Bayesian Determination of Threshold for Identifying Differentially Expressed Genes in Microarray Experiments
|
|
|
- Jared McKenzie
- 5 years ago
- Views:
Transcription
1 Bayesian Determination of Threshold for Identifying Differentially Expressed Genes in Microarray Experiments Jie Chen 1 Merck Research Laboratories, P. O. Box 4, BL3-2, West Point, PA 19486, U.S.A. Telephone: (484) Sanat K. Sarkar 2 Department of Statistics, Temple University, Philadelphia, PA 19422, U.S.A. Telephone: (215) Short title: Bayesian Threshold for Differential Gene Expression 1. jie chen@merck.com sanat@surfer.sbm.temple.edu. The research is supported by NSF Grant DMS
2 Abstract The original definition of false discovery rate (FDR) can be understood as the frequentist risk of false rejections conditional on the unknown parameter, while the Bayesian posterior FDR is conditioned on the data, a particular realization of an experiment. From a Bayesian point of view, it seems natural to take into account the uncertainty in both the parameter and the data. In this spirit, we propose the Average FDR (AFDR) and Average FNR (AFNR) approaches in which the frequentist risks of false rejections and false non-rejections are averaged out with respect to some prior distribution of parameter. A linear combination of the AFDR and AFNR, called the Average Bayes Error Rate (ABER), is considered as an overall risk. Some useful formulas for the AFDR, AFNR and ABER are developed for normal samples with hierarchical mixture priors. The idea of finding threshold values minimizing the ABER is illustrated using a gene expression data set. Simulation studies show that the proposed approaches are more powerful and relatively robust than the widely used FDR method. Keywords: Average false discovery rate, Average false non-discovery rate, Average Bayes error rate, Hierarchical mixture model, Microarray experiment. 1 Introduction The emergence of DNA microarray technology allows for the study of sequence, structure and expression of thousands of genes simultaneously. Microarrays are being used increasingly in a wide variety of areas ranging from toxicological research (toxicogenomics), gene discovery, disease diagnosis, to drug discovery (pharmacogenomics) (Nuwaysir, Bittner, Trent, Barrett, and Afshari 1999; Afshari, Nuwaysir, and Barrett 1999; Callow, Dudoit, Gong, Speed, and Rubin 2
3 2000). A typical DNA microarray generates massive amount of data concerning gene regulations and interactions. A natural question that arises in such a study is whether differential expression of a gene is associated with certain condition, such as tumor type of breast cancer. This question is commonly posed as a multiple testing problem with the null hypothesis for each gene representing no association of expression level with the condition. With a large number of hypothesis tests performed simultaneously, the probability of misidentifying a gene as differentially expressed when it is not can increase sharply. The traditional concept of familywise error rate (FWER) is too restrictive to adopt in such a multiple testing situation. Instead, the false discovery rate (FDR) of Benjamini and Hochberg (1995) (hence BH-FDR hereafter) and related measures seem more appropriate as they offer less stringent criteria and thus provide more powerful methods in dealing with this magnitude of multiplicity. For an overview of multiple hypothesis testing in gene expression analysis, the reader is referred to Dudoit, Shaffer, and Boldrick (2003), Reiner, Yekutieli, and Benjamini (2003), and Ge, Dudoit, and Speed (2003). Suppose we have k genes, with the ith gene having the test statistic or differential expression measurement D i which has a probability distribution depending on an unknown parameter θ i, i = 1,..., k. Let D = {D 1,..., D k } and θ = {θ 1,..., θ k }. The underlying hypotheses of interest are H i : θ i = θ 0 against H i : θ i θ 0, i = 1,..., k, for some known θ 0. Decisions on these k hypotheses, that is, rejection or acceptance of the null hypotheses, are usually based on the magnitudes of the corresponding test statistics D i, i = 1,..., k. Table 1 shows all possible outcomes for the k hypothesis tests. The FDR is defined as the expected proportion of false rejections among the set of rejected hypotheses, i.e., F DR = E{V/(R 1)}. Storey (2002, 2003) introduces a modified version of FDR, called the positive false discovery rate 3
4 (pfdr), defined as pf DR = E [V/R R > 0], and argues that the pfdr is a more appropriate and useful error measure, as the probability of at least one rejection can be less than 1. Storey (2003) showed that the pfdr can be written as a Bayesian posterior probability, which is asymptotically true under fairly general conditions. An analog of FDR in terms of false non-rejections, called the false non-discovery rate (FNR), is introduced by Genovese and Wasserman (2002b). It is the expected proportion of false non-rejections among the set of non-rejected hypotheses, i.e., F NR = E{T/(A 1)}. Sarkar (2003) developed some results on the FDR and FNR in single-step procedures for dependent test statistics, under both a model where the number of true null hypotheses is assumed fixed and a mixture model where different configurations of true and false null hypotheses are assumed to have certain probabilities. He extended some previously known results developed under the assumption of independence and explained precisely how an FDR- or FNR-controlling single-step procedure, such as Bonferroni or Šidák procedure, can potentially be improved using an estimate of k 0. Multiple testing can be viewed as a decision making process through which one rejects or retains the null hypotheses by controlling some error or risk. While a frequentist measures this error or risk by considering a loss function and averaging it over different possible realizations of the data D conditional on the parameter θ; a Bayesian, on the other hand, defines this risk by averaging the loss over different possible states of nature conditional on D. For instance, the FDR and FNR can be understood as frequentist risks of false rejections and false nonrejections, respectively. Genovese and Wasserman (2002a) gave the first fully Bayesian exposition of false discovery rate and introduced the Bayesian posterior FDR (PFDR), defined as P F DR = E θ D [V/(R 1)], assuming certain prior distribution of θ. They also introduced the posterior FNR (PFNR), defined as P F NR = E θ D [T/(A 1)]. 4
5 In this article, we address the multiple testing problem in a microarray experiment by determining a threshold or critical value for each gene from a Bayesian perspective. It seems natural that, when a Bayesian approach is taken, one should take into account the uncertainty in both parameter and data in determining risks. In this spirit, we propose the Average FDR (AFDR) approach in which the frequentist risk of false rejections is averaged out with respect to the prior distribution of θ. This is also seen as the expected Bayesian posterior risk of false rejections with respect to the marginal distribution of data D. The AFDR approach provides an alternate view on controlling error rate involving false positives and hence is useful in multiple testing problems, including those arising in gene expression data analysis. An analog of the AFDR in terms of false non-rejections, called Average FNR (AFNR), is also developed to control error rate involving false negatives from a Bayesian viewpoint. An overall Bayes risk is defined as a linear combination of the AFDR and AFNR. We call this an Average Bayes Error Rate (ABER), and propose to determine the threshold value of test statistics by minimizing the ABER. Our simulation studies show that the proposed approaches are more powerful (in terms of average power) and relatively robust to influential data points as well as to the choices of priors than the BH-FDR method. The paper is organized as follows. In Section 2, we describe a hierarchical mixture model that will be used in identifying differentially expressed genes between two sample types. The concept of AFDR is formally defined in Section 3 and some formulas for this measure in a single-step procedure are developed. Section 4 is devoted to the similar development of the AFNR. The ABER is described in Section 5. In Section 6, we apply our AFDR, AFNR and ABER approaches to a breast cancer data set and illustrate how the threshold is determined for detection of differentially expressed genes in terms of minimum ABER. Section 7 5
6 is devoted to simulation studies investigating the average power of AFDR, ABER and BH-FDR as well as the robustness of these methods to influential data points as well as to the choice of priors. Some other possible alternatives defining false positives and false negatives are discussed in Section 8. 2 Hierarchical Mixture Model for Gene Expressions We will present in this section a hierarchical mixture model for identifying differentially expressed genes. The mixture model approaches have been taken by Efron, Tibshirani, Storey, and Tusher (2001), Storey (2003) and Sarkar (2003); we will extend these approaches by using hyper prior distributions on which our proposed procedure is based. We will assume that the microarray data have been pre-processed or normalized to adjust for any bias and systematic variation other than the factor under consideration, and are ready for statistical analysis of significance. For discussions on pre-treatment of microarray data, the reader is referred to Finkelstein, Gollub, and Cherry (2001), Yang, Dudoit, Luu, and Speed (2001), and Chen, Kodell, Sistare, Thompson, Morris, and Chen (2003) We restrict attention to two-sample comparisons, i.e., to compare gene expression levels from two different types of samples (treatment vs control, disease vs non-disease, two different tumor types, etc.). Let X ijl be the lth (normalized) expression measurement for the ith gene from the jth type of sample and X ijl have N(η ij, σ 2 ) distribution, l = 1,..., n ij, j = 1, 2 and i = 1,..., k. One often assigns a prior distribution to σ 2. However, for simplicity we will obtain the unbiased estimator ˆσ 2 of σ 2, from which the variance of difference in average expression levels between two sample types can be easily derived. The unbiased 6
7 estimator ˆσ 2 of σ 2 is simply where X ij ˆσ 2 = 1 n 2k k i=1 n 2 ij ( Xijl X ) 2 ij, (2.1) j=1 l=1 = n ij l=1 X ijl/n ij and n = k i=1 2 j=1 n ij. Let D i = X i1 X i2 and θ i = η i1 η i2 be, respectively, the differences of sample and population means of expression levels for gene i between two sample types, i = 1,..., k. Suppose n i1 = n 1 and n i2 = n 2 for all i = 1,..., k, i.e., the sample sizes within sample type are the same for all of the genes. This gives the estimated variance of D i as ˆσ 2 D = ˆσ2 (1/n 1 + 1/n 2 ). The problem of identifying differentially expressed genes that are associated with certain condition is typically approached by simultaneously testing the null hypotheses H i : θ i = 0 against the complementary alternatives H i : θ 0, i = 1,..., k. Towards this goal, we assume the following hierarchical mixture models D i θ i N ( θ i, ˆσ 2 D), i = 1,..., k; θ i µ, τ 2 π 0 I (θ i = 0) + (1 π 0 ) N ( µ, τ 2) I (θ i 0), i = 1,..., k; µ ξ, τ 2 N ( 0, ξτ 2), where π 0 is the prior probability of the null hypothesis being true and (ξ, τ 2 ) may follow some distribution g 1 (ξ, τ 2 ) or sometimes are assigned arbitrary values. That is, the D i s are conditionally independent given θ i s which are also conditionally independent given µ, τ 2 and ξ. Under this distributional setup, it can be shown that the marginal distribution of the data D = {D 1,..., D k }, conditional on τ 2 and ξ, is given by D N k (0, ψ(τ, ξ)), (2.2) 7
8 where ψ(τ, ξ) = v(ξ, τ 2 ) c(ξ, τ 2 )... c(ξ, τ 2 ) c(ξ, τ 2 ) v(ξ, τ 2 )... c(ξ, τ 2 ) c(ξ, τ 2 ) c(ξ, τ 2 )... v(ξ, τ 2 ), v(ξ, τ 2 ) = ˆσ D 2 +(1 π 0) 2 (1 + ξ) τ 2 and c(ξ, τ 2 ) = (1 π 0 ) 2 ξτ 2. Berger (1985) and Schervish (1995) provide a detailed proof for normal hierarchical models with a similar structure. The property of positive correlation among D in (2.2) is useful in establishing some characteristics of the AFDR and AFNR, as will be seen in Sections 3 and 4. Some recent articles adopt the Bayesian hierarchical mixture model approaches to identifying differentially expressed genes from microarray experiments (Baldi and Long 2001; Broët, Richardson, and Radvanyi 2002; Ibrahim, Chen, and Gray 2002; Ishwaran and Rao 2003). These procedures, however, are based solely on the posterior distribution of the parameter and such derived FDR s are the Bayesian posterior FDR s conditional on the data (Ishwaran and Rao 2003; Newton, Noueiry, Sarkar, and Ahlquist 2003). To account for the uncertainty in both parameter and data, the concepts of the AFDR and AFNR are introduced and some useful formulas under the above hierarchical mixture models are developed in the next two sections. 3 Average False Discovery Rate We define the AFDR in the similar way as in Benjamini and Hochberg (1995), except that an additional expectation is taken with respect to some prior distribution of parameter. Definition 1. The Average False Discovery Rate (AFDR) among the set of 8
9 rejected hypotheses is defined to be AF DR = E θ [ E D θ ( V R 1 )]. (3.1) This quantity is the Bayes risk of false rejections among the set of rejected hypotheses. Noting that reversing the order of integration in (3.1), we obtain the alternate form AF DR = E D [ E θ D ( V R 1 )], (3.2) which is the expected posterior risk with respect to some marginal distribution of the data D. Therefore, AF DR = E D (P F DR). Suppose that a large absolute value of D i compared to a threshold value c leads to the rejection of H i, identifying the corresponding gene to be either under- or over-expressed. Let D ( i) 1:k 1,..., D ( i) k 1:k 1 be the ordered components of { D j : j J ( i) } with J ( i) = J {i} and J = {1,..., k}. Define D 0:k = and D k+1:k =. Then, the AFDR of this procedure can be written as AF DR = 1 k k P { D i c, θ i = 0} i=1 [ k 1 P 1 + k j=1 { D ( i) j:k 1 < c D i c, θ i = 0 (k j)(k j + 1) } ]. (3.3) If D is a multivariate normal with non-negative correlations, as it is the case under the above hierarchical mixture models, then the AFDR is no more than the right hand side of (3.3); see Sarkar (2003). If (D i, θ i ), i = 1,..., k, are iid, (3.3) reduces to AF DR = P {R > 0}P {θ 1 = 0 D 1 c} ; (3.4) 9
10 see, also Storey (2003). Under the hierarchical mixture model, notice that (D i, θ i ), i = 1,..., k are iid conditional on µ, ξ and τ 2. Therefore, conditional on µ, ξ and τ 2, the AFDR can be written as AF DR = [ 1 ν k] 2π 0 [1 Φ (c 0 )], 1 ν k 1 = 2π 0 [1 Φ (c 0 )] ν j, (3.5) where Φ is the c.d.f. of standard normal, j=0 c 0 = ν = π 0 [2Φ (c 0 ) 1] + (1 π 0 ) [Φ(c 2 ) Φ(c 1 )], c, c 1 = c µ ˆσ 2 D ˆσ D + τ, and c 2 2 = ˆσ c µ 2 D + τ. 2 The AFDR is the integral of (3.5) with respect to µ, ξ and τ 2. It is a nonincreasing function of c. This can be seen from the following two results, conditionally given µ, ξ and τ 2. First, P {R > 0} = 1 [P { D 1 < c}] k is nonincreasing in c. Second, P {θ 1 = 0 D 1 c} = = [ 1 + (1 π 0) π 0 [ 1 + (1 π 0) π 0 P { D 1 c θ 1 } P { D 1 c θ 1 = 0} φ(θ 1; µ, τ 2 )dθ 1 ] P {χ 2 1(λ) > c 2 1 0} P {χ 2 1 > c 2 0} φ(θ 1; µ, τ 2 )dθ 1, ] 1 (3.6) where φ(x; µ, τ 2 ) is the density of N(µ, τ 2 ), χ 2 1 is the central chi-squared random variable with 1 degree of freedom, and χ 2 1(λ) is the non-central chi-squared random variable with 1 degree of freedom and the non-centrality parameter λ = θ1/ˆσ 2 D 2, which is also nonincreasing in c because the ratio P {χ 2 1(λ) > c 2 0} P {χ 2 1 > c 2 0} is known to be nondecreasing in c 2 0 (Gupta and Sarkar 1984) and hence in c. 10
11 4 Average False Non-discovery Rate The AFDR defined above is only one part of the Bayes risk of misclassifications. Another quantity measuring the error rate of false non-rejections is the average false non-discovery rate which is defined as follows. Definition 2. The Average False Non-discovery Rate (AFNR) among the set of non-rejected hypotheses is defined to be AF NR = E θ [ E D θ ( T A 1 )]. (4.1) In words, the AFNR is the average risk of non-rejections when the hypotheses are false. This quantity is seen as the expected posterior risk of false nonrejections with respect to the marginal distribution of the data D. The AFNR of a single-step procedure that rejects H i if D i is large compared to the threshold value c can be written as AF NR = 1 k P { D i < c, θ i 0} k i=1 [ { } k 1 P D ( i) j:k 1 c D ] i < c, θ i k. (4.2) j(j + 1) j=1 If D is a multivariate normal with non-negative correlations, as it is the case under the above hierarchical mixture models, then the AFNR is no more than the right hand side of (4.2); see Sarkar (2003). If (D i, θ i ), i = 1,..., k, are iid, then (4.2) reduces to AF NR = P {A > 0}P {θ 1 0 D 1 < c}, (4.3) see also Storey (2003). Thus, under the above hierarchical mixture model and conditional on µ, ξ, and τ 2, the AFNR can be written as [ AF NR = 1 {1 ν} k] (1 π 0) [Φ (c 2 ) Φ (c 1 )], (4.4) ν 11
12 where ν, c 1 and c 2 are as defined in (3.5). The AFNR is the integral of (4.4) with respect to µ, ξ, and τ 2. It is a nondecreasing function of c, which can be proved using the same arguments as used in the case of AFDR. 5 Combining the AFDR and AFNR The AFDR and AFNR together constitute the Bayes risk of misclassifications. Our idea here is to determine the threshold that will minimize the Bayes risk in some sense. We consider a weighted linear combination of the AFDR and AFNR, defined as the Average Bayes Error Rate (ABER) of false rejections and false non-rejections, i.e., ABER = waf DR + (1 w)af NR, (5.1) with the weight 0 w 1 to the AFDR being determined by the importance of false rejections relative to false non-rejections, and find the threshold that minimizes the ABER. This is in the spirit of Storey (2003) and Genovese and Wasserman (2002b) who considered similar combinations in terms of the FDR and FNR. Storey (2003) points out there are two approaches that can be taken for the FDR: fix the FDR at the acceptable level α first and estimate the rejection region, or fix the rejection region first and provide an estimate of the FDR over that region. These approaches are practically useful, since the FDR is a monotonic function of c. Although we focus here on minimizing the ABER and finding its corresponding threshold, one can alternatively consider fixing the ABER, AFDR or AFNR and then estimating the threshold. By doing this, however, one may not be able to achieve the minimum of the ABER since the minimization process requires the input of threshold values. In what follows we will illustrate the 12
13 ABER minimization approach in a gene expression example and a simulation study where the thresholds at which the ABER is minimized will be provided. 6 An Application to Gene Expression Data Hereditary breast cancer is known to be associated with mutations in BRCA1 and BRCA2 proteins. Hedenfalk et al. (2001) report that a group of genes are differentially expressed between tumors with BRCA1 mutations and tumors with BRCA2 mutations. The data, which are publicly available from the web site consist of 22 breast cancer samples, among which n 1 = 7 are BRCA1 mutants, n 2 = 8 are BRCA2 mutants, and n 3 = 7 are sporadic (not used in this illustration). Expression levels, in terms of fluorescent intensity ratios of a tumor sample to a common reference sample, are measured for 3226 genes using cdna microarrays. As usual, the base 2 logarithmic transformation of the ratios was performed, from which ˆσ 2 and ˆσ 2 D were estimated to be and , respectively. We then computed the common two-sample t test statistic (t = D/ˆσ D, with 13 d.f.) and its corresponding raw p-value for each gene. Without multiplicity adjustment, there are 378 genes (out of 3226) whose raw p-values However, the most conservative Bonferroni-adjustment method suggests only 2 rejections at FWER 0.05 and the BH-FDR procedure declares 15 differentially expressed genes (adjusted p-value 0.05). Before applying our procedures to this data set, we first assume π 0 = 0.90, which, as Ishwaran and Rao (2003) pointed out, represents a fairly realistic scenario for gene expression data. Then we define the prior distribution g 1 (ξ, τ 2 ) = (1/τ 2 )g 2 (ξ); thus τ 2 is given the usual noninformative prior and ξ > 0 is given g 2 (ξ) = 1 ( ξ 3/2 exp 1 ). (6.1) 2π 2ξ 13
14 This prior results in f(µ τ 2 ) = Cauchy(0, τ) by integrating over ξ; see Berger, Boukai, and Wang (1997) for more discussion on the choice of this prior. The integrations with respect to µ, τ 2 and ξ in the calculations of the AFDR and AFNR under the hierarchical mixture model are carried out using Monte Carlo integration method. Specifically, we sample τ 2, ξ, and µ from their respective prior distributions and substitute these values into (3.5) and (4.4). The AFDR, AFNR and ABER are then obtained at a given c value by averaging over 5000 iterations. The AFDR and AFNR across c-values for the breast cancer data are graphically displayed (Figure 1). As one expected, the AFDR is decreasing and the AFNR is increasing in c. The ABER s were obtained and plotted against c for various weights w from 0.5, 0.6, 0.7, 0.8 to 0.9 (Figure 2). Clearly, one can always find a c value that minimizes the ABER for a given w. Note that all the ABER s for various w s cross at c = at which the AFDR and AFNR are approximately equal. We estimated critical value c that minimizes the ABER and then applied these critical values to the breast cancer data (Table 2). For instance, with π 0 = 0.9 and w = 0.9 there are 28 genes whose D i 1.48 that are declared differentially expressed between BRCA1 mutation tumors and BRCA2 mutation tumors. The AFDR and AFNR at c = 1.48 are and , respectively. The mean differences in gene expression levels between the two cancer types, together with the results of our approach and BH-FDR procedure, are shown in Figure 3. It is noted that the ABER approach picks up 13 more genes than the BH-FDR method, which is due to a higher power of the approach, as will be shown in the simulation studies of the next section. Since the ABER minimization results are dependent on the variance of D i and prior probability π 0, we provided the critical value c and the corresponding 14
15 ABER for ˆσ D 2 from 0.05 to 0.15 and π 0 = 0.85, 0.90, 0.95 (Tables 3). It can be seen that for the breast cancer data if ˆσ D 2 decreases from to 0.08, i.e., the number of expression measurements increases to 10 for each tumor type of each gene, then the critical value c decreases from 1.48 to 1.26, resulting more rejections and smaller misclassification rate (ABER drops from to ). Thus, Tables 3 can also be used in designing a microarray experiment. 7 Simulations We compare in this section our proposed AFDR and ABER approaches with the BH-FDR method in terms of some power using simulation studies. Specifically, we study the average power of AFDR, ABER and BH-FDR under the setup of hierarchical model, and then investigate the influence of outliers or prior (hyperprior) parameters on the performance of the methods, i.e., the robustness of the procedures to influential data points and prior information. 7.1 Power comparison The average power is defined in the frequentist context as the expected proportion of false null hypotheses that are correctly rejected and has been widely used in comparing multiple testing procedures (Benjamini and Hochberg 1995; Shaffer 1999; Benjamini and Liu 1999; Storey 2002). The setup of this simulation for average power comparison is as follows: k = 10, 50, 100, 5000, 1000; σd 2 = 0.1, 0.2, 0.3; π 0 = 0.2, 0.5, 0.8, 15
16 and a simulation is conducted for each combination of k, σ 2 D and π 0. Under the null hypothesis, i.e., θ i = 0, a data point is drawn from N(0, σd 2 ) and under the alternative hypothesis, i.e., θ i 0, a data point is drawn from N(θ i, σd 2 ), where θ i is an independent draw from N(µ, τ 2 ) distribution and µ follows Cauchy(0, τ 2 ) distribution given τ 2. A total number of kπ 0 null data points and k(1 π 0 ) alternative data points are randomly drawn for each set of k hypotheses. The average powers are obtained over 5000 simulations and plotted against k for AFDR 0.05, ABER with w =0.5, 0.7, 0.9 and BH-FDR 0.05 (Figure 4). It is clear that, as expected, the average power for all methods decreases with the increase in the number of hypotheses k, the variance of observed data σ 2 D, and the proportion of true null hypotheses π 0. The main point of the plot, however, is that our proposed approach, either AFDR 0.05 or minimum ABER with different w s, is more powerful than BH-FDR method. This is due to the fact that the BH-FDR procedure assigns the same FDR to all rejected hypotheses, i.e., the FDR is the same for all hypotheses with test statistics in the rejection region. On the other hand, the AFDR or ABER is the weighted average of the BH-FDR (and FNR) with the prior density of the parameter as the weight. By controlling the error rate at the same level, the AFDR or ABER approach would result in more rejections and hence more powerful than the BH-FDR method. 7.2 Robustness We study the robustness of AFDR, ABER and BH-FDR approaches to some influential data points as well as to various choices of hyper-prior information. To simplify the investigation, we first fix k = 1000, σd 2 = 0.1, π 0 = 0.80 and let the prior for ξ vary. The prior for τ 2 is still conventionally non-informative 1/τ 2, as it is not only computationally convenient but also practically indistinguishable from subjective prior when there are sufficient data (Berger and Deely 1988). 16
17 Towards this goal, we first generate influential or outlying data points, D i δn(θ i, σ 2 D) + (1 δ)n(θ i, σ 2 D), i = 1,..., k, (7.1) where δ is a random binary variable with P (δ = 1) = γ and θ i follows N(µ, c τ 2 ) distribution with some pre-specified c > 1. The hyper-prior for ξ is chosen as an inverse gamma density IG ( 1 2, β) with β being specified below. Note that IG(ξ; 1 2, β) leads to f(µ τ 2 ) = Cauchy(0, τ 2/β). The following setup is considered for the robustness simulation: γ = 0.90, 0.95, 0.99; c = 2, 4, 8; β = 0.25, 0.5, 1. As in the previous section, we considered all configurations of γ, c and β. The resulting average powers are obtained from 5000 simulations and plotted against β for each combination of γ and c (Figure 5). Since a large β makes the IG ( 1 2, β) density skew to the left, leading to an inflated variance of θ and consequently a decrease in average power, which is seen for all of the approaches. However, the average power for AFDR and ABER s is relatively flat and hence these approaches are more robust to the choice of ξ, as compared with BH-FDR method. Also, there is no practical impact of influential data points on the average power for all procedures, which is due to that all of the influential data points are generated from alternative population and are more likely to be rejected. 8 Discussion A different Bayesian approach is taken for measuring the risks of false rejections and false non-rejections in multiple testing. Although we illustrated this approach 17
18 to identifying differentially expressed genes from a microarray experiment, it can also be applied to other multiple testing situations. In an attempt to come up with a Bayesian measure of Type I error rate, we have started with the proportion of Type I errors among the total number of rejections, i.e., the proportion of false discoveries, before averaging it with respect to the distributions of data and prior. There is, however, another way one can measure Type I error rate, which is to consider the proportion of Type I errors among the hypotheses that are true and average it over data and parameters. In other words, one might consider the following alternative measure of Type I error rate, we call Bayesian False Positive Rate (BFPR): [ ( )] V BF P R = E θ E D θ. (8.1) k 0 1 It seems that a Bayesian would prefer (8.1) to the AFDR as a measure of Type I error rate, since it is based on the ratio measuring how many of the null hypotheses believed to be true are rejected by the data. Similarly, the Bayesian False Negative Rate (BFNR), defined as BF NR = E θ [ E D θ ( T k 1 1 )], (8.2) seems to be a more appropriate measure of Type II error rate to a Bayesian than the AFNR. If we use the hierarchical mixture models considered in Section 2 with the same definitions of parameters (i.e., π 0 = 0.9 and w = 0.9), then a linear combination of the BFPR and BFNR is minimized at c = 1.24, resulting in 54 rejections. Some issues surrounding the AFDR, AFNR and ABER, such as robustness of the results, remain to be investigated. We have arbitrarily chosen the prior distributions for convenience; other choices of priors could be considered and the impact of prior input on the results of rejections and non-rejections could also be evaluated. 18
19 Acknowledgement We would like to thank A. Lawrence Gould of Merck Research Laboratories for practical suggestions on the simulation, the Associate Editor and the anonymous referee for constructive comments that have improved the presentation of this paper. References Afshari, C. A., Nuwaysir, E. F., and Barrett, J. C. (1999). Application of complementary dna microarray technology to carcinogen identification, toxicology, and drug safety evaluation. Cancer Research 59, Baldi, P. and Long, A. D. (2001). A Bayesian framework for the analysis of microarray expression data: Regularized t-test and statistical inferences of gene changes. Bioinformatics 17, Benjamini, Y. and Hochberg, Y. (1995). Controlling the false discovery rate: A practical and powerful approach to multiple testing. Journal of the Royal Statistical Society, Series B 57, Benjamini, Y. and Liu, W. (1999). A step-down multiple hypothesis testing procedure that controls the false discovery rate under independence. Journal of Statistical Planning and Inference 82, Berger, J. O. (1985). Statistical Decision Theory and Bayesian Analysis (2nd ed.). New York: Springer-Verlag. Berger, J. O., Boukai, B., and Wang, Y. (1997). Unified frequentist and Bayesian testing of a precise hypothesis. Statistical Science 12, Berger, J. O. and Deely, J. (1988). A Bayesian approach to ranking and selection of related means with alternatives to analysis of variance methodology. 19
20 Journal of the American Statistical Association 83, Broët, P., Richardson, S., and Radvanyi, F. (2002). Bayesian hierarchical model for identifying changes in genes expression from microarray experiments. Journal of Computational Biology 9, Callow, M. J., Dudoit, S., Gong, E. L., Speed, T. P., and Rubin, E. M. (2000). Microarrays expression profiling identifies genes with altered expression in HDL-deficient mice. Genome Research 10, Chen, Y. J., Kodell, R., Sistare, F., Thompson, K. L., Morris, S., and Chen, J. J. (2003). Normalization methods for analysis of microarray geneexpression data. Journal of Biopharmaceutical Statistics 13, Dudoit, S., Shaffer, J. P., and Boldrick, J. C. (2003). Multiple hypothesis testing in microarray experiments. Statistical Science 18, Efron, B., Tibshirani, R., Storey, J. D., and Tusher, V. (2001). Empirical Bayes analysis of a microarray experiment. Journal of the American Statistical Association 96, Finkelstein, D. B., Gollub, J., and Cherry, J. M. (2001). Normalization and systematic measurement error in cdna microarray data. In ASA Proceedings of the Joint Statistical Meetings. Ge, Y., Dudoit, S., and Speed, T. P. (2003). Resampling-based multiple testing for microarray data analysis. Test 12, Genovese, C. and Wasserman, L. (2002a). Bayesian and frequentist multiple testing. Technical Report, Carnegie Mellon University. Genovese, C. and Wasserman, L. (2002b). Operating characteristics and extentions of the false discovery rate procedure. Journal of the Royal Statistical Society B. 64,
21 Gupta, S. D. and Sarkar, S. K. (1984). On tp 2 and log-concavity. In Y. L. Tong (Ed.), Inequalities in Statistics and Probability, pp Institute of Mathematical Statistics. Hedenfalk, I., Duggan, D., Chen, Y., Radmacher, M., Bittner, M., amd P. Meltzer, R. S., Gusterson, B., Esteller, M., Kallioniemi, O. P., Wilfond, B., Borg, A., and Trent, J. (2001). Gene-expression profiles in hereditary breast cancer. The New England Journal of Medicine 344, Ibrahim, J. G., Chen, M. H., and Gray, R. J. (2002). Bayesian models for gene expression with DNA microarray data. Journal of the American Statistical Association 97, Ishwaran, H. and Rao, J. S. (2003). Detecting differentially expressed genes in microassays usins Bayesian model selection. Journal of the American Statistical Association 98 (462), Newton, M. A., Noueiry, A., Sarkar, D., and Ahlquist, P. (2003). Detecting differential gene expression with a semiparametric hierarchical mixture method. Technical report, Department of Statistics, University of Wisconsin Madison. Technical Report #1074. Nuwaysir, E. F., Bittner, M., Trent, J., Barrett, J. C., and Afshari, C. A. (1999). Microarrays and toxicology: the advent of toxicogenomics. Molecular Carcinogenesis 24, Reiner, A., Yekutieli, D., and Benjamini, Y. (2003). Identifying differentially expressed genes using false discovery rate controlling procedures. Bioinformatics 19, Sarkar, S. K. (2003). False discovery and false non-discovery rates in single-step multiple testing procedures. Technical report, Temple University. Schervish, M. J. (1995). Theory of Statistics. New York: Springer-Verlag. 21
22 Shaffer, J. P. (1999). A semi-bayesian study of Duncan s Bayesian multiple comparison procedures. Journal of Statistical Planning and Inference 82, Storey, J. D. (2002). A direct approach to false discovery rates. Journal of the Royal Statistical Society, Series B 64, Storey, J. D. (2003). The positive false discovery rate: A Bayesian interpretation and the q-value. Annals of Statistics 31, Yang, Y. H., Dudoit, S., Luu, P., and Speed, T. P. (2001, January). Normalization for cdna microarray data. Technical Report 589, Department of Statistics, UC Berkeley. 22
23 AFNR 0.03 AFDR AFNR 0.15 AFDR c 0.00 Figure 1: The AFDR and AFNR as a function of critical value c for the breast cancer data with π 0 = ABER W= c Figure 2: The ABER as a function of critical value c for the breast cancer data with π 0 =
24 B B B B A A Mean Difference, di A B B B A B A A B AAA A B AA B -3.0 B Gene Figure 3: Detection of differentially expressed genes for the breast cancer data with π 0 = 0.90 and w = 0.9: B declared differentially expressed by both the ABER and BH FDR methods; A declared by the ABER method but not BH FDR method; not declared by either method. The two dashed horizontal lines represent the critical values c = ±1.48. Table 1: Outcomes of k Hypothesis Tests True State Accepted Rejected Total Null Hypotheses U V k 0 Alternative Hypotheses T S k 1 A R k 24
25 Table 2: Critical value c at which the ABER is minimized and the corresponding estimated AFDR and AFNR as well as the number of rejections for breast cancer data with π 0 = 0.9 w c AFDR AFNR ABER No. rejections
26 Table 3: Critical value c and the corresponding ABER for various combinations of π 0 and ˆσ 2 D c (ABER) π 0 ˆσ D 2 w = 0.5 w = 0.6 w = 0.7 w = 0.8 w = (0.0158) 0.84 (0.0129) 0.87 (0.0099) 0.90 (0.0068) 0.95 (0.0035) (0.0170) 0.92 (0.0139) 0.95 (0.0107) 0.99 (0.0073) 1.04 (0.0038) (0.0181) 0.99 (0.0149) 1.03 (0.0114) 1.06 (0.0078) 1.12 (0.0041) (0.0192) 1.06 (0.0157) 1.10 (0.0121) 1.14 (0.0083) 1.20 (0.0043) (0.0201) 1.12 (0.0165) 1.16 (0.0127) 1.21 (0.0087) 1.27 (0.0045) (0.0210) 1.19 (0.0172) 1.22 (0.0132) 1.27 (0.0090) 1.34 (0.0047) (0.0218) 1.24 (0.0179) 1.29 (0.0137) 1.33 (0.0094) 1.41 (0.0049) (0.0226) 1.30 (0.0185) 1.34 (0.0142) 1.40 (0.0097) 1.47 (0.0050) (0.0233) 1.35 (0.0191) 1.40 (0.0146) 1.45 (0.0100) 1.53 (0.0052) (0.0240) 1.40 (0.0196) 1.45 (0.0151) 1.51 (0.0103) 1.59 (0.0053) (0.0247) 1.46 (0.0202) 1.51 (0.0155) 1.57 (0.0106) 1.65 (0.0055) (0.0107) 0.90 (0.0087) 0.92 (0.0067) 0.95 (0.0046) 0.99 (0.0024) (0.0116) 0.98 (0.0094) 1.01 (0.0072) 1.04 (0.0050) 1.09 (0.0026) (0.0123) 1.06 (0.0101) 1.09 (0.0077) 1.12 (0.0053) 1.18 (0.0027) (0.0131) 1.13 (0.0107) 1.16 (0.0082) 1.20 (0.0056) 1.26 (0.0029) (0.0137) 1.20 (0.0112) 1.23 (0.0086) 1.27 (0.0059) 1.33 (0.0030) (0.0143) 1.26 (0.0117) 1.30 (0.0090) 1.34 (0.0061) 1.41 (0.0032) (0.0149) 1.32 (0.0122) 1.36 (0.0093) 1.41 (0.0064) 1.48 (0.0033) (0.0154) 1.38 (0.0126) 1.43 (0.0096) 1.47 (0.0066) 1.55 (0.0034) (0.0159) 1.44 (0.0130) 1.49 (0.0100) 1.54 (0.0068) 1.61 (0.0035) (0.0164) 1.50 (0.0134) 1.54 (0.0103) 1.60 (0.0070) 1.67 (0.0036) (0.0169) 1.55 (0.0138) 1.60 (0.0105) 1.66 (0.0072) 1.74 (0.0037) (0.0056) 0.98 (0.0045) 1.00 (0.0035) 1.03 (0.0024) 1.07 (0.0012) (0.0060) 1.07 (0.0049) 1.09 (0.0038) 1.12 (0.0026) 1.17 (0.0013) (0.0065) 1.15 (0.0053) 1.18 (0.0040) 1.21 (0.0027) 1.26 (0.0014) (0.0068) 1.23 (0.0056) 1.26 (0.0043) 1.30 (0.0029) 1.35 (0.0015) (0.0072) 1.31 (0.0058) 1.34 (0.0045) 1.38 (0.0030) 1.43 (0.0016) (0.0075) 1.38 (0.0061) 1.41 (0.0047) 1.45 (0.0032) 1.51 (0.0016) (0.0078) 1.45 (0.0064) 1.48 (0.0049) 1.53 (0.0033) 1.59 (0.0017) (0.0081) 1.51 (0.0066) 1.55 (0.0050) 1.60 (0.0034) 1.66 (0.0018) (0.0084) 1.58 (0.0068) 1.62 (0.0052) 1.67 (0.0035) 1.73 (0.0018) (0.0086) 1.64 (0.0070) 1.68 (0.0053) 1.73 (0.0036) 1.80 (0.0019) (0.0088) 1.70 (0.0072) 1.74 (0.0055) 1.80 (0.0037) 1.87 (0.0019) 26
27 σ = = = D σ D σ D π 0 = Average Power (%) π 0 = π 0 = Number of Hypotheses Tested AFDR ABERw = 0.5 ABERw = 0.7 ABERw = 0.9 BH-FDR Figure 4: Average power (the proportion of the false null hypotheses which are correctly rejected) for ABER s with different weights w(= 0.5, 0.7, 0.9), AFDR and BH-FDR at various combinations of π 0 and σ 2 D. 27
28 γ=0.90 γ=0.95 γ= c'=8 90 Average Power (%) c'= c'= β AFDR ABERw = 0.5 ABERw = 0.7 ABERw = 0.9 BH-FDR Figure 5: Average power for ABER s with different weights w(= 0.5, 0.7, 0.9), AFDR and BH-FDR at various combinations of the choice of β, γ and c. 28
A Bayesian Determination of Threshold for Identifying Differentially Expressed Genes in Microarray Experiments
 A Bayesian Determination of Threshold for Identifying Differentially Expressed Genes in Microarray Experiments Jie Chen 1 Merck Research Laboratories, P. O. Box 4, BL3-2, West Point, PA 19486, U.S.A. Telephone:
A Bayesian Determination of Threshold for Identifying Differentially Expressed Genes in Microarray Experiments Jie Chen 1 Merck Research Laboratories, P. O. Box 4, BL3-2, West Point, PA 19486, U.S.A. Telephone:
Statistical Applications in Genetics and Molecular Biology
 Statistical Applications in Genetics and Molecular Biology Volume 5, Issue 1 2006 Article 28 A Two-Step Multiple Comparison Procedure for a Large Number of Tests and Multiple Treatments Hongmei Jiang Rebecca
Statistical Applications in Genetics and Molecular Biology Volume 5, Issue 1 2006 Article 28 A Two-Step Multiple Comparison Procedure for a Large Number of Tests and Multiple Treatments Hongmei Jiang Rebecca
STEPDOWN PROCEDURES CONTROLLING A GENERALIZED FALSE DISCOVERY RATE. National Institute of Environmental Health Sciences and Temple University
 STEPDOWN PROCEDURES CONTROLLING A GENERALIZED FALSE DISCOVERY RATE Wenge Guo 1 and Sanat K. Sarkar 2 National Institute of Environmental Health Sciences and Temple University Abstract: Often in practice
STEPDOWN PROCEDURES CONTROLLING A GENERALIZED FALSE DISCOVERY RATE Wenge Guo 1 and Sanat K. Sarkar 2 National Institute of Environmental Health Sciences and Temple University Abstract: Often in practice
A GENERAL DECISION THEORETIC FORMULATION OF PROCEDURES CONTROLLING FDR AND FNR FROM A BAYESIAN PERSPECTIVE
 A GENERAL DECISION THEORETIC FORMULATION OF PROCEDURES CONTROLLING FDR AND FNR FROM A BAYESIAN PERSPECTIVE Sanat K. Sarkar 1, Tianhui Zhou and Debashis Ghosh Temple University, Wyeth Pharmaceuticals and
A GENERAL DECISION THEORETIC FORMULATION OF PROCEDURES CONTROLLING FDR AND FNR FROM A BAYESIAN PERSPECTIVE Sanat K. Sarkar 1, Tianhui Zhou and Debashis Ghosh Temple University, Wyeth Pharmaceuticals and
Modified Simes Critical Values Under Positive Dependence
 Modified Simes Critical Values Under Positive Dependence Gengqian Cai, Sanat K. Sarkar Clinical Pharmacology Statistics & Programming, BDS, GlaxoSmithKline Statistics Department, Temple University, Philadelphia
Modified Simes Critical Values Under Positive Dependence Gengqian Cai, Sanat K. Sarkar Clinical Pharmacology Statistics & Programming, BDS, GlaxoSmithKline Statistics Department, Temple University, Philadelphia
Controlling Bayes Directional False Discovery Rate in Random Effects Model 1
 Controlling Bayes Directional False Discovery Rate in Random Effects Model 1 Sanat K. Sarkar a, Tianhui Zhou b a Temple University, Philadelphia, PA 19122, USA b Wyeth Pharmaceuticals, Collegeville, PA
Controlling Bayes Directional False Discovery Rate in Random Effects Model 1 Sanat K. Sarkar a, Tianhui Zhou b a Temple University, Philadelphia, PA 19122, USA b Wyeth Pharmaceuticals, Collegeville, PA
Chapter 1. Stepdown Procedures Controlling A Generalized False Discovery Rate
 Chapter Stepdown Procedures Controlling A Generalized False Discovery Rate Wenge Guo and Sanat K. Sarkar Biostatistics Branch, National Institute of Environmental Health Sciences, Research Triangle Park,
Chapter Stepdown Procedures Controlling A Generalized False Discovery Rate Wenge Guo and Sanat K. Sarkar Biostatistics Branch, National Institute of Environmental Health Sciences, Research Triangle Park,
FDR-CONTROLLING STEPWISE PROCEDURES AND THEIR FALSE NEGATIVES RATES
 FDR-CONTROLLING STEPWISE PROCEDURES AND THEIR FALSE NEGATIVES RATES Sanat K. Sarkar a a Department of Statistics, Temple University, Speakman Hall (006-00), Philadelphia, PA 19122, USA Abstract The concept
FDR-CONTROLLING STEPWISE PROCEDURES AND THEIR FALSE NEGATIVES RATES Sanat K. Sarkar a a Department of Statistics, Temple University, Speakman Hall (006-00), Philadelphia, PA 19122, USA Abstract The concept
Controlling the False Discovery Rate: Understanding and Extending the Benjamini-Hochberg Method
 Controlling the False Discovery Rate: Understanding and Extending the Benjamini-Hochberg Method Christopher R. Genovese Department of Statistics Carnegie Mellon University joint work with Larry Wasserman
Controlling the False Discovery Rate: Understanding and Extending the Benjamini-Hochberg Method Christopher R. Genovese Department of Statistics Carnegie Mellon University joint work with Larry Wasserman
Procedures controlling generalized false discovery rate
 rocedures controlling generalized false discovery rate By SANAT K. SARKAR Department of Statistics, Temple University, hiladelphia, A 922, U.S.A. sanat@temple.edu AND WENGE GUO Department of Environmental
rocedures controlling generalized false discovery rate By SANAT K. SARKAR Department of Statistics, Temple University, hiladelphia, A 922, U.S.A. sanat@temple.edu AND WENGE GUO Department of Environmental
MIXTURE MODELS FOR DETECTING DIFFERENTIALLY EXPRESSED GENES IN MICROARRAYS
 International Journal of Neural Systems, Vol. 16, No. 5 (2006) 353 362 c World Scientific Publishing Company MIXTURE MOLS FOR TECTING DIFFERENTIALLY EXPRESSED GENES IN MICROARRAYS LIAT BEN-TOVIM JONES
International Journal of Neural Systems, Vol. 16, No. 5 (2006) 353 362 c World Scientific Publishing Company MIXTURE MOLS FOR TECTING DIFFERENTIALLY EXPRESSED GENES IN MICROARRAYS LIAT BEN-TOVIM JONES
Comparison of the Empirical Bayes and the Significance Analysis of Microarrays
 Comparison of the Empirical Bayes and the Significance Analysis of Microarrays Holger Schwender, Andreas Krause, and Katja Ickstadt Abstract Microarrays enable to measure the expression levels of tens
Comparison of the Empirical Bayes and the Significance Analysis of Microarrays Holger Schwender, Andreas Krause, and Katja Ickstadt Abstract Microarrays enable to measure the expression levels of tens
The miss rate for the analysis of gene expression data
 Biostatistics (2005), 6, 1,pp. 111 117 doi: 10.1093/biostatistics/kxh021 The miss rate for the analysis of gene expression data JONATHAN TAYLOR Department of Statistics, Stanford University, Stanford,
Biostatistics (2005), 6, 1,pp. 111 117 doi: 10.1093/biostatistics/kxh021 The miss rate for the analysis of gene expression data JONATHAN TAYLOR Department of Statistics, Stanford University, Stanford,
Summary and discussion of: Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing
 Summary and discussion of: Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing Statistics Journal Club, 36-825 Beau Dabbs and Philipp Burckhardt 9-19-2014 1 Paper
Summary and discussion of: Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing Statistics Journal Club, 36-825 Beau Dabbs and Philipp Burckhardt 9-19-2014 1 Paper
FALSE DISCOVERY AND FALSE NONDISCOVERY RATES IN SINGLE-STEP MULTIPLE TESTING PROCEDURES 1. BY SANAT K. SARKAR Temple University
 The Annals of Statistics 2006, Vol. 34, No. 1, 394 415 DOI: 10.1214/009053605000000778 Institute of Mathematical Statistics, 2006 FALSE DISCOVERY AND FALSE NONDISCOVERY RATES IN SINGLE-STEP MULTIPLE TESTING
The Annals of Statistics 2006, Vol. 34, No. 1, 394 415 DOI: 10.1214/009053605000000778 Institute of Mathematical Statistics, 2006 FALSE DISCOVERY AND FALSE NONDISCOVERY RATES IN SINGLE-STEP MULTIPLE TESTING
False discovery rate and related concepts in multiple comparisons problems, with applications to microarray data
 False discovery rate and related concepts in multiple comparisons problems, with applications to microarray data Ståle Nygård Trial Lecture Dec 19, 2008 1 / 35 Lecture outline Motivation for not using
False discovery rate and related concepts in multiple comparisons problems, with applications to microarray data Ståle Nygård Trial Lecture Dec 19, 2008 1 / 35 Lecture outline Motivation for not using
High-throughput Testing
 High-throughput Testing Noah Simon and Richard Simon July 2016 1 / 29 Testing vs Prediction On each of n patients measure y i - single binary outcome (eg. progression after a year, PCR) x i - p-vector
High-throughput Testing Noah Simon and Richard Simon July 2016 1 / 29 Testing vs Prediction On each of n patients measure y i - single binary outcome (eg. progression after a year, PCR) x i - p-vector
A moment-based method for estimating the proportion of true null hypotheses and its application to microarray gene expression data
 Biostatistics (2007), 8, 4, pp. 744 755 doi:10.1093/biostatistics/kxm002 Advance Access publication on January 22, 2007 A moment-based method for estimating the proportion of true null hypotheses and its
Biostatistics (2007), 8, 4, pp. 744 755 doi:10.1093/biostatistics/kxm002 Advance Access publication on January 22, 2007 A moment-based method for estimating the proportion of true null hypotheses and its
PROCEDURES CONTROLLING THE k-fdr USING. BIVARIATE DISTRIBUTIONS OF THE NULL p-values. Sanat K. Sarkar and Wenge Guo
 PROCEDURES CONTROLLING THE k-fdr USING BIVARIATE DISTRIBUTIONS OF THE NULL p-values Sanat K. Sarkar and Wenge Guo Temple University and National Institute of Environmental Health Sciences Abstract: Procedures
PROCEDURES CONTROLLING THE k-fdr USING BIVARIATE DISTRIBUTIONS OF THE NULL p-values Sanat K. Sarkar and Wenge Guo Temple University and National Institute of Environmental Health Sciences Abstract: Procedures
Research Article Sample Size Calculation for Controlling False Discovery Proportion
 Probability and Statistics Volume 2012, Article ID 817948, 13 pages doi:10.1155/2012/817948 Research Article Sample Size Calculation for Controlling False Discovery Proportion Shulian Shang, 1 Qianhe Zhou,
Probability and Statistics Volume 2012, Article ID 817948, 13 pages doi:10.1155/2012/817948 Research Article Sample Size Calculation for Controlling False Discovery Proportion Shulian Shang, 1 Qianhe Zhou,
A Large-Sample Approach to Controlling the False Discovery Rate
 A Large-Sample Approach to Controlling the False Discovery Rate Christopher R. Genovese Department of Statistics Carnegie Mellon University Larry Wasserman Department of Statistics Carnegie Mellon University
A Large-Sample Approach to Controlling the False Discovery Rate Christopher R. Genovese Department of Statistics Carnegie Mellon University Larry Wasserman Department of Statistics Carnegie Mellon University
Step-down FDR Procedures for Large Numbers of Hypotheses
 Step-down FDR Procedures for Large Numbers of Hypotheses Paul N. Somerville University of Central Florida Abstract. Somerville (2004b) developed FDR step-down procedures which were particularly appropriate
Step-down FDR Procedures for Large Numbers of Hypotheses Paul N. Somerville University of Central Florida Abstract. Somerville (2004b) developed FDR step-down procedures which were particularly appropriate
Parametric Empirical Bayes Methods for Microarrays
 Parametric Empirical Bayes Methods for Microarrays Ming Yuan, Deepayan Sarkar, Michael Newton and Christina Kendziorski April 30, 2018 Contents 1 Introduction 1 2 General Model Structure: Two Conditions
Parametric Empirical Bayes Methods for Microarrays Ming Yuan, Deepayan Sarkar, Michael Newton and Christina Kendziorski April 30, 2018 Contents 1 Introduction 1 2 General Model Structure: Two Conditions
Two-stage stepup procedures controlling FDR
 Journal of Statistical Planning and Inference 38 (2008) 072 084 www.elsevier.com/locate/jspi Two-stage stepup procedures controlling FDR Sanat K. Sarar Department of Statistics, Temple University, Philadelphia,
Journal of Statistical Planning and Inference 38 (2008) 072 084 www.elsevier.com/locate/jspi Two-stage stepup procedures controlling FDR Sanat K. Sarar Department of Statistics, Temple University, Philadelphia,
Table of Outcomes. Table of Outcomes. Table of Outcomes. Table of Outcomes. Table of Outcomes. Table of Outcomes. T=number of type 2 errors
 The Multiple Testing Problem Multiple Testing Methods for the Analysis of Microarray Data 3/9/2009 Copyright 2009 Dan Nettleton Suppose one test of interest has been conducted for each of m genes in a
The Multiple Testing Problem Multiple Testing Methods for the Analysis of Microarray Data 3/9/2009 Copyright 2009 Dan Nettleton Suppose one test of interest has been conducted for each of m genes in a
Large-Scale Hypothesis Testing
 Chapter 2 Large-Scale Hypothesis Testing Progress in statistics is usually at the mercy of our scientific colleagues, whose data is the nature from which we work. Agricultural experimentation in the early
Chapter 2 Large-Scale Hypothesis Testing Progress in statistics is usually at the mercy of our scientific colleagues, whose data is the nature from which we work. Agricultural experimentation in the early
A Sequential Bayesian Approach with Applications to Circadian Rhythm Microarray Gene Expression Data
 A Sequential Bayesian Approach with Applications to Circadian Rhythm Microarray Gene Expression Data Faming Liang, Chuanhai Liu, and Naisyin Wang Texas A&M University Multiple Hypothesis Testing Introduction
A Sequential Bayesian Approach with Applications to Circadian Rhythm Microarray Gene Expression Data Faming Liang, Chuanhai Liu, and Naisyin Wang Texas A&M University Multiple Hypothesis Testing Introduction
Statistical testing. Samantha Kleinberg. October 20, 2009
 October 20, 2009 Intro to significance testing Significance testing and bioinformatics Gene expression: Frequently have microarray data for some group of subjects with/without the disease. Want to find
October 20, 2009 Intro to significance testing Significance testing and bioinformatics Gene expression: Frequently have microarray data for some group of subjects with/without the disease. Want to find
Comments on: Control of the false discovery rate under dependence using the bootstrap and subsampling
 Test (2008) 17: 443 445 DOI 10.1007/s11749-008-0127-5 DISCUSSION Comments on: Control of the false discovery rate under dependence using the bootstrap and subsampling José A. Ferreira Mark A. van de Wiel
Test (2008) 17: 443 445 DOI 10.1007/s11749-008-0127-5 DISCUSSION Comments on: Control of the false discovery rate under dependence using the bootstrap and subsampling José A. Ferreira Mark A. van de Wiel
The Pennsylvania State University The Graduate School A BAYESIAN APPROACH TO FALSE DISCOVERY RATE FOR LARGE SCALE SIMULTANEOUS INFERENCE
 The Pennsylvania State University The Graduate School A BAYESIAN APPROACH TO FALSE DISCOVERY RATE FOR LARGE SCALE SIMULTANEOUS INFERENCE A Thesis in Statistics by Bing Han c 2007 Bing Han Submitted in
The Pennsylvania State University The Graduate School A BAYESIAN APPROACH TO FALSE DISCOVERY RATE FOR LARGE SCALE SIMULTANEOUS INFERENCE A Thesis in Statistics by Bing Han c 2007 Bing Han Submitted in
High-Throughput Sequencing Course. Introduction. Introduction. Multiple Testing. Biostatistics and Bioinformatics. Summer 2018
 High-Throughput Sequencing Course Multiple Testing Biostatistics and Bioinformatics Summer 2018 Introduction You have previously considered the significance of a single gene Introduction You have previously
High-Throughput Sequencing Course Multiple Testing Biostatistics and Bioinformatics Summer 2018 Introduction You have previously considered the significance of a single gene Introduction You have previously
Doing Cosmology with Balls and Envelopes
 Doing Cosmology with Balls and Envelopes Christopher R. Genovese Department of Statistics Carnegie Mellon University http://www.stat.cmu.edu/ ~ genovese/ Larry Wasserman Department of Statistics Carnegie
Doing Cosmology with Balls and Envelopes Christopher R. Genovese Department of Statistics Carnegie Mellon University http://www.stat.cmu.edu/ ~ genovese/ Larry Wasserman Department of Statistics Carnegie
Fall 2017 STAT 532 Homework Peter Hoff. 1. Let P be a probability measure on a collection of sets A.
 1. Let P be a probability measure on a collection of sets A. (a) For each n N, let H n be a set in A such that H n H n+1. Show that P (H n ) monotonically converges to P ( k=1 H k) as n. (b) For each n
1. Let P be a probability measure on a collection of sets A. (a) For each n N, let H n be a set in A such that H n H n+1. Show that P (H n ) monotonically converges to P ( k=1 H k) as n. (b) For each n
FDR and ROC: Similarities, Assumptions, and Decisions
 EDITORIALS 8 FDR and ROC: Similarities, Assumptions, and Decisions. Why FDR and ROC? It is a privilege to have been asked to introduce this collection of papers appearing in Statistica Sinica. The papers
EDITORIALS 8 FDR and ROC: Similarities, Assumptions, and Decisions. Why FDR and ROC? It is a privilege to have been asked to introduce this collection of papers appearing in Statistica Sinica. The papers
POSITIVE FALSE DISCOVERY PROPORTIONS: INTRINSIC BOUNDS AND ADAPTIVE CONTROL
 Statistica Sinica 18(2008, 837-860 POSITIVE FALSE DISCOVERY PROPORTIONS: INTRINSIC BOUNDS AND ADAPTIVE CONTROL Zhiyi Chi and Zhiqiang Tan University of Connecticut and Rutgers University Abstract: A useful
Statistica Sinica 18(2008, 837-860 POSITIVE FALSE DISCOVERY PROPORTIONS: INTRINSIC BOUNDS AND ADAPTIVE CONTROL Zhiyi Chi and Zhiqiang Tan University of Connecticut and Rutgers University Abstract: A useful
EMPIRICAL BAYES METHODS FOR ESTIMATION AND CONFIDENCE INTERVALS IN HIGH-DIMENSIONAL PROBLEMS
 Statistica Sinica 19 (2009), 125-143 EMPIRICAL BAYES METHODS FOR ESTIMATION AND CONFIDENCE INTERVALS IN HIGH-DIMENSIONAL PROBLEMS Debashis Ghosh Penn State University Abstract: There is much recent interest
Statistica Sinica 19 (2009), 125-143 EMPIRICAL BAYES METHODS FOR ESTIMATION AND CONFIDENCE INTERVALS IN HIGH-DIMENSIONAL PROBLEMS Debashis Ghosh Penn State University Abstract: There is much recent interest
A Unified Computational Framework to Compare Direct and Sequential False Discovery Rate Algorithms for Exploratory DNA Microarray Studies
 Journal of Data Science 3(2005), 331-352 A Unified Computational Framework to Compare Direct and Sequential False Discovery Rate Algorithms for Exploratory DNA Microarray Studies Danh V. Nguyen University
Journal of Data Science 3(2005), 331-352 A Unified Computational Framework to Compare Direct and Sequential False Discovery Rate Algorithms for Exploratory DNA Microarray Studies Danh V. Nguyen University
Journal of Statistical Software
 JSS Journal of Statistical Software MMMMMM YYYY, Volume VV, Issue II. doi: 10.18637/jss.v000.i00 GroupTest: Multiple Testing Procedure for Grouped Hypotheses Zhigen Zhao Abstract In the modern Big Data
JSS Journal of Statistical Software MMMMMM YYYY, Volume VV, Issue II. doi: 10.18637/jss.v000.i00 GroupTest: Multiple Testing Procedure for Grouped Hypotheses Zhigen Zhao Abstract In the modern Big Data
Resampling-Based Control of the FDR
 Resampling-Based Control of the FDR Joseph P. Romano 1 Azeem S. Shaikh 2 and Michael Wolf 3 1 Departments of Economics and Statistics Stanford University 2 Department of Economics University of Chicago
Resampling-Based Control of the FDR Joseph P. Romano 1 Azeem S. Shaikh 2 and Michael Wolf 3 1 Departments of Economics and Statistics Stanford University 2 Department of Economics University of Chicago
Probabilistic Inference for Multiple Testing
 This is the title page! This is the title page! Probabilistic Inference for Multiple Testing Chuanhai Liu and Jun Xie Department of Statistics, Purdue University, West Lafayette, IN 47907. E-mail: chuanhai,
This is the title page! This is the title page! Probabilistic Inference for Multiple Testing Chuanhai Liu and Jun Xie Department of Statistics, Purdue University, West Lafayette, IN 47907. E-mail: chuanhai,
Stat 206: Estimation and testing for a mean vector,
 Stat 206: Estimation and testing for a mean vector, Part II James Johndrow 2016-12-03 Comparing components of the mean vector In the last part, we talked about testing the hypothesis H 0 : µ 1 = µ 2 where
Stat 206: Estimation and testing for a mean vector, Part II James Johndrow 2016-12-03 Comparing components of the mean vector In the last part, we talked about testing the hypothesis H 0 : µ 1 = µ 2 where
Multiple Testing. Hoang Tran. Department of Statistics, Florida State University
 Multiple Testing Hoang Tran Department of Statistics, Florida State University Large-Scale Testing Examples: Microarray data: testing differences in gene expression between two traits/conditions Microbiome
Multiple Testing Hoang Tran Department of Statistics, Florida State University Large-Scale Testing Examples: Microarray data: testing differences in gene expression between two traits/conditions Microbiome
Quick Calculation for Sample Size while Controlling False Discovery Rate with Application to Microarray Analysis
 Statistics Preprints Statistics 11-2006 Quick Calculation for Sample Size while Controlling False Discovery Rate with Application to Microarray Analysis Peng Liu Iowa State University, pliu@iastate.edu
Statistics Preprints Statistics 11-2006 Quick Calculation for Sample Size while Controlling False Discovery Rate with Application to Microarray Analysis Peng Liu Iowa State University, pliu@iastate.edu
Aliaksandr Hubin University of Oslo Aliaksandr Hubin (UIO) Bayesian FDR / 25
 Presentation of The Paper: The Positive False Discovery Rate: A Bayesian Interpretation and the q-value, J.D. Storey, The Annals of Statistics, Vol. 31 No.6 (Dec. 2003), pp 2013-2035 Aliaksandr Hubin University
Presentation of The Paper: The Positive False Discovery Rate: A Bayesian Interpretation and the q-value, J.D. Storey, The Annals of Statistics, Vol. 31 No.6 (Dec. 2003), pp 2013-2035 Aliaksandr Hubin University
A BAYESIAN STEPWISE MULTIPLE TESTING PROCEDURE. By Sanat K. Sarkar 1 and Jie Chen. Temple University and Merck Research Laboratories
 A BAYESIAN STEPWISE MULTIPLE TESTING PROCEDURE By Sanat K. Sarar 1 and Jie Chen Temple University and Merc Research Laboratories Abstract Bayesian testing of multiple hypotheses often requires consideration
A BAYESIAN STEPWISE MULTIPLE TESTING PROCEDURE By Sanat K. Sarar 1 and Jie Chen Temple University and Merc Research Laboratories Abstract Bayesian testing of multiple hypotheses often requires consideration
CHOOSING THE LESSER EVIL: TRADE-OFF BETWEEN FALSE DISCOVERY RATE AND NON-DISCOVERY RATE
 Statistica Sinica 18(2008), 861-879 CHOOSING THE LESSER EVIL: TRADE-OFF BETWEEN FALSE DISCOVERY RATE AND NON-DISCOVERY RATE Radu V. Craiu and Lei Sun University of Toronto Abstract: The problem of multiple
Statistica Sinica 18(2008), 861-879 CHOOSING THE LESSER EVIL: TRADE-OFF BETWEEN FALSE DISCOVERY RATE AND NON-DISCOVERY RATE Radu V. Craiu and Lei Sun University of Toronto Abstract: The problem of multiple
Hierarchical Mixture Models for Expression Profiles
 2 Hierarchical Mixture Models for Expression Profiles MICHAEL A. NEWTON, PING WANG, AND CHRISTINA KENDZIORSKI University of Wisconsin at Madison Abstract A class of probability models for inference about
2 Hierarchical Mixture Models for Expression Profiles MICHAEL A. NEWTON, PING WANG, AND CHRISTINA KENDZIORSKI University of Wisconsin at Madison Abstract A class of probability models for inference about
Improved Statistical Tests for Differential Gene Expression by Shrinking Variance Components Estimates
 Improved Statistical Tests for Differential Gene Expression by Shrinking Variance Components Estimates September 4, 2003 Xiangqin Cui, J. T. Gene Hwang, Jing Qiu, Natalie J. Blades, and Gary A. Churchill
Improved Statistical Tests for Differential Gene Expression by Shrinking Variance Components Estimates September 4, 2003 Xiangqin Cui, J. T. Gene Hwang, Jing Qiu, Natalie J. Blades, and Gary A. Churchill
Control of Directional Errors in Fixed Sequence Multiple Testing
 Control of Directional Errors in Fixed Sequence Multiple Testing Anjana Grandhi Department of Mathematical Sciences New Jersey Institute of Technology Newark, NJ 07102-1982 Wenge Guo Department of Mathematical
Control of Directional Errors in Fixed Sequence Multiple Testing Anjana Grandhi Department of Mathematical Sciences New Jersey Institute of Technology Newark, NJ 07102-1982 Wenge Guo Department of Mathematical
Exceedance Control of the False Discovery Proportion Christopher Genovese 1 and Larry Wasserman 2 Carnegie Mellon University July 10, 2004
 Exceedance Control of the False Discovery Proportion Christopher Genovese 1 and Larry Wasserman 2 Carnegie Mellon University July 10, 2004 Multiple testing methods to control the False Discovery Rate (FDR),
Exceedance Control of the False Discovery Proportion Christopher Genovese 1 and Larry Wasserman 2 Carnegie Mellon University July 10, 2004 Multiple testing methods to control the False Discovery Rate (FDR),
Applying the Benjamini Hochberg procedure to a set of generalized p-values
 U.U.D.M. Report 20:22 Applying the Benjamini Hochberg procedure to a set of generalized p-values Fredrik Jonsson Department of Mathematics Uppsala University Applying the Benjamini Hochberg procedure
U.U.D.M. Report 20:22 Applying the Benjamini Hochberg procedure to a set of generalized p-values Fredrik Jonsson Department of Mathematics Uppsala University Applying the Benjamini Hochberg procedure
Statistical Data Analysis Stat 3: p-values, parameter estimation
 Statistical Data Analysis Stat 3: p-values, parameter estimation London Postgraduate Lectures on Particle Physics; University of London MSci course PH4515 Glen Cowan Physics Department Royal Holloway,
Statistical Data Analysis Stat 3: p-values, parameter estimation London Postgraduate Lectures on Particle Physics; University of London MSci course PH4515 Glen Cowan Physics Department Royal Holloway,
arxiv: v3 [math.st] 15 Jul 2018
![arxiv: v3 [math.st] 15 Jul 2018 arxiv: v3 [math.st] 15 Jul 2018](/thumbs/94/118060858.jpg) A New Step-down Procedure for Simultaneous Hypothesis Testing Under Dependence arxiv:1503.08923v3 [math.st] 15 Jul 2018 Contents Prasenjit Ghosh 1 and Arijit Chakrabarti 2 1 Department of Statistics, Presidency
A New Step-down Procedure for Simultaneous Hypothesis Testing Under Dependence arxiv:1503.08923v3 [math.st] 15 Jul 2018 Contents Prasenjit Ghosh 1 and Arijit Chakrabarti 2 1 Department of Statistics, Presidency
Statistical Applications in Genetics and Molecular Biology
 Statistical Applications in Genetics and Molecular Biology Volume 6, Issue 1 2007 Article 28 A Comparison of Methods to Control Type I Errors in Microarray Studies Jinsong Chen Mark J. van der Laan Martyn
Statistical Applications in Genetics and Molecular Biology Volume 6, Issue 1 2007 Article 28 A Comparison of Methods to Control Type I Errors in Microarray Studies Jinsong Chen Mark J. van der Laan Martyn
On Methods Controlling the False Discovery Rate 1
 Sankhyā : The Indian Journal of Statistics 2008, Volume 70-A, Part 2, pp. 135-168 c 2008, Indian Statistical Institute On Methods Controlling the False Discovery Rate 1 Sanat K. Sarkar Temple University,
Sankhyā : The Indian Journal of Statistics 2008, Volume 70-A, Part 2, pp. 135-168 c 2008, Indian Statistical Institute On Methods Controlling the False Discovery Rate 1 Sanat K. Sarkar Temple University,
29 Sample Size Choice for Microarray Experiments
 29 Sample Size Choice for Microarray Experiments Peter Müller, M.D. Anderson Cancer Center Christian Robert and Judith Rousseau CREST, Paris Abstract We review Bayesian sample size arguments for microarray
29 Sample Size Choice for Microarray Experiments Peter Müller, M.D. Anderson Cancer Center Christian Robert and Judith Rousseau CREST, Paris Abstract We review Bayesian sample size arguments for microarray
Familywise Error Rate Controlling Procedures for Discrete Data
 Familywise Error Rate Controlling Procedures for Discrete Data arxiv:1711.08147v1 [stat.me] 22 Nov 2017 Yalin Zhu Center for Mathematical Sciences, Merck & Co., Inc., West Point, PA, U.S.A. Wenge Guo Department
Familywise Error Rate Controlling Procedures for Discrete Data arxiv:1711.08147v1 [stat.me] 22 Nov 2017 Yalin Zhu Center for Mathematical Sciences, Merck & Co., Inc., West Point, PA, U.S.A. Wenge Guo Department
Rejoinder on: Control of the false discovery rate under dependence using the bootstrap and subsampling
 Test (2008) 17: 461 471 DOI 10.1007/s11749-008-0134-6 DISCUSSION Rejoinder on: Control of the false discovery rate under dependence using the bootstrap and subsampling Joseph P. Romano Azeem M. Shaikh
Test (2008) 17: 461 471 DOI 10.1007/s11749-008-0134-6 DISCUSSION Rejoinder on: Control of the false discovery rate under dependence using the bootstrap and subsampling Joseph P. Romano Azeem M. Shaikh
ON STEPWISE CONTROL OF THE GENERALIZED FAMILYWISE ERROR RATE. By Wenge Guo and M. Bhaskara Rao
 ON STEPWISE CONTROL OF THE GENERALIZED FAMILYWISE ERROR RATE By Wenge Guo and M. Bhaskara Rao National Institute of Environmental Health Sciences and University of Cincinnati A classical approach for dealing
ON STEPWISE CONTROL OF THE GENERALIZED FAMILYWISE ERROR RATE By Wenge Guo and M. Bhaskara Rao National Institute of Environmental Health Sciences and University of Cincinnati A classical approach for dealing
On Procedures Controlling the FDR for Testing Hierarchically Ordered Hypotheses
 On Procedures Controlling the FDR for Testing Hierarchically Ordered Hypotheses Gavin Lynch Catchpoint Systems, Inc., 228 Park Ave S 28080 New York, NY 10003, U.S.A. Wenge Guo Department of Mathematical
On Procedures Controlling the FDR for Testing Hierarchically Ordered Hypotheses Gavin Lynch Catchpoint Systems, Inc., 228 Park Ave S 28080 New York, NY 10003, U.S.A. Wenge Guo Department of Mathematical
Sample Size Estimation for Studies of High-Dimensional Data
 Sample Size Estimation for Studies of High-Dimensional Data James J. Chen, Ph.D. National Center for Toxicological Research Food and Drug Administration June 3, 2009 China Medical University Taichung,
Sample Size Estimation for Studies of High-Dimensional Data James J. Chen, Ph.D. National Center for Toxicological Research Food and Drug Administration June 3, 2009 China Medical University Taichung,
This paper has been submitted for consideration for publication in Biometrics
 BIOMETRICS, 1 10 Supplementary material for Control with Pseudo-Gatekeeping Based on a Possibly Data Driven er of the Hypotheses A. Farcomeni Department of Public Health and Infectious Diseases Sapienza
BIOMETRICS, 1 10 Supplementary material for Control with Pseudo-Gatekeeping Based on a Possibly Data Driven er of the Hypotheses A. Farcomeni Department of Public Health and Infectious Diseases Sapienza
Non-specific filtering and control of false positives
 Non-specific filtering and control of false positives Richard Bourgon 16 June 2009 bourgon@ebi.ac.uk EBI is an outstation of the European Molecular Biology Laboratory Outline Multiple testing I: overview
Non-specific filtering and control of false positives Richard Bourgon 16 June 2009 bourgon@ebi.ac.uk EBI is an outstation of the European Molecular Biology Laboratory Outline Multiple testing I: overview
The optimal discovery procedure: a new approach to simultaneous significance testing
 J. R. Statist. Soc. B (2007) 69, Part 3, pp. 347 368 The optimal discovery procedure: a new approach to simultaneous significance testing John D. Storey University of Washington, Seattle, USA [Received
J. R. Statist. Soc. B (2007) 69, Part 3, pp. 347 368 The optimal discovery procedure: a new approach to simultaneous significance testing John D. Storey University of Washington, Seattle, USA [Received
Permutation Test for Bayesian Variable Selection Method for Modelling Dose-Response Data Under Simple Order Restrictions
 Permutation Test for Bayesian Variable Selection Method for Modelling -Response Data Under Simple Order Restrictions Martin Otava International Hexa-Symposium on Biostatistics, Bioinformatics, and Epidemiology
Permutation Test for Bayesian Variable Selection Method for Modelling -Response Data Under Simple Order Restrictions Martin Otava International Hexa-Symposium on Biostatistics, Bioinformatics, and Epidemiology
Multiple testing: Intro & FWER 1
 Multiple testing: Intro & FWER 1 Mark van de Wiel mark.vdwiel@vumc.nl Dep of Epidemiology & Biostatistics,VUmc, Amsterdam Dep of Mathematics, VU 1 Some slides courtesy of Jelle Goeman 1 Practical notes
Multiple testing: Intro & FWER 1 Mark van de Wiel mark.vdwiel@vumc.nl Dep of Epidemiology & Biostatistics,VUmc, Amsterdam Dep of Mathematics, VU 1 Some slides courtesy of Jelle Goeman 1 Practical notes
New Approaches to False Discovery Control
 New Approaches to False Discovery Control Christopher R. Genovese Department of Statistics Carnegie Mellon University http://www.stat.cmu.edu/ ~ genovese/ Larry Wasserman Department of Statistics Carnegie
New Approaches to False Discovery Control Christopher R. Genovese Department of Statistics Carnegie Mellon University http://www.stat.cmu.edu/ ~ genovese/ Larry Wasserman Department of Statistics Carnegie
Sanat Sarkar Department of Statistics, Temple University Philadelphia, PA 19122, U.S.A. September 11, Abstract
 Adaptive Controls of FWER and FDR Under Block Dependence arxiv:1611.03155v1 [stat.me] 10 Nov 2016 Wenge Guo Department of Mathematical Sciences New Jersey Institute of Technology Newark, NJ 07102, U.S.A.
Adaptive Controls of FWER and FDR Under Block Dependence arxiv:1611.03155v1 [stat.me] 10 Nov 2016 Wenge Guo Department of Mathematical Sciences New Jersey Institute of Technology Newark, NJ 07102, U.S.A.
Zhiguang Huo 1, Chi Song 2, George Tseng 3. July 30, 2018
 Bayesian latent hierarchical model for transcriptomic meta-analysis to detect biomarkers with clustered meta-patterns of differential expression signals BayesMP Zhiguang Huo 1, Chi Song 2, George Tseng
Bayesian latent hierarchical model for transcriptomic meta-analysis to detect biomarkers with clustered meta-patterns of differential expression signals BayesMP Zhiguang Huo 1, Chi Song 2, George Tseng
Semiparametric Bayes Multiple Testing: Applications to Tumor Data
 Semiparametric Bayes Multiple Testing: Applications to Tumor Data 1 Lianming Wang and David B. Dunson Biostatistics Branch, MD A3-03 National Institute of Environmental Health Sciences U.S. National Institutes
Semiparametric Bayes Multiple Testing: Applications to Tumor Data 1 Lianming Wang and David B. Dunson Biostatistics Branch, MD A3-03 National Institute of Environmental Health Sciences U.S. National Institutes
Spiked Dirichlet Process Prior for Bayesian Multiple Hypothesis Testing in Random Effects Models
 Bayesian Analysis (2009) 4, Number 4, pp. 707 732 Spiked Dirichlet Process Prior for Bayesian Multiple Hypothesis Testing in Random Effects Models Sinae Kim, David B. Dahl and Marina Vannucci Abstract.
Bayesian Analysis (2009) 4, Number 4, pp. 707 732 Spiked Dirichlet Process Prior for Bayesian Multiple Hypothesis Testing in Random Effects Models Sinae Kim, David B. Dahl and Marina Vannucci Abstract.
STATS 200: Introduction to Statistical Inference. Lecture 29: Course review
 STATS 200: Introduction to Statistical Inference Lecture 29: Course review Course review We started in Lecture 1 with a fundamental assumption: Data is a realization of a random process. The goal throughout
STATS 200: Introduction to Statistical Inference Lecture 29: Course review Course review We started in Lecture 1 with a fundamental assumption: Data is a realization of a random process. The goal throughout
discovery rate control
 Optimal design for high-throughput screening via false discovery rate control arxiv:1707.03462v1 [stat.ap] 11 Jul 2017 Tao Feng 1, Pallavi Basu 2, Wenguang Sun 3, Hsun Teresa Ku 4, Wendy J. Mack 1 Abstract
Optimal design for high-throughput screening via false discovery rate control arxiv:1707.03462v1 [stat.ap] 11 Jul 2017 Tao Feng 1, Pallavi Basu 2, Wenguang Sun 3, Hsun Teresa Ku 4, Wendy J. Mack 1 Abstract
arxiv: v1 [stat.me] 25 Aug 2016
![arxiv: v1 [stat.me] 25 Aug 2016 arxiv: v1 [stat.me] 25 Aug 2016](/thumbs/94/119227727.jpg) Empirical Null Estimation using Discrete Mixture Distributions and its Application to Protein Domain Data arxiv:1608.07204v1 [stat.me] 25 Aug 2016 Iris Ivy Gauran 1, Junyong Park 1, Johan Lim 2, DoHwan
Empirical Null Estimation using Discrete Mixture Distributions and its Application to Protein Domain Data arxiv:1608.07204v1 [stat.me] 25 Aug 2016 Iris Ivy Gauran 1, Junyong Park 1, Johan Lim 2, DoHwan
Estimation of Operational Risk Capital Charge under Parameter Uncertainty
 Estimation of Operational Risk Capital Charge under Parameter Uncertainty Pavel V. Shevchenko Principal Research Scientist, CSIRO Mathematical and Information Sciences, Sydney, Locked Bag 17, North Ryde,
Estimation of Operational Risk Capital Charge under Parameter Uncertainty Pavel V. Shevchenko Principal Research Scientist, CSIRO Mathematical and Information Sciences, Sydney, Locked Bag 17, North Ryde,
arxiv: v1 [stat.me] 13 Dec 2017
![arxiv: v1 [stat.me] 13 Dec 2017 arxiv: v1 [stat.me] 13 Dec 2017](/thumbs/78/77255504.jpg) Local False Discovery Rate Based Methods for Multiple Testing of One-Way Classified Hypotheses Sanat K. Sarkar, Zhigen Zhao Department of Statistical Science, Temple University, Philadelphia, PA, 19122,
Local False Discovery Rate Based Methods for Multiple Testing of One-Way Classified Hypotheses Sanat K. Sarkar, Zhigen Zhao Department of Statistical Science, Temple University, Philadelphia, PA, 19122,
arxiv: v1 [math.st] 31 Mar 2009
![arxiv: v1 [math.st] 31 Mar 2009 arxiv: v1 [math.st] 31 Mar 2009](/thumbs/72/66691762.jpg) The Annals of Statistics 2009, Vol. 37, No. 2, 619 629 DOI: 10.1214/07-AOS586 c Institute of Mathematical Statistics, 2009 arxiv:0903.5373v1 [math.st] 31 Mar 2009 AN ADAPTIVE STEP-DOWN PROCEDURE WITH PROVEN
The Annals of Statistics 2009, Vol. 37, No. 2, 619 629 DOI: 10.1214/07-AOS586 c Institute of Mathematical Statistics, 2009 arxiv:0903.5373v1 [math.st] 31 Mar 2009 AN ADAPTIVE STEP-DOWN PROCEDURE WITH PROVEN
Family-wise Error Rate Control in QTL Mapping and Gene Ontology Graphs
 Family-wise Error Rate Control in QTL Mapping and Gene Ontology Graphs with Remarks on Family Selection Dissertation Defense April 5, 204 Contents Dissertation Defense Introduction 2 FWER Control within
Family-wise Error Rate Control in QTL Mapping and Gene Ontology Graphs with Remarks on Family Selection Dissertation Defense April 5, 204 Contents Dissertation Defense Introduction 2 FWER Control within
A Simple, Graphical Procedure for Comparing Multiple Treatment Effects
 A Simple, Graphical Procedure for Comparing Multiple Treatment Effects Brennan S. Thompson and Matthew D. Webb May 15, 2015 > Abstract In this paper, we utilize a new graphical
A Simple, Graphical Procedure for Comparing Multiple Treatment Effects Brennan S. Thompson and Matthew D. Webb May 15, 2015 > Abstract In this paper, we utilize a new graphical
Optimal Tests Shrinking Both Means and Variances Applicable to Microarray Data Analysis
 Statistics Preprints Statistics 4-2007 Optimal Tests Shrinking Both Means and Variances Applicable to Microarray Data Analysis J.T. Gene Hwang Cornell University Peng Liu Iowa State University, pliu@iastate.edu
Statistics Preprints Statistics 4-2007 Optimal Tests Shrinking Both Means and Variances Applicable to Microarray Data Analysis J.T. Gene Hwang Cornell University Peng Liu Iowa State University, pliu@iastate.edu
STAT 461/561- Assignments, Year 2015
 STAT 461/561- Assignments, Year 2015 This is the second set of assignment problems. When you hand in any problem, include the problem itself and its number. pdf are welcome. If so, use large fonts and
STAT 461/561- Assignments, Year 2015 This is the second set of assignment problems. When you hand in any problem, include the problem itself and its number. pdf are welcome. If so, use large fonts and
Estimation of a Two-component Mixture Model
 Estimation of a Two-component Mixture Model Bodhisattva Sen 1,2 University of Cambridge, Cambridge, UK Columbia University, New York, USA Indian Statistical Institute, Kolkata, India 6 August, 2012 1 Joint
Estimation of a Two-component Mixture Model Bodhisattva Sen 1,2 University of Cambridge, Cambridge, UK Columbia University, New York, USA Indian Statistical Institute, Kolkata, India 6 August, 2012 1 Joint
arxiv: v1 [math.st] 17 Jun 2009
![arxiv: v1 [math.st] 17 Jun 2009 arxiv: v1 [math.st] 17 Jun 2009](/thumbs/72/67454030.jpg) The Annals of Statistics 2009, Vol. 37, No. 3, 1518 1544 DOI: 10.1214/08-AOS616 c Institute of Mathematical Statistics, 2009 arxiv:0906.3082v1 [math.st] 17 Jun 2009 A NEW MULTIPLE TESTING METHOD IN THE
The Annals of Statistics 2009, Vol. 37, No. 3, 1518 1544 DOI: 10.1214/08-AOS616 c Institute of Mathematical Statistics, 2009 arxiv:0906.3082v1 [math.st] 17 Jun 2009 A NEW MULTIPLE TESTING METHOD IN THE
Supplementary Materials for Molecular QTL Discovery Incorporating Genomic Annotations using Bayesian False Discovery Rate Control
 Supplementary Materials for Molecular QTL Discovery Incorporating Genomic Annotations using Bayesian False Discovery Rate Control Xiaoquan Wen Department of Biostatistics, University of Michigan A Model
Supplementary Materials for Molecular QTL Discovery Incorporating Genomic Annotations using Bayesian False Discovery Rate Control Xiaoquan Wen Department of Biostatistics, University of Michigan A Model
Bayesian linear regression
 Bayesian linear regression Linear regression is the basis of most statistical modeling. The model is Y i = X T i β + ε i, where Y i is the continuous response X i = (X i1,..., X ip ) T is the corresponding
Bayesian linear regression Linear regression is the basis of most statistical modeling. The model is Y i = X T i β + ε i, where Y i is the continuous response X i = (X i1,..., X ip ) T is the corresponding
Statistical Inference
 Statistical Inference Liu Yang Florida State University October 27, 2016 Liu Yang, Libo Wang (Florida State University) Statistical Inference October 27, 2016 1 / 27 Outline The Bayesian Lasso Trevor Park
Statistical Inference Liu Yang Florida State University October 27, 2016 Liu Yang, Libo Wang (Florida State University) Statistical Inference October 27, 2016 1 / 27 Outline The Bayesian Lasso Trevor Park
SOME STEP-DOWN PROCEDURES CONTROLLING THE FALSE DISCOVERY RATE UNDER DEPENDENCE
 Statistica Sinica 18(2008), 881-904 SOME STEP-DOWN PROCEDURES CONTROLLING THE FALSE DISCOVERY RATE UNDER DEPENDENCE Yongchao Ge 1, Stuart C. Sealfon 1 and Terence P. Speed 2,3 1 Mount Sinai School of Medicine,
Statistica Sinica 18(2008), 881-904 SOME STEP-DOWN PROCEDURES CONTROLLING THE FALSE DISCOVERY RATE UNDER DEPENDENCE Yongchao Ge 1, Stuart C. Sealfon 1 and Terence P. Speed 2,3 1 Mount Sinai School of Medicine,
Parameter estimation and forecasting. Cristiano Porciani AIfA, Uni-Bonn
 Parameter estimation and forecasting Cristiano Porciani AIfA, Uni-Bonn Questions? C. Porciani Estimation & forecasting 2 Temperature fluctuations Variance at multipole l (angle ~180o/l) C. Porciani Estimation
Parameter estimation and forecasting Cristiano Porciani AIfA, Uni-Bonn Questions? C. Porciani Estimation & forecasting 2 Temperature fluctuations Variance at multipole l (angle ~180o/l) C. Porciani Estimation
False discovery control for multiple tests of association under general dependence
 False discovery control for multiple tests of association under general dependence Nicolai Meinshausen Seminar für Statistik ETH Zürich December 2, 2004 Abstract We propose a confidence envelope for false
False discovery control for multiple tests of association under general dependence Nicolai Meinshausen Seminar für Statistik ETH Zürich December 2, 2004 Abstract We propose a confidence envelope for false
Resampling-based Multiple Testing with Applications to Microarray Data Analysis
 Resampling-based Multiple Testing with Applications to Microarray Data Analysis DISSERTATION Presented in Partial Fulfillment of the Requirements for the Degree Doctor of Philosophy in the Graduate School
Resampling-based Multiple Testing with Applications to Microarray Data Analysis DISSERTATION Presented in Partial Fulfillment of the Requirements for the Degree Doctor of Philosophy in the Graduate School
Adaptive Filtering Multiple Testing Procedures for Partial Conjunction Hypotheses
 Adaptive Filtering Multiple Testing Procedures for Partial Conjunction Hypotheses arxiv:1610.03330v1 [stat.me] 11 Oct 2016 Jingshu Wang, Chiara Sabatti, Art B. Owen Department of Statistics, Stanford University
Adaptive Filtering Multiple Testing Procedures for Partial Conjunction Hypotheses arxiv:1610.03330v1 [stat.me] 11 Oct 2016 Jingshu Wang, Chiara Sabatti, Art B. Owen Department of Statistics, Stanford University
Statistical analysis of microarray data: a Bayesian approach
 Biostatistics (003), 4, 4,pp. 597 60 Printed in Great Britain Statistical analysis of microarray data: a Bayesian approach RAPHAEL GTTARD University of Washington, Department of Statistics, Box 3543, Seattle,
Biostatistics (003), 4, 4,pp. 597 60 Printed in Great Britain Statistical analysis of microarray data: a Bayesian approach RAPHAEL GTTARD University of Washington, Department of Statistics, Box 3543, Seattle,
SIGNAL RANKING-BASED COMPARISON OF AUTOMATIC DETECTION METHODS IN PHARMACOVIGILANCE
 SIGNAL RANKING-BASED COMPARISON OF AUTOMATIC DETECTION METHODS IN PHARMACOVIGILANCE A HYPOTHESIS TEST APPROACH Ismaïl Ahmed 1,2, Françoise Haramburu 3,4, Annie Fourrier-Réglat 3,4,5, Frantz Thiessard 4,5,6,
SIGNAL RANKING-BASED COMPARISON OF AUTOMATIC DETECTION METHODS IN PHARMACOVIGILANCE A HYPOTHESIS TEST APPROACH Ismaïl Ahmed 1,2, Françoise Haramburu 3,4, Annie Fourrier-Réglat 3,4,5, Frantz Thiessard 4,5,6,
Single gene analysis of differential expression
 Single gene analysis of differential expression Giorgio Valentini DSI Dipartimento di Scienze dell Informazione Università degli Studi di Milano valentini@dsi.unimi.it Comparing two conditions Each condition
Single gene analysis of differential expression Giorgio Valentini DSI Dipartimento di Scienze dell Informazione Università degli Studi di Milano valentini@dsi.unimi.it Comparing two conditions Each condition
Weighted Adaptive Multiple Decision Functions for False Discovery Rate Control
 Weighted Adaptive Multiple Decision Functions for False Discovery Rate Control Joshua D. Habiger Oklahoma State University jhabige@okstate.edu Nov. 8, 2013 Outline 1 : Motivation and FDR Research Areas
Weighted Adaptive Multiple Decision Functions for False Discovery Rate Control Joshua D. Habiger Oklahoma State University jhabige@okstate.edu Nov. 8, 2013 Outline 1 : Motivation and FDR Research Areas
On adaptive procedures controlling the familywise error rate
 , pp. 3 On adaptive procedures controlling the familywise error rate By SANAT K. SARKAR Temple University, Philadelphia, PA 922, USA sanat@temple.edu Summary This paper considers the problem of developing
, pp. 3 On adaptive procedures controlling the familywise error rate By SANAT K. SARKAR Temple University, Philadelphia, PA 922, USA sanat@temple.edu Summary This paper considers the problem of developing
Lecture 8: Information Theory and Statistics
 Lecture 8: Information Theory and Statistics Part II: Hypothesis Testing and I-Hsiang Wang Department of Electrical Engineering National Taiwan University ihwang@ntu.edu.tw December 23, 2015 1 / 50 I-Hsiang
Lecture 8: Information Theory and Statistics Part II: Hypothesis Testing and I-Hsiang Wang Department of Electrical Engineering National Taiwan University ihwang@ntu.edu.tw December 23, 2015 1 / 50 I-Hsiang
Simultaneous Testing of Grouped Hypotheses: Finding Needles in Multiple Haystacks
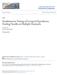 University of Pennsylvania ScholarlyCommons Statistics Papers Wharton Faculty Research 2009 Simultaneous Testing of Grouped Hypotheses: Finding Needles in Multiple Haystacks T. Tony Cai University of Pennsylvania
University of Pennsylvania ScholarlyCommons Statistics Papers Wharton Faculty Research 2009 Simultaneous Testing of Grouped Hypotheses: Finding Needles in Multiple Haystacks T. Tony Cai University of Pennsylvania
Semi-Penalized Inference with Direct FDR Control
 Jian Huang University of Iowa April 4, 2016 The problem Consider the linear regression model y = p x jβ j + ε, (1) j=1 where y IR n, x j IR n, ε IR n, and β j is the jth regression coefficient, Here p
Jian Huang University of Iowa April 4, 2016 The problem Consider the linear regression model y = p x jβ j + ε, (1) j=1 where y IR n, x j IR n, ε IR n, and β j is the jth regression coefficient, Here p
arxiv:math/ v1 [math.st] 29 Dec 2006 Jianqing Fan Peter Hall Qiwei Yao
![arxiv:math/ v1 [math.st] 29 Dec 2006 Jianqing Fan Peter Hall Qiwei Yao arxiv:math/ v1 [math.st] 29 Dec 2006 Jianqing Fan Peter Hall Qiwei Yao](/thumbs/74/69806335.jpg) TO HOW MANY SIMULTANEOUS HYPOTHESIS TESTS CAN NORMAL, STUDENT S t OR BOOTSTRAP CALIBRATION BE APPLIED? arxiv:math/0701003v1 [math.st] 29 Dec 2006 Jianqing Fan Peter Hall Qiwei Yao ABSTRACT. In the analysis
TO HOW MANY SIMULTANEOUS HYPOTHESIS TESTS CAN NORMAL, STUDENT S t OR BOOTSTRAP CALIBRATION BE APPLIED? arxiv:math/0701003v1 [math.st] 29 Dec 2006 Jianqing Fan Peter Hall Qiwei Yao ABSTRACT. In the analysis
