Diffusion-Based Analysis of. in High-Dimensional Data
|
|
|
- Dwight Potter
- 5 years ago
- Views:
Transcription
1 Raymond and Beverly Sackler Faculty of Exact Science Blavatnik School of Computer Science Diffusion-Based Analysis of Locally Low-Dimensional Geometries in High-Dimensional Data Thesis submitted for the degree of Doctor of Philosophy by Guy Wolf The thesis was carried out under the supervision of Prof. Amir Averbuch Submitted to the Senate of Tel-Aviv University October, 2013
2
3 Dedicated to the memory of my grandfather J.A. Wolf ( ) who passed away during my Ph.D. studies and never got to witness their completion.
4
5 Abstract Data availability has increased rapidly in recent years and massive highdimensional data have become common in many parts of our life. Typically, dozens of parameters are either measured or sensed, and massive volumes of data are collected for analysis. The size and dimensionality of such datasets pose practical challenges on how to extract information from them and perform data analysis on them. Processing massive volumes of collected data requires learning methods to consider local clusters or patches in the analysis. Such learning methods include, for example, dictionary constructions and coarse-graining approaches that reduce the volume of the analyzed data. The high-dimensionality of the data, due to the number of observable (i.e., measured or sensed) parameters, generates the curse of dimensionality phenomenon. To cope with it, the analysis should utilize dimensionality reduction methods with related methodologies, and then process the resulting embedded space. The Diffusion Maps (DM) methodology, which constitutes an important foundation for this thesis, has been recently utilized for many modern data analysis applications. DM is based on constructing a Markovian diffusion process that quantifies the connectivity between data points and follows the dominant geometric patterns in the data. The DM embedding is obtained by spectral analysis of this process (i.e., eigenvalues and eigenvectors of its transition operator). This embedding usually provides a faithful low-dimensional representation of patterns and trends in the analyzed data. This thesis has three parts that enhance and expand the DM methodology (i.e., theories, algorithms, and software). The first two parts follow the manifold assumption. This assumption uses a manifold geometry to model locally low-dimensional patterns and structures in the data. The third part replaces the manifold assumption with a different data model that is based on measure theory. These parts utilize the properties of the DM diffusion process together with underlying geometric properties of the data, which originate from either the manifold geometry or the measure-based model. Each part focuses on a different aspect of DM and provides relevant theories and methods to augment it. i
6 The first part of the thesis focuses on the relations between DM and the underlying manifold geometry of the data under the manifold assumption. This part presents a relation between the intrinsic volume of the manifold and the numerical rank of Gaussian kernels, which play an important role in DM. It also introduces an enhanced patch-based version of DM, which is called Patch-to-Tensor Embedding (PTE). This method considers manifold patches instead of analyzing individual data points. It utilizes the tangential spaces on the manifold together with distances between data points to provide non-scalar affinities between the analyzed patches. Spectral analysis of these affinities is then used to embed the patches into a tensor space that encompasses more information than what the DM embedded space provides. The PTE methodology can also be used to provide patch-based versions of other kernel methods. The second part of the thesis focuses on the DM diffusion process. It provides data analysis methods that preserve the DM diffusion properties while reducing the required volume of the analyzed data. This is done by either coarse-graining and pruning local data clusters, or by constructing a dictionary of representatives that are sufficient for representing the embedded DM diffusion geometry. The suggested coarse-graining preserves the main stochastic properties of the diffusion process, such as ergodicity, which are essential for providing faithful embedding of data points or pruned data clusters. The dictionary construction, on the other hand, is based on preserving the metric space that is defined by the diffusion distance metric. This metric measures proximities and distances between data points according to their connectivity in the underlying DM diffusion process. It is computed by considering all viable (i.e., relatively short and probable) diffusion paths between them. This distance corresponds to Euclidean distances in the DM embedded space and provides an important insight into its geometry. The last part of the thesis enhances the DM method to consider measurebased data modeling instead of the manifold-based model. The suggested model assumes that the sampled data distribution correlates with a measure that represents locally low-dimensional structures in the analyzed data. This model fits well in many modern data analysis scenarios since data densities define a suitable measure for it. These densities originate naturally from collected datasets, which are not necessarily sampled from a manifold. The presented measure-based DM is robust to perturbations, varying local dimensionality, high-curvature geometries, and similar challenging patterns, due to its ability to express and model them as part of the measure assumption. These three parts together extend the DM methodology to provide a diffusion-based framework for analyzing modern big high-dimensional data. ii
7 Acknowledgments First and foremost, I would like to express my deepest gratitude to my adviser, Prof. Amir Averbuch, for all the help and guidance throughout my Ph.D. studies. Amir has guided me through my first steps into academic research since I first started working on my M.Sc. thesis. During our mutual work, I always found his advices to be priceless. He always knew when to provide constructive criticism, when to push harder to get better results, and when to reassure that we are on the right track. I can honestly say I could not have hoped for a better experience than working with him, and I consider him a friend and a mentor. I would like to thank all the people with whom I have worked during my studies. In particular, many thanks are reserved to Moshe Salhov and Dr. Amir Bermanis, with whom I worked closely together during this time. Researching together with them was satisfying and productive, but more importantly, it was fun. I learned a lot from this experience and I hope to continue working with them in the future. I wish to thank Aviv Rotbart, who has been my friend since before I started my Ph.D. studies, and working with him was a pleasure, both personally and professionally. I also wish to thank current and former lab members Yaniv Shmueli, Gil Shabat, Dr. Shahar Harussi, Dr. Gil David and Dr. Neta Rabin for the experience of working together with them. I wish to express my great appreciation to Prof. Yoel Shkolinsky and Prof. Raphi Coifman for fruitful discussions that had an important impact on the nature of my research. I would also like to thank Prof. Pekka Neittaanmäki for a fruitful collaboration and for his great hospitality during my visits to his lab. I am certain that without all these people my research and my Ph.D. studies would not have been so enjoyable, enriching and rewarding. On a personal note, I would like to thank my family for accompanying me through my academic journey. I am grateful to my parents, Amos and Ruthi Wolf, for always guiding me in the right direction. Finally, most importantly, I owe a debt of gratitude to my wife, Karina Wolf, for always bearing with me and for her endless love and support. iii
8 iv
9 Contents Introduction and Preliminaries 1 Contributions and structure of the thesis Published & submitted papers Preliminaries: Diffusion Maps Diffusion Maps Overview Historical background of Diffusion Maps and related works.. 15 I Manifold Learning 17 2 Cover-based bounds on Gaussian numerical ranks Introduction Problem setup Ambient box-based bounds Cover-based bounds Subadditivity of the Gaussian numerical rank The Gaussian convolution operator Examples and discussion Strict inequality and equality in Proposition Cover-based bounds of plain curves Discussion Conclusion Patch-to-tensor embedding Introduction Benefits of patch processing Overview Problem setup DM on discrete data Super-kernel Linear projection super-kernel v
10 3.3.1 Global linear-projection (GLP) super-kernel Local linear-projection super-kernel Diffusion super-kernel Linear-projection diffusion super-kernel Numerical examples Data analysis using patch-to-tensor embedding Electrical impedance breast tissue classification Image segmentation Conclusion Appendix 3.A Technical proofs Linear-projection diffusion Introduction Manifold representation Naïve extensions of DM operators to vector-field settings Extended diffusion operator Linear-projection diffusion Infinitesimal generator Stochastic diffusion process Linear-projection diffusion process demonstration Conclusion II Diffusion-based Learning Coarse-grained localized diffusion Introduction Problem Setup Stochastic view of DM Localized diffusion folders (LDF) Localized Diffusion Process Pruning algorithm Relation to LDF Conclusion Appendix 5.A Proof of Lemma Approximately-isometric diffusion maps Introduction Problem formulation Algorithmic view of DM Diffusion Maps of a partial set vi
11 6.4 An out-of-sample extension that preserves the PDM geometry An efficient computation of the SVD of  The µ-isometric construction Distance accuracy of µidm A spectral bound for the kernel approximation Computational complexity Experimental results Discussion and conclusions Appendix 6.A Proof of the invertibility of the matrix Φ κ (S κ ) Appendix 6.B Proof of Theorem III Measure-based Learning Diffusion-based kernel methods on metric measure spaces Introduction Problem setup DM technicalities in general nonmanifold settings Measure-based diffusion and affinity kernels Spectral properties Infinitesimal generator Geometric examples Noisy spiral curve Uniform grid with a fish-shaped measure Conclusion Appendix 7.A Technical Proofs A.1 Proof of Lemma A.2 Proof of Theorem Appendix 7.B Color maps Conclusions 189 vii
12 viii
13 List of Figures 1 Illustration of the structure of the thesis Covering a manifold (or a compact set) using a set of small (e.g., unit-size) boxes vs. a single box with discretized (e.g., integer) side-lengths An illustration of the flexibility of (l, d)-covers: (a) in lowcurvature areas, the long sides can be set on tangent directions; (b) in high-curvature areas, they can be set on normal (i.e., orthogonal to the tangent) directions Due to the curvature of the unit-circle, for two adjacent data points x and y in a finite dataset (sampled from the unitdiameter circle) there is a ξ-wide band that is not necessary when only covering the dataset, since there are no data points on the arc between them. This band is necessary when the entire (continuous) unit-diameter circle is covered Illustrations of the relations that are used to provide the lengthbased bound of the Gaussian numerical rank of plain curves An illustration of viewing an arbitrary vector in R nd as a n d matrix An illustration of the application of the matrix [a(x, z)o T x a(y, z)o T y ] to the unit vector u Examined manifolds The 100 largest eigenvalues of the LPD-super kernels that corresponds to (a) Sphere, (b) Swiss roll, and (c) Mobius band The CDFs of the eigenvalues of the LPD-super kernels of (a) Sphere, (b) Swiss Roll and (c) Mobius band The PTE segmentation results for the image Cubes when l = 10 and d = 10. The results are shown at several diffusion times t The PTE segmentation results for the image Hand when l = 10 and d = 10. The results are shown at several diffusion times t. 76 ix
14 3.7.3 The PTE segmentation result for the image Sport when l = 10 and d = 17. The results are shown at several diffusion times t The PTE segmentation results for the image Fabric when l = 10 and d = 10. The results are shown at several diffusion times t An illustration of an open set Ũ M around x M, its projection U T x (M) on the tangent space T x (M), and the exponential map exp x, which maps each point y Ũ on the manifold to y U on the tangent space T x (M) The jump of the LPD discrete process goes from time t 0 to time t 1 = t 0 + τ. The jump starts with a vector v x T x (M) that is attached to the manifold at x M. First, a point y T x (M) is chosen according to the transition probabilities of the diffusion operator P (Eq ). Then, the exponential map is used to translate this point to a point y M on the manifold. Finally, the vector v x T x (M) is projected to v y T y (M) and attached to the manifold at y M The results from performing 100 independent iterations of a single transition of the local process defined by u and v (see Section 4.4.2) from x M (in red), with a tangent vector (1, 1) in local coordinates of the tangent space T x (M). The starting point x is marked in red and the destinations of the transitions are marked in blue The results from performing 100 independent iterations of a single transition of the LPD process defined by ũ and ṽ (see Section 4.4.2) starting at x M with a tangent vector (1, 1) in local coordinates of the tangent space T x (M). The points in the area around x on the paraboloid M are presented. The starting point x is marked in orange, the destinations of the transitions are marked in red, and other points in this area are marked in blue Two additional perspectives of the transitions shown in Fig Two independent trajectories of the LPD process defined by ũ and ṽ (see Section 4.4.2) starting at x M with the tangent vector (1, 1) in local coordinates of the tangent space T x (M). The starting point x is marked in orange The projection of the trajectories in Fig on the tangent space T x (M) at the starting point x M of these trajectories. The starting point x is marked in orange Additional perspectives of the trajectory shown in Fig (a).104 x
15 4.4.9 Additional perspectives of the trajectory shown in Fig (b) Illustration of the difference between localized and non-localized paths Illustration of non-trivial localized paths Construction of the pruned-kernel K ij between the clusters C i and C j The relation between Algorithm 2 and the LDF pruning algorithm Examined manifolds The embedding of a Swiss roll via DM and µidm The CDFs of (a) pairwise distance error and (b) coordinates mapping error between µidm and DM embeddings The eigenvalues of DM (denoted by +) and µidm (denoted by o) for each example An illustration of the MGC affinities. It shows a measure that surrounds a straight line (marked in magenta), and the Gaussians around two examined data points (marked in red and blue). The MGC affinity is based on the intersection (marked in dark purple) between the supports of these three functions An illustration of the MGC affinities in two common data analysis scenarios. For every pair of compared data points, the significant values of the integration variable r, from Definition or Proposition 7.3.1, are marked A spiral curve with 5000 noisy data points concentrated around it, and 10 4 points that represent an exponentially-decaying measure around the curve. The colorscale color map from Fig. 7.B.1(a) is used to represent the measure values Two neighborhoods from the Gaussian kernel (K) and the MGC kernels ( K c and K v ) on the spiral curve, using the heatmap in Fig. 7.B.1(b) to represent the kernel values The stationary distributions of: (a) the Gaussian-based diffusion process, and (b) the MGC-based diffusion process. Both use the grayscale color map from Fig. 7.B.1(c) to represent the distribution values The first two diffusion coordinates of the Gaussian-based and MGC-based DM embeddings xi
16 7.4.5 The first three diffusion coordinates of the Gaussian-based and MGC-based DM embeddings Fish shape measure Diffusion degrees and stationary distribution The MGC-based DM embedding of the grid based on the first four MGC-based diffusion coordinates. It is presented in two pairs: 1st-2nd and 3rd-4th The three-dimensional presentation of the embedded grid based on the second, third and fourth MGC-based diffusion coordinates B.1The color maps that are used in Section 7.4, where low values are on the left side and high values are on the right side of the map/scale xii
17 List of Tables The six classes of tissues that are represented in the analyzed dataset Performance summary of the PTE-based classification algorithm Performance summary of the original classification performances from [dsdsj00] µidm computational complexity: m is the size of the ambient space, n is the number of samples, n κ is the dictionary size µidm characterization summary where µ = µidm: spectral difference, bound and empirical measurements 150 xiii
18 xiv
19 Introduction and Preliminaries 1
20
21 Introduction Over the last decade, data collection and storage technologies advanced rapidly and, as a result, the availability of data has significantly increased. Modern datasets consist of massive volumes of observations that are described and quantified by many features that are measured, collected, streamed, and computed. In their raw form, these observations are represented by high-dimensional vectors that contain these features. Such datasets are often referred to as big high-dimensional data. They have become common in many data analysis applications. Analyzing and processing the raw high-dimensional representation of such data becomes impractical, if not impossible, due to the amount of features in the data. The term curse of dimensionality is used as a general name 1 for the problems that cause many learning methods to fail for such data. The main common theme of curse of dimensionality problems is the relation between the volume of a given raw feature space and its dimensionality, which is determined by the number of features in it (whether measured, collected, streamed, or computed). As the number of features increases, the volume of the feature space grows rapidly compared to the concentration areas of the data, which occupy a small portion of this volume. As a result, the data in its raw form becomes too sparse to obtain practical useful knowledge. Common analyzed phenomena usually originate from a small set of hidden (or unknown) underlying factors that generate the larger set of observable (i.e., measured, collected, streamed, or computed) features via nonlinear mappings. This underlying low-dimensional structure in the data provides the foundation and motivation for utilizing dimensionality reduction techniques for data analysis. These techniques embed the analyzed data from their original high-dimensional representation (i.e., the observable features) to low-dimensional representations that correlate with the hidden underlying 1 To the author s best knowledge, this term was first used by Richard E. Bellman in [Bel57] in the context of dynamic optimization. It has since been used in various other contexts and has become a popular name for the described challenges in analyzing highdimensional data. 3
22 4 factors. For example, the Independent Component Analysis (ICA) approach aims to find the independent components (i.e., the underlying factors) of observed data by applying a known model to the nonlinear maps that generate the analyzed data [Sin06b, SC08]. Furthermore, data points with similar low-dimensional representations (i.e., according to the underlying factors of the analyzed phenomena) typically have similar high-dimensional representations (i.e., according to the observable features) and vice versa. This crucial property is utilized by kernel methods in order to perform nonparametric data analysis of massive highdimensional datasets. These methods extend the well known MDS [CC94, Kru64] method. They are based on a construction of an affinity kernel that encapsulates the relations (distances, similarities, or correlations) between data points. Spectral analysis of this kernel provides an efficient representation of the data that simplifies its analysis. The nonparametric nature of this analysis eliminates the redundancies of the observable features, uncovers the important underlying factors, and reveals the geometry of the data. The MDS method uses the eigenvectors of a Gram matrix, which contains the inner products between the data points in the analyzed dataset, to define a mapping of these data points into an embedded space that preserves (or approximates) most of these inner products. This method is equivalent to PCA [Jol86, Hot33], which projects the data onto the span of the principal directions of the variance of the data. Both of these methods capture linear structures in the data. They separate between meaningful directions, which represent the distribution of the data, and noisy uncorrelated directions. The former ones are associated with significant eigenvalues (and eigenvectors) of the Gram matrix, while the latter ones are associated with small eigenvalues. Kernel methods, such as Isomap [TdSL00], LLE [RS00], Laplacian Eigenmaps [BN03], Hessian Eigenmaps [DG03] and Local Tangent Apace Alignment [YXZ08, ZZ02], extend the MDS paradigm by considering locally-linear structures in the data. A convenient interpretation of these structures is provided by the assumptions that they form a low-dimensional manifold that captures the dependencies between the observable features in the data. This is called the manifold assumption. Namely, the data is assumed to be sampled from this manifold. The resulting spectrally-embedded space in these methods preserves the intrinsic geometry of the manifold, which incorporates and correlates with the underlying factors of the analyzed phenomena in the data. Kernel methods are also inspired from spectral graph theory [Chu97]. The defined kernel can be interpreted as a weighted adjacency matrix of a graph whose vertices are the data points. The edges of this graph are defined and weighted by the local relations (or similarities) in the kernel
23 matrix. The analysis of the eigenvalues and the corresponding eigenvectors of this matrix reveals many qualities and connections in this graph, which serves as a discretization of the continuous manifold geometry. The diffusion maps (DM) kernel method [CL06a] utilizes a stochastic diffusion process to analyze data. It defines diffusion affinities via symmetric conjugation of a transition probability operator. These probabilities are based on local distances between data points. The Euclidean distances in the DM embedded space correspond to a diffusion distance metric in the observable space. This distance metric quantifies the connectivity between data points by incorporating all the diffusion paths between them. When the data is sampled from a low-dimensional manifold, these diffusion paths follow the intrinsic geometry of the manifold (i.e., geodesic paths on it). Therefore, the resulting diffusion distances capture the underlying manifold geometry of the data. The diffusion distance metric was utilized for clustering & classification [DA12], parametrization of linear systems [TKC + 12], and shape recognition [BB11]. Furthermore, the DM method was used in a wide variety of data analysis and pattern recognition applications. Examples include audio quality improvement by suppressing transient interference [TCG13], moving vehicle detection [SAR + 10], scene classification [JYS09], gene expression analysis [RDW07] and source localization [TCG11]. In addition, several enhancements and enrichments of the DM methodology were presented recently. For example, in [LKC06, KCLZ10], the DM methodology is enhanced to consider several data sources and fuse the generated datasets into a single embedded representation. In [SW11, SW12, SSHZ11], the DM affinities were enhanced to consider the orientation and local neighborhoods of the underlying manifold, and the resulting embedded space was used for Cryo EM applications. This thesis presents further enhancements of the DM methodology by utilizing the underlying manifold structure of the data, the nature of the diffusion process on it, and a formulation of the distribution of data using measure theory. These enhancements expand DM to provide an extensive diffusion framework for analyzing the locally low-dimensional structures that usually exist in modern high-dimensional data. 5 Contributions and structure of the thesis This thesis explores the properties of the diffusion-based embedding provided by the DM methodology from [CL06a, Laf04] and presents several theoretical and practical enhancements of this methodology. For completeness, a brief
24 6 overview of DM is presented in Chapter 1. The presented enhancements utilize the properties of DM together with the properties of the assumed models and structures from which the analyzed data are sampled. These enhancements provide novel theories and methods for dealing with highdimensional data in order to extract additional intelligence and information from them using the DM methodology. Part III: Measure-based learning Modeled-geometry properties Chap. 7: Diffusionbased kernel methods on metric measure spaces Part I: Manifold learning Chap. 2: Cover-based bounds on Gaussian numerical ranks Chap. 3: Patch-to-Tensor Embedding Chap. 4: Linear-projection diffusion Diffusion Maps Underlying-manifold assumption Chap. 5: Coarse-grained localized diffusion Chap. 6: Approximately-isometric diffusion maps Part II: Diffusion-based learning Figure 1: Illustration of the structure of the thesis The thesis has three parts, as illustrated in Fig. 1. The first part of the thesis considers the underlying-manifold model of the data. The presented theories in this part utilize the intrinsic properties of the assumed manifold to introduce additional information to kernel methods in general, and the diffusion process of DM in particular. The second part of the thesis considers diffusion-specific properties (e.g., transition probabilities and diffusion distances). It provides a coarse-graining method for pruning data clusters and a diffusion-based dictionary construction for reducing the required volume of analyzed data. While the first two parts work under the manifold assumption, the third part provides an alternative modeling of the data based on
25 measure theory. Using a measure structure is natural in many cases when the data distribution is available due to the large number of collected observations. This third and final part of the thesis presents a diffusion process that follows the distribution of the data and explores its properties without requiring the restrictive (and sometimes artificial and inapplicable) manifold assumption. The rest of this section presents brief overviews of each of these parts. Manifold learning (Part I) Common approaches to process high-dimensional data assume that such data lie on a low-dimensional manifold immersed in a high-dimensional ambient space. The geometric structure of these manifolds can then be used to represent redundancies and dependencies in the data. Kernel methods use this geometry to introduce the notion of local affinities between data points via the construction of a suitable kernel. Spectral analysis of the kernel yields an efficient (preferably low-dimensional) representation of the data, which encompasses its intrinsic properties and reveals the patterns that govern it. The underlying manifold geometry provides a powerful tool for deducing information from high-dimensional data. The underlying manifold structure is utilized in Chapter 2, which is based on [BWA13a], to achieve an upper bound on the numerical rank of Gaussian affinity kernels based on a discretized volume of the underlying manifold. This numerical rank determines the dimensionality of the embedded space of kernel-based dimensionality reduction methods that are often based on Gaussian affinities. The discretized volume, which bounds this numerical rank, is based on covering the manifold with a set of small boxes that cover its local neighborhoods. Each box bounds the numerical rank in its local neighborhood. Then, the subadditivity of the numerical rank is used to combine their volumes and achieve the numerical rank bound. The presented bound extends a previous result from [BAC13] that is based on the extrinsic volume of the data in the high-dimensional ambient space. Essentially, the achieved cover-based bound provides an insight into the relations between the manifold that is used to model the data and the number of underlying factors that govern it. Most standard kernel methods use scalar affinities to define the kernel. Chapter 3, which is based on [SWA12], presents a new methodology called Patch-to-Tensor Embedding (PTE). The PTE methodology extends these scalar relations to matrix relations that encompass multidimensional similarities between local neighborhoods on the manifold. The resulting superkernel is a block matrix that contains these matrix relations as its blocks. 7
26 8 Spectral analysis of this super-kernel provides a representation that embeds the patches (i.e., local neighborhoods) of the manifold to tensors, instead of embedding data points from the manifold to vectors. These tensors can represent and express many additional properties of the analyzed data by considering local areas and incorporating directional information instead of just taking a single data point into consideration. Several constructions of super-kernels for the PTE methodology are presented in Chapter 3 and in [SWA12]. They mainly rely on weighted linearprojections between the tangent spaces of the manifold in order to define the matrix relations between patches. These constructions are shown to preserve important spectral properties of the scalar weights that are used in conjunction with the projections. Of particular importance is the Linear- Projection Diffusion (LPD) super-kernel, which extends DM to the PTE scheme. This construction is further discussed in Chapter 4, which is based on [WA13], where its infinitesimal properties are examined and related to a vector-propagating diffusion process over the manifold. Diffusion-based learning (Part II) The kernel in the DM method is based on a Markovian diffusion process that follows the geometry of the manifold and the distribution of the data over it. The affinities in this case result from comparing between random-walk distributions around data points. The resulting distances in the embedded low-dimensional space according to DM correspond to diffusion distances in the original data (or in the underlying manifold). The diffusion distances were enhanced in the PTE framework [SWA12], which is presented in Part I of the thesis, to consider directional information and patch-based relations. The underlying diffusion process in DM can be used to provide additional information beyond the geometric information from the manifold. This information is used in Chapter 5, which is based on [WRDA12], to provide a coarse graining method for the diffusion process. This method can be utilized to obtain hierarchical clustering of the data. The coarse-grained diffusion process, which describes the relations between clusters, is shown to have the same stochastic properties (e.g., ergodicity) as the original diffusion process in DM. The relations between clusters in this process incorporate localized random walks while eliminating the global stationary distribution, which usually becomes more dominant as the diffusion time (i.e., number of steps in the random walks) becomes larger, in order to encompass wider clusters. The resulting localized diffusion kernel provides a diffusion-based representation of clusters and can be used for embedding purposes or for further applications of clustering and coarse graining phases in order to pro-
27 vide a hierarchy of clusters, as done in the Localized Diffusion Folders (LDF) method [DA12]. In fact, this approach provides a theoretical foundation for the LDF method. Another utilization of the diffusion properties is shown in Chapter 6, which is based on [SBWA13b], for diffusion-based dictionary construction. The presented construction algorithm selects a small dictionary set of representative data points that are sufficient for approximating the metric structure of the DM embedded space. The constructed dictionary is then utilized to compute an efficient embedding of the entire input dataset. The resulting embedded space is approximately isometric to the embedded space of DM. Namely, the Euclidean distances in this space are approximately the same as the distances in the DM embedded space and the diffusion distances between data points. Measure-based learning (Part III) The underlying manifold structure is convenient and powerful but it is also restrictive. A manifold has the same dimensionality in every local area. In order to conform to the manifold structure, the number of dominant directions should be the same in every patch (or neighborhood) in the data. Furthermore, in order for these dominant directions to represent tangential or intrinsic directions on the manifold (as opposed to normal directions, which are orthogonal to the manifold), a bounded curvature should be assumed for it. Finally, theoretical results that are obtained under the assumption that the data is sampled from a manifold are not necessarily applicable in practice when the underlying manifold is not known (or does not even exist). One example for such a case is when analyzing noisy data that lies around the manifold and not on it. The noise in such data can create false links between intrinsically unrelated areas. Another example is when the data contains anomalies that do not lie on the main normal-behavior manifold. Thus, the intrinsic representation does not apply to these abnormal data points. Chapter 7, which is based on [BWA12], presents a new approach for utilizing a diffusion-based analysis with a less restrictive assumption of having a measure-based underlying structure. The diffusion kernels in this approach are defined by incorporating local distances in the data together with measure-based information, which represents the data distribution or its density. The generalized construction does not require an underlying manifold to provide a meaningful kernel interpretation. Instead, it works under a more relaxed assumption that the measure and its support are related to a locally low dimensional nature of the analyzed phenomena. This kernel is shown to 9
28 10 satisfy the necessary spectral properties that are required in order to provide a low-dimensional embedding of the data. Additionally, the associated diffusion process is analyzed via its infinitesimal generator. Published & submitted papers Journal papers: 1. A. Bermanis, G. Wolf, and A. Averbuch. Cover-based bounds on the numerical rank of Gaussian kernels. Applied and Computational Harmonic Analysis, DOI: /j.acha G. Wolf and A. Averbuch. Linear-projection diffusion on smooth Euclidean submanifolds. Applied and Computational Harmonic Analysis, 34(1):1 14, G. Wolf, A. Rotbart, G. David, and A. Averbuch. Coarse-grained localized diffusion. Applied and Computational Harmonic Analysis, 33(3): , Y. Shmueli, G. Wolf, and A. Averbuch. Updating kernel methods in spectral decomposition by affinity perturbations. Linear Algebra and its Applications, 437(6): , M. Salhov, G. Wolf, and A. Averbuch. Patch-to-tensor embedding. Applied and Computational Harmonic Analysis, 33(2): , M. Salhov, A. Bermanis, G. Wolf, and A. Averbuch. Approximately-isometric diffusion maps. Submitted to Applied and Computational Harmonic Analysis, M. Salhov, A. Bermanis, G. Wolf, and A. Averbuch. Approximate patch-to-tensor embedding via dictionary construction. Submitted to Pattern Analysis and Machine Intelligence, A. Bermanis, G. Wolf, and A. Averbuch. Diffusion-based kernel methods on Euclidean metric measure spaces. Submitted to Applied and Computational Harmonic Analysis, 2012.
29 11 Conference papers: 1. A. Bermanis, G. Wolf, and A. Averbuch. Measure-based diffusion kernel methods. In SampTA 2013: 10th international conference on Sampling Theory and Applications, Bremen, Germany, M. Salhov, G. Wolf, A. Bermanis, and A. Averbuch. Constructive sampling for patch-based embedding. In SampTA 2013: 10th international conference on Sampling Theory and Applications, Bremen, Germany, G. Wolf, Y. Shmueli, S. Harussi, and A. Averbuch. Polar classification of nominal data. In S. Repin, T. Tiihonen, and T. Tuovinen, editors, Numerical Methods for Differential Equations, Optimization, and Technological Problems, volume 27 of Computational Methods in Applied Sciences, pages Springer Netherlands, M. Salhov, G. Wolf, A. Bermanis, A. Averbuch, and P. Neittaanmäki. Dictionary construction for patch-to-tensor embedding. In J. Hollmén, F. Klawonn, and A. Tucker, editors, Advances in Intelligent Data Analysis XI, volume 7619 of Lecture Notes in Computer Science, pages Springer Berlin Heidelberg, M. Salhov, G. Wolf, A. Averbuch, and P. Neittaanmäki. Patchbased data analysis using linear-projection diffusion. In J. Hollmén, F. Klawonn, and A. Tucker, editors, Advances in Intelligent Data Analysis XI, volume 7619 of Lecture Notes in Computer Science, pages Springer Berlin Heidelberg, G. Wolf, Y. Shmueli, S. Harussi, and A. Averbuch. Polar clustering. In ECCOMAS Thematic Conference on Computational Analysis and Optimization, Funding acknowledgments The author of this thesis was supported by the Eshkol Fellowship from the Israeli Ministry of Science & Technology. The presented researches in the thesis were also partially supported by the Israel Science Foundation (Grant No. 1041/10) and the Ministry of Science & Technology (Grant No ).
30 12
31 Chapter 1 Preliminaries: Diffusion Maps The DM method plays a fundamental role in this thesis since most of the presented theories, tools, and results in it explore and enhance the properties of this method. For completeness, a brief overview of DM and its main properties is presented in this chapter. More detailed descriptions of specific aspects or variations of this methodology are presented in the relevant chapters of the thesis. 1.1 Diffusion Maps Overview Let M R m be a dataset that is sampled from a manifold M that lies in the ambient space R m. For the purpose of this chapter, the dataset M will be considered as an infinite contiguous subset of M, which may even consist of the entire manifold. Let d m be the intrinsic dimension of M, thus, it has a d-dimensional tangent space T x (M), which is a subspace of R m, at every point x M. If the manifold is densely sampled, the tangent space T x (M) can be approximated by a small enough neighborhood around x M. The DM method [CL06a, Laf04] analyzes the dataset M by exploring the geometry of the manifold M from which it is sampled. This method is based on defining an isotropic kernel operator Kf(x) = k(x, y)f(y)dy (for M f : M R) that consists of the affinities k(x, y), x, y M. The affinities in this kernel represent similarity, or proximity, between data-points in the dataset and on the manifold. The kernel K can be viewed as a construction of a weighted graph over the dataset M. The points in M are used as vertices in this graph and the weights of the edges are defined by the affinities in K. These affinities in K are assumed (or required) to satisfy the following properties: Each data-point has positive self-affinity: k(x, x) > 0, x M; 13
32 14 Affinities are non-negative: k(x, y) 0, x, y M; Affinities are symmetric: k(x, y) = k(y, x), x, y M; The graph defined by the weighted adjacencies in K is connected. A popular affinity kernel, which is used in most of the chapters in this thesis, is the Gaussian kernel k(x, y) e x y ε, x, y M, with a suitable ε > 0. This kernel is also used in other dimensionality reduction methods (e.g., Laplacian Eigenmaps [BN03]) as well as out-of-sample extension methods (e.g., Geometric Harmonics [CL06b] and MSE [BAC13]). Other alternatives include slight variations of this kernel (e.g., k(x, y) exp( x y 2 /2ε), which is used in Chapter 7) and the shake & bake kernel from [DA12], which is further discussed in Chapter 5. The graph that is represented by K represents the intrinsic structure of the manifold and is used by DM to construct a Markovian (random-walk) diffusion process that follows it. The degree of each data point (i.e., vertex) x M in this graph is defined as q(x) k(x, y)dy. The diffusion process M is defined by normalizing the kernel K with these to obtain the transition probabilities p(x, y) k(x, y)/q(x), from x M to y M. These probabilities constitute the row stochastic transition operator P f = p(x, y)f(y)dy M of the diffusion process. The diffusion maps method computes an embedding of data points on the manifold into a Euclidean space whose dimensionality is usually significantly lower than the original data dimensionality. This embedding is a result of spectral analysis of the diffusion kernel. Thus, it is preferable to work with a symmetric conjugate to P, which is denoted by A and its elements are a(x, y) = k(x,y) = q 1/2 (x)p(x, y)q 1/2 (y), x, y M. The operator A q(x)q(y) is usually referred to as the diffusion affinity kernel or as the symmetric diffusion kernel. This construction is also known as the (normalized) graph Laplacian in spectral graph theory [Chu97]. In addition, the DM operators were shown in [Laf04, CL06a, NLCK06a, NLCK06b] to be related (under certain density normalizations) to the Fokker-Plank diffusion operators and to the Laplace-Beltrami operator of the underlying manifold M via their infinitesimal generators. This aspect of DM is further discussed in Chapters 4 and 7. Under mild conditions on the kernel K, which are satisfied for the kernels in this thesis, the resulting diffusion affinity kernel A has a discrete decaying spectrum of eigenvalues 1 = λ 0 λ 1 λ 2... (see [Laf04]). These eigenvalues are used in DM together with their corresponding eigenvectors 1 = φ 0, φ 1, φ 2,... to define the DM embedding of the data. Each
33 data point x M is embedded by DM to the diffusion coordinates Φ(x) ( λ 1 φ 1 (x),..., λ δ φ δ (x)), where the exact value of δ depends on the spectrum of A. In most cases, δ is significantly smaller than the original dimensionality of the observable data. As a result of the spectral theorem, the Euclidean distances in the embedded space correspond to the diffusion distance metric of the diffusion process defined by P [CL06a, Laf04]. This metric quantifies the connectivity between data points in the diffusion process by considering the diffusion paths between them. It is defined as either p(x, ) p(y, ) or a(x, ) a(y, ) (depending on the used variation of DM) for x, y M. Therefore, the resulting embedded space of DM follows the geometry that is defined by the underlying diffusion process of DM. When the data is sampled from a lowdimensional manifold, this diffusion geometry reveals the intrinsic structure of the underlying manifold and the DM embedding provides a meaningful representation of the data. Further analysis techniques can then be applied to the embedded space to perform common learning tasks, such as clustering, classification and anomaly detection [Dav09, Rab10]. 1.2 Historical background of Diffusion Maps and related works The DM embedding approach originates from the Laplacian Eigenmaps method, which was first introduced by M. Belkin and P. Niyogi in [BN03], in relation to manifold learning. The interpretation of a graph Laplacian in terms of the infinitesimal generator for the diffusion as well as the choice of weights that are based on the heat equation were also given there. The formal result for the convergence of data graph Laplacian to the Laplace-Beltrami operator on the manifold for the case of uniform distribution appeared in the thesis [Bel03] of M. Belkin, which was also published as [BN08]. This convergence result was generalized in the thesis [Laf04] of S. Lafon (parts of which were also publishes in [CL06a, CL06b]), where density dependent analysis (similar to Theorem in this thesis) and certain normalizations were introduced. Lafon s thesis also introduced the rescaling of the Laplacian eigenmap based on the eigenvalues of the diffusion kernel, the actual Diffusion Map and the corresponding notion of Diffusion Distance. There have consequently been a number of studies, such as [GK06, HJAvL07, Sin06a, NLCK06a, NLCK06b], expanding and generalizing these results as well as providing better bounds and more general analysis. Specific related works will be discussed in more details in the appropriate chapters. 15
34 16
35 Part I Manifold Learning 17
36
37 Chapter 2 Cover-based bounds on the numerical rank of Gaussian kernels A popular approach for analyzing high-dimensional datasets is to perform dimensionality reduction by applying nonparametric affinity kernels. Usually, it is assumed that the represented affinities are related to an underlying lowdimensional manifold from which the data is sampled. This approach works under the assumption that, due to the low-dimensionality of the underlying manifold, the kernel has a low numerical rank. Essentially, this means that the kernel can be represented by a small set of numerically-significant eigenvalues and their corresponding eigenvectors. In this chapter, an upper-bound is presented for the numerical rank of Gaussian convolution operators. These operators are commonly used as kernels by spectral manifold-learning methods. The achieved bound is based on the underlying geometry that is provided by the manifold from which the dataset is assumed to be sampled. The bound can be used to determine the number of significant eigenvalues/eigenvectors that are needed for spectral analysis purposes. Furthermore, the results in this chapter provide a relation between the underlying geometry of the manifold (or dataset) and the numerical rank of its Gaussian affinities. The presented bound in this chapter is called cover-based bound because it is computed by using a finite set of small constant-volume boxes that cover the underlying manifold (or the dataset). This chapter presents coverbased bounds for finite Gaussian-kernel matrices as well as for the continuous Gaussian convolution operator. The relations between the achieved bounds in the finite and continuous cases are explored and demonstrated. The coveroriented methodology is also used to provide a relation between the geodesic 19
38 20 length of a curve and the numerical rank of Gaussian kernels of datasets that are sampled from it. The results in this chapter appear in [BWA13a]. 2.1 Introduction The rapid development of data collection techniques together with high availability of data and storage space introduce increasingly big high-dimensional datasets that fit data analysis tasks. In many cases the quantity of data does not reflect on its quality. Usually, it contains many redundancies that do not add important information over a limited set of representatives. Furthermore, more often than not, the distribution of samples (also called data points) is significantly affected by the sampling techniques that are used. These problems affect both the massive size of the sampled datasets and their high dimensionality, which in turn prevents many methods from being effective tools to analyze these datasets due to the curse of dimensionality phenomenon. Due to the vast number of observable quantities that can be measured, sensed and used as parameters or features, the raw representation of the data is usually high-dimensional. Recent dimensionality reduction methods use manifolds to cope with this problem. Under this manifold existence assumption, a dataset is assumed to be sampled from an Euclidean submanifold that has a relatively small intrinsic dimension. The ambient high dimensional Euclidean space of the manifold is defined by the raw parameters (or features) of the dataset. These parameters are mapped via non-linear functions to low-dimensional coordinates of the manifold, which represent the independent factors that control the behaviors of the analyzed phenomenon. Several methods have been suggested to provide a low-dimensional representation of data points by preserving the intrinsic structure of their underlying manifold. Kernel methods such as k-pca [MSS + 99, SSM98], LLE [RS00], Isomap [TdSL00], Laplacian Eigenmaps [BN03], Hessian Eigenmaps [DG03], Local Tangent Space Alignment [YXZ08, ZZ02] and Diffusion Maps [CL06a] have been used for this task. These methods extend the classical PCA [Jol86, Hot33] and MDS [CC94, Kru64] methods that project the data on a lowdimensional hyperplane that preserves most of the variance in the dataset. Kernel methods substitute the linear relations (i.e., inner-products) that are preserved by PCA and MDS with a kernel construction that introduces the synonymous notion of similarity, proximity, or affinity between data points. Spectral analysis of this kernel is used to obtain an embedding of the data points into a Euclidean space while preserving the kernel s qualities, which are based on non-linear local qualities of the underlying manifold.
39 Beside the high-dimensionality of the data, its size (i.e., number of sampled data-points) is usually very big. The massive size of the dataset is mostly due to the ease of obtaining data points. For example, most systems nowadays collect detailed logs of every action, event and operation that occur with high frequency over long periods of time. However, most of the collected data points are redundant, either because they are near-duplicates of other already-measured data points, or because their properties can be interpolated by suitable subsets of representatives. Therefore, a combination of subsampling and out-of-sample extension techniques can alleviate performance issues that massive datasets entail, and provide a more suitable representation of the analyzed data. Optimally, such a representation would not be affected by the availability of the data or by a sampling method but only rely on the behavior of the observed and analyzed phenomena. The kernel approach, which is used for dimensionality reduction, has been applied for the described out-of-sample extension tasks. A classical kernel-based technique is the Nyström extension [PTVF92, Bak77]. More recent methods are Geometric Harmonics (GH) [CL06b] and the Multiscale Extension (MSE) in [BAC13]. These methods use the spectral decomposition of the kernel (i.e., its eigenvalues and eigenvectors) as a basis of its range. The eigenfunctions are shown to be easily extended to new data points, thus any function in its range, which can be expressed as a linear combination of these eigenfunctions, is also easily extended. Functions that are not in the range of the kernel are extended by projecting them on the kernel s range and using the resulting function (and extension) as an approximation of the original function. There are many kernel methods that work under the assumption that the used kernel has a small set of significant eigenvalues that should be considered for the analysis, and the rest are negligible in the sense that they are numerically zero. This can be phrased as a low numerical rank assumption, where the numerical rank is the number of numerically nonzero eigenvalues or singular values (see Definition for an explicit formulation). While in practice this assumption is usually satisfied, most papers do not present rigorous mathematical support (beyond intuition) for it. In this chapter, which is based on [BWA13a], we present upper bounds for the numerical rank of affinity kernels. We focus on Gaussian kernels, which are popular in many spectral kernel methods (e.g. [CL06a, BN03]). Such an upper bound was achieved in [BAC13] based on a bounding box volume of the analyzed dataset in the observable ambient space. We refine this bound by considering the underlying geometry that is provided by the underlying manifold from which the dataset is assumed to be sampled. Instead of using a single large bounding box, we use a finite set of small constant-volume boxes 21
40 22 that cover the dataset (or its underlying manifold), and use the minimal cover to provide a cover-based bound. When the constant size of the boxes is large enough to cover the whole dataset with one box, this bound converges to the one in [BAC13]. Thus, it is at least as tight as this already established one. The analysis of the numerical rank in this chapter is related to the spectral convergence of the Gaussian kernel (i.e., convergence of its eigenvalues and eigenvectors) as the size of the data tends to infinity and the dataset tends to the entire manifold. Such convergence was discussed in [BN06, BN08] for a Gaussian-based graph Laplacian kernel, which is obtained via appropriate normalization of the Gaussian kernel. There, the eigenvalues and eigenvectors of the graph Laplacian are shown to converge to the eigendecomposition of the Laplace-Beltrami operator on the underlying manifold. The techniques used in this chapter are also closely related to the techniques used in [vlbb08] for examining the spectral convergence of similar kernels in order to analyze the consistency of spectral clustering as the number of analyzed data samples increases. However, while these related works examine the graph Laplacian kernel, the results in this chapter are proved for the Gaussian kernel without requiring its normalization. This unnormalized kernel is used, for example, in spectral out-of-sample extension techniques such as GH [CL06b, Laf04] and MSE [BAC13]. Furthermore, the results in this chapter focus on establishing a clear bound (in terms of the underlying data manifold geometry of the data) on the numerical ranks of Gaussian kernel, both as a finite matrix and as a continuous operator, in addition to the examination of its spectral convergence. The chapter has the following structure. The problem setup and a previously-established bound are described in Section 2.2. The refined cover-based bounds are established in Section 2.3. Section 2.4 demonstrates various nuances and concepts of cover-based bounds, as well as their theoretical application for proving relations between the geodesic length of curves and the numerical rank of datasets that are sampled from these curves. 2.2 Problem setup Let M be a low-dimensional compact manifold that lies in the high-dimensional ambient space R m that has an Euclidean metric. In addition, let β be the Borel σ-algebra on M and let µ be a probability measure on (M, β). Finally, let M R m be a set of n data points (i.e., n = M ) sampled from the manifold M. Define the affinity between two data points x, y M to be g ε (x, y) = e x y 2 /ε where ε is a positive parameter. Let G M ε be an n n affinity
41 23 kernel between the data points in M, where each row and each column of G M ε corresponds to a single data point in the dataset M, and each cell contains the affinity g ε (x, y) between the row s data point x M and the column s data point y M. The matrix G M ε is called the Gaussian kernel over the dataset M. This kernel introduces the notion of affinities and local neighborhoods of data points in the dataset M (or on the manifold M) due to the exponential decay of it s values in relation to the distances between data points. The Gaussian kernel with its spectral analysis and its spectral decomposition are utilized for dimensionality reduction in [CL06a, CLL + 05, BN03] and for outof-sample function extension in [CL06b, BAC13]. We denote the rank of the Gaussian kernel G M ε (i.e., the dimension of its range or, equivalently, the number of its nonzero eigenvalues) by ρ(g M ε ). Usually, the kernel will not have strictly nonzero eigenvalues. However, since its spectrum decays rapidly (i.e., exponentially), most of its eigenvalues will have negligible (albeit nonzero) values, and it will only have a limited number of numerically significant eigenvalues from a practical analysis point of view. Therefore, the algebraic rank is insufficient to characterize the spectral properties of G M ε. A more desirable characteristic needs to consider the number of numerically nonzero (i.e., numerically significant) eigenvalues based on a predetermined significance threshold δ. Definition introduces the numerical rank of G M ε for this purpose 1. This definition is standard in many papers that use spectral analysis. Definition The numerical rank of the Gaussian kernel G M ε precision δ 0 is { ρ δ (G M ε ) # j : σ } j(g M ε ) σ 1 (G M ε ) > δ, up to where σ j (G M ε ) denotes the j-th largest singular value of the matrix G M ε. The numerical rank ρ δ (G M ε ) determines the dimension of the embedded space that is achieved by dimensionality reduction methods such as Diffusion Maps [CL06a, CLL + 05] and Laplacian Eigenmaps [BN03], and the number of harmonics or sampled representatives that are used by out-of-sample methods such as Geometric Harmonics [CL06b] and the Multiscale Extension in [BAC13]. We notice that when the significance threshold δ is zero then 1 Specifically, for the discussed Gaussian kernel, its singular values and eigenvalues are the same and the definition can be based on either of them equivalently. The presented definition (using singular values) is also valid for any general matrix, and not just for the Gaussian kernel.
42 24 the numerical rank converges to the algebraic rank ρ(g M ε ) = ρ 0 (G M ε ). For the rest of the chapter, unless mentioned otherwise, we consider the parameters ε and δ to be predetermined and constant, such that ε > 0 and 0 δ < 1. For clarity, we will refer to the numerical rank ρ δ (G M ε ) of the Gaussian kernel over the dataset M as the Gaussian numerical rank of the dataset M Ambient box-based bounds The relation between the numerical rank of G M ε and the observable ambient space R m of the manifold M, from which the dataset M was sampled, is shown in [BAC13]. This relation was expressed by an upper-bound on the numerical rank, which was expressed by the volume of a bounding box of the dataset in the ambient space. However, the geometry of the manifold M is ignored by this bound. In this chapter, we refine the bounds achieved in [BAC13] by considering a small set of boxes that cover the manifold and any dataset that is sampled from it. First, we reiterate the results from [BAC13], then, in Section 2.3, we use these results to prove the new manifold-related bound. Let Q R m be a box in the observable space, where q 1... q m R are the lengths of its sides (listed, without loss of generality, in a descending order). Thus, the volume of Q is m i=1 q i. Let X Q be a finite dataset that is contained within the box Q, and let G X ε be the Gaussian kernel over this dataset. Then, according to [BAC13], the numerical rank of G X ε is bounded from above by m ρ δ (G X ε ) { κq j + 1}, (2.2.1) where j=1 κ 2 π ε 1 ln(δ 1 ). (2.2.2) We examine now an arbitrary side length q j (j = 1,..., m) of the box Q. If q j < 1, then the j-th term of the product in Eq is κq κ j + 1 = 1 and the side length q j does not affect the bound in this equation. A side of Q whose length q j < 1 (j = 1,..., m), which is too short to affect the bound in κ Eq , is called a short side. A side whose length q j 1 does affect this κ bound is called a long side. We call Q a d-box if it has exactly d m long sides and its other m d sides are short sides. Since we assumed (without loss of generality) that the side lengths of Q are listed in descending order, then for a d-box we have q 1... q d 1/κ > q d+1... q m. Since, in this case, q d+1,..., q m are short-side lengths that do not affect the bound in Eq , then for d-box Q (and any finite dataset X Q) the following
43 25 bound is satisfied: d ρ δ (G X ε ) { κq j + 1}. (2.2.3) j=1 2.3 Cover-based bounds The bound in Eq is based on a single box that covers the whole dataset (or the whole manifold). The volume (or, more accurately, the product of the discretized side lengths) of this box determines the value of this upper bound. If the dataset is sampled from a flat manifold (e.g., a hyperplane), the long sides of the bounding box can be set on the principal direction of this manifold while the remaining short sides on other directions (see Fig (a)). In this case, the bound in Eq considers the intrinsic geometry of the data and measures the volume on the approximately linear area of the manifold from which the data is sampled. However, when the manifold is not flat and contains curved areas, a single box, which contains the whole dataset, is expected to be unnecessarily large (see Fig (b)). (a) A single bounding box is sufficient for a relatively flat manifold. (b) For a non-flat manifold, a single bounding box is unnecessarily large. Figure 2.3.1: Covering a manifold (or a compact set) using a set of small (e.g., unit-size) boxes vs. a single box with discretized (e.g., integer) side-lengths. Instead of covering the whole dataset (or its underlying manifold) with a single large box, we use a set of small boxes to obtain a cover. Since each box covers a small area on the manifold, and due to the locally low dimensional
44 26 nature of the manifolds, each box is expected to have a small number d m of long sides. It is convenient to have all the boxes of approximately the same size by setting a constant length l to their long sides. This way, the size of the cover can be easily determined by l, d and the number of boxes in the cover. Definition introduces the type of boxes that will be used to cover a manifold or a dataset that is sampled from it. Definition ((l, d)-box). Let l 1 (where κ is defined in Eq ) be κ a real number and 1 d m be a positive integer. An (l, d)-box in R m is a d-box whose length of each of its d long sides is l. The boxes from Definition are the building blocks for the cover that will be used to set a bound on the numerical rank of Gaussian kernels of manifolds and datasets. Definition presents this cover for any subset in the ambient space R m. In particular, it defines the cover for a manifold that lies in this ambient space and for any dataset that is sampled from such a manifold. Definition ((l, d)-cover). Let C be a finite set of (l, d)-boxes in R m, l 1, and a positive integer 1 d m. Denote the number of boxes in it κ by #(C). The set C is called a (l, d)-cover of an arbitrary set X R m if for every data point x X there is at least one (l, d)-box Q C such that x Q. The size of a (l, d)-cover C is the number #(C) of boxes in C. We will use the notation C (l,d) (X ) for the set of all (l, d)-covers, l 1 κ and a positive integer 1 d m, of a subset X R m of the ambient space. Specifically, C (l,d) (M) is the set of all (l, d)-covers of a manifold M that lies in this ambient space, and C (l,d) (M) is the set of all (l, d)-covers of the dataset M that is sampled from this manifold. When the exact values of l and d are irrelevant, we will use the term box-cover for a (l, d)-cover with arbitrary values of the length l 1 and the integer 1 d m. The exact κ values of l and d will be referred to as the scale of the box-cover. The sets of all box-covers of X, M and M will be denoted by C(X ), C(M) and C(M), respectively. Definition specifies the conditions that define a box-cover of a set in the ambient space. Not all the sets have a box-cover since by definition only a finite number of boxes can be used in it. In this chapter, we are only interested in their existence for compact manifolds and finite datasets that are sampled from such manifolds. The existence of box-covers for finite datasets is immediate. Proposition shows the existence of box-covers (for every scale) for a compact manifold in R m.
45 27 (a) Low curvatures (b) High curvatures Figure 2.3.2: An illustration of the flexibility of (l, d)-covers: (a) in lowcurvature areas, the long sides can be set on tangent directions; (b) in highcurvature areas, they can be set on normal (i.e., orthogonal to the tangent) directions. Proposition Let M R m be a compact manifold in the ambient space. Then, C(M), and for every length l 1 and every integer κ 1 d m, C (l,d) (M). Proof. Consider an arbitrary scale (l, d). Surely, we can construct an open (l, d)-box around every data point x M on the manifold. This infinite set of open boxes covers the entire manifold. Since the manifold is compact, there must be a finite subset of these boxes that is sufficient for covering the entire manifold. This set constitutes an (l, d)-cover of M. This argument is valid for every scale that proves the proposition. Let C C(M) be a box-cover of the manifold M. Notice that there are no limitations or conditions set on the orientations and positions of the boxes in C. In low-curvature areas, it seems beneficial to set the long sides of the covering boxes to be tangent to the manifold (see Fig (a)). However, in high-curvature areas, it might be more efficient (depending on the scale of the box-cover) to set the long sides along the normal of the tangent space (see Fig (b)). The definition of the box-covers allows us to use this flexibility to consider efficient coverings of the manifold. Theorem introduces an
46 28 upper bound on the numerical rank of Gaussian kernels on datasets that are sampled from the manifold. Corollary extends this result to set a mutual upper bound on any dataset that is sampled from the given manifold. Theorem Let M R m be a manifold in the ambient space R m and let M M be a dataset sampled from this manifold. The numerical rank of the Gaussian kernel G M ε over the dataset M is bound by ρ δ (G M ε ) r(m), where r(m) min { #(C) h(l, d) l 1/κ, 1 d m, C C (l,d) (M) }, (2.3.1) and h(l, d) = d j=1 { κl + 1}. Theorem shows that any box-cover of a dataset provides an upper bound on the numerical rank of the Gaussian kernel over this dataset. We use the term cover-based bound for the upper bounds in the set {#(C) h(l, d) l 1/κ, 1 d m, C C (l,d) (M)} from Theorem We call their minimum r(m) as the tightest cover-based bound of the dataset M. The proof of Theorem is based on the subadditivity of the Gaussian numerical rank, which is shown in Section Proposition shows that box-covers exist for any compact manifold. Thus, it is reasonable to consider cover-based bounds that are set by the underlying manifold for any dataset that is sampled from it. Such bounds are not dependent on any specific sampling, but rather on the geometry of the analyzed phenomena. Corollary extends the result of Theorem and introduces the cover-based bounds, as well as the tightest cover-based bound of any compact manifold. Corollary Let M R m be a manifold in the ambient space R m. The numerical rank of the Gaussian kernel G M ε over any sampled dataset M M is bounded by ρ δ (G M ε ) r(m), where r(m) min { #(C) h(l, d) l 1/κ, 1 d m, C C (l,d) (M) }, and h(l, d) = d j=1 { κl + 1}. Proof. The existence of box-covers for the manifold M is established by Proposition and by the definition #(C) h(l, d) for any l 1/κ, 1 d m and C C (l,d) (M), is a positive integer. Therefore, the minimum r(m) exists and is well defined. Every box-cover of the manifold is also a box-cover of any dataset M M that is sampled from the manifold. Thus, the set of bounds in the corollary is a subset of the set in Theorem and thus ρ δ (G M ε ) r(m) r(m).
47 We call the bound r(m) in Corollary the tightest cover-based bound of the manifold M, and any bound in the set {#(C) h(l, d) l 1/κ, 1 d m, C C (l,d) (M)} is called a cover-based bound of the manifold. Proposition shows that the tightest cover-based bound of the manifold is indeed the tightest cover-based bound of some large enough dataset that is sampled from it. Therefore, no tighter cover-based bound can be set for every possible dataset that are sampled from M. Proposition Let M R m be a manifold in the ambient space R m. The tightest cover-based bound of the manifold satisfies r(m) max{r(m) M M, M < }. Proof. Surely, every box-cover of the manifold M also covers any subset of the manifold. Specifically, it is true for every finite dataset that is sampled from it. Therefore, we must have r(m) r(m) for every dataset M M and the weak inequality in the proposition is proved. The existence of the maximum is due to the discreteness of the tightest cover-based bounds. Proposition justifies the name tightest cover-based bound that we used for r(m) by showing that it indeed serves as a maximal tightest cover-based bound for all the finite sampled dataset from the manifold. Section provides examples for equality and strict inequality cases. In addition to examining finite datasets and defining Gaussian kernel matrices over them, we can also define a continuous Gaussian kernel operator G M ε : C(M) C(M) over the whole manifold M. This operator is defined by G M ε f(x) = g ε (x, y)f(y)dµ(y), f : M R, x M (2.3.2) M and it represents the affinities between all the data points on the manifold. Due to the compactness of M and the continuity of g ε, then according to the Hilbert-Schmidt theorem, the Gaussian kernel operator G M ε has a discrete set of real eigenvalues that forms a decaying spectrum [CL06a, CL06b], which is similar to the spectrum of Gaussian kernel matrices over datasets that are sampled from the manifold. Therefore, we can also examine the numerical rank of this operator that considers the manifold itself instead of considering a finite sampling of data points from it. Theorem shows that the tightest cover-based bound r(m) also serves as an upper bound for the Gaussian numerical rank ρ δ (G M ε ) of the manifold M and not only as an upper bound on Gaussian numerical ranks of finite datasets that are sampled from it. 29
48 30 Theorem Let M R m be a compact manifold in the ambient space R m. The numerical rank of the Gaussian kernel operator G M ε over the manifold M is bounded by ρ δ (G M ε ) r(m), where r(m) is the tightest cover-based bound (from Corollary 2.3.3) of the manifold M. Theorem 2.3.5, which will be proved in Section 2.3.2, shows that the achieved upper bound of the Gaussian numerical rank is a property of the embedded manifold itself and not just a result of finite samplings of the manifold. In some sense, it also provides an insight for the finite datasets usage to represent properties of the manifold. Together with Proposition 2.3.4, it shows a relation between the maximal tightest cover-based bound that is achieved by a finite dataset and the upper bound on the Gaussian numerical rank of the continuous manifold itself. Some implications and nuances of the results in this section are demonstrated on simple manifolds (i.e., curves and surfaces) in Section 2.4. The rest of this section deals first with proving the two main theorems. In Section 2.3.1, we prove Theorem by showing that the Gaussian numerical rank is subadditive. In Section 2.3.2, we prove Theorem by showing a series of finite matrices whose numerical ranks converge to the numerical rank of the continuous operator in Eq Subadditivity of the Gaussian numerical rank Theorem is a result of the subadditivity of the numerical rank of Gaussian kernels. In this section, we will prove this property and then prove the theorem by using this result. Lemma shows the relation between the numerical rank of Gaussian kernels of two sets and the numerical rank of their Gaussian kernel union. In order to prove it, we first show a technical result in Lemma 2.3.6, about the relations between the numerical rank and the algebraic rank of principal submatrices. Lemma Let G C n n be a nonsingular complex matrix and let G C q q (q < n) be a principal submatrix of G. If ρ δ (G) = ρ(g) then ρ δ ( G) = ρ( G). Proof. If ρ δ (G) = ρ(g) then, by the definition of the numerical rank, σn(g) σ 1 (G) δ. From Cauchy s interlacing theorem [SS90] we get σ q ( G) σ n (G) and σ 1 ( G) σ 1 (G), thus, σq( G) σ 1 ( G) rank again, we finally get ρ δ ( G) = ρ( G). δ. By using the definition of the numerical Lemma Let X = {x 1, x 2,..., x p 1, x p } and Y = {y 1, y 2,..., y q 1, y q } be two sets in R m. Then, for any ε > 0 and 0 δ < 1 ρ δ (Gε X Y ) ρ δ (G X ε ) + ρ δ (G Y ε ).
49 Proof. Suppose that ρ δ (Gε X Y ) = r and let Z X Y be a subset of r data points such that ρ δ (G Z ε ) = r. Additionally, let X = X Z and Ỹ = Y Z. According to Bochner s theorem [Wen05], ρ(g Z ε ) = r. Thus we get, r = ρ(g Z ε ) X + Ỹ. According to Lemma 2.3.6, ρ δ(g X ε ) = X and ρ δ (GỸε ) = Ỹ. Since X X and Ỹ Y, ρ δ(g X ε ) ρ δ (G X ε ) and ρ δ (GỸε ) ρ δ (G Y ε ). As a consequence, ρ δ (G X Y ε ) ρ δ (G X ε ) + ρ δ (G Y ε ). Lemma shows that the Gaussian numerical rank of a union of two sets is at most the sum of their Gaussian numerical ranks. This result can be easily extended to unions of any number of sets by applying Lemma as many times as needed. Therefore, we get Corollary that states the subadditivity of the Gaussian numerical rank. Corollary (Subadditivity of the Gaussian numerical rank). Let X 1, X 2,..., X q be q finite subsets of R m, and let M = q j=1 X j. Then, ρ δ (G M ε ) q j=1 ρ δ(g X j ε ). We are now ready to prove Theorem by combining Corollary and the results from [BAC13]. In essence, each box provides an upper bound on the Gaussian numerical rank of a local subset according to the result from [BAC13], and these bounds can be combined according to Corollary 2.3.8, thus achieving an upper bound on the Gaussian numerical rank of the whole dataset. Proof of Theorem Let M M be a finite dataset that is sampled from the compact manifold M R m. Since the dataset is finite, there exists a box-cover C C (l,d) (M) for some l 1/κ and 1 d m. By Definition 2.3.2, the cover-based bound #(C) h(l, d) for any such box-cover is a positive integer. Therefore, the minimum r(m) min { #(C) h(l, d) l 1/κ, 1 d m, C C (l,d) (M) } (2.3.3) of a nonempty set of positive integers exists and it is well defined. By definition 2.3.2, any arbitrary (l, d)-cover C of M (for appropriate values of l and d) is a set of (l, d)-boxes Q 1,..., Q q, q = #(C), such that M Q 1 Q q. Therefore, we can define the q sets M j Q j M, j = 1,..., q, and get that M = M 1 M q where each set M j, j = 1,..., q, is bounded by the corresponding (l, d)-box. Each of these boxes is a d-box where all its long-sides have the length l. Therefore, according to Eq , the Gaussian numerical rank of every M j, j = 1,..., q, is bounded by ρ δ (G M j ε ) d { κl + 1} = ( κl + 1) d = h(l, d), i=1 31
50 32 thus, together with Corollary we get that the Gaussian numerical rank of M = M 1 M q is bounded by ρ δ (G M ε ) q j=1 #(C) ρ δ (G M j ε ) h(l, d) = #(C) h(l, d). j=1 Therefore, each arbitrary (l, d)-cover C C (l,d) (M) provides a cover-based upper-bound #(C) h(l, d) on the Gaussian numerical rank of M. In particular, the tightest (i.e., minimum) cover-based bound r(m) (see Eq ) is indeed an upper-bound for this numerical rank as the theorem states. Notice that, in fact, the proof of Lemma does not rely on any specific inter-subset affinities values of g ε (x, y), x X Y, y Y X, in the context of Lemma As a result, both Lemma and Corollary 2.3.8, also apply when these inter-subset affinities are not directly determined. Therefore, one can measure the affinities in local areas on the manifolds, and either ignore (i.e., set to zero) or deduce (e.g., by random-walks or diffusion) the affinities between farther data points. The resulting kernel will still abide by Lemma and Corollary If the local neighborhoods, in which the affinities are directly measured, are determined by a box-cover that achieves the tightest cover-based bound r(m), then, according the proof of Theorem 2.3.2, the numerical rank of the resulting locally-measured kernel will not exceed the bound r(m) of the Gaussian numerical rank The Gaussian convolution operator In this section, we focus on the continuous kernel operator G M ε from Eq This kernel is in fact a Gaussian convolution operator that acts on the manifold M in R m. The main goal of this section is to bound its numerical rank (from above) and prove Theorem The notations and the techniques in the rest of this section are similar to the ones that were presented in [vlbb08]. These notations are slightly different from the rest of this chapter, but they are more suitable for the purposes of the following discussion. Let M be the manifold defined in Section 2.2. Assume, without loss of generality, that M dµ(x) = 1. Let X = {x i} i N be a discrete set of data points that are drawn independently from M according to the probability distribution µ. Let X n = {x i } n i=1 be the subset consisting of the first n data points in X. We define the empirical measure µ n (M) = 1 n n i=1 δ x i, where δ x is the Dirac delta function centered at x X. Thus, for any function f : M R we have M f(x)dµ n(x) = 1 n n i=1 f(x i).
51 Let (C(M), ) be the Banach space of all real continuous functions defined on M with the infinity norm, and B is the unit ball in this space. Let g ε : M M R be the Gaussian affinity g ε (x, y) = exp{ x y 2 /ε}, where denotes the Euclidean norm in R m. Define the integral operator G ε : C(M) C(M) to be the convolution operator G ε f(x) = g M ε(x, y)f(y)dµ(y). According to Hilbert-Schmidt theorem G ε, as an operator from L 2 (M, µ) to itself, is a compact operator. Additionally, G ε is positive-definite, due to Bochner s theorem. Therefore, the spectrum of G ε consists of isolated eigenvalues. For brevity, since ε is constant throughout this section, we omit the ε subscript. We will call the operator G the full convolution operator, as opposed to the partial convolution operators that will be defined later in this section. Notice that the defined operator G is the same as the operator G M ε from Eq We denote the spectrum of the operator G by σ(g). For every positive integer n N we define a n n matrix Ḡ n 1, n GXn where G Xn is the Gaussian kernel matrix over the dataset X n. We also define for n N the partial convolution operator G n : C(M) C(M) that computes the convolution over the data points in X n instead of computing it over the whole manifold as done for G n f(x) g M ε(x, y)f(y)dµ n (y). Finally, we define the restriction operator R n : C(M) R n to be R n (f) (f(x 1 ), f(x 2 ),..., f(x n )) T. We will use these constructions to show the relation between the Gaussian numerical rank of a manifold and the Gaussian numerical rank of finite datasets that are sampled from it. Proposition shows the relations between the defined constructions. 33 Proposition The operators G and G n, n N, are compact, uniformly bounded in (C(M), ), and Ḡ n R n = R n G n. Proof. Since the dimension of the range of G n is finite then G n is compact for any n N. In order to prove that G is compact, we will prove that for any sequence of functions {f n } n N B, the sequence {Gf n } n N is relatively compact. Due to Arzela-Ascoli Theorem (e.g., Section I.6 in [RS80]), it suffices to prove that the set {Gf n } n N is pointwise bound and equicontinuous. Since g ε = 1, f n = 1 and µ(m) = 1, we get Gf n = g ε (x, y)f n (y)dµ(y) 1, namely the set {Gf n } n N is pointwise bound. M
52 34 In addition, Gf n (x) Gf n (x ) = M (g ε (x, y) g ε (x, y))f n (y)dµ(y) g ε (x, ) g ε (x, ) 2 eε x x. This proves the equicontinuity of {Gf n }, which completes the proof of compactness of G. It remains to show that G and G n, n N are uniformly bounded. G n = sup G n f f B 1 n = sup g ε (x, x i )f(x i ) n 1. f B,x M Due to the first part of the proof, G 1. Therefore, G and G n, n N, are uniformly bounded by 1. The last part of the Lemma is a direct result from the definitions of Ḡ n, R n and G n. The definition of the numerical rank of a compact self-adjoint operator G is identical to the definition of the numerical rank on matrices (see Definition 2.2.1), where instead of singular values we use eigenvalues 2. For diagonalizable operators, and specifically for compact self-adjoint operators, the numerical rank is the dimensionality of the significant eigen-subspaces, namely, the subspaces that correspond to the significant eigenvalues. Therefore, Definition is an equivalent definition of the numerical rank definition of a compact operator G. We use the term Gaussian numerical rank of a manifold M to denote the numerical rank of the Gaussian convolution operator that acts on that manifold. Definition Let G be a compact operator in a Banach space. The numerical rank of G up to precision δ 0 is ρ δ (G) dim(proj λ G), (2.3.4) λ δλ max 2 The singular values and eigenvalues of Gaussian kernel matrices are anyway equal. Therefore, the results achieved for them in this chapter are also valid when using this eigenvalue-based definition. i=1
53 where λ max is the largest eigenvalue of G, proj λ G is the projection operator on the eigenspace corresponding to λ, and dim(proj λ G) is the dimension of this eigenspace. 35 Our goal is to prove that the Gaussian numerical rank ρ δ (G) of a manifold M is bounded by ρ δ (G) r(m). For this purpose, we take a linear-operator approximation approach. First, in Section 2.3.2, we prove that ρ δ (G n ) = ρ δ (Ḡ n ) for any n N. Therefore, due to Proposition 2.3.4, the numerical rank of each partial convolution operator is bounded by ρ δ (G n ) r(m). Then, in Section 2.3.2, we show that the full convolution operator G is the limit operator of the partial convolution operators {G n } n N and as a consequence ρ δ (G n ) ρ δ (G), which completes the proof. The numerical rank of G n, n N Due to Bochner s theorem, the matrix Ḡ n is strictly positive definite, hence all its eigenvalues are positive. Lemma shows that Ḡ n and G n have the same nonzero eigenvalues with the same geometric multiplicities. Lemma The following relations between the eigen-systems of the matrix Ḡ n and the partial convolution operator G n are satisfied: 1. Let v = (v 1, v 2,..., v n ) t be an eigenvector of Ḡ n that corresponds to an eigenvalue λ. Then, the continuous function f v : M R, defined by n f v (x) = 1 k(x, x nλ j )v j is an eigenfunction of G n, corresponding to j=1 the same eigenvalue λ. 2. If f is an eigenfunction of G n that corresponds to an eigenvalue λ then R n f is an eigenvector of g n that corresponds to the same eigenvalue λ. 3. Let λ be an eigenvalue of Ḡ n with the geometric multiplicity m. Then, the geometric multiplicity of λ as an eigenvalue of G n is m. 4. ρ δ (G n ) = ρ δ (Ḡ n ) for any n N. Proof. 1. Since Ḡ n v = λv, then λf v (x i ) = 1 n n g ε (x i, x j )v j = λv i for all j=1
54 36 i = 1, 2,..., n. Therefore, G n f v (x) = 1 n = 1 n = 1 n n g ε (x, x i )f v (x i ) i=1 n 1 n [g ε (x, x i ) g ε (x i, x j )v j ] nλ j=1 n g ε (x, x i )v i = λf v (x). i=1 i=1 2. If G n f = λf then, due to Proposition 2.3.9, Ḡ n R n f = R n G n f = λr n f. 3. Let v 1,..., v m be a basis for the eigenspace of Ḡ n that corresponds to the eigenvalue λ. Since v 1,..., v m are linearly independent, then the functions f v1,..., f vn are linearly independent. Therefore, dim(proj λ G n ) dim(proj λ Ḡ n ). Since the ranges of G n and Ḡ n are both of dimension n, we get dim(proj λ G n ) = dim(proj λ Ḡ n ) for any nonzero eigenvalue λ. 4. The equality ρ δ (G n ) = ρ δ (Ḡ n ) is a direct consequence of the above. Corollary is an immediate result of Theorem and Lemma This proposition provides an upper bound for the numerical rank of the partial convolution operators G n, n N. This bound will be used in Section to provide an upper bound for the numerical rank of the full convolution operator G. Corollary The numerical rank of G n, for any n N, is bounded by ρ δ (G n ) r(m). The numerical rank of G In this section, we prove that the sequence {G n } n N converges to G compactly as defined in Definition Proposition shows that this convergence also guarantees the convergence of the corresponding eigenspaces of the sequence {G n } n N to those of G. Definition (Convergence of operators). Let (F, F ) be a Banach space, B its unit ball and {S n } n N is a sequence of bounded linear operators on F: The set {S n } n N converges pointwise, denoted by S n Sf F 0 for all f F. p S, if Sn f
55 c p The set {S n } n N converges compactly, denoted by S n S, if S n S and if for every sequence {f n } n N in B, the sequence {(S S n )f n } n N is relatively compact (has a compact closure) in (F, F ). Proposition (Proposition 6 in [vlbb08]). Let (F, F ) be a Banach c space, and {S n } n N and S are bounded linear operators on F such that S n S. Let λ σ(s) be an isolated eigenvalue with finite multiplicity m, and M C an open neighborhood of λ such that σ(s) M = {λ}. Then: 1. Convergence of eigenvalues: There exists an N N such that, for all n > N the set σ(s n ) M is an isolated part of σ(s n ) that consists of at most m different eigenvalues, and their multiplicities sum up to m. Moreover, the sequence of the sets σ(s n ) M converges to the set {λ} in the sense that every sequence {λ n } n N with λ n σ(s n ) M satisfies lim n λ n = λ. 2. Convergence of spectral projections: Let P r be the spectral projection of S that corresponds to λ, and for n > N, let P r n be the spectral p projection of S n that corresponds to σ(s n ) M. Then, P r n P r. 37 Lemma The full and partial convolution operators satisfy G n in (C(M), ). p G Proof. Let f C(M) then G n f Gf = sup x M M 1 = sup n x M g ε (x, y)f(y)dµ n (y) g ε (x, y)f(y)dµ(y) M n g ε (x, x i )f(x i ) E(g ε (x, )f( )), i=1 where E(g ε (x, )f( )) is the expected value of g ε (x, y)f(y) as a function of y for a fixed x. As n, this expression converges to zero due to the uniform law of large numbers, and therefore the convergence in the Lemma is proved. Lemma The partial and the full convolution operators satisfy G n G in (C(M), ). p Proof. Due to Lemma , we already have G n G. It remains to show that for every sequence {f n } n N in the unit ball B in C(M), the sequence {(G G n )f n } n N is relatively compact in (C(M), ). Due to Arzela-Ascoli c
56 38 Theorem, it is suffices to show that {(G G n )f n } n N is pointwise bounded and equicontinuous. As for the first property, according to the proof of Proposition 2.3.9, (G G n )f n Gf n + G n f n 2. The second property is a result of the bounded derivative of the Gaussian function: (G G n )f n (x) (G G n )f n (x ) G(f n (x) f n (x )) + G n (f n (x) f n (x )) = (g ε (x, y) g ε (x, y))f n (y)dµ(y) M 1 n + (g ε (x, x i ) g ε (x, x i ))f n (x i ) n i=1 2 max g ε(x, y) g ε (x, y) y M 2 2 εe x x. Proposition shows that the full convolution operator G is compact. This operator is also strictly positive definite due to Bochner s theorem. Therefore, all the eigenvalues of this operator are positive and isolated. Theorem shows the relation between the numerical rank of G and the numerical ranks of the partial convolution operators G n, n N. This theorem is a immediate result of Corollary , Proposition and Lemma Theorem The operators G n, n N, and G satisfy lim ρ δ(g n ) = ρ δ (G). n Theorem essentially states that ρ δ (G) r(m), which we proceed to prove in this section, is also a direct result of this discussion, and can be considered as a corollary of Theorem Therefore, the tightest coverbased bound of the manifold bounds the numerical rank of the affinity kernel operator that considers all the data points on the manifold. This property of the tightest cover-based bound shows that it can be regarded as a property of the embedded manifold, and not just a bound for the purpose of analyzing sampled datasets.
57 Examples and discussion Strict inequality and equality in Proposition Example 1: the unit-diameter circle curve (strict inequality) x y } ξ short-side { }} { }{{} long-side Figure 2.4.1: Due to the curvature of the unit-circle, for two adjacent data points x and y in a finite dataset (sampled from the unit-diameter circle) there is a ξ-wide band that is not necessary when only covering the dataset, since there are no data points on the arc between them. This band is necessary when the entire (continuous) unit-diameter circle is covered. Let the parameters δ and ε have values such that κ = 1, and consider a circle M (as a plain curve) with unit-diameter in R 2. The (l, d)-covers have two parameters (l and d) that need to be considered. In this case, there are two possible values for d: If d = 1, then each box in the cover has one side (i.e., the long side) of length l 1, and the other side (i.e., the short side) is of length 1 ξ (for an arbitrarily small 0 < ξ < 1) since it has to be strictly less than one. In any case, an (l, 1)-cover of M must consist of at least two (l, 1)- boxes, since the short side of a single box is shorter than the diameter of the circle (see Fig ). The resulting bound (from Theorem 2.3.2) in this case is 2 ( 1 l +1) 4. On the other hand, for any finite dataset M M, we can select two adjacent data points x, y M and set the
58 40 long-side of the box to be parallel to the straight line between x and y as illustrated in Fig We can assume, without loss of generality, that l = 1 and that ξ is small enough for this single (1, 1)-box to cover of M, and thus the bound in this case is 1 ( ) = 2. If d = 2, then clearly we can use a single (1, 2)-box to form a (1, 2)-cover of both M and M, thus the resulting bound is 1 ( ) 2 = 4. Any larger value of l will achieve the same (or larger) bound. As a consequence of the above, we get r(m) = 2 < 4 = r(m) for any finite dataset M M. Example 2: a two-dimensional unit square (equality) Let M be the unit square curve 3 in R 2. We use the same parameters δ and ε as in Example 1 such that κ = 1. Using arguments similar to the ones in the previous example, we need at least two (l, 1)-boxes to cover M, or exactly one (l, 2)-box, for any l 1. Both resulting bounds are again at least four, so in this case r(m) = 4. Let M M be the dataset that contains the four corners of the square. This dataset cannot be covered by a single (l, 1)-box, since its short side must be shorter than one, and therefore the (l, 2)-covers are anyway similar for the dataset and the manifold in this case, therefore, we can use the same arguments that we used for M and get r(m) = 4 = r(m) Cover-based bounds of plain curves In this section, we examine the curves (i.e., one-dimensional manifolds) in a two-dimensional ambient plane R 2. We apply the cover-based methodology to introduce the relation between the Gaussian numerical rank of a curve (or datasets sampled from it) and its geodesic arc-length. Specifically, we show that the Gaussian numerical rank of datasets that are sampled from a finite-length curve is bounded by a function of its length. Proposition 2.4.1, which is illustrated in Fig (a), presents a relation between the geodesic length of a curvature-bounded curve section γ and the dimensions of a tangent bounding box of that section. The presented relation provides a method to determine the size of the local boxes that can be used to construct a box-cover of the entire curve. 3 The manifold in this example is not differentiable at the four corners of the square, but the corners of the square can be slightly rounded by conformal mapping to become smooth in a way that preserves the validity of the presented results.
59 41 γ = l t } {{ } l (a) The relation from Proposition between the arc-length l and the bounding l t box. T } {{ } L γ j = L (b) A single local bounding box (of the curve section γ j, j = 1,..., k) from the box-cover in Corollary Figure 2.4.2: Illustrations of the relations that are used to provide the lengthbased bound of the Gaussian numerical rank of plain curves. Proposition Let γ R 2 be a smooth plain curve and let t and r be positive constants such that t r. Let γ be a section of γ with arc-length ( γ = l = r arccos 1 t ). (2.4.1) r Let γ(s), 0 s l, be an arc-length parametrization of γ and assume that the curvature c(s) is bounded from above by 1 r. Then, the section γ R2 can be bounded in a two-dimensional box whose dimensions are l t. Proof. Suppose that γ : R R 2 is parameterized by arc length such that γ(s) = γ(s) for 0 s l. Let {e 1, e 2 } be the standard coordinates system for R 2 such that γ(0) = 0 and the derivative γ (0) = e 1. Let γ(s) = (x(s), y(s)) be the parametrization of γ in these coordinates, i.e., x(s) and y(s) are the orthogonal projections of γ(s) on e 1 and e 2, respectively. Let θ : [0, l] [0, 2π), θ(s) = arctan ( y (s) x (s) ) be the angle that γ (s) makes with e 1. Thus, (see [dc76]), θ (s) = c(s) and y (s) = sin(θ(s)) or, equivalently, y(s) = s sin(θ(s))ds and θ(s) = s c(z)dz s for any 0 s l. Thus, due to 0 0 r Eq , we get l l ( s ) sin ds r y(l) = sin(θ(s))ds = 0 ( ) l r r cos = t r Obviously, x(l) l, therefore, γ can be bounded in an l t box. Corollary 2.4.2, which is illustrated in Fig (b), uses Proposition to provide a relation between the geodesic length of a finite length curve and its Gaussian numerical rank. Specifically, it shows that this Gaussian numerical rank is bounded in proportion to the arc-length of the curve. 0
60 42 Corollary Let t and r satisfy the conditions of Proposition 2.4.1, such that t 1 and let 1 L = 2r arccos ( 1 t 2κ κ r). Assume that γ is a plain curve of finite length γ whose curvature is bounded from above by 1. Then, r for any finite configuration X γ, ρ δ (G X ε ) h(l, 1). Proof. Divide γ to k = γ L γ L sub-curves such that γ = k j=1 γ j where each is of length L except, perhaps, γ k. Let T = 2t. For each sub curve γ j, construct an L T bounding box B j, whose center c j is the midpoint of γ j, such that its long side is parallel to γ (c j ). This construction is possible due to Proposition since t 1 and L 1, k 2κ κ j=1 B j constitutes an (l, 1)-cover of X. Therefore, according to Theorem 2.3.2, for any finite configuration X γ, ρ δ (G X ε ) h(l, 1). γ L It should be noted that extending these results to volumes of higher dimensional manifolds (e.g., geodesic areas of surfaces) is not trivial. This type of analysis depends on the exact volume form of the manifold and is beyond the scope of this chapter. However, in practical cases, manifold characterizations in general, and its volume form specifically, are anyway not known. From a practical data point of view, the box-covers used in this chapter provide a sufficient volume metric that incorporates the low-dimensional locality nature of the manifold together with possible high-curvature singularities and noisy sampling techniques Discussion In many cases, although not in all of them, the subadditivity of the Gaussian numerical rank, which is presented in Proposition 2.3.8, enables to provide a much tighter bound than the one presented in [BAC13]. This bound considers the intrinsic dimensionality of the data, rather than its extrinsic dimensionality. For example, consider a dataset that was sampled from a one-dimensional square-shaped manifold, whose side-length is q, embedded in the real plane. Then, the bound on the Gaussian numerical rank provided by [BAC13] is, due to Eq , quadratic in q (i.e., ( κq + 1) 2 ). On the other hand, by covering the data with four (q, 1)-boxes, a linear bound is provided by Proposition (i.e., 4( κq + 1)). This bound is tighter than the quadratic one for sufficiently large q (i.e., q > 4/κ). In any case, the definition of the proposed bound r(m) (Eq ) considers all the (l, d)-covers of the data, including single-box covers. As such, this bound is at least as tight as the bound presented in [BAC13].
61 Conclusion In this chapter we presented a relation between the numerical rank of Gaussian affinity kernels of low-dimensional manifolds (and datasets that are sampled from them) and the local-geometry of these manifolds. Specifically, we introduced an upper-bound for this numerical rank based on the properties of a box-cover of the manifold. The used cover is based on a set small boxes that contain local areas of the manifold. Together, this set of boxes incorporates the non-linear nature of the manifold while coping with varying curvatures and possible sampling noise. The presented relation validates one of the fundamental assumptions in kernel-based manifold learning techniques that local low-dimensionality of the underlying geometry yields a low numerical rank of the used affinities, thus, spectral analysis of these affinities provides a dimensionality reduction of the analyzed data. The results in this chapter support this assumption by showing that, in the Gaussian affinity case, its numerical rank is indeed bounded by properties of the underlying manifold geometry.
62 44
63 Chapter 3 Patch-to-tensor embedding In this chapter, we extend the scalar relations that are used in manifold-based kernel methods, such as DM, to matrix relations that encompass multidimensional similarities between local neighborhoods of points on the manifold. We utilize the diffusion maps methodology together with linear-projection operators between tangent spaces of the manifold to construct a super-kernel that represents these relations. The properties of the presented super-kernels are explored and their spectral decompositions are utilized to embed the patches of the manifold into a tensor space in which the relations between them are revealed. We present two applications that utilize the patch-to-tensor embedding framework: data classification and data clustering. The results in this chapter appear in [SWA12, SWAN12]. 3.1 Introduction High-dimensional datasets have become increasingly common in many areas due to high availability of data and continuous technological advances. For many such datasets, curse of dimensionality problems cause learning methods to fail when they are applied to the raw (high-dimensional) data points. Recent machine learning methods in general, and kernel methods in particular, seek to obtain and analyze simplified representations of these datasets. Such methods typically assume that the observable parameters in these datasets are related to a small set of underlying factors via nonlinear mappings. Mathematically, this assumption is often characterized by a manifold structure on which data points are assumed to lie. This underlying manifold is immersed (or submersed) in an ambient space that is defined by the observable parameters of the data. Usually, the intrinsic dimension of the underlying manifold is significantly smaller than the dimension of the 45
64 46 ambient space. A recent work [SW11] suggests to enrich the information represented by a simplified version of the kernel used in the Diffusion Maps method. The original kernel expresses the notion of proximity or the neighborhood structure of the manifold. The enriched kernel also maintains the information about the orientation of the coordinate systems in each neighborhood. This information allows the resulting eigenmap (i.e., the map constructed by the eigenvalues and eigenvectors of the kernel) to be used for determining the orientability of the underlying manifold. In cases when the manifold is orientable, this method finds a suitable global orientation together with the global coordinate system of the embedded space. If the manifold is not orientable, a modification of the used kernel can be utilized to find a double-cover of this manifold. In this chapter, which is based on [SWA12], we extend the original Diffusion Maps method in particular and kernel methods in general by suggesting the concept of a super-kernel. We aim at analyzing patches of the manifold instead of analyzing single points on the manifold. Each patch is defined as a local neighborhood of a point in a dataset sampled from an underlying manifold. The relation between two patches is described by a matrix rather than by a scalar value. This matrix represents both the affinity between the points at the centers of these patches and the similarity between their local coordinate systems. The constructed matrices between all patches are then combined in a block matrix, which we call a super-kernel. We suggest a few methods for constructing super-kernels. In particular, linear-projection operators between tangent spaces of data points are suggested for expressing the similarities between the local coordinate systems of their patches. Similar relations were also used in [LHZJ13] to present an ismoretric embedding that preserves the intrinsic metric structure of the manifold. There, it was shown that by constructing an embedding that preserves (or approximates) parallel vector fields, the geodesic structure of the manifold can be isometrically approximated by Euclidean distances in the embedded space. In our case, however, we do not embed single data points, or parallel vector fields that follow geodesics on the manifold, but we are interested in embedding patches of data points. Furthermore, the presented embedding is aimed to consider diffusion metrics and geometries on the manifold, which are different from the geodesic ones by considering all paths between analyzed patches (or data points) instead of just the shortest path. Therefore, we also suggest using the original diffusion kernel for expressing the affinities between points on the manifold. We examine and determine the bounds for the spectra (i.e., the eigenvalues) of the suggested constructions. Then, the eigenvalues and the eigenvectors of the constructed
65 super-kernels are used to embed the patches of the manifold into a tensor space. We relate the Frobenius distance metric between the coordinate matrices of the embedded tensors to a new distance metric between the patches in the original space. We show that this metric can be regarded as an extension of the diffusion distance metric, which is related to the original Diffusion Maps method [CL06a]. An alternative super-kernel construction was presented in [SW12], where parallel transport operators on the underlying manifold were utilized to define the similarities between the patches of the manifold. The resulting superkernel was utilized there to construct a Vector Diffusion Map (VDM) via spectral analysis. The continuous parallel transport operators were approximated there, in the finite case, by orthogonal transformations that achieve minimal Frobenius distances from the linear-projection operators that are used in this chapter. Algorithmically, this orthogonalization step seems like a small difference between projection-based super-kernels, which are presented here, and the ones presented in [SW12]. However, the theoretical implications of this additional step are significant. While the linear-projections incorporate the effects of the curvature of the manifold on the relations between patches in the super-kernel, these effects are canceled in the orthogonalization process, and only intrinsic quantities to the compared patches (i.e., not the general manifold) are preserved. The resulting VDM embedding shares many of the qualities of the original diffusion maps embedding [CL06a], when the scalar (i.e., 0-form) operators translated to 1-forms setting. Specifically, the infinitesimal generator of the VDM super-kernel converges to the connection- Laplacian, which is related to the heat kernel on 1-forms. One important quality of the VDM construction [SW12] is that it does not require an ambient space in which the underlying manifold lies. Therefore, it can be utilized to analyze general graphs. The linear-projection approach, on the other hand, relies on the existence of an ambient space. In practice, most analyzed datasets inherently define an ambient space by the measured features of the data,and thus its existence is well established. However, there are image-processing applications in which the VDM approach would be preferable, since the orthogonal transformations used there can be interpreted as isometries that achieve the best fitting between pairs of images. One such example that utilizes the VDM for the analysis of images in cryo-electron microscopy is presented in [SW12, SSHZ11]. In this example [SSHZ11], noisy two-dimensional EM snapshots of molecules were gathered from many unknown viewing angles, and the embedding performed by VDM was utilized to order the analyzed snapshots according to these angles. Once the viewing angles are known, a three-dimensional illustration of the analyzed molecule can be constructed from these 2D snapshots, but this step is not relevant to 47
66 48 the presented methods in here and in [SW12]. Another approach for applying spectral analysis of non-scalar affinities to data-analysis tasks is to consider non-pairwise relations between data-points. One example of this approach is shown in [ALZM + 05], where a hyper-graph was used to model the relations between data-points. Each hyper-edge in this hyper-graph represents a relation between an unordered set of datapoints, and is assigned a weight that quantifies this relation. By expanding the hyper-edges to cliques of related data-points, this hyper-graph can be reduced to a standard graph on which well-known partitioning algorithms can be performed to achieve clustering of the original data. A different example of the utilization of non-pairwise affinities for clustering is presented in [SZH06]. Instead of constructing a hyper-graph to represent the non-pairwise affinities, and then reducing it to a standard graph, an affinity tensor (i.e., a N-way array) is constructed and analyzed directly. This super-symmetric tensor replaces the standard affinity matrix that is usually used in kernel methods. The data clusters are achieved by probabilistic clustering that is performed on the constructed affinity tensor. The approach used in [ALZM + 05, SZH06] to extend kernel methods to use non-scalar affinities is significantly different from the one presented in this chapter and in [SW11, SW12]. First, this approach does not utilize the locally-linear structure of the underlying manifold when defining the relations between data-points. Secondly, the analyzed items in this approach are still individual data-points, even though the considered relations between them are more complex than in classical kernel methods. The patch-processing approach (used in here and in [SW11, SW12]), on the other hand, considered pairwise affinities between local patches on the manifold. While the complexity (i.e., non-scalarity) of the affinities in [ALZM + 05, SZH06] comes from the nature of the relations between individual data points, the complexity in our case comes from the analyzed items themselves, which are patches instead of data points. This property is best seen by considering the structure of the extended affinity kernel (or super-kernel), which is a block matrix in our case and a N-way array (i.e., not a matrix) in [SZH06]. The chapter has the following structure: The benefits of patch processing are discussed in Section Section 3.2 contains an overview that includes the problem setup (Section 3.2.1), a description of Diffusion Maps (Section 3.2.2) and a description of the general patch-to-tensor embedding scheme based on the construction of a super-kernel (Section 3.2.3). Linear projection super-kernels are discussed in Section 3.3. Description of the diffusion super-kernel is given in Section 3.4. Description of the linear-projection diffusion super-kernel is given in Section 3.5. Numerical examples, which demonstrate some aspects of the above constructions, are presented in Sec-
67 tion 3.6. The application of the proposed patch-to-tensor embedding for data-analysis tasks is demonstrated in Section 3.7. Technical proofs are given in the appendix in Section 3.A Benefits of patch processing In this section we provide additional motivation and justification for the approach of analyzing patches rather than individual points. The two main questions that should be addressed for such a justification are: 1. Why is patch processing, which is also called vector processing, the right way to go when we want to manipulate high-dimensional data? 2. Do these patches exist in real-life datasets? We will provide brief answers to both questions here. We assume that the processed data have been generated by some physical phenomenon, which is governed by an underlying potential [NLCK06a, NLCK06b]. Therefore, the affinity kernel will reveal clustered areas that correspond to neighborhoods of the local minima of this potential. In other words, these high-dimensional data points reside on several patches located on the low dimensional underlying manifold. On the other hand, if the data is spread sparsely over the manifold in the high-dimensional ambient space, then the application of an affinity kernel to the data will not reveal any patches/clusters. In this case, the data is too sparse to represent or detect the underlying manifold structure, and the only available processing tools are variations of nearest-neighbor algorithms. Therefore, data points on a low-dimensional manifold in a high-dimensional ambient space can either reside in locally-defined patches, and then the method in this chapter is applicable to it, or scattered sparsely all over the manifold and thus there is no detectable coherent physical phenomenon that can provide an underlying structure for it. Since the algorithm in this chapter is based on a manifold learning approach, it is inapplicable in the latter case. In general, all the tools that extract intelligence from high-dimensional data assume that under some affinity kernel there are data points that reside on locally-related patches, otherwise no intelligence (or correlations) will be extracted from the data and it can be classified as noise of uncorrelated data points. Therefore, the local patches, and not the individual points, are the basic building blocks for correlations and underlying structures in the dataset, and their analysis can provide a more natural representation of meaningful insights to the patterns that govern the analyzed phenomenon. The proposed methodology in this chapter is classified as a spectral method. Spectral methods are global in the sense that they usually require the relations between all the samples in the dataset. This global considera- 49
68 50 tion hinders their use in practical large-scale problems due to high memory (e.g., fitting the kernel matrix in memory) and computational costs. However, in many cases there are many duplicities, or near duplicities, in massive datasets and a the number of different patches of closely-related data-points is significantly less then the number of samples in the dataset. Processing patches, instead of individual data points, reduces the many redundancies that usually occur in massive datasets, thus, it enables also to localize spectral processing and reduce these overheads and impracticalities. 3.2 Overview Problem setup Let M R m be a set of n points sampled from a manifold M that lies in the ambient space R m. Let d m be the intrinsic dimension of M, thus, it has a d-dimensional tangent space T x (M), which is a subspace of R m, at every point x M. If the manifold is densely sampled, the tangent space T x (M) can be approximated by a small enough patch (i.e., neighborhood) N(x) M around x M. Let o 1 x,..., o d x R m, where o i x = (o i1 x,..., o im x ) T, i = 1,..., d, form an orthonormal basis of T x (M) and let O x R m d be a matrix whose columns are these vectors: O x o 1 x o i x o d x x M. (3.2.1) We will assume from now on that vectors in T x (M) are expressed by their d coordinates according to the presented basis o 1 x,..., o d x. For each vector u T x (M), the vector ũ = O x u R m is the same vector as u represented by m coordinates, according to the basis of the ambient space. For each vector v R m in the ambient space, the vector v = O T x v T x (M) is the linear projection of v on the tangent space T x (M). Section explains the application of the original diffusion maps method for the analysis of the dataset M. Then, Section describes the new construction we propose for embedding patches of the manifold M based on the points in the dataset M DM on discrete data The DM method from [CL06a, Laf04] was briefly discussed in Chapter 1. In this chapter we will use this method in a setting that analyzes the finite
69 dataset M. In this setting, we use the Gaussian isotropic kernel matrix K R n n, whose elements are k(x, y) e x y ε, x, y M, where ε is a meta-parameter of the algorithm. Kernel normalization of K with the degrees q(x) y M k(x, y), x M, produces the diffusion affinity kernel A R n n whose elements are 51 a(x, y) k(x, y) = 1 q(x)p(x, y) x, y M, (3.2.2) q(x)q(y) q(y) where p(x, y) k(x, y)/q(x) are the transition probabilities of the underlying diffusion process of DM. The matrix A is a symmetric conjugate of the rowstochastic transition matrix P R n n whose elements are these transition probabilities. The eigenvalues 1 = σ 0 σ 1... of A and their corresponding eigenvectors ψ 0, ψ 1,... are used by DM to embed each data point x M onto the point Ψ(x) = (σ i ψ i (x)) δ i=0 for a sufficiently small δ. The value of δ determines the dimension of the embedded space and it depends on the decay of the spectrum of A. The DM method uses scalar values to describe the affinities between points on the manifold. We extend this method by considering affinities, or relations, between patches (i.e., neighborhoods of points) on the manifold. These relations cannot be expressed by mere scalar values, since the similarity between patches must contain information about their relative positions in the manifold, their orientations and the correlations between their coordinates. We suggest to use the tangent spaces of the manifold M (i.e., similarities between them) together with scalar affinities between their tangential data points, to construct a block matrix, where each block represents the affinity between two patches. The rest of this section describes the construction of such block matrices that we call super-kernels Super-kernel Let Ω R n n be an affinity kernel defined on M R m, i.e., each row or each column in Ω corresponds to a data point in M, and each element in it, [Ω] xy = ω(x, y) x, y M, represents an affinity between x and y. We will require, by definition, that Ω will be symmetric and positive semi-definite. Furthermore, we will require that its elements satisfy ω(x, y) 0 x, y M. The exact definition of Ω can vary. We will present few ways to define it in the following sections. For x, y M, let O xy R d d be a d d matrix that represents the similarity between the matrices O x and O y, which were defined in Eq The matrices O x and O y represent bases of the tangent spaces T x (M) and
70 52 T y (M), respectively. Thus, the matrix O xy represents, in some sense, the similarity between these tangent spaces. We will refer to it as a tangent similarity matrix. We will require that the tangent similarity matrices satisfy the following condition: O xy = O T yx x, y M. (3.2.3) Specific definitions of such tangent similarity matrices will be presented in the following sections. Defintion uses the affinity kernel Ω and the tangent similarity matrices O xy in the following definition to introduce the concept of a super-kernel. Definition (Super-kernel). A super-kernel is a matrix G R nd nd where in terms of blocks, it is a block matrix of size n n and each block in it is a d d matrix. Each row and each column of blocks in G corresponds to a point in M, and a single block G xy (where x, y M) represents an affinity or similarity between the patches N(x) and N(y). Each block G xy R d d is defined as G xy ω(x, y)o xy, x, y M. It is convenient to consider each single cell in G as an element in a block, i.e., [G xy ] ij where x, y M and i, j {1,..., d}. We can also use the vectors o i x and o j y to apply this indexing scheme and use the following notation: g(o i x, o j y) [G xy ] ij x, y M, i, j {1,..., d}. (3.2.4) In this notation, it is easy to see that G is symmetric since [G xy ] ij = [G T yx] ij = [G yx ] ji x, y M, i, j {1,..., d}, where the first equality is due to Eq , to the symmetry of Ω and the definition of G xy. It is important to note that g(o i x, o j y) is only a notation for convenience reasons and a single element of a block in G does not necessarily have any special meaning. The block itself, as a whole, holds meaningful similarity information. We will use spectral decomposition for analyzing a super-kernel G, and utilize it to embed the patches N(x) of the manifold (for x M) into a tensor space. Let λ 1 λ 2... λ l be the l most significant eigenvalues of G and let φ 1, φ 2,..., φ l be their corresponding eigenvectors. According to the spectral theorem, if l is greater than the numerical rank of G, then G l λ i φ i φ T i, (3.2.5) i=1
71 where the eigenvectors are treated as column vectors. For convenience reasons, we will treat this approximation as an equality, since, from a theoretical point of view, l can always be chosen to be large enough for actual equality to hold. In practice, the exact value of l depends on the numerical rank of G, the decay of its spectrum, and the exact application of the construction. Usually, however, the affinity kernel and the tangent similarity matrices can be chosen in such a way that a small l will obtain sufficient accuracy for the desired task. Each eigenvector φ i, i = 1,..., l, is a vector of length nd. We use a similar notation to Eq to denote each of its elements as φ i (o j x) where x M and j = 1,..., d. An eigenvector φ i can also be regarded as a vector of n sections, each of which is a vector of length d that corresponds to a point x M on the manifold. To express this notion we use the notation ϕ j i (x) = φ i(o j x) x M, i = 1,..., l, j = 1,..., d. (3.2.6) Thus, the section that corresponds to x M in the eigenvector φ i is the vector (ϕ 1 i (x),..., ϕ d i (x)) T. We use the eigenvalues and eigenvectors of G to construct a spectral map whose definition is similar to the standard (i.e., classic) diffusion map: 53 λ µ 1φ 1 (o j x) Φ(o j x) =., (3.2.7) λ µ l φ l(o j x) where µ is a meta-parameter of the embedding. It depends on the specific affinity kernel and on tangent similarity matrices that are used. In Section 3.3, we will use the value µ = 1 (for a positive semi-definite G), and in 2 Section 3.4, we will use the value µ = 1. By using this construction, we get nd vectors of length l. Each x M corresponds to d of these vectors, i.e., Φ(o j x), j = 1,..., d. We use these vectors to construct the tensor T x R l R d for each x M, which is represented by the following l d matrix: T x Φ(o 1 x) Φ(o d x) x M. (3.2.8) In other words, the coordinates of T x (i.e., the elements in this matrix) are [T x ] ij = λ µ i ϕj i (x) x M, i = 1,..., l, j = 1,..., d, (3.2.9)
72 54 where µ is the meta-parameter that is used in Eq Each tensor T x represents an embedding of the patch N(x), x M, into the tensor space R l R d. In the following sections, we will present several constructions for a superkernel G and the properties of the embedded tensors, which result from its spectral analysis, are examined. Specifically, we will relate the Frobenius distance between the embedded tensors, regarded as their coordinate matrices, to the relations between their corresponding patches in the original manifold. 3.3 Linear projection super-kernel The proposed construction of a super-kernel (see Definition 3.2.1) encompasses both the affinities between points on the manifold M and the similarities between their tangent spaces. The latter are expressed by the tangent similarity matrices, which can be defined in several ways. In this chapter, we will use linear projection operators to define these similarity matrices. Specifically, for x, y M, assume that T x (M) and T y (M) are two tangent spaces of the manifold. The operator O T x O y, which defines a linear projection from T y (M) to T x (M) via the ambient space R m, is used to describe the similarity between them. The obvious extreme cases are an identity matrix, which indicates on complete similarity and a zero matrix, which indicates on orthogonality (i.e. complete dissimilarity). The following definition formalizes the use of these linear projections as tangent similarities in the construction of a super-kernel. Definition (LP super-kernel). A Linear Projection (LP) super-kernel is a super-kernel G, as defined in Definition 3.2.1, where the tangent similarity matrices are defined by the linear projection operators O xy = O T x O y x, y M, i.e., for every x, y M, the blocks of G are defined as G xy = ω(x, y)o T x O y. The linear projection operators, which define the tangent similarity matrices by a LP super-kernel, express some important properties of the manifold structure, e.g., curvatures between patches and differences in orientation. While there might be other ways to construct a super-kernel that expresses these properties, LP super-kernels do have an important property, which is given by the following theorem: Theorem A LP super-kernel G is positive semi-definite and its spectral norm satisfies G Ω, where Ω is the spectral norm of the affinity kernel.
73 To prove this theorem, we first need to introduce some notations. Let u R nd be an arbitrary vector of length nd. We can view u as having n subvectors of length d, where each subvector u x corresponds to a point x M on the manifold. Let U R n d be a n d matrix such that for every x M its rows are the subvectors u x and let u 1,..., u d be the columns of this matrix. Figure illustrates these notations. An element in U, which is in a row u x, x M and a column u j, j = 1,..., d, is denoted by u j x. 55 u R nd n subvectors d cells d cells u x1. u xn nd cells U R n d n rows u x1. u xn } {{ } d columns = u 1 u d }{{} d columns n rows Figure 3.3.1: An illustration of viewing an arbitrary vector u R nd as a matrix U R n d. Note that x 1,..., x n are used here to denote all the points in M. Each subvector u x, x M, has d elements, therefore, it can be seen as a vector on the tangent space T x (M). We define the same vector, presented by m coordinates of the ambient space R m, as ũ x = O x u x, x M. Since both u x and ũ x represent the same vector (in two different orthonormal coordinate systems), their norms have the same value. Indeed, ũ x 2 = ũ T x ũ x = u T x O T x O x u x = u T x u x = u x 2, x M. (3.3.1) We denote by Ũ Rn m the n m matrix whose rows are ũ x for every
74 56 x M and we denote its columns by ũ 1,..., ũ m. Each element in Ũ, which is in a row ũ x, x M, and a column ũ i, i = 1,..., m, is denoted as ũ i x. Lemma Let G be a LP super-kernel and let u R nd be an arbitrary vector of length nd. Then, u T Gu = m (ũ i ) T Ωũ i, where Ω is the affinity kernel, always holds. The proof of Lemma is technical and it is given in Appendix 3.A. We can now prove Theorem Proof of Theorem Let G R nd nd be a LP super-kernel and let u R nd be an arbitrary vector of length nd. First, we recall that we require the affinity kernel Ω to be positive semi-definite, thus, from Lemma we get therefore, i=1 v T Ωv 0 v R n, (3.3.2) u T Gu = m (ũ i ) T Ωũ i 0. (3.3.3) i=1 Since u is an arbitrary vector of length nd, Eq shows that G is positive semi-definite. This proves the first part of the theorem. Next, we denote the spectral norm of Ω by σ = Ω, thus, therefore, from Lemma we get m m u T Gu = (ũ i ) T Ωũ i σ ũ i 2 = σ i=1 v T Ωv σ v 2 v R n (3.3.4) i=1 Then, by using Eq we get z M ũ z 2 = z M u z 2 = z M By combining Eqs and 3.3.5, we get m ũ i z 2 = σ ũ z 2. z M i=1 z M (3.3.5) d u j z 2 = u 2. (3.3.6) j=1 u T Gu σ u 2. (3.3.7) Since u is an arbitrary vector of length nd, Eq shows that the Raleigh quotient of G is at most σ. We have already shown that G is positive semi-definite, hence, its spectral norm is its largest eigenvalue, which is also the maximal value of its Raleigh quotient. Therefore, the spectral norm of G is at most σ, and the second part of the theorem is also proved.
75 In sections and 3.3.2, we present two constructions of LP superkernels. The first construction preserves global tangent similarities by ignoring the affinity between the points in M. The second construction uses binary affinities (i.e., 0 or 1) that preserves local tangent similarities. In Section 3.5, we will present our final construction, which uses the diffusion affinity kernel to define a LP super-kernel that is used to define the patch-to-tensor embedding Global linear-projection (GLP) super-kernel A simple way to construct a LP super-kernel is to ignore the affinity kernel completely. In other words, we can use an all-ones matrix as the affinity kernel, thus, the resulting super-kernel will contain only the information about the tangent similarities between patches. While this approach may not be useful in practice, it will provide an insight into the effect the linear projection operators have on the embedding achieved by using a LP super-kernel. The following definition formalizes the described construction of a global LP super-kernel. Definition (GLP super-kernel). A Global Linear-Projection (GLP) super-kernel is a LP super-kernel G, as was defined in Definition 3.3.1, where the affinity kernel is defined as a constant ω(x, y) 1 x, y M, i.e., the affinity kernel Ω, in this case, is an all-ones matrix, and the blocks of G are defined as G xy = O T x O y, x, y M. By definition, a GLP super-kernel G is a LP super-kernel, thus, Theorem applies to it and G is positive semi-definite. Therefore, all the eigenvalues of G are non-negative, and a spectral map Φ (Eq ) can be defined using µ = 1. The defined spectral map can then be used to embed 2 each patch N(x), x M, to a tensor T x (Eq ). In fact, such an embedding can be defined for every LP super-kernel. The following lemma shows an important relation between the blocks of a LP super-kernel G and the embedded tensors resulting from this construction. Lemma Let x, y M be two points on the manifold and let T x and T y be their embedded tensors (Eq ). If the embedding is done by using the spectral map Φ (Eq ) of a LP super-kernel G with the meta-parameter µ = 1 2, then G xy = T T x T y x, y M, where the tensors are treated as matrices (i.e., their coordinate matrices). 57
76 58 Lemma is a result from the construction of the embedded tensors, the definition of LP super-kernels and the application of the spectral theorem to them. A detailed proof of this lemma is given in Appendix 3.A. A GLP super-kernel preserves global tangent similarities, which are defined as linear projection operators, between patches. The resulting embedded tensors can be regarded as l d matrices and their distances can be defined by a matrix norm. Let D be a matrix norm. The distance between two tensors T x and T y, x, y M, is defined as D(T x T y ). Theorem shows that for matrix norms of a certain form, this distance is equivalent to the distance between the basis matrices O x and O y under the same norm. Theorem Let D be a matrix norm, defines as D(S) = f(s T S) for every matrix S of arbitrary size, where f is a suitable function from the set of all matrices (of all sizes) to R. Let x, y M be two points on the manifold and let T x and T y be their embedded tensors (Eq ). If the embedding is done by using the spectral map Φ (Eq ) of a GLP super-kernel G with the meta-parameter µ = 1 2, then D(T x T y ) = D(O x O y ) x, y M, where the tensors are treated as matrices (i.e., their coordinate matrices). Proof. For x, y M, let O x and O y be the matrices defined in Eq and let D be the matrix norm described in the theorem. Then, by definition, D(O x O y ) = f((o x O y ) T (O x O y )) x, y M. (3.3.8) We recall the definitions of the blocks in a GLP super-kernel G, thus, the matrix product in the right-hand side Eq is (O x O y ) T (O x O y ) = G xx G xy G yx + G yy x, y M, therefore, according to Lemma 3.3.3, (O x O y ) T (O x O y ) = T T x T x T T x T y T T y T x + T T y T y = (T x T y ) T (T x T y ) x, y M. (3.3.9) By combining Eqs and we get D(O x O y ) = f((t x T y ) T (T x T y )) = D(T x T y ) x, y M, as stated in the theorem.
77 Theorem shows a relation between matrix distances in the original space and the same type of distances in the embedded space. The distance metrics covered by this theorem are defined by the matrix norms of the form D(S) = f(s T S). In fact, two popular matrix norms (i.e., the Frobenius norm and the spectral norm) satisfy this property, and are thus covered by this theorem. The following corollary states that this fact in a formal way. Corollary Let x, y M be two points on the manifold and let T x and T y be their embedded tensors (Eq ). If the embedding is done by using the spectral map Φ (Eq ) of a GLP super-kernel G, with the meta-parameter µ = 1 2, then: 1. The Frobenius distances, defined by the Frobenius (also called Hilbert- Schmidt) norm, in the embedded tensor space satisfy T x T y F = O x O y F ; 2. The spectral distances, defined by the spectral (also called operator) norm, in the embedded tensor space satisfy T x T y = O x O y. Proof. The Frobenius norm is defined by S F = tr(s T S) and the spectral norm is defined by S = λ max (S T S) (where λ max is a the largest eigenvector of a square matrix). Both definitions fit the form of the matrix norm in Theorem 3.3.4, thus its result applies for the distances defined by these norms Local linear-projection super-kernel We presented an important property (Theorem and Corollary 3.3.5) of the GLP super-kernel construction, but it also has a critical flaw. Manifolds are based on local structures and the similarities between tangent spaces of far-away points are meaningless. The next construction introduces the notion of locality in a LP super-kernel. We use the notion of neighboring points to define a simple local affinity kernel. We use the notation x y to denote the fact that two points x, y M on the manifold are considered neighbors of one another. It means that x y [N(x) N(y) ], i.e., x and y are neighbors if their patches have mutual points. A more restrictive definition requires neighbors to be in the patches of one another, i.e., x y [x, y N(x) N(y)]. The exact definition of neighboring points is not crucial for the presented construction.
78 60 The following definition uses the concept of neighboring points to construct a local LP super-kernel by using a binary affinity kernel, which indicates whether two points are neighbors (i.e., their affinity is 1) or not (i.e., their affinity is 0). Definition (LLP super-kernel). A Local Linear-Projection (LLP) superkernel is a linear-projection super-kernel G, as was defined in Definition 3.3.1, where the affinity kernel is defined as { 1 x y ω(x, y) x, y M, 0 otherwise i.e., the blocks of G are defined as G xy = O T x O y for x y M and as the zero matrix for non-neighboring points in M. Since, by definition, a LLP super-kernel is a LP super-kernel, both Theorem and Lemma are applicable for it. Thus, we can use it to embed patches on the manifolds to tensors by using a spectral map Φ (Eq ), with µ = 1, to construct the tensors in Eq Theorem showed that 2 for a wide range of matrix distance metrics, when the embedding is done with a GLP super-kernel, the distance between embedded tensors is equal to the distance between the basis matrices (Eq ) of the original patches. While the result in this theorem is not globally true when the embedding is done with a LLP super-kernel, Theorem shows that a similar result does apply to neighboring points in this embedding. Theorem Let D be a matrix norm of the same form as in Theorem 3.3.4, let x y M be two neighboring points on the manifold and let T x and T y be their embedded tensors (Eq ). If the embedding is done by using the spectral map Φ (Eq ) of a LLP super-kernel G, with the meta-parameter µ = 1 2, then D(T x T y ) = D(O x O y ) x, y M, where the tensors are treated as matrices (i.e., their coordinate matrices). Proof. Let G be the LLP super-kernel that is used to embed the data points in the theorem. According to Definition 3.3.3, G xy = O T x O y and G yx = O T y O x. Also, according to the same definition, since any point is a neighbor of itself then we get G xx = O T x O x and G yy = O T y O y. Therefore, (O x O y ) T (O x O y ) = G xx G xy G yx + G yy,
79 and by combining this result with Eq (from the proof of Theorem 3.3.4), which still applies here (since matrix norms of the same form are considered in both theorems), we get D(O x O y ) = f(g xx G xy G yx + G yy ). Since Lemma applies for LLP super-kernels, a calculation similar to the one in Eq gives D(O x O y ) = f((t x T y ) T (T x T y )) = D(T x T y ), as stated in the theorem. Theorem extends Theorem to the case of LLP super-kernels and it shows that the embedding achieved by it is locally similar to the one achieved by a GLP super-kernel. Locally similar means that the distances between the embedded tensors are equivalent in both cases of neighboring points. Corollary stated that the result of Theorem applies, in particular, to the Frobenius distance and to the spectral distance. A similar corollary can be stated for the result of Theorem and its proof is the same as in Corollary Corollary Let x y M be two neighboring points on the manifold and let T x and T y be their embedded tensors (Eq ). If the embedding is done by using the spectral map Φ (Eq ) of a LLP super-kernel G with the meta-parameter µ = 1. Then, the Frobenius distances, defined by 2 the Frobenius norm in the embedded tensor space and the spectral distances, defined by the spectral norm, in the embedded tensor space satisfy T x T y F = O x O y F and T x T y = O x O y, respectively. The presented construction of a LLP super-kernel takes us one step closer to our final construction of a LP super-kernel that will be used to define the desired patch-to-tensor embedding, since it considers the local nature of the manifold. Section 3.4 will further examine this aspect by utilizing the diffusion affinity kernel to introduce the notion of locality in the construction of a super-kernel Diffusion super-kernel The definition of a super-kernel (Definition 3.2.1) is based on an affinity kernel, which describes the relations between points on the manifold, and a set of tangent similarity matrices, which describe the relations between
80 62 tangent spaces of the manifold. Section 3.3 explored mainly the latter part of this construction (i.e., the matrices O xy for x, y M), and proposed two simple definitions of an affinity kernel to use in conjunction with the proposed LP super-kernel (see Definitions and 3.3.3). In this section, we set aside the exact definition of the tangent similarity matrices and focus on the affinity kernel that is used. Specifically, Definition suggests to use the classic diffusion affinity kernel A (defined in Eq ) to describe the affinities in the construction of a super-kernel. Definition (Diffusion super-kernel). A diffusion super-kernel is a superkernel G, as was defined in Definition 3.2.1, where the affinity kernel is defined as ω(x, y) = a(x, y), x, y M, i.e., the affinity kernel is the symmetric diffusion kernel. The Euclidean distance between data points in the embedded space, which results from the application of the usual diffusion maps, is equal to a diffusion distance in the original ambient space. This diffusion distance measures the distance between two diffusion bumps a(x, ) and a(y, ), each of which is a row in the symmetric diffusion kernel that defines the diffusion map. From a technical point of view, this relation means that the Euclidean distance between two arbitrary points in the range of a diffusion map is equal to the Euclidean distances between the corresponding rows of its symmetric diffusion kernel. Lemma establishes the same technical relation between the spectral map Φ (Eq ) of a diffusion super-kernel G and the rows of the super-kernel itself. Lemma Let G be a diffusion super-kernel and let Φ be a spectral map (Eq ) of this kernel with the meta-parameter µ = 1. For every x, y M and j = 1,..., d, Φ(o j x) Φ(o j y) = g(o j x, ) g(o j y, ), where g(o j x, ) (or g(o j y, )) is a vector whose elements are g(o j x, o ξ z) (or g(o j y, o ξ z)), which are defined in Eq for every z M and ξ = 1,..., d. The proof of Lemma 3.4.1, which appears in the Appendix 3.A, is based on the spectral theorem. It is similar to the corresponding result regarding the standard diffusion maps method. The relation provided by Lemma is useful from a technical point of view, but it does not provide meaningful information about the relation between the embedded tensors and the original patches. Theorem shows a relation between tensor distances (in the embedded space), defined using the Frobenius norm, to an extended diffusion distance. The extended diffusion distance encompasses the information
81 about similarities between tangent spaces, as well as the affinities between points on the manifold in a fashion similar to the definition of the original diffusion distance. Theorem Let x, y M be two points on the manifold and let T x and T y be their embedded tensors (Eq ). If the embedding is done by using the spectral map Φ (Eq ) of a diffusion super-kernel G with the meta-parameter µ = 1, then T x T y 2 F = z M a(x, z)o xz a(y, z)o yz 2 F, 63 where the tensors are treated as matrices (i.e., their coordinate matrices) when computing the Frobenius distance between them. Proof. First, we use the definition of the Frobenius norm and the construction of the embedded tensor space to get T x T y 2 F = = l i=1 d λ i ϕ j i (x) λ iϕ j i (y) 2 = j=1 d j=1 l λ i φ i (o j x) λ i φ i (o j y) 2 d Φ(o j x) Φ(o j y) 2. (3.4.1) j=1 Next, we combine this result with Lemma to get T x T y 2 F = d g(o j x, ) g(o j y, ) 2 = j=1 = z M d j=1 ξ=1 as states in the theorem. d i=1 j=1 z M ξ=1 d a(x, z)[o xz ] jξ a(y, z)[o yz ] jξ 2 = z M a(x, z)o xz a(y, z)o yz 2 F, d g(o j x, o ξ z) g(o j y, o ξ z) 2 Corollary reinforces our argument that the presented metric is indeed an extension of the original diffusion distance by presenting a case in which both metrics converge up to multiplication by a constant (i.e., d). Corollary In the context of Theorem 3.4.2, if all the tangent similarity matrices are orthogonal, and for every x, y, z M, the product O T xzo yz is symmetric and positive semidefinite, then T x T y 2 F = d a(x, ) a(y, ) 2,
82 64 where a(u, ) denotes a vector of length n with the entries a(u, z) for every z M. In other words, the extended diffusion distance in this case is the original diffusion distance multiplied by d. Proof. According to Theorem 3.4.2, T x T y 2 F = z M a(x, z)o xz a(y, z)o yz 2 F x, y M, and since the Frobenius norm comes from the Frobenius inner product (denoted by : ), a(x, z)o xz a(y, z)o yz 2 F = a(x, z)o xz 2 F 2 (a(x, z)o xz : a(y, z)o yz ) + a(y, z)o yz 2 F = a(x, z) 2 tr(o T xzo xz ) 2a(x, z)a(y, z) tr(o T xzo yz ) + a(y, z) 2 tr(o T yzo yz ) x, y, z M. If O xz and O yz are both d d orthogonal matrices, as the corollary assumes, then so are O T xzo xz, O T yzo yz, and O T xzo yz. In fact, the first two are the d d identity matrix, whose trace is d. The product O T xzo yz is also symmetric and positive semidefinite, by the assumption in the corollary, thus, its d eigenvalues are all ones and its trace is d. The traces of these matrices are all d, thus, a(x, z)o xz a(y, z)o yz 2 F = da(x, z) 2 2da(x, z)a(y, z) + da(y, z) 2, therefore, if we combine this result with Theorem 3.4.2, we get T x T y 2 F = d z M ( a(x, z) 2 2a(x, z)a(y, z) + a(y, z) 2) as stated in the corollary. = d( a(x, ), a(x, ) 2 a(x, ), a(y, ) + a(y, ), a(y, ) ) = d a(x, ) a(y, ) 2 x, y M, 3.5 Linear-projection diffusion super-kernel In Section 3.4, we utilized the diffusion affinity kernel to construct a superkernel without defining the tangent similarity matrices. In Section 3.3, we presented a general construction of a super-kernel that is based on linearprojection tangent similarity matrices (see Definition 3.3.1) without defining
83 the affinity kernel. Definition combines these constructions and introduces our construction of a linear-projection diffusion super-kernel. We will construct a patch-to-tensor embedding, which maps patches of the manifold into a meaningful tensor space by using the spectral map of this super-kernel. Definition (LPD super-kernel). A Linear-Projection Diffusion (LPD) super-kernel G is both a diffusion super-kernel as was defined in Definition and a LP super-kernel as was defined in Definition 3.3.1, i.e., its blocks are defined as G xy = a(x, y)o T x O y, x, y M. Since a LPD super-kernel is a LP super-kernel, Theorem applies to it. We recall that the spectral norm of the symmetric diffusion kernel is A = 1, therefore, we get Corollary for the case of LPD super-kernels, whose proof is an immediate result of this discussion. Corollary A LPD super-kernel G is positive semi-definite and its operator norm satisfies G 1. Another immediate result of Theorem in this case, or rather of Corollary 3.5.1, has to do with the eigenvalues of a LPD super-kernel: Corollary All the eigenvalues of a LPD super-kernel are between 0 and 1, i.e., its eigenvalues are 1 λ 1 λ Proof. According to Corollary 3.5.1, an LPD super kernel G is positive semidefinite, thus, its eigenvalues are non-negative and the largest one is equal to the spectral norm of G, which satisfies G 1, according to the same corollary. Therefore, every eigenvalue of G is at least 0 and at most is We recall that the original diffusion distance between two points x, y M a(x, ) a(y, ) 2 = z M(a(x, z) a(y, z)) 2. According to Theorem and Definition 3.5.1, when the spectral map Φ (Eq ) of a LPD super-kernel is used to embed the patches of these points, the Frobenius distance between the resulting embedded tensors (regarded as their coordinate matrices) satisfies T x T y 2 F = z M a(x, z)o T x O z a(y, z)o T y O z 2 F = z M d (a(x, z)ox T a(y, z)oy T )o j z 2. (3.5.1) j=1
84 66 The vectors o j z in this equation are unit vectors that form an orthonormal basis of the tangent space T x (M) at the point z M. For each point z M, the matrix [a(x, z)o T x a(y, z)o T y ] is applied to each of these unit vectors and the squared lengths of the resulting vectors are summed. These terms can be seen as extensions of the terms (a(x, z) a(y, z)) of the original diffusion distance, which only consider the differences between scalar affinities. Let u be a unit vector. We examine the result of applying the matrix [a(x, z)o T x a(y, z)o T y ], x, y, z M, to such a vector (see Fig ). First, since u is a unit vector, a(x, z)u and a(y, z)u are vectors of lengths a(x, z) and a(y, z), respectively, in the same direction as u (see Fig (a)). The vector a(x, z)o T x u is a linear projection of the vector a(x, z)u on the tangent space T x (M), where the resulting vector is represented by the d coordinates of this a(x, z) a(y, z) (a) Two vectors in the direction of u. (b) Projection of the two vectors on two tangent spaces. a(x, z)o T x u a(y, z)o T y u a(x, z)o T x u a(y, z)o T y u (c) The difference between the local coordinates of the projected vectors. Figure 3.5.1: An illustration of the application of the matrix [a(x, z)o T x a(y, z)o T y ] to the unit vector u.
85 tangent space (see Fig (b)). Similarly, a(y, z)o T y u is the projection of a(y, z)u on T y (M), represented by the d local coordinates of this tangent space. The resulting vector a(x, z)o T x u a(y, z)o T y u contains the difference between the two resulting vectors of length d (see Fig (c)). If the lengths of vectors in the direction of u are not changed by these projections (e.g., u is on both T x (M) and T y (M)), and if the coordinate systems of these tangent spaces are equivalent, in the sense that the direction of these projections is the same in both of them, then the length of the resulting vector will simply be the scalar difference a(x, z) a(y, z). This is an extreme scenario. In most cases, these differences (i.e., the scalar difference and the length of the difference vector) will not coincide due to the curvature of the manifold and the difference in coordinate systems on the manifold. We have shown that the embedding achieved by spectral analysis of a LPD super-kernel is similar, in some sense, to the one achieved by the original diffusion maps method. We use the name Patch-to-Tensor Embedding (PTE) for the presented embedding, 67 N(x) P T E T x x M, (3.5.2) which maps each patch of the manifold to the corresponding tensor, defined by Eq , by using the spectral map Φ (Eq ) with µ = 1, of a LPD super-kernel that is constructed over the input set M of points on the manifold. In section 3.6, a simple (yet not optimal) algorithm for constructing a Patch-to-Tensor Embedding is presented. The results of its applications on synthetic datasets are presented in Section 3.6 and its utilizations for data clustering & classification and for image segmentation are presented in Section 3.7. The finite LPD super-kernel that we presented here is further explored in Chapter 4, where its properties when it becomes continuous are examined and analyzed. Specifically, the infinitesimal generator of this super-kernel and the stochastic process defined by it are explored. It is shown there that the resulting infinitesimal generator of this super-kernel converges to a natural extension of the original diffusion operator from scalar functions to vector fields. This operator is shown to be locally equivalent to a composition of linear projections between tangent spaces and the vector-laplacians on them. A LPD process can then be defined by using the LPD super-kernel as a transition operator while extending the process to be continuous. The LPD process propagates tangent vectors over the manifold (see Chapter 4). Since it is a stochastic process, it has an inherent time parameter, which we will refer to as the diffusion time in the rest of this chapter. In the finite discrete case, this parameter is interpreted as the number of steps performed by the diffusion. The original DM algorithm also has such diffusion
86 68 time parameter, which is expressed as powers of the diffusion operator or, more conveniently, as powers of the used eigenvalues in the embedding process. In Section 3.7, we use different diffusion times (also expressed as powers of the used spectrum in the embedding) to obtain ideal data clustering in the resulting embedded space of the LPD-based PTE. 3.6 Numerical examples This section presents several numerical results that demonstrate the PTE characteristics on synthetically produced datasets. Specifically, the following demonstration relates Theorem and the corresponding corollary (Corollary 3.5.1) to the LPD super-kernels. Algorithm 1 is used to construct a PTE for the analysis of three exemplary manifolds. Algorithm 1: Patch-to-Tensor Embedding Construction (PTEC) Input: Data points: x 1,..., x n R m and parameters: Patch size ρ and l For each x M estimate an orthonormal basis O x R m d of the local tangent space based on ρ points uniformly distributed over a small neighborhood of x; Construct a diffusion affinity kernel A according to Eq ; Construct a LPD super kernel G according to Definition 3.5.1; Construct spectral map Φ(o j x) for j = 1,..., d according to Eq utilizing the SVD decomposition of the constructed LPD super-kernel G; Construct a Tensor T x R l R d for each x M according to Eq The examined manifolds are illustrated in Fig They include: the unit sphere S 2, the three dimensional Swiss roll and the three dimensional Mobius band. The analyzed datasets were produced using the following steps. We sample 2000 points uniformly from each manifold embedded in R 3. Each set of points was extended to an ambient space of 17 by a linear transformation operator Q R The linear operator Q was chosen randomly with uniform distribution under the constraint Q T Q 0. The positive definiteness constraint guaranty that Q is non-singular. Algorithm 1 with parameters ρ = 30 and l = 3 was utilized to find the LPD super-kernel and the corresponding mapping for each example. The choice of the value l = 3 was calculated by aggregating all the estimated local dimensions of each tangent space T x (M) following the footsteps of [SW11].
87 69 (a) A Sphere (b) A Swiss Roll Figure 3.6.1: Examined manifolds (c) A Mobius Band Figure describes the numerical rank of the LPD super-kernel for increasing values of µ. The resulting eigenvalues for all of the examples are decaying for µ = 1 and the decay increases as µ increases. In order to analyze the support interval that the eigenvalues span, we compute the Cumulative Distribution Function (CDF) of the eigenvalues of the LPD super-kernel. The corresponding CDF is the probability that any real-valued eigenvalue of the LPD super-kernel will have a value less than or equal to a threshold τ. More rigorously, the CDF is defined as F (τ, f (λ i )) = P (λ i τ), (3.6.1) where f (λ i ) is the distribution function of λ i and τ is a given threshold. The utilization of the CDF enables a compact and informative presentation of the characterization of the relevant eigenvalues. The CDF describes the interval on which there is a positive probability to find eigenvalues and what is the percentage of non-negligible eigenvalues from all the eigenvalues distributions. The estimated CDFs of the eigenvalues for all the manifold examples are
88 70 (a) (b) (c) Figure 3.6.2: The 100 largest eigenvalues of the LPD-super kernels that corresponds to (a) Sphere, (b) Swiss roll, and (c) Mobius band
89 71 (a) (b) (c) Figure 3.6.3: The CDFs of the eigenvalues of the LPD-super kernels of (a) Sphere, (b) Swiss Roll and (c) Mobius band
90 72 presented in Fig For each LPD super-kernel instance, we estimated the distribution function f (λ i ) by integrating the corresponding histogram of the resulted eigenvalues. According to the calculated CDFs the examined LPD-super kernels have positive probability only on the [0, 1] interval as was suggested by Theorem and by the corresponding Corollary for the LPD-super kernel. Furthermore, the CDFs calculated probabilities, which an eigenvalue will have a value which is less than 0.1 on the Sphere, Swiss roll and the Mobius examples, are 0.996, and 0.990, respectively. The CDFs have high probabilities to have a small eigenvalue, hence, only a small number of eigenvalues and their corresponding eigenvectors are required to preserve the structure and variability in the LPD super-kernel matrices. 3.7 Data analysis using patch-to-tensor embedding PTE provides a general framework that can be utilized in a wide collection of data analysis tasks such as clustering, classification, anomaly detection and related manifold learning tasks. In this section, we demonstrate the application of the PTE method to two data analysis challenges: 1. Classification of breast tissue impedance measurements. 2. Data clustering that is based on image segmentation Electrical impedance breast tissue classification Biological tissues have complex electrical impedance related to the tissue dimension, the internal structure and the arrangement of the constituent cells. Therefore, the electrical impedance can provide useful information based on heterogeneous tissue structures, physiological states and functions [Sch59]. Electrical impedance techniques have long been used for tissue characterization [KPW70]. Recently, an interesting dataset of breast tissue impedance measurements was published [FA10]. The dataset consisted of 106 spectra recorded in samples of breast tissue from 64 patients undergoing breast surgery. Each spectrum consisted of twelve impedance measurements taken at different frequencies ranging from 488 Hz to 1 MHz. Detailed description of the data collection procedure as well as classification of the cases and frequencies used are given in [Jos96, dsdsj00]. Table shows the six classes of tissue that are represented in the given dataset.
91 73 Normal Tissue Classes Pathological Tissue Classes Number of Measurements Connective tissue (con) 14 Adipose tissue (adi) 22 Glandular tissue (gla) 16 Carcinoma (car) 21 Fibro-adenoma (fad) 15 Mastopathy (mas) 18 Table 3.7.1: The six classes of tissues that are represented in the analyzed dataset. Several extracted features from the impedance measurements for the classification preprocessing step were described in [dsdsj00]: I 0 - Impedivity at zero frequency (low frequency limit resistance) P A Phase angle at 500 khz S HF - High frequency slope of phase angle (at 250, 500 and 1000 KHz points) D 4 - Impedance distance between spectral ends AREA - Area under spectrum AREA D4 - Area normalized by D 4 IP Max - Maximum of the spectrum DR - Distance between I 0 and real part of the maximum frequency point P ERIM - Length of spectral curve The computed attributes are given in the dataset. More details are given in [dsdsj00]. A tissue classification method for the given impedance attributes is given in [dsdsj00]. The suggested method is based on a hierarchal architecture in which in the first stage a classifier is used to discriminated fatty tissue (Connective and Adipose tissue) from the non-fatty tissue (Carcinoma, Fibroadenoma, Mastopathy and Glandular tissue). At the second stage, additional classifier is used to discriminate Carcinoma tissue from other non-fatty tissue categories. The performances of the algorithm in [dsdsj00] are 100% success in discriminating the fatty from the non-fatty tissue at the first stage. The Carcinoma can be discriminated from the other non-fatty
92 74 tissue types with more than 86% success. The success in identifying the FMG class (Fibroadenoma + Mastopathy + Glandular tissue) is 94.5%. In this section, we follow the foot steps of [dsdsj00] in classifying the postprocessing attributes into the same tissue categories using PTE. Initially, the given dataset was normalized to have zero mean and a unit standard deviation for each attribute. Then, the PTE construction, detailed in Algorithm 1, was used to construct the LPD super kernel followed by embedding of the measurements into a tensor space. The affinity kernel is computed by Eq where ε was chosen as the mean Euclidean distance between all the pairs of data points in the given dataset. The parameters in the PTE construction were l = 5 and ρ = 66. They were chosen in an exhaustive search to optimize the classification accuracy. The classification performance is based on a leave-one-out methodology in which each of the measurements was labeled according to its nearest neighbor in the embedded tensor space. The Frobenius norm was used as the distance metric. The classification performance is described in Table Tissue category Correct detection False detection Miss-detection Fatty 97.2% 0 2.7% Carcinoma 86.36% 13.6% 9.5% FMG 93.9% 6.1% 6.1% Performance summary of the PTE-based classification algo- Table 3.7.2: rithm. Tissue category Correct detection False detection Miss-detection Fatty 100% 0% 0% Carcinoma 86.36% 13.6% 13.6% FMG 94.54% 5.45% 5.45% Table 3.7.3: Performance summary of the original classification performances from [dsdsj00]. The achieved classification performances, which were obtained by PTE with a single classification stage, are competitive to the ones in [dsdsj00], which are presented in Table The optimization of the classifier was done with respect to only two parameters: ρ the number of points per patch and l the number of eigenvectors from the application of the SVD procedure. This optimization was performed using leave-one-out cross-validations.
93 Image segmentation Image segmentation clusters pixels into image regions corresponding to individual surfaces, objects, or natural parts of objects. It plays a key role in many computer vision tasks such as object recognition, image compression, image editing and image retrieval. It has been extensively studied in computer vision [BF70, Pav77, HS85] and statistics with a vast number of different algorithms [JD88, KR90, JTLB04, NMA05]. Early techniques utilized region splitting or merging [BF70, OPR78, PL90], which correspond to divisive and agglomerative algorithms in the clustering literature [JTLB04]. More recent algorithms often optimize some global criterion such as intra-region consistency and inter-region boundary lengths or dissimilarity [MS89, MS01, FH04, CRD07]. Graph cut techniques from combinatorial optimization are used for image segmentation [BK04, RKB04, SM00]. Graph cut methods view the image as a graph weighted to reflect intensity changes and performs a max-flow/mincut analysis to find the minimum-weight cut between the source and the sink. One of the features of this algorithm is that an arbitrary segmentation may be obtained with enough user interaction and it generalizes easily to 3D and beyond. The PTE framework enables to view the image via a LPD super-kernel that reflects the affinities between pixels and the projection of the related tangent spaces. The PTE construction translates the given pixel-related features into tensors in the embedded space. The image segmentation into similar sets is achieved by clustering the tensors in the embedded space. For our image segmentation examples, we utilized pixel color information and its spatial (x,y) location multiplied by scaling factor w = 0.1. Hence, given an RGB image with I x I y pixels, we generated a 5 (I x I y ) dataset X. Algorithm 1 embeds X into a tensor space. The first step in Algorithm 1 constructs local patches. Each generated patch captures the relevant neighborhood and considers both color similarity and spatial similarity. Hence, a patch is more likely to include attributes related to spatially close pixels. It is important to note that the affinity kernel is computed according to Eq where ε equals the mean Euclidean distance between all the pairs in X. The PTE parameters l and ρ were chosen to generate the most homogenous segments. The k-means algorithm with sum of square differences was used to cluster the tensors into similar sets. Figures present the segmentation results from the application of the PTE algorithm, where for each figure, (a) is the original image. All of the images are of size except for the Sport (Fig ) image, which
94 76 (b) t = 1 (c) t = 2 (d) t = 3 (e) t = 4 (f) t = 5 (g) t = 6 (h) t = 7 Figure 3.7.1: The PTE segmentation results for the image Cubes when l = 10 and d = 10. The results are shown at several diffusion times t. Im- (a) Original age im- (a) Original age (b) t = 1 (c) t = 2 (d) t = 3 (e) t = 4 (f) t = 5 (g) t = 6 (h) t = 7 Figure 3.7.2: The PTE segmentation results for the image Hand when l = 10 and d = 10. The results are shown at several diffusion times t.
95 77 im- (a) Original age (b) t = 1 (c) t = 2 (d) t = 3 (e) t = 4 (f) t = 5 (g) t = 6 (h) t = 7 Figure 3.7.3: The PTE segmentation result for the image Sport when l = 10 and d = 17. The results are shown at several diffusion times t.
96 78 im- (a) Original age (b) t = 1 (c) t = 2 (d) t = 3 (e) t = 4 (f) t = 5 (g) t = 6 (h) t = 7 Figure 3.7.4: The PTE segmentation results for the image Fabric when l = 10 and d = 10. The results are shown at several diffusion times t. is of size Each figure describes the segmentation result at several diffusion times t. The impact of the diffusion time on the segmentation quality was significant for the Cubes (Fig ) and Fabric (Fig ) images. For example, as can be seen in Fig , the first two images (Fig (b) and Fig (c)), which correspond to t = 1 and t = 2 respectively, show poor segmentation qualities. As t increases, the segmentation becomes more homogeneous and the main structures in the original image can be separated as we see, for example, in (e) where t = 4. Another interesting aspect related to the diffusion time parameter t is the smoothing effect it has, when it increases, on the pairwise distances between data points in the embedded space. By increasing t, the pairwise distances between similar tensors decrease while the distances between dissimilar tensors increase. In the segmentation case, the result will be pixel-label change. For example, Fig presents the Cubes image segmentation as a function of t = 1, 2,..., 7. The rightmost cube in the segmented images becomes more homogeneous as t increases. 3.8 Conclusion In this chapter, we presented an extension of the scalar-affinity kernels that are used in kernel methods. We used mainly a linear-projection-based con-
97 struction of this extension, which we call a super-kernel. Other constructions such as ones based on orthogonal transformations can also be used. Such constructions will be explored in future works. The linear-projection diffusion (LPD) super-kernel that was introduced in this chapter is further explored in Chapter 4, which is based on [WA13]. There, its properties in the continuous case are examined and the generated diffusion process that propagates tangent vectors along the manifold is presented. This LPD process can also be utilized for patch-based dictionary constructions and out-of-sample extensions of vector fields [SBWA13a, SWB + 12, SWBA13]. Future works will present these utilizations together with a complete patch-processing data-mining framework that combines coarse-graining, dictionary-based subsampling, dimensionality reduction and smooth interpolation techniques. Among other benefits, the patch-processing approach introduced here will enable the reduction of wide redundancies in many large-scale datasets. It provides a meaningful representation of the essential intelligence from the analyzed data without any superfluous information that does not benefit the sought-after patterns and can thus be regarded as noise from the analysis point of view. 79 Appendix 3.A Technical proofs Lemma Let G be a LP super-kernel and let u R nd be an arbitrary vector of length nd. Then, u T Gu = m (ũ i ) T Ωũ i, where Ω is the affinity kernel, always holds. Proof. Let G be a LP super-kernel as was defined in Definition 3.3.1, and let u R nd be an arbitrary nd vector. The product u T Gu can be expressed block-wise as u T Gu = u T x G xy u y. (3.A.1) x M y M Vy using the definition of the blocks of G, we get therefore, i=1 u T x G xy u y = ω(x, y)u T x O T x O y u y = ω(x, y)ũ T x ũ y x, y M, (3.A.2) u T Gu = ω(x, y)ũ T x ũ y = x M y M x M ( ) m = ũ i xω(x, y)ũ i y = i=1 x M y M ω(x, y) y M m ũ i xũ i y i=1 (3.A.3) m (ũ i ) T Ωũ i, (3.A.4) i=1
98 80 as stated in the lemma. Lemma Let x, y M be two points on the manifold and let T x and T y be their embedded tensors (Eq ). If the embedding is done by using the spectral map Φ (Eq ) of a LP super-kernel G with the meta-parameter µ = 1 2, then G xy = T T x T y x, y M, where the tensors are treated as matrices (i.e., their coordinate matrices). Proof. Let x, y M be two points on the manifold, and let G be the LP super-kernel that is used to embed the points in the lemma. According to Eq (with µ = 1 T ), the elements of the matrix product T 2 x T y are [T T x T y ] ij = l [T x ] ξi [T y ] ξj = ξ=1 l ξ=1 λξ ϕ i ξ(x) λ ξ ϕ j ξ (y), i, j = 1,..., d. According to the spectral theorem, Eqs and 3.2.4, we have l λ ξ ϕ i ξ(x)ϕ j ξ (y) = ξ=1 l λ ξ φ ξ (o i x)φ ξ (o j y) = g(o i x, o j y) i, j = 1,..., d, ξ=1 thus, [T T x T y ] ij = g(o i x, o j y) = [G xy ] ij i, j = 1,..., d. Therefore, T T x T y = G xy as the lemma states. Lemma Let G be a diffusion super-kernel and let Φ be a spectral map (Eq ) of this kernel with the meta-parameter µ = 1. For every x, y M and j = 1,..., d, Φ(o j x) Φ(o j y) = g(o j x, ) g(o j y, ), where g(o j x, ) (or g(o j y, )) is a vector whose elements are g(o j x, o ξ z) (or g(o j y, o ξ z)), which are defined in Eq for every z M and ξ = 1,..., d. Proof. According to the definition of the spectral map Φ (Eq ) with µ = 1, Φ(o i x), Φ(o j y) = l λ 2 ζφ ζ (o i x)φ ζ (o j y) x, y M, i, j = 1,..., d. (3.A.5) ζ=1
99 81 We recall that {λ 2 ζ }l ζ=1 spectral theorem, are the eigenvalues of G2, thus, according to the l λ 2 ζφ ζ (o i x)φ ζ (o j y) = g(o i x, ), g(o j y, ) ζ=1 x, y M, i, j = 1,..., d, (3.A.6) since the right side of the equation is a cell in G 2. Therefore, Φ(o i x ) Φ(o j y) 2 = Φ(o i x), Φ(o i x) 2 Φ(o i x), Φ(o j y) + Φ(o j y), Φ(o j y) as stated in the lemma. = g(o i x, ), g(o i x, ) 2 g(o i x, ), g(o j y, ) + g(o j y, ), g(o j y, ) = g(o i x, ) g(o j y, ) 2,
100 82
101 Chapter 4 Linear-projection diffusion on smooth Euclidean submanifolds In order to process massive high-dimensional datasets, this chapter utilizes the underlying assumption that data on a manifold is approximately linear in sufficiently small patches (or neighborhoods of points), which are sampled with sufficient density from the manifold. Under this assumption, each patch can be represented (up to a small approximation error) by a tangent space of the manifold in its area and the tangential point of this tangent space. This chapter extends the results that were obtained in Chapter 3 (and in [SWA12]) for the finite construction of the LPD super-kernel by exploring its properties when it becomes continuous. Specifically, its infinitesimal generator and the stochastic process defined by it are explored. We show that the resulting infinitesimal generator of this super-kernel converges to a natural extension of the original diffusion operator from scalar functions to vector fields. This operator is shown to be locally equivalent to a composition of linear projections between tangent spaces and the vector-laplacians on them. We define a LPD process by using the LPD super-kernel as a transition operator while extending the process to be continuous. The obtained LPD process is demonstrated on a synthetic manifold. The results in this chapter appear in [WA13, SWAN12]. 4.1 Introduction A fundamental, well-based, assumption of kernel methods in general, and DM in particular, is that, locally, the manifold is approximately linear in sufficiently small patches (or neighborhoods of points). Under this assumption, each patch can, in fact, be represented (up to a small approximation 83
102 84 error) by a tangent space of the manifold in its area and the tangential point of this tangent space. Local PCA was suggested in [SW11, SW12] to compute an approximation of suitable tangent spaces and their tangential points for patches that define neighborhoods of points that are sampled with sufficient density from the manifold. An alternative method using a multiscale PCA algorithm was suggested in [LLJM09]. Using the suggested representations, the relations between patches can be modeled by the usual affinity between tangential points and an operator that translates vectors from one tangent space to another. The structure of the ambient space was utilized in Chapter 3 to define linear-projection operators between tangent spaces and utilize them to construct a super-kernel that represents the affinity/similarity between patches. The structure of the underlying manifold was utilized for a similar purpose in [SW12] to define continuous parallel transport operators between tangent spaces and to define such a super-kernel by using discrete approximations of these operators. In fact, algorithmically, the approximations in [SW12] are achieved by orthogonalization of the linear-projection operators in Chapter 3. Although these constructions differ by only a small modification of the construction algorithm, the resulting super-kernels have very different properties and different derived theories. The construction of a super-kernel via orthogonal transformations between tangent spaces, which yield discrete approximations of the parallel transport operator on the underlying manifold, was utilized in [SW12] to define a vector diffusion map. The vector diffusion map is defined by the constructed super-kernel in a manner similar to the diffusion map, which is defined by the diffusion kernel. The infinitesimal generator of the superkernel constructed in [SW12] converges to the connection-laplacian on the manifold. A variation of this super-kernel was also utilized in [SW11] to encompass information about the orientability of the underlying manifold. When the manifold is orientable, the resulting orientable diffusion map gives an orientation over the manifold (in addition to the embedded coordinates). When it is not orientable, the double-cover of the manifold can be computed using this method. In addition to defining the constructions of vector diffusion maps (VDM) and orientable diffusion maps (ODM), [SW12, SW11] utilized a local-pca process to approximate the tangent spaces that represent the analyzed patches on the manifold. The bounds of these approximation are thoroughly explored there and optimal values for the meta-parameters for this process are presented. In this chapter, we assume that these tangent spaces can be approximated, e.g., by methods similar to the one described in [SW12, SW11]. In this chapter, which is based on [WA13], we focus and extend the prop-
103 erties of linear-projection diffusion (LPD) super-kernels that were presented in Chapter 3, which is based on [SWA12]. These LPD super-kernels are a specific type of linear-projection super-kernels, whose spectra (i.e., eigenvalues) were shown to be non-negative. In case of LPD super-kernels, all the eigenvalues are between zero and one. This super-kernel was utilized in Chapter 3 (and in [SWA12]) to define an embedding of the patches on the manifold to a tensor space. The Frobenius distance between the coordinate matrices of the resulting tensors can be seen as an extension of the original diffusion distance, which was defined in [CL06a]. This extension includes both the data about the proximities of tangential points in the diffusion process and the projections between the corresponding tangent spaces that represent the patches (see Chapter 3 and [SWA12]). The results from Chapter 3 and [SWA12] are extended in this chapter by exploring the properties of the LPD super-kernel when it becomes continuous. We show that the resulting infinitesimal generator of this super-kernel converges to a natural extension of the original diffusion operator from scalar functions to vector fields. This operator is shown it be locally equivalent to a composition of linear projections between tangent spaces and the vector- Laplacians on them. We define a Linear-Projection Diffusion (LPD) process by using the LPD super-kernel as a transition operator while extending the process to be continuous. The chapter has the following structure. The manifold representation is defined in Section 4.2. The original diffusion operator, the resulting diffusion map and a natural extension of the diffusion operator to work on vectorfields are presented in Section 4.3. Section 4.4 describes the properties of the LPD diffusion operator. Specifically, its infinitesimal generator is explored in Section and the stochastic process defined by it is described in Section Finally, Section demonstrates the LPD process on a synthetic manifold Manifold representation Let M R m be a d-dimensional smooth Euclidean submanifold, which lies in the ambient space R m. For every point x M, the manifold M has a d-dimensional tangent space T x (M), which is a subspace of R m. Assume o 1 x,..., o d x T x (M) form a d-dimensional orthonormal basis of T x (M). These d vectors can also be regarded as vectors in the ambient space R m, thus, we can represent them by using m coordinates by a basis of R m. Assume that O x R m d is a matrix whose columns are these vectors represented by the
104 86 ambient coordinates O x o x o i x o d x x M. (4.2.1) We will assume from now on that vectors in T x (M) are expressed by their d coordinates according to the presented basis o 1 x,..., o d x. For each vector v T x (M), the vector ṽ = O x v R m is the same vector as v represented by m coordinates according to the basis of the ambient space. For each vector u R m in the ambient space, the vector u = O T x u T x (M) is the linear projection of u on the tangent space T x (M). 4.3 Naïve extensions of DM operators to vectorfield settings Chapter 1 briefly reviewed the DM method from [CL06a, Laf04]. In this chapter, we use a setting of DM that analyzes the geometry of the manifold M using the Gaussian affinities k(x, y) e x y ε, x, y M, where ε is a meta-parameter of the algorithm. These affinities are used to obtain a stochastic transition operator P f(x) = f(y)p(x, y)dy (f : M R), where p(x, y) = k(x, y) q(x) x, y M, (4.3.1) and q(x) k(x, y)dy, x M. M The transition operator P defines a Markovian diffusion process over the points on the manifold M. The diffusion affinity operator is defined as a symmetric conjugate to P, which is denoted by A and its elements are a(x, y) = k(x, y) = 1 q(x)p(x, y) x, y M. (4.3.2) q(x)q(y) q(y) The DM method embeds each data point x M onto the point Ψ(x) = (σ i ψ i (x)) δ i=0, where 1 = σ 0 σ 1... are the eigenvalues of A, ψ 0, ψ 1,... are their corresponding vectors, and δ > 0 is a sufficiently small integer. The exact value of δ depends on the decay of the spectrum of A and it determines the dimension of the embedded space Extended diffusion operator The original diffusion kernel operates on scalar functions. Its extension to vector fields, which are expressed in local coordinates, is not trivial, since the
105 local coordinates vary from point to point. However, in global coordinates (i.e., the coordinates of the ambient space R m ), a simple extension can be defined as P ν(x) = ν(x)p(x, y)dy, (4.3.3) where ν : M R m is a vector field expressed in the global coordinates of the ambient space. The relation between a tangent vector field ν : M R d, expressed in local coordinates and its corresponding global vector field ν : M R m is given by ν(x) = O x ν(x), x M. It should be noted that the vector field P ν, which results from the extended diffusion operator, is not necessarily a tangent vector field, i.e., the resulting vectors may not be tangent to the manifold at their assigned points. While the operator P might not be useful by itself (since its operation does not result in a tangent vector field), it does allow us to extend the infinitesimal generator of the diffusion kernel in a meaningful way. The infinitesimal generator of the diffusion kernel is defined by L(P ) lim ε 0, I P ε and it is shown in [CL06a] that if the manifold has a uniform density, it satisfies L(P ) =, where is the Laplace-Beltrami operator. If the density of the manifold is not uniform, a simple correction to the diffusion kernel can be used to maintain the same result. Therefore, for every function f : M R, L(P )f(x) lim ε 0 f(x) P f(x) ε = f(x). An extension of the described infinitesimal generator can be defined such that every vector field ν : M R m can be expressed in global coordinates of the ambient space to be L( P ν(x) ) ν(x) lim P ν(x). ε 0 ε The vector field ν can be defined by using m scalar functions that determine its (global) coordinates at any point on the manifold, i.e. ν(x) = ( ν 1 (x),..., ν m (x)) T. By using these functions, the extended infinitesimal generator takes the form L( P ) ν(x) = lim ε 0 ν 1 (x) P ν 1 (x) ε. lim ε 0 ν m(x) P ν m(x) ε 87 ν 1 (x) =., (4.3.4) ν m (x)
106 88 which resembles the vector Laplacian in Cartesian coordinates, where the Laplace-Beltrami operator replaces the standard Laplacian on each coordinate. In this chapter, we will show a different extension for the diffusion operator that uses the linear-projection operator to maintain the tangency of the vector fields on which this extension operates. We will show that the infinitesimal generator of both extensions are equivalent. 4.4 Linear-projection diffusion In this section, we extend the original diffusion operator (Eq ). This extended operator is introduced in Definition 4.4.1, which uses both the scalar values from Eq , which can be seen as transition probabilities between points on the manifold, and the linear-projection matrices between tangent spaces of the manifold, which are defined using the basis matrices from Eq Definition (LPD operator). Let ν : M R d be a smooth tangent vector field on M that assigns for each x M a vector ν(x) T x (M) represented in the d local coordinates of T x (M). A Linear-Projection Diffusion (LPD) operator G operates on such vector fields in the following way: G ν(x) = G xy ν(y)dy, where G xy p(x, y)o T x O y, x, y M. The LPD operator in Definition operates on tangent vector fields expressed in local coordinates (of the tangent spaces) and it results in tangent vector fields as well. Proposition shows that this operator is independent of the global coordinates of the ambient space, i.e., it does not change under a change of basis of the ambient space. This is intuitively reasonable since the linear projections, which are used to define it, depend only on the relations between tangent spaces (i.e., their local bases) of the manifold and the scalar components from the diffusion operator (Eq ) depend only on distances between points on the manifold (and not on the coordinates used to express these points). Proposition The LPD operator G is independent of the coordinates of the ambient space. Proof. Every change of basis in the ambient space R m is represented and defined by an orthogonal m m matrix. Assume that B is such a matrix.
107 Assume that O x, x M, are the matrices from Eq expressed in the original basis. Then, in the new basis (i.e., after a change was made), they are expressed by BO x, x M, thus, under the new basis we have G xy = p(x, y)(bo x ) T (BO y ) = p(x, y)o T x B T BO y = p(x, y)o T x O y x, y M, (4.4.1) where the d d matrices G xy are used in Definition to represent the LPD operator G. Therefore, the LPD operator G does not change under a change of basis of the ambient space. In fact, it is expressed by the same matrices G xy, x, y M, in every basis of the ambient space. A symmetric Linear-Projection Diffusion (LPD) super-kernel was constructed in Chapter 3 for a finite dataset of points sampled from an Euclidean submanifold. In the finite case, this super-kernel was a block matrix, where each block was defined by the diffusion affinities (see Eq in this chapter) and linear-projection matrices between the tangent spaces of the manifold. An extension of this construction to the continuous case is given by the symmetric conjugate Ĝ of the LPD operator. The continuous LPD super-kernel Ĝ is defined by its operation on the tangent vector field ν : M R d as Ĝ ν(x) = Ĝ xy ν(y)dy, where Ĝ xy = a(x, y)ox T O y = q(x)p(x, y)ox T 1 O y q(y) = 1 q(x)g xy x, y M. (4.4.2) q(y) 89 Therefore, the relation between the LPD operator G and the LPD superkernel Ĝ is similar to the one between the diffusion operator P (Eq ) and the diffusion affinity kernel A (Eq ). While the eigen-decompositions of operators that operate on a vector fields are not well-studied as the ones of operators that operate on scalar functions, in the finite case these operators become block matrices and their eigen-decompositions follow from these matrices. As a block matrix, the LPD super-kernel is positive semi-definite and all its eigenvalues are between zero and one. Since the LPD super-kernel is symmetric conjugate of the LPD operator, then, in the finite case, the spectrum of the LPD operator is also between zero and one. Therefore, for all practical purposes, the LPD operator, which is presented in this chapter, can be treated as positive semidefinite with all of its eigenvalues less than or equal to one. The eigenvectors of the LPD operator and the LPD super-kernel are related via conjugation in a manner similar to the relation between the original diffusion operator and
108 90 the affinity kernel. The reader is referred to [CL06a] for more information about these relations in the original diffusion-maps case Infinitesimal generator This section is devoted to the infinitesimal generator study of the LPD operator presented in Definition Theorem shows that this infinitesimal generator is equivalent to that of the extended diffusion operator presented in Section (specifically, by Eq ). Corollary uses this result to explain the resulting operator in terms of vector-laplacian operators on the tangent spaces of the manifold. Theorem Let P be an extended diffusion operator (Eq ), G be a LPD operator (Definition 4.4.1) and L( P ) and L(G) be the infinitesimal generators of these operators. In addition, let ν be a vector field expressed in the local coordinates of the tangent spaces and let ν be the same vector field expressed in global coordinates. Then, L( P ) ν(x) = O x L(G) ν(x) x M, where the matrices O x are defined in Eq , i.e., the infinitesimal generators of P and G are equivalent, where P and G operate in global and in local coordinates, respectively. Proof. The infinitesimal generator of P is L( P ) = limε 0 (I P )/ε. x M be an arbitrary point on the manifold, then Let L( P ν(x) ) ν(x) = lim P ν(x), (4.4.3) ε 0 ε where, by definition, P ν(x) = ν(y)p(x, y)dy. Since the tangent space T x (M) is a subspace of the ambient space R m, every vector ν(y) R m, y M, can be expressed as the sum of a vector on the subspace T x (M) and by a orthogonal vector to it. In other words, for every y M, we can define a vector ν (y) T x (M) (expressed in the global coordinates of the ambient space) and a vector ν (y) T x (M) such that ν(y) = ν (y) + ν (y). Therefore, Eq can be rewritten as L( P ν(x) ( ν (y) + ν (y))p(x, y)dy ) ν(x) = lim. (4.4.4) ε 0 ε Since T x (M) is a d-dimensional subspace of R m, its basis o 1 x,..., o d x can be treated as an orthonormal set of d vectors in R m. As such, this set can
109 be expanded with the m d additional orthonormal vectors b 1 x,..., b m d x T x (M) to form a basis for R m. Every vector in R m can be expressed by m coordinates c 1,..., c m R where c 1,..., c d correspond to o 1 x,..., o d x and c d+1,..., c m correspond to b 1 x,..., b m d x. Thus, a vector in T x (M) has c d+1 = c d+2... = c m = 0, while a vector orthogonal to T x (M) has c 1 = c 2... = c d = 0. From here on we will assume w.l.o.g. that these coordinates are the global coordinates used to express the vectors in the ambient space. According to the presented coordinate system, the vectors of ν take the form ν 1 (y) ν 1 (y).. 0. ν ν = d (y) ν = d (y) 0 + = ν ν d+1 (y) ν d+1 (y) (y) + ν (y) y M,. 0.. ν m (y) 0 ν m (y) 91 where ν 1,..., ν m are the coordinate functions of the vector field ν according to this system. Thus, we get 0. 0 ν (y)p(x, y)dy =. (4.4.5) νd+1 (y)p(x, y)dy. νm (y)p(x, y)dy Let us examine one of the nonzero coordinates in the vector ξ {d + 1,..., m} (Eq ). The function ν ξ is a scalar function. Thus, the results shown in [CL06a] can be applied to it. Specifically, the integration νξ (y)p(x, y)dy over the entire manifold can be approximated by the integration ν y x <ε ξ(y)p(x, y)dy on an open ball of radius ε around x on the manifold (i.e., the distance y x is a geodesic distance). Also, for a small enough ε, all the points in this ball are in the same coordinate neighborhood, where their coordinates can be expressed by the d orthogonal geodesics s 1,..., s d that meet at x. Every point y M in this ball (i.e., y x < ε in terms of geodesic distances) can be represented by a vector s y = (s y 1,..., s y d ) such that s y < ε. By using this representation, we can apply Taylor expan-
110 92 sion in this ball to the function ν ξ to get ν ξ (y) = ν ξ (x) + d j=1 ν ξ s j s y j + d i=1 d j=1 2 ν ξ s i s j s y i sy j +... sy < ε, y M. (4.4.6) Since ν(x) T x (M), the orthogonal component ν (x) is zero, thus ν ξ (x) = 0 and this term is canceled in Eq We combine the above arguments to get ν ξ (y)p(x, y)dy d ν ξ j s j j=1 and by canceling odd terms we get ν ξ (y)p(x, y)dy d s y j p(x, y)dy + d i=1 2 ν ξ s 2 i i=1 d j=1 2 ν ξ s i s j (s y i )2 p(x, y)dy. s y i sy j p(x, y)dy, According to [CL06a], the approximation error of this calculation is of order ε 2, or higher powers of ε, for a small enough meta-parameter 1 > ε > 0. In addition, since the integration can be taken to be within an open ball of radius ε, we can have (s y i )2 < ε 2, thus, for a small 1 > ε > 0, νξ (y)p(x, y)dy ε ε 2 γ ε = εγ, (4.4.7) where γ is the sum d 2 ν ξ i=1 i, which is a suitable constant coefficient for s 2 i bounding the approximation error. Combining Eqs and we get ν (y)p(x, y)dy ε m d εγ. Thus, when ε 0, the length of the vector ν (y)p(x, y)dy/ε becomes zero. Therefore, by using Eq we get L( P ν(x) ν (y)p(x, y)dy ν (y)p(x, y)dy ) ν(x) = lim + ε 0 ε ε ν(x) ν (y)p(x, y)dy = lim. (4.4.8) ε 0 ε Finally, we notice that for y M, the vector ν (y) is in fact the projection of ν(y) on T x (M) expressed in the global coordinates of the ambient space,
111 which is given by the matrix O x Ox T. Also, the relation between ν and ν, which expresses the same vector field in local and global coordinates respectively, is given by ν(y) = O y ν(y) for every y M. Therefore, we have L( P ) ν(x) = lim ε 0 ν(x) = lim ε 0 O x ν(x) = O x lim ε 0 ν(x) G ν(x) ε ν (y)p(x, y)dy ε Ox Ox T O y ν(y)p(x, y)dy ε = L(G) ν(x), and since x M was chosen arbitrarily, the equality is satisfied at every point on the manifold and the theorem is proved. Intuitively, the infinitesimal generator of an operator such as P or G considers the effects of the operator on the values of vector fields (i.e., vectorvalued functions) in infinitesimal neighborhoods on the manifold. In the case of P, the ambient directions of the vectors are not changed by the operator and the measured effects are determined by the scalar affinities (i.e., transition probabilities). The LPD operator G, however, also projects the vectors on the corresponding tangent spaces of the manifold so the vector field remains tangent to it. Thus, in the case of G, the measured effects are determined by both the scalar affinities (i.e., transition probabilities) and the curvatures of the manifold, which are intuitively manifested as the angles between its tangent spaces in the considered area. Thus, the difference between the two infinitesimal generators comes from the curvature component in the latter case. However, as the proof of Theorem shows, when only an infinitesimal area is considered, the manifold converges to its locally-linear nature and the distances (i.e., angles) between the tangent spaces of the manifold diminish and converge to zero. This result also gives an insight into the role of the scaling meta-parameter ε in the LPD construction. In scalar diffusion maps (and in the extended diffusion operator P ) it controls the sizes of the considered neighborhoods. In the LPD operator, it controls both these sizes and the effects of the curvatures that are taken in consideration. Smaller sizes of ε consider smaller neighborhoods and less effect of the curvatures, and when ε 0, neighborhoods converge to single points and the effects of the curvatures are canceled. Theorem shows that the LPD operator G maintains the same infinitesimal generator as the extended diffusion operator P while operating in local coordinates instead of global ones. This result shows that the LPD construction maintains, to some degree, the infinitesimal behavior (or nature) of the original diffusion operator and of the extended one. In the scalar 93
112 94 case, the infinitesimal generator of the diffusion operator can be expressed by Laplace operators (specifically, the graph Laplacian and the Laplace-Beltrami operator on manifolds). Corollary utilizes the relation shown in Theorem to provide an expression for the resulting infinitesimal generator using the vector-laplacian, which extends the Laplacian from scalar functions to vector fields. Corollary Let G be the LPD operator with the infinitesimal generator L(G). Let ν be a tangent vector field expressed by the local coordinates of the tangent spaces of the manifold M. Then, L(G) ν(x) = (proj x ν)(x) x M, where the operator proj x projects a vector field on the tangent space T x (M), and is the vector-laplacian on this tangent space. Proof. Let x M be an arbitrary point on the manifold and let ν expresses the tangent vector field ν by the ambient coordinates resulting from expanding the basis o 1 x,..., o d x of the tangent space T x (M) with additional m d orthonormal vectors, as was explained in the proof of Theorem Let ν 1,..., ν m be the coordinate functions of ν, where the first d vectors correspond to o 1 x,..., o d x and the rest correspond to the other m d vectors, which are orthogonal to T x (M). The projection of the vector field on T x (M) can now be written as proj x ν(y) = ( ν 1 (y),..., ν d (y)) T y M. (4.4.9) According to Theorem 4.4.2, we have and by using Eq we get O T x L( P ) ν(x) = O T x O x L(G) ν(x) = L(G) ν(x) L(G) ν(x) = ( ν 1 (x),..., ν d (x)) T, (4.4.10) where is regarded as the Laplace-Beltrami operator (at x) on the manifold or, equivalently, the Laplacian in a small open set on T x (M) around x whose points are related to those on the manifold via the exponential map. Using the second interpretation and by recalling Eq , the right-hand side expression in Eq is in fact the vector-laplacian (in Cartesian coordinates of T x (M)) at x of the projection of the vector field ν on the tangent space T x (M), as stated in the corollary.
113 4.4.2 Stochastic diffusion process In this section, we define the Brownian motion (BM) on a d-dimensional manifold in R m, d m. Let ũ : R M R m be a stochastic process on the manifold M, such that at time t 0 the process is at x = ũ(t 0 ) M. The d- dimensional manifold is defined locally at each point x M. Let Ũ M be a sufficiently small open set around x defined as Ũ = {z M x z < ζ} for a small ζ. We can choose ζ to be sufficiently small such that all the points in Ũ have the same coordinate neighborhood in M, and furthermore we can set it so that the coordinates in Ũ are given by the bijective exponential map exp x : U Ũ, where U T x(m) is the projection of Ũ on the tangent space T x (M) (see Fig ). Let t M be sufficiently small such that almost surely ũ(t) Ũ for every t (t 0 t, t 0 + t). Therefore, the stochastic process can be expressed, in the time segment T t (t 0 ) = (t 0 t, t 0 + t), by the local coordinates of U, i.e., we define the process u : T t (t 0 ) U such that for each t T t (t 0 ), it satisfies exp x (u(t)) = ũ(t). We define the Brownian trajectories of the local process u (and thus its global version ũ) by u(t 0 + τ) u(t 0 ) + u(τ) T x (M) τ < t, (4.4.11) where the transition vector u T x (M) is d-dimensional stochastic vector given by u(τ) B w τ < t, (4.4.12) where w N (0, τi) is a vector of d i.i.d. normal zero-mean random variables with variance τ, and B is a d d diffusion coefficients matrix. To stay on T x M, the vector u(τ) has to satisfy the orthogonality condition u(τ), n(x) = 0 where n(x) is the m-dimensional unit normal to T x (M). 95 Ũ y M (a) The set Ũ M and its projection U T x (M) on the tangent space of the manifold x M. U T x (M) x y (b) The exponential map exp x maps the points y Ũ M and y U T x (M) to each other. Figure 4.4.1: An illustration of an open set Ũ M around x M, its projection U T x (M) on the tangent space T x (M), and the exponential map exp x, which maps each point y Ũ on the manifold to y U on the tangent space T x (M).
114 96 The global process ũ can be discretized by setting a time unit τ < t and expressing the transition probabilities from x = ũ(t 0 ) to each possible y = ũ(t 0 + τ) by a probability distribution function p x : M [0, 1]. According to our choice of t, almost surely ũ(t 0 + τ) Ũ and therefore, since there is a bijection between U and Ũ (i.e., the exponential map exp x), a restriction of p x to Ũ should yield the transition probabilities of the local process u. In fact, the row-stochastic diffusion operator P (Eq ), with a suitable meta-parameter ε, defines such probability distributions by setting p x ( ) = p(x, ) [CL06a, Laf04]. The processes u and ũ represent transitions between points on the manifold. However, while the Brownian trajectories defined by ũ give points on the manifold itself, the points on Brownian trajectories defined by u are just approximations that lie on the tangent T x (M) (see Fig ). The exponential map exp x raises these approximations to lie on the manifold, thus providing the bijective relation between the local process u and the global process ũ. It was shown in [CL06a] that in a sufficiently small neighborhood around x, all quantities concerning the diffusion operator in Eq , and the resulting diffusion process, can be expressed in terms of the tangent space T x (M). This representation entails an infinitesimal approximation error that is canceled when the process becomes continuous in the limit τ 0 (ε 0 in terms of the diffusion operator). This result justifies our definition of the process ũ via its local approximation u. Further justification comes from examining the difference between the point y = ũ(t 1 ) and its tangential approximation y = u(t 1 ), t 1 = t 0 + τ. Since x, y T x (M), we have y x T x (M), and since y y = (y x) (y x) we get that y = y + ρ n(x), (4.4.13) where n(x) R m is the normal of T x (M) (as a subspace of R m ). Since the difference is in a direction orthogonal to the tangent space T x (M), it is bounded by the distance between x and y, and by the angle θ between the tangent space T x (M) and the vector y x, which goes from x to y. The distance between x and y is bound by the radius ζ of Ũ, which can be chosen to be as infinitesimally small (as long as t is set accordingly). Also, as y gets infinitesimally close to x, the angle between the vector y x and the tangent T x (M) vanishes, where the rate of the decrement is given by the curvature of the manifold M around x. Therefore, both error terms are canceled when the process becomes continuous (i.e., by taking ζ 0, t 0 and τ 0). Equation has d + 1 unknowns: the d local coordinates of u(t 1 ) and ρ, which is the Euclidean distance from y T x (M) to y M - see Fig To solve the system (4.4.13), we linearize it locally by setting
115 u(t 0 + τ) = u(t 0 ) + u and expanding everything to leading order in τ. We obtain ũ(t 0 ) + ũ u u + ρ n(x) = y + O( u 2 ) + O(ρ u), u=u(t0 ) and by using Eqs and , we get ũ u u + ρ n(x) = B w + O( u 2 ) + O(ρ u). (4.4.14) u=u(t0 ) The system (4.4.14) consists of m linear equations for the d+1 unknowns u and ρ. The term ρ n(x) can be dropped, because ρ ũ(t 1 ). For m = d+1 the Euler scheme for the BM on M becomes [ ] 1 ũ u(t + τ) = u(t 0 ) + B w. (4.4.15) u In the limit τ 0, ũ(t 1 ) converges in the usual way to a continuous trajectory on M. The PDF of ũ(t 1 ) satisfies the Laplace-Beltrami equation [CL06a, Laf04] on M. In addition to the processes defined by u and ũ, which govern the movement from one point to another on the manifold M, we define the vector functions v : T t (t 0 ) T x (M) and ṽ : R R d that define the propagation of a vector along the route determined by the diffusion process. Let v x = v(t 0 ) T x (M) be a tangent vector attached to the diffusion process at x in time t 0. In the discrete case, when the diffusion advances from time t 0 to time t 1 = t 0 + τ, it goes from x = u(t 0 ) (since the tangential point x is on both M and T x (M)) to y = u(t 1 ). Since this step is done entirely in the tangent space T x (M), we can propagate the vector v(t 0 ) to v(t 1 ) = v(t 0 ) without change, and thus we attach the same vector v x = v y T x (M) to the point y T x (M). However, when we move back to the manifold using the exponential map to get y = ũ(t 1 ) = exp x (y ) M, this vector cannot be directly propagated as-is to y M since v y / T y (M) (unless the manifold is flat). To deal with this problem, we use the linear projection operator Oy T O x and define ṽ y = ṽ(t 1 ) = Oy T O x v(t 1 ). Thus, at time t 1, the vector ṽ(t 1 ) consists of d-coordinates that represent the closest vector in T y (M) to v(t 1 ). The linear projection, which is used to transform the vector v y to ṽ y, does not preserve the length of the vector. In fact, the resulting vector becomes shorter. Eventually, at t, the vectors, which were propagated by this discrete process, will converge to 0. However, this is only a property of the discretization and not of the continuous case. Since ṽ y is the projection 97
116 98 of v y on T y (M), then ṽ y = vy cos θ, where θ is the angle between v y and T y (M). Also, θ is bounded from above by the angle between the tangent spaces T x (M) and T y (M). Therefore, smaller angles between tangent spaces yield less decrement of the length by the projection. In the continuous case, we can take y to be infinitesimally close to x. Therefore, the angle between their tangent spaces T x (M) and T y (M) gets infinitesimally small (where the rate of the decrement is given by the curvature of the manifold M), thus, θ 0 and ṽ y vy. We conclude the discussion in this section by providing a short summary of the LPD process properties. The transitions performed by this process, as discussed in this section and summarized below, are illustrated in Fig Summery of the LPD process properties Let G be a LPD operator with a sufficiently small ε such that if x, y M are not in the same neighborhood then p(x, y) 0 with infinitesimal approximation error. The operator G is a transition operator of a discrete stochastic process that propagates vectors along the manifold. Each d d block G xy = p(x, y)ox T O y describes a transition from the tangent vector v x T x (M) based at x M to the vector v y T y (M) based at y M. This discrete transition is done by the following steps: 1. A destination point y M is randomly chosen with probability p(x, y); y v y M T y (M) v y = O T y O x v x x v x x y T x (M) y y = exp x (y ) M T x (M) Figure 4.4.2: The jump of the LPD discrete process goes from time t 0 to time t 1 = t 0 + τ. The jump starts with a vector v x T x (M) that is attached to the manifold at x M. First, a point y T x (M) is chosen according to the transition probabilities of the diffusion operator P (Eq ). Then, the exponential map is used to translate this point to a point y M on the manifold. Finally, the vector v x T x (M) is projected to v y T y (M) and attached to the manifold at y M.
117 2. The direction and the length of the transition are represented by a vector u x y T x (M) from x to the projection of y on T x (M); 3. The vector u x y and the exponential map around x are used to perform the transition to y = exp x (x + u x y ) = x + u x y + ρ n(x), where n(x) is the normal of T x (M) in R m and ρ M is the distance of the projection from the manifold; 4. The vector v x (treated as a column vector) is projected on T y (M) to get vy T = vx T Ox T O y, thus v y = v x η n(y), where n(y) is the normal of T y (M) in R m and η M is determined by the length of v x and the angle it makes with T y (M); 5. The transition ends with the achieved tangent vector v y T y (M) at y M. As the process becomes continuous, ε 0, ρ 0 and η 0, thus the process remains on the manifold Linear-projection diffusion process demonstration To demonstrate the stochastic process described in Section 4.4.2, we implemented it on a two-dimensional paraboloid lying in a three dimensional Euclidean ambient space. We sampled 8101 points from the paraboloid defined by the equation z 3 = (z 1 /4) 2 + (z 2 /4) 2 for z = (z 1, z 2, z 3 ) R 3. We will refer to this paraboloid from now on as the manifold and denote it M. Assume x = (0, 0, 0) M and the vector (1, 1, 0) is tangent to the paraboloid M at the origin x. We will demonstrate the local and the global processes defined in Section by propagating this vector from x using the stochastic transitions of these processes along the points that were sampled from the paraboloid. In this case, the parametrization of the manifold is known, therefore there is no need to approximate the (known) tangent spaces of the manifold. We set the basis of the tangent space at x to be the vectors (1, 0, 0) and (0, 1, 0). The bases at every other point were set via parallel transport, which is computed using the known parameterization of the paraboloid, of the basis at x to each of the 8100 other sampled point. Once the bases of the tangent spaces 99
118 100 were calculated we constructed the matrices O y for every samples y M using Eq Finally, we computed the diffusion operator (Eq ) and constructed the LPD operator from Definition We use the constructed operator to perform the LPD process transitions and propagate the vector we set at x on the resulting trajectories. The stochastic nature of a single transition will be demonstrated first, and then the resulting trajectories will be demonstrated. In order to show the stochastic nature of a single transition, we performed 100 iterations that perform a single transition of the LPD process from x. The LPD transition, which was explained in details in Section 4.4.2, consists of two main phases. First, a transition of the local process u is performed on the tangent space T x (M). Then, the resulting point and its vector are projected on the manifold to show the transition of the global process ũ. The results from the iterations of the local transitions of u are presented in Fig The ones from the global LPD transitions of ũ are presented in Figs Figure 4.4.3: The results from performing 100 independent iterations of a single transition of the local process defined by u and v (see Section 4.4.2) from x M (in red), with a tangent vector (1, 1) in local coordinates of the tangent space T x (M). The starting point x is marked in red and the destinations of the transitions are marked in blue.
119 101 Figure 4.4.4: The results from performing 100 independent iterations of a single transition of the LPD process defined by ũ and ṽ (see Section 4.4.2) starting at x M with a tangent vector (1, 1) in local coordinates of the tangent space T x (M). The points in the area around x on the paraboloid M are presented. The starting point x is marked in orange, the destinations of the transitions are marked in red, and other points in this area are marked in blue. and These results demonstrate the locality of the transition, as well as its similarity to Brownian transitions over points on the manifold. Also, the vectors attached to the points on the manifold have similar magnitudes and directions while still remaining tangent to the manifold. After a single transition was demonstrated, we perform several iterations that generate a trajectory of the LPD process over the manifold. We will demonstrate two trajectories that were generated by this process. For this demonstration, we will only show the first 10 transitions of each trajectory. Figure shows these two trajectories on the manifold. Additional perspectives of the first trajectory are shown in Fig and the second one in Fig The vectors are propagated over the diffusion trajectory and they maintain similar directions and magnitude while remaining tangent to the manifold at their corresponding points. To see this more clearly, we projected each of the trajectories on the initial tangent space T x (M) at the starting point x. The projected trajectories are shown in Fig The LPD operator in Chapter 3, which generates the demonstrated stochastic process, was utilized for two data-analysis tasks. Specifically, it was utilized for classification of breast tissue impedance measurements to detect
120 102 (a) The area around x as seen in the ambient space. (b) The area around x is magnified here to see the directions of the tangent vectors Figure 4.4.5: Fig Two additional perspectives of the transitions shown in
121 103 (a) (b) Figure 4.4.6: Two independent trajectories of the LPD process defined by ũ and ṽ (see Section 4.4.2) starting at x M with the tangent vector (1, 1) in local coordinates of the tangent space T x (M). The starting point x is marked in orange. (a) (b) Figure 4.4.7: The projection of the trajectories in Fig on the tangent space T x (M) at the starting point x M of these trajectories. The starting point x is marked in orange. cancerous growth and for image segmentation. The latter application showed that various time-scales of the diffusion process provide different resolutions of the segmentation based on color shades and light levels. We refer the reader to Chapter 3 for more information on the implementation and on the applicative results of the LPD-based data analysis. 4.5 Conclusion The chapter enhances the properties of the linear-projection diffusion (LPD) super-kernels in Chapter 3 two-folds: 1. We showed that the infinitesimal generator of the LPD super-kernel
122 104 (a) The trajectory on the paraboloid as seen in the ambient space. (b) The area containing the trajectory is magnified here to show the propagation of the tangent vectors more clearly. Figure 4.4.8: Additional perspectives of the trajectory shown in Fig (a).
123 105 (a) The trajectory on the paraboloid as seen in the ambient space. (b) The area containing the trajectory is magnified here to see the propagation of the tangent vectors more clearly. Figure 4.4.9: Additional perspectives of the trajectory shown in Fig (b).
124 106 converges to a natural extension of the original diffusion operator from scalar functions to vector fields. This operator was shown to be locally equivalent to a composition of linear projections between tangent spaces and the vector-laplacians on them. 2. We introduced the stochastic process defined by the LPD super-kernels and demonstrated it on a synthetic manifold. Future research plans include: utilization of the presented LPD super-kernels methodology to provide out-of-sample extension, adapting large kernel-based methods to computing environments with limited resources by applying the patch-based methodologies that were described in Chapter 3 and in this chapter (based on [SWA12, WA13]) while processing real massive datasets.
125 Part II Diffusion-based Learning 107
126
127 Chapter 5 Coarse-grained localized diffusion Data-analysis methods nowadays are expected to deal with increasingly large amounts of data. Such massive datasets often contain many redundancies. One effect from these redundancies is the high-dimensionality of datasets, which is handled by dimensionality reduction techniques. Another effect is the duplicity of very similar observations (or data points) that can be analyzed together as a cluster. In this chapter, we propose an approach for dealing with both effects by coarse-graining the DM dimensionality reduction framework from the data-point level to the cluster level. This way, the size of the analyzed dataset is decreased by only referring to clusters instead of individual data points. Then, the dimensionality of the dataset can be decreased by the DM embedding. We show that the essential properties (e.g., ergodicity) of the underlying diffusion process of DM are preserved by the coarse-graining. The affinity that is generated by the coarse-grained process, which we call Localized Diffusion Process (LDP), is strongly related to the recently introduced Localized Diffusion Folders (LDF) [DA12] hierarchical clustering algorithm. We show that the LDP coarse-graining is in fact equivalent to the affinity-pruning that is achieved at each folder-level in the LDF hierarchy. The results in this chapter appear in [WRDA12]. 5.1 Introduction Massive high-dimensional datasets have become an increasingly common input for data-analysis tasks. When dealing with such datasets, one requires a method that reduces the complexity of the data while preserving the essential information for the analysis. One approach for obtaining this goal is 109
128 110 to analyze sets of closely-related data points, instead of directly analyzing the raw data points. A recent approach for obtaining such an analysis is the Localized Diffusion Folders (LDF) method [DA12]. This method recursively prunes closely-related clusters, while preserving the information about local relations between the pruned clusters. The DM framework [CL06a, CLL + 05] provides an essential foundation for LDF to succeed. This framework is based on defining similarities between data points by using an ergodic Markovian diffusion process on the dataset. The ergodicity of this process ensures it has a stationary distribution and numerically-stable spectral properties. The transition probability matrix of this process can be used to define diffusion affinities between data points. The first few eigenvectors of this diffusion affinity kernel represent the long-term behavior of the process and they can be used to obtain a lowdimensional representation of the dataset, in which the Euclidean distances between data points correspond to diffusion distances between their original (high-dimensional) counterparts. We present a coarse-graining of this diffusion process, while preserving its essential properties (e.g., ergodicity). We show that this coarse-graining is equivalent to the pruning method that appeared in the LDF. The LDF method performs an iterative process that obtains a folder hierarchy that represents the points in the dataset. Each level in the hierarchy is constructed by pruning clusters of folders (or data points) from the previous level. The iterative process has two main phases in each iteration: 1. Clustering phase: the shake & bake method is used to cluster the folders (or data points) of the current level in the hierarchy by using a diffusion affinity matrix. 2. Pruning phase: the clusters of the current level are pruned and given as folders of the next level in the hierarchy. The diffusion affinity is also pruned to represent affinities between pruned clusters (i.e., folders of the next level in the hierarchy) instead of folders in the current hierarchial level. In this chapter, which is based on [WRDA12], we focus on exploring the pruning that is performed in the second phase of this process. We consider the clustering of the data, which may be performed by shake & bake process [DA12] or by another clustering algorithm, as prior knowledge. Essentially, LDF provides an hierarchical data clustering with additional affinity information for each level in the hierarchy. Other examples of hierarchical clustering methods can be found in [ZRL96, KR90]. However, these methods are not related to DM and to its underlying diffusion process. Since
129 111 we are mainly concerned with the pruning phase of the algorithm, the clustering aspect of LDF and its relation with these methods is beyond the scope of this chapter. A detailed survey of clustering algorithms and their relation to LDF is provided in [DA12, Section 2]. While there are many empirical justifications for the merits of LDF and its utilization in various fields (e.g., unsupervised learning and image processing), it lacked theoretical justifications. In this chapter, we introduce a coarse-graining of the underlying diffusion process of DM. The resulting coarse-grained process, which we call Localized Diffusion Process (LDP), preserves essential properties of the original process, which enable its utilization for dimensionality reduction tasks. We relate this process, or rather the diffusion affinity generated by it, to the one achieved by the LDF pruning phase. This relation adds the needed complimentary foundations for the LDF framework by providing theoretical justifications for its alreadyobtained empirical support. Additionally, the presented relation shows that the applications presented in [DA12] in fact demonstrate the utilization of the LDP for data-analysis tasks and the results presented there provide empirical support of its benefits. A similar coarse-graining approach was presented in [LL06]. The approach there is based on a graph representation of the diffusion random-walk process. The clustering of data points was performed by graph partitioning. Then, transition probabilities between partitions were achieved by averaging transition probabilities between their vertices. The resulting random-walk process maintains most of the spectral properties of the original diffusion process and its eigendecomposition can be approximated by the original spectral decomposition. However, the approximation error strongly depends on the exact partitioning used. In addition, since all the random-walk paths are considered in the averaging process, there is a limited number of viable time-scales (in the diffusion process) that can be used by this process before it converges to the averaging of the stationary distribution. The presented coarse-graining process in this chapter copes with the rapid convergence toward the stationary distribution by only preserving localized paths between clusters while ignoring paths that are global from the cluster point-of-view. While it is desirable that the clusters will be sufficiently coherent to consist of a continuous partitioning of the dataset and its underlying manifold, the properties of the presented coarse-graining process are neither depend on such assumptions nor on the exact clustering method used. An alternative approach for local sets considerations of data points is to analyze them as patches on the underlying manifold of the dataset [SWA12, WA13, SW11, SW12]. The relations between patches are represented by nonscalar affinities that combine the information about both geodesic proximity
130 112 of the patches and the alignment of their tangent spaces. This approach was used in [SW11] to modify DM to preserve the orientation of the manifold through the embedding process. A more comprehensive utilization of this approach was presented in [SWA12] and [SW12], where affinities between patches were defined as matrices that transform vectors between tangent spaces. Parallel transport operators were used in [SW12], with the resulting affinity block matrix being related to the connection-laplacian. Linearprojections were used in [SWA12] and further explored in [WA13], where the resulting diffusion process was shown to propagate tangent vectors on the manifold. The results from [SWA12] and [WA13] are presented in Chapters 3 and 4 (accordingly) of this thesis. Both discussed methods in [SWA12, SW12] lead to an embedded tensor space instead of a vector space. Also, the resulting diffusion process is not necessarily ergodic and may not have a stationary distribution. Therefore, they do not preserve one of the crucial properties of the diffusion process used in DM. The approach used in this chapter produces a scalar-affinity matrix between closely-related clusters of data points. It neither depends explicitly on the existence nor on the knowledge of the (usually unknown) underlying manifold of the dataset. The resulting diffusion process is similar to the one used in DM (for data points), and it preserves the essential properties of that diffusion process. Finally, the same spectral analysis, which is performed in DM, can be used to obtain an embedding that is based on the coarse-grained process presented here, which results with an embedding of clusters to vectors (and not tensors). The chapter has the following structure. The problem setup is described in Section 5.2. Specifically, the DM method is discussed in Section and the LDF method is discussed in Section Section 5.3 introduces the localized diffusion process (LDP), which is the main construction in this chapter. The pruning algorithm for constructing the LDP is presented in Section Finally, the strong relation between LDF and LDP is presented in Section Problem Setup Let X R d be a dataset of n data points that are sampled from a low dimensional manifold that lies in a high dimensional Euclidean ambient space. Assume the data consists of n coherent disjoint clusters, which correspond to dense local neighborhoods that were generated by an affinity kernel. Assume that C 1, C 2,..., C n are these clusters in the underlying manifold, where
131 113 X = n i=1c i and C i C j = for i j {1, 2,..., n}. Assume that maps each data point x X to its cluster C(x). C : X {C 1, C 2,..., C n } (5.2.1) Remark about matrix notation: In this chapter, we will deal with several matrices that represent relations between data points or clusters of data points. Let M be such a matrix where every row and column of M corresponds to a data point in the dataset X or a subset of this dataset. It is convenient in this case to use the lowercase notation m(x, y) to denote the cell in the x s row and the y s column in M. For t Z, the notation m t (x, y) denotes cells in M t, which is the t-th power of M. Similar notation will also be used for matrices with rows and columns that correspond to clusters of data points Stochastic view of DM The DM methodology [CL06a, Laf04], which was briefly reviewed in Chapter 1, is based on constructing a Markovian diffusion process P over a dataset. This process essentially defines random walks over data points in the dataset. It consists of paths between these data points, where each path P P is a series of transitions (steps on the data points), denoted by P 0 P 1... P l, where P 0, P 1,..., P l X, l 1. Each path has a probability, which is defined by the probabilities of its transitions and will be discussed later. The length of the path P, denoted by len(p) = l, is its number of transitions. The source (i.e., the starting data point) of the path is denoted by s(p) = P 0 and its destination is denoted by t(p) = P len(p) = P l. When only paths of specific length l = 1, 2,..., are considered, the notation P P l will be used to denote that P P and len(p) = l. For example, a single transition in P is a path of unit length P P 1. In order to assign transition probabilities between data points, an n n affinity kernel K is defined on the dataset. Each cell k(x, y), x, y X, in this kernel represents similarity, or proximity, between data points. A popular affinity kernel is the isotropic Gaussian diffusion kernel k(x, y) = exp( x y /ε) with a suitable ε > 0. An alternative kernel, which is based on clustering patterns in the dataset, is the shake-and-bake kernel [DA12] that will be discussed in more details in Section Next, a degree matrix Q is defined as a diagonal matrix whose main diagonal holds the degrees q(x, x) = q(x) y X k(x, y), x X. Kernel normalization by these degrees yields a row-stochastic matrix P Q 1 K that defines the transition
132 114 probabilities p(x, y) = k(x,y), x, y X, between data points. These transition q(x) probabilities define the Markovian diffusion process P over the dataset. The diffusion process P specifies the probability of moving from one data point to another via paths of any given integer length l 1. We denote this probability by Pr[x Pl y] Pr[t(P) = y s(p) = x P P l ] x, y X. (5.2.2) Since the diffusion process is a Markovian process with single-transition probabilities defined by P, Eq becomes Pr[x Pl y] = p l (x, y) x, y X, l = 1, 2,..., where in particular Pr[x P1 y] = p(x, y). The diffusion process P is an ergodic Markov process. This means that P has a stationary distribution in the limit l of the path lengths. Spectral analysis of this kernel yields a decaying spectrum 1 = λ 0 λ 1 λ , where λ i, i = 0, 1, 2,..., are the eigenvalues of P. When an isotropic Gaussian kernel is used, the decay of the spectrum can be used to approximate the intrinsic dimension of underlying manifold of the dataset. This relation between the decay of the spectrum and the underlying geometry of the data was discussed in [CL06a] and in Chapter 2 of this thesis. Dimensionality reduction can be achieved by spectral analysis of P or, more conveniently, its symmetric conjugate A = Q 1/2 P Q 1/2 that is referred to as the diffusion-affinity matrix. Let φ 0, φ 1, φ 2,... be the eigenvectors of A that correspond to the eigenvalues λ 0, λ 1, λ 2,... (conjugation maintains the same eigenvalues of A), then the DM embedding is defined as x Φ(x) ( λ 0 φ 0 (x), λ 1 φ 1 (x), λ 2 φ 2 (x),...) T x X. A subset of these coordinates can be used by ignoring the eigenvectors with sufficiently small eigenvalues, which will anyway result with approximatelyzero embedded coordinates. A simple coarse-graining of the original diffusion process can be done by cluster pruning while defining transition probability between two clusters by considering all the paths between them. However, due to the decay of the diffusion kernel s spectrum, this method will converge fast (especially when applied several times) to the stationary distribution of the diffusion process. An alternative coarse graining method, which excludes paths that are considered global from clusters point-of-view, will be presented in Section 5.3. One aspect of any diffusion process coarse-graining is to translate a data point terminology to a cluster terminology. This aspect must be addressed
133 115 regardless of the paths that are considered when computing the transitional probabilities between clusters, since any path in the diffusion process is defined in terms of data points. The probability of reaching every data point on a path is determined by its starting data point and by suitable powers of P. Since the clusters are disjoint, these probabilities can be easily interpreted as the probability of reaching a destination cluster. Specifically, it can be done by using the function C (Eq ) and by summing the appropriate probabilities. Paths that start in a source cluster, denoted by s(p) C i, P P, i = 1,..., n, require a nontrivial interpretation in terms of a source data point s(p) = x C i. This interpretation should be defined by probability terms. We will use the same intuition that was used to construct the transitional probability matrix P in order to define the probability Pr[s(P) = x C i s(p) C i ], i = 1,..., n, P P. The kernel K was interpreted as weighted adjacencies of a graph whose vertices are the data points in X. According to this interpretation, the degree of each data point x X is a sum of the edges weights k(x, y), y X that begin at x. To measure the occurrence probability of the transition x y when starting at x, the weight of the edge (x, y) is divided by the total weight of the edges starting at x, which gives the probability measure p(x, y) = k(x, y)/q(x). Assume the volume of the cluster C i, i = 1,..., n, is defined as vol(c i ) x C i q(x). Therefore, the volume of a cluster is the total sum of the degrees of the data points in this cluster, which is the sum of the weights of all the edges that start in C i. According to the same reasoning as before, the occurrence probability of the transition x y, x C i, y X, which started at the cluster C i, is k(x,y). vol(c i ) Therefore, the transition probability, which starts at C i to actually starts at a specific data point x C i, is Pr[s(P) = x C i s(p) C i ] = y X k(x, y) vol(c i ) = q(x) vol(c i ), (5.2.3) because the transitions to different designated data points are independent events. Notice that the choice of the first transition in a path is independent of its length. Thus, the presented probability is independent of the length of the path P and the assumption that P P l for some l 1 does not affect it Localized diffusion folders (LDF) As described in the DM brief overview in Section 5.2.1, P is the affinity matrix of the dataset and it is used to find the diffusion distances between data
134 116 points. This distance metric can be used to cluster data points according to the diffusion distances propagation that is controlled by the time parameter t. In addition, it can be used to construct a bottom-up hierarchical data clustering. For t = 1, the affinity matrix reflects direct connections between data points. These connections can be interpreted as local adjacencies between data points. The resulting clusters preserve the local neighborhood of each data point. These clusters are the bottom level in the hierarchy. By raising t, which means time advancement, the affinity matrix is changed accordingly and it reflects indirect rare connections between data points in the graph. The diffusion distance between data points in the graph accounts for all possible paths of length t between these data points at a given time step. The more we advance in time the more we increase indirect and global connections. Therefore, by raising t we can construct the upper levels of the clustering hierarchy. In each time step, it is possible to merge more and more low-level clusters since there are more and more new paths between them. The resulting clusters reflect global neighborhood of each data point that is highly affected by the advances of the parameter t. The major risk in this global approach is that increasing t will also increase noise, which is classified as connections between data points that are not closely related in the affinity matrix. Moreover, clustering errors in the lower levels of the hierarchy will diffuse to the upper levels of the hierarchy and hence will significantly affect the correctness of the upper levels clustering. As a result, some areas in the graph, which are assumed to be separated, will be connected by the new noise-result and error-result paths. Thus, erroneous clusters will be generated (a detailed description of this situation is given in [DA12]). This type of noise significantly affects the diffusion process and eventually the resulting clusters will not reflect the correct relations among the data points. Although these clusters consist of data points that are adjacent according to their diffusion distances, the connections among these data points in each cluster can be classified as too global and too loose that generate inaccurate clusters. A hierarchical clustering method of high-dimensional data via the localized diffusion folders (LDF) methodology is introduced in [DA12]. This methodology overcomes the problems that were described above. It is based on the key idea that clustering of data points should be achieved by utilizing the local geometry of the data and the by local neighborhood of each data point and by constructing a new local geometry every advance in time. The new geometry is constructed according to local connections and according to diffusion distances in previous time steps. This way, as we advance in time, the geometry from the induced affinity reflects better the data locality while the affinity noise in the new localized matrix decreases and the accuracy
135 117 of the resulting clusters is improved. LDF is introduced to achieve the described local geometry and to preserve it along the hierarchical construction. The LDF framework provides a multilevel partitioning (similar to Voronoi diagrams in diffusion metric) of the data into local neighborhoods that are initiated by several random selections of data points or folders of data points in the diffusion graph and by defining local diffusion distances between them. Since every different selection of initial data points yields a different set of diffusion folders (DF), it is crucial to repeat this selection process several times. The multiple system of folders, which we get at the end of this random selection process, defines a new affinity and this reveals a new geometry in the graph. This localized affinity is a result of what is called the shake & bake process in [DA12]. First, we shake the multiple Voronoi diagrams together in order to get rid of the noise in the original affinity. Then, we bake a new cleaner affinity that is based on the actual geometry of the data while eliminating rare connections between data points. This affinity is more accurate than the original affinity since instead of defining a general affinity on the graph, we let the data define its localized affinity on the graph. In every time step, this multi-level partitioning defines a new localized geometry of the data and a new localized affinity matrix that is used in the next time step. In every time step, we use the localized geometry and the LDF that were generated in the previous time step to define the localized affinity between DF. The affinity between two DF is defined by the localized diffusion distance metric between data points in the two DF. In order to define this distance between these DF, we construct a local sub-matrix that contains only the affinities between data points (or between DF) of the two DF. This sub-matrix is raised to the power of the current time step (according to the current level in the hierarchy) and then it is used to find the localized diffusion distance between the two DF. The result of this clustering method is a bottom-up hierarchical data clustering where each level in the hierarchy contains DF of DF from lower levels. Each level in the hierarchy defines a new localized affinity (geometry) that is dynamically constructed and it is used by the upper level. This methodology preserves the local neighborhood of each data point while eliminating the noisy connections between distinct points and areas in the graph. In summary, [DA12] deals with new methodologies to denoise empirical graphs. Usually, in applications data is connected through spurious connections. One of the goals of [DA12] is to introduce a notion of consistency of connections in order to repair a noisy network. This consistency is achieved through the construction of a forest of partition trees, which redefine the connectivity in the network. This opens the door to robust processing of
136 118 data clouds in which group consistency is exploited. 5.3 Localized Diffusion Process In this section, we present a coarse-graining diffusion process between clusters in a dataset. The transitions between clusters, which are considered as vertices in this process, will be defined by certain paths in the original diffusion process. Definition introduces the notion of a localized path, which will be used to define these transitions. Then, in Definition 5.3.3, these localized paths will be used to define the localized diffusion process between clusters. Definition (localized l-path). A localized l-path in a diffusion process P is the path P P l of length l that traverses solely through data points in its source and destination clusters, i.e., P 0, P 1,..., P l C(s(P)) C(t(P)). The difference between localized and non-localized paths is demonstrated in Fig The path in Fig (a) traverses through data points in its source cluster, then passes via a single transition to its destination cluster and then traverses in it to its destination data point. Therefore, it does not pass through any cluster other than its source and destination clusters and thus it is localized. On the other hand, the non-localized path in Fig 5.3.1(b), traverses through a third intermediary cluster, thus it is not localized. Notice that a localized path does not necessarily contain a single transition between its source and destination clusters. Figure illustrates two (a) A localized path. (b) A non-localized path Figure 5.3.1: Illustration of the difference between localized and non-localized paths
137 119 (a) Localized path with identical source and destination clusters (b) Localized path with multiple transitions between the source and destination clusters Figure 5.3.2: Illustration of non-trivial localized paths such non-trivial paths. A path that traverses solely in a single cluster (see Fig (a)) is in fact a localized path from the cluster to itself. A localized path can also alternate between its source and destination clusters a few times before reaching its final destination, as shown in Fig (b). As long as the cluster involves only its source and destination cluster/s (whether they are identical or not) without passing through any intermediary cluster, then it is considered to be localized. We denote the set of all localized l-paths in the diffusion process P by L(P l ) P l. The usual diffusion transition probabilities between data points in a dataset via paths of a given length l 1 were described in Section These probabilities consider all the paths of length l between two data points. The construction presented in this chapter only considers localized paths and ignores other paths, which are considered global from a cluster pointof-view. Therefore, we define the localized transition probabilities, which describe the probabilities of a transition from x C i to y C j, i, j = 1,..., n, via localized l-paths, as Pr[x L(Pl ) y] Pr[t(P) = y P L(P l ) s(p) = x P P l ]. (5.3.1) Similarly, the localized transition probability from x C i to the cluster C j is defined as Pr[x L(Pl ) C j ] Pr[t(P) C j P L(P l ) s(p) = x P P l ]. (5.3.2) Finally, the localized transition probability from the cluster C i to the cluster C j is defined as Pr[C i L(P l ) C j ] Pr[t(P) C j P L(P l ) s(p) C i P P l ]. (5.3.3)
138 120 The original transition probabilities are clearly related to the diffusion operator P via Eq The defined localized transition probabilities do not have such a direct relation with the diffusion operator. They will be further explored in Section The localized transition probabilities in Eq consider localized paths from a source cluster to a destination cluster. Not all the paths from the source cluster are localized. Therefore, only a portion of the paths from a given cluster (to any other cluster) are actually viable for consideration with these probabilities. Definition provides a measure for the portion of viable paths going out from a cluster from all the paths starting in it. Definition (l-path localization probability). The l-path localization probability (lpr) of a cluster C i, i = 1,..., n, is lpr l (C i ) = Pr[p L(P l ) s(p) C i P P l ]. It is the probability that a path of length l, which starts at this cluster, is a localized path. Definition uses the defined localized paths in the original diffusion process to define a localized diffusion process between clusters. Definition (l-path localized diffusion process). Let P be a diffusion (random walk) process defined on the data points of the dataset X. An l-path localized diffusion process P is a random walk on the clusters C 1, C 2,..., C n where a transition from C i to C j, i, j = 1,..., n, represents all the localized l-paths in the diffusion process P from data points in C i to data points in C j. The probability of such a transition, according to P, is the probability to reach the destination cluster C j when starting at the source cluster C i and traveling solely via localized l-paths. The l-path localized diffusion process is a Markovian random-walk process. Thus, its transition probabilities are completely governed by its singlestep transition probabilities. These probabilities can be computed, by definition, according to the transition probabilities of the original diffusion process. Using notations similar to the ones used for the original diffusion process, we get the single-step transition probabilities P1 Pr[C i C j ] Pr[t(P) C j s(p) C i P L(P l )], i, j = 1,..., n, (5.3.4) for the l-path localized diffusion process P. Notice that these differ from the probabilities in Eq The former considers the term P L(P l ) in
139 121 the hypothesis part, since it considers only the localized paths of the original diffusion process. The latter considers this term in the condition part, since it computes the probability over all the paths in the original diffusion. Ergodicity is one of the main properties in the diffusion process that is used by DM [CL06a]. Ergodicity means that the eigenvalues of P have a magnitude of at most one, and therefore its spectrum decays with time as a function of the numerical rank of the transitional kernel. As we advance the diffusion process in time, it converges to a stationary distribution and therefore its long term state can be represented by a low-dimensional space. Proposition shows that the coarse graining suggested here preserves this property, i.e., the l-path localized diffusion process is ergodic and its transition matrix has a decaying spectrum. Proposition The localized diffusion process P, which is defined by Definition 5.3.3, is an ergodic Markov process. Proof. According to [CL06a], the original diffusion process P is aperiodic and irreducible. We will show that P is also aperiodic and irreducible process. The Ergodicity follows from these properties. From the aperiodicity of P we have p(x, x) > 0 for every x X. Let P P l be a path with P i = x, i = 0,..., l. Obviously, this path is a localized l-path from C(x) to itself. The probability of this path is (p(x, x)) l > 0. P Therefore, the transition probability Pr[C(x) l C(x)], which sums the probabilities of all the localized l-paths from C(x) to itself, must be nonzero. This argument holds for every x C i X, i = 1,..., n, and thus holds for every cluster C i = C(x). Therefore, the process P is aperiodic. Due to the irreducibility of the original diffusion process P, there exists a path P P with nonzero probability between every pair of data points x y X. For each transition P i P i+1, i = 0,..., len(p) 1 in this path, the following localized l-path P = P i P i... P }{{} i P i+1, l 1 transitions between C(P i ) and C(P i+1 ) is constructed. Due to the aperiodicity of P, the first l 1 transitions have nonzero probability. The last transition of P is the same transition P i P i+1 from P, which also has nonzero probability. Therefore, the path P is a localized l-path between C(P i ) and C(P i+1 ) with nonzero probability. This holds for every transition in P and thus the transition probability from C(P i ) to C(P i+1 ) via the localized l-paths is nonzero for each i = 0,..., len(p) 1. Thus, the path C(x) = C(P 0 ) C(P 1 )... C(P len(p) ) = C(y)
140 122 has nonzero probability in P. Since x, y were chosen arbitrarily, this holds for every pair of clusters and thus P is irreducible. Together with the aperiodicity that was shown above, P is ergodic. In this section, we introduced a coarse-grained diffusion process (i.e., the l-path localized diffusion process) that preserves the crucial properties of the DM. A coarse-graining algorithm, which constructs this process, is presented in Section In Section 5.3.2, we will show that this process is directly related to the construction presented in [DA12]. Specifically, the presented coarse graining is related to the pruning done at the transition between levels in the LDF hierarchy Pruning algorithm Algorithm 2: transition matrix pruning Input: Dataset X of n data points; Clustering function C : X {C 1, C 2,..., C n } of the data into n clusters; Transition probability matrix P between data points in X; Parameter l of the path length. Output: A row-stochastic n n matrix P that represents transitions between clusters; A diagonal matrix Q that contains the degrees of the clusters on its diagonal. foreach i, j = 1,..., n do P C i C j C i C j square matrix ; // Denote by p(x, y) the cell of P in the row of x // and the column of y (x, y C i C j ). // Denote by p l (x, y) the same cell in P l. foreach x, y C i C j do p(x, y) p(x, y) ; end K ij x C i,y C j q(x)p l (x, y) ; end foreach i = 1,..., n do Q ii n j=1 K ij ; end P Q 1 K ;
141 123 P C i C j C i C j P C i {}}{ C j {}}{ }Ci } C j (a) Construction of the submatrix P from the matrix P P l P l C j {}}{ }Ci Kij (b) Computation of K ij as a weighted sum of cells in P l Figure 5.3.3: Construction of the pruned-kernel Kij between the clusters C i and C j The l-path localized diffusion process described in Section 5.3 is a coarsegrained version of the original diffusion process. As a Markovian process, it defines a transition probability matrix between clusters. Algorithm 2 shows how to construct this transition probability matrix, denoted by P, based on the transition probability matrix of the original diffusion process. In addition to P, the algorithm outputs the degree matrix Q that holds the degrees of the clusters on its diagonal. Theorem shows that the resulting matrix P defines a localized diffusion process. Algorithm 2 performs a coarse-graining of the original diffusion process by pruning the clusters into vertices of a Markovian random-walk process. The transition probabilities of this process are determined by the row-stochastic transition matrix P. For each pair of clusters, C i and C j, the algorithm considers the sub-matrix P of P, which contains only rows and columns of data points in C i C j (see Fig (a)). The algorithm then calculates the affinity, denoted by K ij, between the clusters C i and C j. First, It raises the sub-matrix P to the l-th power in order to generate the l-path localized tran-
142 124 sition probabilities between points in C i and C j. Then, the affinity between these clusters is a weighted sum of the elements in P l, where the weight of the element p l (x, y) is q(x) (see Fig (b)). Finally, the degree of each cluster is calculated by summing its affinities Q ii = n K j=1 ij with all the clusters. Notice that Algorithm 2 is similar to the pruning algorithm described in [DA12, Section 3.3]. Both algorithms get an input matrix of relations between data points and a clustering function that assigns each point to its cluster. Then, for each pair of clusters, these algorithms consider a sub-matrix that contains the relations between the data points in the two considered clusters. In order to achieve a scalar representation of the relation between the considered clusters, both algorithms aggregate the elements of a suitable power of the considered sub-matrix. However, these algorithms differ in the input matrix itself and in the aggregation function that is being used. Algorithm 2 gets a transition probability matrix as an input and uses a weighted sum for the aggregation, while the algorithm in [DA12, Section 3.3] gets an affinity matrix as an input and suggests three different aggregations of the sub-matrix elements. Theorem shows that the resulting Markov process in Algorithm 2 is in fact a transition probability matrix of the l-path localized diffusion process that was defined in Definition Theorem The output matrix P from Algorithm 2 is a transition probability matrix of an l-path localized diffusion process. In order to prove Theorem 5.3.2, we need Lemmas and that relate the matrices used in Algorithm 2 to the original diffusion process. Lemma Let P be the sub-matrix of P defined in a single iteration of Algorithm 2 for specific i, j = 1,..., n. P is related to the localized transition probabilities of P in the following ways: 1. p l (x, y) = Pr[x L(Pl ) y]; 2. y C j p l (x, y) = Pr[x L(Pl ) C j ]; 3. x C i y C j q(x) vol(c i ) pl L(P (x, y) = Pr[C l ) i C j ]. The proof of Lemma is given in 5.A. Lemma Let q(c i ) Q ii n j=1 K ij, i = 1,..., n, be the degree (i.e., row sum) defined in Algorithm 2. Then, q(c i ) = vol(c i ) lpr l (C i ).
143 125 Proof. According to Definition lpr l (C i ) = Pr[p L(P l ) s(p) C i P P l ] = n j=1 Pr[C i L(P l ) C j ], Combining with property (3) of Lemma yields i = 1,..., n. lpr l (C i ) = n j=1 x C i y C j q(x) vol(c i ) pl (x, y) i = 1,..., n, where P depends on the choice of i and j. By using the matrix K from Algorithm 2 we get lpr l (C i ) = n j=1 K ij vol(c i ) = q(c i) vol(c i ) i = 1,..., n, and multiplying by vol(c i ) yields the desired result. Lemmas and relate the localized transition probabilities from the diffusion process to the original diffusion transition probabilities via the matrices constructed in Algorithm 2. These relations can now be used to prove Theorem Proof of Theorem Consider two clusters C i and C j, i, j = 1,..., n. According to Algorithm 2, Pij K ij Q ii and K ij x C i,y C j q(x)p l (x, y). Using Lemmas and we obtain P ij = vol(c i) Pr[C i L(P l ) C j ] vol(c i ) lpr l (C i ) By Definition and Eq , L(P l ) = Pr[C i C j ]. lpr l (C i ) P ij = Pr[t(P) C j P L(P l ) s(p) C i P P l ], Pr[p L(P l ) s(p) C i P P l ] and by conditional probability considerations we get P ij = Pr[t(P) C j p L(P l ) s(p) C i P P l ]. (5.3.5) The term P P l in the hypothesis of Eq is redundant by the localized l-path. Thus, by combining with Eq we get P P1 ij = Pr[C i C j ] and the theorem is proved.
144 Relation to LDF In the original diffusion, it is assured that the magnitude of the eigenvalues of P is between zero and one. Another important property of P is the existence of a symmetric conjugate A. Being a symmetric matrix, the eigenvalues of A are all real and its left and right eigenvectors are identical. The matrix A has the same eigenvalues as P and its eigenvectors are related to those of P by the same conjugation that relates A to P. The additional information provided by the symmetric conjugate A allows for a simple spectral analysis to be used to achieve dimensionality reduction as described in [CL06a]. Theorem shows that these properties also apply to the ergodic l-path localized diffusion process P. Theorem Let P be the transition probability matrix of a localized l-path diffusion process, which resulted from Algorithm 2. Let Q be the corresponding degree matrix. Then the conjugate matrix  = Q 1/2 P Q 1/2 is symmetric. Furthermore,  is equivalent to the result from the weighted-sum LDF runner in [DA12, Section 3.3]. Proof. Consider two clusters C i and C j, i, j = 1,..., n. Let P be the matrix defined for them in the corresponding iteration of Algorithm 2. Let Q be a diagonal C i C j C i C j matrix where each cell on its diagonal corresponds to a data point x C i C j and holds its degree q(x) = q(x). As discussed in Section 5.2.1, the diffusion affinity matrix A = Q 1/2 P Q 1/2 is a symmetric conjugate of the diffusion operator P. Its cells are q(x) a(x, y) = p(x, y) x, y X. q(y) Let A = Q 1/2 PQ 1/2 be a C i C j C i C j conjugate of P, then its cells are a(x, y) = q(x) q(x) p(x, y) = q(y) q(y) p(x, y) = a(x, y) x, y C i C j. (5.3.6) From the symmetry of A, we get a(x, y) = a(y, x), x, y C i C j and A is symmetric. The powers of a symmetric matrix are also symmetric, thus A l, l 1, which was given as a parameter to Algorithm 2, is also symmetric and since the terms Q 1/2 Q 1/2 = I are canceled, then A l = Q 1/2 PQ 1/2 Q 1/2 PQ 1/2 Q 1/2 PQ }{{ 1/2 = Q } 1/2 P l Q 1/2. (5.3.7) l times
145 127 The symmetry is maintained by multiplying the symmetric matrix A l from left and right by the diagonal matrix Q 1/2. Thus the resulting matrix is Q 1/2 A l Q 1/2 = Q 1/2 Q 1/2 P l Q 1/2 Q 1/2 = QP l. According to Algorithm 2 and since q(x) = q(x) for x C i C j, then K ij = q(x)p l (x, y). x C i,y C j According to the symmetry of QP l, we obtain K ij = q(y)p l (y, x). x Ci y C j The same matrices P and Q are also used in the iteration that computes K ji in Algorithm 2, thus K ji = q(y)p l (y, x) = K ij y Cj x C i holds and K is symmetric. Since P Q 1 K by Algorithm 2, then  = Q 1/2 P Q 1/2 = Q 1/2 K Q 1/2. (5.3.8) Multiplication by the diagonal matrix Q 1/2 from both sides maintains the symmetry of K, thus  is also symmetric. Combining Eq and the definition of K in Algorithm 2, yields  ij = K ij = q(x)pl (x, y), Q ii Q ii x C i y C j q(c(x)) q(c(y)) Together with Eq , we receive  ij = q(x) q(y)a l (x, y). x C i q(c(x)) q(c(y)) Let y C j q(x)q(y) w xy q(c(x)) q(c(y)) then the following weighted sum is obtained:  ij = x C i y C j w xy a l (x, y). x, y X,
146 128 Finally, according to Eq , the matrix A is a sub-matrix of A, which contains cells in rows and columns that correspond to data points in C i C j. This is exactly the sub-matrix used in the corresponding iteration (for C i and C j ) in the LDF algorithm [DA12, Section 3.3], and thus Âij contains a weighted sum of the cells that are combined by the LDF runners [DA12, Section 3.3]. Therefore, the matrix Â, which is a symmetric conjugate of P, can be directly obtained by a weighted-sum LDF runner with the defined weights w xy, x, y X. From Theorem 5.3.5, the symmetric matrix  can be used for spectral analysis of the localized diffusion since it has the same spectrum as P and its eigenvectors are related to the eigenvectors of P by the same conjugation that relates  to P. In fact,  is a result of the LDF runner, which is used to prune a level in the LDF hierarchy to the next (higher) level, and thus, it can be constructed directly from A without using P, P and the conjugations (see Fig ). P Conjugation A Alg. 2 P Conjugation  LDF Figure 5.3.4: The relation between Algorithm 2 and the LDF pruning algorithm If we denote the eigenvalues of  by 1 = λ 0 λ 1 λ 2... and the corresponding eigenvectors by φ 0, φ 1, φ 2,..., we can define a coarse-grained DM, which we call the l-path localized diffusion map (LDM). This map embeds each cluster C i, i = 1,..., n, to a point Φ(C i ) = ( λ 0 φ 0 (C i ), λ 1 φ 1 (C i ), λ 2 φ 2 (C i ),...) T. According to the above discussion, this embedding has the same properties as the DM embedding, which was presented in [CL06a]. By combining the original DM, which embeds data points, and the presented LDM, which embeds clusters, we obtain a two-level embedding (i.e., a data point level and a cluster level). Moreover, the LDM is defined by spectral analysis of the affinity constructed by LDF. Therefore, it can be defined for each level of the LDF hierarchy. Thus, we get a multilevel embedding of the data where, in each level, the corresponding DF are embedded. Furthermore, each coarse-graining iteration (between LDF levels) prunes longer
147 129 paths to single transitions, and thus, a wider time scale of the diffusion is considered. Therefore, the multilevel embedding, which results from the LDM and from the LDF hierarchy, provides a multiscale coarse-grained DM. 5.4 Conclusion The presented l-path localized diffusion process introduces a coarse-grained version of the diffusion process that is used in DM for high-dimensional data analysis and dimensionality reduction. This coarse-grained process preserves the locality of the data by pruning previously detected clusters while considering only localized paths between them. A simple pruning algorithm can be used to perform the described coarse-graining while maintaining the essential algebraic and spectral properties of the DM process as was introduced in [CL06a]. Furthermore, this pruning is equivalent (via conjugation) to the one performed by the LDF algorithm when it computes the LDF hierarchy. By combining the results of this chapter with the ones in [DA12], the LDF hierarchy is shown to provide the foundations for a multi-scale coarse-grained DM-based embedding of data points and clusters/folders to a low-dimensional space. Appendix 5.A Proof of Lemma Proof. Since P is a Markovian random walk process with a transition probability matrix P, then the probability of a path P P l is l ξ=1 p(p ξ 1, P ξ ). The probability Pr[x L(Pl ) y], which is defined in Eq , considers only paths with P 0 = s(p) = x, P l = t(p) = y and P ξ C i C j, ξ = 1,..., l 1, thus Pr[x L(Pl ) y] = l 1 [p(x, P 1 ) p(p ξ 1, P ξ ) p(p l 1, y)] x C i, y C j. P 1,...,P l 1 C i C j ξ=2 By Algorithm 2, p(p ξ 1, P ξ ) = p(p ξ 1, P ξ ), ξ = 1,..., l when P 1,..., P l 1 C i C j, P 0 = x C i and P l = y C j. Therefore, Pr[x L(Pl ) y] = l 1 [p(x, P 1 ) p(p ξ 1, P ξ ) p(p l 1, y)] P 1,...,P l 1 C i C j ξ=2 = p l (x, y) x C i, y C j,
148 130 and the first part of the lemma is proved. The probability Pr[x L(Pl ) C j ], which was defined in Eq , combines all the probabilities in Eq with y C j. Different paths are considered independent events, thus Pr[x L(Pl ) C j ] = y C j Pr[x L(Pl ) y] = y C j p l (x, y) x C i, and the second part of the lemma is proved. The probability Pr[C i L(P l ) C j ], which was defined in Eq , combines all the probabilities in Eq with x C i. Since x is part of the condition in these probabilities, we get Pr[C i L(P l ) C j ] = x C i Pr[s(P) = x s(p) C i ] Pr[x L(Pl ) C j ] Using Eq we get = x C i Pr[C i L(P l ) C j ] = x C i y C j Pr[s(P) = x s(p) C i ] p l (x, y). y C j and the final part of the lemma is proved. q(x) vol(c i ) pl (x, y),
149 Chapter 6 Approximately-isometric diffusion maps In this chapter, we present an efficient approximation of the DM embedding. The presented approximation algorithm produces a dictionary of data points by identifying a small set of informative representatives. Then, based on this dictionary, the entire dataset is efficiently embedded into a low dimensional space. The Euclidean distances in the resulting embedded space approximate the diffusion distances. The properties of the presented embedding and its relation to DM method are analyzed and demonstrated. The results in this chapter appear in [SBWA13b]. 6.1 Introduction For a sufficiently small dataset, kernel methods can be implemented and executed on relatively standard computing devices. However, even for moderate size datasets, the necessary computational requirements to process them are unreasonable and, in many cases, impractical. For example, a segmentation of a medium size image with pixels requires a kernel matrix. The size of such a matrix necessitated about 270 GB of memory assuming double precision. Furthermore, the spectral decomposition procedure applied to such a matrix will be a formidable slow task. Hence, there is a growing need to have more computationally efficient methods that are practical for processing large datasets. The main computational load associated with kernel methods is generated by the application of a spectral decomposition to a kernel matrix. Sparsification by a sparse eigensolver such as Lanczos, which computes the relevant eigenvectors [CW02] of the kernel matrix, is widely used to reduce the compu- 131
150 132 tational load involved in processing a kernel matrix. Another sparsification approach is to transform the dense kernel matrix into a sparse matrix by selectively truncating elements outside a given neighborhood radius of each dataset member. Other approaches to achieve matrix sparsification are described in [vl07]. Given a dataset with n data points, common methods including the one described in this chapter for processing kernel methods require at least O(n 2 ) operations to determine which entries to either calculate or to threshold. While there are methods to alleviate these computational complexities [AM01], kernel sparsification might result in a significant loss of intrinsic geometric information such as distances and similarities. A prominent approach to reduces the discussed computational load is based on the Nyström extension method [FBCM04], which estimates the eigenvectors needed for an embedding. This approach is based on three phases: 1. The dataset is subsampled uniformly over the set of indices that are randomly chosen without repetition. 2. The subsamples define a smaller (than the dataset size) kernel. SVD is applied to the small kernel. 3. Spectral decomposition of a small kernel is extended by the application of the Nyström extension method to the entire dataset. This three-phase approach reduces the computational load, but the approximated spectral decomposition output suffers from several major problems. Subsampling affects the quality of the spectral approximation. In addition, the Nyström extension method exhibits ill-conditioned behavior that also affects the spectral approximation [BAC13]. Uniform subsampling of a sufficient number of data points captures most of the data probability distribution. However, rare events, compared to the subsampled size, might get lost. The results from this loss of information degrades the quality of the estimated embedded distances. The Nyström extension method is based on inverting a kernel matrix that was derived from a uniform sampling. This kernel does not necessarily has a full rank. Therefore, a direct kernel matrix inversion is ill-conditioned. The Moor-Penrose pseudo-inverse operator can overcome the ill-conditioned effect in Nyström extension. However, this solution may generate an inaccurate extension. Therefore, combining Nyström extension with random sampling can result in inaccurate approximations of spectral decomposition. Recently, the multiscale extension (MSE) scheme was suggested in [BAC13] as an alternative to the Nyström extension. This scheme, which samples
151 133 scattered data and extends functions defined on sampled data points, overcomes some of the limitations of the Nyström method. The MSE method is based on mutual distances between data points. It uses a coarse-to-fine hierarchy of a multiscale decomposition of a Gaussian kernel to overcome the ill-conditioned phenomenon and to speed the computations. In this chapter, which is based on [SBWA13b], we focus on alleviating the computational complexity of the DM method and enabling its application for large datasets. This kernel method utilizes a Markovian diffusion process to define and represent nonlinear relations between data points. It provides a diffusion distance metric that correlates with the intrinsic geometry of the data. It is defined by the pairwise connectivity of the data points in the DM diffusion process [LKC06]. Unlike the geodesic distance metric of manifolds, the diffusion distance metric is very robust to noise. The diffusion distance metric proved to be useful in clustering [DA12], parametrization of linear systems [TKC + 12] and even shape recognition [BB11]. The performances of the DM method depend on the size of the constructed kernel for the analyzed dataset. This size imposes severe limitations on the physical computational abilities to process it. In this chapter, we efficiently approximate the DM method by modifying the Nyström extension. This approximation, called µidm, guaranties that the difference between the diffusion distances in DM embedding and the Euclidean distances in µidm embedding, is preserved isometrically up to a given controllable error µ. The µidm utilizes the low dimensional geometry from the DM embedding to constructively design a dictionary that approximates the geometry of the entire DM embedding. The members of this dictionary are tailored to reduce the worst case approximation errors between the different embeddings. Additionally, we prove the convergence of the µidm spectrum to the respected DM spectrum. We bound the spectral convergence error as a function of the controllable error µ. The chapter has the following structure. Section 6.2 describes the general setup of the problem that includes a review of DM. Section 6.3 shows how a subset of distances in the DM space can be exactly computed via a spectral decomposition of a small kernel. Section 6.4 presents a variant of the Nyström method and analyzes the conditions that are required for the resulting mapping to preserve the diffusion distances of the relevant subset. Section 6.5 presents the dictionary construction and the µ-isometric approximation. In addition, this section analyzes the resulting approximation accuracy, its spectral convergence to DM spectrum and provides a computation complexity estimation as a function of the dataset and of the dictionary size. Finally, Section 6.6 examines the proposed method on data.
152 Problem formulation Let M be a low-dimensional manifold that lies in the high-dimensional Euclidean ambient space R m and let d m be its intrinsic dimension. Let M M be a dataset of M = n data points that are sampled from this manifold. The DM method [CL06a, Laf04] analyzes datasets such as M by exploring the geometry of the manifold M from which they are sampled. DM embeds the data into a space where the Euclidean distances between data points in the embedded space correspond to diffusion-based distances on the manifold M. A detailed construction of the DM is given in Section DM is a kernel method, which is based on the spectral analysis of a n n kernel matrix that holds the affinities between all the data points in M. For large datasets, derivation of the exact spectral decomposition of such a kernel is impractical due to the O(n 3 ) operations required by SVD. One way to reduce the computational complexity is to approximate this spectral decomposition such as in [AM01, vl07]. However, such SVD-based distances approximations in the embedded space is in general inaccurate and does not allow a direct control of the incurred approximation error. In this chapter, we efficiently approximate the DM embedding Φ : M R δ by a map Φ : M R δ. In order to quantify the error between the two maps, we introduce the notion of µ-isometric maps, which is given in Definition Definition (µ-isometric Maps). The maps Φ : M R δ and Φ : M R δ are µ-isometric if for every x, y M, Φ(x) Φ(y) Φ(x) Φ(y) µ. The notation denotes Euclidean norm in the respective space. The proposed method identifies a dictionary of data points in M that are sufficient to describe the pairwise distances between DM embedded data points. Then, the approximated map Φ is computed by an out-of-sample extension that preserves the pairwise diffusion distances in the dictionary. This is a modified version of Nyström extension that is used to compute the µ-isometric maps Algorithmic view of DM The DM method from [CL06a, Laf04] was briefly discussed in Chapter 1. In this chapter, we efficiently approximate the application of this method to the finite dataset M. We begin by reviewing the notations and main steps in the application of DM to M. Let K be the isotropic kernel matrix, whose elements are k(x, y) e x y ε x, y M, (6.2.1)
153 135 where ε is a meta-parameter. The degrees of data points based on this kernel are defined as q(x) y M k(x, y) x M. (6.2.2) Kernel normalization with this degree produces the row stochastic transition matrix P whose elements are p(x, y) = k(x, y)/q(x), x, y M. This transition matrix defines a Markov process over the data points in M. The DM embedding is obtained using spectral decomposition of the diffusion affinity kernel matrix A, which is a symmetric conjugate matrix of P. The elements of A are the diffusion affinities [A] (x,y) = a(x, y) k(x, y) = 1 q(x)p(x, y) x, y M. (6.2.3) q(x)q(y) q(y) The eigenvalues 1 = σ 1 σ 2... σ n > 0 of A and their corresponding eigenvectors φ 1, φ 2,..., φ n, are used to construct the diffusion map Φ : M R δ that maps each data point x M to Φ(x) = [q 1/2 (x)(σ 1 φ 1 (x)),..., q 1/2 (x)(σ δ φ δ (x))] (6.2.4) for a sufficiently small δ > 0. The value of δ depends on the decay of the spectrum of A and it determines the dimension of the embedded space. Typically, the application of DM to a dataset M of size n involves the following steps: 1. Use Eq to construct the n n kernel K; 2. Compute a diagonal matrix Q that holds for the data points in M the degrees q i n j=1 K ij for all i = 1,..., n; 3. Normalize K by Q to get a n n symmetric diffusion affinity kernel A = Q 1/2 KQ 1/2 by using Eq ; 4. Obtain the eigenvalues and the eigenvectors of A by the application of SVD to A = φσφ T to get the matrices σ 1 0 Σ =....., φ = φ 1 φ n. 0 σ n that hold the eigenvalues and the eigenvectors of A, respectively;
154 Use the matrix Q 1/2 φσ to embed each data point x i M, i = 1,..., n, to the i-th row of this matrix. Under the manifold assumption, the spectrum of the matrix A decays fast and only a small number of eigenvectors are required to obtain a reliable low dimensional embedding space. The diffusion distances between data points x, y M are defined as p(x, ) p(y, ), where p(x, ) and p(y, ) are the transition probabilities that are defined by the stochastic transition matrix P. The use of the spectral theorem in [CL06a] shows that the Euclidean distances in the embedded space of DM correspond to the diffusion distances in the manifold. Namely, Φ(x) Φ(y) = p(x, ) p(y, ). 6.3 Diffusion Maps of a partial set The computation of the DM embedding from Eq requires the spectral decomposition of the full n n symmetric diffusion kernel. Performing this decomposition on large datasets is computationally expensive. In this section, we describe an efficient method to compute the pairwise diffusion distances between data points of a partial dataset S M. We assume that, without loss of generality, M = {x 1,..., x n } and S = {x 1,..., x s }, s < n. Define the partial kernel K as the upper s n submatrix of the Gaussian kernel K from Eq Also let Q be the s s diagonal matrix whose diagonal entries are the degrees q(x i ) = n j=1 k(x i, x j ), i = 1, 2,..., s. Finally, we define the s n diffusion affinity kernel à between the partial set S and the dataset M by à Q 1/2 KQ 1/2. (6.3.1) Let φ Σ 2 φt be the SVD of the s s symmetric matrix ÃÃT. The eigenvalues of the matrix Σ 2, which are located on its diagonal, are σ σ s while its eigenvectors φ 1,..., φ s are located as its columns in φ. Definition uses the SVD-based decomposition to define a Partial Diffusion Map (PDM) on the partial set S. In what follows, the notation q(x) and φ j (x) will stand for q i and the i-th coordinate of φ j, respectively, where x = x i. Definition (Partial Diffusion Map). The Partial Diffusion Map (PDM) Φ : S R s of the partial set S is Φ(x) [ q 1/2 (x)( σ 1 φ1 (x)),..., q 1/2 (x)( σ s φs (x))].
155 137 Definition takes into consideration the entire spectrum of the decomposed partial kernel. In the rest of the chapter, we will assume that DM also considers the entire spectrum (i.e., δ = n in Eq ). However, for practical purposes, we can modify Definition so that PDM will only use a small number δ s < n of eigenvalues, similarly to the truncation of the number of eigenvalues as done in the DM embedding by Eq Theorem shows that the geometry of S under the DM embedding is preserved by the PDM embedding. Theorem The geometry of S under the DM embedding is preserved by the PDM applied to S. Formally, for every x, y S, Φ(x) Φ(y) = Φ(x) Φ(y) and Φ(x), Φ(y) = Φ(x), Φ(y). Due to Theorem 6.3.1, an embedding that preserves the diffusion distances of a partial set of size s can be computed by decomposing only a s s matrix instead of using a much bigger n n matrix. Lemma is needed for the proof of Theorem Lemma The matrix Q 1/2 ÃÃT Q 1/2 is the s s upper left matrix of Q 1/2 A 2 Q 1/2, i.e., it satisfies (Q 1/2 A 2 Q 1/2 ) (x,y) = ( Q 1/2 ÃÃT Q 1/2 ) (x,y) for every x, y S. Proof. According to Eq , Q 1/2 A 2 Q 1/2 = Q 1 KQ 1 KQ 1. Due to Eq and definitions of Q and K, the restriction of the matrix Q 1/2 A 2 Q 1/2 to the s s upper left matrix yields for every x, y S, (Q 1/2 A 2 Q 1/2 ) (x,y) = ( Q 1 KQ 1 KT Q 1 ) (x,y) = ( Q 1/2 ÃÃT Q 1/2 ) (x,y). Lemma shows the relation between the partial affinities and the full affinities and their associated degrees. The proof of Theorem uses this relation. Proof of Theorem By definition 6.3.1, for any x, y S, s Φ(x), Φ(y) = q 1/2 (x) σ j φj (x) q 1/2 (y) σ j φj (y). j=1 By using the spectral theorem, we get (ÃÃT ) (x,y) = s j=1 σ2 j φ j (x) φ j (y). Since the diagonal matrix Q holds the partial degrees q( ), we get Φ(x), Φ(y) = [ Q 1/2 ÃÃT Q 1/2 ] (x,y). Finally, we use Lemma to replace Q 1/2 ÃÃT Q 1/2 with Q 1/2 A 2 Q 1/2, thus Φ(x), Φ(y) = [Q 1/2 A 2 Q 1/2 ] (x,y).
156 138 On the other hand, by the DM definition we have n Φ(x), Φ(y) = q 1/2 (x)σ j φ j (x) q 1/2 (y)σ j φ j (y) = [Q 1/2 A 2 Q 1/2 ] (x,y). j=1 Therefore, Φ(x), Φ(y) = Φ(x), Φ(y) as the theorem states. Distance preservation in the theorem follows immediately since u v 2 = u, u 2 u, v + v, v for every u, v in both embedded spaces. 6.4 An out-of-sample extension that preserves the PDM geometry PDM provides an embedding Φ : S R s of a partial dataset S where s = S. In order to extend this embedding to the entire dataset M, an out-of-sample extension method is applied such that Φ is preserved over S. This is called an extended map. In this section, we utilize the Nyström extension [AM01, FH04] to compute the extended map for the entire dataset. In addition, we will constrain the extended map to have the same pairwise distances as PDM has. Therefore, the extended map will preserve the diffusion distances in S. Given a partial set S M of size s and its complement S = M \ S of size n s, then a diffusion affinity kernel A (Eq ) can be described as having the following block structure [ ] A(S,S) A A = (S, S) A T, (6.4.1) (S, S) A ( S, S) where the block A (S,S) R s s holds the diffusion affinities between data points in S, the block A ( S, S) R (n s) (n s) holds the affinities between data points in S, and the block A (S, S) R s (n s) holds the affinities between data points in S and data points in S. Under this formulation, Eq becomes à = [ A (S,S) A (S, S) ]. (6.4.2) Let A (S,S) = ψ Λ ψ T be the spectral decomposition of the positive-definite upper left block of A, where Λ is a diagonal matrix that contains the eigenvalues λ 1 λ 2... λ s, and ψ contains their corresponding eigenvectors as its columns. To extend this decomposition to the entire dataset M, the Nyström extension uses the property that every eigenvector ψ and every eigenvalue λ satisfy ψ = A ψ λ 1 (S,S) in the following way: [ ] ψ ψ = A T 1 ψ Λ (S, S).
157 It results in an n n approximated affinity matrix  = ψ Λ ψ T = 139 [ ] A(S,S) A (S, S) A T (S, S) A T (S, S) (A (S,S)) 1. (6.4.3) A (S, S) Therefore, the diffusion affinity matrix can be approximated by the extension given in Eq The DM embedding is based on the spectral decomposition of the diffusion affinity matrix A, which is approximated by Â. Therefore, in order to approximate DM embedding using the discussed extension, the matrix  has to be decomposed as  = φλ φ T, (6.4.4) where φ is a n s matrix with orthonormal columns and Λ is a s s diagonal matrix. A numerically efficient scheme for obtaining such a decomposition is presented in Section Definition presents the corresponding Nyström-based approximated DM based on this discussion. Definition Let φ and Λ be the matrices from Eq The Orthogonal Nyström-based Map (ONM) is the map Φ : M R s, given by Φ(x) [q 1/2 (x)λ 1 φ1 (x),..., q 1/2 (x)λ s φs (x)], where φ 1,..., φ s R n are the columns of the matrix φ and λ i, i = 1,..., s, is the ith diagonal elements in Λ. In other words, ONM embeds each data point in M into R s by the corresponding row of the matrix Q 1/2 φλ. The ONM in Definition embeds the entire dataset M into R s. As described in Section 6.4.1, ONM requires a spectral decomposition of a s s matrix rather than performing a spectral decomposition of n n matrix. Proposition shows that the geometry of S under the PDM is preserved by the ONM embedding. Proposition Let Φ and Φ be the PDM and the ONM embedding functions, respectively. Then, for every x, y S, Φ(x) Φ(y) = Φ(x) Φ(y) and Φ(x), Φ(y) = Φ(x), Φ(y). Proof. Since  = φλ φ T (Eq ) and φ T φ = I, we have  2 = φλ 2 φt, thus, the inner products in the embedded space of the ONM satisfy for every x, y M Φ(x), Φ(y) = [Q 1/2 φλ 2 φt Q 1/2 ] (x,y) = q 1/2 (x)[â2 ] (x,y) q 1/2 (y).
158 140 Furthermore, due to the structure of  in Eq , the upper left block of  2 is ÃÃT, thus, we have for every x, y S Φ(x), Φ(y) = q 1/2 (x)[ããt ] (x,y) q 1/2 (y) = Φ(x), Φ(y). The equality holds due to the spectral decomposition of ÃÃT and Definition Since the inner products in both embedded spaces are equal, the distance preservation follows immediately. Recall that according to Theorem 6.3.1, the geometry of S under the DM embedding is preserved by the PDM embedding. By combining Theorem with Proposition 6.4.1, we get Corollary Corollary The geometry of S under DM embedding is preserved by the ONM embedding, i.e., for every x, y S, Φ(x) Φ(y) = Φ(x) Φ(y) and Φ(x), Φ(y) = Φ(x), Φ(y) An efficient computation of the SVD of  Recall that the upper left submatrix A (S,S) is positive definite [Laf04]. Hence, it can be used to formulate an alternative Nyström approximation, which was presented in [FBCM04]. It can be verified that for every orthogonal s s matrix ψ and for any s s non-singular matrix Λ, the matrix [ ] A(S,S) φ = A 1/2 (S,S) ψλ 1/2 (6.4.5) A T (S, S) satisfies  = φλ φ T. Furthermore, according to [FBCM04], the matrix φ can be designed to decompose the matrix  in Eq as  = φλ φ T while having orthogonal columns (i.e., φ T φ = I). In our case, the extension is aimed to preserve the pairwise diffusion distances, and the related inner products, according to the PDM of S. Technically, the matrices φ and Λ are required to satisfy A 1/2 (S,S) ψλψt A 1/2 (S,S) = A (S,S)A (S,S) + A (S, S) A T (S, S), (6.4.6) where the LHS consists of the inner products of the Nyström approximation (Eq ) of S and the RHS consists of the inner products ÃÃT of the PDM of the same set S. This formulation dictates the definition of φ and Λ as the SVD C = ψλψ T, (6.4.7) where C is defined as C A (S,S) + A 1/2 (S,S) A (S, S)A T (S, S) A 1/2 (S,S). (6.4.8)
159 The µ-isometric construction In this section, we describe a constructive method to choose a partial set S M such that the resulting ONM from Definition will be µ-isometric to DM, which utilizes the full diffusion kernel. The proposed method uses a single scan of the entire dataset M and optimizes the dictionary selected set S for each processed data point. The construction of S is designed such that the geometry of M under the DM embedding is approximated by the ONM embedding applied to S. The proposed algorithm is iterative and it gradually constructs the dictionary subset S and the associated ONM. For its description, the following notations are used: The dataset M is assumed to be enumerated such that M = {x 1,..., x n }. Since the algorithm scans M only once, where in each iteration it examines a unique data point, the indices of the data points will indicate the current iteration number. That is, in iteration j (j = 1,..., n) of the algorithm the j-th data point is examined. In the j-th iteration, the algorithm holds a subdictionary S j = {y 1, y 2,..., y nj }. The subdictionary S j is a subset of M j = {x 1,..., x j } where n j j. Our algorithm constructs a monotonically increasing sequence of subdictionaries, i.e., S j 1 S j for any j = 2,..., n. The final dictionary S n is denoted by S. The notation Φ j denotes the ONM Φ j : M R n j applied to S j. Let κ < l < n, then according to Corollary 6.4.2, for all x, y S κ, Φ κ (x) Φ κ (y) = Φ(x) Φ(y) = Φ l (x) Φ l (y), i.e., the geometry of S κ under the DM embedding is identical to its geometries under the ONM embedding, applied to S l. Thus, there exists T : R nκ R n l that maps Φ κ (S κ ) onto Φ l (S κ ) isometrically. Definition defines such a map. This definition uses the invertibility of Φ κ (S κ ), which is proved in Appendix 6.A. Definition (Map-to-Map (MTM) Transformation). Assume the matrices Φ κ (y 1 ) Φ l (y 1 ) [ Φ κ (S κ )] =., [ Φ l (S κ )] =. } Φ κ (y nκ ) {{ } } Φ l (y nκ ) {{ } n κ n κ n κ n l hold the coordinates of data points in the dictionary S κ according to the maps Φ κ and Φ l, respectively. The linear Map-to-Map (MTM) transformation T κ,l : R nκ R n l is defined by the application 1 of the matrix [T κ,l ] [ Φ κ (S κ )] 1 [ Φ l (S κ )] to vectors u R nκ such that T κ,l (u) = u[t κ,l ] R n l. 1 Vectors in this definition are considered as row vectors and the matrix [T κ,l ] is applied to their right hand side.
160 142 It is clear from Definition that the MTM transformation of every Φ κ (x), x S κ, satisfies Φ l (x) = T κ,l Φ κ (x). (6.5.1) Therefore, the geometry of S κ is preserved in R n l under Tκ,l. Theorem shows that this transformation is an isometry between R nκ and its image in R n l. For data points in Sl \ S κ, the maps T κ,l Φ κ and Φ l may provide different embeddings. For all x S l \ S κ, the error β = T κ,l Φ κ (x) Φ l (x) (6.5.2) evaluates how well DM embeddings of data points in the set S l are approximated by ONM applied to S κ. We will base our dictionary membership criterion on this evaluation, and whether it is sufficiently small compared to a desired error bound. Algorithm 3: The µ-isometric DM (µidm) Input: data points: x 1,..., x n R m. Parameters: Distance error bound µ, Gaussian width ε Output: The approximated DM coordinates Φ(x i ), i = 1,..., n Initialize the dictionary: S 1 {x 1 } and s S 1 = 1 Initialize Q (Eq ) and à given S 1 (Eq ) Initialize the embedding: Φ ONM (Definition 6.4.1) of S1. for κ = 1 to n 1 do Set S S κ {x κ+1 } Compute Q and à (Eq ) given S Compute Φ ONM (Definition 6.4.1) of S Membership Test: Compute T MTM (Def ) from Φ( ) to Φ ( ) Compute β T ( Φ(x κ+1 )) Φ (x κ+1 ) If β > µ 2 Set S κ+1 S Set Q Q and à à Set Φ Φ Output the approximated diffusion coordinates Φ(x 1 ),..., Φ(x n ) The µidm construction in Algorithm 3 sequentially scans the data points x 1, x 2,..., x n M to check if their embeddings can be approximated by the dictionary or they have to be added to it. Initially, the dictionary is set to
161 143 contain a single data point x 1. Then, at each iteration κ, data points in M κ = {x 1,..., x κ }, which were already scanned, are approximated by the constructed dictionary S κ M κ. The algorithm processes the next data point x κ+1 and checks if the approximation of its embedding by dictionary S κ is sufficiently accurate. If it is, then the algorithm proceeds to the next iteration and the dictionary remains unchanged (i.e., S κ+1 = S κ ). Otherwise, this data point is added to the dictionary S κ. In the next iteration, S κ+1 = S κ {x κ+1 }. In Definition and in the accompanied discussion, we assumed without loss of generality that M contains the first κ and l data points that are the sets M κ and M l, respectively. This assumption simplifies the presentation. The dictionary membership criterion is based on comparing the approximation error Φ (x κ+1 ) T Φ(x κ+1 ) between the examined data point x κ+1 and a given adjustable threshold µ. In Section 6.5.1, we will show that this criterion guaranties that at the end of the dictionary construction process, the ONM embedding of every data point in M \ S is µ-isometric to DM. The rest of this section analyzes the accuracy of the resulting embedding and the computational complexity of its construction Distance accuracy of µidm Algorithm 3 constructs an optimized dictionary. Then, it uses the ONM of this dictionary to approximate the DM embedding. Corollary 4.2 guaranties that the ONM-based embedding preserves the diffusion distances between the dictionary members. Equivalently, it preserves the corresponding diffusion distances. The dictionary membership criterion guarantees that the distances from every data point not in the dictionary to the dictionary members approximate well the DM embedded distances up to the accuracy threshold µ. Theorem shows that the resulting dictionary-based ONM embedding preserves all the DM embedded diffusion distances in M, up to accuracy µ. Theorem Let Φ be the DM embedding (see Section 6.2.1) of M. Let S M be the dictionary constructed by Algorithm 3 and let Φ be the ONM, based on this dictionary. Then, for all x, y M, Φ(x) Φ(y) Φ(x) Φ(y) with an approximation error of at most µ. Theorem shows that the parameter µ in Algorithm 3 dictates the worst-case error of the approximated pairwise distances of the µidm. In order to prove this theorem, we first present Theorem and Lemma Theorem The MTM transformation T κ,l from Definition embeds R nκ isometrically in R n l, i.e. it satisfies for every u, v R n κ, T κ,l (u) T κ,l (v) = u v.
162 144 The proof of Theorem appears in Appendix 6.B. This theorem is used to prove Lemma 6.5.3, which shows that the µidm and the MTM isometry can be used to approximate the embedded diffusion coordinates of every data point up to an approximation error of µ 2. Lemma Assume we have Φ, S, Φ from Theorem Let T be the MTM isometry (Definition 6.5.1) between the µidm embedded space Φ( ) and the DM embedded space Φ( ). Then, every data point x M satisfies Φ(x) (T Φ)(x) µ 2. Proof. Recall that by Definition of the MTM isometry, Φ(x) = T Φ(x), x S. Then, we only have to consider data points that are not in the dictionary S. Consider such a data point x M \ S. Let S = S {x } and let Φ be the ONM of S. Assume also that T is the MTM isometry between Φ( ) and Φ ( ). Let T be the MTM isometry between Φ ( ) and Φ( ). The dictionary membership criterion in Algorithm 3 guarantees that for x / S, Φ (x ) (T Φ)(x ) µ. By the application of Theorem to the MTM 2 isometry T we get (T Φ )(x ) (T T Φ)(x ) = Φ (x ) (T Φ)(x ) µ 2. According to Definition 6.5.1, we have T = (T T ) and Φ(x ) = (T Φ )(x ). Therefore, we get Φ(x) (T Φ)(x) = (T Φ )(x ) (T T Φ)(x ) µ 2. The dictionary construction in Algorithm 3 compares the ONM approximations of each data point x κ+1 based on the dictionary S κ with the PDM of S κ {x κ+1 }. This comparison is done by utilizing a MTM isometry (see Definition 6.5.1). The result in Lemma shows that this membership criterion guarantees that the µidm embedding followed by the MTM transformation is sufficiently close to the DM embedding (up to a perturbation of size µ/2). Lemma is used to prove Theorem 6.5.1, which shows that the µidm embedding of M is µ-isometric to the application of DM embedding to M. Proof of Theorem Consider two data points x, y M. Then, by using Lemma we get Φ(x) (T Φ)(x) < µ 2 and Φ(y) (T Φ)(y) < µ 2.
163 Therefore, from the triangle inequality (T Φ)(x) (T Φ)(y) Φ(x) (T Φ)(x) + Φ(x) Φ(y) + Φ(y) (T Φ)(y) Φ(x) Φ(y) + µ, 145 and Φ(x) Φ(y) + Φ(x) (T Φ)(x) + (T Φ)(x) (T Φ)(y) Φ(y) (T Φ)(y) (T Φ)(x) (T Φ)(y) + µ. According to Theorem 6.5.2, the isometry in these equations satisfy (T Φ)(x) (T Φ)(y) = Φ(x) Φ(y). Thus, we get Φ(x) Φ(y) µ Φ(x) Φ(y) Φ(x) Φ(y) + µ A spectral bound for the kernel approximation In this section, we quantify the approximation quality of the diffusion kernel A from Eq by  from Eq Lemma provides a bound for the difference between the associated spectra, while Proposition shows the similarity between these operators. Recall that the eigenvalues of A are the diagonal elements of Σ, 1 = σ 1... σ n > 0 and the eigenvalues of à are the diagonal elements of Λ, λ 1... λ S > λ S +1 =... = 0. For the proofs, we consider a full SVD of Â, rather than its s SV D from Eq Let Θ be the n n orthogonal matrix, whose n s leftmost submatrix is φ, and the rest n (n s) constitute orthonormal basis for the orthogonal complement of the subspace spanned by the columns of φ. Additionally, let Λ be the diagonal n n matrix, whose upper left s s block is Λ and the rest are zeros. Lemma The difference between the spectra of A and  are bounded by µ 2 n s Q 1/2, i.e. for any j = 1,..., n, σ j λ j µ 2 n s Q 1/2. Proof. Due to Lemma 6.5.3, there is an orthogonal transformation T, for which Q 1/2 φλt Q 1/2 φσ µ 2 n s. Thus, according to Weyl s inequality, for every j = 1,..., n λ j σ j φλt φσ Q 1/2 Q 1/2 φλt Q 1/2 φσ µ 2 n s Q 1/2.
164 146 The last step assumes a worst-case scenario in which the difference between Q 1/2 φσ and Q 1/2 φλt is concentrated in a single coordinate with absolute value of µ. The spectrum of A is of great importance, since it indicates the dimensionality of the embedding for which the lost information is 2 negligible. Lemma states that the spectrum of A can be approximated with an error controlled by µ. More specifically, the diagonal matrix Q holds the degrees of the data points (see Eq ) on its diagonal, and it satisfies Q = max x M q(x). Thus, µ can be fixed such that the bound from Lemma is sufficiently tight and the numerical ranks of these operators are similar. Proposition is a direct consequence of Lemma It shows that A and à are almost similar, namely they act almost the same, up to orthogonal change of basis. It shows that  is a rank-s approximation of A, where, as in Lemma the error is controlled by µ. Proposition There exists an orthogonal n n matrix P for which P AP T  µ 2 n s Q 1/2. Proof. Obviously,  = Θ ΛΘ T. Define the n n matrix P φθ T. Then, P is orthogonal, and P AP T = φ Λφ T. Thus, due to Lemma 6.5.3, P AP T  µ 2 n s Q 1/ Computational complexity The analysis of the computational complexity is divided between the three main parts of µidm: 1. Initialization. 2. Membership Test, and 3. Update. This section assumes that the µidm is applied to M of size n. 1. Initialization: µidm computes the pairwise affinity matrix A, the corresponding degree matrix Q and the first mapping approximation Φ. An accurate computation of the degree requires O(mn 2 ) operations where m is the dimension of the ambient space. Additionally, the mapping initialization Φ requires an additional O(n) operations. This step is done once. 2. Membership Test: At the κ-th iteration, we have D κ = n κ κ. For a new data point x κ+1 in M, the µidm computes the matrix C (Eq ) that takes O(n κ n) operations. The matrix C is decomposed by the application of a SVD. It takes O(n κ 3 ) operations. Furthermore, the new mapping Φ is computed according to Eq It takes O(n κ 3 ) operations. The MTM computation is based on the already computed
165 147 [ Φ κ (S κ )] 1 (according to Definition 6.5.1). It takes an order of O(n κ 3 ) operations. Computation of β (Eq ) takes O(n κ 3 ) operations. Therefore, the total computational complexity of this step is O(n κ n 2 + n κ 3 n) operations assuming n iterations. 3. Update Step: For each new member in the dictionary, the µidm updates the relevant matrices and recompute the ONM mapping and [ Φ κ (S κ )] 1 with a total of O(nn κ 3 ) operations. Table summarizes the estimated complexity for computing µidm. The most expensive task is the computation of the affinity matrix and the degree matrix, which takes approximately O(mn 2 ) operations. Under the assumption that µ was chosen such that the dictionary size is smaller than n, then the µidm is more computationally efficient by an order of magnitude in comparison to DM computation. Operation Operations Initialization (done once) O(mn 2 ) Membership Test O(n 2 n κ + n 3 κ ) Update O(nn 3 κ ) Table 6.5.1: µidm computational complexity: m is the size of the ambient space, n is the number of samples, n κ is the dictionary size 6.6 Experimental results This section displays the µidm characteristics for three manifolds given in Fig Specifically, the presented examples in this section validate Theorem and Lemma Algorithm 3 is used for constructing the dictionary in the demonstrated analysis. The examined manifolds, which reside in R 3 and illustrated in Fig , include the unit sphere S 2 (a), the three dimensional Swiss-roll (b) and the three dimensional Mobius band (c). Each dataset was embedded in a highdimensional space, then it was uniformly sampled in 2000 data points. These datasets were embedded in R 17 by a random full-rank linear transformation, whose representative matrix is a 17 3 matrix where its entries are uniformly i.i.d. in [0, 1]. Its full rank guaranties the preservation of the intrinsic dimensionality of the manifolds. Algorithm 3 finds the µidm of each dataset. Figure compares between the first three coordinates of µidm and DM embeddings. µidm completes the scanning of the 2000 data points with a dictionary of size 164
166 148 (a) A Sphere (b) A Swiss Roll Figure 6.6.1: Examined manifolds (c) A Mobius Band where µ = It is clear from the figure that both maps are similar even though µidm utilized a SVD of a matrix of size instead of SVD of size In order to analyze the approximation errors in terms of pairwise distances and coordinates, the Cumulative Distribution Function (CDF) of each error of µidm relative to DM are computed. The corresponding CDF is the probability that any approximated coordinate of a data point or approximated distance in the embedded space is less than or equal to a threshold τ. More rigorously, the CDF is defined by F (τ, f(error)) = Pr[Error τ], (6.6.1) where f(error) is the distribution function of the respective error. The CDF describes an interval on which there is a positive probability to find an error and the percentage of non-negligible errors from all the error distributions. The estimated CDFs of the two errors from the Swiss Roll example are presented in Fig In each case, f(error) is estimated by integrating the corresponding histogram of the relevant error. For the
167 149 (a) The µidm of a swiss-roll (b) The DM of a Swiss roll (c) DM (+) and µidm (O) Figure 6.6.2: The embedding of a Swiss roll via DM and µidm (a) (b) Figure 6.6.3: The CDFs of (a) pairwise distance error and (b) coordinates mapping error between µidm and DM embeddings
168 150 coordinates error calculation, which were caused by the µidm embedding, the MTM between µidm and DM is utilized. According to the calculated CDFs that are shown in Fig , the coordinates errors from the application of µidm have positive probabilities only on the interval [0, ] as proved by Lemma In addition, the pairwise distance errors from the application of µidm have positive probabilities only on the interval [0, ]. This error is smaller than the error derived in Theorem Figure shows that in 50% of the cases, the calculated CDF probabilities of both errors are smaller by approximately one order of magnitude than their worst-cases. Table summarizes the measured error for the three datasets. Dataset ε S Max error between µidm and DM embeddings Max pairwise distance error Sphere Swiss Roll Mobius Band Table 6.6.1: µidm characterization summary where µ = Lemma discusses the convergence of the µidm spectrum to the associated DM spectrum. Figure compares the spectral decays of DM and µidm for the three datasets. Table provides the estimated bound and the measured difference for each dataset. Figure and Table suggests that the bound is not tight. The empirical difference is at least one order better than the corresponding bound. Furthermore, for a sufficiently small µ, µidm has a similar spectral decay as DM. Thus, when DM generates a low dimensional embedding due to its spectral decay, µidm embedding uses the same number of eigenvectors. Additionally, the empirical difference between the spectra suggests that a diffusion time greater than 1 can also be efficiently approximated by µidm. Dataset Q 1 2 S Approximation Bound (Lemma 6.5.4) max λ j σ j Sphere Swiss Roll Mobius Band Table 6.6.2: µidm: spectral difference, bound and empirical measurements
169 151 (a) The spectrum of a Sphere (b) The spectrum of a Swiss Roll (c) The spectrum of a Mobius band Figure 6.6.4: The eigenvalues of DM (denoted by +) and µidm (denoted by o) for each example
170 Discussion and conclusions This chapter presents a computationally efficient embedding scheme that approximately preserves the diffusion distances between embedded data points. The presented method scans the entire dataset once and validates the embedding approximation accuracy for each data point. This validation compares between a dictionary-based embedding and the exact DM embedding, which is efficiently computed over a subset of data points. The single scan of the dataset uses an iterative approach, and each iteration utilizes several techniques. In each iteration, a newly processed data point is considered for inclusion in the dictionary that was constructed from previously scanned data points and does not include the new data point. First, the Nyström extension is applied to the dictionary in order to approximate the embedding of the newly processed data point. Then, the PDM embedding of this data point, together with the dictionary, is efficiently computed. Finally, an MTM transformation is designed between the Nyström approximated embedded spaces and the exact PDM embedded spaces. This MTM transformation is used to measure the approximation accuracy of the embedding map. The entire computational complexity of this iterative process is lower than computational complexity of DM. The exact number of required operations depends on the dictionary size and on the dimensionality m of the original ambient space. The proposed method utilizes the exact pairwise affinities between data points in a given dataset. This computation limits the ability to reduce the computational complexity. However, future work will explore how to efficiently approximate this computation and quantify the resulting embedding errors. In order to demonstrate the effectiveness of the proposed method, we analyzed several synthetic datasets. This analysis showed that any choice of an approximation bound µ leads to a mapping that is similar to DM up to a pairwise distance error µ. Furthermore, the spectral properties of µidm and their relation to the spectral characteristics of DM were explored and proved. This spectral characterization suggests that the proposed method allows to have an effective dimensionally reduction that is similar to DM. Appendix 6.A Proof of the invertibility of the matrix Φ κ (S κ ) This appendix is dedicated to the presentation and proof of Lemma 6.A.1. It uses notations that were presented in Section 6.5. The presented lemma shows (and proves) the invertibility of the matrix Φ κ (S κ ), which consists of
171 153 the ONM embedding of the data points in the set S κ as its rows. Lemma 6.A.1. The matrix Φ κ (S κ ) in invertible. Proof. By definition 6.4.1, Φ κ (S κ ) is the upper s s submatrix of the s n matrix Q 1/2 φλ. Obviously, is suffices to prove that the upper s s submatrix of φλ is invertible. Due to Eqs , and 6.4.7, this submatrix equals to A 1/2 (S,S) ψλ 1/2, where ψλψ is the SVD of C = A (S,S) +A 1/2 (S,S) A (S, S)A T A 1/2 (S, S) (S,S). Since A is strictly positive definite, C is invertible and, as a consequence, so is Λ. Appendix 6.B Proof of Theorem This appendix presents the proof of Theorem 6.5.2, which states that the MTM transformation in Definition is an isometry. In order to prove this theorem, we first prove Lemma 6.B.1. The notation used in this section are the same as those used in Definition and Theorem Lemma 6.B.1. The matrix [T κ,l ] of size n κ n l, which defines the MTM in Definition 6.5.1, satisfies [T κ,l ][T κ,l ] T = I, where I is the n κ n κ identity matrix. Proof. By Definition 6.5.1, we have [T κ,l ] [ Φ κ (S κ )] 1 [ Φ l (S κ )]. Thus, [T κ,l ][T κ,l ] T = [ Φ κ (S κ )] 1 [ Φ l (S κ )][ Φ l (S κ )] T ([ Φ κ (S κ )] 1 ) T. (6.B.1) The entries in the matrix [ Φ l (S κ )][ Φ l (S κ )] T of size n κ n κ are the inner products between the embeddings of data points in S κ that were generated by the application of ONM to S l. Since S κ S l, Corollary is applied to these inner products. They are equal to the inner products generated by DM embedding. In addition, these inner products are also preserved by the application of ONM to S κ, which are the entries of the matrix [ Φ κ (S κ )][ Φ κ (S κ )] T. Therefore, we can replace [ Φ l (S κ )][ Φ l (S κ )] T with [ Φ κ (S κ )][ Φ κ (S κ )] T in Eq. 6.B.1 to get [T κ,l ][T κ,l ] T = ( [ Φ κ (S κ )] 1 [ Φ ) ( κ (S κ )] [ Φ κ (S κ )] 1 [ Φ T κ (S κ )]) = II T = I.
172 154 Proof. Proof of Theorem Consider two arbitrary data points u, v R nκ and their MTM-based transformed versions T κ,l (u), T κ,l (v) R n l, respectively. Then, by using Definition 6.5.1, we can write the inner product of the transformed data points as T κ,l (u), T κ,l (v) = u[t κ,l ], v[t κ,l ] = u[t κ,l ][T κ,l ] T v T. Due to Lemma 6.B.1, we get T κ,l (u), T κ,l (v) = uv T = u, v. Since all the inner products are preserved by the MTM transformation (recall u, v are arbitrary) we also get T κ,l (u) T κ,l (v) 2 = T κ,l (u v), T κ,l (u v) = u v, u v = u v 2, which proves Theorem
173 Part III Measure-based Learning 155
174
175 Chapter 7 Diffusion-based kernel methods on Euclidean metric measure spaces This chapter presents a generalized approach for defining diffusion-based kernels by incorporating measure-based information, which represents the density or distribution of the data, together with its local distances. The generalized construction does not require an underlying manifold to provide a meaningful kernel interpretation but assumes a more relaxed assumption that the measure and its support are related to a locally low dimensional nature of the analyzed phenomena. This kernel is shown to satisfy the necessary spectral properties that are required in order to provide a low dimensional embedding of the data. The associated diffusion process is analyzed via its infinitesimal generator and the provided embedding is demonstrated in two geometric scenarios. The results in this chapter appear in [BWA12, BWA13b]. 7.1 Introduction The original DM methodology from [CL06a, Laf04] is based on local distances in the data. When applied on data that are sampled from low-dimensional manifolds, these distances and the resulting diffusion neighborhoods capture the local (intrinsic) structure of the manifold. In this chapter, which is based on [BWA12], this methodology is enhanced by incorporating information about the distribution of the data, in addition to the distances on which DM is based. This distribution is expressed in term of a measure over the observable space. The measure (and its support) replace the manifold assumption. We assume that the measure quantifies the likelihood for the 157
176 158 presence of data over the geometry of the space. This assumption is significantly less restrictive than the need to have a manifold present. In practice this measure can either be provided as an input (e.g., by a-priori knowledge or a statistical model), or deduced from a given training set (e.g., by a density estimator). The manifold assumption can be expressed in terms of the measure assumption by setting the measure to be concentrated around an underlying manifold or (in the extremely restrictive case), to be supported by the manifold. Therefore, the measure assumption is not only less restrictive than the manifold assumption but it also generalizes it. The densities of the data were considered in two related variations of the DM framework. The anisotropic DM in [CL06a, Laf04] approximates these densities to separate the distribution of the data from the geometry of the underlying manifold. The adaptive-scale DM in [Dav09, DA12] uses these densities to adjust the local neighborhoods of the data by considering their connectivity. In [THJ10], a generalized diffusion process was considered, which encompasses these two variations in a single kernel definition. This kernel uses location dependent weight and bandwidth functions to normalize the kernel function (e.g., the Gaussian kernel). Under the manifold assumption, this kernel is shown to yield graph Laplacian matrices that converge to the Laplace-Beltrami operator on the underlying manifold. Other generalized kernels were also discussed in [SMC06, SMC08], where the diffusion kernel from [Laf04] was extended to consider function- and graph-based metrics to measure local distances and define diffusion transition probabilities. In the case of [CL06a, Laf04] and [Dav09, DA12], the used densities are deduced directly from the analyzed data by the application of a density estimator, which is based on the distances of the data. Specifically, when the sampled dataset is discrete, only the estimated densities at the sampled data points are used. The kernels in [SMC06, SMC08] and in [THJ10] allow for more flexible anisotropic normalizations, and in the latter case, the kernel can also be used for density estimation. However, these anisotropic considerations are done by normalization of the local distances (or the resulting kernel values) between data points. In the suggested construction, the used measure, which can represent densities, is separated from the distances and from the analyzed dataset. Therefore, when dealing with discrete data, this construction can utilize two different sets of samples: the analyzed dataset and the measure-related set with attached empirical measure values. Furthermore, from theoretical point of view, this construction combines continuous measures with either discrete or continuous datasets. Finally, most of the methods in these related works operate under and rely on the manifold assumption. Namely, they assume the data is sampled from an underlying low-dimensional manifold. As explained above, the presented results in this
177 159 chapter are obtained under a more relaxed measure assumption, without requiring the restrictive manifold assumption. The DM method and its variations are based on spectral analysis of transition probabilities operator, which is an integral operator over a measure space, determined by the distribution of the analyzed data. In our setup, due to the separation between the analyzed data and the underlying measure, which is encapsulated by the kernel, the integral transition operator can be defined without considering this distribution. Thus, the resulting kernel is still based on the distribution that is represented by the underlying measure, without any assumptions on the distribution of data points in the analyzed dataset. The structure of this chapter is as follows. Section 7.2 describes the problem setup. A brief description of the DM method is presented in Section Then, in Section 7.3, the measure-based kernel is formulated. Its spectral properties are presented in Section and its infinitesimal generator is analyzed in Section Finally, two geometric examples that demonstrate the proposed method are presented in Section Problem setup Let Ω R n, for some natural n, be a metric space with the Euclidean distance metric. For simplicity, we assume that Ω is a Euclidean subspace of R n. The integration notation dy in this chapter will refer to the Lebesgue integral Ω dy over the subspace Ω, instead of the whole space Rn. Let µ be a probability measure defined on Ω and let q(x) be the distribution function of µ, i.e., dµ(x) = q(x)dx. The measure µ and the distribution q are assumed to represent the distribution of data in Ω. Furthermore, we assume that q is sufficiently smooth 1 and it has a compact support supp(q) Ω, which is approximately locally low-dimensional. We aim to combine the distance metric of Ω and the measure µ to define a kernel function k(x, y), x, y Ω, which represents the affinities between data points in Ω. Then, these affinities can be used to construct a diffusion map, as described in Section 7.2.1, and utilize it to embed the data into a lowdimensional representation that considers both proximities and distributions of the data points. 1 Specifically, we assume q C 4 (Ω).
178 DM technicalities in general nonmanifold settings Technically, the DM method, which was briefly reviewed in Chapter 1, is based on an affinity kernel k and the associated integral operator that is defined as Kf(x) = k(x, y)f(y)dy, x Ω, for any function f L 2 (Ω). The affinity kernel k is normalized by a set of degrees ν(x) k(x, y)dy, x Ω, to obtain the transition probabilities p(x, y) k(x, y)/ν(x), from x Ω to y Ω, of the Markovian diffusion process. Under mild conditions on the kernel k, the resulting transition probability operator has a discrete decaying spectrum of eigenvalues 1 = λ 0 λ 1 λ 2..., which are used together with their corresponding eigenvectors 1 = φ 0, φ 1, φ 2,... to achieve the diffusion map of the data. Each data point x Ω is embedded by this diffusion map to the diffusion coordinates (λ 1 φ 1 (x),..., λ δ (x)φ δ (x)), where the exact value of δ depends on the spectrum of the transition probabilities operator P. The relation between the diffusion distance metric p(x, ) p(y, ) and the Euclidean distances in the embedded space, is a result of the spectral theorem [CL06a, Laf04]. Since the embedding is based on spectral analysis of the diffusion transition operator, it is usually comfortable to work with its symmetric conjugate that is defined by a(x, y) ν(x) 1/2 p(x, y)ν(y) 1/2 = k(x, y)/ ν(x)ν(y). This symmetric conjugate is called the diffusion affinity kernel, and the values a(x, y), x, y Ω, are the diffusion affinities of the data. Usually, the Gaussian affinities k ε (x, y) = exp( x y 2 /2ε), for some suitable ε > 0, are used for the construction of the diffusion map. When the data in Ω lies on a low dimensional manifold, its tangent spaces can be utilized to express the infinitesimal generator of the diffusion affinity kernel A in terms of the Laplacian operators on the manifold. In this chapter, we do not assume any underlying manifold structure. Instead, we assume we have access to a measure that represents the locally low dimensional distribution of the analyzed data. This measure can be supported by a low-dimensional manifold, but it can also represent nonmanifold structures that have no tangent spaces. Another benefit of using a smooth measure instead of a strict underlying structure is that it can gradually dissipate, thus accounting for possible noise that results in data points being spread around an underlying structure, instead of strictly lying on that structure. The standard DM method, which is based on the Gaussian kernel, is unsuitable for this case since it only utilizes distances and does not inherently consider the measure µ. In this chapter, we will present an enhanced kernel that incorporates the measure information together with distance information to define affinities and utilize them to obtain the DM
179 161 representation of the data in Ω. 7.3 Measure-based diffusion and affinity kernels In this section, we define and analyze an affinity kernel that is based on the distances in Ω and on the measure µ. We use this kernel together with the DM method, which was briefly described in Section 7.2.1, to obtain a measure-based diffusion affinity kernel and its resulting diffusion map. In Section 7.3.1, we explore the spectral properties of the associated integral operator, which are crucial for the spectral analysis that provides the embedded diffusion coordinates. Then, in Section 7.3.2, we show the relations between the infinitesimal generator of the resulting diffusion operator and the Laplacian operator on the space Ω and the measure µ. In order to define the desired kernel, we first define the function for some constant ρ 1. Its rescaled version is { g 1 (t) e t2 t ρ 0 otherwise, (7.3.1) g ε (t) g 1 ( t ε ), (7.3.2) for any ε > 0. The support of g ε ( t ) is B ερ(0), which is the closed ball of radius ερ centered at the origin. Notice that for a sufficiently large ρ, the Gaussian kernel, which is usually used in the DM method, can be defined as k ε (x, y) g 2ε ( x y ), and this definition will be used in the rest of the chapter. Definition uses the function g ε to define an alternative kernel that incorporates both local distance information, as the Gaussian kernel does, and measure information, which the Gaussian kernel lacks. Definition (Measure-based Gaussian Correlation kernel). The Measurebased Gaussian Correlation (MGC) affinity function k ε : Ω Ω R is defined as k ε (x, y) g ε ( x r ) g ε ( y r )dµ(r).
180 162 The MGC integral operator is defined by this function as K ε f(x) = k ε (x, y)f(y)dy for every function f L 2 (Ω) and data point x Ω. Figure 7.3.1: An illustration of the MGC affinities. It shows a measure that surrounds a straight line (marked in magenta), and the Gaussians around two examined data points (marked in red and blue). The MGC affinity is based on the intersection (marked in dark purple) between the supports of these three functions. The MGC affinity k ε (x, y), x, y Ω, from Definition 7.3.1, is in fact the inner product in L 2 (Ω, µ) between two Gaussians of width ε that are centered at x and y, respectively. This affinity is based on the correlation, which also takes into consideration the measure µ, between the described Gaussians around at the examined data points as illustrated in Fig The numerically significant positions of r in this correlation must be close enough to x and to y (based on their Gaussians of radius ε), but they must also be in an area with a high enough concentration of the measure µ. Notice that the measure information is considered and incorporated in the affinity definitions. It is not required any more in the application of the kernel
181 163 operator K to functions over Ω. An alternative formulation of the MGC affinities is presented in Proposition Proposition The MGC affinities from Definition can also be expressed by ( ) x + y k ε (x, y) = k ε (x, y) g ε/2 r 2 dµ(r). (7.3.3) Proof. Using the identity x r 2 + y r 2 = 1 x 2 y x+y r 2, 2 we get g ε ( x r )g ε ( y r ) = g 2ε ( x y )g ε/2 ( x+y r 2 ), which satisfies Eq Proposition shows the relation between the MGC kernel and the Gaussian kernel. While the Gaussian affinity only considers the distances between the examined data points, the MGC affinity also considers the region in which this distance is measured by using a Gaussian around the midpoint between them. This midpoint represents the direct path that determines the distance between the two data points. For a given distance between two data points, the MGC affinity increases when its path lies in an area with a high concentration of the measure µ, and decreases when it lies in an area with a low concentration of µ. If the measure µ is uniform over Ω, then the MGC kernel becomes the same as the Gaussian kernel up to a constant term that depends only on ε and can be easily normalized. Consider the case of uniform distribution { 1 x Ω q(x) = 0 otherwise, where Ω Ω is an open and connected set of unit volume, i.e., vol(ω ) = dr = 1. In this case, for every x, y Ω, the measure term of the MGC Ω affinities according to Proposition becomes g ε/2 ( x + y 2 ) r dµ(r) = Ω g ε 2 ( ) x + y r 2 dr, thus it does not represent any meaningful information about the data points x and y. Indeed, whenever the midpoint of x and y is far from the boundary of Ω (with respect to ε) Ω g ε 2 ( ) x + y r 2 dr ( πε ) n/2. 2
182 164 Therefore, we can normalize the MGC affinities k ε to get the normalized MGC affinity ˆk ε (x, y) ε n/2 kε (x, y) (7.3.4) that converges to the Gaussian affinity k ε when the measure is uniform, but incorporates the measure in the affinity when it is not uniform. The MGC affinity k ε and its normalized version ˆk ε only differ by a normalization term, thus they can be used interchangeably and the achieved results are equivalently valid for both of them. (a) When the data lies around a curve, the MGC affinities consider paths that follow the curve. (b) When the data lies in two separate clusters, the affinities between data points within a cluster are higher than data points from a different cluster. Figure 7.3.2: An illustration of the MGC affinities in two common data analysis scenarios. For every pair of compared data points, the significant values of the integration variable r, from Definition or Proposition 7.3.1, are marked. The dual representation of the MGC kernel in Definition and Proposition can be used to detect and consider several common patterns in data analysis directly from the initial construction of the kernel. Figure 7.3.2(a) uses the formulation in Definition to illustrate a case when the data is concentrated in areas around a curve with significant curvatures. In this case, the affinity will be more affected by the distances over the path that follows the noisy curve and not by the directions that follow sparse areas and bypass the curve. Figure 7.3.2(b) uses the formulation in Proposition to illustrate the affinities when the data is concentrated in two distinct clusters. In this case, we can see that the affinity between data points
183 165 from different clusters is significantly reduced due to the measure even if they are relatively close. Notice that in both illustrated cases, the density around the examined data points is similar, and the important information comes from considering the densities in the areas between them. This emphasizes a significant difference between the MGC kernel, the anisotropic kernel in [Laf04] and the adaptive kernel from [DA12]. The latter two approximate the densities around the compared data points and use these densities to normalize or adjust the affinity between them. However, when these data points lie in similarly significant densities, these adjustments do not take into account the areas between them. In practice, when dealing with finite sampled datasets, the MGC kernel does not require knowledge of the densities (or measure values) at the compared data points (x and y in Definition 7.3.1), which can be sampled independently from the inner integrand values (r in Definition 7.3.1), for which the densities are required. In fact, we can use two different sets: the analyzed dataset and the measure representing set. The utilization of these two sets of samples will be demonstrated in Section 7.4 together with additional examples. Section shows that the presented MGC affinity kernel satisfies the spectral properties that are required (and assumed) in [CL06a, Laf04] for its utilization with the DM framework. These properties enable us to define a diffusion process that is based on the MGC affinities. Then, the resulting diffusion map is used to embed the data in a way that considers the distances and the measure distribution. Section analyzes the properties of the resulting diffusion process by examining the infinitesimal generator of its transition probabilities and relating it to the infinitesimal generator in [CL06a] Spectral properties The DM embedding is based on spectral analysis of a normalized version of the used affinity kernel. Therefore, in order to use the MGC kernel with the DM analysis framework, the spectral properties of the associated integral operator have to be established first. In this section, we show that this kernel satisfies the assumptions (or conditions) in [CL06a], thus, the achieved DM results are applicable when the MGC kernel is utilized to provide the affinities of the data. We define the symmetric and positive kernel ã ε : Ω Ω R as ã ε (x, y) k ε (x, y) νε (x)ν ε (y), (7.3.5)
184 166 where ν ε (x) = kε (x, y)dy. The normalization values ν ε (x), x Ω, are referred to as the diffusion degrees of the data. The associated integral operator is à ε f(x) ã ε (x, y)f(y)dy. (7.3.6) This operator consists of the diffusion affinities of the data, when the diffusion is based on the MGC kernel. We will refer to it as the MGC diffusion affinities kernel. The operator Ãε is the symmetric conjugate of a stochastic operator that consists of the transition probabilities of the underlying diffusion process as was explained for the general DM setup in Section Its symmetry eases the investigation of its spectral properties, which are (up to conjugacy) the properties of the conjugate stochastic one. Proposition shows that à ε is a Hilbert-Schmidt operator. Proposition The diffusion affinity operator Ãε is a Hilbert-Schmidt operator from L 2 (Ω) into itself where its norm is Ãε = 1. It is L 2 (Ω) achieved by the square root of the stationary distribution of the underlying diffusion process. Corollary is a direct consequence of Proposition It essentially means that the spectral analysis of Ãε results in a small number of significant eigenvalues (and eigenvectors). Therefore, this operator enables the utilization of the DM framework for dimensionality reduction based on the MGC affinities. Corollary As a Hilbert-Schmidt operator, Ãε is compact self-adjoint, therefore its spectrum is discrete, it decays to zero and it is bounded from above by 1. The proof of Proposition is based on Lemma This Lemma establishes a crucial property of ã ε, which is required in order to show that à ε is a Hilbert-Schmidt operator. Lemma The MGC affinity function ã ε (Eq ) has a compact support in Ω Ω. Lemma is proved by using the compactness of the support of g ε. This proof is essentially technical, and it appears in Appendix 7.A.1. We can now prove Proposition Proof of Proposition The kernel function k ε (see Definition 7.3.1) is positive and continuous on its support, therefore, ν ε (x) satisfies the same
185 167 properties. As a consequence, ã ε (x, y) is a continuous function in Ω Ω whose support is compact (see Lemma 7.3.4) that satisfies ã 2 ε(x, y)dxdy <. Consequently, à ε is a Hilbert-Schmidt operator from L 2 (Ω) into itself. Additionally, k ε (x, y) f(y) ( ) 1/2 ( dy k ε (x, y)dy k ε (x, y) f ) 2 1/2 (y) ν(y) ν ε (y) dy = ( ν ε (x) k ε (x, y) f ) 2 (y) ν ε (y) dy, therefore, Ãε f, f = k ε (x, y) νε (x)ν ε (y) f(x)f(y)dxdy ( f(x) [ f k ε (x, y) f ) 2 1/2 (y) ν ε (y) dy dx k ε (x, y) f 2 (y) ν ε (y) dydx = f 2, hence, Ãε 1. Applying Ãε to ν ε (x) yields L 2 (Ω) ] 1/2 ( ) kε (x, y) à ε νε (x) = νε (x) dy = ν ε (x). The stationary distribution is achieved by normalizing the degrees of the data by the volume ν(x)dx. The last result remains valid even after normalization by this volume, thus the square root of the resulting stationary distribution is also an eigenvector of Ãε, associated with eigenvalue 1, as the proposition states. Corollary ensures that the DM can be utilized using the MGC kernel for dimensionality reduction. Furthermore, since the spectrum of à ε is bounded from above by 1, the diffusion process converges over time. Proposition shows that Ãε is positive definite, therefore, the discrete spectrum of Ãε lies in the interval [0, 1].
186 168 Proposition The operator Ãε is positive definite in L 2 (Ω) Proof. For any f L 2 (Ω) using Definition and Eq Ãε f, f = k ε (x, y) f(y) νε (y) dy f(x) νε (x) dx = g ε ( x r )g ε ( y r )q(r)dr f(x) f(y) νε (x) νε (y) dxdy { = g ε ( x r ) q(r)f(x)dx g ε ( y r ) } q(r)f(y)dy dr ( = g ε ( x r ) 2 q(r)f(x)dx) dr 0 To conclude this section, we summarize the spectral properties of the MGC integral operator Ãε. The spectrum of this operator is discrete, positive, bounded form above by 1 and decays to zero. Therefore, the eigenvalues of Ãε are denoted by 1 = λ 0 λ These properties enable the utilization of the DM for dimensionality reduction, by using the MGC affinities kernel. More specifically, considering the eigensystem of à ε, which satisfies Ãε φ j = λ j φj, j = 0, 1,..., the map 2 x ( λ t φ 1 1 (x),..., λ t φ δ δ (x)) is well defined and converges as t tends to infinity Infinitesimal generator The DM framework is based on Markovian diffusion process, which is defined and represented by a transition probability operator denoted by P ε. The infinitesimal generator of this operator encompasses the nature of the diffusion process. In [CL06a, Laf04], it was shown that when the data is sampled from a low dimensional underlying manifold, the infinitesimal generator of P ε has the form of Laplacian+Potential. In this section, we show a similar result, when using the MGC-based diffusion without requiring the underlying manifold assumption to hold. The MGC affinity function k ε is symmetric and positive, i.e., k ε (x, y) > 0 for any pair of data points x, y Ω. To convert it to be a transition kernel 2 The value of δ is determined by the numerical rank of the MGC operator, and it plays the same role here as in the original DM framework (see Section 7.2.1)
187 of a Markov chain on Ω, we normalize it as follows: thus, 169 p ε (x, y) k ε (x, y) ν ε (x), (7.3.7) p ε (x, y)dy = 1. (7.3.8) We define the corresponding stochastic operator P ε f(x) p ε (x, y)f(y)dy. (7.3.9) This operator is conjugate to Ã, defined in Eq , as their kernels satisfy the conjugacy relation ã ε (x, y) = νε 1/2 (x) p ε (x, y)νε 1/2 (y). Therefore, their spectral qualities are identical up to conjugacy. More specifically, their spectra are identical, and the eigenfunctions are conjugated, i.e., if ψ ε (x) is an eigenfunction of P ε corresponding to eigenvalue λ ε, then νε 1/2 (x)ψ ε (x) is an eigenfunction of Ãε, corresponding to the same eigenvalue. Similar relations between the diffusion affinities kernel a(x, y) and the transition probabilities kernel p(x, y) were already introduced in Section as the DM building blocks. The infinitesimal generator of the diffusion transition operator P ε is defined as I L lim P ε. ε 0 ε We use the notation ε = (I P ε )/ε, thus the infinitesimal generator takes the form L = ε 0. Theorem shows that the operator L takes the form Laplacian+potential, which is similar to the result shown in [Laf04, Corollary 2]. The expression, which Theorem provides for L, characterizes the differential equation for diffusion processes [CKL + 08, CLL + 05]. The rest of this section deals with the proof of Theorem Theorem If the density function q is in C 4 (Ω), then the infinitesimal generator L of the MGC-based diffusion operator is Lf = m ( ) 2 q f + m 0 q, f, f C 4 (Ω), where, m 0 = m 2 = g 1 ( x )dx, g 1 ( x )(x (j) ) 2 dx.
188 170 The proof of Theorem contains two parts. The first part of the proof, in Lemma 7.3.7, examines the application of the diffusion transition operator P ε to an arbitrary function. The second part, in Proposition 7.3.8, examines the asymptotic infinitesimal behavior of the operator ε, which results in the infinitesimal generator L. Lemma For any x, y Ω and for any positive ε P ε f(x) = (ε d m 2 0f(x)q(x) + m 0 m 2 ε d+1 (f(x) q(x) + (q(x) f(x))) + O(ε d+2 ))/ν ε (x). The proof of Lemma is based on Taylor expansions of the function f and the density function q (of the measure). It is similar to the approach taken by [Laf04, CL06a], but instead of using tangential structures (of a manifold), we use measure based considerations. The complete proof is rather technical and it appears in Appendix 7.A.2. Proposition uses the result in Lemma 7.3.7, to examine asymptotic behavior of the transition operator P ε. Proposition For any x Ω, f(x) lim P ε f(x) ε 0 ε = m 2 (q(x) f(x)). m 0 q(x) The proof of Proposition relies on Lemma and some technical limit calculations. The complete proof of this proposition appears in Appendix 7.A.2. Theorem is a direct result of Proposition Indeed, (q f) = f + q, f, which gives the expression for L in the theorem. 7.4 Geometric examples In this section, we demonstrate by two examples the MGC kernel and the resulting diffusion map. The first example analyzes noisy data that is spread around a spiral curve. In this case, we compare the MGC kernel and its diffusion to the classic DM [CL06a]. The second example presents a case when only the measure is given, and the analyzed data points are given by a uniform grid around the support of the measure. This case can occur, for example, when only statistical information about the distributions of the data is given, or when dealing with massive datasets where the analysis of individual data points is unfeasible. In this case, the original DM method from [CL06a] cannot be applied at all since the distances of the uniform grid are meaningless. However, the MGC kernel is also based on measure
189 171 (a) Noisy data around the curve (b) An exponentially-decaying measure around the curve Figure 7.4.1: A spiral curve with 5000 noisy data points concentrated around it, and 10 4 points that represent an exponentially-decaying measure around the curve. The colorscale color map from Fig. 7.B.1(a) is used to represent the measure values. information, therefore, it reveals the underlying geometry that is represented by this measure. Note: The figures in this section use three color maps. For reference, these color maps are presented in Fig. 7.B.1 in Appendix 7.B Noisy spiral curve In this section, we compare between the Gaussian-based DM embedding from [CL06a] and the embedding achieved by the MGC-based DM presented in this chapter. We use a noisy spiral curve (see Fig (a)) for the comparison. The dataset was produced by sampling 500 equally spaced points from the curve and then sampling 10 normally distributed data points around each of these curve points. The resulting data has 5000 data points that lie in areas around the curve, as shown in Fig (a), where the curve is marked in red and the noisy data points are marked in blue. We used the same scale meta-parameter ε to the compared DM applications. This meta-parameter was set to be sufficiently high to overcome the noise and to detect the high affinity between data points that originated from the same position (out of the 500 curve points) on the curve. The application of the Gaussian-based DM is straightforward, as explained in [CL06a]. The Gaussian kernel k is constructed and then nor-
190 172 malized by the degrees to obtain the diffusion transition matrix P and the diffusion affinity matrix A. Spectral analysis of these matrices yields an embedding that is based on their most significant eigenvalues and eigenvectors. The MGC kernel from Definition requires to define a measure over the area where the data lies. Notice that the measure of the actual data points is not required. We can define a completely different set of points r from Definition and then define their weights, which represent their measure values. We use two different measures for this definition. The first measure is based on 10 4 equally spaced data points from the curve and all the weights are set to one. This measure is essentially an indicator function of the spiral curve denoted by µ c. The second measure is based on 10 4 points that are sampled around the curve by adding Gaussian noise to the data points that were used for defining µ c. The weights of the point decay exponentially in relation to their distance from the curve. The resulting measure is denoted by µ v and it is presented in Fig (b) where the 10 4 measure points are colored according to their measure weights. We use the notations K c, Pc and Ãc to denote the matrices that result from Definition 7.3.1, Eq and Eq , respectively, with the measure µ c. The notations K v, Pv and Ãv are used in a similar way for the measure µ v. Notice that in both cases, even though the measure is based on 10 4 positions of the integration variable r (from Definition 7.3.1), the kernel and its normalized versions are of size , since the data has only 5000 data points. Figure compares the neighborhoods that are represented by the three kernels K, Kc and K v. We examine the neighborhoods of two data points on two different levels of the curve. In both cases, the Gaussian kernel captures inter-level affinities (i.e., it links different levels of the spiral) while both versions of the MGC kernel only capture relations in the same level of the spiral, thus, they are able to separate between these levels. In addition, the shape of the neighborhoods of the MGC kernels form ellipses whose major axes clearly follow the significant tangential directions of the curve. The Gaussian kernel, however, captures circular neighborhoods that do not express any information about the significant directions of the data. Since both K c and K v show similar neighborhoods and they indeed capture similar relations, we will only present from now on the comparison between the Gaussian-based diffusion and the MGC-based diffusion that is based on µ c. Similar results are also achieved by using µ v. The embedding, which is achieved by DM, is based on a diffusion process whose time steps are represented by powers of the diffusion transition matrix or the diffusion affinity matrix. The resulting Markov process has a stationary distribution when the time steps are taken to infinity. This stationary
191 173 (a) 1st K neighborhood (b) 2nd K neighborhood (c) 1st K c neighborhood (d) 2nd K c neighborhood (e) 1st K v neighborhood (f) 2nd K v neighborhood Figure 7.4.2: Two neighborhoods from the Gaussian kernel (K) and the MGC kernels ( K c and K v ) on the spiral curve, using the heat-map in Fig. 7.B.1(b) to represent the kernel values.
192 174 (a) Gaussian-based stationary distribution (b) MGC-based stationary distribution Figure 7.4.3: The stationary distributions of: (a) the Gaussian-based diffusion process, and (b) the MGC-based diffusion process. Both use the grayscale color map from Fig. 7.B.1(c) to represent the distribution values. (a) Gaussian-based DM (b) MGC-based DM Figure 7.4.4: The first two diffusion coordinates of the Gaussian-based and MGC-based DM embeddings.
193 175 (a) Gaussian-based DM (b) MGC-based DM Figure 7.4.5: The first three diffusion coordinates of the Gaussian-based and MGC-based DM embeddings. distribution reveals the concentrations and the underlying potential of the diffusion process. It is represented by the first eigenvector of the diffusion affinity matrix 3. Figure compares the stationary distributions of the Gaussian-based diffusion with the MGC-based diffusion as represented by the first eigenvector of the corresponding diffusion affinity matrix A or Ãc. This comparison shows that the Gaussian-based diffusion considers the entire spiral as one pit of potential. At infinity, the diffusion is distributed over the entire region of the curve. The MGC-based diffusion, on the other hand, separates different levels of the spiral. At infinity, this diffusion is concentrated on the curve levels themselves and not on the areas between them. Finally, we compare between the embedded spaces of the Gaussian-based DM and the MGC-based DM. Figure presents these embedded spaces based on the first two diffusion coordinates and Fig presents these spaces based on the first three diffusion coordinates (i.e., the two/three most significant eigenvectors of the diffusion transition operator). The comparison in Fig clearly shows that the MGC-based embedding results in a better separation between the spiral levels. Figure further establishes this observation by showing that, in fact, the Gaussian-based diffusion considers the 3 More accurately, the first eigenvector of the diffusion affinity is the square root of the stationary distribution, but it is sufficient to use these values for the purposes of these demonstrations
194 176 whole noisy spiral as a two-dimensional disk. The MGC-based embedding, on the other hand, uses the third diffusion coordinate to completely separate the levels of the spiral by stretching it apart in the three-dimensional embedded space. The superior results (e.g., separation between the spiral levels) of the MGC-based DM demonstrate its robustness to noise. The reason for this robustness is because the noise is part of the model on which the MGC construction is based. The Gaussian-based DM assumes that the data lies on (or it is sampled from) an underlying manifold, and any significant noise outside this manifold may violate this assumption. The MGC-based DM, on the other hand, already assumes variable concentrations and distributions of the data, which are represented by the measure and incorporated into the affinities. Therefore, this setting is more natural when dealing with data that is concentrated around an underlying manifold structure but does not necessarily lie on the manifold Uniform grid with a fish-shaped measure In this section, we demonstrate a case when the Gaussian-based DM is inapplicable but the MGC-based DM can be applied for the analysis. Instead of using a discrete dataset of samples to represent the analyzed data, we use a measure, which holds the meaningful information about the analyzed phenomenon. This scenario can occur, for example, when dealing with massive datasets where it is unfeasible to analyze individual data points but one can obtain a density estimator over the observable space by using the massive number of samples. We will use a uniform grid or arbitrary size, which does not depend on the measure or its representative points, and utilize the MGC-based DM to analyze this grid in relation to the input measure. We use a measure that is concentrated around a fish shape in two dimensions (see Fig ). It is represented by approximately 25, 000 points. These points are sampled from areas around the support of the measure, and they are weighted according to their measure value. Figure shows the representative points and their measure-representing weights. In order to analyze the measure, we generate a square grid in the bounding box of the support of the measure, and use the resulting 10, 000 grid points as a dataset for the analysis. Since the grid is uniform, the distances between its grid points do not hold any meaningful information. Therefore, the Gaussian-based DM cannot be applied to analyze it. The MGC-based DM, on the other hand, can incorporate the measure information (based on the 25, 000 representative data points) in the grid analysis. Thus, the resulting embedding will consider the meaningful information of the measure and not
195 177 Figure 7.4.6: Fish shape measure. just the meaningless distances. We use Definition to construct the MGC kernel K of the grid and the measure. The values of the integration variable r (in Definition 7.3.1) are taken from the 25, 000 measure representatives, while the values of the compared points x and y (in Definition 7.3.1) are taken from the 10, 000 grid points. The resulting kernel size is , and it does not depend on the number of measure representatives. Therefore, we can use an arbitrarily large number of points for representing the measure without affecting the MGC kernel size, which is only determined by the grid size. In order to apply the DM scheme to the MGC kernel K, we normalize it to (a) Diffusion degrees. (b) Stationary distribution Figure 7.4.7: Diffusion degrees and stationary distribution.
196 178 obtain the transition matrix P (see Eq ) and the diffusion affinity à (see Eq ). The normalization values of the kernel are the degrees of the data points in a graph that is represented by K as its weighted adjacency matrix. These degrees measure the centrality of each data point in this graph and the resulting diffusion process. Figure 7.4.7(a) shows the degrees of the grid data points. Even though the grid is uniform and its distances are meaningless, this figure shows that the data points that lie in concentrated areas of the measure, are more central than others. This property of the MGC-based construction is a result of the measure information being considered and incorporated in the MGC kernel. Another property of the diffusion process is its stationary distribution. This distribution represents the underlying potential of the diffusion. It governs the concentrations of the diffusion process as it converges to an equilib- (a) The 1st and 2nd diffusion coordinates (b) The 3rd and 4th diffusion coordinates Figure 7.4.8: The MGC-based DM embedding of the grid based on the first four MGC-based diffusion coordinates. It is presented in two pairs: 1st-2nd and 3rd-4th.
197 179 Figure 7.4.9: The three-dimensional presentation of the embedded grid based on the second, third and fourth MGC-based diffusion coordinates.
Measure-based diffusion grid construction and high-dimensional data discretization
 Measure-based diffusion grid construction and high-dimensional data discretization Amit Bermanis, Moshe Salhov, Guy Wolf, Amir Averbuch School of Computer Sciene, Tel Aviv University, Tel Aviv 69978, Israel
Measure-based diffusion grid construction and high-dimensional data discretization Amit Bermanis, Moshe Salhov, Guy Wolf, Amir Averbuch School of Computer Sciene, Tel Aviv University, Tel Aviv 69978, Israel
Contribution from: Springer Verlag Berlin Heidelberg 2005 ISBN
 Contribution from: Mathematical Physics Studies Vol. 7 Perspectives in Analysis Essays in Honor of Lennart Carleson s 75th Birthday Michael Benedicks, Peter W. Jones, Stanislav Smirnov (Eds.) Springer
Contribution from: Mathematical Physics Studies Vol. 7 Perspectives in Analysis Essays in Honor of Lennart Carleson s 75th Birthday Michael Benedicks, Peter W. Jones, Stanislav Smirnov (Eds.) Springer
Learning Eigenfunctions: Links with Spectral Clustering and Kernel PCA
 Learning Eigenfunctions: Links with Spectral Clustering and Kernel PCA Yoshua Bengio Pascal Vincent Jean-François Paiement University of Montreal April 2, Snowbird Learning 2003 Learning Modal Structures
Learning Eigenfunctions: Links with Spectral Clustering and Kernel PCA Yoshua Bengio Pascal Vincent Jean-François Paiement University of Montreal April 2, Snowbird Learning 2003 Learning Modal Structures
Nonlinear Dimensionality Reduction
 Nonlinear Dimensionality Reduction Piyush Rai CS5350/6350: Machine Learning October 25, 2011 Recap: Linear Dimensionality Reduction Linear Dimensionality Reduction: Based on a linear projection of the
Nonlinear Dimensionality Reduction Piyush Rai CS5350/6350: Machine Learning October 25, 2011 Recap: Linear Dimensionality Reduction Linear Dimensionality Reduction: Based on a linear projection of the
Nonlinear Dimensionality Reduction. Jose A. Costa
 Nonlinear Dimensionality Reduction Jose A. Costa Mathematics of Information Seminar, Dec. Motivation Many useful of signals such as: Image databases; Gene expression microarrays; Internet traffic time
Nonlinear Dimensionality Reduction Jose A. Costa Mathematics of Information Seminar, Dec. Motivation Many useful of signals such as: Image databases; Gene expression microarrays; Internet traffic time
Non-linear Dimensionality Reduction
 Non-linear Dimensionality Reduction CE-725: Statistical Pattern Recognition Sharif University of Technology Spring 2013 Soleymani Outline Introduction Laplacian Eigenmaps Locally Linear Embedding (LLE)
Non-linear Dimensionality Reduction CE-725: Statistical Pattern Recognition Sharif University of Technology Spring 2013 Soleymani Outline Introduction Laplacian Eigenmaps Locally Linear Embedding (LLE)
Focus was on solving matrix inversion problems Now we look at other properties of matrices Useful when A represents a transformations.
 Previously Focus was on solving matrix inversion problems Now we look at other properties of matrices Useful when A represents a transformations y = Ax Or A simply represents data Notion of eigenvectors,
Previously Focus was on solving matrix inversion problems Now we look at other properties of matrices Useful when A represents a transformations y = Ax Or A simply represents data Notion of eigenvectors,
Nonlinear Dimensionality Reduction
 Outline Hong Chang Institute of Computing Technology, Chinese Academy of Sciences Machine Learning Methods (Fall 2012) Outline Outline I 1 Kernel PCA 2 Isomap 3 Locally Linear Embedding 4 Laplacian Eigenmap
Outline Hong Chang Institute of Computing Technology, Chinese Academy of Sciences Machine Learning Methods (Fall 2012) Outline Outline I 1 Kernel PCA 2 Isomap 3 Locally Linear Embedding 4 Laplacian Eigenmap
Nonlinear Methods. Data often lies on or near a nonlinear low-dimensional curve aka manifold.
 Nonlinear Methods Data often lies on or near a nonlinear low-dimensional curve aka manifold. 27 Laplacian Eigenmaps Linear methods Lower-dimensional linear projection that preserves distances between all
Nonlinear Methods Data often lies on or near a nonlinear low-dimensional curve aka manifold. 27 Laplacian Eigenmaps Linear methods Lower-dimensional linear projection that preserves distances between all
Dimension Reduction Techniques. Presented by Jie (Jerry) Yu
 Dimension Reduction Techniques Presented by Jie (Jerry) Yu Outline Problem Modeling Review of PCA and MDS Isomap Local Linear Embedding (LLE) Charting Background Advances in data collection and storage
Dimension Reduction Techniques Presented by Jie (Jerry) Yu Outline Problem Modeling Review of PCA and MDS Isomap Local Linear Embedding (LLE) Charting Background Advances in data collection and storage
Unsupervised Learning Techniques Class 07, 1 March 2006 Andrea Caponnetto
 Unsupervised Learning Techniques 9.520 Class 07, 1 March 2006 Andrea Caponnetto About this class Goal To introduce some methods for unsupervised learning: Gaussian Mixtures, K-Means, ISOMAP, HLLE, Laplacian
Unsupervised Learning Techniques 9.520 Class 07, 1 March 2006 Andrea Caponnetto About this class Goal To introduce some methods for unsupervised learning: Gaussian Mixtures, K-Means, ISOMAP, HLLE, Laplacian
Kernel-Based Contrast Functions for Sufficient Dimension Reduction
 Kernel-Based Contrast Functions for Sufficient Dimension Reduction Michael I. Jordan Departments of Statistics and EECS University of California, Berkeley Joint work with Kenji Fukumizu and Francis Bach
Kernel-Based Contrast Functions for Sufficient Dimension Reduction Michael I. Jordan Departments of Statistics and EECS University of California, Berkeley Joint work with Kenji Fukumizu and Francis Bach
Advances in Manifold Learning Presented by: Naku Nak l Verm r a June 10, 2008
 Advances in Manifold Learning Presented by: Nakul Verma June 10, 008 Outline Motivation Manifolds Manifold Learning Random projection of manifolds for dimension reduction Introduction to random projections
Advances in Manifold Learning Presented by: Nakul Verma June 10, 008 Outline Motivation Manifolds Manifold Learning Random projection of manifolds for dimension reduction Introduction to random projections
Graph Metrics and Dimension Reduction
 Graph Metrics and Dimension Reduction Minh Tang 1 Michael Trosset 2 1 Applied Mathematics and Statistics The Johns Hopkins University 2 Department of Statistics Indiana University, Bloomington November
Graph Metrics and Dimension Reduction Minh Tang 1 Michael Trosset 2 1 Applied Mathematics and Statistics The Johns Hopkins University 2 Department of Statistics Indiana University, Bloomington November
Connection of Local Linear Embedding, ISOMAP, and Kernel Principal Component Analysis
 Connection of Local Linear Embedding, ISOMAP, and Kernel Principal Component Analysis Alvina Goh Vision Reading Group 13 October 2005 Connection of Local Linear Embedding, ISOMAP, and Kernel Principal
Connection of Local Linear Embedding, ISOMAP, and Kernel Principal Component Analysis Alvina Goh Vision Reading Group 13 October 2005 Connection of Local Linear Embedding, ISOMAP, and Kernel Principal
March 13, Paper: R.R. Coifman, S. Lafon, Diffusion maps ([Coifman06]) Seminar: Learning with Graphs, Prof. Hein, Saarland University
![March 13, Paper: R.R. Coifman, S. Lafon, Diffusion maps ([Coifman06]) Seminar: Learning with Graphs, Prof. Hein, Saarland University March 13, Paper: R.R. Coifman, S. Lafon, Diffusion maps ([Coifman06]) Seminar: Learning with Graphs, Prof. Hein, Saarland University](/thumbs/93/111953007.jpg) Kernels March 13, 2008 Paper: R.R. Coifman, S. Lafon, maps ([Coifman06]) Seminar: Learning with Graphs, Prof. Hein, Saarland University Kernels Figure: Example Application from [LafonWWW] meaningful geometric
Kernels March 13, 2008 Paper: R.R. Coifman, S. Lafon, maps ([Coifman06]) Seminar: Learning with Graphs, Prof. Hein, Saarland University Kernels Figure: Example Application from [LafonWWW] meaningful geometric
Lecture 10: Dimension Reduction Techniques
 Lecture 10: Dimension Reduction Techniques Radu Balan Department of Mathematics, AMSC, CSCAMM and NWC University of Maryland, College Park, MD April 17, 2018 Input Data It is assumed that there is a set
Lecture 10: Dimension Reduction Techniques Radu Balan Department of Mathematics, AMSC, CSCAMM and NWC University of Maryland, College Park, MD April 17, 2018 Input Data It is assumed that there is a set
Preprocessing & dimensionality reduction
 Introduction to Data Mining Preprocessing & dimensionality reduction CPSC/AMTH 445a/545a Guy Wolf guy.wolf@yale.edu Yale University Fall 2016 CPSC 445 (Guy Wolf) Dimensionality reduction Yale - Fall 2016
Introduction to Data Mining Preprocessing & dimensionality reduction CPSC/AMTH 445a/545a Guy Wolf guy.wolf@yale.edu Yale University Fall 2016 CPSC 445 (Guy Wolf) Dimensionality reduction Yale - Fall 2016
Diffusion Wavelets and Applications
 Diffusion Wavelets and Applications J.C. Bremer, R.R. Coifman, P.W. Jones, S. Lafon, M. Mohlenkamp, MM, R. Schul, A.D. Szlam Demos, web pages and preprints available at: S.Lafon: www.math.yale.edu/~sl349
Diffusion Wavelets and Applications J.C. Bremer, R.R. Coifman, P.W. Jones, S. Lafon, M. Mohlenkamp, MM, R. Schul, A.D. Szlam Demos, web pages and preprints available at: S.Lafon: www.math.yale.edu/~sl349
Geometric Component Analysis and its Applications to Data Analysis
 Geometric Component Analysis and its Applications to Data Analysis Amit Bermanis 1, Moshe Salhov 2, and Amir Averbuch 2 1 Department of Mathematics, University of Toronto, Canada 2 School of Computer Science,
Geometric Component Analysis and its Applications to Data Analysis Amit Bermanis 1, Moshe Salhov 2, and Amir Averbuch 2 1 Department of Mathematics, University of Toronto, Canada 2 School of Computer Science,
Data-dependent representations: Laplacian Eigenmaps
 Data-dependent representations: Laplacian Eigenmaps November 4, 2015 Data Organization and Manifold Learning There are many techniques for Data Organization and Manifold Learning, e.g., Principal Component
Data-dependent representations: Laplacian Eigenmaps November 4, 2015 Data Organization and Manifold Learning There are many techniques for Data Organization and Manifold Learning, e.g., Principal Component
Diffusion/Inference geometries of data features, situational awareness and visualization. Ronald R Coifman Mathematics Yale University
 Diffusion/Inference geometries of data features, situational awareness and visualization Ronald R Coifman Mathematics Yale University Digital data is generally converted to point clouds in high dimensional
Diffusion/Inference geometries of data features, situational awareness and visualization Ronald R Coifman Mathematics Yale University Digital data is generally converted to point clouds in high dimensional
LECTURE NOTE #11 PROF. ALAN YUILLE
 LECTURE NOTE #11 PROF. ALAN YUILLE 1. NonLinear Dimension Reduction Spectral Methods. The basic idea is to assume that the data lies on a manifold/surface in D-dimensional space, see figure (1) Perform
LECTURE NOTE #11 PROF. ALAN YUILLE 1. NonLinear Dimension Reduction Spectral Methods. The basic idea is to assume that the data lies on a manifold/surface in D-dimensional space, see figure (1) Perform
Certifying the Global Optimality of Graph Cuts via Semidefinite Programming: A Theoretic Guarantee for Spectral Clustering
 Certifying the Global Optimality of Graph Cuts via Semidefinite Programming: A Theoretic Guarantee for Spectral Clustering Shuyang Ling Courant Institute of Mathematical Sciences, NYU Aug 13, 2018 Joint
Certifying the Global Optimality of Graph Cuts via Semidefinite Programming: A Theoretic Guarantee for Spectral Clustering Shuyang Ling Courant Institute of Mathematical Sciences, NYU Aug 13, 2018 Joint
Statistical Machine Learning
 Statistical Machine Learning Christoph Lampert Spring Semester 2015/2016 // Lecture 12 1 / 36 Unsupervised Learning Dimensionality Reduction 2 / 36 Dimensionality Reduction Given: data X = {x 1,..., x
Statistical Machine Learning Christoph Lampert Spring Semester 2015/2016 // Lecture 12 1 / 36 Unsupervised Learning Dimensionality Reduction 2 / 36 Dimensionality Reduction Given: data X = {x 1,..., x
Apprentissage non supervisée
 Apprentissage non supervisée Cours 3 Higher dimensions Jairo Cugliari Master ECD 2015-2016 From low to high dimension Density estimation Histograms and KDE Calibration can be done automacally But! Let
Apprentissage non supervisée Cours 3 Higher dimensions Jairo Cugliari Master ECD 2015-2016 From low to high dimension Density estimation Histograms and KDE Calibration can be done automacally But! Let
Beyond Scalar Affinities for Network Analysis or Vector Diffusion Maps and the Connection Laplacian
 Beyond Scalar Affinities for Network Analysis or Vector Diffusion Maps and the Connection Laplacian Amit Singer Princeton University Department of Mathematics and Program in Applied and Computational Mathematics
Beyond Scalar Affinities for Network Analysis or Vector Diffusion Maps and the Connection Laplacian Amit Singer Princeton University Department of Mathematics and Program in Applied and Computational Mathematics
Graphs, Geometry and Semi-supervised Learning
 Graphs, Geometry and Semi-supervised Learning Mikhail Belkin The Ohio State University, Dept of Computer Science and Engineering and Dept of Statistics Collaborators: Partha Niyogi, Vikas Sindhwani In
Graphs, Geometry and Semi-supervised Learning Mikhail Belkin The Ohio State University, Dept of Computer Science and Engineering and Dept of Statistics Collaborators: Partha Niyogi, Vikas Sindhwani In
Statistical Pattern Recognition
 Statistical Pattern Recognition Feature Extraction Hamid R. Rabiee Jafar Muhammadi, Alireza Ghasemi, Payam Siyari Spring 2014 http://ce.sharif.edu/courses/92-93/2/ce725-2/ Agenda Dimensionality Reduction
Statistical Pattern Recognition Feature Extraction Hamid R. Rabiee Jafar Muhammadi, Alireza Ghasemi, Payam Siyari Spring 2014 http://ce.sharif.edu/courses/92-93/2/ce725-2/ Agenda Dimensionality Reduction
Robust Laplacian Eigenmaps Using Global Information
 Manifold Learning and its Applications: Papers from the AAAI Fall Symposium (FS-9-) Robust Laplacian Eigenmaps Using Global Information Shounak Roychowdhury ECE University of Texas at Austin, Austin, TX
Manifold Learning and its Applications: Papers from the AAAI Fall Symposium (FS-9-) Robust Laplacian Eigenmaps Using Global Information Shounak Roychowdhury ECE University of Texas at Austin, Austin, TX
Unsupervised dimensionality reduction
 Unsupervised dimensionality reduction Guillaume Obozinski Ecole des Ponts - ParisTech SOCN course 2014 Guillaume Obozinski Unsupervised dimensionality reduction 1/30 Outline 1 PCA 2 Kernel PCA 3 Multidimensional
Unsupervised dimensionality reduction Guillaume Obozinski Ecole des Ponts - ParisTech SOCN course 2014 Guillaume Obozinski Unsupervised dimensionality reduction 1/30 Outline 1 PCA 2 Kernel PCA 3 Multidimensional
PCA, Kernel PCA, ICA
 PCA, Kernel PCA, ICA Learning Representations. Dimensionality Reduction. Maria-Florina Balcan 04/08/2015 Big & High-Dimensional Data High-Dimensions = Lot of Features Document classification Features per
PCA, Kernel PCA, ICA Learning Representations. Dimensionality Reduction. Maria-Florina Balcan 04/08/2015 Big & High-Dimensional Data High-Dimensions = Lot of Features Document classification Features per
Face Recognition Using Laplacianfaces He et al. (IEEE Trans PAMI, 2005) presented by Hassan A. Kingravi
 Face Recognition Using Laplacianfaces He et al. (IEEE Trans PAMI, 2005) presented by Hassan A. Kingravi Overview Introduction Linear Methods for Dimensionality Reduction Nonlinear Methods and Manifold
Face Recognition Using Laplacianfaces He et al. (IEEE Trans PAMI, 2005) presented by Hassan A. Kingravi Overview Introduction Linear Methods for Dimensionality Reduction Nonlinear Methods and Manifold
MACHINE LEARNING. Methods for feature extraction and reduction of dimensionality: Probabilistic PCA and kernel PCA
 1 MACHINE LEARNING Methods for feature extraction and reduction of dimensionality: Probabilistic PCA and kernel PCA 2 Practicals Next Week Next Week, Practical Session on Computer Takes Place in Room GR
1 MACHINE LEARNING Methods for feature extraction and reduction of dimensionality: Probabilistic PCA and kernel PCA 2 Practicals Next Week Next Week, Practical Session on Computer Takes Place in Room GR
From graph to manifold Laplacian: The convergence rate
 Appl. Comput. Harmon. Anal. 2 (2006) 28 34 www.elsevier.com/locate/acha Letter to the Editor From graph to manifold Laplacian: The convergence rate A. Singer Department of athematics, Yale University,
Appl. Comput. Harmon. Anal. 2 (2006) 28 34 www.elsevier.com/locate/acha Letter to the Editor From graph to manifold Laplacian: The convergence rate A. Singer Department of athematics, Yale University,
Learning gradients: prescriptive models
 Department of Statistical Science Institute for Genome Sciences & Policy Department of Computer Science Duke University May 11, 2007 Relevant papers Learning Coordinate Covariances via Gradients. Sayan
Department of Statistical Science Institute for Genome Sciences & Policy Department of Computer Science Duke University May 11, 2007 Relevant papers Learning Coordinate Covariances via Gradients. Sayan
Data dependent operators for the spatial-spectral fusion problem
 Data dependent operators for the spatial-spectral fusion problem Wien, December 3, 2012 Joint work with: University of Maryland: J. J. Benedetto, J. A. Dobrosotskaya, T. Doster, K. W. Duke, M. Ehler, A.
Data dependent operators for the spatial-spectral fusion problem Wien, December 3, 2012 Joint work with: University of Maryland: J. J. Benedetto, J. A. Dobrosotskaya, T. Doster, K. W. Duke, M. Ehler, A.
Global (ISOMAP) versus Local (LLE) Methods in Nonlinear Dimensionality Reduction
 Global (ISOMAP) versus Local (LLE) Methods in Nonlinear Dimensionality Reduction A presentation by Evan Ettinger on a Paper by Vin de Silva and Joshua B. Tenenbaum May 12, 2005 Outline Introduction The
Global (ISOMAP) versus Local (LLE) Methods in Nonlinear Dimensionality Reduction A presentation by Evan Ettinger on a Paper by Vin de Silva and Joshua B. Tenenbaum May 12, 2005 Outline Introduction The
Principal Component Analysis
 Machine Learning Michaelmas 2017 James Worrell Principal Component Analysis 1 Introduction 1.1 Goals of PCA Principal components analysis (PCA) is a dimensionality reduction technique that can be used
Machine Learning Michaelmas 2017 James Worrell Principal Component Analysis 1 Introduction 1.1 Goals of PCA Principal components analysis (PCA) is a dimensionality reduction technique that can be used
THE HIDDEN CONVEXITY OF SPECTRAL CLUSTERING
 THE HIDDEN CONVEXITY OF SPECTRAL CLUSTERING Luis Rademacher, Ohio State University, Computer Science and Engineering. Joint work with Mikhail Belkin and James Voss This talk A new approach to multi-way
THE HIDDEN CONVEXITY OF SPECTRAL CLUSTERING Luis Rademacher, Ohio State University, Computer Science and Engineering. Joint work with Mikhail Belkin and James Voss This talk A new approach to multi-way
Manifold Regularization
 9.520: Statistical Learning Theory and Applications arch 3rd, 200 anifold Regularization Lecturer: Lorenzo Rosasco Scribe: Hooyoung Chung Introduction In this lecture we introduce a class of learning algorithms,
9.520: Statistical Learning Theory and Applications arch 3rd, 200 anifold Regularization Lecturer: Lorenzo Rosasco Scribe: Hooyoung Chung Introduction In this lecture we introduce a class of learning algorithms,
Diffusion Geometries, Diffusion Wavelets and Harmonic Analysis of large data sets.
 Diffusion Geometries, Diffusion Wavelets and Harmonic Analysis of large data sets. R.R. Coifman, S. Lafon, MM Mathematics Department Program of Applied Mathematics. Yale University Motivations The main
Diffusion Geometries, Diffusion Wavelets and Harmonic Analysis of large data sets. R.R. Coifman, S. Lafon, MM Mathematics Department Program of Applied Mathematics. Yale University Motivations The main
Introduction to Machine Learning
 10-701 Introduction to Machine Learning PCA Slides based on 18-661 Fall 2018 PCA Raw data can be Complex, High-dimensional To understand a phenomenon we measure various related quantities If we knew what
10-701 Introduction to Machine Learning PCA Slides based on 18-661 Fall 2018 PCA Raw data can be Complex, High-dimensional To understand a phenomenon we measure various related quantities If we knew what
L26: Advanced dimensionality reduction
 L26: Advanced dimensionality reduction The snapshot CA approach Oriented rincipal Components Analysis Non-linear dimensionality reduction (manifold learning) ISOMA Locally Linear Embedding CSCE 666 attern
L26: Advanced dimensionality reduction The snapshot CA approach Oriented rincipal Components Analysis Non-linear dimensionality reduction (manifold learning) ISOMA Locally Linear Embedding CSCE 666 attern
Bi-stochastic kernels via asymmetric affinity functions
 Bi-stochastic kernels via asymmetric affinity functions Ronald R. Coifman, Matthew J. Hirn Yale University Department of Mathematics P.O. Box 208283 New Haven, Connecticut 06520-8283 USA ariv:1209.0237v4
Bi-stochastic kernels via asymmetric affinity functions Ronald R. Coifman, Matthew J. Hirn Yale University Department of Mathematics P.O. Box 208283 New Haven, Connecticut 06520-8283 USA ariv:1209.0237v4
Chapter 3. Riemannian Manifolds - I. The subject of this thesis is to extend the combinatorial curve reconstruction approach to curves
 Chapter 3 Riemannian Manifolds - I The subject of this thesis is to extend the combinatorial curve reconstruction approach to curves embedded in Riemannian manifolds. A Riemannian manifold is an abstraction
Chapter 3 Riemannian Manifolds - I The subject of this thesis is to extend the combinatorial curve reconstruction approach to curves embedded in Riemannian manifolds. A Riemannian manifold is an abstraction
Laplacian Eigenmaps for Dimensionality Reduction and Data Representation
 Introduction and Data Representation Mikhail Belkin & Partha Niyogi Department of Electrical Engieering University of Minnesota Mar 21, 2017 1/22 Outline Introduction 1 Introduction 2 3 4 Connections to
Introduction and Data Representation Mikhail Belkin & Partha Niyogi Department of Electrical Engieering University of Minnesota Mar 21, 2017 1/22 Outline Introduction 1 Introduction 2 3 4 Connections to
Lecture: Some Practical Considerations (3 of 4)
 Stat260/CS294: Spectral Graph Methods Lecture 14-03/10/2015 Lecture: Some Practical Considerations (3 of 4) Lecturer: Michael Mahoney Scribe: Michael Mahoney Warning: these notes are still very rough.
Stat260/CS294: Spectral Graph Methods Lecture 14-03/10/2015 Lecture: Some Practical Considerations (3 of 4) Lecturer: Michael Mahoney Scribe: Michael Mahoney Warning: these notes are still very rough.
Discriminative Direction for Kernel Classifiers
 Discriminative Direction for Kernel Classifiers Polina Golland Artificial Intelligence Lab Massachusetts Institute of Technology Cambridge, MA 02139 polina@ai.mit.edu Abstract In many scientific and engineering
Discriminative Direction for Kernel Classifiers Polina Golland Artificial Intelligence Lab Massachusetts Institute of Technology Cambridge, MA 02139 polina@ai.mit.edu Abstract In many scientific and engineering
Unsupervised learning: beyond simple clustering and PCA
 Unsupervised learning: beyond simple clustering and PCA Liza Rebrova Self organizing maps (SOM) Goal: approximate data points in R p by a low-dimensional manifold Unlike PCA, the manifold does not have
Unsupervised learning: beyond simple clustering and PCA Liza Rebrova Self organizing maps (SOM) Goal: approximate data points in R p by a low-dimensional manifold Unlike PCA, the manifold does not have
EECS 275 Matrix Computation
 EECS 275 Matrix Computation Ming-Hsuan Yang Electrical Engineering and Computer Science University of California at Merced Merced, CA 95344 http://faculty.ucmerced.edu/mhyang Lecture 23 1 / 27 Overview
EECS 275 Matrix Computation Ming-Hsuan Yang Electrical Engineering and Computer Science University of California at Merced Merced, CA 95344 http://faculty.ucmerced.edu/mhyang Lecture 23 1 / 27 Overview
Laplacian Eigenmaps for Dimensionality Reduction and Data Representation
 Laplacian Eigenmaps for Dimensionality Reduction and Data Representation Neural Computation, June 2003; 15 (6):1373-1396 Presentation for CSE291 sp07 M. Belkin 1 P. Niyogi 2 1 University of Chicago, Department
Laplacian Eigenmaps for Dimensionality Reduction and Data Representation Neural Computation, June 2003; 15 (6):1373-1396 Presentation for CSE291 sp07 M. Belkin 1 P. Niyogi 2 1 University of Chicago, Department
CSE 291. Assignment Spectral clustering versus k-means. Out: Wed May 23 Due: Wed Jun 13
 CSE 291. Assignment 3 Out: Wed May 23 Due: Wed Jun 13 3.1 Spectral clustering versus k-means Download the rings data set for this problem from the course web site. The data is stored in MATLAB format as
CSE 291. Assignment 3 Out: Wed May 23 Due: Wed Jun 13 3.1 Spectral clustering versus k-means Download the rings data set for this problem from the course web site. The data is stored in MATLAB format as
Spectral Clustering. by HU Pili. June 16, 2013
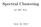 Spectral Clustering by HU Pili June 16, 2013 Outline Clustering Problem Spectral Clustering Demo Preliminaries Clustering: K-means Algorithm Dimensionality Reduction: PCA, KPCA. Spectral Clustering Framework
Spectral Clustering by HU Pili June 16, 2013 Outline Clustering Problem Spectral Clustering Demo Preliminaries Clustering: K-means Algorithm Dimensionality Reduction: PCA, KPCA. Spectral Clustering Framework
SPECTRAL CLUSTERING AND KERNEL PRINCIPAL COMPONENT ANALYSIS ARE PURSUING GOOD PROJECTIONS
 SPECTRAL CLUSTERING AND KERNEL PRINCIPAL COMPONENT ANALYSIS ARE PURSUING GOOD PROJECTIONS VIKAS CHANDRAKANT RAYKAR DECEMBER 5, 24 Abstract. We interpret spectral clustering algorithms in the light of unsupervised
SPECTRAL CLUSTERING AND KERNEL PRINCIPAL COMPONENT ANALYSIS ARE PURSUING GOOD PROJECTIONS VIKAS CHANDRAKANT RAYKAR DECEMBER 5, 24 Abstract. We interpret spectral clustering algorithms in the light of unsupervised
Statistical Learning. Dong Liu. Dept. EEIS, USTC
 Statistical Learning Dong Liu Dept. EEIS, USTC Chapter 6. Unsupervised and Semi-Supervised Learning 1. Unsupervised learning 2. k-means 3. Gaussian mixture model 4. Other approaches to clustering 5. Principle
Statistical Learning Dong Liu Dept. EEIS, USTC Chapter 6. Unsupervised and Semi-Supervised Learning 1. Unsupervised learning 2. k-means 3. Gaussian mixture model 4. Other approaches to clustering 5. Principle
Intrinsic Structure Study on Whale Vocalizations
 1 2015 DCLDE Conference Intrinsic Structure Study on Whale Vocalizations Yin Xian 1, Xiaobai Sun 2, Yuan Zhang 3, Wenjing Liao 3 Doug Nowacek 1,4, Loren Nolte 1, Robert Calderbank 1,2,3 1 Department of
1 2015 DCLDE Conference Intrinsic Structure Study on Whale Vocalizations Yin Xian 1, Xiaobai Sun 2, Yuan Zhang 3, Wenjing Liao 3 Doug Nowacek 1,4, Loren Nolte 1, Robert Calderbank 1,2,3 1 Department of
A Cross-Associative Neural Network for SVD of Nonsquared Data Matrix in Signal Processing
 IEEE TRANSACTIONS ON NEURAL NETWORKS, VOL. 12, NO. 5, SEPTEMBER 2001 1215 A Cross-Associative Neural Network for SVD of Nonsquared Data Matrix in Signal Processing Da-Zheng Feng, Zheng Bao, Xian-Da Zhang
IEEE TRANSACTIONS ON NEURAL NETWORKS, VOL. 12, NO. 5, SEPTEMBER 2001 1215 A Cross-Associative Neural Network for SVD of Nonsquared Data Matrix in Signal Processing Da-Zheng Feng, Zheng Bao, Xian-Da Zhang
PARAMETERIZATION OF NON-LINEAR MANIFOLDS
 PARAMETERIZATION OF NON-LINEAR MANIFOLDS C. W. GEAR DEPARTMENT OF CHEMICAL AND BIOLOGICAL ENGINEERING PRINCETON UNIVERSITY, PRINCETON, NJ E-MAIL:WGEAR@PRINCETON.EDU Abstract. In this report we consider
PARAMETERIZATION OF NON-LINEAR MANIFOLDS C. W. GEAR DEPARTMENT OF CHEMICAL AND BIOLOGICAL ENGINEERING PRINCETON UNIVERSITY, PRINCETON, NJ E-MAIL:WGEAR@PRINCETON.EDU Abstract. In this report we consider
Manifold Learning: Theory and Applications to HRI
 Manifold Learning: Theory and Applications to HRI Seungjin Choi Department of Computer Science Pohang University of Science and Technology, Korea seungjin@postech.ac.kr August 19, 2008 1 / 46 Greek Philosopher
Manifold Learning: Theory and Applications to HRI Seungjin Choi Department of Computer Science Pohang University of Science and Technology, Korea seungjin@postech.ac.kr August 19, 2008 1 / 46 Greek Philosopher
Manifold Coarse Graining for Online Semi-supervised Learning
 for Online Semi-supervised Learning Mehrdad Farajtabar, Amirreza Shaban, Hamid R. Rabiee, Mohammad H. Rohban Digital Media Lab, Department of Computer Engineering, Sharif University of Technology, Tehran,
for Online Semi-supervised Learning Mehrdad Farajtabar, Amirreza Shaban, Hamid R. Rabiee, Mohammad H. Rohban Digital Media Lab, Department of Computer Engineering, Sharif University of Technology, Tehran,
High-Dimensional Pattern Recognition using Low-Dimensional Embedding and Earth Mover s Distance
 High-Dimensional Pattern Recognition using Low-Dimensional Embedding and Earth Mover s Distance Linh Lieu,a, Naoki Saito a a Department of Mathematics, University of California, Davis, CA 95616, USA Abstract
High-Dimensional Pattern Recognition using Low-Dimensional Embedding and Earth Mover s Distance Linh Lieu,a, Naoki Saito a a Department of Mathematics, University of California, Davis, CA 95616, USA Abstract
Notes on Latent Semantic Analysis
 Notes on Latent Semantic Analysis Costas Boulis 1 Introduction One of the most fundamental problems of information retrieval (IR) is to find all documents (and nothing but those) that are semantically
Notes on Latent Semantic Analysis Costas Boulis 1 Introduction One of the most fundamental problems of information retrieval (IR) is to find all documents (and nothing but those) that are semantically
Signal processing methods have significantly changed. Diffusion Maps. Signal Processing
 [ Ronen Talmon, Israel Cohen, Sharon Gannot, and Ronald R. Coifman ] Diffusion Maps for Signal Processing Advances in Kernel-based Learning for Signal Processing istockphoto.com/ aleksandar velasevic [
[ Ronen Talmon, Israel Cohen, Sharon Gannot, and Ronald R. Coifman ] Diffusion Maps for Signal Processing Advances in Kernel-based Learning for Signal Processing istockphoto.com/ aleksandar velasevic [
15 Singular Value Decomposition
 15 Singular Value Decomposition For any high-dimensional data analysis, one s first thought should often be: can I use an SVD? The singular value decomposition is an invaluable analysis tool for dealing
15 Singular Value Decomposition For any high-dimensional data analysis, one s first thought should often be: can I use an SVD? The singular value decomposition is an invaluable analysis tool for dealing
Welcome to Copenhagen!
 Welcome to Copenhagen! Schedule: Monday Tuesday Wednesday Thursday Friday 8 Registration and welcome 9 Crash course on Crash course on Introduction to Differential and Differential and Information Geometry
Welcome to Copenhagen! Schedule: Monday Tuesday Wednesday Thursday Friday 8 Registration and welcome 9 Crash course on Crash course on Introduction to Differential and Differential and Information Geometry
Laplace-Beltrami Eigenfunctions for Deformation Invariant Shape Representation
 Laplace-Beltrami Eigenfunctions for Deformation Invariant Shape Representation Author: Raif M. Rustamov Presenter: Dan Abretske Johns Hopkins 2007 Outline Motivation and Background Laplace-Beltrami Operator
Laplace-Beltrami Eigenfunctions for Deformation Invariant Shape Representation Author: Raif M. Rustamov Presenter: Dan Abretske Johns Hopkins 2007 Outline Motivation and Background Laplace-Beltrami Operator
Nonlinear Manifold Learning Summary
 Nonlinear Manifold Learning 6.454 Summary Alexander Ihler ihler@mit.edu October 6, 2003 Abstract Manifold learning is the process of estimating a low-dimensional structure which underlies a collection
Nonlinear Manifold Learning 6.454 Summary Alexander Ihler ihler@mit.edu October 6, 2003 Abstract Manifold learning is the process of estimating a low-dimensional structure which underlies a collection
Learning from patches by efficient spectral decomposition of a structured kernel
 Mach Learn (2016) 103:81 102 DOI 10.1007/s10994-015-5538-4 Learning from patches by efficient spectral decomposition of a structured kernel Moshe Salhov 1,2 Amit Bermanis 1 Guy Wolf 1,2 Amir Averbuch 1,2
Mach Learn (2016) 103:81 102 DOI 10.1007/s10994-015-5538-4 Learning from patches by efficient spectral decomposition of a structured kernel Moshe Salhov 1,2 Amit Bermanis 1 Guy Wolf 1,2 Amir Averbuch 1,2
Supplemental Materials for. Local Multidimensional Scaling for. Nonlinear Dimension Reduction, Graph Drawing. and Proximity Analysis
 Supplemental Materials for Local Multidimensional Scaling for Nonlinear Dimension Reduction, Graph Drawing and Proximity Analysis Lisha Chen and Andreas Buja Yale University and University of Pennsylvania
Supplemental Materials for Local Multidimensional Scaling for Nonlinear Dimension Reduction, Graph Drawing and Proximity Analysis Lisha Chen and Andreas Buja Yale University and University of Pennsylvania
II. DIFFERENTIABLE MANIFOLDS. Washington Mio CENTER FOR APPLIED VISION AND IMAGING SCIENCES
 II. DIFFERENTIABLE MANIFOLDS Washington Mio Anuj Srivastava and Xiuwen Liu (Illustrations by D. Badlyans) CENTER FOR APPLIED VISION AND IMAGING SCIENCES Florida State University WHY MANIFOLDS? Non-linearity
II. DIFFERENTIABLE MANIFOLDS Washington Mio Anuj Srivastava and Xiuwen Liu (Illustrations by D. Badlyans) CENTER FOR APPLIED VISION AND IMAGING SCIENCES Florida State University WHY MANIFOLDS? Non-linearity
Iterative Laplacian Score for Feature Selection
 Iterative Laplacian Score for Feature Selection Linling Zhu, Linsong Miao, and Daoqiang Zhang College of Computer Science and echnology, Nanjing University of Aeronautics and Astronautics, Nanjing 2006,
Iterative Laplacian Score for Feature Selection Linling Zhu, Linsong Miao, and Daoqiang Zhang College of Computer Science and echnology, Nanjing University of Aeronautics and Astronautics, Nanjing 2006,
The Curse of Dimensionality for Local Kernel Machines
 The Curse of Dimensionality for Local Kernel Machines Yoshua Bengio, Olivier Delalleau & Nicolas Le Roux April 7th 2005 Yoshua Bengio, Olivier Delalleau & Nicolas Le Roux Snowbird Learning Workshop Perspective
The Curse of Dimensionality for Local Kernel Machines Yoshua Bengio, Olivier Delalleau & Nicolas Le Roux April 7th 2005 Yoshua Bengio, Olivier Delalleau & Nicolas Le Roux Snowbird Learning Workshop Perspective
Theorem 2. Let n 0 3 be a given integer. is rigid in the sense of Guillemin, so are all the spaces ḠR n,n, with n n 0.
 This monograph is motivated by a fundamental rigidity problem in Riemannian geometry: determine whether the metric of a given Riemannian symmetric space of compact type can be characterized by means of
This monograph is motivated by a fundamental rigidity problem in Riemannian geometry: determine whether the metric of a given Riemannian symmetric space of compact type can be characterized by means of
Clustering in kernel embedding spaces and organization of documents
 Clustering in kernel embedding spaces and organization of documents Stéphane Lafon Collaborators: Raphy Coifman (Yale), Yosi Keller (Yale), Ioannis G. Kevrekidis (Princeton), Ann B. Lee (CMU), Boaz Nadler
Clustering in kernel embedding spaces and organization of documents Stéphane Lafon Collaborators: Raphy Coifman (Yale), Yosi Keller (Yale), Ioannis G. Kevrekidis (Princeton), Ann B. Lee (CMU), Boaz Nadler
Linear Algebra. Preliminary Lecture Notes
 Linear Algebra Preliminary Lecture Notes Adolfo J. Rumbos c Draft date May 9, 29 2 Contents 1 Motivation for the course 5 2 Euclidean n dimensional Space 7 2.1 Definition of n Dimensional Euclidean Space...........
Linear Algebra Preliminary Lecture Notes Adolfo J. Rumbos c Draft date May 9, 29 2 Contents 1 Motivation for the course 5 2 Euclidean n dimensional Space 7 2.1 Definition of n Dimensional Euclidean Space...........
Is Manifold Learning for Toy Data only?
 s Manifold Learning for Toy Data only? Marina Meilă University of Washington mmp@stat.washington.edu MMDS Workshop 2016 Outline What is non-linear dimension reduction? Metric Manifold Learning Estimating
s Manifold Learning for Toy Data only? Marina Meilă University of Washington mmp@stat.washington.edu MMDS Workshop 2016 Outline What is non-linear dimension reduction? Metric Manifold Learning Estimating
COMS 4721: Machine Learning for Data Science Lecture 19, 4/6/2017
 COMS 4721: Machine Learning for Data Science Lecture 19, 4/6/2017 Prof. John Paisley Department of Electrical Engineering & Data Science Institute Columbia University PRINCIPAL COMPONENT ANALYSIS DIMENSIONALITY
COMS 4721: Machine Learning for Data Science Lecture 19, 4/6/2017 Prof. John Paisley Department of Electrical Engineering & Data Science Institute Columbia University PRINCIPAL COMPONENT ANALYSIS DIMENSIONALITY
EE Technion, Spring then. can be isometrically embedded into can be realized as a Gram matrix of rank, Properties:
 5/25/200 2 A mathematical exercise Assume points with the metric are isometrically embeddable into Then, there exists a canonical form such that for all Spectral methods We can also write Alexander & Michael
5/25/200 2 A mathematical exercise Assume points with the metric are isometrically embeddable into Then, there exists a canonical form such that for all Spectral methods We can also write Alexander & Michael
Exploiting Sparse Non-Linear Structure in Astronomical Data
 Exploiting Sparse Non-Linear Structure in Astronomical Data Ann B. Lee Department of Statistics and Department of Machine Learning, Carnegie Mellon University Joint work with P. Freeman, C. Schafer, and
Exploiting Sparse Non-Linear Structure in Astronomical Data Ann B. Lee Department of Statistics and Department of Machine Learning, Carnegie Mellon University Joint work with P. Freeman, C. Schafer, and
Lecture Notes 1: Vector spaces
 Optimization-based data analysis Fall 2017 Lecture Notes 1: Vector spaces In this chapter we review certain basic concepts of linear algebra, highlighting their application to signal processing. 1 Vector
Optimization-based data analysis Fall 2017 Lecture Notes 1: Vector spaces In this chapter we review certain basic concepts of linear algebra, highlighting their application to signal processing. 1 Vector
Filtering via a Reference Set. A.Haddad, D. Kushnir, R.R. Coifman Technical Report YALEU/DCS/TR-1441 February 21, 2011
 Patch-based de-noising algorithms and patch manifold smoothing have emerged as efficient de-noising methods. This paper provides a new insight on these methods, such as the Non Local Means or the image
Patch-based de-noising algorithms and patch manifold smoothing have emerged as efficient de-noising methods. This paper provides a new insight on these methods, such as the Non Local Means or the image
Wolf-Tilo Balke Silviu Homoceanu Institut für Informationssysteme Technische Universität Braunschweig
 Multimedia Databases Wolf-Tilo Balke Silviu Homoceanu Institut für Informationssysteme Technische Universität Braunschweig http://www.ifis.cs.tu-bs.de 14 Indexes for Multimedia Data 14 Indexes for Multimedia
Multimedia Databases Wolf-Tilo Balke Silviu Homoceanu Institut für Informationssysteme Technische Universität Braunschweig http://www.ifis.cs.tu-bs.de 14 Indexes for Multimedia Data 14 Indexes for Multimedia
Principal Component Analysis
 Principal Component Analysis Yingyu Liang yliang@cs.wisc.edu Computer Sciences Department University of Wisconsin, Madison [based on slides from Nina Balcan] slide 1 Goals for the lecture you should understand
Principal Component Analysis Yingyu Liang yliang@cs.wisc.edu Computer Sciences Department University of Wisconsin, Madison [based on slides from Nina Balcan] slide 1 Goals for the lecture you should understand
Manifold Learning and it s application
 Manifold Learning and it s application Nandan Dubey SE367 Outline 1 Introduction Manifold Examples image as vector Importance Dimension Reduction Techniques 2 Linear Methods PCA Example MDS Perception
Manifold Learning and it s application Nandan Dubey SE367 Outline 1 Introduction Manifold Examples image as vector Importance Dimension Reduction Techniques 2 Linear Methods PCA Example MDS Perception
APPENDIX A. Background Mathematics. A.1 Linear Algebra. Vector algebra. Let x denote the n-dimensional column vector with components x 1 x 2.
 APPENDIX A Background Mathematics A. Linear Algebra A.. Vector algebra Let x denote the n-dimensional column vector with components 0 x x 2 B C @. A x n Definition 6 (scalar product). The scalar product
APPENDIX A Background Mathematics A. Linear Algebra A.. Vector algebra Let x denote the n-dimensional column vector with components 0 x x 2 B C @. A x n Definition 6 (scalar product). The scalar product
Machine Learning (BSMC-GA 4439) Wenke Liu
 Machine Learning (BSMC-GA 4439) Wenke Liu 02-01-2018 Biomedical data are usually high-dimensional Number of samples (n) is relatively small whereas number of features (p) can be large Sometimes p>>n Problems
Machine Learning (BSMC-GA 4439) Wenke Liu 02-01-2018 Biomedical data are usually high-dimensional Number of samples (n) is relatively small whereas number of features (p) can be large Sometimes p>>n Problems
Dimensionality Reduc1on
 Dimensionality Reduc1on contd Aarti Singh Machine Learning 10-601 Nov 10, 2011 Slides Courtesy: Tom Mitchell, Eric Xing, Lawrence Saul 1 Principal Component Analysis (PCA) Principal Components are the
Dimensionality Reduc1on contd Aarti Singh Machine Learning 10-601 Nov 10, 2011 Slides Courtesy: Tom Mitchell, Eric Xing, Lawrence Saul 1 Principal Component Analysis (PCA) Principal Components are the
8.1 Concentration inequality for Gaussian random matrix (cont d)
 MGMT 69: Topics in High-dimensional Data Analysis Falll 26 Lecture 8: Spectral clustering and Laplacian matrices Lecturer: Jiaming Xu Scribe: Hyun-Ju Oh and Taotao He, October 4, 26 Outline Concentration
MGMT 69: Topics in High-dimensional Data Analysis Falll 26 Lecture 8: Spectral clustering and Laplacian matrices Lecturer: Jiaming Xu Scribe: Hyun-Ju Oh and Taotao He, October 4, 26 Outline Concentration
Manifold Approximation by Moving Least-Squares Projection (MMLS)
 Manifold Approximation by Moving Least-Squares Projection (MMLS) Barak Sober David Levin School of Applied Mathematics, Tel Aviv University, Israel 2018-01-17 arxiv:1606.07104v4 [cs.gr] 16 Jan 2018 Abstract
Manifold Approximation by Moving Least-Squares Projection (MMLS) Barak Sober David Levin School of Applied Mathematics, Tel Aviv University, Israel 2018-01-17 arxiv:1606.07104v4 [cs.gr] 16 Jan 2018 Abstract
CALCULUS ON MANIFOLDS. 1. Riemannian manifolds Recall that for any smooth manifold M, dim M = n, the union T M =
 CALCULUS ON MANIFOLDS 1. Riemannian manifolds Recall that for any smooth manifold M, dim M = n, the union T M = a M T am, called the tangent bundle, is itself a smooth manifold, dim T M = 2n. Example 1.
CALCULUS ON MANIFOLDS 1. Riemannian manifolds Recall that for any smooth manifold M, dim M = n, the union T M = a M T am, called the tangent bundle, is itself a smooth manifold, dim T M = 2n. Example 1.
Lecture 2: Linear Algebra Review
 EE 227A: Convex Optimization and Applications January 19 Lecture 2: Linear Algebra Review Lecturer: Mert Pilanci Reading assignment: Appendix C of BV. Sections 2-6 of the web textbook 1 2.1 Vectors 2.1.1
EE 227A: Convex Optimization and Applications January 19 Lecture 2: Linear Algebra Review Lecturer: Mert Pilanci Reading assignment: Appendix C of BV. Sections 2-6 of the web textbook 1 2.1 Vectors 2.1.1
22 : Hilbert Space Embeddings of Distributions
 10-708: Probabilistic Graphical Models 10-708, Spring 2014 22 : Hilbert Space Embeddings of Distributions Lecturer: Eric P. Xing Scribes: Sujay Kumar Jauhar and Zhiguang Huo 1 Introduction and Motivation
10-708: Probabilistic Graphical Models 10-708, Spring 2014 22 : Hilbert Space Embeddings of Distributions Lecturer: Eric P. Xing Scribes: Sujay Kumar Jauhar and Zhiguang Huo 1 Introduction and Motivation
4 Bias-Variance for Ridge Regression (24 points)
 Implement Ridge Regression with λ = 0.00001. Plot the Squared Euclidean test error for the following values of k (the dimensions you reduce to): k = {0, 50, 100, 150, 200, 250, 300, 350, 400, 450, 500,
Implement Ridge Regression with λ = 0.00001. Plot the Squared Euclidean test error for the following values of k (the dimensions you reduce to): k = {0, 50, 100, 150, 200, 250, 300, 350, 400, 450, 500,
An indicator for the number of clusters using a linear map to simplex structure
 An indicator for the number of clusters using a linear map to simplex structure Marcus Weber, Wasinee Rungsarityotin, and Alexander Schliep Zuse Institute Berlin ZIB Takustraße 7, D-495 Berlin, Germany
An indicator for the number of clusters using a linear map to simplex structure Marcus Weber, Wasinee Rungsarityotin, and Alexander Schliep Zuse Institute Berlin ZIB Takustraße 7, D-495 Berlin, Germany
Distance Preservation - Part 2
 Distance Preservation - Part 2 Graph Distances Niko Vuokko October 9th 2007 NLDR Seminar Outline Introduction Geodesic and graph distances From linearity to nonlinearity Isomap Geodesic NLM Curvilinear
Distance Preservation - Part 2 Graph Distances Niko Vuokko October 9th 2007 NLDR Seminar Outline Introduction Geodesic and graph distances From linearity to nonlinearity Isomap Geodesic NLM Curvilinear
A Statistical Look at Spectral Graph Analysis. Deep Mukhopadhyay
 A Statistical Look at Spectral Graph Analysis Deep Mukhopadhyay Department of Statistics, Temple University Office: Speakman 335 deep@temple.edu http://sites.temple.edu/deepstat/ Graph Signal Processing
A Statistical Look at Spectral Graph Analysis Deep Mukhopadhyay Department of Statistics, Temple University Office: Speakman 335 deep@temple.edu http://sites.temple.edu/deepstat/ Graph Signal Processing
Machine Learning. B. Unsupervised Learning B.2 Dimensionality Reduction. Lars Schmidt-Thieme, Nicolas Schilling
 Machine Learning B. Unsupervised Learning B.2 Dimensionality Reduction Lars Schmidt-Thieme, Nicolas Schilling Information Systems and Machine Learning Lab (ISMLL) Institute for Computer Science University
Machine Learning B. Unsupervised Learning B.2 Dimensionality Reduction Lars Schmidt-Thieme, Nicolas Schilling Information Systems and Machine Learning Lab (ISMLL) Institute for Computer Science University
Dimensionality Reduction:
 Dimensionality Reduction: From Data Representation to General Framework Dong XU School of Computer Engineering Nanyang Technological University, Singapore What is Dimensionality Reduction? PCA LDA Examples:
Dimensionality Reduction: From Data Representation to General Framework Dong XU School of Computer Engineering Nanyang Technological University, Singapore What is Dimensionality Reduction? PCA LDA Examples:
Functional Maps ( ) Dr. Emanuele Rodolà Room , Informatik IX
 Functional Maps (12.06.2014) Dr. Emanuele Rodolà rodola@in.tum.de Room 02.09.058, Informatik IX Seminar «LP relaxation for elastic shape matching» Fabian Stark Wednesday, June 18th 14:00 Room 02.09.023
Functional Maps (12.06.2014) Dr. Emanuele Rodolà rodola@in.tum.de Room 02.09.058, Informatik IX Seminar «LP relaxation for elastic shape matching» Fabian Stark Wednesday, June 18th 14:00 Room 02.09.023
