Hongsheng Dai, Jianxin Pan & Yanchun Bao First version: 14 December 29 Research Report No. 16, 29, Probability and Statistics Group
Sequential Monitoring of Clinical Trials Session 4 - Bayesian Evaluation of Group Sequential Designs
 Sequential Monitoring of Clinical Trials Session 4 - Bayesian Evaluation of Group Sequential Designs Presented August 8-10, 2012 Daniel L. Gillen Department of Statistics University of California, Irvine
Sequential Monitoring of Clinical Trials Session 4 - Bayesian Evaluation of Group Sequential Designs Presented August 8-10, 2012 Daniel L. Gillen Department of Statistics University of California, Irvine
STATS 200: Introduction to Statistical Inference. Lecture 29: Course review
 STATS 200: Introduction to Statistical Inference Lecture 29: Course review Course review We started in Lecture 1 with a fundamental assumption: Data is a realization of a random process. The goal throughout
STATS 200: Introduction to Statistical Inference Lecture 29: Course review Course review We started in Lecture 1 with a fundamental assumption: Data is a realization of a random process. The goal throughout
Power and sample size calculations for designing rare variant sequencing association studies.
 Power and sample size calculations for designing rare variant sequencing association studies. Seunggeun Lee 1, Michael C. Wu 2, Tianxi Cai 1, Yun Li 2,3, Michael Boehnke 4 and Xihong Lin 1 1 Department
Power and sample size calculations for designing rare variant sequencing association studies. Seunggeun Lee 1, Michael C. Wu 2, Tianxi Cai 1, Yun Li 2,3, Michael Boehnke 4 and Xihong Lin 1 1 Department
RESEARCH ARTICLE. Likelihood-based Approach for Analysis of Longitudinal Nominal Data using Marginalized Random Effects Models
 Journal of Applied Statistics Vol. 00, No. 00, August 2009, 1 14 RESEARCH ARTICLE Likelihood-based Approach for Analysis of Longitudinal Nominal Data using Marginalized Random Effects Models Keunbaik Lee
Journal of Applied Statistics Vol. 00, No. 00, August 2009, 1 14 RESEARCH ARTICLE Likelihood-based Approach for Analysis of Longitudinal Nominal Data using Marginalized Random Effects Models Keunbaik Lee
Repeated ordinal measurements: a generalised estimating equation approach
 Repeated ordinal measurements: a generalised estimating equation approach David Clayton MRC Biostatistics Unit 5, Shaftesbury Road Cambridge CB2 2BW April 7, 1992 Abstract Cumulative logit and related
Repeated ordinal measurements: a generalised estimating equation approach David Clayton MRC Biostatistics Unit 5, Shaftesbury Road Cambridge CB2 2BW April 7, 1992 Abstract Cumulative logit and related
Semi-Nonparametric Inferences for Massive Data
 Semi-Nonparametric Inferences for Massive Data Guang Cheng 1 Department of Statistics Purdue University Statistics Seminar at NCSU October, 2015 1 Acknowledge NSF, Simons Foundation and ONR. A Joint Work
Semi-Nonparametric Inferences for Massive Data Guang Cheng 1 Department of Statistics Purdue University Statistics Seminar at NCSU October, 2015 1 Acknowledge NSF, Simons Foundation and ONR. A Joint Work
Bayesian Quantile Regression for Longitudinal Studies with Nonignorable Missing Data
 Biometrics 66, 105 114 March 2010 DOI: 10.1111/j.1541-0420.2009.01269.x Bayesian Quantile Regression for Longitudinal Studies with Nonignorable Missing Data Ying Yuan and Guosheng Yin Department of Biostatistics,
Biometrics 66, 105 114 March 2010 DOI: 10.1111/j.1541-0420.2009.01269.x Bayesian Quantile Regression for Longitudinal Studies with Nonignorable Missing Data Ying Yuan and Guosheng Yin Department of Biostatistics,
Multilevel Statistical Models: 3 rd edition, 2003 Contents
 Multilevel Statistical Models: 3 rd edition, 2003 Contents Preface Acknowledgements Notation Two and three level models. A general classification notation and diagram Glossary Chapter 1 An introduction
Multilevel Statistical Models: 3 rd edition, 2003 Contents Preface Acknowledgements Notation Two and three level models. A general classification notation and diagram Glossary Chapter 1 An introduction
Model Selection for Semiparametric Bayesian Models with Application to Overdispersion
 Proceedings 59th ISI World Statistics Congress, 25-30 August 2013, Hong Kong (Session CPS020) p.3863 Model Selection for Semiparametric Bayesian Models with Application to Overdispersion Jinfang Wang and
Proceedings 59th ISI World Statistics Congress, 25-30 August 2013, Hong Kong (Session CPS020) p.3863 Model Selection for Semiparametric Bayesian Models with Application to Overdispersion Jinfang Wang and
Structural Nested Mean Models for Assessing Time-Varying Effect Moderation. Daniel Almirall
 1 Structural Nested Mean Models for Assessing Time-Varying Effect Moderation Daniel Almirall Center for Health Services Research, Durham VAMC & Dept. of Biostatistics, Duke University Medical Joint work
1 Structural Nested Mean Models for Assessing Time-Varying Effect Moderation Daniel Almirall Center for Health Services Research, Durham VAMC & Dept. of Biostatistics, Duke University Medical Joint work
A simulation study for comparing testing statistics in response-adaptive randomization
 RESEARCH ARTICLE Open Access A simulation study for comparing testing statistics in response-adaptive randomization Xuemin Gu 1, J Jack Lee 2* Abstract Background: Response-adaptive randomizations are
RESEARCH ARTICLE Open Access A simulation study for comparing testing statistics in response-adaptive randomization Xuemin Gu 1, J Jack Lee 2* Abstract Background: Response-adaptive randomizations are
