CSCI1950 Z Computa4onal Methods for Biology Lecture 4. Ben Raphael February 2, hhp://cs.brown.edu/courses/csci1950 z/ Algorithm Summary
|
|
|
- Dorothy Price
- 5 years ago
- Views:
Transcription
1 CSCI1950 Z Computa4onal Methods for Biology Lecture 4 Ben Raphael February 2, 2009 hhp://cs.brown.edu/courses/csci1950 z/ Algorithm Summary Parsimony Probabilis4c Method Input Output Sankoff s & Fitch s Alg. Characters, T A, B Perfect Phylogeny Characters A, B, T Felsenstein Characters, T, B A T = tree topology B = branch lengths A = ancestral states 1
2 Pairwise Compa4bility Test (Wilson 1965) Binary characters i and j are pairwise compa4ble if and only if: j is homogenous w.r.t i 0 or i 1. Equivalently: i 1 and j 1 are disjoint or one contains the other Equivalently: all 4 rows do not exist (0,0), (0,1), (1,0), (1,1) i 0 i 1 i A 0 B 0 C 0 D 1 E 1 j A 0 B 0 C 1 D 0 E 0 k A 0 B 0 C 1 D 1 E 0 Pairwise Compa4bility Theorem (Estabrook et al. 1976) A set S of binary characters is mutually compa4ble if and only if all pairs c and c of characters in S are pairwise compa4ble. Pairwise compa4bility mutual compa4bility. 2
3 Perfect Phylogeny A set of mutually compa4ble binary characters gives a perfect phylogeny: 1. Evolu4onary model Binary characters {0,1} Each character changes state only once in evolu4onary history (no homoplasy!). 2. Tree in which every muta4on is on an edge of the tree. All the species in one sub tree contain a 0, and all species in the other contain a 1. For simplicity, assume root = (0, 0, 0, 0, 0) species A B C D E traits Last )me: algorithm to reconstruct a tree. 1 0 Trees and Splits Given a set X, a split is a par44on of X into two non empty subsets A and B. X = A B. For a phylogene4c tree T with leaves L, each edge e defines a split L e = A B, where A and B are the leaves in the subtrees obtained by removing e. i In perfect phylogeny, edges where binary character changes state gave split i 0 and i 1. i 1 i 0 We will return to splits in a future lecture. 3
4 Splits Equivalence Theorem A phylogene4c tree T defines a collec4on of splits Σ(T) = { L e e is edge in T}. Splits A 1 B 1 and A 2 B 2 are pairwise compa3ble if at least one of A 1 A 2, A 1 B 2, B 1 A 2, and B 1 B 2 is the empty set. Splits Equivalence Theorem: Let Σ be a collec4on of splits. There is a phylogene4c tree such that Σ(T) = Σ if and only if the splits in Σ are pairwise compa4ble. The Pairwise Compa4bility Theorem (for binary characters) follows from this theorem. Outline Distance based methods for phylogene4c tree reconstruc4on. Review of distances/metrics. Tree distances and addi4ve distances Small and large phylogeny problems. Non addi4ve distances and clustering UPGMA and ultrametric distances. 4
5 Distances A distance on a set X is a func4on d: X R sa4sfying: d(x, y) 0, with equality iff x = y. For all x, y X, d(x, y) = d(y, x) [symmetry] For all x, y, z X, d(x, z) d(x, y) + d(y, z) [triangle inequality] Examples: X = real numbers, d(x, y) = x y is distance. X = strings over some alphabet. d H (s, t) = number of posi4ons where s and t differ is called Hamming distance. Distances in Biological Data String distances (e.g. Hamming distance, edit distance) on DNA/protein sequence data Subs4tu4on model (Jukes Cantor, Kimura, etc.): scores for par4cular changes A T, C G, etc. Rat: ACAGTCACGCCCCACACGT Mouse: ACAGTGACGCCACACACGT Gorilla: CCTGTGACGTAACAAACGA Chimpanzee: CCTGTGAGGTAGCAAACGA Human: CCTGTGAGGTAGCACACGA 5
6 Distance Matrix For n species, form n x n distance matrix D ij Example: D ij = edit distance between a gene in species i and species j. Mouse: ACAGTGACGCCACACACGT Gorilla: CCTGCGACGTAACAAACGC Chimpanzee: CCTGCCAGTTAGCAAACGC Human: CCTGCCAGTTAGCACACGA Alignment vs. Distance Matrix Mouse: ACAGTGACGCCACACACGT Gorilla: CCTGCGACGTAACAAACGC Chimpanzee: CCTGCCAGTTAGCAAACGC Human: CCTGCCAGTTAGCACACGA Sequence a gene of length m in n species n x m alignment matrix. Reverse transforma4on not possible due to loss of informa4on. Transform into n x n distance matrix 6
7 Distances in Trees Given a tree T with a posi4ve weight w(e) on each edge, we define the tree distance d T on the set L of leaves by: d T (i, j) = sum of weights of edges on unique path from i to j. In evolu4onary biology, weights are some4mes called branch lengths. Distance in Trees: an Example j i d T (1,4) = = 69 7
8 Distance vs. Tree Distance n x n distance matrix for n species Note that d T (i, j), tree distance between i and j, not necessarily equal to D ij as given by distance matrix. Rat: ACAGTGACGCCCCAAACGT Mouse: ACAGTGACGCTACAAACGT Gorilla: CCTGTGACGTAACAAACGA Chimpanzee: CCTGTGACGTAGCAAACGA Human: CCTGTGACGTAGCAAACGA Fivng a Distance Matrix Given n species, we can compute the n x n distance matrix D ij Evolu4on of these species is described by a tree that we don t know. We need an algorithm to construct a tree that best fits the distance matrix D ij Find a tree T such that: Lengths of path in an (unknown) tree T D ij = d T (i,j) Distance between species (known) 8
9 Distance Based Phylogeny Problem Goal: Reconstruct an evolu4onary tree from a distance matrix Input: n x n distance matrix D ij Output: weighted tree T with n leaves fivng D Unknown topology of tree makes evolu4onary tree reconstruc4on hard! # unrooted binary trees n leaves: T(n) = (2n 3)! / ((n 2)! 2 n 2 ) n = 24: T(n) = 5.74 x If D is addi3ve, this problem has a solu4on and there is a simple algorithm to solve it Distance based vs. character based Key difference: Distance based methods do not reconstruct ancestral states. A B C D A B C D Note that C and D are iden4cal. 9
10 Reconstruc4ng a 3 Leaved Tree Tree reconstruc4on for a 3x3 matrix is straighxorward We have 3 leaves i, j, k and a center vertex c Observe: d ic + d jc = D ij d ic + d kc = D ik d jc + d kc = D jk Reconstruc4ng a 3 Leaved Tree (cont d) d ic + d jc = D ij + d ic + d kc = D ik 2d ic + d jc + d kc = D ij + D ik 2d ic + D jk = D ij + D ik d ic = (D ij + D ik D jk )/2 Similarly, d jc = (D ij + D jk D ik )/2 d kc = (D ki + D kj D ij )/2 10
11 Trees with > 3 Leaves A binary tree with n leaves has 2n 3 edges Fivng a given tree to a distance matrix D requires solving a system with n(n 1)/2 equa4ons and 2n 3 variables Solu4on not always possible for n > 3. Addi4ve Distance Matrices Matrix D is ADDITIVE if there exists a tree T with d ij (T) = D ij NON-ADDITIVE otherwise 11
12 Addi4ve Distance Phylogeny Small Addi>ve Distance Phylogeny: Given phylogene4c tree T and distance matrix D, determine branch lengths such that d T (i,j) = D ij. Large Addi>ve Distance Phylogeny: Given distance matrix D, find T and branch lengths such that d T (i,j) = D ij. Both of these problems can be solved efficiently. Reconstruc4ng Addi4ve Distances Given T x D v w x y z v w x z y T w v y 0 14 z 0 If we know T and D, but do not know the length of each edge, we can reconstruct those lengths 12
13 Reconstruc4ng Addi4ve Distances Given T x D v w x y z y T v w z w x v y 0 14 z 0 Reconstruc4ng Addi4ve Distances Given T D v w x y z v w x y 0 14 z 0 z y x Find neighbors v, w (common parent) a w a x y z d ax = ½ (d vx + d wx d vw ) v D 1 a x y 0 14 z 0 d ay = ½ (d vy + d wy d vw ) d az = ½ (d vz + d wz d vw ) 13
14 Reconstruc4ng Addi4ve Distances Given T D 1 a x y z a x y 0 14 z 0 z y 7 4 x 5 b 3 Neighbors x, y (common parent) c 3 a 4 w D 2 a b z a b 0 10 z 0 D 3 a c a 0 3 c 0 6 d(a, c) = 3 d(b, c) = d(a, b) d(a, c) = 3 d(c, z) = d(a, z) d(a, c) = 7 d(b, x) = d(a, x) d(a, b) = 5 d(b, y) = d(a, y) d(a, b) = 4 d(a, w) = d(z, w) d(a, z) = 4 d(a, v) = d(z, v) d(a, z) = 6 Correct!!! v Trees and Neighbors Previous algorithm relied only on finding neighboring leaves: 1. Find neighboring leaves i and j with parent k 2. Remove the rows and columns of i and j 3. Add a new row and column corresponding to k, where the distance from k to any other leaf m can be computed as: D km = (D im + D jm D ij )/2 Compress i and j into k, iterate algorithm for rest of tree 14
15 Finding Neighboring Leaves To find neighboring leaves we simply select a pair of closest leaves. i j k l i j k 0 13 l 0 WRONG! i and j are neighbors, but (d ij = 13) > (d jk = 12). Finding a pair of neighboring leaves is a nontrivial problem! Degenerate Triples A degenerate triple is a set of three dis4nct elements 1 i, j, k n where D ij + D jk = D ik Element j in a degenerate triple i,j,k lies on the evolu4onary path from i to k (or is ahached to this path by an edge of length 0). 15
16 Looking for Degenerate Triples If distance matrix D has a degenerate triple i,j,k then j can be removed from D thus reducing the size of the problem. If distance matrix D does not have a degenerate triple i,j,k, one can create a degenera4ve triple in D by shortening all hanging edges (in the tree). Shortening Hanging Edges to Produce Degenerate Triples Shorten all hanging edges (edges that connect leaves) un4l a degenerate triple is found 16
17 Finding Degenerate Triples If there is no degenerate triple, all hanging edges are reduced by the same amount δ, so that all pair wise distances in the matrix are reduced by 2δ. Eventually this process collapses one of the leaves (when δ = length of shortest hanging edge), forming a degenerate triple i,j,k and reducing the size of the distance matrix D. The ahachment point for j can be recovered in the reverse transforma4ons by saving D ij for each collapsed leaf. Reconstruc4ng Trees for Addi4ve Distance Matrices Trim(D, δ) for all 1 i j n D ij = D ij 2δ 17
18 Addi4vePhylogeny Algorithm AdditivePhylogeny(D) if D is a 2 x 2 matrix T = tree of a single edge of length D 1,2 return T if D is non-degenerate Compute trimming parameter δ Trim(D, δ) Find a triple i, j, k in D such that D ij + D jk = D ik x = D ij Remove j th row and j th column from D T = AdditivePhylogeny(D) Traceback Addi4vePhylogeny (cont d) Traceback Add a new vertex v to T at distance x from i to k Add j back to T by creating an edge (v,j) of length 0 for every leaf l in T if distance from l to v in the tree D l,j output matrix is not additive return Extend all hanging edges by length δ return T Ques>on: How to compute δ? 18
19 Addi4ve Distance How to tell if D is addi4ve? Addi4vePhylogeny provides a way to check if distance matrix D is addi4ve An even more efficient addi>vity check is the four point condi>on The Four Point Condi4on (Zaretskii 1965, Buneman 1971) Let 1 i,j,k,l n be four dis4nct leaves in a tree Compute: 1. D ij + D kl, 2. D ik + D jl, 3. D il + D jk 2 and 3 represent the same number: (length of all edges) + 2 * (length middle edge) represents a smaller number: (length of all edges) (length middle edge) 19
20 The Four Point Condi4on Four point condi>on: Every four leaves (quartet) can be labeled as i,j,k,l such that: D ij + D kl D ik + D jl = D il + D jk Theorem : An n x n matrix D is addi4ve if and only if the four point condi4on holds for every quartet 1 i,j,k,l n The Four Point Condi4on Theorem : An n x n matrix D is addi4ve if and only if the four point condi4on holds for every quartet 1 i,j,k,l n. Proof: Since D addi4ve, D = d T. Find split such that: i, j S 1 and k, l S 2. Define λ m to be weights in tree below. D ik + D jl = (λ 1 + λ 3 + λ 4 ) + (λ 2 + λ 3 + λ 5 ) = D il + D jk >= (λ 1 + λ 2 ) + ( λ 4 + λ 5 ). i λ 1 λ 4 k λ 3 j λ 2 λ 5 l 20
21 Non addi4ve Distances What if there is no tree T such that D ij = d T (i,j). Approaches: 1. Find tree such that minimizes error 2. Heuris4c approaches: clustering. Least Squares Distance Phylogeny Problem If the distance matrix D is NOT addi4ve, then we look for a tree T that approximates D the best: Squared Error : i,j (d ij (T) D ij ) 2 Squared Error is a measure of the quality of the fit between distance matrix and the tree: we want to minimize it. Least Squares Distance Phylogeny Problem: Find approxima4on tree T with minimum squared error for a non addi4ve matrix D. (NP hard) 21
22 Tree construc4on as clustering Pair Group Methods Itera4vely combine closest leaves/groups into larger groups C { {1},, {n} } While C > 2 do [Find closest clusters.] d(c i, C j ) = min d(c i, C j ). C k C i C j [Replace C i and C j by C k.] C (C \ C i \ C j ) C k
23 Pair Group Methods What is d? 1 4 How to define branch lengths? C { {1},, {n} } While C > 2 do [Find closest clusters.] d(c i, C j ) = min d(c i, C j ). C k C i C j [Replace C i and C j by C k.] C (C \ C i \ C j ) C k UPGMA Unweighted Pair Group Method with Averages Distance between clusters defined as average pairwise distance Assigns height to every vertex in the tree, effec4vely da4ng every vertex
24 UPGMA Unweighted Pair Group Method with Averages Distance between clusters defined as average pairwise distance Given two disjoint clusters C i, C j of sequences, 1 d ij = Σ {p Ci, q C j } d pq C i C j UPGMA Unweighted Pair Group Method with Averages Assigns height to every vertex in the tree, effec4vely da4ng every vertex Add a vertex connec4ng C i, C j and place it at height d ij /
25 UPGMA Algorithm Ini>aliza>on: Assign each x i to its own cluster C i Define one leaf per sequence, each at height 0 Itera>on: Find two clusters C i and C j such that d ij is min Let C k = C i C j Add a vertex connec4ng C i, C j and place it at height d ij /2 Delete C i and C j Termina>on: When a single cluster remains Trees from UPGMA UPGMA produces an ultrametric tree; distance from the root to any leaf is the same The Molecular Clock: The evolu4onary distance between species x and y is twice the Earth 4me to reach the nearest common ancestor That is, the molecular clock has constant rate in all species years The molecular clock results in ultrametric distances 25
26 UPGMA s Weakness: Example Correct tree UPGMA Ultrametrics D ij is an ultrametric provided for all species i, j, k (dis4nct leaves of tree) two of the distances D ij, D jk and D ik are equal and the third. Ex. d(i,k) = d(j, k) d(i, j) i j λ 1 λ 1 λ 2 λ k 1 + λ 2 2 λ 1 Thus λ 2 λ 1 Proposi>on: If d is ultrametric, then d is addi4ve. 26
27 Ultrametrics Both addi4ve distance phylogeny and perfect phylogeny can be reduced to the ultrametric phylogeny problem. Let v = row of D containing largest entry m v. Define D ij = m v + (D ij D vi D vj ) / 2 i = m v λ 3 λ 1 λ 3 v Theorem: D is addi4ve if and only if D is ultrametric. (See Gusfield, Ch. 17) j λ 2 27
CSCI1950 Z Computa4onal Methods for Biology Lecture 5
 CSCI1950 Z Computa4onal Methods for Biology Lecture 5 Ben Raphael February 6, 2009 hip://cs.brown.edu/courses/csci1950 z/ Alignment vs. Distance Matrix Mouse: ACAGTGACGCCACACACGT Gorilla: CCTGCGACGTAACAAACGC
CSCI1950 Z Computa4onal Methods for Biology Lecture 5 Ben Raphael February 6, 2009 hip://cs.brown.edu/courses/csci1950 z/ Alignment vs. Distance Matrix Mouse: ACAGTGACGCCACACACGT Gorilla: CCTGCGACGTAACAAACGC
Molecular Evolution and Phylogenetic Tree Reconstruction
 1 4 Molecular Evolution and Phylogenetic Tree Reconstruction 3 2 5 1 4 2 3 5 Orthology, Paralogy, Inparalogs, Outparalogs Phylogenetic Trees Nodes: species Edges: time of independent evolution Edge length
1 4 Molecular Evolution and Phylogenetic Tree Reconstruction 3 2 5 1 4 2 3 5 Orthology, Paralogy, Inparalogs, Outparalogs Phylogenetic Trees Nodes: species Edges: time of independent evolution Edge length
Evolutionary Tree Analysis. Overview
 CSI/BINF 5330 Evolutionary Tree Analysis Young-Rae Cho Associate Professor Department of Computer Science Baylor University Overview Backgrounds Distance-Based Evolutionary Tree Reconstruction Character-Based
CSI/BINF 5330 Evolutionary Tree Analysis Young-Rae Cho Associate Professor Department of Computer Science Baylor University Overview Backgrounds Distance-Based Evolutionary Tree Reconstruction Character-Based
Phylogeny: traditional and Bayesian approaches
 Phylogeny: traditional and Bayesian approaches 5-Feb-2014 DEKM book Notes from Dr. B. John Holder and Lewis, Nature Reviews Genetics 4, 275-284, 2003 1 Phylogeny A graph depicting the ancestor-descendent
Phylogeny: traditional and Bayesian approaches 5-Feb-2014 DEKM book Notes from Dr. B. John Holder and Lewis, Nature Reviews Genetics 4, 275-284, 2003 1 Phylogeny A graph depicting the ancestor-descendent
Additive distances. w(e), where P ij is the path in T from i to j. Then the matrix [D ij ] is said to be additive.
![Additive distances. w(e), where P ij is the path in T from i to j. Then the matrix [D ij ] is said to be additive. Additive distances. w(e), where P ij is the path in T from i to j. Then the matrix [D ij ] is said to be additive.](/thumbs/92/110439366.jpg) Additive distances Let T be a tree on leaf set S and let w : E R + be an edge-weighting of T, and assume T has no nodes of degree two. Let D ij = e P ij w(e), where P ij is the path in T from i to j. Then
Additive distances Let T be a tree on leaf set S and let w : E R + be an edge-weighting of T, and assume T has no nodes of degree two. Let D ij = e P ij w(e), where P ij is the path in T from i to j. Then
CS5238 Combinatorial methods in bioinformatics 2003/2004 Semester 1. Lecture 8: Phylogenetic Tree Reconstruction: Distance Based - October 10, 2003
 CS5238 Combinatorial methods in bioinformatics 2003/2004 Semester 1 Lecture 8: Phylogenetic Tree Reconstruction: Distance Based - October 10, 2003 Lecturer: Wing-Kin Sung Scribe: Ning K., Shan T., Xiang
CS5238 Combinatorial methods in bioinformatics 2003/2004 Semester 1 Lecture 8: Phylogenetic Tree Reconstruction: Distance Based - October 10, 2003 Lecturer: Wing-Kin Sung Scribe: Ning K., Shan T., Xiang
Phylogeny: building the tree of life
 Phylogeny: building the tree of life Dr. Fayyaz ul Amir Afsar Minhas Department of Computer and Information Sciences Pakistan Institute of Engineering & Applied Sciences PO Nilore, Islamabad, Pakistan
Phylogeny: building the tree of life Dr. Fayyaz ul Amir Afsar Minhas Department of Computer and Information Sciences Pakistan Institute of Engineering & Applied Sciences PO Nilore, Islamabad, Pakistan
Page 1. Evolutionary Trees. Why build evolutionary tree? Outline
 Page Evolutionary Trees Russ. ltman MI S 7 Outline. Why build evolutionary trees?. istance-based vs. character-based methods. istance-based: Ultrametric Trees dditive Trees. haracter-based: Perfect phylogeny
Page Evolutionary Trees Russ. ltman MI S 7 Outline. Why build evolutionary trees?. istance-based vs. character-based methods. istance-based: Ultrametric Trees dditive Trees. haracter-based: Perfect phylogeny
EVOLUTIONARY DISTANCES
 EVOLUTIONARY DISTANCES FROM STRINGS TO TREES Luca Bortolussi 1 1 Dipartimento di Matematica ed Informatica Università degli studi di Trieste luca@dmi.units.it Trieste, 14 th November 2007 OUTLINE 1 STRINGS:
EVOLUTIONARY DISTANCES FROM STRINGS TO TREES Luca Bortolussi 1 1 Dipartimento di Matematica ed Informatica Università degli studi di Trieste luca@dmi.units.it Trieste, 14 th November 2007 OUTLINE 1 STRINGS:
 Tree of Life iological Sequence nalysis Chapter http://tolweb.org/tree/ Phylogenetic Prediction ll organisms on Earth have a common ancestor. ll species are related. The relationship is called a phylogeny
Tree of Life iological Sequence nalysis Chapter http://tolweb.org/tree/ Phylogenetic Prediction ll organisms on Earth have a common ancestor. ll species are related. The relationship is called a phylogeny
Phylogenetics: Distance Methods. COMP Spring 2015 Luay Nakhleh, Rice University
 Phylogenetics: Distance Methods COMP 571 - Spring 2015 Luay Nakhleh, Rice University Outline Evolutionary models and distance corrections Distance-based methods Evolutionary Models and Distance Correction
Phylogenetics: Distance Methods COMP 571 - Spring 2015 Luay Nakhleh, Rice University Outline Evolutionary models and distance corrections Distance-based methods Evolutionary Models and Distance Correction
Phylogenetic Tree Reconstruction
 I519 Introduction to Bioinformatics, 2011 Phylogenetic Tree Reconstruction Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Evolution theory Speciation Evolution of new organisms is driven
I519 Introduction to Bioinformatics, 2011 Phylogenetic Tree Reconstruction Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Evolution theory Speciation Evolution of new organisms is driven
Phylogenetic trees 07/10/13
 Phylogenetic trees 07/10/13 A tree is the only figure to occur in On the Origin of Species by Charles Darwin. It is a graphical representation of the evolutionary relationships among entities that share
Phylogenetic trees 07/10/13 A tree is the only figure to occur in On the Origin of Species by Charles Darwin. It is a graphical representation of the evolutionary relationships among entities that share
9/30/11. Evolution theory. Phylogenetic Tree Reconstruction. Phylogenetic trees (binary trees) Phylogeny (phylogenetic tree)
 I9 Introduction to Bioinformatics, 0 Phylogenetic ree Reconstruction Yuzhen Ye (yye@indiana.edu) School of Informatics & omputing, IUB Evolution theory Speciation Evolution of new organisms is driven by
I9 Introduction to Bioinformatics, 0 Phylogenetic ree Reconstruction Yuzhen Ye (yye@indiana.edu) School of Informatics & omputing, IUB Evolution theory Speciation Evolution of new organisms is driven by
Constructing Evolutionary/Phylogenetic Trees
 Constructing Evolutionary/Phylogenetic Trees 2 broad categories: istance-based methods Ultrametric Additive: UPGMA Transformed istance Neighbor-Joining Character-based Maximum Parsimony Maximum Likelihood
Constructing Evolutionary/Phylogenetic Trees 2 broad categories: istance-based methods Ultrametric Additive: UPGMA Transformed istance Neighbor-Joining Character-based Maximum Parsimony Maximum Likelihood
CSCI1950 Z Computa3onal Methods for Biology Lecture 24. Ben Raphael April 29, hgp://cs.brown.edu/courses/csci1950 z/ Network Mo3fs
 CSCI1950 Z Computa3onal Methods for Biology Lecture 24 Ben Raphael April 29, 2009 hgp://cs.brown.edu/courses/csci1950 z/ Network Mo3fs Subnetworks with more occurrences than expected by chance. How to
CSCI1950 Z Computa3onal Methods for Biology Lecture 24 Ben Raphael April 29, 2009 hgp://cs.brown.edu/courses/csci1950 z/ Network Mo3fs Subnetworks with more occurrences than expected by chance. How to
Phylogene)cs. IMBB 2016 BecA- ILRI Hub, Nairobi May 9 20, Joyce Nzioki
 Phylogene)cs IMBB 2016 BecA- ILRI Hub, Nairobi May 9 20, 2016 Joyce Nzioki Phylogenetics The study of evolutionary relatedness of organisms. Derived from two Greek words:» Phle/Phylon: Tribe/Race» Genetikos:
Phylogene)cs IMBB 2016 BecA- ILRI Hub, Nairobi May 9 20, 2016 Joyce Nzioki Phylogenetics The study of evolutionary relatedness of organisms. Derived from two Greek words:» Phle/Phylon: Tribe/Race» Genetikos:
A Phylogenetic Network Construction due to Constrained Recombination
 A Phylogenetic Network Construction due to Constrained Recombination Mohd. Abdul Hai Zahid Research Scholar Research Supervisors: Dr. R.C. Joshi Dr. Ankush Mittal Department of Electronics and Computer
A Phylogenetic Network Construction due to Constrained Recombination Mohd. Abdul Hai Zahid Research Scholar Research Supervisors: Dr. R.C. Joshi Dr. Ankush Mittal Department of Electronics and Computer
Algorithms in Bioinformatics
 Algorithms in Bioinformatics Sami Khuri Department of Computer Science San José State University San José, California, USA khuri@cs.sjsu.edu www.cs.sjsu.edu/faculty/khuri Distance Methods Character Methods
Algorithms in Bioinformatics Sami Khuri Department of Computer Science San José State University San José, California, USA khuri@cs.sjsu.edu www.cs.sjsu.edu/faculty/khuri Distance Methods Character Methods
Phylogeny Tree Algorithms
 Phylogeny Tree lgorithms Jianlin heng, PhD School of Electrical Engineering and omputer Science University of entral Florida 2006 Free for academic use. opyright @ Jianlin heng & original sources for some
Phylogeny Tree lgorithms Jianlin heng, PhD School of Electrical Engineering and omputer Science University of entral Florida 2006 Free for academic use. opyright @ Jianlin heng & original sources for some
Phylogenetic inference
 Phylogenetic inference Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, March 7 th 016 After this lecture, you can discuss (dis-) advantages of different information types
Phylogenetic inference Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, March 7 th 016 After this lecture, you can discuss (dis-) advantages of different information types
Inferring Phylogenetic Trees. Distance Approaches. Representing distances. in rooted and unrooted trees. The distance approach to phylogenies
 Inferring Phylogenetic Trees Distance Approaches Representing distances in rooted and unrooted trees The distance approach to phylogenies given: an n n matrix M where M ij is the distance between taxa
Inferring Phylogenetic Trees Distance Approaches Representing distances in rooted and unrooted trees The distance approach to phylogenies given: an n n matrix M where M ij is the distance between taxa
Mul$ple Sequence Alignment Methods. Tandy Warnow Departments of Bioengineering and Computer Science h?p://tandy.cs.illinois.edu
 Mul$ple Sequence Alignment Methods Tandy Warnow Departments of Bioengineering and Computer Science h?p://tandy.cs.illinois.edu Species Tree Orangutan Gorilla Chimpanzee Human From the Tree of the Life
Mul$ple Sequence Alignment Methods Tandy Warnow Departments of Bioengineering and Computer Science h?p://tandy.cs.illinois.edu Species Tree Orangutan Gorilla Chimpanzee Human From the Tree of the Life
A few logs suce to build (almost) all trees: Part II
 Theoretical Computer Science 221 (1999) 77 118 www.elsevier.com/locate/tcs A few logs suce to build (almost) all trees: Part II Peter L. Erdős a;, Michael A. Steel b,laszlo A.Szekely c, Tandy J. Warnow
Theoretical Computer Science 221 (1999) 77 118 www.elsevier.com/locate/tcs A few logs suce to build (almost) all trees: Part II Peter L. Erdős a;, Michael A. Steel b,laszlo A.Szekely c, Tandy J. Warnow
BINF6201/8201. Molecular phylogenetic methods
 BINF60/80 Molecular phylogenetic methods 0-7-06 Phylogenetics Ø According to the evolutionary theory, all life forms on this planet are related to one another by descent. Ø Traditionally, phylogenetics
BINF60/80 Molecular phylogenetic methods 0-7-06 Phylogenetics Ø According to the evolutionary theory, all life forms on this planet are related to one another by descent. Ø Traditionally, phylogenetics
X X (2) X Pr(X = x θ) (3)
 Notes for 848 lecture 6: A ML basis for compatibility and parsimony Notation θ Θ (1) Θ is the space of all possible trees (and model parameters) θ is a point in the parameter space = a particular tree
Notes for 848 lecture 6: A ML basis for compatibility and parsimony Notation θ Θ (1) Θ is the space of all possible trees (and model parameters) θ is a point in the parameter space = a particular tree
Math 239: Discrete Mathematics for the Life Sciences Spring Lecture 14 March 11. Scribe/ Editor: Maria Angelica Cueto/ C.E.
 Math 239: Discrete Mathematics for the Life Sciences Spring 2008 Lecture 14 March 11 Lecturer: Lior Pachter Scribe/ Editor: Maria Angelica Cueto/ C.E. Csar 14.1 Introduction The goal of today s lecture
Math 239: Discrete Mathematics for the Life Sciences Spring 2008 Lecture 14 March 11 Lecturer: Lior Pachter Scribe/ Editor: Maria Angelica Cueto/ C.E. Csar 14.1 Introduction The goal of today s lecture
Theory of Evolution Charles Darwin
 Theory of Evolution Charles arwin 858-59: Origin of Species 5 year voyage of H.M.S. eagle (83-36) Populations have variations. Natural Selection & Survival of the fittest: nature selects best adapted varieties
Theory of Evolution Charles arwin 858-59: Origin of Species 5 year voyage of H.M.S. eagle (83-36) Populations have variations. Natural Selection & Survival of the fittest: nature selects best adapted varieties
CSCI1950 Z Computa3onal Methods for Biology* (*Working Title) Lecture 1. Ben Raphael January 21, Course Par3culars
 CSCI1950 Z Computa3onal Methods for Biology* (*Working Title) Lecture 1 Ben Raphael January 21, 2009 Course Par3culars Three major topics 1. Phylogeny: ~50% lectures 2. Func3onal Genomics: ~25% lectures
CSCI1950 Z Computa3onal Methods for Biology* (*Working Title) Lecture 1 Ben Raphael January 21, 2009 Course Par3culars Three major topics 1. Phylogeny: ~50% lectures 2. Func3onal Genomics: ~25% lectures
ALGORITHMS FOR RECONSTRUCTING PHYLOGENETIC TREES FROM DISSIMILARITY MAPS
 ALGORITHMS FOR RECONSTRUCTING PHYLOGENETIC TREES FROM DISSIMILARITY MAPS DAN LEVY, FRANCIS EDWARD SU, AND RURIKO YOSHIDA Manuscript, December 15, 2003 Abstract. In this paper we improve on an algorithm
ALGORITHMS FOR RECONSTRUCTING PHYLOGENETIC TREES FROM DISSIMILARITY MAPS DAN LEVY, FRANCIS EDWARD SU, AND RURIKO YOSHIDA Manuscript, December 15, 2003 Abstract. In this paper we improve on an algorithm
Phylogenetic Analysis. Han Liang, Ph.D. Assistant Professor of Bioinformatics and Computational Biology UT MD Anderson Cancer Center
 Phylogenetic Analysis Han Liang, Ph.D. Assistant Professor of Bioinformatics and Computational Biology UT MD Anderson Cancer Center Outline Basic Concepts Tree Construction Methods Distance-based methods
Phylogenetic Analysis Han Liang, Ph.D. Assistant Professor of Bioinformatics and Computational Biology UT MD Anderson Cancer Center Outline Basic Concepts Tree Construction Methods Distance-based methods
Phylogenetic Trees. Phylogenetic Trees Five. Phylogeny: Inference Tool. Phylogeny Terminology. Picture of Last Quagga. Importance of Phylogeny 5.
 Five Sami Khuri Department of Computer Science San José State University San José, California, USA sami.khuri@sjsu.edu v Distance Methods v Character Methods v Molecular Clock v UPGMA v Maximum Parsimony
Five Sami Khuri Department of Computer Science San José State University San José, California, USA sami.khuri@sjsu.edu v Distance Methods v Character Methods v Molecular Clock v UPGMA v Maximum Parsimony
"Nothing in biology makes sense except in the light of evolution Theodosius Dobzhansky
 MOLECULAR PHYLOGENY "Nothing in biology makes sense except in the light of evolution Theodosius Dobzhansky EVOLUTION - theory that groups of organisms change over time so that descendeants differ structurally
MOLECULAR PHYLOGENY "Nothing in biology makes sense except in the light of evolution Theodosius Dobzhansky EVOLUTION - theory that groups of organisms change over time so that descendeants differ structurally
Hierarchical Clustering
 Hierarchical Clustering Some slides by Serafim Batzoglou 1 From expression profiles to distances From the Raw Data matrix we compute the similarity matrix S. S ij reflects the similarity of the expression
Hierarchical Clustering Some slides by Serafim Batzoglou 1 From expression profiles to distances From the Raw Data matrix we compute the similarity matrix S. S ij reflects the similarity of the expression
Plan: Evolutionary trees, characters. Perfect phylogeny Methods: NJ, parsimony, max likelihood, Quartet method
 Phylogeny 1 Plan: Phylogeny is an important subject. We have 2.5 hours. So I will teach all the concepts via one example of a chain letter evolution. The concepts we will discuss include: Evolutionary
Phylogeny 1 Plan: Phylogeny is an important subject. We have 2.5 hours. So I will teach all the concepts via one example of a chain letter evolution. The concepts we will discuss include: Evolutionary
Reconstruction of certain phylogenetic networks from their tree-average distances
 Reconstruction of certain phylogenetic networks from their tree-average distances Stephen J. Willson Department of Mathematics Iowa State University Ames, IA 50011 USA swillson@iastate.edu October 10,
Reconstruction of certain phylogenetic networks from their tree-average distances Stephen J. Willson Department of Mathematics Iowa State University Ames, IA 50011 USA swillson@iastate.edu October 10,
MOLECULAR EVOLUTION AND PHYLOGENETICS SERGEI L KOSAKOVSKY POND CSE/BIMM/BENG 181 MAY 27, 2011
 MOLECULAR EVOLUTION AND PHYLOGENETICS If we could observe evolution: speciation, mutation, natural selection and fixation, we might see something like this: AGTAGC GGTGAC AGTAGA CGTAGA AGTAGA A G G C AGTAGA
MOLECULAR EVOLUTION AND PHYLOGENETICS If we could observe evolution: speciation, mutation, natural selection and fixation, we might see something like this: AGTAGC GGTGAC AGTAGA CGTAGA AGTAGA A G G C AGTAGA
Minimum Edit Distance. Defini'on of Minimum Edit Distance
 Minimum Edit Distance Defini'on of Minimum Edit Distance How similar are two strings? Spell correc'on The user typed graffe Which is closest? graf gra@ grail giraffe Computa'onal Biology Align two sequences
Minimum Edit Distance Defini'on of Minimum Edit Distance How similar are two strings? Spell correc'on The user typed graffe Which is closest? graf gra@ grail giraffe Computa'onal Biology Align two sequences
Consistency Index (CI)
 Consistency Index (CI) minimum number of changes divided by the number required on the tree. CI=1 if there is no homoplasy negatively correlated with the number of species sampled Retention Index (RI)
Consistency Index (CI) minimum number of changes divided by the number required on the tree. CI=1 if there is no homoplasy negatively correlated with the number of species sampled Retention Index (RI)
Phylogenetic Trees. What They Are Why We Do It & How To Do It. Presented by Amy Harris Dr Brad Morantz
 Phylogenetic Trees What They Are Why We Do It & How To Do It Presented by Amy Harris Dr Brad Morantz Overview What is a phylogenetic tree Why do we do it How do we do it Methods and programs Parallels
Phylogenetic Trees What They Are Why We Do It & How To Do It Presented by Amy Harris Dr Brad Morantz Overview What is a phylogenetic tree Why do we do it How do we do it Methods and programs Parallels
Phylogenetics: Building Phylogenetic Trees
 1 Phylogenetics: Building Phylogenetic Trees COMP 571 Luay Nakhleh, Rice University 2 Four Questions Need to be Answered What data should we use? Which method should we use? Which evolutionary model should
1 Phylogenetics: Building Phylogenetic Trees COMP 571 Luay Nakhleh, Rice University 2 Four Questions Need to be Answered What data should we use? Which method should we use? Which evolutionary model should
A (short) introduction to phylogenetics
 A (short) introduction to phylogenetics Thibaut Jombart, Marie-Pauline Beugin MRC Centre for Outbreak Analysis and Modelling Imperial College London Genetic data analysis with PR Statistics, Millport Field
A (short) introduction to phylogenetics Thibaut Jombart, Marie-Pauline Beugin MRC Centre for Outbreak Analysis and Modelling Imperial College London Genetic data analysis with PR Statistics, Millport Field
Lecture 18 Molecular Evolution and Phylogenetics
 6.047/6.878 - Computational Biology: Genomes, Networks, Evolution Lecture 18 Molecular Evolution and Phylogenetics Patrick Winston s 6.034 Somewhere, something went wrong 1 Challenges in Computational
6.047/6.878 - Computational Biology: Genomes, Networks, Evolution Lecture 18 Molecular Evolution and Phylogenetics Patrick Winston s 6.034 Somewhere, something went wrong 1 Challenges in Computational
Bioinformatics 1. Sepp Hochreiter. Biology, Sequences, Phylogenetics Part 4. Bioinformatics 1: Biology, Sequences, Phylogenetics
 Bioinformatics 1 Biology, Sequences, Phylogenetics Part 4 Sepp Hochreiter Klausur Mo. 30.01.2011 Zeit: 15:30 17:00 Raum: HS14 Anmeldung Kusss Contents Methods and Bootstrapping of Maximum Methods Methods
Bioinformatics 1 Biology, Sequences, Phylogenetics Part 4 Sepp Hochreiter Klausur Mo. 30.01.2011 Zeit: 15:30 17:00 Raum: HS14 Anmeldung Kusss Contents Methods and Bootstrapping of Maximum Methods Methods
BMI/CS 776 Lecture 4. Colin Dewey
 BMI/CS 776 Lecture 4 Colin Dewey 2007.02.01 Outline Common nucleotide substitution models Directed graphical models Ancestral sequence inference Poisson process continuous Markov process X t0 X t1 X t2
BMI/CS 776 Lecture 4 Colin Dewey 2007.02.01 Outline Common nucleotide substitution models Directed graphical models Ancestral sequence inference Poisson process continuous Markov process X t0 X t1 X t2
Phylogenetics: Building Phylogenetic Trees. COMP Fall 2010 Luay Nakhleh, Rice University
 Phylogenetics: Building Phylogenetic Trees COMP 571 - Fall 2010 Luay Nakhleh, Rice University Four Questions Need to be Answered What data should we use? Which method should we use? Which evolutionary
Phylogenetics: Building Phylogenetic Trees COMP 571 - Fall 2010 Luay Nakhleh, Rice University Four Questions Need to be Answered What data should we use? Which method should we use? Which evolutionary
Let S be a set of n species. A phylogeny is a rooted tree with n leaves, each of which is uniquely
 JOURNAL OF COMPUTATIONAL BIOLOGY Volume 8, Number 1, 2001 Mary Ann Liebert, Inc. Pp. 69 78 Perfect Phylogenetic Networks with Recombination LUSHENG WANG, 1 KAIZHONG ZHANG, 2 and LOUXIN ZHANG 3 ABSTRACT
JOURNAL OF COMPUTATIONAL BIOLOGY Volume 8, Number 1, 2001 Mary Ann Liebert, Inc. Pp. 69 78 Perfect Phylogenetic Networks with Recombination LUSHENG WANG, 1 KAIZHONG ZHANG, 2 and LOUXIN ZHANG 3 ABSTRACT
POPULATION GENETICS Winter 2005 Lecture 17 Molecular phylogenetics
 POPULATION GENETICS Winter 2005 Lecture 17 Molecular phylogenetics - in deriving a phylogeny our goal is simply to reconstruct the historical relationships between a group of taxa. - before we review the
POPULATION GENETICS Winter 2005 Lecture 17 Molecular phylogenetics - in deriving a phylogeny our goal is simply to reconstruct the historical relationships between a group of taxa. - before we review the
Tree-average distances on certain phylogenetic networks have their weights uniquely determined
 Tree-average distances on certain phylogenetic networks have their weights uniquely determined Stephen J. Willson Department of Mathematics Iowa State University Ames, IA 50011 USA swillson@iastate.edu
Tree-average distances on certain phylogenetic networks have their weights uniquely determined Stephen J. Willson Department of Mathematics Iowa State University Ames, IA 50011 USA swillson@iastate.edu
Bioinformatics 1 -- lecture 9. Phylogenetic trees Distance-based tree building Parsimony
 ioinformatics -- lecture 9 Phylogenetic trees istance-based tree building Parsimony (,(,(,))) rees can be represented in "parenthesis notation". Each set of parentheses represents a branch-point (bifurcation),
ioinformatics -- lecture 9 Phylogenetic trees istance-based tree building Parsimony (,(,(,))) rees can be represented in "parenthesis notation". Each set of parentheses represents a branch-point (bifurcation),
Chapter 3: Phylogenetics
 Chapter 3: Phylogenetics 3. Computing Phylogeny Prof. Yechiam Yemini (YY) Computer Science epartment Columbia niversity Overview Computing trees istance-based techniques Maximal Parsimony (MP) techniques
Chapter 3: Phylogenetics 3. Computing Phylogeny Prof. Yechiam Yemini (YY) Computer Science epartment Columbia niversity Overview Computing trees istance-based techniques Maximal Parsimony (MP) techniques
Neighbor Joining Algorithms for Inferring Phylogenies via LCA-Distances
 Neighbor Joining Algorithms for Inferring Phylogenies via LCA-Distances Ilan Gronau Shlomo Moran September 6, 2006 Abstract Reconstructing phylogenetic trees efficiently and accurately from distance estimates
Neighbor Joining Algorithms for Inferring Phylogenies via LCA-Distances Ilan Gronau Shlomo Moran September 6, 2006 Abstract Reconstructing phylogenetic trees efficiently and accurately from distance estimates
Week 5: Distance methods, DNA and protein models
 Week 5: Distance methods, DNA and protein models Genome 570 February, 2016 Week 5: Distance methods, DNA and protein models p.1/69 A tree and the expected distances it predicts E A 0.08 0.05 0.06 0.03
Week 5: Distance methods, DNA and protein models Genome 570 February, 2016 Week 5: Distance methods, DNA and protein models p.1/69 A tree and the expected distances it predicts E A 0.08 0.05 0.06 0.03
Phylogeny and Evolution. Gina Cannarozzi ETH Zurich Institute of Computational Science
 Phylogeny and Evolution Gina Cannarozzi ETH Zurich Institute of Computational Science History Aristotle (384-322 BC) classified animals. He found that dolphins do not belong to the fish but to the mammals.
Phylogeny and Evolution Gina Cannarozzi ETH Zurich Institute of Computational Science History Aristotle (384-322 BC) classified animals. He found that dolphins do not belong to the fish but to the mammals.
Notes on the Matrix-Tree theorem and Cayley s tree enumerator
 Notes on the Matrix-Tree theorem and Cayley s tree enumerator 1 Cayley s tree enumerator Recall that the degree of a vertex in a tree (or in any graph) is the number of edges emanating from it We will
Notes on the Matrix-Tree theorem and Cayley s tree enumerator 1 Cayley s tree enumerator Recall that the degree of a vertex in a tree (or in any graph) is the number of edges emanating from it We will
Theory of Evolution. Charles Darwin
 Theory of Evolution harles arwin 858-59: Origin of Species 5 year voyage of H.M.S. eagle (8-6) Populations have variations. Natural Selection & Survival of the fittest: nature selects best adapted varieties
Theory of Evolution harles arwin 858-59: Origin of Species 5 year voyage of H.M.S. eagle (8-6) Populations have variations. Natural Selection & Survival of the fittest: nature selects best adapted varieties
Multiple Sequence Alignment. Sequences
 Multiple Sequence Alignment Sequences > YOR020c mstllksaksivplmdrvlvqrikaqaktasglylpe knveklnqaevvavgpgftdangnkvvpqvkvgdqvl ipqfggstiklgnddevilfrdaeilakiakd > crassa mattvrsvksliplldrvlvqrvkaeaktasgiflpe
Multiple Sequence Alignment Sequences > YOR020c mstllksaksivplmdrvlvqrikaqaktasglylpe knveklnqaevvavgpgftdangnkvvpqvkvgdqvl ipqfggstiklgnddevilfrdaeilakiakd > crassa mattvrsvksliplldrvlvqrvkaeaktasgiflpe
2 (n 1). f (i 1, i 2,..., i k )
 Math Time problem proposal #1 Darij Grinberg version 5 December 2010 Problem. Let x 1, x 2,..., x n be real numbers such that x 1 x 2...x n 1 and such that x i < 1 for every i {1, 2,..., n}. Prove that
Math Time problem proposal #1 Darij Grinberg version 5 December 2010 Problem. Let x 1, x 2,..., x n be real numbers such that x 1 x 2...x n 1 and such that x i < 1 for every i {1, 2,..., n}. Prove that
The Generalized Neighbor Joining method
 The Generalized Neighbor Joining method Ruriko Yoshida Dept. of Mathematics Duke University Joint work with Dan Levy and Lior Pachter www.math.duke.edu/ ruriko data mining 1 Challenge We would like to
The Generalized Neighbor Joining method Ruriko Yoshida Dept. of Mathematics Duke University Joint work with Dan Levy and Lior Pachter www.math.duke.edu/ ruriko data mining 1 Challenge We would like to
Rela%ons and Their Proper%es. Slides by A. Bloomfield
 Rela%ons and Their Proper%es Slides by A. Bloomfield What is a rela%on Let A and B be sets. A binary rela%on R is a subset of A B Example Let A be the students in a the CS major A = {Alice, Bob, Claire,
Rela%ons and Their Proper%es Slides by A. Bloomfield What is a rela%on Let A and B be sets. A binary rela%on R is a subset of A B Example Let A be the students in a the CS major A = {Alice, Bob, Claire,
Phylogeny Jan 5, 2016
 גנומיקה חישובית Computational Genomics Phylogeny Jan 5, 2016 Slides: Adi Akavia Nir Friedman s slides at HUJI (based on ALGMB 98) Anders Gorm Pedersen,Technical University of Denmark Sources: Joe Felsenstein
גנומיקה חישובית Computational Genomics Phylogeny Jan 5, 2016 Slides: Adi Akavia Nir Friedman s slides at HUJI (based on ALGMB 98) Anders Gorm Pedersen,Technical University of Denmark Sources: Joe Felsenstein
C3020 Molecular Evolution. Exercises #3: Phylogenetics
 C3020 Molecular Evolution Exercises #3: Phylogenetics Consider the following sequences for five taxa 1-5 and the known outgroup O, which has the ancestral states (note that sequence 3 has changed from
C3020 Molecular Evolution Exercises #3: Phylogenetics Consider the following sequences for five taxa 1-5 and the known outgroup O, which has the ancestral states (note that sequence 3 has changed from
Phylogenetics. BIOL 7711 Computational Bioscience
 Consortium for Comparative Genomics! University of Colorado School of Medicine Phylogenetics BIOL 7711 Computational Bioscience Biochemistry and Molecular Genetics Computational Bioscience Program Consortium
Consortium for Comparative Genomics! University of Colorado School of Medicine Phylogenetics BIOL 7711 Computational Bioscience Biochemistry and Molecular Genetics Computational Bioscience Program Consortium
arxiv: v1 [q-bio.pe] 1 Jun 2014
![arxiv: v1 [q-bio.pe] 1 Jun 2014 arxiv: v1 [q-bio.pe] 1 Jun 2014](/thumbs/90/101381538.jpg) THE MOST PARSIMONIOUS TREE FOR RANDOM DATA MAREIKE FISCHER, MICHELLE GALLA, LINA HERBST AND MIKE STEEL arxiv:46.27v [q-bio.pe] Jun 24 Abstract. Applying a method to reconstruct a phylogenetic tree from
THE MOST PARSIMONIOUS TREE FOR RANDOM DATA MAREIKE FISCHER, MICHELLE GALLA, LINA HERBST AND MIKE STEEL arxiv:46.27v [q-bio.pe] Jun 24 Abstract. Applying a method to reconstruct a phylogenetic tree from
66 Bioinformatics I, WS 09-10, D. Huson, December 1, Evolutionary tree of organisms, Ernst Haeckel, 1866
 66 Bioinformatics I, WS 09-10, D. Huson, December 1, 2009 5 Phylogeny Evolutionary tree of organisms, Ernst Haeckel, 1866 5.1 References J. Felsenstein, Inferring Phylogenies, Sinauer, 2004. C. Semple
66 Bioinformatics I, WS 09-10, D. Huson, December 1, 2009 5 Phylogeny Evolutionary tree of organisms, Ernst Haeckel, 1866 5.1 References J. Felsenstein, Inferring Phylogenies, Sinauer, 2004. C. Semple
Reconstructing Trees from Subtree Weights
 Reconstructing Trees from Subtree Weights Lior Pachter David E Speyer October 7, 2003 Abstract The tree-metric theorem provides a necessary and sufficient condition for a dissimilarity matrix to be a tree
Reconstructing Trees from Subtree Weights Lior Pachter David E Speyer October 7, 2003 Abstract The tree-metric theorem provides a necessary and sufficient condition for a dissimilarity matrix to be a tree
Thanks to Paul Lewis and Joe Felsenstein for the use of slides
 Thanks to Paul Lewis and Joe Felsenstein for the use of slides Review Hennigian logic reconstructs the tree if we know polarity of characters and there is no homoplasy UPGMA infers a tree from a distance
Thanks to Paul Lewis and Joe Felsenstein for the use of slides Review Hennigian logic reconstructs the tree if we know polarity of characters and there is no homoplasy UPGMA infers a tree from a distance
Evolutionary Models. Evolutionary Models
 Edit Operators In standard pairwise alignment, what are the allowed edit operators that transform one sequence into the other? Describe how each of these edit operations are represented on a sequence alignment
Edit Operators In standard pairwise alignment, what are the allowed edit operators that transform one sequence into the other? Describe how each of these edit operations are represented on a sequence alignment
Constructing Evolutionary/Phylogenetic Trees
 Constructing Evolutionary/Phylogenetic Trees 2 broad categories: Distance-based methods Ultrametric Additive: UPGMA Transformed Distance Neighbor-Joining Character-based Maximum Parsimony Maximum Likelihood
Constructing Evolutionary/Phylogenetic Trees 2 broad categories: Distance-based methods Ultrametric Additive: UPGMA Transformed Distance Neighbor-Joining Character-based Maximum Parsimony Maximum Likelihood
Did you know that Multiple Alignment is NP-hard? Isaac Elias Royal Institute of Technology Sweden
 Did you know that Multiple Alignment is NP-hard? Isaac Elias Royal Institute of Technology Sweden 1 Results Multiple Alignment with SP-score Star Alignment Tree Alignment (with given phylogeny) are NP-hard
Did you know that Multiple Alignment is NP-hard? Isaac Elias Royal Institute of Technology Sweden 1 Results Multiple Alignment with SP-score Star Alignment Tree Alignment (with given phylogeny) are NP-hard
Phylogenetics: Parsimony and Likelihood. COMP Spring 2016 Luay Nakhleh, Rice University
 Phylogenetics: Parsimony and Likelihood COMP 571 - Spring 2016 Luay Nakhleh, Rice University The Problem Input: Multiple alignment of a set S of sequences Output: Tree T leaf-labeled with S Assumptions
Phylogenetics: Parsimony and Likelihood COMP 571 - Spring 2016 Luay Nakhleh, Rice University The Problem Input: Multiple alignment of a set S of sequences Output: Tree T leaf-labeled with S Assumptions
Evolutionary trees. Describe the relationship between objects, e.g. species or genes
 Evolutionary trees Bonobo Chimpanzee Human Neanderthal Gorilla Orangutan Describe the relationship between objects, e.g. species or genes Early evolutionary studies The evolutionary relationships between
Evolutionary trees Bonobo Chimpanzee Human Neanderthal Gorilla Orangutan Describe the relationship between objects, e.g. species or genes Early evolutionary studies The evolutionary relationships between
Phylogenetic Networks, Trees, and Clusters
 Phylogenetic Networks, Trees, and Clusters Luay Nakhleh 1 and Li-San Wang 2 1 Department of Computer Science Rice University Houston, TX 77005, USA nakhleh@cs.rice.edu 2 Department of Biology University
Phylogenetic Networks, Trees, and Clusters Luay Nakhleh 1 and Li-San Wang 2 1 Department of Computer Science Rice University Houston, TX 77005, USA nakhleh@cs.rice.edu 2 Department of Biology University
Lecture 4: Evolutionary Models and Substitution Matrices (PAM and BLOSUM)
 Bioinformatics II Probability and Statistics Universität Zürich and ETH Zürich Spring Semester 2009 Lecture 4: Evolutionary Models and Substitution Matrices (PAM and BLOSUM) Dr Fraser Daly adapted from
Bioinformatics II Probability and Statistics Universität Zürich and ETH Zürich Spring Semester 2009 Lecture 4: Evolutionary Models and Substitution Matrices (PAM and BLOSUM) Dr Fraser Daly adapted from
3. Coding theory 3.1. Basic concepts
 3. CODING THEORY 1 3. Coding theory 3.1. Basic concepts In this chapter we will discuss briefly some aspects of error correcting codes. The main problem is that if information is sent via a noisy channel,
3. CODING THEORY 1 3. Coding theory 3.1. Basic concepts In this chapter we will discuss briefly some aspects of error correcting codes. The main problem is that if information is sent via a noisy channel,
j=1 [We will show that the triangle inequality holds for each p-norm in Chapter 3 Section 6.] The 1-norm is A F = tr(a H A).
![j=1 [We will show that the triangle inequality holds for each p-norm in Chapter 3 Section 6.] The 1-norm is A F = tr(a H A). j=1 [We will show that the triangle inequality holds for each p-norm in Chapter 3 Section 6.] The 1-norm is A F = tr(a H A).](/thumbs/88/117636161.jpg) Math 344 Lecture #19 3.5 Normed Linear Spaces Definition 3.5.1. A seminorm on a vector space V over F is a map : V R that for all x, y V and for all α F satisfies (i) x 0 (positivity), (ii) αx = α x (scale
Math 344 Lecture #19 3.5 Normed Linear Spaces Definition 3.5.1. A seminorm on a vector space V over F is a map : V R that for all x, y V and for all α F satisfies (i) x 0 (positivity), (ii) αx = α x (scale
Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut
 Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic analysis Phylogenetic Basics: Biological
Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic analysis Phylogenetic Basics: Biological
RECOVERING NORMAL NETWORKS FROM SHORTEST INTER-TAXA DISTANCE INFORMATION
 RECOVERING NORMAL NETWORKS FROM SHORTEST INTER-TAXA DISTANCE INFORMATION MAGNUS BORDEWICH, KATHARINA T. HUBER, VINCENT MOULTON, AND CHARLES SEMPLE Abstract. Phylogenetic networks are a type of leaf-labelled,
RECOVERING NORMAL NETWORKS FROM SHORTEST INTER-TAXA DISTANCE INFORMATION MAGNUS BORDEWICH, KATHARINA T. HUBER, VINCENT MOULTON, AND CHARLES SEMPLE Abstract. Phylogenetic networks are a type of leaf-labelled,
Estimating Phylogenies (Evolutionary Trees) II. Biol4230 Thurs, March 2, 2017 Bill Pearson Jordan 6-057
 Estimating Phylogenies (Evolutionary Trees) II Biol4230 Thurs, March 2, 2017 Bill Pearson wrp@virginia.edu 4-2818 Jordan 6-057 Tree estimation strategies: Parsimony?no model, simply count minimum number
Estimating Phylogenies (Evolutionary Trees) II Biol4230 Thurs, March 2, 2017 Bill Pearson wrp@virginia.edu 4-2818 Jordan 6-057 Tree estimation strategies: Parsimony?no model, simply count minimum number
Stochastic Errors vs. Modeling Errors in Distance Based Phylogenetic Reconstructions
 PLGW05 Stochastic Errors vs. Modeling Errors in Distance Based Phylogenetic Reconstructions 1 joint work with Ilan Gronau 2, Shlomo Moran 3, and Irad Yavneh 3 1 2 Dept. of Biological Statistics and Computational
PLGW05 Stochastic Errors vs. Modeling Errors in Distance Based Phylogenetic Reconstructions 1 joint work with Ilan Gronau 2, Shlomo Moran 3, and Irad Yavneh 3 1 2 Dept. of Biological Statistics and Computational
3D Data, Phylogenetics, and Trees In this lecture.
 3D Data, Phylogenetics, and Trees In this lecture. 1. Analysis of 3D landmarks 2. Controversies about use of morphometrics in phylogene>cs 3. Empirical data on phylogene>c component of morphometric varia>on
3D Data, Phylogenetics, and Trees In this lecture. 1. Analysis of 3D landmarks 2. Controversies about use of morphometrics in phylogene>cs 3. Empirical data on phylogene>c component of morphometric varia>on
Dr. Amira A. AL-Hosary
 Phylogenetic analysis Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic Basics: Biological
Phylogenetic analysis Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic Basics: Biological
Cartesian Tensors. e 2. e 1. General vector (formal definition to follow) denoted by components
 Cartesian Tensors Reference: Jeffreys Cartesian Tensors 1 Coordinates and Vectors z x 3 e 3 y x 2 e 2 e 1 x x 1 Coordinates x i, i 123,, Unit vectors: e i, i 123,, General vector (formal definition to
Cartesian Tensors Reference: Jeffreys Cartesian Tensors 1 Coordinates and Vectors z x 3 e 3 y x 2 e 2 e 1 x x 1 Coordinates x i, i 123,, Unit vectors: e i, i 123,, General vector (formal definition to
(Stevens 1991) 1. morphological characters should be assumed to be quantitative unless demonstrated otherwise
 Bot 421/521 PHYLOGENETIC ANALYSIS I. Origins A. Hennig 1950 (German edition) Phylogenetic Systematics 1966 B. Zimmerman (Germany, 1930 s) C. Wagner (Michigan, 1920-2000) II. Characters and character states
Bot 421/521 PHYLOGENETIC ANALYSIS I. Origins A. Hennig 1950 (German edition) Phylogenetic Systematics 1966 B. Zimmerman (Germany, 1930 s) C. Wagner (Michigan, 1920-2000) II. Characters and character states
The Strong Largeur d Arborescence
 The Strong Largeur d Arborescence Rik Steenkamp (5887321) November 12, 2013 Master Thesis Supervisor: prof.dr. Monique Laurent Local Supervisor: prof.dr. Alexander Schrijver KdV Institute for Mathematics
The Strong Largeur d Arborescence Rik Steenkamp (5887321) November 12, 2013 Master Thesis Supervisor: prof.dr. Monique Laurent Local Supervisor: prof.dr. Alexander Schrijver KdV Institute for Mathematics
Integer Programming for Phylogenetic Network Problems
 Integer Programming for Phylogenetic Network Problems D. Gusfield University of California, Davis Presented at the National University of Singapore, July 27, 2015.! There are many important phylogeny problems
Integer Programming for Phylogenetic Network Problems D. Gusfield University of California, Davis Presented at the National University of Singapore, July 27, 2015.! There are many important phylogeny problems
Lecture 10: Phylogeny
 Computational Genomics Prof. Ron Shamir & Prof. Roded Sharan School of Computer Science, Tel Aviv University גנומיקה חישובית פרופ' רון שמיר ופרופ' רודד שרן ביה"ס למדעי המחשב,אוניברסיטת תל אביב Lecture
Computational Genomics Prof. Ron Shamir & Prof. Roded Sharan School of Computer Science, Tel Aviv University גנומיקה חישובית פרופ' רון שמיר ופרופ' רודד שרן ביה"ס למדעי המחשב,אוניברסיטת תל אביב Lecture
Constructing Evolutionary Trees
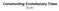 Constructing Evolutionary Trees 0-0 HIV Evolutionary Tree SIVs (monkeys)! HIV (human)! human infection! human HIV/M human HIV/M chimpanzee SIV chimpanzee SIV human HIV/N human HIV/N chimpanzee SIV chimpanzee
Constructing Evolutionary Trees 0-0 HIV Evolutionary Tree SIVs (monkeys)! HIV (human)! human infection! human HIV/M human HIV/M chimpanzee SIV chimpanzee SIV human HIV/N human HIV/N chimpanzee SIV chimpanzee
DNA Phylogeny. Signals and Systems in Biology Kushal EE, IIT Delhi
 DNA Phylogeny Signals and Systems in Biology Kushal Shah @ EE, IIT Delhi Phylogenetics Grouping and Division of organisms Keeps changing with time Splitting, hybridization and termination Cladistics :
DNA Phylogeny Signals and Systems in Biology Kushal Shah @ EE, IIT Delhi Phylogenetics Grouping and Division of organisms Keeps changing with time Splitting, hybridization and termination Cladistics :
394C, October 2, Topics: Mul9ple Sequence Alignment Es9ma9ng Species Trees from Gene Trees
 394C, October 2, 2013 Topics: Mul9ple Sequence Alignment Es9ma9ng Species Trees from Gene Trees Mul9ple Sequence Alignment Mul9ple Sequence Alignments and Evolu9onary Histories (the meaning of homologous
394C, October 2, 2013 Topics: Mul9ple Sequence Alignment Es9ma9ng Species Trees from Gene Trees Mul9ple Sequence Alignment Mul9ple Sequence Alignments and Evolu9onary Histories (the meaning of homologous
Lecture 2: Pairwise Alignment. CG Ron Shamir
 Lecture 2: Pairwise Alignment 1 Main source 2 Why compare sequences? Human hexosaminidase A vs Mouse hexosaminidase A 3 www.mathworks.com/.../jan04/bio_genome.html Sequence Alignment עימוד רצפים The problem:
Lecture 2: Pairwise Alignment 1 Main source 2 Why compare sequences? Human hexosaminidase A vs Mouse hexosaminidase A 3 www.mathworks.com/.../jan04/bio_genome.html Sequence Alignment עימוד רצפים The problem:
Arrangements, matroids and codes
 Arrangements, matroids and codes first lecture Ruud Pellikaan joint work with Relinde Jurrius ACAGM summer school Leuven Belgium, 18 July 2011 References 2/43 1. Codes, arrangements and matroids by Relinde
Arrangements, matroids and codes first lecture Ruud Pellikaan joint work with Relinde Jurrius ACAGM summer school Leuven Belgium, 18 July 2011 References 2/43 1. Codes, arrangements and matroids by Relinde
Distances and similarities Based in part on slides from textbook, slides of Susan Holmes. October 3, Statistics 202: Data Mining
 Distances and similarities Based in part on slides from textbook, slides of Susan Holmes October 3, 2012 1 / 1 Similarities Start with X which we assume is centered and standardized. The PCA loadings were
Distances and similarities Based in part on slides from textbook, slides of Susan Holmes October 3, 2012 1 / 1 Similarities Start with X which we assume is centered and standardized. The PCA loadings were
Multiple Sequence Alignment, Gunnar Klau, December 9, 2005, 17:
 Multiple Sequence Alignment, Gunnar Klau, December 9, 2005, 17:50 5001 5 Multiple Sequence Alignment The first part of this exposition is based on the following sources, which are recommended reading:
Multiple Sequence Alignment, Gunnar Klau, December 9, 2005, 17:50 5001 5 Multiple Sequence Alignment The first part of this exposition is based on the following sources, which are recommended reading:
Example 3.3 Moving-average filter
 3.3 Difference Equa/ons for Discrete-Time Systems A Discrete-/me system can be modeled with difference equa3ons involving current, past, or future samples of input and output signals Example 3.3 Moving-average
3.3 Difference Equa/ons for Discrete-Time Systems A Discrete-/me system can be modeled with difference equa3ons involving current, past, or future samples of input and output signals Example 3.3 Moving-average
A PARSIMONY APPROACH TO ANALYSIS OF HUMAN SEGMENTAL DUPLICATIONS
 A PARSIMONY APPROACH TO ANALYSIS OF HUMAN SEGMENTAL DUPLICATIONS CRYSTAL L. KAHN and BENJAMIN J. RAPHAEL Box 1910, Brown University Department of Computer Science & Center for Computational Molecular Biology
A PARSIMONY APPROACH TO ANALYSIS OF HUMAN SEGMENTAL DUPLICATIONS CRYSTAL L. KAHN and BENJAMIN J. RAPHAEL Box 1910, Brown University Department of Computer Science & Center for Computational Molecular Biology
Phylogeny. November 7, 2017
 Phylogeny November 7, 2017 Phylogenetics Phylon = tribe/race, genetikos = relative to birth Phylogenetics: study of evolutionary relationships among organisms, sequences, or anything in between Related
Phylogeny November 7, 2017 Phylogenetics Phylon = tribe/race, genetikos = relative to birth Phylogenetics: study of evolutionary relationships among organisms, sequences, or anything in between Related
Inferring phylogeny. Today s topics. Milestones of molecular evolution studies Contributions to molecular evolution
 Today s topics Inferring phylogeny Introduction! Distance methods! Parsimony method!"#$%&'(!)* +,-.'/01!23454(6!7!2845*0&4'9#6!:&454(6 ;?@AB=C?DEF Overview of phylogenetic inferences Methodology Methods
Today s topics Inferring phylogeny Introduction! Distance methods! Parsimony method!"#$%&'(!)* +,-.'/01!23454(6!7!2845*0&4'9#6!:&454(6 ;?@AB=C?DEF Overview of phylogenetic inferences Methodology Methods
Algebraic Statistics Tutorial I
 Algebraic Statistics Tutorial I Seth Sullivant North Carolina State University June 9, 2012 Seth Sullivant (NCSU) Algebraic Statistics June 9, 2012 1 / 34 Introduction to Algebraic Geometry Let R[p] =
Algebraic Statistics Tutorial I Seth Sullivant North Carolina State University June 9, 2012 Seth Sullivant (NCSU) Algebraic Statistics June 9, 2012 1 / 34 Introduction to Algebraic Geometry Let R[p] =
Week 15-16: Combinatorial Design
 Week 15-16: Combinatorial Design May 8, 2017 A combinatorial design, or simply a design, is an arrangement of the objects of a set into subsets satisfying certain prescribed properties. The area of combinatorial
Week 15-16: Combinatorial Design May 8, 2017 A combinatorial design, or simply a design, is an arrangement of the objects of a set into subsets satisfying certain prescribed properties. The area of combinatorial
