Conformational Searching using MacroModel and ConfGen. John Shelley Schrödinger Fellow
|
|
|
- Nancy Deirdre Cunningham
- 6 years ago
- Views:
Transcription
1 Conformational Searching using MacroModel and ConfGen John Shelley Schrödinger Fellow
2 Overview Types of conformational searching applications MacroModel s conformation generation procedure General features supporting conformational searches Conformation generation methods Methods which accumulate changes Methods which exhaustively sample local minima General guidelines Closing points
3 Types of conformational searching applications Types of systems Ligands Proteins and protein-ligand complexes Other Characteristics Size of molecules Scale of conformational changes expected Thoroughness Collectivity of conformational changes Time/disk space usage
4 MacroModel s conformation generation procedure...for Each molecule Read the input structure 2. Characterize the molecule Stereochemistry: chiralities and double bond geometries (E/Z) Molecule/search method specific features* Symmetry Comparison atoms Rotatable bonds Ring systems (and ring opening bonds) Restraints Number of steps Etc. 3. Optionally energy minimize the input structure 4. Generate conformers one at a time Generate a new conformation for the molecule* Optionally post process the conformation Estimate energy or minimize Filter by energy Eliminate redundant conformers 4. Save results * These vary with the search method employed
5 General features supporting conformational searches 1. Force fields (all atom) OPLS_2005, OPLS_2001 MMFF, MMFFs 2. Solvent models Vacuum (or constant dielectric) Distance dependent dielectric (1/4r) GB/SA 3. para_bmin For distributing the separate searches of different molecules across multiple processors Fault tolerant 4. dbmin For distributing the search of a single structure across multiple processors Useful for protein-ligand complexes.
6 Conformation generation methods that accumulate changes
7 Methods which accumulate changes A collection of conformations is built up during the search. Conformations generated earlier in the search are used as starting points for subsequent search steps. Methods: 1. MCMM, Monte Carlo Multiple Minimum Random torsional search plus minimization 2. Low MODe (LMOD) and Large-scale Low Mode (LLMOD) Use low frequency modes to construct conformational changes 3. Mixed Mode A combination of MCMM with one of LMOD or LLMOD
8 MCMM (Monte Carlo Multiple Minimum) Each MCMM step consists of: 1. Select an existing conformer 2. Introduce changes by randomly: - adjusting the dihedral angles for rotatable bonds - Translating and rotating molecules (if there are two or more molecules) Automatic Setup 3. Sampling rings: - Pretend to break a bond in the ring (the ring opening bond) - Treat the remaining single bonds as rotatable. - If the atoms in the broken bond end up lying close to each other reform the bond. Otherwise, try another set of bond rotations 4. Post process the structure: - Minimization - Stereochemical checks - Energy window - Redundancy check - If OK, add structure to collection of conformers
9 MCMM: Strengths and Weaknesses Strengths 1. General method which can be applied to both small and large structures 2. Very effective if rotations about a small number of bonds will yield a large unhindered conformational change Weaknesses 1. Need to limit the number of simultaneous rotations 2. Large conformational changes may need a series of localized changes. Searches can be long and CPU-intensive. 3. Not so good for environments in which explicit atoms fill up most of the space because changes need to be collective.
10 Low Mode and Large scale Low Mode For each LMOD or LLMOD step: 1. Select an existing conformer 2. Generate normal modes 3. Select a normal mode - Amplify the normal mode so as to generate a very different structure 4. Post process the structure - Minimization - Stereochemical checks - Energy window - Redundancy check - If OK, add structure to collection of conformers
11 LMOD and LLMOD: Strengths and Weaknesses Strengths 1. General method which can be applied to both small and large structures 2. Useful for environments in which explicit atoms fill up most of the space causing changes to be collective Weaknesses 1. Large conformational changes may need a series of localized changes (more so than MCMM). Searches can be long and CPU-intensive. 2. Individual search steps use more computer time than MCMM. 3. Sometimes this is not effective at sampling translations and rotations of molecules in complexes.
12 Mixed mode searches Idea Use the strengths of both methods by performing a search in which some steps are LMOD or LLMOD steps and others are MCMM steps. Implementation Just like either of these search methods, but at each step randomly choose which type of step to use Can set the probability MCMM vs LMOD or LLMOD (usually 50/50 works fine) Mixed mode searches are recommended over searches employing one search technique. Exploration: Can we also usefully employ fairly new methods like FLAP (ring corner reflection) and MCRC (ring sampling using pre-generated conformations) in mixed mode searches?
13 GUI for methods which accumulate changes Accessing Maestro Applications (menu) MacroModel Conformational Search CSearch (tab) Recommendation: For ligands, the default settings are appropriate.
14 Automatic setup What is automatic setup doing? To find out: 1. Place an interesting molecule in the workspace 2. Turn off the Multi-ligand toggle 3. Turn off Perform automatic setup during calculation 4. Hit Perform Automatic Setup Before After How can I modify the setup? Most features can be changed using the Search variables pull-down menu and then hitting the Edit button.
15 Hints: mixed mode searches of protein-ligand complexes 1. Be selective about what bonds to rotate using MCMM (see previous slide) - Typically only a few such rotations can be changed at one simultaneously. - Focus those changes on key rotatable bonds that look like they can have a significant chance of resulting in larger changes. 2. Use substructures (see next slide)
16 Defining a substructure region for conformational sampling Substructure flexible atoms Shell 1 constrained atoms Shell 2 frozen atoms Substructure Options: - Write.sbc file for a future use - ASL formatted.sbc file
17 Conformation generation methods that exhaustively sample local minima
18 Methods which exhaustively sample local minima - ConfGen Overall goal: systematically explore all or nearly all conformations Key features: 1. Identify low energy ring conformations I. Flexible ring systems are identified II. - Mutually connected flexible rings and aromatic rings that share at least two atoms with one of the flexible rings in the ring system A corresponding template is sought from a large template library i. Template matching is strict; element and hybridization for all atoms in the template must match the target ring system. ii. Low energy conformers for each template ring system have already been generated and stored in the template library 2. Identify invertible nitrogen atoms - Non-ring pyramidal sp3 nitrogen atoms that are bonded to only three other atoms (i.e., unprotonated sp3 nitrogen not very common) 3. Identify rotatable bonds - Single bonds whose rotations result in non-trivial changes in the molecules
19 ConfGen: Concept of Molecular core O H N S rotamer group N rotamer group O- O
20 ConfGen: Core geometry sampling For all combinations of low energy ring system conformations and nitrogen atom inversions: 1. Identify minima for rotating about each rotatable bond I. Generate tabulated potentials for each rotatable bonds i. Use OPLS_2001 dihedral ii. iii. Lennard-Jones non-bonded interactions for a subset of the atoms on either side of the bond that are deemed key to avoiding topologically local vdw clashes Enforce rotational symmetry Option: If the range of the force field based potential is small replace it with a cosine potential (typically 6 fold) II. Identify minima in the tabulated potentials i. Limit to at most 10 minima per rotatable bond ii. Eliminate equivalent minima for bonds with rotationally symmetry. iii. Determine the relative energies of the minima 2. Generate conformations for all combinations of minima for rotatable bonds within the core I. Eliminate high energy conformations on the fly based upon relative ring energies and the relative energies of the minima for the rotatable bonds. II. III. Minimize the energy by adjusting the dihedral angles for the rotatable bonds. Score core geometries and prune by score if excessive core conformations are produced.
21 ConfGen: Rotamer geometry sampling For each core geometry sample rotamers of the peripheral groups: 1. Use one of two user selectable approaches: Sample all combinations of peripheral group rotamers Excite each peripheral group in turn while keeping the other groups in their lowest energy minima. 2. Eliminate high energy conformations (as for core conformation sampling)
22 ConfGen: Conformation filtering Goal: Eliminate undesirable conformations Filter out conformations with: 1. vdw overlap I. Conformations in which heavy atoms approach each other more closely than 60% of the sum of their vdw radii are eliminated. II. III. If the previous step eliminates all of the conformations then reduce the closest approach distance by 15% and try again. If the previous step eliminates all of the conformations then reduce the closest approach distance by another 15% and try again (this is rarely needed). 2. Close approach of atoms with formal charges of the same sign 3. A hydrogen bonds pointing towards nearby atoms that have a net positive formal charge 4. high local concentrations of heavy atoms (e.g. stacked rings)
23 ConfGen: Selecting a subset of conformers Limit the number of conformers to at most: N target = (N mult )(Effective number of degrees of freedom) N mult is a parameter Uniformly selecting a subset of all of the conformers Conformers ranked by score 1. Select the first conformer 2. Select the last conformer 3. Select conformers ½ way between those already selected in the ordered list of conformers 4. Repeat 3 until sufficient conformers have been selected
24 ConfGen: conformation generation Sample Rotatable Bonds Sample Ring Conformations Resulting Conformations
25 ConfGen and MacroModel ConfGen functionality was originally developed for Glide and has undergone further development for use with other products. ConfGen searches can be performed using Phase or MacroModel. ConfGen/MacroModel calculations: are supported by ConfGen Panels in Maestro permit MacroModel s pre- and post-processing functionalities to work synergistically with ConfGen s. Namely: the use of a range of force fields solvation models minimization filtering (energy, redundancy) Net result is a reduction in the number of conformers along with an improvement in the quality of the conformations.
26 ConfGen/MacroModel search optimization study Ligands Used: All studies were conducted on a set of 667 ligands, including 36 from Boström (Comp. Aided Mol. Design 2001, 15, ), the 100 public structures from Perola and Charifson (J. Med. Chem. 2004, 47, ), and an additional 538 selected from the Protein Data Bank Defined 4 protocols Very fast - no energy evaluations (or minimizations) in MacroModel - RMS for redundant conformer elimination: N mult = 5 Fast - Eliminate conformers with high all atom relative energies > 12 kj/mol - RMS for redundant conformer elimination: N mult = 5 Intermediate - Eliminate conformers with high all atom relative energies > kj/mol - RMS for redundant conformer elimination: N mult = 75 Comprehensive - Eliminate conformers with high all atom relative energies > 500 kj/mol - RMS for redundant conformer elimination: N mult = 75
27 ConfGen/MacroModel search optimization study Systematic Exploration of how to use ConfGen and MacroModel together Method Root Mean Square distance in Å* # of Time** <0.5 <1.0 < 1.5 <2.0 Confs Very Fast 16% 52% 84% 96% Fast 19% 55% 82% 95% Intermediate 21% 64% 85% 97% Comprehensive 34% 70% 90% 98% * Percent of ligands with a generated conformer with an RMS relative to the bioactive conformer less than value listed **Time in seconds on a AMD Opteron 1.6 GHz processors with 2 GB memory.
28 ConfGen/MacroModel search optimization study Comprehensive has the best results at all levels of accuracy. At the standard level used in the literature for matching bioactive conformations (<2.0 Å ) the coverage is nearly complete for all 4 protocols. Searches are fast enough for use in generating databases of conformers Very fast, Fast, and Intermediate produce fewer conformers than is typically used/required (approximately ) for this level of coverage at (<2.0 Å). - Lower disk space usage for large databases - Computations using these conformers should use less CPU time. - Fewer irrelevant and perhaps high energy conformers that may generate spurious results in subsequent processing (e.g. pharmacophore generation) These are done without minimization. Minimization generally increases CPU time by a factor of 4-12, while increasing the matches by 1-5 percent. Versatile with a broad range of speed/quality trade offs.
29 ConfGen GUI Accessing: Maestro Applications (menu) ConfGen Standard or Advanced *Recommended *
30 ConfGen: Strengths and Weaknesses Strengths 1. Fast enough to be used for large databases of ligands 2. Exhaustive sampling of local minima ensures comprehensive coverage of conformer space 3. Small subsets of conformers provide good coverage of conformer space Weaknesses 1. Limited to ligand-sized molecules 2. Even exhaustive sampling of local minima may miss some minima that arise from non-local interactions.
31 General Guidelines For Ligands Fairly accurate yet quite fast (seconds/ligand) ConfGen without minimization e.g. conformer database generation Quite accurate yet fairly fast (1 minute/ligand) ConfGen with minimization Very accurate yet slow (1 hr/ ligand) MCMM/LMOD mixed searches e.g. generating conformers for using in creating pharmacophores Protein/Ligand complexes Use substructures Use mixed mode MCMM/LLMOD or MCMM/LMOD (< 250 moving atoms) searches Consider selecting MCMM torsional moves carefully Other systems Approach usually needs to be tailored to the nature of the problem For large system mixed mode MCMM/LLMOD is usually the appropriate choice Use larger ranges for LLMOD moves Careful selection of MCMM torsional moves Use larger energy window (reminimize output later) Increase RMS values somewhat
32 Closing points Together ConfGen and MacroModel can be used to search a wide range of molecules from ligands to protein/ligand complexes and other types of molecules. Maestro makes setting up and running these calculations easy. Wide range of speed/quality trade offs possible. Small subsets of conformers generated by ConfGen can cover conformational space quite well. Searching multiple structures in a single calculation automatically is supported. Distributing calculations is supported.
33 Acknowledgments MacroModel team dozens of people over the years ConfGen Development and Application studies Pranav Dalal Rich Friesner Robert Murphy Woody Sherman Shawn Watts
Schrodinger ebootcamp #3, Summer EXPLORING METHODS FOR CONFORMER SEARCHING Jas Bhachoo, Senior Applications Scientist
 Schrodinger ebootcamp #3, Summer 2016 EXPLORING METHODS FOR CONFORMER SEARCHING Jas Bhachoo, Senior Applications Scientist Numerous applications Generating conformations MM Agenda http://www.schrodinger.com/macromodel
Schrodinger ebootcamp #3, Summer 2016 EXPLORING METHODS FOR CONFORMER SEARCHING Jas Bhachoo, Senior Applications Scientist Numerous applications Generating conformations MM Agenda http://www.schrodinger.com/macromodel
Kd = koff/kon = [R][L]/[RL]
![Kd = koff/kon = [R][L]/[RL] Kd = koff/kon = [R][L]/[RL]](/thumbs/96/127564193.jpg) Taller de docking y cribado virtual: Uso de herramientas computacionales en el diseño de fármacos Docking program GLIDE El programa de docking GLIDE Sonsoles Martín-Santamaría Shrödinger is a scientific
Taller de docking y cribado virtual: Uso de herramientas computacionales en el diseño de fármacos Docking program GLIDE El programa de docking GLIDE Sonsoles Martín-Santamaría Shrödinger is a scientific
Analyzing Molecular Conformations Using the Cambridge Structural Database. Jason Cole Cambridge Crystallographic Data Centre
 Analyzing Molecular Conformations Using the Cambridge Structural Database Jason Cole Cambridge Crystallographic Data Centre 1 The Cambridge Structural Database (CSD) 905,284* USOPEZ a natural product intermediate,
Analyzing Molecular Conformations Using the Cambridge Structural Database Jason Cole Cambridge Crystallographic Data Centre 1 The Cambridge Structural Database (CSD) 905,284* USOPEZ a natural product intermediate,
The PhilOEsophy. There are only two fundamental molecular descriptors
 The PhilOEsophy There are only two fundamental molecular descriptors Where can we use shape? Virtual screening More effective than 2D Lead-hopping Shape analogues are not graph analogues Molecular alignment
The PhilOEsophy There are only two fundamental molecular descriptors Where can we use shape? Virtual screening More effective than 2D Lead-hopping Shape analogues are not graph analogues Molecular alignment
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/309/5742/1868/dc1 Supporting Online Material for Toward High-Resolution de Novo Structure Prediction for Small Proteins Philip Bradley, Kira M. S. Misura, David Baker*
www.sciencemag.org/cgi/content/full/309/5742/1868/dc1 Supporting Online Material for Toward High-Resolution de Novo Structure Prediction for Small Proteins Philip Bradley, Kira M. S. Misura, David Baker*
ICM-Chemist-Pro How-To Guide. Version 3.6-1h Last Updated 12/29/2009
 ICM-Chemist-Pro How-To Guide Version 3.6-1h Last Updated 12/29/2009 ICM-Chemist-Pro ICM 3D LIGAND EDITOR: SETUP 1. Read in a ligand molecule or PDB file. How to setup the ligand in the ICM 3D Ligand Editor.
ICM-Chemist-Pro How-To Guide Version 3.6-1h Last Updated 12/29/2009 ICM-Chemist-Pro ICM 3D LIGAND EDITOR: SETUP 1. Read in a ligand molecule or PDB file. How to setup the ligand in the ICM 3D Ligand Editor.
Creating a Pharmacophore Query from a Reference Molecule & Scaffold Hopping in CSD-CrossMiner
 Table of Contents Creating a Pharmacophore Query from a Reference Molecule & Scaffold Hopping in CSD-CrossMiner Introduction... 2 CSD-CrossMiner Terminology... 2 Overview of CSD-CrossMiner... 3 Features
Table of Contents Creating a Pharmacophore Query from a Reference Molecule & Scaffold Hopping in CSD-CrossMiner Introduction... 2 CSD-CrossMiner Terminology... 2 Overview of CSD-CrossMiner... 3 Features
Using Phase for Pharmacophore Modelling. 5th European Life Science Bootcamp March, 2017
 Using Phase for Pharmacophore Modelling 5th European Life Science Bootcamp March, 2017 Phase: Our Pharmacohore generation tool Significant improvements to Phase methods in 2016 New highly interactive interface
Using Phase for Pharmacophore Modelling 5th European Life Science Bootcamp March, 2017 Phase: Our Pharmacohore generation tool Significant improvements to Phase methods in 2016 New highly interactive interface
Molecular Modeling and Conformational Analysis with PC Spartan
 Molecular Modeling and Conformational Analysis with PC Spartan Introduction Molecular modeling can be done in a variety of ways, from using simple hand-held models to doing sophisticated calculations on
Molecular Modeling and Conformational Analysis with PC Spartan Introduction Molecular modeling can be done in a variety of ways, from using simple hand-held models to doing sophisticated calculations on
5.1. Hardwares, Softwares and Web server used in Molecular modeling
 5. EXPERIMENTAL The tools, techniques and procedures/methods used for carrying out research work reported in this thesis have been described as follows: 5.1. Hardwares, Softwares and Web server used in
5. EXPERIMENTAL The tools, techniques and procedures/methods used for carrying out research work reported in this thesis have been described as follows: 5.1. Hardwares, Softwares and Web server used in
The Schrödinger KNIME extensions
 The Schrödinger KNIME extensions Computational Chemistry and Cheminformatics in a workflow environment Jean-Christophe Mozziconacci Volker Eyrich Topics What are the Schrödinger extensions? Workflow application
The Schrödinger KNIME extensions Computational Chemistry and Cheminformatics in a workflow environment Jean-Christophe Mozziconacci Volker Eyrich Topics What are the Schrödinger extensions? Workflow application
User Guide for LeDock
 User Guide for LeDock Hongtao Zhao, PhD Email: htzhao@lephar.com Website: www.lephar.com Copyright 2017 Hongtao Zhao. All rights reserved. Introduction LeDock is flexible small-molecule docking software,
User Guide for LeDock Hongtao Zhao, PhD Email: htzhao@lephar.com Website: www.lephar.com Copyright 2017 Hongtao Zhao. All rights reserved. Introduction LeDock is flexible small-molecule docking software,
Performing a Pharmacophore Search using CSD-CrossMiner
 Table of Contents Introduction... 2 CSD-CrossMiner Terminology... 2 Overview of CSD-CrossMiner... 3 Searching with a Pharmacophore... 4 Performing a Pharmacophore Search using CSD-CrossMiner Version 2.0
Table of Contents Introduction... 2 CSD-CrossMiner Terminology... 2 Overview of CSD-CrossMiner... 3 Searching with a Pharmacophore... 4 Performing a Pharmacophore Search using CSD-CrossMiner Version 2.0
NMRPredict Functional Block Diagram
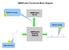 NMRPredict Functional Block Diagram 2 Submit to server. NMRPredict Server NMRPredict DeskTop Software Review results. 3 1 Input structure. Starting the NMRPredict Interface Click on the NMRPredict icon.
NMRPredict Functional Block Diagram 2 Submit to server. NMRPredict Server NMRPredict DeskTop Software Review results. 3 1 Input structure. Starting the NMRPredict Interface Click on the NMRPredict icon.
How to Create a Substance Answer Set
 How to Create a Substance Answer Set Select among five search techniques to find substances Since substances can be described by multiple names or other characteristics, SciFinder gives you the flexibility
How to Create a Substance Answer Set Select among five search techniques to find substances Since substances can be described by multiple names or other characteristics, SciFinder gives you the flexibility
Docking. GBCB 5874: Problem Solving in GBCB
 Docking Benzamidine Docking to Trypsin Relationship to Drug Design Ligand-based design QSAR Pharmacophore modeling Can be done without 3-D structure of protein Receptor/Structure-based design Molecular
Docking Benzamidine Docking to Trypsin Relationship to Drug Design Ligand-based design QSAR Pharmacophore modeling Can be done without 3-D structure of protein Receptor/Structure-based design Molecular
Ligand Scout Tutorials
 Ligand Scout Tutorials Step : Creating a pharmacophore from a protein-ligand complex. Type ke6 in the upper right area of the screen and press the button Download *+. The protein will be downloaded and
Ligand Scout Tutorials Step : Creating a pharmacophore from a protein-ligand complex. Type ke6 in the upper right area of the screen and press the button Download *+. The protein will be downloaded and
Assignment 2: Conformation Searching (50 points)
 Chemistry 380.37 Fall 2015 Dr. Jean M. Standard September 16, 2015 Assignment 2: Conformation Searching (50 points) In this assignment, you will use the Spartan software package to investigate some conformation
Chemistry 380.37 Fall 2015 Dr. Jean M. Standard September 16, 2015 Assignment 2: Conformation Searching (50 points) In this assignment, you will use the Spartan software package to investigate some conformation
Generating Small Molecule Conformations from Structural Data
 Generating Small Molecule Conformations from Structural Data Jason Cole cole@ccdc.cam.ac.uk Cambridge Crystallographic Data Centre 1 The Cambridge Crystallographic Data Centre About us A not-for-profit,
Generating Small Molecule Conformations from Structural Data Jason Cole cole@ccdc.cam.ac.uk Cambridge Crystallographic Data Centre 1 The Cambridge Crystallographic Data Centre About us A not-for-profit,
MM-GBSA for Calculating Binding Affinity A rank-ordering study for the lead optimization of Fxa and COX-2 inhibitors
 MM-GBSA for Calculating Binding Affinity A rank-ordering study for the lead optimization of Fxa and COX-2 inhibitors Thomas Steinbrecher Senior Application Scientist Typical Docking Workflow Databases
MM-GBSA for Calculating Binding Affinity A rank-ordering study for the lead optimization of Fxa and COX-2 inhibitors Thomas Steinbrecher Senior Application Scientist Typical Docking Workflow Databases
The Conformation Search Problem
 Jon Sutter Senior Manager Life Sciences R&D jms@accelrys.com Jiabo Li Senior Scientist Life Sciences R&D jli@accelrys.com CAESAR: Conformer Algorithm based on Energy Screening and Recursive Buildup The
Jon Sutter Senior Manager Life Sciences R&D jms@accelrys.com Jiabo Li Senior Scientist Life Sciences R&D jli@accelrys.com CAESAR: Conformer Algorithm based on Energy Screening and Recursive Buildup The
Evaluating the Use of Spartan in Studying the Effects of Charged Lysine Residues
 Evaluating the Use of Spartan in Studying the Effects of Charged Lysine Residues Mark Lazari Abstract Spartan by Wavefunction, Inc. is a powerful computational modeling tool that is used in both the research
Evaluating the Use of Spartan in Studying the Effects of Charged Lysine Residues Mark Lazari Abstract Spartan by Wavefunction, Inc. is a powerful computational modeling tool that is used in both the research
Exercises for Windows
 Exercises for Windows CAChe User Interface for Windows Select tool Application window Document window (workspace) Style bar Tool palette Select entire molecule Select Similar Group Select Atom tool Rotate
Exercises for Windows CAChe User Interface for Windows Select tool Application window Document window (workspace) Style bar Tool palette Select entire molecule Select Similar Group Select Atom tool Rotate
Introduction to Structure Preparation and Visualization
 Introduction to Structure Preparation and Visualization Created with: Release 2018-4 Prerequisites: Release 2018-2 or higher Access to the internet Categories: Molecular Visualization, Structure-Based
Introduction to Structure Preparation and Visualization Created with: Release 2018-4 Prerequisites: Release 2018-2 or higher Access to the internet Categories: Molecular Visualization, Structure-Based
Garib N Murshudov MRC-LMB, Cambridge
 Garib N Murshudov MRC-LMB, Cambridge Contents Introduction AceDRG: two functions Validation of entries in the DB and derived data Generation of new ligand description Jligand for link description Conclusions
Garib N Murshudov MRC-LMB, Cambridge Contents Introduction AceDRG: two functions Validation of entries in the DB and derived data Generation of new ligand description Jligand for link description Conclusions
Assignment 1: Molecular Mechanics (PART 1 25 points)
 Chemistry 380.37 Fall 2015 Dr. Jean M. Standard August 19, 2015 Assignment 1: Molecular Mechanics (PART 1 25 points) In this assignment, you will perform some molecular mechanics calculations using the
Chemistry 380.37 Fall 2015 Dr. Jean M. Standard August 19, 2015 Assignment 1: Molecular Mechanics (PART 1 25 points) In this assignment, you will perform some molecular mechanics calculations using the
Exercise 2: Solvating the Structure Before you continue, follow these steps: Setting up Periodic Boundary Conditions
 Exercise 2: Solvating the Structure HyperChem lets you place a molecular system in a periodic box of water molecules to simulate behavior in aqueous solution, as in a biological system. In this exercise,
Exercise 2: Solvating the Structure HyperChem lets you place a molecular system in a periodic box of water molecules to simulate behavior in aqueous solution, as in a biological system. In this exercise,
Physical Chemistry Final Take Home Fall 2003
 Physical Chemistry Final Take Home Fall 2003 Do one of the following questions. These projects are worth 30 points (i.e. equivalent to about two problems on the final). Each of the computational problems
Physical Chemistry Final Take Home Fall 2003 Do one of the following questions. These projects are worth 30 points (i.e. equivalent to about two problems on the final). Each of the computational problems
Conformational Sampling of Druglike Molecules with MOE and Catalyst: Implications for Pharmacophore Modeling and Virtual Screening
 J. Chem. Inf. Model. 2008, 48, 1773 1791 1773 Conformational Sampling of Druglike Molecules with MOE and Catalyst: Implications for Pharmacophore Modeling and Virtual Screening I-Jen Chen* and Nicolas
J. Chem. Inf. Model. 2008, 48, 1773 1791 1773 Conformational Sampling of Druglike Molecules with MOE and Catalyst: Implications for Pharmacophore Modeling and Virtual Screening I-Jen Chen* and Nicolas
Flexibility and Constraints in GOLD
 Flexibility and Constraints in GOLD Version 2.1 August 2018 GOLD v5.6.3 Table of Contents Purpose of Docking... 3 GOLD s Evolutionary Algorithm... 4 GOLD and Hermes... 4 Handling Flexibility and Constraints
Flexibility and Constraints in GOLD Version 2.1 August 2018 GOLD v5.6.3 Table of Contents Purpose of Docking... 3 GOLD s Evolutionary Algorithm... 4 GOLD and Hermes... 4 Handling Flexibility and Constraints
Ultra High Throughput Screening using THINK on the Internet
 Ultra High Throughput Screening using THINK on the Internet Keith Davies Central Chemistry Laboratory, Oxford University Cathy Davies Treweren Consultants, UK Blue Sky Objectives Reduce Development Failures
Ultra High Throughput Screening using THINK on the Internet Keith Davies Central Chemistry Laboratory, Oxford University Cathy Davies Treweren Consultants, UK Blue Sky Objectives Reduce Development Failures
Dictionary of ligands
 Dictionary of ligands Some of the web and other resources Small molecules DrugBank: http://www.drugbank.ca/ ZINC: http://zinc.docking.org/index.shtml PRODRUG: http://www.compbio.dundee.ac.uk/web_servers/prodrg_down.html
Dictionary of ligands Some of the web and other resources Small molecules DrugBank: http://www.drugbank.ca/ ZINC: http://zinc.docking.org/index.shtml PRODRUG: http://www.compbio.dundee.ac.uk/web_servers/prodrg_down.html
ICM-Chemist How-To Guide. Version 3.6-1g Last Updated 12/01/2009
 ICM-Chemist How-To Guide Version 3.6-1g Last Updated 12/01/2009 ICM-Chemist HOW TO IMPORT, SKETCH AND EDIT CHEMICALS How to access the ICM Molecular Editor. 1. Click here 2. Start sketching How to sketch
ICM-Chemist How-To Guide Version 3.6-1g Last Updated 12/01/2009 ICM-Chemist HOW TO IMPORT, SKETCH AND EDIT CHEMICALS How to access the ICM Molecular Editor. 1. Click here 2. Start sketching How to sketch
Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015,
 Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015, Course,Informa5on, BIOC%530% GraduateAlevel,discussion,of,the,structure,,func5on,,and,chemistry,of,proteins,and, nucleic,acids,,control,of,enzyma5c,reac5ons.,please,see,the,course,syllabus,and,
Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015, Course,Informa5on, BIOC%530% GraduateAlevel,discussion,of,the,structure,,func5on,,and,chemistry,of,proteins,and, nucleic,acids,,control,of,enzyma5c,reac5ons.,please,see,the,course,syllabus,and,
Application Note. U. Heat of Formation of Ethyl Alcohol and Dimethyl Ether. Introduction
 Application Note U. Introduction The molecular builder (Molecular Builder) is part of the MEDEA standard suite of building tools. This tutorial provides an overview of the Molecular Builder s basic functionality.
Application Note U. Introduction The molecular builder (Molecular Builder) is part of the MEDEA standard suite of building tools. This tutorial provides an overview of the Molecular Builder s basic functionality.
NMR Predictor. Introduction
 NMR Predictor This manual gives a walk-through on how to use the NMR Predictor: Introduction NMR Predictor QuickHelp NMR Predictor Overview Chemical features GUI features Usage Menu system File menu Edit
NMR Predictor This manual gives a walk-through on how to use the NMR Predictor: Introduction NMR Predictor QuickHelp NMR Predictor Overview Chemical features GUI features Usage Menu system File menu Edit
SeeSAR 7.1 Beginners Guide. June 2017
 SeeSAR 7.1 Beginners Guide June 2017 Part 1: Basics 1 Type a pdb code and press return or Load your own protein or already existing project, or Just load molecules To begin, let s type 2zff and download
SeeSAR 7.1 Beginners Guide June 2017 Part 1: Basics 1 Type a pdb code and press return or Load your own protein or already existing project, or Just load molecules To begin, let s type 2zff and download
Pipelining Ligands in PHENIX: elbow and REEL
 Pipelining Ligands in PHENIX: elbow and REEL Nigel W. Moriarty Lawrence Berkeley National Laboratory Physical Biosciences Division Ligands in Crystallography Drug design Biological function studies Generate
Pipelining Ligands in PHENIX: elbow and REEL Nigel W. Moriarty Lawrence Berkeley National Laboratory Physical Biosciences Division Ligands in Crystallography Drug design Biological function studies Generate
Molecular Visualization. Introduction
 Molecular Visualization Jeffry D. Madura Department of Chemistry & Biochemistry Center for Computational Sciences Duquesne University Introduction Assessments of change, dynamics, and cause and effect
Molecular Visualization Jeffry D. Madura Department of Chemistry & Biochemistry Center for Computational Sciences Duquesne University Introduction Assessments of change, dynamics, and cause and effect
Ranking of HIV-protease inhibitors using AutoDock
 Ranking of HIV-protease inhibitors using AutoDock 1. Task Calculate possible binding modes and estimate the binding free energies for 1 3 inhibitors of HIV-protease. You will learn: Some of the theory
Ranking of HIV-protease inhibitors using AutoDock 1. Task Calculate possible binding modes and estimate the binding free energies for 1 3 inhibitors of HIV-protease. You will learn: Some of the theory
Lewis Structures and Molecular Shapes
 Lewis Structures and Molecular Shapes Rules for Writing Lewis Structures 1. Determine the correct skeleton structure (connectivity of atoms). Usually, the most electronegative atoms go around the edges
Lewis Structures and Molecular Shapes Rules for Writing Lewis Structures 1. Determine the correct skeleton structure (connectivity of atoms). Usually, the most electronegative atoms go around the edges
Exploring symmetry related bias in conformational data from the Cambridge Structural Database: A rare phenomenon?
 Exploring symmetry related bias in conformational data from the Cambridge Structural Database: A rare phenomenon? Aim To explore some well known cases where symmetry effects bias the distribution of conformational
Exploring symmetry related bias in conformational data from the Cambridge Structural Database: A rare phenomenon? Aim To explore some well known cases where symmetry effects bias the distribution of conformational
Introduction to Hartree-Fock calculations in Spartan
 EE5 in 2008 Hannes Jónsson Introduction to Hartree-Fock calculations in Spartan In this exercise, you will get to use state of the art software for carrying out calculations of wavefunctions for molecues,
EE5 in 2008 Hannes Jónsson Introduction to Hartree-Fock calculations in Spartan In this exercise, you will get to use state of the art software for carrying out calculations of wavefunctions for molecues,
Transition states and reaction paths
 Transition states and reaction paths Lab 4 Theoretical background Transition state A transition structure is the molecular configuration that separates reactants and products. In a system with a single
Transition states and reaction paths Lab 4 Theoretical background Transition state A transition structure is the molecular configuration that separates reactants and products. In a system with a single
Distance Constraint Model; Donald J. Jacobs, University of North Carolina at Charlotte Page 1 of 11
 Distance Constraint Model; Donald J. Jacobs, University of North Carolina at Charlotte Page 1 of 11 Taking the advice of Lord Kelvin, the Father of Thermodynamics, I describe the protein molecule and other
Distance Constraint Model; Donald J. Jacobs, University of North Carolina at Charlotte Page 1 of 11 Taking the advice of Lord Kelvin, the Father of Thermodynamics, I describe the protein molecule and other
Scuola di Chimica Computazionale
 Societa Chimica Italiana Gruppo Interdivisionale di Chimica Computazionale Scuola di Chimica Computazionale Introduzione, per Esercizi, all Uso del Calcolatore in Chimica rganica e Biologica Modellistica
Societa Chimica Italiana Gruppo Interdivisionale di Chimica Computazionale Scuola di Chimica Computazionale Introduzione, per Esercizi, all Uso del Calcolatore in Chimica rganica e Biologica Modellistica
Searching Substances in Reaxys
 Searching Substances in Reaxys Learning Objectives Understand that substances in Reaxys have different sources (e.g., Reaxys, PubChem) and can be found in Document, Reaction and Substance Records Recognize
Searching Substances in Reaxys Learning Objectives Understand that substances in Reaxys have different sources (e.g., Reaxys, PubChem) and can be found in Document, Reaction and Substance Records Recognize
CSD Conformer Generator User Guide
 CSD Conformer Generator User Guide 2018 CSD Release Copyright 2017 Cambridge Crystallographic Data Centre Registered Charity No 800579 Conditions of Use The CSD Conformer Generator is copyright work belonging
CSD Conformer Generator User Guide 2018 CSD Release Copyright 2017 Cambridge Crystallographic Data Centre Registered Charity No 800579 Conditions of Use The CSD Conformer Generator is copyright work belonging
CAMBRIDGE STRUCTURAL DATABASE SYSTEM 2010 RELEASE WORKED EXAMPLES
 CAMBRIDGE STRUCTURAL DATABASE SYSTEM 2010 RELEASE WORKED EXAMPLES Copyright 2009 The Cambridge Crystallographic Data Centre Registered Charity No 800579 Table of Contents 1 Introduction.................................................................
CAMBRIDGE STRUCTURAL DATABASE SYSTEM 2010 RELEASE WORKED EXAMPLES Copyright 2009 The Cambridge Crystallographic Data Centre Registered Charity No 800579 Table of Contents 1 Introduction.................................................................
Physical Chemistry Analyzing a Crystal Structure and the Diffraction Pattern Virginia B. Pett The College of Wooster
 Physical Chemistry Analyzing a Crystal Structure and the Diffraction Pattern Virginia B. Pett The College of Wooster L. W. Haynes and his Senior Independent Study students conducted the 2 + 2 photo addition
Physical Chemistry Analyzing a Crystal Structure and the Diffraction Pattern Virginia B. Pett The College of Wooster L. W. Haynes and his Senior Independent Study students conducted the 2 + 2 photo addition
Towards Physics-based Models for ADME/Tox. Tyler Day
 Towards Physics-based Models for ADME/Tox Tyler Day Overview Motivation Application: P450 Site of Metabolism Application: Membrane Permeability Future Directions and Applications Motivation Advantages
Towards Physics-based Models for ADME/Tox Tyler Day Overview Motivation Application: P450 Site of Metabolism Application: Membrane Permeability Future Directions and Applications Motivation Advantages
CMPS 3110: Bioinformatics. Tertiary Structure Prediction
 CMPS 3110: Bioinformatics Tertiary Structure Prediction Tertiary Structure Prediction Why Should Tertiary Structure Prediction Be Possible? Molecules obey the laws of physics! Conformation space is finite
CMPS 3110: Bioinformatics Tertiary Structure Prediction Tertiary Structure Prediction Why Should Tertiary Structure Prediction Be Possible? Molecules obey the laws of physics! Conformation space is finite
Docking with Water in the Binding Site using GOLD
 Docking with Water in the Binding Site using GOLD Version 2.0 November 2017 GOLD v5.6 Table of Contents Docking with Water in the Binding Site... 2 Case Study... 3 Introduction... 3 Provided Input Files...
Docking with Water in the Binding Site using GOLD Version 2.0 November 2017 GOLD v5.6 Table of Contents Docking with Water in the Binding Site... 2 Case Study... 3 Introduction... 3 Provided Input Files...
CMPS 6630: Introduction to Computational Biology and Bioinformatics. Tertiary Structure Prediction
 CMPS 6630: Introduction to Computational Biology and Bioinformatics Tertiary Structure Prediction Tertiary Structure Prediction Why Should Tertiary Structure Prediction Be Possible? Molecules obey the
CMPS 6630: Introduction to Computational Biology and Bioinformatics Tertiary Structure Prediction Tertiary Structure Prediction Why Should Tertiary Structure Prediction Be Possible? Molecules obey the
Computational Chemistry Lab Module: Conformational Analysis of Alkanes
 Introduction Computational Chemistry Lab Module: Conformational Analysis of Alkanes In this experiment, we will use CAChe software package to model the conformations of butane, 2-methylbutane, and substituted
Introduction Computational Chemistry Lab Module: Conformational Analysis of Alkanes In this experiment, we will use CAChe software package to model the conformations of butane, 2-methylbutane, and substituted
3. An Introduction to Molecular Mechanics
 3. An Introduction to Molecular Mechanics Introduction When you use Chem3D to draw molecules, the program assigns bond lengths and bond angles based on experimental data. The program does not contain real
3. An Introduction to Molecular Mechanics Introduction When you use Chem3D to draw molecules, the program assigns bond lengths and bond angles based on experimental data. The program does not contain real
Self-Organizing Superimposition Algorithm for Conformational Sampling
 Self-Organizing Superimposition Algorithm for Conformational Sampling FANGQIANG ZHU, DIMITRIS K. AGRAFIOTIS Johnson & Johnson Pharmaceutical Research & Development, L.L.C., 665 Stockton Drive, Exton, Pennsylvania
Self-Organizing Superimposition Algorithm for Conformational Sampling FANGQIANG ZHU, DIMITRIS K. AGRAFIOTIS Johnson & Johnson Pharmaceutical Research & Development, L.L.C., 665 Stockton Drive, Exton, Pennsylvania
1.b What are current best practices for selecting an initial target ligand atomic model(s) for structure refinement from X-ray diffraction data?!
 1.b What are current best practices for selecting an initial target ligand atomic model(s) for structure refinement from X-ray diffraction data?! Visual analysis: Identification of ligand density from
1.b What are current best practices for selecting an initial target ligand atomic model(s) for structure refinement from X-ray diffraction data?! Visual analysis: Identification of ligand density from
Large Scale FEP on Congeneric Ligand Series Have Practical Free Energy Calculations arrived at Last?
 Large Scale FEP on Congeneric Ligand Series Have Practical Free Energy Calculations arrived at Last? Thomas Steinbrecher, Teng Lin, Lingle Wang, Goran Krilov, Robert Abel, Woody Sherman, Richard Friesner
Large Scale FEP on Congeneric Ligand Series Have Practical Free Energy Calculations arrived at Last? Thomas Steinbrecher, Teng Lin, Lingle Wang, Goran Krilov, Robert Abel, Woody Sherman, Richard Friesner
Softwares for Molecular Docking. Lokesh P. Tripathi NCBS 17 December 2007
 Softwares for Molecular Docking Lokesh P. Tripathi NCBS 17 December 2007 Molecular Docking Attempt to predict structures of an intermolecular complex between two or more molecules Receptor-ligand (or drug)
Softwares for Molecular Docking Lokesh P. Tripathi NCBS 17 December 2007 Molecular Docking Attempt to predict structures of an intermolecular complex between two or more molecules Receptor-ligand (or drug)
Conformational sampling of macrocycles in solution and in the solid state
 Conformational sampling of macrocycles in solution and in the solid state Paul Hawkins, Ph.D. Head of Scientific Solutions Stanislaw Wlodek, Ph.D. Senior Scientific Developer 6/6/2018 https://berkonomics.com/?p=2437
Conformational sampling of macrocycles in solution and in the solid state Paul Hawkins, Ph.D. Head of Scientific Solutions Stanislaw Wlodek, Ph.D. Senior Scientific Developer 6/6/2018 https://berkonomics.com/?p=2437
M E R C E R W I N WA L K T H R O U G H
 H E A L T H W E A L T H C A R E E R WA L K T H R O U G H C L I E N T S O L U T I O N S T E A M T A B L E O F C O N T E N T 1. Login to the Tool 2 2. Published reports... 7 3. Select Results Criteria...
H E A L T H W E A L T H C A R E E R WA L K T H R O U G H C L I E N T S O L U T I O N S T E A M T A B L E O F C O N T E N T 1. Login to the Tool 2 2. Published reports... 7 3. Select Results Criteria...
Protein-Ligand Docking Evaluations
 Introduction Protein-Ligand Docking Evaluations Protein-ligand docking: Given a protein and a ligand, determine the pose(s) and conformation(s) minimizing the total energy of the protein-ligand complex
Introduction Protein-Ligand Docking Evaluations Protein-ligand docking: Given a protein and a ligand, determine the pose(s) and conformation(s) minimizing the total energy of the protein-ligand complex
General Chemistry Lab Molecular Modeling
 PURPOSE The objectives of this experiment are PROCEDURE General Chemistry Lab Molecular Modeling To learn how to use molecular modeling software, a commonly used tool in chemical research and industry.
PURPOSE The objectives of this experiment are PROCEDURE General Chemistry Lab Molecular Modeling To learn how to use molecular modeling software, a commonly used tool in chemical research and industry.
CSD Conformer Generator User Guide
 CSD Conformer Generator User Guide 2017 CSD Release Copyright 2016 Cambridge Crystallographic Data Centre Registered Charity No 800579 Conditions of Use The CSD Conformer Generator is copyright work belonging
CSD Conformer Generator User Guide 2017 CSD Release Copyright 2016 Cambridge Crystallographic Data Centre Registered Charity No 800579 Conditions of Use The CSD Conformer Generator is copyright work belonging
3. An Introduction to Molecular Mechanics
 3. An Introduction to Molecular Mechanics Introduction When you use Chem3D to draw molecules, the program assigns bond lengths and bond angles based on experimental data. The program does not contain real
3. An Introduction to Molecular Mechanics Introduction When you use Chem3D to draw molecules, the program assigns bond lengths and bond angles based on experimental data. The program does not contain real
OECD QSAR Toolbox v.4.1. Tutorial illustrating new options for grouping with metabolism
 OECD QSAR Toolbox v.4.1 Tutorial illustrating new options for grouping with metabolism Outlook Background Objectives Specific Aims The exercise Workflow 2 Background Grouping with metabolism is a procedure
OECD QSAR Toolbox v.4.1 Tutorial illustrating new options for grouping with metabolism Outlook Background Objectives Specific Aims The exercise Workflow 2 Background Grouping with metabolism is a procedure
MassHunter Software Overview
 MassHunter Software Overview 1 Qualitative Analysis Workflows Workflows in Qualitative Analysis allow the user to only see and work with the areas and dialog boxes they need for their specific tasks A
MassHunter Software Overview 1 Qualitative Analysis Workflows Workflows in Qualitative Analysis allow the user to only see and work with the areas and dialog boxes they need for their specific tasks A
Tutorial 1: Setting up your Skyline document
 Tutorial 1: Setting up your Skyline document Caution! For using Skyline the number formats of your computer have to be set to English (United States). Open the Control Panel Clock, Language, and Region
Tutorial 1: Setting up your Skyline document Caution! For using Skyline the number formats of your computer have to be set to English (United States). Open the Control Panel Clock, Language, and Region
Section III - Designing Models for 3D Printing
 Section III - Designing Models for 3D Printing In this section of the Jmol Training Guide, you will become familiar with the commands needed to design a model that will be built on a 3D Printer. As you
Section III - Designing Models for 3D Printing In this section of the Jmol Training Guide, you will become familiar with the commands needed to design a model that will be built on a 3D Printer. As you
Dock Ligands from a 2D Molecule Sketch
 Dock Ligands from a 2D Molecule Sketch March 31, 2016 Sample to Insight CLC bio, a QIAGEN Company Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.clcbio.com support-clcbio@qiagen.com
Dock Ligands from a 2D Molecule Sketch March 31, 2016 Sample to Insight CLC bio, a QIAGEN Company Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.clcbio.com support-clcbio@qiagen.com
Type II Kinase Inhibitors Show an Unexpected Inhibition Mode against Parkinson s Disease-Linked LRRK2 Mutant G2019S.
 Type II Kinase Inhibitors Show an Unexpected Inhibition Mode against Parkinson s Disease-Linked LRRK2 Mutant G219S. Min Liu@&*, Samantha A. Bender%*, Gregory D Cuny@, Woody Sherman, Marcie Glicksman@ Soumya
Type II Kinase Inhibitors Show an Unexpected Inhibition Mode against Parkinson s Disease-Linked LRRK2 Mutant G219S. Min Liu@&*, Samantha A. Bender%*, Gregory D Cuny@, Woody Sherman, Marcie Glicksman@ Soumya
How is molecular dynamics being used in life sciences? Davide Branduardi
 How is molecular dynamics being used in life sciences? Davide Branduardi davide.branduardi@schrodinger.com Exploring molecular processes with MD Drug discovery and design Protein-protein interactions Protein-DNA
How is molecular dynamics being used in life sciences? Davide Branduardi davide.branduardi@schrodinger.com Exploring molecular processes with MD Drug discovery and design Protein-protein interactions Protein-DNA
Lewis Structures and Molecular Shapes
 Lewis Structures and Molecular Shapes Rules for Writing Lewis Structures 1. Determine the correct skeleton structure (connectivity of atoms). Usually, the most electronegative atoms go around the edges
Lewis Structures and Molecular Shapes Rules for Writing Lewis Structures 1. Determine the correct skeleton structure (connectivity of atoms). Usually, the most electronegative atoms go around the edges
Using AutoDock 4 with ADT: A Tutorial
 Using AutoDock 4 with ADT: A Tutorial Ruth Huey Sargis Dallakyan Alex Perryman David S. Goodsell (Garrett Morris) 9/2/08 Using AutoDock 4 with ADT 1 What is Docking? Predicting the best ways two molecules
Using AutoDock 4 with ADT: A Tutorial Ruth Huey Sargis Dallakyan Alex Perryman David S. Goodsell (Garrett Morris) 9/2/08 Using AutoDock 4 with ADT 1 What is Docking? Predicting the best ways two molecules
Comparing whole genomes
 BioNumerics Tutorial: Comparing whole genomes 1 Aim The Chromosome Comparison window in BioNumerics has been designed for large-scale comparison of sequences of unlimited length. In this tutorial you will
BioNumerics Tutorial: Comparing whole genomes 1 Aim The Chromosome Comparison window in BioNumerics has been designed for large-scale comparison of sequences of unlimited length. In this tutorial you will
Polypeptide Folding Using Monte Carlo Sampling, Concerted Rotation, and Continuum Solvation
 Polypeptide Folding Using Monte Carlo Sampling, Concerted Rotation, and Continuum Solvation Jakob P. Ulmschneider and William L. Jorgensen J.A.C.S. 2004, 126, 1849-1857 Presented by Laura L. Thomas and
Polypeptide Folding Using Monte Carlo Sampling, Concerted Rotation, and Continuum Solvation Jakob P. Ulmschneider and William L. Jorgensen J.A.C.S. 2004, 126, 1849-1857 Presented by Laura L. Thomas and
ENERGY MINIMIZATION AND CONFORMATION SEARCH ANALYSIS OF TYPE-2 ANTI-DIABETES DRUGS
 Int. J. Chem. Sci.: 6(2), 2008, 982-992 EERGY MIIMIZATI AD CFRMATI SEARC AALYSIS F TYPE-2 ATI-DIABETES DRUGS R. PRASAA LAKSMI a, C. ARASIMA KUMAR a, B. VASATA LAKSMI, K. AGA SUDA, K. MAJA, V. JAYA LAKSMI
Int. J. Chem. Sci.: 6(2), 2008, 982-992 EERGY MIIMIZATI AD CFRMATI SEARC AALYSIS F TYPE-2 ATI-DIABETES DRUGS R. PRASAA LAKSMI a, C. ARASIMA KUMAR a, B. VASATA LAKSMI, K. AGA SUDA, K. MAJA, V. JAYA LAKSMI
Using Web-Based Computations in Organic Chemistry
 10/30/2017 1 Using Web-Based Computations in Organic Chemistry John Keller UAF Department of Chemistry & Biochemistry The UAF WebMO site Practical aspects of computational chemistry theory and nomenclature
10/30/2017 1 Using Web-Based Computations in Organic Chemistry John Keller UAF Department of Chemistry & Biochemistry The UAF WebMO site Practical aspects of computational chemistry theory and nomenclature
Alchemical free energy calculations in OpenMM
 Alchemical free energy calculations in OpenMM Lee-Ping Wang Stanford Department of Chemistry OpenMM Workshop, Stanford University September 7, 2012 Special thanks to: John Chodera, Morgan Lawrenz Outline
Alchemical free energy calculations in OpenMM Lee-Ping Wang Stanford Department of Chemistry OpenMM Workshop, Stanford University September 7, 2012 Special thanks to: John Chodera, Morgan Lawrenz Outline
The Schrödinger KNIME extensions
 The Schrödinger KNIME extensions Computational Chemistry and Cheminformatics in a workflow environment Jean-Christophe Mozziconacci Volker Eyrich KNIME UGM, Berlin, February 2015 The Schrödinger Extensions
The Schrödinger KNIME extensions Computational Chemistry and Cheminformatics in a workflow environment Jean-Christophe Mozziconacci Volker Eyrich KNIME UGM, Berlin, February 2015 The Schrödinger Extensions
VCell Tutorial. Building a Rule-Based Model
 VCell Tutorial Building a Rule-Based Model We will demonstrate how to create a rule-based model of EGFR receptor interaction with two adapter proteins Grb2 and Shc. A Receptor-monomer reversibly binds
VCell Tutorial Building a Rule-Based Model We will demonstrate how to create a rule-based model of EGFR receptor interaction with two adapter proteins Grb2 and Shc. A Receptor-monomer reversibly binds
Structural Bioinformatics (C3210) Molecular Mechanics
 Structural Bioinformatics (C3210) Molecular Mechanics How to Calculate Energies Calculation of molecular energies is of key importance in protein folding, molecular modelling etc. There are two main computational
Structural Bioinformatics (C3210) Molecular Mechanics How to Calculate Energies Calculation of molecular energies is of key importance in protein folding, molecular modelling etc. There are two main computational
Molecular Modeling Lecture 7. Homology modeling insertions/deletions manual realignment
 Molecular Modeling 2018-- Lecture 7 Homology modeling insertions/deletions manual realignment Homology modeling also called comparative modeling Sequences that have similar sequence have similar structure.
Molecular Modeling 2018-- Lecture 7 Homology modeling insertions/deletions manual realignment Homology modeling also called comparative modeling Sequences that have similar sequence have similar structure.
Namur, December 3 rd, 2007
 University of Namur Laboratoire de Physico-Chimie Informatique Daniel P. Vercauteren Director, Professor at the Sciences Faculty Rue de Bruxelles, 61 B-5000 Namur, Belgium Phone +32 (0)81 72 45 34 Fax
University of Namur Laboratoire de Physico-Chimie Informatique Daniel P. Vercauteren Director, Professor at the Sciences Faculty Rue de Bruxelles, 61 B-5000 Namur, Belgium Phone +32 (0)81 72 45 34 Fax
PROTEIN-PROTEIN DOCKING REFINEMENT USING RESTRAINT MOLECULAR DYNAMICS SIMULATIONS
 TASKQUARTERLYvol.20,No4,2016,pp.353 360 PROTEIN-PROTEIN DOCKING REFINEMENT USING RESTRAINT MOLECULAR DYNAMICS SIMULATIONS MARTIN ZACHARIAS Physics Department T38, Technical University of Munich James-Franck-Str.
TASKQUARTERLYvol.20,No4,2016,pp.353 360 PROTEIN-PROTEIN DOCKING REFINEMENT USING RESTRAINT MOLECULAR DYNAMICS SIMULATIONS MARTIN ZACHARIAS Physics Department T38, Technical University of Munich James-Franck-Str.
Building small molecules
 Building small molecules Use the Builder (right panel) to build up molecules. Start building clicking a fragment/atom in the builder and it will appear to the workspace. Continue modifying the molecule
Building small molecules Use the Builder (right panel) to build up molecules. Start building clicking a fragment/atom in the builder and it will appear to the workspace. Continue modifying the molecule
Chem 310, Organic Chemistry Lab Molecular Modeling Using Macromodel
 Chem 310, Organic Chemistry Lab Molecular Modeling Using Macromodel This is a molecular modeling experiment, and should be written up in your lab notebook just as if it were a normal "wet-chemistry" experiment.
Chem 310, Organic Chemistry Lab Molecular Modeling Using Macromodel This is a molecular modeling experiment, and should be written up in your lab notebook just as if it were a normal "wet-chemistry" experiment.
Protein-Ligand Docking Methods
 Review Goal: Given a protein structure, predict its ligand bindings Protein-Ligand Docking Methods Applications: Function prediction Drug discovery etc. Thomas Funkhouser Princeton University S597A, Fall
Review Goal: Given a protein structure, predict its ligand bindings Protein-Ligand Docking Methods Applications: Function prediction Drug discovery etc. Thomas Funkhouser Princeton University S597A, Fall
The Schrödinger KNIME extensions
 The Schrödinger KNIME extensions Computational Chemistry and Cheminformatics in a workflow environment Jean-Christophe Mozziconacci Volker Eyrich KNIME UGM, Zurich, February 2014 The Schrödinger extensions
The Schrödinger KNIME extensions Computational Chemistry and Cheminformatics in a workflow environment Jean-Christophe Mozziconacci Volker Eyrich KNIME UGM, Zurich, February 2014 The Schrödinger extensions
Learning to Use Scigress Wagner, Eugene P. (revised May 15, 2018)
 Learning to Use Scigress Wagner, Eugene P. (revised May 15, 2018) Abstract Students are introduced to basic features of Scigress by building molecules and performing calculations on them using semi-empirical
Learning to Use Scigress Wagner, Eugene P. (revised May 15, 2018) Abstract Students are introduced to basic features of Scigress by building molecules and performing calculations on them using semi-empirical
Preparing a PDB File
 Figure 1: Schematic view of the ligand-binding domain from the vitamin D receptor (PDB file 1IE9). The crystallographic waters are shown as small spheres and the bound ligand is shown as a CPK model. HO
Figure 1: Schematic view of the ligand-binding domain from the vitamin D receptor (PDB file 1IE9). The crystallographic waters are shown as small spheres and the bound ligand is shown as a CPK model. HO
Design of a Novel Globular Protein Fold with Atomic-Level Accuracy
 Design of a Novel Globular Protein Fold with Atomic-Level Accuracy Brian Kuhlman, Gautam Dantas, Gregory C. Ireton, Gabriele Varani, Barry L. Stoddard, David Baker Presented by Kate Stafford 4 May 05 Protein
Design of a Novel Globular Protein Fold with Atomic-Level Accuracy Brian Kuhlman, Gautam Dantas, Gregory C. Ireton, Gabriele Varani, Barry L. Stoddard, David Baker Presented by Kate Stafford 4 May 05 Protein
A Brief Guide to All Atom/Coarse Grain Simulations 1.1
 A Brief Guide to All Atom/Coarse Grain Simulations 1.1 Contents 1. Introduction 2. AACG simulations 2.1. Capabilities and limitations 2.2. AACG commands 2.3. Visualization and analysis 2.4. AACG simulation
A Brief Guide to All Atom/Coarse Grain Simulations 1.1 Contents 1. Introduction 2. AACG simulations 2.1. Capabilities and limitations 2.2. AACG commands 2.3. Visualization and analysis 2.4. AACG simulation
Conformation Searching Applications
 Chemistry 380.37 Fall 2015 Dr. Jean M. Standard September 23, 2015 Conformation Searching Applications Cycloheptadecane Redux So what would a conformation search on cycloheptadecane using Spartan find?
Chemistry 380.37 Fall 2015 Dr. Jean M. Standard September 23, 2015 Conformation Searching Applications Cycloheptadecane Redux So what would a conformation search on cycloheptadecane using Spartan find?
Chemistry 14CL. Worksheet for the Molecular Modeling Workshop. (Revised FULL Version 2012 J.W. Pang) (Modified A. A. Russell)
 Chemistry 14CL Worksheet for the Molecular Modeling Workshop (Revised FULL Version 2012 J.W. Pang) (Modified A. A. Russell) Structure of the Molecular Modeling Assignment The molecular modeling assignment
Chemistry 14CL Worksheet for the Molecular Modeling Workshop (Revised FULL Version 2012 J.W. Pang) (Modified A. A. Russell) Structure of the Molecular Modeling Assignment The molecular modeling assignment
All-atom Molecular Mechanics. Trent E. Balius AMS 535 / CHE /27/2010
 All-atom Molecular Mechanics Trent E. Balius AMS 535 / CHE 535 09/27/2010 Outline Molecular models Molecular mechanics Force Fields Potential energy function functional form parameters and parameterization
All-atom Molecular Mechanics Trent E. Balius AMS 535 / CHE 535 09/27/2010 Outline Molecular models Molecular mechanics Force Fields Potential energy function functional form parameters and parameterization
ONETEP PB/SA: Application to G-Quadruplex DNA Stability. Danny Cole
 ONETEP PB/SA: Application to G-Quadruplex DNA Stability Danny Cole Introduction Historical overview of structure and free energy calculation of complex molecules using molecular mechanics and continuum
ONETEP PB/SA: Application to G-Quadruplex DNA Stability Danny Cole Introduction Historical overview of structure and free energy calculation of complex molecules using molecular mechanics and continuum
Fast, Intuitive Structure Determination IV: Space Group Determination and Structure Solution
 Fast, Intuitive Structure Determination IV: Space Group Determination and Structure Solution November 25, 2013 Welcome I I Dr. Michael Ruf Product Manager Crystallography Bruker AXS Inc. Madison, WI, USA
Fast, Intuitive Structure Determination IV: Space Group Determination and Structure Solution November 25, 2013 Welcome I I Dr. Michael Ruf Product Manager Crystallography Bruker AXS Inc. Madison, WI, USA
2D Clausius Clapeyron equation
 2D Clausius Clapeyron equation 1/14 Aim: Verify the Clausius Clapeyron equation by simulations of a 2D model of matter Model: 8-4 type potential ( Lennard-Jones in 2D) u(r) = 1 r 8 1 r 4 Hard attractive
2D Clausius Clapeyron equation 1/14 Aim: Verify the Clausius Clapeyron equation by simulations of a 2D model of matter Model: 8-4 type potential ( Lennard-Jones in 2D) u(r) = 1 r 8 1 r 4 Hard attractive
